Abstract
Osteoarthritis is a common degenerative joint disease, in which mechanical overloading disrupts subchondral bone remodeling before cartilage degeneration and the osteocytes in the subchondral bone are mainly responsible for mechanosensing. However, their functional role in the early osteoarthritis is still unclear. Here we show that mechanical stress induces osteocytes in subchondral bone to secrete extracellular vesicles that accelerate cartilage metabolic dysregulation in patients with both sexes and male mice. The miR-23b-3p in extracellular vesicles promotes cartilage catabolism and inhibits anabolism by targeting OTUD4, disrupting mitophagy in chondrocytes. Inhibiting miR-23b-3p in osteocytes or chondrocytes reduces cartilage degeneration and osteoarthritis progression in male mice. Together, our findings highlight that osteocyte-derived extracellular vesicles mediate communication with chondrocytes and suggest miR-23b-3p as a potential therapeutic target for osteoarthritis.
Similar content being viewed by others
Introduction
Osteoarthritis (OA) is the most common degenerative joint disease1, and it is characterized by a range of joint changes, including articular cartilage damage, synovitis, subchondral bone (SCB) sclerosis and osteophyte formation2,3. As a leading cause of disability, OA is a major clinical and socioeconomic burden across the globe3. However, due to its poorly understood pathomechanisms, there currently is no effective clinical intervention for OA.
Over the last decade or so OA has been recognized as a disease of the whole joint, implicating pathological alterations in the articular cartilage, SCB, ligaments, capsule, synovial membrane and periarticular muscles4,5. While cartilage degeneration plays a crucial role in OA pathogenesis, a series of studies have suggested that SCB remodeling precedes such degeneration6,7,8. Emerging evidence indicates that abnormal mechanical stress during the early stages of OA progression leads to increased vascularity, porosity, microcracks and fissures at the bone-cartilage interface, resulting in the release of various cytokines and extracellular vesicles (EVs) from the SCB that likely communicate with the covered articular cartilage, aggravating cartilage degeneration and OA progression7,9. For instance, Jung et al. reported that cartilage can acquire calcium-phosphate complexes from the SCB, leading to increased production of MMP13, a key extracellular protease know to contribute to OA progression, in chondrocytes10. Further, Liu et al. showed that osteoclast-derived exosomes contain miRNAs that impair chondrocyte function in OA11. Thus, the crosstalk between the SCB and the cartilage plays an important role in OA pathogenesis and progression.
Osteocytes constitute 90–95% of the total skeletal cell population, which is approximately 20 times the number of osteoblasts, and are considered the primary cell type responsive to mechanical loading12. Previous studies have reported striking morphological and phenotypic changes in the osteocytes of the SCB from OA samples, including rounded cell bodies with reduced dendrites and dysregulated expression of osteocyte markers, which may be a response to excessive mechanical loading12,13. Mazur et al. reported that deficiency of MMP13 in osteocytes had a protective effect on early OA progression through its impact on perilacunar/canalicular remodeling14. Importantly, another study identified a highly co-expressed signature of mechano-responsive osteocytes in SCB lesions that are associated with OA13. However, the potential role of SCB osteocytes under mechanical stress in OA development has not been reported.
EVs are nanosized vesicles with diameters ranging from 40-160 nm that are actively secreted by cells under physiological or pathological conditions15. They transport various proteins, mRNAs and miRNAs to recipient cells, altering their functions15,16. EVs play a crucial role in intercellular and interorgan communication, exerting pleiotropic effects on nearby or distant cells, tissues and organs16. For example, myostatin has been shown to suppress miR-218 levels in an osteocyte cell line and its derived exosomes inhibit osteoblast differentiation in vitro17. Also, Wang et al. showed that EVs from an aged bone matrix promoted the calcification of vascular smooth muscle cells18. In summary, osteocyte-derived EVs play crucial roles in mediating various normo- and pathophysiological processes via their encapsulated miRNAs, highlighting their strong potential for targeted interventions and therapeutic applications. Therefore, we hypothesized that SCB osteocytes under mechanical stress secret EVs to regulate chondrocyte function during OA progression.
Here, we demonstrate that SCB osteocytes secrete EVs to skew chondrocyte metabolism during OA progression. Subsequently, we show that miR-23b-3p is the key miRNA in the osteocyte-derived EVs responsible for the effects on cartilage metabolism. Mechanistically, we show that miR-23b-3p inhibits mitophagy by targeting Otud4 to promote catabolism and inhibit anabolism in chondrocytes. Further, we show that inhibition of miR-23b-3p in either osteocytes or chondrocytes alleviates cartilage degeneration during OA progression in a mouse model. Together, our results unveil a previously unknown mechanism of intercellular communication between SCB osteocytes and cartilage chondrocytes during OA, while identifying a novel therapeutic intervention for OA.
Results
Mechanosensitive SCB osteocytes secrete EVs to impair chondrocyte function during OA progression
As the SCB plays a vital role in OA, we collected clinical specimens from patients with knee OA and investigated the histological and molecular features of the cartilage and the SCB. We found that the medial tibial plateau exhibited more advanced cartilage deterioration and greater SCB sclerosis, whereas the lateral tibial plateau showed minimal cartilage degeneration and slight SCB sclerosis (Fig. S1A, B). By qPCR analysis we found that there was significantly higher mRNA expression of genes encoding catabolic enzymes (ADAMTS5 and MMP13) and significantly lower expression of anabolism-related marker genes (COL2A1, ACAN and SOX9) in human articular cartilage at the medial tibial plateau compared to the lateral tibial plateau (Fig. S1A). Moreover, by finite element analysis (FEA) we found evidence of higher mechanical stress at the medial region than that at the lateral region within the knee joint of the patients with OA compared with that in healthy controls (Fig. S1C). Hence, the lateral and medial tibial plateaus were respectively referred to as knee joint regions experiencing low stress (LS) and high stress (HS) for subsequent analysis.
Considering that osteocytes are the predominant cell type in the skeleton and primarily responsible for mechanosensing in the SCB, we hypothesized that SCB osteocytes in HS area impair local chondrocyte function during OA progression. Therefore, we conducted a co-culture experiment involving primary human osteoarthritic chondrocytes and SCB tissues exclusively containing osteocytes, followed by a series of analyses as described below (Fig. 1A). Compared with the LS group, we found significantly higher levels of pro-inflammatory factors (IL-1β, IL-6, and TNF-α) in the supernatant of human chondrocytes from the HS area (Fig. 1B). Moreover, the intensity of Alcian blue staining of proteoglycans was markedly lower in the HS area (Fig. 1C). We also found significantly higher mRNA and protein levels of ADAMTS5/ADAMTS5 and MMP13/MMP13, as well as significantly lower expression of COL2A1/COLII, ACAN/ACAN and SOX9/SOX9, in the HS area compared to the LS area (Figs. 1D, E and S1D). These data suggest that SCB osteocytes may release mediators that negatively impact articular cartilage function during OA progression.
A A schematic diagram illustrating the workflow utilizing primary human osteoarthritic chondrocytes co-cultured with osteocytes located in high stress (HS) and low stress (LS) region of the subchondral bone (SCB). B Levels of IL-1β, IL-6 and TNF-α measured by ELISA assays (n = 6). C The Alcian blue staining. Scale bar, 1 mm. D qPCR analysis of COL2A1, ACAN, SOX9, MMP13 and ADAMTS5 levels in primary human osteoarthritic chondrocytes co-cultured with LS/HS tissues (n = 3). E Western blot of COLII, ACAN, SOX9, MMP13 and ADAMTS5 protein expression in primary human osteoarthritic chondrocytes co-cultured with LS/HS tissues (n = 3). F RNA-sequencing and Gene Ontology (GO) enrichment analysis of the genes in LS/HS tissues. G A heat map (top) and its quantitation (bottom) indicating the expression of genes associated with extracellular vesicles (EVs) biogenesis within LS/HS tissues. H A schematic diagram summarizing the biology of EV biogenesis. I Representative electron microscopy images of intraluminal vesicles (ILVs) in LS/HS tissues. Scale bar, 500 nm. J Illustration of the animal experimental protocol for osteoarthritis (OA) modeling using osteocyte-specific ILVs/EVs reporter mice. K Safranin O staining. Scale bar, 200 μm. L The Osteoarthritis Research Society International (OARSI) scores (n = 6). M Localization of Cd63-GFP (green) in SCB osteocytes. Scale bar, 50 μm. SCB Subchondral bone, BM Bone marrow. N The semi-quantitative analysis of Cd63-GFP+ positive cells in SCB osteocytes (n = 6). O Localization of Cd63-GFP (green) in articular cartilage. The white arrows indicated Cd63-GFP (green). Scale bar, 50 μm. P The semi-quantitative analysis of Cd63-GFP+ positive cells in articular cartilage (n = 6). Data are presented as the mean ± SD. P values are from paired two-tailed Student’s t-test (B, D), Kruskal–Wallis test followed by two-tailed Mann–Whitney U test (L), one-way analysis of variance (ANOVA) with Tukey’s multiple comparisons test (N, P) or Hypergeometric test (F). Source data are provided as a Source Data file.
To investigate the type of mediator released by SCB osteocytes under mechanical stress that regulates cartilage metabolism during OA progression, we performed RNA sequencing analysis on SCB osteocytes from the LS and HS regions of knee joint from patients with OA. Notably, by GO analysis focusing on cellular components we found a strong enrichment in the ‘extracellular exosome’ category and a striking increase in genes related to EV synthesis and secretion among the samples from the HS area compared to the LS area (Fig. 1F–H). Consistent with these results, the expression of RAB27B, which is known to play a crucial role in EV secretion19, was significantly higher in the HS area compared to that in the LS area (Fig. S1E). Moreover, by transmission electron microscopy (TEM) analysis we found a greater number of intraluminal vesicles (ILVs) within the SCB osteocytes in the HS area compared to the LS area (Fig. 1I).
Next, we explored if the findings in human OA can be extended to a mouse model. Thus, we induced OA in 8-week-old male C57BL/6 mice by surgically destabilizing the medial meniscus (DMM). By micro-computed tomography (micro-CT) analysis we found that there was progressive SCB deformity and sclerosis during 8 weeks of OA progression after surgery (Fig. S1F, G). By Safranin O staining and Osteoarthritis Research Society International (OARSI) scoring we found that there was an obvious proteoglycans loss and progressive articular cartilage degeneration in the DMM mice from 2 to 4 weeks after surgery, which was markedly exacerbated at 8 weeks after surgery (Fig. S1H, I). And consistent with the findings from the above human study, by immunohistochemistry (IHC) we found that the expression of Rab27b was progressively higher during the progression of OA in the mouse DMM model (Fig. S1J, K).
Next, we generated transgenic mice expressing a reporter gene specific to osteocyte-derived ILVs/EVs (Dmp1cre;Cd63loxp-eGFP) (Fig. S2A). We detected no eGFP signals in the SCB and cartilage of Dmp1cre mice and Cd63loxp-eGFP control mice (Fig. S2B). To verify whether EVs from SCB osteocytes can reach the overlying cartilage layer during OA progression, we performed DMM surgery in 8-week-old male reporter mice and followed them for 8 weeks (Fig. 1J). By Safranin O staining and OARSI scoring we found that there was progressive cartilage degeneration after DMM in the reporter mice (Fig. 1K, L). In addition, we observed a progressively greater number of Cd63+ osteocytes in the SCB over time following DMM surgery (Fig. 1M, N). Notably, we found a progressively greater eGFP signal mainly in the deep zone of the cartilage and the SCB marrow cavity that are associated with endomucin+ vessels in the DMM mice (Figs. 1O, P and S2C). These findings suggest that osteocyte-derived EVs may reach the calcified and deep articular cartilage during OA development directly through microcracks and vascular channels.
Osteocyte-derived EVs skew the metabolism of chondrocytes
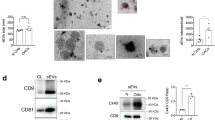
To further clarify the regulatory effect of osteocyte-derived EVs on chondrocytes in vitro, we firstly isolated EVs from the SCB at LS (LS-EVs) and HS (HS-EVs) regions from the knee joints of patients with OA and then used them to treat human primary chondrocytes (Fig. 2A). At first we observed the morphology of isolated osteocyte-derived EVs by TEM (Fig. S3A) and particle size distribution by nanoparticle tracking analysis (NTA) (Fig. 2B), as well as by Western blotting for expression of EV markers TSG101, CD81, CD9, CD63 and HSP70 (Fig. S3B). These results showed that EV secretion was increased in the HS region compared to the LS region. Next, PKH26 labeled the isolated osteocyte-derived EVs and found that they were taken up by human OA primary articular chondrocytes after we treated the cells with the labeled EVs (Fig. 2C). Notably, by Alcian blue staining we found that the intensity of proteoglycans was strikingly lower in the chondrocytes treated with the HS-EVs than those treated with the LS-EVs (Fig. 2D). We also found significantly higher levels of pro-inflammatory cytokines (IL-1β, IL-6 and TNF-α) in the supernatant of the human primary OA articular chondrocytes after treatment with HS-EVs compared to those treated with LS-EVs (Fig. S3C). Furthermore, the gene and protein levels of ADAMTS5/ADAMTS5 and MMP13/MMP13 were significantly higher, while the levels of COL2A1/COLII, ACAN/ACAN and SOX9/SOX9 were significantly lower, in chondrocytes after treatment with HS-EVs compared to those treated with LS-EVs (Figs. S3D, 2E and S3E).
A A schematic diagram illustrated that primary human osteoarthritic chondrocytes were treated with extracellular vesicles (EVs) extracted from low stress/high stress tissues (LS/HS-EVs). B The nanoparticle tracking analysis (NTA) analysis. C Representative confocal images showing the uptake of LS/HS-EVs in chondrocytes. Scale bar, 30 μm. D Alcian blue staining. Scale bar, 1 mm. E Western blot analysis of COLII, ACAN, SOX9, MMP13 and ADAMTS5 protein expression (n = 3). F A schematic diagram illustrating the administration of intra-articular injection of LS/HS-EVs to destabilization of the medial meniscus (DMM) mice. G Representative confocal images of PKH26-labeled osteocyte-EVs in the cartilage. Scale bar, 50 μm. H The Safranin O images (n = 8). Scale bar in upper panel, 200 μm. Scale bar in lower panel, 100 μm. I A schematic diagram illustrated that primary mouse chondrocytes were treated with Static/Load-EVs. J The NTA analysis. K Representative confocal fluorescence micrographs showing the uptake of EVs in chondrocytes. Scale bar, 30 μm. L Alcian blue staining. Scale bar, 2 mm. M Western blot analysis of ColII, Acan, Sox9, Mmp13 and Adamts5 protein level (n = 3). N A schematic diagram illustrating the administration of intra-articular injection of Static-EVs and Load-EVs to DMM mice. O Representative confocal images of PKH26-labeled MLO-Y4-EVs in the cartilage. Scale bar, 50 μm. P The Safranin O images of cartilage (n = 8). Scale bar in upper panel, 200 μm. Scale bar in lower panel, 100 μm. Q A schematic diagram illustrating the strategy for reducing osteocyte-derived EVs in DMM mice. R Representative immunohistochemical staining of Cd63 in subchondral bone (SCB) osteocytes (n = 8). Scale bars: 50 μm. S The Safranin O images (n = 8). Scale bar in upper panel, 200 μm. Scale bar in lower panel, 100 μm. Data are presented as the mean ± SD. Data was analyzed by unpaired two-tailed Student’s t-test (R). Source data are provided as a Source Data file.
A A schematic diagram illustrating the co-culturing system. Scale bar, 30 μm. B miR-223 expression in primary mouse chondrocytes (n = 3). C Images of primary mouse osteocytes isolated from wild-type (WT)- or osteocytes-specific Dicer knockout mice (cKO). Scale bar, 100 μm. D Western blot (left panel) and semi-quantitative analysis (right panel) of Dicer protein level in WT/cKO osteocytes (n = 3). E A schematic diagram illustrating the coculture of primary mouse chondrocytes and extracellular vesicles (EVs) from WT/cKO osteocytes subjected to mechanical loading (WT-EVs / cKO-EVs). F Alcian blue staining. Scale bar, 2 mm. G qPCR analysis of the Col2a1, Acan, Sox9, Mmp13 and Adamts5 levels in chondrocytes treated with WT-EVs or cKO-EVs (n = 3). H Western blot analysis (the left panel) and quantitative analysis (the right panel) of ColII, Acan, Sox9, Mmp13 and Adamts5 protein level in chondrocytes treated with WT-EVs or cKO-EVs (n = 3). I A schematic diagram showed that WT/cKO mice were performed Sham/destabilization of the medial meniscus (DMM) surgery. J Western blot (left panel) and semi-quantitative analysis (right panel) of Dicer protein level in cKO mice (n = 5). K Heat map of miRNA qPCR array (n = 3). L Safranin O images (n = 8). Scale bar in upper panel, 200 μm. Scale bar in lower panel, 100 μm. M The Osteoarthritis Research Society International (OARSI) scores (n = 8). N Representative confocal images of Acan, Sox9, Mmp13 and Adamts5 expression in cartilage from the indicated groups. Scale bars: 50 μm. O The quantitative analysis of Acan, Sox9, Mmp13 and Adamts5 protein (n = 8). Data are presented as the mean ± SD. P values are from unpaired two-tailed Student’s t-test (B, D, G, H and J), Kruskal–Wallis test followed by two-tailed Mann–Whitney U test (M), one-way analysis of variance (ANOVA) with Tukey’s multiple comparisons test (O). Source data are provided as a Source Data file.
To further determine whether human HS-EVs exacerbate OA progression in vivo, we intraarticularly administered the isolated LS-EVs or HS-EVs from the human samples to DMM mice for 8 weeks as shown in the flowchart (Figs. 2F and S3F). The injected PKH26-labeled human osteocyte-derived EVs were successfully taken up by the mouse chondrocytes in vivo, as evidenced by detected fluorescent signals along the surface of the cartilage of DMM mice (Fig. 2G). Compared with the LS-EVs injection group, we found by Safranin O staining and OARSI score that cartilage degeneration was further aggravated after HS-EVs injection (Figs. 2H and S3G). In addition, by immunofluorescent analysis we found a greater number of Adamts5+ and Mmp13+ cells and a lower number of Acan+ and Sox9+ cells in cartilage from the HS-EVs injected mice compared to the LS-EV-injected controls (Fig. S3H, I).
To mimic abnormal mechanical loading during OA progression in vitro, mouse long-bone osteocyte Y4 (MLO-Y4) cells were subjected to excessive mechanical stress at a frequency of 0.5 Hz and cyclic tensile strain of 20% for 24 hours. We then subsequently co-cultured these loaded MLO-Y4 cells with primary mouse chondrocytes. Furthermore, we simultaneously inhibited EV secretion from loaded MLO-Y4 cells using the neutral sphingomyelinase inhibitor GW4869 (Fig. S4A). By a nanoparticle tracking analysis (NTA) assay we found that loading of the MLO-Y4 cells promoted greater EV release, which was partially blunted by GW4869 treatment (Fig. S4B). By Alcian blue staining we found a lower intensity of proteoglycans in the chondrocytes co-cultured with loaded MLO-Y4 cells, which was strikingly normalized in chondrocytes that were co-cultured with GW4869-treated loaded MLO-Y4 cells (Fig. S4C). We found similar results, including normalization, for the expression of catabolic and anabolic genes and proteins (Fig. S4D–F).
To clarify the regulatory effect of MLO-Y4-derived EVs (MLO-Y4-EVs) induced by excessive mechanical stress on chondrocytes, we extracted MLO-Y4-EVs from MLO-Y4 cell lines under excessive mechanical stress (Load-EVs) or under a static condition (Static-EVs) and used each of these EVs to treat primary mouse chondrocytes (Fig. 2I). Before using the Static-EVs/Load-EVs we observed the morphology of isolated osteocyte-derived EVs by TEM (Fig. S4G) and particle size distribution by NTA (Fig. 2J), as well as by Western blotting for expression of EV markers Tsg101, Cd81, Cd9, Cd63 and Hsp70 (Fig. S4H). These results showed that EV secretion increased in the Load group compared to the Static group. We firstly isolated MLO-Y4-EVs from stable transgenic MLO-Y4 cell lines expressing the Cd63-GFP fusion protein and then treated primary mouse chondrocytes in vitro with the labeled MLO-Y4-EVs, observing their successful uptake by the chondrocytes (Fig. 2K). By Alcian blue staining we found that the intensity of proteoglycans was markedly lower in the chondrocytes after treatment with Load-EVs compared to those treated with Static-EVs (Fig. 2L). Furthermore, we found the mRNA and protein expression levels of catabolic markers (Adamts5/Adamts5 and Mmp13/Mmp13) were significantly higher, while the levels of anabolic markers (Col2a1/ColII, Acan/Acan and Sox9/Sox9) were significantly lower, in chondrocytes after treatment with Load-EVs compared to those treated with Static-EVs (Figs. S4I, 2M and S4J). Furthermore, to simulate the inflammatory environment of primary mouse chondrocyte in OA, IL-1β was used to co-treat primary mouse chondrocytes along with the EVs (Fig. S5A). By qPCR analysis and immunofluorescence staining we found co-treatment with IL-1β and Load-EVs exacerbated the positive and negative effects on the expression of various catabolic (Adamts5/Adamts5 and Mmp13/Mmp13) and anabolic (Col2a1/ColII, Acan/Acan and Sox9/Sox9) markers, respectively, in the chondrocytes compared to IL-1β plus Static-EVs co-treatment (Fig. S5B–D).
To determine whether MLO-Y4-derived EVs after mechanical loading exacerbate OA progression in vivo, equal amounts of Static-EVs and Load-EVs were intraarticularly administered to DMM mice for 8 weeks (Figs. 2N and S4K). We observed fluorescent signals of PKH26-labeled MLO-Y4-derived EVs in chondrocytes at the cartilage surface in the DMM mice, indicating the injected EVs were successfully taken up by chondrocytes in vivo (Fig. 2O). Compared with the Static-EVs injection group, by Safranin O staining and OARSI scoring we found that cartilage degeneration was further aggravated in the Load-EVs-treated mice (Figs. 2P and S4L). In addition, by immunofluorescent analysis we found a greater number of Adamts5+ and Mmp13+ cells and a lower number of Acan+ and Sox9+ positive cells in the cartilage of Load-EVs injected mice compared to the Static-EVs-injected mice (Fig. S4M, N).
As our results so far demonstrated that osteocyte-derived EVs induced by excessive mechanical stress (HS-EVs or Load-EVs) during OA development exacerbate cartilage degeneration, we wondered whether specific blockade of EV secretion from osteocytes could antagonize retrograde degeneration of cartilage during OA progression. To test this possibility, we encapsulated GW4869 in our previously established bone-targeting drug delivery system (liposome-DSS6)20 to develop an osteocyte-targeting EV secretion inhibitor, i.e. lipoGW4869@DSS6 (Fig. S6A). The morphology of liposomes by TEM and particle size distribution by NTA (Fig. S6B). Consistent with our previous data, the liposome-DSS6 could specifically deliver DiR to bone tissues via systematic administration, as we found the femur and tibia exhibited high-intensity signals in mice injected with lipoDiR@DSS6 but not in mice injected with lipoDiR (without DSS6) (Fig. S6C). Subsequently, we administered lipoGW4869@DSS6 or lipoVehicle@DSS6 to DMM mice via tail vein injection every 3 days for 8 weeks (Figs. 2Q and S6D). We found that there were a fewer number of Cd63+ osteocytes in the SCB in mice treated with GW4869 compared to the vehicle-treated group, indicating EV secretion from osteocytes in the SCB was successfully blocked (Fig. 2R). Importantly, by Safranin O staining and OARSI scoring we found that there was significantly less cartilage degeneration in the GW4869-treated group compared to the control-treated group (Figs. 2S and S6E). In addition, GW4869 treatment attenuated cartilage catabolism and promoted cartilage anabolism in DMM mice (Fig. S6F, G).
Collectively, these data suggest that EVs secreted from osteocytes induced by excessive mechanical stress during OA development promoted a metabolic skewing in chondrocytes that was associated with their dysfunction.
Osteocyte-derived EVs containing miRNAs are responsible for chondrocyte dysfunction during OA progression
As miRNAs in EVs play a pivotal role in the regulation of recipient cells, we investigated whether the osteocytes can secrete extracellular miRNAs and transport them into chondrocytes to regulate chondrocyte function during OA progression. We firstly transfected MLO-Y4-Cd63-GFP stable cell lines with a Cy3-labeled miR-223 mimic, a myeloid cell-selective miRNA21, and subsequently co-cultured the labeled MLO-Y4 cells with primary mouse chondrocytes in a transwell plate (Fig. 3A). We found that the osteocyte-derived EVs could deliver the Cy3-miR-223 mimic to the cocultured chondrocytes, as evidenced by colocalization of the Cy3 red fluorescent signal with the GFP green fluorescent signal in chondrocytes (Fig. 3A). Quantitatively, the miR-223 expression level in chondrocytes was increased by ~6-fold after co-culturing with the above indicated labeled MLO-Y4 cells (Fig. 3B).
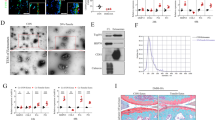
It has been previously shown that knocking out Dicer1, which encodes for the endoribonuclease Dicer that is necessary for miRNA biogenesis, can abolish the expression of most intracellular miRNAs22. To further clarify whether miRNAs are involved in the inhibitory effects of osteocyte-derived EVs on chondrocytes, we extracted EVs from either primary mouse osteocytes from wide-type (WT-EVs) or osteocyte-specific Dicer1 knockout (cKO-EVs) mice (Fig. 3C). The flow cytometry analysis showed that 85.5% of the isolated cells were osteocytes (Fig. S7A). Moreover, the western blot indicated that the protein expression of Dicer was markedly lower in osteocytes isolated from Dicer-cKO mice (Fig. 3D). Then, loaded them and used them to treat primary mouse chondrocytes (Fig. 3E). By Alcian blue staining for proteoglycan in chondrocytes we found that loaded WT-EVs resulted in their strikingly lower levels, which was not observed with the loaded cKO-EVs (Fig. 3F). Likewise, the mRNA and protein expression levels of catabolic markers (Adamts5/Adamts5 and Mmp13/Mmp13) was significantly lower, while the levels of anabolic markers (Col2a1/ColII, Acan/Acan and Sox9/Sox9) was significantly higher in chondrocytes after treatment with loaded cKO-EVs than those treated with loaded WT-EVs (Fig. 3G, H).
To further investigate the contribution of osteocyte-derived miRNAs to OA progression in vivo, we established a DMM OA model in WT and Dicer-cKO mice (Fig. 3I). By Western blot analysis we confirmed that the protein expression of Dicer was markedly lower in osteocytes in Dicer-cKO mice (Fig. 3J). Further, by qPCR analysis we confirmed that most of the intracellular miRNA expression in osteocytes was strikingly lower in Dicer-cKO mice compared to WT mice (Fig. 3K). By Safranin O staining and OARSI scoring we found that cartilage degeneration in the Dicer-cKO mice was partially alleviated compared to those in WT mice (Fig. 3L, M). Consistent with these findings, by immunofluorescence staining we found that there was significantly less Adamts5+ and Mmp13+ cells and a significantly greater number of Acan+ and Sox9+ cells in the cartilage from Dicer-cKO mice compared to these from WT mice (Fig. 3N, O). These findings suggested that SCB osteocyte-derived miRNAs promote chondrocyte catabolism and inhibit their anabolism during OA progression.
MiR-23b-3p expression in osteocyte-derived EVs regulates cartilage metabolism during OA progression
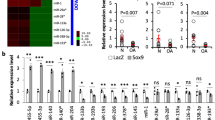
To identify the miRNA responsible for the inhibitory effects of osteocyte-derived EVs on cartilage metabolism, we performed microRNA sequencing analysis in LS-EVs and HS-EVs isolated from patients with OA, as well as plasma EVs from Dicer conditional knock out (Dicer-cKO) and WT mice (Fig. 4A). By doing so we identified 17 upregulated miRNAs in HS-EVs compared to LS-EVs from the patients and 23 miRNAs in plasma EVs from Dicer cKO mice that were lower than those from WT mice (Fig. 4B). Analysis of the overlapping miRNAs revealed that only miR-23b-3p was in common between the two groups (Fig. 4C). Subsequent qPCR analysis further verified the upregulated expression level of miR-23b-3p in HS-EVs compared to LS-EVs and that its levels were significantly lower in SCB osteocytes and cartilage in Dicer-cKO mice after DMM surgery compared to WT mice (Figs. 4D, E and S8A). Moreover, the miR-23b-3p levels in MLO-Y4 cells and MLO-Y4 derived EVs were also higher after loading compared with those in the Static group (Fig. S8B).
A A schematic diagram illustrating the separation of extracellular vesicles (EVs) and their enrichment from human tissues and mouse plasma. Heatmaps of relative miRNA differences (B) and enrichment analysis (C). D The miR-23b-3p levels in LS-EVs and HS-EVs (n = 5). E The miR-23b-3p levels in subchondral bone (SCB) osteocytes from WT and Dicer-cKO mice with destabilization of the medial meniscus (DMM) surgery (n = 5). F The miR-23b-3p levels in cartilage and SCB osteocytes from human LS/HS tissues (n = 20). G The miR-23b-3p levels in cartilage and SCB osteocytes from mice with DMM surgery (n = 6). H The pre-miR-23b-3p levels in cartilage and SCB osteocytes from human LS/HS tissues (n = 20). I The pre-miR-23b-3p levels in cartilage and SCB osteocytes from mice with DMM surgery (n = 6). J Alcian blue staining. Scale bar in upper panel, 1 mm. Scale bar in lower panel, 2 mm. K Western blot analysis of COLII, ACAN, SOX9, MMP13 and ADAMTS5 protein expression in primary human and mouse chondrocytes transfected with nc or mimic (n = 3). L The schematic depicted the overexpression of miR-23b-3p in cartilage through intraarticular injection of rAAV5-miR-23b-3p-mcherry (rAAV5-miR-23b-3p) or negative control (rAAV5-nc) in 8-week-old male mice following DMM surgery (n = 8). M The Safranin O images of cartilage. Scale bar in upper panel, 200 μm. Scale bar in lower panel, 100 μm. Data are presented as the mean ± SD. Data was analyzed by unpaired two-tailed Student’s t-test (D, E, G, I and K) and paired two-tailed Student’s t-test (F and H). Source data are provided as a Source Data file.
To further address whether this pathologically elevated miR-23b-3p in cartilage during OA originates from other tissues, we also measured the levels of the miRNA in different tissues apart from bone and cartilage of DMM mice. We found that there was no significant change of expression of miR-23b-3p in the liver, skeletal muscles, heart, lungs, spleen or kidneys between DMM mice and Sham control mice (Fig. S8C).
With regard to human relevance we found that miR-23b-3p levels in SCB osteocytes and chondrocytes were significantly higher in the HS region of knee joints from patients with OA compared to the LS region (Fig. 4F). Consistent with these clinical results, we also found that miR-23b-3p levels in SCB osteocytes and chondrocytes were significantly higher in DMM-induced OA mice compared with those in Sham mice (Fig. 4G). To examine whether the higher miR-23b-3p levels resulted from greater chondrocytic transcription, we measured primary miR-23b-3p expression in SCB osteocytes and cartilage. We found that the level of primary miR-23b-3p was significantly greater in SCB osteocytes in the HS region of human OA samples or DMM mice compared to the LS region or Sham control mice, respectively, but there was no difference in the cartilage in either species (Fig. 4H, I). Next, we detected the expression level of miR-23b-3p in cartilage from mice after intra-articular injection of EVs isolated from knee joint SCB of patients with OA or EVs isolated from MLO-Y4 cells. We found that the miR-23b-3p levels in cartilage from HS-EVs- or Load-EVs treated mice were significantly higher than those from LS-EVs- or Static-EVs-treated mice after injection, respectively (Fig. S8D, E). In addition, the injection of GW4869 (to inhibit osteocyte-EV release) led to significantly lower miR-23b-3p levels in the cartilage of DMM mice (Fig. S8F). These data indicate that the levels of miR-23b-3p in the cartilage during OA progression is determined by osteocyte-derived EVs that harbor the miRNA.
To investigate the functional role of miR-23b-3p in chondrocytes in vitro, we transfected primary human osteoarthritic chondrocytes or primary mouse chondrocytes with either a negative control (nc) or a miR-23b-3p mimic (mimic) (Fig. S9A). We detected a ~ 33-fold increase of miR-23b-3p expression in primary human osteoarthritic chondrocytes and a ~ 37-fold increase of miR-23b-3p expression in primary mouse chondrocytes after transfection, respectively (Fig. S9B). By Alcian blue staining we found that the intensity of proteoglycan was significantly lower in chondrocytes transfected with the mimic compared to the nc group in both species (Fig. 4J). In both species the mRNA and protein levels of ADAMTS5/ADAMTS5 and MMP13/MMP13 were significantly higher, while the levels of COL2A1/COLII, ACAN/ACAN and SOX9/SOX9 in chondrocytes were lower after transfection with mimic compared to transfection with nc (Figs. 4K, S9C and S9D).
To determine whether miR-23b-3p in chondrocytes exerted harmful effects on cartilage metabolism in vivo, we specifically overexpressed miR-23b-3p in chondrocytes by intra-articular injection of adeno-associated virus 5-miR-23b-3p-mCherry (rAAV5-miR-23b-3p) in C57BL/6 mice and the tissue was collected for analysis 8 weeks after surgery (Fig. 4L). We found that there was red fluorescence in the injected mice and that the levels of miR-23b-3p expression were higher in the cartilage of the rAAV5-miR-23b-3p-injected DMM mice compared to the nc-injected control DMM mice, indicating successful transfection of rAAV5 into cartilage (Fig. S9E, F). By Safranin O staining and OARSI scoring we found that there was more severe cartilage degeneration in both the Sham and DMM mice after injection with rAAV5-miR-23b-3p compared with their corresponding control groups, indicating that even in the absence of wounding overexpression of the miRNA can alter chondrocyte function (Figs. 4M and S9G). Furthermore, by immunofluorescent analysis we found that greater miR-23b-3p expression was associated with greater cartilage catabolism and less anabolism based on relevant protein expression (Fig. S9H, I). These findings suggested that elevated miR-23b-3p in chondrocytes exacerbates cartilage degeneration during OA progression.
Gain- and loss-of function studies confirm that miR-23b-3p in osteocyte-derived EVs leads to chondrocyte dysfunction during OA progression
To further investigate the functional role of miR-23b-3p delivered to chondrocytes by osteocyte-derived EVs in vitro, we isolated EVs from MLO-Y4 cells transfected with miR-23b-3p mimics (mimic-EVs) under static conditions, miR-23b-3p inhibitor (inhibitor-EVs) following loading and their corresponding negative controls (static-nc-EVs or load-nc-EVs) (Fig. 5A). We found that the levels of miR-23b-3p in MLO-Y4 cells and EVs, as well as in chondrocytes transfected with miR-23b-3p mimics, were significantly higher than the cells treated with negative controls (Fig. S10A, B). By Alcian blue staining we found that the intensity of proteoglycan was significantly lower in chondrocytes after transfection with mimic-EVs compared to nc-EVs, while transfection with the inhibitor-EVs had the opposite effect (Fig. 5B). Moreover, the mRNA and protein levels of Adamts5/Adamts5 and Mmp13/Mmp13 were significantly higher, while the levels of Col2a1/ColII, Acan/Acan and Sox9/Sox9 were significantly lower, in chondrocytes after exposure to mimic-EVs than nc-EVs, while treatment with the inhibitor-EVs had the opposite effects (Figs. 5C, D and S10C).
A A schematic diagram illustrating the use of extracellular vesicles (EVs) derived from MLO-Y4 cells transfected with either a negative control (nc mimic-EVs) or a miR-23b-3p mimic (mimic-EVs), as well as EVs obtained from loaded MLO-Y4 cells transfected with either a negative control (nc inhibitor-EVs) or a miR-23b-3p inhibitor (inhibitor-EVs), which were subsequently added to primary mouse chondrocytes. B Alcian blue staining. Scale bar, 2 mm. C Western blot analysis of ColII, Acan, Sox9, Mmp13 and Adamts5 protein expression in primary mouse chondrocytes (n = 3). D qPCR analysis of the Col2a1, Acan, Sox9, Mmp13 and Adamts5 levels in primary mouse chondrocytes (n = 3). E A schematic diagram illustrating mouse study in which the Dmp1cre mice were administered tail vein injection with rAAV9DSS6-FLEx-miR-23b-3p (rAAV9DSS6-miR-23b-3p) or rAAV9DSS6-FLEx-nc (rAAV9DSS6-nc) in destabilization of the medial meniscus (DMM) or sham surgery mice, to specifically overexpress miR-23b-3p expression in osteocytes (n = 7). F Safranin O images (n = 7). Scale bar in upper panel, 200 μm. Scale bar in lower panel, 100 μm. G The Osteoarthritis Research Society International (OARSI) scores (n = 7). H A schematic diagram illustrating the mouse study in which Dmp1cre mice were administered tail vein injection with rAAV9DSS6-FLEx-miR-23b-3p-RNAi (rAAV9DSS6-miR-23b-3p-RNAi) or rAAV9DSS6-FLEx-nc (rAAV9DSS6-nc) in DMM or sham surgery mice, to specifically antagonize miR-23b-3p expression in osteocytes (n = 7). I Safranin O images. Scale bar in upper panel, 200 μm. Scale bar in lower panel, 100 μm. J The OARSI scores (n = 7). Data are presented as the mean ± SD. P values are from unpaired two-tailed Student’s t-test (D), Kruskal–Wallis test followed by two-tailed Mann–Whitney U test (G and J). Source data are provided as a Source Data file.
To further investigate the impact of miR-23b-3p-containing osteocyte-derived EVs on cartilage degeneration in vivo, we utilized the cre-loxp system to generate mice with osteocyte-specific overexpression/inhibition of miR-23b-3p. To specifically overexpress miR‑23b‑3p in osteocytes, the Dmp1cre mice with Sham/DMM surgery were injected with rAAV9DSS6-miR-23b-3p or rAAV9DSS6-nc (Figs. 5E and S10D). By qPCR analysis we found greater miR‑23b‑3p expression in chondrocytes and SCB osteocytes in rAAV9DSS6-miR‑23b‑3p mice compared to the rAAV9DSS6-nc mice (Fig. S10E). Following DMM surgery, we found by Safranin O staining and OARSI scoring that there was more severe cartilage degeneration in mice injected with rAAV9DSS6miR‑23b‑3p compared to those injected with rAAV9DSS6-nc (Fig. 5F, G). Furthermore, by immunofluorescent analysis we found that osteocyte-specific miR-23b-3p overexpression promoted cartilage catabolism and inhibited anabolism compared to the control group after DMM surgery (Figs. S10F and S10G).
To specifically antagonize the expression of miR-23b-3p in osteocytes, the Dmp1cre mice with Sham/DMM surgery were injected with rAAV9DSS6-miR-23b-3p RNAi-GFP or rAAV9DSS6-nc (Fig. 5H). We found green fluorescent staining in osteocytes, indicating the successful transfection of rAAV9DSS6-miR-23b-3p RNAi into the osteocytes (Fig. S10H). Following DMM surgery, by Safranin O staining and OARSI scoring we found that there was less severe cartilage degeneration in mice injected with rAAV9DSS6-miR-23b-3p RNAi compared to those injected with rAAV9DSS6-nc at 8 weeks (Fig. 5I, J). Furthermore, by immunofluorescent analysis we found significantly less Adamts5+ and Mmp13+ cells and a significantly greater number of Acan+ and Sox9+ cells in cartilage from mice injected with rAAV9DSS6-miR-23b-3p RNAi compared to those injected with rAAV9DSS6-nc after DMM surgery (Fig. S10I, J). Collectively, these findings suggest that osteocytes-derived miR‑23b‑3p aggravates OA progression by enhancing catabolic activity and inhibiting anabolic activity of chondrocytes in vivo.
MiR-23b-3p inhibits mitophagy via targeting Otud4 to skew chondrocyte metabolism
To gain further insights into the underlying mechanisms by which miR-23b-3p regulates chondrocyte function during OA progression, we conducted RNA-sequencing analysis on human primary osteoarthritic chondrocytes infected with negative control (nc) or a miR-23b-3p mimic (mimic). Notably, by Kyoto Encyclopedia of Genes and Genomes (KEGG) and GO analysis we found that most downregulated genes were enriched in the mitophagy pathway (Fig. 6A). Previous studies have suggested that the impaired mitophagy could contribute to cartilage degradation by promoting the accumulation of defective mitochondria23,24. By TEM analysis we found that there was a greater number of autolysosome-like structures in chondrocytes in the HS region compared to those in the LS region of cartilage from knee joints of patients with OA, indicating a potential association between OA progression and inhibition of mitophagy processes (Fig. 6B).
A Gene Ontology (GO) enrichment and the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis of potential targets for regulating cartilage metabolism from downregulated genes from human primary osteoarthritic chondrocytes transfected with negative control (nc) or miR-23b-3p mimics (mimic) using RNAseq. B Representative electron microscopy images of mitophagy in chondrocytes from low stress (LS) and high stress (HS) areas of clinical samples. Scale bars, 1 μm. C Mitophagy detection. Scale bars, 100 μm. D JC-1 staining. Scale bars, 50 μm. E Alcian blue staining. Scale bar, 1 mm. F Western blot analysis of PARKIN, PINK, MFN2, LC3B, DRP1, COLII, ACAN, SOX9, MMP13 and ADAMTS5 protein expression in human primary osteoarthritic chondrocytes (n = 3). G Venn diagram of miR-23b-3p targets. H qPCR analysis of OTUD4 levels in human samples (n = 12), Sham/destabilization of the medial meniscus (DMM) mice samples (n = 6), chondrocytes with nc/mimic transfection (n = 3) and chondrocytes with nc/inhibitor transfection (n = 3). I The representative confocal images of OTUD4 of cartilage in human samples (n = 10). Scale bars, 100 μm. J The representative confocal images of Otud4 of cartilage in Sham or DMM mice (n = 6). Scale bars, 50 μm. K Dual luciferase reporter activities after transfecting 239 T cells with miR-23b-3p and reporter carrying 3’ UTR in the long form of OTUD4 (n = 6). L Mitophagy detection. Scale bars, 100 μm. M JC-1 staining. Scale bars, 50 μm. N Alcian blue staining. Scale bar, 1 mm. O Western blot analysis of OTUD4, PARKIN, PINK, MFN2, LC3B, DRP1, COLII, ACAN, SOX9, MMP13 and ADAMTS5 protein expression in vehicle- and Lv-OTUD4-treated human primary osteoarthritic chondrocytes transfected with nc or mimic (n = 3). Data are presented as the mean ± SD. Data analyzed by paired two-tailed Student’s t-test (I), unpaired two-tailed Student’s t-test (H), one-way analysis of variance (ANOVA) with Tukey’s multiple comparisons test (J and K) or Hypergeometric test (A). Source data are provided as a Source Data file.
To further investigate whether miR-23b-3p promotes cartilage catabolism and inhibits cartilage anabolism by impairing mitophagy in chondrocytes, we transfected human primary osteoarthritic chondrocytes with either a miR-23b-3p mimic or a negative control (nc) followed by treatment with or without the mitophagy agonist Urolithin A (UA). We observed less colocalization of the mitophagy dye and the lysosome dye in human primary osteoarthritic chondrocytes transfected with the miR-23b-3p mimic compared to the nc group (Fig. 6C). In addition, we performed JC-1 staining to assess mitochondrial membrane potential (MMP) in chondrocytes, by which the red fluorescence represents JC-1 aggregates appearing in mitochondria after potential-dependent aggregation, while the green fluorescence represents JC-1 monomers appearing in the cytosol after mitochondrial membrane depolarization. Human primary osteoarthritic chondrocytes transfected with the miR-23b-3p mimic exhibited a greater extent of JC-1 aggregates (red) and a lower number of JC-1 monomers (green) compared to those in the nc group (Fig. 6D). By Alcian blue staining we found that the intensity of proteoglycans was significantly lower in chondrocytes transfected with the miR‑23b‑3p mimic compared to the nc control, which was rescued by UA treatment (Fig. 6E). Furthermore, by Western blot analysis we found that the expression of mitophagy-related markers, including PARKIN, PINK, MFN2 and LC3B, were significantly lower, while DRP1 was significantly higher, in human primary osteoarthritic chondrocytes transfected with the miR‑23b‑3p mimic compared with the nc-transfected group (Figs. 6F and S11A). However, UA treatment alleviated the inhibitory effects of miR‑23b‑3p on mitophagy and reversed the decrease in expression of COLII, ACAN and SOX9 proteins, as well as attenuating the increase in ADAMTS5 and MMP13 by miR‑23b‑3p overexpression (Figs. 6F and S11A).
To further elucidate the target genes of miR-23b-3p that lead to the impairment of mitophagy in chondrocytes, we identified four genes (OTUD4, VCAN, NRK and GREM1) by combining down-regulated differentially expressed genes (Basemean >1000) and multiple databases, including Targetscan, miRbase and miRDB (Fig. 6G). Subsequently, we examined the expression levels of these four genes in various samples. Of note, only OTUD4 exhibited significantly lower expression not only in chondrocytes infected with the mimic but also in chondrocytes incubated with HS-EVs or Load-EVs (Figs. 6H and S11B–E). These results suggest that miR-23b-3p targets OTUD4 to regulate chondrocyte function. Furthermore, by immunofluorescent analysis we found a notable reduction in the number of OTUD4+ cells within the cartilage at the HS region compared to the LS region in knee joints from patients with OA (Fig. 6I). Consistent with these clinical results, we found that in DMM mice there was progressively lower OTUD4 expression at 8 weeks after DMM surgery compared to 2 weeks and 4 weeks after surgery (Fig. 6J).
To confirm whether miR-23b-3p directly targets OTUD4 in chondrocytes, we utilized a dual luciferase assay to demonstrate efficient binding of miR-23b-3p to the 3’ UTR of OTUD4 (Fig. 6K). To further investigate whether miR-23b-3p regulates cartilage metabolism by targeting OTUD4 to impair mitophagy, we transfected human primary osteoarthritic chondrocytes with nc or mimic following treatment with a lentivirus carrying either non-targeting control sequence (Lv-nc) or lentivirus overexpressing OTUD4 (Lv-OTUD4). Notably, the supplementation of OTUD4 alleviated the inhibition of mitophagy by miR-23b-3p as evidenced by mitophagy detection assays (Fig. 6L), JC-1 staining (Fig. 6M). Additionally, by Alcian blue staining the intensity of proteoglycans was strikingly lower in chondrocytes transfected with the mimic compared to the nc-transfected cells, which was rescued by Lv-OTUD4 treatment (Fig. 6N). More importantly, Western blot analysis indicated that the replenishment of OTUD4 alleviated the decrease in expression of COLII, ACAN and SOX9 proteins and the increase in expression of ADAMTS5 and MMP13 induced by miR-23b-3p (Figs. 6O and S11F). Taken together, these results suggest that miR-23b-3p in osteocytes-derived EVs directly targets OTUD4 to suppress chondrocyte function during OA progression.
Therapeutic inhibition of miR-23b-3p in chondrocytes or osteocytes alleviates cartilage degeneration during OA progression
To assess the beneficial effects of miR-23b-3p inhibition in chondrocytes on cartilage degeneration during OA progression, we intra-articularly administered antagomiR-23b-3p into the knee joints of 8-week-old DMM mice every two weeks starting at day 2 after the surgery (Fig. 7A). By immunofluorescent and PCR analysis, the increased Otud4 expression in the cartilage in the antagomiR-treated mice compared to the control-treated mice confirmed the successful antagonistic effect of antagomiR-23b-3p (Fig. 7B–D). By determining mitophagy number, as well as by Safranin O staining and OARSI scoring, we found that antagomiR treatment resulted in a greater degree of mitophagy and a more normal cartilage matrix at 8 weeks after surgery compared to the control-treated mice (Fig. 7E–H). Furthermore, by immunofluorescent analysis we found that there was a significantly greater number of Acan+ and Sox9+ cells and a significantly lower number of Adamts5+ and Mmp13+ cells in the cartilage in the antagomiR-treated mice compared to the control-treated mice (Fig. 7I, J). To further evaluate whether specific inhibition of miR-23b-3p in osteocytes by antagomiR-23b-3p could attenuate cartilage degeneration, we intravenously injected antagomiR‑23b‑3p encapsulated in liposome-DSS6, i.e. lipoantagomiR‑23b‑3p@DSS6, into DMM or Sham mice using the same treatment regimen as for the chondrocyte targeting above (Fig. 7K). By immunofluorescent and PCR analysis, the increased Otud4 expression in the cartilage in the lipoantagomiR‑23b‑3p@DSS6-treated mice compared to the control-treated mice confirmed the inhibition effect of antagomiR-23b-3p (Fig. 7L–N). After a 8-week treatment period we observed greater mitophagy and less cartilage degeneration in chondrocytes from the lipoantagomiR‑23b‑3p-treated DMM mice compared to control-treated group (Fig. 7O–R). Additionally, by immunofluorescent analysis we found that there was an elevation in Acan+ and Sox9+ cells, as well as fewer Adamts5+ and Mmp13+ cells, within the articular cartilage of DMM micinjected with lipoantagomiR‑23b‑3p@DSS6 compared to the control mice (Fig. 7S, T).
A A schematic diagram illustrating the intra-articular injection of antagomiR-23b-3p or antagomiR-nc in sham/destabilization of the medial meniscus (DMM) mice. Representative confocal images (B) and the quantitative analysis (C) of Otud4 in the articular cartilage (n = 7). Scale bar, 50 μm. D The qPCR analysis of Otud4 mRNA expression in chondrocytes (n = 3). E Representative electron microscopy images of mitophagy in chondrocytes. Scale bars, 500 nm. F The numbers of mitophagy in chondrocytes (n = 3). G Safranin O images. Scale bar in upper panel, 200 μm. Scale bar in lower panel, 100 μm. H The Osteoarthritis Research Society International (OARSI) scores (n = 7). Representative confocal images (I) and the quantitative analysis (J) of Acan, Sox9, Mmp13 and Adamts5 in the articular cartilage. Scale bar, 50 μm. (n = 7). K A schematic diagram illustrating the workflow by which osteocyte-targeted antagomiR-23b-3p (LipoantagomiR-23b-3p@DSS6) or nc (Liponc@DSS6) was performed in DMM or sham mice. Representative confocal images (L) and the quantitative analysis (M) of Otud4 in the articular cartilage. Scale bar, 50 μm. (n = 7). N The qPCR analysis of Otud4 mRNA expression in the articular cartilage (n = 3). O Representative electron microscopy images of mitophagy in chondrocytes. Scale bars, 500 nm. P The numbers of mitophagy in chondrocytes (n = 3). Q Safranin O images. Scale bar in upper panel, 200 μm. Scale bar in lower panel, 100 μm. R The OARSI scores (n = 7). Representative confocal images (S) and the quantitative analysis (T) of Acan, Sox9, Mmp13 and Adamts5 in the articular cartilage (n = 7). Scale bar, 50 μm. Data are presented as the mean ± SD. P values are from Kruskal–Wallis test followed by two-tailed Mann–Whitney U test (H and R) and one-way analysis of variance (ANOVA) with Tukey’s multiple comparisons test (C, D, F, J, M, N, P and T). Source data are provided as a Source Data file.
Discussion
The degeneration of articular cartilage is the primary concern during OA progression, yet its metabolic homeostasis and structural integrity rely on the biochemical and biomechanical interplay with the SCB and the other joint tissues25. Here, we uncovered a previously unrecognized cross-talk between the SCB and cartilage in OA, in which the disrupted mechanical loading induces osteocytes in the SCB to secret EVs, which transfer their cargo of miRNAs, especially miR-23b-3p, to chondrocytes, subsequently promoting cartilage catabolism while inhibiting their anabolism during OA progression (Fig. 8).
Created in BioRender. Shi (2025) https://BioRender.com/9xrn222.
Long-term abnormal mechanical stress can lead to changes in the bone remodeling process of the SCB. Excessive or uneven loading may cause bone sclerosis, further exacerbating OA progression26. Clinically, most OA patients have knee varus alignment, and the load on the medial side of the knee joint is higher than that on the lateral side. To clarify the stress distribution of the SCB, we choose a typical OA patient (varus alignment) with characteristic X-ray images, MRI features, and pathological markers, reflecting the general pathological process of OA and aligning with the core objectives of our study. The Finite element analysis in this study is used to reveal that the force on the medial side of the knee joint in OA patients is higher than that on the lateral side. It is worth noting that several studies have shown that the finite element modelling method was based on data from one patient27,28,29,30.
As the key mechanosensory and endocrine cell in the bone, osteocytes can sense and transmit biomechanical signals and secrete a variety of bioactive factors that mediate inter-organ crosstalk between the bone and other organs, including the cartilage31. For instance, osteocyte-derived factors, such as Dmp1, Sost, Mmp13 and Igf1, are involved in the regulation of SCB remodeling and cartilage degeneration12,14,32,33. In this study, we found enhanced cartilage catabolism and reduced cartilage anabolism in primary human chondrocytes after co-culturing with SCB osteocytes from the HS region compared to the LS region of knee joints from patients with OA. It is noteworthy that the human primary osteocyte cultures used in our co-culture system were osteocyte-enriched SCB fragments after removal of surface cells using collagenase, a culture method that has been reported in previous studies34,35. The results of the coculture assay encouraged us to further investigate whether the osteocytes in the SCB could release certain mediators that affect articular cartilage function during OA progression. Subsequent RNA-sequencing and GO enrichment analysis revealed the enrichment of the EV biogenesis pathway within osteocytes in the SCB at the HS region, which is consistent with previous findings that mechanical stress stimulation, including fluid shear and tension, can promote EV secretion in different cells36,37,38. We further used the Flexcell Tension Plus system to construct a mechanical overload model in vitro to mimic abnormal mechanical loading in OA progression and observed a marked rise in the secretion of EVs from loaded osteocytes. Therefore, we speculated that osteocytes in the SCB under mechanical stress may secrete EVs to impair chondrocyte function as well as to mediate cross-talk between the SCB and cartilage during OA progression.
EVs, which are highly heterogeneous in their biomolecular composition, act as insoluble mediators to specifically regulate cell-cell crosstalk39,40. There is growing evidence that osteocyte-derived EVs can mediate the communication from the bone to adjacent tissues or distal organs, such as the brain41, blood vessels18 and the prostate42. However, to the best of our knowledge no studies have been conducted to determine whether osteocyte-derived EVs are involved in the crosstalk between the SCB and cartilage, which is of great challenge and interest to study. Our findings here demonstrate that Cd63-labeled EVs derived from osteocytes in the SCB could reach the overlying cartilage, particularly in the deep zone of cartilage in DMM mice but not in sham mice. Interestingly, it has been reported that neovascularization as well as microcracks occurs in the SCB, which could facilitate the transfer of osteocyte-derived EVs from the SCB to cartilage during OA progression43. Consistent with this notion, Liu et al. showed that osteoclast-derived EVs in the SCB can carry miR-214 through blood vessels and microcracks to the calcified cartilage layer11. Interestingly, we also found the Cd63-labeled osteocyte-EVs in the SCB marrow cavity associated with SCB endomucin+ vessels, particularly the vessels at the osteochondral junction in DMM mice. Furthermore, we obtained LS- and HS-EVs or Static- and Load-EVs and further confirmed their purity via the detection of their morphology, diameter distribution and surface marker expression profiles. A previous study reported that SCB osteocyte dysfunction reduced cartilage proteoglycan content and promoted cartilage degeneration, consistent with early OA14. In this study, we demonstrated that HS-EVs or Load-EVs could impair chondrocyte function and accelerate cartilage degradation during OA progression. Moreover, specific blockade of osteocyte-EVs generation alleviated OA progression. In our study, GW4869 was used for inhibiting the osteocyte-EVs generation. A study has shown that the lipid composition of EVs is similar to that of lipid rafts and is rich in cholesterol, sphingolipids, saturated lecithin and ceramide. Ceramide can promote the budding of exosomal vesicles into multivesicular endosomes44. GW4869 specifically inhibits nSMase2, an enzyme responsible for hydrolyzing sphingomyelin into ceramide. Ceramide is a key lipid molecule involved in the budding of multivesicular bodies (MVBs), the precursors of EVs44. By inhibiting nSMase2, GW4869 reduces ceramide production, thereby affecting EV formation and secretion. Several studies have used GW4869 as an inhibitor of EVs secretion45,46,47. In summary, GW4869 is specific for inhibiting EVs secretion due to its selective inhibition of nSMase2, and it does not broadly affect other sphingomyelinases or vesicle types like microvesicles, making it a precise tool for studying EV-related processes.
EVs are now recognized not only as important carriers of proteins, lipids and nucleic acids in the plasma, but also as being capable of transferring such cargo to recipient cells48,49. More importantly, most EVs can selectively encapsulate miRNAs, a class of single-stranded non-coding RNAs of 19 to 22 nucleotides, and then release them into the circulation to mediate interorgan communication by regulating gene expression in distant cells50,51,52. In our study, we demonstrated that the miRNAs encapsulated in osteocyte-EVs are mainly responsible for the functional depression of chondrocytes in OA progression, as evidenced by the findings from Dicer1 cKO mice. It is worth noting that our flow cytometry analysis stated 85.5% of the cells were osteocytes and only 0.45% were osteoblast in the extraction of primary osteocytes. Therefore, there may be a slight effect from osteoblasts. Following miRNA sequencing, we identified miR-23b-3p, a highly conserved vertebrate miRNAs involved in various biological processes53, as a miRNA that is upregulated in HS-EVs from patients with OA, indicating that miR-23b-3p may be the key miRNA in osteocyte-derived EVs responsible for their inhibitory effects on cartilage metabolism in OA progression. We further verified that miR-23b-3p levels in both HS-EVs and Load-EVs were significantly upregulated. Recently, it has been reported that mechanical stimulation can induce EVs of Schwann cells to selectively package miR-23b-3p and transfer them to neurons to promote their regeneration54. Combined with our results, there may be a selective mechanism by which EVs can encapsulate miR-23b-3p after mechanical stimulation, which will be of keen interest to explore in a future study. Furthermore, we detected the elevation of miR-23b-3p in OA damaged cartilage tissues related to matched smooth cartilage tissues, which is in accord with previous studies55,56 and large-scale microarray miRNA analysis57. Importantly, inhibition of miR-23b-3p in osteocytes prevented cartilage degeneration in DMM-induced OA mice. These results indicate that miR-23b-3p may function as an important mediator in osteocyte-EVs with mechanical loading that promotes the early phases of OA progression. Interestingly, in previous studies, miR-23 cluster knockout did not accelerate the severity of OA in mouse aging and post-traumatic models58. The inconsistencies across the published studies and ours may be partly explained by different animal models.
It has been shown that miRNAs exhibit their function mainly based on binding their target genes59. To further understand the mechanisms underlying miR-23b-3p mediated OA progression, we analyzed the transcriptome of human primary osteoarthritic chondrocytes infected with negative control (nc) or miR-23b-3p mimics (mimic) using RNA-sequencing. By subsequent GO and KEGG analyses we found an enrichment of the mitophagy pathway. A previous study has indicated that mitophagy is impaired in several age‐related diseases, such as degenerative joint disorders24,60. Mitophagy impairment gives rise to the progressive accumulation of defective mitochondria, leading to cartilage degeneration61. In our study, a miR-23b-3p mimic repressed the mitophagy of cartilage, which was rescued by UA treatment. A study has reported that UA improves mitochondrial health, reduces cartilage degeneration, and alleviates pain in OA62. Interestingly, we observed an increase in mitochondrial membrane potential after miR-23b-3p mimic treatment. Previous studies have suggested that excessive mitochondrial membrane potential can result from an overly polarized inner mitochondrial membrane63,64. This excess membrane potential may, in fact, inhibit mitophagy, particularly in the PINK1/PARKIN pathway65. Further combined analysis of downregulated genes belonging to these biological functions and online bioinformatics databases confirmed the role of miR-23b-3p in regulating Otud4 expression. Previous studies have shown that reduction of miR-23b-3p can promote cartilage degeneration via targeting other genes, such as COL11A254 or HS6ST255. The differences of target genes across the published studies and ours may be partly explained by the use of different cells in vitro. Our study has demonstrated that miR-23b-3p inhibits mitophagy of chondrocytes by targeting Otud4, thereby promoting metabolic skewing of chondrocytes. It is now well established that the mitophagy machinery undergoes extensive post-translational modifications, such as phosphorylation/dephosphorylation, ubiquitination/deubiquitination, and acetylation/deacetylation, involving a series of enzymes66. The OTUD family (OTU domain-containing deubiquitinases family) is a type of deubiquitinating enzyme that has the function of removing ubiquitin tags on proteins and regulates many key physiological processes in cells, including protein degradation, signal transduction, stress response, inflammation and autophagy67,68,69,70,71. In our study, we found that overexpression of OTUD4 activated mitophagy to regulate cartilage metabolism. Interestingly, previous study has shown that OTUD5 inhibited ubiquitination-mediated degradation of VDAC2 and maintained mitochondrial homeostasis, subsequently ameliorating MASLD progression72. It is worth investigating in future research whether OTUD4 participates in the regulation of mitochondria through a similar signaling pathway.
Regarding potential translational to the clinic, the first small-interfering RNA, Patisiran, was recently granted FDA approval in 2018, and studies focused on the preclinical and clinical applications of microRNAs have expanded73. miRNA repressors have strong potential clinical utility as they act to silence the transcript function in peripheral tissues74. In the present study, we also delivered antagomiR-23b-3p via a bone-targeting system. The results showed that antagomiR-23b-3p effectively slowed the progression of OA in DMM mice, although the long-term systemic toxicity and potential off-target effects of antagomiR-23b-3p need to be further evaluated in future studies. As the SCB is altered earlier than cartilage during OA progress6,7,8, the abnormal upregulation of miR-23b-3p in osteocyte-EVs from the SCB could be used as a means for early diagnosis of OA, which is necessary to be further verified in a larger cohort of patients in the future. Moreover, inhibitors suppressing the expression of miR-23b-3p in osteocytes could be a novel direction for OA therapy. Taken together, these findings suggest that interfering with osteocyte-cartilage interactions by blocking osteocyte-derived EVs may be a potential therapeutic strategy for OA treatment.
Limitations of study
There are several limitations to this study that warrant further investigation. Firstly, despite our validation that miR-23b-3p is a key component of osteocyte-EVs that promotes cartilage degeneration, we cannot exclude the potential role of other secreted factors that are present in the EVs, which may also contribute to OA progression. Secondly, while we found that disrupted mechanical loading could affect the generation and secretion of EVs in SCB osteocytes, we have not yet investigated the specific molecular mechanism by which mechanical stress mediates the generation and secretion of EVs in osteocytes. Thirdly, future studies are planned to incorporate data from additional patients to further validate the findings and expand the generalizability of the model. Fourthly, while our current findings suggest a potential role for osteocyte-derived EVs in cartilage metabolism, further studies are needed to confirm the in vivo delivery of miR-23b-3p to chondrocytes. Moreover, we have no way to analyze the specific content of miRNA in each EV. Future studies may employ single EV analysis techniques to gain a deeper understanding of the distribution of miRNAs in different EVs. Lastly, considering that osteocyte-EVs can enter blood vessels, the role of osteocyte-derived exosomal miRNAs in mediating the cross-talk between osteocyte and osteoblast, and between osteocyte and macrophages, remains to be investigated. Collectively, we highlight and discuss current experimental limitations that need to be resolved and the state of the art in the cell biology of osteocyte-EVs and their possible functions.
Methods
Clinical samples
The tibial plateaus were collected from patients with OA who were hospitalized for Total knee arthroplasty (TKA) (age range from 56 to 68 years) in the Affiliated Nanjing Drum Tower Hospital of Nanjing University Medical School. All the samples were each separated by disease stage into macroscopically distinct sclerotic zones and non-sclerotic areas per sample. Morphologically, non-sclerotic regions were covered by cartilage without surface irregularities, whereas sclerotic regions were denuded or covered by severely degraded cartilage. For histology analysis, the human samples were fixed in 4% paraformaldehyde. For mRNA or protein analyses, samples are snap-frozen in liquid nitrogen prior to storing at −80 °C. For primary chondrocyte isolation and tissue EVs extraction, the fresh samples were immediately transferred from the operating room on ice and processed. Patients with OA were excluded if they have metabolic diseases. The human studies were conducted in accordance with the principles of the Declaration of Helsinki. All procedures were approved by the Nanjing Drum Tower Hospital Ethics Committees (No. NSFC81991514). Written informed consent was provided by all participants.
Animal models
All animals were kept in a specific-pathogen-free facility in individually ventilated cages in a room with a controlled environment (temperature, 20–24 °C; relative humidity, 30–70%; positive pressure) at five or six mice per cage, and the animals were in a 12-h day–night cycle. 6-8 weeks old mice underwent a surgically-induced destabilization of the medial meniscus (DMM) to model OA. In the DMM model, the medial meniscus was destabilized by transecting the medial meniscotibial ligament according to previously established protocols75. Age-matched mice were fed a standard normal diet (1010084, Xietong Shengwu) for the same weeks as control. The mice were randomly assigned in the DMM or Sham groups with various treatments. For the experiments in Figs. 1 and S1, the operated mice were euthanized at 2, 4 and 8 weeks after DMM surgery. For all other experiments, the operated mice were euthanized at 8 weeks after DMM surgery. All C57BL/6J male mice were purchased from Hangzhou Ziyuan Laboratory Animal Technology. All the animal experimental procedures were performed in accordance with the Laboratory Animal Care Guidelines approved by the Model Animal Research Center of Nanjing University. ARRIVE guidelines were adhered to in conducting this study. Regarding that male and female mice have physiological differences, including differences in hormonal levels. A study has indicated that most DMM-induced OA studies perform surgery on male mice76. In addition, young female mice were found to be less prone to developing OA following DMM compared to their male counterparts due to the perceived protective effect of sex hormone77. Hence, only male mice were used in our study.
Osteocyte-specific Cd63(loxp-eGFP) mice were constructed by using a Cd63-GFP floxed mouse mated with mice expressing Dmp1-Cre to generate Dmp1Cre; Cd63(loxp-eGFP) mice. Osteocyte-specific Dicer conditional knockout (Dicer-cKOfl/fl) mice were constructed by using a Dicer-cKO-floxed mouse mated with mice expressing Dmp1-Cre to generate Dmp1Cre; Dicer-cKOfl/fl mice. Both mice are on the C57BL/6 background. Eight-week-old Dmp1Cre; Cd63(loxp-eGFP) mice, Dmp1Cre; Dicer-cKOfl/fl mice and widetype (WT) littermate mice were used for experiments. Dmp1Cre; Cd63(loxp-eGFP) mice and Dmp1Cre; Dicer-cKOfl/fl mice were genotyped by PCR. Primers for Dmp1, Dicer and Cd63 were listed in Supplementary Table S1. Cd63(loxp-eGFP) mice were purchased from Shanghai Model Organisms. Dicer-cKO mice were purchased from GemPharmatech. Dmp1-Cre mice were a gift from Professor Junjie Gao.
To assess the effect of osteocyte-derived EVs on cartilage degeneration, EVs derived from human SCB osteocytes in low stress or high stress areas (LS/HS EVs) or MLO-Y4 (Procell Life Science & Technology) with or without loaded (Static/Load EVs) resuspended in PBS were sterilized with a 0.22 μm filter and intra-articularly injected into joint cavities (20 μg per mouse at each injection, 2 times a week) for 8 weeks post DMM surgery. To inhibit EVs biogenesis, mice were injected with neutral sphingomyelinase-2 (nSMase2) inhibitor GW4869 (2.5 mg/kg body weight, 3 times a week, 6823-69-4, Sigma) with the bone-targeted delivery system (LipoGW4869@DSS6) via tail vein injection for 8 weeks post DMM surgery.
For osteocyte-specific knockdown and overexpression of miR-23b-3p, the bone-targeting peptide motif DSS(Asp–Ser–Ser)6 was used to construct the bone-targeting recombinant AAV9 vector (rAAV9DSS6). rAAV9DSS6-miR-23b-3p and rAAV9DSS6-miR-23b-RNAi were provided by Genechem (Shanghai, China). Briefly, pre-miR-23b-3p and miR-23b-3p-RNAi were cloned into the CMV bGlobin-FLEX-MCS-WPREhGH polyA and CMV bGlobin-FLEX-EGFP-MCS-WPRE-hGH polyA vectors, respectively. After sequencing confirmation, then the vectors were packaged into capsids from rAAV9DSS6. A total of 200 μL of rAAV9DSS6-miR-23b-3p (4 × 1011 VG; 2 × 1013 VG/kg) and rAAV9DSS6-miR-23b-RNAi (8 × 1011 VG; 4 × 1013 VG/kg) were injected into Dmp1cre mice. The corresponding control mice were injected with the same amount of NC vector.
For cartilage-specific overexpression of miR-23b-3p, rAAV5-miR-23b-3p was intra-articularly injected into joint cavities. rAAV5-miR-23b-3p were provided by OBiO Technology. Briefly, pri-miR-23b-3p was cloned into the CMV-MCS-EF1amCherry-3xFLAG-WPRE. After sequencing confirmation, then the vectors were packaged into capsids from rAAV5. 10 μL of rAAV5-miR-23b-3p (1 × 1011 VG; 5 × 1012 VG/kg) was intra-articularly injected into knee joints of 8-week-old male mice.
For inhibition of miR-23b-3p in vivo, negative control antagomir (antagomir-NC) or antagomir-23b-3p were obtained from Generay (Shanghai, China). To antagonize the expression of miR-23b-3p in cartilage, the mice were intra-articularly injected into joint cavities with 10 µL antagomir-NC or antagomir-23b-3p for 8 weeks (2.5 mg/kg; once a fortnight) post DMM surgery. To antagonize the expression of miR-23b-3p in osteocytes, the mice were injected with 200 µL antagomir-23b-3p or antagomir-NC (10 mg/kg, once a fortnight) with the bone-targeted delivery system by tail vein injection.
Isolation and characterization of extracellular vesicles (EVs)
For tissue-EVs isolation, each human sample (consisted of 3 males and 3 females) from low stress (LS) or high stress (HS) SCBs were washed with HBSS to remove bone marrow and fragmented into 1 to 2 cm3 segments. After further crushing with a tissue homogenizer, these bone species were subjected to digestion by collagenase IA for overnight at 37 °C. Then the digested tissues were filtered through 70-µm nylon mesh membranes and centrifuged at 4500 x g for 30 min and 10000 x g for 30 min at 4 °C. The supernatant was collected for ultracentrifugation (UC) at 100000 x g for 100 min at 4 °C using L-80 XP ultracentrifuge (Beckman Coulter) and filtered sequentially through 0.22 μm filters (Millipore, Billerica, USA). Then, the resuspended solution was subjected to be further purified using Exosome Purification Kit (Ome-01E, Omiget) to harvest tissue-EVs (LS-EVs or HS-EVs). For osteocyte-EVs collection, the culture supernatants of MLO-Y4 cells and primary osteocytes were collected sequential centrifugation (300 x g for 10 min, 2000 x g for 30 min, and 10000 x g for 30 min) at 4 °C, then the supernatant was centrifuged at 100000 x g for 100 min at 4 °C using L-80 XP ultracentrifuge. PBS was used to resuspend the pellets.
For EVs characterization, western blotting was used to identify the expression of EV-specific proteins (CD81, CD9, CD63, TSG101 and HSP70). The number and size distribution of EVs were tested by nanoparticle tracking analysis (NTA). The morphologies of EVs were detected by a transmission electron microscope (TEM).
For EVs uptake assay in vitro, LS/HS-EVs were dyed with the PKH26 red fluorescent (P0096, Merck) following the manufacturer’s protocol. Static/Load-EVs were collected from the supernatant of Cd63-eGFP-MLO-Y4 stable cell lines (OBiO Technology) with or without loading. EVs were then added to primary chondrocytes and incubated at 37 °C for 12 h. After discarding the culture supernatant and washing the cells with PBS, the cells were fixed in 4% paraformaldehyde (PFA) for 15 min and incubated with DAPI to stain nuclei. Green phalloidin and red phalloidin were used to stain the cytoskeleton of primary human chondrocytes and primary mouse chondrocytes, respectively. For EVs uptake assay in vivo, PKH26 labeled EVs were intra-articularly injected into joint cavities of mice. 12 hours after EVs injection, articular cartilages were collected for further detection.
Alcian blue staining
To evaluate proteoglycan synthesis of the chondrocytes, the cells were washed with PBS and fixed with 4% formaldehyde for 10 min at room temperature. Then, the cells were washed three times with PBS and stained with Alcian Blue (ALCB-10001, OriCell) for 1 h. Cells were washed in PBS and images were captured. Qualitative analysis of staining results was conducted based on color depth. The darker the color, the higher the synthesis of proteoglycans. The same magnified field of view was selected for scanning.
Cell culture and treatment
All cells were grown at 37 °C in a humid atmosphere with 5% CO2.
Primary culture of mice osteocytes
Osteocytes were isolated from the mouse femurs and tibias as described previously78. Briefly, the bones (femurs and tibias) dissected were placed in α-MEM (SH30265.01, HyClone) + 1% penicillin/streptomycin (P/S). After removing the periosteum, epiphyses and bone marrow, the bones were further cut into 1 to 2 cm3 segments in DPBS, followed by digestion with 2 mg/mL collagenase type IA (9001-12-1, Sigma) in α-MEM supplemented with 1% P/S at 37 °C + 5% CO2 for 25 min and subsequent three washes with DPBS. The collagenase type IA-DPBS treatment was repeated twice again. The bone pieces were then incubated with 5 mM EDTA solution for 25 min and washed with DPBS, followed by digestion with collagenase type IA for another 25 min and subsequent three washes with DPBS. After EDTA–DPBS–collagenase type IA–DPBS treatment for another 2 times, the fractions containing abundant osteocytes could be obtained. After mincing the remaining bone pieces, the resulting bone particles and the osteocyte-enriched fractions were seeded into the collagen I-coated plates and incubated in α-MEM + 10% FBS + 1% P/S. Each primary osteocytes sample was derived from six male mice.
Mechanical strain loading on osteocytes
MLO-Y4 cells and primary osteocytes from WT or Dice-cKO mice were seeded into 6-well Bioflex plates (Flexcell International, Burlington, NC) at a density of 1 × 105 cells/well and subjected to cyclic stretch with 20% shape variable at a frequency of 0.5 Hz for 24 h by the Flexcell Tension Plus system (FX-4000 T, Flexcell International, Burlington, NC). The cell culture supernatant was collected for EVs extraction after 24 h.
Primary culture of human chondrocytes
Primary human chondrocytes were isolated and cultured according to previous protocols79. Cartilages of knee joints were obtained from OA patients undergoing TKA. Minced cartilage fragments were washed three times with PBS and digested with 0.2% collagenase II (17101015, Gibco) in Dulbecco’s Modified Eagle Medium/F12 (DMEM/F12, 11320033, Gibco) containing 10% FBS and 1% P/S at 37 °C + 5% CO2 overnight. The digested cell suspension was filtered through a 70 μm filter and cells were collected using centrifugation at 300 x g for 5 min. The chondrocytes were cultured in DMEM/F12 containing 10% FBS and 1% P/S. The medium was changed every 2 days. Cells at passage 1 were used to avoid phenotype loss. Each human chondrocytes sample was derived from three OA patients (2 males and 1 female).
For coculture experiment, human SCB samples from LS/HS areas were alternately digested with collagenase type IA and EDTA to obtain osteocyte-enriched bone fragments (LS/HS tissues). The LS/HS tissues were then cultured in a 0.4-μm pore sized filter which was placed above primary human chondrocytes. The primary human chondrocytes and LS/HS tissues share the same culture medium without direct cell-cell contact.
For LS-EVs or HS-EVs treatment, 1 × 105 primary human chondrocytes were incubated with 5 μg of LS-EVs or HS-EVs for qPCR assay (after 24 h), Alcian Blue staining and protein extraction (after 48 h).
For miRNA transfection, primary human chondrocytes were seeded into 12-well or 6-well plates and reached to 70–80% confluency. Then primary human chondrocytes were subjected to individual treatments of lentivirus-mediated overexpression of OTUD4 (Lv-OTUD4) (OBiO, China) or miR-23b-3p mimics, as well as a combined treatment of Lv-OTUD4 and miR-23b-3p mimics.
For experiments studying the effect of miR-23b-3p in chondrocytes, 50 uM Urolithin A (HY-100599, MedChemExpress) was used in our study to induce mitophagy. The primary human chondrocytes were subjected to individual treatments of UA or miR-23b-3p mimics, as well as a combined treatment of UA and miR-23b-3p mimics.
Primary culture of mice chondrocytes
The articular cartilage of C57BL/6 newborn mice were used to isolate primary mice chondrocytes as described previously80. Cartilage slices were first digested with trypsin for 15 min and then with 0.2% collagenase type II in DMEM/F12 containing 10% FBS and 1% P/S at 37 °C + 5% CO2 for 4 h. After isolation, the chondrocytes were cultured in DMEM/F12 containing 10% FBS and 1% P/S. The medium was changed every 2 days. Cells were used at passage 1 to avoid phenotype changes of primary mice chondrocytes. Each primary mice chondrocytes sample was derived from six newborn mice.
For coculture experiment, MLO-Y4 cells were seeded into 6-well Bioflex plates at a density of 1 × 105 cells/well and were pretreated with or without GW4869 (10 mM) 24 hours prior to co-culture. The same density of primary mice chondrocytes was seeded in the top chamber of the insert apparatus 6-well plate 6 hour prior to co-culture. When coculture, the MLO-Y4 cells were subjected to cyclic stretch with 20% shape variable at a frequency of 0.5 Hz for 24 h by the Flexcell Tension Plus system. qPCR assay (after 24 h), Alcian Blue staining (after 48 h) and protein extraction (after 48 h) were performed subsequently.
For static-EVs or load-EVs treatment, 5 μg of static-EVs or load-EVs per wells was added into primary mice chondrocytes with or without 10 ng/mL recombinant mouse IL-1β protein treated. qPCR assay (after 24 h), Alcian Blue staining (after 48 h), IF staining (after 48 h) and protein extraction (after 48 h) were performed subsequently. For WT-EVs or Dicer-cKO-EVs treatment, 5 μg of WT-EVs or Dicer-cKO-EVs was added into 1 × 105 primary mouse chondrocytes. qPCR assay (after 24 h), Alcian Blue staining and protein extraction (after 48 h) were performed subsequently.
For nc/mimic-EVs or nc/inhibitor-EVs treatment, static MLO-Y4 cells were transfected with miR-23b-3p mimics or nc while loaded MLO-Y4 cells were transfected with miR-23b-3p inhibitors or nc at 100 nM using Lipofectamine 2000 (11668019, Invitrogen). After 6–8 h, the culture medium was replaced and incubated for another 24 h for EVs extraction. Static nc/mimic-EVs or loaded nc/inhibitor-EVs were added into primary mice chondrocytes. qPCR assay (after 24 h), Alcian Blue staining and protein extraction (after 48 h) were performed subsequently.
For miRNA transport and transfection assay, Cd63-eGFP-MLO-Y4 stable cell lines were transfected with Cy3-miR-223 mimic using Lipofectamine 2000 and then co-cultured with primary mice chondrocytes using a trans-well plate for 12 h, with primary mice chondrocytes placed in the lower chamber and CD63-eGFP-MLO-Y4 cells in the upper chamber. After washing with PBS twice, the co-localization of Cy3 red fluorescence and eGFP green fluorescence as well as the expression of miR-223 on primary mice chondrocytes were examined.
To overexpress miR-23b-3p in chondrocytes in vitro, primary mice chondrocytes were transfected with either miR-23b-3p mimics at 100 nM or NC using Lipofectamine 2000. After 6–8 h, the culture medium was replaced and incubated for qPCR assay (after 24 h), Alcian Blue staining and protein extraction (after 48 h).
Flow cytometry analysis
After preparing a single-cell suspension from the extracted primary cells, resuspend the cell pellet in permeabilization buffer and centrifuge at 500 x g for 5 mins at room temperature (RT). Following a wash step, resuspend the pellet in 100 µL of flow cytometry staining buffer. Add the primary antibodies anti-Dmp1 (PA5-47621, Invitrogen, 1:50 dilution) and anti-Runx2 (ab192256, abcam, 1:50 dilution) to the staining buffer, and incubate the cells in the dark at RT for 20-60 mins. Following a wash step, then the cells were incubated with the corresponding secondary antibodies at RT for 30 mins. After centrifuging at 500 x g for 5 mins at 4 °C, discard the supernatant, resuspend the cells in 500 µL of staining buffer and load the sample onto the flow cytometer for analysis.
Mitophagy detection
TEM was performed to visualize mitochondrial morphology within human or mice cartilage samples. Mitophagy Detection Kit (MD01, DOJINDO, Japan) was used to observe the colocation of mitophagy and lysosomal mitophagy in primary human chondrocytes. JC-1 MitoMP Detection Kit (MT09, DOJINDO, Japan) was used to measure the mitochondrial membrane potential in primary human chondrocytes.
Western blotting and enzyme-linked immunosorbent assay (ELISA)
Total protein was extracted from the collected cells with pre-cooled RIPA lysis buffer (Thermo Fisher Scientific) supplemented with protease/phosphatase inhibitor cocktail and PMSF (KGP2100, KeyGEN). The protein concentrations were measured using BCA protein assay kits. For western blotting, after electrophoresis on 7% or 10% SDSPAGE gels (Bio-Rad, USA), the proteins were transferred to PVDF membranes (Merck Millipore) and incubated in blocking buffer (P0023B, Beyotime) for 1 h. The PVDF membranes were then incubated with primary antibodies overnight at 4 °C. After incubating with a horseradish peroxidase (HRP)-conjugated secondary antibody (1:10000 dilution) and visualized using a commercial enhanced chemiluminescence kit (ECL, P2300, New Cell & Molecular Biotech). Quantitative analysis was performed using ImageJ software and normalized to expressions of β-actin as loading control. The following commercial antibodies were used: anti-HSP70 (A12948, ABclonal, 1:1000 dilution), anti-TSG101 (ab125011, abcam, 1:1000 dilution), anti-CD63 (A19023, ABclonal, 1:1000 dilution), anti-CD81 (ab109201, abcam, 1:1000 dilution), anti-CD9 (1:1000 dilution), anti-Aggrecan (NB600-504, Novus, 1:1000 dilution), anti-SOX9 (82630, CST, 1:1000 dilution), anti-MMP13 (18165-1-AP, Proteintech, 1:1000 dilution), anti-ADAMTS5 (NBP2-15286, Novus, 1:1000 dilution), anti-COLII (A00517-1, BOSTER, 1:1000 dilution), anti-β-actin (20536-1-AP, Proteintech, 1:2000 dilution), anti-OTUD4 (25070-1-AP, Proteintech, 1:1000 dilution), anti-PARKIN (A0968, ABclonal, 1:1000 dilution), anti-PINK (A7131, ABclonal, 1:1000 dilution), anti-MFN2 (A19678, ABclonal, 1:1000 dilution), anti-DRP1 (A21968, ABclonal, 1:1000 dilution), anti-LC3B (A7198, ABclonal,1:1000 dilution).
ELISA was utilized to quantify concentrations of pro-inflammatory factors IL-1β (ml058059, mlbio), IL-6 (RK00004, ABclonal), and TNF-α (RK00030, ABclonal) in the cell culture supernatant, using commercial kits according to the manufacturer’s instructions.
RNA extraction, RT–PCR and miRNA qPCR array
Total RNA or tissue miRNAs were isolated using TRIzol reagent (Takara) following the standard protocol provided by the manufacturer. EVs miRNA was isolated using miRNeasy Mini kit (217004, Qiagen). For mRNA qPCR, the first-strand cDNA was synthesized from the total RNA with the HiScript III RT SuperMix (R323-01, Vazyme). Real-time PCR was carried out using the ABI ViiA 7 system (Thermo) with the ChamQ SYBR qPCR Master Mix (Q711, Vazyme). The mRNA primer sequences (Generay biotech, Shanghai) were shown in Supplementary Table S2. The RT-qPCR cycling conditions were as follows: initial denaturation at 50 °C for 2 minutes and 95 °C for 10 minutes, followed by 40 cycles of 95 °C for 15 seconds, 60 °C for 1 minute, and 95 °C for 15 seconds. For mature miRNA detection, the expression of mature miRNAs was detected by miRNA 1st Strand cDNA Synthesis Kit (by tailing A) (MR201-01, Vazyme). The mature miRNA primer sequences were obtained from RIBOBIO. For primary miRNA detection, the expression of primary miRNAs was detected using miRNA 1st Strand cDNA Synthesis Kit (by stem-loop) (MR101, Vazyme) and miRNA Universal SYBR qPCR Master Mix (MQ101, Vazyme). The primary miRNA primer sequences were shown in Supplementary Table S3. The relative expression of mRNA or miRNA was calculated by the 2−ΔΔCt method with β-actin (mRNA), U6 (intracellular miRNA control), cel-miR39 (exosomal miRNA control in cell culture medium). For miRNA profiling by qPCR array, the mmu-miRNome MicroRNA Profiling Kit (QM043, igenebio) were used for profiling the miRNAs in osteocytes from Dicer-cKO mice and WT mice according to the manufacturer’s recommendations. miRNA array analyze (z-score) was performed by using Heatmap Plot tools in Hiplot Pro (https://hiplot.com.cn/), a comprehensive web service for biomedical data analysis and visualization. A miRNA was plotted only if the raw Ct Value was ≤35 in all the samples.
Finite element analysis
The magnetic resonance data were obtained from a male patient with OA or a healthy male volunteer by a 3-T clinical MR scanner (uMR 770; United Imaging, Shanghai, China) using the 12-channel knee send-receive radio frequency coil. The 3D reconstruction was performed by MIMICS 19.0 (Materialise, Leuven, Belgium). The tibial plateaus above the growth plate were selected from the μCT images, thresholded, segmented into cartilage and bone, and converted into finite element models by setting each image voxel to an a 10-node quadratic tetrahedron (C3D10H). The bone material behaviour was linear, with an elastic modulus (E) as of 7300 MPa and a Poisson ratio (ν) of 0.329. Stresses and strains in the SCB were calculated using in the Abaqus 2017 (SIMULIA, Rhode Island, USA).
MicroCT analysis
The mouse knee joints free of soft tissue were fixed in 4% paraformaldehyde for 24 h immediately after death. The fixed samples were imaged by micro-CT (vivaCT80, SCANCO Medical AG, Switzerland). We defined the region of interest as covering the whole SB medial compartment and subsequently preformed 3D reconstruction and analysis. 3D structural parameters analyzed were bone mineral density (BMD) and bone volume to total volume (BV/TV) were analyzed with the built-in software.
Histology assays
Decalcified human samples and mouse knee joints were fixed in 4% paraformaldehyde for 24 h following by dehydration in gradient ethanol and xylene before embedding in paraffin. The 5-μm paraffin sections were made and mounted onto the slides. For histology assay, three of the sequential sections were proceeded to Safranin O/Fast Green staining (Sigma) according to standard protocols. Histologic grading of cartilage degeneration was performed using the Osteoarthritis Research Society International (OARSI) scoring system81.
Immunohistochemistry (IHC) and immunofluorescence analyses (IF)
The deparaffinized and rehydrated sections were immersed in pepsin buffer for antigen retrieval and incubated with 3% hydrogen peroxide for 15 min. The cryostat sections were placed at room temperature and washed 3 times with PBS for 30 min. All sections were blocked using Instant-Using Goat-serum at room temperature containing 0.15% Triton X-100 for 1 h followed by incubation with the primary antibodies at the indicated dilutions at 4 °C overnight, including anti-CD63 (1:200 dilution), anti-Aggrecan (1:200 dilution), anti-SOX9 (1:200 dilution), anti-MMP13 (1:200 dilution), anti-ADAMTS5 (1:200 dilution), anti-COLII (1:200 dilution), anti-OTUD4 (1:200 dilution) and anti-Endomucin (AF4666, R&D Systems, 1:200 dilution). For IHC staining, sections were incubated with HRP-conjugated secondary antibodies (AS014/ AS003, ABclonal, 1:200 dilution). For negative control, goat non-immune serum at the same concentration as the primary antibody was used to control for any non-specific binding from the secondary antibody. Isotype-matched control antibodies at the same concentration as the primary antibody was used to ensure that observed staining was antibody-specific. After stained with 3,3′-diaminobenzidine, IHC images were captured with an Olympus BX53 microscope. For IF staining, the sections were washed and incubated with fluorescent secondary antibodies (abcam, 1:400 dilution) for 1 h. Negative control where no primary antibody was applied, and only secondary antibodies were used to check for any secondary antibody cross-reactivity. The sections were then mounted by cover slides with Fluoroshield Mounting Medium with DAPI.
For cell IF staining, cells were fixed with 4% formaldehyde for 10 min, washed in PBS and treated with Triton X-100 for 10 min. After washed again with PBS, cells were blocked 1 h with blocking buffer followed by incubation with the primary antibodies: anti-Aggrecan (1:200 dilution), anti-SOX9 (1:200 dilution), anti-MMP13 (1:200 dilution), anti-ADAMTS5 (1:200 dilution), anti-COLII (1:200 dilution) overnight at 4 °C. Next, the cells were washed three times with PBS, incubated with fluorescent secondary antibodies (1:400 dilution) for 1 h. Nuclei were counterstained with DAPI. IF images were captured with Leica DMi8 THUNDER Imaging Systems or Olympus FV3000 laser confocal microscope.
Luciferase reporter assays
The 3′ UTR segments of the OTUD4 genes were amplified by polymerase chain reaction (PCR) and inserted into the vector. Co-transfections of OTUD4 3′ UTR plasmids with miR-23b-3p mimics or nc into the HEK293T cells (CL-0005, Procell Life Science & Technology) were accomplished by using Lipofectamine 2000 (Invitrogen). Luciferase activity was measured 48 h after transfection by the Dual-Luciferase Reporter Assay System (E1910, Promega). The absorbance values were read using the GloMax 96 Microplate Luminometer.
Laser-capture microdissection
The mice knee joints were decalcified in 10% EDTA, embedded in OCT and cutted into frozen sections (20 μm). Adjacent sections were mounted on polyethylene membrane– equipped slides (Leica, Germany). SCB osteocytes in adjacent sections were isolated with microdissection system (Leica CTR6500), which were harvested for subsequent miRNA extraction using reaction tubes with the lysis buffer added in advance.
RNA transcriptome sequencing (RNA-seq)
RNA-seq experiment and data analysis were conducted by Seqhealth Technology Co., LTD (Wuhan, China). The details of the sequencing depth were shown in Fig. S12. Briefly, total RNA was isolated from osteocyte-enriched human tissues (LS/HS tissues) or primary human chondrocytes with miR-23b-3p mimic or NC transfected. 2 μg total RNAs were used for stranded RNA sequencing library preparation using KCTM Stranded mRNA Library Prep Kit for Illumina® following the manufacturer’s instruction. PCR products corresponding to 200-500 bps were enriched, quantified and finally sequenced on DNBSEQ-T7 sequencer with PE150 model. A log2fold change of <0 and an adjust p value of <0.05 were used to judge the statistical significance of gene expression differences. Different gene expressions (DGEs) between groups were identified using the edgeR package (version 3.40.2). Gene ontology (GO) analysis and Kyoto encyclopedia of genes and genomes (KEGG) enrichment analysis for differentially expressed genes were implemented by KOBAS software (version: 2.1.1) with a p value of <0.05 to judge statistically significant enrichment.
Small RNA deep sequencing and bioinformatics analysis
For LS-EVs or HS-EVs miRNA sequencing, each mixed samples were consisted of six OA patient samples (consisted of 3 males and 3 females). miRNA components in either control LS-EVs or HS-EVs were profiled by single-end sequencing (1 × 50 bp) on an Illumina platform (NEB, USA) at Novogene (Beijing, China). The raw data were processed through custom perl and python scripts. Known miRNAs were identified by comparison to the miRBase 20.0 database (http://www.mirbase.org/). Differential expression analysis of two conditions/ groups was performed using the DESeq2 R package (v1.24.0). The P values was adjusted using the Benjamini and Hochberg method. Corrected P value of 0.05 was set as the threshold for significantly differential expression by default.
For WT-EVs or Dicer cKO-EVs miRNA sequencing, each mixed samples were consisted of four Dicer cKO mice samples. Sequencing libraries were generated using NEBNext@ Multiplex Small RNA Library Prep Set for Illumina at LC Sciences (Houston, Texas, USA). The libraries were quality-assessed on an Agilent Bioanalyzer 2100 system using DNA High Sensitivity Chips and sequenced on an Illumina HiSeq 2500/2000 platform. Known miRNAs were identified by comparison to the miRBase 20.0 database. Data analysis was performed by using ACGT101-miR (LC Sciences).
Synthesis and characterization of bone-targeted delivery system
A bone-targeted delivery approach as described by our team in previously published articles20,82. Briefly, the lipid nanoparticle composed of DSPE-PEG2000 was conjugated with repetitive sequences of aspartate, serine, serine ((DSS)6), effectively encapsulating GW4869 or antagomir-23b-3p within lipid nanoparticle. Liposomes contain nothing were referred to as blank liposomes. LipoGW4869, lipoDiR or lipoantagomir-23b-3p did not conjugated with DSS6. LipoDiR@DSS6 contained DiR at a concentration of 1 mg/mL. To characterize these liposomes, the particle size distribution and morphology of liposomes were measured by brookhaven 90 Plus Laser Particle Sizer and TEM, respectively. To identify the targeting effect of DSS6, 200 μL lipoDiR@DSS6 and lipoDiR was injected via mice tail vein for 48 h, respectively. Mice were sacrificed and the major organs (heart, liver, spleen, lung, kidney, femur) were collected to determine fluorescence signal of DiR by in vivo Imaging Instrument.
Statistics and reproducibility
Statistical analyses were performed using SPSS 23 software (IBM, NY, USA) or GraphPad Prism 8.0 software (GraphPad, La Jolla, CA, USA). All data are shown as mean ± SD and a P value < 0.05 was considered as significant. For differences between two groups, two-tailed unpaired or paired Student’s t-tests were used. For comparison of more than two groups, one-way analysis of variance (ANOVA) with Tukey’s multiple-comparison test were used. Data quantified based on the OARSI scores, whose data points are not continuous and do not follow a normal distribution, were analyzed using a non-parametric test based on two-tailed Mann–Whitney U test. Data collection and statistical analyses were blinded by replacing group names with blind codes, and no data were excluded from the analyses. All results from the representative experiments described in this research were conducted at least 3 times in independent experiments. We determined the sample size of mice used in our experiment based on the Resource Equation Approach method83,84 and the design of our previous experience85.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The authors declare that the data supporting the findings of this study are available within the paper and its Supplementary Information files, and that all additional data are publicly available. The high throughput data reported in this paper have been deposited in the GEO database under accession codes: GSE284357, GSE284358, GSE284359, GSE284361. Source data are provided with this paper.
References
Bijlsma, J. W. J., Berenbaum, F. & Lafeber, F. P. J. G. Osteoarthritis: an update with relevance for clinical practice. Lancet 377, 2115–2126 (2011).
Glyn-Jones, S. et al. Osteoarthritis. Lancet 386, 376–387 (2015).
Martel-Pelletier, J. et al. Osteoarthritis. Nat. Rev. Dis. Prim. 2, 16072 (2016).
Goldring, S. R. Alterations in periarticular bone and cross talk between subchondral bone and articular cartilage in osteoarthritis. Ther. Adv. Musculoskelet. Dis. 4, 249–258 (2012).
Mobasheri, A. et al. The role of metabolism in the pathogenesis of osteoarthritis. Nat. Rev. Rheumatol. 13, 302–311 (2017).
Karsdal, M. A. et al. The coupling of bone and cartilage turnover in osteoarthritis: opportunities for bone antiresorptives and anabolics as potential treatments? Ann. Rheum. Dis. 73, 336–348 (2014).
Goldring, S. R. & Goldring, M. B. Changes in the osteochondral unit during osteoarthritis: structure, function and cartilage-bone crosstalk. Nat. Rev. Rheumatol. 12, 632–644 (2016).
Burr, D. B. & Gallant, M. A. Bone remodelling in osteoarthritis. Nat. Rev. Rheumatol. 8, 665–673 (2012).
Hu, W., Chen, Y., Dou, C. & Dong, S. Microenvironment in subchondral bone: predominant regulator for the treatment of osteoarthritis. Ann. Rheum. Dis. 80, 413–422 (2021).
Jung, Y.-K. et al. Calcium-phosphate complex increased during subchondral bone remodeling affects earlystage osteoarthritis. Sci. Rep. 8, 487 (2018).
Liu, J. et al. Exosomal transfer of osteoclast-derived miRNAs to chondrocytes contributes to osteoarthritis progression. Nat. Aging 1, 368–384 (2021).
Jaiprakash, A. et al. Phenotypic characterization of osteoarthritic osteocytes from the sclerotic zones: a possible pathological role in subchondral bone sclerosis. Int. J. Biol. Sci. 8, 406–417 (2012).
Zhou, J. et al. Identification of mechanics-responsive osteocyte signature in osteoarthritis subchondral bone. Bone Jt. Res. 11, 362–370 (2022).
Mazur, C. M. et al. Osteocyte dysfunction promotes osteoarthritis through MMP13-dependent suppression of subchondral bone homeostasis. Bone Res. 7, 34 (2019).
Kalluri, R. & LeBleu, V. S. The biology, function, and biomedical applications of exosomes. Science 367, https://doi.org/10.1126/science.aau6977 (2020).
Isaac, R., Reis, F. C. G., Ying, W. & Olefsky, J. M. Exosomes as mediators of intercellular crosstalk in metabolism. Cell Metab. 33, 1744–1762 (2021).
Qin, Y. et al. Myostatin inhibits osteoblastic differentiation by suppressing osteocyte-derived exosomal microRNA-218: a novel mechanism in muscle-bone communication. J. Biol. Chem. 292, 11021–11033 (2017).
Wang, Z.-X. et al. Aged bone matrix-derived extracellular vesicles as a messenger for calcification paradox. Nat. Commun. 13, 1453 (2022).
Ostrowski, M. et al. Rab27a and Rab27b control different steps of the exosome secretion pathway. Nat. Cell Biol. 12, https://doi.org/10.1038/ncb2000 (2010).
He, Y. et al. Elimination of senescent osteocytes by bone-targeting delivery of β-galactose-modified maytansinoid prevents age-related bone loss. Adv. Health. Mater. 13, e2302972 (2024).
Johnnidis, J. B. et al. Regulation of progenitor cell proliferation and granulocyte function by microRNA-223. Nature 451, 1125–1129 (2008).
Kim, Y.-K., Kim, B. & Kim, V. N. Re-evaluation of the roles of DROSHA, Export in 5, and DICER in microRNA biogenesis. Proc. Natl Acad. Sci. USA 113, E1881–E1889 (2016).
Blanco, F. J., Valdes, A. M. & Rego-Pérez, I. Mitochondrial DNA variation and the pathogenesis of osteoarthritis phenotypes. Nat. Rev. Rheumatol. 14, 327–340 (2018).
Sun, K., Jing, X., Guo, J., Yao, X. & Guo, F. Mitophagy in degenerative joint diseases. Autophagy 17, 2082–2092 (2021).
Lories, R. J. & Luyten, F. P. The bone-cartilage unit in osteoarthritis. Nat. Rev. Rheumatol. 7, 43–49 (2011).
Ziemian, S. N. et al. Low bone mass resulting from impaired estrogen signaling in bone increases severity of load-induced osteoarthritis in female mice. Bone 152, 116071 (2021).
Kozaki, T. et al. Medial meniscus extrusion and varus tilt of joint line convergence angle increase stress in the medial compartment of the knee joint in the knee extension position-finite element analysis. J. Exp. Orthop. 9, 49 (2022).
Steer, J. W., Worsley, P. R., Browne, M. & Dickinson, A. Key considerations for finite element modelling of the residuum-prosthetic socket interface. Prosthet. Orthot. Int. 45, 138–146 (2021).
Li, L. et al. 3D printing individualized heel cup for improving the self-reported pain of plantar fasciitis. J. Transl. Med. 16, 167 (2018).
Li, L. et al. Biomechanical analysis of the effect of medial meniscus degenerative and traumatic lesions on the knee joint. Am. J. Transl. Res. 11, 542–556 (2019).
Dallas, S. L., Prideaux, M. & Bonewald, L. F. The osteocyte: an endocrine cell… and more. Endocr. Rev. 34, 658–690 (2013).
Gluhak-Heinrich, J. et al. Mechanical loading stimulates dentin matrix protein 1 (DMP1) expression in osteocytes in vivo. J. Bone Min. Res. 18, 807–817 (2003).
Zhang, L. & Wen, C. Osteocyte dysfunction in joint homeostasis and osteoarthritis. Int. J. Mol. Sci. 22, https://doi.org/10.3390/ijms22126522 (2021).
Ilas, D. C. et al. The simultaneous analysis of mesenchymal stem cells and early osteocytes accumulation in osteoarthritic femoral head sclerotic bone. Rheumatology 58, 1777–1783 (2019).
Pathak, J. L. et al. Systemic inflammation affects human osteocyte-specific protein and cytokine expression. Calcif. Tissue Int. 98, 596–608 (2016).
Bratengeier, C. et al. Mechanical loading intensities affect the release of extracellular vesicles from mouse bone marrow-derived hematopoietic progenitor cells and change their osteoclast-modulating effect. FASEB J. 38, e23323 (2024).
Patel, D. B., Luthers, C. R., Lerman, M. J., Fisher, J. P. & Jay, S. M. Enhanced extracellular vesicle production and ethanol-mediated vascularization bioactivity via a 3D-printed scaffold-perfusion bioreactor system. Acta Biomater. 95, 236–244 (2019).
Guo, S. et al. Stimulating extracellular vesicles production from engineered tissues by mechanical forces. Nano Lett. 21, 2497–2504 (2021).
Mathieu, M., Martin-Jaular, L., Lavieu, G. & Théry, C. Specificities of secretion and uptake of exosomes and other extracellular vesicles for cell-to-cell communication. Nat. Cell Biol. 21, https://doi.org/10.1038/s41556-018-0250-9 (2019).
Lim, C. Z. J., Zhang, L., Zhang, Y., Sundah, N. R. & Shao, H. New sensors for extracellular vesicles: insights on constituent and associated biomarkers. ACS Sens. 5, https://doi.org/10.1021/acssensors.9b02165 (2020).
Jiang, Y.-L. et al. The protective effects of osteocyte-derived extracellular vesicles against Alzheimer’s disease diminished with aging. Adv. Sci. 9, e2105316 (2022).
Wang, Y.-Y., Xia, K., Wang, Z.-X., Xie, H. & Xu, R. Osteocyte exosomes accelerate benign prostatic hyperplasia development. Mol. Cell Endocrinol. 531, 111301 (2021).
Findlay, D. M. & Kuliwaba, J. S. Bone-cartilage crosstalk: a conversation for understanding osteoarthritis. Bone Res. 4, 16028 (2016).
Trajkovic, K. et al. Ceramide triggers budding of exosome vesicles into multivesicular endosomes. Science 319, 1244–1247 (2008).
Menck, K. et al. Neutral sphingomyelinases control extracellular vesicles budding from the plasma membrane. J. Extracell. Vesicles 6, 1378056 (2017).
Sun, W. et al. Osteoclast-derived microRNA-containing exosomes selectively inhibit osteoblast activity. Cell Discov. 2, 16015 (2016).
Jiang, N. et al. Exosomes mediate epithelium-mesenchyme crosstalk in organ development. ACS Nano 11, 7736–7746 (2017).
Kowal, J., Tkach, M. & Théry, C. Biogenesis and secretion of exosomes. Curr. Opin. Cell Biol. 29, 116–125 (2014).
Raposo, G. & Stoorvogel, W. Extracellular vesicles: exosomes, microvesicles, and friends. J. Cell Biol. 200, 373–383 (2013).
Ratajczak, J., Wysoczynski, M., Hayek, F., Janowska-Wieczorek, A. & Ratajczak, M. Z. Membrane-derived microvesicles: important and underappreciated mediators of cell-to-cell communication. Leukemia 20, 1487–1495 (2006).
Mori, M. A., Ludwig, R. G., Garcia-Martin, R., Brandão, B. B. & Kahn, C. R. Extracellular miRNAs: from biomarkers to mediators of physiology and disease. Cell Metab. 30, 656–673 (2019).
Garcia-Martin, R. et al. MicroRNA sequence codes for small extracellular vesicle release and cellular retention. Nature 601, 446–451 (2022).
Suzuki, H. I. et al. Regulation of TGF-β-mediated endothelial-mesenchymal transition by microRNA-27. J. Biochem. 161, 417–420 (2017).
Xia, B. et al. Mechanical stimulation of Schwann cells promote peripheral nerve regeneration via extracellular vesicle-mediated transfer of microRNA 23b-3p. Theranostics 10, 8974–8995 (2020).
Guo, Y. et al. Downregulation of HS6ST2 by miR-23b-3p enhances matrix degradation through p38 MAPK pathway in osteoarthritis. Cell Death Dis. 9, 699 (2018).
Yang, Q. et al. Downregulation of microRNA-23b-3p alleviates IL-1β-induced injury in chondrogenic CHON-001 cells. Drug Des. Dev. Ther. 13, 2503–2512 (2019).
Iliopoulos, D., Malizos, K. N., Oikonomou, P. & Tsezou, A. Integrative microRNA and proteomic approaches identify novel osteoarthritis genes and their collaborative metabolic and inflammatory networks. PLoS ONE 3, e3740 (2008).
Fujiwara, Y. et al. miR-23a/b clusters are not essential for the pathogenesis of osteoarthritis in mouse aging and post-traumatic models. Front. Cell Dev. Biol. 10, 1043259 (2022).
Guo, H., Ingolia, N. T., Weissman, J. S. & Bartel, D. P. Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature 466, 835–840 (2010).
Loeser, R. F., Collins, J. A. & Diekman, B. O. Ageing and the pathogenesis of osteoarthritis. Nat. Rev. Rheumatol. 12, 412–420 (2016).
Ansari, M. Y., Khan, N. M., Ahmad, I. & Haqqi, T. M. Parkin clearance of dysfunctional mitochondria regulates ROS levels and increases survival of human chondrocytes. Osteoarthr. Cartil. 26, 1087–1097 (2018).
D’Amico, D. et al. Urolithin A improves mitochondrial health, reduces cartilage degeneration, and alleviates pain in osteoarthritis. Aging Cell 21, e13662 (2022).
Perl, A., Gergely, P., Nagy, G., Koncz, A. & Banki, K. Mitochondrial hyperpolarization: a checkpoint of T-cell life, death and autoimmunity. Trends Immunol. 25, 360–367 (2004).
Menges, S. et al. Alpha-synuclein prevents the formation of spherical mitochondria and apoptosis under oxidative stress. Sci. Rep. 7, 42942 (2017).
Yang, N., Zhang, R., Zhang, H., Yu, Y. & Xu, Z. ZIP7 contributes to the pathogenesis of diabetic cardiomyopathy by suppressing mitophagy in mouse hearts. Cardiovasc. Diabetol. 23, 399 (2024).
Wang, L., Qi, H., Tang, Y. & Shen, H.-M. Post-translational modifications of key machinery in the control of mitophagy. Trends Biochem. Sci. 45, 58–75 (2020).
Zhu, Q., Fu, Y., Li, L., Liu, C. H. & Zhang, L. The functions and regulation of Otubains in protein homeostasis and diseases. Ageing Res. Rev. 67, 101303 (2021).
Liu, W. et al. OTUD1 stabilizes PTEN to inhibit the PI3K/AKT and TNF-alpha/NF-kappaB signaling pathways and sensitize ccRCC to TKIs. Int. J. Biol. Sci. 18, 1401–1414 (2022).
Zhou, N. et al. Deubiquitinase OTUD3 regulates metabolism homeostasis in response to nutritional stresses. Cell Metab. 34, https://doi.org/10.1016/j.cmet.2022.05.005 (2022).
Dinallo, V. et al. The deubiquitinating enzyme OTUD5 sustains inflammatory cytokine response in inflammatory Bowel disease. J. Crohns Colitis 16, 122–132 (2022).
Chu, Y. et al. LUBAC and OTULIN regulate autophagy initiation and maturation by mediating the linear ubiquitination and the stabilization of ATG13. Autophagy 17, 1684–1699 (2021).
Dai, J. et al. Hepatocyte deubiquitinating enzyme OTUD5 deficiency is a key aggravator for metabolic dysfunction-associated steatohepatitis by disturbing mitochondrial homeostasis. Cell Mol. Gastroenterol. Hepatol. 17, 399–421 (2024).
Yin, W. & Rogge, M. Targeting RNA: a transformative therapeutic strategy. Clin. Transl. Sci. 12, https://doi.org/10.1111/cts.12624 (2019).
Hanna, J., Hossain, G. S. & Kocerha, J. The potential for microRNA therapeutics and clinical research. Front. Genet. 10, 478 (2019).
Glasson, S. S., Blanchet, T. J. & Morris, E. A. The surgical destabilization of the medial meniscus (DMM) model of osteoarthritis in the 129/SvEv mouse. Osteoarthr. Cartil. 15, 1061–1069 (2007).
Huang, H., Skelly, J. D., Ayers, D. C. & Song, J. Age-dependent changes in the articular cartilage and subchondral bone of C57BL/6 mice after surgical destabilization of medial meniscus. Sci. Rep. 7, 42294 (2017).
Ma, H. L. et al. Osteoarthritis severity is sex dependent in a surgical mouse model. Osteoarthr. Cartil. 15, 695–700 (2007).
Marahleh, A. et al. Obtaining primary osteocytes through murine calvarial fractionation of GFP-expressing osteocytes. J. Vis. Exp. https://doi.org/10.3791/61513 (2020).
Thirion, S. & Berenbaum, F. Culture and phenotyping of chondrocytes in primary culture. Methods Mol. Med. 100, 1–14 (2004).
Gosset, M., Berenbaum, F., Thirion, S. & Jacques, C. Primary culture and phenotyping of murine chondrocytes. Nat. Protoc. 3, 1253–1260 (2008).
Pritzker, K. P. H. et al. Osteoarthritis cartilage histopathology: grading and staging. Osteoarthr. Cartil. 14, 13–29 (2006).
Zhang, G. et al. A delivery system targeting bone formation surfaces to facilitate RNAi-based anabolic therapy. Nat. Med. 18, 307–314 (2012).
Charan, J. & Kantharia, N. D. How to calculate sample size in animal studies? J. Pharm. Pharmacother. 4, 303–306 (2013).
Arifin, W. N. & Zahiruddin, W. M. Sample size calculation in animal studies using resource equation approach. Malays. J. Med. Sci. 24, 101–105 (2017).
Chen, X. et al. METTL3-mediated m6A modification of ATG7 regulates autophagy-GATA4 axis to promote cellular senescence and osteoarthritis progression. Ann. Rheum. Dis. 81, 87–99 (2022).
Acknowledgements
This work was supported by the National Key Research and Development Project (2021YFA1201404, 2023YFC3605800), the National Science Foundation of China (82272530, 82372459, 81972124), the Major Project of NSFC (81991514), Jiangsu Province Medical Innovation Center of Orthopedic Surgery (CXZX202214), Jiangsu Provincial Key Medical Center Foundation, Jiangsu Provincial Medical Outstanding Talent Foundation, Jiangsu Provincial Medical Youth Talent Foundation and Jiangsu Provincial Key Medical Talent Foundation, the Fundamental Research Funds for the Central Universities (14380493, 14380494), Natural science foundation of Jiangsu province (Grant No. BK20220780, BE2022818), Fundamental Research Funds for the Central Universities (Grant No. 14380532, 2024300419).
Author information
Authors and Affiliations
Contributions
Baosheng Guo and Qing Jiang conceived and designed the work; Na Liu obtained the data; Na Liu, Yuze Ma, Wang Gong, Tianshu Shi, Xiaoyan Shao, Xiang Chen, Wenshu Wu and Yong Shi performed in vivo studies; Lan Li performed the FEA test; Pu Wang, Wentian Gao, Rui Du, Jianghui Qin and Dongyang Chen collected clinical samples; Na Liu, Pan Zhang, Jiaquan Lin, Chengzhi Wang, Yi He, Depeng Fang and Bin Liu performed in vitro studies; Lin Yang and Xueying An performed the synthesis and characterization of bone-targeted delivery system; Na Liu wrote the paper; Ying Chen and Depeng Fang performed IF/IHC staining and flow cytometric analysis during the revision; Pingqiang Cai provided additional financial support during the revision; Baosheng Guo and Qing Jiang revised the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Masaru Ishii, Junichi Kikuta and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, N., Ma, Y., Gong, W. et al. Osteocyte-derived extracellular vesicles mediate the bone-to-cartilage crosstalk and promote osteoarthritis progression. Nat Commun 16, 4746 (2025). https://doi.org/10.1038/s41467-025-59861-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-59861-5
This article is cited by
-
Intravital Microscopy of Osteocytes: A New Era in Bone Cell Biology
Current Osteoporosis Reports (2025)