Abstract
Due to low availability of CO2 in aquatic environment, microalgae have evolved a CO2 concentrating mechanism (CCM). It has long been thought that operation of CCM would suppress photorespiration by increasing the CO2 concentration at the Rubisco active site, but experimental evidence is scarce. To better explore the function of photorespiration in algae, we first characterized a Chlamydomonas reinhardtii mutant defected in low-CO2 inducible 20 (LCI20) and show that LCI20 is a chloroplast-envelope glutamate/malate transporter playing a role in photorespiration. By monitoring growth and glycolate excretion in mutants deficient in either CCM or photorespiration, we conclude that: (i.) CCM induction does not depend on photorespiration, (ii.) glycolate excretion together with glycolate dehydrogenase down-regulation prevents the toxic accumulation of non-metabolized photorespiratory metabolites, and (iii.) photorespiration is active at low CO2 when the CCM is operational. This work provides a foundation for a better understanding of the carbon cycle in the ocean where significant glycolate concentrations have been found.
Similar content being viewed by others
Introduction
Microalgae account for nearly half of the photosynthetic CO2 fixation on Earth1,2. They contribute to climate change mitigation by assimilating CO2 and are promising candidates for the production of biobased compounds3,4,5. Due to the low availability of CO2 in water, most algal cells have developed biophysical CO2 concentrating mechanisms (CCMs), thus increasing the CO2 concentration at the catalytic site of Rubisco and favoring the carboxylase reaction at the expense of the oxygenase. Microalgal biophysical CCMs generally involve carbonic anhydrases located in different cellular compartments, bicarbonate and/or CO2 channels or transporters, and in most cases the pyrenoid (a specific cellular compartment where Rubisco is packed in a liquid phase separated organelle and where CO2 is concentrated)6,7,8,9,10. As a consequence of the CCM operation, the production of 2-phosphoglycolate (2-PG) resulting from the oxygenase activity of Rubisco is diminished11,12. Because of its inhibitory effect on the Calvin cycle, 2-PG is metabolized and recycled through photorespiration. Photorespiration consists of multiple metabolic reactions distributed across different subcellular compartments thus requiring inter-organelle communication13. During photorespiration, the 2-PG is converted in the chloroplast into glycolate by the 2-PG phosphatase PGP1. In vascular plants lacking a biophysical CCM, glycolate is converted into glycerate through a series of reactions in peroxisomes and mitochondria. Although photorespiration occurs in both plants and microalgae, its subcellular organization seems to differ. For example, in vascular plants lacking a biophysical CCM, photorespiration requires tight cooperation of chloroplast, peroxisome and mitochondria13; whereas in Chlamydomonas, photorespiration depends mainly on cooperation between chloroplast and mitochondria14,15.
In the model green microalga Chlamydomonas reinhardtii (hereafter Chlamydomonas), part of the glycolate is transferred to mitochondria and further metabolized into glyoxylate, glycine and serine, the other part is excreted out of the cell16,17,18. During acclimation of Chlamydomonas to low CO2, CCM and photorespiratory genes, which share the same master regulator CIA5, are simultaneously induced19,20,21. Such a feature is intriguing since the activity of CCM supposedly inhibits photorespiration. It was proposed that photorespiration is transiently operational during the acclimation from high to low CO2 conditions until the CCM is fully induced7,12, photorespiratory metabolites such as 2-PG possibly acting as signaling molecules triggering CCM induction12,14,22,23,24.
Two types of CCM regimes have been described in Chlamydomonas depending on external CO2 levels, the low-CO2-based CCM (CO2 air level), in which cells preferentially transport CO2, and the very-low-CO2-based CCM, in which specific transporters are induced, such as the ATP-binding cassette transporter High Light Activated 3 (HLA3) involved in active bicarbonate uptake25,26. During CCM induction, mitochondria migrate from the chloroplast cup to the periphery of the cell at the vicinity of the plasma membrane27. The contribution of mitochondria to the energy supply of the algal CCM has been recently evidenced, and energy trafficking between chloroplast and mitochondria was suggested to supply ATP to CCM bicarbonate transporters28,29. Actually, energy exchange between subcellular organelles was evidenced decades ago in microalgae30,31,32,33 and has been re-examined more recently34,35,36,37. The exchange of reducing power between organelles may operate via “malate shuttles” composed of malate dehydrogenases (MDHs) and membrane transporters of dicarboxylic acids38,39,40,41.
Despite the existence of several genes encoding putative chloroplast malate transporters41,42,43, none of them has been characterized in Chlamydomonas. Based on the increased expression of Low-CO2 Inducible 20 (LCI20) gene, which encodes a putative malate/2-oxoglutarate transporter, during acclimation to limiting CO219,20,21, it was proposed that LCI20 may be involved as a malate shuttle in the energy trafficking between chloroplast and mitochondria to feed CCM bicarbonate transporters44. By another way, photorespiration may also contribute to the energy supply to external CCM transporters44,45. Indeed, during photorespiration NADH is produced during the mitochondrial condensation of two glycine molecules into one serine, the NADH being converted into ATP by the mitochondrial respiratory chain. Until now, the nature of metabolic pathways and the identity of proteins involved in the energy network transferring photosynthetic energy from the chloroplast towards mitochondria to feed the external CCM transporters remain largely unexplored.
In this study, we aimed at better characterizing the relationship between CCM and photorespiration in green algae. For this purpose, we a Chlamydomonas mutant deficient in LCI20 and showed that LCI20 is a malate/glutamate transporter involved in photorespiration by supplying amino groups for the mitochondrial conversion of glyoxylate into glycine. By comparing growth properties and glycolate excretion in mutants affected in photorespiration or CCM under various photorespiration regimes, we conclude that photorespiratory metabolites do not contribute to CCM induction, that glycolate excretion avoids toxicity of non-metabolized photorespiratory intermediates and that downstream glycolate metabolism, although occurring when the CCM is functioning under very low CO2, is not required for CCM operation.
Results
LCI20 encodes a putative chloroplast malate transporter whose expression is modulated by light and CO2 levels
Three putative chloroplast 2-oxoglutarate/malate transporters (annotated as OMT1, OMT2 and LCI20) orthologues of previously characterized Arabidopsis DiT1 and DiT2 malate transporters were identified in the Chlamydomonas genome42,43. Previous transcriptomic analysis performed during a day-night cycle or during limiting CO2 acclimation showed that the expression of LCI20 (but not that of OMT1 and OMT2) is strongly induced at the beginning of the light period46 or after the transition from H-CO2 (2%) to L-CO2 (0.04%) or VL-CO2 (0.01%)19,20,21 (Supplementary Fig. 1a, b). A phylogenetic analysis revealed that LCI20, which belongs to the GreenCut247, is widespread in Chlorophyta, closely related to DiT2 transporters, and evolutionarily divergent from OMT family transporters (Supplementary Figs. 1c and 2). Furthermore, LCI20 has recently been localized in the chloroplast but the enrichment in the envelope was not clearly shown48,49. Here, we reexamined the strain expressing LCI20-mVenus under the control of PSAD promoter and we can see clearly that LCI20 is localized to the chloroplast envelope (Fig. 1a and Supplementary Fig. 3).
a Subcellular localization of LCI20 protein fused with the Venus fluorescent reporter. False colours were used to represent Venus (green) and chlorophyll (magenta) fluorescence signals. Scale bar, 5 µm. The localization of LCI20-Venus in the chloroplast envelope was confirmed by three independent experiments. b Genomic structure of LCI20 gene and insertion site of the paromomycin resistance cassette in the CLiP lci20 mutant. Gray boxes at both extremities represent the 5’ and 3’UTR, respectively. Exons are colored blue, and the position of cassette insertion is indicated by red arrows. c Immunoblot analysis using anti-LCI20 antibodies. Uncropped immunoblots are provided as a Source Data file. Due to the limited amount of LCI20 antibodies, this immunoblot was repeated twice with a similar result. Photoautotrophic growth of lci20, CC4533 (i.e. WT in this study all throughout) and complemented lines (C1, C2) on agar plates kept under various CO2 regimes. Cells were grown photo-autotrophically in liquid culture under air CO2 level (d), or air supplemented with 2% CO2 (e) prior to spot testing. Images were taken 3 days (H-CO2 and L-CO2) or 5 days (VL-CO2) after growing under 80 μmol photons m−2 s−1.
The growth of lci20 is impaired during a transition from high to very low CO2
A mutant harboring an insertion in the sixth intron of the LCI20 locus was obtained from the Chlamydomonas library project (CLiP) collection50 and we named the mutant here as lci20 (Fig. 1b). We confirmed the insertion of the paromomycin resistance cassette and the absence of the LCI20 transcript in lci20 (Supplementary Fig. 4a, b). We then performed genetic complementation of lci20 by expressing the full-length genomic LCI20 coding sequence driven by the PSAD promoter (Supplementary Fig. 4a, b). A LCI20 antibody was produced in this study and used to show the absence of LCI20 in the mutant and its presence in WT and the complemented lines (Fig. 1c).
The growth of lci20 was then investigated under photoautotrophic conditions in both solid and liquid cultures under various CO2 levels (Fig. 1d, e; Supplementary Fig. 4c, d). On solid media, lci20 grew normally under H-CO2, L-CO2 or VL-CO2 when cells were previously acclimated to these conditions (Fig. 1d). However, the growth of lci20 was severely impaired when transitioning from H-CO2 to VL-CO2 (Fig. 1e), with smaller decreases in growth observed after a transition from H-CO2 to L-CO2 or from L-CO2 to VL-CO2 (Fig. 1e). In liquid media, while lci20 grew normally in H-CO2 or during transition from H-CO2 to L-CO2, its growth was reduced when expressed on the basis of cell volume during transition from H-CO2 to VL-CO2 (Supplementary Fig. 4c, d). Based on these results, we conclude that LCI20 is required for photoautotrophic growth when cells are subjected to a sudden and severe CO2 limitation.
lci20 is affected in photorespiratory glycolate metabolism
To determine whether the mutant phenotype is due to the role of LCI20 in the induction of the CCM or in photorespiration, we performed growth assays on agar plates by transferring H-CO2 grown cells into VL-CO2 under photorespiratory (21% O2) or non-photorespiratory (2% O2) conditions (Fig. 2a). As a control, we used the CCM1 transcription factor knockout mutant cia5, which is defective in the transcriptional induction of CCM and of the photorespiratory pathway11,45,51,52. cia5 grew normally under H-CO2 but growth was completely abolished when cells were transferred to VL-CO2, irrespective of O2 levels (Fig. 2a), thus indicating that a defect in the CCM, but not in photorespiration, prevented growth of the cia5 mutant under VL-CO2. Note that similar growth in both solid and liquid cultures was observed when wild-type cells were exposed to VL-CO2 either under photorespiratory conditions (21% O2) or non-photorespiratory conditions (2% O2) (Supplementary Fig. 5), indicating that photorespiration is not required for CCM induction. In contrast, lci20 grew poorly after transition under photorespiratory conditions (VL-CO2; 21% O2), but the growth was unaffected under non-photorespiratory conditions (VL-CO2; 2% O2) (Fig. 2a), indicating that lci20 is defective in photorespiration rather than in CCM. To further distinguish phenotypes between photorespiration and CCM deficiency, we have analyzed the growth of two CCM mutants bsti and cah3 deficient in bestrophins (BST1-3)53 and lumenal carbonic anhydrase CAH354, respectively (Supplementary Fig. 6). Note that BST1-3 and CAH3were reported as low CO2 inducible genes in the same transcriptomic study that has also reported LCI20 expression19,21. In contrast to lci20 but similar to cia5 (Fig. 2a), both bsti and cah3 were unable to grow under VLC regardless of the photorespiratory status (Supplementary Fig. 6a), indicating a defect in CCM but not in photorespiration.
a Growth performance of cia5, lci20 mutants and their respective controls when transferring photo-autotrophically grown cells in liquid cultures under H-CO2 to agar plates exposed to VL-CO2 either under 21% or 2% O2 under 80 μmol photons m−2 s−1. Images were taken after 3 days (H-CO2) or 5 days (VL-CO2) of growth. b Quantification of the glycolate concentration in the liquid medium after 20 h growth of cultures either under H-CO2 or upon transition for H-CO2 to VL-CO2 either in the presence of 21% or 2% O2. Bars show the average and dots show data from independent biological replicates (n = 3 ± SD). Asterisks represent statistically significant difference compared to the wild-type strains (*p ≤ 0.05 and ****p ≤ 0.0001) using one-way ANOVA.
Glycolate is excreted into the culture medium by Chlamydomonas cells when either the CCM or the glycolate metabolism is defective16,18,45,55,56. Glycolate excretion was measured after 20 h acclimation of H-CO2 grown cells to VL-CO2 either at 21% or 2% O2. At 21% O2, both cia5 and lci20 mutants showed a significantly higher excretion compared to their respective controls (Fig. 2b). Similarly, bsti and cah3 showed higher and continuous glycolate excretion during acclimation to VL-CO2 at 21% O2 (Supplementary Fig. 6b). Note that we did not detect glycolate excretion under H-CO2 or VL-CO2 at 2% O2, conditions when photorespiration is strongly reduced (Fig. 2b). We conclude that lci20 is defective in the photorespiratory glycolate metabolism, which is triggered when Chlamydomonas cells are transferred from H-CO2 to VL-CO2 at 21% O2.
Changes in abundance of photorespiration and CCM proteins during cells’ response to different CO2 and O2 levels
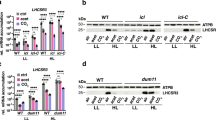
To identify possible causes for the growth defect and high glycolate excretion observed in lci20, we probed the abundance of different photorespiratory enzymes including the glycolate dehydrogenase (GYD1), hydroxypyruvate reductase (HPR1) and glycine cleavage system P (GCSP) proteins together with CCM-related proteins HLA3, low CO2-inducible 1 (LCI1) and low CO2-inducible C (LCIC) following a transfer from H-CO2 to VL-CO2 either at 21% or at 2% O2 (Fig. 3a and Supplementary Fig. 7). We observed a lower abundance of GYD1 in lci20 compared to control strains when the transfer was performed under photorespiratory conditions (21% O2), but no difference was observed under non-photorespiratory conditions (2% O2) (Fig. 3a, c) indicating that the down-regulation of GYD1 in lci20 is caused by the activity of photorespiration. The abundances of other photorespiration or CCM proteins were not consistently different in all three biological replicates between lci20 and control strains (Fig. 3a and Supplementary Fig. 7).
a Immunoblot analysis of representative CCM and photorespiration related proteins in lci20, its wild-type control CC4533 and one complemented line (C1). b Immunoblot analysis of representative CCM and photorespiration related proteins in cia5 and its wild-type control CC125. α-tubulin was used as a loading control. Numbers after protein names represent the calculated molecular weights. Relative abundance of GYD1 protein in lci20 (c) and cia5 (d) grown under various CO2 and O2 levels. The GYD1 immunoblots were normalized by α-tubulin signals. Data are means of three independent biological replicates (n = 3 ± SD). ns, not significant. Asterisks represent statistically significant difference compared to the wild-type CC4533 and/or CC125 under VL-CO2 21% O2 (*p ≤ 0.05, **p ≤ 0.01, and ****p ≤ 0.0001) using one-way ANOVA. Cells were cultivated photo-autotrophically under H-CO2 and 80 μmol photons m−2 s−1 and then acclimated for 20 h at the indicated CO2 and O2 levels prior to immunoblot analysis. Uncropped immunoblots are provided as a Source Data file.
In Chlamydomonas, photorespiration and CCM genes are co-regulated during acclimation to sub-optimal CO2 and controlled by a common transcription factor CIA5. We then monitored the accumulation of photorespiratory enzymes (GYD1, HPR1 and GCSP) and of CCM-related proteins (HLA3, LCI1 and LCIC) following acclimation to different CO2 and O2 levels in cia5 and its wild-type control (Fig. 3b and Supplementary Fig. 8). Whereas CCM and photorespiratory proteins were induced after 20 h of acclimation to VL-CO2 in a CIA5-dependent manner, only photorespiratory proteins were detected under H-CO2 (Fig. 3b, Supplementary Fig. 8), indicating that photorespiratory enzymes and CCM proteins are regulated differently at the protein level. Interestingly, photorespiratory proteins GCSP and GYD1 levels were more abundant in wild-type CC125 and cia5 under VL-CO2 at 21% O2 compared to VL-CO2 2% O2 whereas CCM related proteins were unaffected by O2 levels (Fig. 3b, d, Supplementary Fig. 8). HPR1 abundance was only affected by O2 levels in the cia5 (Supplementary Fig. 8b). This effect of O2 levels on the abundance of photorespiratory proteins was not observed in the wild-type CC4533 strain (Supplementary Fig. 7b), which could indicate a feedback regulation of photorespiratory proteins.
To better understand the reasons for the growth defect observed in lci20, and further determine whether it may be caused by the observed decrease of GYD1 abundance in lci20 mutant (Fig. 3c), we have isolated and characterized two CLiP mutants of Chlamydomonas harboring insertions in introns of the GYD1 gene (Fig. 4a) and validated to be deficient in the GYD1 protein by immunoblot (Fig. 4b). Both mutants showed higher glycolate excretion than the control strain (Fig. 4c). However, the growth of both gyd1 mutants was similar to the control wild-type strain after a transition from H-CO2 to VL-CO2 regardless of O2 levels (Fig. 4d, Supplementary Fig. 9). gyd1 mutants did not require H-CO2 to grow, which is in contrast to a previous work where the H-CO2 requiring 89 mutant (HCR89) is reported to harbor a mutation at the GYD1 locus45. Therefore, the growth defect of lci20 observed during H-CO2 to VL-CO2 is unlikely due to a decrease in GYD1 abundance but rather results from the accumulation of photorespiratory metabolites downstream the GYD1 reaction step. Indeed, the lci20 mutants excreted half as much glycolate as gyd1 mutants (Figs. 2b and 4c), which likely explain why their growth is more affected during a transition from H-CO2 to VL-CO2 under photorespiratory conditions.
a Genomic structure of GYD1 gene and insertion sites of the paromomycin resistance cassettes in the two CLiP mutants gyd1-1 and gyd1-2. Gray boxes at both extremities represent the 5’ and 3’UTR, respectively. Exons are colored blue and the position of cassette insertion is indicated by red arrows. b Immunoblot analysis using anti-GYD1 antibodies performed on both gyd1 mutants and their WT control grown under H-CO2 or VL-CO2. α-tubulin was used as a loading control. Uncropped immunoblots are provided as a Source Data file. c Quantification of the glycolate concentration in the culture medium after a transition from H-CO2 to VL-CO2 at 21% or 2% O2. Bars show the average and dots show data from independent biological replicates (n = 3 ± SD). d Photoautotrophic growth of both gyd1 mutants, their wild-type control CC4533 and cia5 and its wild-type control CC125 on agar plates exposed to various CO2 and O2 concentrations. Cells were grown in liquid culture in flasks photo-autotrophically under H-CO2 prior to spot test. Images were taken after 3 days (H-CO2) or 5 days (VL-CO2) of growth under 80 μmol photons m−2 s−1.
Photosynthesis of lci20 is affected under photorespiratory conditions
To investigate the consequence of impaired photorespiration on the photosynthetic capacity of lci20, we performed room temperature chlorophyll fluorescence measurements to assess the PSII quantum yield and PQ redox state (1-qL) in H-CO2 grown cells and following their acclimation to VL-CO2 (Fig. 5, Supplementary Fig. 10). Under H-CO2, lci20 showed a slightly higher effective PSII quantum yield and lower PQ redox state compared to wild-type but was mainly similar to the complemented line (Fig. 5a, d). Following VL-CO2 acclimation under photorespiratory conditions (21% O2), lci20 showed a significantly reduced effective PSII quantum yield and elevated PQ redox state compared to the control strains (wild-type and complemented line) (Fig. 5b, e). These effects were suppressed under non-photorespiratory conditions (VL-CO2, 2% O2) (Fig. 5c, f), thus indicating that the accumulation of non-metabolized photorespiratory compounds may be responsible for the inhibition.
PSII operating yields measured under actinic light in cells grown photo-autotrophically (a) under H-CO2, (b) during acclimation from H-CO2 to VL-CO2 at 21% O2 or (c) at 2% O2 for 20 h. d–f PQ redox state measured as the (1-qL) parameter in the same samples as in (a–c). Cells were exposed for 1 min at the indicated light intensity. Bars show the average and dots show data from independent biological replicates (n = 3 ± SD). Asterisks represent statistically significant difference compared to the wild-type strain CC4533 (*p ≤ 0.05 and **p ≤ 0.01) using two-way ANOVA.
Metabolomic analyses reveal an increased accumulation of malate and glutamate in lci20
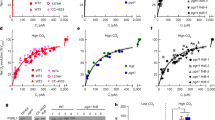
To investigate the metabolic changes occurring in lci20 and obtain clues to the metabolites transported by LCI20, we performed a metabolomic analysis in the different strains grown under photoautotrophic H-CO2, and during transitions from H-CO2 to L-CO2 and from H-CO2 to VL-CO2 (Fig. 6; Supplementary Fig. 11; Supplementary Data 1). The metabolite profile of H-CO2-grown lci20 cells was comparable to that of control strains (Fig. 6a Supplementary Fig. 11; Supplementary Data 1). After transition from H-CO2 to L-CO2 or to VL-CO2, we observed a higher accumulation of malic acid and glutamic acid in lci20 when cells were cultivated under photorespiratory condition (21% O2), the effect being suppressed under non-photorespiratory conditions (2% O2) (Fig. 6b, d). Accumulation of almost all the other metabolites detected was not significantly affected in lci20 under the conditions tested (Supplementary Fig. 11; Supplementary Data 1). Based on these results, we propose that LCI20 is involved in the transport of malate and glutamate during photorespiration and that disruption of glutamate transport impairs photorespiratory glycolate metabolism at the step of glyoxylate conversion to glycine thus explaining the lci20 phenotypes observed during acclimation to VL-CO2.
Accumulation of intracellular glutamic acid and malic acid in cells grown (a) under H-CO2, (b) or after transition from H-CO2 to L-CO2 for 20 h, (c) after transition from H-CO2 to VL-CO2 under 21% O2 or (d) under 2% O2 for 20 h. Bars show the average and dots show data from independent biological replicates (n = 8 ± SD). Asterisks represent statistically significant difference compared to the wild-type strain CC4533 (*p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, and ****p ≤ 0.0001) using two-way ANOVA. A.U., arbitrary unit; ns, not significant.
Discussion
In this study, we demonstrate the role of LCI20 in the photorespiratory glycolate metabolism in Chlamydomonas (Fig. 7), as evidenced by a high glycolate excretion and the marked decline in growth of the lci20 mutant during a transition from H-CO2 to VL-CO2 at ambient O2 levels (21%), these effects being suppressed at 2% O2 (Figs. 1, 2). Many photorespiratory mutants have been previously reported in land plants and only very few in algae. In Arabidopsis, a defect in genes encoding photorespiratory enzymes generate a high-CO2 requiring phenotype, which is attributed to the accumulation of non-metabolized photorespiratory intermediates13. In Chlamydomonas, the situation is not that clear, and this is further complicated by the occurrence of CCM in algae. A phosphoglycolate phosphatase 1 (pgp1) mutant, impaired in the conversion of 2-PG to glycolate, exhibits a growth defect under stationary conditions of limiting CO2 levels24. The high-CO2 growth requirement of pgp1 was attributed to the inhibitory effect of non-metabolized 2-PG accumulating upon ribulose bisphosphate oxygenation24. After the conversion of 2-PG into glycolate by PGP1, glycolate can follow two different fates in Chlamydomonas, it can be either excreted out of the cell or metabolized into glyoxylate by GYD1 in the mitochondria. A Chlamydomonas gyd1 mutant isolated from a screening for an H-CO2 requirement showed a high glycolate excretion during transition from H-CO2 to L-CO245. However, the H-CO2 requiring phenotype of this mutant was not clearly evidenced. We have shown here that impairing photorespiration at the level of GYD1 does not lead to an H-CO2 requirement but instead to a high glycolate excretion under VL-CO2 (Fig. 4, Supplementary Fig. 9). It was recently reported that growth of Chlamydomonas hpr1 mutant is slightly impaired during the transition from H-CO2 to L-CO2, accompanied by a high glycolate excretion57. The slight growth defect of the hpr1 mutant being interpreted as a consequence of glycolate hyper-excretion57. However, the gyd1 mutants characterized here show high glycolate excretion rates but no growth defect in any CO2 conditions tested (Fig. 4, Supplementary Fig. 9). We therefore conclude that inhibition of photorespiration at different steps of the pathway can lead to contrasting phenotypes. While inhibition of glycolate conversion does not impair growth (i.e. in gyd1), the accumulation of intermediate metabolites such as 2-PG (i.e. in pgp1) or downstream glycolate conversion (i.e. in hpr1) is deleterious. The strong down-regulation of GYD1 observed in lci20 under photorespiratory conditions (Fig. 3a) may result from a protective mechanism aiming at limiting the toxicity of non-metabolized photorespiratory intermediates by decreasing their production while promoting glycolate excretion. Consequently, glycolate excretion together with GYD1 down-regulation could be seen as complementary protection mechanisms preventing the toxic accumulation of photorespiratory intermediates during acclimation to VL-CO2. Such mechanisms would be particularly needed until CCM components and photorespiratory enzymes are fully induced, thus allowing photorespiratory metabolism to be tuned to the residual Rubisco oxygenase activity which remains at VL-CO2 despite the presence of CCM.
Photorespiration is initiated by the oxygenase activity of Rubisco producing phosphoglycolate (2-PG), which is converted into glycolate by a phospho-glycolate phosphatase (PGP), and then glyoxylate by the glycolate dehydrogenase (GYD). Glyoxylate is transaminated into glycine by a glutamate-glyoxylate aminotransferase (GGAT) using glutamate as a -NH2 donor. During the condensation of two molecules of glycine into one serine by the glycine decarboxylase (GDC), CO2 and NH3 are released. Photorespiratory NH3 is likely reassimilated by the GS/COGAT as proposed in Arabidopsis59,87 and Chlamydomonas17,68. LCI20 would function as a glutamate transporter exporting glutamate out of the chloroplast in counter-exchange with malate, thus supplying a -NH2 donor for glyoxylate conversion into glycine. The growth phenotype of lci20 observed during a transition from H-CO2 to VL-CO2 would result from the accumulation of glyoxylate. The absence of lci20 phenotype under steady state L-CO2 or VL-CO2 would result from the induction of an alternative recycling pathway for the photorespiratory NH3, for instance the mitochondrial glutamate dehydrogenase (GDH).
Unlike the above-mentioned proteins (PGP1, GYD1 and HPR1), which are all involved in the core photorespiratory pathways, LCI20 turns out to be an auxiliary protein involved in the supply of amino (-NH2) donor as glutamate for the conversion of glyoxylate into glycine (Fig. 7). In Arabidopsis chloroplasts, the export of glutamate in counter-exchange with malate is mediated by the dicarboxylate transporter DiT2.1 that is required for photorespiratory nitrogen assimilation and displays a 60% homology with the Chlamydomonas LCI2058,59,60,61. Based on our metabolomic analysis, we suggest that LCI20 mediates the export of glutamate in counter exchange with malate across the chloroplast inner envelope (Fig. 6; Supplementary Fig. 11). Impairing glutamate export by knocking out LCI20 likely disrupts glycolate metabolism at the step of glyoxylate transamination. The higher glycolate excretion observed in lci20 (Fig. 2b) would then result from an inefficient conversion of glycolate into glyoxylate, which would in turn result in a decrease in the GYD1 abundance (Fig. 3a, c). Both glycolate and glyoxylate, which could not be determined in our metabolomic analysis, are intermediate metabolites whose accumulation has been shown to be toxic for plants62,63. Because Chlamydomonas is unlikely to accumulate glycolate due to the existence of an excretion pathway, we conclude that the growth impairment of lci20 does not result from glycolate excretion per se, but rather from poisoning by non-metabolized glyoxylate. In line with our data, mutants deficient in the LCI20 homologues DiT2.1 and OMT1 in tobacco and Arabidopsis showed photoinhibition of PSII caused by the negative feedback of non-metabolized glyoxylate on the activation state of the Calvin cycle enzymes60,64,65. Glyoxylate was shown to be over accumulated in omt1 mutants in both Arabidopsis and tobacco60,64. The negative effects of glyoxylate on the activation state of Rubisco was shown in several studies66,67. Overall, LCI20 plays a vital role during the acclimation of Chlamydomonas from H-CO2 to VL-CO2 by exporting glutamate from chloroplast toward mitochondria which is required for the function of photorespiratory pathway to efficiently recycle Rubisco oxygenation product 2-PG back to the Calvin cycle in the chloroplast.
The fact that lci20 growth is unaffected under steady-state L-CO2 or VL-CO2 could be explained in two different ways. On the one hand, we could consider that photorespiration is only transiently active before the activation of CCM during acclimation to sub-optimal CO2 levels but is suppressed or strongly reduced upon CCM activation under steady-state conditions. A similar conclusion was drawn from the observation that the glycolate excretion is suppressed in air acclimated Chlamydomonas wild-type cells when CCM is fully induced18. However, we can also imagine that the arrest of glycolate excretion is caused by the upregulation of enzymes of the glycolate metabolic pathway both at transcript and protein levels. The later hypothesis is supported by the fact that the transaminase inhibitor aminooxyacetate (AOA), which blocks the photorespiratory pathway, induces a higher glycolate excretion in air-adapted Chlamydomonas wild-type cells with a fully activated CCM45. Alternatively, we could consider that during photorespiration the need for LCI20-mediated -NH2 recycling is only transient. Indeed, two -NH2 are needed for the conversion of two molecules of glyoxylate into two glycines. One -NH2 is likely supplied by the conversion of one serine into one hydroxypyruvate. Since the conversion of two molecules of glycine into one serine is accompanied by the release of NH3, the later needs to be re-assimilated to balance the nitrogen budget. In Chlamydomonas, inhibition of glutamine synthetase induces excretion of NH3 in an O2-dependent manner therefore showing that at least part of the NH3 produced by photorespiration is recycled within chloroplasts by the GS-GOGAT pathway68. In Arabidopsis, the re-assimilation of photorespiratory NH3 is mediated by the chloroplast GS/GOGAT cycle, which produces glutamate in turn re-imported into mitochondria; this process is essential for photorespiration as evidenced by the lethal phenotype of both chloroplast and mitochondrial glutamate transporters59,69,70. In this context, the transient phenotype of lci20 may result from the induction of an alternative pathway of NH3 reassimilation for instance through a mitochondrial isoform of the glutamate dehydrogenase71 which may compensate for the lack of LCI20 under steady state limiting CO2 (Fig. 7).
An hypothesis often put forward regarding the physiological role of photorespiration in microalgae is that photorespiratory metabolites could be the metabolic signal responsible for the induction of CCM during acclimation to sub-optimal CO2 levels11,14,22,23,24,72,73. Our data clearly establish that both the accumulation of CCM components (Fig. 3b, Supplementary Fig. 8) and the CCM functioning (Fig. 2a) are independent of O2 levels, thus ruling out this hypothesis. In C3 plants, it has recently been proposed that photorespiration helps to maintain cellular redox balance by generating malate in the chloroplast, with malate export to the cytosol providing NADH for nitrogen assimilation74. Such a role in redox balance or any other physiological function of photorespiration remains to be demonstrated in microalgae.
This work paves the way for a more detailed understanding of photorespiration in microalgae and how it interacts with the CCM functioning. Significant amounts of glycolate have been found in the ocean, particularly during algal blooms75,76,77,78,79, suggesting that photorespiration can be active in phytoplankton despite the occurrence of CCMs in most species9,80,81. This apparent discrepancy could be due to the fact that CCMs from different marine species show quite different efficiencies9,80,81. For instance, Nannochloropsis has been reported to harbour a quite leaky CCM82. It is also possible that algal species with an efficient CCM excrete significant glycolate amounts in particular conditions, for instance when the CO2 concentration drops during algal blooms, or when the incident light suddenly increases. Further studies are needed to better understand how algal photorespiration contributes to the carbon footprint of oceans.
Methods
Growth conditions and strains
The lci20 (LMJ.RY0402.205944), gyd1-1 (LMJ.RY0402.206272), gyd1-2 (LMJ.RY0402.232044), cia5 (CC-2702), and wild-types CC4533 and CC125 were purchased from the Chlamydomonas resource center. Cells were cultivated in an incubation shaker (INFORS Multitron pro) maintained at 25 °C, with 120 rpm shaking and constant illumination at 80 µmol m−2 s−1 supplied by fluorescent tubes delivering white light enriched in red wavelength. Cells were grown in MOPS-buffered (20 mM MOPS, pH 7.2) minimal medium (MM) exposed to various CO2 levels. Photorespiration was induced by transferring 2% (H) CO2-grown cells into 0.01% (VL) CO2 under atmospheric O2 level (21%) and can be suppressed by 2% O2. Growth kinetics were monitored with a Multisizer 3 Coulter counter (Beckman Coulter). For the wild-type and gyd1-1 growth assays performed in liquid culture, cells were cultivated photoautotrophically in 1-L photobioreactors operated as turbidostats. Cell density was monitored through turbidity measurement (OD 800 nm) and maintained at 0.4 O.D by addition of fresh media. The temperature was maintained at 25 °C, the pH at 7.2, at the light intensity at 125 µmol photons m−2 s−1. Cultures were bubbled with gas mixtures containing CO2 at 4%, 0.1%, or 0.04% in air, and supplied at a constant flow rate of 0.5 L min−1. The dissolved CO2 concentration in the photobioreactor medium was measured by membrane inlet mass spectrometry83 upon rapid filtration of the culture. For spot test, cells were grown in liquid culture with MM, harvested during active growth at around 10 µg mL−1 of chlorophyll and resuspended in fresh MM medium to make series of dilutions to 0.5, 1, and 2 µg mL−1 chlorophyll per spot. Eight-microliter drops were spotted on 1.5% of MM agar plates at pH 7.2 buffered with 20 mM MOPS and exposed to various CO2 and light regimes. Homogeneous light was supplied by a panel of fluorescent tubes.
Protein extraction and immunoblot analysis
Total protein was extracted as previously described in ref. 26. Exponentially grown cells (equivalent of 20 µg of chlorophyll) were harvested by centrifugation at 4000 g for 3 min at 4 °C. Pellets were resuspended in 200 µL of PBS with a complete protease inhibitor EDTA-free mixture tablet (Roche) by vortexing and sonicated for 15 s 30% pulses on ice using a sonicator (product number UR-21P; TOMY). 200 μL of Novex™ Nupage™ LDS buffer 2x (Invitrogen™) containing 1x reducing agent DTT was added to the solution, and the total protein was solubilized by incubation at 37 °C for 20 min. Incubated samples were subsequently centrifuged at 4000 g for 3 min. 10 μL (1X) of protein samples were loaded on Novex™ Nupage™ Bis tris 10% or tris-acetate 3-8% (Invitrogen™) gel, migrated 1 h at 190 V in Novex™ Nupage™ MOPS or tris-acetate (Invitrogen™) buffer according to protein molecular weight and transferred to nitrocellulose membrane using semidry transfer technique. Immuno-detection was performed using antibodies raised against LCI20 (1/250), GYD1 (1/500), HPR1 (1/500), GCSP (1/500), HLA3 (1/500), LCI1 (1/500) and LCIC (1/500). Antibody raised against α-Tubulin 1/2000 (Sigma-Aldrich ref. T6074) was used as control. Secondary anti-rabbit peroxidase-conjugated antibodies (Sigma-Aldrich; no. AQ132P) (1/10,000) were used for the detection with the G:BOX Chemi XRQ system (Syngene) using ECL detection reagents (GE Healthcare). Images were captured with a CCD camera equipped with a GeneSys Image Acquisition Software (Syngene).
Glycolate quantification
Excreted glycolic acid was analyzed from the supernatant after centrifugation of 1 mL culture at 4000 g for 2 min at 4 °C. The supernatant was diluted 10x in acetonitrile and analyzed using a Vanquish UHPLC/Q Exactive Plus (ThermoScientific). Glycolic acid was separated using a HILIC stationary phase (SeQuant ZIC HILIC, 100 × 2.1 mm, 3.5 µm), heated to 35 °C. A binary solvent system was used, in which mobile phase A consisted of acetonitrile: water (95:5, v/v) with 5 mM ammonium acetate and mobile phase B consisted of water:acetonitrile (95:5, v/v) with 5 mM ammonium acetate. Separations were made over a 32 min period following the gradient: 0-0.5 min: 5% B, 0.5–24.5 min: 5-95% B, 24.5–26.5 min: 95% B, 26.5–26.6 min: 95-5% B, 26.6–32 min: 5% B. The flow rate was set to 0.3 mL min–1 and the injection volume was 5 µL. After separation, glycolic acid was directed into the ESI source of the Q Exactive Plus (Orbitrap-mass spectrometer). The ESI source was set as following: negative mode ion spray voltage at –2.5 kV, capillary temperature at 300 °C, S-lens RF level at 50. Data was acquired using a targeted t-SIM method with glycolic acid in inclusion list (formula C2H4O3, m/z 75.00877).
Measurement of chlorophyll fluorescence
Chlorophyll fluorescence measurements were performed using a Dual Pulse Amplitude Modulated Fluorometer (DUAL-PAM-100; Walz) equipped with a red LED source of actinic light. Samples (2 mL of cells at 10 µg mL–1 chlorophyll) were placed into a cuvette under constant stirring at room temperature (25 °C) and dark adapted for 15 min prior to measurement. Both effective PSII quantum yield and 1-qL were calculated from the light curve. The latter was obtained by increasing red actinic light stepwise every 1 min starting from 17, 110, 190, 390, 610 and 940 μmol photons m−2 s−1, each being separated by a saturating pulse 10,000 μmol photons m−2 s−1, 600 ms duration. The effective PSII quantum yield and 1-qL were calculated as ΦPSII = (Fm’-Fs)/Fm’ and qL = ((Fm’-Fs)/(Fm’-F0))*(F0/Fs) respectively with Fm’ the fluorescence value after saturating pulses, Fs the stationary fluorescence at each actinic light and F0 the fluorescence value in the dark84.
Metabolomics analysis
Cells were grown to exponential phase under photoautotrophic H-CO2 conditions in the flask with 80 μmol photons m−2 s−1. For L-CO2 and VL-CO2 conditions, H-CO2 grown cells were transferred to either L-CO2 or VL-CO2 levels for 20 h prior to sampling. About 60 million cell suspensions were injected into a −70 °C cold quenching solution composed of 70% methanol in water using a thermoblock above dry ice to obtain a final concentration of 35% methanol. Centrifuge tubes containing the cell suspensions were cooled in a pre-chilled cooling box to keep the sample temperature below −20 °C. Cell pellets were collected by centrifugation at 4000 g for 2 min at −10 °C. The supernatant was decanted and residual liquid together with cells carefully transferred to a 2 mL Eppendorf tube and re-centrifuged 13,000 g for 1 min at −10 °C. The pellet was flash-frozen in liquid nitrogen and lyophilized at −50 °C. The extraction and derivation of the metabolites were performed as previously85. The analytic and quantification methods were exactly as reported in ref. 37. Data is reported following recently updated metabolomics standards86.
Statistics
All statistical tests used are noted in figure legends. One-way or two-way ANOVA using GraphPad Prism (GraphPad Software) was used to perform statistical analysis. The P-values were computed by One-way or two-way ANOVA test (uncorrected p values). Statistical significance (α = 0.05) according to p values is indicated by asterisk (*p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001 and ****p ≤ 0.0001).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Genes studied in this article can be found on Phytozome 13 (Chlamydomonas genome v6.1 or 5.6) under the loci Cre06.g260450 [LCI20], Cre06.g295450 [HPR1], Cre06.g288700 [GYD1], Cre12.g534800 [GCSP], Cre03.g162800 [LCI1], Cre02.g097800 [HLA3], Cre06.g307500 [LCIC] and Cre02.g096300 [CIA5]. All antibodies used are available upon request to corresponding authors. Metabolites data are provided as supplementary data 1. All the other methods are reported in the Supplementary Information file . Source data are provided with this paper.
References
Field, C. B., Behrenfeld, M. J., Randerson, J. T. & Falkowski, P. Primary production of the biosphere: integrating terrestrial and oceanic components. Science 281, 237–240 (1998).
Smetacek, V. Diatoms and the ocean carbon cycle. Protist 150, 25–32 (1999).
Fernández, F. G. A. et al. The role of microalgae in the bioeconomy. N. Biotechnol. 61, 99–107 (2021).
Ullmann, J. & Grimm, D. Algae and their potential for a future bioeconomy, landless food production, and the socio-economic impact of an algae industry. Org. Agr. 11, 261–267 (2021).
Reisoglu, Ş., Aydin, S., Reisoglu, Ş. & Aydin, S. Microalgae as a promising candidate for fighting climate change and biodiversity loss. (IntechOpen, 2023). https://doi.org/10.5772/intechopen.1002414.
Moroney, J. V. et al. The carbonic anhydrase isoforms of Chlamydomonas reinhardtii: intracellular location, expression, and physiological roles. Photosynth Res. 109, 133–149 (2011).
Wang, Y., Stessman, D. J. & Spalding, M. H. The CO2 concentrating mechanism and photosynthetic carbon assimilation in limiting CO2: how Chlamydomonas works against the gradient. Plant J. 82, 429–448 (2015).
Mackinder, L. C. M. The Chlamydomonas CO2-concentrating mechanism and its potential for engineering photosynthesis in plants. N. Phytologist 217, 54–61 (2018).
Barrett, J., Girr, P. & Mackinder, L. C. M. Pyrenoids: CO2-fixing phase separated liquid organelles. Biochimica et. Biophysica Acta (BBA) - Mol. Cell Res. 1868, 118949 (2021).
He, S., Crans, V. L. & Jonikas, M. C. The pyrenoid: the eukaryotic CO2-concentrating organelle. Plant Cell 35, 3236–3259 (2023).
Marek, L. F. & Spalding, M. H. Changes in photorespiratory enzyme activity in response to limiting CO2 in Chlamydomonas reinhardtii. Plant Physiol. 97, 420–425 (1991).
Spalding, M. H. Chapter 8 - The CO2-concentrating mechanism and carbon assimilation. In The Chlamydomonas Sourcebook (Second Edition) (eds. Harris, E. H., Stern, D. B. & Witman, G. B.) 257–301 (Academic Press, London, 2009). https://doi.org/10.1016/B978-0-12-370873-1.00016-2.
Eisenhut, M., Roell, M.-S. & Weber, A. P. M. Mechanistic understanding of photorespiration paves the way to a new green revolution. N. Phytologist 223, 1762–1769 (2019).
Santhanagopalan, I., Wong, R., Mathur, T. & Griffiths, H. Orchestral manoeuvres in the light: crosstalk needed for regulation of the Chlamydomonas carbon concentration mechanism. J. Exp. Bot. 72, 4604–4624 (2021).
Shi, M., Zhao, L. & Wang, Y. Identification and characterization of genes encoding the hydroxypyruvate reductases in Chlamydomonas reveal their distinct roles in photorespiration. Front Plant Sci. 12, 690296 (2021).
Nelson, E. B. & Tolbert, N. E. The regulations of glycolate metabolism in Chlamydomonas reinhardtii. Biochimica et. Biophysica Acta (BBA) - Gen. Subj. 184, 263–270 (1969).
Peltier, G. & Thibault, P. Light-dependent oxygen uptake, glycolate, and ammonia release in l-methionine sulfoximine-treated Chlamydomonas. Plant Physiol. 77, 281–284 (1985).
Moroney, J. V., Wilson, B. J. & Tolbert, N. E. Glycolate metabolism and excretion by Chlamydomonas reinhardti. Plant Physiol. 82, 821–826 (1986).
Yamano, T., Miura, K. & Fukuzawa, H. Expression analysis of genes associated with the induction of the carbon-concentrating mechanism in Chlamydomonas reinhardtii. Plant Physiol. 147, 340–354 (2008).
Brueggeman, A. J. et al. Activation of the carbon concentrating mechanism by CO2 deprivation coincides with massive transcriptional restructuring in Chlamydomonas reinhardtii. Plant Cell 24, 1860–1875 (2012).
Fang, W. et al. Transcriptome-wide changes in Chlamydomonas reinhardtii gene expression regulated by carbon dioxide and the CO2-concentrating mechanism regulator CIA5/CCM1. Plant Cell 24, 1876–1893 (2012).
Marcus, Y., Harel, E. & Kaplan, A. Adaptation of the cyanobacterium Anabaena variabilis to low CO2 concentration in their environment. Plant Physiol. 71, 208–210 (1983).
Spalding, M.H., Spreitze, R. J. & Ogren, W. L. Use of mutants in analysis of the CO2-concentrating pathway of Chlamydomonas reinhardtii. In Inorganic Carbon Uptake by Photosynthetic Organisms (eds Lucas, W. & Berry, J.) 361–375 (American Society of Plant Physiologists, Rockville, Maryland, 1985).
Suzuki, K., Marek, L. F. & Spalding, M. H. A photorespiratory mutant of Chlamydomonas reinhardtii. Plant Physiol. 93, 231–237 (1990).
Wang, Y. & Spalding, M. H. Acclimation to very low CO2: contribution of limiting co2 inducible proteins, LCIB and LCIA, to inorganic carbon uptake in Chlamydomonas reinhardtii. Plant Physiol. 166, 2040–2050 (2014).
Yamano, T., Sato, E., Iguchi, H., Fukuda, Y. & Fukuzawa, H. Characterization of cooperative bicarbonate uptake into chloroplast stroma in the green alga Chlamydomonas reinhardtii. Proc. Natl. Acad. Sci. 112, 7315–7320 (2015).
Geraghty, A. M. & Spalding, M. H. Molecular and structural changes in Chlamydomonas under limiting CO2 (a possible mitochondrial role in adaptation). Plant Physiol. 111, 1339–1347 (1996).
Burlacot, A. et al. Alternative photosynthesis pathways drive the algal CO2-concentrating mechanism. Nature 1–6 https://doi.org/10.1038/s41586-022-04662-9 (2022).
Burlacot, A. Quantifying the roles of algal photosynthetic electron pathways: a milestone towards photosynthetic robustness. N. Phytologist 240, 2197–2203 (2023).
Evans, E. H. & Carr, N. G. The interaction of respiration and photosynthesis in microalgae. In Photosynthesis II: Photosynthetic Carbon Metabolism and Related Processes (eds. Gibbs, M. & Latzko, E.) 163–173 (Springer, Berlin, Heidelberg, 1979). https://doi.org/10.1007/978-3-642-67242-2_13.
Peltier, G. & Thibault, P. O2 uptake in the light in Chlamydomonas. Plant Physiol. 79, 225–230 (1985).
Lemaire, C., Wollman, F. A. & Bennoun, P. Restoration of phototrophic growth in a mutant of Chlamydomonas reinhardtii in which the chloroplast atpB gene of the ATP synthase has a deletion: an example of mitochondria-dependent photosynthesis. Proc. Natl. Acad. Sci. USA 85, 1344–1348 (1988).
Cardol, P. et al. Photosynthesis and state transitions in mitochondrial mutants of Chlamydomonas reinhardtii affected in respiration. Plant Physiol. 133, 2010–2020 (2003).
Peltier, G. et al. Alternative electron pathways of photosynthesis power green algal CO2 capture. Plant Cell koae143 https://doi.org/10.1093/plcell/koae143 (2024).
Dang, K.-V. et al. Combined increases in mitochondrial cooperation and oxygen photoreduction compensate for deficiency in cyclic electron flow in Chlamydomonas reinhardtii. Plant Cell 26, 3036–3050 (2014).
Bailleul, B. et al. Energetic coupling between plastids and mitochondria drives CO2 assimilation in diatoms. Nature 524, 366–369 (2015).
Kong, F. et al. Interorganelle communication: peroxisomal malate dehydrogenase 2 connects lipid catabolism to photosynthesis through redox coupling in Chlamydomonas. Plant Cell 30, 1824–1847 (2018).
Fridlyand, L. E., Backhausen, J. E. & Scheibe, R. Flux control of the malate valve in leaf cells. Arch. Biochem. Biophys. 349, 290–298 (1998).
Scheibe, R. Malate valves to balance cellular energy supply. Physiologia Plant. 120, 21–26 (2004).
Selinski, J. & Scheibe, R. Malate valves: old shuttles with new perspectives. Plant Biol. (Stuttg.) 21, 21–30 (2019).
Dao, O., Kuhnert, F., Weber, A. P. M., Peltier, G. & Li-Beisson, Y. Physiological functions of malate shuttles in plants and algae. Trends Plant Sci. 27, 488–501 (2022).
Merchant, S. S. et al. The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 318, 245–250 (2007).
Johnson, X. & Alric, J. Central carbon metabolism and electron transport in Chlamydomonas reinhardtii: metabolic constraints for carbon partitioning between oil and starch. Eukaryot. Cell 12, 776–793 (2013).
Burlacot, A. & Peltier, G. Energy crosstalk between photosynthesis and the algal CO2-concentrating mechanisms. Trends Plant Sci. https://doi.org/10.1016/j.tplants.2023.03.018 (2023).
Nakamura, Y., Kanakagiri, S., Van, K., He, W. & Spalding, M. H. Disruption of the glycolate dehydrogenase gene in the high-CO2-requiring mutant HCR89 of Chlamydomonas reinhardtii. Can. J. Bot. 83, 820–833 (2005).
Zones, J. M., Blaby, I. K., Merchant, S. S. & Umen, J. G. High-resolution profiling of a synchronized diurnal transcriptome from Chlamydomonas reinhardtii reveals continuous cell and metabolic differentiation. Plant Cell 27, 2743–2769 (2015).
Karpowicz, S. J., Prochnik, S. E., Grossman, A. R. & Merchant, S. S. The GreenCut2 resource, a phylogenomically derived inventory of proteins specific to the plant lineage. J. Biol. Chem. 286, 21427–21439 (2011).
Mackinder, L. C. M. et al. A spatial interactome reveals the protein organization of the algal CO2-concentrating mechanism. Cell 171, 133–147.e14 (2017).
Wang, L. et al. A chloroplast protein atlas reveals punctate structures and spatial organization of biosynthetic pathways. Cell 186, 3499–3518.e14 (2023).
Li, X. et al. An indexed, mapped mutant library enables reverse genetics studies of biological processes in Chlamydomonas reinhardtii. Plant Cell 28, 367–387 (2016).
Moroney, J. V. et al. Isolation and characterization of a mutant of Chlamydomonas reinhardtii deficient in the CO2 concentrating mechanism. Plant Physiol. 89, 897–903 (1989).
Fukuzawa, H. et al. CCM1, a regulatory gene controlling the induction of a carbon-concentrating mechanism in Chlamydomonas reinhardtii by sensing CO2 availability. PNAS 98, 5347–5352 (2001).
Mukherjee, A. et al. Thylakoid localized bestrophin-like proteins are essential for the CO2 concentrating mechanism of Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 116, 16915–16920 (2019).
Terentyev, V. V. & Shukshina, A. K. CAH3 from Chlamydomonas reinhardtii: unique carbonic anhydrase of the thylakoid lumen. Cells 13, 109 (2024).
Taubert, A., Jakob, T. & Wilhelm, C. Glycolate from microalgae: an efficient carbon source for biotechnological applications. Plant Biotechnol. J. 17, 1538–1546 (2019).
Yun, E. J. et al. Glycolate production by a Chlamydomonas reinhardtii mutant lacking carbon-concentrating mechanism. J. Biotechnol. 335, 39–46 (2021).
Shi, M., Zhao, L. & Wang, Y. Identification and Characterization of Genes Encoding the Hydroxypyruvate Reductases in Chlamydomonas Reveal Their Distinct Roles in Photorespiration. Front. Plant Sci. 12, 690296 (2021).
Weber, A. & Flügge, U. Interaction of cytosolic and plastidic nitrogen metabolism in plants. J. Exp. Bot. 53, 865–874 (2002).
Renné, P. et al. The Arabidopsis mutant dct is deficient in the plastidic glutamate/malate translocator DiT2. Plant J. 35, 316–331 (2003).
Kinoshita, H. et al. The chloroplastic 2-oxoglutarate/malate transporter has dual function as the malate valve and in carbon/nitrogen metabolism. Plant J. 65, 15–26 (2011).
Weber, A. P. M. & Linka, N. Connecting the plastid: transporters of the plastid envelope and their role in linking plastidial with cytosolic metabolism. Annu. Rev. Plant Biol. 62, 53–77 (2011).
Lu, Y. et al. Suppression of glycolate oxidase causes glyoxylate accumulation that inhibits photosynthesis through deactivating Rubisco in rice. Physiologia Plant. 150, 463–476 (2014).
Dellero, Y., Jossier, M., Schmitz, J., Maurino, V. G. & Hodges, M. Photorespiratory glycolate–glyoxylate metabolism. J. Exp. Bot. 67, 3041–3052 (2016).
Schneidereit, J., Häusler, R. E., Fiene, G., Kaiser, W. M. & Weber, A. P. M. Antisense repression reveals a crucial role of the plastidic 2-oxoglutarate/malate translocator DiT1 at the interface between carbon and nitrogen metabolism. Plant J. 45, 206–224 (2006).
Takahashi, S., Bauwe, H. & Badger, M. Impairment of the photorespiratory pathway accelerates photoinhibition of photosystem ii by suppression of repair but not acceleration of damage processes in Arabidopsis. Plant Physiol. 144, 487–494 (2007).
Campbell, W. J. & Ogren, W. L. Glyoxylate inhibition of ribulosebisphosphate carboxylase/oxygenase activation in intact, lysed, and reconstituted chloroplasts. Photosynth Res 23, 257–268 (1990).
Häusler, R. E., Bailey, K. J., Lea, P. J. & Leegood, R. C. Control of photosynthesis in barley mutants with reduced activities of glutamine synthetase and glutamate synthase. Planta 200, 388–396 (1996).
Peltier, G. & Thibault, P. Ammonia exchange and photorespiration in Chlamydomonas. Plant Physiol. 71, 888–892 (1983).
Eisenhut, M. et al. Arabidopsis A BOUT DE SOUFFLE is a putative mitochondrial transporter involved in photorespiratory metabolism and is required for meristem growth at ambient CO2 levels. Plant J. 73, 836–849 (2013).
Porcelli, V. et al. Molecular identification and functional characterization of a novel glutamate transporter in yeast and plant mitochondria. Biochimica et. Biophysica Acta (BBA) - Bioenerg. 1859, 1249–1258 (2018).
Moyano, E., Ramazanov, Z., Cárdenas, J. & Muñoz-Blanco, J. Intracellular localization of three l-glutamate dehydrogenase isozymes from Chlamydomonas reinhardtii. Plant Physiol. 100, 1575–1579 (1992).
Neofotis, P. et al. The induction of pyrenoid synthesis by hyperoxia and its implications for the natural diversity of photosynthetic responses in Chlamydomonas. eLife 10, e67565 (2021).
Adler, L. et al. New horizons for building pyrenoid-based CO2-concentrating mechanisms in plants to improve yields. Plant Physiol. 190, 1609–1627 (2022).
Shi, X., Hannon, N. M. & Bloom, A. J. Metals and other ligands balance carbon fixation and photorespiration in chloroplasts. Physiologia Plant. 176, e14463 (2024).
Leboulanger, C., Descolas-Gros, C. & Jupin, H. HPLC determination of glycolic acid in seawater. An estimation of phytoplankton photorespiration in the Gulf of Lions, western Mediterranean Sea. J. Plankton Res. 16, 897–903 (1994).
Leboulanger, C., Oriol, L., Jupin, H. & Desolas-gros, C. Diel variability of glycolate in the eastern tropical atlantic ocean. Deep Sea Res. 44, 2131–2139 (1997).
Leboulanger, C., Martin-Jézéquel, V., Descolas-Gros, C., Sciandra, A. & Jupin, H. J. Photorespiration in continuous culture of dunaliella tertiolecta (chlorophyta): relationships between serine, glycine, and extracellular glycolate. J. Phycol. 34, 651–654 (1998).
Schnitzler Parker, M., Armbrust, E. V., Piovia-Scott, J. & Keil, R. G. Induction of photorespiration by light in the centric diatom Thalassiosira weissflogii (bacillariophyceae): molecular characterization and physiological consequences. J. Phycol. 40, 557–567 (2004).
Schada von Borzyskowski, L. et al. Marine proteobacteria metabolize glycolate via the β-hydroxyaspartate cycle. Nature 575, 500–504 (2019).
Badger, M. R. et al. The diversity and coevolution of Rubisco, plastids, pyrenoids, and chloroplast-based CO2 -concentrating mechanisms in algae. Can. J. Bot. 76, 1052–1071 (1998).
Tsuji, Y., Mahardika, A. & Matsuda, Y. Evolutionarily distinct strategies for the acquisition of inorganic carbon from seawater in marine diatoms. J. Exp. Bot. 68, 3949–3958 (2017).
Gee, C. W. & Niyogi, K. K. The carbonic anhydrase CAH1 is an essential component of the carbon-concentrating mechanism in Nannochloropsis oceanica. Proc. Natl Acad. Sci. 114, 4537–4542 (2017).
Burlacot, A., Burlacot, F., Li-Beisson, Y. & Peltier, G. Membrane inlet mass spectrometry: a powerful tool for algal research. Front Plant Sci. 11, 1302 (2020).
Maxwell, K. & Johnson, G. N. Chlorophyll fluorescence—a practical guide. J. Exp. Bot. 51, 659–668 (2000).
Lisec, J., Schauer, N., Kopka, J., Willmitzer, L. & Fernie, A. R. Gas chromatography mass spectrometry–based metabolite profiling in plants. Nat. Protoc. 1, 387–396 (2006).
Alseekh, S. et al. Mass spectrometry-based metabolomics: a guide for annotation, quantification and best reporting practices. Nat. Methods 18, 747–756 (2021).
Coschigano, K. T., Melo-Oliveira, R., Lim, J. & Coruzzi, G. M. Arabidopsis gls mutants and distinct Fd-GOGAT genes. Implications for photorespiration and primary nitrogen assimilation. Plant Cell 10, 741–752 (1998).
Acknowledgements
O.D. thanks the French Atomic Energy and Alternative Energy Commission (CEA) for a PhD scholarship. We thank Shiyan Zheng for assistance in performing the genetic complementation experiment; Gaurav Kumar for technical assistance during microscopic observation; Stephanie Blangy for trials on generating antibodies against LCI20; Franziska Kuhnert for trials on the transport assays. We thank the ZoOM Microscopy facility (CEA Cadarache) and the University of York Biosciences Technology Facility for confocal microscopy access. This work is financially supported by the “L’économie Circular du Carbone” program of the CEA (CO2storage), the ANR project “AlgalCCM” (n°ANR-366 22-CE44-0023-01), the France 2030 initiative project “CO2_CMPhi” (ANR-23-PEXF-0002) and the Region Sud (“AlgalCO2” project). We acknowledge the European Union Regional Developing Fund (ERDF), the Région Provence Alpes Côte d’Azur, the French Ministry of Research and the CEA for funding the HelioBiotec platform. C.M. is grateful to the INRAE MIGALE bioinformatics facility (MIGALE, INRAE, 2020) for providing computing resources. AKL was supported by the Labex Saclay Plant Sciences-SPS (ANR-17-EUR-0007), the platform of Biophysics of the I2BC supported by the French Infrastructure for Integrated Structural Biology (FRISBI; grant number ANR-10-INSB-05). A.P.M.W. acknowledges funding by the European Union’s H2020 research and innovation and the Horizon programs (grants GAIN4CROPS, GA No. 862087 and BEST-CROP, GA No. 101082091) and the Deutsche Forschungsgemeinschaft (Cluster of Excellence for Plant Sciences (CEPLAS) under Germany’s Excellence Strategy EXC-2048/1 under project ID 390686111). S.A. and A.R.F. acknowledge the European Union’s Horizon 2020 research and innovation program, project PlantaSYST (SGA-CSA No. 739582 under FPA No. 664620) and the BG05M2OP001-1.003-001-C01 project, financed by the European Regional Development Fund through the Bulgarian ‘Science and Education for Smart Growth’ Operational Program. Both authors acknowledge the support by the Max Planck Society. A.B. acknowledges support from the Carnegie Institution for Science.
Author information
Authors and Affiliations
Contributions
Y.L-B., G.P., O.D. conceived the study. Y.L-B. and G.P. supervised the work. O.D. performed most of the experiments. A.B., A.K.L. and G.P. supervised photosynthesis measurement performed by O.D. P.A., M.B., S.C. and O.D. carried out immunoblots. O.D. and V.E. isolated and characterized gyd1 mutants with contribution from A.M. C.M. performed phylogenetic analysis. M.B., F.V. and O.D. performed genetic complementation of lci20 mutant. S.A. and A.R.F. performed metabolomics. O.D., P.-C.N. and B.L. performed glycolate analysis. O.D., G.P. and Y.L-B. drafted the manuscript with contributions from L.C.M.M., A.B., A.K.L., A.R.F. and A.P.W.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dao, O., Bertrand, M., Alseekh, S. et al. The green algae CO2 concentrating mechanism and photorespiration jointly operate during acclimation to low CO2. Nat Commun 16, 5296 (2025). https://doi.org/10.1038/s41467-025-60525-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-60525-7
This article is cited by
-
Integrative multi-omic and phenotypic analysis of open raceway pond production of Monoraphidium minutum 26B-AM reveals distinct stress signatures for scale-up and infection
Biotechnology for Biofuels and Bioproducts (2026)
-
Harnessing microalgae for bioproducts: innovations in synthetic biology
World Journal of Microbiology and Biotechnology (2025)