Abstract
Anhydromuropeptide permease (AmpG) is a transporter protein located in the inner membrane of certain gram -negative bacteria, involved in peptidoglycan (PG) recycling and β-lactamase induction. Decreased AmpG function reduces resistance of antibiotic-resistant bacteria to β-lactam antibiotics. Therefore, AmpG-targeting inhibitors are promising ‘antibiotic adjuvants’. However, as the tertiary structure of AmpG has not yet been identified, the development of targeted inhibitors remains challenging. We present four cryo-electron microscopy (cryo-EM) structures: the apo-inward and apo-outward state structures and the inward-occluded and outward states complexed with the substrate GlcNAc-1,6-anhMurNAc. Through functional analysis and molecular dynamics (MD) simulations, we identified motif A, which stabilizes the outward state, substrate-binding pocket, and protonation-related residues. Based on the structure of AmpG and our experimental results, we propose a muropeptide transport mechanism for AmpG. A deeper understanding of its structure and transport mechanism provides a foundation for the development of antibiotic adjuvants.
Similar content being viewed by others
Introduction
Antibiotic development, which began with the discovery of penicillin, not only significantly improved human health but also provided important inspiration for the research and understanding of microorganisms. However, owing to widespread antibiotic use and misuse, the emergence of antimicrobial-resistant (AMR) microorganisms is increasing1. The future is not promising2. If this problem is not resolved, we will not be free from diseases caused by pathogens. Development of new antibiotics is costly and time-consuming, and antibiotic-resistant strains have rapidly emerged. Only a few antibiotics have been approved since 2010s3. Attempts have been made to reduce the incidence of AMR microorganisms by using antibiotic combination therapy4,5.
However, the development of antibiotics alone cannot effectively suppress the emergence of AMR microorganisms. Therefore, antibiotic adjuvants are being explored as a complementary strategy to mitigate antimicrobial resistance6. These substances do not directly inhibit the growth of microorganisms but instead neutralize their antibiotic resistance mechanisms or enhance the effectiveness of antibiotics. Substances such as sulbactam and tazobactam, which act as β-lactamase inhibitors, are commonly used as β-lactam antibiotic adjuvants7. However, the types and applications of antibiotic adjuvants are limited, with FDA-approved options restricted to a few β-lactam antibiotics combined with β-lactamase inhibitors such as clavulanic acid, sulbactam, and avibactam8. Recently, research on compounds that inhibit efflux pumps and teichoic acid biosynthesis has been actively conducted9,10.
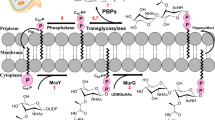
AmpG is a transporter protein found in the inner membrane of certain Gram-negative bacteria, such as Escherichia coli11 and Pseudomonas aeruginosa12, and plays a key role in peptidoglycan (PG) recycling. The PG layer undergoes continuous breakdown and reconstruction throughout bacterial growth and division. Its building blocks, muropeptides, are transported into the cytosol by AmpG. During this process, AmpG specifically recognizes the N-acetylglucosamine (GlcNAc)−1,6-anhydroMurNAc moiety of muropeptides and transports the entire muropeptide into the cytosol (Fig. 1a), functioning as a secondary active transporter that utilizes a proton concentration gradient13. Generally, AmpG prefers substrates in the anhydro form14, although homologs may exhibit slight differences in substrate specificity. It can transport muropeptides with one to up to five consecutive peptides attached to the sugar moiety15. Notably, AmpG can also transport GlcNAc-1,6-anhydroMurNAc even in the absence of an attached peptide, but it has no transport activity for the peptide alone13. Additionally, AmpG can transport substrates where the peptide is replaced with a fluorescent label16. However, the presence of a negative charge on the substrate significantly reduces its binding affinity17. Structurally, AmpG typically consists of 12 or 14 transmembrane helices arranged into two pseudosymmetric bundles18.
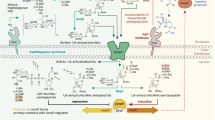
a Chemical structures of GlcNAc-1,6-anhydroMurNAc and its pentapeptide derivative. The red circle marks the peptide conjugation site. b Schematic of the AmpC induction pathway of AmpG. c Overall structure of AmpG in the apo outward state (green)(PDB:8ZBB), the outward state complexed with GlcNAc-1,6-anhMurNAc (cyan)(PDB:9J9Z), the inward-occluded state complexed with GlcNAc-1,6-anhMurNAc (orange)(PDB:8ZKE), and the apo inward state (magenta)(PDB:8ZGZ). AmpG: Anhydromuropeptide permease G, AmpR: AmpC regulator, ampC: β-lactamase gene, LT: Lytic transglycosylase, Ami: N-acetylmuramoyl-L-alanine amidase, PBP: Penicillin-binding protein, NagZ: N-acetyl-β-D-glucosaminidase Z.
Beyond its role in PG recycling, AmpG is crucial for the induction of β-lactamase (AmpC)19 in β-lactam-resistant Enterobacteriaceae (Fig. 1b). AmpC, a class C β-lactamase predominantly found in Enterobacteriaceae, hydrolyzes various β-lactam antibiotics, including penicillin. It shows activity against several classes of antibiotics, such as cephamycins (e.g., cefoxitin and cefotetan), oxyiminocephalosporins (e.g., ceftazidime, cefotaxime, and ceftriaxone), and the benzylpenicillin class of monobactams19,20.
In antibiotic-resistant bacteria, the transcriptional regulator AmpR controls AmpC expression. When β-lactams inhibit penicillin-binding proteins (PBPs) in the periplasm, the accumulation of GlcNAc-1,6-anhydromuropeptides triggers their transport into the cytoplasm by AmpG. PBPs are essential for bacterial cell wall biosynthesis, and their inhibition compromises cell wall integrity, initiating a stress response. Once inside the cytoplasm, these muropeptides are processed by NagZ and other enzymes, generating intermediates that activate AmpR. Conversely, under normal conditions, UDP-MurNAc-pentapeptide, a peptidoglycan precursor, binds to AmpR and suppresses ampC transcription21. Activated AmpR upregulates AmpC expression, leading to β-lactam degradation and increased antibiotic resistance. This feedback loop links AmpG-mediated PG recycling to the AmpR-AmpC regulatory pathway controlling β-lactam resistance22,23. Mutations or deficiencies in AmpG significantly reduce AmpC induction, as demonstrated by minimum inhibitory concentration (MIC) tests and β-lactamase activity assays24,25. Targeting AmpG could be a promising strategy for developing antibiotic adjuvants. However, designing potent inhibitors requires a detailed understanding of the tertiary structure and transport mechanisms of AmpG.
Using cryo-electron microscopy (cryo-EM), we identified four critical structures of AmpG from Yokenella regensburge that represent sequential states within its transport mechanism: the outward and inward states of apo AmpG and the outward and inward-occluded states of AmpG complexed with GlcNAc-1,6-anhMurNAc with resolutions ranging from 2.96 to 3.87 Å. Key residues observed in the structures were functionally validated using mutagenesis-based biological and biochemical experiments. In addition, molecular dynamics (MD) simulations were performed to investigate the effect of protonation on transport. Based on our experiments, we present an alternating access transport model for AmpG.
Results
Design and Functional Validation of AmpG Constructs for Cryo-EM Structural Determination
In this study, we designed two constructs to determine the cryo-electron microscopy (cryo-EM) structures of Yokenella regensburgei AmpG (Supplementary Fig. 1a). The first construct, AmpG (cryo), was generated by fusing a fiducial marker, engineered BRIL26,27,28, into the intracellular loop 3 (ICL3) between TM6 and 7 of the wild-type AmpG (AmpG(WT)) (Supplementary Fig. 1b). To determine the outward state structure, we introduced mutations at residues G50 and L269 (Supplementary Fig. 1h), identified at the periplasmic interface in the inward state, substituting them with tryptophan to induce the outward state29. This construct, termed AmpG(ww), was further engineered by fusing BRIL to create AmpG(cryo-ww) for structural determination.
To assess whether AmpG(cryo) retained its function, it was transformed into the Klebsiella aerogenes (KE-Y6) strain, previously used in our study25, which was derived from the β-lactam-resistant Klebsiella aerogenes (KE-Y1) strain by knocking out AmpG. AmpG(cryo) successfully restored antibiotic resistance to levels comparable to those of the wild type (Supplementary Fig. 1c) and conferred resistance to a broad spectrum of β-lactam antibiotics (Supplementary Fig. 1d). Furthermore, thermal shift assay (TSA) and microscale thermophoresis (MST) confirmed a concentration-dependent increase in thermal stability and affinity for GlcNAc-1,6-anhMurNAc (Supplementary Figs. 1e and 2b), indicating proper substrate recognition and functionality. In contrast, AmpG(ww), which was designed to lock the transporter in the outward state, failed to restore antibiotic resistance in Klebsiella aerogenes (KE-Y6) (Supplementary Fig. 1c, d). However, TSA and MST experiments confirmed its ability to bind GlcNAc-1,6-anhMurNAc(Supplementary Figs. 1e and 2c), consistent with previous reports showing that mutations locking major facilitator superfamily (MFS) transporters in a single state disrupt transport function while maintaining substrate binding29,30.
To optimize the performance of the BRIL fiducial marker in AmpG(cryo) and AmpG(cryo-ww), we utilized a Fab (BAG2) that specifically recognizes BRIL and a nanobody targeting BAG2 for cryo-EM grid preparation31. As a result, we determined the inward and outward state structures of apo AmpG at resolutions of 3.87 and 3.11 Å, respectively. Additionally, we identified the inward-occluded state in complex with GlcNAc-1,6-anhMurNAc at 3.72 Å and the outward state complex at 2.96 Å (Fig. 1b, Supplementary Figs. 3–6).
Periplasmic-cytoplasmic interface and motif A of AmpG
In the inward-state structure of AmpG, the N-lobe (TM1–6) and the C-lobe (TM7–12) open toward the cytoplasm (Fig. 2a). TM13 and TM14 are structurally distinct and exist independently of the N-lobe and C-lobe. On the periplasmic side, the two helical bundles were in contact, particularly between TM1 and TM7 (Fig. 2b). Mutating residues such as Thr40 and Thr247 in these helices to alanine did not affect antibiotic resistance, as shown by MIC tests. However, when these residues were replaced with bulkier residues like tryptophan or tyrosine, resistance to β-lactam antibiotics such as cefotaxime and ceftazidime was lost (Fig. 2e). Additionally, residues located at the contact surface on the periplasmic side exhibited relatively low conservation (Supplementary Fig. 7). Meanwhile, in the outward-state structure, the N- and C-lobes were open toward the periplasm.
a Structure of AmpG in the inward state: top view (left) and side view (right). b The contact surface of the inward state between the N-lobe and C-lobe on the periplasmic side. c Structure of AmpG in the outward state: top view (left) and side view (right). d The contact surface of the outward state between the N-lobe and C-lobe on the cytoplasmic side. e MIC (minimum inhibitory concentration) test results for the wild type and mutations on the contact surface. blue: resistance, yellow: intermediate, red: susceptible. f TSA (thermal shift assay) results for the wild type and mutants with GlcNAc-1,6-anhMurNAc Data are shown as mean ± SD from n = 3 independent experiments.
At the cytoplasmic contact surface, the conserved motif A32,33 residues (Supplementary Fig. 7), located between TM2 and TM3, interact with the terminal region of TM11. In this region, Asp70, Arg79, and Arg80 from the N-lobe, along with Asp134, form an interaction network with Ser348 and Ser350 from the C-lobe. Arg79, in particular, forms weak hydrogen bonds with Arg80 and Ser350 at distances of 3.2 and 3.6 Å, respectively, and is also suggested to form a potential salt bridge with Asp134 at a distance of 4.1 Å. When residues such as Asp70, Arg79, and Arg80 were mutated to alanine, a decrease in resistance to β-lactam antibiotics was observed. Additionally, mutations of Ser348 or Ser350 to alanine in the C-lobe resulted in only a slight decrease in antibiotic resistance. However, when these mutations were made simultaneously, a significant decrease in resistance was observed (Fig. 2e).
TSA results for the D70A, R79A, and R80A mutants showed a concentration-dependent increase in the thermal stability of GlcNAc-1,6-anhMurNAc. motif A maintains the outward state of the MFS proteins and is therefore unrelated to substrate binding event33. However, the binding affinity of the mutants, as measured using MST, was approximately six times lower than that of the wild-type (WT) or outward mutants (Supplementary Fig. 2a–f). This difference in binding affinity was also consistent with the dissociation constant (kd_tm)34 calculated using the TSA data (Supplementary Table 1). This difference in binding affinity may result from a change in the outward-inward state ratio of AmpG caused by the mutation. In the MFS protein, which transports substances from the outside to the inside of the cell, it has been observed that the outward state has a higher affinity for the substrate than the inward state30,35. In the cryo-EM structure of AmpG in the inward state, the interaction between Arg79 of motif A and Glu4 of BRIL (Supplementary Fig. 1d) provided a clear explanation for the relatively low binding affinity of AmpG(cryo) compared with that of the WT. This interaction might disrupt the function of motif A, favoring the inward state and consequently affecting the binding affinity.
Substrate-binding pocket of AmpG
To elucidate the complex structure of AmpG with its substrates GlcNAc-1,6-anhMurNAc, we prepared grids by adding GlcNAc-1,6-anhMurNAc to the samples used to determine the inward and outward apo structures. In the data collected from both samples, we observed electron microscopy (EM) density maps of GlcNAc-1,6-anhMurNAc between the N- and C-lobes (Supplementary Figs. 5e, f, 6e, and f). The structure of inward-state AmpG complexed with GlcNAc-1,6-anhMurNAc exhibited more occlusion on the cytoplasmic side than on the apo-inward side (Figs. 1b and 3a). In this state, AmpG primarily interacts with GlcNAc-1,6-anhMurNAc via residues in the N-lobe. Specifically, Lys62 forms a hydrogen bond with the O3 of the glucosamine moiety at a distance of 2.75 Å, whereas Asp125 also forms a hydrogen bond with this O3 at a distance of 2.73 Å. Tyr152 and Arg153 interact with the anhydrous muramic acid moiety. Specifically, Arg153 formed hydrogen bonds with the anhydro linkage (OA6) at distances of 2.76 Å, whereas Tyr152 created a hydrophobic pocket (Fig. 3b).
a Surface of the inward-occluded state AmpG and the substrate GlcNAc-1,6-anhMurNAc. b Substrate binding mode of the inward-occluded state AmpG. c Surface of the outward state AmpG and the substrate GlcNAc-1,6-anhMurNAc. d Substrate binding mode of the outward state AmpG. e Minimum inhibitory concentration (MIC) results for mutants of residues interacting with the substrate. blue: resistance, yellow: intermediate, red: susceptible. f Thermal shift assay (TSA) results for mutations of binding pocket residues with GlcNAc-1,6-anhMurNAc. Data are shown as mean ± SD from n = 3 independent experiments.
In contrast, in the outward state complexed with GlcNAc-1,6-anhMurNAc, the interactions remain mostly consistent with those in the inward-occluded state, although slight differences are observed (Fig. 3c). Asp125 now interacts with the O4 of the glucosamine moiety at a distance of 2.94 Å, instead of O3. Additionally, Arg365 was observed to form a hydrogen bond with the acetyl group of the anhydrous muramic acid moiety at a distance of 2.95 Å. Despite these differences, the interactions of Lys62 and Arg153 were largely maintained, with Lys62 binding to the glucosamine moiety at a distance of 3.12 Å and Arg153 forming hydrogen bonds with the anhydrous muramic acid moiety at 3.16 Å (Fig. 3d).
The residues observed in the structures were mutated to alanine and an MIC test was conducted to confirm their roles. The results showed that mutations K62A, D125A, R153A, and R365A led to a loss of antibiotic resistance to both cefotaxime and ceftazidime (Fig. 3e). The mutated proteins were purified and subjected to TSA to confirm their direct involvement in substrate binding. The K62A, D125A, D125N, R153A, and R365A mutants showed only a slight increase in thermal stability at varying substrate concentrations compared to the WT (Fig. 3f). Moreover, MST experiments indicated that these mutations did not yield a significant curve, suggesting that the key residues identified in the structure are crucial for substrate binding (Supplementary Fig. 2g–k). In sugar-transporting MFS, hydrophobic residues in the substrate-binding pocket are common36,37,38,39. Mutations of Tyr152 to different residues showed no significant change or only minimal effects on antibiotic resistance in MIC tests (Fig. 3e and Supplementary Data 1). However, TSA and MST assays conducted on the Y152A mutation indicated a significant decrease in substrate binding affinity (Fig. 3f and Supplementary Fig. 2j). These findings indicate that Tyr152 is critical for maintaining high binding affinity with GlcNAc-1,6-anhMurNAc. Despite being mutated to alanine, the protein retained its ability to transport muropeptides and induce AmpC, thereby conferring antibiotic resistance. A similar observation has been reported for the lactose transporter LacY40, where the mutation of Trp151 in the substrate-binding pocket reduced substrate binding affinity, yet the catalytic activity remained intact. This suggests that while substrate binding affinity may decrease with certain mutations, the overall transport mechanism can still function effectively.
Proton binding site of AmpG
AmpG requires a proton gradient to actively transport substrates into the cells13; therefore, protonation is essential. The acidic residues in the transmembrane regions of this protein are Asp125, Asp129, Asp238, and Glu326, all of which are highly conserved (Supplementary Fig. 7). Mutations D125A, D238A, and E326A resulted in decreased antibiotic resistance in MIC tests, whereas the D238E and E326D mutations retained antibiotic resistance (Figs. 3e and 4a). Although Asp125 is crucial for substrate binding, mutations D238A and E326A did not affect the binding to GlcNAc-1,6-anhMurNAc (Fig. 4b and Supplementary Fig. 8a–c). Interestingly, the E326A mutant maintained its binding affinity for the substrate, even at high pH, in contrast to the wild-type protein (Supplementary Fig. 8d–h). These findings are similar to those observed with the LacY41, where experiments involving a mutation of Glu325, the final proton acceptor42, also indicated that binding affinity was preserved at high pH43. Protonation of this residue may influence the dissociation constant of the LacY protein44.
a Minimum inhibitory concentration (MIC) results for mutants of acidic residues located in a transmembrane helix. blue: resistance, yellow: intermediate, red: susceptible. b Thermal shift assay (TSA) results for mutants of acidic residues. Data are shown as mean ± SD from n = 3 independent experiments. c The final snapshot at 500 ns in the Asp125 and Asp238 protonated condition of the outward complex state (replica 1). d The distance change between Asp125, Arg153, and GlcNAc-1,6-anhMurNAc in the Asp125 and Asp238 protonated condition. e The snapshot at 0 ns in the deprotonated condition of the outward complex state (replica 1). f The distance change between Asp125, Arg153, and GlcNAc-1,6-anhMurNAc in the deprotonated condition. g the final snapshots at 500 ns in the deprotonated condition of the outward complex state. MD trajectories in panels d and f are shown with smoothed bold lines to emphasize the overall trend.
MD simulations were used to analyze the movement of acidic residues in AmpG, focusing on their behavior under different protonation states (Supplementary Figs. 9 and 10). Protonation of Asp125 and Asp238 influenced their interactions with Lys62 and Arg365, key residues involved in substrate binding. Protonation of Asp125 increased its average distance from Lys62, while protonation of Asp238 increased its distance from Arg365 in both the outward state and inward-occluded state simulations. The salt bridge between Asp238 and Arg365 is not observed in the cryo-EM structures. Specifically, for Arg365 to interact with Asp238, it must be oriented away from the substrate-binding pocket (Supplementary Figs. 9b, d, 10b, d), positioning the Asp238-Arg365 interaction as a crucial link between the substrate-binding site and the proton-binding site.
In simulations using the outward GlcNAc-1,6-anhMurNAc complex structure, protonation of both Asp125 and Asp238 stabilized substrate binding by maintaining key interactions within the binding pocket (Fig. 4c, d). In contrast, under the deprotonated condition, AmpG failed to stabilize substrate binding, resulting in substrate displacement across all three replicas (Fig. 4e, f). This instability led to different modes of substrate displacement: flipping within the binding pocket (replica 1), dissociation from the protein (replica 2), or drifting away from the binding pocket (replica 3) (Fig. 4g).
Protonation of Asp125 and Asp238 not only affected the flexibility of Lys62 and Arg365, which are crucial for substrate binding, but also helped maintain the overall charge balance within the AmpG binding pocket. Such charge-dependent substrate binding regulation via protonation of acidic residues has also been observed in other MFS transporters45,46, supporting the idea that protonation plays a key role in stabilizing substrate interactions. As previously mentioned, AmpG exhibits high substrate selectivity based on charge17, while simultaneously requiring charge neutrality within the binding pocket for stable substrate binding. When Asp125 and Asp238 were not protonated, disruptions in charge balance destabilized substrate binding, causing substrate destabilization and conformational shifts. These results suggest that substrate binding in AmpG is not solely dictated by physical interactions but is also highly dependent on the electrostatic environment within the binding pocket.
Meanwhile, simulations using the inward occluded complex structure with GlcNAc-1,6-anhMurNAc exhibited structural differences depending on protonation states. Under the Asp125 and Asp238 protonated condition, the inward occluded state was maintained throughout the 500 ns simulation, with the N-lobe and C-lobe remaining tightly closed (Fig. 5a, b). In replica 1, the Cα distance between Ala146 and Gly280 remained at 7.0 Å, indicating a narrow protein pore. Notably, in replica 2, Ala146 and Gly280 came into direct contact, ultimately reversing the transition toward the outward state (Fig. 5c). In contrast, in simulations where Glu326 was protonated, the N-lobe and C-lobe opened toward the cytoplasm, resembling an inward state. In replica 1, the distance between Ala146 and Gly280 increased to 12.0 Å, and the cytoplasmic pore also expanded (Fig. 5d, e). A comparison of pore radius in the final snapshots further supported these observations. Replica 1 under the Asp125 and Asp238 protonated condition closely resembled the inward occluded state, whereas replica 2 showed notable differences on the periplasmic side. Similarly, Glu326 protonation resulted in a pore structure highly similar to the inward state (Fig. 5e). Structural transition toward the inward state was also observed under deprotonated conditions (Supplementary Fig. 10e).
a The final snapshot at 500 ns in the Asp125 and Asp238 protonated condition of the inward-occluded state (replica 1). b The distance change between Ala146 and Gly280 in the Asp125 and Asp238 protonated condition. c Inward-occluded state to outward state transition (replica 3), left: snapshot at 0 ns, right: snapshot at 50 ns in the Asp125 and Asp238 protonated condition. d The final snapshot at 500 ns in the Glu326 protonated condition of the inward-occluded state (replica 1). e The distance change between Ala146 and Gly280 in the Glu326 protonated condition. f A graph analyzing the pores of the inward and inward-occluded state structures and the pores from the final stages of the Asp125 and Asp238 protonated condition and the Glu326 protonated condition, displaying the pore size along the vertical z-axis (Å) of the protein from the center of the membrane. MD trajectories in (b) and (e) are shown with smoothed bold lines to emphasize the overall trend.
These findings demonstrate that protonation is essential for both substrate binding and structural transitions in AmpG. In the outward state, Asp125 and Asp238 protonation stabilized substrate binding. In the inward occluded state, their protonation maintained the occluded conformation, whereas Glu326 protonation promoted the inward transition by facilitating the opening of the cytoplasmic pore.
Muropeptide transport cycle of AmpG
We propose a muropeptide transport mechanism for AmpG based on the results of the structural analysis of AmpG combined with biological and biochemical experiments and MD simulations (Fig. 6). The substrate transport mechanism of AmpG involves the following processes: (i) In the outward state, AmpG is stabilized by interactions between motif A of the N-lobe and the end of TM 11. This configuration complicates substrate binding because of the formation of salt bridges between Lys62 and Asp125 and between Asp238 and Arg365. (ii) Protonation of Asp125 and Asp238 releases Lys62 and Arg365, respectively, which are key for substrate binding and enhanced helical flexibility while also contributing to charge neutralization within the binding pocket. (iii) Binding of the substrate to the protonated outward state of AmpG triggers a conformational change to the occluded state. (iv) In the occluded state, Asp125 and Asp238 were deprotonated, initiating the transition toward the inward state. This deprotonation likely requires proton transfer to Glu326, which acts as a proton acceptor. (v) As the inward transition progresses, Asp125 and Asp238 tend to reform salt bridges with Lys62 and Arg365, destabilizing substrate binding and promoting its release. (vi) Following disassociation of the substrate and deprotonation, AmpG returns to the outward state (i) and is ready to initiate another transport cycle.
The alternating access model of the anhydromuropeptide permease AmpG. AmpG utilizes proton concentration gradients to transport substrates and specifically recognizes the sugar portion of the peptidoglycan (PG) block. Key residues critical to the alternating access mechanism are highlighted, with protonated states shown in red (H). Yellow double-circle shapes represent the substrate, and salt bridges between residues are depicted with black dotted lines.
Discussion
In this study, we report the structure of Yokenella regensburgei AmpG in four distinct states, which likely represent sequential steps in the transport cycle. Structural analysis enabled the identification of key residues in AmpG that facilitate alternating access transport. The motif A residue, located between TM2 and TM3, is crucial for stabilizing the outward state (Fig. 2d). The residues Lys62, Asp125, Arg153, and Arg365, which were observed in the substrate complex structure, play critical roles in substrate recognition (Fig. 3b, d). MD simulations demonstrated the potential of Lys62 and Arg365 to form salt bridges with Asp125 and Asp238 (Supplementary Figs. 9 and 10). These salt bridges restrict the flexibility of key residues and helices, making substrate binding less favorable. However, protonation weakens these interactions and neutralizes charges within the binding pocket, stabilizing a conformation more conducive to substrate binding. Specifically, in the outward-state MD simulations, protonation of both Asp125 and Asp238 was essential for stable substrate binding (Fig. 4c–g). In contrast, in the inward-occluded state, transition to the inward state did not occur until these residues were deprotonated (Fig. 5a–e).
Mutation of Asp238 and Glu326 to alanine led to a significant loss of resistance to several β-lactam antibiotics, including cefotaxime, ceftazidime, aztreonam, and ertapenem. However, the D238E mutant restored antibiotic resistance to wild-type levels across all tested antibiotics. Similarly, the E326D mutant also showed substantial recovery of resistance, particularly against ceftazidime and aztreonam (Fig. 4a). In contrast, mutation of Asp125 to glutamate resulted in a loss of β-lactam antibiotic resistance (Supplementary Data 1). Additionally, TSA revealed that the Asp125 to asparagine mutation caused subtle shifts in the thermal stability of GlcNAc-1,6-anhMurNAc, whereas MIC tests showed that the D125N mutant retained partial antibiotic resistance (Fig. 3e, f). These findings suggest that Asp125 protonation plays a dual role, being essential for direct substrate interactions while also influencing conformational transitions in the transport cycle. MD simulations further support this idea, demonstrating that the protonation state of Asp125, particularly in the outward conformation is crucial for stable substrate binding. PROPKA3.147,48 predictions corroborate this, indicating that the pKa of Asp125 is significantly higher in the outward state than in the inward states (Supplementary Fig. 11). Specifically, the pKa was 6.84 in the apo outward state and 7.97 in the substrate-bound outward state, whereas in the inward states, it dropped to 5.66 in the inward-occluded state and 3.80 in the apo inward state. This shift suggests that Asp125 remains protonated under physiological conditions in the outward state, promoting stable substrate binding, but readily deprotonates to facilitate the conformational transition. Interestingly, TM4, where Asp125 is located, also contains two highly conserved acidic residues, Asp129 and Asp134 (Supplementary Fig. 7). While mutations at Asp129 had minimal impact on antibiotic resistance, mutations at Asp134, which interacts with motif A in the outward state, caused a slight reduction in resistance (Supplementary Data 1). These findings suggest that Asp125 not only interacts with the substrate but also plays a regulatory role in transporter dynamics through the rocker-switch mechanism, contributing to the alternating access model of AmpG. Based on this, its role in the transport cycle can be interpreted in two possible ways. The first possibility is that Asp125 functions similarly to the EXXEK motif in POT transporters49, serving as an additional protonation site beyond the extracellular region and influencing substrate transport. In this model, Asp125 actively participates in proton-coupled conformational changes, acting as a relay in the transport cycle. Alternatively, Asp125 may remain protonated for most of the transport cycle, similar to E206 in the XylE transporter, as suggested by hydrogen-deuterium exchange mass spectrometry (HDX-MS)50. In this scenario, its protonation state may influence the energetic landscape of the alternating access mechanism of AmpG, which could affect transitions between outward and inward states. Further biochemical and structural studies are required to elucidate how Asp125 protonation contributes to the alternating access model of AmpG.
AmpG homologs generally consist of 12 transmembrane (TM) helices, but structural variations exist among different species. For instance, Yokenella regensburgei, Escherichia coli, and Klebsiella aerogenes possess two additional helices (6 + 6 + 2), while Pseudomonas aeruginosa has a different configuration (6 + 4 + 6). In contrast, Vibrio cholerae and Tannerella forsythia lack these extra helices (Supplementary Fig. 12a). Despite these structural differences, the precise function of the extra helices remains unclear. Experimental deletion of the extra helix in Yokenella regensburgei AmpG resulted in a significant loss of antibiotic resistance, as shown by MIC testing (Supplementary Data 1). However, protein purification of the truncated variant was also challenging, suggesting that the extra helix may contribute to protein folding stability. Given that the extra helix contains numerous hydrophobic residues, exhibits minimal interaction with the transmembrane domain, and its removal hinders protein purification, it is likely involved in protein folding, membrane targeting, or stability. However, its potential role in substrate specificity cannot be ruled out. Interestingly, the absence of extra helices in Vibrio cholerae and Tannerella forsythia correlates with their ability to transport noncanonical51 or non-anhydro muropeptides14, suggesting that extra helices might influence the substrate selectivity of AmpG. Further studies are needed to elucidate the specific function of the extra helix. Nevertheless, key residues in both the substrate-binding and proton-binding pockets of AmpG are highly conserved across homologs, indicating that while extra helices may contribute to structural and functional diversity, the core transport mechanism remains well preserved (Supplementary Fig. 12b). To further examine the functional role of these conserved residues, MIC tests were conducted using the Klebsiella aerogenes strain (KE-Y6). The results showed that AmpG from Klebsiella aerogenes and Escherichia coli, which share an extra-helix structure with Yokenella regensburgei, successfully restored antibiotic resistance to KE-Y6. In contrast, mutation of key residues led to the loss of resistance, supporting the importance of these residues in AmpG function. Furthermore, AmpG homologs from Vibrio cholerae, Tannerella forsythia, and Pseudomonas aeruginosa, which have different extra-helix structures, failed to restore antibiotic resistance even in their wild-type forms (Supplementary Fig. 12c). These findings suggest that AmpG homologs from different species may not function efficiently within the membrane environment of Klebsiella aerogenes, potentially due to differences in membrane composition or structural requirements.
Sverak et al. recently reported the Escherichia coli AmpG structure (PDB: 9C3F)17. While the fusion location of the fiducial marker (BRIL) and the detergent used for solubilization differed between the two studies, the overall structures remained highly similar (outward Apo RMSD = 1.13 Å, outward complex RMSD = 1.58 Å). A structural comparison between the Escherichia coli and Yokenella regensburgei AmpG outward states revealed a highly conserved transmembrane arrangement (Supplementary Fig. 13a, b), as well as similar substrate-binding residues and protonation sites (Supplementary Fig. 13d). Despite the BRIL fusion in ICL3, the motif A residue network, which stabilizes the outward state, remained well conserved (Supplementary Fig. 13c). Several MFS transporters that mediate substrate uptake favor the outward state, stabilized by motif A52. This trend is consistent with the Escherichia coli AmpG structure, which was resolved in an outward conformation. Similarly, AlphaFold3 predictions also favor the outward state. Notably, AlphaFold3 predicted that mutating Asp70 of motif A to alanine shifts the conformation to inward, suggesting its role in outward-state stabilization17. A similar trend was observed in Yokenella regensburgei AmpG. In Yokenella regensburgei AmpG(cryo), where BRIL was fused to ICL3, interactions between Glu4 of BRIL and Arg79 of motif A biased the equilibrium toward inward-state particles both in vitro and on the cryo-electron microscopy grid (Supplementary Figs. 4 and 6). In contrast, the Escherichia coli AmpG structure, where BRIL was fused to the N-terminus, was resolved in the outward state without requiring mutations. This difference is likely due to the lack of BRIL-induced interference in motif A and the role of the detergent used, specifically n-dodecyl-β-D-maltoside (DDM). In the Yokenella regensburgei AmpG-GlcNAc-1,6-anhMurNAc complex structure, the substrate is positioned horizontally between the N-lobe and C-lobe. However, in the Escherichia coli AmpG structure, a single DDM molecule was observed in a vertical orientation within the substrate-binding pocket (Supplementary Fig. 13b). In this conformation, the maltose head group of DDM interacts with Lys62 and Asp125 in the substrate-binding site, while the dodecyl tail extends toward the extra-helical region between the N-lobe and C-lobe (Supplementary Fig. 13d). This hydrophobic interaction likely contributes to the stabilization of the outward state. For our structural determination, we used lauryl maltose neopentyl glycol (LMNG) instead of DDM, which may explain the absence of a similar interaction in our dataset.
Inhibiting AmpG to increase the susceptibility of antibiotic-resistant bacteria to β-lactam antibiotics could be an effective strategy to combat broad-spectrum β-lactam resistance. In our study, the Klebsiella aerogenes (KE-Y6) strain25 used in the MIC tests is an AmpG knockout derivative of Klebsiella aerogenes (KE-Y1), which originally exhibited broad-spectrum resistance. This strain was resistant not only to third-generation cephalosporins, such as cefotaxime and ceftazidime, but also to carbapenems, including ertapenem and imipenem. However, upon AmpG knockout, the strain lost its antibiotic resistance. Reintroducing AmpG from Yokenella regensburgei via a vector fully restored resistance (Supplementary Fig. 1d), whereas mutations in key AmpG residues abolished resistance (Supplementary Data 1). AmpG plays a critical role in peptidoglycan (PG) recycling, a function also facilitated by other peptide permeases53. Interestingly, knocking out AmpG had minimal impact on cell growth rate but significantly increased sensitivity to β-lactam antibiotics. These findings suggest that targeting AmpG imposes minimal selective pressure on microorganisms while restoring sensitivity to broad-spectrum β-lactam antibiotics. Therefore, AmpG inhibitors represent promising candidates for antibiotic adjuvants.
In conclusion, the proposed muropeptide transport mechanism and binding pocket of AmpG provide a blueprint for the development of antibiotic adjuvants targeting AmpG through structure-based drug discovery (SBDD). These adjuvants may contribute to addressing the challenge of antimicrobial resistance (AMR).
Methods
Construction of AmpG (cryo) and outward mutant (cryo-ww) and protein expression
The AmpG from Yokenella regensburgei (UniProtKB: G9Z488) was cloned into a modified pET28a vector with an N-terminal 6× His tag and a TEV protease cleavage site, referred to as AmpG(WT). For structural studies, AmpG(cryo) was created by inserting BRIL (thermostabilized apocytochrome b562 from Escherichia coli) between residues Ser199 and Val203 of AmpG(WT). To generate the outward-open mutant AmpG(ww), G50W and L269W mutations were introduced into AmpG(WT) using site-directed mutagenesis. The AmpG(cryo-ww) construct was derived from AmpG(ww) by inserting BRIL at the same Ser199-Val203 site as in AmpG(cryo). Additionally, Glu4 of BRIL was mutated to alanine. All DNA sequences were synthesized commercially. All AmpG constructs were expressed in C41(DE3) cells (Sigma). Cells were cultivated in LB Broth (Cellconic) in a shaking incubator at 150 rpm and 37 °C. The culture was grown until reaching an OD600 of ~0.6, typically requiring 4–5 h. The temperature was then reduced to 17 °C, and protein expression was induced with 0.1 mM IPTG overnight. The cells were harvested by centrifugation, and the resulting pellets were stored at −80 °C for further use.
Protein purification for thermal shift assay and microscale thermophoresis assay
Wild type and mutants AmpG expressing cells were disrupted via sonication and ultracentrifuge with lysis buffer (20 mM HEPES, 300 mM NaCl; pH 7.0) containing protease inhibitors (once; 500 μM AEBSF, 1 μM E-64, 1 μM leupeptin, and 150 nM aprotinin). The purified membranes were resuspended in lysis buffer containing protease inhibitors. Additionally, to solubilize, 1% (w/v) n-dodecyl-β-D-maltopyranoside (DDM)(Anatrace), and 0.2% (w/v) cholesteryl hemisuccinate (CHS)(Anatrace) were added for 2 h at 4 °C. Insoluble materials were removed via centrifugation at 150,000 × g for 30 min, followed by incubation with TALON IMAC resin (Clontech) overnight at 4 °C. Next, the resin was washed with 20 column volumes (CVs) of wash buffer (20 mM HEPES (pH 7.0), 300 mM NaCl, 0.05% (w/v) DDM, and 0.01% (w/v) CHS). The proteins were eluted in 5–6 CVs elution buffer (20 mM HEPES (pH 7.0), 300 mM NaCl, 0.05% (w/v) DDM, 0.01% (w/v) CHS, and 300 mM imidazole). The eluted AmpG was desalted using a desalting column (Cytiva) in a buffer containing 20 mM HEPES (pH 7.0), 300 mM NaCl, 0.05% (w/v) DDM, and 0.01% (w/v) CHS to remove imidazole. Subsequently, TEV protease was added and incubated for 16 h. To remove the 6x His tag and impurity proteins, talon beads were allowed to bind to the sample at 4 °C for 2 h and flow-through was collected. Subsequently, the protein was concentrated using a 100 kDa cutoff centricon (Vivaspin). Gel filtration was performed in a buffer containing 20 mM HEPES (pH 7.0), 300 mM NaCl, 0.05% (w/v) DDM, and 0.01% (w/v) CHS. Finally, the purified protein was used for TSA and MST assays.
AmpG(cryo) or AmpG(cryo-ww) – BAG2 – nanobody complex formation for cryo-EM grid
The BAG2 and nanobody clones were obtained from Mukherjee. The proteins were purified using a previously published method31. AmpG(cryo) and AmpG(cryo-ww) expressing cells were disrupted by sonication and ultracentrifugation in lysis buffer (20 mM HEPES, 300 mM NaCl, pH 7.0) containing protease inhibitors. The purified membranes were resuspended in lysis buffer containing protease inhibitors. Additionally, to solubilize 0.5% (w/v) lauryl maltose neopentyl glycol (LMNG)(anatrace) and 0.05% (w/v) CHS were added for 2 h at 4 °C. Insoluble materials were removed via centrifugation at 150,000 × g for 30 min, followed by incubation with TALON IMAC resin overnight at 4 °C. Next, the resin was washed with 20 column volume (CVs) of wash buffer (20 mM HEPES (pH 7.0), 300 mM NaCl, 0.02% (w/v) LMNG and 0.002% (w/v) CHS). The proteins were eluted in 5–6 CVs elution buffer (20 mM HEPES (pH 7.0), 300 mM NaCl, 0.02% (w/v) LMNG, 0.002% (w/v) CHS, and 300 mM imidazole). The eluted AmpG was desalted using a desalting column in a buffer containing 20 mM HEPES (pH 7.0), 300 mM NaCl, 0.02% (w/v) LMNG, and 0.002% (w/v) CHS to remove imidazole. Subsequently, TEV protease was added and incubated for 16 h. To remove the 6x His tag and impurity proteins, talon beads were allowed to bind to the sample at 4 °C for 2 h and flow-through was collected. To make the complex, AmpG(cryo) or AmpG(cryo-ww), BAG2, and nanobody were mixed at a molar ratio of 1:2:4, followed by a 30 min incubation at 4 °C. Finally, gel filtration was performed using a buffer containing 20 mM HEPES (pH 7.0), 300 mM NaCl, 0.000075% (w/v) LMNG, 0.000025% (w/v) GDN (Anatrace), and 0.000001% (w/v) CHS.
Grid preparation for cryo-EM and data collection
The AmpG(cryo) complex was concentrated to approximately 8 mg/ml, and two types of self-wicking grids (SPT Labtech) were prepared using the Chameleon system (SPT Labtech) at IMP (Korea). These grids featured an inward-state structure and an inward-occluded state, with the latter generated by the addition of 10 mM GlcNAc-1,6-anhMurNAc. GlcNAc-1,6-anhMurNAc was synthesized as previously described54 and was provided by LigaChem Biosciences (Korea). For the AmpG(cryo-ww) complex, which was concentrated to approximately 11 mg/mL, two additional grids were prepared: one for the outward structure and the other for the outward structure with 10 mM GlcNAc-1,6-anhMurNAc. The grids were prepared using a Vitrobot Mark IV device (Thermo Scientific) at 15 mA for 60 s on glow-discharged Quan Mfoil R1.2/1.3 copper grids (mesh 300). Subsequently, frozen grids were screened on a Glacios microscope operating at 200 kV (Thermo Fisher Scientific), followed by data acquisition on a Titan Krios microscope (Thermo Fisher Scientific) equipped with either a Falcon4i (Thermo Fisher Scientific, BaobabAIBio) or Bioquantum K3 (Gatan, IBS). Two AmpG(cryo) grids, four AmpG(cryo) GlcNAc-1,6-anhMurNAc grids, one AmpG(cryo-ww) grid, and two AmpG(cryo-ww) GlcNAc-1,6-anhMurNAc grids were selected and automatically collected using EPU software (Thermo Fisher Scientific) in the electron counting mode and correlated double sampling mode (Gatan). Supplementary Table 2 provides detailed information on each dataset.
Cryo-EM image processing
Cryo-EM image processing was performed using the CryoSPARC v3.152. The initial image-processing approach was consistent for both datasets. Comprehensive details of image processing for the individual datasets are described in Supplementary Figs. 3–6. The micrograph movies were corrected for beam-induced motion, followed by CTF estimation. Particle picking commenced with a blob picker and was subjected to three rounds of 2D classification, followed by 3D classification using ab initio. The selected particles were used as inputs for the neural network particle picker embedded in Topaz55. These topaz-picked particles were classified by reference-free 2D classification and re-extracted with finer sampling for multi-reference-guided 3D classification56. A poor reference map from the initial 3D classification was used as a decoy to remove the junk particles. The selected particles were then subjected to additional 3D-classified and non-uniform refinement. Focused refinement was implemented for 14 TM domains (apo, 1-14) and 12 TM domains (sub, 1–12) to enhance the molecular features of the transmembrane domain during refinement. Finally, a deep learning-based automatic post-processing suite, DeepEMhancer, was applied to both datasets to improve the molecular features of the transmembrane domain57. The initial model of AmpG was generated using AlphaFold258, and the BAG2–nanobody complex was modeled with reference to the cryo-EM structure of CCR6-BAG2-nanobody(PDB: 9D3E)59.
Minimum inhibitory concentration (MIC) test
The pADY123 vector, containing either wild-type or mutant AmpG, was transformed into the Klebsiella aerogenes strain (KE-Y6) with an AmpG knockout25. Colonies grown on Mueller-Hinton agar plates were tested for antimicrobial susceptibility using the Vitek2 system (bioMérieux, Marcy l’Etoile, France). All procedures followed the manufacturer’s instructions. Minimum inhibitory concentrations were classified as susceptible, intermediate, or resistant, according to CLSI interpretive criteria60. MIC breakpoints (µg/mL) were defined as follows: Beta-lactams (Ampicillin ≤8/16/ ≥ 32, Amoxicillin+Clavulanic acid ≤8 + 4/16 + 8/ ≥ 32 + 16, Piperacillin+Tazobactam ≤8 + 4/16 + 4/ ≥ 32 + 4); Cephalosporins (Cefazolin ≤2/4/ ≥ 8, Cefoxitin ≤8/16/ ≥ 32, Cefotaxime ≤1/2/ ≥ 4, Ceftazidime ≤4/8/ ≥ 16, Cefepime ≤2/4–8/ ≥ 16, Aztreonam ≤4/8/ ≥ 16); Carbapenems (Ertapenem ≤0.5/1/ ≥ 2, Imipenem ≤1/2/ ≥ 4); Aminoglycosides (Amikacin ≤4/8/ ≥ 16, Gentamicin ≤2/4/ ≥ 8); Fluoroquinolone (Ciprofloxacin ≤0.25/0.5/ ≥ 1); and Others (Tigecycline 2/4/8, Trimethoprim+Sulfamethoxazole 2/38/—/4/76).
The Ceftazidime growth curve experiment was performed as follows. First, a 2X dilution of ceftazidime was prepared in MHB, and 50 µL was aliquoted into each well. The Klebsiella aerogenes strain (KE-Y6) was diluted in distilled water to achieve 108 CFU/mL (200X dilution) using McF standard 0.5, and further diluted to 106 CFU/mL (2X dilution) in MHB. Then, 50 µL of the diluted bacteria was added to each well. The plates were incubated at 37 °C for 16–20 h. OD at 600 nm for each well was measured using a Varioskan LUX, and the growth curves were plotted.
Thermal shift assay (TSA)
Purified AmpG proteins, both wild-type and mutants (3–5 µM), were prepared by mixing them in a 1:1 ratio with various GlcNAc-1,6-anhMurNAc concentrations. The resulting samples were loaded onto TY-C001 capillaries (NanoTemper Technologies) for the measurements. The measurements were performed using a Tycho NT.6 (NanoTemper Technologies) at room temperature. The Ti value changes of wild-type and mutant proteins, dependent on various concentrations of GlcNAc-1,6-anhMurNAc, were analyzed using the kd_tm program34.
Microscale thermophoresis assay (MST)
Purified WT or mutant AmpG protein was diluted to 200 nM using a dilution buffer composed of 20 mM HEPES (pH 7.0), 300 mM NaCl, and 0.05% (w/v) DDM. The proteins were labeled using the His-Tag Labeling Kit RED-tris-NTA 2nd Generation (NanoTemper Technologies) following the manufacturer’s protocol. GlcNAc-1,6-anhMurNAc prepared in the same buffer was serially diluted from 1000 µM to 7.81 µM. The diluted samples were mixed in a 1:1 ratio with the stained protein and loaded onto MO-Z022 capillaries (NanoTemper Technologies) for analysis. The MST analysis was conducted using a Monolith NT115 instrument (NanoTemper Technologies) at room temperature. The settings for the analysis were adjusted to 100% LED power and 40% MST power. All measurements were performed at least three times to ensure consistency and accuracy. The collected MST data were analyzed using MO Affinity Analysis Software (NanoTemper Technologies).
Isothermal titration calorimetry (ITC) assay
ITC measurements were performed using a MicroCal Auto-iTC200 instrument (GE Healthcare, Little Chalfont, UK). WT AmpG was prepared in a buffer containing 20 mM HEPES (pH 7.0), 300 mM NaCl, and 0.05%(w/v) DDM. GlcNAc-1,6-anhMurNAc was dissolved in the same buffer. The protein concentration in the measurement cell was 55 µM, whereas the ligand concentration in the titration procedure was 2 mM. Titrations were performed at 25 °C, and data were fitted to the one-binding-site model using the software Origin 7.0 (MicroCal).
Molecular dynamics simulations
Molecular dynamics (MD) simulations were conducted for AmpG from Yokenella regensburgei. The structural models used in this study were the outward-state complexed with GlcNAc-1,6-anhMurNAc (PDB: 9J9Z) and the inward-occluded state complexed with GlcNAc-1,6-anhMurNAc (PDB: 8ZKE). Missing residues at the N- and C-termini were modeled, and the BRIL fusion region was restored to the Yokenella regensburgei AmpG sequence. The N- and C-termini were capped with acetyl and N-methyl groups, respectively. To build the system, the protein-ligand complex was embedded in a 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphoethanolamine (POPE) bilayer within a 105 × 105 × 104 ų simulation box using CHARMM-GUI61. The system was then solvated using the TIP3P water model, and 150 mM NaCl was added to neutralize the system. Details of the system composition are provided in Supplementary Table 3. MD simulations were performed using GROMACS 2024.462 with the CHARMM3663,64,65 force field applied to proteins, lipids, and ions. The ligand topology was generated using the CHARMM General Force Field (CGenFF)66. After energy minimization using the steepest descent algorithm, the system was equilibrated in the NVT ensemble for 2 ns, allowing relaxation of the lipid tails, lipid heads, and solvent molecules. Further equilibration was conducted in the NPT ensemble, where positional restraints were progressively released. Initially, restraints were removed from the lipids and solvent, followed by the protein sidechains. This was then extended to the ligand, and finally, the Cα atoms of the protein were released over 2.5 ns per stage. After equilibration, production simulations were carried out for 200–500 ns with a 2 fs time step. Periodic boundary conditions (PBC) were applied, and the Nose-Hoover thermostat67 and Parrinello-Rahman barostat68 maintained the system at 310 K and 1 atm, respectively. Long-range electrostatic interactions were calculated using the particle mesh Ewald (PME)69 method with a 12 Å cutoff, and van der Waals interactions were smoothly switched to zero between 10–12 Å. All bonds involving hydrogen atoms were constrained by the LINCS algorithm70, enabling a 2 fs integration time step. Each system was simulated in triplicate with different initial velocities. The data collected during the molecular dynamics simulation included trajectory and structural data, which were analyzed using VMD71. Protein pore sizes were calculated using HOLE72 and visualized using Moleonline73.
Sequence and structural models of AmpG homologs
The genes for Klebsiella aerogenes (UniProt: A0A0F1TIN5), Escherichia coli (strain K12) (UniProt: P0AE16), Vibrio cholerae (UniProt: A0A0H3Q0D6), Tannerella forsythia (UniProt: A0A1D3UBL0), and Pseudomonas aeruginosa (UniProt: Q9X3S5) were synthesized and cloned into the pADY123 vector for use in MIC tests. Additionally, structural models of the homologs were predicted using AlphaFold258 for further structural analysis.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Cryo-EM maps and structural coordinates were deposited in the Protein Data Bank (PDB) and Electron Microscopy Data Bank (EMDB) under the following accession numbers: 8ZGZ and EMD-60093 (AmpG apo-inward state), 8ZBB and EMD-39900 (AmpG apo-outward state), 8ZKE and EMD-60190 (AmpG inward-occluded state complexed with GlcNAc-1,6-anhMurNAc), 9J9Z and EMD-61285 (AmpG outward state complexed with GlcNAc-1,6-anhMurNAc). In addition, previously published structures 9C3F and 9D3E were referenced and used for comparison. The MD source data are available at Zenodo (https://doi.org/10.5281/zenodo.15628553). Source data are provided as a Source Data file. Source data are provided with this paper.
Change history
07 July 2025
In the Acknowledgements section of this article the grant numbers relating to the Basic Science Research Program through the National Research Foundation of Korea (NRF), funded by the Ministry of Science and ICT were incorrectly given as 'NRF-2021R1A2C3010506, NRF-2017M3A9F6029755, and NRF-2019M3E5D6063903' and should have been 'RS-2024-00351254, RS-2024-00344154, RS-2021-NR056553, NRF-2021R1A2C3010506, and NRF-2019M3E5D6063903'. In addition, the following information was missing from the Acknowledgement section: 'Additional support was provided by the Brain Korea 21 (BK21) FOUR program. C.N., H.K., U.K., Y.C., and Y.Y. are BK21 FOUR fellowship recipients. Use of the cryo-EM facilities of the NEXUS consortium was supported by the NRF (RS-2024-00440289)'. The original article has been corrected.
References
WHO. Antimicrobial resistance. (WHO, 2021).
O’Neill, J. Antimicrobial resistance: tackling a crisis for the health and wealth of nations. (WHO, 2014).
Browne, K. et al. A new era of antibiotics: the clinical potential of antimicrobial peptides. Int. J. Mol. Sci. 21, 7047 (2020).
Hawkins, S. M. & Nephew, K. P. Unintended consequences of antibiotic therapy on the microbiome delivers a gut punch in ovarian cancer. Cancer Res. 82, 4511–4512 (2022).
Petrosillo, N., Capone, A., Di Bella, S. & Taglietti, F. Management of antibiotic resistance in the intensive care unit setting. Exp. Rev. Anti-infective Ther. 8, 289–302 (2010).
Wright, G. D. Antibiotic adjuvants: rescuing antibiotics from resistance. Trends Microbiol. 24, 862–871 (2016).
Kumar, V. et al. Antibiotic adjuvants: synergistic tool to combat multi-drug resistant pathogens. Front. Cell. Infect. Microbiol. 13, 1293633 (2023).
Dhanda, G., Acharya, Y. & Haldar, J. Antibiotic adjuvants: a versatile approach to combat antibiotic resistance. ACS Omega 8, 10757–10783 (2023).
Du, D. et al. Multidrug efflux pumps: structure, function and regulation. Nat. Rev. Microbiol. 16, 523–539 (2018).
Sewell, E. W. C. & Brown, E. D. Taking aim at wall teichoic acid synthesis: new biology and new leads for antibiotics. J. Antibiotics 67, 43–51 (2014).
Simpson, B. W., Gilmore, M. C., McLean, A. B., Cava, F. & Trent, M. S. Escherichia coli utilizes multiple peptidoglycan recycling permeases with distinct strategies of recycling. Proc. Natl Acad. Sci. USA 120, e2308940120 (2023).
Zhang, Y. et al. ampG gene of pseudomonas aeruginosa and its role in β-lactamase expression. Antimicrob. Agents Chemother. 54, 4772–4779 (2010).
Cheng, Q. & Park, J. T. Substrate specificity of the AmpG permease required for recycling of cell wall anhydro-muropeptides. J. Bacteriol. 184, 6434–6436 (2002).
Mayer, V. M. T. et al. Utilization of different MurNAc sources by the oral pathogen Tannerella forsythia and role of the inner membrane transporter AmpG. BMC Microbiol. 20, 352 (2020).
Gilmore, M. C. & Cava, F. Bacterial peptidoglycan recycling. Trends Microbiol. 33, 340–353 (2025).
Perley-Robertson, G. E. et al. A fluorescent transport assay enables studying AmpG permeases involved in peptidoglycan recycling and antibiotic resistance. ACS Chem. Biol. 11, 2626–2635 (2016).
Sverak, H. E. et al. Cryo-EM characterization of the anydromuropeptide permease AmpG central to bacterial fitness and β-lactam antibiotic resistance. Nat. Commun. 15, 9936 (2024).
Chahboune, A., Decaffmeyer, M., Brasseur, R. & Joris, B. Membrane topology of the escherichia coli ampG permease required for recycling of cell wall anhydromuropeptides and AmpC β-Lactamase Induction. Antimicrob. Agents Chemother. 49, 1145–1149 (2005).
Jacoby, G. A. AmpC β-lactamases. Clin. Microbiol. Rev. 22, 161–182 (2009).
Tamma, P. D. et al. A primer on AmpC β-lactamases: necessary knowledge for an increasingly multidrug-resistant world. Clin. Infect. Dis. 69, 1446–1455 (2019).
Mark, B. L., Vocadlo, D. J. & Oliver, A. Providing β-lactams a helping hand: targeting the Ampc β-lactamase induction pathway. Future Microbiol. 6, 1415–1427 (2011).
Sonnabend, M. et al. Identification of drug resistance determinants in a clinical isolate of pseudomonas aeruginosa by high-density transposon mutagenesis. Antimicrob. Agents Chemother. 64, e01771 (2019).
Korfmann, G. & Sanders, C. C. ampG is essential for high-level expression of AmpC beta-lactamase in Enterobacter cloacae. Antimicrob. Agents Chemother. 33, 1946–1951 (1989).
Li, P. et al. Structure-function analysis of the transmembrane protein AmpG from pseudomonas aeruginosa. PLoS One 11, e0168060 (2016).
D’Souza, R. et al. Role of AmpG in the resistance to β-lactam agents, including cephalosporins and carbapenems: candidate for a novel antimicrobial target. Ann. Clin. Microbiol. Antimicrob. 20, 45 (2021).
Dang, Y. et al. Molecular mechanism of substrate recognition by folate transporter SLC19A1. Cell Discov. 8, 141 (2022).
Tsutsumi, N. et al. Structure of human Frizzled5 by fiducial-assisted cryo-EM supports a heterodimeric mechanism of canonical Wnt signaling. Elife 9, e58464 (2020).
Wright, N. J. et al. Methotrexate recognition by the human reduced folate carrier SLC19A1. Nature 609, 1056–1062 (2022).
Kumar, H., Finer-Moore, J. S., Kaback, H. R. & Stroud, R. M. Structure of LacY with an α-substituted galactoside: connecting the binding site to the protonation site. Proc. Natl. Acad. Sci. USA 112, 9004–9009 (2015).
Jiang, X. et al. Engineered XylE as a tool for mechanistic investigation and ligand discovery of the glucose transporters GLUTs. Cell Discov. 5, 14 (2019).
Mukherjee, S. et al. Synthetic antibodies against BRIL as universal fiducial marks for single−particle cryoEM structure determination of membrane proteins. Nat. Commun. 11, 1598 (2020).
Paulsen, I. T., Brown, M. H. & Skurray, R. A. Proton-dependent multidrug efflux systems. Microbiol. Rev. 60, 575–608 (1996).
Jiang, D. et al. Structure of the YajR transporter suggests a transport mechanism based on the conserved motif A. Proc. Natl. Acad. Sci. USA 110, 14664–14669 (2013).
Kotov, V. et al. Plasticity of the binding pocket in peptide transporters underpins promiscuous substrate recognition. Cell Rep. 42, 112831 (2023).
Smirnova, I. et al. Outward-facing conformers of LacY stabilized by nanobodies. Proc. Natl. Acad. Sci. USA 111, 18548–18553 (2014).
Sun, L. et al. Crystal structure of a bacterial homologue of glucose transporters GLUT1–4. Nature 490, 361–366 (2012).
Dang, S. et al. Structure of a fucose transporter in an outward-open conformation. Nature 467, 734–738 (2010).
Nomura, N. et al. Structure and mechanism of the mammalian fructose transporter GLUT5. Nature 526, 397–401 (2015).
Deng, D. et al. Molecular basis of ligand recognition and transport by glucose transporters. Nature 526, 391–396 (2015).
Guan, L., Hu, Y. & Kaback, H. R. Aromatic stacking in the sugar binding site of the lactose permease. Biochemistry 42, 1377–1382 (2003).
Abramson, J. et al. Structure and mechanism of the lactose permease of escherichia coli. Science 301, 610–615 (2003).
Kimanius, D., Lindahl, E. & Andersson, M. Uptake dynamics in the Lactose permease (LacY) membrane protein transporter. Sci. Rep. 8, 14324 (2018).
Smirnova, I., Kasho, V., Sugihara, J., Choe, J.-Y. & Kaback, H. R. Residues in the H+ translocation site define the pka for sugar binding to LacY. Biochemistry 48, 8852–8860 (2009).
Grytsyk, N., Sugihara, J., Kaback, H. R. & Hellwig, P. pKa of Glu325 in LacY. Proc. Natl Acad. Sci. USA 114, 1530–1535 (2017).
Li, J. et al. Proton-coupled transport mechanism of the efflux pump NorA. Nat. Commun. 15, 4494 (2024).
Dmitrieva, N. et al. Transport mechanism of DgoT, a bacterial homolog of SLC17 organic anion transporters. EMBO J. 43, 6740–6765 (2024).
Søndergaard, C. R., Olsson, M. H. M., Rostkowski, M. & Jensen, J. H. Improved Treatment of ligands and coupling effects in empirical calculation and rationalization of pKa values. J. Chem. Theory Comput. 7, 2284–2295 (2011).
Olsson, M. H. M., Søndergaard, C. R., Rostkowski, M. & Jensen, J. H. PROPKA3: consistent treatment of internal and surface residues in empirical pKa predictions. J. Chem. Theory Comput. 7, 525–537 (2011).
Parker, J. L. et al. Cryo-EM structure of PepT2 reveals structural basis for proton-coupled peptide and prodrug transport in mammals. Sci. Adv. 7, eabh3355 (2021).
Jia, R. et al. Hydrogen-deuterium exchange mass spectrometry captures distinct dynamics upon substrate and inhibitor binding to a transporter. Nat. Commun. 11, 6162 (2020).
Hernández, S. B., Dörr, T., Waldor, M. K. & Cava, F. Modulation of peptidoglycan synthesis by recycled cell wall tetrapeptides. Cell Rep. 31, 107578 (2020).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Uehara, T. & Park, J. T. Peptidoglycan recycling. EcoSal Plus 3, 4.7.1.5 (2008).
Paulsen, H., Himpkamp, P. & Peters, T. Bausteine von Oligosacchariden, LXIX. Synthese von 1,6-Anhydromuramylpeptiden. Liebigs Ann. der Chem. 1986, 664–674 (1986).
Bepler, T., Kelley, K., Noble, A. J. & Berger, B. Topaz-Denoise: general deep denoising models for cryoEM and cryoET. Nat. Commun. 11, 5208 (2020).
Yan, R. et al. A structure of human Scap bound to Insig-2 suggests how their interaction is regulated by sterols. Science 371, eabb2224 (2021).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Wasilko, D. J. et al. Structural basis for CCR6 modulation by allosteric antagonists. Nat. Commun. 15, 7574 (2024).
Wayne, P. C. a. L. S. I. Performance standards for antimicrobial susceptibility testing. 32th ed. CLSI M100., (Clinical and Laboratory Standards Institute, 2022).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1-2, 19–25 (2015).
Huang, J. & MacKerell, A. D. Jr CHARMM36 all-atom additive protein force field: Validation based on comparison to NMR data. J. Comput. Chem. 34, 2135–2145 (2013).
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010).
Mark, P. & Nilsson, L. Structure and dynamics of the TIP3P, SPC, and SPC/E water models at 298 K. J. Phys. Chem. A 105, 9954–9960 (2001).
Vanommeslaeghe, K., Raman, E. P. & MacKerell, A. D. Jr. Automation of the CHARMM general force field (CGenFF) II: assignment of bonded parameters and partial atomic charges. J. Chem. Inf. Model.52, 3155–3168 (2012).
Hoover, W. G. Canonical dynamics: equilibrium phase-space distributions. Phys. Rev. A 31, 1695–1697 (1985).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Hess, B. P-LINCS: a parallel linear constraint solver for molecular simulation. J. Chem. Theory Comput. 4, 116–122 (2008).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. P. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360 (1996).
Pravda, L. et al. MOLEonline: a web-based tool for analyzing channels, tunnels and pores (2018 update). Nucleic Acids Res. 46, W368–W373 (2018).
Acknowledgements
We express our sincere gratitude to LigaChem Biosciences (Korea) for providing the compound (GlcNAc-1,6-anhMurNAc) essential for this study. Their support played a crucial role in conducting experiments and obtaining valuable results. We also extend our appreciation to the Institute of Membrane Proteins (Korea) for their technical assistance in generating cryo-EM grids using the Chameleon (Spt Labtech), as well as for supporting the initial screening and processing of the AmpG(cryo) complex. Data processing was performed using the GPGPU cluster at the Institute (NFEC-2025-03-304437). Data screening and collection for the AmpG(cryo-ww) apo and substrate complexes were conducted at the microscope facility at BaobabAIBio (Songdo, South Korea), whereas AmpG(cryo) data were collected at the IBS Research Solution Center (Daejeon, South Korea) and processed using the Olaf supercomputing system at the same facility. Computational resources for molecular dynamics simulations were provided by Nam-Chul Ha at the Research Institute of Agriculture and Life Sciences, Seoul National University. This study was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF), funded by the Ministry of Science and ICT (RS-2024-00351254, RS-2024-00344154, RS-2021-NR056553, NRF-2021R1A2C3010506, and NRF-2019M3E5D6063903, all awarded to H.C.; applicable to C.N., H.K., U.K., Y.C., and Y.Y.). This work was also supported by the Strategic Initiative for Microbiomes in Agriculture and Food, funded by the Ministry of Agriculture, Food and Rural Affairs (RS-2018-IP918012). Additional support was provided by the Brain Korea 21 (BK21) FOUR program. C.N., H.K., U.K., Y.C., and Y.Y. are BK21 FOUR fellowship recipients. Use of the cryo-EM facilities of the NEXUS consortium was supported by the NRF (RS-2024-00440289).
Author information
Authors and Affiliations
Contributions
Conceptualization: C.N. and H.K. Methodology: C.N., H.K., U.K., Y.C., Y.Y., J.W.K., H.L., K.M. and K.K. Investigation: C.N., H.K. and Y.Y. Visualization: C.N., H.K., U.K. and Y.Y. Supervision: D.Y. and H.C. Writing—original draft: C.N. and H.K. Writing, reviewing, and editing: C.N., H.K., J.L., J.L., Y.C., D.Y. and H.C.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jan Steyaert, Gabriel Torrens, Bert van den Berg, and the other, anonymous, reviewer for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chang, N., Kim, H., Kim, U. et al. Structural and functional insights of AmpG in muropeptide transport and multiple β-lactam antibiotics resistance. Nat Commun 16, 5744 (2025). https://doi.org/10.1038/s41467-025-61169-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61169-3