Abstract
SARS-CoV-2 and its emerging variants pose continuing threats to public health. SARS-CoV-2 assembles at the ER–Golgi intermediate compartment (ERGIC), where the viral membrane (M) protein highly accumulates to act as the central driver. However, how M is concentrated in the ERGIC, which hosts factor(s), may be involved, and whether they could be exploited as broad-spectrum antiviral targets remains unclear. Here, we identify an M-interacting host protein, ARF1, as a proviral factor that bolsters the propagation of SARS-CoV-2 and its variants in cultured cells and the viral infection and pathogenicity in female K18-hACE2 mice. By its N-terminal helix, ARF1 interacts with M and facilitates M’s ERGIC accumulation and thus M-driven virion production. Consistently, pharmacological ARF1 inhibition by small molecules disrupts both ARF1 and M concentration at the ERGIC, blocking virion assembly and propagation. Furthermore, a designed peptide mimicking the M-targeted motif of ARF1 competitively blocks M-ARF1 interaction, M accumulation at the ERGIC, and viral assembly and propagation in vitro. Moreover, the peptidomimetic inhibitor exhibits therapeutic efficacy against SARS-CoV-2 infection and pathogenicity in vivo. These findings provide critical insights into the basic biology of SARS-CoV-2 and demonstrate the potential to develop pan-SARS-CoV-2 therapeutics by targeting ARF1 and/or the ARF1-M interaction interface.
Similar content being viewed by others
Introduction
Coronavirus Disease 2019 (COVID-19) caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has resulted in unprecedented medical and socioeconomic impacts globally1,2. Moreover, SARS-CoV-2 has been undergoing continuous evolution, leading to the emergence of many variants of concern (VOCs), such as the Delta and Omicron strains3,4. The evolution of mutant strains that could reduce the efficacy of existing vaccines and drugs poses a persistent threat to public health, necessitating the development of new broad-spectrum antiviral strategies2,3,4.
SARS-CoV-2 consists of four structural proteins: spike (S), nucleocapsid (N), envelope (E), and membrane (M) proteins5. Despite the extensive studies conducted in the past few years, SARS-CoV-2 remains a newly emerging pathogen with numerous aspects that are yet to be fully understood, including its basic life cycle. Most of the attention on the virus life cycle has been focused on the S-mediated invasion6, viral enzyme-catalyzed RNA and protein production7,8, and regulations of host processes (especially immune responses)9,10,11. However, studies of virion assembly, a fundamental step in propagation, largely lag behind. As with other coronaviruses, SARS-CoV-2 assembles and buds at the ER-Golgi intermediate compartment (ERGIC) to produce progeny virions, achieving the viral propagation12,13,14,15,16. Therein, M is highly accumulated at the ERGIC and acts as the central driver of the virus assembly and morphogenesis14,15,16,17,18. Correspondingly, M is the most abundant structural constituent of SARS-CoV-2 virions and shapes the virions19,20,21. At a high local concentration, M creates a protein lattice across the ERGIC (or virion envelope) membranes that integrates the other viral membrane proteins S and E into the structural matrix and packages the N-RNA complex (ribonucleoprotein, RNP) by multiple molecular interactions16,20,21,22,23,24. However, it is unclear how M is enriched at the ERGIC. Particularly, which host factors may be involved in the ERGIC concentration of M, the prerequisite for efficient viral assembly, remains a mystery. Understanding the involvement of host factors in M accumulation and hence the driven virion production at the ERGIC would not only advance the knowledge of basic viral infection mechanisms but also may guide the development of host-directed antiviral therapies. In comparison with the conventional viral component-targeting approaches, antiviral interventions targeting the host factors critical for virus infection tend to have broad-spectrum antiviral properties and higher barriers to drug-resistant mutations. Thus, they are promising next-generation antiviral therapeutics against the evolving SARS-CoV-2 and COVID-19, deserving urgent research efforts.
In this study, through systematic functional experiments conducted in cellular and animal models, we identified a host protein, ADP-ribosylation factor 1 (ARF1), as an M-interacting proviral factor that bolsters the propagation and pathogenicity of SARS-CoV-2 and its variants in vitro and in vivo. Mechanistically, by the interaction with M, ARF1 facilitates the accumulation of M at the ERGIC and hence promotes M-driven virion assembly and propagation. Moreover, as proof of concept, a series of interventions by small molecules, dominant negative mutation, and designed peptidomimetics that targeted ARF1 or the ARF1-M interaction interface not only validated the function/mechanism findings but also showed broad-spectrum anti-SARS-CoV-2 potential in cellular and animal models.
Results
SARS-CoV-2 M protein interacts and colocalizes with host factor ARF1
Previous studies have reported the cellular interactome of SARS-CoV-2 proteins. However, for specific viral proteins, including M, there are great discrepancies in the identification results of different studies25,26,27,28,29. We thus further analyzed the potential cellular targets of M by Stag-pulldown assay, a highly specific and efficient affinity purification (AP) method29,30, coupled with mass spectrometry (MS). HEK293T cells were transfected with a Stag-fused M expressing plasmid or the control vector, followed by Stag-pulldown-MS analysis. Interestingly, many peptides of a cellular protein ARF1 that together cover 58.6% of the molecular length were specifically identified with high confidence in the cellular co-precipitates of SARS-CoV-2 M but not in the control (Supplementary Fig. S1). ARF1 has been suggested as a cellular protein potentially involved in several virus infections31,32,33,34,35; however, its role in SARS-CoV-2 infection is unknown. Moreover, the dysregulation of ARF1 is implicated in various human cancers, attracting many efforts to target ARF1 pharmacologically by small molecules for anti-tumor treatments36,37,38,39. Additionally, ARF1 has not been found in previous AP-MS-based studies of the potential M cellular interactome25,26,27,28, which also motivated our further validation of the interaction.
First, an independent pulldown and immunoblotting analysis confirmed the robust coprecipitation of ARF1 with SARS-CoV-2 M (Fig. 1a). Then, a reciprocal interaction assay with ARF1 as the bait demonstrated that M including its glycosylated form18,21 can be strongly coprecipitated by ARF1 (Fig. 1b). Furthermore, the interaction of M with endogenous ARF1 was also corroborated by additional co-precipitation (Co-IP) assays in the contexts of both transfection (Fig. 1c) and SARS-CoV-2 infection (Fig. 1d). Protein colocalization was next analyzed by immunofluorescence assays (IFA). As shown in Fig. 1e, both M and ARF1 were mainly localized in accumulational punctate regions in the cytoplasm and, importantly, exhibited remarkable colocalization in cells expressing the proteins by transfection. In addition, similar subcellular localization and colocalization of M and ARF1 could also be observed in SARS-CoV-2-infected cells (Fig. 1f), confirming the strong M-ARF1 association.
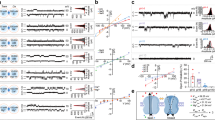
a Validation of the ARF1 co-precipitation with SARS-CoV-2 M by pulldown and WB analyses. HEK293T cells were co-transfected with the plasmids encoding M-Stag and ARF1-Flag, or the corresponding control vectors. At 24 h post-transfection (hpt), cells were lysed for Stag-pulldown assays, followed by WB analyses of the cell lysates (lysate input) and pulldown products with the indicated antibodies. b A reciprocal co-precipitation assay confirms the strong M-ARF1 interaction. HEK293T cells were co-transfected with the plasmids encoding ARF1-Stag and M-Flag, or control vectors. At 24 hpt, cells were lysed for additional pulldown and WB analyses. c Confirmation of the interaction of M with endogenous ARF1 by Co-IP. HEK293T cells were transfected with the control (vector) or M-encoding plasmids, followed by Co-IP assays using the ARF1-specific antibody or control IgG, and WB analyses. d Interaction of endogenous ARF1 with M in the context of SARS-CoV-2 infection. Huh-ACE2 cells were mock-infected or infected with SARS-CoV-2 at a multiplicity of infection (MOI) of 3. At 24 h post-infection (hpi), cell lysates were delivered to Co-IP and WB analyses as in (c). e Colocalization of M with endogenous ARF1 in the context of transfection. HeLa cells were transfected with the plasmids expressing M-Stag or the control vector. Twenty-four hours later, cells were fixed for IFA with the antibodies against Stag and ARF1, respectively. Nuclei were stained with Hoechst. f Colocalization of M with endogenous ARF1 in the context of SARS-CoV-2 infection. HeLa cells were transfected with the ACE2 expression plasmid, followed by SARS-CoV-2 infection (MOI, 1). At 24 hpi, cells were fixed for IFA with the antibodies against M and ARF1. Bars, 5 μm. These data are representative of three independent experiments with similar results. See also Fig. S1. Source data are provided as a Source data file.
ARF1 is a pro-viral host factor for SARS-CoV-2 and its variants
Subsequently, to uncover the role of ARF1 in SARS-CoV-2 infection, we first silenced ARF1 expression by RNAi. ARF1 knockdown (KD) significantly inhibited SARS-CoV-2 infection in HEK293 cells expressing human angiotensin I-converting enzyme-2 (ACE2), as manifested by the decreased virion and viral protein yields (Fig. 2a–c and Supplementary Fig. S2a). Moreover, the siRNA with relatively higher silence efficiency (siARF1-2) exhibited a stronger inhibitory activity against the virus, showing a dose dependence (Fig. 2a–c and Supplementary Fig. S2a). To further confirm the significance of ARF1 in SARS-CoV-2 infection, we generated ARF1-knockout (ARF1-KO) HEK293 cell lines by using the CRSPR-Cas9 system, which were confirmed with DNA sequencing and immunoblotting (Supplementary Fig. S3a–c). Interestingly, the viral proliferation and protein production were more severely blocked in all three ARF1-KO cell lines (Fig. 2d, e and Supplementary Fig. S2b). Furthermore, a significant increase in the viral propagation was observed in cells overexpressing ARF1, although the ARF1 overexpression seemed to only slightly enhance the viral protein production (Fig. 2f, g and Supplementary Fig. S2c). The effects of ARF1 overexpression were overall limited, indicating that the endogenous expression level of ARF1 is likely sufficient to support the virus infection. Thus, in the following study, we evaluated the influence of ARF1 reconstitution in the KO cells by transfection of the ARF1 expression plasmid. Indeed, ARF1 reconstitution led to substantial recovery of the virus propagation (Fig. 2h, i and Supplementary Fig. S2d). Collectively, these results establish that ARF1 is a significant host factor bolstering SARS-CoV-2 propagation.
a–c HEK293 cells were co-transfected with the ACE2-Stag expression plasmid and ARF1-specific siRNAs or siRNA negative control (siNC). At 24 hpt, cells were infected with SARS-CoV-2 (MOI, 0.01) for 24 h. Following the confirmation of the ARF1 mRNA KD (a), SARS-CoV-2 propagation was then determined by plaque-forming unit (PFU) counting of infectious progeny virions (b). The viral N and ARF1 protein production was also analyzed by WB (c). d, e Wildtype (WT) or ARF1-KO (three clones) HEK293 cells were transfected with the ACE2 expression plasmid, followed by SARS-CoV-2 infection. At 24 hpi, the viral propagation (d) and protein production (e) were analyzed by plaque assays and WB, respectively. f–i The effects of ARF1 overexpression and reconstitution on SARS-CoV-2 propagation. WT cells (f, g) or the ARF1-KO (HEK293-ARF1-KO-2) cells (h, i) co-transfected with the ACE2 and ARF1 expression plasmids were similarly infected with SARS-CoV-2 at 24 hpt, followed by the viral propagation (f, h) and protein yield detections (g, i). j–m WT or ARF1-KO HEK293 cells transfected with the ACE2 expression plasmid were similarly infected with the indicated SARS-CoV-2 variants, followed by the viral propagation (j–l) and protein yield (m) analyses. Data are representative of three independent experiments with similar results (c, e, g, i, m). Graphs are presented as means ± SD, n = 3 (a, b, d, h, j–l) or 4 (f) independent biological replicates. Comparisons were performed with two-tailed Student’s t-test (f, h, j, k, l) or one-way ANOVA with Dunnett’s test (a, b, d). ***p < 0.001; ****p < 0.0001. See also Figs. S2 and S3. Source data with detailed p-values is provided as a Source data file.
M is highly conserved among SARS-CoV-2 variants40. We speculated that host ARF1 could be a common requirement for the efficient infection of SARS-CoV-2 and its variants. Thus, the effect of ARF1 on several SARS-CoV-2 variants, including Delta and Omicron (BA.4 and BA.5), was also assessed. Indeed, ARF1 KO greatly impaired the propagation of all the tested variants (Fig. 2j–m and Supplementary Fig. S2e–g), suggesting that the requirement for ARF1 is conserved in infection of SARS-CoV-2 and its variants.
ARF1 supports SARS-CoV-2 infection and pathogenicity in vivo
Since ARF1 showed remarkable pro-SARS-CoV-2 activity in cultured cells, we asked whether ARF1 affects the virus infection and pathogenicity in vivo. As ARF1 KO is embryonically lethal in mice41, we here exploited an ARF1-KD mouse model by transient transduction of a lentiviral vector expressing specific mouse ARF1 (mARF1)-targeting shRNA via intravenous injection (iv)42,43. Adult K18-hACE2 transgenic mice were transduced with the control or ARF1-targeting shRNA expression vectors, followed by intranasal instillation with SARS-CoV-2 suspension (5 × 102 TCID50) under deep anesthesia at a week post-transduction. Consistent with previous reports44,45,46, the humanized mouse model was highly susceptible to the virus, and all infected animals indistinguishably exhibited severe weight loss from 4 days post-infection (dpi) (Supplementary Fig. S4a). One animal in the control group (i.e., 1/5) was dead at 4 dpi, and the others, losing 20% or more of their body weight, had to be sacrificed at 5 dpi for further analyses (Supplementary Fig. S4a, b). By qPCR and WB analyses, ARF1 expression was confirmed to be partially knocked down in the mouse lungs (Fig. 3a, c). Then, compared with the control group, the viral titers and N protein levels in the lung tissues of mice with ARF1 KD were significantly decreased (Fig. 3b, c). Consistently, immunohistochemical (IHC) staining showed that the viral infection positive area in lung tissue was reduced in the ARF1-KD animals (Fig. 3d). Moreover, histopathological changes in lungs, including inflammatory cell infiltration and alveolar wall congestion and thickening, were also attenuated in the ARF1-KD group as indicated by H&E staining analyses (Fig. 3e). Collectively, these findings suggest that in line with the data obtained in cultured cells, ARF1 likely also contributes to SARS-CoV-2 infection and pathogenicity in vivo.
a–d ARF1 KD inhibits SARS-CoV-2 infection in K18-hACE2 transgenic mice. K18-hACE2 mice were transduced with the viral vectors encoding control or mARF1-targeting shRNAs (5 × 107 transduction units, 100 μL) via intravenous injection, followed by SARS-CoV-2 (500 TCID50, 50 μL) infection through intranasal inoculation at 7 d post-transduction. Five days post infection (dpi), mice were sacrificed for analyses of the ARF1 mRNA levels (a) or infectious virion titers (b) in the lungs by plaque assays or the protein production by WB (c) and IHC (d). Data are presented as means ± SD (a) or median with interquartile range (b), n = 5 mice/group. Statistical significance was determined by two-tailed Student’s t-test (a) or Mann–Whitney U-test (b). **p < 0.01. Representative IHC analyses using the anti-N antibody identified SARS-CoV-2-infected cells in the lung tissue sections (d). e ARF1-KD mice exhibited ameliorated pathological injury. Lung H&E staining shows evident histopathological changes, including alveolar wall congestion and thickening (red arrows) and inflammatory cell infiltration (black arrows) in the control group, which were milder in the ARF1-KD group. Data are representative of similar results from testing five animal samples per group. Scale bars, 100 μm. Source data are provided as a Source data file.
In the K18-hACE2 mouse model, disproportionately high brain infection has been documented following SARS-CoV-2 challenge47,48. This severe brain infection is considered a major reason for mortality and a drawback of the model for SARS-CoV-2 infection and intervention research, as it is not typically seen in human patients47,48. We also analyzed the effects of ARF1 KD in the animal brains. A significant reduction in the virus loads was observed in the ARF1-KD group (Supplementary Fig. S4c, d), supporting the proviral role of ARF1. However, consistent with previous studies, brain infections were generally severe, and even in the ARF1-KD group, infectious virus titers remained very high (Supplementary Fig. S4d). This may account for the lack of stronger survival protection observed after the partial depletion of ARF1 expression, in addition to the limited KD efficiency.
ARF1 enables M accumulation to the ERGIC
Coronavirus morphogenesis takes place at the ERGIC, where M accumulates to a high local concentration to propel the virion assembly13,14,15,16,17,18,19. However, the mechanism for M’s concentration in the ERGIC remains unclear. ARF1 localization at ERGIC has been reported previously49,50. Consistently, we observed that ARF1 itself exhibited evident ERGIC localization irrespective of M expression, while it co-accumulated with M to the ERGIC in cells co-expressing the two proteins, exhibiting a notable colocalization (Supplementary Fig. S5). Based on these observations, together with the strong interaction between M and ARF1, we hypothesized that ARF1 might be involved in the accumulation of M in the ERGIC, thereby facilitating viral assembly and propagation. To validate the hypothesis, we first analyzed M localization in ARF1-KO HeLa cells (Supplementary Fig. S3d, e) by IFA. As shown in Fig. 4a, compared with the agminated ERGIC localization pattern in wild-type cells, M exhibited a more diffused cytoplasmic distribution in ARF1-KO cells, and its accumulation at the ERGIC was significantly reduced (Fig. 4a, b). This result indicates that ARF1 is likely required for the efficient concentration of M at the ERGIC. Consistently, similar findings were obtained in the context of SARS-CoV-2 infection (Fig. 4c, d). Further, we reconstituted ARF1 expression in ARF1-KO cells by transfection of the plasmid encoding GFP-tagged ARF1 (ARF1-GFP) for additional IFA analysis. Interestingly, M was indeed repositioned to accumulate in the ERGIC, along with ARF1-GFP, compared to the diffused distribution in the control cells with expression of GFP alone (Fig. 4e, f). Together, these data suggest that ARF1 interacts with M and facilitates the accumulation of the viral protein at the ERGIC.
a, b WT or ARF1-KO HeLa cells transfected with the M expression plasmid were fixed at 24 hpt for IFA with antibodies against the tag and ERGIC53. PCC (ME), Pearson’s correlation coefficient (PCC) values for M and ERGIC colocalization. c, d WT or ARF1-KO cells expressing ACE2 were infected with SARS-CoV-2 (MOI, 1). At 24 hpi, cells were fixed for IFA and PCC analyses. e, f ARF1-KO HeLa cells were co-transfected with the M expression plasmid together with the plasmid expressing ARF1-GFP or GFP control, followed by IFA and PCC analyses. g, h HeLa cells were co-transfected with the plasmids expressing M and Flag-tagged ARF1-T31N, or the control vector, followed by IFA and PCC analyses. Bars, 10 μm. i HEK293T cells were co-transfected with the VLP packaging plasmids along with the indicated siRNAs. At 36 hpt, VLPs produced in culture medium were collected for Nano-Luc luciferase signal collection. Relative VLP levels were calculated by normalization to the siNC control group. j WT or ARF1-KO HEK293 cells were co-transfected with the packaging plasmids, followed by Nano-Luc signal collection of the produced VLPs at 36 hpt. k HEK293T cells were co-transfected with the packaging plasmids, along with the ARF1-Flag expression plasmids or the vector control, followed by VLP luciferase signal quantification. l ARF1-KO HEK293 cells were co-transfected with the packaging plasmids, together with the ARF1-Flag expression plasmid or the control vector, followed by VLP signal analyses. Data are medians with interquartile ranges, n = 10 cells (b, d, f, h), or means ± SD, n = 4 biological replicates (i–l). The p-values were determined using the Mann–Whitney U-test (b, d, f, h), one-way ANOVA with Dunnett’s test (i, j), or two-tailed Student’s t-test (k, l). ***p < 0.001; ****p < 0.0001. Source data are provided as a Source data file.
ARF1(T31N) is an inactive ARF1 mutant in which Thr is replaced by Asn at position 31 of the ARF1 amino acid sequence51. Biochemical analyses suggest that the mutation likely converts ARF1 to a form with a preferential affinity for GDP51. In the GDP-bound conformation, the membrane-binding features of ARF1 could be masked, resulting in its dispersed localization to the cytosol49,52,53. Interestingly, expression of ARF1(T31N), which, consistent with the previous studies54, loses the capacity to be efficiently localized at ERGIC, could also destroy the accumulational distribution pattern of M and more specifically, the efficient concentration of M to the ERGIC (Fig. 4g, h). Protein interaction analysis showed that the mutation did not impair the interaction with M (Fig. S6a), suggesting that ARF1(T31N) retains the ability to interact with M and thus likely competitively inhibits endogenous wild-type ARF1-directed accumulation of M at the ERGIC. These observations support the role of ARF1 in facilitating M localization at the ERGIC as well.
ARF1 bolsters M-driven viral assembly
Given the important role of ARF1 in M accumulation at the ERGIC, a central process for virion production, we considered that ARF1 deficiency likely would impair the virus assembly and hence propagation. Indeed, a scarcity of virus particles assembled in ARF1-KO cells was observed under transmission electron microscopy, in stark contrast to the abundant presence of virions within vesicular/tubular compartments that bear resemblance to the ERGIC morphology55 in SARS-CoV-2-infected WT cells (Supplementary Fig. S7). Previous studies have shown that depletion of ARF1 itself had no noticeable effect on cellular secretion despite the potential regulatory role in protein trafficking56,57,58. Consistently, ARF1 KO did not significantly interfere with the secretion pathway, either in our analysis (Supplementary Fig. S8), ruling out a potential effect caused by disruption of global secretion. In agreement, we observed decreased virion assembly but not accumulation and detention of assembled virions in ARF1-KO cells (Supplementary Fig. S7). To further assess the effect of ARF1 on virion assembly, we constructed a built-in luminescence-based reporting system for M-driven virus-like particle (VLP) production. The system allows the quantification of produced VLP, which is labeled by a HiBiT tag through co-expression of SARS-CoV-2 structural proteins, including M, HiBiT-tagged S (S-HiBiT), N, and E59,60,61,62. We observed that, indeed, VLP yield was significantly reduced by ARF1 KD (Fig. 4i). Then, ARF1 KO led to more dramatic blockade of the VLP production (Fig. 4j). Next, expression of ARF1(T31N) by transfection with the ARF1(T31N)-encoding plasmid was also shown to decrease VLP yield (Supplementary Fig. S6b) and the viral propagation (Supplementary Fig. S6c), in agreement with its disruption of M accumulation at the ERGIC (Fig. 4g, h). Conversely, wildtype ARF1 overexpression resulted in a moderate but significant promotion of the VLP production (Fig. 4k). It further supports the role of ARF1 in M-driven virion assembly and also reflects that endogenous ARF1 could be basically sufficient to prop up virion production, consistent with the moderate enhancement of the virus propagation by ARF1 overexpression (Fig. 2f). Furthermore, we observed more substantially recovered VLP production when ARF1 was reconstituted by transfection in the ARF1-KO cells (Fig. 4l). We also exploited another VLP reporter system with HiBiT-tagged N63,64 for similar analyses and obtained consistent results (Supplementary Fig. S9), validating the role of ARF1 in facilitating M-directed assembly. Altogether, these results substantiate that by facilitating M accumulation at the ERGIC, ARF1 enhances M-driven virion assembly and hence propagation.
ARF1 small-molecule inhibitors disrupt ARF1 and M localization at the ERGIC and abate virion assembly, inhibiting SARS-CoV-2 propagation
There are existing small-molecule inhibitors against ARF1, including Brefeldin A (BFA) and Golgicide A (GCA) that freeze ARF1 in the inactive state and destroy ARF1 anchoring to the ERGIC65,66,67. Interestingly, we found that treatments with the inhibitors BFA and GCA at concentrations of no noticeable cytotoxicity both greatly inhibited SARS-CoV-2 propagation (Figs. 5a, b and Supplementary Fig. S10a, b). Further, we analyzed the effects of the drugs on ARF1 and M cellular localization. As shown in the following IFA results, the localization patterns of ARF1-GFP along with M were obviously changed from accumulated puncta to diffused distribution by the treatments with BFA and GCA (Fig. 5c and Supplementary Fig. S10c). More specifically, these agents disrupted the ERGIC localization of ARF1 and consequently, also deprived M of its efficient accumulation in the ERGIC, compared with the control group (Fig. 5c, d). Consistently, we further demonstrated that treatments with BFA and GCA both significantly inhibited VLP production (Fig. 5e and Supplementary Fig. S10d, e), indicating that by disrupting the accumulation of ARF1 and hence M to the ERGIC, the drugs interfere with M-driven virion production. These results of pharmacological interventions not only support the notable role of ARF1 in sustaining M accumulation at the ERGIC and promoting virion propagation but also suggest the possibility of targeting ARF1 for antiviral therapy.
a, b ARF1 small-molecule inhibitors BFA and GCA inhibit SARS-CoV-2 propagation. Huh7-ACE2 cells were infected with SARS-CoV-2 at an MOI of 0.01 in the presence of the indicated drugs or DMSO control for 16 h or 24 h. The viral yields in the cell supernatants were then quantified by plaque assays. No significant cytotoxicity of these drugs to Huh7-ACE2 cells or to the other cells (Supplementary Fig. S10a–e) used below was observed under the indicated concentrations as measured by the CCK-8 assays. c, d BFA and GCA disrupt the accumulated localization patterns of ARF1 and M at the ERGIC. HeLa cells were co-transfected with the plasmids encoding ARF1-GFP and M-Stag. At 24 hpt, cells were treated with BFA, GCA, or DMSO for 20 min, followed by IFA analyses to visualize the distribution of ARF1-GFP and M. PCC(ME), PCC colocalization efficiency between M and ERGIC. Nuclei were stained with Hoechst. Bars, 5 μm. e BFA and GCA inhibit M-driven VLP production. HEK293T cells were co-transfected with the VLP packaging plasmids. Six hours later, the culture medium containing liposome-plasmid complexes was replaced with fresh medium supplemented with the indicated doses of drugs or DMSO for a further 24 h, followed by the relative quantification of VLPs based on luciferase activities. Data are presented as means ± SD, n = 3 (a, b) or 5 (e) independent biological replicates, or medians with interquartile ranges, n = 15 cells (d). One-way ANOVA with Dunnett’s test (a, b, e) and Kruskal–Wallis H-test followed by Dunn’s Q-test (d): ****p < 0.0001. Source data are provided as a Source data file.
The N-terminal helix of ARF1 (ARF11–17) is the target domain of SARS-CoV-2 M, and when expressed alone, disrupts M enrichment at the ERGIC
To further characterize M hijacking of ARF1 and uncover clues for additional intervention strategies targeting the ARF1-M interaction interface, we then mapped the pivotal domain(s) of ARF1 targeted by M. A series of ARF1-truncated mutants tagged with GFP were constructed for the following protein interaction analysis (Fig. 6a). As shown in Fig. 6b, the full-length ARF1-GFP, but not the GFP control, was specifically pulled down by M, consistent with the observations above (Fig. 1a–d). Further, all the C-terminally truncated ARF1 mutants that contain the N-terminal helix of 17 amino acids (ARF11–17), including even the ARF1–17 alone, could be notably co-precipitated by M, although the protein expression or stability appeared to be affected by the truncation to different extents (Fig. 6b). In line with the results, an N-terminal truncation with deletion of the helix (ARF1Δ17) greatly disrupted the interaction with M (Fig. 6b). These data indicate that the N-terminal helix of ARF1, i.e., ARF11–17, is the major target domain of M.
a Schematic diagram of GFP-tagged ARF1 truncated mutants used in this study. Several structural domains are indicated. Hel helix, SW switch, In SW inter switch. b Mapping of the ARF1 domain targeted by M. HEK293T cells was co-transfected with the M-Stag expression plasmid and the plasmids encoding full-length or truncated ARF1 proteins fused with GFP or the pEGFP-N3 control plasmid. At 24 hpt, protein interactions were analyzed by Stag-pull-down and WB analyses. *The major bands of ARF1 and its truncated mutants tagged with GFP in lysate inputs. Data are representative of three independent replicates with similar results. c–e Colocalization analyses of M with representative ARF1 mutants (ARF11–17 and ARF1Δ17) and the effects of the mutant expression on M accumulation in the ERGIC. HeLa cells were co-transfected with the M expression plasmid and the plasmids encoding GFP (pEGFP-N3) or GFP-tagged full-length or truncated ARF1 proteins. At 24 hpt, cells were fixed for IFA analysis. Nuclei were stained with Hoechst. Bars, 5 μm. PCC (MG), PCC between M and GFP or GFP-fused proteins; PCC (ME), PCC between M and ERGIC. The statistical diagrams are presented as medians with interquartile ranges, n = 10 cells (d, e). Kruskal–Wallis H-test followed by Dunn’s Q-test for pairwise comparisons (e): ns not significant; **p < 0.01. Source data are provided as a Source data file.
Since ARF11–17 is critical for the interaction with M, we further investigated its potential effects on the protein subcellular localization and colocalization. By contrast to the controls (GFP and ARF1Δ17-GFP), both ARF1-GFP and ARF11–17-GFP exhibited specific colocalization with M (Fig. 6c, d), in agreement with the protein interaction analyses (Fig. 6b). However, compared with the evident ERGIC localization of M in the GFP-expressing control cells, expression of ARF11–17-GFP, but not ARF1Δ17-GFP, markedly interfered with M localization to the ERGIC (Fig. 6c, e), while ARF1-GFP seemed to slightly promote the ERGIC localization of M, albeit not statistically significant (Fig. 6c, e). That is to say, although ARF11–17, like the full-length ARF1, has the ability to interact with M, its expression leads to a converse effect on the M’s accumulation in the ERGIC (Fig. 6c–e). The data suggest that ARF11–17 likely competes with endogenous ARF1 to interact with M and therefore affects the endogenous ARF1-mediated ERGIC localization of M. These findings corroborate the important role of ARF1-17 in M usurpation of ARF1 and indicate a potential capacity of the short domain to interfere with the ARF1-M interface when expressed alone.
Additionally, we also attempted to map the potential domain(s) of M required for the interaction with ARF1 using a series of M mutant expression constructs. Further interaction analyses showed that various deletions of the M domains tested all led to the loss of interaction with ARF1 (Supplementary Fig. S11a, b), indicating that the integrity of M appears to be crucial for its interaction with ARF1. In agreement with the interaction mapping, all the M mutants with the structural domains deleted evidently lost the strong colocalization with ARF1 and correspondingly, lost the highly efficient concentration to the ERGIC (Supplementary Fig. S11c). These results also consistently support the importance of the interaction with ARF1 for M accumulation in the ERGIC.
A synthesized peptide mimicking ARF11–17 blocks M hijacking of ARF1, suppressing viral assembly and propagation
To further evaluate the effects of ARF11–17, we designed a mimic peptide (referred to as PEP17) that contains ARF11–17 to aim to competitively obstruct the M-ARF1 interface and is conjugated with a cell membrane-penetrating sequence containing 14 amino acids (briefly named CPS14) for intracellular uptake68,69. As seen in Fig. S10f–k, neither PEP17 nor the control peptide CPS14 had any noticeable cytotoxicity at the tested concentrations. The peptides did not affect cellular secretion either (Supplementary Fig. S12). Interestingly, treatment of cells with PEP17 evidently changed the localization pattern of M to diffused distribution and more specifically, reduced the M-ARF1 colocalization and M (but not ARF1) accumulation to the ERGIC (Fig. 7a). Indeed, we further showed that PEP17, but not the control, could inhibit the M-ARF1 interaction by the following pulldown analysis (Fig. 7b). Consistently, VLP reporter assays next revealed that PEP17 dose-dependently suppressed VLP assembly with a half-maximal inhibitory concentration (IC50) of 1.53 μM, while the control peptide CPS14 did not exhibit such inhibitory effect even at higher concentrations (Fig. 7c, d). Furthermore, we examined the potential of PEP17 to affect SARS-CoV-2 propagation. Indeed, PEP17 exhibited a notable anti-SARS-CoV-2 activity, dose-dependently inhibiting the viral propagation with an IC50 of 2.15 μM as determined by qPCR analysis of genomic copies (Fig. 7e). More notably, its IC50 for inhibiting the infectious virion production measured by plaque assays reached a sub-micromolar level (0.82 μM) (Fig. 7g). In contrast, the control peptide treatment did not result in any apparent inhibition to SARS-CoV-2 (Fig. 7f, h). Taken together, these consistent results suggest that by disrupting M usurpation of ARF1, the synthesized PEP17 can significantly inhibit SARS-CoV-2 assembly and propagation.
a PEP17 inhibits the M-ARF1 colocalization and M’s ERGIC accumulation. HeLa cells were transfected with the M expression plasmid. Six hours later, specifically at the time of transfection medium exchange, cells were treated with 5 μM PEP17 or CPS14 (control) for 24 h, followed by IFA. Bars, 5 μm. PCC (MA), PCC between M and ARF1; PCC (ME), PCC between M and ERGIC; PCC (AE), PCC between ARF1 and ERGIC. b PEP17 inhibits the M-ARF1 interaction. HEK293T cells were co-transfected with the plasmids encoding M-Stag and ARF1-Flag or control vectors. Six hours later, cells were treated with 5 μM PEP17 or CPS14 control for 24 h, followed by Stag-pulldown and WB. Data are representative of three independent replicates with similar results. c, d PEP17 inhibits M-driven VLP production. HEK293T cells were co-transfected with the VLP packaging plasmids and treated with the indicated doses of PEP17 (c) or CPS14 (d) for 24 h, followed by VLP signal quantification. IC50 was calculated by GraphPad Prism 9 (c, right panel). e–h PEP17 suppresses SARS-CoV-2 propagation. Huh7-ACE2 cells were infected with SARS-CoV-2 (MOI, 0.01) in the presence of different doses of PEP17 (e, g) or CPS14 control (f, h) for 24 h. The virus yields in the cell supernatants were quantified by qRT-PCR (e, f) and plaque assays (g, h). i–n PEP17 restricts SARS-CoV-2 variants’ propagation. Huh7-ACE2 cells were infected with the indicated SARS-CoV-2 variants (MOI, 0.01) in the presence of PEP17 (i–k) or CPS14 control (l–n) for 24 h, followed by progeny quantification. Data are medians with interquartile ranges, n = 15 cells (a), or means ± SD, n = 3 (c, e–n) or 4 (d) independent biological replicates. Mann–Whitney U-test (a) or one-way ANOVA with Dunnett’s test (c–n): ns not significant; *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001. Source data are provided as a Source data file.
Additionally, the antiviral activity was evaluated against pandemic variants of SARS-CoV-2. Also, the propagation of the viruses was remarkably repressed by PEP17 in a dose-dependent manner, whereas no marked suppressive influence was observed in the CPS14 control group even with higher concentrations (Fig. 7i–n).
These data from mutual validation of the function/mechanism findings and the function/mechanism-based antiviral design not only further underline the significance of M hijacking of ARF1 in SARS-CoV-2 propagation, but also support a potential, broad-spectrum antiviral strategy targeting the M-ARF1 interface.
ARF1 N-myristoylation, which can occur co-translationally within ARF11–17, is involved in the protein physiology by enhancing its membrane association49,70. However, based on our above analyses of PEP17 without N-myristoylation, the modification is likely not required for the interaction with M. To provide further evidence, we constructed GFP-tagged mutants, ARF1(G2A) and ARF1-17(G2A), in which the glycine at position 2 was replaced by an alanine to delete the N-myristoylation modification71. Protein interaction analyses showed that these mutants could still evidently interact with M (Supplementary Fig. S13a), confirming that the modification is not essential for the interaction with M. Consistently, treatment with a potent inhibitor of N-myristoylation modification, PCLX-00172, did not abolish ARF1-M interaction either (Supplementary Fig. S13b). Furthermore, we also synthesized a peptide with the N-myristoylation modification (Myr-PEP17) and found that it could inhibit SARS-CoV-2 propagation as well (Supplementary Fig. S13c). However, the antiviral activity was not significantly enhanced, compared to the unmodified PEP17 (Supplementary Fig. S13c). These additional analyses consistently suggest that the modification is not essential for the interaction with M or for the antiviral activity of the peptide.
PEP17 inhibits SARS-CoV-2 infection and pathogenicity in hamsters
Syrian hamsters are generally used as a SARS-CoV-2 infection model to evaluate the drug and vaccine effects73,74,75,76. These animals are considered to be able to better recapitulate common human infections, showing evident but non-lethal respiratory injury, in comparison with the excessive vulnerability of the ACE2 humanized mouse model. We thus exploited the model to further evaluate the antiviral efficacy of PEP17 in vivo. First, 6-week-old hamsters were infected with 104 TCID50 of SARS-CoV-2 by intranasal instillation. Then, 2 h post-infection, they were injected with PEP17 intraperitoneally (30 mg/kg body weight) once a day for three consecutive days. In the following 3 days after the viral challenge, all the infected animals exhibited good tolerance, showing no discernible adverse effects except slight weight loss, and moreover, no significant difference in the weight loss was detected between the PEP17 and CPS14 treated groups. Subsequently, the hamsters were euthanized at 3 dpi, and the lung, trachea, and brain tissues were collected for the following analyses. RT-qPCR results showed that the lung tissues had high viral RNA copies, followed by the trachea, while the brains displayed the lowest levels in comparison, which is consistent with the previous reports74,75. Intriguingly, the viral titers in all these tested organs were significantly decreased by the PEP17 treatment (Fig. 8a–c). Viral antigens (the N proteins) could be readily detected in the lung and trachea tissues by WB and in the lungs by IHC staining (Fig. 8d–f). Consistently, compared with the control groups, the viral antigen levels in the trachea and lung tissues of hamsters treated with PEP17 were also decreased as indicated in the WB analyses (Fig. 8d, e). Moreover, the IHC staining showed less viral antigen positive areas in the lung tissues of hamsters treated with PEP17 (Fig. 8f). These results suggest the significant inhibition of viral propagation and spread in vivo by the PEP17 treatment. Furthermore, histopathological analyses by H&E staining showed that pathological changes in the lung tissues including the alveolar structure damage and inflammatory cell infiltration were reduced by the PEP17 treatment (Fig. 8g). Moreover, the treatment with PEP17 also evidently mitigated exfoliation, necrosis, and hydropic degeneration of mucosal epithelial cells and inflammatory cell infiltration in submucosa of the trachea (Fig. 8h). Overall, the histopathological damages induced by SARS-CoV-2 in both the lungs and trachea were ameliorated after PEP17 treatment, as suggested by the quantitative analyses (Fig. 8i, j). Together, these data validate the anti-SARS-CoV-2 therapeutic efficacy of PEP17 in vivo and suggest the potential to target the virus-host (M-ARF1) interaction interface for the development of novel antiviral intervention strategies against COVID-19.
Syrian hamsters (n = 5/group) were infected with SARS-CoV-2 and then treated with PEP17 or CPS14 control for 3 days. At three dpi, the animals were sacrificed, and the indicated tissues were collected to detect the infectious virus loads (a–c) and antigen (N) yields in tissue homogenates (d, e) and the antigen (N) distribution in lung sections by IHC (f). Data are representative of five biological replicates with similar results (d–f). Representative H&E staining analyses of lung (g) and trachea (h) tissues show alleviated tissue injuries and inflammation by the PEP17 treatment. Noticeable pathological changes are indicated with arrows. Scale bars, 500 μm (f), 100 μm (g) and 20 μm (h). Black arrow, inflammatory cell infiltration around interstitial blood vessels and bronchus (g) or in the tracheal submucosa (h); blue arrow, alveolar wall thickening and alveolar collapse or disappearance of alveolar cavity structure (g); green arrow, exfoliation of mucosal epithelial cells (g, h); red arrow, hydropic degeneration of mucosal epithelial cells (h). i, j Quantitative analyses of histopathological damages in the lung and trachea tissues. Lesion scores were ranked on a scale of 1–4, indicating mild, moderate, moderate to severe, and severe, respectively, while 0 indicates no detectable lesion. Cartograms are presented as medians with interquartile ranges, n = 5 animals/group (a–c and i–j); p-values were determined using the Mann–Whitney U-test. *p < 0.05; **p < 0.01. Source data are provided as a Source data file.
Discussion
As obligate intracellular parasites, viruses depend on their host cells to propagate. Virion assembly is fundamental to completing the infection cycle. However, the virus-host interactions during SARS-CoV-2 assembly remain unclear, hindering the understanding of the viral infection mechanisms and the design of antiviral interventions targeting this essential step of the viral life cycle. SARS-CoV-2 M is the major structural protein that plays a central role in orchestrating the virion assembly and propagation. Here, we identified the host protein ARF1 as an M-interacting proviral factor that supports the infection and pathogenicity of SARS-CoV-2 and its variants in vitro and in vivo. By the ARF1-M interaction, ARF1 facilitates M accumulation at the ERGIC and thus M-driven viral assembly and propagation. Furthermore, small-molecule inhibitors that disrupt the ERGIC accumulation of ARF1 and hence M exhibit strong anti-SARS-CoV-2 activities. Otherwise, the dominant negative mutant ARF1(T31N) that maintains the interaction with M but loses the ability to accumulate at the ERGIC could, as expected, competitively interfere with M enrichment at the ERGIC and reduce the viral assembly. Moreover, an artificially synthesized peptide mimicking the M-targeted motif of ARF1 could block M hijacking of ARF1 and also significantly inhibit SARS-CoV-2 propagation and pathogenicity in cellular and animal models. Overall, the study identified ARF1 as a proviral factor hijacked by M for efficient viral assembly and as a candidate broad-spectrum anti-SARS-CoV-2 target that could be intervened by multiple strategies. These findings (summarized in Fig. 9) not only shed light on the knowledge gap of virus-host interactions during the fundamental assembly step of the SARS-CoV-2 life cycle, but also provide clues for the development of new pan-SARS-CoV-2 therapeutic methods.
SARS-CoV-2 assembles at the ERGIC to produce progeny virions, achieving its propagation. By interacting with SARS-CoV-2 M, the host protein ARF1, which itself is localized at the ERGIC, enables the accumulation of M, the central driver of the viral assembly, to the ERGIC. At a high local concentration, M then drives and nucleates the virion morphogenesis at the ERGIC by complex molecular interactions with the adjacent M, E, S, and RNP. Small-molecule inhibitors against ARF1, such as BFA and GCA, can disrupt the ERGIC localization of ARF1 and consequently reduce M accumulation at ERGIC, inhibiting the viral propagation. Otherwise, interventions targeting the ARF1-M interaction interface, such as the peptidomimetic PEP17, can block the ARF1-M interaction, i.e., M hijacking of ARF1, and thus ablate M accumulation at the ERGIC, also leading to the inhibition of viral propagation. M matrix/membrane protein, E envelope protein, S spike, RNP ribonucleoprotein, ERGIC the ER-Golgi intermediate compartment. Created with BioRender (Min Y. 2025; https://BioRender.com/ja67qc5).
Coronaviruses assemble at the ERGIC, where M is highly concentrated to drive the virion formation. However, how coronavirus M accumulates in the ERGIC is a long-standing unanswered question. Considering the lack of typical location or retention signal sequences in M itself, host factors are likely involved in the viral protein concentration in the ERGIC. Using SARS-CoV-2 M as a model, the present study demonstrated that ARF1 is an M-interaction host factor required for M accumulation at the ERGIC, providing insights into the virus-host interactions during the viral assembly. The M protein sequences of SARS-CoV-2 and its variants are almost identical40. Moreover, M proteins are also highly conserved among the members of Sarbecovirus subgenus of β-Coronavirus. For instance, SARS-CoV-2 and SARS-CoV M proteins share a sequence identity of >90%. Given the high conservativeness of M, the hijacking of ARF1 for viral assembly and propagation could be a common mechanism employed by many coronaviruses, especially the sarbecoviruses, which merits further testing and validation. Likewise, as exemplified by our proof-of-concept studies, targeting ARF1 or the ARF1-M interaction may lead to broad-spectrum antiviral strategies against the evolving SARS-CoV-2 variants and other emerging coronaviruses in the future.
Co-expression of other coronaviral structural proteins with M generates VLPs driven by the M protein. When the S protein is present, it is incorporated into the VLPs, forming corona-like structures similar to those of authentic viruses16,61,62,63,77,78. In this study, we first employed a VLP reporter system using S protein C-terminally fused to the HiBiT tag (for S detection)59, which confirms the significant role of ARF1 in M-driven assembly. Extensive published literature has demonstrated that C-terminally tagged S proteins can be used for efficient production of VLPs, and moreover, these VLPs are widely applicable to studies not only on assembly, but also on entry, antibody, and vaccine development that all strictly require both S incorporation and functionality16,59,60,61,62,78,79,80,81,82,83. These studies consistently indicate that the C-terminal tag fusion does not significantly impair or disrupt VLP packaging or functionality. However, considering the potential role of the SARS-CoV-2 S protein’s C-terminus in regulating its trafficking64,84, more or less effects of tagging on S incorporation into VLPs (particularly with larger tags) might not be entirely ruled out. Although such effects (even if present) do not notably compromise the utility of VLPs as valuable research tools (as shown in numerous studies), we also validated our findings using an independent N-detecting VLP system with untagged S to further address the possible concern. This system yielded consistent results and conclusions, further supporting the critical role of ARF1 in promoting M-driven assembly.
ARF1 belongs to the Ras superfamily of small guanosine triphosphatases and is a regulator of vesicular traffic49,85,86,87. Active ARF1 in a GTP-bound conformation that is catalyzed by guanine nucleotide-exchange factors (GEFs) is anchored to the ERGIC/cis-Golgi membranes, while the interaction of inactive ARF1 form with membranes is much weaker, leading to diffusion to cytosol49,85. ARF1 can recruit soluble cytosolic effectors to the membranes, such as coat protein complex I (COPI) coatomer, clathrin cargo adapters, and lipid metabolism enzymes, to regulate the vesicular assembly or remodel the membranes88,89,90,91. Previous work has shown that ARF1 might be involved in multiple virus infections with unknown or several proposed mechanisms31,32,33,34,35,92,93,94,95,96, while whether and how ARF1 has a role in SARS-CoV-2 infection was unknown before our study. Particularly, Faure et al. have reported that the HIV-1 Nef protein can interact with both cluster of differentiation 4 (CD4) and ARF1, and hijack ARF1 onto the endosomal membranes to downregulate CD4 by accelerated endocytosis and lysosomal degradation31. Moreover, HIV-1 Nef can also downregulate the cell surface major histocompatibility class I (MHC-I) and tetherin (a viral restriction factor) by lysosomal degradation, which relies on ARF1 to unlock the adapter proteins required for endocytosis via clathrin-coated vesicles32,33. Additionally, the 3 A protein of Coxsackievirus B3 (CVB3) binds to ARF1 and GBF1 (a pivotal GEF of ARF1) and promotes preferential recruitment of phosphatidylinositol-4-kinase IIIb (PI4KIIIβ) to membranes over coat proteins34. The recruited PI4KIIIb then catalyzes the formation of an uncoated PI4P-enriched lipid microenvironment that is shown to be required for both CVB3 and hepatitis C virus (HCV) infection34. In the present study, we showed that SARS-CoV-2 M can interact with and hijack ARF1 for its efficient concentration to the ERGIC and thus the viral assembly, presenting a notable function/mechanism for the potential involvement of ARF1 in viral infections. The membrane anchoring of ARF1 can be strengthened by GBF153,97. Consistently, we also found that indeed, GBF1 KD could significantly impair SARS-CoV-2 propagation (Supplementary Fig. S14a, b), indicating that the cellular regulatory factors of ARF1 (known or unknown) might also be potential targets for future anti-SARS-CoV-2 research. However, interestingly, the PI4KIIIb seems not to be required for SARS-CoV-2 propagation (Supplementary Fig. S14c, d), reflecting the differential requirements of different viruses for their infection environment.
Of note, despite the potential role of ARF1 in vesicular traffic and cellular secretion, previous studies have found that deletion of ARF1 does not affect cellular transport/secretion homeostasis, due to functional overlap and redundancy among homologous molecules56,57. Consistently, our additional analyses also show that ARF1 KO seems not to affect cellular secretion and COPI trafficking/localization to ERGIC. Furthermore, previous studies have demonstrated that deletion of ARF1 does not nonspecifically cause significant inhibition against various viral infections58, despite its potential roles in some viruses as aforementioned. Additionally, we observed a dramatic reduction of assembled virions in cells with ARF1 KO, rather than intracellular retention of assembled virions that would likely be seen following disruption of secretion. These also rule out the possibility that the inhibition of virus proliferation under the condition of ARF1 deficiency is due to the nonspecific impact of globally disrupted secretion homeostasis. Consistent with these observations, in addition to not causing noticeable cytotoxicity, PEP17 (which specifically disrupts M but not ARF1 accumulation at ERGIC through interaction with M, thereby inhibiting virus proliferation) does not significantly impair cellular secretion either. The data further indicate that PEP17 specifically inhibits SARS-CoV-2 propagation by disrupting M-ARF1 interaction rather than causing cytotoxicity or impacting cellular secretion homeostasis.
ARF1 is also implicated in the progression and invasiveness of various human cancers, such as non-small cell lung cancer, colorectal cancer, breast cancer, epithelial ovarian cancer, and pancreatic adenocarcinoma, which has attracted great interest in developing ARF1-targeting intervention strategies for oncotherapy37,38,98,99,100,101. Consequently, dozens of small drugs have been identified as inhibitors against ARF1 and/or its interactors and hence as potential antitumor agents. In addition, by sequence-based AI virtual screening, Chang et al. recently identified a natural product demethylzeylasteral (DMZ), extracted from Tripterygium wilfordii Hook F, as a potent ARF1 inhibitor that can suppress tumor growth in a mouse model36. Interestingly, the same group then determined ARF1 as a target for anti-tumor drugs, proton pump inhibitors (PPIs), by an inverse application of the sequence-based computational drug screening pipeline39. These studies provide a significant pool of ARF1 inhibitors. As a proof-of-concept, we validated that two representative compounds, BFA and GCA, could indeed disrupt ARF1 localization at the ERGIC and thus deprive SARS-CoV-2 M of its ERGIC concentration, inhibiting the viral assembly and propagation. However, more existing agents with great clinical application potential are likely merited to be further tested in the future for re-purposing usage as possible host-directed broad-spectrum anti-SARS-CoV-2 drugs. Additionally, our loss-of-function assays with RNAi or CRISPR-Cas9 editing showed the blockade of SARS-CoV-2 propagation after ARF1 KD or KO, which supports the possibility of reducing ARF1 expression for anti-SARS-CoV-2 therapies, for instance, by targeted protein degradation (TPD). Moreover, based on the identification of the M-targeted ARF1 motif, ARF11–17, we designed a mimetic peptide that significantly blocked SARS-CoV-2 and its variants’ propagation and attenuated the viral pathogenicity in vivo. As another proof-of-concept, the peptide treatment targeting the M-ARF1 interaction interface presents an additional intervention avenue, which, mechanistically, is similar to the effects of the dominant negative mutant but different from those of the small molecule inhibitors or loss-of-function. Together, the dependence of SARS-CoV-2 and its variants on ARF1 likely represents a notable weak point that could be targeted by numerous strategies for developing broad-spectrum anti-SARS-CoV-2 therapies, meriting more studies. As ARF1 is conserved across mammals, including human, mice, and bats, it is likely a common requirement for SARS-CoV-2 infection and likewise a basis for interspecies transmission. Such interventions targeting ARF1 and/or the ARF1-M interface for veterinary use may also help to control the spread among animals or spillover to humans of SARS-CoV-2 or other related coronaviruses in the future.
In summary, we identified ARF1 as a host proviral factor for SARS-CoV-2 and its variants through a series of loss/gain-of-function analyses in cultured cells and animal models. By interaction with the viral M protein, ARF1 enables M accumulation at the ERGIC and hence M-driven viral assembly and propagation, bolstering the viral infection and pathogenicity. Multiple intervention experiments targeting ARF1 or the ARF1-M interaction interface, including the use of the peptidomimetic for in vitro and in vivo treatments, corroborate the function/mechanism model and present proof-of-concept illustrations for developing potential intervention strategies based on the current findings. Overall, the study not only provides critical insights into the fundamental biology of SARS-CoV-2 propagation but also may help expand our tactics to design new broad-spectrum anti-SARS-CoV-2 therapies.
Methods
Ethics and biosafety statement
All animal experiments have been approved by the Animal Ethics Committee of the Wuhan Institute of Virology, Chinese Academy of Sciences (approval numbers: WIVA42202004, WIVA17202005, and WIVA13202401) and performed in accordance with the guidelines for the Care and Use of Medical Laboratory Animals (Ministry of Health, China). All of the infection-related experiments were conducted with the biosafety level 3 facilities in the National Biosafety Laboratory, Wuhan, Chinese Academy of Sciences.
Cells, viruses, and animals
HEK293 (ATCC, CRL-1573), HeLa (ATCC, CCL-2), and Vero-E6 (ATCC, CCL-81) cell lines were maintained in minimum Eagle’s medium (MEM; Gibco) supplemented with 10% Fetal Bovine Serum (FBS; Gibco) and 1% Sodium Pyruvate (Gibco). HEK293T (ATCC, CRL-11268) and Huh7-ACE2 (Huh7 stably expressing hACE229,30) were cultured in Dulbecco’s modified Eagle’s medium (DMEM; Gibco) supplemented with 10% FBS. All of the cell lines were cultured at 37 °C with 5% CO2 atmosphere.
For generation of ARF1-KO HEK293 or HeLa cell lines by CRISPR-Cas9, the ARF1-specific guide RNA (gRNA) sequences (Supplementary Table S1) were designed by the online CRISPR Design tool (http://crispr.mit.edu/) and cloned into the sgRNA scaffold of pSpCas9 (BB)-2A-Puro (PX459) V2.0 vector (Addgene, 62988)102. The selection and validation of ARF1-KO HEK293 or HeLa cell lines were conducted through puromycin selection, single-cell cloning, DNA sequencing of the targeted locus (Primers are listed in Table S1), and Western blotting (WB) analysis using ARF1-specific antibodies42,103.
The prototypic SARS-CoV-2 WIV04 strain (IVCAS 6.7512)1 and its Delta (IVCAS 6.7585) and Omicron sublineage BA.4 (IVCAS 6.8980) and BA.5 (IVCAS 6.8981) variants were obtained from the National Virus Resource Center, Wuhan Institute of Virology, Chinese Academy of Sciences, and expanded in Vero-E6 cells29. Viral titer was determined by the 50% tissue culture infective dose (TCID50) method using cytopathic effect assay (CPE) and plaque-forming unit (PFU) counting as correspondingly indicated29.
Six- to eight-week-old female H11-K18-hACE2 transgenic mice (strain, C57BL/6JGpt-H11em1Cin(K18-ACE2)/Gpt) were purchased from GemPharmatech (Jiangsu, China). The Syrian hamsters (LVG Golden Hamsters, also known as Mesocricetus auratus; strain, Crl:LVG (SYR); 6-week-old, female) were purchased from Beijing Vital River Laboratory Animal Technology Co., Ltd. (Beijing, China). Animals were housed under controlled conditions: temperature maintained at 22–24 °C, relative humidity at 40–60 %, and a 12-h light/dark cycle.
Plasmids and siRNAs
The expression plasmids for C-terminally Flag-tagged ARF1 and ARF1(T31N) were constructed by cloning the coding sequences into the pCAGGSP7 plasmid. The plasmids expressing GFP-fused ARF1 and truncated mutants were constructed using the pEGFP-N3 vector. The open reading frame of ACE2 was cloned into the pLVX vector. The coding sequences for SARS-CoV-2 M, N, E, and S with or without C-terminally Flag, Stag, or HiBiT-tag were also cloned into the pCAGGSP7 plasmid. For ARF1 RNAi constructs, double-stranded oligonucleotides corresponding to specific target sequences (shmARF1 or the negative control shNC, Table S1) were cloned into the shRNA-expressing vector pLKO.1. All the cloning constructs were confirmed by DNA sequencing. Small interfering RNAs (siRNA) were designed and synthesized by GenePharma. The sequences of two specific ARF1 siRNAs and a nontargeting siRNA control were shown in Table S1.
Antibodies, peptides, and reagents
Primary mouse or rabbit monoclonal antibodies (mAb) include: mouse mAb to Flag (ABclonal, AE005), β-actin (Proteintech, 66009-1-Ig), and ERGIC-53 (Santa Cruz Biotechnology, sc-365158), and rabbit mAb to GFP tag (ABclonal, AE078) and SARS-CoV-2 N (ABclonal, A20021, for IHC analysis; Sino Biological, 40413-R019, for WB analysis). Primary polyclonal antibodies (pAb) include: rabbit pAb to Stag (Sino Biological, 101290-T38), ERGIC-53 (ABclonal, A10440), ARF1 (Proteintech, 10790-1-AP), and IgG control (Proteintech, 30000-0-AP), goat pAb to ARF1 (Abcam, ab58578), and sheep pAb to SARS-CoV-2 M (MRC PPU Reagents and Services, DA107). Fluorescence-labeled secondary antibodies used were Alexa Fluor 488, 555, or 647 labeled donkey anti-sheep, goat, rabbit, or mouse IgG (Thermo Fisher Scientific, A11015, A11055, A332794, A21447). Horseradish peroxidase (HRP)-labeled goat anti-rabbit (AS014) or mouse (AS003) IgG antibodies and donkey anti-mouse (AS033), rabbit (AS038), or goat (AS031) IgG were purchased from ABclonal. HRP-labeled donkey anti-sheep IgG was purchased from Thermo Fisher Scientific (A16041).
The PEP17 peptide (YGRKKRRQRRRGSGMGNIFANLFKGLFGKKE) is composed of ARF11–17 (MGNIFANLFKGLFGKKE), an N-terminal conjugate cell membrane-penetrating sequence (YGRKKRRQRRR), and a Gly-Ser-Gly linker68,104. PEP17, Myr-PEP17 (Myr-GNIFANLFKGLFGKKEGSGYGRKKRRQRRR), and the control CPS14 (YGRKKRRQRRRGSG), consisting of the cell membrane-penetrating sequence and linker, were synthesized by GenScript Biochem. The synthetic peptides were amidated at the C terminus, and PEP17 and CPS14 were acetylated at the N terminus for capping. They were then dissolved in PBS at a concentration of 2 mM as stock solutions.
BFA (MedChemExpress, HY-16592), GCA (MedChemExpress, HY-100540), Lipofectamine 2000/3000/RNAiMAX reagents (Invitrogen, 11668019, L3000015, 13778150), Trizol (Invitrogen, 15596018), Stag-pulldown agarose (Merck Millipore, 69704-4), GFP-trap beads (AlpaLife, KTSM1301), Hoechst 33258 (Beyotime, C1011), MiniBEST Viral RNA Extraction Kit (Takara, RR047A), and PrimeScript RT reagent Kit with gDNA Eraser (Takara, RR047A) and TB Green Realtime PCR Master Mix (Takara, RR820A) were purchased from the indicated manufacturers.
Protein interaction, MS, and WB analyses
Stag-pulldown, Co-IP, and GFP-nanotrap assays were used for the protein interaction analyses29,42. Briefly, cells expressing the indicated proteins were lysed with a lysis buffer (20 mM Tris, pH 7.5, 150 mM NaCl, 1 mM EDTA, 1% Triton X-100; Beyotime, P0013) supplemented with protease inhibitor cocktail (Roche, 05892791001). After centrifugation, supernatants of the cell lysates were incubated with the Stag-trap agarose or the anti-GFP nanobody-coated beads at 4 °C for 4 h. For Co-IP, protein A/G magnetic beads (MedChemExpress, HY-K0202) were pretreated with the anti-ARF1 antibody or control IgG and then mixed with the cell lysate supernatants at 4 °C for 4 h. After extensive washing, the binding proteins were eluted from the beads by boiling in 1× SDS-loading buffer, followed by SDS-PAGE and WB analysis. For MS analysis, Stag-pulldown products of M and vector (control) groups (n = 3 biological replicates per group) were denatured, reduced, alkylated, and eluted in the reaction buffer (1% sodium deoxycholate/100 mM Tris-HCl, pH 8.5/10 mM Tris (2-carboxyethyl) phosphine/40 mM chloroacetamide) at 95 °C for 10 min. The supernatants were then subjected to trypsin digestion in solution, followed by peptide desalting and liquid chromatography-tandem MS (LC-MS/MS) analysis using the nano-LC-equipped TripleTOF 5600 system (AB SCIEX)29,30,43. Chromatographic separation was performed on a C18 analytical column (Eksigent; 300 μm × 150 mm, 3 μm) with a 30-min gradient (5 μL/min; mobile phase A: 3% DMSO/0.1% FA in H₂O; B: 3% DMSO/0.1% FA in acetonitrile), ramping from 6% to 80% phase B. MS data were acquired in data-dependent mode (ESI+, 2.3 kV spray voltage) with one MS1 scan (m/z 350–1500, 250 ms) followed by 40 MS2 scans (m/z 100–1500, 50 ms; dynamic exclusion: 18 s). Spectrum files were processed in MaxQuant 1.6.2.10 (Max Planck Institute of Biochemistry) against UniProt database, allowing tryptic digestion (≤2 missed cleavages), carbamidomethylation (fixed), methionine oxidation (variable), 20 ppm/20 ppm mass tolerances (MS1/MS2), and 1% false discovery rate (FDR)29,30,43. ARF1 peptides were detected in all biological replicates of the M group but were not absent in the control samples, and the representative tandem spectra are shown in Figure S1. For WB detection, cell or homogenized tissue samples were mixed with SDS loading buffer and then boiled for 10 min followed by SDS-PAGE separation. Then, the proteins in the gel were transferred to a polyvinylidene difluoride (PVDF) membrane (Millipore). After blocking with 5% skimmed milk, the PVDF membranes were treated with primary antibodies and then with corresponding HRP-conjugated secondary antibodies. The protein signal was detected by using a chemiluminescence kit (Millipore) on BG-gdsAUTO 710 mini (Baygene Biotech Company Limited).
IFA and confocal microscopy
Cells were fixed with 4% paraformaldehyde (PFA) for 10 min at room temperature (RT), permeabilized with 0.5% Triton X-100 for 10 min at 4 °C, and blocked with 5% BSA for 1 h at RT. Then, cells were incubated with primary antibody overnight at 4 °C, and stained with the fluorescein-labeled secondary antibodies for 1 h at RT. Nuclei of cells were stained with Hoechst 33258 for 5 min at RT. Images were taken on Dragonfly 202 confocal microscopy (Andor Technology PLC), and analyzed by Image J 2.0.0 (National Institutes of Health, http://imagej.nih.gov/ij). For the co-localization assay, Pearson’s correlation coefficient (PCC) was calculated using the coloc-2 plugin of ImageJ43.
Transmission electron microscopy assay
WT or ARF1-KO HEK293 cells were transfected with the hACE2 expression plasmid for 24 h, followed by SARS-CoV-2 infection (MOI = 3) for another 24 h. Cells were then fixed with 2.5% glutaraldehyde for 30 min, and subsequently with 4% paraformaldehyde for 2 h at room temperature. After collection, cells were washed in PBS at 4 °C and post-fixed with 1% osmium tetroxide for 2–3 h on ice under dark conditions. Following PBS washes, the cells were dehydrated through a graded ethanol series (30%, 50%, 75%, 95%, and 100%) and then processed for resin infiltration using acetone/epoxy resin (2:1, 1:1) mixtures before final embedding in pure epoxy resin. Ultrathin sections (80–100 nm) were then prepared and stained with uranyl acetate and lead citrate, and examined using transmission electron microscopy (Tecnai G2 20 TWIN, FEI).
RNA extraction and real-time quantitative reverse-transcription PCR (qRT-PCR)
The host gene mRNA and viral genomic or N RNA levels were assessed by qRT-PCR analyses29,105. Briefly, total cellular RNA, viral genome in cell supernatants, and RNA from homogenized tissue suspension were isolated by using TRIzol (Invitrogen, 15596018), viral RNA/DNA Extraction kit (TaKaRa, 9766), or TRIzol LS (Invitrogen, 10296028CN), respectively. The RNAs were then subjected to gDNA remover and reverse transcription for cDNA synthesis. The cDNA was used as template in quantitative PCR (qPCR) containing SYBR Green Realtime PCR Master Mix and specific primers. The real-time qPCRs were run on CFX Connect (Bio-Rad) using the standard cycling conditions. Relative mRNA levels were calculated by the comparative CT method with GAPDH mRNA level as the internal control. Alternatively, viral RNA copies were calculated by the standard curve method as previously described29. Primers for qPCR are listed in Table S1.
Quantification of the virion-like particles (VLPs)
HiBiT-tagged VLP reporter systems were employed for evaluation of M-driven viral assembly59,60,61,62,63,64. Briefly, the SARS-CoV-2 M, E, N, and S-HiBiT encoding plasmids (ratio 1:1:1:1) for the S-tagging VLP system were transfected into HEK293T or HEK293 cells using the Lipofectamine 2000 reagent (Invitrogen, 11668019). In the N-tagging VLP system, HiBiT-tagged N (N-HiBiT) and untagged S were used to replace the N and S-HiBiT. The supernatants containing HiBiT-labeled VLPs were collected for luminescence signal analysis using Nano-Glo HiBiT Extracellular Detection reagents (Promega, N2420) on Glomax Multi (Promega).
ARF1 KD and SARS-CoV-2 infection in K18-hACE2 mice
For the production of the viral vectors, HEK293T cells were co-transfected with the pLKO.1-based plasmids encoding ARF1-targeting or control shRNAs, together with the packaging plasmids (psPAX2 and pMD2.G)42,43. At 48 h and 72 hpt, the cell culture media were harvested and cleared by centrifugation, followed by concentration using Lenti-X Concentrator (Takara, 631231) and resuspension in PBS. Viral vector titers were measured by using Quick Titer Lentivirus Titer Kit (CELL BIOLABS, INC., VPK-107-T).
For evaluation of SARS-CoV-2 infection in ARF1-KD K18-hACE2 mice, K18-hACE2 mice (6–8-weeks-old; n = 5/group) were injected with 5 × 107 transducing units (TU) of the viral vector particles via intravenous injection (iv). Seven days post-transduction, mice were intranasally instilled with a liquid SARS-CoV-2 suspension (5 × 102 TCID50, 50 μL inoculum per mouse) under deep anesthesia. At 5 dpi, mice were sacrificed for tissue sample collection and the subsequent virological and histopathological analyses.
SARS-CoV-2 challenge and PEP17 administration in Syrian hamsters
Six-week-old female Syrian hamsters were randomly divided into the PEP17 and CPS14 control groups (n = 5/group). At day 0, all hamsters were challenged with SARS-CoV-2 via intranasal routes (104 TCID50, 50 μL inoculum per hamster). Two hours post-infection, the animals were treated with PEP17 or the control peptide (CPS14) via intraperitoneal injection (dissolved in PBS, 30 mg/kg) once daily for three days. At day 3, the hamsters were sacrificed. The brain, trachea, and lung tissues were collected for the subsequent virological and histopathological analyses.
Histological analysis
Tissues were adequately fixed in 4% PFA and then embedded in paraffin for sectioning106. The sections were delivered to hematoxylin and eosin (H&E) staining to assess tissue morphology or to an IHC assay for the viral antigen (N) detection. Slides were scanned by PANNORAMIC DESK/MIDI/250/1000 (3Dhistech Pannoramic Scan, Hungary) and viewed by CaseViewer 2.4 (3Dhistech Pannoramic Scan, Hungary).
CCK8 assay
The potential cytotoxicity of BFA, GCA, and peptides on Huh7-ACE2, HeLa, or HEK293T cells was examined with the CCK-8 assay kit (MedChemExpress, HY-K0301) according to the manufacturer’s instructions. In brief, cells plated on 96-well culture plates were treated with different concentrations of the inhibitors or peptides, followed by adding CCK-8 solution (1/10 dilution) into the cells. After incubation for 1 h, the culture plates were delivered for formazan detection at 450 nm absorbance on a microplate reader (BioTek Synergy). Cell viability (%) = [(As − Ab)/(Ac − Ab)] × 100. As = absorbance of the tested drug/peptide well, Ab = absorbance of blank well, Ac = absorbance of solvent control well.
Statistical analysis
Graphs were drawn using Prism 9 (GraphPad Software). Data are shown as means ± SD or medians with interquartile ranges for non-parametric data, with n ≥ 3 biological replicates as indicated. Statistical analyses were performed using Prism 9 with Student’s t-test for the two-group comparisons or one-way analysis of variance (ANOVA) for the multiple comparisons, followed by Dunnett’s test. For the non-normal distribution data, Mann–Whitney U-tests for the two-group comparisons or Kruskal–Wallis H-tests for the multiple comparisons, followed by Dunn’s Q-test, were used. Detailed statistical results with p-values are provided in the Source data file.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The MS proteomics data generated in this study have been deposited in the Science Data Bank (CSTR: 31253.11.sciencedb.25947; https://www.scidb.cn/en/s/Yvmeaq) and in the ProteomeXchange Consortium via the iProX partner repository with the dataset identifier PXD065198 (https://www.iprox.cn/page/MSV022.html). Other data are all contained within the article/Supplementary information. Source data are provided with this paper.
References
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270 (2020).
Li, G. D., Hilgenfeld, R., Whitley, R. & De Clercq, E. Therapeutic strategies for COVID-19: progress and lessons learned. Nat. Rev. Drug Discov. 22, 449–475 (2023).
Carabelli, A. M. et al. SARS-CoV-2 variant biology: immune escape, transmission and fitness. Nat. Rev. Microbiol. 21, 162–177 (2023).
Markov, P. V. et al. The evolution of SARS-CoV-2. Nat. Rev. Microbiol. 21, 361–379 (2023).
Finkel, Y. et al. The coding capacity of SARS-CoV-2. Nature 589, 125–U254 (2021).
Jackson, C. B., Farzan, M., Chen, B. & Choe, H. Mechanisms of SARS-CoV-2 entry into cells. Nat. Rev. Mol. Cell Biol. 23, 3–20 (2022).
Yan, L. et al. Cryo-EM structure of an extended SARS-CoV-2 replication and transcription complex reveals an intermediate state in Cap synthesis. Cell 184, 184–193 e110 (2021).
Sabbah, D. A., Hajjo, R., Bardaweel, S. K. & Zhong, H. A. An Updated review on SARS-CoV-2 main proteinase (M): protein structure and small-molecule inhibitors. Curr. Top. Med Chem. 21, 442–460 (2021).
Schultze, J. L. & Aschenbrenner, A. C. COVID-19 and the human innate immune system. Cell 184, 1671–1692 (2021).
Min, Y. Q. et al. Immune evasion of SARS-CoV-2 from interferon antiviral system. Comput Struct. Biotechnol. J. 19, 4217–4225 (2021).
Lowery, S. A., Sariol, A. & Perlman, S. Innate immune and inflammatory responses to SARS-CoV-2: implications for COVID-19. Cell Host Microbe 29, 1052–1062 (2021).
Krijnse-Locker, J., Ericsson, M., Rottier, P. J. & Griffiths, G. Characterization of the budding compartment of mouse hepatitis virus: evidence that transport from the RER to the Golgi complex requires only one vesicular transport step. J. Cell Biol. 124, 55–70 (1994).
de Haan, C. A. M. & Rottier, P. J. M. Molecular interactions in the assembly of coronaviruses. Adv. Virus Res. 64, 165–230 (2005).
Klumperman, J. et al. Coronavirus M-proteins accumulate in the Golgi-complex beyond the site of virion budding. J. Virol. 68, 6523–6534 (1994).
Stertz, S. et al. The intracellular sites of early replication and budding of SARS-coronavirus. Virology 361, 304–315 (2007).
Boson, B. et al. The SARS-CoV-2 envelope and membrane proteins modulate maturation and retention of the spike protein, allowing assembly of virus-like particles. J. Biol. Chem. 296, 100111 (2021).
Neuman, B. W. et al. A structural analysis of M protein in coronavirus assembly and morphology. J. Struct. Biol. 174, 11–22 (2011).
Voss, D. et al. Studies on membrane topology, N-glycosylation and functionality of SARS-CoV membrane protein. Virol. J. 6, 79 (2009).
Fehr, A. R. & Perlman, S. Coronaviruses: an overview of their replication and pathogenesis. Methods Mol. Biol. 1282, 1–23 (2015).
Dolan, K. A. et al. Structure of SARS-CoV-2 M protein in lipid nanodiscs. Elife. https://doi.org/10.7554/eLife.81702 (2022).
Zhang, Z. K. et al. Structure of SARS-CoV-2 membrane protein essential for virus assembly. Nat. Commun.https://doi.org/10.1038/s41467-022-32019-3 (2022).
Siu, Y. L. et al. The M, E, and N structural proteins of the severe acute respiratory syndrome coronavirus are required for efficient assembly, trafficking, and release of virus-like particles. J. Virol. 82, 11318–11330 (2008).
Tseng, Y. T. et al. Self-assembly of severe acute respiratory syndrome coronavirus membrane protein. J. Biol. Chem. 285, 12862–12872 (2010).
Narayanan, K., Chen, C. J., Maeda, J. & Makino, S. Nucleocapsid-independent specific viral RNA packaging via viral envelope protein and viral RNA signal. J. Virol. 77, 2922–2927 (2003).
Nabeel-Shah, S. et al. SARS-CoV-2 nucleocapsid protein binds host mRNAs and attenuates stress granules to impair host stress response. Iscience 25, 103562 (2022).
Stukalov, A. et al. Multilevel proteomics reveals host perturbations by SARS-CoV-2 and SARS-CoV. Nature 594, 246–252 (2021).
Li, J. et al. Virus-host interactome and proteomic survey reveal potential virulence factors influencing SARS-CoV-2 pathogenesis. Medicine. https://doi.org/10.1016/j.medj.2020.07.002 (2021).
Gordon, D. E. et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583, 459–468 (2020).
Min, Y. Q. et al. A new cellular interactome of SARS-CoV-2 nucleocapsid protein and its biological implications. Mol. Cell Proteomics. https://doi.org/10.1016/j.mcpro.2023.100579 (2023).
Feng, K. et al. Interactome profiling reveals interaction of SARS-CoV-2 NSP13 with host factor STAT1 to suppress interferon signaling. J. Mol. Cell Biol. https://doi.org/10.1093/jmcb/mjab068 (2021).
Faure, J. et al. ARF1 regulates Nef-induced CD4 degradation. Curr. Biol. 14, 1056–1064 (2004).
Shen, Q. T., Ren, X., Zhang, R., Lee, I. H. & Hurley, J. H. HIV-1 Nef hijacks clathrin coats by stabilizing AP-1:Arf1 polygons. Science 350, aac5137 (2015).
Morris, K. L. et al. HIV-1 Nefs are cargo-sensitive AP-1 trimerization switches in tetherin downregulation. Cell 174, 659–671 e614 (2018).
Hsu, N. Y. et al. Viral Reorganization of the Secretory Pathway Generates Distinct Organelles for RNA Replication. Cell 141, 799–811 (2010).
Hansen, M. D. et al. Hepatitis C virus triggers Golgi fragmentation and autophagy through the immunity-related GTPase M. Proc. Natl. Acad. Sci. USA 114, E3462–E3471 (2017).
Chang, J. et al. Discovery of ARF1-targeting inhibitor demethylzeylasteral as a potential agent against breast cancer. Acta Pharm. Sin. B 12, 2619–2622 (2022).
Ma, H. D., Fang, W. Q., Li, Q. M., Wang, Y. T. & Hou, S. X. Arf1 Ablation in colorectal cancer cells activates a super signal complex in DC to enhance anti-tumor immunity. Adv Sci. https://doi.org/10.1002/advs.202305089 (2023).
Wang, G. H. et al. Arf1-mediated lipid metabolism sustains cancer cells and its ablation induces anti-tumor immune responses in mice. Nat. Commun. https://doi.org/10.1038/s41467-019-14046-9 (2020).
Chen, L. F. et al. Sequence-based drug design as a concept in computational drug design. Nat. Commun. https://doi.org/10.1038/s41467-023-39856-w (2023).
Marques-Pereira, C. et al. SARS-CoV-2 membrane protein: from genomic data to structural new insights. Int. J. Mol. Sci. https://doi.org/10.3390/ijms23062986 (2022).
Hayakawa, N. et al. The ADP-ribosylation factor 1 gene is indispensable for mouse embryonic development after implantation. Biochem. Biophs. Res. Co. 453, 748–753 (2014).
Mo, Q., Xu, Z., Deng, F., Wang, H. L. & Ning, Y. J. Host restriction of emerging high-pathogenic bunyaviruses via MOV10 by targeting viral nucleoprotein and blocking ribonucleoprotein assembly. PLoS Pathog. https://doi.org/10.1371/journal.ppat.1009129 (2020).
Dai, S. et al. Interactome profiling of Crimean-Congo hemorrhagic fever virus glycoproteins. Nat. Commun. 14, 7365 (2023).
Dong, W. J. et al. The K18-human ACE2 transgenic mouse model recapitulates non-severe and severe COVID-19 in response to an infectious dose of the SARS-CoV-2 virus. J. Virol. https://doi.org/10.1128/JVI.00964-21 (2022).
Winkler, E. S. et al. SARS-CoV-2 infection of human ACE2-transgenic mice causes severe lung inflammation and impaired function (vol 51, pg 831, 2020). Nat. Immunol. 21, 1470–1470 (2020).
Zheng, J. et al. COVID-19 treatments and pathogenesis including anosmia in K18-hACE2 mice. Nature 589, 603 (2021).
Fumagalli, V. et al. Administration of aerosolized SARS-CoV-2 to K18-hACE2 mice uncouples respiratory infection from fatal neuroinvasion. Sci. Immunol. 7, eabl9929 (2022).
Da Costa, C. B. P. et al. Using in vivo animal models for studying SARS-CoV-2. Expert Opin. Drug Dis. 17, 121–137 (2022).
Donaldson, J. G. & Jackson, C. L. ARF family G proteins and their regulators: roles in membrane transport, development and disease. Nat. Rev. Mol. Cell Biol. 12, 362–375 (2011).
Chun, J., Shapovalova, Z., Dejgaard, S. Y., Presley, J. F. & Melancon, P. Characterization of class I and II ADP-ribosylation factors (Arfs) in live cells: GDP-bound class II Arfs associate with the ER-Golgi intermediate compartment independently of GBF1. Mol. Biol. Cell 19, 3488–3500 (2008).
Dascher, C. & Balch, W. E. Dominant inhibitory mutants of Arf1 block endoplasmic-reticulum to Golgi transport and trigger disassembly of the Golgi-apparatus. J. Biol. Chem. 269, 1437–1448 (1994).
Liu, Y. Z., Kahn, R. A. & Prestegard, J. H. Structure and membrane interaction of myristoylated ARF1. Structure 17, 79–87 (2009).
Muccini, A. J., Gustafson, M. A. & Fromme, J. C. Structural basis for activation of Arf1 at the Golgi complex. Cell Rep. https://doi.org/10.1016/j.celrep.2022.111282 (2022).
Hirschenberger, M. et al. ARF1 prevents aberrant type I interferon induction by regulating STING activation and recycling. Nat. Commun. 14, 6770 (2023).
Appenzeller-Herzog, C. & Hauri, H. P. The ER-Golgi intermediate compartment (ERGIC): in search of its identity and function. J. Cell Sci. 119, 2173–2183 (2006).
Volpicelli-Daley, L. A., Li, Y., Zhang, C. J. & Kahn, R. A. Isoform-selective effects of the depletion of ADP-ribosylation factors 1-5 on membrane traffic. Mol. Biol. Cell 16, 4495–4508 (2005).
Farhat, R. et al. Identification of class II ADP-ribosylation factors as cellular factors required for hepatitis C virus replication. Cell Microbiol 18, 1121–1133 (2016).
Ferlin, J. et al. Investigation of the role of GBF1 in the replication of positive-sense single-stranded RNA viruses. J. Gen. Virol. 99, 1086–1096 (2018).
Miura, K. et al. Distinct motifs in the E protein are required for SARS-CoV-2 virus particle formation and lysosomal deacidification in host cells. J. Virol. https://doi.org/10.1128/jvi.00426-23 (2023).
Plescia, C. B. et al. SARS-CoV-2 viral budding and entry can be modeled using BSL-2 level virus-like particles. J. Biol. Chem. https://doi.org/10.1074/jbc.RA120.016148 (2021).
Xu, R. D., Shi, M. F., Li, J., Song, P. & Li, N. Construction of SARS-CoV-2 virus-like particles by mammalian expression system. Front. Bioeng. Biotech. https://doi.org/10.3389/fbioe.2020.00862 (2020).
Yang, Q. et al. Role for glycans and calnexin-calreticulin chaperones in SARS-CoV-2 Spike maturation and viral infectivity. Sci. Adv. https://doi.org/10.1126/sciadv.abq8678 (2022).
Kumar, B. et al. Assembly and Entry Of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV2): Evaluation Using Virus-like Particles. Cells. https://doi.org/10.3390/cells10040853 (2021).
Dey, D. et al. A single C-terminal residue controls SARS-CoV-2 spike trafficking and incorporation into VLPs. Nat. Commun. https://doi.org/10.1038/s41467-023-44076-3 (2023).
Vasudevan, C. et al. The distribution and translocation of the G protein ADP-ribosylation factor 1 in live cells is determined by its GTPase activity. J. Cell Sci. 111, 1277–1285 (1998).
Saenz, J. B. et al. Golgicide A reveals essential roles for GBF1 in Golgi assembly and function. Nat. Chem. Biol. 5, 157–165 (2009).
van der Linden, L., van der Schaar, H. M., Lanke, K. H., Neyts, J. & van Kuppeveld, F. J. Differential effects of the putative GBF1 inhibitors Golgicide A and AG1478 on enterovirus replication. J. Virol. 84, 7535–7542 (2010).
Guidotti, G., Brambilla, L. & Rossi, D. Cell-penetrating peptides: from basic research to clinics. Trends Pharm. Sci. 38, 406–424 (2017).
Fang, Y. et al. Antiviral peptides targeting the helicase activity of Enterovirus nonstructural protein 2C. J. Virol. https://doi.org/10.1128/JVI.02324-20 (2021).
Zhang, Y. et al. Myr-Arf1 conformational flexibility at the membrane surface sheds light on the interactions with ArfGAP ASAP1. Nat. Commun. https://doi.org/10.1038/s41467-023-43008-5 (2023).
Johnson, D. R., Bhatnagar, R. S., Knoll, L. J. & Gordon, J. I. Genetic and biochemical-studies of protein N-myristoylation. Annu. Rev. Biochem. 63, 869–914 (1994).
Beauchamp, E. et al. Targeting N-myristoylation for therapy of B-cell lymphomas. Nat. Commun. https://doi.org/10.1038/s41467-020-18998-1 (2020).
Imai, M. et al. Syrian hamsters as a small animal model for SARS-CoV-2 infection and countermeasure development. Proc. Natl. Acad. Sci. USA 117, 16587–16595 (2020).
Sia, S. F. et al. Pathogenesis and transmission of SARS-CoV-2 in golden hamsters. Nature 583, 834 (2020).
Chan, J. F. W. et al. Simulation of the clinical and pathological manifestations of coronavirus disease 2019 (COVID-19) in a Golden Syrian Hamster model: implications for disease pathogenesis and transmissibility. Clin. Infect. Dis. 71, 2428–2446 (2020).
Chu, H. et al. Coronaviruses exploit a host cysteine-aspartic protease for replication. Nature 609, 785 (2022).
Gourdelier, M. et al. Optimized production and fluorescent labeling of SARS-CoV-2 virus-like particles. Sci. Rep. 12, https://doi.org/10.1038/s41598-022-18681-z (2022).
Yuan, Z. et al. The E3 ubiquitin ligase RNF5 facilitates SARS-CoV-2 membrane protein-mediated virion release. mBio 13, e0316821 (2021).
Naskalska, A. et al. Functional severe acute respiratory syndrome Coronavirus 2 virus-like particles from insect cells. Front. Microbiol. https://doi.org/10.3389/fmicb.2021.732998 (2021).
Yilmaz, I. C. et al. Development and preclinical evaluation of virus-like particle vaccine against COVID-19 infection. Allergy 77, 258–270 (2022).
Li, Y. H. et al. A substitution at the cytoplasmic tail of the spike protein enhances SARS-CoV-2 infectivity and immunogenicity. Ebiomedicine. https://doi.org/10.1016/j.ebiom.2024.105437 (2024).
Gullberg, R. C. & Frydman, J. Novel mode of nanoluciferase packaging in SARS-CoV-2 virions and VLPs provides versatile reporters for virus production. Viruses. https://doi.org/10.3390/v15061335 (2023).
Jaron, M. et al. Baculovirus-free SARS-CoV-2 virus-like particle production in insect cells for rapid neutralization assessment. Viruses. https://doi.org/10.3390/v14102087 (2022).
Sultana, R. & Stahelin, R. V. Strengths and limitations of SARS-CoV-2 virus-like particle systems. Virology. https://doi.org/10.1016/j.virol.2024.110285 (2025).
D’Souza-Schorey, C. & Chavrier, P. ARF proteins: roles in membrane traffic and beyond. Nat. Rev. Mol. Cell Biol. 7, 347–358 (2006).
Kahn, R. A. et al. Arf family GTPases: roles in membrane traffic and microtubule dynamics. Biochem. Soc. T 33, 1269–1272 (2005).
Thomas, L. L., Highland, C. M. & Fromme, J. C. Arf1 orchestrates Rab GTPase conversion at the trans-Golgi network. Mol. Biol. Cell 32, 1104–1120 (2021).
Donaldson, J. G. & Jackson, C. L. Regulators and effectors of the ARF GTPases. Curr. Opin. Cell Biol. 12, 475–482 (2000).
Gamara, J., Chouinard, F., Davis, L., Aoudjit, F. & Bourgoin, S. G. Regulators and effectors of Arf GTPases in neutrophils. J. Immunol. Res. 2015, 235170 (2015).
Shin, H. W. & Nakayama, K. Dual control of membrane targeting by PtdIns(4)P and ARF. Trends Biochem. Sci. 29, 513–515 (2004).
Pelham, H. R. B. & Rothman, J. E. The debate about transport in the Golgi—Two sides of the same coin?. Cell 102, 713–719 (2000).
Banerjee, S. et al. Hijacking of multiple phospholipid biosynthetic pathways and induction of membrane biogenesis by a picornaviral 3CD protein. PLoS Pathog. 14, e1007086 (2018).
Lanke, K. H. W. et al. GBF1, a guanine nucleotide exchange factor for Arf, is crucial for Coxsackievirus B3 RNA replication. J. Virol. 83, 11940–11949 (2009).
Gimenez, M. C. et al. Rab1b-GBF1-ARF1 secretory pathway axis is required for birnavirus replication. J. Virol. 96, e0200521 (2022).
Iglesias, N. G. et al. Dengue virus uses a non-canonical function of the host GBF1-Arf-COPI system for capsid protein accumulation on lipid droplets. Traffic 16, 962–977 (2015).
Zhang, L. et al. ARF1 with Sec7 domain-dependent GBF1 activates coatomer protein I To support classical swine fever virus entry. J. Virol. 96, e0219321 (2022).
Kaczmarek, B., Verbavatz, J. M. & Jackson, C. L. GBF1 and Arf1 function in vesicular trafficking, lipid homoeostasis and organelle dynamics. Biol. Cell 109, 391–399 (2017).
Jiang, Z. B. et al. Plumbagin suppresses non-small cell lung cancer progression through downregulating ARF1 and by elevating CD8 T cells. Pharmacol. Res. https://doi.org/10.1016/j.phrs.2021.105656 (2021).
Schlienger, S., Campbell, S. & Claing, A. ARF1 regulates the Rho/MLC pathway to control EGF-dependent breast cancer cell invasion. Mol. Biol. Cell 25, 17–29 (2014).
Zhao, L. G., Wang, J., Li, J. & Li, Q. F. miR-744-5p inhibits cellular proliferation and invasion via targeting ARF1 in epithelial ovarian cancer. Kaohsiung J. Med. Sci. 36, 799–807 (2020).
Cheng, J., Lou, Y. H. & Jiang, K. Downregulation of long non-coding RNA LINC00460 inhibits the proliferation, migration and invasion, and promotes apoptosis of pancreatic cancer cells via modulation of the miR-320b/ARF1 axis. Bioengineered 12, 96–107 (2021).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Chang, M. et al. Host factor MxA restricts Dabie bandavirus infection by targeting the viral NP protein to inhibit NP-RdRp interaction and ribonucleoprotein activity. J. Virol. https://doi.org/10.1128/jvi.01568-23 (2023).
Chen, X. Y., Zaro, J. L. & Shen, W. C. Fusion protein linkers: property, design and functionality. Adv. Drug Deliv. Rev. 65, 1357–1369 (2013).
Min, Y. Q., Ning, Y. J., Wang, H. & Deng, F. A RIG-I-like receptor directs antiviral responses to a bunyavirus and is antagonized by virus-induced blockade of TRIM25-mediated ubiquitination. J. Biol. Chem. 295, 9691–9711 (2020).
Ning, Y. J. et al. Ebola virus mucin-like glycoprotein (Emuc) induces remarkable acute inflammation and tissue injury: evidence for Emuc pathogenicity in vivo. Protein Cell 9, 389–393 (2018).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2022YFC2303300, Y.-J.N.), the National Natural Science Foundation of China (grant number 32170171, Y.-J.N.), the Youth Innovation Promotion Association of Chinese Academy of Sciences (2020333, Y.-J.N.), and the Guangzhou Laboratory (EKPG21-30-2, X.S. and Y.-Q.M). We thank the running team of the National Biosafety Laboratory and the Institutional Center for Shared Technologies and Facilities of Wuhan Institute of Virology, Chinese Academy of Sciences, for technical assistance. We also would like to thank all team members of the following institutions for providing SARS-CoV-2 and the variants: National Virus Resource Center, Wuhan Institute of Virology; Institute of Laboratory Animals Science, Chinese Academy of Medical Sciences and Peking Union Medical College; National Pathogen Resource center, Chinese Center for Disease Control and Prevention, Beijing, China.
Author information
Authors and Affiliations
Contributions
X.S. and Y.-Q.M. supervised the project. Y.-J.N. and Y.-Q.M. conceived the study. Y.-Q.M., C.Z. and Y.-J.N. designed the experiments. C.Z. and Y.-Q.M. performed the majority of the experiments and acquired the data. H.X., H.Z., K.L., Y.T., Z.Y., Z.Z., H.Y. and C.S. provided experimental assistance. C.Z., Y.-Q.M., Y.-J.N. and X.S. analyzed and visualized the data. Y.-Q.M., Y.-J.N. and C.Z. wrote the original draft. Y.-J.N., Y.-Q.M. and X.S. reviewed and edited the manuscript. X.S., Y.-Q.M. and Y.-J.N. provided funding. All authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Y.-Q.M., C.Z., X.S. and Y.-J.N. have filed a patent application for the antiviral PEP17 (CN116478301A) as inventors, while all other authors declare that they have no conflicts of interest with the contents of this article.
Peer review
Peer review information
Nature Communications thanks Parikshit Bagchi and the other, anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, C., Min, YQ., Xue, H. et al. Host protein ARF1 is a proviral factor for SARS-CoV-2 and a candidate broad-spectrum therapeutic target. Nat Commun 16, 6326 (2025). https://doi.org/10.1038/s41467-025-61431-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61431-8