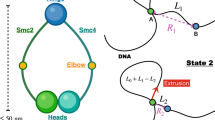
Abstract
Condensin I and topoisomerase IIα (topo IIα) are chromosomal ATPases essential for mitotic chromosome assembly. Mechanistically how the two ATPases cooperate to assemble mitotic chromosomes remains unknown. Here we investigate the interplay between condensin I and topo IIα at single-molecule resolution. While condensin I alone exhibits ATP-dependent DNA loop formation, it generates stable, compact structures (“lumps”) in the presence of topo IIα in a manner dependent on its C-terminal domain. These lumps predominantly contain a single condensin I complex and a single topo IIα dimer. The strand passage activity of topo IIα introduces DNA knots within the lumps, rendering them resistant to protease treatment. An ATP hydrolysis-deficient mutant of condensin I forms smaller lumps, in which the probability of DNA knotting is markedly reduced. Our findings demonstrate that topo IIα-mediated strand passage is functionally coupled with condensin I-mediated loop extrusion, providing insights into the mechanism underlying mitotic chromosome assembly.
Similar content being viewed by others
Introduction
Mitotic chromosome assembly is a fundamental cellular process that ensures the faithful segregation of genetic information during the mitotic cell cycle. Extensive studies over the past decades led to a consensus that two distinct classes of ATPases, condensins and topoisomerase IIα (topo IIα), play central roles in this process1,2. For example, the chromosome scaffold, originally characterized as a proteinaceous structure observed in histone-depleted metaphase chromosomes, was later found to contain topo IIα3,4 and the subunits of condensin I5,6. More recent studies have successfully reconstituted the core reaction of this process in vitro by mixing a simple substrate with only six purified proteins, including core histones, three histone chaperones, topo IIα, and condensin I7,8.
Eukaryotic topo IIα, which belongs to the type IIA topoisomerase family, introduces a double-strand break in one DNA strand and passes through a second DNA strand before rejoining the break in an ATP hydrolysis-dependent manner9,10,11. This strand passage reaction enables intermolecular catenation/decatenation and intramolecular knotting/unknotting of circular DNA in vitro (Fig. 1a). The direction of the reactions (i.e., topological simplification vs. topological complication) depends on the topological states of the DNA substrates and the reaction environment. Such enzymology of topo IIα is reasonably well understood.
a The strand passage activity of topo IIα catalyzes the decatenation of catenated circular DNA or the catenation of free circular DNA. It also catalyzes the knotting of free circular DNA or the unknotting of knotted DNA. b At an early stage of chromosome assembly, the strand passage activity of topo IIα catalyzes inter-chromatid disentanglement, thereby promoting chromosome individualization and sister chromatid resolution. At a late stage of chromosome assembly, it catalyzes intra-chromatid entanglement, making each chromatid more compact. c Frequency of lumps and loops on U-shape DNA in the presence of condensin I (cond I) alone, cond I and topo IIα, or topo IIα alone. Percentages of DNAs per condition are plotted (red: lump, blue: loop, white: N.D., which denotes no significant structure was identified, mean ± SEM). Proteins were simultaneously introduced into U-shape DNA-tethered flow cells with imaging buffer at 20 μl/min flow for 100 s. Unbound proteins were washed out during the observation. Data were collected from 3 trials. d Snapshots of loop and lump on a single lambda DNA. DNA was stained with SYTOX Orange and condensin I was labeled with Halo-Alexa 488 (Bar: 2 μm). Note that all DNA molecules are at least bound by single condensin I complexes. e Stability of lumps in the presence or absence of topo IIα. The maximum duration is the observation time 20 min. The translucent and solid points denote the individual lump and median of each trial, respectively. The bars represent the mean of the median ± SEM. Data were collected from 3 trials. Single-molecule tracking of condensin I and compacted lump (f, g) or extruded loop (h) in the presence (g) or absence (f, h) of topo IIα. Time 0 indicates the moment when condensin I bound to DNA. In all experiments in this figure, DNA was stained with SYTOX Orange. All pvalues are calculated by a two-tailed Mann–Whitney U test. Source data are provided as a Source Data file. Sample sizes are stated in the Source Data file.
Much less is known about the mechanisms of action of condensin I. Early studies showed that condensin I has the ability to introduce positive superhelical tension into double-stranded DNA (dsDNA) in vitro in an ATP hydrolysis-dependent manner: relaxed circular DNA is converted into positively supercoiled DNA in the presence of type I topoisomerases12, whereas nicked circular DNA is converted into knotted DNA in the presence of a type II topoisomerase13. Because Cdk1 phosphorylation of condensin I strongly stimulates these activities13,14, they are thought to be physiologically relevant activities underlying mitotic chromosome assembly. Recent single-molecule studies have shown that condensins have the so-called loop extrusion activity that forms and expands DNA loops in an ATP hydrolysis-dependent manner15,16. The loop extrusion activity has been proposed to be a fundamental activity that organizes mitotic chromosomes by generating consecutive loops17.
As a natural consequence of the double-stranded nature of DNA, sister DNAs become entangled after DNA replication (Fig. 1b, left)18. Substantial evidence has demonstrated that the action of condensins helps topo II to facilitate the resolution of such entanglements (inter-chromatid disentanglement) in preparation for complete sister chromatid separation during mitosis (Fig. 1b, center)19,20. Interestingly, evidence is now accumulating for a second function of topo IIα in mitotic chromosome assembly: DNA strands within individualized chromatids become self-entangled (intra-chromatid entanglement) through the action of topo IIα (Fig. 1b, right)8,21,22. It is speculated that this previously underappreciated action of topo IIα is facilitated by the crowded environment created during the process of mitotic chromosome assembly8.
Although the individual activities of topo IIα and condensin I have been studied extensively, only a limited number of efforts have been made to address the question of how the two ATPases cooperate to assemble mitotic chromosomes at a mechanistic level23.
In the current study, we have used total internal reflection fluorescence microscopy to analyze the interplay between condensin I and topo IIα at single-molecule resolution. To this end, topo IIα is added to a standard condensin I-mediated loop extrusion assay. We find that inclusion of topo IIα converts the condensin I-mediated DNA structures into stable compact structures in a manner that is dependent on the C-terminal domain (CTD) of topo IIα. Moreover, the strand passage activity of topo IIα induces DNA knotting within the stable lumps, rendering them resistant to protease treatment. ATP hydrolysis by condensin I stimulates the topo IIα-mediated knotting reaction. Our results demonstrate how topo IIα-mediated strand passage is coupled with condensin I-mediated loop extrusion to generate a compact DNA structure. The functional implications of these observations for mitotic chromosome assembly and stabilization are discussed.
Results
Condensin I forms stable lumps in the presence of topo IIα
To address the question of how condensin I and topo IIα cooperate to change the conformation of DNA at a single molecule resolution, we introduced topo IIα to our condensin I-mediated loop extrusion assay. In brief, we utilized a flow cell-based assay system, which was visualized under a total internal reflection fluorescence microscope (TIRFM) as previously reported24. 48.5-kbp λDNA was biotinylated at both ends and tethered onto a streptavidin-coated coverslip to make U-shape DNA substrates. A mammalian condensin I complex and Xenopus laevis topo IIα were expressed using the baculovirus expression system and purified as previously reported8,25. The purified condensin I was then fluorescently labeled with Alexa488. Unless otherwise indicated, we used unlabeled topo IIα.
When condensin I alone (1 nM) was injected into the flow cells, U-shape DNA was either converted into a loop (35.6%) as reported previously15,16,25,26 or a compact structure that consisted of a DNA mass without extruded loop (27.3%), hereafter referred to as a “lump” (Fig. 1c, see cartoon in Fig. 1d). The lumps were quantitatively distinguished from loops by measuring the roundness of DNA structures strongly stained with SYTOX Orange. To quantify roundness, the SYTOX Orange-positive DNA areas were approximated as ellipses and the ratio of the major axis (a) to the minor axis (b), i.e., a/b, was calculated. We defined loops as structures in which the a/b ratio was greater than 1.6 at least two consecutive frames during the observation and defined all other compacted structures as lumps (Supplementary Fig. 1a–c). Injection of topo IIα alone at a low concentration (0.125 nM) did not change the DNA shape. However, when condensin I and topo IIα were injected together at concentrations of 1 nM and 0.125 nM, respectively, lump formation dominated: lumps were formed on ~70% of U-shape DNA (see an example in Fig. 1d, “lump”), whereas loops were formed on less than 10% of DNA (see an example in Fig. 1d, “loop”). Importantly, we found that the lumps formed by condensin I and topo IIα were sustained for far longer time (in most cases, sustained throughout the observation time [20 min]) than those formed by condensin I alone (Fig. 1e), although the kinetics of lump formation were similar under the two conditions (Fig. 1f, g). Time-lapse imaging revealed that DNA began to be converted into lumps immediately after condensin I binding (Fig. 1f, g, and Supplementary Fig. 1d–g, Supplementary Movie 1), and the lumps grew more slowly than the loops (Fig. 1f–h). We did not observe examples of loop-to-lump conversion unless a second condensin is bound to the loops (Supplementary Movie 2), suggesting that the lump formation is initiated before loop growth. In the absence of ATP, we observed only limited numbers of lumps with reduced size (Fig. 2a, b, and Supplementary Fig. 2a, b) and reduced rates of formation (Fig. 2c), indicating that the stable lump formation is ATP-dependent.
a Time trace of loop or lump size associated with single condensin I (cond I) complexes in the presence or absence of topo IIα and ATP. The line and shaded area denote the Exponential fits and interquartile ranges (red: condensin I alone, blue: condensin I and topo IIα, black: condensin I and topo IIα without ATP). Time 0 indicates the moment when condensin I bound to DNA. Data were collected from 3 trials. b Size of lump or loop formed by condensin I in the presence or absence of topo IIα and ATP. The bars denote the median and quartiles. Data were collected from 3 trials. c Rate of loop or lump formation by condensin I in the presence or absence of topo IIα and ATP. The bars denote the median and quartiles. d A snapshot of condensin I (cond I)A488 and topo IIαA546 on a lump. After the detection of proteins, DNA was stained with SYTOX Green (Bar: 2 μm). Time trace of condensin IA488 (e) and topo IIαA546 (f) signals binding non-specifically to the glass surface. Condensin IA488 or topo IIαA546 particles were bleached by 488 nm or 561 nm laser and images were acquired every 0.5 s for 15 s. Note that both signals were bleached in one step (left). Fluorescence intensities of particles bleached in one step were plotted (right, mean ± SD). Binding kinetics of condensin IA488 (g) and topo IIαA546 (h) on DNA. Median (solid line) and interquartile ranges (shaded area) are shown. Dashed lines denote fluorescence intensities of single condensin IA488 complexes or single topo IIαA546 dimers. Time 0 indicates the moment when the first proteins bound to DNA. Data were collected from 2 trials. All pvalues are calculated by a two-tailed Mann–Whitney U test. Source data are provided as a Source Data file. Sample sizes are stated in the Source Data file.
To identify how many molecules of condensin I and topo IIα are involved in the formation of stable lumps, we measured the intensities of the fluorescently labeled condensin I (Alexa488) and topo IIα (Alexa546) that bind to each lump (Supplementary Fig. 2c–e). The fluorescence intensities of single molecules of condensin IA488 and topo IIαAlexa546 were determined by photobleaching experiments, in which individual molecules were bleached in one step (Fig. 2e, f). By comparing these single-molecule intensities with those of condensin I and topo IIα bound to lumps, we estimated that condensin I was predominantly present as a single complex on each lump (48.1%), with smaller populations of two (24.7%) and three (11.2%) complexes (Supplementary Fig. 2c, d). Similarly, topo IIα was predominantly present as a single dimer (75%), with a minority existing as two dimers (25%) (Fig. 2g, h and Supplementary Fig. 2e–j).
Condensin I/topo IIα-mediated lumps are resistant to protease treatment
Although the lumps formed by condensin I alone were far less stable than those formed by both condensin I and topo IIα (Fig. 1e), we found that condensin I was continued to be detectable on DNA throughout the observation time [20 min] even after the lumps were dissolved (Supplementary Fig. 3a). Thus, condensin I binds stably to DNA even in the absence of topo IIα.
To further clarify the differences between condensin I-mediated and condensin I/topo IIα-mediated lumps, we then treated the two types of lumps with a protease (Proteinase K). As expected, the fluorescent signals of both condensin I and topo IIα disappeared from DNA upon protease treatment (Supplementary Fig. 3b, c). We found that the lumps and loops formed by condensin I alone were immediately dissolved upon this treatment (Fig. 3a upper, and 3b, c, Supplementary Movie 3). Remarkably, however, the lumps formed by both condensin I and topo IIα remained even after protease treatment although their size decreased slightly (Fig. 3a lower, and 3b, c). These lumps are hereafter referred to as “protease-resistant lumps”. We also found that the small, condensin I/topo IIα-mediated lumps observed in the absence of ATP (Fig. 2b) were protease-sensitive (Fig. 3c). The protease-resistant lump formation appeared to be sensitive to DNA tension, as its efficiency was decreased under continuous buffer flow, during which the DNA was continuously extended (Supplementary Fig. 3d) As the flow conditions were identical in the experiments and the distance between the tethered ends of the DNA molecules did not significantly vary, differences in DNA tension are not expected to account for the observed changes (Supplementary Fig. 3e, f). Although ~80% of the lumps formed by condensin I and topo IIα were protease-resistant, ~20% were judged to be protease-sensitive (Fig. 3c). We then compared the DNA binding kinetics of condensin I and topo IIα between the protease-resistant and protease-sensitive lumps, and found a tendency for the lumps to become more protease-resistant when condensin I bound to DNA before topo IIα (Supplementary Fig. 4a). These results suggested that condensin I creates a preferred DNA structure for topo IIα to bind and thereby generates the protease-resistant lumps. This idea was further supported by experiments involving the sequential addition of condensin I and topo IIα (Supplementary Fig. 4b).
a Snapshots of the lumps formed by condensin I (cond I) alone or cond I and topo IIα before (−20 s) and after (160 s, 340 s) Proteinase K (ProK) treatment. Time 0 indicates the moment when the first injections of ProK. Cond I and topo IIα were injected simultaneously (−200 s), washed out with buffer (IB) (−100 s), and then ProK (0.5 mg/ml) was injected (0 s). b The size of the lumps formed in the experiment shown in (a). The translucent and solid points denote the individual lump and median of each trial, respectively. The bars represent the mean of the median ± SEM. Data were collected from 3 trials. c Frequency of protease-resistant and -sensitive lumps associated with single condensin I complexes in the presence or absence of ATP and topo IIα. Percentages of lumps per condition are plotted (black: protease-resistant, light gray: protease-sensitive, mean ± SEM). Data were collected from 3 trials. In all experiments in this figure, DNA was stained with SYTOX Orange. All p values are calculated by a two-tailed Mann–Whitney U test. Source data are provided as a Source Data file. Sample sizes are stated in the Source Data file.
Finally, we examined changes in the size of protease-resistant lumps (after deproteinization) in response to DNA tension imposed by different flow rates. If the DNA within the lumps was topologically constrained, then it was expected that (1) the lumps would be maintained even as DNA tension increased; and (2) the apparent lump size would decrease as DNA tension increased, but it would recover as tension decreased. We evaluated the apparent lump size by measuring the cross-sectional intensity of the lumps and comparing either the highest DNA intensity on the lump (Max) or the sum of the DNA intensities on the lump (Area Under the Curve [AUC]), and found that both Max and AUC values decreased when the flow rate was increased, but they were restored when the flow rate was decreased again (Supplementary Fig. 5). The lump size was approximately 2–3-fold greater in compacted tips than that in uncompacted tips both before and after extension. These results strongly suggested that the condensin I/topo IIα-mediated lump, but not the condensin I-mediated lump, contains a topologically constrained DNA structure.
DNA strand passage by topo IIα is essential for protease-resistant lump formation
We next directly tested whether the DNA strand passage activity of topo IIα is required for protease-resistant lump formation. We expressed and purified Xenopus laevis topo IIα harboring the strand passage-deficient mutation Y803F (Supplementary Fig. 6a)27,28. We confirmed that the mutant topo IIα (topo IIαY803F) was catalytically null in the standard decatenation assay in vitro (Supplementary Fig. 6b). As expected, topo IIαY803F bound to chromatin but failed to assemble mitotic chromosomes in Xenopus egg extracts (Supplementary Fig. 6c).
Topo IIαY803F was then subjected to our single-molecule assay. We found that topo IIαY803F supported lump formation similarly to wild-type topo IIα (topo IIαWT) in terms of formation efficiency (Fig. 4a), kinetics (Fig. 4b), stability (Fig. 4c), size (Fig. 4d), and formation rate (Fig. 4e). These results are consistent with the observation that topo IIαY803F retains its ability to bind to chromatin in Xenopus egg extracts as described above. Remarkably, however, the lumps formed by topo IIαY803F were largely protease-sensitive, as opposed to those formed by topo IIαWT (Fig. 4f). Thus, the strand passage activity is essential for protease-resistant lump formation, but not for stable lump formation. However, despite the fact that the topo IIαY803F is catalytically null, a non-negligible fraction (~10%) of lumps were judged to be protease-resistant under the current condition (Fig. 4f). We reasoned that the DNA present in the residual protease-resistant lumps might be constrained in a non-topological manner. In fact, when a higher extension force was imposed on the DNA by increasing the flow rate (from 20 μl/min to 1 ml/min; Supplementary Fig. 7), the residual protease-resistant lumps formed by topo IIαY803F completely disappeared (Supplementary Fig. 6d, YF), although the protease-resistant lumps formed by topo IIαWT were maintained (Supplementary Fig. 6d, WT).
a Frequency of lumps and loops on U-shape DNA formed by condensin I (cond I) and either topo IIαWT (WT) or topo IIαY803F (YF). Percentages of DNAs per condition are plotted (red: lump, blue: loop, white: N.D., mean ± SEM). b Time trace of the size of a lump formed by condensin I in the presence of either topo IIαWT or topo IIαY803F. The line and shaded area denote the Exponential fits and interquartile ranges (black: condensin I and topo IIαWT, blue: condensin I and topo IIαY803F). Time 0 indicates the moment when condensin I bound to DNA. Data were collected from 3 trials. c Stability of lump in the presence of condensin I and either topo IIαWT or topo IIαY803F. The translucent and solid points denote the individual lump and median of each trial, respectively. The bar represents the mean of the median ± SEM. d Size of lump formed in the presence of condensin I and either topo IIαWT or topo IIαY803F. The bars denote the median and quartiles. e Rate of lump formation in the presence of condensin I and either topo IIαWT or topo IIαY803F. The bars denote the median and quartiles. f Frequency of protease-resistant and -sensitive lump in the presence of condensin I and either topo IIαWT or topo IIαY803F. Percentages of lumps per condition are plotted (black: protease-resistant, light gray: protease-sensitive, mean ± SEM). g Changes in protease-resistant lumps after the 2nd topo IIα treatment were categorized into 4 groups and the percentages are plotted (white: increased or no change, light gray: decreased by no step, dark gray: decreased by one step, black: disappeared by one step, mean ± SEM). In all experiments in this figure, DNA was stained with SYTOX Orange. All data in this figure were collected from 3 trials. All pvalues are calculated by a two-tailed Mann–Whitney U test. Source data are provided as a Source Data file. Sample sizes are stated in the Source Data file.
We next treated deproteinized lumps with freshly added topo IIα and found that most of the lumps diminished or disappeared (Fig. 4g). It is noteworthy that in most cases, DNA intensities were decreased in a stepwise manner (Fig. 4g), implying that these lumps were dissolved by one or a few step(s) of the strand passage reaction. Based on these results, we concluded that the protease-resistant lumps generated by condensin I and topo IIαWT contain DNA knots, as reported in the previous bulk biochemical studies13,29.
The CTD of topo IIα is required for protease-resistant lump formation
To get additional insights into the mechanism of protease-resistant lump formation, we next asked if stable binding of topo IIα to DNA is required for this process. Previous studies have shown that the intrinsically disordered CTD of topo IIα is required for stable localization of topo IIα to reconstituted mitotic chromosomes and their thickening8. We purified topo IIα lacking its CTD (topo IIαΔCTD; Fig. 5a) and found that the addition of topo IIαΔCTD had little, if any, impact on condensin I-mediated structural changes of DNA in our single-molecule assay (Fig. 5b). Similar to the lumps formed with condensin I alone, the lumps formed with condensin I and topo IIαΔCTD were all protease-sensitive (Fig. 5c). Thus, stable DNA binding mediated by the CTD of topo IIα is required for protease-resistant lump formation, even though topo IIαΔCTD is catalytically fully active as judged by the standard DNA decatenation assay8. Consistently, we confirmed that freshly added topo IIαΔCTD can dissolve deproteinized lumps containing knots (Supplementary Fig. 8).
a Schematic representation of topo IIαWT (WT) and topo IIαΔCTD (ΔCTD). b Frequency of lumps and loops on U-shape DNA formed by condensin I (cond I) in the presence of either topo IIαWT and topo IIαΔCTD. Percentages of DNAs per condition are plotted (red: lump, blue: loop, white: N.D., mean ± SEM). c Frequency of protease-resistant and -sensitive lump formation by condensin I and either topo IIαWT or topo IIαΔCTD. Percentages of lumps per condition are plotted (black: protease-resistant, light gray: protease-sensitive, mean ± SEM). All data in this figure were collected from 3 trials. In all experiments in this figure, DNA was stained with SYTOX Orange. Source data are provided as a Source Data file. Sample sizes are stated in the Source Data file.
ATP hydrolysis by condensin I strongly stimulates protease-resistant lump formation
Next, we investigated how condensin I’s ATPase activity affects lump formation. For this purpose, we prepared a fluorescently labeled, condensin I mutant complex deficient in ATP hydrolysis (Smc2[E1114Q] and Smc4 [E1218Q], henceforth referred to as condensin IEQ)25. Consistent with previous studies in all types of eukaryotic SMC complexes16,30,31,32, condensin IEQ alone showed no loop extrusion activity (Fig. 6a). Moreover, topo IIα-independent lump formation was also significantly suppressed (Fig. 6a). On the contrary, in the presence of topo IIα, condensin IEQ formed lumps with a high frequency and stability indistinguishable from those observed with condensin IWT (Fig. 6b, c).
a Frequency of lumps and loops on U-shape DNA formed by condensin IWT (WT) or condensin IEQ (EQ). Percentages of DNAs per condition are plotted (red: lump, blue: loop, white: N.D., mean ± SEM). b Frequency of lumps and loops on U-shape DNA formed in the presence of topo IIαWT and either condensin IWT or condensin IEQ. Percentages of DNAs per condition are plotted (red: lump, blue: loop, white: N.D., mean ± SEM). c Stability of lumps formed by condensin IWT or condensin IEQ in the presence of topo IIαWT. The translucent and solid points denote the individual lump and median of each trial, respectively. The bars represent the mean of the median ± SEM. d Time trace of the size of a lump formed by condensin IWT or condensin IEQ in the presence of topo IIαWT. The line and shaded area denote the Exponential fits and interquartile ranges (black: condensin IWT and topo IIα, amber: condensin IEQ and topo IIα). Time 0 indicates the moment when condensin I bound to DNA. e Rate of lump formation in the presence of topo IIαWT and either condensin IWT or condensin IEQ. The bars denote the median and quartiles. f Size of lumps formed in the presence or absence of either condensin IWT or condensin IEQ and topo IIαWT. The bars denote the median and quartiles. g Frequency of protease-resistant and -sensitive lumps formed by condensin IWT or condensin IEQ in the presence of topo IIαWT. Percentages of lumps per condition are plotted (black: protease-resistant, light gray: protease-sensitive, (mean ± SEM). All data in this figure were collected from 3 trials. DNA was stained with SYTOX Orange. All pvalues are calculated by a two-tailed Mann–Whitney U test. Source data are provided as a Source Data file. Sample sizes are stated in the Source Data file.
Whereas condensin IEQ, like condensin IWT, bound to DNA as single complexes (Supplementary Fig. 9a, b), we noticed that condensin IEQ formed lumps more slowly than condensin IWT (Fig. 6d). The formation rate (Fig. 6e) and the size (Fig. 6f) of the lumps formed by condensin IEQ were significantly smaller than those formed by condensin IWT. Furthermore, most of the lumps formed by condensin IEQ in the presence topo IIαWT ( ~ 90%) were protease-sensitive (Fig. 6g). To test whether the DNA present in the residual protease-resistant lumps (~10%) is topologically constrained, we applied a higher extension force by increasing the flow rate. We found that the residual protease-resistant lumps formed by condensin IEQ and topo IIαWT were fully tolerant to the high extension force (Supplementary Fig. 9c), in contrast to those formed by condensin IWT and topo IIαY803F (Supplementary Fig. 6d). This observation suggests that condensin IEQ can introduce DNA knots in the presence of topo IIαWT, but at a much lower frequency than condensin IWT.
Discussion
The establishment of a chromatid reconstitution assay has allowed us to identify the minimal set of protein components required for the core reaction of mitotic chromosome assembly7,8. Among the six essential components identified, condensin I and topo IIα are the only two ATPases. Although the two ATPases have extensively been studied using various single-molecule assays, they have never been studied in combination. The current study represents an attempt to study the interplay between condensin I and topo IIα at single-molecule resolution, with the aim of bridging the gap in our knowledge between the reconstitution and single-molecule assays.
Our results reported in the current study are summarized in Fig. 7a. In our current experimental setup, condensin I alone generates loops and unstable lumps on the U-shape DNA substrates (Fig. 1). We speculate that the observed unstable lumps are a precursor of the loops (while these precursors were also observed in previous studies, their frequency may vary depending on experimental conditions, including the concentration of SYTOX dye, differences in protein preparations, and the level of torsional tension imposed on the DNA15,33,34). Topo IIα alone, at the low concentration used in the current study, has little or no impact on the conformation of the DNA substrates. However, when condensin I and topo IIα are added together, stable lump formation dominates over loop formation (Fig. 1). Importantly, this condensin I/topo IIα-mediated lump formation depends on the CTD of topo IIα (Fig. 5), but not on its catalytic activity (Fig. 4). Although we still do not know how the condensin I/topo IIα-mediated lumps are formed, the requirement for the CTD suggests that topo IIα must stay on the DNA for a long time through CTD-mediated binding to a specific DNA structure created by condensin I (e.g., DNA crossover or bent DNA). It is also possible that topo IIα recognizes a chiral DNA structure generated by condensin I, as human topo IIα has been proposed to preferentially relax positive supercoils in a CTD-dependent manner35. We do not exclude the possibility that topo IIα physically interacts with condensin I on DNA, although there is no direct evidence for condensin-topo II interactions in solution (i.e., without DNA) in eukaryotic systems (e.g., ref. 32).
a Observations in the current study. Wild-type condensin I (cond IWT) forms a stable lump in the presence of wild-type topo IIα (topo IIαWT). Stable lump formation per se does not require DNA strand passage by topo IIα (topo IIαY803F), but depends on the CTD of topo IIα (topo IIαΔCTD). However, the lump formed in the presence of topo IIαWT becomes protease-resistant because the DNA within the lump is converted to a knotted form. A mutant condensin I complex deficient in ATP hydrolysis (cond IEQ), in the presence of topo IIαWT, forms a small lump in which DNA is rarely knotted. b Functional implications in mitotic chromosome assembly. On the one hand, the entanglement within single loops (i.e., intra-loop entanglement) would facilitate lateral compaction and stabilization of mitotic chromosomes. On the other hand, the entanglement between adjacent loops (i.e., inter-loop entanglement) would promote axial shortening and stabilization of mitotic chromosomes. These two possibilities, which are not mutually exclusive, would readily occur in the highly crowded environment created during mitotic chromosome assembly.
The condensin I/topo IIα-mediated lumps become protease-resistant when topo IIα is catalytically active (Fig. 4). The following two lines of evidence strongly suggest that the protease-resistant lumps contain knotted DNA (Fig. 7a). First, the topo IIαY803F mutant supports the formation of stable lumps but fails to render them protease-resistant. Second, the protease-resistant lumps, after deproteinization, can be resolved with freshly added topo IIα. Most of the protease-resistant lumps are dissolved in a single step, suggesting that most of the knots generated under the current conditions are simple knots, such as 3-noded knots, similar to those observed in the previous bulk biochemistry using condensin I and a type II topoisomerase13.
A recent preprint using magnetic tweezers has reported that a low concentration (~0.1 nM or less) of human topo IIα displays the ability to compact DNA, possibly through a mechanism of a polymer-collapse phase transition36. The compaction activity is fully dependent on the CTD of topo IIα, but is independent of ATPase activity of condensin I. Thus, the topo IIα-mediated DNA compaction reported in the preprint is clearly distinct from the condensin I/topo IIα-mediated lump formation described in the current study. Nevertheless, it is important to note that both studies highlight the hitherto underappreciated role of the topo IIα CTD in the conformational changes of DNA that is separable from its strand passage activity.
How might condensin I-mediated loop extrusion be coupled to topo IIα-mediated knotting? A mutant condensin I complex harboring the ATP hydrolysis-deficient SMC subunits (condensin IEQ) generates lumps at the same frequency as condensin IWT in the presence of topo IIα (Fig. 6). We notice that the size of condensin IEQ /topo IIα-mediated lumps was substantially smaller than that of condensin IWT /topo IIα-mediated lumps and that the former was far more protease-sensitive than the latter. Here, we interpret our observations based on the recently updated model of condensin-mediated loop extrusion37. According to this model, upon ATP binding and head-head engagement, a small loop is formed in the SMC lumen (i.e., the “feeding” loop). Upon ATP hydrolysis and head-head disengagement, the feeding loop is transported into the kleisin lumen, where it is merged into the “extruding” loop (Supplementary Fig. 10). We speculate that the extruding loop generated by condensin IWT would become an excellent substrate for topo IIα-mediated knotting. In contrast, condensin IEQ forms a feeding loop but is unable to convert it into an extruding loop. This could explain why condensin IEQ generates a small lump in which topo IIα-mediated knotting occurs only at a low frequency. The question of how the condensin-mediated loop extrusion process imposes superhelical tension on DNA has only recently been recognized in the field29,38,39. The currently available data are somewhat fragmentary, and our interpretation is partially, if not completely, consistent with the emerging picture. Clearly, future studies are needed to address this important question.
What are the functional implications of the current observations for mitotic chromosome assembly and stabilization? Our finding that the CTD of topo IIα is essential for the formation of protease-resistant lumps, hence DNA knotting, would provide an important clue (Fig. 1a). It has been shown that topo IIαΔCTD is proficient in topo IIα-catalyzed decatenation of circular DNA but is deficient in its efficient catenation (Fig. 1a)8,40. In addition, topo IIαΔCTD is proficient in inter-chromatid disentanglement but is deficient in intra-chromatid entanglement that is predicted to support chromatid thickening (axial shortening) in the reconstitution assay (Fig. 1b)8. Importantly, unlike topo IIαWT, topo IIαΔCTD is not detectable on chromosomes during the assembly processes. These observations have been interpreted to mean that the long residence time imposed by the CTD helps topo IIα to catalyze catenation8,40 and intra-chromatid entanglement in the reconstitution assay8. We speculate that the same is true for the topo IIα-stimulated lump formation and DNA knotting reported in the current study. Although direct evidence for the occurrence of intra-chromatid entanglements (self-entanglements) in mitotic chromosomes is lacking, an early study using micromanipulation approach21 and a recent study using high-throughput chromosome conformation capture (Hi-C) and polymer simulations22 have provided data consistent with this idea.
We consider two scenarios for the role of intra-chromatid entanglements within mitotic chromosomes (Fig. 7b). The first is the entanglement within single loops (i.e., intra-loop entanglement), which would facilitate lateral compaction and stabilization of mitotic chromosomes. The second is the entanglement between adjacent loops (i.e., inter-loop entanglement), which would promote axial shortening and stabilization of mitotic chromosomes. The products observed in the current study (DNA knots) are roughly equivalent to those resulting from intra-loop entanglements. However, we consider that it may be possible to recapitulate inter-loop entanglements by modifying the current setup of single-molecule analyses. In any case, it is reasonable to speculate that the two reactions hypothesized here, which are not mutually exclusive, would readily occur in highly crowded environments created during mitotic chromosome assembly. Our current results do not exclude the possibility that topo IIα also uses a mechanism of protein-based “locking” of two DNA duplexes to stabilize mitotic chromosomes after their assembly41,42.
Finally, it should be added that, although the DNA present within the protease-resistant lumps is most likely a simple form of DNA knots (i.e., three-noded knots), it is uncertain whether they have specific chiralities. In this sense, it is worth mentioning that a mitotically phosphorylated form of native condensin I complex purified from Xenopus egg extracts generates DNA knots with a specific chirality (i.e., positive 3-noded knots) in the presence of a type II topoisomerase13. Because the recombinant condensin I complexes used in the current study are not subjected to mitotic phosphorylation, the knots discussed in this study may not have specific chiralities. A better characterization of the interplay between condensins and topo IIα in vitro and on mitotic chromosomes in situ is an important direction in the future.
Methods
Preparation of recombinant condensin I holocomplexes
For the expression of recombinant condensin complexes in insect cells, the Bac-to-Bac Baculovirus Expression System (Thermo Fisher Scientific) was used as described previously25. The ATP hydrolysis-deficient mutations of mSMC2 (E1114Q) and mSMC4 (E1218Q) were introduced as described previously43. These mutations were referred to as transition-state (TR) mutations in Ref#38, whereas the same mutations are referred to as EQ mutations in the current study. Purification and labeling of Halo-tagged versions of the condensin I holocomplexes were performed as described previously25.
To determine fluorescence labeling efficiency of condensin IA488, anti-FLAG M2 antibodies (Sigma Aldrich, F1804) were nonspecifically attached to the glass, then 1 nM FLAG-tagged condensin IA488 in ELB + + was introduced at 30 μl/min for 1 min and incubated for 5 min with 5 μl/min flow. After incubation, the flow cells were washed with ELB + + at 30 μl/min for 2 min, and anti-mSmc4 antibody (AfR326-3L) in ELB-Casein (10 mM HEPES-KOH [pH7.7], 50 mM KCl, 2.5 mM MgCl2, 0.4 mg/ml Casein) was introduced at 10 μl/min for 18 min, and washed with ELB + + at 20 μl/min for 5 min. Then 2 μg/ml Alexa Fluor 647-conjugated anti-rabbit IgG antibodies (Invitrogen) in ELB + + were introduced at 10 μl/min for 10 min and washed with ELB + + at 20 μl/min for 5 min. Finally, proteins were visualized with the 488 and 640 lasers, and the percentage of signal colocalization was calculated.
Preparation of recombinant topo IIα proteins
Plasmid DNAs encoding Xenopus laevis topo IIα (topo IIαWT [residues 1-1579] and topo IIαΔCTD [residues 1-1,205]) with an N-terminal 3xFLAG-tag and a C-terminal Strep II-tag, were constructed as described previously8. To prepare topo IIαY803F, an amino acid substitution of Y803F was introduced into the original plasmid DNAs using the QuikChange XL site-directed mutagenesis kit (Agilent, 200517). An N-terminally SNAP-tagged topo IIα construct was created with a synthetic DNA fragment encoding the SNAP tag (codon-optimized for Trichoplusia ni). Using these plasmid DNAs, recombinant bacmid DNAs and baculoviruses were produced as described previously8. Recombinant proteins were expressed in HighFive insect cells by infecting them with amplified baculovirus. For purification of the full-length versions of topo IIα (topo IIαWT and topo IIαY803F), cell lysate was subjected to tandem chromatography using Strep-Tactin Sepharose resin (IBA, 2-1201-010) and a Capto HiRes S 5/50 cation exchanger column (Cytiva, 29275877). For purification of topo IIαΔCTD, a combination of Strep-Tactin Sepharose resin and HiTrap Q HP anion exchanger column (Cytiva, 17115301) was used. Purified SNAP-topo IIα derivatives were fluorescently labeled with the SNAP-Surface AlexaFluor 546 ligand (New England Biolabs, S9132) according to the manufacturer’s instructions. Unconjugated ligand was removed by an additional round of cation exchange chromatography. Protein and conjugated ligand concentrations were determined by measuring absorbance at 280 nm and 558 nm, respectively. The labeling efficiency of Alexa Fluor 546–topo IIα used in this study, calculated as the ratio of the conjugated ligand concentration to the protein concentration, was estimated to be 62.5 %. All purified topo IIα proteins were dialyzed against KHG200/50 buffer (20 mM HEPES-KOH [pH 7.7], 200 mM KCl, 50% glycerol), aliquoted, snap-frozen in liquid nitrogen, and stored at −80 °C until use. The enzymatic activity of each recombinant topo IIα derivative was validated through kinetoplast DNA decatenation assays.
Preparation of microfluidic device
Single-molecule imaging was performed in cruciform flow-cell channels. For preparation of coverslips (Matsunami 22 × 22 mm No.1-S), the coverslips were sonicated in 1 M KOH for 20 min, rinsed with water, sonicated again in methanol, and left to stand for 1–3 nights in 2% 3-Aminopropyltriethoxysilane (Sigma Aldrich, 440140) in methanol. The coverslips were then treated with 100 mM NaHCO3 containing 100 mg/ml PEG (Laysan Bio, #MPEG-SVA-5K) and 6.6 mg/ml BioPEG (Laysan Bio, # BIO-PEG-SVA-5K) overnight. The coverslips were then gently washed with water, dried, and stored at −80° C. Before use, the coverslips were treated with 80 μM Streptavidin (Sigma Aldrich, S4762) in ELB + + (10 mM HEPES-KOH [pH7.7], 50 mM KCl, 2.5 mM MgCl2, 1 mg/ml BSA [Sigma Aldrich, 2905-OP, OmniPur BSA, Fraction V]) for at least 30 min. Excess unbound streptavidin was removed by washing with water.
The cruciform flow-cell channels were developed by putting glass slides and coated coverslips with double-sided tape (Grace Bio Labs GBL620001-1EA) with a cruciform channel of 2-mm width. Inlet (Natsume KN-392-1-SP-19) and outlet tubes (Natsume KN-392-1-SP-55) were connected to each channel, and the outlet tubes were connected to a syringe pump (Harvard Apparatus Pump 11 Elite) to withdraw solutions. The channels were washed with ELB + +.
Biotin-conjugation at both ends of λDNA (N3011, New England Biolabs) was performed as follows: 0.15 pmol λDNA and two biotinylated oligonucleotides, which are complementary to the cos sites (oligo #1, 5’-GGGCGGCGACC[biotinylated-T]−3’, 2 pmol, custom made by Integrated DNA Technologies); oligo #2, 5’-AGGT[biotinylated-T]CGCCGCCC-3’, 10 pmol, custom made by Eurofins Genomics, were phosphorylated with 0.5 μl T4 polynucleotide kinase (M0201, New England Biolabs) in the presence of 1 mM ATP at 37°C in 12 μl volume for 3 h. Oligo #1 (13.3x molar excess) and λDNA was diluted with 84 μl T4 ligation buffer (B0202, New England Biolabs), annealed by cooling the mixture gradually from 65 to 10 °C, and then ligated with 0.75 μl T4 DNA ligase (M0202, New England Biolabs) and 10 mM ATP at 16 °C for 2 h. Finally, oligo #2 (71.4 x molar excess) was annealed to another end of λDNA and ligated in the same way.
The resultant biotin-conjugated λDNA was injected into the perpendicular channel of the flow-cell at a final concentration of 11.5 pM in DNA buffer (50 mM Tris-HCl [pH7.5], 50 mM NaCl, 5 mM MgCl2) with either 22 nM SYTOX Orange (Thermo Fisher Scientific, S11368, inFigs. 1c–h, 2a–c, 3–6, Supplementary Figs. 1, 2a–d, 3a, c–f, 4b, 5, 6d, 8, 9) or SYTOX Green (Thermo Fisher Scientific, S7020, in Fig. 2d, g, h, Supplementary Figs. 2e, 3b, 4a). The linear distance between both ends of the DNA was found to be 1 to 3 μm. Unbound λDNA was washed out first with DNA buffer and then with imaging buffer (IB: 50 mM Tris-HCl [pH7.5], 50 mM NaCl, 5 mM MgCl2, 3 mM ATP, 1 mM DTT, 22 nM SYTOX Orange). In the experiments using Alexa546-labeled topo IIα (topo IIαA546), high salt-imaging buffer (50 mM Tris-HCl [pH7.5], 100 mM NaCl, 5 mM MgCl2, 3 mM ATP, 1 mM DTT, 22 nM SYTOX Green) was used to prevent non-specific binding of topo IIαA546 (Fig. 2d, g, h, Supplementary Figs. 2e, 4a). To observe U-shape DNA, the flow direction was switched from the perpendicular channel to the horizontal channel at a flow rate of 20 μl/min (estimated tension is <0.2 pN).
Lump formation assay
Condensin I and unlabeled topo IIα were simultaneously introduced into U-shape DNA-tethered flow cells with imaging buffer at a flow rate of 20 μl/min for 100 s. Condensin I was introduced at 1 nM in all experiments and topo IIα was introduced at either 125 pM (Figs. 1c–h, 2a–c except sample without ATP, 3, 6, Supplementary Figs. 1, 2a–d, 3a, b (left), c, e, f, 9a-b) or 88 pM (Fig. 2a–c, sample without ATP, Supplementary Figs. 3d, 4b, 5, 6d, 8, 9c). Unbound proteins were washed out with imaging buffer. To prevent air from entering the flow cells, the buffer flow was transiently stopped for approximately ten seconds when exchanging buffers. When ATP dependency was tested, we used imaging buffer without ATP instead (Figs. 2a–c, 3c). When both condensin IA488 and topo IIαA546 were visualized, due to the overlap of the fluorescent wavelengths of Alexa488 and SYTOX Green, DNA was first detected by SYTOX Green to identify the places of U-shape DNA. After washing out SYTOX dye, DNA binding of condensin I A488 and topo IIαA546 was monitored in real time. At the end of monitoring, DNA was stained again with SYTOX Green to confirm lump formation. To test protease resistance, lumps were treated with 0.5 mg/ml Proteinase K (ProK, Roche, #03115873001) in imaging buffer at a flow rate of 20 μl/min. When topo IIαA546 was visualized, the lumps were treated by ProK after the 2nd SYTOX staining (Fig. 2d, g, h Supplementary Figs. 2e, 4a). In topo IIα reinjection assay (Fig. 4g, Supplementary Fig. 8), ProK was washed out with imaging buffer at 20 μl/min for 100 s, and then 250 pM topo IIαWT or topo IIαΔCTD was reinjected with imaging buffer at 20 μl/min. In the intense wash assay (Supplementary Figs. 6d, 9c), after washing out ProK, the flow rate was increased from 20 μl/min to 1 ml/min for 50 s, and then decreased to 20 μl/min again to see whether the lumps remained. To calculate the number of folds of DNA intensity, protease-resistant lumps were formed as described above, then the flow rate was increased in a stepwise manner: at 20, 100, 200, 400 μl/min for 100 s, then at 1000 μl /min for 25 s, and finally at 20 μl/min for 100 s. To clarify the effect of flow on lump formation, proteins and DNA were incubated with and without flow. Condensin I at a concentration of 1 nM and unlabeled topo IIα at 88 pM were simultaneously introduced into the U-shape DNA binding flow cell with imaging buffer at a flow rate of 20 μl/min for 100 s. The flow cells were incubated with or without flow for another 100 s. Without changing the flow rate, the inlet tube carrying the proteins was quickly switched to a protein-free imaging buffer, and the flow cells were washed for 100 s. Similarly, the inlet tube was quickly switched to imaging buffer containing ProK without changing the flow rate. Finally, the percentage of ProK-resistant lump was measured.
In all imaging experiments except the bleaching assay, condensin IA488 and SYTOX Green-stained DNA were visualized using a 350 μW 488-nm laser (Coherent Sapphire LP 488 ± 2, 150 mW), and topo IIαA546 and SYTOX Orange-stained DNA were visualized using a 100 μW 561-nm laser (Coherent Sapphire LPX 561 ± 2, 480 mW). All experiments were performed at room temperature (25 ± 2 °C).
DNA lengthening using optical tweezers
Optical tweezers experiments were performed using a C-trap system (LUMICKS) to estimate the extension force on DNA. Two streptavidin-coated polystyrene beads (4.35 μm in diameter; Spherotech SVP-40-5) were laser-trapped, and topologically unconstrained biotin-conjugated λDNA was captured between the beads. After trapping, the bead-DNA complex was placed to a channel filled with imaging buffer containing SYTOX Orange (22 nM). The beads were first settled at a relaxed position (5 μm) and then moved to an extended position till double-strand denaturing (25 μm) at a rate of 0.5 μm/s. DNA force-extension (FE) curves were measured based on the external force acting on the beads and the distance between them. Experiments and data acquisition were performed under the control of BlueLake software (LUMICKS). Under extensive flow conditions (1000 μl/min), DNA extension was measured to be 13.8 μm. From the obtained FE curve, the corresponding force was estimated to be 1.2 ± 0.33 pN (Supplementary Fig. 7a).
To estimate the force under gentle flow conditions (20 μl/min), the obtained FE curve was fitted using the worm-like chain (WLC) model44, with or without polynomial correction45, assuming a persistence length (Lp) of 50 nm.
where, the external force, Boltzmann constant, absolute temperature, the DNA extension, and contour length are denoted as F, kB, T, z, and LC, respectively. The order of the polynomial is denoted as arbitrary i, where α2 = -0.5164228, α3 = -2.737418, α4 = 16.07497, α5 = -38.87607, α6 = 39.49944, and α7 = -14.17718. The best fit was obtained using the least squares method with i = 4 and L0 = 16.7 μm (Supplementary Fig. 7b). DNA extensions (z) under buffer flow at 20 μl/min was measured to be 6.0 ± 0.7 μm. By substituting these values into the Eq. (1), the force was calculated to be 0.07 ± 0.02 pN.
For extensible WLC (eWLC) model (2)46,47, we tested to fit FE curve with Lc = 16.7 μm, Lp = 50 nm, and different K values (either 500, 1000 or 10,000 pN).
where, K denotes the elasticity modulus. We realized it was less fit the FE curve compared to WLC with 4th polynomial correction (Supplementary Fig. 7c). All calculation and visualization were performed using Anaconda and Spyder software. For curve fitting, the curve_fit function from the scipy.stats package in the Scipy library was used. The source code used for the analysis is available at https://doi.org/10.5281/zenodo.15789118.
Bleaching assay
For the bleaching assay to detect single-molecule intensities of condensin IA488 and topo IIαA546 (Figs. 2e, f), 1 nM condensin IA488 or 75 pM topo IIαA546 was non-specifically attached to coverslips. The fluorescently labeled proteins were then bleached by either a 3.5 mW 488-nm laser or 5.0 mW 561-nm laser, the images were taken every 0.5 s, and the bleaching step was detected. The step size was determined by calculating the difference in average intensity of five frames before and after the bleaching.
DNA decatenation assay
One hundred nanograms of a catenated DNA substrate (kinetoplast DNA; TG2013, TopoGEN) were mixed with 40 ng of recombinant topo IIα in a 10-µl volume (20 mM HEPES-KOH [pH 7.7], 80 mM KCl, 5 mM MgCl2, and 2 mM ATP), and incubated at 22 °C. At various times, aliquots were taken and treated with SDS (0.5%) and proteinase K (1.0 mg/ml; P-4032, Sigma) at 37 °C for 1 h. The resultant DNAs were purified with phenol and separated by gel electrophoresis on a 0.8% agarose gel in TAE. After being stained with ethidium bromide, fluorescent images were acquired with an image analyzer (Amersham Imager 680 [version 2.0.0], Cytiva).
Depletion and add-back assays using recombinant topo IIα in Xenopus egg extracts
The high-speed supernatant of metaphase-arrested Xenopus egg extracts (M-HSS) and demembranated Xenopus sperm nuclei were prepared as described previously48,49. A rabbit polyclonal antibody was raised against a synthetic peptide corresponding to the C-terminal amino acid sequence of Xenopus laevis topo IIα (GRQKKPVTYLEDSDDDF) and affinity-purified (referred to here as anti-topo IIα, in-house identifier AfR474-2). One hundred microliters of M-HSS were incubated with Dynabeads Protein A (Thermo Fisher Scientific, 10002D) conjugated with 20 µg of anti-topo IIα. After two successive rounds of incubation on ice (30 min each), the supernatants were isolated from the beads under a magnetic field and used as topo IIα-depleted M-HSS for subsequent assays. Control mock-depleted M-HSS was prepared using non-immune rabbit IgG (Sigma-Aldrich, I-5006) instead of anti-topo IIα. To assess the ability of recombinant topo IIα (wild-type and Y803F mutant) to support mitotic chromatid assembly in egg extracts, sperm nuclei were incubated with either mock-depleted or topo IIα-depleted M-HSS that had been supplemented with buffer (KHG200/50) or recombinant topo IIα at 22 °C for 120 min. The resultant chromatin structures were fixed, stained with DAPI, labeled with anti-topo II monoclonal antibody (M052-3, MBL, RRID: AB_592894), and analyzed by fluorescence microscopy as described previously8.
Immunofluorescence labeling of FLAG-topo IIα
After lump formation by condensin I and FLAG-tagged topo IIα, the lumps were washed with imaging buffer in the presence or absence of ProK for 100 s, and then ProK was washed out by another 100-s wash. The channels were washed with ELB + + at 20 μl/min for 5 min, 1 μg/ml anti-FLAG M2 antibodies (Sigma Aldrich, F1804) in ELB + + was introduced at 10 μl/min for 18 min, and washed with ELB + + at 20 μl/min for 5 min. Then 2 μg/ml Alexa Fluor 647-conjugated anti-mouse IgG antibodies (Thermo Fisher Scientific, A-21235) in ELB + + were introduced at 10 μl/min for 10 min and washed with ELB + + at 20 μl/min for 5 min. Finally, DNAs were visualized with 11 nM SYTOX Green in ELB + +, and the localization and intensity of FLAG-topo IIα on the lump were analyzed.
Data collection and processing
Experiments were performed on a total internal reflection fluorescence microscope (Nikon Ti) and images were acquired every 0.5 or 5 s using a CCD camera (Andor DU-888 X-7952) under the control of NIS software (NIS-Elements AR 4.40 and NIS-Elements AR Analysis 5.40.01). All images were analyzed by Fiji (Image J ver 1.52p). The intensities of condensin I and topo IIα on all U-shape DNA were measured, and only those with intensities equivalent to a single molecule were used for the analyses of lump formation. Lumps and loops on U-shape DNA were classified by visual observation. The difference in shape between loops and lumps was also determined as follows. A threshold was set to distinguish loops and lumps from other parts of DNA, and these regions were selected using the wand tool of Fiji. An ellipse was then fitted to each selected region, and the ratio of the major axis to the minor axis was measured. Measurements were taken every 5 s for 60 s following the initial binding of condensin I to the DNA. Lump stability was defined as the time from the initiation of lump formation to either the disappearance of the lump or the end of the observation (up to 20 min) under a flow rate of 20 μl/min. The fluorescent intensities of DNA-bound condensin I and topo IIα were measured every 5 s, and the background was subtracted. The fluorescence intensitiy (30–50 s after the initial binding of condensin I to the DNA, as shown in Fig. 2a) was measured to estimate the number of condensin I complexes and topo IIα dimers forming each lump. The distribution of fluorescence intensities was fitted using a five-component Gaussian mixture model, with mean and variance corresponding to 1–5 fluorescent molecules, as determined by bleaching experiments. We also simulated the expected distribution of fluorescence intensities, based on the labeling efficiency, assuming different ratios of molecule numbers. All calculations and visualizations were performed using Microsoft® Excel® for Microsoft 365 MSO (ver. 2412), GraphPad Prism (Prism 9.5.1), Anaconda (conda 24.11.3), and Spyder software (Spyder IDE 5.5.1). For fitting, the norm function from the scipy.stats package in the Scipy library and the GaussianMixture function from from sklearn.mixture package in Scikit-learn library were used (https://doi.org/10.5281/zenodo.15789118). Visualization was performed using the Matplotlib library. The lump size was calculated by normalizing the intensity of lumps to the intensity of full-length λDNA and converting it to DNA length by multiplying by 48.5 kbp. The tip of U-shape DNA, where a lump is typically formed, exhibits naturally higher DNA intensity compared to the dsDNA in the side arm of U-shape DNA. Hence, the size of the tip without protein was also calculated for comparison. The lump and loop formation rates were determined based on the average growth rate as long as they continued to increase. Relative DNA length was calculated by dividing the DNA length at each frame by the average DNA length from the first five frames taken before protein injection. To determine the fold increase in lump intensities, line profiles were taken across the DNA cross-sections at the tip regions with or without lump, and at the side arms. From the resulting line profiles, the relative sum intensities (Area Under the Curve, AUC) and the relative maximum DNA intensities (Max) were calculated by subtracting background signals from the integrals of the tip and side arm DNA profiles. All images shown in the figures were background-subtracted, smoothed, and resized for clarity.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data analyzed in this study will be made available by the authors upon request. Source data are provided with this paper.
Code availability
The Python-based data analysis code is available at https://doi.org/10.5281/zenodo.1578911850.
Change history
27 August 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41467-025-63490-3
References
Hirano, T. Condensin-based chromosome organization from bacteria to vertebrates. Cell 164, 847–857 (2016).
Paulson, J. R., Hudson, D. F., Cisneros-Soberanis, F. & Earnshaw, W. C. Mitotic chromosomes. Semin. Cell Dev. Biol. 117, 7–29 (2021).
Earnshaw, W. C., Halligan, B., Cooke, C. A., Heck, M. M. & Liu, L. F. Topoisomerase II is a structural component of mitotic chromosome scaffolds. J. Cell Biol. 100, 1706–1715 (1985).
Gasser, S. M., Laroche, T., Falquet, J., Boy de la Tour, E. & Laemmli, U. K. Metaphase chromosome structure. Involvement of topoisomerase II. J. Mol. Biol. 188, 613–629 (1986).
Saitoh, N., Goldberg, I. G., Wood, E. R. & Earnshaw, W. C. ScII: an abundant chromosome scaffold protein is a member of a family of putative ATPases with an unusual predicted tertiary structure. J. Cell Biol. 127, 303–318 (1994).
Maeshima, K. & Laemmli, U. K. A two-step scaffolding model for mitotic chromosome assembly. Dev. Cell 4, 467–480 (2003).
Shintomi, K., Takahashi, T. S. & Hirano, T. Reconstitution of mitotic chromatids with a minimum set of purified factors. Nat. Cell Biol. 17, 1014–1023 (2015).
Shintomi, K. & Hirano, T. Guiding functions of the C-terminal domain of topoisomerase IIalpha advance mitotic chromosome assembly. Nat. Commun. 12, 2917 (2021).
Nitiss, J. L. DNA topoisomerase II and its growing repertoire of biological functions. Nat. Rev. Cancer 9, 327–337 (2009).
Vos, S. M., Tretter, E. M., Schmidt, B. H. & Berger, J. M. All tangled up: how cells direct, manage and exploit topoisomerase function. Nat. Rev. Mol. Cell Biol. 12, 827–841 (2011).
Pommier, Y., Nussenzweig, A., Takeda, S. & Caroline, A. Human topoisomerases and their roles in genome stability and organization. Nat. Rev. Mol. Cell Biol. 23, 407–427 (2022).
Kimura, K. & Hirano, T. ATP-dependent positive supercoiling of DNA by 13S condensin: a biochemical implication for chromosome condensation. Cell 90, 625–634 (1997).
Kimura, K., Valentin, V. R., Crisona, N. J., Hirano, T. & Cozzarelli, N. R. 13S Condensin actively reconfigures DNA by introducing global positive Writhe: implications for chromosome condensation. Cell 98, 239–248 (1999).
Kimura, K., Hirano, M., Kobayashi, R. & Hirano, T. Phosphorylation and activation of 13S condensin by Cdc2 in vitro. Science 282, 487–490 (1998).
Ganji, M. et al. Real-time imaging of DNA loop extrusion by condensin. Science 360, 102–105 (2018).
Kong, M. et al. Human condensin I and II drive extensive ATP-dependent compaction of nucleosome-bound DNA. Mol. Cell 79, 99–114 (2020).
Goloborodko, A., Imakaev, M. V., Marko, J. F. & Mirny, L. Compaction and segregation of sister chromatids via active loop extrusion. Elife 5, e14864 (2016).
Postow, L., Crisona, N. J., Peter, B. J., Hardy, C. D. & Cozzarelli, N. R. Topological challenges to DNA replication: Conformations at the fork. Proc. Natl. Acad. Sci. USA 98, 8219–8226 (2001).
Baxter, J. et al. Positive supercoiling of mitotic DNA drives decatenation by topoisomerase II in eukaryotes. Science 331, 1328–1332 (2011).
Dyson, S., Segura, J., Martinez-Garcia, B., Valdes, A. & Roca, J. Condensin minimizes topoisomerase II-mediated entanglements of DNA in vivo. EMBO J. 40, e105393 (2021).
Kawamura, R. et al. Mitotic chromosomes are constrained by topoisomerase II-sensitive DNA entanglements. J. Cell Biol. 188, 653–663 (2010).
Hildebrand, E. M. et al. Mitotic chromosomes are self-entangled and disentangle through a topoisomerase-II-dependent two-stage exit from mitosis. Mol. Cell 84, 1422–1441 (2024).
Baxter, J. & Aragon, L. A model for chromosome condensation based on the interplay between condensin and topoisomerase II. Trends Genet 28, 110–117 (2012).
Sakata, R. et al. Opening of cohesin’s SMC ring is essential for timely DNA replication and DNA loop formation. Cell Rep. 35, 108999 (2021).
Kinoshita, K. et al. A loop extrusion-independent mechanism contributes to condensin I-mediated chromosome shaping. J. Cell Biol. 221, e202109016 (2022).
Tane, S. et al. Cell cycle-specific loading of condensin I is regulated by the N-terminal tail of its kleisin subunit. Elife. 11, e84694 (2022).
Berger, J. M., Gamblin, S. J., Harrison, S. C. & Wang, J. C. Structure and mechanism of DNA topoisomerase II. Nature 379, 225–232 (1996).
Liu, Q. & Wang, J. C. Identification of active site residues in the “GyrA” half of yeast DNA topoisomerase II. J. Biol. Chem. 273, 20252–20260 (1998).
Martinez-Garcia, B. et al. Condensin pinches a short negatively supercoiled DNA loop during each round of ATP usage. EMBO J. 42, e111913 (2023).
Davidson, I. F. et al. DNA loop extrusion by human cohesin. Science 366, 1338–1345 (2019).
Pradhan, B. et al. The Smc5/6 complex is a DNA loop-extruding motor. Nature 616, 843–848 (2023).
Ryu, J. K. et al. The condensin holocomplex cycles dynamically between open and collapsed states. Nat. Struct. Mol. Biol. 27, 1134–1141 (2020).
Golfier, S., Quail, T., Kimura, H. & Brugues, J. Cohesin and condensin extrude DNA loops in a cell cycle-dependent manner. Elife 9, e53885 (2020).
Shaltiel, I. A. et al. A hold-and-feed mechanism drives directional DNA loop extrusion by condensin. Science 376, 1087–1094 (2022).
McClendon, A. K. et al. Bimodal recognition of DNA geometry by human topoisomerase II alpha: preferential relaxation of positively supercoiled DNA requires elements in the C-terminal domain. Biochemistry 47, 13169–13178 (2008).
Wu, M. et al. Human topoisomerase iialpha promotes chromatin condensation via a phase transition. bioRxiv, https://doi.org/10.1101/2024.10.15.618281 (2024).
Dekker, C., Christian, H. H., Peters, J.-M. & Rowland, B. D. How do molecular motors fold the genome?. Science 382, 646–648 (2023).
Davidson, I. F. et al. Cohesin supercoils DNA during loop extrusion. Cell Rep. 44, 115856 (2025).
Janissen, R. et al. All eukaryotic SMC proteins induce a twist of -0.6 at each DNA-loop-extrusion step. Sci. Adv. 10, eadt1832 (2024).
Kawano, S. et al. DNA-binding activity of rat DNA topoisomerase II alpha C-terminal domain contributes to efficient DNA catenation in vitro. J. Biochem. 159, 363–369 (2016).
Nielsen, C. F., Zhang, T., Barisic, M., Kalitsis, P. & Hudson, D. F. Topoisomerase IIalpha is essential for maintenance of mitotic chromosome structure. Proc. Natl. Acad. Sci. USA 117, 12131–12142 (2020).
Meijering, A. E. C. et al. Nonlinear mechanics of human mitotic chromosomes. Nature 605, 545–550 (2022).
Kinoshita, K., Kobayashi, T. J. & Hirano, T. Balancing acts of two HEAT subunits of condensin I support dynamic assembly of chromosome axes. Dev. Cell 33, 94–106 (2015).
Marko, J. F. & Siggia, E. D. Stretching DNA. Macromolecules 28, 8759–8770 (1995).
Bouchiat, C. et al. Estimating the persistence length of a worm-like chain molecule from force-extension measurements. Biophys. J. 76, 409–413 (1999).
Wang, M. D., Yin, H., Landick, R., Gelles, J. & Block, S. M. Stretching DNA with optical tweezers. Biophys. J. 72, 1335–1346 (1997).
Patkowski, J. B. et al. The F-pilus biomechanical adaptability accelerates conjugative dissemination of antimicrobial resistance and biofilm formation. Nat. Commun. 14, 1879 (2023).
Shintomi, K. & Hirano, T. A sister chromatid cohesion assay using xenopus egg extracts. Methods Mol. Biol. 1515, 3–21 (2017).
Shintomi, K. & Hirano, T. Reconstitution of mitotic chromatids in vitro. Curr. Protoc. Cell Biol. 79, e48 (2018).
YukoTsubota YukoTsubota/condtopo: Source code of paper (ver1.01).Zenodo ta. https://doi.org/10.5281/zenodo.15789119 (2025).
Acknowledgements
We are grateful to Katsunori Fujiki and Katsuhiko Shirahige for technical support in the use of the optical tweezer system. We also thank Keiko Kimura and Mai Shimoura for the preparation of the flow-cell devices, Ryota Sakata and Yoshimi Kinoshita for technical advice on microscopic work, and Makoto M. Yoshida for discussions. The work in the authors’ laboratory was supported by JSPS Grants-in-Aid for Scientific Research/KAKENHI (#19H05755 and #22H02551 to K.S.; #23K05649 to K.K.; #18H05276 and #20H05938 to T.H.; #20H05933 and #20H05937 to T.N.). Y.T was supported by Grant-in-Aid for JSPS Fellows (#24KJ1514).
Author information
Authors and Affiliations
Contributions
Y.T., K.S., K.K., T.H., and T.N. designed experiments. Y.T. performed the single-molecule experiments. K.S., K.K., Y.M.-N., Y.A., and M.S. purified topo IIα and condensin I complexes. Y.T., K.S., and T.N., analyzed data. Y.T., T.H., and T.N. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tsubota, Y., Shintomi, K., Kinoshita, K. et al. Functional interplay between condensin I and topoisomerase IIα in single-molecule DNA compaction. Nat Commun 16, 7239 (2025). https://doi.org/10.1038/s41467-025-62600-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-62600-5
This article is cited by
-
Motorized chromosome models of mitotic chromosome folding
Nature Communications (2025)