Abstract
Obesity-related conditions are among the leading causes of preventable death and are increasing in prevalence worldwide. Body size and composition are complex traits that are challenging to characterize due to environmental and genetic influences, longitudinal variation, heterogeneity between sexes, and differing health risks based on adipose distribution. Here, we construct a 4-factor genomic structural equation model using 18 measures, unveiling shared and distinct genetic architectures underlying birth size, abdominal size, adipose distribution, and adiposity. Multivariate genome-wide associations reveal the adiposity factor is enriched specifically in neural tissues and pathways, while adipose distribution is enriched more broadly across physiological systems. In addition, polygenic scores for the adiposity factor predict many adverse health outcomes, while those for body size and composition predict a more limited subset. Finally, we characterize the factors’ genetic correlations with obesity-related traits and examine the druggable genome by constructing a bipartite drug-gene network to identify potential therapeutic targets.
Similar content being viewed by others
Introduction
Human body size and body composition vary throughout an individual’s lifecourse and among individuals in a population. The strong associations linking excess fat stores with a constellation of morbidities have highlighted the importance of understanding how various anthropometric traits are connected to the broad and multifaceted biological systems underpinning human health. Obesity prevalence has increased markedly in the United States between 1999 and 2020 from 30.5% to 41.9%1. On a global scale, the increasing rates of obesity observed among children and adults are a widespread source of concern2; obesity-related conditions such as heart disease, stroke, type 2 diabetes (T2D), and some cancers are among the leading causes for preventable death3. Although family-based studies2 and genome-wide association studies (GWASs)4 point to substantive genetic influences on obesity, the broader landscape of what characterizes this genetic signal across different measures of adiposity remains poorly understood.
The phenotypic and genetic signal of adiposity traits is remarkably difficult to characterize due to heterogeneity between sexes and longitudinal variation across the lifespan5. The genetic architecture for adipose distribution is notably different between males and females6, and women exhibit a greater ratio of subcutaneous-to-visceral adipose tissue than men7. Moreover, the amount of visceral adipose tissue tends to increase with age for both males and females, but men tend to lose relatively more visceral adipose tissue due to calorie restriction than women7,8,9. Body mass index (BMI) – an easily obtainable clinical measure (diagnosing obesity as BMI ≥ 30 kg/m2) – falls short when differentiating between masses of visceral adipose, subcutaneous adipose, muscle, or bone, leading to its criticism as a misleading metric of body composition and cardiometabolic health10,11,12. Waist circumference adjusted for BMI (WCadjBMI), hip circumference adjusted for BMI (HCadjBMI), and waist-to-hip circumference ratio adjusted for BMI (WHRadjBMI)6 are proxy measures of body fat distribution. Notably, the genetic drivers of BMI and WHRadjBMI are distinct: genetic associations for BMI and obesity are linked to enriched gene expression in the central nervous system (CNS), implicating a relationship between obesity and the brain2,6,13,14, whereas genes associated with WHRadjBMI demonstrate less enrichment for tissue-specific expression in the CNS and more with gene expression in preadipocytes and adipocytes6,15. Similarly, the genetic contributors to metabolic syndrome (MetSyn) – a cluster of often comorbid risk factors (e.g., hypertension, elevated triglycerides, and hyperglycemia) that link adiposity with cardiovascular disease and T2D – strongly overlap with the genetic associations for waist circumference (WC)16. However, the alleles associated with a higher subcutaneous-to-visceral adipose distribution (increased capacity for adipose tissue expansion)17 are protective for T2D, heart disease, and high blood pressure18,19. These findings highlight the complexity of body composition and genetic influences, with sometimes contrasting effects on health outcomes.
Given this complex and intertwined landscape of anthropometric measurements, we speculated that the genetic associations for human body size and body composition would be more suitably represented as latent variables in a genomic structural equation modeling (Genomic SEM) framework20. Genomic SEM estimates how strongly the genetic associations of various observed traits are related to a number of underlying and unobserved genetic constructs (latent factors). It does so by estimating the strengths of the relationships (loadings) of each trait with the factors, which themselves can be related to one another (genetic correlations). A primary characteristic of Genomic SEM is its ability to include different sets of traits from various participant samples; this enabled us to incorporate a diverse range of anthropometric traits from across the lifespan and stratified by biological sex into the same statistical model. Through this modeling process, we balanced model complexity and parsimony to unveil the shared versus distinct genetic components underlying differences in birth size, abdominal size, body size/composition, and adiposity. We found the enrichment of biological pathways and tissue types to be distinct among the 4 genetic factors in the model, and the factors showed different associations with adverse health outcomes in an independent dataset with electronic medical records. In addition, we contextualized the genome-wide signal for each of the factors by identifying differing patterns of genetic correlations with behavioral and obesity-related traits. Together, our results particularly highlighted the adiposity genetic factor for its distinct enrichment in nervous systems, substantial genetic correlations with related traits, and predictive capability for adverse health outcomes across broad phenotypic domains. Finally, we examined the druggable genome and constructed a bipartite drug-gene network to identify possible mechanistic explanations for weight-related side effects and the potential for repurposing therapeutics to address adiposity.
Results
A four-factor model of anthropometric and adiposity genetics
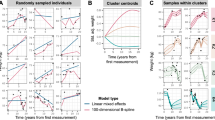
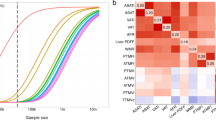
We began by bringing together GWAS summary statistics for 18 adiposity and anthropometric measures from different points in the lifespan and stratified by sex (Supplementary Table 1). The Genomic SEM model in Fig. 1 revealed an overall structure with 4 latent genetic factors referred to as F1-F4 and had adequate model fit21,22 (comparative fit index [CFI] = 0.94 and a standardized root mean square residual [SRMR] = 0.11). The genetic covariance and correlation matrices are shown in Supplementary Figs. 1–2 and Supplementary Data 61–64 along with further description of the modeling techniques and considerations in the methods section. The model estimated differing strengths of relationships between the 18 genetic indicator variables and their underlying latent constructs, as represented by their factor loadings (Fig. 1 one-directional arrows). F1 included 3 loadings for traits related to birth size, F2 included 3 loadings for traits relating to abdominal size, F3 included 7 loadings for traits relating to body size and adipose distribution, and F4 included 7 loadings for traits relating to adiposity. These latent genetic variables (F1-F4) represent the shared genetic effects underlying a cluster of genetically similar traits (e.g., F1 is an unobserved variable that captures the genetic influences underlying a set of observed traits relating to birth size). The 4 factors generally exhibited small genetic correlations (Fig. 1; |rg | ≤ 0.15 among the between-factor standardized covariance relationships indicated by two-directional arrows above the factors). The only sizable genetic correlation was for F1 and F3 (rg = 0.44), likely reflecting the shared genetic effects of birth length and adult height (rg = 0.49). Together, this emphasized the unique subclusters of genetic signal across traits relating to anthropometry and adiposity. Our analysis incorporated direct replication of the genomic structural equation model’s 4-factor structure through an exploratory factor analysis (EFA) using odd chromosomes followed by a confirmatory factor analysis (CFA) using even chromosomes to serve as a hold-out sample and protect from model overfitting. Within this replicated 4-factor model structure, only indicator variables with substantial loadings were permitted to load onto factors (representing considerable non-zero genetic covariances among indicator variables loading onto the same factor). This ensured that the associations that we identified in each factor’s multivariate GWAS were representative of shared effects across the indicator variables for that factor, even if those shared genetic effects were not large enough to be detected in the original trait specific GWASs.
The standardized measurement model derived using genomic structural equation modeling (SEM) comprised of 4 latent genetic factors and 18 indicator variables. The 4 genetic factors are shaded yellow, the traits with combined males and females are shaded gray, and the traits stratified by males and females are shaded in color-matched pairs. The one-directional arrows signify standardized factor loadings and describe the strength and direction of the relationships between genetic indicators and their underlying latent constructs. Standardized covariance relationships (i.e., correlations) between the factors are represented by two-directional arrows, and the two-directional arrows pointing from a variable to itself denote the standardized residuals (the unique genetic variance not reflected through other paths in the model). Uncertainties in standardized parameter estimates are indicated by standard errors provided in parentheses.
Among the 6 sex-stratified traits, each male-female pair generally loaded onto the same factor, highlighting the largely shared genetic associations within males and females. HCadjBMI male and female had similar loadings on F3, and BMI male and female had similar loadings on F4 – however, across the other sex-stratified traits there were some notable differences. More genetic variance of WHRadjBMI was explained by F2 in females relative to males (see loadings in Fig. 1), and F4 explained more variance of female than male arm fat ratio (AFR). In addition, the variance in female trunk fat ratio (TFR) was mostly explained by F3, but male TFR had modest cross-loadings between F3 and F4, with substantial residual genetic variance (0.83) and generally low genetic covariance (Supplementary Fig. 1) with other anthropometric traits, suggesting a more divergent genetic influence on male TFR. WCadjBMI female cross-loaded substantially onto both F2 and F3, while WCadjBMI male only loaded on F3. One primary advantage of our SEM is its ability to estimate these sex-specific differences and relationships within the landscape of anthropometric traits across the lifespan. The 4 factors in our model provide latent constructs that are less prone to measurement error and can discern the genetic components relating to body size and body composition; as such, this valuable genetic representation goes beyond any single indicator variable, such as BMI.
We subsequently used our 4-factor Genomic SEM to perform multivariate GWASs, which leveraged improved power over the constituent indicator GWASs. We identified multiple genome-wide significant (GWS; p < 5 × 10−8) variants that were unique to each factor and were not identified in the underlying GWASs after removing SNPs with heterogeneous effects (QSNPs; Supplementary Table 2). F1, F2, F3, and F4 respectively uncovered 103; 1318; 8; and 6206 GWS SNPs within the 7; 35; 1; and 139 independent loci that were not identified by the individual indicator GWASs for each factor. Manhattan plots for each multivariate factor-GWAS are shown in Supplementary Figs. 3–6. Supplementary Table 2 summarizes the number of independent association signals for each factor and how many were novel relative to each factor’s indicator traits. The lack of independent datasets for all 18 indicator variables precluded us from performing a formal replication analysis of the novel loci which would require constructing a comparable Genomic SEM using independent data and performing 4 corresponding multivariate GWASs. However, within the independent All of Us dataset23,24 there were GWASs for 3 traits that were indicator variables for F2 (WHRadjBMI; N = 102,746), F3 (height; N = 111,755), and F4 (BMI; N = 111,482). We used these independent GWASs to test for concordant statistical significance and consistent direction of effect for the novel identified loci, though we anticipated much reduced power for a single indicator compared to our multivariate factors20. See the methods section for further details. Complete summary statistic information is summarized in Supplementary Data 41–43 for the lead SNPs of novel loci, and an overview of the loci for each factor is included in Supplementary Table 2. Both of the 2 novel loci for F2 (relative to WHRadjBMI), showed consistent direction of effect, but neither had concordant significance. The 2 novel loci for F3 had consistent direction of effect when compared to the All of Us height GWAS, and 1 of the 2 loci had concordant significance. For F4, there were 28 novel loci relative to BMI, 25 of which had consistent effect direction (binomial test p = 1.52 × 10−6) and 5 of which had concordant significance. This comparison between our multi-variate GWASs and independent univariate GWASs highlighted broadly consistent effect directions and a couple novel loci had evidence for concordant significance in All of Us, providing a confirmatory context for our multivariate factor GWASs’ novel associations. In Supplementary Note 1 we have outlined certain aspects of the modeling process that guard against false positives to provide additional context for these multivariate GWAS associations.
We next characterized these multivariate GWASs in multiple downstream analyses. First, we implemented DEPICT25 to identify significantly prioritized genes (false discovery rate [FDR] <0.05) from the 88; 344; 1173; and 675 independent GWS loci for F1, F2, F3, and F4 respectively, and assessed the enrichment of those loci across functional gene sets (p < 4.56 × 10−6, the Bonferroni-corrected significance threshold) and tissue-specific expression profiles (FDR < 0.05). Next, we used FOCUS26 to perform transcription-level analyses (transcriptome wide association studies [TWASs]) for each of the latent factor GWASs, and we extracted genes that were fine-mapped to non-null 90% credible sets (CSs) with a posterior inclusion probability (PIP) > 0.1. Gene set overlap across the GWASs and TWASs is shown in Supplementary Fig. 7. We then used the factors’ multivariate GWAS effect estimates and LDpred227 to develop 4 polygenic risk scores (PGSs) and applied them to an external dataset (Colorado Center for Personalized Medicine [CCPM] Biobank freeze2; N = 25,240). These PGSs were tested for association with 1514 phecode-based phenotypes (FDR < 0.10 Bonferroni-corrected significance threshold, due to the highly correlated structure of the phecodes) in a phenome-wide association study (pheWAS). Next, we estimated the genetic correlations with comorbidity-related traits to contextualize each factor within a broader genomic landscape using Linkage Disequilibrium Score Regression (LDSC)28,29. Finally, we constructed drug-gene interaction networks for the factors’ DEPICT- and FOCUS-identified genes to advance existing, proposed, and novel therapeutic targets for adiposity-related conditions.
F1 – birth size
F1 characterized the genetic signal underlying size at birth with loadings from 3 indicator variables (Fig. 2a). The DEPICT analysis highlighted 88 independent GWS loci with 24 significantly prioritized genes and 3 enriched gene sets including ‘incomplete somite formation’ and ‘decreased embryo size’ gene sets. The GWS loci for F1 were not enriched for expression profiles across physiological systems, cell types, or tissue types (Fig. 2b). In a tissue-agnostic TWAS analysis using FOCUS, however, we identified 158 fine-mapped genes with PIP > 0.1 across 69 non-null CSs (Supplementary Fig. 8). These putatively causal gene-expression mediated effects consisted of SNP-expression weights from 27 general tissues including the brain (43 genes), adipose (17 genes), and esophagus (16 genes). The F1 PGSs that were applied in an external dataset (N = 25,240) were negatively associated with acute sinusitis, insomnia, renal failure, T2D, and hypertension (Fig. 2c). The F1 GWAS, DEPICT enrichment, TWAS, and pheWAS results are summarized in Supplementary Data1–3, 16–20, 44–46, and 56.
The 3 indicator variables relating to birth size and their standardized loadings on F1, the 1st latent genetic factor, are shown in (a). This genetic factor did not have any genetic enrichment across physiological systems, cell types, or tissue types (FDR < 0.05) in the DEPICT enrichment analysis (b). The polygenic score (PGS) weights for F1 were applied in an external sample (CCPM Biobank, N = 25,240) and implemented in a phenome wide association study (pheWAS); The significant logistic regression pheWAS associations between F1 PGS and phenotypes are shown in (c), with phenotype labels for the points to the right of the vertical dashed red line denoting the FDR < 0.10 Bonferroni-corrected significance threshold, and triangle direction (up/down) indicating F1 PGS direction of effect (+/−). Source data are provided as a Source Data file.
F2 – abdominal size
F2 had 3 loadings from indicator variables relating to adult abdominal size (Fig. 3a) and 319 significant DEPICT-prioritized genes from 344 independent GWS loci. We observed significant physiological system enrichment across 7 of the 10 categories (Fig. 3b), including adipocytes, subcutaneous adipose tissue, and abdominal adipose tissue. Beyond those adipose-related tissues, F2’s genetic signal was broadly enriched throughout the body including the musculoskeletal, urogenital, cardiovascular, digestive, and endocrine systems (Supplementary Data 24). Using tissue-agnostic FOCUS TWAS we identified 676 fine-mapped genes with PIP > 0.1 across 243 non-null CSs (Supplementary Fig. 9). These prioritized TWAS associations spanned 28 general tissues but primarily consisted of brain (160 genes) and adipose tissue weights (78 genes). F2 PGS-pheWAS showed positive associations with T2D, peripheral angiopathy, and hypertension (Fig. 3c) suggesting a genetic propensity for larger abdominal size was predictive of these circulatory and metabolic health outcomes. These phenotypic associations were aligned with the DEPICT gene-set analysis which identified 185 significantly enriched gene sets relating to insulin resistance and organ development/morphology (particularly within the cardiovascular system). The F2 GWAS, DEPICT enrichment, TWAS, and pheWAS results are summarized in Supplementary Data 4-6, 21–25, 47–49 and 57.
The 3 indicator variables relating to abdominal size and their standardized loadings on F2, the 2nd latent genetic factor, are shown in (a). This genetic factor showed gene expression enrichment across a variety of physiological systems, cell types, and tissue types (orange coloring, FDR < 0.05) in the DEPICT enrichment analysis (b). The polygenic score (PGS) weights for F2 were applied in an external sample (CCPM Biobank, N = 25,240) and implemented in a phenome wide association study (pheWAS); The significant logistic regression pheWAS associations between F2 PGS and phenotypes are shown in (c), with phenotype labels for the points to the right of the vertical dashed red line denoting the FDR < 0.10 Bonferroni-corrected significance threshold, and triangle direction (up/down) indicating F2 PGS direction of effect (+/−). Source data are provided as a Source Data file.
F3 – body size and adipose distribution
The third genetic factor, F3, captured the shared variance among 7 indicator variables describing body size and adipose distribution (Fig. 4a), with notable differences between the loadings for male and female traits, especially for TFR (described above). The DEPICT analysis for F3 identified 1864 significantly prioritized genes for 1173 independent GWS loci and enrichment in 8 of the 10 physiological system categories (Fig. 4b; musculoskeletal, urogenital, cardiovascular, endocrine, digestive, respiratory, hemic and immune, integumentary), exemplifying the multifaceted physiology underlying variation in adult body size and adipose distribution (Supplementary Data 29). We found 1127 gene sets significantly enriched for F3, including many gene sets relating to embryonic development and protein‑protein interaction subnetworks. In a tissue-agnostic FOCUS TWAS, we identified 2266 fine-mapped genes with PIP > 0.1 across 689 non-null CSs (Supplementary Fig. 10), spanning 28 general tissues, particularly brain (571 genes), esophagus (242 genes), adipose (218 genes), and artery (202 genes). Interestingly, APOE, a gene linked to Alzheimer’s disease and catabolism of lipoprotein constituents, was significantly associated via prostate expression weights (Z-score = −5.35, PIP = 0.61). The PGS-pheWAS analysis revealed that F3 was predictive of a few health outcomes including negative associations with abdominal pain, hyperlipidemia, and hypertension, but a positive association with atrial fibrillation (Fig. 4c). The F3 GWAS, DEPICT enrichment, TWAS, and pheWAS results are summarized in Supplementary Data 7–9, 26–30, 50–52, and 58.
The 7 indicator variables relating to body size and adipose distribution and their standardized loadings on F3, the 3rd latent genetic factor, are shown in (a). This genetic factor showed gene expression enrichment across a variety of physiological systems, cell types, and tissue types (FDR < 0.05) in the DEPICT enrichment analysis (b). The polygenic score (PGS) weights for F3 were applied in an external sample (CCPM Biobank, N = 25,240) and implemented in a phenome wide association study (pheWAS); The significant logistic regression pheWAS associations between F3 PGS and phenotypes are shown in (c), with phenotype labels for the points to the right of the vertical dashed red line denoting the FDR < 0.10 Bonferroni-corrected significance threshold, and triangle direction (up/down) indicating F3 PGS direction of effect (+/−). Source data are provided as a Source Data file.
F4 – adiposity
F4 had 7 adiposity-related indicator variables loading onto it relating to excess fat tissue and obesity (Fig. 5a). The associated loci were enriched only in one physiological system (nervous; Fig. 5b, Supplementary Data 34). Broad regions across the CNS were enriched, including the hindbrain (cerebellum) and the forebrain (cerebral cortex, temporal lobe, occipital lobe, frontal lobe, parietal lobe, basal ganglia) – regions responsible for complex perceptual, cognitive, and behavioral processes involving learning, emotion, and memory. The F4 DEPICT analysis identified 437 significantly prioritized genes for the 675 independent GWS loci and 62 enriched gene sets; upon comparing these gene sets to the other 3 factors, they were much more specific to the CNS, relating to brain development, neurons, synaptosomes, and dendrites. In a brain-tissue-prioritized FOCUS TWAS, we identified 850 fine-mapped genes with PIP > 0.1 across 335 non-null CSs (Supplementary Fig. 11). These prioritized TWAS associations spanned 28 general tissues but the majority corresponded to brain tissue weights (498 genes). The PGS-pheWAS analysis for F4 uncovered many more associations with adverse health outcomes, spanning a wide range of domains (Fig. 5c): chronic pain, fatigue, asthma, shortness of breath, sleep apnea, benign skin neoplasm, cancer of kidney and renal pelvis, osteoarthritis, substance use disorders, anxiety, depression, sepsis, allergy to medications, skin/nail fungal infections, anemia, renal disease/failure, obesity, T2D, liver disease/cirrhosis, bariatric surgery, esophageal diseases, acid reflux, cellulitis, long-term anticoagulants, and hypertension. The F4 GWAS, DEPICT enrichment, TWAS, and pheWAS results are summarized in Supplementary Data 10−12, 31−35, 53−55, and 59.
The 7 indicator variables relating to adiposity and their standardized loadings on F4, the 4th latent genetic factor, are shown in (a). This genetic factor showed gene expression enrichment only in nervous physiological systems and cell types (FDR < 0.05) in the DEPICT enrichment analysis (b). The polygenic score (PGS) weights for F4 were applied in an external sample (CCPM Biobank, N = 25,240) and implemented in a phenome wide association study (pheWAS); The significant logistic regression pheWAS associations between F4 PGS and phenotypes are shown in (c), with phenotype labels for the points to the right of the vertical dashed red line denoting the FDR < 0.10 Bonferroni-corrected significance threshold, and triangle direction (up/down) indicating F4 PGS direction of effect (+/−). Source data are provided as a Source Data file.
Comparison of F4 and BMI genetic signals
The male and female BMI indicator variables both had large standardized loadings of 0.95 with F4; therefore, we explored the shared versus distinct aspects of the genetic signals for F4 (a highly predictive latent factor) compared to BMI. There were 6578 GWS SNPs common between the F4 and BMI GWASs, but 6206 SNPs that were novel to F4 (i.e., not GWS in any of the indicator GWASs loading onto F4, including BMI male and BMI female). Overall, the GWS SNPs for F4 and BMI (combined males and females) resided in 675 and 1035 independent significant loci, respectively, which only partially overlapped (624 of the 675 F4 loci had genomic positional overlap with the BMI loci; Supplementary Figs. 6, 12, and 13). Notably, while 392 DEPICT-prioritized genes were common to BMI and F4, 45 genes were unique to only F4 (Supplementary Data 13, 15, and 36–37; Supplementary Fig. 14). In addition, while 339 putatively causal genes with expression mediated effects (FOCUS-identified genes; Supplementary Figs. 14–15, Supplementary Data 14, 40) were common to BMI and F4, 511 genes were unique to F4. Only 21 genes were common to all 4 analyses (identified by DEPICT and FOCUS for both F4 and BMI). Beyond these distinguishing overlaps at the gene level, the DEPICT gene set and tissue enrichment analyses (Fig. 5b, Supplementary Fig. 16, Supplementary Data 33–34 and 38–39) pinpointed a key difference between F4 and BMI: the BMI-associated genetic loci were distinctively enriched for the hypothalamus and the hypothalamo hypophyseal system – the brain’s control center for hunger and satiety. The BMI-associated loci were therefore enriched in the canonical energy homeostasis-related areas of the brain whereas the F4-associated loci were not. Thus, F4 was characterized by a salient partitioning of the genetic architecture of adiposity; F4 disentangles a neural and behavioral component of adiposty that is rooted in sensory processing, learning, memory, and experience.
The genetic differences between F4 and BMI motivated us to perform an additional pheWAS controlling for BMI to investigate the conditionally independent associations of F4’s PGS with heath outcomes (Supplementary Fig. 17, Supplementary Data 60). We observed an attenuation of the F4-pheWAS associations, as expected, after conditioning on BMI (Supplementary Fig. 18); several health outcomes including chronic pain, sleep apnea, depression, and acid reflux dropped below the significance threshold, implicating BMI as a potential mediator for some disease associations30,31,32. However, F4 clearly captured additional and unique contributions to health outcomes beyond BMI alone, with F4 still positively and significantly predicting adverse health outcomes for the vast majority of associations after adjusting for BMI. These results illustrate the utility of F4 as a polygenic predictor beyond BMI, and they showcase the added value of our model for disentangling the genetics of adiposity and anthropometrics across the lifespan.
Genetic correlations with related traits
Following the characterization of each of the 4 factors with regard to their genome-, transcriptome-, and phenome-wide associations, we estimated LDSC-based genetic correlations between each factor and 75 related traits (Supplementary Table 3 and Supplementary Data 65), including metabolism, substance use, psychopathology, neuroticism, risk tolerance, diet, sleep, exercise, pain, frailty, dementia, inflammatory disease, autoimmune disease, and cardiovascular disease. The full genetic covariance and correlation matrices are shown in Supplementary Figs. 19–20 and Supplementary Data 66–69 (with 95% confidence intervals and standard errors), and pairwise genetic correlations with F1, F2, F3, and F4 are shown in Supplementary Figs. 21–30. Figure 6 depicts the prominent genetic correlations (|rg| > 0.15) with each of the factors in our Genomic SEM; F1 and F3 were the only factors with a notable inter-factor genetic correlation (rg = 0.44). The genetic link between F3 and atrial fibrillation recapitulated the F3 pheWAS result (Fig. 4c) highlighting the shared genetics underlying an association between taller stature and increased risk of atrial fibrillation33. In addition, F1’s genetic correlations mirrored the pheWAS results (Fig. 2c), exhibiting negative genetic correlations with cardiovascular traits and T2D. F2 had positive genetic correlations with the components of MetSyn, reflecting the F2 pheWAS associations with T2D and hypertension and emphasizing shared genetic influences on visceral adipose deposits and metabolic abnormalities34. F2 also had positive genetic correlations relating to substance use, internalizing behaviors, and frailty. F4 was again the most central factor in terms of the strength and quantity of genetic correlations, including positive correlations with metabolic disorders, pain, internalizing disorders, general risk-tolerance, attention-deficit hyperactivity disorder, substance use disorders, frailty, adult-onset asthma, coronary artery disease, and gout. F4 was negatively correlated with measures of fitness/exercise, compulsive disorders, high-density lipoprotein (HDL) cholesterol, alcohol consumption frequency, and sleep efficiency. Interesting and nuanced relationships emerged between adipose genetic factors and mental health traits: general neuroticism was more genetically correlated with F2 (rg = 0.18) compared to F4 (−0.01), but the depressed affect and worry subtypes of neuroticism were more genetically correlated with F4 (0.20 and −0.21, respectively) compared with F2 (0.12 and 0.10, respectively). Thus, we found opposite directionality of the genetic correlation between F4 and the neuroticism subtypes and also between F4 and internalizing disorders (e.g., anxiety disorders [rg = 0.12] and major depressive disorder [rg = 0.14]) versus compulsive disorders (e.g., obsessive compulsive disorder [rg = −0.25] and anorexia nervosa [rg = −0.27]). Together, this suggests that the relationship of adiposity and mental health outcomes depends in part on which aspect of body composition is evaluated, and in turn, the possible physiological and neurological systems involved.
A network with edges depicting the strength of genetic correlations between the 4 Genomic SEM factors and an array of genetically related traits. Small pairwise correlations ≤ 0.15 were pruned from the network highlighting this subset of 33 genetically correlated traits from a broader set of 75 considered traits (see Supplementary Table 3). The 4 factors had weak between-factor genetic correlations (besides F1 and F3) and their unique genetic signals are characterized by each factor’s grouping with a distinct, but not entirely exclusive, set of phenotypes. Trait names and abbreviations in alphabetical order: Alcohol Consumption Frequency (ACF); Attention-Deficit Hyperactivity Disorder (ADHD); Age of Smoking Initiation (AgeSI); Anorexia Nervosa (AN); Anxiety Disorders (ANX); Adult-Onset Asthma (AsAO); Atrial Fibrillation (AtrFib); Coronary Artery Disease (CAD); Cigarettes Per Day (CPD); Cannabis Use Disorder (CUD); Personality - Extraversion (Extrv); F1 - Birth Size (F1); F2 - Abdominal Size (F2); F3 - Body Size, Adipose Distribution (F3); F4 - Adiposity (F4); Fasting Glucose (FastGluc); Cardiorespiratory Fitness - Heart Rate (FitHR); Cardiorespiratory Fitness - VO2 Max (FitVO2); Frailty Index (Frail); High-Density Lipoprotein Cholesterol (HDL); Metabolic Syndrome (MetSyn); Neuroticism (Neur); Neuroticism - Depressed Affect (NeurD); Neuroticism - Worry (NeurW); Obsessive Compulsive Disorder (OCD); Pain - General (PainG); Pain - Musculoskeletal (PainM); Problematic Alcohol Use (PAU); Physical Activity (PhysA); General Risk-Tolerance (Risk); Systolic Blood Pressure (SBP); Smoking Cessation (SC); Smoking Initiation (SI); Sleep Efficiency (SleepE); Type 2 Diabetes (T2D); Triglycerides (Triglyc); Serum Urate - Gout (Urate). Source data are provided as a Source Data file.
Drug-gene network
Our final downstream analysis aimed to identify potential therapeutics that might ameliorate or prevent adipostiy by querying the significantly prioritized GWAS and TWAS genes across two drug-gene interaction databases (Drug Repurposing Hub [DRH]35 and Drug-Gene Interaction Database [DGIdb]36). We constructed a bipartite drug-gene network for each of the latent factors to assess the druggable genome in the context of our 4-factor model (Supplementary Data 70–77). Given the extensive phenotypic associations we observed for the PGS trained on the 4th factor (Fig. 5c), we primarily focused on F4’s 1239 DEPICT- or FOCUS-identified genes (Supplementary Data 1–2: 48 genes identified by both DEPICT and FOCUS, 389 genes identified by DEPICT only, and 802 genes identified by FOCUS only). Our bipartite network for F4 included 733 drug-gene pairs (90 identified by both DRH and DGIdb, 451 identified by DRH only, 192 identified by DGIdb only), consisting of 151 genes and 529 drugs with regulatory approval. Of these 529 drugs, a substantial number (148) had prior descriptions of weight-related adverse drug events (wADEs) in the OnSIDES database37. The 381 drugs without wADEs typically interacted with genes that were connected to drugs with known wADEs (Supplementary Figs. 31–35).
The drug-gene network (Fig. 7) had groups of drugs clustered around high-degree genes, and drugs that served as links between different modules. Upon annotation of these drug clusters, we identified parts of the network that were specific to psychiatry, neurology, cardiology, oncology, endocrinology, and gastroenterology illustrating the diversity of therapeutics with potential wADEs based on interactions with F4-associated target genes. This analysis identified drug-gene pairs for serotonergic (e.g., trazodone) and dopaminergic agents (e.g., quetiapine) – well-known psychiatric medication classes with wADEs, sulfonylureas – diabetes medications with known wADEs, and tirzepatide – a potent weight loss and diabetes medication that interacts with GIPR. In addition, the drug-gene network for F3 recapitulated the function of fenofibrate as a therapeutic for MetSyn components16 via interactions with two significant genes (SCARB1 and GCKR). These confirmatory results support the utility of our approach to identify novel and salient drug targets or existing drugs that might be repurposed to target adiposity. Moreover, genes interacting with drugs with known wADEs—e.g., antihistamines interacting with HRH1 – frequently interacted with numerous other medications of the same drug class, suggesting weight-related drug effects may be under-recognized among medications with a common mechanism of action. Our bipartite network results can also be used to explore direct mechanisms for the drug-induced bodyweight changes that are commonly listed as adverse side effects of treatment and are observed in routine clinical care. For example, olanzapine (a psychiatric drug for schizophrenia and bipolar disorder), interacts with the same gene target as tirzepatide – GIPR – and this could explain the adverse weight gain often associated with olanzapine administration38,39,40,41. In addition, the DEPICT GWAS identified muscarinic cholinergic receptor gene CHRM4 and the FOCUS TWAS identified histamine receptor gene HRH1 for F4 – these genes provide potential explanations for the wADEs of drugs that are used to treat mental disorders42 and antihistamine medications43. Similarly, the identification of several receptor tyrosine kinases as having potentially causal effects on adiposity from the DEPICT and FOCUS analyses provides a mechanistic explanation for the wADEs of tyrosine kinase inhibitors44. We also uncovered potentially high-impact drug-gene pairs that may inform studies of drug repurposing. One of the 45 genes that was identified by our DEPICT analyses for F4 but not for BMI was PDE5A on chromosome 12; this gene is targeted by dipyridamole (a medication used to prevent blood clots), which has been implicated as a potential therapeutic for weight loss via stimulating brown fat energy expenditure45.
The bipartite approved drug-gene network for significant genes in the GWAS DEPICT gene prioritization analysis or the TWAS FOCUS fine mapping analysis for F4 (the latent genetic factor relating to adiposity). For visualization of this network we removed drugs that did not have ‘launched’ clinical phase in the Drug Repurposing Hub (DRH) or ‘approved’ status in the Drug-Gene Interaction Database (DGIdb), for a total of 733 drug-gene pairs (451 identified in the DRH [purple edges], 192 identified in the DGIdb [orange edges], and 90 identified by both [red edges]) between 529 drugs and 151 genes significant for F4. The gene vertices are colored grey, and the drug vertices are colored by their most frequent indication category in the MEDI-C database. Drugs vertices with weight-related adverse drug events (wADEs) listed in the OnSIDES database have a black border. Source data are provided as a Source Data file.
Discussion
Our 4-factor structural equation model serves as an informative and parsimonious representation of the genetic relationships among anthropometrics and adiposity across the lifespan. While many different measurements aim to quantify aspects of body size and body composition, our approach using correlated latent factors is less prone to the measurement error introduced by a singular phenotype definition, such as BMI. Furthermore, our modeling approach leveraged the combined power across indicator GWASs to identify novel genomic associations and provided a comprehensive mapping of the genetic architecture underlying birth size, abdominal size, body size/composition, and adiposity. Our model highlighted differing genetic effects and loadings between males and females, and we characterized the distinct polygenic signals underlying each of the 4 genetic factors through various downstream analyses: multivariate GWASs, SNP-to-gene mapping, gene set enrichment, tissue enrichment, fine-mapped TWASs, PGS-based pheWASs, genetic correlations, and drug-gene interaction networks.
All of these analyses recapitulated the importance of F4, the adiposity factor, as the primary genetic culprit predisposing individuals to adverse health outcomes. Compared to the other 3 factors, F4 showed distinct enrichment for neuronal tissues and gene sets, stronger genetic correlations with related traits, broad health associations across numerous phenotypic domains, and relevant drug-gene pairings across diverse fields of medicine. Furthermore, F4 showed distinct genetic signal compared to BMI. The link between F4 and substance use traits is further accentuated by our identification of GIPR and tirzepatide in the drug-gene network because of the growing evidence for GIP and GLP-1 receptor agonists as potential anti-addiction treatments (beyond their primary indication for diabetes and weight loss)46,47. In the context of our ongoing search for more effective treatments, F4 provided possible mechanistic explanations for weight-related side effects across many medications and identified the potential for repurposed therapeurics to address adiposity (e.g., dipyridamole, an antiplatelet medication, which has been shown to target inosine as a stimulant of energy expenditure in brown adipocytes)45,48. The findings from our downstream analyses triangulated F4’s close relationship with behavioral traits through disentangling the genetic architecture of adiposity; the neuronal and behavioral context of F4 emphasized that the genetic loci associated with increased adiposity are underlain by complex relationships with environmental and lifestyle influences. F4 implicated a broad and cascading network of adiposity-mediated diseases49 and the underlying physiology of excess fat storage50, adipokines (e.g., leptin and adiponectin)51,52, chronic inflammation from adipocyte apoptosis53, MetSyn16, and diabetes subtypes54,55.
Anthropometrics and adiposity across the lifespan have important health implications amidst a complex landscape of various patterns of inheritance (e.g., rare-vs-common genetic variants, high-vs-low penetrance, large-vs-small effect sizes)2 and diverse environmental contexts (e.g., food availability, physical activity, exposure to pollutants)56,57,58. The present analyses were limited to individuals of European ancestry, and future work will aim to characterize anthropometrics for additional ancestry groupings. In addition, our analyses share the strengths, assumptions, and limitations of the underlying methods including Genomic SEM20, LDSC28,29, DEPICT25, and FOCUS26. Another limitation to our study is the potential for collider bias among some of the indicator variables. Waist and hip measurements are often adjusted for BMI to be used as proxies for abdominal adipose deposition across the strata of overall body mass. However, the adjustment for BMI can result in biased genetic effects59, and this adjustment could have contributed to the low negative genetic correlations observed between F2 and F4 and between F3 and F4. In addition, uneven sample sizes and/or precision of effect sizes among indicator GWASs present an important consideration when interpreting Genomic SEMs. Indicator GWASs with large sample sizes tend to have more precise estimates of SNP heritability and genetic covariances, thereby influencing model structure, factor loading estimates, and power when estimating SNP effects in the multivariate GWASs20. The inclusion of multiple well-powered indicator GWASs in combination with precise phenotyping (bioelectrical impedance measurements) may explain why F4 produced a notable number of novel associations relative to its indicator variables. Our model broadly disentangled the genetic associations for size at birth (F1) from size in adulthood (F2, F3, and F4), however, it did not provide the same granularity as longitudinal growth trait analyses regarding genomic associations with anthropometrics across the lifespan. The observation that childhood BMI loaded onto F4 rather than F1 was consistent with a prior longitudinal study5 which identified strong overlap between the genetics of child and adult BMI, but differing genetic factors that control infant and child BMI. Extending these genetic insights into multi-omics60 frameworks will enable the identification of biological markers beyond the genome and further disentangle the etiology of adipose-related diseases. While F4 had the strongest and most widespread health implications, the other three genetic factors characterized important aspects of body size and adipose distribution, reflecting unique influences on additional health outcomes, including respiratory illness61, renal failure62, hypertension63, kidney stones64, T2D, and hyperlipidemia65,66,67. Future directions might involve further exploration of the negative pheWAS association for F3 with hyperlipidemia, especially in the context of F3’s evidence for sex differences regarding depot-specific genetic architectures of adipose distribution68.
Our model describing the genetic associations for variation in human body size and body composition across the lifespan recapitulates the notion that food intake is not merely an unconditioned response to an energy deficiency, nor is it restricted to the canonical energy homeostasis areas in the brain (e.g., the hypothalamus)69. Instead, the involvement of brain areas performing the functions of sensory processing, learning, emotion, and memory indicates a broader neuro-centric genetic relationship with obesity. In this context, this neural component carries significant influence on diverse health outcomes; and from a personalized medicine perspective, F4 has the promising capability to improve the prediction, diagnosis, treatment, and prevention of morbidities such as obesity, diabetes, adult persistent asthma, heart disease, chronic pain, substance use, and mental disorders.
Methods
Ethics
The Ethics Board at the University of Colorado Boulder deemed that institutional review board approval was not necessary for our analyses as GWAS summary data do not include individual-level results; the studies that published the incorporated summary statistics obtained written informed consent from participants and were approved by local ethics committees. Our study design and conduct complied with all relevant regulations regarding the use of human study participants and was conducted in accordance with the criteria set by the Declaration of Helsinki.
Genomic structural equation modeling
Structural equation modeling is a widely used methodology for understanding the correlation and covariance patterns of interconnected variables. The resulting models are useful for explaining the variance of measurable variables, latent variables, and the relationships between those latent variables70. We constructed an SEM describing the genetic associations of body size and body composition using a set of publicly available GWASs for various anthropometric traits. The measurement model that we constructed consisted of 18 individual GWAS summary statistics for 12 different phenotypes (described in Supplementary Table 1)15,71,72,73,74,75,76,77,78,79. Given our interest in investigating the sex-specific genetic architecture of body size and body composition, we included male and female GWASs independently for 6 of the 12 traits. The GWAS summary statistics were formatted using the munge function in the GenomicSEM R package after specifying the effect alleles, effect sizes, standard errors, and sample sizes for each dataset. All 18 GWASs passed heritability-based quality control (QC) with heritability Z-statistics > 4, signifying they were well powered and had measurable effects across 954,086 overlapping genetic variants. These GWASs were comprised of European ancestry populations and the corresponding SNP reference file and linkage disequilibrium (LD) scores and were downloaded from the Genomic SEM data repository (https://github.com/GenomicSEM/GenomicSEM).
The only binary trait included in the analysis was childhood obesity, which consisted of 9116 cases and 13,292 controls; because this GWAS was a meta-analysis of multiple cohorts, the sum of effective sample sizes was used along with a sample prevalence of 0.5 (per the Genomic SEM multivariable LDSC function guidelines) and a population prevalence of 0.20 for liability scale conversion72. We implemented the standard parameters for Genomic SEM, and QC criteria ensured the included SNPs were common (maf.filter = 0.01) and that the SNPs with lower imputation quality were removed from the analysis (info.filter = 0.9). When initially attempting to include all 3 bio-electrical impedance fat distribution GWASs (arm-fat-ratio [AFR], leg-fat-ratio [LFR], and trunk-fat-ratio [TFR]), the model showed poor fit and spurious standardized loadings greater than 1. This was due to the linear dependency among these 3 traits (the ratios of AFR, LFR, and TFR sum to 1, and therefore one ratio is predictable by the other two) which was problematic when inverting the sample covariance matrix in the process of computing the model estimates. LFR and TFR are inversely genetically correlated (rg < −0.9) and are largely representative of the same trait (i.e., the distribution of adipose between those two compartments)71. Given the well-established relationship between visceral adipose tissue and adverse health outcomes34, we retained TFR in the analysis, thereby omitting LFR.
We implemented Genomic SEM in a 2-stage modeling process to fit an SEM to the genetic association estimates20. We used multivariate LDSC28,29 to construct the genetic covariance (SLDSC) and sampling covariance (\({V}_{{S}_{{LDSC}}}\)) matrices for the 18 GWAS summary statistics. Then, we fit an SEM using diagonally weighted least squares (DWLS) estimation. An important feature of Genomic SEM is that it is designed to handle varying degrees of sample overlap among the incorporated GWASs.
We first performed an exploratory factor analysis (EFA) by using odd chromosomes then a confirmatory factor analysis (CFA) using the even chromosomes to serve as a hold-out sample and protect from model overfitting. We used the Kaiser rule80, the acceleration factor, and optimal coordinates criteria81 to assess the EFA and determine which eigenvalues of the genetic covariance matrix were most pronounced; all 3 criteria indicated that specifying 4 latent factors was a judicious choice for the SEM. The factanal R package was used to perform a promax (i.e., correlated factor) rotation preceding the estimation of the unstandardized and standardized loadings from the nearest positive definite genetic covariance matrix via the nearPD function from the matrix R package. Variables with standardized loadings greater than 0.3 were specified to load onto each of the 4 latent factors, and the model structure was notably consistent for any threshold choice between 0.3 and 0.5. Heywood cases were handled for indicator variables with loadings close to 1 by constraining the residuals to be greater than 0.0001. The resulting fit of the SEM was evaluated using the comparative fit index (CFI) and the standardized root mean square residual (SRMR). Generally, CFI > 0.9 and SRMR < 0.1 are indicative of acceptable model fit for Genomic SEM models21,22. WCadjBMI females and TFR males showed notable genetic correlations with indicator variables loading onto the 3rd and 4th factors, respectively; including these cross-loadings improved model fit and resolved warnings regarding the covariance matrix of the residuals of the observed variables being non-positive definite. Ultimately, the CFA showed consistent factor structure with the EFA, and the overall measurement model achieved a reasonable balance between model fit and model parsimony. The resulting Genomic SEM model contained 4 factors and 127 degrees of freedom with a CFI = 0.94 and an SRMR = 0.11. After observing the generally distinct signals exhibited by these 4 factors and the poor model fit from a common factor model, we refrained from fitting a hierarchical factor model to the data.
Genetic factors: multivariate genome wide association study
After defining the measurement model, we estimated SNP effects for the 4 genetic factors. This analysis was run in parallel for 954,086 SNPs that were common across the indicator GWASs and passed QC criteria. For each factor, we fit an independent pathways model for each SNP to test for heterogeneity of effect sizes among the indicator variables loading onto the same factor. The Genomic SEM QSNP methods included a fix_measurement parameter which was used to specify that the measurement model should be fixed across all SNPs, and we used the differences in the two models’ χ2 test statistics and degrees of freedom to identify SNPs with evidence for significant differences in model fit (QSNP p < 5 × 10−8)20. While these QSNPs are of interest because their indicator-specific effects might explain phenotypic divergence, for the purposes of constructing latent genetic factors that represent shared variance we removed these QSNPs along with nearby SNPs in LD. A European ancestry LD reference panel from the thousand genomes project (1KGP)82 consisting of 503 unrelated individuals and 13.6 million genetic variants was implemented with PLINK83,84 to identify and filter variants within 1 mega-base and LD r2 ≥ 0.2 with the QSNPs. F1, F2, F3, and F4 respectively had 23; 335; 1525; and 969 significant QSNPs, and after considering LD structure 79; 1284; 6909; and 4183 SNPs were removed. The allele frequencies and the standard errors of the effect estimates were used to estimate the effective sample size for each of the 4 latent factors via the method described in the supplement of Mallard et al.85. F1, F2, F3, and F4 had estimated effective sample sizes of 52,404; 176,820; 690,110; and 393,268 respectively.
We used DEPICT25 v1.194 (https://github.com/perslab/depict) to identify independent, associated genomic loci using default parameters of p < 5 × 10−8, LD pairwise r2 < 0.1, and physical distance <1 Mb (Supplementary Data 16, 21, 26, and 31). These significantly associated independent loci were used as input for the following analyses included in the DEPICT framework. First, we performed DEPICT SNP-to-gene mapping to identify likely causal genes based on the assumption that genes within an associated locus have functional similarity to genes from other associated loci. This consisted of a scoring step (to quantify the similarity of gene set membership of genes near associated loci), a bias adjustment step (to control for gene length and data structure), and an FDR estimation step. Significantly prioritized genes with FDR < 0.05 were retained as likely causal genes for our downstream analyses and are listed in Supplementary Data 1, 4, 7, and 10 for each factor GWAS. Next, DEPICT was used to identify functional or phenotypic gene sets that were enriched for genes within associated loci. This was performed using DEPICT’s 10,968 reconstituted gene sets with membership Z-scores representing the likelihood of membership of a gene in a gene set based on similarities (i.e., co-regulation) across gene expression data. These reconstituted gene sets were representative of a broad spectrum of biological annotations (Kyoto encyclopedia of genes and genomes [KEGG] pathways86, Gene Ontology [GO] terms87, Mammalian Phenotype [MP] ontology88, Reactome gene sets89, and protein-protein interaction [PPI] subnetworks90). DEPICT quantified enrichment (via the gene set scoring step, bias correction step, and FDR estimation step) was considered significant for gene sets with nominal p-values less than the Bonferroni-corrected significance threshold (p < 4.56×10-6) for each factor GWAS (Supplementary Data 18, 23, 28, and 33). Finally, DEPICT was implemented to test for enrichment (FDR < 0.05) of the associated loci across 210 annotations of relative gene expression in physiological systems, tissues, or cell types. Thus, the DEPICT gene set scoring step, bias correction step, and FDR estimation step were used to assess if genes in associated loci were highly expressed in certain tissues or cell types (Supplementary Data 19, 24, 29, and 34).
We tested for concordance of associated GWAS loci in an external sample to reduce the risk of false positives and increase the reliability of our results. In the absence of publicly available replication summary statistics for each trait included as an indicator variable, it was unfeasible to perform a complete replication analysis for each factor (i.e., construct a comparable Genomic SEM using independent data and perform a corresponding multivariate GWASs). In order to provide some confirmatory context, however, we evaluated the concordance of F2, F3, and F4 novel loci using primary indicator variables for each factor and the All of Us database23,24 as an independent dataset. We used the All by All (All-x-All) GWAS tables available through the Researcher Workbench. In this context, and throughout the manuscript, we defined novel loci as independent associated loci with no genomic positional overlap for the GWASs being compared. WHRadjBMI male and female indicators had strong loadings on F2 (0.90 and 0.71), and the All of Us database provided an independent dataset (European WHRadjBMI GWAS N = 102,746) to test for concordance of the lead SNPs of the 2 loci that were novel relative to the WHRadjBMI GWAS (combined males and females from the GIANT consortium)76. Similarly, we compared F3 to height, which had a strong loading of 0.88, to assess concordance in the All of Us European height GWAS (N = 111,755) for the lead SNPs of the 2 loci that were novel relative to the indicator height GWAS (combined males and females from the GIANT consortium)77,78. BMI male and female indicators both had strong loadings of 0.95 on F4, and we used BMI (All of Us European BMI GWAS N = 111,482) to evaluate concordance of the lead SNPs of the 28 loci that were novel relative to BMI (combined males and females from the GIANT consortium)79. All of Us did not have a large enough GWAS sample size for birth or infant anthropometrics to evaluate the concordance of F1 loci. To assess the concordance of statistical significance for each lead SNP we applied Bonferroni correction criteria (p < 0.025 for F2 and F3, and p < 1.79 × 10−3 for F4). We also evaluated concordance of effect direction after matching effect versus non-effect alleles for each lead SNP. Specifically, we used the pbinom function in R to test whether the observed concordance of effect directions was significantly greater than expected by chance under the null hypothesis of random effect direction (i.e., probability = 0.5).
Genetic factors: transcriptome wide association study
TWAS methods provided a natural extension of the multivariate GWASs to highlight genes with predicted expression that are putatively causal for the latent factors. We implemented TSEM91 and FOCUS26,92 softwares to perform transcription-level analyses of the previously discussed latent factor GWASs. Briefly, FOCUS uses a Bayesian framework to fine-map gene-trait TWAS associations by accounting for the induced correlation structure of predicted gene expression that is due to LD between SNPs and shared expression quantitative trait loci (eQTLs; i.e., pleiotropic effects). TWAS fine-mapping aims to prioritize genes with heritable variation in gene expression that causally impact the trait by assigning each gene a PIP. Within a region, genes are rank ordered by their PIPs to compute minimal 90%-credible gene sets that contain the causal gene with 90% probability; concentrating on the 90% CSs that do not contain the null model enables the identification of regions with stronger evidence for gene expression driving trait variation (as opposed to regions where the association between expression and trait variation is due to chance). Ultimately, we prioritized the FOCUS results over the TSEM results in our TWAS analysis because the software’s fine-mapping approach handled the underlying correlation structure for predicted gene expression and provided credible sets of putatively causal genes with PIPs. Although we do not discuss the TSEM associations here, they are included in Supplementary Data 44–55.
We ran FOCUS v0.9 (https://github.com/mancusolab/ma-focus/) with data for SNP LD structure, prediction eQTL weights, and the factor GWAS summary statistics, and FOCUS provided 90%-credible gene sets that excluded the null model (Supplementary Data 20, 25, 30, 35). We used the FOCUS repository’s recommended European ancestry reference LD plink-formatted files from LDSC and the FOCUS repository’s multiple tissue, multiple eQTL reference panel weight database. First, the FOCUS munge functionality was used to format the factors’ GWAS summary statistics, and then each chromosome was run in parallel using independent genomic regions across European ancestry identified by LDetect93 and the prior probability for a gene to be causal as 0.001. The tissue-enrichment results from our prior DEPICT analysis revealed that the 4th factor was the only factor with enrichment in a singular physiological system (enriched only for nervous tissues and cell types); thus, FOCUS was run tissue-agnostic for the first 3 factors (F1, F2, and F3) and was run tissue-prioritized for the ‘brain’ for F4. F1, F2, F3, and F4 respectively had 86, 290, 737, and 562 LD blocks with identified 90%-credible gene sets, and 69, 243, 690, and 335 of those did not contain the null model. Among those gene sets that did not contain the null model, we retained genes with PIP > 0.1 to filter out low probability genes from our downstream analyses. Given the polygenic architecture of these latent traits (many genes with small effect sizes at the level of transcription) and that our aims were largely exploratory, a PIP threshold of 0.1 allowed genes with moderate statistical support to be considered. This thresholding step resulted in 158; 676; 2266; and 850 respective genes with putatively causal predicted gene expression effects for each of the 4 factors (Supplementary Data 2, 5, 8, and 11).
Genetic factors: genetic correlations
We evaluated the genetic correlations of the four factors with a broad range of obesity-related traits, using multivariate LDSC. Given the far-reaching spectrum of obesity-related health outcomes, we compiled a list of traits relating to psychopathology, risky behavior, neuroticism, diet, sleep, exercise, substance use, pain, frailty, dementia, inflammatory disease, autoimmune disease, cardiovascular disease, and metabolism. The full set of considered traits is described in Supplementary Table 3 and Supplementary Data 65 along with LDSC parameters for sample sizes, population prevalence, and heritability Z-statistics. The multivariate LDSC function in the Genomic SEM R package (https://github.com/GenomicSEM/GenomicSEM) was used to estimate genetic covariances and correlations. Prior to visualizing the prominent correlations (Fig. 6) we filtered out 10 traits with heritability Z-statistics <4 to avoid misinterpretation due to small sample size or minimal genetic effects.
Genetic factors: phenome wide association study of polygenic scores
PGSs estimate an individual’s genetic predisposition to a trait, based on the weighted sum of genetic variant effects across the genome. We derived PGS SNP weights for the 4 factor GWASs using LDpred2 (https://privefl.github.io/bigsnpr/articles/LDpred2.html)27. Briefly, LDpred2 estimates model hyperparameters (SNP-based heritability and the fraction of causal variants) from GWAS data and uses an iterative Bayesian Gibbs sampler to adjust for LD between SNPs and update effect size estimates. We used a random subset of 5000 unrelated individuals of European ancestry from the UK Biobank for the LD reference panel. This LD reference panel was sufficiently large (>1000 individuals per the LDpred2 guidelines), and we defined unrelated individuals using gcta64 --grm-singleton 0.0594. Standard QC processes95 involved filtering SNPs based on Hardy-Weinberg equilibrium p > 1 × 10−6, genotyping rate >0.99, non-ambiguous alleles, minor allele frequency (MAF) > 1%, and filtering individuals based on heterozygosity within 3 standard deviations of the mean and sample missingness <0.02. The ancestry matched remarkably well between the LD panel and the multivariate GWAS summary statistics, and nearly all SNPs were retained when applying the LDpred2 standard deviation filter (Supplementary Fig. 36). The snp_ldpred2_auto function in the bigsnpr package (v.1.12.15) was used to generate LD-adjusted PGS weights for a sequence of causal variant thresholds (30 evenly spaced values on a logarithmic scale ranging from 1×10−4 to 0.9). The average of the betas for the models that converged were used for the PGSs resulting in 710,801; 710,489; 709,195; and 709,830 SNP weights for F1, F2, F3, and F4 respectively. Visualization of the raw GWAS effect sizes compared to the attenuated LDpred2-adjusted PGS weights is shown in Supplementary Fig. 37.
These PGS weights were applied in an external dataset with no sample overlap with the included GWASs. We conducted 4 phenome-wide association studies (pheWASs) to investigate the associations between each of the 4 PGSs and all 1514 phecode-based phenotypes in a cohort of unrelated Europeans from the Colorado Center for Personalized Medicine (CCPM) Biobank freeze2 (N = 25,240). Ancestry information was inferred based on the grouping of individuals’ genetic proximity to reference populations via PCA-UMAP (Principal Component Analysis-Uniform Manifold Approximation and Projection) projection as input for k-nearest neighbors clustering (using the UMAP coordinates of reference panel individuals to train the clusters). We excluded individuals with second-degree or closer relatedness identified through KING-robust kinship estimates greater than 2 × 10−3.5, using the bigsnpr package in R96. Details regarding the recruitment of CCPM Biobank participants, data processing, and the inference of population structures are described in Wiley et al.97.
Our pheWAS association model corrected for age, sex, batch, and the first 10 genetic principal components. Participant age, sex, and batch were standard covariates delivered in freeze2 from the CCPM Biobank institutional data warehouse which harmonized health information from the Epic-based electronic health record (EHR)97. To achieve unbiased estimates in the presence of case-control imbalance, we utilized the Saddlepoint approximation method from the SPAtest package in R98. Due to the highly correlated structure of phecodes in the CCPM Biobank EHR97, we considered associations with p-values below the FDR < 0.10 Bonferroni-corrected significance threshold (6.61 × 10−5) significant when characterizing the predictive signal of the factor PGSs. To evaluate the predictive utility of the F4 PGS conditioned on BMI, we ran an auxiliary pheWAS with BMI included as an additional covariate. To estimate BMI for each participant, we used the median of BMI measurements across the EHR. The median BMI was used over the most recent or mean BMI, as it provided greater robustness to outlier events such as pregnancy or bariatric surgery. However, while the median mitigates the influence of extreme values, it may not fully eliminate the impact of outlier events. For each encounter with documented height (measured in inches) and weight (measured in ounces) we performed unit conversions and calculated the BMI as height/weight2. BMI values less than 13 kg/m2 or greater than 60 kg/m2 were removed before finding the median. These outlier thresholds were based on the empirical BMI distribution for the CCPM Biobank and were similar to outlier thresholds applied to other large-scale biobanks (15 kg/m2 and 60 kg/m2)99.
Genetic factors: drug-gene network
We queried two large drug repurposing databases (Drug Repurposing Hub [DRH; 3/24/2020 version]35 and the Drug-Gene Interaction Database [DGIdb; 12/2023 version])36 for the genes that were either significantly prioritized by DEPICT (independent GWAS loci; FDR < 0.05) or FOCUS (fine-mapped TWAS 90% credible sets without the null model; PIP > 0.1). There were 24; 319; 1864; and 437 significantly prioritized DEPICT genes and 215; 862; 2944; and 864 FOCUS fine-mapped genes for F1, F2, F3, and F4, respectively. The DGIdb contained drug-gene interaction scores reflecting strength of supporting publications and the relative drug-gene specificity. We filtered out drug-gene pairs with low interaction scores (<0.50) based on the QC procedures described in similar studies100,101. To map gene identifiers between datasets, we used the custom download feature from https://www.genenames.org/ to map the official gene symbol approved by the HGNC to the Ensembl Gene IDs. There were 14,472 drug-gene pairs for 6798 drugs in the DRH and 19,819 drug-gene pairs for 8037 drugs in the DGIdb. For visualization102 of the drug-gene network for F4 we removed drugs that did not have ‘launched’ clinical phase in the DRH or ‘approved’ status in the DGIdb. Drug indications were extracted from the ensemble MEDication Indication resource (MEDI-C)103 containing 38,378 high precision drug-indication pairs. The PheWAS R package104 was used to map the indication ICD10CM codes to phecodes and their corresponding phenotype domains. The ON-label SIDE effectS resource (OnSIDES, v2.0.0_20231113)37 was used to identify wADEs for the drugs in the network. This database contained 2020 ingredients and 4302 unique adverse reactions that were assigned using natural language processing models of drug labels. We considered drug-ADE pairs for which the adverse reaction was extracted from at least 75% of labels, and defined wADEs based on the following list of drug events: ‘Obesity’, ‘Central obesity’, ‘Weight increased’, ‘Weight decreased’, ‘Weight fluctuation’, ‘Abnormal loss of weight’, ‘Abnormal weight gain’, ‘Weight loss poor’, ‘Decreased appetite’, ‘Increased appetite’, ‘Appetite disorder’, ‘Hunger’, ‘Early satiety’, ‘Binge eating’, ‘Sleep-related eating disorder’, ‘Eating disorder’. Of these 16 terms, ‘Decreased appetite’, ‘Weight increased’, ‘Weight decreased’, and ‘Increased appetite’ were the most prominent and frequently observed (comprising 98% of wADE instances). The annotations provided by MEDI-C and OnSIDES aided the interpretation of the bipartite drug-gene networks through providing context regarding the drugs’ indicated medical domains and wADEs.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The multivariate GWAS summary statistics generated in this study have been deposited in the GWAS Catalog database under accession codes GCST90624101 (F1), GCST90624103 (F2), GCST90624105 (F3), and GCST90624107 (F4). The LDpred2-derived PGS weights generated in this study have been deposited in the PGS Catalog database under under publication ID PGP000739 and score IDs PGS005232 (F1), PGS005233 (F2), PGS005234 (F3), and PGS005235 (F4). The CCPM genetic and EHR datasets are available under restricted access due to the sensitive nature of these datasets and HIPAA compliance, and access can be obtained through an Access to Biobank Committee (ABC) study proposal request (https://medschool.cuanschutz.edu/cobiobank/contact); consult with Health Data Compass and the CCPM biobank team to understand team regarding logistical requirements and the timeframe for data access after an initial request. All GWAS summary statistics included in this study were publicly available and citations linking downloads for the GWAS summary statistic files are included in the Supplementary Information. Data on birth weight traits were contributed by the EGG Consortium and were downloaded from www.egg-consortium.org. Data on anthropometric traits were contributed by the GIANT Consortium and were downloaded from https://portals.broadinstitute.org/collaboration/giant/index.php/GIANT_consortium_data_files. Data on BMI trajectory were downloaded from https://ucla.app.box.com/v/trajgwassummary. Data on bio-electrical impedance were downloaded from https://myfiles.uu.se/ssf/s/readFile/share/3993/1270878243748486898/publicLink/GWAS_summary_stats_ratios.zip. The DGIdb and DRH datasets were publicly available and were downloaded from https://www.dgidb.org/downloads and https://repo-hub.broadinstitute.org/repurposing#download-data. The MEDI-C and OnSIDES datasets were publicly available and were downloaded from https://www.vumc.org/wei-lab/medi and https://github.com/tatonetti-lab/onsides/releases. Source Data are provided as a Source Data file. Source data are provided with this paper.
Code availability
The code used to perform the analyses in this study is available at https://github.com/char4816/AdiposityGSEM and at Zenodo [https://doi.org/10.5281/zenodo.15733864]105.
Change history
12 September 2025
In this article the funding from the University of Colorado Boulder Libraries Open Access Fund was omitted. The original article has been corrected.
References
Bryan, S. et al. NHSR. National Health and Nutrition Examination Survey 2017–March 2020 Pre-Pandemic Data Files. https://stacks.cdc.gov/view/cdc/106273 (2021).
Loos, R. J. F. & Yeo, G. S. H. The genetics of obesity: from discovery to biology. Nat. Rev. Genet. 23, 120–133 (2022).
CDC. Obesity is a common, serious, and costly disease. Cent. Dis. Control Prev. https://www.cdc.gov/obesity/data/adult.html (2022).
Khera, A. V. et al. Polygenic prediction of weight and obesity trajectories from birth to adulthood. Cell 177, 587–596.e9 (2019).
Couto Alves, A. et al. GWAS on longitudinal growth traits reveals different genetic factors influencing infant, child, and adult BMI. Sci. Adv. 5, eaaw3095 (2019).
Hansen, G. T. et al. Genetics of sexually dimorphic adipose distribution in humans. Nat. Genet. 55, 461–470 (2023).
Ibrahim, M. M. Subcutaneous and visceral adipose tissue: structural and functional differences. Obes. Rev. 11, 11–18 (2010).
Aronica, L., Rigdon, J., Offringa, L. C., Stefanick, M. L. & Gardner, C. D. Examining differences between overweight women and men in 12-month weight loss study comparing healthy low-carbohydrate vs. low-fat diets. Int. J. Obes. 45, 225–234 (2021).
Doucet, E. et al. Reduction of visceral adipose tissue during weight loss. Eur. J. Clin. Nutr. 56, 297–304 (2002).
Okorodudu, D. O. et al. Diagnostic performance of body mass index to identify obesity as defined by body adiposity: a systematic review and meta-analysis. Int. J. Obes. 34, 791–799 (2010).
Tomiyama, A. J., Hunger, J. M., Nguyen-Cuu, J. & Wells, C. Misclassification of cardiometabolic health when using body mass index categories in NHANES 2005–2012. Int. J. Obes. 40, 883–886 (2016).
De Lorenzo, A. et al. Normal-weight obese syndrome: early inflammation?. Am. J. Clin. Nutr. 85, 40–45 (2007).
Abdellaoui, A. & Verweij, K. J. H. Dissecting polygenic signals from genome-wide association studies on human behaviour. Nat. Hum. Behav. 5, 686–694 (2021).
Morys, F. et al. Neuroanatomical correlates of genetic risk for obesity in children. Transl. Psychiatry 13, 1 (2023).
Shungin, D. et al. New genetic loci link adipose and insulin biology to body fat distribution. Nature 518, 187–196 (2015).
Van Walree, E. S. et al. Disentangling genetic risks for metabolic syndrome. Diabetes 71, 2447–2457 (2022).
Virtue, S. & Vidal-Puig, A. Adipose tissue expandability, lipotoxicity and the Metabolic Syndrome-an allostatic perspective. Biochim. Biophys. Acta 1801, 338–349 (2010).
Yaghootkar, H. et al. Genetic evidence for a link between favorable adiposity and lower risk of type 2 diabetes, hypertension, and heart disease. Diabetes 65, 2448–2460 (2016).
Yaghootkar, H. et al. Genetic evidence for a normal-weight ‘metabolically obese’ phenotype linking insulin resistance, hypertension, coronary artery disease, and type 2 diabetes. Diabetes 63, 4369–4377 (2014).
Grotzinger, A. D. et al. Genomic structural equation modelling provides insights into the multivariate genetic architecture of complex traits. Nat. Hum. Behav. 3, 513–525 (2019).
Grotzinger, A. D. et al. Multivariate genomic architecture of cortical thickness and surface area at multiple levels of analysis. Nat. Commun. 14, 946 (2023).
Kaplan, D. Structural Equation Modeling: Foundations and Extensions (2nd edn) (SAGE Publications, Inc., 2009). https://doi.org/10.4135/9781452226576.
The All of Us Research Program Genomics Investigators et al. Genomic data in the All of Us Research Program. Nature 627, 340–346 (2024).
The All of Us Research Program Investigators. The “All of Us” Research Program. N. Engl. J. Med. 381, 668–676 (2019).
Pers, T. H. et al. Biological interpretation of genome-wide association studies using predicted gene functions. Nat. Commun. 6, 5890 (2015).
Mancuso, N. et al. Probabilistic fine-mapping of transcriptome-wide association studies. Nat. Genet. 51, 675–682 (2019).
LDpred2: better, faster, stronger | Bioinformatics | Oxford Academic. https://academic.oup.com/bioinformatics/article/36/22-23/5424/6039173
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Cherny, S. S., Livshits, G. & Williams, F. M. K. A genetic and environmental analysis of inflammatory factors in chronic widespread pain using the TwinsUK Cohort. Biomolecules 15, 155 (2025).
Speed, M. S., Jefsen, O. H., Børglum, A. D., Speed, D. & Østergaard, S. D. Investigating the association between body fat and depression via Mendelian randomization. Transl. Psychiatry 9, 184 (2019).
Zhou, J. et al. Causal relationship between cheese intake and risk of gastroesophageal reflux disease and Barrett’s esophagus: findings from multivariable Mendelian randomization and mediation analysis. Eur. J. Nutr. 64, 49 (2025).
Levin, M. G. et al. Genetics of height and risk of atrial fibrillation: A Mendelian randomization study. PLOS Med 17, e1003288 (2020).
Wajchenberg, B. L. Subcutaneous and visceral adipose tissue: their relation to the metabolic syndrome. Endocr. Rev. 21, 697–738 (2000).
Corsello, S. M. et al. The drug repurposing hub: a next-generation drug library and information resource. Nat. Med. 23, 405–408 (2017).
Cannon, M. et al. DGIdb 5.0: rebuilding the drug–gene interaction database for precision medicine and drug discovery platforms. Nucleic Acids Res. 52, D1227–D1235 (2024).
Tanaka, Y. et al. OnSIDES database: Extracting adverse drug events from drug labels using natural language processing models. Med 6, 100642 (2025).
Komossa, K. et al. Olanzapine versus other atypical antipsychotics for schizophrenia. Cochrane Database Syst. Rev. https://doi.org/10.1002/14651858.CD006654.pub2 (2010).
Praharaj, S. K., Jana, A. K., Goyal, N. & Sinha, V. K. Metformin for olanzapine-induced weight gain: a systematic review and meta-analysis. Br. J. Clin. Pharmacol. 71, 377–382 (2011).
Ono, S. et al. Association between the GIPR gene and the insulin level after glucose loading in schizophrenia patients treated with olanzapine. Pharmacogenom. J. 12, 507–512 (2012).
Ono, S. et al. GIPR gene polymorphism and weight gain in patients with schizophrenia treated with olanzapine. J. Neuropsychiatry Clin. Neurosci. 27, 162–164 (2015).
Jeon, W. J., Dean, B., Scarr, E. & Gibbons, A. The role of muscarinic receptors in the pathophysiology of mood disorders: a potential novel treatment?. Curr. Neuropharmacol. 13, 739–749 (2015).
Simons, F. E. R. & Simons, K. J. Histamine and H1-antihistamines: celebrating a century of progress. J. Allergy Clin. Immunol. 128, 1139–1150.e4 (2011).
Zhao, M., Jung, Y., Jiang, Z. & Svensson, K. J. Regulation of energy metabolism by receptor tyrosine kinase ligands. Front. Physiol. 11, 354 (2020).
Niemann, B. et al. Apoptotic brown adipocytes enhance energy expenditure via extracellular inosine. Nature 609, 361–368 (2022).
Klausen, M. K., Thomsen, M., Wortwein, G. & Fink-Jensen, A. The role of glucagon-like peptide 1 (GLP-1) in addictive disorders. Br. J. Pharmacol. 179, 625–641 (2022).
Tsermpini, E. E., Goričar, K., Kores Plesničar, B., Plemenitaš Ilješ, A. & Dolžan, V. Genetic variability of incretin receptors and alcohol dependence: a pilot study. Front. Mol. Neurosci. 15, 908948 (2022).
Willemsen, N., Kotschi, S. & Bartelt, A. Fire up the pyre: inosine thermogenic signaling for obesity therapy. Signal Transduct. Target. Ther. 7, 1–2 (2022).
Huang, J. et al. Genomics and phenomics of body mass index reveals a complex disease network. Nat. Commun. 13, 7973 (2022).
Emdin, C. A. et al. Genetic association of waist-to-hip ratio with cardiometabolic traits, type 2 diabetes, and coronary heart disease. JAMA 317, 626–634 (2017).
Nakao, K. Adiposcience and adipotoxicity. Nat. Clin. Pract. Endocrinol. Metab. 5, 63–63 (2009).
Whitehead, J. P., Richards, A. A., Hickman, I. J., Macdonald, G. A. & Prins, J. B. Adiponectin-a key adipokine in the metabolic syndrome. Diabetes Obes. Metab. 8, 264–280 (2006).
Röszer, T. Adipose tissue immunometabolism and apoptotic cell clearance. Cells 10, 2288 (2021).
Fuchsberger, C. et al. The genetic architecture of type 2 diabetes. Nature 536, 41–47 (2016).
Kim, H. et al. High-throughput genetic clustering of type 2 diabetes loci reveals heterogeneous mechanistic pathways of metabolic disease. Diabetologia 66, 495–507 (2023).
Qasim, A. et al. On the origin of obesity: identifying the biological, environmental and cultural drivers of genetic risk among human populations. Obes. Rev. 19, 121–149 (2018).
Alderete, T. L. et al. Ambient and traffic-related air pollution exposures as novel risk factors for metabolic dysfunction and type 2 diabetes. Curr. Epidemiol. Rep. 5, 79–91 (2018).
Alderete, T. L. et al. Prenatal traffic-related air pollution exposures, cord blood adipokines and infant weight. Pediatr. Obes. 13, 348–356 (2018).
Aschard, H., Vilhjálmsson, B. J., Joshi, A. D., Price, A. L. & Kraft, P. Adjusting for heritable covariates can bias effect estimates in genome-wide association studies. Am. J. Hum. Genet. 96, 329–339 (2015).
Watanabe, K. et al. Multiomic signatures of body mass index identify heterogeneous health phenotypes and responses to a lifestyle intervention. Nat. Med. https://doi.org/10.1038/s41591-023-02248-0 (2023).
Walter, E. C., Ehlenbach, W. J., Hotchkin, D. L., Chien, J. W. & Koepsell, T. D. Low birth weight and respiratory disease in adulthood. Am. J. Respir. Crit. Care Med. 180, 176–180 (2009).
Lillås, B. S., Qvale, T. H., Richter, B. K. & Vikse, B. E. Birth weight is associated with kidney size in middle-aged women. Kidney Int. Rep. 6, 2794–2802 (2021).
Eriksson, J., Forsén, T., Tuomilehto, J., Osmond, C. & Barker, D. Fetal and childhood growth and hypertension in adult life. Hypertens. Dallas Tex. 1979 36, 790–794 (2000).
Abufaraj, M. et al. Association between body fat mass and kidney stones in US adults: analysis of the national health and nutrition examination survey 2011-2018. Eur. Urol. Focus 8, 580–587 (2022).
Després, J. P. The insulin resistance-dyslipidemic syndrome of visceral obesity: effect on patients’ risk. Obes. Res. 6, 8S-17S (1998).
Sironi, A. M. et al. Visceral fat in hypertension: influence on insulin resistance and beta-cell function. Hypertens. Dallas Tex. 1979 44, 127–133 (2004).
Goswami, B., Reang, T., Sarkar, S., Sengupta, S. & Bhattacharjee, B. Role of body visceral fat in hypertension and dyslipidemia among the diabetic and nondiabetic ethnic population of Tripura—A comparative study. J. Fam. Med. Prim. Care 9, 2885–2890 (2020).
Agrawal, S. et al. Inherited basis of visceral, abdominal subcutaneous and gluteofemoral fat depots. Nat. Commun. 13, 3771 (2022).
Timshel, P. N., Thompson, J. J. & Pers, T. H. Genetic mapping of etiologic brain cell types for obesity. eLife 9, e55851 (2020).
Kline, R. B. Principles and Practice of Structural Equation Modeling (The Guilford Press, 2023).
Rask-Andersen, M., Karlsson, T., Ek, W. E. & Johansson, Å Genome-wide association study of body fat distribution identifies adiposity loci and sex-specific genetic effects. Nat. Commun. 10, 339 (2019).
Bradfield, J. P. et al. A trans-ancestral meta-analysis of genome-wide association studies reveals loci associated with childhood obesity. Hum. Mol. Genet. 28, 3327–3338 (2019).
Vogelezang, S. et al. Genetics of early-life head circumference and genetic correlations with neurological, psychiatric and cognitive outcomes. BMC Med. Genom. 15, 124 (2022).
van der Valk, R. J. P. et al. A novel common variant in DCST2 is associated with length in early life and height in adulthood. Hum. Mol. Genet. 24, 1155–1168 (2015).
EGG Consortium et al. Maternal and fetal genetic effects on birth weight and their relevance to cardio-metabolic risk factors. Nat. Genet. 51, 804–814 (2019).
Pulit, S. L. et al. Meta-analysis of genome-wide association studies for body fat distribution in 694,649 individuals of European ancestry. Hum. Mol. Genet. 28, 166–174 (2019).
Yengo, L. et al. A saturated map of common genetic variants associated with human height. Nature 610, 704–712 (2022).
The Electronic Medical Records and Genomics (eMERGE) Consortium et al. Defining the role of common variation in the genomic and biological architecture of adult human height. Nat. Genet 46, 1173–1186 (2014).
Ko, S. et al. GWAS of longitudinal trajectories at biobank scale. Am. J. Hum. Genet. 109, 433–445 (2022).
Kaiser, H. F. The application of electronic computers to factor analysis. Educ. Psychol. Meas. 20, 141–151 (1960).
Ruscio, J. & Roche, B. Determining the number of factors to retain in an exploratory factor analysis using comparison data of known factorial structure. Psychol. Assess. 24, 282–292 (2012).
The 1000 Genomes Project Consortium et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience 4, 7 (2015).
Mallard, T. T. et al. Multivariate GWAS of psychiatric disorders and their cardinal symptoms reveal two dimensions of cross-cutting genetic liabilities. Cell Genom.2, 100140 (2022).
Kanehisa, M., Goto, S., Sato, Y., Furumichi, M. & Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 40, D109–D114 (2012).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Blake, J. A. et al. The Mouse Genome Database: integration of and access to knowledge about the laboratory mouse. Nucleic Acids Res. 42, D810–D817 (2014).
Croft, D. et al. Reactome: a database of reactions, pathways and biological processes. Nucleic Acids Res. 39, D691–D697 (2011).
Lage, K. et al. A human phenome-interactome network of protein complexes implicated in genetic disorders. Nat. Biotechnol. 25, 309–316 (2007).
Grotzinger, A. D., de la Fuente, J., Davies, G., Nivard, M. G. & Tucker-Drob, E. M. Transcriptome-wide and stratified genomic structural equation modeling identify neurobiological pathways shared across diverse cognitive traits. Nat. Commun. 13, 6280 (2022).
Lu, Z. et al. Multi-ancestry fine-mapping improves precision to identify causal genes in transcriptome-wide association studies. Am. J. Hum. Genet. 109, 1388–1404 (2022).
Berisa, T. & Pickrell, J. K. Approximately independent linkage disequilibrium blocks in human populations. Bioinformatics 32, 283–285 (2016).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Choi, S. W., Mak, T. S.-H. & O’Reilly, P. F. Tutorial: a guide to performing polygenic risk score analyses. Nat. Protoc. 15, 2759–2772 (2020).
Privé, F., Aschard, H., Ziyatdinov, A. & Blum, M. G. B. Efficient analysis of large-scale genome-wide data with two R packages: bigstatsr and bigsnpr. Bioinformatics 34, 2781–2787 (2018).
Wiley, L. K. et al. Building a vertically integrated genomic learning health system: the biobank at the Colorado Center for Personalized Medicine. Am. J. Hum. Genet. 111, 11–23 (2024).
Dey, R., Schmidt, E. M., Abecasis, G. R. & Lee, S. A fast and accurate algorithm to test for binary phenotypes and its application to PheWAS. Am. J. Hum. Genet. 101, 37–49 (2017).
Di Angelantonio, E. et al. Body-mass index and all-cause mortality: individual-participant-data meta-analysis of 239 prospective studies in four continents. Lancet 388, 776–786 (2016).
Olla, S. et al. Combining human genetics of multiple sclerosis with oxidative stress phenotype for drug repositioning. Pharmaceutics 13, 2064 (2021).
Grotzinger, A. D. et al. Transcriptome-wide structural equation modeling of 13 major psychiatric disorders for cross-disorder risk and drug repurposing. JAMA Psychiatry 80, 811–821 (2023).
Csárdi, G. et al. igraph for R: R interface of the igraph library for graph theory and network analysis. Zenodo https://doi.org/10.5281/ZENODO.7682609 (2024).
Zheng, N. S. et al. An updated, computable MEDication-Indication resource for biomedical research. Sci. Rep. 11, 18953 (2021).
Carroll, R. J., Bastarache, L. & Denny, J. C. R. PheWAS: data analysis and plotting tools for phenome-wide association studies in the R environment. Bioinformatics 30, 2375–2376 (2014).
Arehart, C. H. et al. Modeling the genomic architecture of adiposity and anthropometrics across the lifespan. AdiposityGSEM https://doi.org/10.5281/zenodo.15733864 (2024).
Acknowledgements
We thank the generous feedback from members of the Institute for Behavioral Genetics (IBG) statistical genetics group. Data storage for this project was supported by the PetaLibrary and computational analysis was supported by the Blanca and Alpine high performance computing resources at the University of Colorado Boulder (funded by the University of Colorado Boulder, the University of Colorado Anschutz, and Colorado State University). CHA was supported by the IBG NIMH T32MH016880 training grant and by the Interdisciplinary Quantitative Biology program. RAG was supported by the University of Colorado Boulder’s Summer Multicultural Access to Research Training program, part of the Colorado Diversity Initiative, which is funded internally by the University of Colorado Boulder Graduate School. MAS was supported by grant K01 HL157658. ADG was supported by NIH Grants R01MH120219 and RF1AG073593. LME was supported by R01AG046938. This research has been conducted using the UK Biobank Resource under Application Number 1665. We thank the participants of the Colorado Center for Personalized Medicine Biobank. In addition, we gratefully acknowledge All of Us participants for their contributions and thank the National Institutes of Health’s All of Us Research Program for making the All by All (All-x-All) GWAS tables available through the Researcher Workbench. Publication of this article was funded by the University of Colorado Boulder Libraries Open Access Fund.
Author information
Authors and Affiliations
Consortia
Contributions
C.H.A. and L.M.E. conceived and designed the study. C.H.A., A.D.G., and L.M.E. developed the 4-factor Genomic SEM. C.H.A. collected and curated the data and performed the formal analyses. M.L. performed the pheWAS in the CCPM Biobank and R.A.G. performed the supporting TSEM analysis. C.H.A. generated all figures and data visualizations. C.H.A., M.L., S.R., C.R.G., M.A.S., A.D.G., and L.M.E. interpreted the results. C.H.A. wrote the original draft, and M.L., S.R., C.R.G., M.A.S., A.D.G., and L.M.E. reviewed and edited the manuscript. L.M.E. supervised the project and provided overall direction. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Arehart, C.H., Lin, M., Gibson, R.A. et al. Modeling the genomic architecture of adiposity and anthropometrics across the lifespan. Nat Commun 16, 7494 (2025). https://doi.org/10.1038/s41467-025-62730-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-62730-w