Abstract
As central cells involved in osteoimmunology, bone niche macrophages possess diverse functions, and their differentiation fate regulates bone homeostasis. Elucidation of the underlying mechanism involved in macrophage differentiation is important for developing new therapeutic targets for osteoporosis. Here, we show that knocking out Raf kinase inhibitor protein (RKIP), either globally or in macrophages, results in dramatically increased bone mass in mice due to synergistic inhibition of bone resorption and promotion of bone formation. Mechanistically, RKIP knockout inhibits differentiation of macrophages into osteoclasts and promotes their differentiation towards pro-angiogenic subclusters, which enhances formation of H-type vessels. RKIP enhances osteoclastogenesis by interacting with ARHGAP to suppress CDC42 inactivation. Intranuclear RKIP suppresses angiogenic genes expression by bridging the association between HIF-1α and VHL to reduce the protein stability of HIF-1α in macrophages. Furthermore, RKIP deletion or inhibitor rescues ovariectomy (OVX)-induced bone loss in vivo. Collectively, this study provides insights into the different roles of extranuclear or intranuclear RKIP in regulating differentiation of bone niche macrophages and could inform potential therapies for bone homeostasis-related diseases.
Similar content being viewed by others
Introduction
Bone is a dynamic tissue which provides structural support for the body, protects vital organs, houses the hematopoietic system, and maintains the homeostasis of calcium and phosphate ions1,2. Bone homeostasis refers to the balance between bone formation and resorption, which is crucial for the normal morphology and function of the skeletal system3. The composition of the bone niche, including osteoblasts, osteoclasts, osteocytes, endothelial cells, mesenchymal stem cells, and stromal cells, collectively influences bone homeostasis. Therefore, gaining a comprehensive understanding of the regulatory mechanisms within the bone niche is of significant importance4,5,6.
In the year 2000, Arron et al. first introduced the concept of osteoimmunology, highlighting the reciprocal regulation between immune cells and bone-related cells, and the potential disruption of bone homeostasis due to immune dysregulation7. Immune cells mainly consist of lymphocytes (T cells and B cells) and granulocytes (monocytes/macrophages, neutrophils), among which macrophages play a central role as essential innate immune cells within the bone-immune regulatory network. Macrophages exhibit remarkable plasticity, and their differentiation fate is regulated by the niche of the tissue. Tissue niche provides physical support and controls the tissue-specific characteristics and proportion of macrophages by generating niche signals8. Various cellular factors in the bone niche, such as M-CSF9, RANKL10, TGF-β11 and BMP12, regulate the maintenance and functional specialization of macrophages. Similarly, the differentiation fate of macrophages within the bone niche is crucial for the regulation of bone homeostasis. Under the treatment of M-CSF, macrophages can differentiate into osteoclast precursors, which promote bone formation by inducing H-type vessel formation through the secretion of PDGF-BB13. Macrophages stimulated by M-CSF and RANKL can differentiate into mature osteoclasts and directly participate in bone resorption14. Bone-resident macrophages can significantly enhance osteoblast differentiation and bone formation by secreting OSM15,16. However, the key molecules that regulate the differentiation fate of bone niche macrophages and their roles in maintaining bone homeostasis are still not fully understood.
Raf kinase inhibitory protein (RKIP) is a member of the phosphatidylethanolamine-binding protein family (PEBP) family, which is widely expressed in prokaryotes and eukaryotes. RKIP regulates multiple intracellular signaling cascades including Raf/MEK/ERK, NF-κB, glycogen synthase kinase-3β (GSK3β), and G-protein-coupled receptors17,18,19,20. Therefore, RKIP is involved in the regulation of various physiological processes, including cell differentiation, cell cycle and apoptosis21. Additionally, RKIP serves as a critical molecule in immune regulation. We have demonstrated that RKIP plays a regulatory role in inflammasome activation22, innate immune responses23,24, inflammatory bowel disease25, allergic asthma26 and autoimmune diseases27. However, whether RKIP contributes to the regulation of macrophage differentiation remains unclear. Furthermore, current research on RKIP mainly focuses on its cytoplasmic functions, whether it can enter the nucleus and its nuclear functions require further clarification.
In this study, we discover that RKIP regulates the differentiation fate of bone niche macrophages. Knocking out RKIP promotes the differentiation of macrophages towards the pro-angiogenic subclusters and inhibits their differentiation into osteoclasts. Mechanistically, RKIP regulates osteoclast differentiation through the RKIP/ARHGAP/CDC42 axis and modulates the differentiation of pro-angiogenic macrophages via the intranuclear RKIP/VHL/HIF-1α axis, thereby influencing H-type vessel formation. This study provides a comprehensive understanding of the nuclear entry mechanism and extranuclear or intranuclear functions of RKIP in differentiation fate regulation of bone niche macrophages.
Result
Global knockout of RKIP suppresses bone resorption and enhances bone formation
To investigate the overall impact of RKIP on bone homeostasis, we initially generated RKIP global knockout (gKO) mice and assessed their bone mass in comparison to littermate controls. Micro-CT revealed that RKIP knockout led to an increase in trabecular bone mass and cortical bone thickness in male and female mice (Fig. 1A, B and Fig. S1A, B). Due to the observation that both male and female mice exhibited a similar trend of increased bone mass, we proceeded with subsequent experiments using male mice. H&E staining showed a significant increase in skeletal mass and narrowing of the bone marrow cavity in the gKO mice (Fig. 1C). Tartrate-resistant acid phosphatase (TRAP) staining demonstrated a decrease in the osteoclast surface per bone surface (Oc.S/BS) and osteoclast number per bone surface (Oc.N/BS) in the gKO mice, which indicating RKIP knockout may inhibit osteoclast differentiation (Fig. 1C, D). To further investigate the bone formation ability, we performed calcein double staining and Von Kossa staining. The gKO mice showed a significant increase in the mineral apposition rate (MAR), bone formation rate (BFR/BS) (Fig. 1E, F) and the number of osteocalcin (OCN) positive cells compared to the control (RKIP+/+) group (Fig. 1G, H). Furthermore, the serum bone resorption marker C-terminal telopeptide of type I collagen (CTX-1) was decreased and the bone formation marker procollagen I N-terminal propeptide (PINP) was increased in the gKO mice (Fig. 1I). In summary, gKO of RKIP significantly inhibits bone resorption and enhances bone formation, leading to osteosclerosis in vivo.
A Micro-CT and three-dimensional (3D) reconstruction images of the distal femur of 3-month-old male RKIP+/+ or RKIP-/- mice are shown. B Quantification of bone volume/tissue volume (BV/TV), trabecular number (Tb.N), trabecular separation (Tb.Sp), trabecular thickness (Tb.Th), and cortical thickness (Ct.Th) (n = 5). C H&E staining and TRAP staining of 3-month-old male RKIP+/+ or RKIP-/- mice femurs sections. D Quantification of osteoclast number per bone surface (N.Oc/BS) and osteoclast surface per bone surface (Oc.S/BS) (n = 6). E Calcein double staining and Von Kossa staining of 3-month-old male RKIP+/+ or RKIP-/- mice femurs sections. F Quantification of mineral apposition rate (MAR) and bone formation rate per bone surface (BFR/BS) (n = 6). G OCN (green) and DAPI (blue) immunofluorescence of 3-month-old male RKIP+/+ or RKIP-/- mice femurs sections; The arrow indicates the cells which express OCN. H Quantification of OCN+ cell number per bone perimeter (N.OCN+/B.Pm) (n = 6). I Serum C-terminal telopeptide of type I collagen (CTX-1) and Procollagen I N-Terminal Propeptide (PINP) concentration measured by ELISA from RKIP+/+ or RKIP-/- male mice (n = 6). Data are presented as mean ± SD. *P < 0.05, **P < 0.01 for a comparison with the control group or as indicated. n value means the number of repetitions in each independent experiment.
Loss of RKIP has no influence on osteoblast lineage differentiation
Due to the role of osteoblasts (OBs) as crucial cellular components of the bone niche, directly involved in bone formation, we further investigated whether loss of RKIP influenced osteoblast lineage differentiation. RKIP protein expression remained unchanged during the osteogenic induction in wild type (WT) murine bone marrow mesenchymal stem cells (BMSCs), while the expression of osteoblast-specific markers, Runx2 and Sp7, were upregulated in a time-dependent manner (Fig. S1C, D). Similarly, there was no significant change in RKIP gene expression in BMSCs after osteogenic induction (Fig. S1, E). Subsequently, we induced osteogenic differentiation in BMSCs isolated from control or gKO group mice for 7 or 21 days and observed that RKIP deletion did not affect alkaline phosphatase and alizarin red staining (Fig. S1F). Additionally, RKIP knockout also did not affect the expression of Runx2 (Fig. S1G, H).
To further investigate the impact of RKIP on osteoblast lineage differentiation in vivo, we generated three conditional knockout mouse models, Prx1-Cre-RKIPf/f, Osx-Cre-RKIPf/f, and OCN-Cre-RKIPf/f, deleting RKIP, respectively, in BMSCs, pre-osteoblasts, and mature osteoblasts. Interestingly, there was no significant difference in bone mass between these three conditional knockout mice and their control littermates, regardless of whether they were female or male (Figs. S2–S3). H&E staining indicated no apparent differences in skeletal structure between control and osteoblast conditional knockout mice (Fig. S3E–G). These findings demonstrate that loss of RKIP has no influence on osteoblast differentiation in vivo and in vitro. Therefore, further investigation is needed to elucidate the mechanisms of increased bone formation and decreased bone resorption observed in gKO mice.
Conditional knockout of RKIP in macrophages impedes osteoclastogenesis in vivo and in vitro
Osteoclasts (OCs) are derived from macrophages and directly participate in the regulation of bone homeostasis. To investigate the role of RKIP in OCs, we induced osteoclast differentiation in WT mouse bone marrow macrophages (BMMs). Gradual increases in the gene and protein expression of RKIP were observed during the process of osteoclast differentiation (Fig. 2A and Fig. S4A, B). Consistently, more expression and co-localization of RKIP and ACP5 were found in the OVX model and rheumatoid arthritis (RA) model (Fig. 2B and Fig. S4C–E). The RKIP inhibitor (Locostatin) impeded osteoclast formation in a dose-dependent manner (Fig. S4F, G), while RKIP agonist exhibited contrasting effect (Fig. S4H, I). To avoid non-specific effects of RKIP agonist, we overexpressed RKIP in BMMs using adenovirus transfection and found RKIP overexpression significantly enhanced osteoclast differentiation (Fig. S4J, K). These results suggest that RKIP play a positive role in osteoclastogenesis.
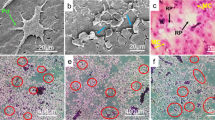
A Western blots of RKIP, c-Fos and NFATc1 protein levels in BMMs under the treatment of M-CSF and RANKL for indicated time. B RKIP (green), ACP5 (red) and DAPI (blue) immunofluorescence staining of femur sections from sham or OVX mice; The arrow indicates the cells which express both RKIP and ACP5. C Micro-CT and three-dimensional (3D) reconstruction images of the distal femur of 3-month-old male RKIPf/f or LysM-Cre-RKIPf/f mice are shown. D Quantification of BV/TV, Tb.N, Tb.Sp, Tb.Th, and Ct.Th (n = 5). E H&E staining and TRAP staining of 3-month-old male RKIPf/f or LysM-Cre-RKIPf/f mice femurs sections. F Quantification of N.Oc/BS and Oc.S/BS (n = 6). G TRAP staining and F-actin immunofluorescence staining to detect osteoclastogenesis of BMMs from RKIPf/f or LysM-Cre-RKIPf/f male mice. H Quantification of size and nuclei numbers of TRAP-positive multinuclear cells (n = 6). I Scanning electron microscopy (SEM) images of bone resorption pits of osteoclast precursors from RKIPf/f or LysM-Cre-RKIPf/f male mice. J Quantification of resorbed area/well (n = 3). K Western blots of RKIP, c-Fos and NFATc1 protein levels in BMMs from RKIPf/f or LysM-Cre-RKIPf/f male mice with or without the treatment of RANKL. Data are presented as mean ± SD. *P < 0.05, **P < 0.01 for a comparison with the control group or as indicated. n value means the number of repetitions in each independent experiment.
To further validate the role of RKIP in vivo, we generated LysM-Cre-RKIPf/f (cKO) mice, which deleted the target gene in the macrophages (osteoclast precursors). Micro-CT analysis revealed increased bone mass and cortical bone thickness in male and female cKO mice compared to littermate controls (Fig. 2C, D and Fig. S5A, B). We used specimens from male mice for the subsequent experiments. H&E and TRAP staining indicated that RKIP knockout in macrophages caused osteosclerosis by inhibiting osteoclast differentiation (Fig. 2E, F). Subsequently, we assessed the osteoclast differentiation capacity of BMMs derived from control and cKO mice. Loss of RKIP in BMMs suppressed osteoclast differentiation (Fig. 2G, H). Following induction of BMMs into osteoclast precursors and subsequent seeding onto bone slices, we observed that RKIP knockout significantly inhibited osteoclast-mediated bone resorption activity in vitro (Fig. 2I, J). Consistently, the elevated expression of osteoclast related-proteins (c-Fos, NFATc1) and genes (ACP5, NFATc1, CTSK and c-Fos) were inhibited in the RKIP knockout group compared with the control group when BMMs were cultured with M-CSF and RANKL (Fig. 2K and Fig. S5C, D).
Additionally, we investigated the potential effects of LysM-Cre-RKIPf/f on bone formation. The cKO mice showed a significant increase in the MAR, BFR/BS (Fig. S5E, F). Immunofluorescent staining showed an unexpected significant increase in the number of OCN+ cells on the bone cortex surface in the cKO group (Fig. S5G, H). Also, decreased CTX-1 and increased PINP serum levels were found in the cKO group compared to the control group (Fig. S5I). These results show that conditional knockout of RKIP in macrophages impedes osteoclastogenesis while coupled bone formation in vivo through unknown mechanisms.
RKIP enhances osteoclastogenesis by interacting with ARHGAP to suppress CDC42 inactivation
Currently, there is a lack of studies about the impact of RKIP on osteoclastogenesis. We observed no significant differences in the expression of MAPK and NF-κB signaling pathway-related proteins in BMMs from RKIP+/+ (control) or RKIP-/- (gKO) mice with RANKL or different concentrations of M-CSF treatment (Fig. S6A–D). Therefore, we isolated BMMs from RKIP+/+ and RKIP-/- mice and performed transcriptome sequencing (RNA-seq) to explore its downstream mechanisms. The RNA-seq revealed 156 upregulated genes and 503 downregulated genes in the gKO group compared to the control group (Fig. S6E, F). GO enrichment analysis revealed that the differentially expressed genes were involved in cell adhesion and positive regulation of cell-cell adhesion processes (Fig. 3A). KEGG enrichment analysis showed that pathways related to osteoclast differentiation and cell adhesion molecules were involved (Fig. S6G). We found that RKIP knockout significantly inhibited the phosphorylation of c-Src and the expression of mRNA for DC-STAMP and OC-STAMP in BMMs stimulated by M-CSF and RANKL (Fig. S6H–J), indicating that RKIP is crucial for the cytoskeletal formation and cell adhesion and migration of osteoclasts. Therefore, using co-immunoprecipitation (Co-IP)-mass spectrometry, we screened potential binding proteins of RKIP and focused on cell adhesion-related proteins. The results identified an interaction between RKIP and CDC42, which played important roles in processes such as cell adhesion, pseudopodia formation, and osteoclast differentiation28,29,30 (Fig. 3B). These findings suggest that CDC42 may be a key molecule through which RKIP regulating osteoclastogenesis.
A GO enrichment analysis of RNA-seq analysis of BMMs from RKIP+/+ and RKIP-/- mice. B CDC42 peptides identified through mass spectrometry are shown. C Western blots of CDC42-GTP and total CDC42 protein levels in BMMs from RKIP+/+ or RKIP-/- male mice with RANKL treatment for indicated time. D RKIP (green), CDC42 (red) and DAPI (blue) immunofluorescence staining of BMMs from RKIP+/+ or RKIP-/- male mice with or without RANKL treatment. E Various truncated fragments of RKIP (R1, 1–90; R2, 91–140; R3, 141–187). R1–R3 and full length of CDC42 plasmid were co-transfected in HEK-293T cells. The interactions were detected through the indicated IP and IB analyses. F Various truncated fragments of CDC42 (C1, 1–47; C2, 48–104; C3, 105–191). C1–C3 and full length of RKIP plasmid were co-transfected in HEK-293T cells. The interactions were detected through the indicated IP and IB analyses. G Overall structure of CDC42 (pink) and RKIP (purple) and key residues of their interaction are shown. H HEK-293T cells were transfected with CDC42-Flag or CDC42-Flag and RKIP-Myc; The interactions were detected through the indicated IP and IB analyses. I HEK-293T cells were transfected with ARHGAP-HA and CDC42-Flag, or ARHGAP-HA, CDC42-Flag and RKIP-Myc; The interactions were detected through the indicated IP and IB analyses. J HEK-293T cells were transfected with RKIP-Myc, ARHGAP-HA or RKIP-Myc and ARHGAP-HA; The interactions were detected through the indicated IP and IB analyses. K The illustration of RKIP inhibiting CDC42 inactivation by interacting with ARHGAP.
CDC42 mediates osteoclastogenesis through its active form (CDC42-GTP)29,30. Therefore, we examined the effect of RKIP on CDC42 activation. Using active CDC42 pull-down and detection kit, we found that RKIP deletion significantly inhibited the upregulation of CDC42-GTP induced by RANKL stimulation (Fig. 3C and Fig. S7A). Conversely, overexpression of RKIP in BMMs significantly promoted CDC42 activation (Fig. S7B, C). Immunofluorescence staining showed co-localization of CDC42 and RKIP at the cellular membrane and within the cell nucleus, with an increased co-localization upon RANKL treatment (Fig. 3D and Fig. S7D). The co-localization at the cellular membrane is likely associated with BMMs cytoskeletal regulation and podosome formation31,32,33. The TRAP staining showed that inhibiting CDC42 or knocking down CDC42 significantly suppressed osteoclastogenesis and rescued the enhanced osteoclast differentiation effect induced by RKIP overexpression (Fig. S7E–H). This indicates that CDC42 is indeed a key downstream molecule involved in RKIP regulating osteoclast differentiation.
To further investigate how RKIP modulates CDC42 activation, we performed Co-IP experiments to confirm the interaction between RKIP and CDC42 in BMMs (Fig. S7I). Additionally, by transfecting CDC42-Flag and RKIP-Myc plasmids into HEK-293T cells, we further validated the interaction between RKIP and CDC42 (Fig. S7J). To map the crucial domains of RKIP and CDC42 that are responsible for their interaction, we constructed truncated fragments of RKIP and CDC42 function domains based on Uniprot database. The truncated fragments were co-transfected with the full-length plasmids of the interacting proteins into HEK-293T cells. R3 region of RKIP (141-187) and the C3 region of CDC42 (105-191) were critical for their interaction (Fig. 3E, F). Molecular docking demonstrated that the key residues (Pro106, Lys107, ASP148) of CDC42 were enriched in C3 region, while the key residues (Ser51, Ser98, Asp105, Asn140) were not within the R3 segment of RKIP (Fig. 3G). We will investigate the reasons behind this finding in future studies.
Further investigation will be conducted to elucidate how the binding of RKIP to CDC42 influences the activation of CDC42. We screened CDC42 activity-related regulatory factors (RACGAP, ARHGAP, C3G, Epac, and Rho GDI) through Co-IP and found that RKIP overexpression specifically promoted the binding between ARHGAP and CDC42 or the CDC42-RKIP complex (Fig. 3H). According to previous researches, ARHGAP promotes the inactivation of CDC4229,30. Therefore, we hypothesized that RKIP might inhibit CDC42 inactivation by competitively binding to ARHGAP. Co-IP showed that RKIP overexpression reduced the binding between ARHGAP and CDC42 (Fig. 3I). Also, we further confirmed the interaction between RKIP and ARHGAP in BMMs and HEK-293T cells (Fig. 3J and Fig. S7K). These results demonstrate that RKIP can competitively bind to ARHGAP, thereby inhibiting CDC42 inactivation to promote osteoclastogenesis (Fig. 3K).
RKIP regulates macrophage differentiation fate and influences H-type vessel formation
As precursors of osteoclasts and participants in bone immune regulation, bone niche macrophages play a crucial role in maintaining bone homeostasis. To further investigate how RKIP knockout in macrophages coupled bone formation in vivo, we selected three male 10-week-old RKIPf/f (control) and LysM-Cre-RKIPf/f (cKO) mice and performed single-cell sequencing (scRNA-seq) on cells isolated from the bone marrow and metaphysis region (Fig. 4A). A total of 9333 cells were captured from the control group, while 8,471 cells were captured from the cKO group. The overall cell dimensionality reduction and clustering analysis revealed that the captured cells could be divided into 23 clusters (Fig. S8A), with characteristic markers and proportions shown in Fig. S8B, C. GO analysis of differentially expressed genes showed significant enrichment in pathways related to regulation of cell-cell adhesion and myeloid cell differentiation (Fig. S8D). Furthermore, differentially expressed genes were found to impact GTPase regulator activity and actin binding molecular function (Fig. S8E), which aligned with our previous findings that RKIP affected osteoclast differentiation by regulating CDC42 activity in macrophages. Similarly, KEGG analysis revealed enrichment in pathways such as focal adhesion and endocytosis, which are closely associated with CDC42 regulation of actin-rich podosome formation (Fig. S8F).
A The illustration of scRNA-seq on cells isolated from the bone marrow and metaphysis. B UMAP dimensionality reduction plot for unsupervised clustering of macrophage subtypes. C Percentage distribution of macrophage subclusters. D Dot plot of the top 5 cluster-specific marker genes of macrophage subclusters. E The box plot of AUCell scores of the pro-angiogenic gene set for each subcluster. F Percentage distribution of non-pro-angiogenic and pro-angiogenic macrophage subcluster in RKIPf/f or LysM-Cre-RKIPf/f male mice. G GO enrichment analysis of pro-angiogenic macrophage subcluster. H The mRNA level of VEGFA-VEGFD and PDGFA-PDGFD in BMMs from RKIP+/+ or RKIP-/- male mice (n = 6). I F-actin immunofluorescence staining of matrigel tube formation assay using conditioned medium (CM) of BMMs from RKIP+/+ or RKIP-/- male mice. J Quantification of cumulative tube length (n = 4). K CD31 (green), Emcn (red) and DAPI (blue) immunofluorescence staining of femur sections from RKIPf/f or LysM-Cre-RKIPf/f male mice. L Quantification of the number of CD31hiEmcnhi cell per trabecular area (N. CD31hiEmcnhi/TB.Ar) and the number of CD31hiEmcnhi per periosteal bone surface (N. CD31hiEmcnhi/P.BS) (n = 6). Data are presented as mean ± SD. *P < 0.05, **P < 0.01 for a comparison with the control group or as indicated. n value means the number of repetitions in each independent experiment.
Then we categorized cells into myeloid cells, B cells, T cells, macrophages, osteoblasts, osteoclasts, endothelial cells, and mesenchymal stem cells using a combination of scMCA auto-annotation and marker annotation (Fig. S8G–I). Subsequently, we examined the proportions of macrophages, mesenchymal stem cells, osteoclasts, osteoblasts, and endothelial cells, which were involved in bone homeostasis regulation. The results indicated that knockout of RKIP in macrophages led to a decrease in the proportions of osteoclasts, while the proportion of macrophages increased (Fig. S8J). There was a slight decrease of the proportion of endothelial cells to total cells in cKO group, however, the proportion of CD31hi Emcnhi H-type endothelial cells (the vascular type promoting bone formation) to total endothelial cells was increased (Fig. S8J, K). This suggests that bone formation may be promoted by an increase in the generation of H-type vessels in the cKO mice.
Considering the crucial regulatory role of macrophages in osteoclastogenesis and vascular formation, we further investigated whether RKIP deletion affected macrophage differentiation fate, leading to an increased generation of H-type vessels in vivo. Developmental trajectory analysis revealed that myeloid progenitor cells could differentiate into two distinct cellular subsets: pro-angiogenic macrophages and osteoclasts (Fig. S9A). Therefore, we performed dimensionality reduction clustering on macrophages, resulting in the identification of 12 subclusters (Fig. 4B), with characteristic markers and proportions shown in Fig. 4C, D. As demonstrated earlier (Fig. S8J), RKIP knockout suppressed the differentiation of macrophages into osteoclasts. Therefore, our focus was on the proportion of the pro-angiogenic subclusters. Due to the low distinguishing ability of identifying macrophage clusters based on pro-angiogenic markers, we used gene set enrichment scoring to identify pro-angiogenic subtypes of macrophages. The box plot shows the AUCell scores for the pro-angiogenic macrophage gene set (VEGFA, TEK, SEMA4D, CXCL8, ANG2, ANGPT1) across different clusters. It can be seen that subclusters 1, 2, 3, 6, 7, 9, and 11 have higher scores for the pro-angiogenic gene set, thus classifying them as pro-angiogenic macrophages (Fig. 4E). The stacked bar plot demonstrated a significant increase in the proportion of pro-angiogenic macrophage subcluster in the cKO group (Fig. 4F), with subcluster 6 exhibiting the most pronounced increase (Fig. 4C). We performed enrichment analysis on differentially expressed genes in pro-angiogenic macrophages between the control and cKO groups. GO analysis revealed enrichment in pathways related to regulation of vasculature development and angiogenesis (Fig. 4G). KEGG analysis revealed that the differentially expressed genes were enriched in to IL-17, TNF, and Toll-like receptor signaling pathways, all of which are closely associated with angiogenesis (Fig. S9B)34,35,36. The scRNA-seq results suggest that RKIP is a key molecule controlling macrophage differentiation fate. RKIP deletion inhibits macrophage differentiation into osteoclasts while promoting its differentiation into the pro-angiogenic subclusters.
To further validate the above findings, we extracted BMMs from the control and gKO mice and found RKIP knockout enhanced the expression of angiogenic genes such as VEGFA, VEGFC, PDGFA, PDGFB, and PDGFD (Fig. 4H). Conditioned medium of BMMs from gKO mice significantly promoted the tube formation of human umbilical vein endothelial cells (HUVECs) (Fig. 4I, J). Significantly higher numbers of H-type endothelial cells (CD31hi Emcnhi) were observed in the trabecular bone and cortical bone surface of gKO mice and cKO mice compared with their respective littermate controls (Fig. 4K, L and Fig. S9C, D). To further elucidate whether the observed phenotype in RKIP knockout mice results from suppressed osteoclast differentiation or enhanced osteoblast differentiation mediated by angiogenesis, we supplemented mouse models treated with angiogenesis inhibitors (Bevacizumab). We also employed the AAV carrying an F4/80 macrophage-specific promoter to knockdown VEGFA in BMMs in vivo. The results demonstrated that angiogenesis inhibition could partially rescue the elevated bone mass caused by macrophage-specific RKIP knockout. However, under the same anti-angiogenic conditions, RKIP-deficient mice still exhibited higher bone mass than wild-type controls (Fig. S10A–F). These findings further confirm that the increased bone mass resulting from RKIP deletion is mediated by both suppressed osteoclast differentiation and the increased differentiation of macrophages into the pro-angiogenic subclusters, leading to upregulated H-type vessel generation.
Intranuclear RKIP modulates angiogenesis by regulating the protein stability of HIF-1α in macrophages
It has been reported that HIF-1α significantly promotes angiogenesis and possible interacts with RKIP37,38. Therefore, we investigated whether RKIP affected the expression of HIF-1α. Loss of RKIP in BMMs significantly upregulated the protein expression of HIF-1α under hypoxia conditions, while it had no effect on the expression of HIF-2α (Fig. 5A and Fig. S11A). Immunofluorescence staining revealed co-localization of RKIP and HIF-1α in the nucleus of BMMs, and knockout of RKIP markedly increased nuclear HIF-1α levels (Fig. 5B and Fig. S11B), indicating an interaction between RKIP and HIF-1α. We further investigated whether RKIP in macrophages regulates angiogenesis through HIF-1α. BMMs from control and gKO mice were treated with the HIF-1α inhibitor (PX-478), and qPCR results showed that inhibiting HIF-1α significantly rescued the upregulation of angiogenesis-related gene expression caused by RKIP deletion (Fig. S11C). Similarly, treatment with PX-478 also significantly suppressed the increased protein expression of VEGF induced by RKIP deletion (Fig. S11D, E). Furthermore, PX-478 significantly reduced the pro-angiogenic ability of gKO BMMs CM (Fig. S11F, G). These findings indicate that RKIP in macrophages regulates angiogenesis through HIF-1α. Previous study has reported that HIF-1α promoted osteoclast differentiation39, and we have proved RKIP knockout inhibited osteoclast differentiation. To reconcile these contrasting phenotypes, we treated BMMs with the CDC42 inhibitor (ZCL278) under normoxic or hypoxic conditions. ZCL278 treatment suppressed the enhanced osteoclast differentiation caused by hypoxia (Fig. S11H, I), suggesting that CDC42 plays a more crucial role in osteoclast differentiation. Inhibition of CDC42 can counteract the promoting effect of HIF-1α on osteoclast differentiation.
A Western blots of HIF-1α, HIF-2α and RKIP protein levels in BMMs from RKIP+/+ or RKIP-/- male mice under hypoxia culture for indicated time. B RKIP (green), HIF-1α (red) and DAPI (blue) immunofluorescence staining of BMMs from RKIP+/+ or RKIP-/- male mice under hypoxia culture for 24 h. C Western blots of HIF-1α and RKIP protein levels in BMMs following the adenovirus transduction of RKIP or the empty vector with the treatment of MG132 or chloroquine (CQ) under hypoxia culture for 48 h. D Western blots of the ubiquitination of HIF-1α in BMMs from RKIP+/+ or RKIP-/- male mice under hypoxia culture for 24 h, which were immunoprecipitated with anti-HIF1α. E BMMs were cultured under normoxia or hypoxia for 48 h; The interactions were detected through the indicated IP and IB analyses. F Western blots of HIF-1α, VHL, and RKIP in cytoplasmic and nuclear of BMMs from RKIP+/+ or RKIP-/- male mice under hypoxia culture for 48 h. G The illustration and images of proximity ligation assays (PLA) of indicated proteins in BMMs under hypoxia culture for 48 h. H Putative murine, rat, human, and canine RKIP NLS sequences, predicted from analysis with the PredictNLS program. I Flag(green), HIF-1α (red) and DAPI (blue) immunofluorescence staining of HEK-293T cells transfected with RKIP-Flag or RKIP-NLS-Del-Flag plasmid, which were under hypoxia culture for 24 h. J Western blots of HIF-1α, VHL, and Flag in cytoplasmic and nuclear of HEK-293T cells transfected with RKIP-Flag or RKIP-NLS-Del-Flag plasmid, which were under hypoxia culture for 48 h. K RKIP(green), HIF-1α (red) and DAPI (blue) immunofluorescence staining of BMMs transfected with NC or KPNB1 siRNA, which were under hypoxia culture for 24 h.
qPCR showed no changes in the gene expression of HIF-1α and HIF-2 in BMMs from control and gKO mice (Fig. S12A). However, RKIP overexpression significantly decreased HIF-1α protein levels, while MG132 (proteasome inhibitor) effectively rescued this phenomenon, whereas CQ (lysosome inhibitor) was ineffective (Fig. 5, C and Fig. S12B). Therefore, we hypothesized that RKIP overexpression triggered proteasome-dependent degradation of HIF-1α protein in BMMs. Ubiquitination detection following MG132 treatment showed that RKIP deletion reduced poly-ubiquitination levels of HIF-1α (Fig. 5D). These results indicate that RKIP promotes HIF-1α degradation through the ubiquitin-proteasome pathway.
It has been indicated that hydroxylated HIF-1α undergoes ubiquitination modification by VHL in the nucleus and is subsequently translocated to the proteasome for degradation40. We observed co-localization of RKIP and HIF-1α in the nucleus, suggesting that RKIP may regulate the degradation of HIF-1α by promoting its ubiquitination by VHL. To validate this hypothesis, we investigated the association of RKIP with the HIF-1α degradation complex. Co-IP showed that under both normoxic and hypoxic conditions, RKIP, VHL, and HIF-1α formed a trimeric complex in BMMs (Fig. 5E). RKIP loss reduced the binding between VHL and HIF-1α (Fig. S12C). Furthermore, RKIP deletion inhibited the nuclear localization of VHL in BMMs, resulting in increasing the stability of HIF-1α (Fig. 5F and Fig. S12D). Proximity ligation assay (PLA) showed that RKIP directly interacted with VHL and HIF-1α in the nuclei of BMMs (Fig. 5, G).
To further confirm the intranuclear function of RKIP, we conducted sequence alignment to confirm the conservation of the nuclear localization sequence (NLS) of RKIP (Fig. 5H). Based on the alignment results, we constructed a nucleus-excluded form of RKIP, in which the NLS is deleted (RKIP-NLS-Del). RKIP overexpression promoted HIF-1α degradation without affecting VHL expression levels, while RKIP-NLS-Del did not exert the same effects (Fig. S12E). RKIP-NLS-Del inhibited VHL nuclear localization and stabilized intranuclear HIF-1α protein in HEK-293T cells (Fig. 5I, J). In addition to the NLS, protein nuclear localization also requires binding to nuclear transport proteins. We reviewed our mass spectrometry results and validated that RKIP can interact with the nuclear transport protein KPNB1 (Fig. S12F). KPNB1 knockdown had no effect on VHL protein expression but inhibited HIF-1α degradation (Fig. S12G). Similarly, KPNB1 knockdown resulted in decreased nuclear localization of RKIP and VHL, thereby increasing the expression of HIF-1α in both the cytoplasm and nucleus (Fig. 5K and Fig. S12H). To further confirm the role of nuclear RKIP in BMMs on angiogenesis, we cultured HUVECs using the CM from BMMs following adenovirus transduction of RKIP or the empty vector, with KPNB1 negative control or siRNA transfection. The results showed that overexpression of RKIP in BMMs significantly inhibits the angiogenic capacity of their CM, while blocking the nuclear translocation of RKIP can rescue this phenotype (Fig. S12I, J). These observations indicate that RKIP bridges the association between VHL and HIF-1α in the nucleus to facilitate HIF-1α degradation.
Knockout of RKIP and RKIP inhibitor prevent OVX-induced bone loss
Given the role of RKIP in regulating differentiation fate of bone niche macrophages, we were interested in whether RKIP deletion would protect the mouse from OVX-induced osteoporosis. RKIP knockout alleviated OVX-induced bone loss compared with the control littermates (Fig. S13A, B). H&E staining of the femurs confirmed the micro-CT results (Fig. S13, C). TRAP staining showed RKIP cKO mice had fewer osteoclasts on the trabecular surface of the femur compared with control mice in the sham and OVX groups (Fig. S13C, D). These findings suggest that RKIP deletion in macrophages inhibits osteoclast differentiation, thereby mitigating OVX-induced bone loss.
Further, we performed intramedullary injection of Sh-RKIP AAV into the femur and tibia of mice using a microsyringe to investigate the effect of RKIP knockdown in the bone marrow cavity on bone mass (Fig. S14A). To verify the knockdown efficiency of RKIP, we collected the femurs of the mice and extracted BMMs 2 months after AAV injection. The results showed a significant decrease in RKIP expression levels in BMMs following Sh-RKIP AAV injection (Fig. S14B, C). Then we found RKIP knockdown alleviated the decrease in bone mass (Fig. 6A, B), and inhibited the enhanced osteoclast differentiation induced by OVX (Fig. S14D, E). OVX increased co-localization of ACP5 and RKIP, while RKIP knockdown significantly suppressed the phenomenon (Fig. S14F, G). Additionally, RKIP knockdown significantly increased MAR and BFR/BS in vivo (Fig. S15A, B). The number of osteoblasts (OCN+) and H-type vessel formation also increased in the RKIP knockdown groups (Fig. S15C–F), which leading to increased bone formation and thereby rescuing OVX-induced bone loss.
A Micro-CT and three-dimensional reconstructed images of distal femurs from intramedullary injection of Sh-NC or Sh-RKIP adeno-associated virus (AAV) mice that underwent either sham or OVX operation. B Quantification of BV/TV, Tb.N, Tb.Sp, Tb.Th, and Ct.Th (n = 6). C Micro-CT and three-dimensional reconstructed images of distal femurs from sham and OVX mice injected with saline or Locostatin (1 mg/kg) for 2 months. D Quantification of BV/TV, Tb.N, Tb.Sp, Tb.Th, and Ct.Th (n = 6). E CD31 (green), Emcn (red) and DAPI (blue) immunofluorescence staining of femur sections from sham and OVX mice injected with saline or Locostatin (1 mg/kg). F Quantification of N. CD31hiEmcnhi/TB.Ar and N. CD31hiEmcnhi/P.BS (n = 6). Data are presented as mean ± SD. *P < 0.05, **P < 0.01 for a comparison with the control group or as indicated. n value means the number of repetitions in each independent experiment.
We also treated mice with Locostatin for 8 weeks following OVX operation. Similar to the results obtained with macrophage conditional knockout of RKIP or in situ RKIP knockdown in the bone marrow cavity, treatment with Locostatin rescued the bone loss induced by OVX (Fig. 6C, D) through mechanisms such as inhibiting osteoclast differentiation (Fig. S16A–D) and enhancing H-type vessel generation (Fig. 6E, F and Fig. S16E, F). In conclusion, knockout of RKIP and RKIP inhibitor can prevent OVX-induced bone loss by inhibiting bone resorption and promoting bone formation.
RKIP regulates osteoclast differentiation in the human
The role of RKIP regulating osteoclast differentiation was verified in human trabecular bone and peripheral blood mononuclear cells (PBMCs). A significant increase in co-localization of RKIP and ACP5 was observed on the trabecular bone surface of patients in the osteoporosis group (Fig. 7A, B). The gene and protein expression of RKIP were higher in human PBMCs of osteoporosis patients compared to those of normal bone mass patients (Fig. 7C–E). Consistently, it was found that RKIP were upregulated in a time-dependent manner with the stimulation of h-RANKL in PBMCs (Fig. 7F and Fig. S17A, B). These findings suggest that RKIP gradually upregulates during the differentiation of human PBMCs into osteoclasts.
A RKIP (green), ACP5 (red) and DAPI (blue) immunofluorescence staining of femur sections from normal or osteoporosis patients; The arrow indicates the cells which express both RKIP and ACP5. B Quantification of the number of ACP5+RKIP+ cells per trabecular perimeter (N. ACP5+RKIP+/TB. Pm) (n = 6). C Western blots of RKIP, c-Fos protein levels in PBMCs from normal or osteoporosis patients. D Quantification and normalization of the gray levels of RKIP and c-Fos proteins to that of GAPDH using Image J (n = 3). E The mRNA level of RKIP in PBMCs from normal or osteoporosis patients (n = 3). F Western blots of RKIP, c-Fos, and NFATc1 protein levels in PBMCs with h-M-CSF and h-RANKL treatment for indicated time. G TRAP staining to detect osteoclastogenesis of PBMCs with various concentrations of Locostatin treatment. H Quantification of size and nuclei numbers of TRAP-positive multinuclear cells (n = 6). I The mRNA level of ACP5, CTSK, NFATc1 in PBMCs with vehicle or Locostatin treatment in the presence or absence of h-RANKL for 10 days (n = 6). J Western blots of RKIP, c-Fos, and NFATc1 protein levels in PBMCs with vehicle or Locostatin treatment in the presence or absence of h-RANKL for 10 days. K TRAP staining to detect osteoclastogenesis of PBMCs with various concentrations of Didymin treatment. L Quantification of size and nuclei numbers of TRAP-positive multinuclear cells (n = 6). M The mRNA level of ACP5, CTSK, NFATc1 in PBMCs with vehicle or Didymin treatment in the presence or absence of h-RANKL for 10 days (n = 6). N Western blots of RKIP, c-Fos, and NFATc1 protein levels in PBMCs with vehicle or Didymin treatment in the presence or absence of h-RANKL for 10 days. Data are presented as mean ± SD. *P < 0.05, **P < 0.01 for a comparison with the control group or as indicated. n value means the number of repetitions in each independent experiment.
We also employed the inhibitor (Locostatin) and agonist (Didymin) of RKIP to treat PBMCs during the osteoclast induction. Osteoclast formation following the Locostatin treatment was significantly inhibited in a dose-dependent manner PBMCs (Fig. 7G, H). The results were further confirmed by the fact the elevated expression of osteoclast related-genes and proteins were inhibited by Locostatin treatment when PBMCs were cultured with h-M-CSF and h-RANKL (Fig. 7I, J and Fig. S17C). Conversely, Didymin treatment promoted osteoclastogenesis (Fig. 7K, L) and osteoclast related-genes and proteins expression (Fig. 7M, N and Fig. S17D). In summary, RKIP regulates osteoclast differentiation in human PBMCs in vivo and in vitro. RKIP is a critical molecule regulating the differentiation fate of bone niche macrophage via the RKIP/ARHGAP/CDC42 and RKIP/VHL/HIF-1α pathways (Fig. 8).
Schematic illustration showing the different roles of extranuclear or intranuclear RKIP in differentiation fate regulation of bone niche macrophages. RKIP is a critical molecule regulating the differentiation fate of bone niche macrophage via the RKIP/ARHGAP/CDC42 and RKIP/VHL/HIF-1α pathways.
Discussion
Bone and immune cells reside within the same microenvironment and engage in reciprocal interactions, collectively contributing to the functions of the ‘osteoimmune system’, which encompasses all cells present in the bone marrow41. Macrophages, serving as pivotal cells in regulating bone immunity, undergo development from hematopoietic stem cells, and their differentiation characteristics are determined by the tissue niche they reside in. Upon migration of circulating monocytes to tissues, they differentiate into tissue-specific macrophages such as osteoclasts in the skeletal system, microglia in the central nervous system, alveolar macrophages in the lungs, and Kupffer cells in the liver42. Bone niche macrophages can exert their influence on bone resorption and formation through various mechanisms, including direct differentiation into osteoclasts, polarization into M1/M2 phenotypes to modulate inflammatory responses, and secretion of relevant factors to impact angiogenesis13,14,43. Thus, the differentiation fate of macrophage plays a crucial role in regulating bone homeostasis.
Our previous studies have indicated that RKIP participates in immune regulation through various pathways, including inhibition of inflammasome activation, promotion of type I interferon production, and positive regulation of the IL-17 signaling pathway22,24,27. Additionally, the loss of RKIP does not affect the response of macrophages to LPS, suggesting that RKIP may not be involved in regulation of classical macrophage M1/M2 polarization23. To further investigate the role of RKIP in bone homeostasis regulation, we first generated RKIP gKO mice and found that they exhibited increased bone mass due to suppressed bone resorption and enhanced bone formation. Osteoblasts are primarily responsible for bone formation and are essential components of the bone niche. To explore the impact of RKIP on osteoblast lineage differentiation, we further generated Prx1-Cre-RKIPf/f, Osx-Cre-RKIPf/f, OCN-Cre-RKIPf/f mice to delete RKIP specifically in early, intermediate, or late-stage of osteoblast differentiation. Research indicates that neither Prx1-Cre nor OCN-Cre has an impact on mouse skeletal development, while Osx-Cre can lead to skeletal defects in mice44,45,46. Therefore, we used different control mice in our subsequent studies. Interestingly, we observed no significant effect on bone mass in those conditional knockout mice. In vitro experiments confirmed that RKIP didn’t affect osteoblast differentiation.
Although RKIP does not influence macrophage M1/M2 polarization, it may still regulate bone homeostasis through modulation of macrophage differentiation fate. Therefore, we generated LysM-Cre-RKIPf/f mice to delete RKIP specifically in macrophages to observe its impact on bone mass. The results revealed that, similar to RKIP gKO mice, LysM-Cre-RKIPf/f mice exhibited a significant phenotype characterized by inhibited bone resorption and enhanced bone formation. Through RNA-seq, mass spectrometry, and in vivo and in vitro experiments, we discovered that RKIP competitively binds to ARHGAP to inhibit CDC42 inactivation and promote osteoclast differentiation. Studies have shown that ARHGAP possesses two key functional domains: the GAP domain and the SH3 domain. The GAP domain is the primary functional domain of ARHGAP, and it interacts with the active center of CDC42, promoting GTP hydrolysis to GDP and consequently inactivating CDC42. Additionally, the SH3 domain of ARHGAP can interact with other proteins, thereby regulating the function of ARHGAP and influencing the activity state of CDC4247,48,49. We have identified the critical region of interaction between RKIP and CDC42 (R3 (141-187) and C3 (105-191)). However, the key regions and amino acid residues involved in the binding of RKIP and ARHGAP, whether their interaction alters the conformation or function of ARHGAP’s GAP/SH3 domains, and whether the binding of RKIP to CDC42 protects its active center from the action of GTP hydrolyzing enzymes to promote CDC42 activation, are all worthy of further investigation.
The loss of RKIP in macrophages inhibits their differentiation into osteoclasts, which can explain the phenotype of suppressed bone resorption in transgenic mice. However, it cannot fully account for the enhanced bone formation phenotype. Therefore, we performed scRNA-seq on bone marrow and metaphyseal region cells of RKIPf/f and LysM-Cre-RKIPf/f mice. The results revealed that the knockout of RKIP in macrophages inhibited their differentiation into osteoclasts and promoted their differentiation towards the pro-angiogenic subcluster, thereby increasing the proportion of H-type endothelial cells. The scRNA-seq results further confirm that RKIP is a critical molecular regulator of differentiation fate of macrophage within the bone niche, and its presence or absence can drive macrophages towards distinct differentiation directions. However, due to the limited discriminatory power of markers for macrophage clustering based on pro-angiogenic properties, we could only utilize gene set scoring to identify the pro-angiogenic subcluster of macrophages. This limitation prevents us from identifying specific surface markers of distinguishing macrophage subclusters between the RKIPf/f and LysM-Cre-RKIPf/f mice.
Hypoxia-inducible factor 1 (HIF-1) is a key transcriptional regulatory factor involved in cellular responses to hypoxic stress. Under normoxic conditions, the proline residues of HIF-1α protein are hydroxylated by prolyl hydroxylase (PHD), followed by binding to von Hippel-Lindau protein (VHL) and rapid degradation through the ubiquitin-proteasome system50,51. In hypoxic conditions, hydroxylation of HIF-1α is inhibited, allowing it to enter the nucleus and bind to HIF-1β, consequently initiating the transcription of VEGF and PDGF to promote angiogenesis and bone formation through vascular-bone coupling52,53. Computational simulations have shown that RKIP can interact with HIF-1α and inhibit its function37. Although some studies have reported the nuclear localization of RKIP54, its nuclear function remains unclear. In this study, we identified for the first time the NLS of murine RKIP and its nuclear transport protein KPNB1. Upon nuclear translocation, RKIP promoted the binding of VHL to HIF-1α, increased the ubiquitination level of HIF-1α, and facilitated HIF-1α degradation through the proteasome pathway. Deletion of RKIP or inhibition of its nuclear translocation in macrophages could increase the protein stability of HIF-1α and promote angiogenesis.
To evaluate the therapeutic potential of targeting RKIP in vivo, we constructed OVX-induced osteolysis models. As expected, deletion or inhibitor of RKIP could evidently rescue bone loss in OVX mice. Furthermore, the role of RKIP regulating osteoclast differentiation was confirmed in human trabecular bone and PBMCs. However, there are still numerous challenges that need to be addressed in order to develop drugs targeting RKIP for the treatment of osteoporosis. For instance, besides its impact on bone mass, it is essential to further investigate whether RKIP inhibitors have toxic side effects on other vital organs. To address the issue, the optimal approach would be to utilize peptide-based or vesicle-based platforms specifically targeting osteoclasts to package or modify the inhibitor, avoiding off-target effects. Additionally, further studies are required to determine the optimal drug concentration, half-life, metabolism, and biosafety of the inhibitor in vivo.
In conclusion, our study is the first to clarify RKIP is a critical molecule regulating the differentiation fate of bone niche macrophage via the RKIP/ARHGAP/CDC42 and RKIP/VHL/HIF-1α pathways. The study systematically elucidates the nuclear translocation mechanism and extranuclear or intranuclear functions of RKIP. This research demonstrates that RKIP can be a potential target for treatment of bone homeostasis-related disorders.
Methods
Reagents
Locostatin (#HY-W013411A, MCE, USA), Didymin (#HY-N2068, MCE, USA), recombinant mouse M-CSF (#CB34, Novoprotein, China) and RANKL (#462-TEC, R&D, USA), recombinant human M-CSF (#C417, Novoprotein, China) and RANKL (#CK63, Novoprotein, China), TRAP staining kits (#387A-1KT, Sigma-Aldrich, USA), Mouse CTX-1 ELISA Kit (#CSB-E12782m, CUSABIO, China), Mouse PINP ELISA Kit (#CSB-E12775m, CUSABIO, China) were used. Alpha-modified Eagle’s medium (α-MEM), Dulbecco’s modified Eagle’s medium (DMEM) and fetal bovine serum (FBS) were purchased from Gibco (USA).
Mice
RKIP gKO, RKIPflox/flox and LysM-Cre mice were provided by Prof. Xiaojian Wang (Institute of Immunology, Zhejiang University School of Medicine, China). Prx1-Cre, Osx-Cre and OCN-Cre mice were purchased from Cyagen Biosciences Inc. In all experiments, RKIP gKO, Prx1-Cre-RKIPf/f, Osx-Cre-RKIPf/f, OCN-Cre-RKIPf/f or LysM-Cre-RKIPf/f mice were compared with their control littermates. Before euthanizing the mice and collecting samples such as femurs, serum, and bone marrow cells, the mice were fasted and water-deprived for 12 h. All animal studies were approved by the Ethics Committee of Zhejiang University School of Medicine (ZJU20230531).
Clinical specimen
Collection of 10 mL peripheral blood and trabecular bone tissue (obtained during procedures such as fracture treatment and joint replacement) from patients with normal bone mass and osteoporosis, along with relevant clinical data (age, gender, bone mass), was performed. The inclusion and exclusion criteria for patient selection are as follows. Inclusion criteria: (1) The patients with normal bone mass and osteoporosis. (2) The patients requiring surgical treatment for fracture and joint replacement. (3) The patients who can understand the informed consent form and willingly sign it. (4) The patients with complete clinical data. Exclusion criteria: (1) The patients with severe blood system disorders unsuitable for blood sampling. (2) The patients with other severe conditions affecting bone mass, such as systemic immune system diseases. (3) Vulnerable populations, including individuals with mental illnesses, cognitive impairments, critically ill patients, minors, and pregnant women. Bone density examination results within the range of −1 ≤ T ≤ 1 indicate normal bone mass, while T < −2.5 indicates osteoporosis. We have obtained informed consent from all participants. All studies were approved by the Ethics Committee of Sir Run Run Shaw Hospital, Zhejiang University School of Medicine (K2024486).
Ovariectomy (OVX)-induced osteoporosis model
Briefly, sham or bilateral OVX operation was performed on 12-week-old female C57BL/6 mice under isofluorane anesthesia. For Locostatin treatment, the mice were randomly divided into three groups: the sham group, the OVX with saline treatment group, and the OVX with Locostatin treatment group. Saline or Locostatin (1 mg/kg) was intraperitoneally injected three times every week for 2 months. For AAV treatment, intramedullary injection of vector or Sh-RKIP AAV in sham or OVX group was performed. All the mice were euthanized 2 months after operation. Femurs and tibias were gathered and fixed in 4% paraformaldehyde for micro-CT and histological staining.
Intramedullary injection of AAV in mice
Mice were anesthetized and their knees were flexed. We removed the hair around the joints and disinfected the joint area. A 1 mL syringe was then inserted into the femoral intercondylar space, gently twisting and applying pressure between the femoral condyles, leaving the 25-gauge needle in place while gently removing the 1 mL syringe. Next, a microsyringe was inserted into the 25-gauge needle, and 10 µL of AAV solution was slowly injected into the femoral marrow cavity. The needle was then slowly withdrawn to minimize backflow. The method for injecting into the tibial marrow cavity was the same.
Rheumatoid arthritis (RA) mouse model
Breeding was conducted between KRN transgenic mice carrying the TCR gene and non-obese diabetic mice. The offspring, known as K/BxN mice, spontaneously develop RA. Non-arthritis littermates from the same breeding were selected as controls. When the K/BxN mice reached 6–8 weeks of age, corresponding to the peak severity of spontaneous arthritis symptoms, euthanasia was performed. The ankle and paw tissues were collected and fixed in a 4% paraformaldehyde for immunohistochemical analysis.
Micro-CT analysis
For micro-CT analysis, mouse distal femurs were scanned by the micro-CT scanner (Skyscan1275, Aartselaar, Belgium) using the X-ray energy of 80 μA/70 kV at a resolution of 9 μm. The analysis of trabecular bone was performed starting from the growth plate with an interval of 60 slices as the starting point, and 100 slices were analyzed downwards. For cortical bone analysis, the starting point was set at an interval of 250 slices from the growth plate, and 100 slices were analyzed downwards. The parameters analyzed included BV/TV (trabecular bone volume per total volume), Tb. N (trabecular number), Tb. Sp (trabecular separation), and Tb. Th (trabecular thickness) and Ct. Th (cortical thickness). Three-dimensional reconstruction of the analysis region was performed using CTvox software.
Cell culture
HEK-293T cells (GNHu17) were obtained from National Collection of Authenticated Cell Cultures, Chinese Academy of Sciences. HUVEC cells (SV40) were obtained from FENGHUISHENGWU, China. The cell line was confirmed to be free of mycoplasma contamination using PCR and cell culture. The identity of the cell line was authenticated with STR profiling. Murine bone marrow-derived macrophages (BMMs) were isolated from the femurs and tibias of 6-week-old C57BL/6 mice and cultured in alpha-MEM with 10% FBS and 25 ng/mL M-CSF for 4 days. BMMs were induced into osteoclasts with 25 ng/mL M-CSF and 50 ng/mL RANKL treatment for 5 days. The human PBMCs were isolated using Ficoll separation liquid (#P9011, Solarbio). For induction of osteoclasts, PBMCs were treated with 25 ng/mL human M-CSF and 50 ng/mL human RANKL for 10 days. Murine BMSCs, were isolated from the femurs and tibias of 6-week-old C57BL/6 mice and cultured in α-MEM with 10% FBS for 4 days. BMSCs were differentiated into osteoblasts with 10 mM vitamin C and 1 M β-glycerophosphate treatment for 21 days.
Single-cell sequencing analysis of bone niche macrophage characteristics
Three male 10-week-old LysM-Cre-RKIPf/f (cKO) mice and their littermate controls (RKIPf/f, control) were selected. Bone marrow and metaphyseal region cells were isolated from these mice and processed into single-cell suspensions. Single-cell RNA sequencing was performed by High Precision (Hangzhou) Life Technology Co., LTD. Briefly, scRNA-Seq libraries were prepared using the Single Cell 3’ Reagent Kit v2 and the samples were sequenced on the Illumina Hiseq XTEN platform. The raw data were standardized and differentially analyzed using the Seurat v3-MAST package in the R software. UMAP analysis was performed on the filtered principal component gene ranking results obtained from principal component analysis to visualize cell populations with high differences. We identified highly differentially expressed genes within each cluster, which were then intersected with marker genes for cell type annotation. The clusters were annotated based on the expression patterns of marker genes. Further analysis focused on the differences in macrophage subclusters between the control and cKO groups. The scoring of pro-angiogenic gene set in different macrophage clusters was evaluated, along with determining the proportions of pro-angiogenic macrophage subcluster. Gene Ontology and KEGG analysis for differentially expressed genes were performed on www.metascape.org/.
RNA-sequencing
BMMs from control and RKIP gKO mice were isolated and then plated at a density of 5 × 105 cells per well in a 6-well plate. The cells were cultured in a medium containing 25 ng/mL M-CSF for 2 days. Then, 1 mL of TRIZOL was added to each well to fully lyse the cells and extract their RNA for subsequent transcriptome sequencing. The raw data from the transcriptome sequencing was preprocessed and normalized using the robust multi-array average (RMA) method in the affy package of R software (version 4.0.1). Differential gene analysis between samples was performed using the limma package, and subsequent multiple hypothesis testing and correction were conducted after obtaining P values. Gene Ontology and KEGG analysis were performed on the differentially expressed genes to identify relevant differential pathways.
Co-immunoprecipitation (co-IP) assay
BMMs or HEK-293T cells were treated differently and immersed in lysis buffer supplemented with 1 mM dithiothreitol, 1 mM PMSF and Protease Inhibitor Cocktail (PIC). The lysates were immunoprecipitated with Flag-Tag mouse mAb (#M1403-2, HUABIO, China, 1:100), HA-Tag mouse mAb (sc-53516, Santa Cruz, USA, 1:50), Myc-Tag mouse mAb (#30601ES60, YEASON, China, 1:100), RKIP rabbit mAb (#ET1610-18, HUABIO, China, 1:100), HIF-1α rabbit mAb (#36169, CST, USA, 1:100), mouse (G3A1) mAb IgG1 isotype control (#5415S, CST, USA) or rabbit (DA1E) mAb IgG isotype control (#3900S, CST, USA) according to experimental design at 4 °C overnight. And then the mixtures were incubated with protein A/G-beads for 3 h at 4 °C. The beads were washed five times with PBS supplemented with PMSF and PIC at 4 °C, resolved by 10% SDS buffer. The samples were heated at 100 °C for 10 min and then detected by western blotting.
Immunoprecipitation and mass spectrum analysis
BMMs were isolated and cultured in 10-cm dishes, respectively transfected with RKIP-Myc adenovirus and its control adenovirus (MOI = 100). The BMMs were lysed and immunoprecipitated with anti-IgG or anti-Myc antibody after 72 h. Following confirmation of RKIP overexpression and IP efficiency, the immunoprecipitated protein samples were sent to Novogene (China) for liquid chromatography with tandem mass spectrometry (LC-MS) analysis. The raw mass spectrometry data has been uploaded to iProX (Project ID: IPX0012888000).
Immunofluorescence staining
For cell immunofluorescence staining, cells are seeded on cell slides and fixed with 4% paraformaldehyde. The fixed cells were blocked with 5% BSA for 1 h at room temperature after permeabilized by 0.5% Triton X-100. Then the cells were incubated with primary antibodies at 4 °C overnight. After being washed with TBST for five times, the cells were incubated with corresponding secondary antibodies (Invitrogen) and DAPI. Confocal microscope (Nikon, Japan) was used to capture the images of immunofluorescence. For tissue immunofluorescence staining, the bone sections were submerged with sodium citrate at 55 °C overnight for antigen retrieving. After being blocked with BSA for 1 h at room temperature, the sections were incubated with primary antibodies at 4 °C overnight and washed with TBST for five times. Then the sections were incubated with the secondary antibodies and DAPI at room temperature for 2 h. After being washed with TBST for five times, the images of sections were captured by confocal microscope (Nikon, Japan).
Proximity ligation assay (PLA)
According to the instructions of the PLA detection kit (#DUO92102, Sigma, USA), the experimental procedure can be summarized as follows: Cells are seeded on cell slides and fixed with 4% paraformaldehyde. After blocking with BSA for 1 h, incubation with primary antibodies (from different species) against the two target proteins is performed. Subsequently, Duolink PLA probes are incubated, followed by ligation, amplification, final washes. The images were captured by confocal microscope (Nikon, Japan).
Western blot analysis
Cells were seeded in 6-well plates (5 × 105 cells/well) and treated differently according to experimental design. The proteins were extracted using RIPA (#R0020, Solarbio, China), which contained 1 mM PMSF (#ST506, Beyotime, China), PIC (#P1005, Beyotime, China) and Phosphatase Inhibitor Cocktail (#P1045, Beyotime, China). The lysates were centrifuged at 12,000 rpm for 5 min and the supernatants were heated at 100 °C for 10 min. We used SDS-PAGE to separate proteins and transferred them to 0.45-μm PVDF membranes according to the previous protocol55. The protein bands were detected using Amersham Imager 600 (GE, USA) and then quantified by ImageJ. The antibodies used for western blotting were listed in the reporting summary.
Quantitative real-time PCR (RT-qPCR)
The cells were seeded (1 × 105 cells/well) in 12-well plates and treated differently according to experimental assignments. We extracted total mRNA using Ultrapure RNA Kit (#CW0581, CWBIO, China) according to the protocol. The cDNA was synthesized using HiFiScript cDNA Synthesis kit (#CW2569, CWBIO, China). RT-qPCR quantification was performed using UltraSYBR Mixture (#CW0957, CWBIO, China) and the relative mRNA expression levels were normalized to the GAPDH expression levels.
Statistical analysis
All data are represented as the mean ± standard deviation (SD). SPSS 22.0 (SPSS, Chicago) served for statistical analyses. Statistical differences were analyzed using two-tailed Student’s t test or one-way ANOVA followed by the Tukey’s post hoc analysis where appropriate. Nonparameter tests were used for the non-normally distributed data. P ≤ 0.05 was considered statistically significant as indicated in the figure legend. All experiments were performed at least three times independently.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data needed to evaluate the conclusions in the paper are present in the paper and/or the Supplementary Information/Source Data file. Additional data related to this paper are available from the authors upon request. Proteomics data has been deposited in the iProX repository with accession code IPX0012888000. Source data are provided with this paper.
References
Clarke, B. Normal bone anatomy and physiology. Clin. J. Am. Soc. Nephrol.3, S131–S139 (2008).
Zaidi, M. Skeletal remodeling in health and disease. Nat. Med. 13, 791–801 (2007).
Zaidi, M., Yuen, T., Sun, L. & Rosen, C. J. Regulation of Skeletal Homeostasis. Endocr. Rev. 39, 701–718 (2018).
Riddle, R. C. & Clemens, T. L. Bone cell bioenergetics and skeletal energy homeostasis. Physiol. Rev. 97, 667–698 (2017).
Kurihara, N., Chenu, C., Miller, M., Civin, C. & Roodman, G. D. Identification of committed mononuclear precursors for osteoclast-like cells formed in long term human marrow cultures. Endocrinology 126, 2733–2741 (1990).
Parfitt, A. M. The mechanism of coupling: a role for the vasculature. Bone 26, 319–323 (2000).
Arron, J. R. & Choi, Y. Bone versus immune system. Nature 408, 535–536 (2000).
van de Laar, L. et al. Yolk sac macrophages, fetal liver, and adult monocytes can colonize an empty niche and develop into functional tissue-resident macrophages. Immunity 44, 755–768 (2016).
Wang, Y. et al. Nonredundant roles of keratinocyte-derived IL-34 and neutrophil-derived CSF1 in Langerhans cell renewal in the steady state and during inflammation. Eur. J. Immunol. 46, 552–559 (2016).
Takayanagi, H. et al. Induction and activation of the transcription factor NFATc1 (NFAT2) integrate RANKL signaling in terminal differentiation of osteoclasts. Dev. Cell 3, 889–901 (2002).
Butovsky, O. et al. Identification of a unique TGF-β-dependent molecular and functional signature in microglia. Nat. Neurosci. 17, 131–143 (2014).
Bonnardel, J. et al. Stellate cells, hepatocytes, and endothelial cells imprint the kupffer cell identity on monocytes colonizing the liver macrophage niche. Immunity 51, 638–654.e639 (2019).
Xie, H. et al. PDGF-BB secreted by preosteoclasts induces angiogenesis during coupling with osteogenesis. Nat. Med. 20, 1270–1278 (2014).
Boyle, W. J., Simonet, W. S. & Lacey, D. L. Osteoclast differentiation and activation. Nature 423, 337–342 (2003).
Chang, M. K. et al. Osteal tissue macrophages are intercalated throughout human and mouse bone lining tissues and regulate osteoblast function in vitro and in vivo. J. Immunol. 181, 1232–1244 (2008).
Wang, J. et al. Osteal tissue macrophages are involved in endplate osteosclerosis through the OSM-STAT3/YAP1 signaling axis in modic changes. J. Immunol. 205, 968–980 (2020).
Van Tongel, A., Karelse, A., Berghs, B., Van Isacker, T. & De Wilde, L. Diagnostic value of active protraction and retraction for sternoclavicular joint pain. BMC Musculoskelet. Disord. 15, 421 (2014).
Yeung, K. C. et al. Raf kinase inhibitor protein interacts with NF-kappaB-inducing kinase and TAK1 and inhibits NF-kappaB activation. Mol. Cell. Biol. 21, 7207–7217 (2001).
Al-Mulla, F. et al. Raf kinase inhibitor protein RKIP enhances signaling by glycogen synthase kinase-3β. Cancer Res. 71, 1334–1343 (2011).
Yeung, K. et al. Suppression of Raf-1 kinase activity and MAP kinase signalling by RKIP. Nature 401, 173–177 (1999).
Lorenz, K., Schmid, E. & Deiss, K. RKIP: a governor of intracellular signaling. Crit. Rev. oncog. 19, 489–496 (2014).
Qin, Q. et al. The inhibitor effect of RKIP on inflammasome activation and inflammasome-dependent diseases. Cell. Mol. Immunol. 18, 992–1004 (2021).
Lai, R. et al. Raf kinase inhibitor protein preferentially promotes TLR3-triggered signaling and inflammation. J. Immunol.198, 4086–4095 (2017).
Gu, M. et al. RKIP and TBK1 form a positive feedback loop to promote type I interferon production in innate immunity. EMBO J. 35, 2553–2565 (2016).
Lin, W. et al. Raf kinase inhibitor protein mediates intestinal epithelial cell apoptosis and promotes IBDs in humans and mice. Gut 66, 597–610 (2017).
Lin, W. et al. Raf kinase inhibitor protein negatively regulates FcεRI-mediated mast cell activation and allergic response. Proc. Natl. Acad. Sci. USA 115, E9859–e9868 (2018).
Lin, W. et al. RKIP mediates autoimmune inflammation by positively regulating IL-17R signaling. EMBO Rep. 19, https://doi.org/10.15252/embr.201744951 (2018).
Ahmed, S. et al. Cryptic Rac-binding and p21(Cdc42Hs/Rac)-activated kinase phosphorylation sites of NADPH oxidase component p67(phox). J. Biol. Chem. 273, 15693–15701 (1998).
Touaitahuata, H., Blangy, A. & Vives, V. Modulation of osteoclast differentiation and bone resorption by Rho GTPases. Small GTPases 5, e28119 (2014).
Itzstein, C., Coxon, F. P. & Rogers, M. J. The regulation of osteoclast function and bone resorption by small GTPases. Small GTPases 2, 117–130 (2011).
Bonfim-Melo, A., Ferreira, ÉR. & Mortara, R. A. Rac1/WAVE2 and Cdc42/N-WASP participation in actin-dependent host cell invasion by extracellular amastigotes of trypanosoma cruzi. Front. Microbiol. 9, 360 (2018).
Amato, C. et al. WASP restricts active rac to maintain cells’ front-rear polarization. Curr. Biol. 29, 4169–4182.e4164 (2019).
Menotti, M. et al. Wiskott-Aldrich syndrome protein (WASP) is a tumor suppressor in T cell lymphoma. Nat. Med. 25, 130–140 (2019).
Cui, G., Liu, H. & Laugsand, J. B. Endothelial cells-directed angiogenesis in colorectal cancer: interleukin as the mediator and pharmacological target. Int. Immunopharmacol. 114, 109525 (2023).
Aggarwal, B. B., Gupta, S. C. & Kim, J. H. Historical perspectives on tumor necrosis factor and its superfamily: 25 years later, a golden journey. Blood 119, 651–665 (2012).
Zhang, M. et al. Ischemia-reperfusion injury: molecular mechanisms and therapeutic targets. Signal Transduct. Target. Ther. 9, 12 (2024).
Srivani, G. et al. HIF-1α and RKIP: a computational approach for pancreatic cancer therapy. Mol. Cell. Biochem. 472, 95–103 (2020).
Pugh, C. W. & Ratcliffe, P. J. Regulation of angiogenesis by hypoxia: role of the HIF system. Nat. Med. 9, 677–684 (2003).
Chen, W. et al. HIF-1α regulates bone homeostasis and angiogenesis, participating in the occurrence of bone metabolic diseases. Cells 11, https://doi.org/10.3390/cells11223552 (2022).
Li, C. et al. Loss of sphingosine kinase 2 promotes the expansion of hematopoietic stem cells by improving their metabolic fitness. Blood 140, 1686–1701 (2022).
Tsukasaki, M. & Takayanagi, H. Osteoimmunology: evolving concepts in bone-immune interactions in health and disease. Nat. Rev. Immunol. 19, 626–642 (2019).
Mosser, D. M. & Edwards, J. P. Exploring the full spectrum of macrophage activation. Nat. Rev. Immunol. 8, 958–969 (2008).
Hu, K., Shang, Z., Yang, X., Zhang, Y. & Cao, L. Macrophage polarization and the regulation of bone immunity in bone homeostasis. J. Inflamm. Res. 16, 3563–3580 (2023).
Chen, J. et al. Osx-Cre targets multiple cell types besides osteoblast lineage in postnatal mice. PloS one 9, e85161 (2014).
Huang, W. & Olsen, B. R. Skeletal defects in Osterix-Cre transgenic mice. Transgenic Res. 24, 167–172 (2015).
Couasnay, G., Madel, M. B., Lim, J., Lee, B. & Elefteriou, F. Sites of Cre-recombinase activity in mouse lines targeting skeletal cells. J. Bone Miner. Res.36, 1661–1679 (2021).
Zhou, Y. T., Chew, L. L., Lin, S. C. & Low, B. C. The BNIP-2 and Cdc42GAP homology (BCH) domain of p50RhoGAP/Cdc42GAP sequesters RhoA from inactivation by the adjacent GTPase-activating protein domain. Mol. Biol. Cell 21, 3232–3246 (2010).
Fidyk, N., Wang, J. B. & Cerione, R. A. Influencing cellular transformation by modulating the rates of GTP hydrolysis by Cdc42. Biochemistry 45, 7750–7762 (2006).
Nassar, N., Hoffman, G. R., Manor, D., Clardy, J. C. & Cerione, R. A. Structures of Cdc42 bound to the active and catalytically compromised forms of Cdc42GAP. Nat. Struct. Biol. 5, 1047–1052 (1998).
Jaakkola, P. et al. Targeting of HIF-alpha to the von Hippel-Lindau ubiquitylation complex by O2-regulated prolyl hydroxylation. Science 292, 468–472 (2001).
Maxwell, P. H. et al. The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature 399, 271–275 (1999).
Zhang, D. et al. Hypoxia promotes osteosarcoma cell proliferation and migration through enhancing platelet-derived growth factor-BB/platelet-derived growth factor receptor-β axis. Biochem. Biophys. Res. Commun. 512, 360–366 (2019).
Deppe, J. et al. Impairment of hypoxia-induced HIF-1α signaling in keratinocytes and fibroblasts by sulfur mustard is counteracted by a selective PHD-2 inhibitor. Arch. Toxicol. 90, 1141–1150 (2016).
Argueta, C. E. et al. RKIP localizes to the nucleus through a bipartite nuclear localization signal and interaction with importin α to regulate mitotic progression. J. Biol. Chem. 299, 103023 (2023).
Zheng, Z. et al. Site-1 protease controls osteoclastogenesis by mediating LC3 transcription. Cell Death Differ. 28, 2001–2018 (2021).
Acknowledgements
We thank Xiaoli Hong from the Core Facilities, Zhejiang University School of Medicine for her technical support. This work was supported by the National Natural Science Foundation of China (82272520, 82272521, 82472472, 82302732); Zhejiang Provincial Natural Science Foundation of China (LQ23H060006, LZ23H060002, LQ23H070004).
Author information
Authors and Affiliations
Contributions
Z.Z., S.T., J.J., Y.L., L.D., and Y.Z. performed the experiments, analyzed the data and prepared the manuscript. Z.H., B.H., Sai, W.L., and X.W. conceived the project and revised the manuscript. M.W., J.C., and F.Z. were responsible for experimental design, project supervision, and manuscript refinement. All authors discussed the results and contributed to the preparation of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Hiroshi Takayanagi, Haibo Zhao, and Gabriel Mbalaviele for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zheng, Z., Tao, S., Jin, J. et al. RKIP regulates bone marrow macrophage differentiation to mediate osteoclastogenesis and H-type vessel formation. Nat Commun 16, 7604 (2025). https://doi.org/10.1038/s41467-025-62972-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-62972-8