Abstract
Severe fever with thrombocytopenia syndrome virus (SFTSV) is a representative high-pathogenic bandavirus (Bandavirus genus, Phenuiviridae family). Inducible expression of interferon-stimulated genes (ISGs) is the foundation of host antiviral defense; however, their roles in bandavirus infection remain elusive. Here, we identify over 200 ISGs potentially inhibiting or promoting bandaviral replication. With SFTSV as the main model, we further systematically uncover the notable antiviral role of one ISG, cyclin D3 (CCND3), against bandaviruses. SFTSV infection induces CCND3 up-regulation and cytoplasmic translocation. CCND3, in turn, inhibits the viral replication in cultured cells and pathogenicity in vivo. The viral nucleoprotein (NP) is the target of CCND3. By its CN domain, CCND3 interacts with NP’s “head” region in an RNA-independent manner, suppressing the ribonucleoprotein (RNP) replication machinery activity. Furthermore, consistent with interaction interface mapping and structural modeling analyses, the CCND3-NP interaction blocks NP multimerization, NP-RNA binding, and NP association with viral polymerase, that is, the NP activities essential to RNP construction and functioning. Conversely, the viral nonstructural protein, NSs, can partially antagonize CCND3 by attenuating its induction and promoting autophagic degradation. These findings provide new insights into bandavirus-host interactions and arms race, advancing the understanding of bandavirus infection and probably informing antiviral therapeutic development.
Similar content being viewed by others
Introduction
Severe fever with thrombocytopenia syndrome virus (SFTSV), also referred to as Dabie bandavirus (DBV), is the causative agent of severe fever with thrombocytopenia syndrome (SFTS), an emerging and life-threatening infectious disease1,2. SFTS is characterized by clinical manifestations such as fever, severe thrombocytopenia, leukopenia, and gastrointestinal symptoms, with a high case-fatality rate of up to 30%1,2. The disease is transmittable through tick bites and human-to-human contact, with its epidemic areas expanding in China and neighboring countries3,4,5,6. Following the identification of SFTSV in China in 2009, several other tick-borne viruses genetically related to SFTSV, such as Heartland virus (HRTV, identified in the United States) and Guertu virus (GTV, discovered in China), were successively isolated around the world7,8. Recently, they have been classified into a new virus genus Bandavirus (Phenuiviridae family)9. There is no licensed antiviral drug or vaccine currently available against them. These bandaviruses, with SFTSV as the highly pathogenic representative, pose a substantial threat to public health, necessitating urgent research.
Bandaviruses are enveloped RNA viruses with a negative-sense, single-stranded RNA genome consisting of three segments. The large (L) and medium (M) genomic segments respectively encode the viral RNA-dependent RNA polymerase (RdRp, i.e., L protein) and envelope glycoproteins (GP), while the small (S) segment encodes the nucleoprotein (NP) and a nonstructural protein (NSs) in an ambisense manner1. NP is a major structural protein component of the virions10,11,12. Through multimerization and complex interactions with viral RNAs and L protein, NP plays crucial roles in virus replication by driving the assembly and participating in the function of the viral ribonucleoprotein complex (RNP)11,12,13,14. RNP is the molecular machinery for transcription and replication11,12,13. Moreover, RNPs encapsulating viral genomic RNAs also can be packaged into progeny virions as the structural core11. In comparison, as a nonstructural protein, bandavirus NSs is not essential to the viral replication but act as an important virulence factor by interfering with multiple host biological processes and in particular, antiviral interferon (IFN) responses15,16,17,18,19,20,21.
Type I IFN response constitutes the first vital line of host defense against various viral infection and pathogenicity by inducing IFN-stimulated gene (ISG) expression22,23. This host antiviral response is initiated by the recognition of viral infections by pattern recognition receptors (PRRs)24,25. During bandavirus infection, retinoic acid-inducible gene I (RIG-I)-like receptors (RLRs) and several Toll-like receptors (TLRs) have proved important for the host recognition in previous studies by us and others20,26. Upon recognition, PRRs activate the downstream signaling cascades for IFN induction, leading to the expression of antiviral IFNs, especially type I IFNs24,27. The produced IFNs then bind to their cell surface receptors and trigger the Janus kinase-signal transducer and activator of transcription (JAK-STAT) pathway, inducing expression of more than 300 ISGs28,29. The induction of these ISGs establishes a strong host antiviral state, although the action mechanisms of most ISGs remain elusive30,31. Many studies have validated that type I IFN system is also critical to restrict bandavirus replication and pathogenicity15,18,26,32,33,34,35,36,37. However, it remains poorly understood which and how ISGs can regulate bandaviruses as IFN effectors.
Here, we analyzed the regulatory potential of ISGs in SFTSV replication by using an ISG cDNA expression library and a previously established minireplicon reporter system38,39,40. Numerous ISGs that likely inhibit or promote the viral replication machinery RNP-associated stage were thus identified, presenting plenty of new clues for further elucidation of the virus-host interactions. Subsequently, we systematically validated the remarkable role of a representative ISG, Cyclin D3 (CCND3), in restricting bandavirus infection and pathogenicity in vitro and in vivo, by using series of loss/gain-of-function assays and SFTSV as the main virus model. Furthermore, we uncovered the detailed molecular mechanism underlying CCND3 restriction of SFTSV, proposing a new antiviral mode employed by CCND3. Additionally, potential antagonizing effects of SFTSV NSs on CCND3 were also analyzed. The findings provide insights into the bandavirus-host interactions, better understanding of which may help advance the development of antiviral therapy against SFTSV and related bandaviruses.
Results
A minireplicon reporter system-based screen for ISGs regulating SFTSV replication
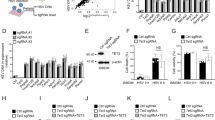
Antiviral IFN response is a pivotal host defense mechanism against bandaviral infection32,33,34,35. However, the specific antiviral potential of ISGs as the IFN effectors against bandaviruses including SFTSV remains to be explored. Therefore, we screened individual ISGs for their ability to regulate SFTSV replication using a cDNA clone library of different ISGs combined with an SFTSV minireplicon reporter system (Fig. 1a). As previously described38,39,40, the minireplicon reporter system constructing viral RNP machinery can indicate the efficiency of the central events in virus life cycle, including RNP assembly and function on driving transcription and replication. Besides, the system facilitated high-throughput screening without the requirements of high-level biosafety facilitates. As shown in the dot plot of mean values from relative reporter activities (Fig. 1b), expression of most ISGs likely inhibited the viral replicon to some extent (averaging ~30% reduction, as indicated by the gray horizontal line), yet a small subset seemed to enhance it. Consistently, MOV10 and MxA that have been reported to inhibit SFTSV infection39,40, were also identified in this screen (Fig. 1b). Furthermore, volcano plot analysis reveals that 234 and 18 ISGs, respectively, inhibit or promote the replicon activity by more than 20%, with adjusted p values of less than 0.05 (Fig. 1c and Supplementary Data 1). Therein, CCND3 was recognized as one of the top candidates with strong inhibitory activities and high statistical significance (Fig. 1b, c). CCND3 is known as a classical protein involved in regulation of cell cycle41,42, whereas its role as an ISG in response to virus infection remains to be further explored. Moreover, there was no report regarding its involvement in bandavirus infection before this study. Deciphering the mechanisms of understudied ISGs like CCND3 in viral infection is imperative for understanding host-virus interplays. We thus selected CCND3 as a candidate for following in-depth function-mechanism analyses.
a Schematic representation of the ISG screen. Individual ISG expression plasmids were co-transfected with the SFTSV minireplicon system plasmids, including an EGFP-based minigenome transcription plasmid and the vectors expressing SFTSV NP and L. After 48 h, cells were fixed and stained with Hoechst 33258, followed by counts of EGFP-positive cells and total cells (nuclei) using Operetta CLSTM high-throughput system. In the screen, three biological replicates for each ISG were conducted to evaluate their potential effects on the replicon, allowing the following statistical significance and volcano plot analyses. b Dot plot of relative SFTSV minireplicon activities. Relative replicon activities were calculated by normalizing the EGFP-positive cell ratio of ISG overexpressing groups to that of the control group co-transfected with the empty vector. Mean values of the relative replicon activities (n = 3) were used in the dot plot. The gray line indicates the ISG population mean. c Volcano diagram of the ISG effects. Points with replicon activity change > 20% and p < 0.05 are highlighted in blue or pink as significant antiviral or pro-viral candidates, respectively. CCND3, as well as the previously reported MOV10 and MxA, is marked with additional circles. See also Supplementary data 1. Comparisons were performed with two-tailed Student’s t-test, followed by p-value adjustments. Source data with detailed p-values are provided as a Source Data file. Elements in (a) were created with BioRender (Xu, Z. (2025) https://BioRender.com/bong7yn).
CCND3 expression and cytoplasmic translocation are enhanced upon SFTSV infection
To better characterize the potential interplays between CCND3 and SFTSV, we first assessed CCND3 expression levels and subcellular localization upon SFTSV infection. Several cell lines previously established as permissive models for SFTSV research were infected with the virus and subjected to analysis of CCND3 expression at different time points. CCND3 mRNA expression was evidently stimulated in all the tested cell types upon SFTSV infection (Fig. 2a–c). Similarly, transcriptional upregulation of CCND3 by SFTSV infection was also observed in isolated primary cells including human peripheral blood mononuclear cells (PBMCs) and mouse bone marrow-derived macrophages (BMDMs) (Fig. 2d, e). Subsequently, the effects of SFTSV infection on CCND3 mRNA transcription were further tested in mice in vivo. Consistently, transient induction of CCND3 expression was detected upon the viral infection in the mouse tissues (Supplementary Fig. S1).
a–e HEK293, THP-1, and MEF cells, as well as primary human PBMCs and mouse BMDMs, were infected with SFTSV (MOI = 5). CCND3 mRNA and SFTSV M RNA levels were subsequently analyzed by qPCR at the indicated time points postinfection. f Localization of endogenous CCND3 upon SFTSV infection. HEK293 cells infected with SFTSV (MOI, 0.5) were fixed at 24 h postinfection (hpi) and subjected to immunofluorescence assays (IFA) and confocal microscopy to visualize cellular CCND3 (red), SFTSV NP (green), and nuclei (blue). Scale bar, 20 μm. g–i Nuclear-cytoplasmic fractionation. THP-1 cells were infected with SFTSV at various MOIs for 24 h. Nuclear and cytoplasmic fractions were separated and protein levels of CCND3 and SFTSV NP were analyzed by Western blot (WB) using the indicated antibodies. Total protein levels from whole cell lysates (WCL) were shown in (g). HDAC1 and β-actin were served as nuclear and cytoplasmic markers and loading controls, respectively (g and h). Relative band intensities of cytoplasmic and nuclear CCND3 (over the corresponding loading controls) were quantified and normalized to the mock groups (i). Results are representative of three independent replicates (f and g). Data are presented as means ± SD, n = 3 biological replicates (a–e and h–i). Source data are provided as a Source Data file.
Then, cell immunofluorescence assays (IFA) showed that CCND3 was predominantly localized in nuclei in the mock-infected group, while SFTSV infection increased the abundance of CCND3 in the cytoplasm at a low multiplicity of infection (MOI, 0.5) (Fig. 2f). Moreover, an overall increase of CCND3 signals could be more evidently observed in both cytoplasm and nuclei at a higher infection dose (MOI, 2) (Supplementary Fig. S2). To further investigate the effect, we analyzed the subcellular localization of CCND3 by nuclear-cytoplasmic fractionation. Total protein analyses with whole cell lysates (WCL) confirmed that CCND3 expression was dose-dependently stimulated by SFTSV infection (Fig. 2g). Consistently, CCND3 was mainly nuclear in the resting state (mock-infected group), but evidently accumulated in the cytoplasm following SFTSV infection (Fig. 2h, i). Slight increase of CCND3 abundance in the nuclei could also be observed with SFTSV infection at high MOIs, but to a much lesser extent compared with that in the cytoplasm (Fig. 2h, i). Given that SFTSV is a cytoplasmic RNA virus replicating exclusively in cytoplasm, the enhancement of CCND3 expression and cytoplasmic localization could be closely associated with the potential regulatory role of the host protein in SFTSV replication.
CCND3 acts as a notable host restriction factor against SFTSV and related bandaviruses
To further confirm the role of CCND3 in modulating SFTSV replication, we next conducted series of loss/gain-of-function assays to evaluate the impact of CCND3 on SFTSV infection. First, CCND3 overexpression indeed significantly inhibited SFTSV replication and propagation (Fig. 3a–e). Conversely, knockdown (KD) of CCND3 by RNAi evidently augmented SFTSV replication and progeny production (Fig. 3f–k). Furthermore, CCND3 knockout (KO) cells were generated by the CRISPR-Cas9 system (Supplementary Fig. S3a) and used for the following analyses of SFTSV replication kinetics. Consistently, CCND3 deletion resulted in marked increase in SFTSV RNA replication, protein expression, and progeny propagation (Fig. 3l–p), corroborating the notable anti-SFTSV role of CCND3.
a–e CCND3 overexpression inhibits SFTSV infection. HEK293 cells were transfected with the CCND3 expression plasmid or control vector. At 24 h post transfection, the cells were infected with SFTSV (MOI = 0.1) and delivered to analyses of the RNA and protein levels at different time points by qPCR (a–c) and WB (d), respectively. The propagation of progeny released into culture medium (growth curve) was also analyzed by TCID50 assay (e). f–k CCND3 knockdown (KD) by RNAi promotes SFTSV infection. HEK293 cells were transfected with the CCND3-targeting or control siRNAs and infected with SFTSV (MOI = 0.1) for 36 h, followed by determination of the RNA levels (f–i), protein expression (j), and viral progeny titers (k), respectively. l–p CCND3 knockout (KO) by CRISPR-Cas9 editing enhances SFTSV infection. CCND3 KO and control cells were infected with SFTSV (MOI = 0.1), followed by analyses of RNA (l–n) and protein (o) levels and progeny propagation (p) at different timepoints. WB results are representative of three independent replicates (d, j, o). Data are shown as means ± SD, n = 3 biological replicates (a–c, e–i, k–n, and p). Statistically significant differences are indicated (two-tailed Student’s t test for each timepoint in a–c, e, l–n, and p; one-way ANOVA in f–i and k). *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; ns non-significant. Source data are provided as a Source Data file.
In addition, the influence of CCND3 on replication of GTV and HRTV, two other bandaviruses genetically related to SFTSV, was also investigated using aforementioned experimental settings. Similarly, CCND3 overexpression dramatically repressed all the three segment-derived RNA replication and progeny propagation of both GTV (Fig. 4a, b) and HRTV (Fig. 4c, d) at different time points. In contrast, loss-of-function assays using the CCND3-KO cells demonstrated that absence of CCND3 led to significant enhancement of both GTV (Fig. 4e, f) and HRTV replication and propagation (Fig. 4g, h). Collectively, these data reveal that CCND3 serves as a conserved host restriction factor against SFTSV and related bandaviruses, meriting further detailed function/mechanism elucidation.
a–d Inhibition of GTV and HRTV infection by CCND3 overexpression. HEK293 cells transfected with the CCND3 expression plasmid or control vector were infected with GTV (a, b) or HRTV (c, d) at 0.1 MOI. At the indicated time points postinfection, the cells and culture media were respectively delivered to analyses of the viral RNA accumulation (a, c) and progeny propagation (b, d). e–h Enhancement of GTV and HRTV infection by CCND3 KO. The CCND3 KO and control cells were infected with GTV or HRTV (MOI = 0.1), followed by the analyses of intracellular viral RNA levels (e, g) and progeny propagation (f, h) at the indicated time points. Data shown are means ± SD, n = 3 biological replicates. Two-tailed t test for the individual timepoints: *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; ns non-significant. Source data are provided as a Source Data file.
CCND3 deficiency increases the susceptibility of mice to SFTSV infection
CCND3, which is critically required for pre-TCR-driven expansion of immature thymocytes, plays an essential role in the normal development of the thymus and T cells in mice43,44,45. To investigate the effect of CCND3 on SFTSV infection and pathogenicity in vivo, we generated a CCND3-deficient adult mouse model by transient transduction of a lentiviral vector expressing CCND3-specific shRNA via intravenous injection, as previously conducted39,46,47. Although adult immunocompetent mice infected with SFTSV do not exhibit obvious clinical symptoms, SFTSV replication and transient tissue lesions can be detected in various organs33,48. Therefore, the knockdown mouse model has been effectively utilized to study the role of specific host factors that may regulate virus replication39,46,47. Consistent with previous reports, no severe clinical manifestations or deaths were observed in all infected mice (n = 6 per group). Thus, viral replication and pathogenicity in various tissues were further analyzed after sacrifice. Firstly, qPCR analysis confirmed a significant decrease of CCND3 expression in CCND3-shRNA treated mice (Fig. 5a). Then, viral RNA levels in the analyzed tissues including spleen, liver, and lung were all significantly higher in the CCND3-KD mice than those in control (Fig. 5b), suggesting the restriction of viral replication by CCND3. Consistently, the CCND3-deficient mice exhibited more severe viremia with higher viral copies in the sera (Fig. 5c). In line with these data, immunohistochemical analysis (IHC) detected more viral antigen-positive foci in the tissues of the CCND3-KD group (Supplementary Fig. S4), further supporting the restrictive effect of CCND3 on SFTSV infection. Thrombocytopenia, leukopenia, and elevated serum biochemical indicators of organ damage are the typical clinical features of SFTSV infection in humans1,2. Interestingly, a significant reduction of platelet counts was observed in the CCND3-KD group but not the control group upon SFTSV infection (Fig. 5d). Moreover, reduction of white blood cells (WBC) counts and increase of alanine aminotransferase (ALT), aspartate aminotransferase (AST), and blood urea nitrogen (UREA) levels caused by the viral infection were more severe after CCND3 depletion (Fig. 5d, e), indicating CCND3 restriction of the viral pathogenicity. Additionally, we examined the pathology of infected tissues through H&E staining. Compared to the control, CCND3-deficient mice infected with SFTSV indeed showed more pronounced histopathological abnormalities (Fig. 5f–h). These mainly included splenic nodules and blurring boundaries between the red pulp and white pulp, local coagulative necrosis of the liver tissues characterized with disappearance of nuclei and visible cytoplasmic fragmentation, and widening of alveolar septa accompanied by interstitial infiltration in the lung tissues (Fig. 5f–h). Together, these results support the notable role of CCND3 in restricting the viral replication and pathogenicity in vivo.
C57BL/6J mice (n = 6/group) were transduced with the viral vectors encoding control (shNC) or CCND3-targeting shRNAs via caudal vein. At 7 d post transduction, the animals were infected with SFTSV (105 TCID50) or mock infected with PBS control via intraperitoneal inoculation. Three days postinfection, the mice were sacrificed for following analyses. CCND3 mRNA levels in the indicated tissues were analyzed by qPCR (a). SFTSV RNA levels in the lung, spleen and liver (b) and serum viral copies (c) were analyzed in SFTSV-infected shNC control and CCND3-deficient mice. Platelet (PLT) and white blood cell (WBC) counts of whole blood (d) and serum ALT, AST, and UREA (e) were also determined as described in Methods. Representative H&E staining of liver (f), spleen (g), and lung (h) sections and cumulative pathological score were respectively shown. Noticeable pathological changes in the tissues are indicated by colored arrows. Liver: blue indicates hepatocellular necrosis, nuclear disappearance, karyomegaly, and nuclear fragmentation; black represents changes and necrosis in hepatic sinusoids; yellow indicates infiltration of inflammatory cells in the liver. Spleen: red indicates indistinct boundaries between the red pulp and white pulp; black denotes dispersed splenic nodules with decreased cellularity of lymphocytes. Lungs: red indicates proliferation of alveolar epithelial cells, alveolar atrophy, and thickening of the alveolar septum; green signifies infiltration of inflammatory cells within the lungs. Scale bar, 100 μm. Data are means ± SD, n = 6 mice. Two-tailed Student’s t test (a–c and f–h) and one-way ANOVA (d and e): *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; ns, non-significant. Source data are provided as a Source Data file.
Unlike immunocompetent adult mice, IFN receptor-deficient mice (e.g., A129) can serve as a fully lethal animal model for SFTSV infection. We also utilized this model to complementally analyze the effect of CCND3 deficiency on viral infection and pathogenesis. Interestingly, in agreement with its anti-SFTSV role, CCND3 KD appeared to worsen SFTSV-induced lethality and weight loss (Supplementary Fig. S5a–b). Consistently, CCND3 KD resulted in higher SFTSV viremia and replication levels across tissues, intensified thrombocytopenia and leukopenia, and elevated biochemical markers of tissue injury (Supplementary Fig. S5c–g). Furthermore, histopathological analysis revealed aggravated tissue damage in CCND3-KD mice (Supplementary Fig. S5h–j). These findings further corroborate the restrictive role of CCND3 in SFTSV infection and pathogenesis.
The antiviral activity of CCND3 is independent of its cell cycle regulatory function and the IFN signaling pathway
Next, we proceeded to methodically elucidate the molecular mechanisms by which CCND3 restricts bandavirus replication. Before investigating if CCND3 directly interferes with viral replication by targeting viral molecular machinery, we first analyzed whether its potential cell cycle regulatory role may indirectly contribute to its antiviral activity. Mutation at the CCND3 T283 can impair its phosphorylation and normal physiological role in cell cycle49. Interestingly, both CCND3 and its T283A mutant dramatically inhibited SFTSV infection and no significant difference was observed (Supplementary Fig. S6a–c). CCND3 participates in cell cycle regulation through the CCND3/CDK4/6 pathway42,50. Therefore, we then constructed CDK4-KO, CDK6-KO, and CDK4/6 double-KO cells and examined the CCND3 antiviral activity in these models. Consistently, deletion of CDK4, CDK6, or both did not impair CCND3’s anti-SFTSV function (Supplementary Fig. S6d–f), demonstrating that the antiviral activity is independent of cycle regulation. Moreover, KO of these CDKs did not lead to any enhancement of the viral replication (Supplementary Fig. S6d–f), different with CCND3 KO (Fig. 3l–p), also supporting the irrelevance of CCND3’s anti-SFTSV activity to the cycle regulation pathways.
Furthermore, we generated IFN receptor-KO cells to test whether the IFN antiviral signaling is directly required for CCND3 inhibition of viral replication. Interestingly, CCND3, as well as the T283A mutant, continued to exert significant inhibitory effects on SFTSV replication in the cells with IFN-signaling KO (Supplementary Fig. S6g–i). This indicates that the antiviral action of CCND3 is also independent of IFN signaling, although its expression can be induced by the signaling as an ISG. Together, these analyses suggest that CCND3 can exert antiviral activity independently of its cell cycle regulatory function and IFN signaling, implying that CCND3 may directly block viral replication by targeting critical viral machinery.
CCND3 targets the viral NP protein to interfere with the RNP machinery
To further uncover the mechanism underlying the antiviral function of CCND3, we identified the potential viral protein targeted by CCND3. HEK293 cells were infected with SFTSV and subsequently subjected to co-immunoprecipitation (Co-IP) assays. Interestingly, NP, but not the other structural proteins, was specifically co-precipitated by CCND3 (Fig. 6a). Additionally, NSs also seemed slightly precipitated but to a much lesser extent. Since NP, not NSs, is the major structural component and plays an essential role in the viral replication, we then primarily focused on characterizing the CCND3-NP interaction. First, the interaction of CCND3 with NP was also validated in the following Flag-pulldown analysis using co-transfected cells (Fig. 6b). Then, we tested whether RNA is involved in the interaction, considering the RNA-binding capacity of NP. Results from the further pulldown assays combined with nuclease treatment39,40 showed that the interaction between CCND3 and NP was likely independent of RNA (Fig. 6c). NP is the core component driving the assembly and participating the function of the replication machinery RNP. Therefore, the targeting of NP by CCND3 is also well linked to CCND3’s inhibitory ability to the minireplicon system indicating the RNP activity as observed in the screening. To further corroborate the interference of CCND3 with the viral RNP activity, we conducted additional evaluation on dose-dependent effects of CCND3 using two minigenome reporter systems based on EGFP or dual-luciferases, respectively. Consistently, CCND3 efficiently inhibited both the SFTSV EGFP (Fig. 6d–f) and dual-luciferase based (Fig. 6g) minireplicons in dose-dependent manners, confirming the blockade of RNP activity by CCND3. These data uncover the direct antiviral action of CCND3 by targeting NP to interfere with viral RNP activity.
a HEK293 cells mock-infected or infected with SFTSV were harvested at 24 hpi for co-immunoprecipitation (Co-IP) and WB analysis of the WCL input and IP products. b HEK293T cells were co-transfected with the Flag-CCND3 and SFTSV-NP expression plasmids or control vector, followed by Flag-pulldown and WB analysis. c Cells were treated as in (b), but the cell lysates were first subjected to UltraNuclease treatment or left untreated, followed by Flag-pulldown and WB analysis, similarly. d–f The dose-dependent inhibitory effect of CCND3 on SFTSV EGFP-based minigenome RNP reporter system. BHK-21 cells were co-transfected with the SFTSV L and NP expression plasmids and the minigenome reporter plasmid (MUTR-EGFP), along with various amounts of the CCND3 expression plasmid for 48 h, followed by fixation and high-content imaging (d) and counting (e). EGFP-positive cell ratios were normalized to the control without CCND3 overexpression (e). In a parallel experiment, the samples were also subjected to WB analysis of the EGFP expression levels (f). g The dose-dependent inhibitory effect of CCND3 on SFTSV firefly luciferase (Fluc)-based minigenome RNP reporter. The SFTSV NP and L expression plasmids and Fluc-based SFTSV minigenome reporter plasmid (MUTR-LUC) were co-transfected together with indicated amounts of the CCND3 expression plasmid and pRL-TK expressing Renilla luciferase (Rluc). Relative luciferase activities (Fluc/Rluc) were calculated and shown. Data in (a–d and f) are representative from three independent replicates with similar results. Data in (e and g) are means ± SD, n = 3 biological replicates. One-way ANOVA (e and g): *p < 0.05, ****p < 0.0001. Source data are provided as a Source Data file.
CCND3 targets the N- and C-lobes of NP to inhibit the RNP activity and hence virus replication via its CN domain
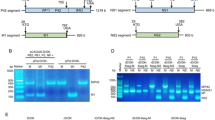
To gain more mechanistic insights on how CCND3 targets NP to interfere with SFTSV replication, we further investigated the critical domains and potential interface involved in CCND3-NP interaction. Previous studies have elucidated the structure of NP, in which the protein is composed of a protruding N-arm and a compact “head” comprising an N-lobe and a C-lobe10,51. The conformation of CCND3 residues 23–254 was also defined in a crystal structure50. Interestingly, structural modeling by the deep learning-based AlphaFold2 program showed that CCND3 and NP potentially form a complex of high confidence (Fig. 7a). The CCND3 N (CN) domain and NP “head” involving both the N- and C-lobes seemed to play major roles in the interaction by constructing a potential interaction interface in the AlphaFold complex structure (Fig. 7a, b). The interaction domains were then experimentally examined by protein interaction analyses using a series of NP and CCND3 truncated mutants (Fig. 7c). As demonstrated in the S.tag pulldown (S-pulldown) results with S.tag-fused NP as the bait, co-precipitation with NP was deprived by the deletions of CCND3 Box (CBox) or even the smaller CN, but not CCND3 C (CC), suggesting that the CN domain is indeed involved in CCND3-NP interaction (Fig. 7d). Additionally, EGFP-NanoTrap assays with EGFP-fused NP mutants indicated that both the NP N- and C-lobes, but not N-arm, could interact with CCND3 (Fig. 7e), also in agreement with the structural modeling analysis. These data suggest that CCND3 likely targets both the N- and C-lobes of NP mainly via its CN domain.
a, b Structural modeling of the CCND3-NP interaction by AlphaFold2. AlphaFold2 modeling reveals a CCND3-NP complex (a) and its interaction interface (a and b, marked in red). The structural domains of CCND3 and SFTSV NP are indicated by different colors. CN, CCND3 N domain; CC, CCND3 C domain. c Schematic diagram of CCND3 and NP truncations used below. CBox, CCND3 Box domain. d Deletion of the CN or CBox, but not CC, disrupts CCND3 targeting of NP. HEK293T cells were co-transfected with the plasmids expressing Flag-tagged CCND3 or mutants and the S-tagged NP expression plasmid, or control vectors (indicated by Vector or “-”), followed by S-pulldown and WB analyses. e Both the N- and C-lobes, but not N-arm, of NP can be targeted by CCND3. Plasmids encoding EGFP or EGFP-tagged NP mutants and CCND3 were co-transfected into HEK293T cells, followed by EGFP-NanoTrap assays and WB analysis. EGFP-fused bait protein bands are indicated by arrowheads. f–h CN is required for CCND3 inhibition on SFTSV RNP activity. Effects of CCND3 or mutant expression on SFTSV EGFP-based minigenome reporter were evaluated by imaging (f), calculation of relative reporter activities (g), and WB (h). i–k CN (but not CC) is essential to CCND3 restriction of SFTSV replication and propagation. Effects of the indicated CCND3 mutants on SFSTV infection were analyzed at 24 hpi by qPCR (i), WB (j), and TCID50 assays (k), respectively. l–p CN expression alone can target NP and inhibit SFTSV infection. Targeting of NP by individual CCND3 domains tagged with EGFP (l) was analyzed by S-pulldown (m). The anti-SFTSV activities were tested by qPCR (n), WB (o), and progeny titration (p). Data in (d–f, h, j, m and o) are representative from three independent replicates with similar results. Data in (g, i, k, n, and p) are presented as means ± SD, n = 3 biological replicates. One-way ANOVA: ****p < 0.0001; ns, non-significant. Source data are provided as a Source Data file.
Next, we tested the effects of the CCND3 mutants on SFTSV minireplicon reporter system. Consistently, removal of the CN or CBox domains abolished the inhibition to SFTSV RNP-driven reporter activity (Fig. 7f–h). Subsequently, influence of the mutation on SFTSV replication was also examined: deletion of the CN or CBox (but not CC) resulted in abolishment of CCND3’s anti-SFTSV activity (Fig. 7i–k). Furthermore, individual expression of CN, and the slightly larger CBox containing CN, could also target NP by protein interaction (Fig. 7l, m). In line with its NP-targeting ability, CN alone exhibited significant antiviral activities, effectively suppressing viral RNA replication, protein expression, and progeny production (Fig. 7n–p). In contrast, the CC domain neither interacted with NP nor displayed the antiviral effects (Fig. 7l–p). Altogether, these consistent data demonstrate that CCND3 targets the N- and C-lobes of NP to interfere with the RNP activity and viral replication by its CN domain.
Following the identification of CN, we further analyzed potential significant motifs or amino acid residues within CN. First, we performed sequential scanning mutagenesis by introducing alanine substitutions in a series of five-residue segments covering the CN domain, followed by functional screening using the NP/RNP reporter system (enabling high-throughput functional assessment). Interestingly, the first four 5-amino acid motifs at the N-terminal region of CN (i.e., residues 62–81) were shown to be important for the suppression of the NP/RNP activity (Supplementary Fig. S7a, b). Subsequently, we conducted single-residue scanning mutagenesis across these 20 amino acids (prioritizing polarity-altering substitutions to maximize phenotypic effects for rapid identification of critical residues). Reporter assays demonstrated that mutations at multiple residues within this region seemed to affect CCND3’s ability to suppress viral NP/RNP activity, with five residues (M64, C68, E70, R72, and P79) exhibiting particularly significant functional importance (Supplementary Fig. S7c, d). Moreover, the structural modeling indeed localized these residues to or adjacent to the predicted interaction interface (Supplementary Fig. S7e). Consistently, protein interaction analysis confirmed that mutations at these sites impaired CCND3 targeting of NP (Supplementary Fig. S7f), correlating with significantly reduced antiviral activities (Supplementary Fig. S7g–i). These findings refine the understanding of CN’s pivotal role in CCND3’s antiviral function and provide residue-level mechanistic insights into the protein targeting.
By the CN domain, CCND3 hinders the NP activities essential for RNP construction and function, including NP multimerization, NP-RNA binding, and NP-L interaction
NP multimerization, NP-RNA binding, and NP-L interactions are critical activities of NP required for viral RNP construction and function11,12,14. Based on characteristics of CCND3-NP interaction and the structural modeling, we considered that CCND3 binding to NP probably obstructs these NP activities (as analyzed below). Therefore, in order to further mechanistically address CCND3 inhibition on the viral RNP by targeting NP, influence of CCND3 on NP multimerization that drives RNP formation was first evaluated by in situ chemical crosslinking with disuccinimidyl suberate (DSS)39,40,52. CCND3 and its T283A mutant that retains the antiviral capability indeed both could significantly hamper NP multimerization (Fig. 8a). However, the CCND3 mutants with CN or CBox deleted lost the ability to interfere with NP multimerization (Fig. 8b), in agreement with the important role of CN in targeting NP and inhibiting the viral replication. According to these experimental data, binding of CCND3 to NP likely leads to a significant steric hindrance obstructing NP multimerization. Interestingly, this is consistent with the structural model, in which the CCND3-bound interface on NP is on the side that would face inward in the supposed NP oligomers, that is, CCND3 binding would sterically hinder the formation of NP oligomers (Fig. 8c).
a CCND3 as well as its T283A mutant inhibits NP multimerization. HEK293T cells were transfected with the plasmids encoding SFTSV NP and Flag-tagged CCND3 or T283A mutant, followed by DSS cross-linking at 24 h post transfection and WB analysis of NP monomeric and oligomers. Gray scale of the monomer or polymer bands in the DSS-treated samples was quantified using ImageJ and normalized to the control (vector) group. b Effects of CCND3 truncations on the inhibition of NP multimerization. c Structural modeling reveals a consistent result in which CCND3 interaction with NP obstructs NP multimerization with the adjacent protomers. NP and CCND3 in the interaction complex were colored in red and gray, respectively. Two adjacent NP protomers that are supposed to be assembled into multimers are indicated in tint colors. d Effects of CCND3 and its mutants on NP-RNA binding. HEK293T cells were transfected with the plasmids encoding Flag-tagged CCND3 (or mutants) and SFTSV NP. At 24 h post transfection, the cell lysates were subjected to RNA-pulldown with beads coupled with S RNA, followed by WB analysis. The intensity of NP bands in the RNA-pulldown products was quantified and normalized to the control. e CCND3 (especially CN) binding to NP seals the CavityR of NP, site for RNA binding, in the CCND3-NP complex structure from AlphaFold modeling. f CCND3 dose-dependently inhibits NP-L interaction. HEK293T cells were co-transfected with the NP (NP-S.tag) and L expression plasmids, along with indicated amounts of the Flag-CCND3 expression plasmid, followed by S-pulldown and WB. g Effects of CCND3 mutants on NP-L interaction. Data are representative from three replicates with similar results (a, b, d, f, and g). Data in the bar graphs (a, b, and d) are means ± SD, n = 3 biological replicates. One-way ANOVA: ****p < 0.0001; ns, non-significant. Source data are provided as a Source Data file.
By interacting with viral RNAs, NP encapsidates the nucleic acid molecules into RNP complexes and is involved in viral transcription and replication11,12. In the “head” region of SFTSV NP, a positively-charged RNA binding cavity, referred to as CavityR, is formed between the N- and C-lobes10,51. Considering that both the two lobes interact with CCND3, we hypothesized that CCND3 may block the RNA binding of NP. RNA-pulldown assays using SFTSV S-segment-derived RNA39 were thus performed. NP was efficiently co-precipitated by the RNA bait; however, this co-precipitation was significantly inhibited by CCND3 (Fig. 8d). In contrast, deleting CN (or CBox) disrupted the inhibitory effect of CCND3 on NP-RNA binding (Fig. 8d), indicating that CN is also required for CCND3 blockade of the NP-RNA interaction. These consistent results are linked to the experimental data characterizing the CCND-NP interaction (Figs. 6 and 7). Moreover, they are also well consistent with the structural modeling of the CCND3-NP complex, in which CCND3 (particularly its CN domain) binding to the two lobes of NP congruously seals the CavityR, providing an interesting structural insight (Figs. 7a, b and 8e).
Finally, S-pulldown assays showed that the interaction between NP and L (the viral RdRp) can also be dose-dependently disrupted by CCND3 (Fig. 8f). However, deleting CN or CBox, but not CC, deprived CCND3 of its disruption of NP-L interaction (Fig. 8g). Taken together, these findings establish that by targeting to the N- and C-lobes of NP, CCND3 blocks the NP multimerization, NP-RNA binding, and NP-L interaction through its CN region, thus interfering with the RNP machinery and virus replication.
NSs counteracts virus/IFN-stimulated CCND3 induction and promotes autophagy-dependent CCND3 degradation
NSs plays multifaceted roles in viral host adaptation, infection, and pathogenesis by antagonizing or manipulating multiple host biological processes, including dampening SFTSV-stimulated PRR and IFN signaling pathways, as well as inducing mTOR-Beclin-1 pro-viral autophagy15,16,17,18,21,53. We therefore considered that CCND3, as an ISG induced by IFNs and SFTSV infection, is likely also subject to NSs-mediated antagonism. First, we observed that NSs did not markedly interfere with the CCND3-mediated suppression of NP activities identified above including the NP multimerization, NP-RNA binding, or NP-L interaction (Supplementary Fig. S8). However, interestingly, the transcriptional upregulation of CCND3, like several other typical ISGs, induced by SFTSV infection was significantly antagonized by NSs, but not a previously reported inactivated mutant NSs-8A16,53,54 (Fig. 9a). Similarly, NSs substantially inhibited IFN induction of CCND3 and other ISGs, whereas the NSs-8A mutant lost this antagonistic activity (Fig. 9b). These findings suggest that NSs could partially counteract the infection- or IFN-induced expression of CCND3, thereby potentially weakening the host’s antiviral response mediated through CCND3. Additionally, we found that treatment with the autophagy inhibitor chloroquine (CQ) or genetic knockout of the autophagy pathway appears to increase endogenous CCND3 protein levels, suggesting that CCND3 itself may undergo constitutive degradation via autophagic flux (Fig. 9c, d). Interestingly, consistent with the previously reported pro-autophagic activity of NSs, NSs (but not the mutant) expression led to a relative reduction of the CCND3 abundance (Fig. 9c, d). Furthermore, the pharmacological inhibition by CQ (but not the proteasome inhibitor MG132) or deletion of the NSs-induced autophagy pathway reported previously abolished NSs-caused CCND3 degradation (Fig. 9c, d), indicating that this effect of NSs is indeed autophagy-dependent. Collectively, these results demonstrate that NSs likely promotes CCND3 degradation by enhancing autophagic activity, which could thereby further contribute to viral counteraction against the host’s CCND3-dependent antiviral response.
a NSs suppression of SFTSV-infection-stimulated CCND3 transcriptional expression. HEK293 cells transfected with NSs or 8A mutant plasmids for 24 h were either left uninfected or infected with SFTSV (MOI = 5). At 24 hpi, induction of the indicated ISGs (CCND3, MxA, OAS2, and ISG15) were analyzed by qPCR. b NSs suppression of IFN-stimulated CCND3 induction. HEK293 cells transfected with the NSs or 8A plasmids were treated with IFN-α for 24 h, followed by qPCR analysis of ISG expression. c NSs expression reduces CCND3 protein abundance, which is abolished by the autophagy inhibitor CQ. HEK293 cells expressing NSs or 8A were treated with the indicated drugs for 48 h and then subjected to WB analysis of CCND3 protein levels. d BECN1-dependent autophagy pathway is required for NSs-induced CCND3 degradation. HEK293 WT or BECN1 KO cells were transfected with the NSs or 8A expression plasmids, followed by WB analysis of CCND3 protein levels at 48 h post transfection. Data are representative from three replicates with similar results (c and d). Data in the bar graphs (a and b) are means ± SD, n = 3 biological replicates. One-way ANOVA: **p < 0.01, ***p < 0.001, ****p < 0.0001. Source data are provided as a Source Data file.
Discussion
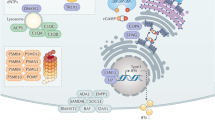
The emerging bandaviruses have posed a significant threat to global public health55. Particularly, SFTSV has been listed by World Health Organization in Pathogens Prioritization Framework (June 2024) as a top priority pathogen which has a high likelihood of causing PHEIC (Public Health Emergencies of International Concern) and requires immediate research and development efforts55. Understanding the intricacies of virus-host interplays is crucial for developing effective antiviral therapies. IFN system contributes significantly to controlling viral infection by mounting the production of ISGs, the effectors of IFN responses. However, it remains unclear which and how ISGs regulate bandavirus infection. Here, we conducted the functional screen of individual ISGs and identified more than 200 candidates potentially regulating SFTSV replication, which may help address the critical knowledge gap regarding the virus-host interactions. Furthermore, with SFTSV as the primary virus model, we uncovered the notable role of CCND3, a top hit from the screen, in restricting bandavirus replication and pathogenicity in vitro and in vivo. Mechanistically, CCND3 targets the “head” (consisting of N- and C-lobes) of the viral NP protein by its CN domain to block NP multimerization, NP-RNA binding, and NP-L interaction, interfering with the RNP machinery and viral replication (as illustrated in a proposed model in Fig. 10). Additionally, CCND3 expression and cytoplasmic localization can be significantly induced by SFTSV infection, which in turn, could further contribute to the antiviral action mediated by CCND3; however, conversely, NSs expression at a relatively later stage of SFTSV infection18 may partially counteract CCND3’s antiviral response by restricting virus/IFN-stimulated upregulation of CCND3 expression and promoting autophagic degradation of CCND3. These findings yield new insights into the virus-host interactions, expanding our understanding of virus infection and host response and potentially aiding future antiviral research.
As with other bandaviruses, SFTSV is an RNA virus that exclusively replicates in the cytoplasm. Upon SFTSV infection, expression of CCND3 as an ISG can be up-regulated by the IFN-mediated JAK-STAT signaling and probably also by the PRR signaling at earlier phase. Particularly, cytoplasmic localization of CCND3 is notably enhanced in response to SFTSV infection. In the cytoplasm, CCND3 exerts its antiviral activity: by its CN domain, CCND3 targets the “head” region, consisting of N- and C-lobes, of the viral NP protein and blocks NP multimerization, NP-RNA binding, and NP-L interaction, inhibiting the RNP activity and viral replication. Conversely, NSs partially counteracts virus and IFN induced upregulation of CCND3 expression and promotes autophagy-dependent CCND3 degradation, likely antagonizing host antiviral response via CCND3. See also text for details. Created in BioRender (Xu, Z. (2025) https://BioRender.com/09o03t1).
Previous studies by large-scale screening have identified antiviral ISGs against several viruses56,57,58,59. In the present study, we screened for the first time the ISGs potentially having regulatory roles in SFTSV infection. Beside CCND3, many other ISG molecules were identified as anti-SFTSV effector candidates which may also merit further function-mechanism studies. Moreover, multiple ISGs were found to have potential pro-viral activities probably enhancing SFTSV replication. It is possible that viruses hijack certain ISGs to indirectly or directly promote their own replication. Exploration of such ISGs could further our knowledge of the overall replication, adaptability and pathogenicity of bandaviruses. However, the expression profile of ISGs that can be induced upon virus infection or IFN stimulation may vary to some extent in different types of cells. Larger-scale screening involving more potentially IFN-stimulated host factors, even in different cells, is pending for future studies to further comprehensively understand the host regulation of bandaviruses. Additionally, we applied a minireplicon reporting system for the screening in this study, which is unrestrained by the demands of high-level biosafety facilities and allows for more ready implementation. Therefore, our study was focused on the RNP machinery-associated transcription and replication process, that is, the central stage of the virus life cycle. In the future, it will be merited to carry out ISG functional screening regarding other stages of bandavirus infection.
Identifying and understanding these pro-viral and antiviral host factors may provide novel perspectives for future antiviral drug development. For antiviral factors like CCND3, potential strategies could involve designing pharmacological agents to enhance their expression levels or strengthen their targeting of viral replication machinery, e.g., reinforcing CCND3 targeting of NP identified here. Alternatively, mechanism-mimicking antiviral peptides could be developed to interfere with viral components like NP. Conversely, for pro-viral factors, approaches might include screening inhibitors to block their virus-supporting interactions, reducing their expression, or leveraging targeted protein degradation technologies (e.g., PROTACs) to eliminate these host factors. These innovative strategies, derived from modulating host-virus interplay, may represent interesting directions for next-generation antiviral therapeutics and warrant further exploration in the future.
As aforementioned, NP-driven RNP construction and RNP functioning are the major and central events during bandavirus infection cycle. Bandavirus RNPs are the viral RNA synthesis machinery catalyzing transcription and replication and those encapsulating viral genome RNAs can be packaged into progeny virion as the structural core11,12. Previous studies by us have demonstrated that host proteins including MOV10 and MxA can target SFTSV RNP and inhibit the viral infection39,40. Here, we further unraveled a clear antiviral mechanism of CCND3 by targeting NP/RNP machinery. These findings therefore support that the viral RNPs and their delicate activities could be vulnerable targets not only for therapeutic design but for host antiviral defense. However, there are several notable differences in the action details of these antiviral molecules. For instance, the N-arm of NP is involved in the targeting of NP by both MOV10 and MxA39,40, but not in the interaction between NP and CCND3. CCND3 specifically targets the NP “head” composed of N- and C-lobes, inhibiting NP-RNA binding. Interestingly, structural modeling analysis coincidently suggested that CCND3 likely binds to the NP “head” and thus seals the RNA binding cavity (CavityR) formed between the N- and C-lobes, blocking the NP-RNA interaction required for encapsulation. In addition to the RNA binding, NP multimerization plays a major role in RNP formation. Although it remains unknown whether MxA affects the NP-RNA binding, previous data showed that MxA has no influence on the NP multimerization40. However, by contrast, the binding of CCND3 to NP “head” also obstructs the multimerization of NP. Again, the AlphaFold complex modeling presented a consistent result showing that CCND3 binding to NP could lead to a significant steric hinderance obstructing NP oligomerization. The divergent targeting mechanisms among these host factors, potentially enabling functional synergy which may merit future investigation. Aside from the targeting of NP by CCND3, we also observed a possible interaction between CCND3 and NSs, but to a less extent. NSs is a nonstructural protein that is not essential for the viral replication but notably, has antagonizing activities against multiple host biological processes including virus-induced PRR/IFN signaling and ISG induction15,16,17,18,20. We thus thought that it, in turn, might counteract the antiviral response mediated by CCND3. First, an additional analysis demonstrated that NSs has no evident antagonizing effects on CCND3 inhibition to the activities of NP, including NP multimerization, NP-RNA binding, or NP-L interaction. Then, as expected, the virus infection- and IFN-stimulated induction of CCND3, as well as several other typical ISGs, was similarly suppressed by NSs. Additionally, we and other groups have demonstrated that NSs induces the autophagy-lysosome pathway that positively supports SFTSV replication21,53, although how the autophagic flux enhances the viral replication remains to be fully addressed. In this study, we found that pharmacological inhibition or genetic deficiency of autophagy led to increased endogenous CCND3 protein abundance, indicating that CCND3 undergoes turnover through the autophagic degradation pathway. Consistent with the previous findings on the pro-autophagic activity of NSs, our data further suggest that NSs likely promotes autophagy-dependent CCND3 degradation. These findings not only demonstrate the potential antagonistic effects of NSs on CCND3, but also advance the understanding of how autophagy facilitates SFTSV replication, expanding the cognizance of virus-host interplays and evolutionary arms races.
CCND3 is one of the D-type cyclins (the other two isoforms are CCND1 and CCND2), which play regulatory roles in the cell cycle42,45,60. These cyclins are expressed in a highly overlapping fashion in different cells and show amino acid similarity and functional redundancy in cell cycle regulation to some extent41,45,61,62. However, in comparison, CCND3 that is expressed in nearly all proliferating cells shows much broader expression pattern than the other two D-type cyclins41,45,63. Interestingly, by a comparative experiment, we found that both CCND1 and CCND2 also inhibit SFTSV replication, which however, is significantly weaker than that mediated by CCND3 (Supplementary Fig. S9). It may be linked with the protein homology but more importantly, suggests the unique antiviral efficacy of CCND3 as an ISG. Apart from the classical cell cycle-associated function, the role of CCND3 in viral infection is poorly understood. According to a few published studies, CCND3 can interact with the M2 protein of influenza virus and the E and M proteins of SARS-CoV-2, inhibiting the viral propagation64,65. In contrast, Ruiz et al. reported that CCND3 may have a positive role in supporting HIV-1 replication66. We here uncovered that by its CN domain, CCND3 targets the pivotal structural and functional regions of SFTSV NP and correspondingly blocks multiple activities of the viral protein required for the RNP machinery, thereby restricting the bandavirus replication and pathogenicity in vitro and in vivo. The current study thus more systematically proposes a clear, new antiviral mode of CCND3, expanding the knowledge of CCND3 biological functions. Additionally, mutation and even the large functional domain truncation did not deprive CCND3 of its antiviral capacity. Consistently, KO of CDK4, CDK6, or both did not impair CCND3’s anti-SFTSV activity or lead to any enhancement of the viral replication, unlike the effects of CCND3 KO, indicating that the antiviral activity of CCND3 is independent of its cell cycle-related physiological functions. Further, although the expression of CCND3 can be induced by viral infection and IFN stimulation, its anti-SFTSV action is independent of the IFN signaling. These observations are also consistent with the interesting antiviral mechanism found in this study, where CCND3 targets NP by a specific protein-protein interaction, directly interfering with the RNP machinery and hence virus replication.
In summary, we identified many ISGs potentially inhibiting or bolstering SFTSV replication. As an exemplification, CCND3 was then validated to be a new host restriction factor against bandavirus infection and pathogenicity. Moreover, a sophisticated antiviral mode exploited by CCND3 was unraveled in details, presenting a typical instance regarding host antiviral mechanism. This study may contribute to comprehensive understanding of the virus-host interactions and promote the development of antiviral therapy in the future.
Methods
Ethics statement
Animal experiments were approved by the Institutional Animal Care and Use Committee and the Ethical Committee of Wuhan Institute of Virology, Chinese Academy of Sciences (Approval No. WIVA23202301), and conducted in accordance to the guidelines for the Care and Use of Medical Laboratory Animals (Ministry of Health, China). All mice were housed under controlled conditions: temperature maintained at 22–24 °C, relative humidity at 40–60 %, and a 12-h light/dark cycle.
Cells and viruses
Human embryonic kidney 293 cells (HEK293, ATCC, CRL-1573) were cultured in minimum Eagle’s medium (MEM, Gibco) supplemented with 10% fetal bovine serum (FBS) at 37 °C under 5% CO2. HEK293T cells (ATCC, CRL-11268), African green monkey kidney cells (Vero, ATCC, CCL-81), mouse embryonic fibroblast cells (MEF; National Virus Resource Center, NVRC), L929 cells (ATCC, CCL-1) and hamster BHK-21 cells (NVRC) were maintained in Dulbecco’s modified Eagle’s medium (DMEM, Hubei NZK) supplemented with 10% FBS. Human monocyte/macrophage cells (THP-1, ATCC, TIB-202) were cultured in Roswell Park Memorial Institute (RPMI) 1640 medium (Gibco) supplemented with 10% FBS, or induced with phorbol 12-myristate 13-acetate (PMA, 100 ng/mL; MCE, Cat#HY-18739) for 48 h for cell differentiation. Human peripheral blood mononuclear cells (PBMCs; ATCC, PCS-800-011) were cultured in RPMI-1640 medium. Mouse Bone Marrow-Derived Macrophages (BMDMs) isolated from the femurs and tibias of mice were cultured in 30% L929 mouse fibroblast supernatant and 70% RPMI-1640 medium with 10% FBS for 7 days67. Severe fever with thrombocytopenia syndrome virus (SFTSV, strain WCH; CSTR: 16533.06. IVCAS 6.6088), Heartland virus (HRTV, strain MO-4; CSTR: 16533.06. IVCAS 6.6330), and Guertu virus (GTV, strain DXM; CSTR: 16533.06. IVCAS 6.6106) were propagated in Vero cells and titrated by the 50% tissue culture infectious dose (TCID50) method15,39,40.
Plasmids and transfection
Expression plasmids for CCND3 and its mutants were constructed using pcDNA3.1(+)-N-Flag, pcDNA3.1(+)-C-HA, or pCAGGS-C-S.tag or HA vectors. Plasmids expressing EGFP-fused NP and NP truncated mutants were constructed using the pEGFP-N1 vector. Plasmids encoding EGFP-fused CCND3 and its individual domains were constructed using the pEGFP-C1 vector. The SFTSV NP, L, or NSs encoding plasmids, EGFP or firefly luciferase-based minigenome reporter plasmids, and Renilla luciferase internal control plasmid (pRL-TK) were described previously15,38,39,40. Plasmids used for RNAi or gene editing were constructed according to the standard procedures as stated in the following. The cDNA expression clones of individual ISGs for functional screening were picked from GIPZ human genome cDNA library (GE Healthcare Dharmacon) or supplementally constructed through standard molecular biology techniques by referencing previous studies57,58, generating a library of 378 ISGs. Plasmid transfection was performed with Lipofectamine 3000 transfection reagent (Invitrogen, Cat#L3000015) according to the manufacturer’s instructions. For RNAi with siRNAs, the gene-specific or control siRNA duplexes (synthesized by Sangon Biotech) were transfected into HEK293 cells using RNATransMate (Sangon Biotech, Cat#E607402-1000).
Antibodies and reagents
Rabbit antisera to SFTSV NP, RdRp, GP (Gn), and NSs, and mouse anti-SFTSV NP serum were described previously15,19,21,39. Mouse anti-Flag mAb (Sigma-Aldrich, Cat#F3165), anti-HA-tag mAb (Proteintech, Cat#66006-2-Ig), anti-CCND3 mAb (Proteintech, Cat#66357-1-Ig) and anti-β-actin mAb (ABclonal, Cat#AC026), and rabbit anti-CDK4 pAb (ABclonal, Cat# A0366), anti-CDK6 mAb (ABclonal, Cat# A0106), anti-S-tag pAb (Sino Biological, Cat#ab101290-T38), and anti-EGFP-tag pAb (Proteintech, Cat#50430-2-AP) were purchased from the indicated manufacturers. For the secondary antibodies, goat anti-mouse IgG conjugated with Alexa Fluor 488 (Thermo Fisher Scientific, Cat#A-11001) and goat anti-rabbit IgG conjugated with Alexa Fluor 647 (Thermo Fisher Scientific, Cat#A32733) were used in immunofluorescence assays; goat anti-mouse or anti-rabbit IgG antibodies conjugated with HRP (Abcam, Cat#ab6789 and Cat#ab6721) were used for Western blot analysis.
Chloroquine (MCE, Cat#HY-17589A) and MG-132(MCE, Cat#HY-13259) were used for cell treatments.
Western blot
For Western blot (WB) analysis, protein samples were subjected to 12 to 15% SDS-polyacrylamide gel electrophoresis (SDS-PAGE) and then transferred to polyvinylidene difluoride membranes (PVDF) (Millipore, Cat#L3000015). After blocking with 5% non-fat milk in Tris-buffered saline and Tween 20 (TBST) (BOSTER, Cat#AR0195-10), the membranes were further incubated with the indicated primary antibodies overnight at 4 °C and the corresponding horseradish peroxidase (HRP)-conjugated secondary antibodies for 2 h at room temperature. Protein bands were detected by SuperSignal West Pico PLUS (Thermo Scientific, Cat#34580) using a chemiluminescence analyzer (Azure Biosystems, lnc C280) and analyzed by ImageJ software. Uncropped scans of the blots are provided in the Source Data file.
Immunofluorescence and confocal microscopy
Transfected or infected cells were fixed by 4% paraformaldehyde fixation (PFA) for 30 min, permeabilized by 0.5% Triton X-100 for 15 min, and blocked with 5% bovine serum albumin for 1 h. The cells were then incubated with primary antibodies overnight at 4 °C and with fluorescence-labeled secondary antibodies for 1 h at room temperature. Nuclei were stained with Hoechst 33258 (Beyotime, Cat#C1011). Confocal analysis was performed using a Leica sp8 laser confocal microscope. Images were analyzed using Leica Application Suite X software.
RNA extraction and qPCR
Total RNA from cells or mouse tissues was extracted using RNAiso Plus (TAKARA, Cat#9109). The cDNA was synthesized using a HiScript II Q RT SuperMix for qPCR kit (Vazyme, Cat#R223-01). qPCR was performed with specific primers as listed in Supplementary Table S1. Relative RNA levels normalized to the mRNA levels of GAPDH were calculated by the 2−ΔΔCt method. RNA in mouse serum was isolated with a TaKaRa MiniBEST Universal RNA Extraction Kit (TAKARA, Cat#9767) for absolute quantification of SFTSV RNA by TaqMan real-time PCR using HiScript II One Step RT-PCR Kit (Vazyme, Cat#P611-01). SFTSV S-segment RNA synthesized with the T7 RNA polymerase transcription kit (Ambion, Cat#AM1314) was used to construct the standard curve20,39.
Minigenome reporter assay and ISG cDNA library Screening
The SFTSV minigenome reporter assays were performed as described previously38. Briefly, the L and NP expression vectors and EGFP-based M-segment minigenome transcription plasmid (pRF42-MUTR-EGFP) were co-transfected into BHK-21 cells in 96-well plates, together with the plasmids encoding CCND3 or its mutants or control vectors. Forty-eight hours post transfection, cells were fixed with 4% PFA and then stained with Hoechst 33258 for 5 min. EGFP-positive cell counts were determined using the Operetta CLSTM high-throughput system (PerkinElmer) and positive ratios (over total cell counts) were then calculated and normalized to the control group. For library screening, the expression plasmids encoding individual ISGs from the cDNA library (Supplementary Data 1) were co-transfected with the minigenome system plasmids, followed by the high-throughput analysis. In the minigenome system with luciferases as reporters, the firefly luciferase-based minigenome transcription plasmid (pRF42-MUTR-LUC) along with the control plasmid (pRF42-TK) were used to replace pRF42-MUTR-EGFP for co-transfection. At 48 h posttransfection, cells were delivered to luciferase activity measurement using a dual-luciferase reporter kit (Promega, Cat#E2940). Relative luciferase activities (Rel. Luc. Act.) were then calculated18,68.
Nuclear-cytoplasmic fractionation
Cells were infected with SFTSV at various MOI or mock infected. Nuclear-cytoplasmic isolation was conducted at 24 hpi using the Nuclear and Cytoplasmic Protein Extraction Kit (Beyotime, Cat#P0028) according to the manufacturer’s protocol, followed by WB analysis. Protein bands were analyzed by ImageJ and normalized to β-actin (cytoplasm) and histone deacetylase1 (HDAC1) (nucleus), respectively.
Protein interaction analysis
Transfected or infected cells were suspended in the immunoprecipitation (IP) lysis buffer (Beyotime, Cat#P0013) supplemented with proteinase inhibitor cocktail (Roche, Cat#04693116001). Supernatants of the cell lysates were then subjected to protein interaction analyses15,18,46,69. For S-pulldown, Flag-IP, or EGFP-NanoTrap assays, the supernatants were respectively incubated with S-protein agarose (Millipore, Cat#69704), anti-Flag Magnetic Beads (MCE, Cat#HY-K0207), or anti-EGFP nanobody-coated agarose beads (AlpaLife, Cat#KTSM1301) for 4 h with gentle rotation at 4 °C. For co-immunoprecipitation (Co-IP) assays, HEK293 cells mock-infected or infected with SFTSV were harvested and lysed using cell lysis buffer (Beyotime, Cat#P0013) at 24 hpi. Subsequently, supernatants of the cell lysates were incubated with the anti-CCND3 antibody or control IgG, together with protein A/G beads, overnight with gentle rotation at 4 °C. After washing, the precipitates were eluted by boiling for 5 min in 1 × SDS sample buffer, followed by WB analyses with specific antibodies. To analyze the effect of nucleic acids on protein-protein interactions, cell lysate samples containing 5 mM MgCl2 were treated with UltraNuclease (Yeasen, Cat# 20157ES25) for 1 h at room temperature before the S-pulldown assay40.
Chemical cross-linking
HEK293T cells were transfected with the indicated expression plasmids or control vectors. At 24 h posttransfection, the cells were harvested and subjected to cross-linking with disuccinimidyl suberate (DSS, Thermo Fisher Scientific, Cat#A39267) at room temperature for 30 min. The reaction was then stopped with 0.1 M Tris-HCl (pH 7.5), before WB analysis of the NP monomer and oligomers39,40.
RNA-protein pulldown assay
RNA-protein pulldown assays were conducted as previously described39. Briefly, SFTSV S-segment RNA was synthesized with a T7 RNA Polymerase Transcription Kit (Ambion, Cat#AM1314), followed by the addition of biotin label to the 3’ end using a Thermo Scientific Pierce RNA 3’-Desthiobiotinylation Kit (Thermo Fisher Scientific, Cat#20163). The labeled RNA was extracted with an equal volume of chloroform/isoamyl alcohol, followed by ethanol precipitation. The biotin-labeled RNA resuspended in nuclease-free water (40 pmol) was incubated with 40 μL of streptavidin magnetic beads (Thermo Fisher Scientific, Cat#20164) at room temperature for 30 min according to manufacturer’s instructions. To facilitate the RNA-protein pulldown analysis, supernatants of indicated cell lysates were rotated with the RNA-bound magnetic beads at 4 °C for 1 h, followed by WB detection of the co-precipitates and lysate inputs.
Construction of knockout (KO) cells by CRISPR-Cas9 gene editing
The human CCND3, CDK4, CDK6 and IFNAR1 gene specific sgRNAs were designed using the online CRISPR Design Tools (https://zlab.bio/guide-design-resources) and cloned into px459 using BbsI restriction sites70. HEK293 cells were transfected with the px459-CCND3, CDK4, or CDK6 sgRNA plasmids and selected for 3 d in the presence of puromycin. Clonal CCND3, CDK4, or CDK6-deficient cell lines were respectively obtained by limiting dilution and validated by WB analysis and sequencing with the specific primers as listed in Supplementary Table S1. For construction of IFNAR1-KO THP-1 cells, the human IFNAR1 gene sgRNA was cloned into lentiCRISPRv2 (Addgene, Cat#52961) and packaged in HEK293T cells with packaging plasmids psPAX2 and pMD2.G39,46. THP-1 cells were infected with the obtained lentivirus vector and selected for 5 d with puromycin. Similarly, the candidate cell lines were validated by WB and sequencing with specific primers (Supplementary Table S1).
CCND3-deficient mice and SFTSV infection experiment
C57BL/6J and IFNAR1−/− (A129) mice were bred in-house at the institutional animal facility under specific pathogen-free (SPF) conditions. The transduction vectors were packaged by co-transfection of HEK293T cell with the CCND3-targeting or control shRNA pLKO.1 plasmids together with the packaging vectors psPAX2 and pMD2.G. At 48 h and 72 h posttransfection, culture supernatants were collected, filtered through a 0.45 μm filter (Millipore, Cat#SLHP033RB), and concentrated by centrifugation (72,100 g, 2 h, 4 °C). The virus particles were suspended in Virus Conservation Solution (PBS with 1% BSA, pH = 7.4) and titrated using the endpoint method71. For in vivo KD analysis in immunocompetent animals, female C57BL/6J mice (6–8 weeks old; n = 6) were injected with 1 × 108 transduction units (TU) of either control or CCND3-targeting viral vectors via the tail vein. Seven days after transduction, the mice were subcutaneously infected with SFTSV (105 TCID50) and three days after infection, euthanized for necropsy and tissue sample collection. Histopathological examination including hematoxylin and eosin (H&E) staining of tissue samples were performed as previously described with the help of Wuhan Servicebio Technology72,73,74,75. Images were obtained by PerkinElmer Vectra Polaris and analyzed by P250 (3D HISTECH). Injury scores for various tissues from each mouse were determined based on specific pathological criteria75,76,77. Platelet (PLT) and white blood cell (WBC) counts in fresh blood treated with EDTA were measured using a hematology analyzer (Drew Scientific, Mascot HEMAVET950). Serum alanine aminotransferase (ALT) (Servicebio, GM1102), aspartate aminotransferase (AST) (Servicebio, GM1103), and blood urea nitrogen (UREA) (Servicebio, GM1110) were measured by ELISA using the indicated kits. CCND3 mRNA levels and viral copies in tissue samples were delivered to qPCR analyses. For KD analysis in the IFNAR1−/− models, female A129 mice (6–8 weeks old) were subjected to the similar CCND3 KD method applied to C57BL/6J. After a week, A129 CCND3-KD or control groups (n = 6) were infected with SFTSV (10³ TCID50), and body weight changes and survival rates were monitored daily. Additionally, in a parallel experiment, infected animals (n = 6) were sacrificed at 3 dpi, followed by tissue sample collection and histopathological analyses as described above. For CCND3 induction analysis, female C57BL/6J mice (n = 4) were intraperitoneally injected with SFTSV (106 TCID50). At indicated time points post infection, tissue samples were collected for qPCR analysis of Ccnd3 mRNA and SFTSV RNA levels.
Statistical analysis
Statistical analysis was performed using Student’s t test or analysis of variance (ANOVA) with GraphPad Prism 9 software (La Jolla, CA, USA). Data are presented as the means ± standard deviations (SD) of n biological replicates. Differences were considered statistically significant if p-value < 0.05. Significance levels are: *p < 0.05; **p < 0.01; *** p < 0.001; *** p < 0.001; **** p < 0.0001; ns, non-significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings of this study are available within the paper and its Supplementary Information. Source data are provided with this paper.
References
Yu, X. J. et al. Fever with thrombocytopenia associated with a novel bunyavirus in China. N. Engl. J. Med. 364, 1523–1532 (2011).
Zhan, J. et al. Current status of severe fever with thrombocytopenia syndrome in China. Virologica Sin. 32, 51–62 (2017).
Yoo, J. R. et al. Family Cluster Analysis of Severe Fever with Thrombocytopenia Syndrome Virus Infection in Korea. Am. J. tropical Med. Hyg. 95, 1351–1357 (2016).
Jiang, X. L. et al. A cluster of person-to-person transmission cases caused by SFTS virus in Penglai, China. Clin. Microbiol Infect. 21, 274–279 (2015).
Matsumoto, C. et al. Investigation of antibody to severe fever with thrombocytopenia syndrome virus (SFTSV) in blood samples donated in a SFTS-endemic area in Japan. Vox sanguinis 113, 297–299 (2018).
Tran, X. C. et al. Endemic Severe Fever with Thrombocytopenia Syndrome, Vietnam. Emerg. Infect. Dis. 25, 1029–1031 (2019).
McMullan, L. K. et al. A New Phlebovirus Associated with Severe Febrile Illness in Missouri. N. Engl. J. Med. 367, 834–841 (2012).
Shen, S. et al. A novel tick-borne phlebovirus, closely related to severe fever with thrombocytopenia syndrome virus and Heartland virus, is a potential pathogen. Emerg. Microbes Infect. 7, 1–14 (2018).
Siddell, S. G. et al. Virus taxonomy and the role of the International Committee on Taxonomy of Viruses (ICTV). J. Gen. Virol. 104, 001840 (2023).
Zhou, H. et al. The nucleoprotein of severe fever with thrombocytopenia syndrome virus processes a stable hexameric ring to facilitate RNA encapsidation. Protein cell 4, 445–455 (2013).
Zhou, H. G., Sun, Y. N., Guo, Y. & Lou, Z. Y. Structural perspective on the formation of ribonucleoprotein complex in negative-sense single-stranded RNA viruses. Trends Microbiol 21, 475–484 (2013).
Sun, Y., Li, J., Gao, G. F., Tien, P. & Liu, W. Bunyavirales ribonucleoproteins: the viral replication and transcription machinery. Crit. Rev. Microbiol. 44, 522–540 (2018).
Malet, H., Williams, H. M., Cusack, S. & Rosenthal, M. The mechanism of genome replication and transcription in bunyaviruses. PLoS Pathog. 19, e1011060 (2023).
Cheng, E. D., Wang, Z. K. & Mir, M. A. Interaction between Hantavirus Nucleocapsid Protein (N) and RNA-Dependent RNA Polymerase (RdRp) Mutants Reveals the Requirement of an N-RdRp Interaction for Viral RNA Synthesis. J. Virol. 88, 8706–8712 (2014).
Ning, Y. J. et al. Viral suppression of innate immunity via spatial isolation of TBK1/IKKε from mitochondrial antiviral platform. J. Mol. cell Biol. 6, 324–337 (2014).
Wu, X. et al. Evasion of antiviral immunity through sequestering of TBK1/IKKε/IRF3 into viral inclusion bodies. J. Virol. 88, 3067–3076 (2014).
Santiago, F. W. et al. Hijacking of RIG-I signaling proteins into virus-induced cytoplasmic structures correlates with the inhibition of type I interferon responses. J. Virol. 88, 4572–4585 (2014).
Ning, Y. J. et al. Disruption of type I interferon signaling by the nonstructural protein of severe fever with thrombocytopenia syndrome virus via the hijacking of STAT2 and STAT1 into inclusion bodies. J. Virol. 89, 4227–4236 (2015).
Ning, Y. J. et al. Interferon-γ-Directed Inhibition of a Novel High-Pathogenic Phlebovirus and Viral Antagonism of the Antiviral Signaling by Targeting STAT1. Front. Immunol. 10, 1182 (2019).
Min, Y.-Q., Ning, Y.-J., Wang, H. & Deng, F. A RIG-I–like receptor directs antiviral responses to a bunyavirus and is antagonized by virus-induced blockade of TRIM25-mediated ubiquitination. J. Biol. Chem. 295, 9691–9711 (2020).
Feng, K. et al. SFTS bunyavirus NSs protein sequestrates mTOR into inclusion bodies and deregulates mTOR-ULK1 signaling, provoking pro-viral autophagy. J. Med. Virol. 95, e28371 (2023).
MacMicking, J. D. Interferon-inducible effector mechanisms in cell-autonomous immunity. Nat. Rev. Immunol. 12, 367–382 (2012).
Darnell, J. E., Kerr, I. M. & Stark, G. R. Jak-stat pathways and transcriptional activation in response to IFNs and other extracellular signaling proteins. Science 264, 1415–1421 (1994).
Chow, K. T., Gale, M. Jr. & Loo, Y. M. RIG-I and other RNA sensors in antiviral immunity. Annu. Rev. Immunol. 36, 667–694 (2018).
Takeuchi, O. & Akira, S. Pattern recognition receptors and inflammation. Cell 140, 805–820 (2010).
Yamada, S. et al. RIG-I-like receptor and toll-like receptor signaling pathways cause aberrant production of inflammatory cytokines/chemokines in a severe fever with thrombocytopenia syndrome virus infection mouse model. J. Virol. 92, e02246-17 (2018).
Lazear, H. M., Schoggins, J. W. & Diamond, M. S. Shared and distinct functions of type I and type III interferons. Immunity 50, 907–923 (2019).
Der, S. D., Zhou, A., Williams, B. R. & Silverman, R. H. Identification of genes differentially regulated by interferon alpha, beta, or gamma using oligonucleotide arrays. Proc. Natl. Acad. Sci. USA 95, 15623–15628 (1998).
Briscoe, J. et al. JAKs, STATs and signal transduction in response to the interferons and other cytokines. Philos. Trans. R. Soc. Lond. B Biol. Sci. 351, 167–171 (1996).
Schneider, W. M., Chevillotte, M. D. & Rice, C. M. Interferon-stimulated genes: a complex web of host defenses. Annu. Rev. Immunol. 32, 513–545 (2014).
Schoggins, J. W. Interferon-stimulated genes: what do they all do?. Annu. Rev. Virol. 6, 567–584 (2019).
Gowen, B. B. et al. Modeling severe fever with thrombocytopenia syndrome virus infection in golden Syrian hamsters: importance of STAT2 in preventing disease and effective treatment with favipiravir. J. Virol. 91, e01942-16 (2017).
Sun, J. et al. Animal model of severe fever with thrombocytopenia syndrome virus infection. Front. Microbiol. 12, 797189 (2021).
Fujii, H. et al. Susceptibility of type I interferon receptor knock-out mice to heartland bandavirus (HRTV) infection and efficacy of favipiravir and ribavirin in the treatment of the mice infected with HRTV. Viruses-Basel 14, 1668 (2022).
Feng, K. et al. Pathogenesis and virulence of Heartland virus. Virulence 15, 2348252 (2024).
Fujii, H. et al. Pathological and virological findings of type I interferon receptor knockout mice upon experimental infection with Heartland virus. Virus Res. 340, 199301 (2024).
Westover, J. B. et al. Modeling Heartland virus disease in mice and therapeutic intervention with 4′-fluorouridine. J. Virol. 98, e0013224 (2024).
Ren, F., Zhou, M., Deng, F., Wang, H. & Ning, Y.-J. Combinatorial minigenome systems for emerging banyangviruses reveal viral reassortment potential and importance of a protruding nucleotide in genome “panhandle” for promoter activity and reassortment. Front Microbiol 11, 599 (2020).
Mo, Q., Xu, Z., Deng, F., Wang, H. & Ning, Y. J. Host restriction of emerging high-pathogenic bunyaviruses via MOV10 by targeting viral nucleoprotein and blocking ribonucleoprotein assembly. PLoS Pathog. 16, e1009129 (2020).
Chang, M. et al. Host factor MxA restricts Dabie bandavirus infection by targeting the viral NP protein to inhibit NP-RdRp interaction and ribonucleoprotein activity. J. Virol. 98, e0156823 (2024).
Bartkova, J., Lukas, J., Strauss, M. & Bartek, J. Cyclin D3: requirement for G1/S transition and high abundance in quiescent tissues suggest a dual role in proliferation and differentiation. Oncogene 17, 1027–1037 (1998).
Sherr, C. J. D-Type Cyclins. Trends Biochem Sci. 20, 187–190 (1995).
Weng, A. P. & Aster, J. C. No T without D3: a critical role for cyclin D3 in normal and malignant precursor T cells. Cancer Cell 4, 417–418 (2003).
Sicinska, E. et al. Requirement for cyclin D3 in lymphocyte development and T cell leukemias. Cancer Cell 4, 451–461 (2003).
Saleban, M., Harris, E. L. & Poulter, J. A. D-type cyclins in development and disease. Genes-Basel 14, 1445 (2023).
Dai, S. et al. Interactome profiling of Crimean-Congo hemorrhagic fever virus glycoproteins. Nat. Commun. 14, 7365 (2023).
Best, S. et al. Interferon-stimulated TRIM69 interrupts dengue virus replication by ubiquitinating viral nonstructural protein 3. PLoS Pathog. 14, e1007287 (2018).
Chen, X. P. et al. Infection and pathogenesis of Huaiyangshan virus (a novel tick-borne bunyavirus) in laboratory rodents. J. Gen. Virol. 93, 1288–1293 (2012).
Casanovas, O., Jaumot, M., Paules, A. B., Agell, N. & Bachs, O. P38SAPK2 phosphorylates cyclin D3 at Thr-283 and targets it for proteasomal degradation. Oncogene 23, 7537–7544 (2004).
Takaki, T. et al. The structure of CDK4/cyclin D3 has implications for models of CDK activation. Proc. Natl. Acad. Sci. USA 106, 4171–4176 (2009).
Jiao, L. Y. et al. Structure of severe fever with thrombocytopenia syndrome virus nucleocapsid protein in complex with suramin reveals therapeutic potential. J. Virol. 87, 6829–6839 (2013).
Alfadhli, A. et al. Hantavirus nucleocapsid protein oligomerization. J. Virol. 75, 2019–2023 (2001).
Li, Z. M. et al. Bunyavirus SFTSV NSs utilizes autophagy to escape the antiviral innate immune response. Autophagy 20, 2133–2145 (2024).
Choi, Y. et al. Severe fever with thrombocytopenia syndrome phlebovirus non-structural protein activates TPL2 signalling pathway for viral immunopathogenesis. Nat. Microbiol. 4, 429–437 (2019).
World Health Organization (WHO). Pathogens prioritization: a scientific framework for epidemic and pandemic research preparedness. https://www.who.int/publications/m/item/pathogens-prioritization-a-scientific-framework-for-epidemic-and-pandemic-research-preparedness (2024).
Schoggins, J. W. et al. A diverse range of gene products are effectors of the type I interferon antiviral response. Nature 472, 481–485 (2011).
Schoggins, J. W. et al. Dengue reporter viruses reveal viral dynamics in interferon receptor-deficient mice and sensitivity to interferon effectors in vitro. Proc. Natl Acad. Sci. USA 109, 14610–14615 (2012).
Schoggins, J. W. et al. Pan-viral specificity of IFN-induced genes reveals new roles for cGAS in innate immunity. Nature 505, 691–695 (2014).
Kuroda, M. et al. Identification of interferon-stimulated genes that attenuate Ebola virus infection. Nat. Commun. 11, 2953 (2020).
Malumbres, M. & Barbacid, M. Cell cycle, CDKs and cancer: a changing paradigm. Nat. Rev. Cancer 9, 153–166 (2009).
Beumer, T. L., Roepers-Gajadien, H. L., Gademan, I. S., Kal, H. B. & de Rooij, D. G. Involvement of the D-type cyclins in germ cell proliferation and differentiation in the mouse. Biol. Reprod. 63, 1893–1898 (2000).
Sherr, C. J. & Roberts, J. M. Living with or without cyclins and cyclin-dependent kinases. Genes Dev. 18, 2699–2711 (2004).
Weng, H. Y., Huang, H. L., Zhao, P. P., Zhou, H. & Qu, L. H. Translational repression of cyclin D3 by a stable G-quadruplex in its 5’ UTR: implications for cell cycle regulation. RNA Biol. 9, 1099–1109 (2012).
Fan, Y. et al. Cell cycle-independent role of cyclin D3 in host restriction of influenza virus infection. J. Biol. Chem. 292, 5070–5088 (2017).
Gupta, R. K. & Mlcochova, P. Cyclin D3 restricts SARS-CoV-2 envelope incorporation into virions and interferes with viral spread. EMBO J. 41, e111653 (2022).
Ruiz, A. et al. Cyclin D3-dependent control of the dNTP pool and HIV-1 replication in human macrophages. Cell Cycle (Georget. Tex.) 14, 1657–1665 (2015).
Toda, G., Yamauchi, T., Kadowaki, T. & Ueki, K. Preparation and culture of bone marrow-derived macrophages from mice for functional analysis. STAR Protoc. 2, 100246 (2021).
Feng, K., Deng, F., Hu, Z., Wang, H. & Ning, Y. J. Heartland virus antagonizes type I and III interferon antiviral signaling by inhibiting phosphorylation and nuclear translocation of STAT2 and STAT1. J. Biol. Chem. 294, 9503–9517 (2019).
Ning, Y. J. et al. Heartland virus NSs protein disrupts host defenses by blocking the TBK1 kinase-IRF3 transcription factor interaction and signaling required for interferon induction. J. Biol. Chem. 292, 16722–16733 (2017).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Geraerts, M., Willems, S., Baekelandt, V., Debyser, Z. & Gijsbers, R. Comparison of lentiviral vector titration methods. BMC Biotechnol. 6, 34 (2006).
Kang, Z. Y. et al. Myocardial mitochondrial antiviral signaling protein promotes heart Ischemia-reperfusion injury via RIG-I signaling in mice. Nat. Commun. 16, 5101 (2025).
Ni, L. Q. et al. Antitumor efficacy of CRISPR/Cas9-engineered ICP6 mutant herpes simplex viruses in a mouse xenograft model for lung adenocarcinoma. J. Med. Virol. 94, 6000–6015 (2022).
Ning, Y. J. et al. Ebola virus mucin-like glycoprotein (Emuc) induces remarkable acute inflammation and tissue injury: evidence for Emuc pathogenicity. Protein Cell 9, 389–393 (2018).
Zhang, C. et al. Host protein ARF1 is a proviral factor for SARS-CoV-2 and a candidate broad-spectrum therapeutic target. Nat. Commun. 16, 6326 (2025).
Lu, J. et al. A full-length glycoprotein mRNA vaccine confers complete protection against severe fever with thrombocytopenia syndrome virus, with broad-spectrum protective effects against bandaviruses. J. Virol. 98, e0076924 (2024).
Min, Y. Q. et al. Developing nucleoside tailoring strategies against SARS-CoV-2 via ribonuclease targeting chimera. Sci. Adv. 10, eadl4393 (2024).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2023YFC2305900 to YJN), National Natural Science Foundation of China (32170171 to YJN), and Youth Innovation Promotion Association of Chinese Academy of Sciences (2020333 to YJN). We thank the National Virus Resource Center (Wuhan Institute of Virology), World Reference Center for Emerging Viruses and Arboviruses (University of Texas Medical Branch), and Prof. Wuchun Cao from State Key Laboratory of Pathogen and Biosecurity (Beijing Institute of Microbiology and Epidemiology) for providing GTV (strain DXM), HRTV (MO-4), and SFTSV (WCH) resources, respectively. We also would like to thank the Core Facility and Technical Support of Wuhan Institute of Virology and the running team of National Biosafety Laboratory, Wuhan, Chinese Academy of Sciences for technical assistants.
Author information
Authors and Affiliations
Contributions
Supervision: H.L.W. and Y.J.N. Funding acquisition: Y.J.N. and H.L.W. Resources: Y.J.N., H.L.W. and F.D. Conceptualization: Y.J.N. Investigation: Z.X. and Z.Y.J. with assistance from H.Y.Z. and C.S. Data curation and raw data validation: Z.X. and K.F. Writing - original draft: Y.J.N. with input on experimental details and results from ZX and other authors. Writing - review & editing: Y.J.N. All authors have read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Emily Yang, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xu, Z., Jiang, Z., Feng, K. et al. Interferon-stimulated gene screening identifies CCND3 as a host restriction factor against emerging high-pathogenic bandaviruses. Nat Commun 16, 7935 (2025). https://doi.org/10.1038/s41467-025-63295-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63295-4