Abstract
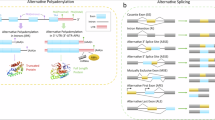
Alternative polyadenylation (APA) generates transcript diversity by producing mRNA isoforms with distinct 3’ ends. Despite the critical roles that APA plays in various biological processes, the mechanisms regulating APA in response to stresses have remained poorly understood in plants. Here, we perform comprehensive analysis of APA in tomato, and focus on a phosphate (Pi)- regulated APA gene SlSPX5, encoding a putative Pi sensor protein. SlSPX5 interacts with and sequesters the transcription factor SlPHL1 in the cytosol, thereby inhibiting the expression of Pi starvation inducible genes. We discover that a cis-natural antisense RNA (cis-NAT) is activated from SlSPX5 to promote its proximal polyadenylation under Pi-depleted conditions. The transcription of this cis-NAT induces RNA Polymerase II pausing, generating Ser2 phosphorylation signals that recruit polyadenylation machinery to the 5’ end of SlSPX5. Our findings demonstrate that a cis-NAT regulates APA of its cognate gene in response to Pi starvation.
Similar content being viewed by others
Introduction
Phosphorus is an essential macronutrient for plant development, playing a crucial role in numerous biological processes, such as nucleic acid biosynthesis, energy metabolism, cellular compartmentalization, and various signaling pathways1,2. However, bioavailable phosphate (Pi) is limited in most soils worldwide, as it is often present as insoluble inorganic phosphates or organic forms that plant roots cannot absorb directly3,4. In response to Pi starvation, plants have evolved several adaptive mechanisms. These include alterations in root system architecture, such as reduced primary root growth and enhanced lateral root initiation5, metabolic adjustments like anthocyanin accumulation to protect nucleic acids6,7, and induction of a series of Pi starvation-inducible (PSI) genes to regulate Pi homeostasis7.
The transcription factor PHOSPHATE RESPONSE1 (PHR1) plays a central role in Pi starvation responses in Arabidopsis thaliana8,9. Under Pi starvation conditions, it binds to the cis-element P1BS (GNATATNC) in the promoters of hundreds of PSI genes, activating their expression9,10. Similarly, in rice, OsPHR2, the orthologue of AtPHR1, binds to the P1BS element to regulate PSI gene expression11. Loss-of-function mutations in PHR genes result in decreased anthocyanin accumulation and reduced induction of PSI genes8,10. In contrast, overexpression of PHR genes enhances Pi uptake and accumulation12,13. Additionally, PHRs are also essential for mycorrhizal symbiosis by regulating the expression of symbiosis-related genes14.
SPX, named after Saccharomyces cerevisiae SYG1/Pho81/mammalian Xpr1, is a hydrophilic and poorly conserved domain found in many proteins related to Pi homeostasis in various species15,16. Crystal structures of the SPX domains from human, yeast, and Chaetomium thermophilum demonstrate that several conserved amino acid residues in the α1, α2 and α4 helices form a positively charged surface, facilitating the binding of inositol polyphosphates17. In Arabidopsis and rice, SPX domain proteins SPX1 and SPX2 are localized in the nucleus where they interact with PHR1 (PHR2 in rice) in a Pi-dependent manner, indicating their roles as Pi sensors18,19. In 2019, we identified inositol pyrophosphate InsP8 as the intracellular Pi signaling molecule20. Under Pi-replete conditions, InsP8 directly binds to the SPX domain, promoting the interaction between SPX1 and PHR1, thereby repressing PHR1 function20,21. Recent studies have shown that InsP8 also functions as a bona fide Pi signaling molecule in mammals and fungi22,23,24. Both Arabidopsis PHR1 and rice PHR2 form a dimer to bind the P1BS element9,25. We determined the structure of the rice SPX1-PHR2 complex and found that the Pi signaling molecule stabilizes helix α1 of SPX1. When two SPX1 molecules bind to the PHR2 dimer, the presence of two α1 helices imposes a steric hindrance, which allosterically decouples the PHR2 dimer and stabilizes the SPX1-PHR2 heterodimer25. SPX4 is another member of the SPX domain-containing protein family that plays a distinct role in Pi homeostasis in plants. Unlike SPX1 and SPX2, which are primarily in the nucleus, SPX4 is localized in the cytoplasm, where it interacts with PHR2, preventing its translocation into the nucleus when Pi levels are sufficient26. However, under Pi deficiency conditions, two E3 ubiquitin ligases, SDEL1 and SDEL2, directly ubiquitinate SPX4 to regulate its degradation via the 26S proteasome, resulting in the release of PHR227. SPX4 also interacts with the nitrate sensor NRT1.1B, and this interaction promotes SPX4 ubiquitination and degradation by NRT1.1B-associated E3 ubiquitin ligase NBIP128. Given the critical roles of SPX proteins in Pi homeostasis, understanding the regulatory mechanisms controlling their expression is essential. One such mechanism is alternative polyadenylation (APA), which modulates gene expression by generating mRNA isoforms.
APA is a widespread regulatory mechanism in eukaryotic gene expression that involves the selection of different polyadenylation [poly(A)] sites on a pre-mRNA molecule to generate mRNA isoforms with varying 3′ termini29,30. Most APA sites are found within the 3′UTRs, which affects post-transcriptional gene regulation by modulating mRNA stability, translation, and nuclear export31,32,33. However, when APA sites are located in introns or coding exons, they can alter the coding sequence of the gene34. Studies have shown that APA occurs in over 70% of genes in Arabidopsis thaliana and rice35,36, indicating its importance in regulating gene expression in plants. The choice of poly(A) sites is influenced by various factors, including developmental cues, environmental stimuli, and the expression levels of polyadenylation factors. APA in plants is a dynamic process that contributes significantly to the diversity and complexity of the transcriptome, allowing plants to fine-tune gene expression in response to various internal and external cues37,38,39. While APA significantly contributes to transcript diversity and gene regulation, the factors influencing APA site selection under nutrient stress conditions like Pi starvation are not fully understood. Emerging evidence suggests that noncoding RNAs, such as cis-natural antisense transcripts (cis-NATs), play important roles in regulating gene expression. However, their potential involvement in modulating APA of their cognate genes remains unexplored. cis-NATs are a class of long noncoding RNAs that overlap with protein-coding genes, typically transcribed from the opposite DNA strand. They are widespread in eukaryotes and play important roles in regulating gene expression at both transcriptional and post-transcriptional levels40. Studies have shown that cis-NATs can impact mRNA stability, splicing, and translation, which are crucial for modulating plant responses under nutrient-deprivation conditions, including Pi starvation41. A cis-NAT associated with the Pi exporter gene Phosphate 1;2 (PHO1;2) enhances mRNA translation without altering the steady-state mRNA level, leading to increased Pi transporter protein accumulation in rice under Pi deficiency42. Similarly, cis-NATs that showed positive or negative correlations with the translation efficiency of their cognate sense mRNA under Pi starvation were also identified in Arabidopsis40.
In tomato, there are five SPX domain proteins: SlSPX1 to SlSPX5. Previous studies have demonstrated that SlSPX1 and SlSPX2 interact with several SlPHRs to regulate Pi homeostasis and arbuscular mycorrhizal symbiosis43,44. However, the functions and regulatory mechanisms of other SlSPX proteins remain largely unexplored. In this study, we used poly(A) tag sequencing (PAT-seq) to investigate the APA profile in response to Pi starvation in tomato, focusing on a putative Pi sensor gene SlSPX5, which undergoes APA. We demonstrate that SlSPX5 interacts with SlPHL1 in the cytosol, preventing its translocation into the nucleus. Moreover, we show that SlPHL1 does not form a dimer and that the interaction between SlSPX5 and SlPHL1 is independent of Pi availability. Finally, we reveal that a cis-NAT is induced by Pi starvation at the SlSPX5 locus, promoting proximal polyadenylation of SlSPX5.
Results
Identification and profiling of tomato poly(A) sites regulated by Pi-starvation
To study the impact of Pi starvation on RNA polyadenylation in tomato, pre-germinated tomato (Ailsa Craig) seedlings were grown under Pi-replete or Pi-depleted conditions for 7 days. Under Pi-depleted conditions, the tomato seedlings accumulated significant amounts of anthocyanin, accompanied by the upregulation of genes involved in anthocyanin biosynthesis (Supplementary Fig. 1a–c). Additionally, Pi content in both the roots and shoots of the 7-day-old seedlings were significantly reduced under Pi-depleted conditions (Supplementary Fig. 1d). The expression of PSI genes was also upregulated under Pi deficient conditions (Supplementary Fig. 1e). Poly(A) tag sequencing (PAT-seq) is an efficient method for analyzing poly(A) site usage, mature transcripts abundance, and functional gene expression on a genome-wide scale45. Therefore, we used PAT-seq to study poly(A) site usage of the tomato samples described above (Supplementary Fig. 2). A summary of the raw reads and mapped poly(A) tags (PATs) for each library can be found in Supplementary Fig. 3a. To ensure high-quality data, we filtered out reads with a quality score below Q20. Using a Perl script45,46 (http://www.bmibig.cn/plantAPAdb/about.php#help=pipeline), we removed sequences lacking a poly(T) tail or those with short or low-quality poly(T) to obtain valid poly(T) sequences. These reads were then mapped to the tomato genome (version Sl3.0). The principal component analysis (PCA) and hierarchical clustering of the samples are shown in Supplementary Fig. 3b.
The mapped reads were counted, and PATs at the same position were merged to obtain effective PATs. About 80% of the PATs were located in 3′UTRs, while 5–15% were found in intergenic regions (excluding the promoter region), and 5–15% were located within introns. The remaining PATs were located in promoter regions. Poly(A) sites within 24 bases of each other were grouped into poly(A) clusters (PACs). To ensure reliable results, we considered only PACs with more than 11 PATs for further analysis. In total, we identified 64173 PACs across 20903 genes, of which 14633 genes (70%) were classified as APA genes, defined as those using two or more poly(A) sites. The number and distribution of PACs for each sample are shown in Fig. 1a, b. Over 60% of the PACs were found in 3′UTRs, 10% in CDSs, 10% in intergenic regions, and 10-15% in introns. The genome-wide proportion of APA genes did not differ significantly between Pi-replete and Pi-depleted conditions (Fig. 1c). To further explore the role of APA genes in regulatory processes under varying Pi concentrations, we calculated the poly(A) usage rate as the ratio of the expression level of a specific transcript to the expression level of all transcripts from the same gene. A cumulative distribution function was then plotted to show the poly(A) site usage ratio (Fig. 1d). The starting point of the cumulative curve for plants grown under Pi-replete conditions is around 14%, while for plants under Pi-depleted conditions, it was around 11%. These findings suggest that there is significant difference in the cumulative poly(A) site profiles between Pi-replete and –depleted conditions.
a Tomato genes grouped according to their number of Poly(A) sites. The X-axis indicates the number of Poly(A) sites. The Y-axis shows the number of genes belonging to each group. b Distribution of PACs among the tomato genome in each sample. The color legend at the bottom indicates the different locations of the PACs among the genome: 3′UTR, 5′UTR, CDS, intergenic and introns. The columns on the X-axis indicate the different samples. The Y-axis shows the percent scale for each location. c Genome-wide percentages of APA and non-APA genes in samples grown under Pi-replete (P+) conditions compared to samples grown under Pi-depleted (Pi-) conditions. The column on the right represents the proportions of APA and non-APA genes among the differentially expressed PAC genes (DE-PAC gene) between the two conditions. d Cumulative distribution function (CDF) curve indicating the Poly(A) site usage ratio between samples grown under Pi-replete (P+) and -depleted (P-) conditions. The X-axis shows the log values for the Poly(A) site usage ratio (PAU). The Y-axis shows the values for the CDF. e Differential expression analysis between samples grown under Pi-replete or Pi-depleted conditions. Green dots indicate the down-regulated (P+ vs P-) PACs, whereas red dots show the up-regulated PACs. f Differential 3′UTR length analysis between samples grown under Pi-replete or Pi-depleted conditions. The X-axis shows the strength of 3′UTR shortening (r < 0) or lengthening (r > 0). Green dots indicate genes with a shorter 3′UTR region under Pi-replete conditions than under Pi-depleted conditions, whereas red dots show genes with a longer 3′UTR region under Pi-replete conditions than under Pi-depleted conditions. Wald test of DEseq2 was used for differential gene expression analysis between P+ vs P- conditions. Padj, Benjamini-Hochberg adjusted p value for multiple test correction; Padj < 0.05 was used as significance threshold for false discovery rate.
Differential poly(A) cluster analyses and GO annotation
We used the DESeq2 package to standardize the number of PATs in each PAC and analyse the differences between conditions, aiming to explore the effect of APA on regulatory processes under varying Pi concentrations. Differentially expressed PACs with an adjusted P < 0.05 were considered significant and labeled as DE-PACs, while the corresponding genes are referred to as DE-PAC genes. DE-PAC genes containing more than two DE-PACs were classified as DE-APA genes. To analyse differential gene expression, all PACs were aggregated into genes, and differential expression analysis was performed. The differential analysis between Pi-replete and Pi-depleted conditions revealed a total of 8651 DE-PACs corresponding to 6328 DE-PAC genes, including 4380 up-regulated (Pi-replete vs Pi-depleted) PACs (across 3432 genes) and 4271 down-regulated PACs (across 2963 genes) (Fig. 1e). Among these DE-PAC genes, 5080 used two or more poly(A) sites, classifying them as DE-APA genes and accounting for 80.3% of the total DE-PAC genes. Of these DE-APA genes, 2694 were up-regulated and 2453 were down-regulated, with 67 DE-APA genes found in both up-regulated and down-regulated groups, indicating that these genes have different PAC expression levels under the two conditions. This indicated that these genes utilize different poly(A) sites under Pi-replete and Pi-depleted conditions, suggesting a dynamic regulation of poly(A) site selection based on Pi availability. Additionally, we compared the global poly(A) site usage between Pi-replete and Pi-depleted conditions (Fig. 1f). The results indicated that 160 genes present a shorter 3′UTR length under Pi-replete condition than under Pi-depleted condition, while 563 genes produced a longer 3′UTR under Pi-replete condition compared to Pi-depleted condition. This observation suggests that under Pi-depleted conditions, most genes tend to produce shorter 3′UTR regions, whereas under Pi-replete conditions, most genes preferentially use the distal poly(A) sites (Fig. 1f). In total, 5718 DE-PAC genes were responsive to variations in Pi levels, yielding 60 significantly enriched GO terms (Supplementary Fig. 4), predominantly associated with cellular processes, metabolic processes, and catalytic activities. Among the 4545 DE-APA genes responsive to Pi starvation, 56 GO terms were significantly enriched (Supplementary Fig. 4), relating to cellular processes, metabolic processes, organic substance metabolism, primary metabolic processes, and catalytic activity.
To gauge how these global trends translate to individual pathways, we inspected the genes with the largest APA shifts (Supplementary Table 1). Extreme distal usage was typified by the PHO1-like transporter Solyc02g088230 (log2FC = –44.0), whereas phosphoethanolamine N-methyl-transferase Solyc12g040790 showed the greatest proximal shift (+29.4). Secondary-metabolism genes, particularly cytochrome P450s (–6 to –42), tended toward longer 3′ UTRs, while primary-metabolism enzymes such as coproporphyrinogen III oxidase favored 3′-UTR shortening (+6.0). Cell-wall remodelers like xyloglucan endotransglucosylase/hydrolase Solyc09g008320 also shifted strongly to proximal sites (+34.3), whereas lipid-transfer proteins (e.g., Solyc10g075150, –35.0) shifted distally. Among the 67 genes previously mentioned as exhibiting antagonistic regulation of different poly(A) sites, the metal-ion-binding protein Solyc01g068590 showed a +1.21 shift at its proximal site but –22.07 at its distal site, and phosphatidate phosphatase Solyc10g084540 shifted +6.19 versus –8.23. Such multi-site switching underscores APA as a finely tuned, pathway-specific layer of post-transcriptional control during Pi starvation.
Our PAT-seq analyses enabled the identification of several genes displaying APA phenotypes that are regulated by external levels of Pi. Some of these genes exhibited a preference for using proximal poly(A) sites, demonstrating that differential poly(A) site usage impacts gene expression in response to low Pi stress. Among the different Pi-regulated APA genes, we selected SlSPX5 for further investigation, as SPX domain-containing proteins are putative sensors involved in Pi homeostasis17.
SlSPX5 regulates Pi homeostasis
SlSPX5 (Solyc02g088210) encodes an SPX domain protein. Multiple sequence alignment of SPX proteins in tomato, Arabidopsis and rice showed that SlSPX5 is closely related to Arabidopsis AtSPX4 and rice OsSPX4 (Supplementary Fig. 5). To investigate whether SlSPX5 is localized in the cytosol, like AtSPX4 and OsSPX4, we transformed tomato protoplasts with a SlSPX5-RFP fusion gene construct. The results showed cytosolic localization of SlSPX5, which was further confirmed by transient transformation of SlSPX5-GFP into tobacco leaves (Fig. 2a). To functionally characterize SlSPX5, we generated CRISPR-Cas9-mediated SlSPX5 knock-out mutants. In slspx5-1 mutant, a single nucleotide “T” insertion in the first exon (132 nucleotides downstream of the start codon), resulted in a frameshift, leading to a premature stop codon. This mutation not only resulted in a truncated SlSPX5 protein, but also decreased the transcript levels of SlSPX5 (Supplementary Fig. 6a). This effect is likely caused by nonsense-mediated mRNA decay triggered by the premature stop codon. In the slspx5-2 mutant, a 124 bp deletion was identified (Fig. 2b). The mutants were grown under Pi-replete or Pi-depleted conditions for 7 days. Under Pi-replete conditions, mutant seedlings exhibited a significant increase in Pi content compared to the wild-type (WT), whereas under Pi-depleted condition, the Pi content in both mutants and WT seedlings was comparable (Fig. 2c). Consistent with the observed increase in Pi content, the high-affinity Pi transporter genes SlPHT1, SlPHT3 and SlPHT5 were constitutively induced in the slspx5 mutants under Pi-replete condition (Fig. 2d, Supplementary Fig. 6b). The relative expression of other PSI genes, such as SlSPX1 and SlPAP10 were also significantly upregulated under Pi-replete conditions (Fig. 2d). However, since the increased Pi can be stored in the vacuole and is not detrimental to plant growth, slspx5 mutants displayed mild phenotypes, if any, such as anthocyanin accumulation, increased biomass, and increased root length (Supplementary Fig. 6c–f). Our findings suggest that SlSPX5 plays a role in regulating PSI gene expression and maintaining Pi homeostasis.
a Subcellular localization of SlSPX5. The RFP-fused SlSPX5 gene driven by the 35S promoter was transformed into tomato protoplasts by transient transformation to observe its localization (upper panel, scale bars = 5 µm). The GFP-fused SlSPX5 driven by its native promoter was transiently expressed in tobacco by Agrobacterium-mediated transformation to observe its localization (lower panel, scale bars = 25 µm). b Information on slspx5 mutants. The upper panel shows the guide RNA targeting position in the first exon of SlSPX5, 132 bp downstream of the ATG start site, and the lower panel shows the two mutant forms of the SlSPX5. c Pi content in shoots of WT and slspx5 mutants grown under Pi-replete (P+) and -depleted (P-) conditions. d RT-qPCR analysis for the relative expression of different PSI genes in WT, slspx5-1 and slspx5-2 mutant lines grown under P+ conditions. In (c, d), data are represented as mean ± SD, with n = 3 biologically independent replicates. Statisticallly significant differences between WT and mutants were calculated using a two-sided unpaired Student’s t-test and are indicated by *(P < 0.05), **(P < 0.01) and ***(P < 0.001). Source data are provided as a Source Data file.
SlSPX5 physically interacts with SlPHL1 independently of the Pi levels
To investigate whether SlSPX5 interacts with PHR protein(s) in tomato, we performed a yeast two-hybrid assay to test the interaction between SlSPX5 and three PHR proteins: SlPHR1, SlPHR2 and SlPHL1. The N terminus transactivation domain of SlPHL1 was removed to avoid self-activation in yeast19. The assay results showed that SlSPX5 specifically interacts with SlPHL1, but not with SlPHR1 or SlPHR2 (Fig. 3a). This interaction between SlSPX5 and SlPHL1 was further validated using luciferase complementary imaging (LCI) in tobacco leaves, where co-expression of SlSPX5 and SlPHL1produced a strong luminescence signal as a consequence of the interaction (Fig. 3b). Moreover, bimolecular fluorescence complementation (BiFC) confirmed that co-expression of SlPHL1-nYFP and SlSPX5-cYFP in tomato protoplasts and tobacco leaves resulted in a fluorescence signal in the cytoplasm, indicating that SlSPX5 interacts with SlPHL1 in the cytoplasm (Fig. 3c, Supplementary Fig. 7).
a Yeast two-hybrid assays for the interaction between SlSPX5 and the C-terminus of SlPHR1 (SlPHR1-C), of SlPHR2 (SlPHR2-C), and of SlPHL1 (SlPHL1-C). SlSPX5 was fused to the GAL4 activation domain (AD-SlSPX5), and SlPHR1-C/SlPHR2-C/SlPHL1-C fused to the GAL4 binding domain (SlPHR1/SlPHR2/SlPHL1-C-BD). Yeast cells co-transformed with both AD-SlSPX5 and BD-SlPHR1/SlPHR2/SlPHL constructs were grown on selective media (the third row in each panel). Co-expression of AD and BD, AD and SlPHR1/SlPHR2/SlPHL1, and AD-SlSPX5 and BD were used as negative controls. The co-expression of AD-AtSPX1 and BD-AtPHR1 was used as the positive control. b Examination of protein interactions between SlSPX5 and SlPHL1 by split luciferase complementation assays in tobacco (Nicotiana benthamiana) leaves. Luciferase activities were detected 48 h after infiltration. The co-transformations of cLUC/nLUC-SlSPX5, SlPHL1-cLUC/nLUC and cLUC/nLUC were used as negative controls. The pseudo color bar indicates the range of luminescence intensity. c BiFC analysis for in vivo interaction between SlSPX5 and SlPHL1 in tomato protoplasts. The N- and C-terminal fragments of GFP (nGFP and cGFP) were fused to the C-terminus of SlPHL1 and to SlSPX5, respectively. Combinations of the cGFP or nGFP with corresponding the SlPHL1-nGFP or SlSPX5-cGFP constructs were used as negative controls. Bars = 5 μm. Co-immunoprecipitation (Co-IP) assay for the interaction between SlSPX5 and SlPHL1 under Pi-replete (d) and –depleted (e) conditions. The SlSPX5-FLAG and SlPHL1-GFP proteins were expressed in tomato leaves, and the GFP protein was used as a negative control. f In vitro pull-down assay for studying the interaction between SlSPX5 and SlPHL1. SlPHL1-GST, SlSPX5-His, and GST were expressed and purified in E. coli and subjected to GST pull-down assays. Source data are provided as a Source Data file.
In Arabidopsis and rice, SPX1 and SPX2 interact with PHR1 (PHR2 in rice) in a Pi-dependent manner18,19. To determine whether the interaction between SlSPX5 and SlPHL1 is also Pi-dependent, we performed coimmunoprecipitation (co-IP) assays using tomato leaves transiently expressing SlSPX5-FLAG together with SlPHL1-GFP or GFP control under Pi-replete and -depleted conditions. The results revealed that SlSPX5-FLAG co-immunoprecipitated with SlPHL1-GFP but not with the GFP control under both Pi-replete (Fig. 3d) and Pi-depleted (Fig. 3e) conditions. To verify this independence, we performed an in vitro pull-down assay, and the results indicated that the His-SlSPX5 fusion protein successfully bound to and interacted with the GST-tagged SlPHL1 in the absence of inositol polyphosphate (Fig. 3f). These results suggest that SlSPX5 interacts with SlPHL1 in the cytoplasm in a Pi-independent manner.
Our previous work indicated that the binding of inositol polyphosphate to SPX1 acts allosterically to decouple the OsPHR2 dimer under Pi-replete condition25. To test whether SlPHL1 forms a dimer, we performed a BiFC analysis in tobacco leaves. When we co-expressed two SlPHR1 (SlPHR1-nGFP and SlPHR1-cGFP) together and two SlPHR2 (SlPHR2-nGFP and SlPHR2-cGFP) together, GFP fluorescence was observed (Supplementary Fig. 8a, b). However, no fluorescence was detected when SlPHL1-nGFP and SlPHL1-cGFP were co-expressed (Supplementary Fig. 8c). We also co-expressed nGFP-SlPHL1 and SlPHL1-cGFP together, and did not detect GFP fluorescence, indicating that SlPHL1 proteins do not form homodimers (Supplementary Fig. 8d). To confirm these results, a split-luciferase assay was conducted to test for dimerization. As a control, SlPHL1 was co-expressed with SlSPX5, producing a strong fluorescence signal due to their interaction. However, no signal was observed when SlPHL1-cLUC and nLUC-SlPHL1 were co-transfected into tobacco leaves (Supplementary Fig. 8e). Furthermore, we performed co-IP assays using tobacco leaves transiently expressing SlPHL1-FLAG together with SlPHL1-GFP. The results revealed that SlPHL1-FLAG could not co-immunoprecipitate with SlPHL1-GFP (Supplementary Fig. 8f). These results confirm that SlPHL1 exists as a monomer and that Pi is not essential for its interaction with SlSPX5.
SlSPX5 prevents SlPHL1 from entering the nucleus to activate gene expression
SlSPX5 interacts with SlPHL1 in the cytoplasm. When SlPHL1-GFP was transformed into tomato protoplasts, GFP fluorescence signal was detected in both the nucleus and cytoplasm (Fig. 4a). However, when SlSPX5-FLAG was overexpressed with SlPHL1-GFP, the GFP signal was predominantly found in the cytoplasm under both Pi-replete and—deficient conditions (Fig. 4a), suggesting that SlSPX5 retains SlPHL1 in the cytoplasm. As a PHR family transcription factor, SlPHL1 can bind to the P1BS sequence (GNATATNC), and this binding was competitively blocked by an unlabeled probe (Fig. 4b). To verify that the SlSPX5-SlPHL1 interaction inhibited the induction of PSI genes, we performed a chimeric trans-activation assay using atphr1/atphl1 Arabidopsis mutant protoplasts. The results showed that SlPHL1 could activate promoters of TPSI1 and PHT1 genes, and overexpression of SlSPX5 significantly reduced the transcriptional activity of SlPHL1, indicating that the restriction of SlPHL1 movement from cytoplasm to nucleus represses PSI gene expression (Fig. 4c, Supplementary Fig. 9).
a Transient expression of 35S::SlPHL1-GFP and p35S::SlSPX5-RFP constructs in tomato protoplasts. Protoplasts were isolated from 14-day-old tomato seedlings grown under P+ or P− conditions. Scale bars = 2 µm. b Electrophoretic mobility shift assay of SlPHL1 binding to the P1BS sequence. c Chimeric trans-activation assay demonstrated the transcriptional activation of SlTPSI1 induced by SlPHL1. The promoter of the PSI gene SlTPSI1 was fused to the LUC gene, and the LUC activity was induced by SlPHL1 in atphr1/atphl1 protoplasts. Data are represented as mean ± SD, with n = 3 biologically independent replicates. Statisticallly significant differences were calculated using a two-sided unpaired Student’s t-test and are indicated by ***(P < 0.001). Source data are provided as a Source Data file.
SlSPX5 is alternatively polyadenylated in response to Pi starvation
Since SlSPX5 interacts constitutively with SlPHL1, its reduction is necessary to relieve the inhibition on SlPHL1 during Pi deficiency. However, previous studies have reported that SlSPX5 is dramatically induced by Pi starvation44,47. To resolve this apparent paradox, we examined SlSPX5 transcript levels in 7-day old tomato seedlings grown under Pi-replete or Pi-depleted conditions using RT-qPCR. We used primers targeting either the 3′ or 5′ end of the SlSPX5 transcript. Interestingly, when primers targeted the 3′ end, SlSPX5 expression was repressed under Pi starvation. In contrast, when primers targeted the 5′ end, SlSPX5 expression under Pi starvation was much higher than under Pi-replete condition (Fig. 5a). This result suggested that SlSPX5 undergoes APA in response to Pi starvation. To investigate further, we conducted a 3′RACE experiment, which revealed that SlSPX5 was expressed as a long transcript corresponding to the full-length mRNA under Pi-replete condition. However, under Pi-depleted conditions, the level of the long transcript dramatically decreased, and even several shorter transcripts increased (Fig. 5b). Our PAT-seq analysis confirmed that SlSPX5 undergoes APA in response to Pi starvation. Under Pi-replete conditions, SlSPX5 transcripts were polyadenylated at the distal 3′ end, encoding the full-length SPX protein. However, under Pi starvation, most of the transcripts were polyadenylated within the first exon (Fig. 5c) indicating that Pi starvation promotes SlSPX5 mRNA proximal polyadenylation, thereby suppressing the production of the functional SlSPX5 protein.
a Quantitative PCR analysis of SlSPX5 transcript using primers on the 3′ (left) or 5′ end (right). SlEF1α was used as the reference gene. Data are represented as mean ± SD, with n = 3 biologically independent replicates. Statisticallly significant differences between P+ and P- were calculated using a two-sided unpaired Student’s t-test and are indicated by ***(P < 0.001). Source data are provided as a Source Data file. b 3′RACE assay was used to analyze the abundance of SlSPX5 transcripts of different lengths under P+/P- conditions. The gel image shows the RACE PCR products. c Regulation of SlSPX5 poly(A) site usage by Pi starvation. Integrative Genomics Viewer (IGV) screenshot of PAT-seq data showing the distinct polyadenylation patterns of SlSPX5 under Pi-replete and Pi-depleted conditions.
A cis-NAT is induced by Pi starvation
To determine how Pi starvation regulates the APA of SlSPX5, we re-analyzed our RNA-seq data47. The analysis revealed some unusual transcripts spanning the first exon and first intron (Supplementary Fig. 10a). Using antisense transcript-specific RT-qPCR, we identified a cis-natural antisense transcript (cis-NAT) induced by Pi starvation, which we named NATSPX5 (Fig. 6a, Supplementary Fig. 10b).
a Quantitative PCR analysis of SlSPX5 full-length transcript and anti-sense transcript NATSPX5 in wild type (WT) grown under Pi-replete (P+) and –depleted (P-) conditions. After reverse transcription with gene specific primers, RT-qPCR was used to analyze the gene expression of SlSPX5 (left) and NATSPX5 (right). b Information on natspx5 mutants. CRISPR vectors containing two sgRNAs were constructed, with the upper panel showing the position of three sgRNAs designed for the first intron of SlSPX5, and the lower panel showing the two mutant forms of NATSPX5. c RT-qPCR analysis of SlSPX5 and NATSPX5 gene expression in WT and the two natspx5 mutants. d 3′RACE assay was used to analyze the abundance of SlSPX5 transcripts in WT, natspx5-1 and natspx5-2 under P+ and P- conditions. The gel image shows the RACE PCR products. e RT-qPCR analysis of PSI gene expression in WT and the two natspx5 mutants. In (a, c, e), data are presented as mean ± SD, with n = 3 biologically independent replicates. Statisticallly significant differences between WT and mutants were calculated using a two-sided unpaired Student’s t-test and are indicated by **(P < 0.01) and ***(P < 0.001). f Pi content quantifications in WT and the two natspx5 mutants under P+ and P- conditions. Data are represented as mean ± SD, with n = 6 biologically independent replicates. Statistical significance was assessed using a two-sided unpaired Student’s t-test. ns not significant. Source data are provided as a Source Data file.
The promoter region of the NATSPX5 gene contains three P1BS sequences, which are potential binding sites for PHR transcription factors (Supplementary Fig. 10b). We performed a chimeric trans-activation assay, and the results showed that SlPHL1 could activate the NATSPX5 promoter, and this activation can also be repressed by SlSPX5 (Supplementary Fig. 10b). Consistent with the trans-activation assay results, the induction of NATSPX5 transcript by Pi starvation was partially repressed in both slphl1 and slphr2 mutants (Supplementary Fig. 10c). These results indicated that PHR proteins bind to the intron of SlSPX5 and activate the expression of a cis-NAT under Pi starvation.
The cis-NAT regulates alternative polyadenylation of SlSPX5
Since NATSPX5 overlaps with the proximal poly(A) site of SlSPX5 under P- conditions (Fig. 5c), we speculated that it impacts the APA of SlSPX5. To this end, we employed CRISPR-mediated genome editing to delete the sequence containing the P1BS site in the first intron of SlSPX5 (Fig. 6b). Our results revealed that the deletion of this promoter region significantly impaired the induction of the NATSPX5 transcript upon Pi starvation (Fig. 6c). Interestingly, we observed that the expression levels of the full-length SlSPX5 transcript in the lines containing the deletion were not significantly down-regulated compared to those under Pi-replete condition, but rather showed a trend of induction by Pi deficiency (Fig. 6c). Consistent with the RT-qPCR results, the 3′RACE experiment showed that SlSPX5 was expressed as a long transcript corresponding to the full-length mRNA under both Pi-replete and Pi-depleted conditions in the natspx5 mutants (Fig. 6d). These results suggest that NATSPX5 plays a role in regulating the expression of the full-length SlSPX5 transcript.
In addition, we examined the expression of PSI genes in the mutant with the deletion in the NATSPX5 promoter and found that PSI genes such as SPX1, PAP10, SPX15 and TPSI1 were significantly down-regulated in the natspx5 mutants compared with the WT under Pi-depleted conditions (Fig. 6e). This suggests that the inhibitory effect of SlSPX5 on SlPHL1 remains intact in the absence of NATSPX5, thus repressing the activation of PSI genes under Pi starvation. However, the mutations did not affect the Pi content under P+ and P- conditions compared to the WT (Fig. 6f). No clear phenotypes were observed in natspx5 mutants, as PHL1 plays a partially redundant role with SlPHR1 and SlPHR2.
The transcription of cis-NAT stalls Pol II and recruits the polyadenylation complex
Given that antisense transcripts often silence sense transcripts by activating the siRNA pathway48, we examined the presence of siRNAs in WT plants grown in Pi-replete and -depleted conditions using 12 different oligonucleotide probes complementary to either the sense or antisense transcript in the overlapped region. However, we found no evidence of siRNA induction at this locus under Pi-depleted condition (Supplementary Fig. 10d). Consistent with this result, the SlSPX5 sense transcript was only shortened but not decreased in overall abundance (Fig. 6a). Our findings indicated that antisense transcription may interfere with sense transcript processing, rather than silencing it through the siRNA pathway. It has been reported that Pol II pausing promotes the Ser2 phosphorylation (Ser2p) at the C-terminal domain (CTD) of RNA polymerase II (Pol II) by Cdk12, and this modification recruits the CPA factor CstF77 for polyadenylation in human cells49. In Arabidopsis, Pol II with Ser2p often pauses downstream of the polyadenylation site50. However, the regulatory mechanisms remain largely elusive. To determine whether SlSPX5 polyadenylation is regulated by Pol II pausing, we performed ChIP-qPCR to examine the genomic enrichment of this well-characterized modification of Pol II under nutrient-rich and Pi-depleted conditions. Based on our PAT-seq analysis, we designed primer sequences flanking the Poly(A) site (Fig. 7a). Our results showed increased Pol II Ser2p enrichment downstream of the poly(A) site under Pi-depleted conditions compared to Pi-replete conditions. In the NATSPX5-induced defective mutants, the enrichment of Ser2p under Pi-replete and -depleted conditions was similar to that of the WT under control conditions. These results suggest that Pi starvation-induced natural antisense transcription temporarily blocks SlSPX5 sense transcription and Pol II pausing.
a ChIP-qPCR analysis of Pol Ⅱ-CTD-S2P protein enrichment in the different sites of the SPX5 5′UTR region, upstream (1) or downstream (2) of the Poly(A) site, in WT and natspx5 mutant plants under P+/P—conditions. Data are presented as the mean ± SD, with n = 3 biologically independent replicates. Statistical significance was assessed using a two-sided unpaired Student’s t-test. **P < 0.01. b Examination of protein interactions between Pol Ⅱ-CTD/Pol Ⅱ-CTD17A with SlCPSF30 or SlHLP1 by split luciferase complementation assays in tobacco leaves. Luciferase activities were detected 48 h after infiltration. c BiFC analysis for in vivo interaction between Pol Ⅱ-CTD/Pol Ⅱ-CTD17A with SlCPSF30 or SlHLP1. The N-terminal fragment of GFP (nGFP) was fused to Pol Ⅱ-CTD/Pol Ⅱ-CTD17A while the C-terminal fragment of GFP (cGFP) was fused to SlCPSF30 or SlHLP1. Scale bars = 20 µM. d Co-immunoprecipitation (Co-IP) assay for the interaction between Pol Ⅱ-CTD, and Pol Ⅱ-CTD17A with SlCPSF30 or SlHLP1. Source data are provided as a Source Data file. e Model for the regulation of SlSPX5 expression by the Pi starvation inducible antisense transcript NATSPX5.
In yeast, Pol II physically and functionally interacts with the cleavage and polyadenylation factors (CPF)51. Although we did not identify the tomato CPF components that directly interact with Pol II in yeast two-hybrid assays, we found that two factors, SlCPSF30 and SlHLP1, could interact with Pol II CTD in split-luciferase and BiFC assays (Supplementary Fig. 11, Fig. 7b, c). Pol II CTD could also co-immunoprecipitate SlHLP1 (Fig. 7d). The interactions between Pol II CTD and CPF factors were compromised by mutation of Ser2 to Ala in the Pol II CTD, indicating that Ser2 phosphorylation is required for the interaction between Pol II and the CPF complex (Fig. 7b, c). These results indicate that cis-NAT transcription stalls Pol II and recruits the polyadenylation complex to regulate SlSPX5 polyadenylation at the proximal site (Fig. 7e).
Dicussion
Genome-wide analysis and dynamic shifts in APA under Pi starvation
Our genome-wide analysis of APA in tomato under phosphate starvation conditions reveals extensive regulation of 3′ end processing in response to these conditions. Similar to studies in Arabidopsis35, rice36 and wheat38, we found that approximately 70% of tomato genes exhibit APA, with a significant portion showing differential poly(A) site usage under Pi starvation. This high prevalence of APA suggests it serves as an important regulatory mechanism for modulating gene expression in response to nutrient availability. A key finding from our study is the shift in poly(A) site usage under Pi starvation conditions. We observed that under Pi deficiency, genes tend to express multiple distinct transcripts with different poly(A) sites, while under Pi-replete conditions, fewer transcript isoforms are expressed but at higher levels. This pattern differs slightly from findings in rice, where tissue-specific APA showed more defined patterns of proximal versus distal poly(A) site usage36. The dynamic regulation we observe may reflect a strategy for expanding transcriptome complexity under stress conditions, potentially allowing for more nuanced responses to Pi limitation.
Extensive remodeling of polyadenylation landscape under Pi deficiency
We identified over 8600 differentially expressed PACs corresponding to more than 6300 genes, with about 80% of these classified as DE-APA genes. This extensive remodeling of the polyadenylation landscape under Pi starvation is reminiscent of what has been observed for other abiotic stresses in plants39. For example, in Arabidopsis, salt stress induces widespread changes in poly(A) site selection through the polyadenylation factor FIP140. The significant enrichment of GO terms related to cellular processes, metabolic processes, and catalytic activities among DE-APA genes suggests that APA-mediated regulation affects core cellular functions during Pi starvation responses. A particularly interesting aspect of our findings is the tendency toward longer 3′UTRs under Pi-replete conditions, with 563 genes showing extended 3′UTRs compared to 160 genes with shortened 3′UTRs. The functional significance of 3′UTR lengthening under Pi starvation may relate to the inclusion of additional regulatory elements that could modulate mRNA stability, localization, or translation efficiency. Previous studies have shown that changes in 3′UTR length can affect the presence of microRNA binding sites or regulatory protein binding sites, thereby impacting post-transcriptional regulation35.
Complex APA regulation provides a multi-layered adaptive response to nutrient deficiency
The differential poly(A) site usage we observed often involved both proximal and distal sites, suggesting complex regulatory mechanisms. Consistent with findings in Arabidopsis, where APA can occur in various genic regions including introns and coding sequences35, our results indicate that Pi starvation-induced APA is not limited to 3′UTRs but occurs throughout gene bodies. This pattern of regulation may provide additional layers of control over gene expression during nutrient stress responses. Our findings also parallel recent work on nitrogen starvation in Arabidopsis, where nutrient deficiency triggers widespread changes in poly(A) site usage52. In line with our observations under Pi starvation, nitrogen limitation promotes the use of non-canonical polyadenylation sites and affects genes involved in metabolic processes and stress responses, suggesting that APA regulation may be a common mechanism in plant responses to different nutrient deficiencies. Interestingly, both nitrogen and phosphate starvation responses involve complex hormone signaling networks—while nitrogen deficiency triggers salicylic acid accumulation to modulate root architecture52, our study reveals that Pi starvation induces significant changes in APA of key regulatory factors. The role of cis-NATs in regulating gene expression during Pi starvation, as demonstrated in rice for PHO1;242, prompted us to examine this aspect in tomato. Among the genes showing differential poly(A) site usage under Pi starvation, our identification of SlSPX5 was particularly significant given that SPX domain-containing proteins are known Pi sensors that interact with PHR transcription factors in other species. This finding not only adds to our understanding of how plants integrate transcriptional and post-transcriptional regulation to fine-tune Pi homeostasis, but also led us to investigate the specific role of SlSPX5 in Pi signaling, ultimately revealing unexpected differences in how tomato regulates Pi responses compared to other well-studied plant models.
Pi-independent interaction of SlPHL1 and SlSPX5
Pi homeostasis is crucial for plant growth and development, and MYB transcription factors, particularly PHRs, play important roles in maintaining Pi homeostasis in plants9,10,12,53. In tomato, 17 putative SlPHR genes have been identified43. Among them, SlPHR3 regulates tomato responses to Pi starvation, including anthocyanin accumulation and increased acid phosphatase activity54,55,56, while SlPHR1 is required to maintain Pi homeostasis as the slphr1 mutant exhibits reduction in Pi content in both shoot and root under Pi deficiency condition43. In Arabidopsis and rice, AtPHR1 and OsPHR2 form homodimers to activate PSI gene expression8,25. SPX domain-containing proteins, such as SPX1 and SPX2, sense the Pi signaling molecule InsP8 and decouple OsPHR2 in the nucleus, inhibiting PHR dimerization and DNA binding to P1BS element in the promoters of PSI genes4125,57. Conversely, under Pi deficiency, decreased levels of InsP8 reduce the SPX-PHR interaction, allowing PHRs to dimerize and activate PSI genes18,19. In this study, however, we demonstrate that SlPHL1 does not form a homodimer and interacts with SlSPX5 in a Pi-independent manner (Supplementary Fig. 9). This contrasts with the Pi-dependent SPX-PHR interactions observed in Arabidopsis and rice25,57, suggesting a unique regulatory mechanism in tomato. Our findings indicate that under Pi deficiency, SlSPX5 must be downregulated to release SlPHL1 from the cytoplasm, allowing it to enter the nucleus and activate PSI gene expression.
Alternative polyadenylation and antisense transcription regulate SlSPX5 during Pi starvation
SlSPX5 is closely related to SPX4 and regulates not only Pi homeostasis but also arbuscular mycorrhiza colonization44 (Supplementary Fig. 5). Although post-translational degradation of SPX4 by SDEL1 and SDEL2 relieves OsPHR2 repression under Pi starvation27, it remains unclear whether SPX4 senses Pi signals and whether this sensing modulates its interaction with OsPHR2 in the cytoplasm preventing its entry into the nucleus26. While previous reports indicated that SlSPX5 is induced by Pi starvation44,47, we found a dramatic decrease in the level of full-length SlSPX5 transcript under Pi deficiency. The increase in transcript levels at this locus comes from the antisense transcript rather than the sense transcript, particularly when Pi is deficient. SlSPX5 harbors a large intron (~3.5 kb) between the first and second exons. PHRs bind the P1BS elements within this intron to activate a cis-NAT. This antisense transcription promotes proximal polyadenylation of SlSPX5, resulting in the suppression of the functional SlSPX5 protein. This regulatory mechanism links antisense RNA to APA and Pi homeostasis in plants.
NAT SPX5-mediated transcriptional regulation through RNA polymerase II pausing and APA
NATs are an important subset of lncRNAs that are widespread across prokaryotic and eukaryotic genomes and regulate their cognate sense genes at transcriptional and/or post-transcriptional level58. Antisense transcripts can regulate transcription, splicing, stability or translation efficiency of the sense transcript, depending on the pattern and degrees of overlap between the sense-antisense pair59. At the transcriptional level, cis-NATs may regulate cognate sense genes through transcriptional interference mechanisms, such as promoter competition, RNA Pol II collision, or promoter occlusion58. In this study, we observed that the induction of cis-NAT NATSPX5 under Pi starvation likely induces Pol II pausing at the 5′ end of SlSPX5, resulting in the phosphorylation on Ser2p at CTD of Pol II (Fig. 7a). The increased Ser2p signal could recruit the cleavage and polyadenylation factor CPF (Fig. 7), facilitating the formation of a functional CPA complex that drives proximal polyadenylation of the shortened SlSPX5 transcript (Fig. 7e). Our findings reveal an important regulatory role for cis-NATs in modulating the polyadenylation of sense transcripts and regulating Pi homeostasis in tomato.
In conclusion, our genome-wide analysis of APA in tomato revealed extensive APA regulation during Pi starvation, with more genes producing shorter 3′UTRs under Pi-depleted conditions. The PAT-seq analysis identified over 14,000 APA genes, with approximately 80% of Pi-responsive genes showing differential poly(A) site usage, highlighting the importance of post-transcriptional regulation in Pi starvation response. Among these, we uncovered a regulatory mechanism where Pi starvation induces a cis-NAT, NATSPX5, which promotes APA of SlSPX5, leading to downregulation of the functional SlSPX5 protein. This process releases SlPHL1 from cytoplasmic retention, allowing it to activate PSI gene expression under Pi-depleted conditions. This mechanism differs from the Pi-dependent SPX-PHR interactions observed in other plants, highlighting the diversity of Pi homeostasis regulation among species. Our study not only reveals the widespread impact of APA in Pi starvation response but also contributes additional insights into the complex network of Pi sensing and signaling in plants, opening avenues for future research on nutrient homeostasis and crop improvement.
Methods
Plant materials and growth condition
All tomato seeds (Ailsa Craig- AC) were germinated in perlite, and seedlings were transferred to Pi-replete or Pi-depleted liquid medium after the emergence of true leaves. The seedlings were grown at a temperature of 25/19 °C (16 h /8 h) day/night cycle and a relative humidity of 60–65%. For all experiments, AC was used as wild-type (WT). The slspx5, slphl1, natspx5 mutants were generated by CRISPR/Cas9-mediated mutagenesis in the WT background. For over-expression lines of SlSPX5 and NATSPX5 genes, the genomic cDNA was cloned into binary vectors and driven by the 35S promoter. For phosphate treatment, seedlings were simultaneously grown under control condition (Pi-replete condition with 0.25 mM KH2PO4) and Pi-depleted conditions (substituting 0.125 mM K2SO4 for KH2PO4) for 7 days.
PAT-seq assay and poly(A) site usage analysis
Total RNA was extracted from 7-day-old seedlings using the RNeasy Plant Mini Kit (74904; Qiagen). PAT-seq was performed as described60. Briefly, 2 µg of total RNA was fragmented at 94 °C for 4 min in 5× first-strand buffer (Invitrogen). RNA fragments containing poly(A) tails were enriched using oligo(dT)25 magnetic beads (New England Biolabs). Reverse transcription was performed with barcoded oligo(dT)18 primers using SMARTSCRIBE enzyme (Clontech) for 2 h, followed by addition of the 5′ adapter for template switching. The generated cDNA was purified, followed by 18 cycles of PCR. The PCR product was electrophoresed on a 2% agarose gel, and fragments of 300–500 bp were purified and sequenced in single-end mode on an Illumina HiSeq 2500 at the core facility of the College of Environment and Ecology, Xiamen University. For PAT-seq data processing, raw reads were pre-treated as previously described60. Poly(A) site usage (PSU) was calculated as the ratio of reads at a specific poly(A) site to the total reads for the corresponding gene. To visualize poly(A) site usage, we used Folded Cumulative Distribution Function from the MOUNTAINPLOT R package for folded cumulative curve plotting, which folds the top half of the cumulative curve over60. The scale of the upslope is shown on the left (from 0 to 0.5), while the downslope is on the right (from 0.5 to 1.0). The top of the folded cumulative curve represents the median of the corresponding dataset. For proximal/distal poly(A) site switching analysis, the top two expressed poly(A) sites were retained for further analysis. The 3′ UTR length was calculated as previously described61, and Padj < 0.05 was considered as the threshold for significant 3′ UTR lengthening or shortening events.
Construction of vectors and generation of transgenic tomato
To generate mutant lines, we inserted target sequences into CRISPR vectors pCAMBIA1300-35S-Cas9. For natspx5 mutants, we designed two target sites, and double mutant vectors were constructed to delete the NATSPX5 promoter region. For the over-expression lines of the SlSPX5 and NATSPX5 genes, the full-length sequences were amplified from cDNA and inserted into pCAMBIA1300-35S-FLAG vectors. Agrobacterium tumefaciens-mediated genetic transformation was performed on somatic cells of AC plants, and transgenic plants were obtained through tissue culture techniques.
RNA extraction, RT-qPCR and RACE
Total RNA was extracted from 7-day-old seedlings using Trizol (Sanon Biotech) reagent and the HiPure Plant RNA Mini Kit (Magen). cDNA synthesis was performed using the Hifair® Ⅲ 1st Strand cDNA Synthesis SuperMix (Yeasen), which contains a mix of random primers and oligo (dT)18 primers for reverse transcription. For gene-specific reverse transcription, the transgene KIT was used. The synthesized cDNA was used in PCR reactions with Hieff® qPCR SYBR Green Master Mix (Yeasen), and RT-qPCR was run on a CFX96 Touch Deep Well Real-Time PCR Detection System (Bio-Rad). Three biological replicates were analyzed. The primers used in this study are listed in table(S). RACE (Rapid Amplification of cDNA Ends) experiments were performed using the 5′/3′-RACE Kit (Sanon Biotech), following the manufacturer’s instructions.
Protein interaction analyses
For the Y2H assays, the C-terminal sequences of SlPHL1, SlPHR1, and SlPHR2 were cloned into the pGBKT7 vectors, and the full-length sequence of SlSPX5 was cloned into the pGADT7 vector to generate AD and BD constructs. The transfected yeasts were cultured in medium lacking Leu and Trp, as well as in media lacking Leu, Trp and His/Ade, and incubated at 28 °C for 3 days. LCI and BiFC were used to demonstrate protein interactions. The full-length coding sequences of SlPHL1, SlPHR1, SlPHR2 and SlSPX5 were cloned into the pDONR221 enter vectors, followed by the insertion of the target sequences into expression vectors pEARLY202-3′-nLUC (2-416aa), pEARLY202-5′-nLUC (1-416aa), pEARLY202-3′-cLUC (298-550aa), pEARLY202-nGFP, and pEARLY202-cGFP, respectively, using Gateway exchange sites. The resulting constructs were introduced into tobacco by transient transformation mediated by Agrobacterium tumefaciens GV3101. LUC fluorescence was captured using a plant in vivo imager (Roper Lumazone PyLoN1300B), and GFP fluorescence signals were observed using a confocal laser scanning microscope (Leica TCS SP8). For the Co-IP experiments, total protein was extracted from the ground tissue, using 2 mL of extraction buffer containing 50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 5 mM EDTA, 0.1% (v/v) Triton X-100, 0.2% (v/v) Nonidet P-40, 5% (v/v) glycerol, and a 1:100 (v/v) dilution of Protease Inhibitor Cocktail (Sigma). After centrifugation, the supernatant was incubated with anti-GFP magnetic beads (Sigma) overnight at 4 °C. The beads was washed five times and mixed with 50 µL of 1× SDS loading buffer and boiled at 98 °C for 10 min to elute bound proteins. The proteins were separated by 10% (w/v) SDS-polyacrylamide gel electrophoresis (SDS-PAGE) and analyzed by immunoblotting using anti-FLAG antibody (Sigma #F1804; 1:10000 dilution) and anti-GFP antibody (Roche, #11814460001, 1:5000 dilution).
Transient expression assays in Arabidopsis protoplasts and chimeric trans-activation assay. Seedlings grown in short-day conditions (8 h light/16 h dark) for approximately 28 days were used for experiments. The lower epidermis of the leaves was peeled off and soaked in enzymolysis solution (1.5% cellulose, 0.3% macerozyme, 0.4 M mannitol, 20 mM MES, 20 mM KCl, 10 mM CaCl2, 5 mM BME, 0.1% BSA) for enzymatic digestion. The leaves were incubated in the dark for 3 h. After enzymatic digestion, protoplasts were collected for plasmid transformation. The transformation included reporter plasmids (pRD29A-proTPSI1-LUC), effector plasmids (pSAT6-35S-SPX5-FLAG, pSAT108-35S-PHL1-eGFP), and internal controls. Protoplast transformation and LUC and GUS activity assays were performed as previously described62.
Subcellular localization assay
atspx4 mutant protoplasts were used for subcellular localization assays. Protoplast cells were separately or co-transformed with 35S-SPX5-eRFP (pSAT108) and 35S-PHL1-eGFP(pSAT108), and fluorescence signals were captured using the confocal laser scanning microscopy (Leica TCS SP8). We performed the same experiment in tobacco leaves by transiently transforming two vectors, pCAMBIA1300-35S-SPX5-FLAG and native promoter-driven pCAMBIA1300-NP-PHL1-GFP. Fluorescence signals were observed using Leica TCS SP8 microscope.
Measurement of Pi concentration
We used seedlings treated in liquid medium for 7 days to measure Pi content. Measurements were performed following the phosphomolybdate colorimetric assay as described63.
Protein expression and purification, pull-down and EMSA
The C-terminal sequence of SlPHL1, containing MYB and CC domains, was amplified from cDNA and inserted into pGEX-4T-GST vector. The full-length SlSPX5 was cloned into the pET-28a(+)-His vector, creating a fusion with a His tag. The plasmids were transformed into E. coli strain BL21(DE3) and Rosetta (DE3). Protein expression was induced with with 0.3 mM IPTG 16 °C overnight. Proteins were purified using GST beads for the GST-tagged proteins and His beads for the His-tagged proteins. For the pull-down assays, GST beads were used to capture the protein complexes, with empty GST protein serving as a control.
Chromatin immunoprecipitation
One gram of Pi-suffcient and Pi-depleted treated seedlings was collected and ground into a fine powder. Chromatin immunoprecipitation (ChIP) was performed following the protocol as described64. For the PolⅡ-CTD-Ser2P- ChIP assays, we used 2 μg of specific antibody: Ser2P-Pol II (Abcam, #ab5095, 1:100 dilution). The antibody was incubated with Dynabeads (Thermo) for 2 h to enrich the protein-chromatin complexes. Purified polymerase complexes were disrupted, and the attached DNA was extracted using Phenol/Chloroform/Isoamyl Alcohol (25:24:1). The extracted DNA was used to assess the amount of enrichment by RT-qPCR. Quantitative analysis was performed on captured DNA by Hieff® qPCR SYBR Green Master Mix (Yeasen). The oligonucleotide sequences can be found in Supplementary Table 2. ChIP enrichments were calculated as the ratio of the IP sample to the product of interest in the corresponding input sample. Error bars represent the average standard error generated from at least three independent replicates.
Statistics and reproducibility
For PAT-seq analysis, three independent biological samples have been analyzed. Statistical analysis was performed using the DESeq2 package with the Wald test (two-sided). Adjusted p values (padj) were calculated using the Benjamini-Hochberg false discovery rate (FDR) method to correct for multiple testing across all poly(A) sites genome-wide. Transcript analyses were conducted using three independent biological samples. For phenotypic analysis, three to six samples were examined. Statistical significance between the two groups was assessed using a two-sided unpaired Student’s t-test. The experiments were repeated independently three times, with similar results. The experimental design incorporated randomization to minimize bias and ensure representative sampling. No data were excluded from the analyses. All analyses were conducted in a blinded manner to prevent experimental bias.
Accession numbers
Gene sequences reported in this article can be found in the Solgenomics databases under the following accession numbers: SlPAP1 (Solyc10g086260), SlF3′H (Solyc02g083860), SlAHRD (Solyc08g080040) SlPHT1 (Solyc09g066410), SlSPX1 (Solyc01g090890), SlSPX15 (Solyc12g009480), SlPAP10 (Solyc03g098010), SlTPSI1 (Solyc10g085850), SlPHR1 (Solyc06g008200), SlPHR2 (Solyc09g072830), SlPHL1 (Solyc05g055940), SlSPX5 (Solyc02g088210), SlHLP1(Solyc02g093340), SlCPSF30(Solyc02g021760), SlRPB1(Solyc02g083350).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The PAT-seq data generated in this study have been deposited in the BioProject database at NCBI under accession number PRJNA1185315. Code and analysis scripts can be found through the following stable website: http://www.bmibig.cn/plantAPAdb/about.php#help=pipeline as well as via Github https://github.com/BMILAB/PAT-seq-pipeline. Source data are provided with this paper.
References
Vance, C. P., Uhde-Stone, C. & Allan, D. L. Phosphorus acquisition and use: critical adaptations by plants for securing a nonrenewable resource. N. Phytol. 157, 423–447 (2003).
Jouhet, J., Maréchal, E. & Block, M. A. Glycerolipid transfer for the building of membranes in plant cells. Prog. Lipid Res. 46, 37–55 (2007).
Raghothama, K. G. Phosphate acquisition. Annu. Rev. Plant Physiol. Plant Mol. Biol. 50, 665–693 (1999).
Holford, I. C. R. Soil phosphorus: its measurement, and its uptake by plants. Soil Res. 35, 227–240 (1997).
López-Bucio, J., Cruz-Ramírez, A. & Herrera-Estrella, L. The role of nutrient availability in regulating root architecture. Curr. Opin. Plant Biol. 6, 280–287 (2003).
Ticconi, C. A. & Abel, S. Short on phosphate: plant surveillance and countermeasures. Trends Plant Sci. 9, 548–555 (2004).
Yuan, H. & Liu, D. Signaling components involved in plant responses to phosphate starvation. J. Integr. Plant Biol. 50, 849–859 (2008).
Bustos, R. et al. A central regulatory system largely controls transcriptional activation and repression responses to phosphate starvation in Arabidopsis. PLoS Genet. 6, e1001102 (2010).
Rubio, V. et al. A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes Dev. 15, 2122–2133 (2001).
Sun, L., Song, L., Zhang, Y., Zheng, Z. & Liu, D. Arabidopsis PHL2 and PHR1 Act redundantly as the key components of the central regulatory system controlling transcriptional responses to phosphate starvation. Plant Physiol. 170, 499–514 (2016).
Ruan, W. et al. Genetic manipulation of a high-affinity PHR1 target cis-element to improve phosphorous uptake in Oryza sativa L. Plant Mol. Biol. 87, 429–440 (2015).
Zhou, J. et al. OsPHR2 is involved in phosphate-starvation signaling and excessive phosphate accumulation in shoots of plants. Plant Physiol. 146, 1673–1686 (2008).
Nilsson, L., Müller, R. & Nielsen, T. H. Increased expression of the MYB-related transcription factor, PHR1, leads to enhanced phosphate uptake in Arabidopsis thaliana. Plant Cell Environ. 30, 1499–1512 (2007).
Shi, J. et al. A phosphate starvation response-centered network regulates mycorrhizal symbiosis. Cell 184, 5527–40.e18 (2021).
Chiou, T. J. & Lin, S. I. Signaling network in sensing phosphate availability in plants. Annu. Rev. Plant Biol. 62, 185–206 (2011).
Secco, D. et al. The emerging importance of the SPX domain-containing proteins in phosphate homeostasis. New Phytol. 193, 842–851 (2012).
Wild, R. et al. Control of eukaryotic phosphate homeostasis by inositol polyphosphate sensor domains. Science 352, 986–990 (2016).
Puga, M. I. et al. SPX1 is a phosphate-dependent inhibitor of Phosphate Starvation Response 1 in Arabidopsis. Proc. Natl. Acad. Sci. USA 111, 14947–14952 (2014).
Wang, Z. et al. Rice SPX1 and SPX2 inhibit phosphate starvation responses through interacting with PHR2 in a phosphate-dependent manner. Proc. Natl. Acad. Sci. USA 111, 14953–14958 (2014).
Dong, J. et al. Inositol pyrophosphate InsP(8) acts as an intracellular phosphate signal in Arabidopsis. Mol. Plant 12, 1463–1473 (2019).
Zhu, J. et al. Two bifunctional inositol pyrophosphate kinases/phosphatases control plant phosphate homeostasis. Elife 8, e43582 (2019).
Li, X. et al. Control of XPR1-dependent cellular phosphate efflux by InsP(8) is an exemplar for functionally-exclusive inositol pyrophosphate signaling. Proc. Natl. Acad. Sci. USA 117, 3568–3574 (2020).
Chabert, V. et al. Inositol pyrophosphate dynamics reveals control of the yeast phosphate starvation program through 1,5-IP(8) and the SPX domain of Pho81. Elife 12, RP87956 (2023).
Yang, S. Y., Lin, W. Y., Hsiao, Y. M. & Chiou, T. J. Milestones in understanding transport, sensing, and signaling of the plant nutrient phosphorus. Plant Cell 36, 1504–1523 (2024).
Zhou, J. et al. Mechanism of phosphate sensing and signaling revealed by rice SPX1-PHR2 complex structure. Nat. Commun. 12, 7040 (2021).
Lv, Q. et al. SPX4 Negatively regulates phosphate signaling and homeostasis through its interaction with PHR2 in rice. Plant Cell 26, 1586–1597 (2014).
Ruan, W. et al. Two RING-finger ubiquitin E3 ligases regulate the degradation of SPX4, an internal phosphate sensor, for phosphate homeostasis and signaling in rice. Mol. Plant 12, 1060–1074 (2019).
Hu, B. et al. Nitrate-NRT1.1B-SPX4 cascade integrates nitrogen and phosphorus signalling networks in plants. Nat. Plants 5, 401–413 (2019).
Tian, B. & Manley, J. L. Alternative polyadenylation of mRNA precursors. Nat. Rev. Mol. Cell Biol. 18, 18–30 (2017).
Lin, J. & Li, Q. Q. Coupling epigenetics and RNA polyadenylation: missing links. Trends Plant Sci. 28, 223–234 (2023).
Nam, J. W. et al. Global analyses of the effect of different cellular contexts on microRNA targeting. Mol. Cell 53, 1031–1043 (2014).
Hoffman, Y. et al. 3’UTR shortening potentiates microRNA-based repression of pro-differentiation genes in proliferating human cells. PLoS Genet. 12, e1005879 (2016).
Neve, J. et al. Subcellular RNA profiling links splicing and nuclear DICER1 to alternative cleavage and polyadenylation. Genome Res. 26, 24–35 (2016).
Yao, P. et al. Coding region polyadenylation generates a truncated tRNA synthetase that counters translation repression. Cell 149, 88–100 (2012).
Wu, X. et al. Genome-wide landscape of polyadenylation in Arabidopsis provides evidence for extensive alternative polyadenylation. Proc. Natl. Acad. Sci. USA 108, 12533–12538 (2011).
Fu, H. et al. Genome-wide dynamics of alternative polyadenylation in rice. Genome Res. 26, 1753–1760 (2016).
Chakrabarti, M., de Lorenzo, L., Abdel-Ghany, S. E., Reddy, A. S. N. & Hunt, A. G. Wide-ranging transcriptome remodelling mediated by alternative polyadenylation in response to abiotic stresses in Sorghum. Plant J. 102, 916–930 (2020). Jun.
Ma, H., Lin, J., Mei, F., Mao, H. & Li, Q. Q. Differential alternative polyadenylation of homoeologous genes of allohexaploid wheat ABD subgenomes during drought stress response. Plant J. 114, 499–518 (2023).
Zhou, J. & Li, Q. Q. Stress responses of plants through transcriptome plasticity by mRNA alternative polyadenylation. Mol. Hortic. 3, 19 (2023).
Deforges, J. et al. Control of cognate sense mRNA translation by cis-natural antisense RNAs. Plant Physiol. 180, 305–322 (2019).
Reis, R. S. et al. An antisense noncoding RNA enhances translation via localized structural rearrangements of its cognate mRNA. Plant Cell 33, 1381–1397 (2021).
Jabnoune, M. et al. A rice cis-natural antisense RNA acts as a translational enhancer for its cognate mRNA and contributes to phosphate homeostasis and plant fitness. Plant Cell 25, 4166–4182 (2013).
Liao, D. et al. SlSPX1-SlPHR complexes mediate the suppression of arbuscular mycorrhizal symbiosis by phosphate repletion in tomato. Plant Cell 34, 4045–4065 (2022).
Singh, N. R. R. et al. Silencing of SlSPX1 and SlSPX2 promote growth and root mycorrhization in tomato (Solanum lycopersicum L.) seedlings. Plant Sci. 333, 111723 (2023).
Lin, J. et al. Alternative polyadenylated mRNAs behave as asynchronous rhythmic transcription in Arabidopsis. RNA Biol. 18, 2594–2604 (2021).
Wu, X., Zhang, Y. & Li, Q. Q. PlantAPA: a portal for visualization and analysis of alternative polyadenylation in plants. Front. Plant Sci. 7, 889 (2016).
Satheesh, V. et al. High transcriptome plasticity drives phosphate starvation responses in tomato. Stress Biol. 2, 18 (2022).
Borsani, O. et al. Endogenous siRNAs derived from a pair of natural cis-antisense transcripts regulate salt tolerance in Arabidopsis. Cell 123, 1279–1291 (2005).
Davidson, L., Muniz, L. & West, S. 3’ end formation of pre-mRNA and phosphorylation of Ser2 on the RNA polymerase II CTD are reciprocally coupled in human cells. Genes Dev. 28, 342–356 (2014).
Zhu, J., Liu, M., Liu, X. & Dong, Z. RNA polymerase II activity revealed by GRO-seq and pNET-seq in Arabidopsis. Nat. Plants 4, 1112–1123 (2018).
Carminati, M. et al. A direct interaction between CPF and RNA Pol II links RNA 3’ end processing to transcription. Mol. Cell 83, 4461–4478 (2023).
Conesa, C. M. et al. Alternative polyadenylation and salicylic acid modulate root responses to low nitrogen availability. Plants 9, 251 (2020).
Wang, J. et al. A phosphate starvation response regulator Ta-PHR1 is involved in phosphate signalling and increases grain yield in wheat. Ann. Bot. 111, 1139–1153 (2013).
Liu, Y. et al. SlPHL1 positively modulates acid phosphatase in response to phosphate starvation by directly activating the genes SlPAP10b and SlPAP15 in tomato. Physiol. Plant 176, e14197 (2024).
Wu, X. et al. SlPHL1 is involved in low phosphate stress promoting anthocyanin biosynthesis by directly upregulation of genes SlF3H, SlF3’H, and SlLDOX in tomato. Plant Physiol. Biochem. 200, 107801 (2023).
Zhang, Y. et al. SlPHL1, a MYB-CC transcription factor identified from tomato, positively regulates the phosphate starvation response. Physiol. Plant 173, 1063–1077 (2021).
Ried, M. K. et al. Inositol pyrophosphates promote the interaction of SPX domains with the coiled-coil motif of PHR transcription factors to regulate plant phosphate homeostasis. Nat. Commun. 12, 384 (2021).
Rosikiewicz, W. & Makałowska, I. Biological functions of natural antisense transcripts. Acta Biochim Pol. 63, 665–673 (2016).
Werner, A., Kanhere, A., Wahlestedt, C. & Mattick, J. S. Natural antisense transcripts as versatile regulators of gene expression. Nat. Rev. Genet. 25, 730–744 (2024).
Yu, Z., Lin, J. & Li, Q. Q. Transcriptome analyses of FY mutants reveal its role in mRNA alternative polyadenylation. Plant Cell 31, 2332–2352 (2019).
Lin, J. et al. HDA6-dependent histone deacetylation regulates mRNA polyadenylation in Arabidopsis. Genome Res. 30, 1407–1417 (2020).
Yoo, S. D., Cho, Y. H. & Sheen, J. Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat. Protoc. 2, 1565–1572 (2007).
Jain, A. et al. Differential effects of sucrose and auxin on localized phosphate deficiency-induced modulation of different traits of root system architecture in Arabidopsis. Plant Physiol. 144, 232–247 (2007).
Chen, X. et al. POWERDRESS interacts with HISTONE DEACETYLASE 9 to promote aging in Arabidopsis. Elife 5, e17214 (2016).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (U24A20385 to M.L., 32270344 to Q.Q.L.), by the Interdisciplinary Research Project of Hangzhou Normal University (2025JCXK01 to M.L.) and by US National Science Foundation (2347540 to Q.Q.L.). We thank Danhui Zhao for helping data submission.
Author information
Authors and Affiliations
Contributions
M.L., V.S., and Q.Q.L. conceived and supervised the project. S.G., J.L., and M.H. performed most of the experiments and analyzed the data with M.L. M.S.B., S.W., Q.L., F.G., J.D., and G.M. assisted in the experiments and discussed the results. M.L., V.S., S.G., and J.L. wrote, and Q.Q.L. revised, the manuscript. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Rodrigo Reis, Hatem Rouached and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ge, S., Li, J., Ma, H. et al. A cis-natural antisense RNA regulates alternative polyadenylation of SlSPX5 under Pi starvation in tomato. Nat Commun 16, 7981 (2025). https://doi.org/10.1038/s41467-025-63406-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63406-1