Abstract
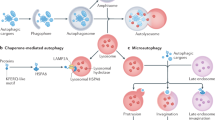
Microautophagy is an intracellular degradation process in which degradatory organelles, such as the lysosome, directly take up substrates by invagination and/or protrusion of their membranes. Here, we provide evidence that Rab32-positive, lysosome-related organelles in macrophages incorporate various other organelles, including endosomes and mitochondria. Our data indicates that, upon exposure to a mitochondria-damaging reagent, mitochondria can be directly engulfed by the lysosome-like organelles independently of macroautophagy or ESCRT machinery. Rab32 GTPase, phosphatidylinositol 3,5-bisphosphates, ubiquitination, and p62/SQSTM1 are crucial for this degradation. Furthermore, the degree of M1 polarization of macrophages, which is facilitated by metabolic reprogramming into increased glycolysis via mitochondrial elimination, is significantly reduced in Rab32/38 double-knockout macrophages. Thus, microautophagy plays a role in the physiological regulation of macrophages.
Similar content being viewed by others
Introduction
Cells maintain proteostasis through several intracellular degradation systems. Macroautophagy involves the formation of double-membrane structures known as autophagosomes, which sequester cytoplasmic components and deliver them to lysosomes for degradation1. Over the past few decades, extensive research has revealed the crucial roles of macroautophagy in various physiological and pathological processes2. In contrast, microautophagy is another intracellular degradation pathway in which degradative organelles—such as the vacuole in yeast and plants—directly engulf cytoplasmic substrates via membrane invagination and/or protrusion3. However, in mammalian cells, lysosomes are generally smaller than other organelles, making them seemingly unsuitable for the degradation of larger structures via microautophagy4,5. Consequently, the specific organelles targeted by microautophagy in mammalian cells have remained largely unclear6.
Macrophages undergo activation, termed M1 polarization, in response to stimuli such as lipopolysaccharides (LPS). Several studies suggest that macroautophagy is involved in this process7,8,9. During M1 polarization, macrophages undergo a metabolic shift characterized by enhanced glycolysis and reduced oxidative phosphorylation10. Furthermore, there is a report indicating that mitochondrial autophagy (mitophagy) is involved in this metabolic reprogramming, though the specific regulatory pathways remain to be fully understood11.
Lysosome-related organelles (LROs) form a distinct class of organelles that share common characteristics with lysosomes, such as the presence of the lysosomal marker Lamp112. They also exhibit cell-type-specific features that differentiate them from conventional lysosomes12. Notable examples of LROs include melanosomes in melanocytes, α-granules in platelets, Weibel-Palade bodies in endothelial cells, and lytic granules in natural killer cells12. In our recent studies, we characterized an LRO marked by the small GTPases Rab32 and Rab38 in mouse bone marrow-derived macrophages (BMDMs) and osteoclasts13,14,15. These LROs share several characteristics with lysosomes, including the expression of the lysosomal membrane protein Lamp1 and acidification via V-ATPase, despite being substantially larger in size (~2000 nm) compared to conventional lysosomes (~100 nm)14.
Here we show that microautophagy mediated invagination plays a central role in mitochondria degradation within these specialized LROs in macrophages.
Results
Rab32-positive lysosome-related organelles degrade the endosome in bone marrow-derived macrophages
Rab32-positive LROs are distinguishable from typical early and late endosomes, as their limiting membranes lack marker proteins of early (Rab5A) and late (Rab7) endosomes14. Unexpectedly, Rab5A and Rab7 signals were occasionally detected within Rab32-positive LROs in BMDMs (Fig. 1A). Correlative light and electron microscopy (CLEM) analysis revealed the presence of Rab5A-associated structures (red arrows) inside Rab32-positive LROs (Supplementary Fig. 1A). Treatment with bafilomycin A1, a V-ATPase inhibitor that neutralizes lysosomal pH and inhibits degradation, resulted in the accumulation of numerous membrane-bound organelles with diverse morphologies—including multivesicular organelles positive for Rab5A (red arrows) and invaginating structures (yellow and white arrows)—inside Rab32-positive single-membrane LROs, accompanied by an increase in the number of LROs (Fig. 1B).
A BMDMs (Bone marrow-derived macrophage) stably expressing Strawberry-Rab5A (Str-Rab5A) or Strawberry-Rab7 (Str-Rab7) were fixed and stained with a Rab32 antibody and observed using confocal fluorescence microscopy. This experiment was repeated twice. Scale bar: 10 μm. B BMDMs expressing GFP-Rab32 and Str-Rab5A were treated with bafilomycin A1 (50 nM) for 24 h. After fixation, organelles exhibiting fluorescence were analyzed using confocal fluorescence microscopy. The targeted regions were further examined by CLEM. Green arrows indicate Rab32-positive organelles, red arrows indicate Str-Rab5A-positive internal structures, white arrows indicate Rab32-engulfed structures, and yellow arrows indicate the investigation structure. Two cells were observed. Scale bar: 5 μm (IF), 2 μm (EM).
These findings suggest that acidification-dependent degradation of organelles occurs within Rab32-positive LROs.
Evidence for Rab32-dependent microautophagy-mediated mitophagy in macrophages
In addition to endosomal components, the mitochondrial marker Tomm20 was internalized into Rab32-positive LROs following bafilomycin A1 treatment (Supplementary Fig. 1B), with this phenomenon becoming more pronounced under mitochondrial damage induced by oligomycin A and antimycin A (OA) treatment (Fig. 2A). To determine whether this mitochondrial transport represented mitophagy—selective autophagy targeting mitochondria—cells were treated with OA and analyzed using Mtphagy Dye, a reagent specific for detecting mitophagy16. The Mtphagy Dye signals were markedly enhanced, indicating active mitophagy under these conditions (Fig. 2B, C). Rab38 is a paralog of Rab32 that functions redundantly in melanocytes and osteoclasts12,13. Although Rab38 expression is minimal in macrophages14, Rab32/38 double knockout (DKO) BMDMs were analyzed to eliminate the potential contribution of Rab38. Rab32/38 DKO BMDMs exhibited attenuated Mtphagy Dye fluorescence signals in response to treatment with either OA or carbonyl cyanide m-chlorophenyl hydrazone (CCCP), another mitochondrial damage-inducing agent (Fig. 2B, C and Supplementary Fig. 1C).
A BMDMs were treated with or without OA for 6 h and then fixed for immunostaining with anti-Rab32 and anti-Tomm20 antibodies. Scale bars: 5 μm (left), 2 μm (insets). Arrows indicate mitochondria-containing Rab32-LRO. B BMDMs isolated from WT and Rab32/Rab38 DKO mice were stained with Mtphagy Dye and treated with OA for 3 h. The cells were observed using confocal microscopy. Scale bar: 10 μm. C The fluorescence signal intensities of Mtphagy Dye in WT and DKO BMDMs from (B) were quantified using ImageJ. Each dot represents a measurement from an individual cell. A.U. denotes arbitrary units. Data are presented as mean values ± SD. Similar experiments were performed more than three times, and a representative result is shown. D WT, Atg7 KO, or Fip200 KO RAW264.7 cells stably expressing GFP-Rab32 were stained with Mtphagy Dye and treated with or without OA for 3 h. Live cells were observed using confocal microscopy. Scale bars: 5 μm (left), 2 μm (insets). E The fluorescence intensity of Mtphagy Dye in the three groups was quantified from (D) using ImageJ. Each dot represents the fluorescence intensity of an individual cell. The experiment was performed twice, and a representative result is shown. Data are presented as mean values ± SD. F Lamp1-GFP-expressing WT, Atg7 KO, Fip200 KO, or Rab32 KO RAW264.7 cells were stained with MitoTracker and treated with OA and bafilomycin A1 for 3 h. Live cells were then observed using confocal microscopy. Scale bars: 5 μm (left) and 2 μm (right). G WT, Atg7 KO, or Rab32 KO RAW264.7 cells were treated with OA in combination with the anti-apoptotic drug z-VAD for 24 h. Cells were then collected for Western blot analysis to detect SDHA, SDHB, and Tomm20 protein levels. Quantitative data were collected from four independent experiments. Data are presented as mean values ± SD. Statistical significance was analyzed using an unpaired two-tailed t-test for (C) and (E) and a paired two-tailed t-test for (G). (*p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001).
Next, we investigated whether macroautophagy is involved in mitophagy in macrophages. In HeLa cells treated with CCCP, the strong colocalization of the autophagosome marker LC317 and Lamp1-associated mitochondria was observed, confirming the occurrence of macroautophagy-dependent mitophagy, as previously reported (Supplementary Fig. 2A)18. In contrast, BMDMs exhibited a larger population of mitochondria that were Lamp1-positive but LC3-negative compared to the LC3-positive population (Supplementary Fig. 2B), and this Lamp1-positive/LC3-negative population was almost absent in Rab32/38 DKO BMDMs (Supplementary Fig. 2B). Although LC3 protein levels appear relatively low in BMDMs, LC3 is still converted to its LC3-II form, supporting the presence of macroautophagy activity (Supplementary Fig. 2C). While macroautophagy contributes to mitophagy, Rab32-positive mitochondria are more abundant than LC3-positive mitochondria in OA-treated BMDMs (Supplementary Fig. 2D).
The mouse macrophage-like cell line RAW264.7 exhibited Mtphagy Dye incorporation into GFP-Rab32-positive LROs following OA treatment, whereas Rab32 knockout (KO) RAW264.7 cells lost Mtphagy Dye signals (Fig. 2D and Supplementary Fig. 3A, B). However, exogenous Rab32 expression restored Mtphagy Dye signals, indicating that a similar phenomenon occurs in RAW264.7 cells. Knockout of the Atg7 or Rb1cc1 (also known as Fip200) genes, both essential for macroautophagy19,20, did not significantly affect Mtphagy Dye signals under OA treatment, with levels comparable to wild-type RAW264.7 cells (Fig. 2D, E and Supplementary Fig. 3C, D). Consistently, other mitochondrial markers, including MitoTracker, Tomm20, and mt-mKeima21, exhibited similar results, and these conclusions were further supported by the strong co-localization of MitoTracker with Tomm20 and the observation of double-positive structures within Rab32-positive LROs. (Supplementary Fig. 3E–J). Furthermore, mitochondria were not transported into Lamp1-GFP-positive LROs in Rab32 KO cells, unlike in wild-type (WT), Atg7 KO, and Fip200 KO cells (Fig. 2F). CCCP treatment also enhanced these mitochondrial transport processes in Atg7 KO and Fip200 KO cells (Supplementary Fig. 4A–D). Moreover, mitochondrial matrix protein levels decreased in WT and Atg7 KO cells but not in Rab32 KO cells (Fig. 2G). Collectively, these findings suggest that macroautophagy plays a less prominent role in mitophagy in these macrophage cells compared to HeLa cells.
Of note, Rab32-positive LROs were sufficiently large to undergo invagination with Lamp1-RFP, allowing the entry of mitochondria, which formed Rab32-positive ring structures (Fig. 3A, B and Supplementary Fig. 5A), but this was not the case for the Rab7-positive late endosome (Supplementary Fig. 5B). CLEM analysis demonstrated that MitoTracker-positive bodies were transported into invaginating Rab32-positive LROs (Fig. 3C and Supplementary Fig. 5C). Additionally, three-dimensional imaging further confirmed that Rab32-positive LROs can fully contain mitochondria in Fip200 KO macrophages (Supplementary Movie 1). These observations indicate that mitophagy in macrophages occurs through microautophagy.
A RAW264.7 cells stably expressing GFP-Rab32 and Lamp1-RFP were observed by confocal microscopy without fixation. Yellow arrowheads indicate invagination sites. This experiment is repeated three times. Scale bar: 5 μm. B GFP-Rab32-expressing Atg7 KO RAW264.7 cells were stained with MitoTracker and then with OA for 1.5 h. This experiment is repeated three times. Scale bar: 2 μm. C WT RAW264.7 cells expressing GFP-Rab32 were stained with MitoTracker and treated with CCCP for 1.5 h. After fixation, cells were observed using confocal microscopy and further analyzed by electron microscopy. Yellow arrows indicate mitochondria stained with MitoTracker. The two arrows on the far-left highlight mitochondria in close association with invaginated membranes. Yellow arrowheads indicate the MitoTracker-positive bodies inside Rab32-LROs, which exhibited poorly defined cristae, suggesting potential damage induced by OA treatment48. Total 22 cells were observed. D RAW264.7 cells expressing GFP-Rab32 WT, Q83L (active form), or T37N (inactive form) were analyzed using confocal microscopy. White arrows indicate invaginated membrane structures positive for LRO. Quantitative data represent the percentage of invagination-positive Rab32-LROs per cell. Any GFP-Rab32 signal detected inside the GFP-Rab32 limiting membrane was classified as an invaginating structure. More than 30 cells per group were analyzed in a blinded manner. No Rab32-ring-like structures were observed in GFP-Rab32TN-expressing cells. The experiments were performed twice, and a representative result is presented. ND: not detectable. Scale bar: 5 μm. Statistical significance was analyzed using an unpaired t-test (****p < 0.0001). E RAW264.7 cells expressing GFP-Rab32 WT or QL were treated with or without Apilimod (AP) for 2 h. This experiment is repeated three times. Scale bar: 2 μm. F GFP-Rab32QL-expressing RAW264.7 cells were stained with MitoTracker and treated with Apilimod (AP) for 30 min followed by washout for 1 or 3 h. After 1-h of washout, cells were treated with OA for the indicated periods, with or without bafilomycin A1. This experiment is repeated three times. Scale bar: 2 μm. G GFP-Rab32QL-expressing Atg7 KO RAW264.7 cells were stained with MitoTracker and treated with Apilimod for 30 min, followed by a 1 h washout. After the washout, cells were treated with OA for 3 h. Time-lapse imaging was performed using confocal super-resolution microscopy (SpinSR10) for 10 min, with an 80 s interval between frames.
Rab32 appears to play an active role as a GTPase in microautophagy, as expression of the constitutively active Rab32 (QL) mutant enhanced invagination and LRO formation (Fig. 3D). Phosphatidylinositol (3,5)-bisphosphate [PtdIns(3,5)P₂] was also critical in this process, as observed in other cases of microautophagy22,23. Treatment with Apilimod, an inhibitor of the PIP kinase, PIKfyve, responsible for PtdIns(3,5)P₂ production, completely abolished Rab32-LRO invagination, even in cells expressing the constitutively active Rab32 (QL) mutant (Fig. 3E). Notably, LROs in Rab32 (QL)-expressing cells remained un-invaginated for at least 3 h following Apilimod washout (Fig. 3F). However, inducing mitochondrial damage with OA treatment 1 h after Apilimod washout effectively triggered LRO invagination and mitochondrial uptake, suggesting that the initiation of microautophagy is tightly regulated (Fig. 3F). This controlled induction enabled real-time imaging of the mitochondrial engulfment process within invaginating LRO membranes (Fig. 3G and Supplementary Movie 2)
The endosomal sorting complex required for transport (ESCRT) system has been implicated in various microautophagy processes22,24,25,26. However, in this study, several ESCRT components, including ALIX, TSG101, CHMP2A, VPS4A, and VPS4B, exhibited minimal association with Rab32-positive LROs, even following OA treatment (Supplementary Fig. 6A). Moreover, knockdown of Tsg101, Chmp3, Vps4A, or Vps4B, as well as exogenous expression of dominant-negative VPS4AE228Q, did not affect this process, suggesting that the contribution of ESCRT machinery is limited (Supplementary Fig. 6B, C).
Ubiquitination/p62 play a role in micromitophagy in macrophage
OA treatment induced mitochondrial ubiquitination and fragmentation (Fig. 4A). The number of ubiquitin-positive puncta was higher in Rab32/38 DKO BMDMs than in wild-type BMDMs (Fig. 4B). However, treatment with bafilomycin A1 did not further increase the number of ubiquitin puncta in DKO cells, indicating that ubiquitinated mitochondria were rarely transported into LROs for degradation in the absence of Rab32 and Rab38 (Fig. 4B). Similarly, Rab32 KO RAW264.7 cells accumulated more ubiquitin than WT cells, with a modest increase in ubiquitin puncta level following bafilomycin A1 treatment (Supplementary Fig. 7A). Ubiquitinated mitochondria were also detected within Rab32-positive LROs in Atg7 KO macrophages after bafilomycin A1 treatment (Supplementary Fig. 7B). Inhibition of ubiquitination with the E1 inhibitor TAK243 severely blocked mitochondrial import into Rab32-LROs, even in macroautophagy-deficient macrophages (Fig. 4C). Furthermore, following Apilimod washout and subsequent OA treatment, microautophagy was reactivated; however, TAK243 treatment suppressed mitochondrial invagination into LROs (Fig. 4D). Interestingly, although expression of the constitutively active Rab32 (QL) mutant promoted LRO invagination, mitochondria were not incorporated into LROs, suggesting that ubiquitination is essential for the attachment of mitochondria to the LRO membrane (Fig. 4D).
A BMDMs were treated with OA for 6 h. Following treatment, cells were fixed, stained with anti-Tomm20 and anti-ubiquitin antibodies. White arrowheads indicate ubiquitin-positive mitochondria. Scale bar: 10 μm. Each data point represents the number of ubiquitin signals per image, normalized to the number of cells. N = 5 images for “-” and 15 images for “OA”. Images were analyzed using the ImageJ ‘Find Maxima’ tool, applying a defined threshold in a blinded manner. The experiments were performed twice, and a representative result is presented. Data are presented as mean values ± SD. B WT or DKO BMDMs were treated and analyzed as in (A). Scale bar: 5 μm. Each point represents the number of ubiquitin signals per image, normalized to the number of cells. N = 4 images for each group. The experiments were performed twice, and a representative result is presented. Data are presented as mean values ± SD. C Atg7 KO or Fip200 KO RAW264.7 cells expressing GFP-Rab32 were pre-stained with MitoTracker and treated with OA and bafilomycin A1, with or without TAK243 (5 μM) for 3 h. Each dot represents the fluorescence intensity of MitoTraker inside Rab32 rings (N = 199, 196, 230, 203 rings from left to right), measured using ImageJ. Similar experiments were conducted at least twice, and a representative result is shown. Box plots indicate the median (center line), the interquartile range (box), and the range (whiskers). D Atg7 KO RAW264.7 cells expressing GFP-Rab32WT or QL were treated with the same condition in Fig. 3F, with or without TAK243 for 1.5 h. White arrows indicate the MitoTracker positive signal engulfed into Rab32-LRO. E Atg7 KO RAW264.7 cells expressing GFP-Rab32 were stained with MitoTracker and then treated with OA and bafilomycin A1 with or without TAK243 for 3 h. After fixation, cells were stained with the anti-p62 antibody. Yellow arrows indicate p62-positive mitochondria inside Rab32-LRO. White arrowheads indicate p62-positive mitochondria associated with Rab32-LRO. Scale bar: 5 μm (upper), 2 μm (bottom). F Scramble control or p62 knockdown (KD) WT or Atg7 KO RAW264.7 cells were transduced with GFP-Rab32 using a retrovirus. After 24 h of transduction, cells were stained with MitoTracker and treated with OA and bafilomycin A1 for 3 h. Scale bar: 2 μm. The percentage of MitoTracker-positive Rab32-LROs per cell was quantified. N = 20, 20, 18, 19 cells from left to right. The experiments were performed twice, and a representative result is presented. Data are presented as mean values ± SD. Statistical significance was analyzed using an unpaired two-tailed t-test (***p < 0.001; ****p < 0.0001).
Regarding ubiquitin adapter proteins, p62/SQSTM127 was recruited to mitochondria in response to OA treatment, whereas NDP52 was not28 (Fig. 4E and Supplementary Fig. 8). p62-positive mitochondria were taken up into Rab32-LROs (Fig. 4E), whereas TAK243 treatment prevented p62 recruitment to mitochondria, and neither p62 nor mitochondria were found inside Rab32-LROs (Fig. 4E). Furthermore, p62 knockdown reduced the efficiency of Rab32-LRO-mediated mitochondrial uptake (Fig. 4F and Supplementary Fig. 8B). These results indicate that the ubiquitin-p62 axis is crucial for Rab32-LROs to recognize mitochondria. Cardiolipin, previously identified as an “eat-me” signal during mitophagy in neuronal cells, was also investigated29. However, the knockdown of PLSCR3, a scramblase that externalizes cardiolipin during mitophagy, had no effect on this process in macrophages (Supplementary Fig. 6B). Likewise, the knockdown of NIPSNAP1 and NIPSNAP2, which form a heterodimeric complex serving as an “eat-me” signal in HeLa cell mitophagy30, did not impact microautophagy in macrophages (Supplementary Fig. 6B).
Collectively, these findings suggest that mitochondrial ubiquitination induced by OA treatment and p62 plays a critical role in promoting microautophagy in macrophages.
Microautophagy plays a role in M1 polarization
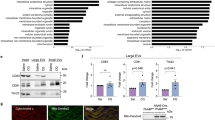
In response to inflammatory stimuli, macrophages polarize toward the M1-activated state, a process accompanied by a metabolic shift from oxidative phosphorylation to glycolysis31. This transition has been reported to involve a reduction in mitochondrial content through programmed mitophagy11. To investigate whether microautophagy contributes to M1 polarization, BMDMs derived from WT and Rab32/38 DKO mice were stimulated with LPS and interferon-γ (IFN-γ). Upon stimulation, mitochondrial ubiquitination (Fig. 5A) and fragmentation (Supplementary Fig. 9A) were observed, followed by the delivery of mitochondria into LROs in WT cells (Fig. 5B). In contrast, in Rab32/38 DKO macrophages, mitochondria failed to localize to LROs (Fig. 5B). After 24 h of treatment, fragmented mitochondria accumulated significantly more in Rab32/38 DKO BMDMs than in WT BMDMs (Supplementary Fig. 9B). Mitochondrial matrix proteins were degraded in WT BMDMs after 24 h of treatment, whereas they largely remained intact in Rab32/38 DKO BMDMs (Fig. 5C). Similar observations were made in Rab32 KO RAW264.7 cells (Supplementary Fig. 9C), whereas Atg7 KO and Fip200 KO cells exhibited only minimal defects (Supplementary Fig. 9D, E). Furthermore, the expression of proinflammatory cytokines IL-6, IL-1β, and TNF-α—markers of M1 polarization—was significantly induced in WT macrophages following LPS and IFN-γ stimulation but was markedly suppressed in Rab32/38 DKO macrophages (Fig. 5D). Correspondingly, cytokine production and secretion, including IL-6 and TNF-α, were significantly suppressed in Rab32/38 DKO macrophages (Fig. 5E). These impairments were not due to defects in the surface expression of LPS receptor TLR4 or IFN-γ receptor IFN-γR, as early signaling events, such as nuclear translocation of NF-κB and STAT1, were unaffected in Rab32/38 DKO cells (Fig. 5F and Supplementary Fig. 9F). However, after 12–24 h, nuclear retention of NF-κB was diminished in DKO cells (Fig. 5F). Additionally, the metabolic shift toward glycolysis in response to LPS/IFN-γ stimulation was impaired in Rab32/38 DKO BMDMs, whereas mitochondrial respiration was reduced to the same level as in WT BMDMs (Fig. 5G and Supplementary Fig. 9G). Collectively, these findings indicate that Rab32-mediated microautophagy plays an essential role in regulating mitochondrial homeostasis in macrophages, which is crucial for M1 polarization.
A BMDMs were treated with LPS/IFN-γ for 6 h, stained with anti-Tomm20 and anti-ubiquitin antibodies, and observed by confocal microscopy. White arrowheads indicate ubiquitin-positive mitochondria. Scale bar: 10 μm. B WT and DKO BMDM cells were stained with MitoTracker and treated as described in (A). Cells were then fixed and stained with anti-Lamp1 antibody. White arrows indicate mitochondria-containing lysosomes. Yellow arrowheads mark fragmented mitochondria undergoing uptake by LRO. Scale bar: 2 μm. C WT or DKO BMDMs treated with or without LPS/IFN-γ for 24 h were subjected to Western blot analysis. Quantitative data were collected from three independent experiments. Data are presented as mean values ± SD. D, E WT or DKO BMDMs were treated with LPS/IFN-γ for 16 or 24 h. Cells were analyzed by qPCR (D), and culture media were analyzed using ELISA (E). ELISA data were normalized to total RNA levels. The experiments were repeated at least three times, and representative data are shown from a technical triplicate. Data are presented as mean values ± SD. F WT or DKO BMDMs were treated with LPS/IFN-γ for the indicated periods, fixed, and stained with anti-NFκB antibody and DAPI (not shown). Cells were observed using confocal microscopy. Scale bar: 10 μm. Quantitative data represent the mean signal intensity of NF-κB in the nucleus per cell among more than 400 cells measured by ImageJ. The blue and yellow background areas indicate standard deviation (SD) ranges. The experiments were performed twice, and representative data are shown. G WT or DKO BMDMs were treated with or without LPS/IFN-γ for 24 h, and the extracellular acidification rate (ECAR) was measured using the Seahorse XF Analyzer. The experiments were performed three times, and representative data are shown from a technical triplicate. Data are presented as mean values ± SD. Statistical significance was analyzed using a paired t-test for (C) and an unpaired two-tailed t-test for (D, E) and (F) (*p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001).
Discussion
Microautophagy in mammalian cells is primarily characterized within typical endosomes or lysosomes. However, even endosomes can be engulfed via microautophagy in the visceral endoderm, where active endocytosis occurs during early mouse embryogenesis32. This process likely functions to prevent the accumulation of endosomes in the cytoplasm by engulfing and degrading excess membrane structures within lysosomal compartments. Macrophages, known for their highly dynamic membrane activities such as phagocytosis33, may also rely on microautophagy to maintain cellular homeostasis. In this context, the relatively large Rab32-positive LROs in macrophages appear to facilitate active microautophagy, promoting efficient membrane turnover, including endosomes.
Macroautophagy in macrophages has been extensively studied, not only in the context of xenophagy7,34 but also for its critical role in mitophagy9,35. However, under our experimental conditions, microautophagy appears to play a more prominent role in mitophagy than macroautophagy. Therefore, this process may be more accurately referred to as micromitophagy. We speculate that macrophages, which have an intrinsic capacity for microautophagy, may preferentially use this pathway for mitophagy due to its lower energy demand compared to macroautophagy. It would be interesting to further investigate its role in macroautophagy because Rab32 localizes to autolysosomes and is required for the recycling of autophagosomal outer membrane components in U2OS human osteosarcoma cells36. Our data indicate that Rab32 activity enhances microautophagy in macrophages. While the ESCRT complex does not appear to be essential for this process, PtdIns(3,5)P₂ plays a key regulatory role in membrane dynamics. Notably, ubiquitination plays a crucial role in micromitophagy, similar to its function in macromitophagy. Therefore, mitochondrial damage may trigger ubiquitination through an “eat-me” signal, independent of cardiolipin or NIPSNAP1/2. In cooperation with ubiquitin, p62 facilitates the capture of mitochondria into LROs, a process that appears to be independent of LC3 family proteins.
Macrophage function is highly influenced by metabolic status, including mitochondrial regulation37, making it plausible that active micromitophagy plays a role in M1 macrophage polarization. However, it remains possible that the absence of Rab32 contributes to phenotypes beyond defects in microautophagy, despite the seemingly unaffected response to LPS and IFN-γ stimulation. Identifying the specific molecular machinery involved in micromitophagy will be essential for clarifying this point. The mechanism underlying the decrease in ECAR levels in DKO cells remains unclear. However, one possible explanation is that the accumulation of fragmented and less active mitochondria in DKO cells leads to cellular dysfunction, which may negatively impact glycolysis and ECAR levels. Furthermore, this could subsequently affect M1 polarization, including NF-κB nuclear translocation, potentially suppressing glycolysis via HIF-1α through a feedback loop. NF-κB has been reported to play a crucial role in the transcriptional regulation of HIF-1α, a key regulator of glycolysis38.
This study suggests that diverse and versatile protein degradation systems are emerging to coordinate various aspects of cellular physiology.
Methods
Plasmids and cells
Plasmids expressing the fluorescently tagged proteins ALIX, TSG101, CHMP2A, VPS4A, and VPS4B were described previously39. Plasmids expressing the doxycycline inducible E228Q-VPS4A-HA, mCherry-Tomm20-N10, and pCHAC-mt-mKeima were obtained from Addgene (#200087, #55146, #72342). pLKO.1-puromycin plasmids expressing shRNA targeting to TSG101, CHMP3, VPS4B, VPS4A, NIPNASP1, NIPNASP2, Plscr3 and p62 were purchased from Sigma-Aldrich (TRC number: TRCN0000-054606, −098066, −101824, −101416, −174717, −192544, −105105, −098619). The pLKO.1-puro-control plasmid was a gift from Professor Makoto Sato, from the Graduate School of Medicine, Osaka University. The pMRX-puro-EGFP-Rab32 and -Rab38 plasmids were prepared as described previously40. Mouse Rab5A and Rab7 cDNA were subcloned into the pMRX-bsr-mStr plasmid or pMRX-puro-mStr vector14,40 from the constructs pMRX-EGFP-C1-Rab5A and pMRX-EGFP-C1-Rab741. To generate the pMRX-puro-Lamp1-EGFP plasmid, EGFP was replaced with RFP in the pMRX-puro-Lamp1-mRFP plasmid42. The genes of the EGFP and pMRX-ires-Lamp1 vector were amplified through PCR with KOD One (KMM 101, TOYOBO). Both contained 20 bps overlapping on both ends. The primers used for insertion were F: GATCCACCGGTCGCCACCATGGTGAGCAAGGGCGAGGA, R: TTTACGTAGCGGCCGCTTTACTTGTACAGCTCGTCCA, and for the vector, F: TAAAGCGGCCGCTACGTAAA, R: CATGGTGGCGACCGGTGGATC. After the reaction, DpnI (1235 A, Takara) was added to the PCR reaction mixture at 37 °C for more than 3 h. These inserts and vectors amplified by PCR were ligated using the Gibson assembly method43. HeLa cells stably expressing myc-Parkin were described previously44. To generate GFP-Rab32-, GFP-Rab38-, Lamp1-GFP, Lamp1-RFP, and mt-mKeima-expressing retrovirus vectors, Platinum-E cells were transfected with the pMRX-puro-EGFP-Rab32, pMRX-puro-EGFP-Rab38, pMRX-puro-Lamp1-EGFP, pMRX-bls-Lamp1-RFP, and pCHAC-mt-mKeima plasmids, respectively, to generate the virus as described previously14. To generate E228Q-VPS4A-HA and shRNA-expressing lentivirus vectors, HEK293 cells were transfected with the pInducer-20-neo-E228Q-VPS4A-HA plasmids or pLKO.1-puro with shRNA to target mRNA genes with psPAX2 and pMDG plasmids by PEI. The virus medium collection was performed in accordance with the retrovirus collection.
To knock out Rab32, Atg7, and Fip200 using the CRISPR/Cas9 system45, guide RNA targeting the exons was designed. The specific recognition sequences of the 20 bp before PAM in each construct were as follows: Rab32, 5′-CACCGAGGGCGAAGTCCACGCCAATGG-3′; Atg7, 5′-CACCGAACTCCAACGTCAAGCGGGT-3′; Fip200, 5′-CACCGCGGTTGATAGCCGCCGCCG −3′. These guide oligos, with BbsI restriction sites at both ends, were annealed with their anti-sense oligos and inserted downstream of the U6 promoter in vector pSpCas9(BB)−2A-GFP (pX458) (#48138, Addgene); the endotoxin (V5910, MIR) was removed from the resultant plasmids, and the plasmids were used for transfection into RAW264.7 cells by electroporation (Voltage, 1680 V; width, 20 ms per pulse, Neon, Thermo Fisher). Single-cell sorting was performed after 24-h transfection. After a one-week culture of single cells with antibiotics, a fresh medium was added to support growth for one more week to allow colony formation. KO cell lines were confirmed by western blotting and genomic DNA sequencing of Rab32, Atg7, and Fip200.
Mice
WT C57BL/6 J mice were purchased from Japan SLC (Shizuoka, Japan), and Rab32/38 double knockout C57BL/6 J mice were described previously13. All mice were maintained under specific pathogen-free conditions in the animal facility of the Graduate School of Dentistry, Osaka University. Eight to twelve-weeks age male mice were used for isolation of bone marrow cells and collection of femurs, tibiae, and fibulae, were appropriately euthanized using carbon dioxide inhalation.
Cell culture
HeLa and RAW264.7 cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM)-high glucose (D6429, Sigma-Aldrich) containing 10% fetal bovine serum (FBS), 100 units/mL penicillin, and 100 µg/mL streptomycin at 37 °C in 5% CO2. WT and Rab32/Rab38 DKO BMDMs were collected from bone, fibula, and femur, and cultured in MEMα medium containing 10% FBS and MCSF13. BMDM cells were infected with a retrovirus to induce fluorescent protein expression within one week of cell collection, followed by selection with puromycin or blasticidin14. BMDMs expressing GFP-Rab32-puro and Str-Rab5A-bsr were generated through sequential transduction with each virus, followed by selection with the respective antibiotics. RAW264.7 cells were infected with a retrovirus to induce fluorescent protein expression, and selected with either puromycin or blasticidin. For shRNA induction, RAW264.7 cells were first infected with a lentivirus and selected with puromycin, and then further infected with the GFP-Rab32-puro retrovirus, with GFP-positive cells subsequently selected under a microscope. For E228Q-VPS4A-HA expression, GFP-Rab32-(puro)-expressing RAW264.7 cells were infected with a retrovirus and selected with G418. The expression of E228Q-VPS4A-HA was induced by doxycycline treatment, confirmed by Western blot analysis, and followed by the indicated treatments. For mt-mKeima expression, RAW264.7 cells were infected with a retrovirus, and fluorescence-positive cells (about 20%) were subsequently selected under a microscope. For the transient expression of genes of interest, plasmids were transfected into RAW264.7 cells by electroporation (Voltage, 1680 V; Width, 20 ms per pulse, Neon, Thermo Fisher).
Cell treatment
For Mtphagy Dye or MitoTracker staining, cells were cultured on glass-bottom dishes (D11530H, Matsunami Glass) and treated with Mtphagy Dye (100 nM, MD01, Dojindo) in serum-free DMEM for 30 min, or MitoTracker CMXRos (200 nM, M7512, Thermo Fisher) in complete medium for 30 min. After three washes, cells were observed without fixation under a TSC SP8 confocal laser scanning microscope (Leica) with a laser wavelength of 568 nm. CCCP (10 μM, 034-16993, Wako) or a combination of oligomycin A (10 μM, 75351, Sigma-Aldrich) and antimycin A (5 μM, A8674, Sigma-Aldrich) was used to induce mitochondrial damage. For long-term treatment of OA, pan caspase inhibitor z-VAD-FMK (20 μM, S7023, Selleck) was used to inhibit apoptotic cell death. Bafilomycin A1 (50 nM for 24-h treatment and 200 nM for 2 or 3-h treatment, BVT-0252-M001, BioViotica) was used for the inhibition of the LRO acidification. Cell treatment with Apilimod (100 nM, STA5326, Selleck) for 30 min or 2 h was used to inhibit PIKfyve activity. Doxycycline (1 μg/ml, D9891, Sigma-Aldrich) was used for 16 h to induce E228Q-VPS4A-HA expression. Macrophages were treated with a combination of LPS (100 ng/ml, L2880, Sigma-Aldrich) and mouse IFN-γ (10 ng/ml, 485-MI, R&D Systems).
Antibodies
The following antibodies and dilutions were used for immunofluorescence (IF) staining and western blotting (WB) in this study: primary antibodies (dilution, catalog number, maker): rabbit anti-LC3 (1/500 for IF and 1/2000 for WB, PM036, MBL and 1/1000, #S9661, CST for WB), rabbit anti-p62 (1/500 for IF and 1/2000 for WB, PM045, MBL), rabbit anti-NDP52 (1/200 for IF, GTX115378, GeneTex), rabbit anti-Atg7 (1/1000 for WB, 8558S, Cell Signaling), rabbit anti-FIP200 (1/2000 for WB, 17250-1-AP, Proteintech), rat anti-Lamp1 (1/200 for IF, sc-19992, Santa Cruz Biotechnology), rabbit anti-Tomm20 (1/500 for IF and 1/5000 for WB, Proteintech, 11802-1-AP), rabbit anti-STAT-1 (1/250 for IF, 10144-2-AP, Proteintech), rabbit anti-NFκB (1/250 for IF, D14E12, CST), rabbit anti-Rab32 polyclonal46 (1/250 for IF in Fig. 1A), mouse anti-Rab32 (1/100 for IF and 1/200 for WB, sc-390178, Santa Cruz Biotechnology), mouse anti-SDHA (1/200 for WB, sc-166909, Santa Cruz Biotechnology), rabbit anti-SDHB (1/5000, 10620-1-AP, Proteintech), mouse anti-ubiquitin (FK2) (1/1000 for IF, BML-PW8810, Enzo Life Sciences), rat anti-HA (1/2000 for WB, clone:3F10, Roche), mouse anti-actin (1/10000 for WB, 66009-1-IG, Proteintech), and mouse anti-tubulin (1/10000 for WB, 66031-1-IG, Proteintech); secondary antibodies: anti-rabbit IgG-Alexa Fluor 488 or 568 (1/1000, A11008, A10042, Thermo Fisher), anti-mouse IgG-Alexa Fluor 488 or 568 (1/1000, A11001, A10037, Thermo Fisher), and anti-rat IgG-Alexa Fluor 488 (1/1000, A21208, Thermo Fisher). Western blotting was performed as reported47.
Immunofluorescence staining and microscopy
Cells were cultured on 12 mm cover glasses (No 1S thickness, C012001, Matsunami Glass), which were placed on 24-well plates. The cells were fixed by adding 4% paraformaldehyde (163-20145, FUJIFILM) for 5 min, and then changed to fresh 4% paraformaldehyde for a further 5 min at room temperature (RT). After a wash with PBS, the cells were permeabilized using 50 µg/mL digitonin (300410, Calbiochem) in PBS for 10 min at RT, followed by blocking with 0.2% gelatin (076-02765, Wako) in PBS for 30 min. Target proteins were labeled using specific primary and secondary antibodies, and samples were mounted with ProLong Glass Antifade Mountant (P36984, Thermo Fisher). For live-cell imaging, cells were cultured on 35 mm glass-bottom dishes (D11350H, Matsunami Glass). Both fixed and live cells were observed under a TSC SP8 confocal laser scanning microscope (Leica). The objective lens used was HC PL APO CS2 63 × 1.40 OIL (Leica). The fluorescence intensity and vesicle number were analyzed using the ImageJ software (NIH). Most of the images were further deconvoluted using the Huygens software (Scientific Volume Imaging). For time-lapse analysis, images were obtained under a SpinSR10 spinning disk confocal super-resolution microscope integrated with the IXplore System (Evident) under an incubation temperature of 37 °C in an INUB-ZILCS-F1 incubator (TOKAI HIT) at 80-second intervals for 10 min. The objective lens used was UAPON100XO. A three-dimensional image was taken using SpinSR10.
CLEM analysis
Cells were cultured in a 3.5 cm gridded glass-bottomed dish (P35G-2-14-CGRD/H, MatTek). The cells were washed twice with PBS and fixed in methanol-free 4% formaldehyde, ultrapure (Polyscience Inc.) / 0.1 M phosphate buffer (19 mM NaH2PO·H2O, 81 mM Na2HPO·7H2O (pH 7.4)) containing 2% sucrose for 30 min at RT. The dishes were subsequently washed thrice with 0.1 M phosphate buffer containing 4% sucrose for a total of 5 min. The cells were observed with a TSC SP8 confocal microscope (Leica). The cells were further processed for electron microscopy, ultrathin-sectioning as described previously47. The images were obtained using the transmission electron microscope JEM-1400plus (JEOL) and an Olympus Veleta CCD camera (Olympus) at 80 kV.
qPCR
Cell pellets were treated with TRIsure (Bioline) to extract total mRNA and the iScript cDNA Synthesis Kit (1725038, Bio-Rad) to generate cDNA. Real-time PCR analysis was performed with the Step One Plus Real-time PCR System (Applied Biosystems) using QuantiTect SYBR Green PCR (204141, Qiagen) and the primers for GAPDH: AAATGGTGAAGGTCGGTGTG (forward), TGAAGGGGTCGTTGATGG (reverse); TNF-α: GCCAGGATGAAAGTTGTATG (forward), CAGGCGTTGTTCTTATTCC (reverse); IL-1β: GCTGGAAACCATGATCACCT (forward), GAGTTGCCACACAGCATCAC (reverse); IL-6: TGTATCGGCTCATCTCCTCCAT (forward), GACTCCTTGGGTTCCTTGCTT (reverse); PLS3: CCTGCTCCTTTCGTGCCATT (forward), CCACTCGTTCAGCCTTCTGAT (reverse); NIPNASP1: ACAACGTGAAGCCCGAATGT (forward), GCGAGCAGGGATAGTCCTCA (reverse); NIPNASP2 (Gbas): AGTCCTTGTTTGTCAGAAAGGTC (forward), GTCTAGGCATTCCGGCTTAAC (reverse); TSG101: TCTAACCGTCCGTCAAACTGT (forward), TTGTACCAGTGAGGTTCACCA (reverse); CHMP3 (VPS24): AGGAAATGAGAGTTGTTGACAGG (forward), TGCATCTTTCACAGACCGTTTTA (reverse); VPS4A: ACGGTGGAATGATGTAGCTGG (forward), CCAAAGAGGAGTATGCCTCGC (reverse); VPS4B: GGCTGCACGGAGAATTAAGAC (forward), TCCAGAACCCAGGGTATATTTGT (reverse). Relative expression levels for each gene were calculated according to the ∆∆Ct method. The fold induction of M1 cytokine marker genes was normalized to the basal levels in BMDMs. The knockdown efficiency of shRNA-targeted genes was normalized to the gene levels in the scramble control.
ELISA
The culture supernatants were subjected to mouse TNFα and IL-6 cytokine detection by ELISA (DY410-05 and DY406-05, R&D Systems). The absorbance at 450 nm and the reference absorbance at 570 nm were measured by iMark (Bio-Rad).
Seahorse analysis
BMDM cells (1 × 105 cells) were cultured on six-well Seahorse XFp cell culture miniplates overnight and then treated with LPS and IFN-γ for 24 h. The culture cells were treated with the Mito Stress Test kit (Kit 103010-100, Agilent Seahorse XFp) and Glycolytic Rate kit (Kit 103017-100, Agilent Seahorse XFp), and analyzed with the Seahorse XFp Analyzer (Agilent). The compound concentrations were: 1 μM oligomycin, 5 μM FCCP (Carbonyl cyanide-p-trifluoromethoxyphenylhydrazone), and 1 μM Rot/AA for the mitochondrial function assay, and 10 mM glucose, 1.5 μM oligomycin, and 40 μM 2-DG for the glycolysis assay.
Statistics & reproducibility
For statics, raw data were imported into GraphPad Prism 10 for macOS and analyzed. Error bars shown in the figures represent standard deviation (SD). P values were calculated using an unpaired or paired two-tailed t-test. Exact p-values were provided in Source Data. For groups with low variations, analysis was performed with Welch’s correction. ***p < 0.001; 0.001 <**p < 0.01; and 0.01 <*p < 0.05 in figures indicate the level of significance. Sample size were determined following our previous study14. We did not exclude any data. Replicated times were described in the figure legends. For quantification analysis, cells and signals were randomly selected. Blinding was performed in a set of experiments and described in each figure legend.
Ethics statement
All animal studies were approved by the Institutional Animal Experiments Committee of The University of Osaka Graduate School of Dentistry (29-008-0). All the experiments related to gene recombination were approved by the Gene Modification Experiments Safety Committee of The University of Osaka. All co-authors of this study fulfill the authorship criteria of Nature Portfolio journals.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data can be available upon reasonable request to the corresponding author. Source data are provided with this paper.
References
Ohsumi, Y. Historical landmarks of autophagy research. Cell Res 24, 9–23 (2014).
Klionsky, D. J. et al. Autophagy in major human diseases. EMBO J. 40, e108863 (2021).
Oku, M. & Sakai, Y. Three distinct types of microautophagy based on membrane dynamics and molecular machineries. BioEssays 40, e1800008 (2018).
Mijaljica, D., Prescott, M. & Devenish, R. J. Microautophagy in mammalian cells: revisiting a 40-year-old conundrum. Autophagy 7, 673–682 (2011).
Klionsky, D. J. & Eskelinen, E.-L. The vacuole vs. the lysosome: when size matters. Autophagy 10, 185–187 (2014).
Wang, L., Klionsky, D. J. & Shen, H.-M. The emerging mechanisms and functions of microautophagy. Nat. Rev. Mol. Cell Biol. 24, 186–203 (2023).
Wu, M.-Y. & Lu, J.-H. Autophagy and macrophage functions: inflammatory response and phagocytosis. Cells 9, 70 (2019).
Wen, J.-H. et al. Macrophage autophagy in macrophage polarization, chronic inflammation and organ fibrosis. Front. Immunol. 13, 946832 (2022).
Zhong, Z. et al. NF-κB restricts inflammasome activation via elimination of damaged mitochondria. Cell 164, 896–910 (2016).
Galván-Peña, S. & O’Neill, L. A. J. Metabolic reprograming in macrophage polarization. Front. Immunol. 5, 420 (2014).
Esteban-Martínez, L. et al. Programmed mitophagy is essential for the glycolytic switch during cell differentiation. EMBO J. 36, 1688–1706 (2017).
Luzio, J. P., Hackmann, Y., Dieckmann, N. M. G. & Griffiths, G. M. The biogenesis of lysosomes and lysosome-related organelles. Cold Spring Harb. Perspect. Biol. 6, a016840–a016840 (2014).
Tokuda, K. et al. Rab32 and Rab38 maintain bone homeostasis by regulating intracellular traffic in osteoclasts. Cell Struct. Funct. 48, 223–239 (2023).
Noda, K. et al. Characterization of Rab32- and Rab38-positive lysosome-related organelles in osteoclasts and macrophages. J. Biol. Chem. 299, 105191 (2023).
Lu, S.-L. & Noda, T. The emerging role of Rab proteins in osteoclast organelle biogenesis and function. Biochem. Soc. Trans. 52, 2469–2475 (2024).
Iwashita, H. et al. Live cell imaging of mitochondrial autophagy with a novel fluorescent small molecule. ACS Chem. Biol. 12, 2546–2551 (2017).
Kabeya, Y. et al. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J. 19, 5720–5728 (2000).
Narendra, D., Tanaka, A., Suen, D.-F. & Youle, R. J. Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J. Cell Biol. 183, 795–803 (2008).
Komatsu, M. et al. Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J. Cell Biol. 169, 425–434 (2005).
Hara, T. et al. FIP200, a ULK-interacting protein, is required for autophagosome formation in mammalian cells. J. Cell Biol. 181, 497–510 (2008).
Katayama, H., Kogure, T., Mizushima, N., Yoshimori, T. & Miyawaki, A. A sensitive and quantitative technique for detecting autophagic events based on lysosomal delivery. Chem. Biol. 18, 1042–1052 (2011).
Kuchitsu, Y. et al. STING signalling is terminated through ESCRT-dependent microautophagy of vesicles originating from recycling endosomes. Nat. Cell Biol. 25, 453–466 (2023).
Klein, A. D., Petruzzi, K. L., Lee, C. & Overholtzer, M. Stress-induced microautophagy is coordinated with lysosome biogenesis and regulated by PIKfyve. MBoC 35, ar70 (2024).
Ogura, M. et al. Microautophagy, regulated by STK38 and GABARAPs, is essential to repair lysosomes and prevent aging. EMBO Rep. 24, e57300 (2023).
Liao, Y.-C. et al. COPII with ALG2 and ESCRTs control lysosome-dependent microautophagy of ER exit sites. Dev. Cell 59, 1410–1424.e4 (2024).
Mejlvang, J. et al. Starvation induces rapid degradation of selective autophagy receptors by endosomal microautophagy. J. Cell Biol. 217, 3640–3655 (2018).
Geisler, S. et al. PINK1/Parkin-mediated mitophagy is dependent on VDAC1 and p62/SQSTM1. Nat. Cell Biol. 12, 119–131 (2010).
Lazarou, M. et al. The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature 524, 309–314 (2015).
Chu, C. T. et al. Cardiolipin externalization to the outer mitochondrial membrane acts as an elimination signal for mitophagy in neuronal cells. Nat. Cell Biol. 15, 1197–1205 (2013).
Princely Abudu, Y. et al. NIPSNAP1 and NIPSNAP2 Act as “Eat Me” Signals for Mitophagy. Dev. Cell 49, 509–525.e12 (2019).
O’Neill, L. A. J. & Pearce, E. J. Immunometabolism governs dendritic cell and macrophage function. J. Exp. Med. 213, 15–23 (2016).
Kawamura, N. et al. Delivery of endosomes to lysosomes via microautophagy in the visceral endoderm of mouse embryos. Nat. Commun. 3, 1071 (2012).
Flannagan, R. S., Jaumouillé, V. & Grinstein, S. The cell biology of phagocytosis. Annu Rev. Pathol. 7, 61–98 (2012).
Bah, A. & Vergne, I. Macrophage autophagy and bacterial infections. Front. Immunol. 8, 1483 (2017).
Zhou, Y. et al. The multifaceted role of macrophage mitophagy in SiO2 -induced pulmonary fibrosis: a brief review. J. Appl. Toxicol. 44, 1854–1867 (2024).
Wu, Z. et al. Rab32 family proteins regulate autophagosomal components recycling. J. Cell Biol. 223, e202306040 (2024).
Xie, J.-H., Li, Y.-Y. & Jin, J. The essential functions of mitochondrial dynamics in immune cells. Cell Mol. Immunol. 17, 712–721 (2020).
Rius, J. et al. NF-κB links innate immunity to the hypoxic response through transcriptional regulation of HIF-1α. Nature 453, 807–811 (2008).
Von Schwedler, U. K. et al. The protein network of HIV budding. Cell 114, 701–713 (2003).
Etoh, K. & Fukuda, M. Rab10 regulates tubular endosome formation through KIF13A/B motors. J. Cell Sci. 132, jcs226977 (2019).
Tsuboi, T. & Fukuda, M. Rab3A and Rab27A cooperatively regulate the docking step of dense-core vesicle exocytosis in PC12 cells. J. Cell Sci. 119, 2196–2203 (2006).
Kuchitsu, Y., Homma, Y., Fujita, N. & Fukuda, M. Rab7 knockout unveiled regulated autolysosome maturation induced by glutamine starvation. J. Cell Sci. 131, jcs215442 (2018).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Yamamoto, Y. et al. ERdj8 governs the size of autophagosomes during the formation process. J. Cell Biol. 219, e201903127 (2020).
Hendel, A. et al. Chemically modified guide RNAs enhance CRISPR-Cas genome editing in human primary cells. Nat. Biotechnol. 33, 985–989 (2015).
Ohbayashi, N., Yatsu, A., Tamura, K. & Fukuda, M. The Rab21-GEF activity of Varp, but not its Rab32/38 effector function, is required for dendrite formation in melanocytes. Mol. Biol. Cell 23, 669–678 (2012).
Lu, S.-L. et al. Endothelial cells are intrinsically defective in xenophagy of Streptococcus pyogenes. PLoS Pathog. 13, e1006444 (2017).
Yoshii, S. R., Kishi, C., Ishihara, N. & Mizushima, N. Parkin mediates proteasome-dependent protein degradation and rupture of the outer mitochondrial membrane. J. Biol. Chem. 286, 19630–19640 (2011).
Acknowledgments
This study was funded by Grants-in-Aid for Scientific Research KAKENHI (20H05326, 22H04647, 23H02475) to T.N. and (20K15789, 25K09629) to S-L.L. This research was conducted as part of the All-Osaka U Research in “The Nippon Foundation - Osaka University Infectious Disease Response Project” to T.N. and Y.W., and Hirose Foundation to S-L.L.
Author information
Authors and Affiliations
Contributions
T.N. supervised this study. S-L.L. and T.N. designed and conceived this study. S-L.L., S.C., and K.N. performed the experiments. Y.L., C-Y.T., Y.K., Z.Z., B.C., K.T., and Z.T. assisted with the experiments. H.O. performed the electron microscopy analysis. C-Y.T., M.W., E.H., M.F., Y.W., E.M., N.U., and S.M. provided resources. All authors reviewed the results and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lu, SL., Chen, S., Noda, K. et al. Evidence that mitochondria in macrophages are destroyed by microautophagy. Nat Commun 16, 8123 (2025). https://doi.org/10.1038/s41467-025-63531-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63531-x