Abstract
Polygenic risk scores hold prognostic value for identifying individuals at higher risk of type 2 diabetes. However, further characterization is needed to understand the generalizability of type 2 diabetes polygenic risk scores in diverse populations across various contexts. We systematically characterize a multi-ancestry type 2 diabetes polygenic risk score among 244,637 cases and 637,891 controls across diverse populations from the Population Architecture Genomics and Epidemiology Study and 13 additional biobanks and cohorts. Polygenic risk score performance is context dependent, with better performance in those who are younger, male, without hypertension, and not obese or overweight. Additionally, the polygenic risk score is associated with various diabetes-related cardiometabolic traits and type 2 diabetes complications, suggesting its utility for stratifying risk of complications and identifying shared genetic architecture between type 2 diabetes and other diseases. These findings highlight the need to account for context when evaluating polygenic risk score as a tool for type 2 diabetes risk prognostication and the potentially generalizable associations of type 2 diabetes polygenic risk score with diabetes-related traits, despite differential performance in type 2 diabetes prediction across diverse populations. Our study provides a comprehensive resource to characterize a type 2 diabetes polygenic risk score.
Similar content being viewed by others
Introduction
The rise in type 2 diabetes (T2D) prevalence is a major public health challenge that shows no signs of abating. Without sufficient efforts to combat this pandemic, it has been estimated that 11.3% of the global population will have diabetes by 20301, at which time its economic burden is projected to be 2.2% of the global gross domestic product2. These predictions provide an urgent incentive to improve diabetes risk prediction, stratification, and prevention.
An emerging and promising approach to quantify T2D risk is the use of polygenic risk scores (PRS), which model the aggregate effect of genetic variants found across the genome on a disease or trait of interest. Such scores, combined with non-genetic predictors, potentially hold utility for identifying individuals at high risk prior to disease onset and could guide positive health behavior changes, screening practices, or therapeutic interventions3,4,5. These PRS have been facilitated by genome-wide association studies (GWAS) that have identified genetic variants conferring T2D susceptibility, with several recent T2D GWAS and fine mapping studies conducted across diverse populations enabling the development of multi-ancestry T2D PRS6,7,8.
Given the large number of risk factors for T2D, there is a clear need to further characterize PRS. Although previous studies have reported that interactions between individual genetic variants and certain lifestyle factors influence T2D risk in European ancestry populations8,9,10,11, little is known about whether the predictive accuracy and utility of T2D PRS are context-dependent and modified by individual characteristics and metabolic parameters, particularly across diverse populations. Moreover, it remains unclear whether PRS for T2D risk could hold prognostic value for diabetes-related complications across numerous organ systems and whether such associations are consistent across populations. While a recent investigation found that a T2D PRS was associated with T2D-related retinopathy, with modest evidence of association with chronic kidney disease, peripheral artery disease, and neuropathy in individuals who were genetically similar to European ancestry7, no study to date has systematically evaluated diabetes-related complications across diverse populations. A thorough investigation of these and other complications across diverse populations could provide biological insights into the pleiotropic nature and prognostic ability of T2D PRS.
The Population Architecture using Genomics and Epidemiology (PAGE) Study, which consists of multiple large and diverse population-based studies, is uniquely situated to address these questions. In PAGE, we previously found that a multi-ancestry T2D PRS outperformed population-specific T2D PRS and explained up to 15.3% of familial relative risk of T2D across populations8. Here, after identifying a multi-ancestry T2D PRS that was optimally predictive of T2D risk across our diverse PAGE populations, we investigated whether the effect of a T2D PRS was context-dependent across PAGE and several independent biobanks and cohorts. To further understand the clinical implications of genetic risk of T2D on other diseases and conditions, we examined whether a T2D PRS was associated with T2D-related phenotypic risk factors and complications and additionally conducted phenome-wide association studies (PheWASs) across diverse populations. The goal of this investigation is to improve precision medicine in diabetes by systematically evaluating the performance and context-dependent effects of a T2D PRS across diverse populations. Our findings serve as a comprehensive resource to characterize T2D PRS.
Results
Participants
An overview of the study design is provided in Fig. 1. In total, 82,944 participants from PAGE and 835,241 participants from 13 additional biobanks and cohorts were included in this study: 244,637 T2D cases, 637,891 controls, and 35,657 individuals with prediabetes. This included individuals who self-reported as African, African American, or Black (AFR); American Indian, Alaskan Native, or Native American (AI); Asian American, East of Southeast Asian, or South Asian/Indian (ASN); White or European American (EUR); Greenlandic; Hispanic/Latino American (HIS); Middle Eastern, North African, or Qatari (MENAQ); and Native Hawaiian or Other Pacific Islander (NH) (Table 1, Supplementary Data 1 and 2, see the “Methods” section, Supplemental Information for population descriptor details).
Four T2D PRS were constructed and evaluated in the Population Architecture using Genomics and Epidemiology (PAGE) study to identify the best-performing score for downstream analyses. The best-performing PRS across populations was used for stratified analyses across demographic, medical, and lifestyle and behavioral characteristics, and association analyses with diabetes-related traits in PAGE. Stratified and association analyses were replicated in 11 independent cohorts. Phenome-wide association studies (PheWAS) were conducted across five independent biobanks to assess the broader clinical impact of the T2D PRS.
Evaluation of T2D PRS performance
Given our unique populations and how rapidly PRS are superseded, our first goal was to identify a T2D PRS that had optimal predictive ability across our populations to be carried forward in subsequent PRS evaluations. In PAGE, we assessed four distinct multi-ancestry PRS (Fig. 1, Supplementary Data 3, see the “Methods” section). The first three PRS were genome-wide PRS developed using 1.259 million HapMap3 variants in PRS-CSx12, an approach that has demonstrated high predictive ability across several traits13,14,15. PRS Method 1: a previously developed T2D PRS16 based on AFR17, ASN18, and EUR19 GWAS summary statistics using meta-analyzed posterior effects with the META = T function in PRS-CSx; PRS Method 2: a new T2D PRS that we developed using a similar approach as PRS Method 1 but leveraging GWAS summary statistics from larger sample sizes of AFR, ASN, and EUR populations and also including HIS and South Asian (SAS) populations6,7; PRS Method 3: a new T2D PRS that we developed using the same summary statistics as PRS Method 2 but calculated from a linear combination of population-specific PRS using the META = F function in PRS-CSx; and PRS Method 4: a previously developed PRS with 582 genome-wide significant variants reported in AFR, ASN, EUR, and HIS populations7,8. PRS derived from each method was standardized to the distribution of controls within each population.
Each of the four T2D PRS methods exhibited good predictive performance in PAGE (Supplementary Data 4). Although confidence intervals of the area under the receiver operating characteristic curve (AUC) overlapped between the methods, PRS Method 1 showed slightly better predictive performance across most populations compared to other methods. PRS Method 1 had AUC improvements of 1.8%, 4.0%, 5.2%, 4.8%, 4.5%, and 4.1% for AFR, AI, ASN, EUR, HIS, and NH populations, respectively, compared to the base model of age, sex, BMI, and the first 10 genetic principal components (PCs) (Fig. 2A and Supplementary Data 4). We used PRS Method 1 for all subsequent analyses.
A Area under the curve (AUC) for four T2D PRS methods in PAGE. PRS Methods 1–4 correspond to those in Fig. 1. Data are presented as AUC with 95% confidence interval (CI). B Association between each SD unit increase in the Ge et al. T2D PRS with T2D risk in PAGE and the additional biobanks and cohorts. Data are presented as odds ratios (OR) with 95% CI. Sample sizes for A and B are provided in Supplementary Data 1.
In PAGE, the PRS was highly predictive of T2D risk across populations but with variable performance (Supplementary Fig. 1). Each standard deviation (SD) unit increase in PRS was associated with 1.78-fold (95% CI: 1.75, 1.82) higher odds of T2D across all populations combined, ranging from 1.45 (95% CI: 1.39, 1.50) in AFR to 3.51 (95% CI: 2.41, 5.10) in AI (Fig. 2B and Supplementary Data 5). Compared to individuals in the 40–60% PRS category, the top 90–100% PRS decile was associated with 2.37 (95% CI: 2.24, 2.51) increased odds of T2D across all populations combined, ranging from 1.75 (95% CI: 1.54, 2.00) in AFR to 6.96 (95% CI: 2.25, 21.56) in AI (Supplementary Fig. 2). Effects were generally similar in the 13 additional biobanks and cohorts within each population (Fig. 2).
As a secondary analysis, we evaluated the association between the T2D PRS and age at T2D diagnosis (Fig. 3 and Supplementary Data 6). Higher T2D PRS was significantly associated with younger age at diagnosis across populations. Individuals with T2D in the top 10% PRS decile were diagnosed, on average, 8.9 years earlier (95% CI: –9.83 to –7.89; P = 1.09 × 10−78) than those in the bottom 10% (Supplementary Data 6). This pattern was consistent across populations, with individuals in the top 10% PRS decile diagnosed on average 3.79 years earlier in AFR (P = 1.48 × 10−9), 10.17 years earlier in ASN (P = 1.67 × 10−3), 2.46 years earlier in EUR (P = 3.33 × 10−4), and 4.86 years earlier in HIS (P = 8.75 × 10−7) populations compared to those in the bottom 10% decile.
Data are presented as mean age at diagnosis with standard error. Sample sizes are provided in Supplementary Data 6.
Context-dependent effects of T2D PRS
We investigated whether the T2D PRS demonstrated context-dependent effects on T2D risk by conducting association analyses stratified by 18 demographic, medical, lifestyle, and behavioral characteristics, and evaluating effect heterogeneity using Cochran’s Q-test and interaction terms in regression models (see the “Methods” section and Supplementary Data 7). Across PAGE and the additional biobanks and cohorts, we conducted stratified analyses in each of four populations (AFR, ASN, EUR, HIS) with sufficient sample sizes, as well as in all populations combined. We consistently found significant differences in T2D PRS effects by age groups, sex, hypertension status, and obesity status within and across populations (Fig. 4 and Supplementary Data 8–12).
Data are presented as odds ratio (OR) with 95% confidence interval (CI). P-values of heterogeneity from a two-sided Cochrane Q-test are indicated when differences were statistically significant (P < 0.05). A Demographic characteristics and medical history factors. B Behavioral and lifestyle factors. C Medication use. D Lipids. Sample sizes for A–D are provided in Supplementary Data 9.
Specifically, in meta-analyses across all populations, the T2D PRS was associated with significantly higher risk of T2D in younger individuals (age ≤50 years: OR = 1.91 (95% CI: 1.88, 1.94); age 50–60 years: OR = 1.85 (95% CI: 1.83, 1.88); age 60–70 years: OR = 1.81 (95% CI: 1.79, 1.83); age >70 years: OR = 1.68 (95% CI: 1.66, 1.71); Phet = 4.20 × 10−28). The T2D PRS also had a significantly larger effect in males compared to females (males: OR = 1.91 (95% CI: 1.90, 1.92); females: OR = 1.74 (95% CI: 1.72, 1.76); Phet = 2.00 × 10−38). The effect of T2D PRS was slightly higher in individuals without hypertension (without hypertension: OR = 1.87 (95% CI: 1.85, 1.90); with hypertension: OR = 1.83 (95% CI: 1.82, 1.85); Phet = 8.30×10−03). The effect of the T2D PRS was attenuated in those with BMI categorized as obese (non-overweight/obese: OR = 1.89 (95% CI: 1.86, 1.92); overweight: OR = 1.96 (95% CI: 1.94, 1.98); obese: OR = 1.76 (95% CI: 1.75, 1.78); Phet = 9.19 × 10−56). Although no significant PRS performance difference by family history (FH) of T2D was observed in meta-analyses across all populations, a stronger effect was observed among those with a FH of T2D in AFR populations (FH: OR = 1.56 (95% CI: 1.50, 1.63); without FH: OR = 1.49 (95% CI: 1.46, 1.52); Phet = 0.04) (Fig. 4, Supplementary Data 9). In analyses performed separately in PAGE versus the additional biobanks and studies, evidence of replication was observed for T2D PRS effects when stratifying by age groups, sex, hypertension and obesity status (Supplementary Fig. 3, Supplementary Data 10 and 11), as summarized in Supplementary Data 8. AUCs calculated separately in each stratum further supported these findings (Supplementary Data 10).
PRS had a stronger effect in never smokers and former smokers compared to current smokers (Phet = 4.08 × 10−18) and in physically active individuals compared to inactive individuals (Phet = 6.14 × 10−3) in meta-analyses across all populations, although these findings were not consistent when examining each population separately. Additionally, the PRS effect was stronger in statin and lipid-lowering medication users than non-users, though the differential association is complicated by higher statin use among diabetes cases and incomplete documentation of lipid medication use. Findings were inconsistent for other medication use, lipids, and current drinking status.
Association between T2D PRS and diabetes-related traits
Next, to investigate the clinical implications of genetic risk of T2D, we examined associations between the T2D PRS and 20 diabetes-related traits across five primary phenotype categories: glycemic traits, cardiometabolic traits, vascular diseases, kidney diseases and functions, and inflammatory biomarkers, stratified by T2D status in AFR, ASN, EUR, HIS, and all populations combined (see the “Methods” section and Supplementary Data 13). Significant associations (applying a P < 2.50 × 10−3 [0.05/20 traits] Bonferroni adjusted threshold) were found across various glycemic and cardiometabolic traits, including HbA1c, fasting glucose, hypertension, systolic blood pressure (SBP), diastolic blood pressure (DBP), triglycerides, non-HDL cholesterol, and HDL cholesterol (Fig. 5, Supplementary Data 14), with consistent results observed in analyses performed separately in PAGE versus the additional biobanks and studies (Supplementary Data 15 and 16). AUC and R2 estimates further supported these findings (Supplementary Data 15).
The X-axis represents the beta estimates and 95% confidence intervals (CIs) of PRS for continuous traits and odds ratios (OR) and 95% CIs of PRS for binary traits. P-values shown on each plot are exact values derived from two-sided linear regression models for continuous traits and two-sided logistic regression models for binary traits. Solid circles indicate significant associations that passed the Bonferroni-adjusted P-value threshold of P < 2.50 × 10−3, while open circles indicate associations that were not significant. Panels show associations for the T2D PRS and A glycemic traits in T2D controls and B cardiometabolic traits in T2D cases (left), controls (middle), and individuals with prediabetes (right). The units of continuous traits are listed in Supplementary Data 13. Sample sizes are provided in Supplementary Data 14. Results for vascular disease, kidney function related traits, and inflammatory biomarkers are shown in Supplementary Fig. 4.
We first examined the associations between T2D PRS and glycemic traits among T2D controls, as glycemic measurements in T2D cases are influenced by diabetes medications. Each SD increase in the T2D PRS was significantly associated with 0.034 (95% CI: 0.031, 0.037) higher units of HbA1c levels and 0.030 mmol/L (95% CI: 0.026, 0.035) higher fasting glucose levels in individuals without T2D (Fig. 5A). These findings were consistent across populations and studies. Additionally, the PRS demonstrated a significant association with log transformed HOMA-IR (P = 5.19 × 10−4), although this was observed only in the EUR population.
Regarding cardiometabolic traits, each SD increase in PRS was associated with a 1.04-fold increased risk (95% CI: 1.03, 1.05) of hypertension in T2D controls across all populations, a 1.04-fold (95% CI: 1.02, 1.06) and 1.03-fold (95% CI: 1.02, 1.04) increased risk in EUR cases and controls, and a 1.10-fold (95% CI: 1.07, 1.13) increased risk in HIS controls (Fig. 5B). PRS was also positively associated with SBP (P = 8.24 × 10−25), DBP (P = 2.72 × 10−6), log-transformed triglycerides (P = 1.13 × 10−68), and non-HDL cholesterol (P = 1.50 × 10−5), and negatively associated with HDL cholesterol (P = 1.50 × 10−5) in T2D controls across all populations.
For kidney diseases, the PRS was associated with a 1.08-fold increased risk (95% CI: 1.03, 1.13) of end stage renal disease (ESRD) in T2D cases across all populations (Supplementary Fig. 4B). However, PRS was inversely associated with a 0.87-fold decreased risk (95% CI: 0.80, 0.95) of chronic kidney disease (CKD) in ASN controls. The PRS was not consistently associated with any vascular diseases or inflammatory markers.
T2D PRS PheWAS
To further investigate the prognostic value of the T2D PRS and clinical phenotypes, we conducted a comprehensive PheWAS meta-analysis of the T2D PRS on 1,815 unique phenotypes representing 17 disease categories tested across five biobanks. We identified significant associations between the T2D PRS and 732 traits (40.3% of all phecodes tested) across all populations combined at Bonferroni adjusted significance (P < 2.75 × 10−5), as well as 213 (12.0%), 43 (3.7%), 689 (38.0%), and 232 (13.7%) significant traits in AFR, ASN, EUR, and HIS populations, respectively (Fig. 6, Supplementary Fig. 5, and Supplementary Data 17). All significant traits identified in AFR, ASN, and HIS and were also significant in the EUR population, which had an additional 428 significant traits that were not significantly associated in any other population (Fig. 7). The PRS-PheWAS effect estimates (ORs) in EUR were reasonably correlated with AFR (r = 0.68), ASN (r = 0.64), and HIS (r = 0.73) populations, with consistent effect directions. Effects had slightly lower correlations between AFR and HIS (r = 0.58), ASN and AFR (0.58), and ASN and HIS (0.59), likely reflecting the larger sample size in the EUR population, which allowed more accurate estimates of effects and thus more overlap with other populations.
The X-axis represents phecodes color-coded by their corresponding phenotype category and is ordered by the category with the most to the category with the least significant hits. The Y-axis represents the −log10(p-value) derived from two-sided logistic regression models. The red horizontal line represents Bonferroni-adjusted p-value threshold P < 2.75 × 10−5 (732 phecodes meet this threshold), and the blue horizontal line represents an unadjusted p-value threshold of P < 0.05 (1120 phecodes meet this threshold). Upward triangles indicate positive associations, while downward triangles indicate negative associations. The top ten most significant associations from the endocrine/metabolic category are annotated, with the single most significant association shown in black and others shown in gray, while the single most significant association from each other category is annotated.
A Venn diagram of significant PheWAS results across populations. B Comparison of odds ratios from T2D PRS PheWAS results across populations.
The 732 significant associations encompassed phenotypes related to the circulatory system (n = 123, 16.8%), endocrine and metabolic functions (n = 90, 12.3%), digestive system conditions (n = 82, 11.2%), sense organs (n = 65, 8.9%), genitourinary system conditions (n = 58, 7.9%), and 12 other disease categories (n = 314, 42.9%). In analyses conducted across all populations combined, T2D (OR = 2.38; P-value < 5.0 × 10−324) demonstrated the most significant association, followed by T2D-related phenotypes from the endocrine/metabolic phenotype group, including diabetic retinopathy, insulin pump user, polyneuropathy in diabetes, and diabetes mellitus (Table 2 and Fig. 6). We observed consistently strong associations overall and across populations between the T2D PRS and type 1 diabetes (T1D) risk, with each SD increase in the T2D PRS associated with 1.63-fold (P = 1.31 × 10−173), 1.47-fold (P = 1.31 × 10−173), 1.83-fold (P = 2.97 × 10−18), 1.63-fold (P < 5.0 × 10−324), and 1.76-fold (P = 4.41 × 10−128), and increased risk of T1D in the overall, AFR, ASN, EUR, and HIS populations, respectively.
The single top phenotype associated with the T2D PRS from other phenotype groups, all of which were positively associated, were essential hypertension, CKD, senile cataract, chronic ulcer of leg or foot, anemia in chronic kidney disease, acute osteomyelitis, other peripheral nerve disorders, open wound of foot, respiratory failure, edema, gastroparesis, tobacco use disorder, diabetes or abnormal glucose tolerance complicating pregnancy disorder, benign neoplasm of colon, dermatophytosis, and cardiac congenital anomalies.
Discussion
In this large-scale T2D investigation of more than 240,000 T2D cases and 630,000 controls from PAGE and 13 independent biobanks and studies, we performed the first comprehensive and systematic cross-population characterization of a multi-ancestry genome-wide T2D PRS. This was achieved using a harmonized framework to evaluate context-dependent T2D PRS effects across different demographic and clinical characteristics, T2D PRS associations with diabetes-related traits, and a PheWAS of the T2D PRS. Our findings demonstrate that the PRS holds predictive value for T2D risk across diverse populations and was associated with younger age at T2D diagnosis. The T2D PRS demonstrated context-dependent associations with T2D risk, with PRS performance varying by certain demographic and clinical characteristics, such as age, sex, hypertension, and obesity status. Additionally, we found that the genetic risk of T2D, as measured by the T2D PRS, was associated with other glycemic and cardiometabolic traits in controls, suggesting that PRS could provide insights into T2D etiology and serve as a valuable tool for predicting dysglycemia and metabolic risk, thereby facilitating early interventions and improving clinical risk assessment. Notably, associations between the T2D PRS and diabetes-related traits and complications were often consistent across populations, despite variable between-population PRS performance in estimating T2D risk, suggesting that the PRS may have clinical and prognostic utility beyond T2D risk prediction. Our PheWAS found associations between the T2D PRS and 40.3% of tested phenotypes, including phenotypes across all disease categories tested, underscoring the pleiotropic effects of T2D genetic risk and suggesting a broad biological impact of T2D-associated variants across organ systems. By providing a comprehensive resource that characterizes T2D PRS across diverse populations, our findings move beyond European-centric discovery and evaluate whether PRS utility—both predictive and prognostic—depends on individual- and population-level contexts. Future applications of T2D PRS in precision medicine should consider not only ancestry-specific calibration but also how environmental, metabolic, and clinical factors modulate the phenotypic expression of genetic risk.
We found that a multi-ancestry PRS was strongly associated with increased risk of T2D in all populations examined, yielding an overall OR of 1.75 (95% CI: 1.71, 1.78) and AUC of 0.641 (95% CI: 0.636, 0.645). The PRS performance across studies was generally comparable, with small variations likely due to study-specific differences, such as study design and phenotype definition. However, we observed larger variations in the performance of T2D PRS across populations, with the highest predictive performance in the AI population (although sample sizes were small), followed by EUR, ASN, NH, HIS, and AFR populations. These findings are consistent with what has been previously reported8,16 and could be attributed to various population-specific factors that are likely not strongly represented in the discovery GWAS used to construct the T2D PRS20. This notably underscores the importance of expanding GWAS and fine-mapping efforts, particularly in AFR populations, to improve the accuracy and transferability of T2D PRS.
Despite the lower predictive ability of the T2D PRS in AFR and HIS populations, we observed the strongest evidence of the PRS being associated with a younger age at T2D diagnosis in these two populations, demonstrating potential clinical utility. AFR and HIS populations tended to develop diabetes earlier than ASN and EUR populations; while these differences could be partially attributed to the younger age at enrollment in these populations (mean age at enrollment: AFR = 54.7 [SD: 10.2], HIS = 53.3 [SD: 11.4], ASN = 54.5 [SD: 12.5], EUR = 60.5 [SD: 12.5]), similar differences in age at T2D diagnosis have been previously reported21. As such, these differences may reflect complex interactions between genetic and non-genetic factors that may warrant further studies to understand the causes and mechanisms of earlier T2D onset in these groups.
Our results show that the prediction accuracy of PRS is context-dependent and can vary even within populations. We consistently observed that the T2D PRS was particularly informative for predicting T2D susceptibility in younger individuals, those without hypertension, males, and individuals who were not obese or overweight, across and within populations. The stronger predictive ability in younger individuals is consistent with published literature, such as the findings from the Framingham Offspring Study22 and the EPIC InterAct Study9, both of which observed a higher relative effect of T2D PRS among individuals under 50 years old compared to those over 50. However, these studies have been limited to T2D PRS developed and evaluated in individuals predominantly of European descent. Similar trends have also been observed in other traits, including prostate cancer23,24,25, colorectal cancer26,27, estrogen receptor-positive breast cancer28, and coronary artery disease29. The stronger genetic influence on disease onset at a younger age is expected, as older individuals have had more time to accumulate environmental risk factors that lead to metabolic impairments, thereby impacting disease risk. The larger effect of T2D PRS in leaner adults and those without hypertension could similarly be attributed to a lower accumulation of environmental risk factors in these individuals, suggesting a higher contribution of genetic risk factors to T2D development, a pattern also shown in several previous studies8,9,30.
Prior research on the performance of T2D PRS stratified by sex is limited9,31, and evidence of sex-based differences in T2D heritability estimates is inconsistent32,33,34,35. However, sex-based differences in PRS prediction have been widely reported across a range of complex traits36,37,38. In this study, we observed stronger predictive performance of the T2D PRS in males. This difference could be attributed to several factors. First, certain genetic variants exhibit sex-dimorphic effects, with some loci demonstrating larger or more penetrant effects in males—for example, variants at IRS1 show male-specific effects on fasting insulin and the 11p15.5 region, which harbors genes involved in insulin production such as KCNQ1, has been implicated in male-specific T2D risk, potentially influenced by maternal inheritance patterns39,40,41,42. Second, sex-specific biological mechanisms, including differences in glucose and lipid metabolism, affect fat distribution and insulin sensitivity, which can modulate the penetrance of these genetic variants41,42. Additionally, diagnosis bias may lead to T2D being more frequently diagnosed in males41. Future sex-specific GWAS and gene-by-sex interaction studies are needed to further investigate sex-specific genetic factors in T2D.
Collectively, these interactions between the T2D PRS and non-genetic factors demonstrate the complex nature of T2D risk and the importance of interpreting the genetic risk of T2D in the context of non-genetic risk factors. Our findings suggest that across diverse populations, genetic effects can vary across contexts and that PRS may exhibit context-specific performance. This indicates that risk prediction models will need to include not only PRS but demographic, medical, lifestyle and behavioral characteristics, and their interactions36. Moreover, any potential application of PRS in clinical care will need to consider such contextual factors.
We found that the T2D PRS was associated with various diabetes-related traits and disease conditions across diverse populations, providing compelling evidence that the T2D PRS could provide insights into T2D prognosis. We observed consistent associations between T2D PRS and glycemic parameters in T2D controls, with significant positive associations for fasting glucose and HbA1c, as well as associations with cardiometabolic traits, including hypertension, SBP, triglycerides, and HDL cholesterol, in both T2D cases and controls. These findings suggest that T2D PRS may serve as a valuable tool for early identification of individuals at risk for developing dysglycemia and cardiometabolic complications, even among those who are currently asymptomatic or undiagnosed. Our findings also have important implications for T2D etiology and pathophysiology. For instance, the positive association between T2D PRS and HOMA-IR in EUR controls provides genetic evidence that complements previous epidemiological studies on the critical role of insulin resistance in T2D development43,44,45. This association suggests that genetic factors contribute to insulin resistance pathways even before clinical diabetes onset, offering mechanistic insights into how genetic predisposition translates into disease risk. The inverse association we observed between T2D PRS and CKD in ASN controls warrants further investigation, as it could reflect selection bias where individuals with higher genetic risk for T2D who remain diabetes-free may represent a healthier subset, possibly benefiting from protective environmental or lifestyle factors that also lower their CKD risk.
Findings from our PheWAS across 1815 diseases and conditions and five biobanks further demonstrated significant associations between T2D PRS and various clinical comorbidities, including those related to the circulatory system, endocrine and metabolic disorders, lipid abnormalities, and kidney diseases. Notably, these associations were consistently observed across populations, with fairly correlated effect estimates. Despite the variability in PRS performance for T2D risk estimation across populations, our findings suggest that genetic mechanisms underlying T2D and diabetes-related traits are shared across populations. As such, the PRS may serve as a valuable tool for stratifying the risk of complications in individuals diagnosed with T2D across diverse populations. The positive association between T2D PRS and micro- and macrovascular complications, such as retinopathy, neuropathy, CKD, cardiovascular diseases, and peripheral artery disease, has also been reported by prior studies7,46,47. The significant positive associations between the T2D PRS and T1D risk from PheWAS may indicate the potential misclassification of T2D and T1D in clinician-recorded diagnoses, given the overlapping clinical features of T2D and T1D48, or it could reflect genetic consistencies between the two diseases. This study complements prior genetics research on T2D by leveraging large sample sizes to identify significant associations between T2D PRS and various traits and conditions not previously reported. Our findings imply that the T2D PRS holds promise as a powerful tool for precision medicine, risk stratification, and disease management in the context of T2D, providing an opportunity to identify individuals at elevated risk of developing T2D and its complications. Further, the PheWAS results reported here serve as a comprehensive catalog of phenotypic associations with T2D PRS, contributing to our understanding of T2D-related comorbidity patterns. These findings could offer a valuable resource for hypothesis generation, disease subtype characterization, and the identification of shared biological pathways and molecular mechanisms.
The results of our study should be interpreted considering several limitations. We used self-identified race and ethnicity (SIRE) population descriptors to determine whether the T2D PRS had discrepancies in performance in the contexts of SIRE. Although other sociodemographic and environmental factors may be more informative than SIRE, because our investigation included a large number of studies, many of which include legacy data collected up to 37 years ago, this study was limited by the variables available across all included studies. Further, SIRE information was collected differently between studies (Supplementary Data 2), which could influence PRS performance between populations. In addition, this study included both incident and prevalent T2D. Future research should replicate these findings by investigating the association between PRS and risk of incident T2D and the other outcomes reported here. Further, most phenotypes and clinical measurements, such as glucose levels and BMI, were collected at baseline, when these measurements are typically available for most individuals. This limits our ability to capture changes over time and assess the dynamic nature of these associations.
In conclusion, our large-scale, systematic and cross-population investigation highlights the predictive value of the T2D PRS across diverse populations, emphasizing its potential as a promising tool for identifying individuals at elevated risk of developing T2D and its associated comorbidities and complications, shedding light on potential avenues for risk prediction and personalized management strategies. Our findings also demonstrate that the T2D PRS has context-dependent effects, indicating the need to account for these characteristics to interpret PRS and improve PRS prediction accuracy. Further research is warranted to evaluate and optimize the clinical utility of T2D PRS across diverse populations.
Methods
Participants
The PAGE Study was developed to conduct genetic epidemiological research in ancestrally diverse populations within the United States49. PAGE includes individuals from the following ongoing population-based biobanks and cohorts: Atherosclerosis Risk in Communities (ARIC) Study50, the Icahn School of Medicine at Mount Sinai BioMe biobank in New York City (BioMe), Coronary Artery Risk Development in Young Adults Study (CARDIA)51, Hispanic Community Health Study/Study of Latinos (HCHS/SOL)52,53, Multiethnic Cohort Study (MEC)54, and Women’s Health Initiative (WHI)55. A detailed description of PAGE has been described49 with additional detail summarized in the Supplemental Information and Supplementary Data 1.
Analyses were also conducted in 13 additional biobanks and cohorts that were not included in the discovery GWAS summary statistics of the best-performing T2D PRS. Studies included All of Us (AoU)56, BioMe (participants were non-overlapping with those from BioMe included in PAGE), BioVU Biobank (BioVU)57, Cameron County Hispanic Study (CCHC)58, Cebu Longitudinal Health and Nutrition Survey (CLHNS)59, China Health and Nutrition Survey (CHNS)60, Colorado Center for Personalized Medicine Biobank (CCPM)61, Greenlandic Health Surveys (Greenlandic)62,63,64, Multi-Ethnic Study of Atherosclerosis Study (MESA)65, Michigan Genomics Initiative (MGI)66, Million Veteran Program (MVP)67,68,69, Qatar Biobank (Qatar)70, and REasons for Geographic and Racial Differences in Stroke Study (REGARDS)71 (Supplementary Data 1). Analyses performed in these additional studies were dependent on the availability of variables and data needed to derive variables.
Our research complies with all relevant ethical regulations regarding the use of human study participants and was conducted in accordance to the criteria set by the Declaration of Helsinki. All participants provided informed consent, and each study was approved by the appropriate Institutional Review Board at their respective institutions. Additional information about each participating study can be found in the Supplementary Information.
Use of population descriptors
Self-identified race and ethnicity (SIRE) population descriptors were used across studies, with the goal of studying how PRS performance may vary across the contexts of SIRE. Although other sociodemographic and environmental factors are expected to be more informative than SIRE, this investigation is limited by the variables that are currently available across all included studies. It is important to note that SIRE does not serve as a proxy for genetic ancestry, holds no biological basis, and does not imply a biological explanation for health disparities72.
We standardized population descriptors across all studies in the analysis by using consistent acronyms. Specifically, AFR refers to African, African American, or Black participants; AI refers to American Indian, Alaskan Native, or Native American; ASN includes Asian American, East or Southeast Asian (e.g., from China, Japan, Korea, Indonesia, the Philippines), as well as South Asian/Indian (e.g., from India, Pakistan); EUR refers to non-Hispanic White American or European individuals; Greenlandic refers to individuals of Greenlandic or mixed Greenlandic-Danish heritage; HIS represents Hispanic/Latino American participants; MENAQ encompasses Qatari and Arab American individuals; and NH refers to Native Hawaiian or Other Pacific Islander. The Greenlandic population was considered separate from others, as this population represents substantial Inuit ancestry63. Detailed definitions of population descriptors used in each study are provided in Supplementary Data 2.
T2D and prediabetes definition
T2D and prediabetes definitions were created based on the American Diabetes Association’s 2021 Standards of Medical Care in Diabetes73. In PAGE, participants were classified as T2D cases if they met at least one of the following criteria: adults ≥25 years old with (1) a T2D diagnosis by a physician/medical professional or use of medication for treatment of diabetes, (2) a fasting (≥8 h) plasma glucose ≥ 126 mg/dl, (3) a random glucose ≥ 200 mg/dl, or (4) an HbA1c ≥ 48 mmol/mol (6.5%). Cases were restricted to those with either an age at diagnosis or most recent age at glucose or HbA1c draw ≥25 years to avoid misclassifying T1D cases as T2D cases. Individuals with prediabetes were defined as adults ≥18 years old who did not meet the definition of T2D and with (1) a fasting plasma glucose between 100–125 mg/dl, (2) an HbA1c between 39 and 46 mmol/mol (5.7–6.4%), or (3) a 2-h oral glucose tolerance test between 140 and 199 mg/dl. T2D controls were defined as adults ≥ 40 years old who did not meet the definitions of T2D or prediabetes. Some studies, however, used their own in-house definitions and did not identify individuals with prediabetes. Definitions used in the additional biobanks and cohorts were similar to those used in PAGE and are summarized in the Supplemental Information.
Genotyping, imputation, and quality control
This study included 36,724 PAGE participants who met the T2D and prediabetes definitions and underwent genotyping using the Multi-Ethnic Genotyping Array (MEGA) panel, specifically designed to capture genetic heterogeneity across diverse populations, with the remaining 46,220 PAGE participants genotyped using a variety of other arrays (Supplementary Data 1)74. Data were imputed using the NHLBI Trans-Omics for Precision Medicine (TOPMed) Imputation Server (https://imputation.biodatacatalyst.nhlbi.nih.gov/#!) using TOPMed-r2 as reference panel, Eagle v2.4 for phasing, and Minimac4 for imputation69,75,76. Variants were excluded if the imputation info score was <0.4 or if the effective sample size was <30. The effective sample size was calculated as 2 × MAF ×(1−MAF)× N × INFO, where MAF represents the minor allele frequency, N is the sample size, and INFO is the imputation quality score. Principal component analysis was performed separately for each study. Principal components (PCs) were generated using EIGENSOFT77,78,79. The top 10 PCs were used to account for population structure in subsequent analyses.
Construction of T2D PRS
Participants included in this investigation were not part of the discovery GWAS used to develop the T2D PRS. PRS for each participant were constructed as the weighted sum of effect alleles, weighted by variant-specific log ORs using “--score” option with “cols = +scoresums” modifier in PLINK 2.080. All T2D PRS were standardized by subtracting the mean PRS and dividing by the standard deviation of PRS calculated in controls within each population. Therefore, 1-unit increase in PRS corresponds to a 1-standard deviation increase within each population.
The R package “rtracklayer”81 was used to align and convert between the GRCh37 (hg19) and the GRCh38 (hg38) genome reference builds across genetic datasets. In the function “GRangesFromDataFrame”, we included a strand field (strand.field = “strand”), which generated the output built with a strand column. If the forward (plus) strand became the reverse (minus) strand in the new build82, the strand column would change from “+” to “−”. The alleles were on the new build, then changed to the complement of the original input build.
PRS-CSx estimates population-specific and multi-ancestry PRS by jointly modeling discovery GWAS summary statistics and pre-calculated linkage disequilibrium (LD) reference panels based on the 1000 Genomes Project (1KG) Phase 3 participants from AFR, Admixed American (AMR), EAS, EUR, and SAS population samples12. The first three PRS methods used the PRS-CSx “auto” option, which automatically learns the global shrinkage parameter ϕ from the discovery summary statistics. The META = T option in PRS-CSx was used to generate multi-ancestry PRS using an inverse-variance-weighted meta-analysis of the population-specific posterior effect size estimates (PRS Method 1 and PRS Method 2). The META = F option was used to generate five population-specific PRS, which were subsequently calculated in PAGE, standardized to a mean of 0 and SD of 1 by population and study, and used to calculate a linear combination PRS (PRS Method 3). In PRS Method 3, for each population and study in PAGE, the dataset underwent a three-fold cross-validation process, where the data was evenly divided into three equally sized subsets. In each iteration, one dataset was selected as the testing dataset, while the remaining two were used as the validation dataset. This process was repeated three times to evaluate all possible combinations of validation and testing datasets, so that each individual participated in both the validation and testing sets.
The following logistic regression model of the standardized population-specific PRS was fit in the validation dataset for each population:
where \({{\rm{l}}}{{\rm{o}}}{{\rm{g}}}{{\rm{i}}}{{\rm{t}}}(p)\) is the log odds of having T2D and β0 is the intercept, with each subsequent β representing the regression coefficient for each population-specific PRS. If any of the regression coefficients were <0, the corresponding population-specific PRS and regression coefficient were removed from the regression model, and the model was refitted until all remaining coefficients were >0. The regression coefficients learned in the validation dataset were then used as weights \(\left({w}_{k}\right)\) in the testing datasets to calculate the final linear combination PRS for each individual:
We assessed the predictive accuracy of the four T2D PRS within each population in PAGE using AUCs. AUC values were calculated using the R package “pROC”83 with logistic regression of T2D risk after adding PRS to a base model of age, sex, BMI, the first ten PCs of ancestry, and study. The best-performing PRS was calculated in the additional biobanks and cohorts (that were independent of the discovery GWAS used to develop the best-performing PRS) and was used for all subsequent analyses.
Association between T2D PRS and age at T2D diagnosis
We evaluated the association between the T2D PRS and age at T2D diagnosis in a case-only analysis in All of Us. This analysis was limited to All of Us due to its large sample size, inclusion of multiple populations, and the ability to estimate age at diagnosis. Age at diagnosis was determined by the earliest occurrence start date of T2D mellitus (SNOMED concept ID: 201826), T2D mellitus without complication (SNOMED concept ID: 4193704), or disorder due to T2D mellitus (SNOMED concept ID: 443732). We tested the association between each PRS category (predictor) and age at diagnosis (outcome) in linear regression models, using the 0–10% PRS decile as the reference. Analyses were performed separately in each population and across all populations.
Stratification analyses
We investigated the context-dependent effects of the best-performing T2D PRS in analyses stratified by demographic characteristics and medical history (age group, self-reported sex at birth, family history (FH) of T2D, and blood pressure categories), behavioral and lifestyle factors (obesity status, current drinking status, smoking status, and physical activity level), medication use (statin, NSAID, fibrates, lipid-lowering medication, and anti-hypertensive medication), and lipid traits (HDL, LDL, triglycerides, total cholesterol (TC), and non-HDL quartiles) (definitions provided in Supplementary Data 7). Analyses were conducted in all populations combined and separately in each population using logistic regression models that predict T2D status, adjusting for age, sex (except in sex-stratified analyses), BMI, the first ten PCs, and study. AUCs were also calculated within each stratum using similar models, as well as models excluding PRS and models of PRS alone. The Cochrane Q-test was used to determine whether effect heterogeneity between subgroups was sufficiently large enough to be attributed to factors beyond sampling error. The P-value from the Cochrane Q-test is referred to as \({P}_{{{\rm{het}}}}\). A P < 0.05 threshold was used to determine statistical significance for stratification analyses. We further evaluated evidence of context-dependent effects of PRS within PAGE by including an interaction term between the PRS and each context variable, also adjusting for the main effects of each. A likelihood ratio test was used to compare models with and without the interaction term.
T2D PRS associations with diabetes-related traits
We next examined the ability of the best performing T2D PRS to predict 20 traits and complications, including glycemic traits (fasting glucose, fasting insulin, HOMA-IR, and HbA1c), cardiometabolic traits (HDL, LDL, triglycerides, TC, non-HDL, systolic and diastolic blood pressure, and hypertension), vascular diseases (heart attack and stroke), kidney diseases and functions (end stage renal disease, chronic kidney disease, urine albumin-creatine ratio, and eGFR), and inflammatory biomarkers (CRP and IL6) (Supplementary Data 13). Analyses were performed in all populations combined and separately in each population using regression models adjusting for age, sex, BMI, the top ten PCs, and study. Analyses of glycemic traits were restricted to T2D controls to mitigate confounding effects from diabetes medications in T2D cases. For the rest of the traits, analyses were conducted separately in T2D cases, T2D controls, and prediabetes individuals. Model performance was evaluated using AUCs and odds ratios (ORs) for dichotomous traits and the proportion of variance explained (R2) and beta coefficients for continuous traits. Statistical significance was defined as P < 2.50 × 10−3 (0.05/20 traits).
PheWAS
To further understand the clinical implications of T2D PRS on the risk of other diseases and conditions, we conducted phenome-wide association studies (PheWASs) in five independent biobanks: BioMe, BioVU, CCPM, MVP, and MGI. Utilizing the PheWAS R package84, we ran logistic regression models to assess PRS-phenotype associations, adjusting for age, sex, BMI, and the first ten PCs. For the mapping of the International Classification of Diseases (ICD) 9 and ICD-10 codes to phecodes, we utilized phecodes version 1.2 for ICD-9-CM and the 2018 beta version of ICD-10-CM. Cases were considered valid if they had a minimum phecode count of two. PheWAS analyses were performed separately in each population and study, and a pooled-sample PheWAS was conducted with adjustments for populations in each study. Only phecodes with at least 10 cases were considered. A total of 1815 unique phenotypes across 17 disease categories were tested across the five biobanks, with 1777, 1171, 1813, and 1695 phenotypes available in AFR, ASN, EUR, and HIS populations, respectively. Significance was defined by correcting for the total number of phenotypes tested in each population using the Bonferroni-adjusted P-value thresholds: P < 2.75 × 10−5 for all populations combined, P < 2.81 × 10−5 for AFR, P < 4.27 × 10−5 for ASN, P < 2.76 × 10−5 for EUR, and P < 2.95 × 10−5 for HIS populations. Any P-values smaller than the smallest non-zero value that R can represent are denoted as <5.0 × 10−324.
Meta-analysis
Results from PAGE and the 13 additional biobanks and cohorts were first meta-analyzed using inverse variance weighted fixed-effect models within populations to obtain population-specific results for AFR, ASN, EUR, and HIS populations. The results from these four populations, along with the results from all other populations (AI, Greenlandic, MENAQ, and NH), were included in the larger meta-analysis conducted among all populations combined. PheWAS results were similarly meta-analyzed across the five biobanks for each of the four populations and across all populations combined. The degree of heterogeneity was evaluated using the τ2 (between-study variance) and I2 (total proportion of variance owing to heterogeneity) statistics. Meta-analyses were conducted only for analyses with at least 100 cases and 100 controls. If only one study was available for an analysis, making it unfeasible to meta-analyze with other studies, we presented results from that single study.
All statistical analyses were performed using R (version 4.1.2).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The Ge et al. T2D PRS [https://doi.org/10.1186/s13073-022-01074-2] and the T2D PRS of 582 variants [https://doi.org/10.1038/s41588-020-0637-y; https://doi.org/10.1016/j.xhgg.2021.100029] are available on the PGS Catalog under accession numbers: PGS002308 and PGS000804). The newly generated PRS in this study are published on the PGS Catalog under accession number: PGP000742). PheWAS results generated in this study are published on Zenodo [https://doi.org/10.5281/zenodo.15998801]. GWAS summary statistics from the Mahajan et al. 2022 study [https://doi.org/10.1038/s41588-022-01058-3] are available on the DIAGRAM consortium website [https://diagram-consortium.org/downloads.html]. GWAS summary statistics from the Vujkovic et al. 2020 study [https://doi.org/10.1038/s41588-020-0637-y] are available on dbGaP under accession numbers: phs001672.v12.p1. LD matrices used for PRS-CSx are available at https://github.com/getian107/PRScsx.
Code availability
All data and statistical analysis tools used in the present study are open source, details of which are available in the “Methods” section and Nature Portfolio Reporting Summary. No customized code was used to process or analyze data. PRS were calculated using PLINK 2.0 [https://www.cog-genomics.org/plink/2.0/] and weights for newly generated T2D PRS were estimated using PRS-CSx [https://github.com/getian107/PRScsx].
References
International Diabetes Federation. IDF Atlas 2021 10th edn (International Diabetes Federation, 2021).
Bommer, C. et al. Global economic burden of diabetes in adults: projections from 2015 to 2030. Diabetes Care 41, 963–970 (2018).
Khera, A. V. et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat. Genet. 50, 1219–1224 (2018).
Frieser, M. J., Wilson, S. & Vrieze, S. I. Behavioral impact of return of genetic test results for complex disease: systematic review and meta-analysis. Health Psychol. 37, 1134–1144 (2018).
Natarajan, P. et al. Polygenic risk score identifies subgroup with higher burden of atherosclerosis and greater relative benefit from statin therapy in the primary prevention setting. Circulation 135, 2091–2101 (2017).
Mahajan, A. et al. Multi-ancestry genetic study of type 2 diabetes highlights the power of diverse populations for discovery and translation. Nat. Genet. 54, 560–572 (2022).
Vujkovic, M. et al. Discovery of 318 new risk loci for type 2 diabetes and related vascular outcomes among 1.4 million participants in a multi-ancestry meta-analysis. Nat. Genet. 52, 680–691 (2020).
Polfus, L. M. et al. Genetic discovery and risk characterization in type 2 diabetes across diverse populations. Hum. Genet. Genom. Adv. 2, 100029 (2021).
Langenberg, C. et al. Gene–lifestyle interaction and type 2 diabetes: the EPIC InterAct Case-Cohort study. PLoS Med. 11, e1001647 (2014).
Wu, P. et al. Smoking-by-genotype interaction in type 2 diabetes risk and fasting glucose. PLoS ONE 15, e0230815 (2020).
Dietrich, S. et al. Gene-lifestyle interaction on risk of type 2 diabetes: a systematic review. Obes. Rev. 20, 1557–1571 (2019).
Ruan, Y. et al. Improving polygenic prediction in ancestrally diverse populations. Nat. Genet. 54, 573–580 (2022).
Darst, B. F. et al. Evaluating approaches for constructing polygenic risk scores for prostate cancer in men of African and European ancestry. Am. J. Hum. Genet. 110, 1200–1206 (2023).
Gunn, S. et al. Comparison of methods for building polygenic scores for diverse populations. Hum. Genet. Genom. Adv. 6, 100355 (2025).
Wang, Y. et al. Polygenic prediction across populations is influenced by ancestry, genetic architecture, and methodology. Cell Genom. 3, 100408 (2023).
Ge, T. et al. Development and validation of a trans-ancestry polygenic risk score for type 2 diabetes in diverse populations. Genome Med. 14, 70 (2022).
Ng, M. C. Y. et al. Meta-analysis of genome-wide association studies in African Americans provides insights into the genetic architecture of type 2 diabetes. PLoS Genet. 10, e1004517 (2014).
Sakaue, S. et al. A cross-population atlas of genetic associations for 220 human phenotypes. Nat. Genet. 53, 1415–1424 (2021).
Mahajan, A. et al. Fine-mapping type 2 diabetes loci to single-variant resolution using high-density imputation and islet-specific epigenome maps. Nat. Genet. 50, 1505–1513 (2018).
Kachuri, L. et al. Principles and methods for transferring polygenic risk scores across global populations. Nat. Rev. Genet. 25, 8–25 (2024).
Wang, M. C., Shah, N. S., Carnethon, M. R., O’Brien, M. J. & Khan, S. S. Age at diagnosis of diabetes by race and ethnicity in the United States from 2011 to 2018. JAMA Intern. Med. 181, 1537–1539 (2021).
de Miguel-Yanes, J. M. et al. Genetic risk reclassification for type 2 diabetes by age below or above 50 years using 40 type 2 diabetes risk single nucleotide polymorphisms. Diabetes Care 34, 121–125 (2011).
Chen, F. et al. Validation of a multi-ancestry polygenic risk score and age-specific risks of prostate cancer: a meta-analysis within diverse populations. eLife 11, e78304 (2022).
Conti, D. V. et al. Trans-ancestry genome-wide association meta-analysis of prostate cancer identifies new susceptibility loci and informs genetic risk prediction. Nat. Genet. 53, 65–75 (2021).
Schaid, D. J., Sinnwell, J. P., Batzler, A. & McDonnell, S. K. Polygenic risk for prostate cancer: Decreasing relative risk with age but little impact on absolute risk. Am. J. Hum. Genet. 109, 900–908 (2022).
Archambault, A. N. et al. Cumulative burden of colorectal cancer-associated genetic variants is more strongly associated with early-onset vs. late-onset cancer. Gastroenterology 158, 1274–1286.e12. (2020).
Thomas, M. et al. Genome-wide modeling of polygenic risk score in colorectal cancer risk. Am. J. Hum. Genet. 107, 432–444 (2020).
Mavaddat, N. et al. Polygenic risk scores for prediction of breast cancer and breast cancer subtypes. Am. J. Hum. Genet. 104, 21–34 (2019).
Marston, N. A. et al. Predictive utility of a coronary artery disease polygenic risk score in primary prevention. JAMA Cardiol. 8, 130–137 (2023).
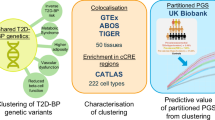
Ghatan, S. et al. Defining type 2 diabetes polygenic risk scores through colocalization and network-based clustering of metabolic trait genetic associations. Genome Med. 16, 10 (2024).
Orozco, G., Ioannidis, J. P., Morris, A., Zeggini, E. & the DIAGRAM consortium. Sex-specific differences in effect size estimates at established complex trait loci. Int. J. Epidemiol. 41, 1376–1382 (2012).
Bernabeu, E. et al. Sex differences in genetic architecture in the UK Biobank. Nat. Genet. 53, 1283–1289 (2021).
Feng, Y. et al. Heritability estimation and environmental risk assessment for type 2 diabetes mellitus in a rural region in Henan, China: family-based and case-control studies. Front. Public Health 9, 690889 (2021).
Ge, T., Chen, C. Y., Neale, B. M., Sabuncu, M. R. & Smoller, J. W. Phenome-wide heritability analysis of the UK Biobank. PLoS Genet. 13, e1006711 (2017).
Zillikens, M. C. et al. Sex-specific genetic effects influence variation in body composition. Diabetologia 51, 2233–2241 (2008).
Mostafavi, H. et al. Variable prediction accuracy of polygenic scores within an ancestry group. eLife 9, e48376 (2020).
Hou, K. et al. Calibrated prediction intervals for polygenic scores across diverse contexts. Nat. Genet. 1–11 https://doi.org/10.1038/s41588-024-01792-w (2024).
Hui, D. et al. Risk factors affecting polygenic score performance across diverse cohorts. eLife 12, https://doi.org/10.7554/eLife.88149.1 (2023).
Berumen, J. et al. Sex differences in the influence of type 2 diabetes (T2D)-related genes, parental history of T2D, and obesity on T2D development: a case–control study. Biol. Sex. Differ. 14, 39 (2023).
Lagou, V. et al. Sex-dimorphic genetic effects and novel loci for fasting glucose and insulin variability. Nat. Commun. 12, 24 (2021).
Kautzky-Willer, A., Harreiter, J. & Pacini, G. Sex and gender differences in risk, pathophysiology and complications of type 2 diabetes mellitus. Endocr. Rev. 37, 278–316 (2016).
Varlamov, O., Bethea, C. L. & Roberts, C. T. Sex-specific differences in lipid and glucose metabolism. Front. Endocrinol. 5, 241 (2015).
Lee, J., Kim, M. H., Jang, J. Y. & Oh, C. M. Assessment HOMA as a predictor for new onset diabetes mellitus and diabetic complications in non-diabetic adults: a KoGES prospective cohort study. Clin. Diabetes Endocrinol. 9, 7 (2023).
Liedtke, T. P. et al. Characterizing trajectories of diabetes-related health parameters before diabetes diagnosis in diabetes subtypes: analysis of a 20-year long prospective cohort study in Sweden. Cardiovasc. Diabetol. 24, 244 (2025).
Ruijgrok, C. et al. Size and shape of the associations of glucose, HbA1c, insulin and HOMA-IR with incident type 2 diabetes: the Hoorn Study. Diabetologia 61, 93–100 (2018).
Tremblay, J. et al. Polygenic risk scores predict diabetes complications and their response to intensive blood pressure and glucose control. Diabetologia 64, 2012–2025 (2021).
Yun, J. S. et al. Polygenic risk for type 2 diabetes, lifestyle, metabolic health, and cardiovascular disease: a prospective UK Biobank study. Cardiovasc. Diabetol. 21, 131 (2022).
Shoaib, M. et al. Evaluation of polygenic risk scores to differentiate between type 1 and type 2 diabetes. Genet. Epidemiol. 47, 303–313 (2023).
Matise, T. C. et al. The next PAGE in understanding complex traits: design for the analysis of population architecture using genetics and epidemiology (PAGE) study. Am. J. Epidemiol. 174, 849–859 (2011).
Wright, J. D. et al. The ARIC (Atherosclerosis Risk In Communities) study: JACC focus seminar 3/8. J. Am. Coll. Cardiol. 77, 2939–2959 (2021).
Friedman, G. D. et al. Cardia: study design, recruitment, and some characteristics of the examined subjects. J. Clin. Epidemiol. 41, 1105–1116 (1988).
Sorlie, P. D. et al. Design and implementation of the Hispanic Community Health Study/Study of Latinos. Ann. Epidemiol. 20, 629–641 (2010).
LaVange, L. M. et al. Sample design and cohort selection in the Hispanic Community Health Study/Study of Latinos. Ann. Epidemiol. 20, 642–649 (2010).
Kolonel, L. N. et al. A multiethnic cohort in Hawaii and Los Angeles: baseline characteristics. Am. J. Epidemiol. 151, 346–357 (2000).
Anderson, G. et al. Design of the Women’s Health Initiative clinical trial and observational study. Control. Clin. Trials 19, 61–109 (1998).
All of Us Research Program Investigators et al. The “All of Us” research program. N. Engl. J. Med. 381, 668–676 (2019).
Roden, D. et al. Development of a large-scale de-identified DNA Biobank to enable personalized medicine. Clin. Pharm. Ther. 84, 362–369 (2008).
Fisher-Hoch, S. P. et al. Socioeconomic status and prevalence of obesity and diabetes in a Mexican American community, Cameron County, Texas, 2004–2007. Prev. Chronic Dis. 7, A53 (2010).
Adair, L. S. et al. Cohort profile: the Cebu longitudinal health and nutrition survey. Int. J. Epidemiol. 40, 619–625 (2011).
Popkin, B. M., Du, S., Zhai, F. & Zhang, B. Cohort Profile: the China Health and Nutrition Survey—monitoring and understanding socio-economic and health change in China, 1989–2011. Int. J. Epidemiol. 39, 1435–1440 (2010).
Wiley, L. K. et al. Building a vertically integrated genomic learning health system: the biobank at the Colorado Center for Personalized Medicine. Am. J. Hum. Genet. 111, 11–23 (2024).
Bjerregaard, P., et al. Inuit health in Greenland: a population survey of life style and disease in Greenland and among Inuit living in Denmark. Int. J. Circumpolar Health 62(Suppl. 1), 3–79 (2003).
Moltke, I. et al. Uncovering the genetic history of the present-day Greenlandic population. Am. J. Hum. Genet. 96, 54–69 (2015).
Stæger, F. F. et al. Genetic architecture in Greenland is shaped by demography, structure and selection. Nature 12, 1–7 (2025).
Bild, D. E. et al. Multi-ethnic study of atherosclerosis: objectives and design. Am. J. Epidemiol. 156, 871–881 (2002).
Zawistowski, M. et al. The Michigan Genomics Initiative: a biobank linking genotypes and electronic clinical records in Michigan Medicine patients. Cell Genom. 3, 100257 (2023).
Gaziano, J. M. et al. Million Veteran Program: a mega-biobank to study genetic influences on health and disease. J. Clin. Epidemiol. 70, 214–223 (2016).
Hunter-Zinck, H. et al. Genotyping array design and data quality control in the Million Veteran Program. Am. J. Hum. Genet. 106, 535–548 (2020).
Taliun, D. et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. Nature 590, 290–299 (2021).
Al Kuwari, H. et al. The Qatar Biobank: background and methods. BMC Public Health 15, 1208 (2015).
Howard, V. J. et al. The reasons for geographic and racial differences in stroke study: objectives and design. Neuroepidemiology 25, 135–143 (2005).
Committee on the Use of Race, Ethnicity, and Ancestry as Population Descriptors in Genomics Research, Board on Health Sciences Policy, Committee on Population, Health and Medicine Division, Division of Behavioral and Social Sciences and Education, National Academies of Sciences, Engineering, and Medicine. Using Population Descriptors in Genetics and Genomics Research: A New Framework for an Evolving Field 26902 (National Academies Press, 2023).
Association A. D. 2. Classification and diagnosis of diabetes: standards of medical care in diabetes—2021. Diabetes Care 2021;44(Supplement_1):S15–S33.
Wojcik, G. L. et al. Genetic analyses of diverse populations improves discovery for complex traits. Nature 570, 514–518 (2019).
Das, S. et al. Next-generation genotype imputation service and methods. Nat. Genet. 48, 1284–1287 (2016).
Fuchsberger, C., Abecasis, G. R. & Hinds, D. A. minimac2: faster genotype imputation. Bioinformatics 31, 782–784 (2015).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Buyske, S. et al. Evaluation of the metabochip genotyping array in African Americans and implications for fine mapping of GWAS-identified loci: the PAGE study. PLoS ONE 7, e35651 (2012).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience 4, s13742-015-0047-0048 (2015).
Lawrence, M., Gentleman, R. & Carey, V. rtracklayer: an R package for interfacing with genome browsers. Bioinformatics 25, 1841–1842 (2009).
Sheng, X. et al. Inverted genomic regions between reference genome builds in humans impact imputation accuracy and decrease the power of association testing. Hum. Genet. Genom. Adv. 4, 100159 (2023).
Robin, X. et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinform. 12, 77 (2011).
Carroll, R. J., Bastarache, L. & Denny, J. C. R PheWAS: data analysis and plotting tools for phenome-wide association studies in the R environment. Bioinformatics 30, 2375–2376 (2014).
Acknowledgements
This work was supported by the National Institutes of Health (R01HL151152 to C.K., C.G., and K.E.N., R00CA246063, R03CA287235, U01CA261339, R01HL174378, P50CA097186, P30CA015704, and U54HG013243 to B.F.D., R01HL143885 and R01DK139598 to P.G.-L. and K.E.N., R01HL163262, R01DK122503, and R01HL142302 to K.E.N., UM1DK078616 to C.K., R01HD30880 and R01AG065357 to P.G.-L., R01DK124097, R01HL156991, and R01DK123019 to R.J.F.L.), an award from the Andy Hill Cancer Research Endowment Distinguished Researchers Program (B.F.D.), and a Fred Hutchinson Cancer Center Translational Data Science New Collaboration Award (B.F.D). R.J.F.L. is supported by grants from the Novo Nordisk Foundation (NNF20OC0059313, NNF23SA0084103, and NNF18CC0034900). Work done by N.A.Y. was made possible by PPM2 grant #PPM2-0226–170020 from the Qatar National Research Fund (QNRF) and Qatar Genome Program (QGP) (members of Qatar Foundation) and by NPRP-S11 grant #NPRP11S-0114–180299 from the Qatar National Research Fund. The full acknowledgments and funding for the individual studies are listed in the Supplementary Information. The findings herein reflect the work and are solely the responsibility of the author. Funders did not contribute to the design or conduct of the study; collection, management, analysis, or interpretation of the data; or preparation, review, or approval of the manuscript.
Author information
Authors and Affiliations
Consortia
Contributions
B.F.D., H.M.H., S.R., and R.A.J.S. contributed to study conception and design. B.G. and B.F.D. wrote the manuscript. C.K., C.G., and K.E.N. contributed to funding acquisition. Central data analysis group: B.G. (PAGE), D.K. (CCHC), Z.W. (BioMe), J.H. (AoU), R.T. (BioVU), K.A.B. (CLHNS), Y.W. (CHNS), N.P. and M.L. (CCPM), F.F.S. (Greenlandic), C.Y. (MESA), B.V. (MGI), K.R.I. and A.T.H. (MVP), N.A.Y. (Qatar), and A.D.P. (REGARDS) contributed to data analysis and interpretation. Study level principal investigators: K.L.M. (CLHNS), P.G.-L. (CHNS), A.A. (Greenlandic), M.B. (MGI), S.S.R. (MESA), A.M. and J.I.R. (MESA), R.M.I. (REGARDS), C.G. (CCPM), K.E.N. (CCHC), R.J.F.L. (BioMe), T.L.A. (MVP), C.A.H., L.L.M., and L.R.W. (MEC), and C.K. (WHI) contributed data and provided resource and administrative support. PAGE, AoU, BioMe, BioVU, CCHC, CCPM, CLHNS, CHNS, Greenlandic, MESA, MGI, MVP, Qatar, and REGARDS provided the data and administrative support. Additional contributions: Y.C., L.S., N.R., J. Shortt, L.W., M. Stanislawski, J. Pattee, L.D., P.S.S., M.M.S., N.J.C., N.R.L., M.E.J., P.B., C.L., T.H., I.M., J.B.M., D.O.S., X.Y., X.Z., K.-M.C., S.L.C., R.G.-S., J.L., P.S.T., S.B., M.G., L.M.R., Q.S., C.S.C., C.B.E., S.L., J.E.M., and U.P. contributed to design, data collection, management, coordination, interpretation, and/or analysis of contributing studies. B.F.D. and H.M.H. supervised the work. B.F.D. takes responsibility for the integrity and accuracy of the manuscript. All authors critically reviewed and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.J.F.L. has acted as a member of advisory boards and as a speaker for Ely Lilly and the Novo Nordisk Foundation, for which she has received fees. L.M.R. is a consultant for the NHLBI TOPMed Administrative Coordinating Center (through Westat). U.P. was a consultant with AbbVie, and her husband holds individual stocks for the following companies: BioNTech SE—ADR, Amazon, CureVac BV, Google/Alphabet Inc Class C, NVIDIA Corp, Microsoft Corp. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Tinashe Chikowore and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Guo, B., Cai, Y., Kim, D. et al. Polygenic risk score for type 2 diabetes shows context-dependent effects across populations. Nat Commun 16, 8632 (2025). https://doi.org/10.1038/s41467-025-63546-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63546-4