Abstract
Embryonic stem cells (ESC) are pluripotent, with the potential to differentiate into multiple cell types, making them a valuable tool for regenerative medicine and disease therapy. However, common culture methods face challenges, including strict operating procedures and high costs. Currently, Leukemia inhibitory factor (LIF), an indispensable bioactive protein for ESC culture, is typically applied to maintain self-renewal and pluripotency, but its instability and high cost limit its effectiveness in stable culture conditions. Hence, we have developed an innovative strategy using a soluble nanomaterial, metal-organic polyhedra (MOPs), to effectively maintain the self-renewal and pluripotency of ESC. The selected amino-modified vanadium-based MOP not only exhibits excellent biocompatibility and high stability but also possesses similar or even superior biological functions compared to commercial LIF. Due to the precise structure of MOPs, the active site responsible for maintaining ESC pluripotency has been identified and regulated at the molecular level. The new ESC culture method significantly reduces costs, simplifies preparation, and enhances the practicality of biopharmaceutical preparation and storage. This represents the first case of using MOPs to maintain self-renewal of ECS, opening an avenue for introducing advanced materials into the development of innovative ESC culture methods.
Similar content being viewed by others
Introduction
Embryonic stem cells (ESC) are characterized by their self-renewal capability and the qualities of pluripotency, the ability to give rise to cell types representative of all the tissues of the embryo and adult when differentiated, and immortality, or unlimited proliferation1. It is these properties that ESC are gradually being modeled for in vitro studies of cell differentiation and developmental regulatory mechanisms and explored for varieties of diseases and injury therapy, such as progressive multiple sclerosis2, Parkinson’s disease3,4, Alzheimer’s disease5,6,7, spinal cord injury8,9,10 and age-related macular degeneration4,10, etc. Furthermore, ESC are highly undifferentiated cells with developmental totipotency and can differentiate into any tissue and organ in the adult animal, including germ cells. However, applying ESC requires optimized methods that maintain their capacity for self-renewal and pluripotency with infinite adjustability and facile processability. Supplementing with the cytokine leukemia inhibitory factor (LIF) or co-culturing with feeder cells are traditional culture strategies that avoid spontaneous differentiation and maintain the therapeutic potential of ESC11,12,13,14. Nevertheless, the multiple supplementary ingredients are expensive and bring various uncertainties that often affect the quality of cells15,16. In addition, protein reagents (e.g., LIF) have to be carefully managed due to their unstable physicochemical properties, and they are highly susceptible to chemical and physical factors, such as disinfecting/sterilizing conditions like high temperatures and alcohol, which significantly impair their stability and viability17,18. Therefore, it is essential to develop effective and convenient methods as an alternative strategy to build the foundation of ESC engineering. Recently, biomaterials have been widely used to maintain the pluripotency of ESC19,20,21. Gold nanoparticle layers22, DNA-silica nanocomposite23, and MgFe-LDH nanoparticles24 have been reported to maintain ESC pluripotency. However, these previous studies did not emphasize the cost of ESC culture and storage conditions of the materials and did not evaluate the performance status of these materials in a strictly sterilized or disinfected state.
The development of crystalline functional materials with unique cavities and defined morphologies, such as metal-organic frameworks (MOFs), covalent organic frameworks (COFs), hydrogen-bonded organic frameworks (HOFs), and cages, has received wide attention over the past several decades. Among them, discrete metal-organic polyhedra (MOPs), unlike crystalline two- or three-dimensional networks (MOFs, COFs, HOFs, etc.), can be dispersed as well-defined single entities in certain solvents, so-called solution processability25. With their adjustable nanoscale architecture and customizable functionality, MOPs exhibit significant potential in biotechnology applications, particularly in areas such as catalysis, biosensors, and drug delivery26,27,28,29,30,31. For instance, MOPs have been applied as enzyme-mimicking nanomaterials or stabilized enzyme by using their large internal cavities to encapsulate proteins and regulate their molecular behavior32,33,34. However, the ability of MOPs to maintain self-renewal and pluripotency of ESC has not been clarified yet.
Changes in the redox state affect the differentiation process of mouse ESC (mESC). Compounds containing some elements have been found to have the ability to change redox state, such as vanadium (V)-containing materials have been reported that participate in glutathione-related physiological processes and have enzyme-mimicking properties35,36,37,38,39,40. Particularly, V-based MOPs are potential candidate, which has attractive features such as precise structure, customizable functionality and solubility in H2O and alcohol, herein, we evaluated the impact of MOPs on mESC pluripotency, which realizes the precise regulation of mESC depending on the specific molecular docking between the NH2 group modified V-based MOP (denoted as MOP-1) and Src homology region 2 domain-containing phosphatase-2 (SHP-2), a key enzyme for mESC pluripotent maintenance. This revolutionary strategy has high resistance against perturbation environments with superior performance that greatly raises the possibilities of in-depth exploration for various scenarios. In addition, this innovative culture method for mESC can dramatically reduce the cost, thereby broadening the applications of both mESC and MOPs, ultimately realizing the prospects of disease therapy.
Results
Structural and stability characterization of MOP-1
MOP-1 was obtained under solvothermal conditions by self-assembly of vanadium chloride and 2-aminoterephthalic acid (H2BDC-NH2)41. The crystals (macroscopic stacking morphology) of MOP-1 presented as dark green octahedrons with a size of ~100 μm under scanning electron microscope (SEM) and optical microscope (Supplementary Fig. 1A). The phase purities of MOP-1 were confirmed by the agreement between the experimental powder X-ray diffraction (PXRD) pattern and the calculated pattern based on structural analysis. As shown in Fig. 1A, the PXRD pattern confirmed the successful preparation of MOP-1 with high purity. MOP-1 is present as an octahedral coordination cage consisting of six clusters as the vertices and twelve H2BDC-NH2 as the linkers41. Each MOP-1 is −12 charges and balanced by twelve [NH2Me2]+ cations, and its zeta potential was −20.4 ± 0.9 mV via dynamic light scattering (DLS) measurement (Supplementary Table S1). The water solubility of MOP-1 also comes from its charged characteristics25. The aqueous solution of monodisperse MOP-1 was clear and dark green (Fig. 1B, Supplementary Data 1). The average diameters of MOP-1 in H2O was 1.95 ± 0.37 nm via DLS measurement (Fig. 1C), whose size was also evident by transmission electron microscopy (TEM) (Fig. 1C). The bonding types were measured via Fourier transform infrared spectroscopy (FT-IR). The absorption band at 3342 cm−1 corresponds to the N-H stretch of the NH2 group of MOP-1 in FT-IR spectroscopy (Supplementary Fig. 1B).
A The photo of MOP-1 aqueous solution (left). Monodisperse structure of MOP-1 in H2O (right). Color codes: V, green; O, red; C, gray; N, blue; Cl, bright blue. The large pink sphere represents the free space inside the molecular cage. For clarity, H atoms were omitted. Schematic (right panel) was created with Diamond software. B Experimental and simulated PXRD patterns of MOP-1. C TEM image of MOP-1 (left). Scale bar, 20 nm. The particle size of MOP-1 in H2O (right). Three experiments were repeated independently with similar results. D The schematic diagram showed that the SHP-2 mediated STAT3 inactivation. Schematic diagram was created with Microsoft Office PowerPoint. E, F An in-depth mechanism investigation of mESC pluripotency control by MOP-1. Binding model from a global view of a complex composed of SHP-2 and MOP-1 illustrated by electrostatic surface potential (E). Binding modes are illustrated by ribbon diagrams of a complex composed of SHP-2 and MOP-1 (the left panel), a complex composed of SHP-2 with MOP (the middle panel) and a complex composed of SHP-2 with ZrMOP (the right panel). The top panel is the global view of the catalytic PTP structure of SHP-2, the bottom panel is the focused view of binding modes illustrated by the Ribbon diagrams (F). G Binding kinetics of MOP-1 (top panel) and MOP-2 (bottom panel) with SHP-2 were measured by the SPR assay. H The ICP-MS analysis of the binding quantity between MOPs and SHP-2 (mean ± s.e.m, n = 6). I The inhibition efficiency of SHP-2, JAK2, JAK1, SHP-1, PTP1B, Cyt c, ACP and lipase by MOP-1 at a concentration of 2 μM using enzyme assay (mean ± s.e.m, n = 3). Data in (H) and (I) are analyzed by one-way ANOVA. ****P < 0.0001, the binding between MOP-1 and SHP-2 vs. the binding between MOP-2 and SHP-2, relative activity of SHP-2 vs. relative activity of JAK2, JAK1, PTP1B, Cyt c, ACP and lipase. ***P < 0.001, relative activity of SHP-2 vs. relative activity of SHP-1.
For biomedical application scenarios, environmental stability is a key factor in practical applications. Stability is not a common characteristic of MOPs, but V-based MOPs with polyoxometalates as metal nodes have been proven to possess good stability in organic solvents42, ionic liquids43 and even buffers44. To further verify the stability of MOP-1 was investigated in various long-term storage conditions (in different buffers) and cell operation processes (e.g., sterilization including high temperature, alcohol and membrane filters). The stability of the crystalline state of MOP-1 was determined by changes in the PXRD pattern. MOP-1 was exposed to room temperature and air atmosphere for 1 h, 1 week, 2 weeks, and 4 weeks, and its PXRD patterns showed no significant changes compared to the initial ones, which revealed that crystalline state MOP-1 can maintain crystallinity and stability of macroscopic stacking morphology at least 4 weeks (Supplementary Fig. 1C). Then, MOP-1 was heated at 100 °C for 1 h and 5 h, and treated with 2 mL alcohol (75% ethanol) for 5 min, respectively. PXRD patterns without significant changes indicated that the crystalline state MOP-1 could survive at least 5 h under 100 °C and remain stable in alcohol (Supplementary Fig. 1D).
Before final application, MOP-1 was fully dissolved in Dulbecco’s modified Eagle medium (DMEM), serum+LIF (LIF) or 2i+LIF (2iL) medium that was frequently used for mESC culture. Therefore, the stability of the monodisperse MOP-1 is also crucial, including long-term stability, thermostability, sterilization stability and buffers and culture media stability. When dissolved in H2O, MOP-1 also presented excellent long-term stability ( > 1 month, Supplementary Fig. 1E), which was monitored by ultraviolet and visible (UV-vis) spectrophotometry. Then, the MOP-1 aqueous solution was heated to 100 °C for 1 h or 5 h and filtered through a 0.22 μm microporous filter membrane. As shown in Supplementary Fig. 1F and G, the characteristic peaks of MOP-1 before and after treatment did not show any changes in displacement or intensity compared to the characteristic peaks of the ligand, which indicated that MOP-1 had enough thermal stability and sterilization stability. Meanwhile, the molecular structure of MOP-1 could be preserved quite well in PBS and DMEM (Supplementary Fig. 2). On the other hand, MOP-1 affects neither the pH nor components of the medium (Supplementary Fig. 3). These results suggested that the well-retained structures and resistance properties of MOP-1 satisfied the majority of needs in adapting different mESC culturing scenarios, it gets rid of the strict storage conditions of traditional mESC culture supplements. The high stability of MOP-1 lays the foundation for subsequent application research.
The mechanism of MOP-1 maintains the pluripotency of ESC
Given the bio-activation of MOP-1 in mESC pluripotency and the potential advantage of MOP-1 instead of LIF-dependent mESC culture, the mechanisms that underlie the pluripotency modulatory effects of MOP-1 were explored. Previous studies45,46 have well documented that signal transducer and activator of transcription 3 (STAT3) plays a central role in maintaining a proper balance of mESC differentiation and pluripotency. Src homology 2 domain-containing protein tyrosine phosphatase (SHP-2), a cytoplasmic tyrosine phosphatase, inhibits STAT3 phosphorylation in mESC by transducing signal relay from the cell surface to the nucleus, which precisely facilitates mESC differentiation11 (Fig. 1D).
To explore how MOP-1 influences the enzymatic activity of SHP-2, we comprehensively evaluated the binding of SHP-2 and MOP-1 in terms of nanoscale size, hydrophobicity, and binding energy. The depth of SHP-2’s catalytic pocket is 0.9 nm, while the size of MOP-1 is 2 nm (1.95 ± 0.37 nm). MOP-1 penetrates the SHP-2 pocket to a depth of 1 nm, which is half the depth of MOP-1 (materials that cannot reach the bottom will not affect the P-loop) (Supplementary Fig. 4). MOP-1 is negatively charged, while the SHP-2 pocket is hydrophobic and positively charged, that confirms the characteristics of MOP-1 meet the conditions for binding to the active site of SHP-2 (Supplementary Table 1). We further performed molecular modeling using the molecular docking method to calculate the binding affinity of MOP-1 and SHP-2. The MOP-1 was docked into the phosphorylation sites (Cys459, Arg465) of SHP-2 via hydrogen bonds, salt bridges, and π-π stackings (Fig. 1E, F) to form a highly stable complex, given the binding free energy of −10. 47 kcal mol−1.
Specifically, the oxygen atoms in the vertex of the MOP-1 were hydrogen-bonded to the hydrogen atoms in the phenolic hydroxyl group of Tyr279 and the hydroxyl group of Ser460, respectively, facilitating the vertex insertion in the pocket. The hydrogen atom in the NH2 group of MOP-1 formed a hydrogen bond with the carbonyl oxygen atom in Gln506, and the other NH2 group formed a hydrogen bond with GLY427. Thus, the NH2 groups play a key role as hydrogen bond donors or acceptors in the interaction between MOP-1 and SHP-2. The benzene rings of the H2BDC-NH2 in MOP-1 formed the π-π stacking interaction with Tyr279 and His426, respectively, which helped MOP-1 fix into the pocket. Molecular docking showed that MOP-1 had a binding affinity to SHP-2 with a free energy of −10.47 kcal mol−1. The morphology and size of MOP-1 match with the catalytic pocket of SHP-2, which revealed that MOP-1 perfectly engaged the catalytic pocket and inactivated SHP-2.
To further explain MOP-1 as a more potent architecture for SHP-2 inhibition, another two MOPs were introduced as a comparison. One has the same vertex but has no NH2 group modification in the linker named MOP-241, and the other possesses the same linker but a different vertex named ZrMOP47. MOP-2 and MOP-1 have the same octahedral structure, while ZrMOP has a different structure (tetrahedral structure) and the same NH2 group as MOP-1. Molecular docking results of MOP-2 showed poorer binding modes and energy than MOP-1, with the free energy of −7.26 kcal mol−1, while ZrMOP did not bind to SHP-2. To consider the random positions of the NH2 group, we built four random models based on MOP-1. These models, which represent possible isomers that could be present in synthesis, formed interactions with SHP-2 through the NH2 groups at positions Arg362/Lys364, respectively. Their binding free energies were −10.87, −10.82, −10.46, and −10.03 kcal mol−1, showing similar affinity to SHP-2 (Supplementary Fig. 5). These results revealed the size and functional groups of MOPs could affect the binding between the material and SHP-2 and MOP-1 was a selective and potent inhibitor of SHP-2 (Fig. 1F).
Based on the results above, we next examined the impact of MOP-1 on SHP-2 activation. The binding affinities of MOP-1 and MOP-2 were analyzed by surface plasmon resonance (SPR). MOP-1 displayed good binding abilities to SHP-2 with a KD value of 3.2×10−7 M, compared to MOP-2 (Fig. 1G). We further investigated the amount of MOP-1 or MOP-2 that could bind per microgram of SHP-2. After co-incubating MOP-1 or MOP-2 with SHP-2 for 6 h at 4 °C, inductively coupled plasma mass spectrometry (ICP-MS) analysis revealed that approximately 15 ng of vanadium element (V, representing MOP-1) was bound per microgram of SHP-2 (Fig. 1H). Having thoroughly demonstrated that MOP-1 could bind to SHP-2, we further validated the inhibitory effect of MOP-1 on SHP-2 activity. As shown in Fig. 1I, MOP-1 effectively binds to and inhibits SHP-2 activity, and shows stronger inhibition compared to other tyrosine phosphatases unrelated to the STAT3 pathway, kinases in the STAT3 pathway, Src homology region 2 domain-containing phosphatase-1 (SHP-1, expressed in blood) and other enzymes, such as protein tyrosine phosphatase-1B (PTP1B), Janus kinase 2 (JAK2), Janus kinase 2 (JAK1), SHP-1, acid phosphatase (ACP), cytochrome c (Cyt c) and lipase, but its inhibition efficiency also increases with higher concentrations (Supplementary Fig. 6). At the 2 μM concentration of MOP-1, the inhibition efficiency of SHP-2 was up to 80%, which conclusively demonstrates that MOP-1 can effectively bind to SHP-2 and inhibit its activity.
Biocompatibility comparison of MOP-1 and other nanoparticles in mESC
To investigate the effect of MOP-1’s morphology or composition on maintaining the pluripotency of mESC, we selected several classical architectures with the same shape (MOP-2), functional group (ZrMOP), similar sizes in buffer (iron oxide nanoparticles (IONPs) and Pt nanoparticles (PtNPs)) and MOPs’ component ligand for comparison. The characterizations of the above materials were tested by PXRD or TEM (Supplementary Figs. 7 and 8A). The survival rate of mESC (J1 cell line) treated with MOP-1 and above nanoparticles, respectively, for 24 h was assessed via MTT assay. The cell viability of mESC cultured with various concentrations of MOP-1, MOP-2, ZrMOP, ligand, IONPs, and PtNPs showed no significant difference compared to mESC cultured with or without LIF (Fig. 2A and Supplementary Fig. 8B). Then, the capacity of mESC self-renewal maintenance was investigated at the same non-cytotoxic concentration (2 μM). As pluripotent stem cells typically have higher levels of alkaline phosphatase (ALP) activity than differentiated cell types, we used ALP staining as a convenient method to identify stem cells48. The mESC incubated with MOP-2 or ZrMOP showed no signs of smooth edges at the colonies’ periphery, and colonies cultured with the ligand also exhibited protrusions along the edges. The mESC incubated with IONPs or PtNPs showed a flattened phenotype, like the mESC group without LIF treatment (denoted as Ctrl mESC), which indeed has a loss of stem cell morphology. In contrast, MOP-1 mESC colonies showed smooth edges and a regular circular shape, with a relatively darker color after ALP staining compared to all test groups, indicating enhanced ALP activity (Fig. 2B and Supplementary Fig. 8C). This suggests that nanoparticles with structures or sizes similar to MOP-1 cannot maintain the self-renewal capability of mESC. In addition, there were no significant differences between MOP-1 mESC and LIF mESC in the spread area of colonies, which indicated MOP-1 exhibited the potential for the conservation of self-renewal ability in mESC (Fig. 2C).
A, B Cell viability assay of MOP-1 (top panel, mean ± s.e.m, n = 3), MOP-2 (second panel, mean ± s.e.m, n = 3), ZrMOP (third panel, mean ± s.e.m, n = 6) and ligand (bottom panel, mean ± s.e.m, n = 5). Schematics (left panel) were created with Diamond software. A MOPs treated mESC (J1 cell line) exhibited flat morphology compared with the mESC cultured in LIF addition medium (denoted as LIF mESC) and mESC cultured without LIF (denoted as Ctrl mESC), which grew in colonies and expressed positive alkaline phosphatase (ALP) staining (B). Scale bar, 100 μm. C Representative images of ALP staining (left panel) and quantification area of LIF and MOP-1 mESC colonies (right panel) (mean ± s.e.m, n = 3). Scale bars, 40 μm). D The cell cycle distribution of MOP-1 mESC was evaluated by flow cytometry. The percentage of G0/G1, S, G2/M phases of the cell cycle (mean ± s.e.m, n = 3). E Plots of FITC-annexin V/PI flow cytometry of mESC cultured with or without MOP-1 for 48 h. F Representative immunoblot images (left panel) and quantification (right panel) of p-SHP-2 in MOP-1 mESC under different concentrations (mean ± s.e.m, n = 3). G Representative immunoblot images of STAT3 in LIF mESC and MOP-1 mESC at different withdrawal times (mean ± s.e.m, n = 4). Data in (C) and (D) are analyzed by a two-tailed unpaired t-test. Data in (F) are analyzed by one-way ANOVA. #P < 0.05, 0 μM MOP-1 mESC vs. LIF mESC; *P < 0.05, 2 μM and 4 μM MOP-1 mESC vs. 0 μM MOP-1 mESC; no statistical significance (NS), LIF mESC vs. 2 μM and 4 μM MOP-1 mESC, MOP-1 mESC vs. LIF mESC or Ctrl mESC.
To determine the effect of MOP-1 on cell cycle arrest and cell death in stem cells, J1 mESC were treated with MOP-1 at 2 μM for 48 h. As shown in Fig. 2D, we found that MOP-1 did not significantly influence cell proliferation in the G0/G1, S and G2/M phases by flow cytometry assay. Moreover, MOP-1 mESC apoptosis was analyzed by fluorescence-activated cell sorting (FACS) (Fig. 2E). No obvious change in the apoptosis rate was identified in mESC incubated with MOP-1 at 2 μM when compared with those incubated in LIF. The ICP results indicate that the intracellular concentration of element V (representing MOP-1 content) peaks at 12 h. No significant difference is observed in the V content of mESC between 0 and 48 h, suggesting that MOP-1 reaches its maximum level in mESC at 12 h and is completely metabolized by 48 h (Supplementary Fig. 9).
With no impact on cell viability or the cell cycle, we further investigated the inhibitory effects of MOP-1 on SHP-2 activity within mESC, as well as the associated changes in STAT3 through experiments. The results indicated that increasing concentrations of MOP-1 led to a reduction in p-SHP-2 expression levels, accompanied by an upregulation of stemness-related genes, organic cation/carnitine transporter 4 (OCT4) and SRY-box transcription factor 2 (SOX2). These changes stabilized at a concentration of 2 μM, showing no significant difference in expression levels compared to mESC cultured with LIF (Fig. 2F and Supplementary Fig. 10). Furthermore, we established an SHP-2-overexpressing E14 mESC cell line. mESC with SHP-2 overexpression lost their self-renewal capacity and exhibited differentiated cell morphology. In contrast, MOP-1 significantly increased the expression of stemness-related proteins, effectively maintaining mESC pluripotency (Supplementary Fig. 11). These results indicate that MOP-1 maintains mESC pluripotency by inhibiting SHP-2. We observed that withdrawal of MOP-1 or LIF, the p-STAT levels in MOP-1 decreased more rapidly over time. After 48 h, the p-STAT expression levels were comparable to those of the control group, which is consistent with the metabolic results of MOP-1 (Fig. 2G and Supplementary Fig. 12). These findings strongly suggest that MOP-1 exhibits a more pronounced ability to maintain mESC pluripotency compared to other MOPs or NPs. Moreover, this preservation of pluripotency is mediated by MOP-1 through the modulation of p-STAT3, achieved by inhibiting SHP-2 activity.
MOP-1 is similar to LIF in supporting ESC self-renewal
Following extensive validation of the mechanism by which MOP-1 inhibits SHP-2, we compared its effectiveness in maintaining mESC self-renewal relative to LIF. The biofunction of MOP-1 at different concentrations in mESC (J1 cell line) culture for 48 h was investigated by observing the clone morphology in each group. Both LIF mESC and MOP-1 mESC had a round shape, whereas clones in the Ctrl mESC were fragmented, presenting an irregular shape and undefined borders, considered typical characteristics of pluripotency loss. It was observed that at the concentration range of 2-4 μM, MOP-1 preserved the morphology of mESC (Supplementary Fig. 13).
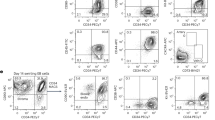
Because both mESC and human embryonic stem cells (hESC) maintain their self-renewal capacity through the inhibition of SHP-2 activity46,49,50, we further performed an analysis of stemness-related genes expression in hESC (RUES2 cell line). The expression of stemness markers, SOX2 and OCT4, in mESC was detected via immunofluorescence staining. The fluorescence signals were greatly downregulated upon LIF withdrawal, whereas MOP-1 treatment enhanced the fluorescence intensity and stemness-related protein expression of SOX2 and OCT4 in mESC (Fig. 3A). Furthermore, we investigated the ability of MOP-1 to maintain the undifferentiated state and pluripotency of hESC in mTeSR1 medium without bFGF. After treating the hESC with MOP-1 for two passages, we assessed the expression of stemness-related genes. The expression levels of SOX2 and NANOG in MOP-1 hESC were significantly higher compared to -bFGF hESC (Fig. 3B). At the end of passage three, the ESC markers of OCT4, SSEA4, NANOG and TRA 1-81 in the undifferentiated state for hESC were analyzed by FACS.
A Representative images showed that the MOP-1 mESC and LIF mESC expressed typical pluripotency-associated markers compared with the Ctrl mESC using immunofluorescence staining. Scale bars, 20 μm. B The expression of stemness-related genes in hESC cultured in mTeSR1 medium (denoted as Ctrl hESC), hESC cultured in mTeSR1 medium without bFGF (denoted as –bFGF hESC) and hESC cultured in 2 μM MOP-1 supplemented mTeSR1 medium without bFGF factor (denoted as MOP-1 hESC) (mean ± s.e.m, n = 5). C hESC were analyzed on Day 9 of passage 3 by FACS for hESC cell markers OCT4 (mean ± s.e.m, n = 3 for Ctrl hESC, n = 4 for –bFGF hESC and MOP-1 hESC), SSEA4 (mean ± s.e.m, n = 3 for –bFGF hESC, n = 4 for Ctrl hESC and MOP-1 hESC), Nango (mean ± s.e.m, n = 3 for MOP-1 hESC, n = 4 for Ctrl hESC and –bFGF hESC) and Tra 1-81 (mean ± s.e.m, n = 4 for each group). D Volcano plot depicting transcriptional profiles of MOP-1 mESC versus LIF mESC. Gray: all no differentially expressed genes (fold change R1, adjusted P < 0.05); red: up genes; blue: down genes. E Heatmap showing genes related to pluripotency related genes in mESC cultured with LIF or MOP-1. HTseq (High-throughput sequence) was used to get the FPKM method to determine gene expression. The analysis was performed for the contrast of MOP-1 mESC vs. LIF mESC; the red color indicated increased gene expression in MOP-1 mESC, while blue indicated decreased gene expression in MOP-1 mESC. Representative images of mESC cultured with MOP-1 and stained with ALP after over 1 week of culturing. ALP staining of J1 mESC (F), E14 mESC (G) and T4 mESC (H) that were treated with MOP-1 at the indicated time points. Three experiments were repeated independently with similar results. Scale bars, 200 μm in (F–H). Data in (B) and (C) are analyzed by one-way ANOVA. *P < 0.05, **P < 0.01, ***P < 0.001, –bFGF hESC vs. Ctrl hESC or MOP-1 hESC; no statistical significance (NS), –bFGF hESC vs. Ctrl hESC or MOP-1 hESC.
When exogenous bFGF was removed, some undifferentiated cells nonetheless persisted, but their proportion significantly decreased. In contrast, the proportion of undifferentiated cells in MOP-1 hESC increased compared to the -bFGF hESC (Fig. 3C). The results indicated that MOP-1 effectively maintains the undifferentiated state of hESC. We further conducted hESC spontaneous embryoid bodies (EB) formation. Immunofluorescence staining was conducted to detect protein expression in the different germ layers formed in the EB on Day 3. Although the marker proteins of the three germ layers were present in all the groups, the expression levels in the Ctrl hESC-derived EB and MOP-1 hESC-derived EB were significantly higher than those in the -bFGF hESC-derived EB (Supplementary Fig. 14). Thus, we concluded that MOP-1 could maintain the pluripotency potential in the absence of bFGF in hESC.
In addition to more effectively assessing changes in the undifferentiated state and pluripotency during an optimal cell state, we also performed global transcriptome sequencing to compare the differences between LIF-cultured mESC and MOP-1-cultured mESC. As shown in the Volcano plot and heatmap, there was no significant difference in stemness-related genes (Nanog, SOX2 and Pou5f1) (Fig. 3D) and cell cycle-related genes (Supplementary Fig. 15) between MOP-1 mESC and LIF mESC. The expression score of stemness-related genes in MOP-1 mESC was higher than that of LIF mESC, while the apoptosis-related genes (Fgf15, Dll1, Sfrp2 and Gadd45g) were downregulated in MOP-1 mESC (Fig. 3D, E). Moreover, the mESC self-renewal maintenance capacity of MOP-1 is not only effective for the J1 cell line, but it also has the same capacity in other cell lines. When mESC were cultured with MOP-1 for over 1 week, all three cell lines (J1 cell line, E14 cell line and T4 cell line) retained their rounded morphologies, and even T4 mESC maintained their rounded morphologies for over 2 weeks (Supplementary Fig. 16). Furthermore, the bright-field microscopy results were verified by the ALP staining. It was observed that MOP-1 could successfully maintain the self-renewal ability of mESC for at least two weeks when treated with a minimal concentration of 2 μM (Fig. 3G, H and Supplementary Figs. 16 and 17). Collectively, these results indicate that MOP-1 is comparable to LIF in maintaining mESC self-renewal and may serve as an alternative to LIF in mESC culture. Additionally, it is worth mentioning that the cost of an additional amount per liter of culture medium for commercial LIF is 719 times that for MOP-1 (Supplementary Tables 2 and 3). Replacing LIF with MOP-1 could significantly reduce the cost of the culture medium.
MOP-1 supports the pluripotency of mESC
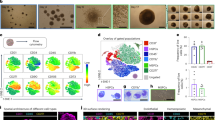
An important feature of mESC is that they can naturally differentiate into three germ layers (ectoderm, mesoderm and endoderm) and assemble various tissues or organs51. In vitro EB52 and in vivo teratomas53,54 are gold standards for assessing their potential in the following application; we thus conducted in vitro and in vivo differentiation assays after MOP-1 withdrawal (denoted as –MOP-1 mESC) or LIF withdrawal (denoted as –LIF mESC). To test the in vitro differential potential of MOP-1 mESC for three germ layers, we aggregated single-cell dissociated –MOP-1 mESC and –LIF mESC in non-adherent plates for 4.5 days to form –LIF EB and –MOP-1 EB, respectively. There was no significant difference sign in size between –LIF and –MOP EB at Day 4.5 (Supplementary Fig. 18). We compared the lineage differentiation levels of three blastoderms, including the representative markers for ectoderm (Pax6, Tubb3, Otx2), mesoderm (Mixl1, Twist2, Branchyary) and endoderm (Gata4, Emoes, Foxa2), indicating that –MOP-1 mESC is potent in successfully differentiation (Supplementary Fig. 19). Subsequently, we compared the lineage differentiation levels of three blastoderms for 4.5 days after withdrawal of LIF or MOP-1 via single cell sequencing and immunofluorescence staining. We annotated clusters according to expression of known cell type markers and clustered the cells isolated from EB to identify eight distinct development-related clusters (Fig. 4A and Supplementary Fig. 20A). These clusters were visualized using Uniform Manifold Approximation and Projection (UMAP), revealing the global structure of cells isolated from –LIF EB and –MOP-1 EB (Fig. 4B). The differentiation proportions of the three germ layers indicate that –MOP-1 EB successfully differentiated into all three germ layers (endoderm, mesoderm, and ectoderm) (Fig. 4C). And the cell number in the epiblast of –MOP-1 EB is greater than that in the epiblast of the –LIF EB, indicating that –MOP-1 EB are differentiating into the ectoderm at Day 4.5 (Supplementary Fig. 20B). To further examine the lineage relationship between undifferentiated stem cells and other germ layer cells, we performed RNA velocity analysis and found that the evolutionary trajectory in both groups begins with undifferentiated stem cells and progresses towards differentiation into either the endoderm and mesoderm, or the ectoderm (Fig. 4D and Supplementary Fig. 21). We also examined the expression of marker genes for different germ layers in both groups. In both –MOP-1 EB or –LIF EB, the expression levels of genes related to cardiac development expressed in the endoderm were similar, indicating that the developmental stages of –MOP-1 EB are comparable to those of –LIF EB (Fig. 4E and Supplementary Fig. 22).
A, B UMAP projection of EB of the atlas data set. In the plot, cells are colored by lineage assignment, and colored by partitioning into 8 clusters or different treatment EB (–LIF EB and –MOP-1 EB). Cells from –LIF EB are blue, cells from –MOP-1 EB are red (B). C Prop Plots showing the proportion of three germ layers for –LIF and –MOP-1 EB differentiation. D Visualization of RNA velocities on the UMAP projection with information for the 8 clusters of –LIF EB and –MOP-1 EB. E Expression of endoderm (Fst, Kdr and Sox17), mesoderm (Hand1) and ectoderm markers (Neto2) in –MOP-1 EB and –LIF EB. F Violin plot of the score of apoptosis in –LIF EB and –MOP-1 EB. G Hematoxylin and eosin (H&E) stained cross-sections of –LIF EB and –MOP-1 EB. Scale bar, 20 μm. H Quantitative analysis of the ratio of dead cells area (left panel, mean ± s.e.m, n = 8 from 4 mice teratoma samples per group) and number of dead cells (right panel, mean ± s.e.m, n = 9 from 4 mice teratoma samples per group) of H&E stained cross-sections of –LIF EB and –MOP-1 EB. Data in (F) and (H) are analyzed by a two-tailed unpaired t-test. *P < 0.05, ****P < 0.0001, –LIF EB vs. –MOP-1 EB.
The EB was then allowed to differentiate for 14 days, and the expression markers of the three germ layers, containing MAP2 (ectoderm), CD31 (mesoderm), and GATA4 (endoderm), were tested by immunostaining. Although both –MOP-1 EB and –LIF EB groups could differentiate into different cell types in all three germ layers, the expression level of CD31 in the –MOP-1 EB group was significantly higher than that in the –LIF EB group (Supplementary Figs. 23A and 24A). Subsequently, the EB was allowed to remain adherent for in vitro spontaneous differentiation to compare the pluripotency of MOP mESC and LIF mESC. The results of MAP2, CD31 and GATA4 marker expression showed that MOP-1 mESC were pluripotent (Supplementary Figs. 23B and 24B). For further differentiation assay, functional beating myocardial cells were acquired (Supplementary Movie 1 and 2). Then, we applied small-molecule compound retinoic acid (RA) for direct neuron differentiation, neuron marker MAP2 was expressed in MOP-1 J1 mESC-derived cells and MOP-1 E14 mESC-derived cells (Supplementary Fig. 25). In addition to the pluripotency of MOP-1 mESC, it is noteworthy that the EB formed by MOP-1 mESC exhibit a lower apoptosis score (Fig. 4F), the ratio of dead cells area and the number of dead cells (Fig. 4G, H and Supplementary Fig. 26) compared to those formed by LIF mESC (J1 and E14 mESC lines). Immunofluorescent analysis was then used to further assess the expression of the proliferation marker Ki67 and the apoptosis marker Caspase3 (Casp3) in the –MOP-1 EB and –LIF EB groups. Both markers were expressed at the edges, but Casp3 expression in –LIF EB was nearly ubiquitous throughout the entire area, significantly more so than in –MOP-1 EB (Supplementary Fig. 27). This indicates that –MOP-1 EB is more effective in reducing cell apoptosis and supporting cell survival and/or proliferation during differentiation than –LIF EB.
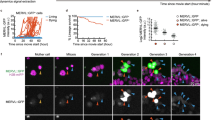
To validate the differentiation ability of MOP-1 mESC under complex physiological conditions in vivo, LIF mESC or MOP-1 mESC were injected into subcutaneous regions of shoulders in nude mice to yield differentiated teratomas (Fig. 5A). The body weight of mice and formed masses derived mESC were carefully recorded (Fig. 5B). The proliferation of teratomas derived from MOP-1 mESC (denoted as –MOP-1 teratoma) was more quickly than teratomas derived from LIF mESC (denoted as –LIF teratoma) at D14 (Fig. 5C-F), while the apoptosis of –MOP-1 teratomas was less than –LIF teratomas (Fig. 5G, H). These results confirm that MOP-1 mESC has differentiation potential, while apoptosis is reduced during the differentiation process of –MOP-1 teratomas. Lastly, we conducted the integrated assessment of germ layer differentiation. H&E staining was used to visualize the structural features of the three germ layers contained squamous epithelium (ectoderm), smooth muscle (mesoderm), and columnar epithelium (endoderm), which revealed the teratoma formation capacity of MOP-1 mESC in vivo is as effective as LIF mESC (Fig. 5I). Ectodermal, mesodermal and endodermal differentiation were also observed by immune-histochemical detection of their representative markers (GFAP, a marker of astrocyte found in the nervous system; CD31, a marker of vascular endothelial cells found in the cardiovascular system; AFP, α-fetoprotein made in the hepatocytes), respectively. MOP-1 mESC exhibited excellent ability of both self-renewal and multidirectional differentiation (Fig. 5J).
A The schematic of LIF mESC or MOP-1 mESC was injected into the subcutaneous regions of the shoulders in nude mice. Schematic diagram was created with Adobe Illustrator. Body weight (B) and tumor volume (C, D) were calculated as indicated in Materials and Methods (mean ± s.e.m, n = 10 in (B) or n = 5 mice per group in (D)). Representative images (E and G) and quantitative analysis (F and H) of proliferation marker Ki67 (E and F) and apoptosis marker Casp3 (G and H) expressed in teratoma derived from –LIF and –MOP-1 mESC (mean ± s.e.m, n = 14 from 4 mice teratoma samples per group). Scale bars, 50 μm. Differentiation of MOP-1 mESC into ectoderm, mesoderm, and endoderm by H&E staining (I) and immunohistochemical staining (J) of teratoma sections. Ectoderm, keratin pearl. Mesoderm, muscle cell. Endoderm, epithelial cell. Three experiments were repeated independently from 4 mice teratoma samples per group with similar results. Scale bars, 20 μm. Data in B, D, F and H are analyzed by a two-tailed unpaired t-test. *P < 0.05, **P < 0.01, ****P < 0.0001, –LIF teratoma vs. –MOP-1 teratoma.
Discussion
Compared to conventional nanoparticles, MOPs have the advantages of good monodispersity, well-defined structural composition, and tunable surface properties. This provides a blueprint to fabricate the MOPs with the desired processability and functions. Furthermore, the precise design of MOP-1 based on application scenarios can facilitate the development of drug screens using mESC or mESC-derived cells. In the present study, we present an excellent example that integrated interdisciplinary expertise from chemistry and biology helps to design subtle functionalized materials for cell fate control and may realize molecularly precise disease therapy in the future.
We develop a facile, efficient, robust and low-cost strategy using MOPs as a revolutionary method to facilitate and simplify mESC culture. Furthermore, we conclusively investigated the differentiation ability of MOP-1 to maintain the self-renewal and pluripotency of mESC. Though the clone morphology, the activity of ALP and the expression levels of self-renewal related genes of MOP-1 mESC were the same as LIF mESC. Subsequent culture experiments with mESC and hESC confirmed that both MOP-1 and LIF exhibit substantial capacity to maintain stemness. EB and teratomas formed by MOP-1 mESC can naturally differentiate into the three germ layers, exhibiting lower expression of apoptosis markers and higher expression of proliferation markers compared to LIF. This indicates that –MOP-1 EB is more effective than –LIF EB in reducing cell apoptosis and promoting cell survival and/or proliferation during differentiation. The underlying mechanism by which MOP-1 maintains mESC pluripotency was explored through enzyme assays and molecular docking. We performed molecular modeling exploration by the molecular docking method. MOP-1 was immobilized in the catalytic pocket of SHP-2 through a variety of hydrogen bonds, salt bridges and π-π stacking interactions and binds perfectly to SHP-2 to inactivate it. In short, MOP-1 could maintain mESC pluripotency via counteracting SHP-2 mediated STAT3 inactivation because the structure of MOP-1 perfectly matches up to SHP-2 inhibition. In summary, we have validated through various approaches, including transcriptomics and single-cell genomics, that MOP-1 can maintain ESC self-renewal and potential pluripotency. However, due to ethical concerns, whether MOP-1 mESCs or hESCs have the potential to form chimeras remains to be further investigated.
In addition to possessing similar biological functions to LIF, MOP-1 has more advantages, as being easy to synthesize and resistant to alcohol and high temperatures, etc. The well-defined structure of MOPs helps to pinpoint the site of action and regulate the pluripotency of stem cells at the molecular level. The high stability of MOPs can greatly broaden the operational conditions, reduce costs, and facilitate the preparation and storage of related biopharmaceuticals. These extraordinary properties surpass traditional materials. Based on the cavities, MOPs have the potential to achieve the dual functionality of maintaining pluripotency and drug delivery in the future.
Methods
Materials
All reagents were purchased from commercial suppliers and used without further purification. 1,4-benzenedicarboxylate (H2BDC), 2-amino-1,4-benzenedicarboxylate (H2BDC-NH2), vanadium(III) chloride (VCl3), bis(cyclopentadienyl)zirconium dichloride (Cp2ZrCl2), chloroauric acid (HAuCl4), trisodium citrate, chloroplantinic acid (H2PtCl6), sodium borohydride (NaBH4) were purchased from Jilin Chinese Academy of Sciences-Yanshen Technology Co., Ltd. N,N’-dimethylformamide (DMF), N,N’-diethylformamide (DEF) were all provided by Aladdin.
Characterization
Powder X-ray diffraction (PXRD) patterns were recorded in the range from 5 to 40° at room temperature on a Rigaku dmax 2500 diffractometer using Cu Kα (λ = 1.5418 Å) radiation, with a scan speed of 1° min−1. Fourier transform infrared spectroscopy (FT-IR) was performed using a Thermo Fisher spectrometer using ATR modes (128 scans, 2 cm−1 resolution). The UV-vis absorption spectra were recorded from 200–800 nm using PerkinElmer (Lambda 950) UV-visible spectrometer. The scanning electron microscopy (SEM) images were obtained on HITACHI SU3500. Dynamic light scattering (DLS), including nanoparticle size and zeta potential analysis, was measured using Malvern Panalytical Nano-ZS in an aqueous dispersion. MOP-1, PtNPs and IONPs morphologies were imaged using the JEM-1400Flash transmission electron microscope (TEM, JEOL).
Synthesis of MOP-1
MOP-1 was synthesized according to the literature with a little modification41. H2BDC-NH2 (20 mg, 0.11 mmol) and VCl3 (30 mg, 0.19 mmol) were dissolved in DMF (2 mL) and ethanol (0.5 mL). Then the mixtures were placed in a Teflon-lined stainless-steel vessel and heated at 160 °C for 2 d. After slow cooling to room temperature, dark green crystals were obtained and they were washed with ethanol and dried in air.
Synthesis of MOP-2
MOP-2 was synthesized according to the literature41. H2BDC (32 mg, 0.19 mmol) and VCl3 (30 mg, 0.19 mmol) were dissolved in DMF (2 mL) and ethanol (0.5 mL). Then, the mixtures were placed in a Teflon-lined stainless-steel vessel and heated at 150 °C for 2 d. After slow cooling to room temperature, dark green crystals were obtained and they were washed with ethanol and dried in air.
Synthesis of ZrMOP
ZrMOP was synthesized according to the literature47. To a 10 mL vial, Cp2ZrCl2 (17.5 mg, 0.06 mmol), H2BDC-NH2 (5.4 mg, 0.03 mmol), DEF (1.0 mL) and water (300 μL) were added. The mixture became a clear yellow solution after sonication. The solution was heated in a preheated oven at 60 °C for 8 h. Yellow cubic crystals were synthesized and washed with fresh DEF. The product was dried at 60 °C in a vacuum overnight.
Synthesis of other nanoparticles
For Au nanoparticle synthesis, 50 mL HAuCl4 solution (1 mM) was obtained by vigorous stirring in a round-bottomed flask and then heated to a boil. A 5 mL trisodium citrate solution (38.8 mM) was rapidly added to the boiling solution, which induced a color change from pale yellow to burgundy. After 10 min, the boiling solution was placed at room temperature and stirred for 15 min. For Pt nanoparticle synthesis, 1 mL H2PtCl6 solution (16 mM) and 1 mL trisodium citrate solution (40 mM) were added into 38 mL deionized H2O with continuous stirring. A 200 μL NaBH4 (50 mM) solution was added dropwise into the above solution. Pt nanoparticles could be obtained by stirring at room temperature for another 1 h.
SHP-2 binding kinetics measured by SPR
The binding kinetics and affinity of MOP-1 or MOP-2 to SHP-2 (MCE, HY-P700618), SHP-1 (MCE, HY-P71141), JAK1 (MCE, HY-P700583), JAK2 (MCE, HY-P701102) and PTP1B (MCE, HY-P73685) were analyzed by SPR (Biacore 8 K, Cytiva). Specifically, proteins were covalently immobilized to CM5 sensor chips via amine groups in 10 mM sodium acetate buffer (pH 5.0). By calculating the coupling amount of proteins, Rmax was less than 100. SPR assays were run at a flow rate of 30 mL min−1 in 1xPBS buffer. Serial dilutions of MOP-1 or MOP-2 were injected, ranging in concentration from 200 to 15.625 nM. The dilution factor was 1:3. The resulting data fit a 1:1 binding model using Biacore Insight Evaluation Software.
Generation of stable cell lines
E14 ESC cell line stably expressing pCMV-PTPN11-mCherry-Neo (SHP-2-mCherry) was generated using the Lipo2000 (Thermo, 11668019) transduction strategy. To prepare the transfection complexes, 250 µL of Opti-MEM was added to an EP tube, followed by the addition of 10 µL of Lipo2000. The mixture was gently mixed and incubated at room temperature for 5 min to create the Lipo2000 dilution. Separately, another EP tube was prepared by adding 250 µL of Opti-MEM and 4 µg of plasmid DNA, which was then gently mixed to form the DNA dilution. The Lipo2000 dilution from the first tube was subsequently added to the DNA dilution in the second tube. This combined solution was gently mixed and allowed to sit at room temperature for 20 min, resulting in the formation of DNA-Lipo2000 complexes. Approximately 1 h before transfection, the cell culture medium was replaced with fresh, antibiotic-free, and serum-free medium to optimize transfection conditions. The prepared DNA-Lipo2000 complexes were then added to the cell culture medium, and the six-well plate was gently rocked to ensure even distribution of the complexes across the cells. The cells were subsequently incubated in a 37 °C incubator with 5% CO2 for 24 h to allow for transfection. Six hours post-transfection, the medium was replaced with complete culture medium to support cell recovery and expression of the transfected genes. After 24 h, the solution was changed and G418 was added to screen positive cells for 7 d.
Mouse embryonic stem cells (mESC) and embryoid bodies (EB) culture
J1 mESC and E14 mESC were cultured in DMEM supplemented with 15% fetal bovine serum (FBS, ExCell, FND500), 1% nonessential amino acids (NEAA, Gibco, 11140050), 1% L-Glutamine (Gibco, 35050061), 1% penicillin/streptomycin (PS, Gibco), 100 µM β-mercaptoethanol Gibco, 21985023) and 1000 U mL−1 LIF (Sigma-Aldrich, ESG1107) at 37 °C and 5% CO2.
T4 mESC, donated by the laboratory of Shuai Ling at Nankai University. T4 mESC were cultured in 2i+LIF (2iL) medium (1:1 mixture of DMEM/F−12 medium (Gibco, C11330500BT) and neurobasal medium (Gibco, 21103049)), which was supplemented with N2 supplement, B27 supplement, 1% nonessential amino acids, 1% L-Glutamine, 1% penicillin/streptomycin, 1 µM PD0325901, 3 µM CHIR99021 and 1000 U mL−1 LIF. mESC colonies were digested with accutase and then gently pipetted with a pipette tip to dissociate them into single cells. The single-cell suspension was counted and diluted to a density of 104 mL−1 for seeding.
Single cells dissociated from mESC aggregated into cellular spheroids in non-adherent Petri dishes, which were also called EB. For further spontaneous differentiation, EB was transferred to plates in N2B27 medium (1:1 mixture of DMEM/F-12 medium and neurobasal medium), which was supplemented with N2 supplement, B27 supplement, and 0.5% L-Glutamine, 1% penicillin/streptomycin, 100 µM β-mercaptoethanol, 2% bovine serum albumin (BSA).
mESC were passaged via accutase after culturing with LIF or MOP-1 and complete dissociation of colonies by pipetting. To begin differentiation (Day 0), cells were trypsinized, removed from LIF or MOP-1 medium, and cultured in suspension in serum-free differentiation medium (N2B27 medium) at a density of 75000 cells mL−1 to form embryoid bodies (EB). At this point (Day 4.5), we collected the EB for single cell sequencing and detected the RNA expression.
For directed differentiation, the EB was transferred to adherent plates in N2B27 medium. The EB were cultured in N2B27 differentiation medium containing retinoic acid (RA) at a concentration of 5 mM for induced neuronal differentiation.
Cell viability assays
mESCs were incubated with MOP-1 (MOP-2/ZrMOP/Ligand) medium containing several concentrations of vanadium element for 48 h. mESC cultured in the medium with LIF was used as the control. The supernatant was replaced with 20 μL Methylthiazolyldiphenyl-tetrazolium bromide (MTT) and incubated for 4 h at 37 °C. Then, 150 μL of DMSO was added to each well and the plate was shaken at 50 g for 10 min. The absorption value of each well was measured using a multi-function microplate reader at 490 nm.
Human embryonic stem cell (hESC) culture and EB culture
RUES2 hESC line-derived and kindly provided by Prof. Leqian Yu’s Laboratory (Key Laboratory of Organ Regeneration and Reconstruction, State Key Laboratory of Stem Cell and Reproductive Biology, Institute of Zoology). RUES2 cells were cultured in mTeSR1 medium (STEMCELL, 85850) or mTeSR1 medium without select factors (STEMCELL, 05896). mTeSR1 without select factors is the normal mTeSR1 formulation but without bFGF, TGF-beta, LiCl, GABA, and pipecolic acid. mTeSR1 without bFGF was prepared by supplementing mTeSR1 without select factors with TGF-beta, LiCl, GABA, and pipecolic acid to evaluate the effects of these supplements on the pluripotency of hESC. Proliferating RUES2 cells grew in adhesion on Geltrex™ (Thermo Fisher Scientific).
EB were formed using AggreWell plates (Stemcell Technologies, 34811) according to the manufacturer’s instructions. In brief, hESC were collected using Gentle Cell Dissociation Reagent, centrifuged at 300 g and re-suspended in AggreWell EB formation medium (Stem Cell Technologies, 5893) with 10 μM ROCK inhibitor (Stem Cell Technologies, 72302). Cells (1.0 × 107) were added to one well of an AggreWell 800ex plate and spun at 100 g for 3 min. After centrifugation, the plates were transferred to an incubator at 37 °C. After 3 d, the embryoid bodies were collected with gentle pipetting.
Flow cytometry analysis of apoptotic cells
The MOP-1 administration was operated according to the previous method. For cell apoptosis detection, the cells were washed twice with PBS and then they were mixed with Annexin V and propidium iodide (PI). Left the mixture in the dark for 15 min at room temperature. Cellular fluorescence was measured by a flow cytometer within 1 h. For cell cycle detection, the cells were washed twice with PBS. We gently mixed cells with 70% ethanol at 4 °C overnight for fixation. Thereafter, RNase was added to the cells, then the cells were incubated with PI in the dark for 30 min at room temperature. Cellular fluorescence was measured using a flow cytometer.
The proportion of undifferentiated cells was analyzed by FACS. The 3-5 × 105 cells were plated on Day 0 into a conditioned medium, TeSR1, TeSR1 minus the bFGF or TeSR1 minus the bFGF containing MOP-1. Cells were cultured for three passages. At the end of passage three, cells were individualized by treatment with Gentle Cell Dissociation Reagen (Stem Cell Technologies, 100-0485) and were analyzed by FACS for ESC markers OCT4 (Santa Cruz Biotechnology, sc-5279), SSEA4 (Abcam, ab16287), NANOG (MCE, HY-P80237) and TRA 1-81 (Thermo, MA1-024X). We used Alexa Fluor® 647-conjugated AffiniPure® Goat Anti-Mouse IgG (H + L) (Jackson, 115-605-003) and Fluorescein (FITC)-conjugated AffiniPure Goat Anti-Rabbit IgG (H + L) (Jackson, 111-095-003) as the secondary antibody.
Alkaline phosphatase staining assay
The MOP-1 administration was operated according to the previous method. Cells (BCIP/NBT Alkaline Phosphatase Color Development Kit, Beyotime, C3206) were fixed with 4% paraformaldehyde and incubated with BCIP/NBT staining work solution reagent in the dark for 30 min. Finally, stop the color development reaction by removing the BCIP/NBT staining solution and washing with PBS 1-2 times, then observe under the Total internal reflection fluorescent microscope TIRF&Thunder (Leica, DMi8).
The metabolism of MOP-1 in mESC
2 μM MOP-1 was added to J1 mESC, and cells were collected at 0 h, 12 h, 24 h, and 48 h for measurement of V element content. The cell supernatant was discarded and then 0.25 % Try was added to collect the cells. After centrifugation at 200 g for 5 min, the cells were washed with PBS and centrifuged at 200 g for 5 min each time. The mESC was counted, the cell precipitate was nitrated, and the content of V element in the cells was measured by ICP-MS (Agilent 7800(MS)).
The binding of MOP-1, MOP-2 and SHP-2
Five groups of SHP-2, MOP-1 or MOP-2, and MOP-1 + SHP-2 or MOP-2 + SHP-2 have been set up. In the MOP-1 + SHP-2 and MOP-2 + SHP-2 groups, 0.2 mL of 1 mg mL−1 MOP-1 or MOP-2 and 6.5 μg SHP-2 were mixed and incubated at 4 °C for 6 h. In the SHP-2 group, only 6.5 μg SHP-2. In the MOP-1 or MOP-2 group, only 0.2 mL of 1 mg mL−1 MOP-1 or MOP-2. After 6 h, these samples were centrifuged at 12000 g for 15 min, and the supernatant was taken to measure vanadium by ICP-MS (Agilent 7800(MS)). Finally, the content of MOP-1 or MOP-2 bound in SHP-2 was calculated by deducting the background element content of MOP-1 or MOP-2.
Real-time quantitative polymerase chain reaction (RT-qPCR)
The MOP-1 administration and EB formation were operated according to the previous method. The formed EB (on the 4th day) were collected for the following detection. Total RNA was extracted using TRIzol Reagent (Invitrogen) according to the manufacturer’s protocol. cDNA was synthesized with TransScript all-in-one first-strand cDNA synthesis SuperMix for qPCR (TransGen Biotech). To evaluate genomic DNA contamination, a negative control was prepared in parallel by treating the same amount of RNA in the same way but without adding the reverse transcriptase enzyme. qPCR was performed with Platinum SYBR Green qPCR SuperMix (TransGen Biotech) in a CFX touch 96 machine. The primer sequences used for qPCR are listed in Supplementary Table 4.
RNA sequencing and analysis
The integrity of RNA extracted from mESC cultured with LIF or MOP-1 for one week was assessed using the RNA Nano 6000 Assay Kit of the Bioanalyzer 2100 system (Agilent Technologies, CA, USA). The clustering of the index-coded samples was performed on a cBot Cluster Generation System using TruSeq PE Cluster Kit v3-cBot-HS (Illumia) according to the manufacturer’s instructions. After cluster generation, the library preparations were sequenced on an Illumina Novaseq platform and 150 bp paired-end reads were generated. Differential expression analysis of two conditions/groups (two biological replicates per condition) was performed using the DESeq2 R package (1.20.0). DESeq2 provides statistical routines for determining differential expression in digital gene expression data using a model based on the negative binomial distribution. The resulting P-values were adjusted using Benjamini and Hochberg’s approach for controlling the false discovery rate. Genes with an adjusted P-value ≤ 0.05 found by DESeq2 were assigned as differentially expressed. HTseq was used to get the FPKM method to determine gene expression. FPKM data was z-scored, transformed and plotted using the Bioconductor pheatmap55,56.
Single-cell association
Single-cell RNA-seq experiment was performed by experimental personnel in the laboratory of NovelBio Co., Ltd. The tissues were removed and kept in MACS Tissue Storage Solution (Miltenyi Biotec) until processing. The tissue samples were processed as described below. Briefly, samples were first washed with phosphate-buffered saline (PBS), minced into small pieces (approximately 1mm3) on ice and enzymatically digested with 0.25% Trypsin (without EDTA) for 5 min at 37 °C, samples were sieved through a 70 µm cell strainer, the supernatant was added to an equal volume of 1640 culture solution containing 10% FBS to terminate digestion; After washing with PBS containing 0.04% BSA, the cell pellets were re-suspended in PBS containing 0.04% BSA and re-filtered through a 35 μm cell strainer. Dissociated single cells were then stained with AO/PI for viability assessment using a Countstar Fluorescence Cell Analyzer.
Single-cell RNA sequencing
The scRNA-Seq libraries were generated using the 10X Genomics Chromium Controller Instrument and Chromium Single Cell 3’ V3 Reagent Kits (10X Genomics, Pleasanton, CA). Briefly, cells were concentrated to approximately 1000 cells µL–1 and loaded into each channel to generate single-cell Gel Bead-In-Emulsions (GEMs). After the RT step, GEMs were broken and barcoded-cDNA was purified and amplified. The amplified barcoded cDNA was fragmented, A-tailed, ligated with adaptors and index PCR amplified. The final libraries were quantified using the Qubit High Sensitivity DNA assay (Thermo Fisher Scientific) and the size distribution of the libraries was determined using a High Sensitivity DNA chip on a Bioanalyzer 2200 (Agilent). All libraries were sequenced by an Illumina sequencer (Illumina, San Diego, CA) on a 150 bp paired-end run.
Single cell RNA data processing and analysis
Data processing
scRNA-seq data analysis was performed by NovelBio Co., Ltd. with NovelBrain Cloud Analysis Platform. We applied fastp57 with default parameter filtering the adaptor sequence and removed the low-quality reads to achieve clean data. Then, the feature-barcode matrices were obtained by aligning reads to the human genome (mm10 Ensemble: version 100) using CellRanger v7.1.0. We applied the downsample analysis among samples sequenced according to the mapped barcoded reads per cell of each sample and finally achieved the aggregated matrix. Cells contained over 200 expressed genes and the mitochondria UMI rate below 20% passed the cell quality filtering and mitochondria genes were removed in the expression table.
Seurat package (version:4.1.1, https://satijalab.org/seurat/) was used for cell normalization and regression based on the expression table according to the UMI counts of each sample and the percent of mitochondria rate to obtain the scaled data. PCA was constructed based on the scaled data with the top 2000 highly variable genes and the top 10 principal components were used for t-SNE construction and UMAP construction. Utilizing the graph-based cluster method, we acquired the unsupervised cell cluster result based on the PCA top 10 principal and we calculated the marker genes by the FindAllMarkers function with the Wilcox rank sum test algorithm under the following criteria: 1. Log2FC > 0.25; 2. p value < 0.05; 3. min. pct > 0.1. To identify the cell type in detail, the clusters of the same cell type were selected for re-t-SNE analysis, graph-based clustering and marker analysis.
Data analysis
Gene ontology (GO) analysis
To elucidate the biological implications of the DEGs and marker genes, GO analysis58 was performed. GO annotations were downloaded from NCBI, the Gene Ontology database (http://www.geneontology.org/) and UniProt (http://www.UniProt.org/). Fisher’s exact test was applied to identify the significant GO categories, and an FDR was used to correct the p values.
Pathway analysis
Pathway analysis59 was used to explore the significant pathways of the DEGs and marker genes based on the Kyoto Encyclopedia of Genes and Genomes (KEGG) database. Fisher’s exact test was used to identify significant pathways, and the threshold of significance was defined by the p value and FDR.
Gene enrichment analysis
For gene enrichment analysis, Fisher’s exact test was applied to calculate the P value for each gene set. Raw P values were adjusted for multiple hypothesis tests using the Benjamini and Hochberg method. Such enrichment analysis was applied to annotations including GO (v2.5.13), the Kyoto Encyclopedia of Genes and Genomes (KEGG, 201900613), the Hallmark (h.all.7.0., https://www.gsea-msigdb.org/gsea/msigdb/index.jsp), Chromosome location gene set (https://ftp.ncbi.nlm.nih.gov/gene/DATA/GENE_INFO/Mammalia/) and customized gene sets. A customized 41-signature gene set including immune-, cytokine- and neurobiology-related terms were collected from the CellphoneDB database. To characterize the relative activation of a given gene set, we performed QUSAGE (2.16.1)60 analysis.
SCENIC analysis
To assess the regulatory strength of TFs, we applied the SCENIC (v0.11.2)61 workflow, which is a computational method for the construction of regulatory networks and identification of different cell states from scRNA-seq data, using the 20-thousand motif database for RcisTarget and GRNboost.
Pseudotime analysis
Single-cell trajectory analysis was performed with Monocle262 (http://cole-trapnell-lab.github.io/monocle-release) utilizing DDR-Tree and default parameters to determine the dramatic translational relationships among cell types and clusters. Before Monocle analysis, marker genes of the Seurat clustering result and raw expression counts of the cell that passed filtering were selected. Branch expression analysis modeling (BEAM analysis) was utilized for branch fate-determined gene analysis based on pseudotime analysis.
RNA velocity
To perform the RNA velocity analysis, we annotated the spliced reads and unspliced reads using the velocity Python package based on previously aligned BAM files of scRNA-seq data. The calculation of RNA velocity values for each gene in each cell and embedding RNA velocity vector in a low-dimension space were done by following the scVelo Python pipeline. The velocity-based cell transition matrix was calculated by using the transition matrix function from scVelo. We estimated the destination of a cell by identifying the highest correlation value in the matrix.
CytoTrace
Cellular Trajectory Reconstruction Analysis using gene Counts and Expression (CytoTRACE, v0.3.3)63, a computational method, was applied to predict the differentiation state of cells from single-cell RNA-sequencing data by counting the number of expressed genes per cell.
Cell-cell communication analysis
We applied cell communication analysis based on the CellPhoneDB64 (v1.1.0), a public repository of ligands, receptors and their interactions. Membrane secreted and peripheral proteins of the cluster were annotated. Significant mean and Cell Communication significance (p-value < 0.05) were calculated based on the interaction and the normalized cell matrix achieved by Seurat Normalization.
Differential gene expression analysis
To identify differentially expressed genes among samples, the function FindMarkers with the Wilcox rank sum test algorithm was used under the following criteria: 1. Log2FC > 0.25; 2. p value < 0.05; 3. min. pct > 0.1, which was more conducive to obtaining different functional genes among samples.
Teratoma assay
All animal studies were conducted under the guidelines set by Tianjin Committee of Use and Care of Laboratory Animals, and the overall project protocols were approved by the Animal Ethics Committee of Nankai University (approval ID: 2021-SYDWLL-000318). mESC at a density of 1×107 mL−1 were injected subcutaneously into nude mice, and the same number of LIF-containing medium-cultured mESC were injected as the control group. The volume of teratomas in mice was calculated by the formula: Volume = (width)2 × length/2. The body weight of the nude mouse was recorded twice a week.
Hematoxylin and eosin staining
Teratomas were fixed in 4% paraformaldehyde at 4 °C for 24 h. The fixed tissues were then transferred to an increasing gradient of ethanol for dehydration. Then the tissues were transferred to dimethylbenzene for transparency. The dehydrated transparent tissues were immersed in liquid paraffin and they were cut into 6 µm slices. Hematoxylin and eosin staining was performed according to the manufacturer’s instructions.
Western blot
Prepared processed protein samples and SDS-PAGE gel. After electrophoresis, proteins were transferred to PVDF membranes. After being blocked with BSA, SDS-PAGE and western blotting analysis were carried out according to standard procedures using the following antibodies: anti-Phospho-Stat3 (Tyr705) (Cell Signaling Technology, 9145 T) at 1:1000, anti-Phospho-SHP-2 (Tyr542) (Cell Signaling Technology, 3751 T) at 1:500, anti-Stat3 (F-2) (Santa Cruz Biotechnology, sc-8019) at 1:500, anti-OCT3/4 (Santa Cruz Biotechnology, sc-5279) at 1:500, anti-SOX2 (abcam, ab97959) at 1:1000, anti-Ki67 (Invitrogen, MA5-14520) at 1:1000, anti-Caspase-3 (Cell Signaling Technology, 9661 T) at 1:1000, anti-β-Actin (EASYBIO, BE0021) at 1:2000. Secondary HRP goat anti-rabbit (EASYBIO, BE0101) antibody and secondary HRP goat anti-mouse (EASYBIO, BE0102) antibody were used at 1:5000.
Immunofluorescence staining of mESC
4% paraformaldehyde was used for fixation, and samples were washed 3 times with PBS. After being blocked with BSA for 15 min, samples were incubated with primary antibodies, diluted at a 1:200 volume ratio with BSA, overnight at 4 °C. Then, the samples were incubated with secondary antibodies. Immunofluorescence staining was carried out according to standard procedures using the following antibodies: anti-OCT3/4 (Santa Cruz Biotechnology, sc-5279) at 1:50, anti-SOX2 (Abcam, ab97959) at 1:200, anti-Ki67 (Invitrogen, MA5-14520) at 1:200, anti-MAP2 (Abcam, ab32454) at 1:200, anti-CD31 (Santa Cruz Biotechnology, sc-20071) at 1:50, anti-Caspase-3 (Cell Signaling Technology, 9661 T) at 1:200. Secondary FITC-conjugated affinipure goat anti-rabbit (Jackson ImmunoResearch, 111-095-003) IgG (H + L) antibody and secondary TRITC-conjugated affinipure goat anti-rabbit (Jackson ImmunoResearch, 111-025-003) IgG (H + L) antibody were used at 1:5000.
Assessment of SHP-2 Inhibition
Assessment of SHP-2 inhibition utilized recombinant human SHP-2 protein (ab42578) and its substrate (disodium 4-nitrophenylphosphate, p-NPP). 10 mL of test compound MOP-1 at decreasing concentrations was spotted into a 96-well plate, followed by the addition of 5 μL of SHP-2 enzyme in assay buffer (50 mM HEPES, pH = 7.4, 2 mM EDTA, 3 mM DTT, 100 mM NaCl). After a 30-minute incubation at room temperature, 50 μL of p-NPP (50 mM) was added to the assay buffer. The final concentration of SHP-2 was 2.5 μg mL−1, and the total volume was 200 μL. The final concentration of the MOP-1 ranged from 0 μM to 20 μM. After a 30-minute incubation, the absorbance of the samples at 405 nm was recorded. Percent inhibitions were calculated based on control wells containing no MOP-1 (0% inhibition). The other two MOPs were assessed for their enzyme inhibitory ability in the same way.
Assessment of other proteins inhibition
The inhibitions of SHP-2, PTP1B, SHP-1, ACP, Cyt c and lipase by MOP-1 were assessed. 10 μL of the test compound MOP-1 at decreasing concentrations was spotted into a 96-well plate, followed by the addition of 5 μL of ACP, lipase, or Cyt c in assay buffer (50 mM HEPES, pH 7.0). After a 30-minute incubation at room temperature, 50 μL of p-NPP (50 mM), p-NPA (50 mM) or ABTS (10 mM), and H2O2 (100 mM) were added to the assay buffer, respectively. Final concentrations of SHP-2, PTP1B, SHP-1, ACP, Cyt c and lipase were 25 μg mL−1, 62.5 μg mL−1 and 0.25 μg mL−1, respectively. The total volume was 200 μL. The final concentration of the test compound MOP-1 ranged from 0.025 μg mL−1 to 5 μg mL−1. After 30 min incubation, the absorbance of the samples at 405 nm, 405 nm, or 420 nm was recorded. Percent inhibitions were calculated based on control wells containing no compound MOP-1 (0% inhibition). The inhibitions of JAK2, JAK1 by MOP-1 were also assessed using the Kinase-Lumi™ Chemiluminescent Kinase Activity Assay Kit (Beyotime, S0150S).
The stability of MOP-1 in H2O and mESC cultured medium
MOP-1 was dissolved in H2O and mESC cultured medium (LIF or 2iL medium) at a concentration of 0.8 mM, respectively. Let them stand at room temperature for 10 d. After diluting 10-fold, a UV-vis spectrophotometer was used to track the spectra of MOP-1 in these solutions. The UV-vis spectra of fresh MOP-1 aqueous solution were used as controls.
The stability of MOP-1 in DMEM medium
MOP-1 was dissolved in a DMEM medium at a concentration of 1 mM. Let it stand at room temperature. After diluting 10-fold, a UV-vis spectrophotometer was used to track the spectra of MOP-1 at different times.
The long-term stability of MOP-1 in H2O
MOP-1 was dissolved in H2O at a concentration of 0.6 mM. Let it stand at room temperature. After diluting 10-fold, a UV-vis spectrophotometer was used to track the spectra of MOP-1 at different times.
The thermostability of MOP-1
MOP-1 was dissolved in H2O at a concentration of 10.0 mM. After being treated by heating (100 °C) for 1 h or 5 h, the spectra of MOP-1 were detected by a UV-vis spectrophotometer from 250 nm to 800 nm.
The stability of MOP-1 after 75% ethanol treatment
30 mg MOP-1 was treated with 2 mL 75% ethanol for 5 min. We centrifuged at 1000 g, collected the powders of MOP-1, and tested the PXRD after drying.
Molecular docking
The SHP-2 structure in the unphosphorylated state was extracted from Protein Data Bank crystal structures (PDB ID: 2SHP)65. Firstly, the SH2 domain and unnecessary small molecules were removed from this crystal structure66. The protonation state for residues at neutral pH was determined using H + +, and the hydrogen atoms and missing side chains of the SHP-2 structure were added by the Leap module of Amber2067. Next, we docked MOP-1 and its variants (MOP-1-Model1 to MOP-1-Model4) into the phosphorylation sites on residues Cys459 and Arg465 using AutoDock468. The MOP-1 structure maintains the atoms in one of the vertices of the hexahedron (V5O9Cl) and four H2BDC-NH2 molecules around the vertex as the “vertex group”, in which other atoms were removed from the structure. The free energy of binding between the SHP-2 and the vertex group of MOP-1 was calculated and given by AutoDock4, which could be used to assess the affinity between them. Rotatable bonds in the vertex group of MOP-1 were restricted, the grid size was set to be 65 Å × 65 Å × 65 Å. The docking mode with the lowest free energy was chosen as the binding mode to gain the MOP-1-bound SHP-2 complex. We applied the same procedure to dock MOP-1-Model1 to MOP-1-Model4 and the other two MOPs with SHP-2 and assessed their binding modes similarly.
Statistics
Statistical analyses were conducted via GraphPad Prism 8.3 software. Statistical comparisons between the two groups were assessed using unpaired t-tests. For multiple comparisons, statistical significance was determined by one-way analysis of variance (ANOVA). Statistical differences in EB data were analyzed by two-way repeated measures ANOVA. Statistical significance was indicated as *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001. All data were presented as mean ± standard error of the mean (SEM).
Ethics
Every experiment involving animal samples has been carried out following a protocol approved by an ethical commission.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings of this study are available within the article and its supplementary files. Any additional requests for information can be directed to and will be fulfilled by the corresponding authors. Source data are provided with this paper.
References
Rathjen, J. & Rathjen, P. Embryonic stem cells. In: Brenner’s encyclopedia of genetics (second edition) (eds Maloy, S. & Hughes, K.). Academic Press (2013).
Genchi, A. et al. Neural stem cell transplantation in patients with progressive multiple sclerosis: an open-label, phase 1 study. Nat. Med. 29, 75–85 (2023).
Wang, Y.-K. et al. Human clinical-grade parthenogenetic ESC-derived dopaminergic neurons recover locomotive defects of nonhuman primate models of Parkinson’s disease. Stem Cell Rep. 11, 171–182 (2018).
Cyranoski, D. Trials of embryonic stem cells to launch in China. Nature 546, 15–16 (2017).
Karvelas, N. et al. Advances in stem cell therapy in Alzheimer’s disease: a comprehensive clinical trial review. Stem Cell Investig. 9, 2 (2022).
Han, F. et al. Stem cell therapy for Alzheimer’s disease. Adv. Exp. Med. Biol. 1266, 39–55 (2020).
Liu, X.-Y. et al. Stem cell therapy for Alzheimer’s disease. World J. Stem Cells 12, 787–802 (2020).
Assunção-Silva, R. C. et al. Hydrogels and cell based therapies in spinal cord injury regeneration. Stem Cells Int 2015, 948040 (2015).
Jin, M. C. et al. Stem cell therapies for acute spinal cord injury in humans: a review. Neurosurg. Focus 46, E10 (2019).
Uzel, S. G. et al. Microfluidic device for the formation of optically excitable, three-dimensional, compartmentalized motor units. Sci. Adv. 2, e1501429 (2016).
Hirai, H. et al. Regulation of embryonic stem cell self-renewal and pluripotency by leukaemia inhibitory factor. Biochem. J. 438, 11–23 (2011).
Takahashi, K. & Yamanaka, S. A decade of transcription factor-mediated reprogramming to pluripotency. Nat. Rev. Mol. Cell Biol. 17, 183–193 (2016).
Chen, G. et al. Small molecules that promote self-renewal of stem cells and somatic cell reprogramming. Stem Cell Rev. Rep. 16, 511–523 (2020).
Graf, U. et al. The role of the leukemia inhibitory factor (LIF) - pathway in derivation and maintenance of murine pluripotent stem cells. Genes 2, 280–297 (2011).
Wang, P.-Y. et al. Pluripotency maintenance of amniotic fluid-derived stem cells cultured on biomaterials. J. Mater. Chem. B 3, 3858–3869 (2015).
Niwa, H. et al. A parallel circuit of LIF signalling pathways maintains pluripotency of mouse ES cells. Nature 460, 118–122 (2009).
Chi, E. Y. et al. Physical stability of proteins in aqueous solution: mechanism and driving forces in nonnative protein aggregation. Pharm. Res. 20, 1325–1336 (2003).
Feng, Y. et al. Antibodies@MOFs: an in vitro protective coating for preparation and storage of biopharmaceuticals. Adv. Mater. 31, e1805148 (2019).
Sarkar, A. & Hochedlinger, K. The Sox family of transcription factors: versatile regulators of stem and progenitor cell fate. Cell Stem Cell 12, 15–30 (2013).
Yang, L. et al. Melatonin restores the pluripotency of long-term-cultured embryonic stem cells through melatonin receptor-dependent m6A RNA regulation. J. Pineal Res. 69, e12669 (2020).
Ross, S. E. & Bogdanovic, O. TET enzymes, DNA demethylation and pluripotency. Biochem. Soc. Trans. 47, 875–885 (2019).
Lyu, Z. et al. Maintaining the pluripotency of mouse embryonic stem cells on gold nanoparticle layers with nanoscale but not microscale surface roughness. Nanoscale 6, 6959–6969 (2014).
Hu, Y. et al. Carbon-nanotube reinforcement of DNA-silica nanocomposites yields programmable and cell-instructive biocoatings. Nat. Commun. 10, 5522 (2019).
He, X. et al. MgFe-LDH nanoparticles: a promising leukemia inhibitory factor replacement for self-renewal and pluripotency maintenance in cultured mouse embryonic stem cells. Adv. Sci. 8, 2003535 (2021).
Percástegui, E. G. et al. Design and applications of water-soluble coordination cages. Chem. Rev. 120, 13480–13544 (2020).
Sepehrpour, H. et al. Biomedically relevant self-assembled metallacycles and metallacages. J. Am. Chem. Soc. 141, 14005–14020 (2019).
Liang, Y. et al. A stable biocompatible porous coordination cage promotes in vivo liver tumor inhibition. Nano Res 14, 3407–3415 (2021).
Liu, J. et al. Post-synthetic modifications of metal–organic cages. Nat. Rev. Chem. 6, 339–356 (2022).
Zhu, W. et al. Modular metal-organic polyhedra superassembly: from molecular-level design to targeted drug delivery. Adv. Mater. 31, 1806774 (2019).
Ahmad, N. et al. Metal–organic molecular cages: applications of biochemical implications. Chem. Soc. Rev. 44, 9–25 (2015).
Dey, N. & Haynes, C. J. E. Supramolecular coordination complexes as optical biosensors. ChemPlusChem 86, 418–433 (2021).
Fujita, D. et al. Protein stabilization and refolding in a gigantic self-assembled cage. Chem. 7, 2672–2683 (2021).
Qin, Y. et al. Copper metal-organic polyhedra nanorods with high intrinsic peroxidase-like activity at physiological pH for bio-sensing. J. Mater. Chem. B 5, 9365–9370 (2017).
Le Ouay, B. et al. Water-soluble ionic metal-organic polyhedra as a versatile platform for enzyme bio-immobilization. J. Am. Chem. Soc. 145, 11997–12006 (2023).
Yanes, O. et al. Metabolic oxidation regulates embryonic stem cell differentiation. Nat. Chem. Biol. 6, 411–417 (2010).
Li, S. et al. The age of vanadium-based nanozymes: Synthesis, catalytic mechanisms, regulation and biomedical applications. Chin. Chem. Lett. 35, 108793 (2024).
Hu, H. et al. Engineering vanadium carbide MXene as multienzyme mimetics for efficient in vivo ischemic stroke treatment. Chem. Eng. J. 440, 135180 (2022).
Fraqueza, G. & Aureliano, M. Polyoxovanadates contribution to pharmacological, antimicrobial and toxicological actions of vanadium. Med. Sci. Forum 11, 8 (2022).
Treviño, S. et al. Vanadium in biological action: chemical, pharmacological aspects, and metabolic implications in diabetes mellitus. Biol. Trace Elem. Res. 188, 68–98 (2018).
Prasad, K. S. & Ramachandrappa, S. U. Potential medicinal applications of vanadium and its coordination compounds in current research prospects: a review. Curr. Bioact. Compd. 16, 201–209 (2020).
Zhang, Y.-T. et al. Polyoxovanadate-based organic-inorganic hybrids: from {V5O9Cl} clusters to nanosized octahedral cages. Dalton Trans. 45, 3698–3701 (2016).
Guo, J. et al. Timing matters: pre-assembly versus post-assembly functionalization of a polyoxovanadate–organic cuboid. Chem. Sci. 13, 5718–5725 (2022).
Zhang, Z. et al. Type II porous ionic liquid based on metal-organic cages that enables l-tryptophan identification. Nat. Commun. 13, 2353 (2022).
Zheng, Y. et al. Self-assembly and antitumor activity of a polyoxovanadate-based coordination nanocage. Chem. Eur. J. 25, 15326–15332 (2019).
Yu, Z. H. & Zhang, Z. Y. Regulatory mechanisms and novel therapeutic targeting strategies for protein tyrosine phosphatases. Chem. Rev. 118, 1069–1091 (2018).
Wu, D. et al. A conserved mechanism for control of human and mouse embryonic stem cell pluripotency and differentiation by shp2 tyrosine phosphatase. PLoS One 4, e4914 (2009).
Liu, J. et al. Self-healing hyper-cross-linked metal–organic polyhedra (HCMOPs) membranes with antimicrobial activity and highly selective separation properties. J. Am. Chem. Soc. 141, 12064–12070 (2019).
Stefkova, K. et al. Alkaline phosphatase in stem cells. Stem Cells Int 2015, 628368 (2015).
Zhang, H. et al. Downregulated nuclear lncRNA NRON inhibits SHP2/Wnt/β-catenin signaling and cardiomyocyte differentiation during the development of Tetralogy of Fallot. Genes Dis. 10, 750–752 (2023).
Humphrey, R. K. et al. Maintenance of pluripotency in human embryonic stem cells is STAT3 independent. Stem Cells 22, 522–530 (2004).
Murry, C. E. & Keller, G. Differentiation of embryonic stem cells to clinically relevant populations: lessons from embryonic development. Cell 132, 661–680 (2008).
International Stem Cell, I. Assessment of established techniques to determine developmental and malignant potential of human pluripotent stem cells. Nat. Commun. 9, 1925 (2018).
Nelakanti, R. V. et al. Teratoma formation: a tool for monitoring pluripotency in stem cell research. Curr. Protoc. Stem Cell Biol. 32, 4A.8.1–4A.8.17 (2015).
Zhang, W. Y. et al. Teratoma formation: a tool for monitoring pluripotency in stem cell research. in StemBook. (Cambridge (MA), 2008).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2014).
Kolde, R. pheatmap. Pretty Heatmaps. R package version 1.0. 12. https://CRAN.R-project.org/package=pheatmap.2019.
Chen, S. et al. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. genet. 25, 25–29 (2000).
Draghici, S. et al. A systems biology approach for pathway level analysis. Genome res 17, 1537–1545 (2007).
Yaari, G. et al. Quantitative set analysis for gene expression: a method to quantify gene set differential expression including gene-gene correlations. Nucleic acids res. 41, e170–e170 (2013).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Gulati, G. S. et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science 367, 405–411 (2020).
Vento-Tormo, R. et al. Single-cell reconstruction of the early maternal-fetal interface in humans. Nature 563, 347–353 (2018).
Hof, P. et al. Crystal structure of the tyrosine phosphatase SHP-2. Cell 92, 441–450 (1998).
Yu, W.-M. et al. Identification of small molecular weight inhibitors of Src homology 2 domain-containing tyrosine phosphatase 2 (SHP-2) via in silico database screening combined with experimental assay. J. Med. Chem. 51, 7396–7404 (2008).
Anandakrishnan, R. et al. H++ 3.0: automating pK prediction and the preparation of biomolecular structures for atomistic molecular modeling and simulations. Nucleic Acids Res. 40, W537–W541 (2012).
Morris, G. M. et al. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 30, 2785–2791 (2009).
Acknowledgements
X.X. acknowledges the financial support from the National Natural Science Foundation of China (82241058). Y.C. acknowledges the financial support from Haihe Laboratory of Synthetic Biology (22HHSWSS00008). X.X. acknowledges the financial support from the Natural Science Foundation of Tianjin Province of China (grant nos. 21JCZDJC00290) and the Key Projects of Tianjin Science and Technology (24ZXZSSS00150).
Author information
Authors and Affiliations
Contributions
Y.C. and X.X. conceived and designed the study, performed analysis and wrote the paper. R.W. and Y.Q. designed and performed experiments, performed analysis. J.L. synthesized and characterized the materials. R.W., Y.Q., J.L., Y.C. and X.X. wrote the paper. Y.L. provided technical assistance with theoretical calculations. H.A., H.W. and J.S. synthesized and characterized the materials. W.H., S.W. and L.Y. analyzed the data and participated in the discussion.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, R., Qi, Y., Liu, J. et al. Metal-organic polyhedra maintain the self-renewal of embryonic stem cells. Nat Commun 16, 8684 (2025). https://doi.org/10.1038/s41467-025-63811-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63811-6