Abstract
Sulfated fucan is one of the most recalcitrant polysaccharides. The molecular mechanism underlying the endo-1,3-fucanase, which plays a critical role in the breakdown of sulfated fucan, remains unexplained. Here, we conduct a comprehensive structure-function relationship investigation on the endo-1,3-fucanases within a family space—GH168. The family can be divided into four subfamilies according to phylogenetic relationship and functional similarities. Subfamily I, Ⅱ and Ⅳ preferentially recognize Fucp2(OSO3−), Fucp2,4(OSO3−) and Fucp units at the +1 subsite, respectively, while consistently recognizing the Fucp2(OSO3−) unit at the −1 subsite. Remarkably, two-thirds of the interacting residues are dedicated to the recognition of sulfate groups along the glycoside chains. This mechanism is distinct from the direct recognition of the sugar backbone employed by neutral polysaccharide hydrolases. These findings unveil a critical enzyme recognition mechanism for sulfate polysaccharides and promote the application of endo-1,3-fucanases in the structural analysis and oligosaccharide production of sulfated fucan.
Similar content being viewed by others
Introduction
Sulfated polysaccharides constitute one of the largest and most varied reservoirs of organic carbon in the ocean. These compounds are decomposed by microorganisms, subsequently releasing carbon back into the marine environment, thereby maintaining the stability of the global marine carbon cycle1,2. Sulfated fucan, also known as fucoidan, is a sulfated polysaccharide predominantly composed of l-fucose and sulfate groups, which is found in brown algae and echinoderm and constitutes one of the important forms of marine organic carbon3. Sulfated fucan is one of the most structurally variable biomolecules in nature, which is recalcitrant and more slowly degraded than other marine polysaccharides such as arabinogalactan, β−1,4-mannan, α−1,4-galacturonan and β−1,3-glucan4. A recent study indicated that hundreds of enzymes are involved in the metabolism of sulfated fucan by ‘Lentimonas’ sp. CC4, belonging to the Verrucomicrobia5. The sulfate group is the most abundant substituent in sulfated fucan, often accounting for over 20% of its composition6. It is recognized as a primary factor contributing to the recalcitrance of sulfated fucan against enzymatic degradation5. Meanwhile, the sulfate group also enhances the physiological activity of sulfated fucan, thereby endowing it with considerable commercial value7. Japan, the United States, and South Korea have developed a variety of functional foods based on sulfated fucan, leveraging its diverse biological activities, such as antitumor, immunomodulatory, and antioxidative properties.
Endo-acting glycoside hydrolases are phylogenetically widely distributed enzymes that are critical for the cleavage of glycosidic bonds8. In the process of breakdown of exogenous polysaccharides, endo-acting glycoside hydrolases are often first released into the extracellular environment to initially hydrolyze the polysaccharide into oligosaccharides. Subsequently, a series of transporters transport these oligosaccharides into the cytoplasm, thereby activating the expression of genes related to the utilization of polysaccharides9. Endo-1,3-fucanase is an enzyme that hydrolyzes α−1,3-fucosidic bonds in an endo-acting manner, playing a crucial role in the breakdown of sulfated fucan. Endo-1,3-fucanase has been applied in the structural analysis and functional investigation of sulfated fucan10,11. The enzyme could also serve as a favorable tool for the preparation of low molecular weight sulfated fucans and oligosaccharides, which exhibit enhanced biological activities compared to their naturally occurring counterparts. The glycoside hydrolases are categorized into “families” based on the sequence homology in the CAZy database12. Currently, a total of 10 characterized endo-1,3-fucanases are present in the CAZy database, distributed across the GH107, GH168, GH174, and GH187 families. Among them, the majority originated from marine Flavobacteriia and Planctomycetota, and remained stable below the temperature of 30 °C.
GH168 is the first family confirmed to contain endo-1,3-fucanase, which was established by our research group13. A recent study confirmed the pivotal role of the GH168 family in the breakdown of sulfated fucan by marine Planctomycetota14. Nevertheless, to date, only three members within the family were characterized15,16,17. The previous study showed that enzymes from this family are sensitive to the sulfate group along the fucan chain. Nevertheless, the diversity of action modes and the underlying molecular mechanisms of the family remain unknown, making endo-1,3-fucanase one of the poorest characterized glycoside hydrolases. A systematic analysis of action modes and molecular mechanisms across the sequence space that remain largely unexplored will facilitate a molecular-level understanding of the core steps involved in the breakdown of sulfated fucans. Concurrently, it is critical for facilitating the biotechnological application of these enzymes.
In this work, we comprehensively characterized 16 endo-1,3-fucanases from the GH168 family and revealed the structure-function relationships of this family by utilizing a combination of bioinformatics, molecular biology, enzymology, mass spectrometry, structural biology, and a variety of structurally well-defined substrates. Notably, the results showed that the GH168 family employs an unusual substrate recognition mechanism, which is different from other glycoside hydrolases acting on neutral polysaccharides. This study is conducive to promoting the biotechnological application of the fucanase, and offers insights into the molecular mechanisms of enzymes that act upon sulfated polysaccharides.
Results
GH168 and other fucan-active enzyme families may originate from a common ancestor
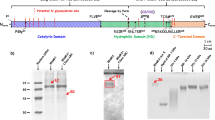
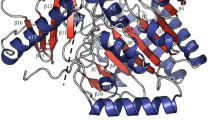
To investigate the overall fold of endo-1,3-fucanase from the GH168 family, the crystal structures of Pb41GH168_Ⅰ, WfGH168_Ⅱ, and WfGH168_Ⅳ were resolved at high resolution (Supplementary Table 1). These enzymes overall adopted a (β/α)8 triosephosphate isomerase (TIM) barrel fold, consisting of eight α-helices and eight parallel β-strands that alternate along the backbone (Fig. 1a). The α5 and α8 were generally composed of only three amino acids and significantly shorten than other α-helices (Supplementary Fig. 1). The substrate-binding interface was a crevice located at the center of the barrel, which had a positive potential that facilitated the binding of the enzymes to negatively charged sulfated fucan (Supplementary Fig. 2). The strictly conserved residues D172 and E243 (number of WfGH168_Ⅱ) located on β4 and β6 strands, respectively, correspond to nucleophilic attack and acid/base catalytic functions. The hydrolysis activities of mutants D172A and E243A were completely lost, consistent with the previous study13.
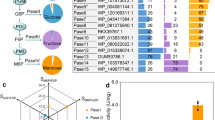
a Overall fold of WfGH168_II. The numbers of the secondary structure elements are indicated. b Hydrophobic interface between the β-sheet subdomain and the catalytic domain, featuring khaki sticks to denote the residues mediating domain association. c Superposition of representative structures from the four fucan-active enzyme family. The similar residues at the −1 subsite are shown as sticks. All residues are conserved, except for the W136 residue within the GH168 family, which may occasionally be substituted by tyrosine, which also possesses an aromatic ring. d Phylogenetic analysis of the GH168 family. e Clustering results of degradation degree of sulfated fucans by the GH168 enzymes.
A subdomain consisting of three β-strands attached to the C-terminus was tightly bound to the α7 of the TIM barrel through hydrophobic interfaces (Fig. 1b) and was present in all resolved crystal structures. The subdomain was distant from the active site and did not directly participate in catalysis. The subdomain also did not exhibit carbohydrate-binding ability (Supplementary Fig. 3); thus, it is not a carbohydrate-binding module. Using WfGH168_Ⅱ as an example, the truncated enzyme without the subdomain was expressed and its activity was assayed by the p-hydroxybenzoic acid hydrazide (pHBH) method. The results showed that the residual activity of the truncated WfGH168_Ⅱ (S1-Y363) was 5.0 ± 1.8%, indicating that the subdomain was critical for the function. Besides, the possibility could not be ruled out that the decrease in activity is attributed to the protein misfolding caused by the loss of structural integrity.
In addition to the GH168 family, the GH29, GH107, and GH151 families are also key enzymes in the breakdown of sulfated fucan that have been extensively studied. All these families adopt the (β/α)8 TIM barrel fold and retaining catalytic mechanism18,19,20, which piqued our interest in exploring the relationships among these families. The structures in these families were superposed and the conserved residues near the active center were compared. Intriguingly, the conserved residues near the −1 subsite exhibited similarity (Fig. 1c). Specifically, the spatial arrangements of the catalytic residues D172 and E243 (number of WfGH168_Ⅱ), as well as the conserved residue Y99, in these families are similar. N218 in the GH168 family also exhibits significant overlap with those in the GH107 and GH151 families. Furthermore, N101 and W332 in the GH168 family also existed in the GH107 family. It is suggested that the four families likely evolved from a common ancestor.
GH168 is divided into four functionally equivalent subfamilies
The homologous sequences of the GH168 family were searched and the phylogenetic tree was constructed. The sequences were divided into four subfamilies according to the phylogenetic relationship, named subfamily I to subfamily IV (Fig. 1d and Supplementary Fig. 4). A total of 50 sequences with dispersed evolutionary positions from these subfamilies were selected for heterologous expression, of which 41 could be expressed in a soluble form (Supplementary Table 2). To explore the diversity of the action mode of the family, five well-characterized α−1.3-linked sulfated fucans from Isostichopus badionotus (Ib-FUC)21, Thelenota ananas (Ta-FUC)22, Acaudina molpadioides (Am-FUC)23, Holothuria tubulosa (Ht-FUC)24 and Stichopus horrens (Sh-FUC)25 were collected as substrates. The distribution of sulfate groups of these substrates showed clear differences (Supplementary Fig. 5). Enzymatic activity assay with these substrates identified 13 previously uncharacterized fucanases: WpGH168_Ⅰ, KbGH168_Ⅰ, VbGH168_Ⅰ, PbGH168_Ⅰ, and PkGH168_Ⅰ from subfamily Ⅰ. PgGH168_II and RiGH168_II from subfamily II. RsGH168_III, Pb41GH168_III, Pb100GH168_III, and VbGH168_III from Subfamily III. Pb41GH168_Ⅳ and WfGH168_Ⅳ from subfamily Ⅳ. All of the enzymes showed a single band in the SDS-PAGE (Supplementary Fig. 6) and exhibited low identities with the previously reported enzymes (Supplementary Table 3). During the initial phase of hydrolysis, these enzymes all resulted in a rapid decrease in the molecular weight of sulfated fucan (Supplementary Fig. 7), and oligosaccharides with different structures were produced (Supplementary Table 4 and Supplementary Data 2), which indicated that they could hydrolyze the α−1,3 glycosidic bonds of sulfated fucan in an endo-acting manner.
The degradation degree of all the GH168 enzymes (including the 13 enzymes discovered in this study and the 3 previously characterized enzymes) on the tested substrates was quantified by measuring the increase in reducing sugars in the hydrolysis products. Cluster analysis showed that enzymes within the same subfamily exhibit similar substrate preferences, while showing differences in substrate preferences between subfamilies (Fig. 1e). It suggested that the phylogenetic analysis successfully grouped enzymes with similar functions together. Moreover, it implied that enzymes from different subfamilies exhibit exquisite differences in the recognition of sulfate group distribution. Subfamilies Ⅰ, Ⅱ and Ⅳ showed significant hydrolysis activities for at least one substrate, while subfamily III only demonstrated weak degradation towards Ib-FUC.
Subsequently, the biochemical properties of the enzymes from subfamilies Ⅰ, Ⅱ and Ⅳ were determined. The optimal temperature was generally 35 °C or 40 °C (Supplementary Fig. 8), and the optimal pH was 6–9 (Supplementary Fig. 9). Furthermore, the members generally exhibited excellent storage stability, retaining over 50% of the activity after being stored at 30 °C for 24 h (Supplementary Fig. 10), and could remain stable within a pH range of 5–10 (Supplementary Fig. 11). Therefore, these enzymes exhibited favorable stability (Supplementary Table 5), which might contribute to their applications in the field of biotechnology. For the majority of the enzymes in this study, alterations in buffer salts did not elicit significant changes in their activity levels. An exception is observed with KbGH168_ I, wherein the enzyme exhibits an activity of 6.7 U/mg when employing a citrate-phosphate buffer solution at a pH of 7.0. In contrast, upon the utilization of PBS buffer with a pH of 7.0, the activity increases to 10.4 U/mg.
The −1 subsite of this family strictly recognizes Fucp2(OSO3 −)
The hydrolysis products of Ib-FUC by all enzymes were analyzed by mass spectrometry to identify the common characteristics of subsite specificity of the whole family, since all of the enzymes generally exhibit hydrolysis activity towards Ib-FUC (Fig. 1e). A variety of fucooligosaccharides were detected in the hydrolytic products of enzymes from subfamilies I, II, and IV (Supplementary Table 4). Notably, a dominant component was noticed to account for over 50% against all oligosaccharides. According to its m/z 460.0269 ([M-2H]2−), the component was confirmed as fucotetraose with four sulfate groups, which is consistent with the structure of Ib-FUC.
The structures of the fucotetraoses were identified according to the further MS/MS analysis and structural model of Ib-FUC. The results showed that the structures of the fucotetraose from subfamilies I and II were F2S-F0S-F2,4S-F2S (F2S represents Fucp2(OSO3−), F0S represents Fucp and F2,4S represents Fucp2,4(OSO3−)) (Fig. 2a, b), while those from subfamily IV were F0S-F2,4S-F2S-F2S (Fig. 2c and Supplementary Fig. 12). Both the fucotetraoses exhibited an F2S unit at their reducing end, indicating that these enzymes recognized F2S at the −1 subsite. A representative member from each subfamily (KbGH168_Ⅰ, RiGH168_Ⅱ, Pb41GH168_Ⅳ) was selected, and the fucotetraose in their hydrolysis products was separated and purified, followed by the nuclear magnetic resonance (NMR) experiments to solve the structure. The NMR results were consistent with the MS/MS results, which further supported the conclusion (Supplementary Fig. 13−15 and Supplementary Table 6–8).
a–c show the MS/MS spectra of fucotetraose in the hydrolysis products of Ib-FUC by representative enzymes of subfamilies Ⅰ, Ⅱ and Ⅳ, respectively (KbGH168_Ⅰ, RiGH168_Ⅱ, Pb41GH168_Ⅳ). The charge number of fragments was provided by the Agilent Masshunter Qualitative Analysis Navigator B.08.00 software. d The conserved polar knuckle of WfGH168_Ⅱ. e The key amino acid residues that interact with the ligand at the −1 subsite. The blue-white sticks and the deep blue sticks represent the amino acid residues and the ligand, respectively. f Residual activities of mutants of key residues at the −1 subsite. Three biological replicates are conducted in this experiment and data points show individual measurements. Data are presented as mean values ± SD. The one-tailed T-test was performed to determine statistical significance between the wild-type (WT) and the variants. ****P value ≤ 0.0001.
To elucidate the structural basis of the recognition for F2S at the −1 subsite, the complex structures of WfGH168_Ⅰ, WfGH168_Ⅱ, and Pb41GH168_Ⅳ with their respective fucotetraose products from Ib-FUC were solved (Supplementary Table 1). A crucial residue, K66 (number of WfGH168_Ⅱ), located near the glycosidic bond linking subsite −1 to −2 of the ligand, is referred to as the “polar knuckle” (Fig. 2d). It interacted with the 4-hydroxyl group and the 3-glycosidic bond of the F2S at −1 subsite through hydrogen bonding. Additionally, the residue formed hydrogen bonds with the 2-O-sulfate group at the −2 subsite. The residue was strictly conserved and thus was one of the most distinctive characteristics of the GH168 family (Supplementary Fig. 16). The mutant K66A of WfGH168_Ⅱ was generated, and subsequent activity assessments revealed a complete loss of catalytic function (Fig. 2f). It demonstrated the indispensable nature of the polar knuckle within the GH168 family.
In all of the complex structures, F2S protruded into a groove that is equipped with a rich array of conserved polar residues at the −1 subsite (Fig. 2e). This groove, which was critical for substrate recognition, left space for the bulky sulfate group. In addition to the polar knuckle, the conserved residues N101 and Y99 formed hydrogen bonds with the 2-O-sulfate group at the subsite. The conserved E65 anchored the 4-hydroxyl group through hydrogen bonding and this residue also anchored the 2-O-sulfate group at the −2 subsite. Additionally, the conserved N218 formed a bidentate hydrogen bond with the 1-hydroxyl group and the 6-oxygen atom. All mutations led to the reduction or even abolition of the enzymatic activity, indicating that these residues play a key role in the substrate binding (Fig. 2f). The recognition at the +1 subsite is also dominated by polar residues, however, most of them are not conserved within the family, suggesting that GH168 members seem to be more tolerant for diverse substrate modifications in the positive subsite region.
Subfamily I preferentially recognizes F2S at the +1 subsite
Members of subfamily I, including WfGH168_Ⅰ, WpGH168_Ⅰ, KbGH168_Ⅰ, VbGH168_Ⅰ, PbGH168_Ⅰ, and PkGH168_Ⅰ, showed hydrolysis activity only towards Ib-FUC, and specifically cleaved the glycosidic bond between two F2S units (Fig. 3a). Since neither Ta-FUC nor Am-FUC contains a F2S-F2S unit, they could not be degraded by members of this subfamily. Although Ht-FUC and Sh-FUC contain F2S-F2S units, they could not be degraded either. It was likely that this subfamily could only accommodate F0S at the +2 subsite; however, the sugar residues in Ht-FUC and Sh-FUC corresponding to the +2 subsite were F2,4S and F2S, respectively. The sulfate groups may cause steric hindrance, preventing effective binding between the substrate and the enzyme. Additionally, the distribution pattern of sulfate groups may also affect the conformation of the sulfated fucan, thereby influencing the interaction between the substrate and the enzyme.
a Cleavage site of subfamily Ⅰ towards different sulfated fucan. The incomplete circles represent the cleavage sites of the enzymes, with the solid circles indicating the preferred cleavage sites, while the dashed circles denote sites where the enzyme can only cleave weakly. b The conformation of the ligand associated with WfGH168_Ⅰ. The number of subsites are labeled. c Critical residues within the negative subsite region of WfGH168_Ⅰ. The conserved key residues of the family and the characteristic residues of subfamily Ⅰ are represented by white sticks and yellow sticks, respectively. d Key residues within the positive subsite region of WfGH168_Ⅰ. Subsites +2 and +3 of WfGH168_Ⅰ interact with the sulfate group via K114, R205, and Q227; however, these residues exhibit high variability and are not a conserved feature of this subfamily. e The +1 subsite of WfGH168_Ⅰ accommodates an open space for the 4-O-sulfate group. f The enzymatic product profile of Ht-FUC by PkGH168_Ⅰ. g MS/MS spectrum of the dominant oligosaccharide in the product of Ht-FUC by PkGH168_Ⅰ.
To elucidate the substrate recognition mechanism of the subfamily Ⅰ, the complex structure of WfGH168_Ⅰ with the fucotetraose product of Ib-FUC was solved at a resolution of 2.04 Å (Supplementary Table 1). Two tetrasaccharides were located at subsites −4 to −1 and subsites +1 to +4 of the enzyme, respectively (Fig. 3b). The interaction between the enzyme and the substrate predominantly occurs at subsites from −2 to +2, which largely dictates the specificity of the enzyme. In contrast, the interactions at subsites −3, −4, +3, and +4 were rather limited. As expected, the polar knuckle K49 and other conserved residues mentioned above were observed at the −1 subsite of WfGH168_Ⅰ (Fig. 3c). Besides, the residue W262 formed an additional hydrogen bond with the 4-O-sulfate group at the −2 subsite, and the residue was only conserved within the subfamily (Supplementary Fig. 17a). Compared to the wild-type WfGH168_Ⅰ, the W262A mutant retained 77.0 ± 4.6% of the activity, which suggested that W262 was not essential for the substrate binding. Unlike the 2-O-sulfate group at the −2 subsite, which pointed towards the wall of the crevice, the 4-O-sulfate group at this subsite pointed outward from the crevice, demonstrating that the group was dispensable for the substrate recognition of the enzyme.
At the +1 subsite of WfGH168_Ⅰ, the 2-O-sulfate group formed a hydrogen bond interaction with Q155, which is only conserved within this subfamily (Fig. 3d). This sulfate group was also anchored by electrostatic interactions to R200 on the opposite side, a residue that was strictly conserved in both subfamilies I and II. The mutants Q155A and R200A exhibited 6.7 ± 2.1% and 7.6 ± 1.4% of activity, indicating that the residues were important for the substrate binding. Additionally, H222, which was only conserved within subfamily I, formed a hydrogen bond with the 4-hydroxyl group.
Although the structure analysis of the hydrolysis products indicated that subfamily Ⅰ could accommodate F2S at the +1 subsite, a space that was able to accommodate a 4-O-sulfate group was observed at this subsite (Fig. 3e). Structural prediction indicated that similar spaces were found in other members in this subfamily (Supplementary Fig. 18). Therefore, we suspected that members of subfamily I could accommodate F2,4S at the +1 subsite. In the substrates used in this study, only Ht-FUC possesses the F2S-F2,4S unit; thus, the activity assay of PkGH168_Ⅰ towards Ht-FUC was conducted. Interestingly, at the substrate concentration of 2 mg/mL, the substrate underwent only a slight depolymerization without the production of oligosaccharides. However, upon increasing the substrate concentration to 20 mg/mL, significant oligosaccharides were produced by the extensive depolymerization of the substrate (Fig. 3f). The dominant oligosaccharide was subjected to MS/MS analysis (Fig. 3g), confirming its structure as F2,4S-F0S-F2S-F2S. It revealed that F2,4S was capable of accessing the +1 subsite, particularly under conditions of high substrate concentration.
Subfamily II preferentially recognizes F2,4S at the +1 subsite
Subfamily II exhibited a broader substrate spectrum compared to subfamily I. Three members, WfGH168_II, PgGH168_II, and RiGH168_II, could hydrolyze Ib-FUC, Ht-FUC and Sh-FUC (Fig. 4a). The dominant oligosaccharide in the hydrolysis products of Sh-FUC was the disaccharide and MS/MS analysis indicated that the structure of the disaccharide was F2S-F2S (Fig. 4b and Supplementary Fig. 19a). It suggested that these enzymes could cleave the glycosidic bond between two F2S units. The dominant oligosaccharide produced by the enzymatic hydrolysis of Ht-FUC was F2,4S-F0S-F2S-F2S (Fig. 4c and Supplementary Fig. 19b), indicating that the cleavage site was the glycosidic bond between F2S and F2,4S. The above conclusions were further confirmed by the NMR spectra of the dominant oligosaccharides produced by RiGH168_Ⅱ from Sh-FUC (Supplementary Fig. 20 and Supplementary Table 9) and Ht-FUC (Supplementary Fig. 21 and Supplementary Table 10). Ht-FUC contains both the disaccharide units F2S-F2S and F2S-F2,4S, yet only the F2S-F2,4S unit was cleaved, suggesting a preference for F2,4S at the +1 subsite of subfamily II.
a Cleavage site of subfamily Ⅱ towards different sulfated fucan. b, c MS/MS spectrum of the dominant oligosaccharide in the product of Sh-FUC and Ht-FUC by RiGH168_Ⅱ, respectively. d Critical residues within the negative subsite region of WfGH168_ Ⅱ. The conserved key residues of the family and the characteristic residues of subfamily Ⅱ are represented by white sticks and blue sticks, respectively. e Surface representation of the complex structure of WfGH168_Ⅱ with the fucotetraose from Ib-FUC. f Key residues within the positive subsite region of WfGH168_ Ⅱ. g Structural superposition of native WfGH168_Ⅱ and its complex. The blue and cyan cartoons represent the structure without substrate and the structure with substrate, respectively.
The complex structure of WfGH168_II with a fucotetraose from Ht-FUC was solved at a resolution of 1.55 Å (Supplementary Table 1). The fucotetraose is bound to subsites −4 to −1 and interactions occur at subsites −2 and −1. Besides conserved residues such as the polar knuckle, the enzyme formed additional electrostatic interactions with the 2-O-sulfate groups at -1 subsite through the characteristic residues W106 and K176 (Fig. 4d). A residue R43, which is only conserved in this subfamily, formed a salt bridge with the 2-O-sulfate group at the −2 subsite. The −2 subsite could bind either F2,4S in Ib-FUC or F2S in Ht-FUC/Sh-FUC (Fig. 4a). Therefore, it was suspected that the 4-O sulfate group at the -2 subsite was not essential for subfamily Ⅱ. To confirm the speculation, the complex structure of WfGH168_II with the fucotetraose product from Ib-FUC was further solved. It confirmed that the sulfate group pointed outward towards the cavity and did not interact with any amino acid residues (Fig. 4e).
To uncover the substrate recognition mechanism in the positive subsite region of subfamily II, attempts were made to co-crystallize inactivated mutants of WfGH168_II with substrates, but the efforts were unsuccessful. Alternatively, WfGH168_II was superposed to the complex structures of WfGH168_I with fucotetraose from Ib-FUC, as both subfamilies produced identical fucotetraose from Ib-FUC. Similar to subfamily I, two characteristic residues at the +1 subsite, K176 and R221 (corresponding to Q155 and R200 of WfGH168_I, respectively), anchored the 2-O-sulfate group on both sides via salt bridges (Fig. 4f). Notably, the 4-hydroxyl group of F2S at this subsite pointed towards an open area, with a characteristic residue Y241 of this subfamily located nearby (Supplementary Fig. 17b). The distance between the Y241 side chain and the 4-hydroxyl group is 6.6 Å, and no interaction was formed. However, if the hydroxyl group were replaced by a bulkier sulfate group, it would likely form a hydrogen bond with Y241, contributing to the preference for F2,4S. The residual activity of the mutant Y241A towards Ht-FUC containing F2S-F2,4S units was merely 4.2 ± 0.8%, confirming that the residue is a key determinant of the recognition preference of F2,4S at the +1 subsite of the subfamily.
Intriguingly, the root mean square deviation (r.m.s.d.) value between the native enzyme and complex of WfGH168_II was 1.27 Å. In contrast, the r.m.s.d. value between the native structure and complex structure of WfGH168_I was only 0.19 Å. It suggested that significant conformational changes occurred in WfGH168_II upon substrate binding. In the native enzyme, the positive subsite region was covered by a long loop consisting of 20 residues (I285-E304). Upon substrate binding, the loop shifted outwards from the substrate binding cavity, and the “hidden pocket” was fully exposed. The residue that experienced the greatest movement on the loop (G290) shifted by a distance of 27.7 Å (Fig. 4g). In addition, the complex structure of WfGH168_II with fucotriose was determined, revealing the same degree of displacement for the loop (Supplementary Fig. 22). Structural predictions indicated that the flexible loop also existed in other members of the subfamily, and thus was a conserved feature of this subfamily (Supplementary Fig. 23). Several negatively charged amino acid residues were present in the loop. It could be speculated that the negatively charged sulfate groups along sulfated fucan may repel the loop and push it out of the cavity, thereby exposing the pocket.
Subfamily Ⅳ strictly recognizes F0S at the +1 subsite
Three members of subfamily Ⅳ, namely Pb41GH168_Ⅳ, Fun168A, and WfGH168_Ⅳ, exhibit efficient degradation of Ib-FUC, Ta-FUC, and Am-FUC (Fig. 5a). The dominant oligosaccharide in the hydrolysis products from Ta-FUC was F0S-F0S-F2,4S-F2S (Fig. 5b and Supplementary Fig. 24a) and the dominant oligosaccharide from Am-FUC was F0S-F2,4S-F0S-F2S (Fig. 5c and Supplementary Fig. 24b). Consequently, subfamily Ⅳ specifically hydrolyzes the glycosidic bond between F2S and F0S. Ht-FUC and Sh-FUC, lacking the F2S-F0S unit, could not be hydrolyzed by these enzymes.
a Cleavage site of subfamily Ⅳ towards different sulfated fucan. b, c MS/MS spectrum of the dominant oligosaccharide in the product of Ta-FUC and Am-FUC by Pb41GH168_Ⅳ, respectively. d Critical residues within the negative subsite region of WfGH168_ Ⅱ. The conserved key residues of the family and the characteristic residues of subfamily Ⅳ are represented by white sticks and red sticks, respectively. e Molecular docking of Pb41GH168_Ⅰ and the ligand F0S-F2,4S-F2S-F2S. The critical residue Q172 is represented by the red stick.
The complex structure of Pb41GH168_IV with the fucotetraose product from Ib-FUC was resolved and the fucotetraose was bound to the subsites −4 to −1 (Supplementary Table 1). The characteristic residues of the subfamily W99 and R135 interacted with the 2-O-sulfate group at the -1 subsite through polar interactions (Fig. 5d). Furthermore, the structure of the inactive mutant D171A of Pb41GH168_IV13 in complex with the oligosaccharide F2S-F0S-F2,4S-F2S was determined at a resolution of 1.40 Å. According to the cleavage site of this subfamily on Ib-FUC, theoretically, the oligosaccharide binds to the subsites −1 to +3. Actually, it is bound to the subsites −4 to −1, suggesting that the occupation of the −2 subsite was critical for substrate binding. The 2-O-sulfate group at the −2 subsite was anchored by the polar knuckle and a critical residue E58. The −2 subsite was occupied by F2,4S, F2S, and F0S, respectively, during the hydrolysis of Ta-FUC, Ib-FUC and Am-FUC. The Km value of the enzyme for Ta-FUC (3.17 ± 0.61 μM) and Ib-FUC (1.05 ± 0.11 μM) was significantly lower than its Km value for Am-FUC (10.28 ± 0.84 μM), showing a lower affinity for Am-FUC. It indicated that the recognition of the 2-O-sulfate group at −2 subsite is conducive to the enzyme-substrate binding. It was noteworthy that the −2 subsite of all GH168 family members recognized F2S or F2,4S, suggesting that the critical recognition for the 2-O-sulfate group at this subsite may be a common feature of the GH168 family. The importance of the −2 subsite had also been observed in the GH16 family26 and this subsite determines whether the enzymes from the family hydrolyze porphyran or agarose. This phenomenon may extend to other CAZyme families that act upon sulfated polysaccharides, even though these enzymes are evolutionarily unrelated.
In the positive subsite region of Pb41GH168_IV, a characteristic residue Q172 (Supplementary Fig. 17c) protruded on the substrate binding interface and formed a “ridge” (sometimes replaced by a structurally similar arginine in this subfamily). Structural superposition of the Pb41GH168_IV and WfGH168_I complexes showed that Q172 created steric hindrance with the 2-O-sulfate group at the +1 subsite. Molecular docking indicated that Q172 established a hydrogen bond with a distance of 2.2 Å to the 2-hydroxyl group of F0S at this subsite (Fig. 5e). The residual activity of the mutant Q172A was 7.2 ± 0.9%, confirming that the residue is important for the subfamily.
GH168 members play a critical role in the breakdown of α-1,3 sulfated fucan
To estimate the distribution of GH168 enzymes in nature, the species origins of endo-fucanase families GH107, GH168, GH174, and GH187 were statistically analyzed. Fucanases were mainly present in four phyla, namely Bacteroidota, Verrucomicrobiota, Planctomycetota, and Proteobacteria. These phyla are well-recognized for their complex polysaccharide metabolic capabilities. Notably, the GH168 family exhibited a higher abundance compared to other families, with the highest proportions in Bacteroidota, Verrucomicrobiota, and Planctomycetota. Particularly in Planctomycetota, the proportion of the GH168 family reached 79.66% (Supplementary Fig. 25). It indicated that the GH168 enzymes extensively participate in the breakdown of sulfated fucan in nature.
Wenyingzhuangia funcanilytica CZ1127T is a specialized sulfated fucan-utilizing strain whose genome encodes a rich array of endo-fucanases, including those belonging to the GH107, GH168, and GH174 families. In addition to the GH168 family, both the GH107 and GH174 families had been confirmed to exhibit endo-1,3-fucanase activity13,27. To evaluate the role of GH168 family enzymes in the depolymerization of α-1,3-linked sulfated fucans, the strain was cultivated in two α-1,3-linked sulfated fucans, namely Ta-FUC and sulfated fucan from Ascophyllum Nodosum (An-FUC)28. The utilization rates of Ta-FUC and An-FUC by the bacterium reached 92.5 ± 2.3% and 90.4 ± 3.9%, respectively. The control group utilized glucose as the sole carbon source in the culture medium, with the bacterial cells at the mid-log phase being harvested for RNA-seq sequencing and subsequent differential expression analysis (Supplementary Fig. 26). The differential expression analysis revealed that six of the seven GH168 genes within the genome exhibited an upregulation, higher than the upregulation fold changes of fucanases from other families (Supplementary Fig. 27 and Supplementary Data 1), which implied that the members from the GH168 family played a key role in the metabolic processes of α-1,3-linked sulfated fucans.
Discussion
In the majority of glycoside hydrolase families acting on neutral polysaccharides, the enzyme-substrate binding is typically achieved through interactions between amino acid residues and the backbone of the polysaccharide. Conversely, the GH168 enzymes achieve recognition of sulfated fucan with different structures through interactions with sulfate groups in different spatial arrangements. The spatial arrangement of sulfate groups on the polysaccharide chain is jointly determined by their substitution positions and the linkage positions of glycosidic bonds, which are the most important structural informations of sulfated fucan. It should be noted that this mechanism also exists within some other glycoside hydrolase families that hydrolyze sulfated polysaccharides29, such as GH16 porphyranases26, GH16 kappa-carrageenases30, and GH82 iota-carrageenases31, despite the absence of evolutionary relationships among these families. Therefore, this mechanism represents a molecular adaptation of glycoside hydrolases to sulfated polysaccharides. It is well known that the CH-π stacking is a key interaction of enzymes on the polysaccharide backbone. It not only plays a role in the substrate recognition of hydrolases of neutral polysaccharides, but also in the hydrolases of acidic polysaccharides, including the GH16 and GH82 families26,30,31,32,33,34. Notably, no obvious CH-π stacking interactions could be observed in the GH168 family (even though the weak stacking interactions at special angles could not be ruled out completely). Conversely, over 2/3 of the interacting residues are dedicated to the recognition of sulfate group in the family. Take WfGH168_Ⅰ as an example, 8/13 interacting residues are involved in the binding of sulfate groups. Therefore, it is suggested that the recognition of sulfate groups dominates substrate binding by the GH168 family. Moreover, it implied that the role of sulfate groups in the substrate recognition of sulfated polysaccharide enzymes seems to be even more crucial than we previously thought. This understanding could guide the design of sulfated polysaccharide-active enzymes and is worthy of further exploration.
At the −1 subsite, the strict recognition for F2S is present throughout the GH168 family. It may contribute to efficient enzyme-substrate binding of the family. The 2-O-sulfate group is the most common substituent in sulfated fucan found in nature, existing in sulfated fucan from nearly all sources. Even some sulfated fucans are predominantly composed of F2S6. The GH168 members achieve specific recognition of the major structural unit of sulfated fucan through the deployment of a series of conserved residues, which likely aid the enzyme in combining structurally complex sulfated fucans with high efficiency. It provided novel ideas into the rational modification and the de novo design of enzymes acting on complex polysaccharides.
The fucanases with different evolutionary statuses in the GH168 family exhibits diverse specificities at the +1 subsite. Subfamilies I, II, and IV preferentially recognize F2S, F2,4S, and F0S at this subsite, respectively. It suggested that the family adapted to sulfated fucan with diverse sulfate groups distribution through divergent evolution. It enhances the degradation efficiency of sulfated fucan and augments the competitiveness of organisms within specific ecological niches. Additionally, the GH168 family, along with other fucanase families GH107, GH174, and GH187, may synergistically contribute to the breakdown of sulfated fucan in nature. This is supported by the fact that the GH174 and GH187 families employ a distinct topological structure dominated by β-sheets, differing from the (β/α)8 TIM barrel fold of the GH168 family35. The structural diversity of fucanases suggested functional diversity. The systematic characterization of other fucanase families would further elucidate this diversity and aid in understanding the mechanisms by which sulfated fucans participate in the carbon cycle. Simultaneously, of greater practical significance, it would greatly facilitate the application of these enzymes in the development of sulfated fucan products.
Subfamily III was phylogenetically close to subfamily II, yet it exhibited significant differences in substrate preference, with only weak hydrolysis activities. Structural prediction of VbGH168_III and superpositions with the complex of WfGH168_Ⅱ indicated that the catalytic pair, polar knuckle, and other key residues were also conserved within this subfamily. However, a residue unique to subfamily III, R122, may occupy part of the space at the −2 subsite (Supplementary Fig. 28). The residue likely prevented the substrate from being anchored in the correct conformation. Therefore, this subfamily may only accommodate unsubstituted fucose residues at the −2 subsite, while the specificities of other subsites are similar to subfamily II. The substrates tested in this study do not meet this characteristic, leading to their difficulty in being hydrolyzed. However, it should be noted that the long side chain of arginine can adopt various conformations and the obstruction of the substrate by R122 is speculative. Alternatively, it is also possible that alterations in distal residues located at a distance from the substrate pocket led to changes in the backbone of the enzyme, thereby affecting the catalysis of the enzymes.
The microbial breakdown of sulfated fucan has always been a topic of interest among researchers. The specialized sulfated fucan-utilization organisms ‘Lentimonas’ sp. CC4 is rich in exo-fucosidases5, while W. funcanilytica CZ1127T boasts a more abundant repertoire of endo-fucanases, indicating the diversity of microbial utilization mechanisms for sulfated fucan. As the discoverers of W. funcanilytica CZ1127T and the annotators of its genome, we elucidated the substrate recognition mechanism of two endo-1,3-fucanases from the strain (WfGH168_Ⅰ and WfGH168_Ⅱ) in the study and confirmed their up-expression under the induction of sulfated fucan. Meanwhile, the global community has taken notice of the strain, and research on endo-1,4-fucanase and exo-fucosidase from this strain has become increasingly abundant36,37,38,39. With its clear genetic background and several characterized sulfated fucan-degrading enzymes, W. funcanilytica CZ1127T is poised to become one of the model organisms for the study of the breakdown of sulfated fucan.
In conclusion, this study unveiled the structure-function relationship of a critical enzyme family involved in the breakdown of sulfated fucan. In this family, subfamily I, Ⅱ and Ⅳ preferentially recognize F2S, F2,4S and F0S at the +1 subsite, respectively, and consistently recognize the F2S unit at the −1 subsite. These enzymes employed a sulfate group-mediated substrate recognition mechanism, which may represent a molecular adaptation of glycoside hydrolases to sulfated polysaccharides. The study lays an important theoretical foundation for the applications of endo-1,3-fucanases in the research and development of sulfated fucan.
METHODS
Preparation of sulfated fucan
Ib-FUC, Ta-FUC, Am-FUC, Ht-FUC and Sh-FUC were extracted following a previously described protocol40. For the extraction of An-FUC, the milled seaweed was suspended in water and heated at 60 °C for 24 h. 20% ethanol was added to the supernatant to precipitate and remove alginate. Crude polysaccharides were then precipitated by the addition of 75% ethanol to the hydrolysate supernatant. Purification was achieved using a HiPrep 26/60 Sephacryl S-500 HR gel filtration column, eluted with 0.2 M NaCl and 50 mM NaH2PO4-Na2HPO4 buffer at 1.3 mL/min. Anionic polysaccharide content in the eluted fractions was assessed with the methylene blue assay. The resulting sulfated fucans were subsequently dialyzed, lyophilized, and characterized by monosaccharide analysis.
Bioinformatics analysis
The sequence homologs were searched by the BLASTp tools in the NCBI non-redundant protein sequence database, with a cut-off expected threshold value of 1 × 10−10, thresholds being an identity > 30%, and a matrix of BLOSUM6241. Multiple sequence alignments were performed using ClustalW and visualized by the WebLogo3 server42,43. A phylogenetic tree was constructed in MEGA v12.0.2 using the neighbor-joining algorithm and iterated 1000 times to calculate the bootstrap value. The glycoside hydrolase domains were annotated by dbCAN344. The signal peptide was predicted by Signal v4.145. The structural prediction of enzymes was conducted by Alphafold346. The RMSD values between the four apo crystal structures of this family and the corresponding AlphaFold3 predicted structures ranged from 0.35 to 0.62 Å, which indicated that AlphaFold3 could accurately predict the structure of this family. To confirm the novelty of the β-sheet subdomain, three structures were randomly selected from each family (if available) that adopted the TIM barrel structure and manually inspected for the presence of similar subdomains.
Protein expression and purification
All of the genes and the mutants were synthesized by Sangon Biotech (Shanghai, China). The genes were inserted into the pET28a(+) and transformed into Escherichia coli BL21(DE3) cells. After incubation with 0.5 mM isopropyl β-D-1-thiogalactopyranoside at 17 °C overnight, cells were collected and disrupted by sonication in 20 mM NaH2PO4-Na2HPO4 buffer (pH 7.5). All of the crude enzymes were loaded onto the HisTrap HP column and the HiTrap Desalting column according to the previous parameters and subjected to the activity assays13.
To ensure the acquisition of highly pure enzymes for crystallization purposes, an extra purification step was implemented. This involved the application of a Hiload Superdex 75 pg column (GE Healthcare, Uppsala, Sweden). The column was eluted using a 10 mM Tris-HCl buffer solution adjusted to a pH of 7.5, supplemented with 0.1 M NaCl and 2 mM dithiothreitol. The concentration of the protein-enriched fractions was achieved through the 10 kDa ultrafiltration device. Enzyme purity was ascertained through SDS-PAGE, while protein concentrations were quantified by measuring the absorbance at a wavelength of 280 nm.
Carbohydrate array assay
The carbohydrate microarray analysis was conducted predominantly following McCartney’s methodology47. Solutions of sulfated fucan at a concentration of 2 mg/mL were applied to the nitrocellulose membranes. To prevent non-specific binding, the membranes were blocked using PBS buffer containing 5% skim milk powder (MPBS) for a period of 1 h. Thereafter, the membranes were incubated with a truncated WfGH168_II protein lacking the subdomain, followed by exposure to an HRP-conjugated (His)6-tag monoclonal antibody with dilution of 1:10000 (Proteintech, Illinois, USA) for 2 h. The membranes were then visualized using the Tanon 5200 imaging system (Shanghai, China).
Enzymatic activity and biochemical property determination
The activities of native enzymes and their mutants were assayed by the pHBH method48. Briefly, Ib-FUC was dissolved in pH 7.5 PBS buffer to 2 mg/mL. The solution containing 0.5 μg enzyme was mixed with 50 μL Ib-FUC and supplemented to a total volume of 100 μL with the same buffer. The solution was incubated at 35 °C for 10 min and then heat-inactivated. Subsequently, 250 μL of p-hydroxybenzoic acid hydrazide solution was added along with 650 μL of H2O. The reaction mixture was then incubated for 5 min at 100 °C, and the OD415 was measured. The standard curve of reducing sugar concentration and OD415 was established with L-fucose as the standard. 1 U of activity was defined as the amount of enzyme that released 1.0 μmol reducing sugar per minute.
To confirm the endo-acting mode of the enzyme, 1 mL Ib-FUC (1 mg/mL, pH 7.5) was mixed with 0.1 U enzyme and incubated at 35 °C for 0 min, 5 min and 30 min, respectively. The global profiles of the hydrolysis products were detected by HPSEC-RID using a TSKgel SuperAW4000 column (Tosoh Corporation, Kanagawa, Japan), which was equilibrated with 0.2 M NaCl. The data were collected in Agilent OpenLAB CDS ChemStation Edition C.01.07.
The biochemical properties were determined by the pHBH method described above. The effect of temperature on the activity was assayed by incubating the enzyme with substrate at 20–55 °C. The thermal stability was determined by measuring the residual activity of the enzyme after pre-treatment at 4 °C, 25 °C, 30 °C, 35 °C and 40 °C for 24 h. The optimal pH was determined by incubating the enzyme at pH 3.0-11.0. To determine pH stability, the enzyme was pretreated in buffers with different pH values (20 mM citrate-phosphate buffer for pH 3.0–7.0, 20 mM PBS buffer for pH 7.0–9.0, and 20 mM Na2CO3-NaHCO3 buffer for pH 9.0–11.0) at 25 C for 1 h, and the residual activity was measured after replacing pH to 7.5. The kinetic characteristics were examined using substrates at concentrations of 0.05 to 5.00 mg/mL of sulfated fucan, mirroring the enzyme activity test conditions except for the differing substrate concentration13.
Mass spectrometry analysis of hydrolysis products
To prepare the final hydrolysis products, 1 mL sulfated fucan (1 mg/mL, pH 7.5) was mixed with 0.1 U enzyme and incubated at 35 °C for 24 h. The hydrolysis products were subjected to mass spectrometry analysis using an ultra-performance size exclusion chromatography coupled with high-resolution mass spectrometry (UPSEC-HRMS) system. The system integrated a Dionex Ultimate 3000 ultra-performance liquid chromatography system (Thermo Fisher Scientific, San Jose, CA, U.S.A.) with a Thermo Scientific Q-Exactive Orbitrap mass spectrometer or a quadrupole time-of-flight mass spectrometer (Thermo Fisher Scientific, San Jose, CA, U.S.A.). The hydrolysis products were introduced onto an Acquity UPSEC BEH 125 SEC column (4.6 × 150 mm, Waters, Milford, MA, U.S.A.) and separated using a mobile phase that comprised 20% (v/v) methanol with 10 mM ammonium acetate. The Q-Exactive Orbitrap mass spectrometer parameters were as follows: negative ionization, capillary temperature of 300 °C, spray voltage of 2000 V, sheath gas pressure of 40 psi, S-lens radio frequency (RF) level of 50 V, and mass range of m/z 200−2000. The quadrupole time-of-flight mass spectrometer parameters were set as follows: negative ionization mode, drying gas temperature 300 °C, drying gas flow 8 L/min, nebulizer 35 psi, sheath gas temperature 300 °C, sheath gas flow 11 L/min, capillary voltage 3500 V, nozzle voltage 1000 V, and fragmentor 380 V. The m/z range was 100–1700. Mass Spectrometry data were collected using Thermo Scientific Xcalibur v4.2.47. The identification of oligosaccharides within the samples was facilitated by a glycoinformatics pipeline. This involved the deconvolution of raw data using DeconTools49 and the construction of a hypothetical database through GlycReSoft v1.050, which offers potential compositions of oligosaccharides. The chemical compositions of the oligosaccharides were determined using GlycReSoft 1.0.
For subsequent MS/MS analysis, the hydrolysis products were reduced and converted to alditol according to a established procedure (Zhang et al., 2006). An aliquot of 100 μL from the hydrolysis products was lyophilized and reconstituted in 20 μL of NaBD4 reagent (0.05 M NaBD4 in 0.01 M NaOH), followed by incubation at 37 °C overnight. The reaction mixture was subsequently neutralized to pH 7.0 and passed over a mini cation exchange column (AG50 W-X8, H form, Bio-Rad, Hemel Hempstead, England). Boric acid was removed by repeated co-evaporation with methanol. The resulting alditols were analyzed by UPSEC-MS/MS. The UPSEC conditions were identical to those used for UPSEC-HRMS. The Q-Exactive Orbitrap mass spectrometer was operated in full-scan ddMS2 mode with an automatic gain control target of 3.0 × e6 and a maximum injection time of 100 ms. The MS/MS collision energy was set to 10 V using stepped normalized collision energy. All data were acquired and processed with the Xcalibur 2.0 software. All mass spectrometry analyses were subjected to three independent replicates.
Preparation of oligosaccharides
The hydrolysis product was subjected to fractionation using two HiLoad 26/600 Superdex 30 pg columns in series (GE healthcare, Uppsala, Sweden) and eluted at a flow rate of 1.3 mL/min with 5 mM ammonium formate. The purity of the oligosaccharides was determined by a Superdex peptide 10/300 GL column (GE healthcare, Uppsala, Sweden). The purified oligosaccharides were collected and subsequently subjected to NMR experiments or the preparation of complex crystals of enzymes.
NMR experiment of oligosaccharide
The oligosaccharides were subjected to lyophilization using deuterated water (D2O, 99.9% purity; Sigma, USA) in a process repeated three times. Following this, 3 mg of oligosaccharides were dissolved in 200 μL of D2O. The NMR experiments, which encompassed one-dimensional (1D) 1H NMR, two-dimensional correlation (COSY), total correlation (TOCSY), and nuclear Overhauser effect (NOESY) spectra, were acquired on a BRUKER AVANCE III 600 MHz NMR spectrometer, equipped with a Probe 5 mm CPTCI 1H-13C/15 N/D Z-GRD, and processed through TopSpin v3.1 software. The spectrometer operated at a temperature of 298.15 K. Standard Bruker pulse sequences were employed, including 1H-zgpr, COSY-cosygpprqf, TOCSY-mlevphpr.2, and NOESY-noesygpphpr. The spectral widths for 1D and 2D NMR were set at 20 ppm and 11 ppm, respectively. The number of scans and acquisition times were provided in Supplementary Table 11. Water suppression was implemented during the experiment. The NMR data were collected in TopSpin3.1. Data analysis was performed in MestReNova v9.0. The chemical shifts for the 1H NMR spectrum were referenced against the compound 4,4-dimethyl-4-silapentane-1-sulfonic acid (DSS) at a δ value of 0.00 ppm.
Protein crystallization, X-ray data collection, and structure determination
All of the enzymes were concentrated to 20–30 mg/mL and crystallized by the vapor diffusion method in both hanging or sitting drops at 17 °C. The native WfGH168_Ⅰ was crystallized under the condition of 0.1 M HEPES pH 7.5, 20% w/v Polyethylene glycol (PEG) 10,000. WfGH168_Ⅰ with fucotetraose from Ib-FUC was co-crystalized in 0.2 M magnesium chloride hexahydrate, 0.1 M HEPES sodium pH 7.5 and 25% w/v PEG 3,350. WfGH168_Ⅱ was crystallized in 0.2 M ammonium acetate, 0.1 M Bis-Tris (pH 6.5) and 25% w/v PEG 3,350. WfGH168_Ⅱ with fucotetraose from Ht-FUC was crystallized in 0.2 M sodium acetate trihydrate, 0.1 M Tris-HCl (pH 8.5) and 30% w/v PEG 4,000. WfGH168_Ⅱ with fucotetraose from Ib-FUC was crystallized in 24% PEG 1,500 and 20% glycerol. WfGH168_Ⅱ with fucotriose from Ib-FUC was crystallized in 10% v/v 2-Propanol, 1 M HEPES sodium (pH 7.5) and 20% w/v PEG 4,000. Pb41GH168_IV was crystallized in 0.2 M lithium sulfate monohydrate, 0.1 M Tris-HCl (pH 8.5) and 25% w/v PEG 3,350. Pb41GH168_IV with fucotetraose from Ib-FUC was crystallized in 0.03 M citric acid, 0.07 M Bis-Tris (pH 7.6) and 20% w/v PEG 3,350. Mutant D171A of Pb41GH168_IV with fucotetraose from Ib-FUC was crystallized in 30% w/v PEG 3,000 and 100 mM CHES (pH 9.5).
The data for diffraction analysis were acquired at the Shanghai Synchrotron Radiation Facility, specifically at beamline BL19U1, within a 100 Kelvin nitrogen environment. The integration of the data was executed utilizing the XDS software v5 202151 and scaled using the Aimless program52. The initial structural model was established through molecular replacement in CCP4 software v7.0.078, guided by a search model predicted by Alphafold253. The generated models were subjected to iterative rounds of structure refinement within the COOT v0.9.8.154 and further optimization using the PHENIX suite v1.21.2-541955. Visual representations of the structural data were generated by PyMOL software (https://www.pymol.org). The analysis of the surface electrostatic potential was conducted using the ABPS tools integrated within PyMOL v3.1.4.1. The r.m.s.d. values between the structures were computed using the US-Align program56. The 2Fo-Fc electron density map for each crystal structure and Fo-Fc omit map for each ligand was provided in Supplementary Figs. 29 and 30, respectively.
Molecular docking
The generation of the oligosaccharide ligand was facilitated through ChemDraw 16.0 and the eLBOW module within the PHENIX suite v1.21.2-541955. The ligand was integrated into the substrate binding crevice of the enzyme by employing the Autodock-Vina software57. To prepare the PDB molecules for docking, they were converted into PDBQT format using AutoDockTools 1.5.758. A total of 14 bonds are rotatable, including C1-O1, C2-O2, C3-O3, and C4-O4, as well as the bonds between the sulfur atom of the sulfate group and the oxygen atom of the fucosyl residues. A Monte-Carlo iterated search combined with the BFGS gradient-based optimizer was used as the search algorithm. The grid (60 × 60 × 60 ų) is centered around the catalytic residue of the protein, with a grid spacing of 0.375 Å. The exhaustiveness parameter was adjusted to a value of 8, and the highest-ranking binding conformation was retained for subsequent analysis within PyMOL. The docking process was independently replicated three times to ensure reproducibility.
mRNA-Seq expression profiling
To conduct transcriptome analysis, the strain W. fucanilytica CZ1127T was cultured in a growth medium59, which contained either 0.2% sulfated fucan or D-glucose (Sigma-Aldrich, St. Louis, MO, USA). The culture was incubated at 25 °C overnight until it achieved the mid-log phase. Total RNA was harvested with the RNAprep pure Cell/Bacteria Kit (TIANGEN, Beijing, China). RNA quality was confirmed using the Agilent 2100 Bioanalyzer (Agilent Technologies, USA). mRNA was purified using the Illumina Ribo-Zero rRNA Removal Kit (Bacteria) and subjected to fragmentation by the addition of a fragmentation buffer. di-tagged cDNAs were synthesized through a series of random priming reactions with terminal-tagging oligonucleotides, followed by purification using AMPure XP beads (Beckman Coulter, Beverly, MA, USA). Then, PCR was performed with Phusion High-Fidelity DNA polymerase, Universal PCR primers and Index (X) Primer, and a final mRNA sequencing library was constructed.
Raw sequencing data was conducted on the Illumina X-Ten platform at Novogene Co., Ltd. (Beijing, China) and converted to Fastq format by using the BCL2Fastq v1.8.4 (C + + and Perl). To obtain clean data, the following steps were employed: Reads containing adapters were removed; Reads with a proportion of undetermined base information greater than 10% were removed; Reads with a quality value of 5 or less accounting for more than 50% of the entire read were removed. At the same time, Q20, Q30 and GC content of the clean data were calculated. All the downstream analysis were based on the clean data with high quality. Clean data were obtained by removing reads containing adapters, reads containing ploy-N and low-quality reads from raw data. Read alignment against the reference genome (RefSeq NZ_CP014224.1) was achieved using Bowtie v2.0.6 (C + +). Gene expression levels were quantified using the HTseq-count function and union model of HTSeq v0.9.1 (Python) to count reads mapped to each gene, and FPKM values were calculated. Normalization was conducted by the TMM methods. Differential expression analysis of the experimental group and control group was performed using the EdgeR v3.24.3 (R language). The resulting P-values were adjusted using Benjamini and Hochberg’s approach for controlling the false discovery rate. A corrected P-value of 0.005 and log2 (Fold change) of 1 was set as the threshold for significantly differential expression.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Unless otherwise stated, all data supporting the results of this study can be found in the article, supplementary, and source data files. The crystal structures presented in this study are deposited in the Protein Data Bank under the following accession numbers: 9JOC, 9JOF, 9JOG, 9JOH, 9JOM, 9JOO, 9JP2, 9JP3 and 9JOS. The RNA-seq data are available in the National Center for Biotechnology Information (NCBI) database under accession code PRJNA1258967 and PRJNA1258907. Source data are provided with this paper.
References
Engel, A., Thoms, S., Riebesell, U., Rochelle-Newall, E. & Zondervan, I. Polysaccharide aggregation as a potential sink of marine dissolved organic carbon. Nature 428, 929–932 (2004).
Arnosti, C. et al. The Biogeochemistry of Marine Polysaccharides: Sources, Inventories, and Bacterial Drivers of the Carbohydrate Cycle (eds Carlson, C. A. & Giovannoni, S. J.) 81–108 (2021).
Zayed, A. & Ulber, R. Fucoidans: downstream processes and recent applications. Mar. Drugs 18, 170 (2020).
Vidal-Melgosa, S. et al. Diatom fucan polysaccharide precipitates carbon during algal blooms. Nat. Commun. 12, 1150 (2021).
Sichert, A. et al. Verrucomicrobia use hundreds of enzymes to digest the algal polysaccharide fucoidan. Nat. Microbiol 5, 1026–1039 (2020).
Zvyagintseva, T. N. et al. Structural diversity of fucoidans and their radioprotective effect. Carbohyd. Polym. 273, 118551 (2021).
Chen, G. et al. A comprehensive review of sulfated fucan from sea cucumber: antecedent and prospect. Carbohydr. Polym. 341, 122345 (2024).
Mccarter, J. D. & Withers, S. G. Mechanisms of enzymatic glycoside hydrolasis. Curr. Opin. Struc. Biol. 4, 885–892 (1994).
Cuskin, F. et al. Human gut Bacteroidetes can utilize yeast mannan through a selfish mechanism. Nature 517, 165–169 (2015).
Chen, G. et al. Utilizing heterologously overexpressed endo-1,3-fucanase to investigate the structure of sulfated fucan from sea cucumber (Holothuria hilla). Carbohyd. Polym. 272, 118480 (2021).
Zhu, Y. et al. Structure–function relationship analysis of fucoidan from sea cucumber (Holothuria tubulosa) on ameliorating metabolic inflammation. J. Food Biochem. 45, e13500 (2021).
Drula, E. et al. The carbohydrate-active enzyme database: functions and literature. Nucleic Acids Res 50, D571–D577 (2022).
Shen, J. J., Chang, Y. G., Zhang, Y. Y., Mei, X. W. & Xue, C. H. Discovery and characterization of an endo-1,3-fucanase from marine bacterium wenyingzhuangia fucanilytica: a novel glycoside hydrolase family. Front. Microbiol. 11, 1674 (2020).
Perez-Cruz, C. et al. Mechanisms of recalcitrant fucoidan breakdown in marine Planctomycetota. Nat. Commun. 15, 10906 (2024).
Shen, J. J. et al. Discovery of a catalytic domain defines a new glycoside hydrolase family containing endo-1,3-fucanase. Carbohyd. Polym. 323, 121442 (2024).
Shen, J. et al. Characterization of a novel endo-1,3-fucanase from Wenyingzhuangia fucanilytica within glycoside hydrolase family 168. Int. J. Biol. Macromol. 281, 136447 (2024).
Chen, G. et al. Structural investigation of Fun168A unraveling the recognition mechanism of endo-1,3-fucanase towards sulfated fucan. Int. J. Biol. Macromol. 271, 132622 (2024).
Vickers, C. et al. Endo-fucoidan hydrolases from glycoside hydrolase family 107 (GH107) display structural and mechanistic similarities to alpha-l-fucosidases from GH29. J. Biol. Chem. 293, 18296–18308 (2018).
Sela, D. A. et al. Bifidobacterium longum subsp infantis ATCC 15697 α-fucosidases are active on fucosylated human milk oligosaccharides. Appl. Environ. Microb. 78, 795–803 (2012).
Lezyk, M. et al. Novel α-L-Fucosidases from a soil metagenome for production of fucosylated human milk oligosaccharides. PLoS One. 11, e0147438 (2016).
Chen, S. et al. Sequence determination and anticoagulant and antithrombotic activities of a novel sulfated fucan isolated from the sea cucumber Isostichopus badionotus. Biochimica et. Biophysica Acta (BBA) - Gen. Subj. 1820, 989–1000 (2012).
Yu, L. et al. Structure elucidation of fucoidan composed of a novel tetrafucose repeating unit from sea cucumber Thelenota ananas. Food Chem. 146, 113–119 (2014).
Yu, L. et al. Structural study of fucoidan from sea cucumber Acaudina molpadioides: a fucoidan containing novel tetrafucose repeating unit. Food Chem. 142, 197–200 (2014).
Chang, Y. et al. Primary structure and chain conformation of fucoidan extracted from sea cucumber Holothuria tubulosa. Carbohyd. Polym. 136, 1091–1097 (2016).
Ustyuzhanina, N. E. et al. A highly regular fucan sulfate from the sea cucumber Stichopus horrens. Carbohyd. Res. 456, 5–9 (2018).
Hehemann, J. et al. Transfer of carbohydrate-active enzymes from marine bacteria to Japanese gut microbiota. Nature 464, 908–912 (2010).
Liu, G. et al. Characterization of an endo-1,3-fucanase from marine bacterium Wenyingzhuangia aestuarii: the first member of a novel glycoside hydrolase family GH174. Carbohyd. Polym. 306, 120591 (2023).
Marais, M. & Joseleau, J. A fucoidan fraction from Ascophyllum nodosum. Carbohyd. Res. 336, 155–159 (2001).
Hehemann, J. H., Boraston, A. B. & Czjzek, M. A sweet new wave: structures and mechanisms of enzymes that digest polysaccharides from marine algae. Curr. Opin. Struc. Biol. 28, 77–86 (2014).
Michel, G. et al. The kappa-carrageenase of P. carrageenovora features a tunnel-shaped active site: a novel insight in the evolution of Clan-B glycoside hydrolases. Structure 9, 513–525 (2001).
Michel, G., Helbert, W., Kahn, R., Dideberg, O. & Kloareg, B. The structural bases of the processive degradation of ι-carrageenan, a main cell wall polysaccharide of red algae. J. Mol. Biol. 334, 421–433 (2003).
Chen, F. Y. et al. Structural insights into the substrate recognition and catalytic mechanism of a GH16 βκ-Carrageenase from Wenyingzhuangia fucanilytica. J. Agr. Food Chem. 72, 20114–20121 (2024).
Luis, A. S. et al. Sulfated glycan recognition by carbohydrate sulfatases of the human gut microbiota. Nat. Chem. Biol. 18, 841–849 (2022).
Pluvinage, B. et al. Molecular basis of an agarose metabolic pathway acquired by a human intestinal symbiont. Nat. Commun. 9, 1043 (2018).
Chen, G. et al. The structure investigation of GH174 endo−1,3-fucanase revealed an unusual glycoside hydrolase fold. Int. J. Biol. Macromol. 280, 135715 (2024).
Zueva, A. O. et al. Expression and biochemical characterization of two recombinant fucoidanases from the marine bacterium Wenyingzhuangia fucanilytica CZ1127. T. Int. J. Biol. Macromol. 164, 3025–3037 (2020).
Zueva, A. O. et al. Production of high- and low-molecular weight fucoidan fragments with defined sulfation patterns and heightened in vitro anticancer activity against TNBC cells using novel endo-fucanases of the GH107 family. Carbohyd. Polym. 318, 121128 (2023).
Dong, S., Chang, Y., Shen, J., Xue, C. & Chen, F. Purification, expression and characterization of a novel α-l-fucosidase from a marine bacteria Wenyingzhuangia fucanilytica. Protein Expres Purif. 129, 9–17 (2017).
Shen, J. et al. Characterization of an alpha-L-fucosidase in marine bacterium Wenyingzhuangia fucanilytica: new evidence on the catalytic sites of GH95 family glycosidases. J. Sci. Food Agric 104, 8240–8247 (2024).
Ribeiro, A., Vieira, R. P., Mourão, P. A. S. & Mulloy, B. A. Sulfated α-l-fucan from sea cucumber. Carbohyd. Res. 255, 225–240 (1994).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25, 3389–3402 (1997).
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res 14, 1188–1190 (2004).
Thompson, J. D., Gibson, T. J. & Higgins, D. G. Multiple sequence alignment using ClustalW and ClustalX. Current protocols in bioinformatics. Chapter 2, 2–3 (2002).
Zheng, J. F. et al. dbCAN3: automated carbohydrate-active enzyme and substrate annotation. Nucleic Acids Res 51, W115–W121 (2023).
Teufel, F. et al. SignalP 6.0 predicts all five types of signal peptides using protein language models. Nat. Biotechnol. 40, 1023 (2022).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
McCartney, L., Gilbert, H. J., Bolam, D. N., Boraston, A. B. & Knox, J. P. Glycoside hydrolase carbohydrate-binding modules as molecular probes for the analysis of plant cell wall polymers. Anal. Biochem. 326, 49–54 (2004).
Lever, M. A new reaction for colorimetric determination of carbohydrates. Anal. Biochem. 47, 273–279 (1972).
Slysz, G. W. et al. The DeconTools framework: an application programming interface enabling flexibility in accurate mass and time tag workflows for proteomics and metabolomics. In Proc 58th ASMS Conf Mass Spectrom Allied Topics (2010).
Maxwell, E. et al. GlycReSoft: a software package for automated recognition of glycans from LC/MS Data. PLoS One. 7, e45474 (2012).
Kabsch, W. XDS. Acta Crystallogr. Sect. D: Biol. Crystallogr. 66, 125–132 (2010).
Potterton, E., McNicholas, S., Krissinel, E., Cowtan, K. & Noble, M. The CCP4 molecular-graphics project. Acta Crystallogr. Sect. D. -Struct. Biol. 58, 1955–1957 (2002).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583 (2021).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. Sect. D: Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. Sect. D. -Struct. Biol. 66, 213–221 (2010).
Zhang, C. X., Shine, M., Pyle, A. M. & Zhang, Y. US-align: universal structure alignments of proteins, nucleic acids, and macromolecular complexes. Nat. Methods 19, 1109 (2022).
Trott, O. & Olson, A. J. Software News and update Autodock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Morris, G. M. et al. AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J. Comput. Chem. 30, 2785–2791 (2009).
Ensor, L. A., Stosz, S. K. & Weiner, R. M. Expression of multiple complex polysaccharide-degrading enzyme systems by marine bacterium strain 2-40. J. Ind. Microbiol. Biot. 23, 123–126 (1999).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (U22A20542 (Y. C.), 324B2061 (G. C.)) and Fundamental Research Funds for the Central Universities (202461048 (G. C.)). We thank the Shanghai Synchrotron Radiation Facility of BL19U1 for the assistance on the X-ray diffraction experiments.
Author information
Authors and Affiliations
Contributions
G.C. performed the most experimental studies. J.S., X.L., and M.S. performed the NMR studies. S.D., Y.S. and A.K. carried out the data analysis. X.M., Y.Z., G.L. and F.C. performed the X-ray data collection. X.S. and K.L. performed the protein expression and purification, and molecular docking. Y.F. and C.X. supervised the work. Y.C. conceived, designed and supervised the experiments. All of the authors revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Stephanann Costello, Mirjam Czjzek, Arun Goyal, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, G., Shen, J., Li, X. et al. Structure-function relationship of the GH168 fucanase reveals an unusual enzyme recognition mechanism for sulfated polysaccharide. Nat Commun 16, 9216 (2025). https://doi.org/10.1038/s41467-025-63891-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63891-4