Abstract
Epidermal stem cells interact with the extracellular matrix (ECM) to regulate their differentiation and maintain skin architecture. Here, we demonstrate a role for basal epidermal stem cells (BECs)-ECM interaction in regulating adhesion molecules expressed by the periderm—the superficial epidermal cells (SECs) of the embryonic bilayered skin. Using the developing zebrafish fin fold, we identify BECs form distinct regions of collagen- versus laminin- enriched basement membranes through integrin-mediated adhesions. Mechanistically, collagen-associated BECs form desmosomes and adherens junctions (AJs) with SECs while laminin-associated BECs display reduced desmosomes but sustain AJs and actomyosin expression with SECs. Notably, we show both in vivo and in a bilayered human keratinocyte model, that laminin, compared to collagen, is sufficient to repress desmosome formation while sustaining AJs specifically at the interlayer cell contacts. In vivo, laminin deficiency enhances desmosome expression across layers and impairs the wound-healing capacity of SECs. This defect was partially rescued by genetic reduction of the desmosome protein Desmoplakin-1a, highlighting the role of ECM-dependent junctional specialization in mediating differences in SEC injury response. Overall, our findings identify that stem cells, through their matrix, establish specialized junctions in the overlying stratified epithelium, which contribute to skin healing properties.
Similar content being viewed by others
Introduction
Stem cells are essential for tissue development, maintenance, and repair due to their capacity for self-renewal and differentiation1,2,3. Their behavior is regulated by cues from the surrounding microenvironment, or niche, which includes neighboring cells and the extracellular matrix (ECM). They interact with specific ECM components via integrins4,5, and form junctions with neighboring cells. For example, adherens junctions (AJs) facilitate intercellular adhesion by connecting the actin cytoskeletons of adjacent cells and modulate tissue tension through actomyosin-driven mechanotransduction6. Additionally, desmosomes provide tissue resistance to tearing or separation by firmly linking adjacent cells through intermediate filaments7. These junctions work synergistically, and altering any of these interactions can affect stem cell proliferation and differentiation8,9,10,11. While the roles of these interactions in stem cell differentiation are well established, it remains unclear whether they also allow stem cells to influence the behavior of tissues independent of fate specification. Exploring this aspect could uncover a critical, underappreciated structural role of stem cells in shaping tissue responses.
Basal epidermal stem cells (BECs) originate from the ectoderm during embryonic development and expand over an ECM that they produce, covering the embryo’s surface as early as four weeks post-gestation in humans12. BECs differentiate to form a stratified epidermis by ~eight weeks of gestation. However, there is a critical period during human embryonic development, before week eight13, when BECs do not undergo differentiation or asymmetric division. Instead, they expand and interact with the periderm, a temporary layer consisting of superficial epidermal cells (SECs) that protects the embryo from the amniotic environment14. Because BECs are undifferentiated, adhesive, and form stable contacts within a bilayered epidermis, they present a unique model for studying how basal cells contribute to tissue architecture and interlayer organization prior to stratification.
Analyzing and manipulating BECs within their intact environment in utero in early mammalian embryos is technically challenging, leaving these functions largely unexplored. Other vertebrate embryos, such as zebrafish, also possess a conserved bilayer epidermis15,16,17. In zebrafish, the fin fold epidermis is bilayered from as early as 2 days post-fertilization (dpf)17,18, and is composed of BECs and SECs, which functionally resemble the mammalian periderm. Notably, the zebrafish fin fold covers the embryonic tailbud and developing limb-like structures, analogous to the periderm coverage of mammalian limb buds19. The BECs remain undifferentiated until around 15 dpf, when they begin to differentiate into a stratified epidermis16. Thus, we used the zebrafish fin fold, and an in vitro human bilayer skin model, to investigate how BECs regulate epidermal architecture and function independently of differentiation.
We discovered that BECs generate distinct ECM zones within the fin fold, characterized by collagen-rich ECM at the center and laminin-rich basement membranes at the periphery. In both zebrafish and human skin models, a laminin-rich ECM beneath BECs was both necessary and sufficient to suppress desmosome formation while maintaining AJs at the BEC–SEC interface. In vivo, this junctional specialization promotes tissue repair after cell injury in the wild-type periderm. While impaired collagen decreases both AJs and desmosomes, reduced laminin expression increases desmosome formation in BECs and across the epidermal bilayer, suppresses actomyosin expression at AJs, and impairs wound healing of the periderm at the fin fold periphery. Partial depletion of desmoplakin 1a (dspa), which encodes for the desmosomal linker protein expressed across epidermal layers, partially rescues the wound closure defects observed at the fin fold periphery of laminin mutants.
Our work suggests that BECs strengthen periderm resilience to tissue injury by forming region-specific ECM zones that direct intercellular junction specialization, revealing a role in skin organization and repair independent of differentiation.
Results
BECs form distinct zones of ECM in zebrafish embryos
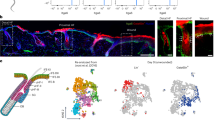
During zebrafish embryogenesis, the entire embryo is enveloped by a bilayer epidermis (Fig. 1A)17,20. The bottom layer consists of basal epidermal stem cells (BECs), which express p6321,22,23, and the top layer of superficial epidermal cells (SECs), called periderm, are p63-negative and express higher levels of keratin 521 (Fig. 1A–C and Extended Data Fig. 1A).
A Zebrafish fin fold 2-2.5 days post fertilization (dpf). Basal epidermal stem cells (BECs) on basement membrane extracellular matrix (ECM) and superficial epidermal cells (SECs) (transverse). B Representative images of TgBAC(ΔNP63:Gal4ff)la213; Tg(UAS:NLS-GFP)el609 BECs at 48hpf, 16-color intensity scale. C Nuclear GFP mean intensity quantification across fin fold. Y-axis: Average normalized intensity/embryo, x-axis: normalized distance from fin fold center (1.0 = fin fold edge). Simple linear regression (red dashed line) (slope=0.2743, n = 42 embryos, two independent experiments). D Representative image of FAM-tagged collagen-hybridizing peptide (left) and Laminin-A1 protein (right). Intensity is displayed using a fire LUT. E Quantification of normalized intensity from fin fold edge (edge = 0 µm). Periphery<50 µm from edge, Center>50 µm. F Violin plot of averaged values for center and periphery/embryo (Kruskal-Wallis test with multiple comparisons, p < 0.0001, n = 41 embryos; 4 independent experiments). G Representative image of TgBAC(ΔNP63:Gal4ff)la213; Tg(UAS:LifeAct-GFP)mu271 in central and peripheral regions as defined in panels E and F. Quantification of (H) BEC area and (I) aspect ratio displayed as a violin plot (distribution=individual cells) with average values/embryo (dots) in central and peripheral regions (two-tailed Mann-Whitney test on violin plots, p < 0.0001 (BEC area and aspect ratio), n = 114-152 cells, dots n = 8–14 embryos; 2 independent experiments). J, N Representative images showing (J) TgBAC(ΔNP63:Gal4ff)la213; Tg(UAS:LifeAct-GFP)mu271 for endogenous E-cadherin labeling, alongside immunofluorescence staining for phosphorylated-myosin light chain and (N) desmoplakin1/2 in central and peripheral regions of fin fold BECs at 48hpf. Z-stack images obtained from SEC-BEC layer. BECs identified by max projection of selected z-slices based on z-position, 16-color intensity scale. K–M, O Quantification of (K) BEC-LifeAct-GFP/cell, junctional (L) E-cadherin, (M) phospho-myosin light chain, and (O) desmoplakin1/2, shown as violin plots (distribution=individual junctions, average values/embryo=dots). Intensity normalized to Center group for each graph. (two-tailed Mann-Whitney test, p < 0.0001 (LifeAct, Desmoplakin), p = 0.000003 (pMyosin), n = 172-175 cells and (BEC-LifeAct), 240 junctions (E-cadherin), 93-120 junctions (pMyosin), and 284-348 junctions (Desmoplakin), dots n = 13-14 (BEC-LifeAct; 2 independent experiments), 18 (E-cadherin; 4 independent experiments), 6 (pMyosin; 1 independent experiment), and 18 (desmoplakin; 2 independent experiments) (P) Summary schematic of intercellular junctions in BECs above laminin- vs. collagen-positive basement membranes. Source data are provided as a Source Data file.
To investigate how undifferentiated BECs contribute to epidermal organization, we first analyzed the composition and spatial distribution of ECM components in the zebrafish fin fold. BECs in the fin express specific types of collagen and laminin24, protein components of the extracellular matrix (ECM) that in turn are required for fin fold development21,25,26 (Extended Data Fig. 1B–E). Collagen is produced by both BECs and embryonic fibroblasts, in contrast, laminins are primarily produced by BECs (Extended Data Fig. 1F)27,28. Confocal imaging of fluorescein-tagged collagen hybridizing peptide, to detect all types of collagens29, revealed fibrillar collagens in the stromal ECM in the fin fold central region, which coincides with the localization of fin mesenchymal cells and interstitial fibroblasts in the dermis30 (Fig. 1D–F, and Extended Data Fig. 1G, H). We also detected cytoplasmic collagens in BECs in the central region (Extended Data Fig. 1H). Unexpectedly, we found that BECs produce a relatively high level of laminin31 only in the peripheral zone of the fin fold compared to the central region (Fig. 1D–F, and Extended Data Fig. 1I).
These findings reveal a region-specific pattern of ECM composition partially produced by BECs depending on their spatial context within the fin fold.
Peripheral BECs show reduced AJs and desmosomes
We hypothesized that regional ECM differences in the fin fold are associated with changes in the cytoskeletal and junctional organization of BECs. To test this, we first analyzed the actin cytoskeleton in BECs. Briefly, we performed confocal live imaging of the actin reporter LifeAct-GFP regulated by the BEC-specific promoter p63. Notably, BEC morphologies and areas were significantly different in the central and peripheral regions of the fin fold (Fig. 1G–I). Specifically, BECs above the peripheral, laminin-rich areas exhibited a larger and more elongated cell area and lower actin levels, compared to those in the central, collagen-dense regions (Fig. 1H-K).
Actin filaments interact with adherens junctions (AJs), intercellular junctions that regulate tissue tension via phosphorylated myosin32. By live confocal microscopy, we found that the expression of the AJ component E-cadherin33 was lower in peripheral BECs compared to the central BECs (Fig. 1J–L). Similarly, immunostaining followed by confocal microscopy revealed reduced junctional expression of phosphorylated-myosin and vinculin34,35 in peripheral compared to central BECs (Fig. 1J–M and Extended Data Fig. 1J, K). To determine if these differences in BECs were unique to AJs, we examined desmosomes. Immunostaining of desmoplakin 1/236 revealed substantially reduced levels of this desmosome component in the peripheral BECs compared to the neighboring central BECs (Fig. 1N, O).
Overall, these data suggest that BECs have differential adhesion profiles correlated with a region-specific pattern of ECM composition (Fig. 1P).
Peripheral SECs sustain AJs and reduce desmosomes
In zebrafish, as in mammals, BECs are tightly connected to SECs10,22,37. Thus, we examined how regional ECM differences beneath the BEC layer might influence the SECs above. Specifically, we analyzed cytoskeletal and junctional organization in SECs overlying the central (collagen-rich) versus peripheral (laminin-rich) zones of the fin fold.
First, we performed confocal live imaging of LifeAct-GFP regulated by the SEC-specific promoter krt5 (Fig. 2A and Extended Data Fig. 1A). Similar to BECs, SECs exhibited differential cytoskeletal organization across the ECM zones. Specifically, in the central, collagen-rich region, SECs exhibited junctional actin and well-organized microridges, which are apical surface protrusions composed of polymerized actin. In contrast, SECs in the peripheral, laminin-positive region showed similar junctional actin but less organized apical actin (Fig. 2A).
A Representative image of Tg(krt5:Gal4ff)la212; Tg(UAS:LifeAct-GFP)mu271 fin fold central region (green) and peripheral boundary (purple) as defined in Fig. 1D–F. Quantifications of (B) SEC apical area and (C) aspect ratio by violin plot (distribution=individual cells, dots=average values/embryo). (apical area: two-tailed unpaired t test p = 0.4548, n = 83-86 cells, dots n = 6-7; aspect ratio: two-tailed Mann-Whitney test p = 0.1672, n = 48-160 cells, dots n = 5-6; both: 3 independent experiments). D, G Representative images showing (D) cdh1-tdTomatoxt18, alongside immunofluorescence staining for phosphorylated-myosin light chain and (G) desmoplakin1/2 at 48hpf. Z-stack images from SEC-BEC layer. SECs identified by max projection of selected z-slices by z-position. 16-color intensity scale. E, F, H Quantification of intensity for junctional (E) E-cadherin, (F) phospho-myosin light chain, and (H) desmoplakin1/2 across embryos, shown as violin plots (distribution=individual junctional measurements, dots=average values/embryo). Intensity normalized to Center group for each graph (two-tailed Mann-Whitney test on violin plots, p = 0.2577 (E-cadherin), 0.0457 (pMyosin), and p < 0.0001 (Desmoplakin), n = 372 junctions (E-cadherin), 99-119 junctions (pMyosin), and 362-366 junctions (Desmoplakin), dots n = 18 (E-cadherin; 4 independent experiments), 6 (pMyosin; 1 independent experiment), and 18 (desmoplakin; 2 independent experiments) embryos). Source data are provided as a Source Data file. I Representative orthogonal view of the fin fold, showing live imaging of cdh1-tdTomatoxt18 or desmoplakin1/2 immunofluorescence staining in collagen-enriched (center) and laminin-enriched (periphery) regions. (Top) Images displayed using 16-color scale. (Bottom) Interface membrane (yellow dashed lines) between SEC and BEC layers, with numbers corresponding to each analyzed cell. J, K Quantification of interlayer (yellow dashed lines) (J) E-cadherin and (K) desmoplakin across embryos, individual measurements distribution shown as violin plots and average values per embryo as dots. Intensity was normalized to the Center group for each graph (two-tailed Mann-Whitney test on violin plots, p = 0.2917 (E-cadherin) and p < 0.0001 (Desmoplakin), n = 70 (E-cadherin), and 60 cells (desmoplakin), dots n = 7 (E-cadherin; 2 independent experiments) and 6 (desmoplakin; 1 independent experiment) embryos). L Summary schematic of junctional characteristics in SECs, SEC-BEC interface, and BECs. Source data are provided as a Source Data file.
Interestingly, SECs in the peripheral and central regions had similar apical cell areas and cell shapes, indicating that the differential distribution of microridges is not associated with cortical contractions38 (Fig. 2B, C). Furthermore, SECs in both regions had equal E-cadherin, and phosphorylated myosin at the junction was elevated in peripheral versus central SECs (Fig. 2D–F). In contrast, desmoplakin junctional expression was reduced in peripheral versus central SECs (Fig. 2G, H). These findings suggest that SECs above laminin-rich ECM maintain epithelial integrity through enhanced actomyosin signaling, potentially compensating for reduced desmosomal adhesion. Thus, SECs exhibit region-specific junctional complexity, characterized by a decoupling of AJs and desmosomes (Fig. 2L).
Since intercellular SEC-SEC and BEC-BEC junctions differ in the peripheral and central regions, we investigated whether these molecular differences extended to interlayer adhesion between BECs and SECs. We performed live imaging of the cdh1-tdTomato; krt5:Gal4ff, UAS:LifeAct-GFP transgenic line, which allowed us to discern the SEC layer (pseudo-colored in grey and yellow) from the BEC layer (pseudo-colored in yellow only) (Extended Data Fig. 2A, Supplementary Movie 1). Notably, the BEC and SEC layers migrated synchronously over the 8 h of imaging, both at the center and periphery regions, with no observable difference in the mean squared displacement between BECs and SECs (Extended Data Fig. 2B). Further tracking of individual BECs and SECs showed sustained interlayer contact over time (Extended Data Fig. 2C), suggesting their fixed connection in both center and periphery. However, when we examined the levels of junction proteins at the membrane interface between BECs and SECs, we found that desmoplakin was more abundant at the central versus peripheral BEC-SEC interface, and E-cadherin levels were consistent across the entire interface (Fig. 2I–K), similar to the organization of SEC-SEC junctions. These findings suggest that the BEC-SEC bilayer is connected via both desmosomes and AJs above the collagen-rich central region, and mainly via AJs above the laminin-rich peripheral region (Fig. 2L).
Together, these findings support a model in which BEC–ECM interactions direct the assembly of regionally specialized junctions across the bilayer, with peripheral SECs relying more heavily on actomyosin contractility and AJs to maintain tissue cohesion in the context of reduced desmosomal support.
BEC-ECM integrin adhesions promote AJs and reduce desmosomes
We hypothesized that BEC-ECM adhesions regulate the organization of intercellular junctions within and between both layers of the epidermis. To test this, we examined mutants with reduced levels of integrin β1, a transmembrane receptor required for cell-ECM adhesions in both collagen- and laminin-rich ECMs39.
In zebrafish, integrin β1 is encoded by itgb1b and itgb1a, and we detected high levels of itgb1b mRNA and protein in BECs in both regions, but especially in the central BECs (Extended Data Fig. 3A–E). As expected, we did not detect high levels of itgb1b and itgb1a in SECs, which do not directly interact with the ECM (Extended Data Fig. 3A-E). To avoid possible compensatory effects between isoforms, we analyzed double heterozygous itgb1b;itgb1a mutants (itgb1b+/-;itgb1a+/-). These mutant embryos did not exhibit any obvious defects in growth or fin fold development (Extended Data Fig. 3F).
A Representative image of wild type (WT) vs. itgb1a+/-, itgb1b+/- fin fold, stained with a FAM-tagged collagen hybridizing peptide and Laminin-A1 at ~48hpf. Intensity displayed using fire LUT. B Quantification of normalized collagen and laminin intensity across fin fold (edge = 0 µm). Each curve is normalized to individual embryo maximum. C Quantification of average normalized collagen and laminin intensity/embryo (Collagen: Ordinary One-Way ANOVA test with multiple comparisons, p = 0.6422 (Center) and 0.4573 (Periphery), n = 18 (WT) and 27 (itgb1a+/-, itgb1b+/-) embryos; 3 independent experiments; Laminin: Kruskal-Wallis test with multiple comparisons, p > 0.9999, n = 6 (WT) and 11 (itgb1a+/-, itgb1b+/-) embryos; 2 independent experiments. D, F,H, J Representative images of cdh1-tdTomatoxt18 ((D) center and (F) periphery and immunofluorescence staining for desmoplakin (H) center and (J) periphery in zebrafish fin folds at 48hpf. Images show SECs, Interlayer (SEC-BEC), and BECs. Z-stack images were taken from the SEC-BEC layer. SECs and BECs were identified by maximum projection of selected z slices based on their z-position. Representative interlayer images are shown in orthogonal view of single z slice. 16-color intensity scale. E, G, I, K Quantification of E-cadherin (E) center and (G) periphery and desmoplakin (I) center and (K) periphery at SEC junctions, interlayer, and BEC junctions shown as violin plot (distribution=individual junctional measurements, dots= average values/embryo). E, G For all graphs, E-cadherin intensity was normalized to the WT Center BEC group (Violin plots tested for two-tailed unpaired t test (Center Interlayer) and two-tailed Mann-Whitney test for all other comparisons, p < 0.0001 (all comparisons), n = 180−200 junctions (SEC and BEC junctions) and 90–100 cells (interlayer), dots n = 9 (WT) and 10 (itgb1a+/-, itgb1b+/-) embryos; 2 independent experiments). I, K For all graphs, desmoplakin intensity was normalized to the WT Center BEC group (Violin plots tested for two-tailed Mann-Whitney test, p < 0.0001 (all comparisons), n = 359 − 395 junctions (SEC and BEC junctions) and 188−190 cells (interlayer), dots n = 20 (WT) and 19 (itgb1a+/-, itgb1b+/-) embryos; 2 independent experiments). Source data are provided as a Source Data file.
Given that integrins can alter the expression of ECM components40,41, we verified that itgb1b+/-;itgb1a+/- mutant embryos show a fin fold morphology, region-specific ECM composition, and overall ECM levels comparable to wild-type (WT) controls (Fig. 3A–C and Extended Data Fig. 3F). This allowed us to isolate the role of integrin-mediated adhesion independent of changes in ECM.
Despite intact ECM composition, there was a profound decrease (p < 0.0001) of the AJ component E-cadherin in both the center and periphery regions of the fin fold of itgb1b+/-;itgb1a+/-, not only between BECs but also at the BEC-SEC interface, and even between SECs (Fig. 3D–G). In contrast, desmoplakin levels increased in both center and periphery regions between BECs, SECs, and at the BEC-SEC interface in mutants compared to WT (Fig. 3H–K). Thus, mutants with reduced integrin β1 exhibit increased desmoplakin levels and decreased E-cadherin across the bilayer epidermis in both central and peripheral regions.
Given that SECs do not directly contact the ECM, these results suggest that reduced BEC–ECM adhesion has non-cell-autonomous effects on junctional organization in overlying SECs.
Together, these findings support a model in which integrin-mediated BEC–ECM interactions coordinate the balance of adherens and desmosomal junctions across both layers of the epidermis.
ECM drives junctional specialization in human epiderm model
Our data suggest that BEC-ECM interactions are required to differentially regulate cell-cell junctions throughout the bilayer epidermis. We hypothesized that the interactions between specific ECM proteins and BECs are sufficient to establish interlayer junctional specialization. To test this hypothesis, we employed a 3D in vitro cell culture model of bilayered human epidermis, enabling precise regulation of the BEC-ECM composition. Specifically, we engineered scaffold materials to mimic the central vs peripheral ECM in the fin fold.
Human keratinocytes cultured in low calcium conditions form a proliferative monolayer. When calcium levels are elevated, these keratinocytes undergo stratification within 24 hours (hrs), accompanied by the accumulation of terminal differentiation markers42,43,44. As expected, we found that treating human basal keratinocytes with Ca2+ for 48 hrs, resulted in two layers: basal keratinocytes positive for P63, and a layer of large suprabasal keratinocytes with significantly lower P63 expression, suggesting successful differentiation45 (Fig. 4A, B).
A Representative immunofluorescence images of P63, F-actin, and DAPI in both layers of the bilayer normal human epidermal keratinocytes (NHEK). DAPI image is z-depth color-coded. B Quantification of P63 intensity (two-tailed unpaired t-test, p < 0.0001, n = 15 (Basal) and 16 (Suprabasal) containing 5–20 cells/FOV (fields of view); 1 independent experiment). Box plot whiskers represent the minimum and maximum value. Bounds mark the 25th and 75th percentiles. Central line is the median. C Schematic of gel constructs containing 100% collagen (Collagen) and 100% collagen gel coated with laminin (Col (LM-coated)). D Representative immunofluorescence z-stack cross sections of NHEKs cultured on each gel system. E Representative immunofluorescence images of β-Catenin (CTNNB1) in culture on each gel type. Images of the Suprabasal, Interlayer (Suprabasal-Basal), and Basal shown. Intensity displayed using a 16-color scale. F Quantification of CTNNB1 intensity at junctions for Suprabasal, Interlayer, and Basal in NHEKs cultured on each gel system. For each graph, intensity was normalized to the Collagen Gel Group (two-tailed unpaired t-test (Suprabasal and Basal) and two-tailed Mann-Whitney test (Interlayer), p = 0.0046 (Suprabasal), 0.2213 (Interlayer), and p < 0.0001 (Basal), n= Suprabasal: 44 (Collagen) and 51 (Col LM coated) cells, Interlayer: 53 cells (Collagen and Col LM coated), and Basal: 41 (Collagen) and 44 (Col LM-coated) cells; 1 independent experiment). Data presented as box plots with whiskers to minimum and maximum value. Bounds mark the 25th and 75th percentiles. Central line is the median. G Representative immunofluorescence images of Desmoplakin (DSP) in culture on each gel type. Layers and intensity displayed as in (E). H Quantification of Desmoplakin intensity at junctions as in (F). (two-tailed unpaired t-test (Suprabasal and Interlayer) and two-tailed Mann-Whitney test (Basal), p = 0.0042 (Suprabasal) and p < 0.0001 (Interlayer and Basal), n= Suprabasal: 41 (Collagen) and 42 (Col LM coated) cells, Interlayer: 48 (Collagen) and 50 (Col LM coated) cells, and Basal: 40 (Collagen) and 42 (Col LM-coated) cells; 1 independent experiment). Data presented as box plots with whiskers to minimum and maximum value. Bounds mark the 25th and 75th percentiles. Central line is the median. Source data are provided as a Source Data file.
We seeded basal keratinocytes on 100% Collagen type I scaffold materials, the major component of the central fin fold and of the dermal ECM in humans, and on a Collagen type I scaffold coated with basement membrane Laminins (containing Laminin A, B1, and B2 chains) mimicking the peripheral fin fold (Fig. 4C). In both conditions, the addition of Ca2+ for 48 hrs still generated a bilayered epidermis (Fig. 4D). Furthermore, we confirmed that basal keratinocytes produced the same types and levels of ECM under both conditions (Extended Fig. 4A–D).
Despite these similarities, the junctional composition differed markedly by ECM scaffolds. Compared to basal keratinocytes seeded on Collagen-1 alone, those seeded on Laminin-coated collagen scaffolds displayed a profound decrease in the junction levels of the AJ component β-Catenin46 and of desmoplakin (Fig. 4E–H, Bottom panels). Suprabasal keratinocytes showed a similar, though less pronounced, Laminin-dependent reduction in β-Catenin and desmoplakin at junctions (Fig. 4E–H, Top panels). Notably, the Laminin-coated scaffold led to reduced levels of desmoplakin without affecting β-Catenin at the junctions between basal and suprabasal keratinocytes (Fig. 4E–H, Middle panels; Extended Data Fig. 4E–F).
These results suggest that ECM composition is sufficient to influence junctional specialization in bilayered epithelia. In particular, Laminins appear to selectively limit desmoplakin enrichment at interlayer contacts, mirroring patterns observed in the zebrafish fin fold.
ECM drives junctional specialization in zebrafish epiderm
Our data thus far suggest that the differential distribution of ECM proteins in the fin fold creates distinct intercellular junctions in the bilayer epidermis. To assess the physiological role of specific ECM components, we generated zebrafish mutants that disrupt laminin a5 (lama5) and collagen type I alpha 1 chain (col1a1a), two of the most enriched ECM proteins in the BEC (Extended Data Fig. 1F).
Homozygous mutants for both lama5 and col1a1a display abnormal fin development25,47, so we used heterozygous embryos, which maintain normal development and intact fin folds, enabling us to examine bilayer architecture (Extended Data Fig. 5A). Heterozygous mutants exhibited reduced levels of their respective ECM proteins throughout the fin fold, with lama5+/- showing a prominent reduction of laminin a5 at the periphery and col1a1a+/- showing a reduction of collagens in the central zone (Fig. 5A–C).
A Representative immunofluorescence image of wild type (WT) vs. lama5+/- vs. col1a1a+/- fin fold, stained with FAM-tagged collagen-hybridizing peptide and Laminin-A1 at ~48hpf. Intensity displayed using fire LUT. B Quantification of average normalized collagen and laminin intensity/embryo. (Collagen: Kruskal-Wallis test with multiple comparisons, p > 0.9999, n = 20 (WT) and 26 (lama5+/-) embryos; 2 independent experiments; Laminin: Ordinary One-Way ANOVA test with multiple comparisons, p = 0.2249 (Center) and 0.0006 (Periphery), n = 18 (WT) and 28 (lama5+/-) embryos; 2 independent experiments. C Quantification as in B. (Collagen: Kruskal-Wallis test with multiple comparisons, p = 0.0246 (Center) and > 0.9999 (Periphery), n = 27 (WT) and 33 (col1a1a +/-) embryos; 3 independent experiments; Laminin. Ordinary One-Way ANOVA test with multiple comparisons, p = 0.8872 (Center) and 0.2019 (Periphery), n = 13 (WT) and 18 (col1a1a +/-) embryos; 2 independent experiments. D, F Representative (D) images of cdh1-tdTomatoxt18 and (F) immunofluorescence staining for desmoplakin in peripheral fin fold region at ~48hpf. Z-stack images taken from SEC-BEC layer. SECs and BECs identified by maximum projection of selected z-slices based on z-position. Representative interlayer images shown in orthogonal view of single z-slice. Intensity displayed using 16-color scale. E, G Quantification of (E) E-cadherin and (G) desmoplakin at junctions in peripheral SEC, interlayer (SEC-BEC), and BEC layers shown as violin plot (distribution=individual junctional measurements; dots= average values/embryo). E For all graphs, E-cadherin intensity was normalized to the WT Center BEC group in panel Extended Data Fig. 5C (two-tailed Mann-Whitney test on violin plots, p = 0.2132 (SEC), 0.4736 (Interlayer), and p < 0.0001 (BEC), n = 348-363 junctions (SEC and BEC junctions) and 160-169F cells (interlayer), dots n = 19 (WT) and 18 (lama5+/-) embryos; 3 independent experiments). G For all graphs, desmoplakin intensity was normalized to the WT Center BEC group in panel Extended Data Fig. 5E (two-tailed Mann-Whitney test on violin plots, p = 0.0014 (SEC), 0.0036 (BEC), and p < 0.0001 (Interlayer), n = 232-240 junctions (SEC and BEC junctions) and 117-120 cells (interlayer), dots n = 12 (WT) and 12 (lama5+/-) embryos; 2 independent experiments) (H, J) Representative (H) images of cdh1-tdTomatoxt18 and (J) immunofluorescence staining for desmoplakin in central fin fold region at ~48hpf as in (D, F). I, K Quantification of (I) E-cadherin and (K) Desmoplakin at central SEC junctions as in (E, G). I For all graphs, E-cadherin intensity was normalized to the WT Center BEC group in bottom panel Fig. 5I (two-tailed unpaired t-test for BEC Center and two-tailed Mann-Whitney test on all other violin plots, p = 0.0001 (SEC), 0.0009 (Interlayer), and 0.04825 (BEC), n = 170 junctions (SEC and BEC junctions) and 110 cells (interlayer), dots n = 11 (WT) and 11 (col1a1a+/-) embryos; 1 independent experiment). K For all graphs, Desmoplakin intensity was normalized to the WT Center BEC group in panel Fig. 5K (two-tailed unpaired t-test on Interlayer, and two-tailed Mann-Whitney test on all other violin plots, p < 0.0001 (SEC), p = 0.01437 (BEC) and 0.0011 (Interlayer); n = 62-82 junctions (SEC and BEC junctions) and 50–60 cells (interlayer), dots n = 6 (WT) and 6 (col1a1a+/-) embryos; 1 independent experiment) (L) Summary schematic of junctional characteristics in the peripheral regions of SECs, SEC-BEC interface, and BECs. Source data are provided as a Source Data file.
First, we analyzed lama5+/- mutant fin folds. For both the central and peripheral regions, E-cadherin levels were lower between BECs, but not between SECs or the SEC-BEC interface, in lama5+/- compared to WT (Fig. 5D, E). By contrast, desmoplakin levels were elevated throughout the lama5+/- mutant versus WT: between BECs, SECs, and BEC-SECs, in the central and peripheral regions (Fig. 5F, G) (Extended Data Fig. 5B–I).
Next, we examined col1a1a+/- mutants. In the central region, we observed a significant reduction in the levels of E-cadherin and desmoplakin across all layers in the mutant compared to WT (Fig. 5H–K). E-cadherin and desmoplakin junction levels were both decreased in the peripheral region of col1a1a+/- fins in BECs, BEC-SECs but not in SECs compared to WT (Extended Fig. 5J–Q).
Together, these findings suggest that BEC-laminin interactions promote AJ formation while limiting desmosomal enrichment, whereas BEC–collagen interactions support both junction types across layers, particularly in the central fin fold (Fig. 5L).
Region-specific ECM requirements for epiderm wound closure
SECs within the periderm in both humans and zebrafish play a critical role in protecting embryonic BECs against external physical forces. We hypothesized that BEC-laminin interactions, and in turn the differential adhesion profiles in the central versus peripheral periderm, might affect its response to mechanical injury. To test this hypothesis, we used a two-photon laser to induce a single-cell ablation in the SEC layer.
Strikingly, SECs in the laminin-enriched peripheral region completed wound closure faster than SECs in the collagen-enriched center region (Fig. 6A and B, Supplementary Movie 2). These observations establish a region-specific difference in epithelial repair and suggest that the ECM composition may influence the wound healing response.
A Representative live confocal images of Tg(krt5:Gal4ff)la212; Tg(UAS:LifeAct-GFP)mu271 for LifeAct-GFP under SEC-specific krt5 promoter, showing the central and peripheral regions of the fin fold at 2dpf before and after single-cell ablation. Shaded border indicates the ablated cell and subsequent wound closure. B Quantification of time (min) required for complete wound closure. Data presented as box plots with whiskers to minimum and maximum value. Bounds mark the 25th and 75th percentiles. Central line is the median. (two-tailed Mann-Whitney test, p = 0.0040, n = 8 (Center) and 6 (Periphery) embryos; 4 independent experiments). C Representative live confocal images of Tg(krt5:Gal4ff)la212; Tg(UAS:LifeAct-GFP)mu271 for LifeAct-GFP under the SEC-specific krt5 promoter in the peripheral regions of fin fold of indicated genotypes at 2dpf before and after single-cell ablation. Shaded border indicates the ablated cell and subsequent wound closure. D Quantification of periphery wound closure time (min), comparing WT and dspa+/- mutants. Data presented as box plots with whiskers to minimum and maximum value. Bounds mark the 25th and 75th percentiles. Central line is the median. (Mann-Whitney test, p = 0.7879, n = 6 (WT) and 5 (dspa+/-) embryos; 4 independent experiments). E Quantification of wound closure time (min), comparing periphery WT, lama5+/-, and lama5+/-, dspa+/- mutants. Data presented as box plots with whiskers to minimum and maximum value. Bounds mark the 25th and 75th percentiles. Central line is the median. (Kruskal-Wallis test with multiple comparisons; p = 0.0273 (WT vs. lama5+/-), > 0.9999 (WT vs. lama5+/-, dspa+/-), and 0.1988 (lama5+/- vs. lama5+/-, dspa+/-); n = 6 (WT), 9 (lama5 +/-), and 5 (lama5+/-, dspa+/-) embryos; 5 independent experiments. Source data are provided as a Source Data file.
To examine the role of laminin more directly, we assessed wound healing in lama5+/− mutants. SECs took longer to complete wound closure after single-cell injury in the peripheral region of the lama5+/− mutant fin fold compared to WT, whereas repair time in the center region was unaffected (Fig. 6C, E, and Extended Data Fig. 6A, B; Supplementary Movie 3, 4). Moreover, compared to WT, lama5+/− peripheral SECs had increased levels of microridge actin, peripheral BECs had elevated cortical actin organization, and both peripheral cell layers had reduced junction levels of phosphorylated myosin (Extended Fig. 6C–H). These changes were not observed in the central region of lama5+/− versus WT (Extended Fig. 6I–N). These findings indicate that laminin a5 is required in the periphery to support efficient wound closure, likely through regulation of actin localization and contractility.
Next, we assessed wound closure in col1a1a+/− fins. In contrast to the lama5+/− mutant fin fold, the time required for peripheral SECs to complete wound healing was comparable in col1a1a+/− and WT (Extended Data Fig. 7A, B, Supplementary Movie 5), whereas centrally positioned SECs took longer to close wounds in col1a1a+/− fin folds compared to WT (Extended Data Fig. 7C, D, Supplementary Movie 6). These findings suggest that collagen plays a more prominent role in promoting repair within the center of the fin fold epithelium.
We propose that the interaction between basal epidermal stem cells (BECs) and laminin influences the junctional organization and injury response of the periderm, via the regulation of interlayer junction dynamics. The developing bilayered skin appendage is organized as collagen-enriched areas, where BECs exhibit increased adherens junction (AJ) actomyosin signaling and desmosome formation. Superficial epidermal cells (SECs) layer (the periderm) above these BECs display reduced motility, characterized by stronger desmosomal junctions and less actomyosin signaling. In laminin-rich areas, BECs show decreased actomyosin activity and reduced AJ and desmosome formation within the intralayer, but they unexpectedly maintain interlayer AJ and repress desmosome formation. SECs above these BECs have faster motility during wound closure. Partial removal of laminin matrices further reduces BEC actomyosin activity, AJ and desmosome formation while increasing desmosome connections at the interfaced membrane with SECs. SECs, connected to these BECs, exhibit increased desmosome overall, and slow wound closure.
Because BEC-laminin interactions limit, whereas BEC-collagen interactions promote, desmoplakin expression, we hypothesized that the increased desmoplakin in the periphery of lama5+/− fin folds underlies impaired wound closure. To test this, we reduced desmoplakin levels by generating dspa mutants. Zebrafish desmoplakin is encoded by two paralogs, dspa and dspb, with dspa being the predominant isoform expressed in BECs (Extended data Fig. 7E-G). We analyzed the fin fold in heterozygotes to avoid the gross morphological defects reported for homozygous mutants48, and observed a small but significant reduction of desmoplakin protein levels in peripheral but not central BECs and SECs of dspa+/- mutants (Extended Data Fig. 7H-M). We found that the time to complete wound closure was similar in dspa+/− mutant and WT peripheral fin folds (Fig. 6C, D and Supplementary Movie 7). Importantly, lama5+/−; dspa+/− double mutants showed partially rescued wound closure times compared to lama5+/− mutants, comparable to WT (Fig. 6C, E, anSupplementary Movie 8). These results suggest that elevated desmoplakin and desmosome accumulation in a laminin-deficient background restricts wound repair to the periphery region.
Overall, our findings suggest that regional ECM composition tunes the wound-healing capacity of the zebrafish fin fold epidermis through its influence on BECs and, in turn, junctional architecture and cytoskeletal organization (Fig. 7).
Discussion
Here, we identify a region-specific adhesion profile within the periderm in the fin fold acquired through interactions between basal epidermal stem cells (BECs) and the extracellular matrix (ECM). The ECM is known to regulate BECs differentiation and fate23,49,50,51,52,53,54. Our findings provide, to our knowledge, the first evidence that BEC-ECM interactions influence injury resilience of an interconnected tissue without affecting their differentiation into stratified cell layers. During embryonic development, superficial epidermal cells (SECs) form a protective layer, the periderm, that extends from the developing limb (fin) to other regions of the embryo body and plays a crucial role in shielding the underlying stem cells from mechanical stresses. We found that the response of the periderm of the fin fold to cell injury is directly dependent on BEC-ECM interactions. BECs organized their junctions with SECs in a region-specific manner, forming both desmosomes and AJs above the central, collagen-enriched ECM, but predominantly AJs above the peripheral, laminin-enriched ECM. We propose that BEC-laminin interactions serve to suppress desmosomes across layers, this shift decreases cellular resistance while preserving SEC actomyosin contractility and tissue integrity, enabling the SECs above the BEC-laminin areas to close wounds faster following cell injury (Fig. 7).
The regulation of cell-cell junctions and cell-ECM interactions has been extensively studied in the context of cell injury and wound healing20,55,56. A profound reorganization of actomyosin contraction typically occurs during wound closure57, and additional junctional forces, including cell resilience generated by desmosomes at the junction can be activated due to substrate stiffness or cell deformation58,59,60,61,62. Overall junctional tension is enhanced through desmosomes and adherens junctions during tissue injury responses. Consequently, the absence of desmosomes is often linked to impaired wound healing, loss of tissue integrity and the formation of blistering in epidermis63. In contrast, our data suggest that interactions between BECs and laminin-rich matrices suppress interlayer desmosomes and, while maintaining the contacts with SEC via AJ, desmosome repression promotes wound closure. This apparent inconsistency may stem from several factors. First, most cell ablation studies have focused on monolayer cell models in direct contact with the ECM, where actin cytoskeleton reorganization must coordinate junctional and focal adhesion tensions to support both actomyosin contraction and cell migration. In contrast, cell injury within stacked cell layers—such as the bilayered epidermis—has, to our knowledge, never been directly evaluated in the context of inter-tissue interactions. Second, based on our model, we propose that desmosomes at the SEC-BEC interface enhance the resistance to cell deformation64 triggered by the death of SECs. Hence, fewer desmosomes at the SEC-BEC interlayer reduce SEC resistance and accelerate wound repair. This is in agreement with the dynamic needs of the developing fin fold, where peripheral regions also have less desmosome presence. This arrangement may balance the simultaneous expansion of skin stem cells and their protection by the periderm during skin repair at the appendages. Despite this reduction in desmosomes, we do not observe increased interlayer movement during wound closure or development, likely because AJs and cortical actin networks maintain coordinated coupling between BECs and SECs. Thus, reduced desmosome levels may enhance cellular dynamics without compromising overall interlayer integrity.
The organization of the ECM plays a crucial role in determining the distribution and type of junctions in monolayered cells, mediated by interactions with various cytoskeletal proteins. ECM distribution is also generally critical for organ morphogenesis. For instance, laminin typically supports basement membrane stability and epithelial integrity, but it is also required to promote osteogenesis25,65,66. Hence, full removal of laminin a5 abolishes fin development25 and thus its effect on epithelial integrity can be tested only with partial loss-of-function models. Similarly, collagen-rich matrices provide stiffness and tensile strength and are key for mesenchymal growth during the initial limb development26,47,67. Collagen is produced by both basal stem cells and embryonic fibroblasts, which are enriched in the collagen-dense central areas of the fin fold. In contrast, laminins are primarily produced by basal epidermal stem cells27 and are concentrated at the fin’s very tip, where fibroblasts are sparse. Since complete deletion of these ECM proteins severely disrupts fin growth, and tissue-specific deletion remains to be investigated, we acknowledge that our data only correlate BEC–ECM interactions with effects on SECs. Thus, other cell types may also be necessary to confer laminin-BEC specialization within the fin fold. Nevertheless, our in vivo studies suggest that central BECs on collagen-enriched areas exhibit increased AJ actomyosin signaling and desmosome formation compared to BECs on laminin-enriched areas. Notably, peripheral BECs in lama5+/- mutants, which interact with reduced levels of laminin, exhibit a different adhesion molecule organization compared to WT central BECs, which also interact with relatively low laminin levels. This suggests that collagen alone in the peripheral regions with decreased laminin is insufficient to recapitulate the BEC architecture observed in the WT center. These differences could also be attributed to ECM properties beyond protein composition, such as the architecture and stiffness of the basement membrane, or other microenvironmental differences such as tissue signaling functions. Further investigation is needed to understand how these properties differentially influence surrounding tissues through their interactions with stem cell compartments.
Communication between skin layers has been observed in both zebrafish embryo (bilayered skin) and multilayered mouse skin10,22,37. For instance, the loss of E-cadherin function in either the periderm or BECs affects E-cadherin levels in the other layer. E-cadherin functions as a transducer of polarity between the periderm and BECs22. It regulates the localization and levels of Lgl2, a basolateral polarity regulator, in both a layer-autonomous and non-autonomous manner. Lgl2 plays a critical role in the formation of hemidesmosomes, which mediate basal cell-to-matrix adhesion through integrin α6β437. Since hemidesmosomes bind primarily to laminin, we currently do not know whether hemidesmosomes play a role in maintaining E-cadherin basolateral polarity in the periphery. However, hemidesmosomes have not formed at 2 dpf37. In mice, intratissue tension generated by the upper epidermal layers through microtubule regulation significantly impacts the behavior of BECs. Increased contractility in these differentiated cells, which can result from microtubule disruption, promotes stem cell hyperproliferation while inhibiting their migration. This highlights that stem cells in stratified tissues respond not only to substrate rigidity but also to mechanical cues from intratissue tension, with differentiated cells playing a critical role in shaping stem cell behavior and maintaining tissue integrity10.
Overall, our findings reinforce existing evidence of intratissue communication and propose a new paradigm for embryonic epidermal stem cells. We propose that periderm near limb appendages have enhanced wound healing properties based on regionally-specific differences in the underlying stem cell-matrix interactions. Possibly by secreting distinct ECMs, and regulating adhesion molecule expression, epidermal stem cells regulate the protective function of the interconnected, differentiated tissue, the periderm. This discovery holds significant implications for our understanding of BEC contributions during epidermal development, and can inform future applications such as tissue engineering and regenerative medicines, including organ repair and skin transplants.
Methods
Zebrafish Husbandry
Zebrafish (Danio rerio) were raised and maintained at 28.5 °C using standard methods and ethical handling according to protocol approved by the Yale University Institutional Animal Care and Use Committee (#2020-11473). AB strain zebrafish (ZDB-GENO-960809-7) were used as wild-type genetic background at larval stages where sex is unspecified. Gonad development is not specified until 13-25 dpf. Therefore, larvae used in this study have no sex influences and sex was not considered. The following zebrafish transgenic lines have been previously described: TgBAC(ΔNP63:Gal4ff)la213 (ZDB-ALT-150424-4)68, Tg(krt5:Gal4ff)la212 (ZDB-ALT-150424-3)68, Tg(UAS:NLS-GFP)el609 (ZDB-ALT-210709-6)69, Tg(krt5:LifeAct-Ruby)la227 (ZDB-ALT-200226-18)38, Tg(UAS:LifeAct-GFP)mu271 (ZDB-ALT-130624-2)70, TgBAC(pdgfrb:eGFP)ncv22Tg (ZDB-ALT-160609-1)71, Tg(Bactin:HRAS-EGFP)vu119 (ZDB-ALT-061107-2)72, and TgBAC(lamC1:lamC1-sfGFP)sk116 (ZDB-TGCONSTRCT-241010-2)31. The following endogenous knock-in lines have been previously described: cdh1-tdTomatoxt18 (ZDB-ALT-190419-2)73 and itgb1b:itgb1b-sfGFPsk133 (ZDB-ALT-241011-7)31. The zebrafish mutants used in this study have been previously described: itgb1ask109 (ZDB-ALT-240116-1) and itgb1bsk110 (ZDB-ALT-240116-2)31. The lama5ya377, col1a1aya392, and dspaya391 mutant lines were generated in this study.
Generation and genotyping of mutant alleles
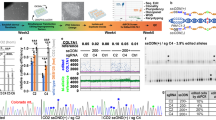
To generate lama5 ya37, col1a1a ya392, and dspa ya391 mutants, we followed previously described CRISPR-Cas9-based gene-editing protocols74. CRISPRScan (https://www.crisprscan.org/) was used to design guide RNAs (gRNAs) to generate loss-of-function mutant of lama5. gRNA preparation was performed as previously described75. Briefly, WT embryos were injected with 100 pg of gRNAs and 200 pg of Cas9 mRNA at the one-cell stage. PCR genome amplification and T7E1 assay was used to validate indels as previously75.
The lama5ya377 mutant contains 23-bp deletion in lama5 exon2 that leads to a pre-mature stop codon in exon3. lama5ya377 homozygous mutant embryos display complete loss of the fin fold, which is consistent with previous reports25. Sequences and primers are listed in supplementary Table 1. To genotype lama5ya377, zebrafish embryos and adults were genotyped using GeneMarker (v2.4.0) as previously described76, with genotyping primers listed in supplementary Table 1.
The col1a1aya392 mutant contains 5-bp deletion in col1a1a exon2 that leads to a pre-mature stop codon in exon3. col1a1aya377 homozygous mutant embryos display damaged fin fold, which is consistent with previous reports47. Sequences and primers are listed in supplementary Table 1. To genotype col1a1aya392, zebrafish embryos and adults were genotyped using GeneMarker (v2.4.0) as previously described76, with genotyping primers listed in supplementary Table 1.
The dspaya391 mutant contains 5-bp deletion in dspa exon2 that leads to a pre-mature stop codon in exon3. Sequences and primers are listed in supplementary Table 1. To genotype dspaya391, zebrafish embryos and adults were genotyped using GeneMarker (v2.4.0) as previously described76, with genotyping primers listed in supplementary Table 1.
To genotype itgb1ask109 and itgb1bsk110, zebrafish embryos and adults were genotyped by PCR as previously described31, with genotyping primers listed in supplementary Table 1.
Zebrafish Embryos Immunofluorescence Staining
Immunofluorescence
Immunofluorescence (IF) was performed as follows for all zebrafish embryos. After overnight 4% paraformaldehyde (Santa Cruz) fixation at 4˚C, 48 hpf embryos were washed with 1X PBS-0.01%Tween-20 (PBSTw) four to five times for 5 minutes, and then permeabilized with 0.125% trypsin (Millipore Sigma T4549) for 5 minutes. Embryos were washed in blocking solution (0.8% Triton-X, 10% normal goat serum, 1% BSA, 0.01% sodium azide in PBSTw) four times for 5 minutes, followed by an additional two-hour incubation in blocking solution with shaking at room temperature. Primary and secondary antibodies incubation were conducted overnight at 4 °C. Following each overnight antibody incubation, six washes for a total of 4 h were performed with washing buffer (1% Triton-X in PBSTw) at room temperature. Primary antibodies used were specific to RFP (1:250, Antibodies Online, ABIN129578), Laminin α1 (1:250, Sigma-Aldrich, L9393), phosphorylated Myosin Light Chain (Ser19) (1:200, Cell Signaling, Cat. #3675), Desmoplakin (undiluted, ProGen, 651155), and Vinculin (1:100, Sigma-Aldrich, V9131). Secondary antibodies for either rabbit (1:250, ThermoFisher, A10040) or mouse (1:250, ThermoFisher, A21202) were used. To stain nuclei, DAPI (1:500, ThermoFisher, D1306) was added during the secondary antibody incubation step. To stain F-actin, Phalloidin-568 (1:100, Invitrogen, A12380) was added during the primary antibody incubation step. Samples were mounted in 1% Low Range Ultra Agarose (Bio-Rad, #1613106) and imaged using the Zeiss LSM980 upright confocal microscope (ZEN 3.4, blue edition, Carl Zeiss) equipped with a water-immersed x10/0,3 or 20/0,5 or x40/1,0 objective lens.
Collagen Hybridizing Peptide
To stain denatured collagen, Collagen hybridizing peptide with 5-FAM or Cy3 conjugate (5 µM, F-CHP/ R-CHP, 3Helix, FLU60/ RED60) was used. After overnight 4% paraformaldehyde (Santa Cruz) fixation at 4˚C, 48 hpf embryos were washed with 1X PBS-0.01%Tween-20 (PBSTw) four to five times for 5 minutes, and then permeabilized with 0.125% trypsin (Millipore Sigma T4549) for 5 minutes. Embryos were washed in blocking solution (0.8% Triton-X, 10% normal goat serum, 1% BSA, 0.01% sodium azide in PBSTw) four times for 5 minutes, followed by an additional two-hour incubation in blocking solution with shaking at room temperature. The CHP stock solution (50 µM) was diluted in PBS buffer to make the working solution (5 µM) and heat-activate by incubating at 80 °C for 5 minutes. The solution was immediately cooled on ice for 30–90 s, to quench it to room temperature, then quickly applied to fixed embryos. Laminin α1 (1:250, Sigma-Aldrich, L9393) primary antibody was added to the diluted CHP solution before applying to the samples. Samples were incubated with the staining solution at 4 °C overnight with shaking. Following each overnight antibody incubation, six washes for a total of 4 h were performed with washing buffer (1% Triton-X in 1x PBSTw) at room temperature.
Cell immunostaining
For immunofluorescence analysis in NHEKs, cells were fixed with 4% paraformaldehyde (Electron Microscopy Sciences) for 20 minutes. Cells were washed with DPBS with calcium and magnesium (Thermo, 21300025) and permeabilized in 0.5% Triton X-100 for 10 minutes. Cells were blocked with 10% serum for 1 h, incubated for either 1 h at room temperature or overnight at 4 °C with primary antibodies, and incubated for 1 h with secondary antibodies. Primary antibodies used were specific to p63 (1:100, BioCare Medical, CM163A), β-catenin (1:500, Sigma, C2206), Desmoplakin (undiluted, ProGen, 651155), Human Collagen Type I (1:50, CedarLane, CL50111AP-1), Collagen Type IV (1:100, ThermoFisher, 19674-1-AP), and Laminin (1:500, ThermoFisher, PA5-143904). Secondary antibodies for either rabbit (1:1000, ThermoFisher, A-32731 or A-21244), mouse (1:1000, ThermoFisher, A-11001 or A-21235), or Rat (1:1000, ThermoFisher, A-11006 or A-21247) were applied. To stain F-actin, phalloidin (1:400, ThermoFisher, A12380) was added during the secondary antibody incubation step. Samples were mounted with DAPI in Fluoromount-G (Thermo, 00-4959-52) and imaged on Zeiss LSM980 Airyscan 2 confocal.
Confocal Live and Time Lapse Imaging of Zebrafish Embryos
Zebrafish embryos were treated with 0.003% 1-phenyl-2-thiourea (PTU, MilliPore Sigma, P7629) starting at 70%/80% gastrulation stage to prevent pigmentation. Embryos imaged live by confocal microscopy were anaesthetized in 0.1% tricaine and mounted in 1% (w/v) Low Range Ultra Agarose (Bio-Rad, #1613106) at 48hpf. Embryos with high transgenic integration were selected with a potential caveat that p63 and krt5 Life-Act drivers do display higher cell-to-cell variability. Confocal live imaging was performed using the Zeiss LSM980 upright confocal microscope (ZEN 3.4, blue edition, Carl Zeiss) equipped with a water-immersed x10/0,3 or 20/0,5 or x40/1,0 objective lens. Z-stacks were acquired at 0.5 µm increments. Confocal time-lapse movies were recorded for wild type embryos in Tg(krt5:Gal4ff )la212, Tg(UAS:LifeAct-GFP)mu271; cdh1-tdTomatoxt18 background, at 28oC incubation stating at 48 hpf. Z-stacks were acquired at 2 µm increments with intervals of every 10 min for a total duration of 8 h. For each confocal imaging session, the laser power was consistently set to the same level across all samples. Cells were tracked using dragon-tail analysis in Imaris software (V.9.9.1, Bitplane).
Two-photon single-cell laser ablation and time lapse imaging
Single cell laser ablation of superficial epidermal cells (SECs) was performed using a Leica Stellaris 8 DIVE multiphoton/confocal microscope equipped with a Spectra Physics MaiTai laser at 2 dpf. Wild type and lama5 +/- embryos, in the Tg(krt5:Gal4ff)la212, Tg(UAS:LifeAct-GFP)mu271 background, were anaesthetized in 0.1% tricaine and mounted in 1% (w/v) Low Range Ultra Agarose (Bio-Rad, #1613106). Prior to ablation, confocal z-stacks were set up at 2 µm increments to capture pre-ablation images. Laser ablation was performed in a 10 µm diameter region of interest with the MaiTai laser at a wavelength of 810 nm with the laser power at 70% for two iterations of 0.743 seconds. Confocal z-stack images were acquired at 1-minute intervals until wound closure.
Cell culture
Normal human epidermal keratinocytes (NHEKs) were purchased from Promo Cell (C-12001) from a 4-year-old Caucasian male donor (Lot #: 494Z030.1). Cells were screened by manufacturer for bacteria, yeast, fungi, mycoplasma, virus and authenticated for positive expression of cytokeratin via flow cytometry. NHEKs were cultured at 37 °C, 5% CO2, and 95% relative humidity according to manufacturer’s instructions up to passage four. Unless otherwise specified, cells were cultured in Keratinocyte Growth Medium 2 (PromoCell, C-20111) which contains no serum, low CaCl2 (0.06 mM), and supplemented with Gentamicin (Thermo, 15710064) at a final concentration of 5 μg/mL.
Production of collagen and laminin gels
Collagen and laminin gels were formed on 35 mm glass-bottom dishes (MatTek, P35G-0-20-C) by first coating dishes with 1.8 mg/mL polydopamine coating solution made from dopamine-HCL (Acros Organics, AC122000100) in 10 mM Tris-HCL (pH 8.5). Dopamine solution was rinsed 3 times with ddH2O, the coated glass was fully dried in a warm oven overnight and cooled to room temperature the next day. To make the neutralized collagen solution (2 mg/mL final concentration of collagen), sterile ddH2O, 10x PBS (Corning, 46-013-CM), Collagen (Corning, 354249), and NaOH were mixed together. All solutions and tubes used for making gels were pre-chilled on ice. Next, 23 uL of the neutralized collagen was dispensed onto the dopamine-HCL coated glass and an autoclaved 8 mm round coverslip (Electron Microscopy Sciences, 7229608) was placed on top of the droplet to ensure even coating. The collagen solution was allowed to solidify in an incubator at 37 C for 40 minutes. After full gelation, the gels were soaked with PBS for 15 minutes and the top coverslip was removed. To make the laminin coated collagen gel, collagen gels were created as described above. A 50%, ~5 mg/mL laminin (Corning, 354259, 10 mg/mL) solution was made in ice-cold PBS and 20 uLs were evenly dispensed on top of the collagen cushion. The laminin layer was allowed to cure at 37 C for 20 minutes. After curing, gels were inspected using a phase microscope to ensure the laminin layer adhered to the collagen gel. Gels were stored in PBS until seeding.
NHEK stratification on collagen and laminin gels
NHEKs were expanded in T-75 flasks (ThermoFisher, 156499) and lifted into a single cell suspension using Trypsin-EDTA (0.05%) with phenol red (ThermoFisher, 25300054). To obtain confluent monolayers on gels within 24 h of seeding, 800,000 cells were seeded into each dish. Cells were seeded and grown overnight in Keratinocyte Growth Medium 2 with low CaCl2 (0.06 mM). 24 h after initial seeding, media was changed to Keratinocyte Growth Medium 2 supplemented with high CaCl2 (1.2 mM). Cells were grown for 48 more hours with a media change every 24 h. Cells were then fixed in 4% paraformaldehyde (Electron Microscopy Sciences) for 20 minutes before proceeding with immunofluorescence staining.
Image analysis and quantifications
Zebrafish confocal fluorescence images
Confocal fluorescence images were analyzed with Imaris software (V.9.9.1, Bitplane, Oxford Instruments) and the Fiji version of ImageJ (NIH)77. P63 expression across the fin fold was quantified using live confocal images of TgBAC(ΔNP63:Gal4ff)la213; Tg(UAS:NLS-GFP)el609. Each basal epidermal stem cell nucleus (GFP + ) was 3D-reconstructed using Imaris surface module based on fluorescence intensity. Mean intensity and x,y,z coordinates (µm) for each BEC nucleus were exported for statistical tests. Distance from the center for each BEC nucleus was calculated using the x,y,z coordinates, and mean intensity was normalized to each embryo’s average intensity.
Confocal z-stack images were taken from the SEC layer to the BEC layer. SECs and BECs were identified by maximum projection of selected z slices based on their z-position. Intralayer junctional fluorescence intensities for E-cadherin, phosphorylated Myosin Light Chain (pMyosin), Desmoplakin, and Vinculin were quantified by drawing segmented lines along junctions and measuring mean gray value. pMyosin and Vinculin junctional intensities were measured using E-cadherin as a landmark in a separate channel. Interlayer E-cadherin and Desmoplakin intensities were quantified in orthogonal slices by drawing segmented lines between the two epidermal layers and measuring the mean gray value for each SEC-BEC interface. Collagen and Laminin fluorescence intensities were quantified by drawing a fixed-size rectangle region of interest starting at the edge of the fin fold, followed by the plot profile function in ImageJ. The region of interest was kept consistent within each experiment. Average intensities of Integrin β1b and BEC actin levels were measured by drawing regions of interest outlining individual epidermal cells using the polygon selection tool in ImageJ. SEC actin microridge length was quantified using ImageJ as previously described38. Briefly, cell outlines were traced using the polygon selection tool manually to measure apical cell area. Brightness and contrast were automatically enhanced, and surrounding area was cleared. Images were blurred with the Smoothen function three times and processed through a Laplacian morphological filter from the MorphoLibJ plugin78 (square element, radius = 1). Filtered images were automatically thresholded using the Triangle method and skeletonized. Microridge length was measured using the Analyze Skeleton (2D/3D) feature.
Cell culture confocal fluorescence images
Confocal fluorescence images were analyzed with the Fiji version of ImageJ (NIH)77. Confocal z-stack images were taken from the suprabasal layer to the basal layer. Suprabasal and basal keratinocytes were identified by maximum projection of selected z slices based on their z-position. Intralayer junctional fluorescence intensity for β-catenin and Desmoplakin was quantified by drawing segmented lines along junctions and measuring mean gray value. Interlayer β-catenin and Desmoplakin intensity were quantified by measuring mean gray value of regions of interest in single z slices at the suprabasal-basal keratinocytes interface. Interlayer Desmoplakin plaque area was quantified by first thresholding based on intensity, followed by the analyze particles function in ImageJ. P63 intensity in suprabasal and basal layers was quantified by measuring the mean gray value of regions of interest created using intensity-thresholded nuclei in maximum projection images of each keratinocyte layer.
Dragon-tail analysis using Imaris software
Epithelial cell migration in the time lapse movies of wild type fin fold development was analyzed using Imaris spot module with manual spot tracing function. Migration distance (Track Length) was visualized as dragon tails. The average speed, track length, and duration of tracks were exported for statistical analysis.
Calculation for mean squared displacement (MSD)
Movement profiles, time and x,y position, for individual cells in the fin fold were used to calculate a mean squared displacement (MSD) value as performed previously79. MSD values are defined as follows:
where \({x}_{i}\) and \({y}_{i}\) are the coordinates of the ith cell and t represents the time interval.
Transcriptome analysis
Transcriptomic data were extracted from Cokus et al. 21. In brief, distinct epithelial cell populations were FACS sorted, quality checked, and prepared with Illumina TruSeq Library. One library was used per replicate, at a sequencing depth of 21.2-45.4 million reads pass filter (PF). CutAdapt 1.8.1 was used for quality control, and reads were aligned to Ensembl release 92 Danio rerio GRCz11 using STAR 2.5.3a. Library distributions were performed with Salmon 0.10.2. Analyses were performed using DeSeq2, using a variation of Sleuth analysis described in Cokus et al. 21 Supplementary File 1.
Enrichment plots for collagen and laminin expression in different cell types were extracted from DanioCell28, an online database of single-cell transcriptomic data across different developmental stages. In brief, cell-type specific clusters were first identified using cell-type specific markers (tp63 for BEC and pdgfrb for Mesenchymal cells). Expression patterns of different ECM proteins were then extracted in the respective clusters.
Statistics and reproducibility
All statistical analyses were performed using Prism version 10.4.1 (Graphpad). Measurements were taken from distinct samples. Sample size and statistical tests are specified in each figure legend. All sample size were determined using power analysis test. For all data sets, outliers were first identified and removed using the ROUT method in prism. Then, data sets were tested for Gaussian distribution using the D’Agostino-Pearson omnibus normality test (significance level = 0.05). For normally distributed datasets, unpaired t-test (two-sided) was used to test mean differences between two groups and ordinary one-way ANOVA with Tukey’s multiple comparisons test was used to test mean differences among multiple groups. For data sets that are not normally distributed, Mann-Whitney test (two-sided) was used for comparison between two groups and Kruskal-Wallis with Dunn’s multiple comparisons test was used for comparisons among multiple groups.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Fish lines are available upon request. The raw data generated in this study are available in the Source Data file provided with this paper, and the Supplementary Information. All data used are available through previously published results in NCBI GEO series GSE132304 and DanioCell (https://daniocell.nichd.nih.gov/) at NCBI GEO series GSE223922. Source data are provided with this paper.
Code availability
No code was generated in this study.
References
Morgner, J. et al. Integrin-linked kinase regulates the niche of quiescent epidermal stem cells. Nat. Commun. 6, 8198 (2015).
Lu, P., Weaver, V. M. & Werb, Z. The extracellular matrix: a dynamic niche in cancer progression. J. Cell Biol. 196, 395–406 (2012).
Gjorevski, N. et al. Designer matrices for intestinal stem cell and organoid culture. Nature 539, 560–564 (2016).
Fuchs, E. & Blau, H. M. Tissue stem cells: architects of their niches. Cell Stem Cell 27, 532–556 (2020).
Watt, F. M. & Hogan, B. L. Out of Eden: stem cells and their niches. Science 287, 1427–1430 (2000).
Charras, G. & Yap, A. S. Tensile forces and mechanotransduction at cell-cell junctions. Curr. Biol. 28, R445–R457 (2018).
Delva, E., Tucker, D. K. & Kowalczyk, A. P. The desmosome. Cold Spring Harb. Perspect. Biol. 1, a002543 (2009).
Song, X., Zhu, C. H., Doan, C. & Xie, T. Germline stem cells anchored by adherens junctions in the Drosophila ovary niches. Science 296, 1855–1857 (2002).
Zhang, J. et al. Identification of the haematopoietic stem cell niche and control of the niche size. Nature 425, 836–841 (2003).
Ning, W., Muroyama, A., Li, H. & Lechler, T. Differentiated daughter cells regulate stem cell proliferation and fate through intra-tissue tension. Cell Stem Cell 28, 436–452 e435 (2021).
Bansaccal, N. et al. The extracellular matrix dictates regional competence for tumour initiation. Nature 623, 828–835 (2023).
Zijl, S., Salameti, V., Louis, B., Negri, V. A. & Watt, F. M. Dynamic regulation of human epidermal differentiation by adhesive and mechanical forces. Curr. Top. Dev. Biol. 150, 129–148 (2022).
De Falco, M., Pisano, M. M. & De Luca, A. in Skin Cancer: A Practical Approach. (eds. Alfonso Baldi, Paola Pasquali & Enrico P. Spugnini) 1–15 (Springer New York, 2014).
McGrath, J. A., Eady, R. & Pope, F. Anatomy and organization of human skin. Rook’s. Textb. Dermatol. 1, 3.2–3.80 (2004).
Eckhart, L., Holthaus, K. B. & Sachslehner, A. P. Cell differentiation in the embryonic periderm and in scaffolding epithelia of skin appendages. Dev. Biol. 515, 60–66 (2024).
Le Guellec, D., Morvan-Dubois, G. & Sire, J. Y. Skin development in bony fish with particular emphasis on collagen deposition in the dermis of the zebrafish (Danio rerio). Int J. Dev. Biol. 48, 217–231 (2004).
Lee, R. T. H., Asharani, P. V. & Carney, T. J. Basal keratinocytes contribute to all strata of the adult zebrafish epidermis. PLOS ONE 9, e84858 (2014).
Guzman, A., Ramos-Balderas, J. L., Carrillo-Rosas, S. & Maldonado, E. A stem cell proliferation burst forms new layers of P63 expressing suprabasal cells during zebrafish postembryonic epidermal development. Biol. Open 2, 1179–1186 (2013).
Richardson, R. J. et al. Periderm prevents pathological epithelial adhesions during embryogenesis. J. Clin. Invest 124, 3891–3900 (2014).
Kennard, A. S., Sathe, M., Labuz, E. C., Prinz, C. K. & Theriot, J. A. Post-injury hydraulic fracturing drives fissure formation in the zebrafish basal epidermal cell layer. Curr. Biol. 33, 2616–2631 e2615 (2023).
Cokus, S. J. et al. Tissue-specific transcriptomes reveal gene expression trajectories in two maturing skin epithelial layers in zebrafish embryos. G3 (Bethesda) 9, 3439–3452 (2019).
Sonawane, M. et al. Zebrafish penner/lethal giant larvae 2 functions in hemidesmosome formation, maintenance of cellular morphology and growth regulation in the developing basal epidermis. Development 132, 3255–3265 (2005).
Liu, Z., Meng, Y., Ishikura, A. & Kawakami, A. Live tracking of basal stem cells of the epidermis during growth, homeostasis and injury response in zebrafish. Development 151, https://doi.org/10.1242/dev.202315 (2024).
Weaver, J. How fins became limbs. Nature https://doi.org/10.1038/news.2010.315 (2010).
Webb, A. E. et al. Laminin alpha5 is essential for the formation of the zebrafish fins. Dev. Biol. 311, 369–382 (2007).
Duran, I., Mari-Beffa, M., Santamaria, J. A., Becerra, J. & Santos-Ruiz, L. Actinotrichia collagens and their role in fin formation. Dev. Biol. 354, 160–172 (2011).
Tsutsui, K. et al. Mapping the molecular and structural specialization of the skin basement membrane for inter-tissue interactions. Nat. Commun. 12, 2577 (2021).
Sur, A. et al. Single-cell analysis of shared signatures and transcriptional diversity during zebrafish development. Developmental Cell 58, 3028–3047.e3012 (2023).
Wyatt, R. A. & Crawford, B. D. Post-translational activation of Mmp2 correlates with patterns of active collagen degradation during the development of the zebrafish tail. Dev. Biol. 477, 155–163 (2021).
Ma, R. C., Kocha, K. M., Mendez-Olivos, E. E., Ruel, T. D. & Huang, P. Origin and diversification of fibroblasts from the sclerotome in zebrafish. Dev. Biol. 498, 35–48 (2023).
Yamaguchi, N. et al. Rear traction forces drive adherent tissue migration in vivo. Nat. Cell Biol. 24, 194–204 (2022).
Lenne, P. F., Rupprecht, J. F. & Viasnoff, V. Cell junction mechanics beyond the bounds of adhesion and tension. Dev. Cell 56, 202–212 (2021).
Arora, P., Dongre, S., Raman, R. & Sonawane, M. Stepwise polarisation of developing bilayered epidermis is mediated by aPKC and E-cadherin in zebrafish. Elife https://doi.org/10.7554/eLife.49064 (2020).
Bays, J. L. et al. Vinculin phosphorylation differentially regulates mechanotransduction at cell-cell and cell-matrix adhesions. J. Cell Biol. 205, 251–263 (2014).
van den Goor, L., Iseler, J., Koning, K. M. & Miller, A. L. Mechanosensitive recruitment of Vinculin maintains junction integrity and barrier function at epithelial tricellular junctions. Curr. Biol. 34, 4677–4691 e4675 (2024).
Peltonen, S., Raiko, L. & Peltonen, J. Desmosomes in developing human epidermis. Dermatol Res Pr. 2010, 698761 (2010).
Sonawane, M., Martin-Maischein, H. & Schwarz, H. & Nusslein-Volhard, C. Lgl2 and E-cadherin act antagonistically to regulate hemidesmosome formation during epidermal development in zebrafish. Development 136, 1231–1240 (2009).
van Loon, A. P., Erofeev, I. S., Maryshev, I. V., Goryachev, A. B. & Sagasti, A. Cortical contraction drives the 3D patterning of epithelial cell surfaces. J. Cell Biol.https://doi.org/10.1083/jcb.201904144 (2020).
Hynes, R. O. Integrins: bidirectional, allosteric signaling machines. Cell 110, 673–687 (2002).
Jülich, D. et al. Cross-scale integrin regulation organizes ECM and tissue topology. Developmental Cell 34, 33–44 (2015).
Yue, J., Zhang, K. & Chen, J. Role of integrins in regulating proteases to mediate extracellular matrix remodeling. Cancer Microenviron. 5, 275–283 (2012).
Deyrieux, A. F., Rosas-Acosta, G., Ozbun, M. A. & Wilson, V. G. Sumoylation dynamics during keratinocyte differentiation. J. Cell Sci. 120, 125–136 (2007).
Arnette, C., Koetsier, J. L., Hoover, P., Getsios, S. & Green, K. J. In vitro model of the epidermis: connecting protein function to 3D structure. Methods Enzymol. 569, 287–308 (2016).
Heenen, M., De Graef, C. & Galand, P. Kinetics of the calcium induced stratification of human keratinocytes in vitro. Cell Prolif. 25, 233–240 (1992).
Bikle, D. D., Xie, Z. & Tu, C. L. Calcium regulation of keratinocyte differentiation. Expert Rev. Endocrinol. Metab. 7, 461–472 (2012).
Huber, A. H. & Weis, W. I. The structure of the beta-catenin/E-cadherin complex and the molecular basis of diverse ligand recognition by beta-catenin. Cell 105, 391–402 (2001).
Gistelinck, C. et al. Zebrafish type I collagen mutants faithfully recapitulate human type I collagenopathies. Proc. Natl. Acad. Sci. USA 115, E8037–E8046 (2018).
Celeghin, R. et al. A novel DSP zebrafish model reveals training- and drug-induced modulation of arrhythmogenic cardiomyopathy phenotypes. Cell Death Discov. 9, 441 (2023).
Chen, S. S., Fitzgerald, W., Zimmerberg, J., Kleinman, H. K. & Margolis, L. Cell-cell and cell-extracellular matrix interactions regulate embryonic stem cell differentiation. Stem Cells 25, 553–561 (2007).
Guilak, F. et al. Control of stem cell fate by physical interactions with the extracellular matrix. Cell Stem Cell 5, 17–26 (2009).
Pinho, S. & Frenette, P. S. Haematopoietic stem cell activity and interactions with the niche. Nat. Rev. Mol. Cell Biol. 20, 303–320 (2019).
Watt, F. M. & Fujiwara, H. Cell-extracellular matrix interactions in normal and diseased skin. Cold Spring Harb. Perspect Biol. https://doi.org/10.1101/cshperspect.a005124 (2011).
De Belly, H., Paluch, E. K. & Chalut, K. J. Interplay between mechanics and signalling in regulating cell fate. Nat. Rev. Mol. Cell Biol. 23, 465–480 (2022).
Biggs, L. C., Kim, C. S., Miroshnikova, Y. A. & Wickstrom, S. A. Mechanical forces in the skin: roles in tissue architecture, stability, and function. J. Invest Dermatol 140, 284–290 (2020).
Ajeti, V. et al. Wound healing coordinates actin architectures to regulate mechanical work. Nat. Phys. 15, 696–705 (2019).
Staddon, M. F. et al. Cooperation of dual modes of cell motility promotes epithelial stress relaxation to accelerate wound healing. PLoS Comput Biol. 14, e1006502 (2018).
Brock, J., Midwinter, K., Lewis, J. & Martin, P. Healing of incisional wounds in the embryonic chick wing bud: characterization of the actin purse-string and demonstration of a requirement for Rho activation. J. Cell Biol. 135, 1097–1107 (1996).
Price, A. J. et al. Mechanical loading of desmosomes depends on the magnitude and orientation of external stress. Nat. Commun. 9, 5284 (2018).
Thomas, M., Ladoux, B. & Toyama, Y. Desmosomal junctions govern tissue integrity and actomyosin contractility in apoptotic cell extrusion. Curr. Biol. 30, 682–690 e685 (2020).
Sadhanasatish, T. et al. A molecular optomechanics approach reveals functional relevance of force transduction across talin and desmoplakin. Sci. Adv. 9, eadg3347 (2023).
Baddam, S. R. et al. The desmosomal cadherin desmoglein-2 experiences mechanical tension as demonstrated by a FRET-based tension biosensor expressed in living cells. Cells https://doi.org/10.3390/cells7070066 (2018).
Broussard, J. A. et al. Scaling up single-cell mechanics to multicellular tissues - the role of the intermediate filament-desmosome network. J. Cell Sci. https://doi.org/10.1242/jcs.228031 (2020).
Ha, J. et al. Intermediate cells of in vitro cellular reprogramming and in vivo tissue regeneration require desmoplakin. Sci. Adv. 8, eabk1239 (2022).
Ton, A. T., MacKeith, A. K., Shattuck, M. D. & O’Hern, C. S. Mechanical plasticity of cell membranes enhances epithelial wound closure. Phys. Rev. Res. 6, L012036 (2024).
Chen, C., Jiang, Z. & Yang, G. Laminins in osteogenic differentiation and pluripotency maintenance. Differentiation 114, 13–19 (2020).
Klees, R. F. et al. Laminin-5 induces osteogenic gene expression in human mesenchymal stem cells through an ERK-dependent pathway. Mol. Biol. Cell 16, 881–890 (2005).
Becerra, J., Montes, G. S., Bexiga, S. R. & Junqueira, L. C. Structure of the tail fin in teleosts. Cell Tissue Res 230, 127–137 (1983).
Rasmussen, J. P., Sack, G. S., Martin, S. M. & Sagasti, A. Vertebrate epidermal cells are broad-specificity phagocytes that clear sensory axon debris. J. Neurosci. 35, 559 (2015).
Barske, L. et al. Evolution of vertebrate gill covers via shifts in an ancient Pou3f3 enhancer. Proc. Natl. Acad. Sci. 117, 24876–24884 (2020).
Helker, C. S. M. et al. The zebrafish common cardinal veins develop by a novel mechanism: lumen ensheathment. Development 140, 2776–2786 (2013).
Ando, K. et al. Clarification of mural cell coverage of vascular endothelial cells by live imaging of zebrafish. Development 143, 1328–1339 (2016).
Dale, R. M. & Topczewski, J. Identification of an evolutionarily conserved regulatory element of the zebrafish col2a1a gene. Developmental Biol. 357, 518–531 (2011).
Cronan, M. R. et al. An explant technique for high-resolution imaging and manipulation of mycobacterial granulomas. Nat. Methods 15, 1098–1107 (2018).
Ghersi, J. J. et al. Haematopoietic stem and progenitor cell heterogeneity is inherited from the embryonic endothelium. Nat. Cell Biol. 25, 1135–1145 (2023).
Kasper, D. M. et al. The N-glycome regulates the endothelial-to-hematopoietic transition. Science 370, 1186–1191 (2020).
Kasper, D. M. et al. MicroRNAs establish uniform traits during the architecture of vertebrate embryos. Developmental Cell 40, 552–565.e555 (2017).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Legland, D., Arganda-Carreras, I. & Andrey, P. MorphoLibJ: integrated library and plugins for mathematical morphology with ImageJ. Bioinformatics 32, 3532–3534 (2016).
Haga, H., Irahara, C., Kobayashi, R., Nakagaki, T. & Kawabata, K. Collective movement of epithelial cells on a collagen gel substrate. Biophys. J. 88, 2250–2256 (2005).
Acknowledgements
We would like to thank Nicole Semanchik and Jared Hintzen for zebrafish husbandry and technical assistance with this project. We are also grateful to Robert Lalonde and Andrew Prendergast of the Yale Zebrafish Research Core for their technical service with CRISPR/Cas9 line genotype. We thank Valentina Greco, Scott Holley, and Anjelica Gonzalez at Yale University for their valuable feedback and suggestions during the development of the project. We also acknowledge Angela Andersen at Life Science Editors for editorial assistance. S.N. discloses support for this work from NHLBI R21HL165342, NHLBI P01HL169168, and AHA award 957692. K.S. discloses support from NIGMS R35GM150645. M.M. acknowledges support from the NIGMS R35GM142875.
Author information
Authors and Affiliations
Contributions
H.M.H., L.C.B., K.S., and S.N. conceptualized and supervised the study. H.M.H. performed and analyzed the zebrafish experiments. L.C.B. performed and analyzed the 3D keratinocyte model experiments in collaboration with X.G. and M.M. J.M.B., S.L.I., and Z.W. contributed to experimental execution and data analysis. H.M.H., M.C., and S.N. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
He, H.M., Boraas, L.C., Bell, J.M. et al. Epidermal stem cells control periderm injury repair via matrix-driven specialization of intercellular junctions. Nat Commun 16, 8967 (2025). https://doi.org/10.1038/s41467-025-64040-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64040-7