Abstract
Cancer immunotherapy using dendritic cells (DC) pulsed ex vivo with tumour antigens is considered safe, but its clinical efficacy is generally modest. Here we engineer DC progenitors (DCP), which can replenish conventional type 1 DCs (cDC1) in mice, to constitutively express IL-12 together with a non-signalling chimeric receptor, termed extracellular vesicle-internalizing receptor (EVIR). By binding to a bait molecule (GD2 disialoganglioside) expressed on cancer cells and their EVs, the EVIR enforces EV internalization by cDC1 to promote their cross-dressing with preformed, tumour-derived MHCI-peptide complexes. Upon systemic deployment to mice, the engineered DCPs cause only mild and transient elevation of liver enzymes, acquire tumour-derived material, engage tumour-specific T cells, and enhance the efficacy of PD-1 blockade in an immunotherapy-resistant melanoma model comprising both GD2-positive and -negative cancer cells, without the need for ex vivo antigen pulsing. These results indicate that EVIR-engineered DCPs may avert the positive selection of antigen-negative cancer cells, potentially addressing a critical limitation of immunotherapies targeting defined tumour antigens.
Similar content being viewed by others
Introduction
Dendritic cells (DC) are myeloid cells that can be grouped, based on their ontogeny and phenotypes, into distinct subsets: conventional dendritic cells (cDC) and plasmacytoid dendritic cells, which largely arise from rare bone marrow (BM) progenitors, as well as monocyte-derived dendritic cells (moDC), which derive from circulating monocytes1,2,3,4. DCs serve as primary antigen-presenting cells that capture, process, and present tumour antigens to T cells, thereby initiating and regulating adaptive immune responses essential for effective anti-tumour activity4,5,6,7. Due to their critical role in modulating the adaptive immune response against tumours, DC therapies have been extensively tested in clinical trials8,9,10,11.
In the traditional DC therapy workflow, moDCs are generated from circulating monocytes, exposed to defined tumour antigens or tumour lysates ex vivo, and activated by means of adjuvants or cytokines prior to injection into patients with cancer1,11,12,13. While such moDC therapies have clearly demonstrated safety and immunogenicity, they have often failed to achieve robust and durable clinical responses in the majority of the treated patients10,14. The limited clinical efficacy of traditional moDC-based therapies is mainly attributed to suboptimal steps in the manufacturing process and the inherent biological attributes of moDCs1. Indeed, ex vivo manufactured moDCs may lack efficient antigen-presentation and migration capabilities. Additionally, the ex vivo antigen loading process may not recapitulate the spectrum of antigenic heterogeneity between independent tumour lesions in patients with metastatic disease or the variability in tumour antigens expressed in tumours of different patients1.
Exploiting the notion that tumour-derived extracellular vesicles (EV) and membrane particles are a source of tumour-associated antigens and neoantigens15,16,17,18, we previously developed a non-signaling chimeric receptor, termed EV-internalizing receptor (EVIR), specifically designed to enforce the uptake and internalization by DCs of tumour-derived EVs/membrane particles displaying a known, tumour-specific “bait” molecule19. One such bait molecule is GD2, a disialoganglioside often over-expressed in tumours of neuroectodermal origin, including melanoma, neuroblastoma, and glioma20. We showed that an anti-GD2 (αGD2) EVIR facilitates the internalization of GD2+ cancer-cell–derived EVs by moDCs and the presentation of EV-associated tumour antigens (called “prey” antigens) to T cells in cell culture assays19. In this context, antigen presentation primarily involved cross-dressing19, the acquisition by the moDCs of preformed peptide-major histocompatibility class I (p-MHCI) complexes from the cancer cells via the direct uptake of tumour-derived EVs or the horizontal transfer of plasma membrane fragments from cancer cells (trogocytosis)21,22,23.
In an effort to develop DC platforms alternative to moDCs, we generated DC progenitors (DCP) that efficiently differentiated into bona fide cDCs upon their transfer to mice24. cDCs, especially type 1 cDCs (cDC1), play key functions in anti-tumour immunity4,25,26,27,28,29. Here, we further engineer the DCPs to co-express interleukin-12 (IL-12) and an αGD2 EVIR. We show that, compared with unmodified cells, the engineered DCPs promote robust immune responses and inhibit the growth of experimental tumours that are otherwise resistant to programmed cell death protein 1 (PD-1) blockade, including melanomas with heterogeneous GD2 expression, without the need for ex vivo antigen loading. These preclinical results emphasize the potential of using gene-modified DCs to improve the efficacy of immunotherapy in patients with cancer.
Results
An αGD2 EVIR enhances tumour antigen uptake by intra-tumoural cDCs
The EVIR facilitates the selective uptake of cancer-cell–derived EVs by DCs in cell culture assays, thereby promoting presentation of tumour antigens19 (Fig. 1a). The EVIR contains an extracellular antibody domain (scFv) directed against GD2 along with a transmembrane and a truncated (non-signaling) intracellular domain derived from the low-affinity nerve growth factor receptor (dLNGFR)19. To investigate the role of the EVIR in the internalization of tumour antigens by myeloid antigen-presenting cells in mice, we generated a transgenic mouse line expressing an αGD2 EVIR specifically in CD11c+ cells (CD11c-EVIR hereon), which comprise cDCs as well subpopulations of monocytes and macrophages30. In CD11c-EVIR mice, the EVIR was expressed in the majority of the cDCs and in a fraction of both Ly6Chigh and Ly6Clow monocytes/macrophages (Supplementary Fig. 1a).
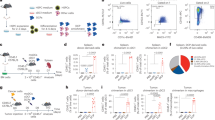
a Schematic illustration of EVIR-mediated uptake and presentation of tumour-associated antigens by DCs. b Procedure to study EVIR-mediated internalization of tumour-derived EVs/particles in CD11c-EVIR transgenic mice (both female and male mice). c Internalization of mCh+ tumour-derived EVs/particles by cDCs, Ly6Chigh and Ly6Clow monocytes/macrophages, and other immune cells (either myeloid or non-myeloid) in tumours of CD11c-EVIR mice (mean ± s.e.m.; n = 1 for MC38-GD2 and n = 10 for MC38-GD2/CD9-mCh). Statistical analysis by two-tailed Mann-Whitney test, comparing only EVIR⁻ and EVIR⁺ cells in mCh⁺ tumours. d Procedure to study the internalization of mCh+ tumour-derived EVs/particles by DCP-EVIR or DCP-CtrlR injected in the tumours of Rag1–/– mice (both female and male mice). DCPs were generated from CD45.1 mice and transferred into CD45.2 recipient mice. e Left: flow cytometry dot plots showing mCh internalization by DCP-derived cells in tumours of CD45.2 mice. Right: quantification of the data (mean ± s.e.m.; n = 1 for B16F10-OVA/GD2; n = 3 for DCP-CtrlR and n = 5 for DCP-EVIR in B16F10-OVA/GD2/CD9-mCh tumour-bearing mice). Statistical analysis by two-tailed Mann-Whitney test comparing DCP-CtrlR and DCP-EVIR in B16F10-OVA/GD2/CD9-mCh tumour-bearing mice. Each data point represents one tumour from an independent mouse. Source data are provided as a Source Data file.
Next, we subcutaneously inoculated CD11c-EVIR mice with MC38 colorectal cancer cells modified to co-express GD2 and a CD9-mCherry fusion protein enabling the tracking of cancer cell-derived EVs/particles (MC38-GD2/CD9-mCh cells; Fig. 1b). Flow cytometry analysis revealed enhanced uptake of mCh+ tumour-derived EVs/particles by EVIR+ (dLNGFR+) myeloid cells, including cDCs and both Ly6Chigh and Ly6Clow monocytes/macrophages, compared to EVIR– myeloid cells (Fig. 1c, Supplementary Fig. 1a). Of note, EVIR-negative non-myeloid immune cells did not appreciably internalize mCh+ tumour-derived EVs/particles. These data indicate that enforced expression of the EVIR enhances the intrinsic ability of myeloid cells to internalize tumour-derived EVs/particles in mice.
Ex vivo-generated DCPs have been shown to efficiently produce professional cDCs in tumours upon systemic delivery to mice24. Therefore, we reasoned that EVIR-expressing DCPs would give rise to cDCs with improved capacity to internalize tumour-derived EVs/particles. To test this hypothesis, we used lentiviral vectors (LV) to transduce DCPs with the αGD2 EVIR (DCP-EVIR) or a control receptor lacking the αGD2 scFv (DCP-CtrlR hereon), both generated from the BM of CD45.1 mice. For these studies, we employed B16F10 melanoma cells modified to express ovalbumin (OVA) and GD2, with or without CD9-mCh (B16F10-OVA/GD2/CD9-mCh and B16F10-OVA/GD2, respectively). We then injected transduced CD45.1+ DCPs in the tumours of CD45.2 Rag1–/– recipient mice (Fig. 1d). Expression of the EVIR and CtrlR, measured by staining dLNGFR (a protein domain present in both receptors), was similar in tumour-infiltrating CD45.1+ cells (Supplementary Fig. 1b). However, only the CD45.1+dLNGFR+ cells derived from DCP-EVIR cells efficiently acquired mCh in the majority of the tumours, whereas DCP-CtrlR–derived cells displayed lower mCh fluorescence (Fig. 1e). These results demonstrate that EVIR expression enhances the uptake of tumour-derived EVs/particles by intra-tumoural DCP-derived cells, potentially facilitating antigen presentation and anti-tumour immune responses.
ɑGD2 DCP-IL-12/EVIR unleash ɑPD-1 efficacy in B16F10-OVA/GD2 melanoma
We previously showed that DCPs engineered to express IL-12 exhibit co-stimulatory activity toward T cells in tumour-bearing mice24. Therefore, we designed LVs to co-express IL-12 with either EVIR or CtrlR in DCPs (DCP-IL-12/EVIR and DCP-IL-12/CtrlR, respectively). To explore the therapeutic potential of the engineered DCPs, we used the B16F10-OVA/GD2 melanoma model, which is resistant to PD-1 blockade despite OVA expression24,31,32. Mice bearing established subcutaneous tumours (mean tumour volume at day 6: 36 mm3) were infused intravenously with transduced DCPs or vehicle, in combination with ɑPD-1 antibodies to unleash the effector functions of the T cells (Fig. 2a). Tumours were analyzed 8 days after the last DCP infusion. In this setting, DCP-IL-12/EVIR significantly inhibited tumour growth compared to both vehicle (PBS) and DCP-IL-12/CtrlR (Fig. 2b). Additionally, flow cytometry analysis of the tumours revealed IL-12–dependent promotion of CD8+ T cell infiltration and EVIR-dependent enhancement of their activation, which may have cooperatively contributed to the observed anti-tumoural response (Fig. 2c). All gating strategies used in flow cytometry analyses are shown in Supplementary Figs. 2–6.
a Procedure to study engineered DCPs plus ɑPD-1 in female tumour-bearing mice. b Volume of B16F10-OVA/GD2 tumours (mean ± s.e.m.; n = 9). Statistical analysis by two-way ANOVA with Tukey’s multiple comparison test. c Frequency of the indicated immune cells in B16F10-OVA/GD2 tumours (mean ± s.e.m.; n = 9). Statistical analysis by one-way ANOVA with Tukey’s multiple comparison test. d Procedure to study engineered DCPs plus ɑPD-1 in female tumour-bearing mice. e Volume of B16F10-OVA/GD2 tumours (mean ± s.e.m.; DCP-IL-12/CtrlR + IgG, n = 10, except n = 9 at the last timepoint; other groups, n = 10). Statistical analysis by two-way ANOVA with Tukey’s multiple comparison test. f Procedure to study the effects of CD8+ T cell depletion in female tumour-bearing mice. g Frequency of CD8β+ T cells in B16F10-OVA/GD2 tumours (mean ± s.e.m.; DCP-IL-12/EVIR + ɑPD-1, n = 6; other groups, n = 8). Data points in brown, also indicated by an arrow, identify mice with unsuccessful CD8+ T cell depletion. The mice in the PBS group were terminated on day 14, whereas those in the other groups were terminated on day 19 (separated by a dotted line). Statistical analysis by one-way ANOVA with Tukey’s multiple comparison test. h Volume of B16F10-OVA/GD2 tumours (mean ± s.e.m.; PBS, n = 8; DCP-IL-12/EVIR + ɑPD-1/CD8a, n = 6; DCP-IL-12/EVIR + ɑPD-1, n = 7). Statistical analysis by two-way ANOVA with Tukey’s multiple comparison test (black p value, comparing all groups until day 14) or Sidak’s multiple comparison test (red p value, comparing DCP-IL-12/EVIR + ɑPD-1 with DCP-IL-12/EVIR + ɑPD-1/ɑCD8a until day 19). The mice with suboptimal CD8+ T cell depletion (see g) were excluded from the analysis. i Frequency of the indicated macrophage states in the tumours (mean ± s.e.m.; DCP-IL-12/EVIR + ɑPD-1, n = 6; other groups, n = 8). Statistical analysis by one-way ANOVA with Tukey’s multiple comparison test. j Pie charts showing the hematopoietic-cell composition of the tumours. The area of each cell population corresponds to the mean percentage value in the total CD45+ cells (DCP-IL-12/EVIR + ɑPD-1, n = 6; other groups, n = 8). Each data point in (c, g, i) represents one tumour from an independent mouse. Each data point in (b, e, h) represents the mean volume of independent tumours. Source data are provided as a Source Data file.
We next examined the contribution of ɑPD-1 to the therapeutic response. To this end, we treated mice bearing early-established B16F10-OVA/GD2 melanomas with either DCP-IL-12/EVIR or DCP-IL-12/CtrlR, with or without ɑPD-1, and monitored the tumours for 14 days after the last DCP infusion (Fig. 2d). Consistent with the previous findings, mice treated with DCP-IL-12/EVIR plus ɑPD-1 exhibited slower tumour growth compared to the other groups (Fig. 2e). Notably, ɑPD-1 therapy did not improve the tumour response to DCP-IL-12/CtrlR, suggesting that expression of the EVIR by DCPs specifically sensitized tumours to PD-1 blockade. In this setting, as compared to the previous experiment, we observed EVIR-dependent enhancement of CD8+ T cell activation only in a fraction of the tumours, possibly due to the longer interval (14 versus 8 days) between the last DCP infusion and tumour analysis (Supplementary Fig. 7a, b).
To investigate the potential involvement of CD8+ T cells in tumour response to combined DCP-IL-12/EVIR plus ɑPD-1 therapy, we used anti-CD8 antibodies to deplete CD8+ T cells in B16F10-OVA/GD2 tumour-bearing mice (Fig. 2f). Efficient elimination of intra-tumoural CD8+ T cells in the majority of the mice (Fig. 2g) moderately rescued tumour growth (Fig. 2h), implicating CD8+ T cells in the anti-tumoural response. Immune profiling of the tumours indicated that DCP-IL-12/EVIR plus ɑPD-1 increased the relative proportions of CD8+ T cells and M1-like (inflammatory) macrophages while decreasing M2-like (pro-tumoural) macrophages33, compared to tumours of vehicle plus ɑPD-1-treated mice (Fig. 2i, j). Notably, CD8+ T cell depletion did not abate DCP-induced M1-like macrophages, potentially explaining why tumour growth was only partially rescued. These data suggest that, beyond promoting CD8+ T cell infiltration and activation, DCP-IL-12/EVIR plus ɑPD-1 skews macrophages toward an anti-tumoural, M1-like phenotype.
IL-12/EVIR-engineered DCPs but not moDCs sensitize B16F10 melanomas to PD-1 blockade
We then asked whether EVIR-expressing moDCs could operationally substitute for DCPs in sensitizing tumours to ɑPD-1 therapy. We used moDC-IL-12/EVIR or control moDC-IL-12/CtrlR, produced as described previously24, and combined them with ɑPD-1 treatment in B16F10-OVA/GD2 tumour-bearing mice.
In a first experiment, mice bearing early-established tumours were treated with moDCs plus ɑPD-1 and were monitored for 14 days after the last moDC infusion (Fig. 3a). In this setting, expression of the EVIR in moDCs failed to further sensitize tumours to PD-1 blockade (Fig. 3b), at variance with results obtained with engineered DCPs (compare with Fig. 2e). This suggests that specific properties of DCPs, likely involving their capacity to efficiently engraft and differentiate into cDCs in mice24, may be critical for enabling EVIR-mediated enhancement of PD-1 blockade in immunotherapy-resistant tumours.
a Procedure to study the combination of engineered moDCs and ɑPD-1 therapy in female tumour-bearing mice. b Volume of B16F10-OVA/GD2 tumours (mean ± s.e.m.; moDC-IL-12/EVIR + ɑPD-1, n = 3; other groups, n = 6). Statistical analysis by two-way ANOVA with Tukey’s multiple comparison test, calculated until day 17. c Procedure to study the tumour response to engineered DCPs and moDCs in female tumour-bearing mice. d Volume of B16F10-OVA/GD2 tumours (mean ± s.e.m.; moDC-IL-12/EVIR + ɑPD-1, n = 3; DCP-IL-12/EVIR + ɑPD-1, n = 7). Statistical analysis by two-way ANOVA with Sidak’s multiple comparison test. e Frequency of indicated immune cell types in B16F10-OVA/GD2 tumours (mean ± s.e.m.; moDC-IL-12/EVIR + ɑPD-1, n = 3; DCP-IL-12/EVIR + ɑPD-1, n = 7). Statistical analysis by two-tailed Mann-Whitney test. f Intracellular IFNγ, GZMB and TNFα in ex vivo-restimulated, CD8+ T cells (mean ± s.e.m.; moDC-IL-12/EVIR + ɑPD-1, n = 3; DCP-IL-12/EVIR + ɑPD-1, n = 7). Statistical analysis by two-tailed Mann-Whitney test. g Procedure to study the combination of engineered DCPs and ɑPD-1 in mice with established MC38-HER2 tumours in WAP transgenic mice (both female and male mice). h Volume of MC38-HER2 tumours (mean ± s.e.m.; PBS + ɑPD-1, n = 7; other groups, n = 8). Statistical analysis by two-way ANOVA with Tukey’s multiple comparison test. Each data point in all panels represents one tumour from an independent mouse except in (b, d, h) in which each data point represents the mean volume of independent tumours. Source data are provided as a Source Data file.
In a second study, we directly compared the therapeutic efficacy of DCPs and moDCs (both with ɑPD-1) in mice bearing established tumours (mean tumour volume at day 7: 49 mm3) (Fig. 3c). In this setting, DCP-IL-12/EVIR achieved more effective tumour control (Fig. 3d) and increased the intra-tumoural infiltration of activated CD8+ (PD-1+TIM3–, IFNγ+, GZMB+, or TNFα+) and CD4+ (IFNγ+) T cells, compared to moDC-IL-12/EVIR (Fig. 3e, f, Supplementary Fig. 8a, b). These findings suggest that engineered DCPs may outperform moDCs in inhibiting tumour growth in an immunotherapy-resistant melanoma model.
DCPs engineered to express an ɑHER2 EVIR and IL-12 improve tumour control in a colorectal cancer model
To test the versatility of our approach, we investigated DCPs engineered to express an anti-human HER2 EVIR. To this aim, we inoculated WAP transgenic mice, which are partly tolerant to human HER234, with MC38-HER2 colon cancer cells19 and treated them with DCP-IL-12/EVIR plus ɑPD-1 (Fig. 3g). DCP-IL-12/EVIR plus ɑPD-1 treatment significantly reduced tumour growth compared to control treatments (Fig. 3h, Supplementary Fig. 8c). The majority of the tumours in mice that received DCP-IL-12/EVIR plus ɑPD-1 showed a reduction in tumour volume at the analysis endpoint (day 15) relative to day 6, when the treatment was initiated (Supplementary Fig. 8d). These results indicate that the therapeutic potential of EVIR-engineered DCPs extends to distinct bait molecules in different tumour models.
DCP-EVIR activate CD8⁺ T cells through antigen cross-dressing
To demonstrate that EVIR-mediated antigen presentation involves DC cross-dressing with preformed p-MHCI complexes19, we generated E0771 mammary tumour cells co-expressing human HER2 as bait molecule relevant to breast cancer and a truncated OVA (tOVA) transgene as surrogate prey antigen (E0771-tOVA/HER2 cells). Unlike the full-length OVA, tOVA lacks the leader sequence required for its extracellular secretion (thus removing potentially confounding effects mediated by secreted OVA) while retaining the ability to generate the OVA-derived, H2Kb/MHCI-restricted SIINFEKL peptide35. Accordingly, E0771-tOVA/HER2 cells displayed the SIINFEKL peptide in association with MHCI/H2Kb (SIINFEKL-H2Kb) on the cell surface (Supplementary Fig. 9a). We also generated β2m knockout (MHCI-deficient) E0771-tOVA/HER2 cells using CRISPR/Cas9 (E0771-tOVA/HER2β2m-KO cells). As expected, these cells did not display SIINFEKL-H2Kb on their surface (Supplementary Fig. 9b).
We next generated mouse BM-derived cDCs comprising both cDC1 and cDC2 (Supplementary Fig. 9c), which are the main differentiation product of DCPs upon their adoptive transfer to mice24. We transduced the cDCs with the EVIR and co-cultured them with E0771-tOVA/HER2 or E0771-tOVA/HER2β2m-KO cells in the presence of recombinant IL-12 and OT-I CD8⁺ T cells, whose TCR specifically recognizes SIINFEKL-H2Kb36. Notably, cDC-EVIR stimulated greater IFNγ production by OT-I CD8⁺ T cells following co-culture with E0771-tOVA/HER2 cells than with E0771-tOVA/HER2β2m-KO cells (Supplementary. Fig. 9d, e). These results suggest that EVIR-enhanced T cell activation was dependent on the acquisition by cDC-EVIR of pre-formed SIINFEKL-H2Kb from the cancer cells, indicative of cross-dressing. Conversely, cDC-EVIR failed to activate OT-II CD4⁺ T cells, whose TCR specifically recognizes the MHCII-restricted OVA peptide ISQAVHAAHAEINEAGR37, in co-culture with E0771-tOVA/HER2 cells (Supplementary. Fig. 9f, g). This was likely because E0771 cells do not express MHCII and, therefore, cannot cross-dress DCs with MHCII-restricted antigens.
Taken together, these data indicate that enhancement of cDC cross-dressing with pre-formed p-MHCI complexes is a mechanism by which EVIR-DCs may stimulate antigen-specific CD8⁺ T cell responses.
αGD2 DCP-EVIR inhibit the growth of bait molecule-negative cancer cells in molecularly heterogenous tumours
The studies described above employed tumours virtually uniformly expressing the bait molecule (i.e., GD2 or HER2) targeted by the EVIR. We then asked whether EVIR-mediated anti-tumour immunity would also extend to bait molecule-negative cancer cells. To this aim, we tested DCP-IL-12/EVIR in mice carrying B16F10 melanomas with heterogenous expression of GD2 and OVA.
In a set or preparative studies aimed to validate the mixed tumour model, we used DCPs engineered to endogenously express and present non-secreted tOVA, with or without IL-12 (DCP-IL-12/tOVA and DCP-tOVA, respectively; Supplementary. Fig. 10a). In tumour-free mice, DCP-tOVA and to a greater extent DCP-IL-12/tOVA, elicited robust OVA-specific T cells in the spleen (Supplementary Fig. 10b). DCP-IL-12/tOVA fully blocked the growth of B16F10-OVA melanomas in a prophylactic setting, a response that was associated with greatly expanded OVA-specific T cells in both blood and spleen (Supplementary Fig. 10c–e). In addition, DCP-IL-12/tOVA robustly controlled B16F10-OVA melanoma growth and elicited OVA-specific T cells also in a therapeutic intervention setting (Supplementary Fig. 10f–h). These data demonstrate that DCPs can elicit robust antigen-specific T cell responses in mice.
We then compared DCP-IL-12/tOVA, which endogenously express and present OVA to directly elicit OVA-specific CD8+ T cells, with DCP-IL-12/EVIR, which can only indirectly elicit antigen-specific T cells—potentially also against OVA—in mice carrying tumours derived from a mixture of B16F10-OVA/GD2 and parental (OVA- and GD2-negative) B16F10 cells (3:1 ratio) (Fig. 4a). Interestingly, DCP-IL-12/EVIR plus ɑPD-1 inhibited the growth and delayed relapse of tumours originating from mixed B16F10-OVA/GD2 and B16F10 cancer cells more efficiently than DCP-IL-12/tOVA plus ɑPD-1 (Fig. 4b, Supplementary Fig. 10i), even though neither OVA nor other tumour antigens were directly targeted. As expected, flow cytometry analysis showed that DCP-IL-12/tOVA plus ɑPD-1 robustly elicited OVA-specific effector T cells in both tumour-draining lymph nodes (tdLNs) and spleen (Fig. 4c, d). Interestingly, DCP-IL-12/EVIR plus ɑPD-1 also induced a small but statistically significant increase in OVA-specific effector T cells relative to ɑPD-1 alone in both tdLNs and spleen. Overall, DCP-IL-12/EVIR elicited more activated (CD44+CD62L–), non-OVA–specific CD8+ T cells than DCP-IL-12/tOVA in both tdLNs and spleen, likely reflecting elicitation of a broader immune response.
a Procedure to study the efficacy of engineered DCPs in female mice bearing tumours with mixed bait antigen-positive and negative cancer cells. b Volume of tumours derived from the injection of mixed B16F10-OVA/GD2 and B16F10 cancer cells (mean ± s.e.m.; PBS + ɑPD-1, n = 5; DCP-IL-12/tOVA + ɑPD-1, n = 9; DCP-IL-12/EVIR + ɑPD-1, n = 8). Statistical analysis by two-way ANOVA with Tukey’s multiple comparison test (black p value, comparing all groups until day 13) or two-tailed Sidak’s multiple comparison test (red p value, comparing DCP-IL-12/tOVA + ɑPD-1 and DCP-IL-12/EVIR + ɑPD-1 until day 19). c, d Frequency of the indicated immune cell types in tdLN (c, mean ± s.e.m.; PBS + ɑPD-1, n = 5; DCP-IL-12/tOVA + ɑPD-1, n = 6; DCP-IL-12/EVIR + ɑPD-1, n = 7) or spleen (d mean ± s.e.m.; PBS + ɑPD-1, n = 5; other groups, n = 8). The dotted line separates tumours analyzed at day 13 (left) and 20 (right). Statistical analysis by one-way ANOVA with Tukey’s multiple comparison test (black p values) or two-tailed Mann-Whitney test (red p values). e Left: Immunofluorescence staining images of tumours originating from mixed B16F10-OVA/GD2 and B16F10 melanoma cells in mice treated as indicated (DAPI nuclear staining: blue; anti-GD2 staining: yellow), analyzed at the experimental endpoints (days 13 and 20). Size-matched tumours are shown. Scale bar, 1 mm. Right: quantification of the data (mean ± s.e.m.; PBS + ɑPD-1, n = 5; other groups, n = 7). Statistical analysis by two-tailed Mann-Whitney test. Each data point in all panels represents one sample from an independent mouse except in (b), in which each data point represents the mean volume of independent tumours. Source data are provided as a Source Data file.
In the context of treatment with DCP-IL-12/tOVA, tumour relapse was likely sustained by the outgrowth of OVA-negative B16F10 cancer cells. Accordingly, immunofluorescence staining of the tumours revealed that virtually all B16F10-OVA/GD2 cancer cells were eliminated in mice treated with DCP-IL-12/tOVA plus ɑPD-1, which contrasted with results in mice treated with DCP-IL-12/EVIR + ɑPD-1, in which (OVA+)GD2+ cancer cells were only in part negatively selected (Fig. 4e). Collectively, these results indicate that the anti-tumour response elicited by EVIR-engineered DCPs is likely multi-antigen–specific.
αGD2 DCP-IL-12/EVIR do not induce liver or kidney toxicity in mice
To evaluate the impact of DCP-IL-12/EVIR on commonly evaluated biomarkers of organ function, we used 8-week–old male and female C57BL/6J mice. The mice were intravenously injected with either PBS or DCP-IL-12/EVIR and monitored over a 36-day period (Fig. 5a). Liver function was evaluated by measuring serum aspartate aminotransferase (AST) and alanine aminotransferase (ALT), whereas kidney function was evaluated by measuring creatinine levels. These parameters were assessed both at day 4, to capture early effects, and day 36, the study endpoint.
a Procedure to study liver and kidney toxicity following systemic DCP-IL-12/EVIR inoculation in tumour-free male and female mice. The experiment was performed once. b Serum creatinine at day 4 post-DCP injection (mean ± s.e.m.; n = 5 male or female mice). c Serum AST and ALT at day 4 post-DCP injection (mean ± s.e.m.; n = 5 male or female mice, except for MCS and MCD, n = 6 male mice). The mice in the MCS and MCD groups originated from an independent experiment (separated by a dotted line). Statistical analysis by two-tailed Mann-Whitney test comparing mice with same sex. d Serum AST and ALT at day 4 and 36 post-injection (mean ± s.e.m.; n = 5 male or female mice). Statistical analysis by repeated measures two-way ANOVA with Sidak’s correction for multiple comparisons (comparing mice with same sex). e Representative H&E-stained kidney sections of female mice from the indicated treatment groups. Scale bar: 0.1 mm. f, Representative H&E-stained liver sections of male or female mice from the indicated treatment groups. Arrows indicate areas of mild inflammatory infiltration in the liver. Scale bar: 0.1 mm. g Frequency of male or female mice (n = 5) exhibiting histological features of inflammation, steatosis, ballooning, or fibrosis in liver sections. Each data point in all panels represents one sample from an independent mouse except in (g), in which values represent the percentage of mice per group showing the indicated pathology grade. Source data are provided as a Source Data file.
At day 4, creatinine levels were similar between groups (Fig. 5b). However, we observed mild elevation in AST and ALT in DCP-treated mice compared to PBS-treated controls (Fig. 5c). Notably, liver enzyme levels in all DCP-treated mice were below 265 IU/mL, substantially lower than those observed in a group of male C57BL/6N mice fed a methionine- and choline-deficient (MCD) diet for 7 weeks in an independent experiment. By day 36, AST and ALT levels had decreased in all DCP-treated mice (Fig. 5d), suggesting recovery of liver function over time.
Hematoxylin/eosin (H&E) staining of day-36 kidney sections showed no histopathological abnormalities (Fig. 5e), consistent with the non-elevated creatinine levels observed in serum analyses. Histological analysis of liver sections by H&E staining revealed mild (grade 1) inflammation and steatosis, without evidence of hepatocyte ballooning or fibrosis, in some of the mice and independent of DCP treatment (Fig. 5f, g). Together, these findings suggest that systemic administration of DCP-IL-12/EVIR only induces mild and transient liver toxicity without significant renal or long-term tissue damage.
Human DCP-IL-12/EVIR promote antigen internalization and presentation
To explore the functionality of the αGD2 EVIR in human DCPs, we generated human DCPs from cord blood (CB)-derived CD34+ cells (Fig. 6a) according to a previously developed protocol24. In parallel, we generated human moDCs from peripheral blood mononuclear cells, as described24. Both DCPs and moDCs were transduced with LVs encoding the control receptor (CtrlR), EVIR, or both human IL-12 (hIL-12) and EVIR. Before use, the DCPs were further differentiated into DCP-derived cells encompassing cDCs and other DC-lineage cells (termed DCP progeny24). As shown in Fig. 6b, c, both CtrlR and EVIR were efficiently expressed in the transduced DCP progeny and moDCs.
a Procedure to produce and differentiate engineered human DCP progeny from CD34+ cord-blood cells. b, c Representative dot plots showing the expression of CtrlR and EVIR in human DCP progeny and moDCs. d Schematic representation of the assay used to study EV internalization by transduced human DCP progeny and moDCs. PKH26 mean fluorescence intensity (MFI) in transduced human DCP progeny (e) or human moDCs (f) measured by flow cytometry from two independent donors and two biological replicates for each (mean ± s.e.m.). g Schematic representation of the assay used to study antigen cross-dressing by transduced human DCP progeny or human moDCs. Antigen presentation by human DCP progeny (h) or human moDCs (i) engineered to express CtrlR, EVIR or hIL-12/EVIR (mean ± s.e.m.; DCP-CtrlR, n = 3; EVIR and hIL-12/EVIR, n = 2; moDC-CtrlR, n = 3; EVIR and hIL-12/EVIR, n = 2 biological replicates from a single donor).Each data point in all panels represents biological replicates from one or two independent donors. Source data are provided as a Source Data file.
We next assessed EVIR-mediated antigen uptake using an EV internalization assay (Fig. 6d). Briefly, we isolated EVs from GD2+ or GD2– human melanoma cell lines38,39, labeled them with PKH26 fluorescent dye, and incubated them with transduced human DCP progeny or moDCs generated from two independent donors. Flow cytometry analysis showed that EVIR-expressing cells exhibited higher PKH26 mean fluorescence intensity (MFI) after incubation with GD2+ EVs, indicating enhanced internalization of labeled EVs, compared to CtrlR-expressing cells or EVIR-expressing cells incubated with GD2– EVs (Fig. 6e, f).
Finally, we evaluated cross-dressing–mediated antigen presentation by the human DCP progeny and moDCs. EVs isolated from either GD2+HLA-A2+, GD2+HLA-A2–, or GD2–HLA-A2+ human melanoma cell lines38,39 were incubated with the CMV/pp65495-504 peptide—a procedure that results in the direct loading of the peptide onto MHCI molecules on the surface of the EVs – and then transferred to transduced DCP progeny or moDCs, both isolated from an HLA-A2− donor. The cells were subsequently cocultured with HLA-A2-restricted, CMV/pp65495-504-specific human CD8+ T cells40 (Fig. 6g). Flow cytometry analysis of IFNγ in CD8+ T cells indicated that co-expression of the EVIR and hIL-12 by the DCP progeny—and, to a lesser extent, moDCs—increased the average magnitude of the T cell response in a GD2 and HLA-A2–dependent manner, demonstrating cross-dressing conducive to T cell stimulation (Fig. 6h, i). Collectively, these results demonstrate that dual EVIR and hIL-12 expression enhances antigen internalization and presentation by human DCs.
Discussion
Here we show that DCPs modified to co-express an ɑGD2 EVIR and IL-12 induce CD8+ T cell activation and unlock the efficacy of PD-1 blockade in mice challenged with immunotherapy-resistant B16F10 melanomas. The engineered DCPs also controlled the growth of non-targeted, bait molecule-negative cancer cells in a tumour model containing both bait molecule-positive and negative cancer cells. This strategy relies on EVIR-mediated tumour antigen uptake by DCP-derived cells and their release of IL-12, a cytokine critical for conditioning the tumour microenvironment and promoting effective anti-tumour immune responses41,42.
In our study, moDCs engineered to co-express the EVIR and IL-12 showed more limited efficacy than DCPs and apparently failed to sensitize tumours to PD-1 blockade. This result may be attributed to several factors. First, the EVIR lacks signaling domains and, unlike traditional chimeric antigen receptors (CAR)43, does not induce cell activation upon binding to the targeted bait molecule. Second, we have previously shown that moDCs exhibit lower engraftment compared to DCPs24, which may limit both the duration of antigen uptake and presentation and the magnitude and temporal window of local IL-12 release. In contrast, DCPs efficiently engraft in the tumour and mainly differentiate into cDC1, a cDC subset involved in priming or boosting tumour-reactive CD8+ T cells44,45,46. Notably, cDC1-derived IL-12 is biologically relevant, as it more efficiently engages T and NK cells in the anti-tumour response compared to IL-12 secreted by other myeloid cells42,47. Overall, our data suggest that DCPs may offer advantages over moDCs in terms of both antigen presentation and IL-12-mediated T cell activation, which are paramount for robust anti-tumour immunity.
Our engineered DCP-based platform harnesses the EVIR to promote tumour antigen uptake through a mechanism known as cross-dressing, which involves the acquisition of preformed, tumour-derived MHCI-bound antigens from tumour-derived membrane particles19,22. This facilitates the presentation by the DC of immunologically relevant tumour antigens, i.e., those effectively displayed by cancer cells. Unlike ex vivo antigen loading, which relies on known antigens or tumour biopsies that may not fully capture tumour heterogeneity and antigen immunogenicity1, the EVIR enforces the acquisition and presentation of MHC-I loaded tumour antigens in vivo, thereby promoting an immune response potentially directed against multiple tumour antigens, including those displayed on bait molecule-negative cancer cells.
Although the precise mechanisms underlying the anti-tumour response induced by DCPs expressing the EVIR and IL-12 remain to be fully identified, our results provide several insights. We show that EVIR expression by DCPs contributes to eliciting T cell responses in combination with IL-12. This is illustrated, for example, by the increased numbers of activated, intra-tumoural CD8+ T cells, which may mediate the tumour response to PD-1 blockade48. Indeed, depletion of CD8+ T cells partially rescued tumour growth in DCP-treated mice, implicating CD8+ T cells in the tumour response. In contrast, depleting CD8+ T cells in mice treated with DCPs engineered to co-express FLT3L and IL-12 – but not the EVIR – did not impair the anti-tumoural response24, suggesting a specific role for the EVIR in the cross-priming of CD8+ T cells. Additionally, DCP-derived IL-12 reprograms the tumour microenvironment by activating NK cells, enhancing IFNγ production, and polarizing macrophages toward an anti-tumoural, M1-like phenotype24. Given the roles of various immune cells and their bioactive mediators in orchestrating anti-tumour immunity, further studies will be required to identify the cellular and molecular underpinnings of the tumour response mediated by EVIR-engineered DCPs. While DCP-IL-12/EVIR elicited OVA-specific T cells in the spleen of treated mice without the need for ex vivo OVA protein pulsing, it remains to be seen whether the engineered DCPs alter the number and composition of T cell clonotypes in the periphery and tumours of treated mice.
The EVIR is a non-signaling CAR. The use of CAR-expressing myeloid cells has been extensively tested in solid tumours. Macrophages have been the primary focus due to some of their inherent features, including their ability to potentially infiltrate solid tumours49,50 and phagocytose cancer cells51,52,53,54. However, early clinical studies with CAR macrophages have delivered largely disappointing results55. CARs have also been used to modify other myeloid cells, such as neutrophils, mainly for directing cytotoxicity56 and drug delivery57 to the tumour site. In contrast, our strategy uses engineered DCPs to enforce tumour antigen acquisition in vivo and amplify potentially broad and diverse immune effector mechanisms. This approach relies on the bait molecule targeted by the EVIR (e.g., GD2) to facilitate tumour EV/particle uptake but not to specifically elicit direct cancer cell killing or phagocytosis. This strategy is less likely to induce immune escape through downregulation of the bait molecule or clonal selection of bait molecule-negative cells, and may result in the destruction of both bait molecule-positive and -negative cancer cells.
The EV internalization capacity of EVIR-engineered human DCPs supports the clinical potential of our platform. Our preclinical studies suggest mild and transient toxicity of DCP-IL-12/EVIR in mice, which is consistent with the short-lived engraftment of the engineered cells24. To further mitigate potential safety concerns associated with constitutive IL-12 expression, non-integrating gene delivery approaches such as mRNA electroporation or lipid nanoparticle (LNP)-mediated mRNA delivery58 may allow for transient cytokine expression. Human DCP-IL-12/EVIR may be conveniently and efficiently produced from mobilized peripheral blood CD34+ cells for autologous cell therapy applications. Alternatively, to facilitate scalable manufacturing of engineered DCPs, induced pluripotent stem cell (iPSC)-derived DCPs could be used for standardized, large-scale production, while the development of allogeneic “off-the-shelf” DCPs combined with genome editing strategies, such as HLA knockout or suppression59, may further broaden clinical applicability. Notably, expression of the EVIR, which facilitates DC cross-dressing with tumour-derived p-MHCI complexes19, may surrogate for the absence of endogenous MHCI molecules in the engineered DCs. Together, these approaches position DCP-EVIR as a promising platform for the development of next-generation DC-based cancer immunotherapies.
Methods
Ethical approvals
Experiments with mice were performed at the EPFL or Agora animal facilities in Lausanne, Switzerland. All studies were conducted in accordance with the guidelines and regulations set by the Cantonal Veterinary Office of Vaud, Switzerland, following authorizations VD3154, VD3154.1, VD3785 and VD3792. αGD2 EVIR transgenic mice were generated at the Netherlands Cancer Institute in accordance with the guidelines of the Animal Ethics Committee of the Netherlands Cancer Institute; protocols were approved by the Dutch national Centrale Commissie Dierproeven (AVD301002016683).
Human cord blood-derived CD34⁺ cells were obtained from STEMCELL Technologies, which collected the samples under informed consent and approved protocols. Human peripheral blood samples from healthy donors were collected with ethical approval from the Blood Transfusion Center (University of Lausanne, Switzerland) under Project P_297.
Design and construction of lentiviral vectors (LV)
We used previously described self-inactivating, third-generation LV constructs to express various transgenes, including GD2 synthase (GD2S)19,24, GD3 synthase (GD23)19,24, human HER219, a CD9-mCherry fusion protein19, CtrlR19,24, EVIR19, or bicistronic IL-12-P2A-CtrlR24.
We also constructed new LVs to express tOVA or a bicistronic IL-12-P2A-tOVA. We used a truncated, non-secreted OVA variant by removing the leader signal:
tOVA (MHCI- and MHCII-restricted peptides are indicated in bold):
MDSTRTQINKVVRFDKLPGFGDSIEAQCGTSVNVHSSLRDILNQITKPNDVYSFSLASRLYAEERYPILPEYLQCVKELYRGGLEPINFQTAADQARELINSWVESQTNGIIRNVLQPSSVDSQTAMVLVNAIVFKGLWEKAFKDEDTQAMPFRVTEQESKPVQMMYQIGLFRVASMASEKMKILELPFASGTMSMLVLLPDEVSGLEQLESIINFEKLTEWTSSNVMEERKIKVYLPRMKMEEKYNLTSVLMAMGITDVFSSSANLSGISSAESLKISQAVHAAHAEINEAGREVVGSAEAGVDAASVSEEFRADHPFLFCIKHIATNAVLFFGRCVSPGGG.
IL-12-P2A-tOVA (MHCI- and MHCII-restricted peptides are indicated in bold):
MCPQKLTISWFAIVLLVSPLMAMWELEKDVYVVEVDWTPDAPGETVNLTCDTPEEDDITWTSDQRHGVIGSGKTLTITVKEFLDAGQYTCHKGGETLSHSHLLLHKKENGIWSTEILKNFKNKTFLKCEAPNYSGRFTCSWLVQRNMDLKFNIKSSSSSPDSRAVTCGMASLSAEKVTLDQRDYEKYSVSCQEDVTCPTAEETLPIELALEARQQNKYENYSTSFFIRDIIKPDPPKNLQMKPLKNSQVEVSWEYPDSWSTPHSYFSLKFFVRIQRKKEKMKETEEGCNQKGAFLVEKTSTEVQCKGGNVCVQAQDRYYNSSCSKWACVPCRVRSGGGGSGGGGSGGGLASGGSMVSVPTASPSASSSSSQCRSSMCQSRYLLFLATLALLNHLSLARVIPVSGPARCLSQSRNLLKTTDDMVKTAREKLKHYSCTAEDIDHEDITRDQTSTLKTCLPLELHKNESCLATRETSSTTRGSCLPPQKTSLMMTLCLGSIYEDLKMYQTEFQAINAALQNHNHQQIILDKGMLVAIDELMQSLNHNGETLRQKPPVGEADPYRVKMKLCILLHAFSTRVVTINRVMGYLSSATGSGATNFSLLKQAGDVEENPGPTRMDSTRTQINKVVRFDKLPGFGDSIEAQCGTSVNVHSSLRDILNQITKPNDVYSFSLASRLYAEERYPILPEYLQCVKELYRGGLEPINFQTAADQARELINSWVESQTNGIIRNVLQPSSVDSQTAMVLVNAIVFKGLWEKAFKDEDTQAMPFRVTEQESKPVQMMYQIGLFRVASMASEKMKILELPFASGTMSMLVLLPDEVSGLEQLESIINFEKLTEWTSSNVMEERKIKVYLPRMKMEEKYNLTSVLMAMGITDVFSSSANLSGISSAESLKISQAVHAAHAEINEAGREVVGSAEAGVDAASVSEEFRADHPFLFCIKHIATNAVLFFGRCVSPGGG.
The tOVA or IL-12-P2A-tOVA sequences were cloned in the pCCL.sin.cPPT.SFFV LV backbone. Bicistronic IL-12-P2A-EVIR and hIL-12-P2A-EVIR sequences were codon-optimized and synthesized by GenScript, before cloning in the pCCL.sin.cPPT.SFFV LV backbone. The mouse and human IL-12 sequences were described previously24.
Production and titration of LVs
LVs were produced in 293 T cells using a five-plasmid co-transfection method, based on previously published protocols60,61 with some modifications to enhance the yield. The plasmid mix included: (i) the envelope plasmid VSV-G, (ii) the packaging plasmid pMDLg/pRRE, (iii) the Rev plasmid, (iv) the pADVANTAGE plasmid, and (v) the transfer plasmid encoding the gene of interest. 293T cells were seeded in T175 flasks 24 h prior to transfection and reached 60–70% confluency at the time of transfection. Transfection was performed using the calcium phosphate (CaCl2-HBS) method60,61.
Sixteen hours after transfection, the medium was replaced and the supernatant collected 24 and 48 h after the medium change. The two harvests were pooled, filtered through a 0.22 μm filter, and concentrated by ultracentrifugation at 82,600 RCF for 2 hours at 20 °C60,61. Lentiviral particles were resuspended in PBS and stored at −80 °C until use.
LV titration was performed on 293T cells, measuring the percentage of marker-positive cells by flow cytometry 4 to 7 days post-transduction, as described61. We used anti-dLNGFR antibody staining to calculate the titer of CtrlR, EVIR, IL-12-P2A-CtrlR, IL-12-P2A-EVIR, and hIL-12-P2A-EVIR LVs. We also used mCherry as a fluorescent marker for titrating the CD9-mCh LV. GD2S and GD3S encoding LVs were utilized directly without titration. To titrate OVA-expressing LVs, MC38 cells were transduced with serially diluted LVs and stained using an anti-H2Kb-SIINFEKL antibody.
Cell lines
The mouse B16F10 melanoma cell line and its OVA-expressing derivative were provided by P. Romero’s laboratory (University of Lausanne, Switzerland). The mouse MC38 colorectal carcinoma cell line was obtained from Kerafast (cat. ENH204-FP). The mouse E0771 mammary tumour cell line was provided by J. Huelsken’s laboratory (EPFL, Switzerland). Human embryonic kidney 293T cells were originally obtained from L. Naldini (San Raffaele Institute, Milan, Italy) and maintained in the De Palma’s laboratory for LV production. Additional stocks were obtained from ATCC (cat. CRL-11268).
All cell lines were cultured in Dulbecco’s Modified Eagle Medium (DMEM; ThermoFisher Scientific) supplemented with 10% fetal bovine serum (FBS; Gibco), 1% penicillin-streptomycin (Pen-Strep; ThermoFisher Scientific), and 2mM L-glutamine (Gibco). Cell cultures were maintained at 37 °C in a humidified incubator with 5% CO2. Each cell line was routinely tested for potential Mycoplasma contamination. Although no authentication was performed in our laboratory, the cancer cell lines appeared authentic based on their morphology, growth behavior in vitro, and tumour formation capability in mice. Additionally, 293 T cells effectively produced high-titer LVs and exhibited the expected morphology and growth patterns in vitro.
Modification of cell lines
To generate GD2-expressing cancer cells, we utilized sequential LV transduction of GD2S and GD3S, as described previously19. B16F10-OVA cells were used to generate B16F10-OVA/GD2 cells. Transduced cells were sorted using a FACS Aria II apparatus (BD Biosciences) to isolate GD2-positive cells, with GD2 expression confirmed by flow cytometry before experiments. To generate HER2-expressing MC38 cancer cells (MC38-HER2), MC38 cells were transduced with a HER2 encoding LV19.
E0771-tOVA/HER2 cells were generated by sequential transduction of E0771 cells with HER2 and tOVA encoding LVs. To generate the β2m-deficient variant (E0771-tOVA/HER2β2m-KO), we used a doxycycline-inducible lentiviral CRISPR-Cas9 system, as described previously19. Briefly, cells were transduced with LVs encoding Cas9 and a β2m-targeting sgRNA19, followed by puromycin selection in the presence of doxycycline for 4 days. Single-cell clones were then isolated by limiting dilution, expanded, and screened for loss of β2m surface expression by flow cytometry. β2m-negative clones were used for downstream functional assays.
The CD9-mCherry-expressing cell lines (B16F10-OVA/GD2/CD9-mCh and MC38-GD2/CD9-mCh) were generated by transducing cells with LVs encoding the CD9-mCherry fusion protein. Transduced cells were sorted to isolate high mCherry-positive cells. Expression of the CD9-mCherry fusion protein in the sorted cells was confirmed by flow cytometry before the experiments.
Mice
We used female CD45.1 (C57Bl/6J, Jackson Laboratory stock #002014) and CD45.2 (C57Bl/6J, Jackson Laboratory stock #000664) C57Bl/6J mice for all experiments involving non-transgenic, wild-type mice, except for the safety study (Fig. 5) where we used both male and female CD45.2 C57Bl/6 J mice (to assess potential sex-dependent effects). We used both female and male Rag1–/–62 (Jackson Laboratory stock #002216), WAP-HER234 (Jackson Laboratory stock #010562) and CD11c-EVIR transgenic mice, depending on availability in the EPFL or Agora animal facilities, and female OT-I36 (Jackson Laboratory stock #003831) and OT-II37 (Jackson Laboratory stock #004194) transgenic mice as T cell donors. All mice were maintained under specific pathogen-free (SPF) conditions at the EPFL or Agora animal facility in Lausanne, Switzerland. All mice were housed in groups of up to 5 mice/cage at 18-24 degrees C (ambient temperature), with 40-60% humidity, and maintained on a 12 h light/dark cycle (6 am to 6 pm). Food and water were available ad libitum.
CD45.2 C57Bl/6 J mice were purchased from Charles River Laboratories (France) while the other strains were bred in-house. We used CD45.1 or CD45.2 C57Bl/6 mice for BM cell isolation. We used OT-I and OT-II C57Bl/6 mice for T cell isolation from the spleen. Rag1–/–, WAP-HER2 and CD11c-EVIR transgenic mice, as well as CD45.2 C57Bl/6J mice, were used as recipient of subcutaneous tumours.
Generation of EVIR transgenic mice
To generate αGD2 EVIR transgenic mice, a targeting construct, containing the isogenic ROSA26 homology arms, the CAGs promoter, a LoxP-Stop-LoxP neomycin resistance selection cassette, and the αGD2 EVIR sequence, was targeted by homologous recombination to the ROSA26 locus of mouse C57BL/6 J embryonic stem cells (ESCs). Briefly, C57BL/6 J ESCs (5 × 106) were electroporated with 20 μg SalI linearized αGD2 EVIR targeting construct DNA (total volume 600 ul) in a 0.4-cm Gene Pulser cuvette (Bio-Rad) at 10 μF, 0.8 kV for 0.1 ms in a Bio-Rad Gene Pulser. Cells were plated on a 10 cm tissue culture dish precoated with 0.1 mg/mL laminin (Sigma). After 24 h, Geneticin (160 ng/mL; Gibco) was added to the 2i culture medium63 to selectively grow clones of transformed ES cells. The medium was refreshed every other day. After 10–14 days, individual clones were picked, dissociated using accutase, and transferred to a 0.1% gelatin-coated 96-well plate. After expansion, genomic DNA was isolated from the clones and used for PCR and Southern blot analysis to identify correctly targeted clones. Cells from selected ESC clones were injected into C57BL/6 N blastocysts and implanted into pseudopregnant B6CBAF1/Ola foster mice. The percentage of chimerism of the resulting chimeras was determined by qPCR. Male chimeras with high ESC contribution were mated with C57BL/6 J females. Germ-line transmission resulted in F1 mice carrying the correctly targeted ROSA26 allele.
To generate mice with EVIR expression specifically in CD11c+ cells (CD11c-EVIR mice), we crossed αGD2 EVIR transgenic mice with Itgax-Cre transgenic mice64. We then used heterozygous Itgax-Cre/homozygous EVIR mice as recipients of subcutaneous tumours.
Generation of mouse DCPs
Mouse DCPs were generated according to a previously published protocol24. Briefly, mouse BM cells were harvested from femurs and tibias of C57Bl/6 J mice. The BM cells were cultured for 2-3 days in complete RPMI medium (containing 10% FBS, 100 U/ml penicillin, 100 μg/ml streptomycin, and 2 mM glutamine) supplemented with 100 ng/ml SCF, 40 ng/ml TPO, 50 ng/ml FLT3L, 30 ng/ml IL-3, 30 ng/ml IL-6, and 30 ng/ml IL-1β (all from PeproTech) to promote the expansion of hematopoietic stem and progenitor cells (HSPC medium). After the expansion phase, floating cells were collected and cultured for an additional 4-5 days in complete RPMI medium supplemented with 200 ng/ml FLT3L and 5 ng/ml GM-CSF (PeproTech), to induce partial differentiation and commitment to the cDC1 lineage (cDC medium). The resulting culture contained DCPs (CD115+, CD11b–, CD11c–, CD103–, MHCII– and CD45R/B220– cells), which were enriched by depleting lineage-positive cells using EasySep Mouse Hematopoietic Progenitor Cell Isolation Kit (STEMCELL Technologies) before transduction, as described24.
Generation of mouse moDCs and cDCs
To produce mouse moDCs, BM cells were cultured in complete RPMI medium supplemented with 100 ng/ml GM-CSF and 40 ng/ml IL-4, at the cell concentration of 2 × 106–3 × 106 cells/ml, as described previously19,65. The cells were cultured for 2 days, after which non-adherent and loosely adherent cells were collected. These cells were then replated at the same concentration and cultured for an additional 4–5 days before transduction.
To generate mouse cDCs, BM cells were first cultured in HSPC expansion medium for 2 days, using the same conditions described for DCPs. After transduction, cells were transferred to cDC1 differentiation medium and seeded at a density of 2 × 106– 3 × 106 cells/ml. Fresh medium (1 ml per well) was added every 3–4 days without adjusting cell density. Cells were maintained for 12–16 days, as previously described24,27.
Transduction of mouse DCPs, moDCs and cDCs with LVs
We used our previously published protocol for the transduction of all DC types24. For DCP transduction, freshly enriched DCPs were plated in cDC1 medium at a concentration of 1.5 × 106 cells/ml in six-well plates. The cells were then transduced with desired LV at multiplicity of infection (MOI) of 350. For moDC transduction, non-adherent or loosely attached cells were collected, plated at a concentration of 1.5 × 106 cells/ml in moDC medium, and transduced with LVs at MOI of 100. For cDC transduction, the cells were transduced on day 2 of the differentiation protocol. Cells were harvested, replated in HSPC medium at 1.5 × 106 cells/ml and transduced with LVs at an MOI of 350. On day 3, cells were collected, washed, and transferred to cDC1 medium; differentiation was continued for 12 to 16 days, as described above. Transduction efficiency was assessed 3 days post-transduction by flow cytometry analysis of the transgene of interest (CtrlR or EVIR, both by anti-dLNGFR antibody staining).
Antigen presentation assays using mouse cDCs and T cells
E0771-HER2, E0771-tOVA/HER2 and E0771-tOVA/HER2β2m-KO cells were seeded at 10,000 cells per well in 96-well flat-bottom plates and cultured overnight. On the following day, cancer cells were treated with 10 μg/ml mitomycin C (Sigma) for 1 hour at 37°C, washed, and used in co-culture assays. Transduced cDCs were resuspended in cDC1 medium and added to each well at 50,000 cells per well. In parallel, OT-I CD8⁺ and OT-II CD4⁺ T cells were isolated from spleens of OT-I and OT-II transgenic mice using magnetic negative selection kits (EasySep, STEMCELL Technologies) and resuspended in T cell medium (complete RPMI supplemented with 50 μM β-mercaptoethanol, minimal non-essential amino acids, 1 mM sodium pyruvate, and 10 mM HEPES). T cells were added at a 1:1 ratio with cDCs (50,000 cells per well). Recombinant mouse IL-12 (Peprotech) was added to all conditions at a final concentration of 20 ng/ml. Co-cultures with OT-I cells were maintained for 3 days, and those with OT-II cells for 5 days, and were analyzed for intracellular IFNγ expression by flow cytometry, as previously described24.
Subcutaneous tumour models
All tumour studies used recipient C57BL/6J mice aged 8–12 weeks. To establish subcutaneous tumour models, cancer cells were passaged at least three times prior to injection. The cancer cells were resuspended in PBS at a concentration of 5 × 106 cells/ml for MC38-HER2 and MC38-GD2/CD9-mCh, and 2 × 106 cells/ml for B16F10-OVA, B16F10-OVA/GD2, and a mixture of B16F10-OVA/GD2 and B16F10 (ratio 3:1). A total of 100 µl of the cell suspension was injected subcutaneously into the right flank of C57Bl/6 mice.
Engineered DCs were administered post-tumour injection unless otherwise specified. Different timepoints for the injection of engineered DCs or antibodies are shown in the figures or explained below. Tumour growth was monitored 2–3 times per week using calipers and tumour volumes were calculated using the formula: (length × width2)/2. Tumours were allowed to grow up to 1 cm3 and the maximal tumour burden was not exceeded. When the tumour volume reached the 1 cm3 limit, the experiment was scheduled for termination within 48 h, provided that all mouse health parameters remained normal based on a detailed health score sheet.
Terminal anaesthesia
At the end of the experiments, the mice were terminated by deep sedation using pentobarbital (150 mg/kg) administered intraperitoneally, followed by cervical dislocation.
Adoptive transfer of mouse DCs
In the experiment involving Rag1–/– mice and the analysis of mCh internalization by adoptively transferred DCPs, transduced DCPs were injected intratumourally. All other tumour experiments and the safety study used intravenously injected DCPs or moDCs. The cells and antibodies were administered at the dosage regimens indicated in each figure. Age- and sex-matched mice injected with PBS were used as controls.
Treatment of the mice with CD8a-depleting or PD-1 blocking antibodies
Mice were treated with ɑPD-1 antibodies (10 mg/kg, clone RMP1-14, Bio X Cell) on days 10 and 13, unless indicated otherwise in the Figures. When combined with ɑCD8a (12 mg/kg, clone 53-6.7, Bio X Cell), four antibody treatments every 3 days were started on day 10. In some experiments, control mice received equivalent doses of rat IgG2a isotype control (10 mg/kg, clone 2A3, Bio X Cell). All antibodies were administered intraperitoneally.
Ex vivo cell restimulation and intracellular staining
Single-cell suspensions obtained from tumours were stimulated with 10 ng/ml PMA and 500 ng/ml ionomycin in complete RPMI medium for 30 minutes. GolgiStop and GolgiPlug protein transport inhibitors (BD Biosciences) were then added according to the manufacturer’s protocol, and the cells were incubated for an additional 3 h. Following incubation, cells were permeabilized and stained for intracellular T cell proteins according to the manufacturer’s instructions. Samples were then acquired using a flow cytometer for analysis. T cells from co-culture assays were processed using the same staining procedure. Samples were acquired using a flow cytometer for analysis.
Mouse serum biochemistry
Blood samples were collected by tail vein puncture at day 4 and cardiac puncture at the endpoint (day 36) into serum separator tubes (MiniCollect, Greiner Bio-One). Samples were centrifuged at 12,000 × g for 15 min at 4 °C. The resulting serum was collected and stored at –80 °C until analysis. Serum levels of alanine aminotransferase (ALT), aspartate aminotransferase (AST), and creatinine were measured using commercially available colorimetric kits: ALT (Abcam, ab105134), AST (Abcam, MAK055), and creatinine (Abcam, ab65340), according to the manufacturers’ protocols. As a positive control for liver injury, 15-weeks old male C57BL/6N mice were fed a methionine- and choline-deficient (MCD) diet (Envigo, TD90262) for 7 weeks. Mice on a matched control diet (Envigo, TD94149) served as negative controls.
Histological analysis of mouse liver and kidney
Liver and kidney tissues were harvested at the study endpoint and fixed overnight in 4% paraformaldehyde (PFA) at 4 °C. Tissues were then embedded in paraffin, sectioned, and stained with hematoxylin and eosin (H&E). Histological evaluation was performed in a blinded manner. Liver sections were scored in a blinded manner for steatosis, lobular inflammation, hepatocellular ballooning, and fibrosis using established Kleiner/NASH-CRN criteria. Renal sections were examined in a blinded fashion using systematic histological scoring, assessing tubular injury patterns (e.g., tubular dilation, epithelial simplification, cellular casts) and glomerular lesions according to standardized morphologic criteria. H&E-stained tissue sections were acquired using a Zeiss Axioscan 7 slide scanner.
Immunofluorescence staining of mouse tumours
Freshly isolated tumours were snap-frozen in OCT Compound and stored at −80 °C. Tissue sections were fixed in 4% paraformaldehyde for 15 min at room temperature, followed by a wash with PBS. The sections were then incubated in blocking buffer (PBS containing Fc block, 1% BSA, and 5% FBS) for 30 minutes at room temperature. Subsequently, the sections were incubated overnight at 4 °C with APC anti-GD2 antibody (1:100 dilution, clone 14G2A, Biolegend; Suppl. Table 1) in the blocking solution. After incubation, the slides were washed with PBS and stained with DAPI (1 µg/ml, Sigma) to reveal cell nuclei. Finally, the slides were mounted with Dako mounting medium (Agilent), dried overnight at room temperature, and images were acquired using an Olympus VS200 slide scanner. To analyze images, we used the Qupath 7 software. A region of interest (ROI) of the tumour was defined by setting a DAPI threshold and adjusted manually to include necrotic areas and exclude adjacent tissue. Pixel intensity-based classification was used to determine the frequency of GD2+ cells, with manual verification against negative controls on 20% of slides in a blinded manner to select the optimal threshold. The number of GD2+ cells was normalized to the total number of cells computed by DAPI staining).
Flow cytometry analysis
Single-cell suspensions were prepared from harvested tumours, spleens, and tumour-draining lymph nodes (tdLNs). Tumours were mechanically dissociated and digested enzymatically with collagenase IV (0.2 mg/ml; Worthington), dispase (2 mg/ml; Life Technologies), and DNaseI (0.1 mg/ml; New England BioLabs), in DMEM medium, for 30 min at 37 °C. The cell suspensions were filtered through a 70 μm cell strainer to remove debris and washed in FACS buffer (PBS + 10% FBS + 2 mM EDTA).
Spleens were thoroughly smashed on a 70 μm cell strainer (Falcon), red blood cells (RBCs) lysed using RBC lysis buffer (Sigma), and splenocytes washed once in FACS buffer. TdLNs were smashed on a 5 ml tube with a cell strainer cap (Falcon) and the cell suspensions washed once in FACS buffer. Blood samples were collected by tail vein puncture, placed into serum separator tubes (MiniCollect, Greiner Bio-One) for RBC lysis with RBC lysis buffer (Sigma), and washed once in FACS buffer before analysis.
Cell suspensions were stained with fluorescently labeled antibodies (Supplementary Table 1). Fixable live/dead staining was performed according to the manufacturer’s protocol. Non-fixable live/dead color, 7-AAD (1 µg/ml; BioLegend) or DAPI (0.1 µg/ml; Sigma-Aldrich), was added before acquisition by flow cytometer. T cell staining with SIINFEKL-bound dextramer was performed according to the manufacturer’s protocol (Immudex). Intracellular staining was carried out according to the manufacturer’s instructions for the GolgiStop and GolgiPlug kits (BD Biosciences). Data were acquired using BD LSRFortessa (Becton Dickinson), BD LSR II SORP (Becton Dickinson), Attune NxT (Invitrogen) or CytoFLEX LX2 (Beckman Coulter) flow cytometers and analyzed with FlowJO v10.1. software. Immune cell populations were identified according to the gating strategies shown in Supplementary Figs. 2–6.
Generation and transduction of human DCPs
Transduced human DCPs were generated according to a previously published protocol24. Cord blood (CB)-derived CD34+ cells (STEMCELL Technologies) were cultured in StemSpan SFEMII medium with a CD34+ Expansion Supplement and 1 μM UM729. On day 1, CD34+ cells were transferred to retronectin-coated wells and treated with dmPGE2 to a final concentration of 10 μM. After 2 h, the cells were transduced with LVs encoding CtrlR, EVIR, or hIL-12-P2A-EVIR, at a MOI of 300. The following day, the medium was replaced, and the cells were cultured for an additional five days. The cells were maintained at a concentration of 5 × 104 cells/ml in U-bottom 96-well plates, with the medium replaced every 2–3 days. On day 7, CD34+CD115+ DCPs were sorted using a FACS Aria II apparatus. The sorted DCPs were then cultured in StemSpan SFEMII medium supplemented with 50 units/ml penicillin, 50 μg/ml streptomycin, 20 ng/ml hGM-CSF, 100 ng/ml hFLT3L, 20 ng/ml hSCF (all from PeproTech), and 1000 IU/ml hIFNa2b (InvivoGen) for another 7 days. On day 14, the cultures contained differentiated cell populations, including DCs and other antigen-presenting cells, which were referred to as DCP progeny.
Generation and transduction of human moDCs
Human moDCs were generated from peripheral blood mononuclear cells (PBMCs) as previously described24,52. PBMCs from healthy donors (Blood Transfusion Center, University of Lausanne, Switzerland) were separated using density gradient centrifugation on Lymphoprep (STEMCELL Technologies). From these PBMCs, CD14+ monocytes were isolated using magnetic bead separation (Miltenyi Biotec). The isolated CD14+ cells were then cultured in human moDC medium, which is complete RPMI medium supplemented with 50 ng/ml hGM-CSF (PeproTech) and 50 ng/ml hIL-4 (PeproTech), at a density of 1 × 106 cells/ml. On day 1, the cells were harvested and transduced with LVs encoding either CtrlR, EVIR, or hIL-12-P2A-EVIR at a MOI of 5 in the presence of Vpx virus like particles66,67. The cells were incubated with the LVs for 24 h. The following day, the medium was replaced with fresh human moDC medium, and the cells were cultured for an additional 6 days to differentiate into moDCs.
Preparation and isolation of EVs from human melanoma cell lines
To isolate EVs from human melanoma cell lines38,39, the cells were cultured in DMEM medium with 5% EV-depleted FBS, prepared as described previously19,68, and 200 ng/mL IFNγ for the last 48 h (Peprotech). Briefly, EV-depleted FBS was prepared by ultracentrifuging regular FBS at 134,000 × g for 16 h at 4 °C, followed by filtration through a 0.1 μm vacuum filtration bottle. The cell culture medium was then processed to remove cells and debris through a series of centrifugation steps: 500 × g for 5 minutes, 2000 × g for 10 min, and 10,000 × g for 30 min, all at 4 °C. Subsequently, the medium underwent ultracentrifugation at 104,000 × g for 70 min at 4 °C using Hitachi ultracentrifuge with a P32ST rotor. The EVs were collected, resuspended in PBS, and stored at –80 °C until further use.
In vitro EV internalization assay with human DCPs and moDCs
For the labeling of EVs with PKH26 (Sigma), we adopted a modified manufacturer’s protocol68. EVs were first diluted in diluent C, and PKH26 dye was added at a ratio of 1 μl dye per 100 μl of the EV solution. The labeling process was conducted for 10 min at room temperature in the dark. To terminate the reaction, 0.1% BSA in PBS was added. The EVs were then washed using Vivaspin® 500 columns with a 300,000 MW cutoff (Sartorius) by centrifuging at 4000 × g for 20 min, repeated three times, to remove any unbound dye. Finally, the concentrated EVs were resuspended in PBS for subsequent use.
Quantification of EVs was performed by measuring the protein content using the BCA protein assay reagent kit (Thermo Fisher Scientific). For co-culture experiments, human moDCs or DCP progeny were plated in U-bottom 96-well plates at a density of 5 × 10^5 cells/ml in the previously described culture medium, with the exception that EV-depleted FBS was used instead of normal FBS. Subsequently, EVs were added to the cultures at a concentration of 5 μg/ml. Uptake and internalization of melanoma-derived EVs were assessed by flow cytometry after overnight incubation.
In vitro antigen cross-dressing assay with human DCPs and moDCs
HLA-A2-restricted CMV/pp65495-504-specific CD8+ T cells40 were cultured in RPMI 1640 supplemented with 8% human serum, antibiotics, glutamine, NEAA, sodium pyruvate, 2-mercaptoethanol, and hIL-2 (PeproTech). EVs isolated from human melanoma cell lines were pulsed with 1 μg/ml CMV/pp65495-504 peptide for one hour at 37 °C. The antigen-loaded EVs were washed with PBS and added to HLA-A2- moDCs or DCP progeny cells at a concentration of 5 μg/ml in RPMI 1640 medium supplemented with 5% EV-depleted FBS, 100 U/ml penicillin, and 100 μg/ml streptomycin and glutamine. Following an overnight incubation at 37 °C and subsequent washing steps to remove unbound EVs, the cells were co-cultured with A2/CMV/pp65495-504-specific CD8+ T cells at a 1:1 ratio in the same medium. After a 30-min incubation at 37 °C, brefeldin A was added to the cultures at a final concentration of 1:1000 (BD Biosciences, GolgiPlug) to block cytokine secretion. The cells were then cultured for an additional 5 h before flow cytometry analysis.
Statistics and reproducibility
Experiments involving independent mouse cohorts were conducted once, with sample sizes selected based on prior experience with various tumour models. No specific statistical tests were used to predetermine the sample size; our previous experience with the different tumour models provided guidance about the adequate number of mice that would provide statistically significant results. Based on these considerations, we typically employed experimental cohorts of 5–10 mice. Although each mouse study was performed once, the reproducibility of the findings was confirmed through independent experiments conducted under varying conditions (e.g., timing of DCP infusion, tumour size at randomization; type of bait molecule) that addressed the same experimental question. In experiments with established tumours, mice were randomized into treatment groups before the start of treatment to ensure consistent tumour sizes across different treatment groups. Experimental endpoints were chosen based on protocols approved by Cantonal Veterinary Office of Vaud for animal experimentation. For studies using human primary blood-derived cells, one to two donors were used to generate human moDCs and DCPs.
Investigators were blinded during tumour measurements, flow cytometry acquisition (but not analysis), and immunofluorescence or H&E staining data analysis. For additional informating, see the Reporting Summary. Occasionally, samples were excluded from specific analyses due to technical issues during sample processing or data collection, for example, in cases of poor cell viability or insufficient cell numbers in flow cytometry analyses. Outliers were always included for data analysis unless specified in the figure legends.
Statistical analyses and graph generation were performed using Prism software (GraphPad Software), with error bars representing the standard error of the mean (s.e.m.) unless otherwise specified. The number of biological replicates and the statistical tests used are detailed in the figure legends. Exact p values are shown in the figures (except when p < 0.0001); statistics were not computed when n < 3. Comparisons between two unpaired groups were conducted using the non-parametric two-tailed Mann-Whitney test. For single-variable multiple comparisons, we used one-way ANOVA followed by Tukey’s multiple comparison test. For two-variable analyses, such as tumour growth over time across multiple groups, two-way ANOVA was used, followed by Tukey’s test for three or more groups or Sidak’s test for two groups. Additional statistical tests were applied in specific cases, as detailed in the figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All reagents used in this study are either commercially available or can be made available from the corresponding author upon request. Source data are provided with this paper.
References
Perez, C. R. & De Palma, M. Engineering dendritic cell vaccines to improve cancer immunotherapy. Nat. Commun. 10, 5408 (2019).
Pittet, M. J., Di Pilato, M., Garris, C. & Mempel, T. R. Dendritic cells as shepherds of T cell immunity in cancer. Immunity 56, 2218–2230 (2023).
Gerhard, G. M., Bill, R., Messemaker, M., Klein, A. M. & Pittet, M. J. Tumor-infiltrating dendritic cell states are conserved across solid human cancers. J. Exp. Med. 218, e20200264 (2021).
Cabeza-Cabrerizo, M., Cardoso, A., Minutti, C. M., Pereira da Costa, M. & Reis e Sousa, C. Dendritic cells revisited. Annu. Rev. Immunol. 39, 131–166 (2021).
Del Prete, A. et al. Dendritic cell subsets in cancer immunity and tumor antigen sensing. Cell. Mol. Immunol. 20, 432–447 (2023).
Marciscano, A. E. & Anandasabapathy, N. The role of dendritic cells in cancer and anti-tumor immunity. Semin. Immunol. 52, 101481 (2021).
Bhandarkar, V., Dinter, T. & Spranger, S. Architects of immunity: how dendritic cells shape CD8+ T cell fate in cancer. Sci. Immunol. 10, eadf4726 (2025).
Hsu, F. J. et al. Vaccination of patients with B–cell lymphoma using autologous antigen–pulsed dendritic cells. Nat. Med. 2, 52–58 (1996).
Yu, J., Sun, H., Cao, W., Song, Y. & Jiang, Z. Research progress on dendritic cell vaccines in cancer immunotherapy. Exp. Hematol. Oncol. 11, 3 (2022).
Heras-Murillo, I., Adán-Barrientos, I., Galán, M., Wculek, S. K. & Sancho, D. Dendritic cells as orchestrators of anticancer immunity and immunotherapy. Nat. Rev. Clin. Oncol. 21, 257–277 (2024).
Harari, A., Graciotti, M., Bassani-Sternberg, M. & Kandalaft, L. E. Antitumour dendritic cell vaccination in a priming and boosting approach. Nat. Rev. Drug Discov. 19, 635–652 (2020).
Anguille, S., Smits, E. L., Lion, E., van Tendeloo, V. F. & Berneman, Z. N. Clinical use of dendritic cells for cancer therapy. Lancet Oncol. 15, e257–e267 (2014).
Saxena, M., van der Burg, S. H., Melief, C. J. M. & Bhardwaj, N. Therapeutic cancer vaccines. Nat. Rev. Cancer 21, 360–378 (2021).
Wculek, S. K. et al. Dendritic cells in cancer immunology and immunotherapy. Nat. Rev. Immunol. 20, 7–24 (2020).
Leary, N. et al. Melanoma-derived extracellular vesicles mediate lymphatic remodelling and impair tumour immunity in draining lymph nodes. J. Extracell. Vesicles 11, e12197 (2022).
Markov, O., Oshchepkova, A. & Mironova, N. Immunotherapy based on dendritic cell-targeted/-derived extracellular vesicles-a novel strategy for enhancement of the anti-tumor immune response. Front. Pharmacol. 10, 1152 (2019).
Ukrainskaya, V. M. et al. CAR-tropic extracellular vesicles carry tumor-associated antigens and modulate CAR T cell functionality. Sci. Rep. 13, 463 (2023).
Wolfers, J. et al. Tumor-derived exosomes are a source of shared tumor rejection antigens for CTL cross-priming. Nat. Med. 7, 297–303 (2001).
Squadrito, M. L., Cianciaruso, C., Hansen, S. K. & De Palma, M. EVIR: chimeric receptors that enhance dendritic cell cross-dressing with tumor antigens. Nat. Methods 15, 183–186 (2018).
Nazha, B., Inal, C. & Owonikoko, T. K. Disialoganglioside GD2 expression in solid tumors and role as a target for cancer therapy. Front. Oncol. 10, 1000 (2020).
MacNabb, B. W. & Kline, J. MHC cross-dressing in antigen presentation. Adv. Immunol. 159, 115–147 (2023).
Martinez-Usatorre, A. & De Palma, M. Dendritic cell cross-dressing and tumor immunity. EMBO Mol. Med. 14, e16523 (2022).
Chatterjee, F. & Spranger, S. MHC-dressing on dendritic cells: Boosting anti-tumor immunity via unconventional tumor antigen presentation. Semin. Immunol. 66, 101710 (2023).
Ghasemi, A. et al. Cytokine-armed dendritic cell progenitors for antigen-agnostic cancer immunotherapy. Nat. Cancer 5, 240–261 (2024).
Salmon, H. et al. Expansion and activation of CD103(+) dendritic cell progenitors at the tumor site enhances tumor responses to therapeutic PD-L1 and BRAF inhibition. Immunity 44, 924–938 (2016).
Spranger, S., Dai, D., Horton, B. & Gajewski, T. F. Tumor-residing Batf3 dendritic cells are required for effector T cell trafficking and adoptive T cell therapy. Cancer Cell 31, 711–723.e4 (2017).
Mayer, C. T. et al. Selective and efficient generation of functional Batf3-dependent CD103+ dendritic cells from mouse bone marrow. Blood 124, 3081–3091 (2014).
Saito, Y., Komori, S., Kotani, T., Murata, Y. & Matozaki, T. The role of type-2 conventional dendritic cells in the regulation of tumor immunity. Cancers 14, 1976 (2022).
Binnewies, M. et al. Unleashing type-2 dendritic cells to drive protective antitumor CD4+ T cell immunity. Cell 177, 556–571.e16 (2019).
Lu, L. et al. Differential expression of CD11c defines two types of tissue-resident macrophages with different origins in steady-state salivary glands. Sci. Rep. 12, 931 (2022).
Francis, D. M. et al. Blockade of immune checkpoints in lymph nodes through locoregional delivery augments cancer immunotherapy. Sci. Transl. Med. 12, eaay3575 (2020).
Garris, C. S. et al. Successful anti-PD-1 cancer immunotherapy requires T cell-dendritic cell crosstalk involving the cytokines IFN-γ and IL-12. Immunity 49, 1148–1161.e7 (2018).
Duval, F. et al. Trajectories of macrophage ontogeny and reprogramming in cancer. iScience 28, 112498 (2025).
Piechocki, M. P., Ho, Y.-S., Pilon, S. & Wei, W.-Z. Human ErbB-2 (Her-2) transgenic mice: a model system for testing Her-2 based vaccines. J. Immunol. 171, 5787–5794 (2003).
Diebold, S. S., Cotten, M., Koch, N. & Zenke, M. MHC class II presentation of endogenously expressed antigens by transfected dendritic cells. Gene Ther. 8, 487–493 (2001).
Hogquist, K. A. et al. T cell receptor antagonist peptides induce positive selection. Cell 76, 17–27 (1994).
Barnden, M. J., Allison, J., Heath, W. R. & Carbone, F. R. Defective TCR expression in transgenic mice constructed using cDNA-based α- and β-chain genes under the control of heterologous regulatory elements. Immunol. Cell Biol. 76, 34–40 (1998).
Neubert, N. J. et al. Broad and conserved immune regulation by genetically heterogeneous melanoma cells. Cancer Res. 77, 1623–1636 (2017).
Neubert, N. J. et al. T cell–induced CSF1 promotes melanoma resistance to PD1 blockade. Sci. Transl. Med. 10, eaan3311 (2018).
Iancu, E. M. et al. Clonotype selection and composition of human cd8 t cells specific for persistent herpes viruses varies with differentiation but is stable over time1. J. Immunol. 183, 319–331 (2009).
Broz, M. L. et al. Dissecting the tumor myeloid compartment reveals rare activating antigen-presenting cells critical for T cell immunity. Cancer Cell 26, 638–652 (2014).
Gardner, A. et al. TIM-3 blockade enhances IL-12-dependent antitumor immunity by promoting CD8+ T cell and XCR1+ dendritic cell spatial co-localization. J. Immunother. Cancer 10, e003571 (2022).
Milone, M. C. et al. Engineering-enhanced CAR T cells for improved cancer therapy. Nat. Cancer 2, 780–793 (2021).
López, L. et al. Dendritic cell-targeted therapy expands CD8 T cell responses to bona-fide neoantigens in lung tumors. Nat. Commun. 15, 2280 (2024).
Pfirschke, C. et al. Macrophage-targeted therapy unlocks antitumoral cross-talk between IFNγ-secreting lymphocytes and IL12-producing dendritic cells. Cancer Immunol. Res. 10, 40–55 (2022).
Ruffell, B. et al. Macrophage IL-10 blocks CD8+ T cell-dependent responses to chemotherapy by suppressing IL-12 expression in intratumoral dendritic cells. Cancer Cell 26, 623–637 (2014).
Ashour, D. et al. IL-12 from endogenous cDC1, and not vaccine DC, is required for Th1 induction. JCI Insight 5, 135143 e135143. (2020).
Hammerich, L. et al. Systemic clinical tumor regressions and potentiation of PD1 blockade with in situ vaccination. Nat. Med. 25, 814–824 (2019).
Hadiloo, K., Taremi, S., Heidari, M. & Esmaeilzadeh, A. The CAR macrophage cells, a novel generation of chimeric antigen-based approach against solid tumors. Biomark. Res. 11, 103 (2023).
Liang, Y., Xu, Q. & Gao, Q. Advancing CAR-based immunotherapies in solid tumors: CAR- macrophages and neutrophils. Front. Immunol. 14, 1291619 (2023).
Anderson, N. R., Minutolo, N. G., Gill, S. & Klichinsky, M. Macrophage-based approaches for cancer immunotherapy. Cancer Res. 81, 1201–1208 (2021).
Klichinsky, M. et al. Human chimeric antigen receptor macrophages for cancer immunotherapy. Nat. Biotechnol. 38, 947–953 (2020).
Liu, M. et al. CAR-Macrophages and CAR-T cells synergistically kill tumor cells in vitro. Cells 11, 3692 (2022).
Pierini, S. et al. Chimeric antigen receptor macrophages (CAR-M) sensitize HER2+ solid tumors to PD1 blockade in pre-clinical models. Nat. Commun. 16, 706 (2025).
Reiss, K. A. et al. CAR-macrophage therapy for HER2-overexpressing advanced solid tumors: a phase 1 trial. Nat. Med. 31, 1171–1182 (2025).
Majumder, A. et al. Generation of GD2-CAR neutrophils from hPSCs for targeted cancer immunotherapy of solid tumors. J. Immunother. Cancer 10, (2022).
Chang, Y. et al. CAR-neutrophil mediated delivery of tumor-microenvironment responsive nanodrugs for glioblastoma chemo-immunotherapy. Nat. Commun. 14, 2266 (2023).
Hou, X., Zaks, T., Langer, R. & Dong, Y. Lipid nanoparticles for mRNA delivery. Nat. Rev. Mater. 6, 1078–1094 (2021).
Tsuneyoshi, N. et al. Hypoimmunogenic human iPSCs expressing HLA-G, PD-L1, and PD-L2 evade innate and adaptive immunity. Stem Cell Res. Ther. 15, 193 (2024).
De Palma, M., Venneri, M. A., Roca, C. & Naldini, L. Targeting exogenous genes to tumor angiogenesis by transplantation of genetically modified hematopoietic stem cells. Nat. Med. 9, 789–795 (2003).
De Palma, M. & Naldini, L. Transduction of a gene expression cassette using advanced generation lentiviral vectors. Methods Enzymol. 346, 514–529 (2002).
Mombaerts, P. et al. RAG-1-deficient mice have no mature B and T lymphocytes. Cell 68, 869–877 (1992).
Huijbers, I. J. et al. Rapid target gene validation in complex cancer mouse models using re-derived embryonic stem cells. EMBO Mol. Med. 6, 212–225 (2014).
Stranges, P. et al. Fas-mediated elimination of antigen-presenting cells and autoreactive T cells contribute to prevention of autoimmunity. Immunity 26, 629–641 (2007).
Helft, J. et al. GM-CSF mouse bone marrow cultures comprise a heterogeneous population of CD11c(+)MHCII(+) macrophages and dendritic cells. Immunity 42, 1197–1211 (2015).
Berger, G. et al. A simple, versatile and efficient method to genetically modify human monocyte-derived dendritic cells with HIV-1–derived lentiviral vectors. Nat. Protoc. 6, 806–816 (2011).
Bobadilla, S., Sunseri, N. & Landau, N. R. Efficient transduction of myeloid cells by a lentiviral vector that packages the Vpx accessory protein. Gene Ther. 20, 514–520 (2013).
Cianciaruso, C. et al. Molecular profiling and functional analysis of macrophage-derived tumor extracellular vesicles. Cell Rep. 27, 3062–3080.e11 (2019).
Acknowledgements
We thank Céline Rmili-Wyser, Bruno Torchia and Alan Guichard for managing mouse colonies and technical help with experiments. This study was supported by grants from the European Research Council (ERC-CoG EVOLVE-725051 to M.D.P.), the Swiss Cancer Research Foundation/Swiss Cancer League (KLS-4505-08-2018 to M.D.P.), the Swiss National Science Foundation (SNSF 310030-188868 to M.D.P. and SNSF 320030-232217 to M.D.P. and D.M.), and the Fondation pour la lutte contre le cancer (to M.D.P.).
Author information
Authors and Affiliations
Contributions
A.G. designed and performed most experiments, collected and analyzed the data, interpreted the results and wrote the paper. A.M.-U. designed, performed and analyzed experiments involving human DCPs and moDCs. H.D. performed and analyzed safety studies in mice. Y.L., L.L., Y.M., A.H. and M.H. contributed to experiments involving mouse models. L.H. and C.E.J.P. generated EVIR transgenic mice. D.E.S. and D.M. provided critical reagents, advised on experimental design, and interpreted the results. M.D.P. designed research and interpreted the results, supervised the study, and wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
M.D.P. has received sponsored research grants from EVIR Therapeutics, Hoffmann La-Roche and Deciphera Pharmaceuticals, serves on the Scientific Advisory Boards of EVIR Therapeutics, Montis Biosciences, Macomics, Deciphera Pharmaceuticals, Light Chain Bioscience/Novimmune, and Genenta. M.D.P., A.G., Y.M. and A.M.-U. are inventors on patents WO/2017/134100, WO/2023/232777, and application No. 24211693.7 filed by EPFL, which cover methods for DC engineering. D.E.S. serves on the Scientific Advisory Board of EVIR Therapeutics. D.M. is a consultant for Limula and MPC Therapeutics, scientific co-founder of Cellula Therapeutics, and inventor on unrelated patents filed by the University of Pennsylvania, Istituto Oncologico della Svizzera Italiana (IOSI) and the University of Geneva. The other authors declare no Competing interests.
Peer review
Peer review information
Nature Communications thanks Zhenyu Ding, Maksim Mamonkin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ghasemi, A., Martinez-Usatorre, A., Liu, Y. et al. Dendritic cell progenitors engineered to express extracellular-vesicle–internalizing receptors enhance cancer immunotherapy in mouse models. Nat Commun 16, 9148 (2025). https://doi.org/10.1038/s41467-025-64172-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64172-w