Abstract
Natural variations provide valuable genetic resources for improving rice grain number per panicle (GNP). Here, our genome-wide association study (GWAS) identifies GNP2 and GNP5 as key regulators of GNP that enhance rice yield. GNP5 encodes a bZIP transcription factor binding to the S5779181 locus in the GNP2 promoter, where natural variation significantly influences GNP. GNP2 encodes a conserved GSK3-like kinase that phosphorylates and stabilizes Gnp4/LAX2. The phosphorylated Gnp4/LAX2T175,262D promotes yield by modulating transcription factors involved in panicle development. Haplotype analysis reveals an elite allele combination (Type I) of GNP5 and GNP2 that significantly increases GNP. Field trials demonstrate that enhanced GNP2 expression raises yield by approximately 10%. Our findings thus uncover a genetic resource with application potential for enhancing rice yield.
Similar content being viewed by others
Introduction
Rice (Oryza sativa L.) serves as a primary staple food for nearly half of the global population1,2. Rice yield is primarily determined by three factors: the number of panicles per plant, grain number per panicle (GNP), and grain weight3,4. Extensive breeding practices have demonstrated that enhancing GNP is one of the most effective approaches to improving rice yield. Consequently, identifying and harnessing genes that regulate GNP in rice, particularly those present in the existing superior rice germplasm, offers the potential to improve rice yield.
The GNP is a complex trait controlled by multiple quantitative trait loci (QTLs), and extensive research has been dedicated to uncovering the genetic basis of GNP in the past few decades. A multitude of genes and QTLs associated with GNP have been identified through traditional genetic mapping, and these can be categorized based on their functions. For instance, GRAIN NUMBER 1a (Gn1a), LONELY GUY (LOG), and GY3 are involved in cytokinin regulation. Gn1a encodes cytokinin oxidase, and a specific allele from the Habataki variety can significantly boost GNP, highlighting the crucial role of hormonal regulation in determining GNP5. LOG and GY3 encode cytokinin-activating enzyme to regulate meristem activity6,7. Genes like LAX PANICLE1 (LAX1), LAX2 (also known as Grain number per-panicle 4, Gnp4), and FRIZZLE PANICLE (FZP) are engaged in the formation of panicle branches and florets. Mutations in LAX1 and Gnp4/LAX2 result in a reduction of secondary branches and GNP, while gene silencing of FZP increases in secondary inflorescence branching and grain number8,9,10,11. DENSE AND ERECT PANICLE1 (DEP1), BRASSINOSTEROID-DEFICIENT DWARF3 (BRD3), and MORE GRAINS 1 (MOG1) are associated with panicle architecture. DEP1 controls the erect and dense panicle phenotype, which enhances the activity of the rice meristem, thereby increasing GNP12. BRD3, participating in the brassinosteroid (BR) metabolism pathway, is activated in the secondary branch meristem of rice. This activation leads to a clustered panicle phenotype, effectively increasing GNP13. Overexpression of MOG1 results in longer and larger panicles with more branches14. Besides these, the GNP in rice tends to increase with a longer growth period, as it facilitates more extensive nutrient accumulation and more thorough panicle development. Enhanced expression of Grain number, plant height, and heading date2 (Ghd2) and its homolog Ghd7 both result in delayed heading and simultaneously increase GNP and plant height, while overexpression of OsMADS15 disrupts panicle development and reduces GNP15,16,17.
While traditional genetic mapping approaches, such as linkage analysis, have been effective in identifying QTLs and genes associated with GNP by leveraging controlled crosses and familial genetic populations, the genome-wide association studies (GWAS) capitalize on the natural genetic variations present in diverse germplasm collections, enabling a more comprehensive exploration of the genetic architecture underlying complex traits like GNP. In rice, GWAS has been successfully applied to study various traits, including grain size, plant height, and so on18. However, its application to understanding the genetic basis of GNP is still being refined, and only several genes directly conferring GNP were isolated by GWAS. Grain number per panicle on chromosome 5 (Gnd5), isolated by GWAS using GNP of japonica population, encodes a GRAS transcription factor (TF) and positively regulates GNP, and mutation of Regulator of grain number 1a (RGN1a) caused significantly reduced GNP19,20. Despite these advances, the genetic architecture of GNP remains complex, with many regulatory genes and interactions yet to be identified.
GLYCOGEN SYNTHASE KINASE 3 (GSK3), an evolutionarily conserved serine/threonine kinase, negatively regulates BR signaling in plants21. Research has identified nine GSK3-like kinases in rice22. Among them, OsGSK1, is highly expressed during panicle development. It is likely to be involved in stress response and flower development23,24. Overexpression of OsGSK2 typically exhibits a typical BR-deficient phenotype. However, in FLAG-aGSK2 overexpression lines, FLAG-aGSK2 accumulates significantly in the pedicels, promoting the clustered growth of multiple spikelets, resulting in dense and compact panicles, which significantly increases the GNP in rice13. Additionally, OsGSK2 also plays a role in regulating rice grain size, stripe mosaic virus resistance, and coleoptile elongation25,26,27,28. In comparison to wild-type, the osgsk3 mutant shows an increase in leaf size, leaf angle, plant height, and tiller number, as well as enhancements in grain length and thousand-grain weight29. OsGSK5 acts as a negative regulator of grain length and weight, and it can coordinately regulate the size and number of cells in the lemma, thereby affecting rice yield and grain development30,31,32. However, current research mainly focuses on the post-transcriptional regulation of GSK3-like kinases, and the mechanism by which GSK3-like kinases regulate rice GNP at the transcriptional level remains poorly understood.
In this work, we perform GWAS on 496 rice germplasms from diverse regions to investigate the genetic basis of phenotypic variation of GNP. We identify two beneficial alleles of two genes designated as GRAIN NUMBER PER PANICLE2 (GNP2) and GNP5. The GNP2 gene encodes a GSK3-like kinase, while GNP5 encodes a basic leucine zipper (bZIP) TF. The results demonstrate that GNP5 binds to the functional site of the GNP2 promoter, thereby positively regulating GNP. Further analysis reveals that GNP2 phosphorylates and stabilizes Gnp4/LAX2, which modulates the transcriptional activity of TFs involved in panicle development, ultimately influencing rice GNP. These findings not only provide insights into the regulatory mechanisms underlying rice GNP but also establish a theoretical foundation for improving panicle architecture and breeding high-yield rice varieties.
Results
Identification of GNP2 and GNP5 by GWAS
To investigate the genetic basis underlying the phenotypic diversity of GNP in rice germplasm, we planted 496 worldwide rice accessions, including 361 indica, 79 temperate japonica, 26 tropical japonica, 21 admixed, 5 aromatic, and 4 aus accessions, in Nanning (latitude 22.8° N, altitude 82 m), Guangxi Province, and Yuxi (latitude 24.4° N, altitude 1862 m), Yunnan Province, China (Supplementary Fig. 1 and Supplementary Data 1). Next, we conducted a GWAS using the GNP values of 496 rice accessions in Nanning and Yuxi with 4,890,951 single-nucleotide polymorphisms (SNPs) under a compressed mixed linear model. We identified 6 and 13 loci significantly associated with GNP in Yuxi and Nanning, respectively. Among these, qGNP1, qGNP4a, and qGNP8c were consistently detected in both environments (Fig. 1a and Supplementary Data 2). To explore the genes underlying qGNP1 and qGNP4a, we performed linkage disequilibrium (LD) analysis of the regions near the lead SNPs. qGNP1 was localized to a 196.395-kb interval (Chr.1: 5,779,181 to Chr.1: 5,975,575) containing eight candidate genes that co-localized with significant loci (Fig. 1b and Supplementary Data 3). Among these, LOC_Os01g10840 showed the highest expression in young panicles (Supplementary Fig. 2a). Similarly, LD analysis of qGNP4a identified five candidate genes in the 450.246-kb interval (Chr.4: 5,337,798 to Chr.4: 5,788,043) (Fig. 1c and Supplementary Data 3). Among these, LOC_Os04g10260 was highly expressed in young panicles (Supplementary Fig. 2b). These results suggest that LOC_Os01g10840 and LOC_Os04g10260 are likely the causal genes for qGNP1 and qGNP4a, respectively, and we named these candidate genes GNP2 and GNP5, respectively.
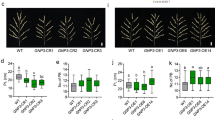
a Manhattan plot of GWAS for GNP in Yuxi and Nanning. Chr, Chromosome. b, c Regional Manhattan plots for qGNP1 (b) and qGNP4a (c), along with pairwise linkage disequilibrium analysis. Significant SNPs are presented as orange dots. d Panicle morphology of Nip, gnp2-knock out plants (ko1, ko2). e–h Statistical results for panicle length (e), grain number on primary branches (pbs., f), grain number on secondary branches (sbs., g), and GNP (h) of Nip and gnp2-ko plants. i Panicle morphology of Nip, GNP2-overexpressing plants (OE1, OE2). j–m Statistical results for panicle length (j), grain number on primary branches (k), grain number on secondary branches (l), and GNP (m) of Nip and GNP2-OE plants. n Panicle morphology of Nip, gnp5-knock out plants (ko1, ko2). o–r Statistical results for panicle length (o), grain number on primary branches (p), grain number on secondary branches (q), and GNP (r) of Nip and gnp5-ko plants. s Panicle morphology of Nip, GNP5-overexpressing plants (OE1, OE2). t–w Statistical results for panicle length (t), grain number on primary branches (u), grain number on secondary branches (v), and GNP (w) of Nip and GNP5-OE plants. In a–c, P values were determined under the compressed mixed linear model using a two-sided Fisher’s exact test. In e–h, j–m, o–r, t–w, different lowercase letters indicate statistically significant differences at P = 0.05 by one-way ANOVA with Duncan’s multiple-range test, and error bars represent means ± SD. Scale bars, 2 cm (d, i, n, s). Source data are provided as a Source Data file.
To determine whether GNP2 and GNP5 regulate GNP in rice, we knocked out (ko) GNP2 and GNP5 in the Nipponbare (Nip) background using CRISPR/Cas9-based mutagenesis. We obtained two independent mutant lines of GNP2, gnp2-ko1 and gnp2-ko2, with frameshift mutations creating a 1-bp insertion and a 1-bp deletion in the GNP2 coding region, respectively (Supplementary Fig. 3b). Compared with the Nip plants, there were no significant changes in plant height, panicle length, number of primary branches, grain number on primary branches, and tiller number in gnp2-ko1 and gnp2-ko2 plants; grain length was significantly increased. However, GNP, number of secondary branches, grain number on secondary branches, grain width, 1000-grain weight, and grain yield per plant were significantly reduced. (Fig. 1d–h and Supplementary Fig. 3c–l). We also obtained two independent mutants of GNP5: gnp5-ko1, with a 1-bp insertion in GNP5; and gnp5-ko2, with a 29-bp deletion in GNP5 (Supplementary Fig. 4a). In comparison to the Nip plants, no significant differences were observed in plant height, number of primary branches, grain number on primary branches, and tiller number in gnp5-ko1 and gnp5-ko2 plants; grain length, grain width, and 1000-grain weight were significantly increased. However, due to the significant reduction in GNP, panicle length, number of secondary branches, and number of grains on secondary branches, the grain yield per plant was significantly decreased. (Fig. 1n–r and Supplementary Fig. 4b, d–l).
In parallel, we introduced the coding sequences of GNP2 and GNP5 driven by the CaMV35S promoter into Nip to generate constitutive GNP2 and GNP5 overexpression lines named GNP2-OE1 and GNP2-OE2, and GNP5-OE1 and GNP5-OE2, respectively (Supplementary Figs. 3a and 4c). In comparison to Nip, the panicle length of GNP2 overexpression lines showed no significant change, but the number of grains on primary branches, the number of grains on secondary branches, and GNP were significantly increased. In GNP5 overexpression lines, panicle length, number of grains on primary branches, number of grains on secondary branches, and GNP were all significantly increased. (Fig. 1i–m, s–w). These results indicate that GNP2 and GNP5 are the causal genes for qGNP1 and qGNP4a, respectively, and play essential roles in the regulation of GNP in rice.
Variation S5779181 in the GNP2 promoter affects GNP
To identify the functional variations of GNP2 that regulate GNP, we identified five significant SNPs (S5776475, S5778082, S5778085, S5779181, and S5780335) in the 2.5-kb promoter and gene body region of GNP2. S5776475, S5778082, and S5778085 are located in the intron, S5779181 and S5780335 are located in the promoter (Fig. 2a and Supplementary Table 1). We classified the GNP2 alleles into four haplotypes based on the presence of the S5779181 and S5780335 variations (Table 1). In indica rice accessions, there was no significant difference in GNP between GNP2-Hap1 and GNP2-Hap2, but the GNP values were much higher in these lines than in GNP2-Hap4. In japonica rice accessions, the GNP was significantly higher in GNP2-Hap1 vs GNP2-Hap4 plants (Table 1). This suggests that GNP2-Hap1 and GNP2-Hap2 are the superior haplotypes for GNP, while GNP2-Hap4 is the inferior haplotype. Further analysis revealed that both GNP2-Hap1 and GNP2-Hap2 have the base G at the S5779181 locus, while GNP2-Hap4 has the base A. In addition, we analyzed the GNP of 657 rice accessions grown in Sanya (latitude 18.3° N, altitude 50 m), Hainan Province, China, based on 40 SNPs in the promoter region and 3 synonymous SNPs in the coding region of GNP2. In indica and japonica rice, accessions with G at S5779181 had a significantly higher GNP than accessions with A at the same locus (Supplementary Table 2). These results suggest that S5779181 in the GNP2 promoter is likely important for regulating GNP.
a Association analysis of variations in GNP2 with GNP. Red dots represent SNP loci with −log₁₀(P) ≥ 4. b Comparison of promoter activities based on the dual-luciferase transient expressing assays in rice protoplasts (n = 3). REN, renilla luciferase. LUC, luciferase. c Statistical results for GNP of different accessions harboring pGNP2(G) or pGNP2(A). d The expression level of GNP2 in young panicle from different accessions harboring pGNP2(G) or pGNP2(A) (n = 3). pGNP2(G) and pGNP2(A), at the GNP2 promoter, S5779181 is either the base G or A. e Panicle morphology of Nip, gnp2-ko, and GNP2 complementation (CM) plants. f Statistical results for GNP of Nip, gnp2-ko, and GNP2-CM plants. In b–d asterisks indicate statistical significance by two-tailed Student’s t-tests (**P < 0.01, ***P < 0.001, data represent means ± SD). In f, different lowercase letters indicate statistically significant differences at P = 0.05 by one-way ANOVA with Duncan’s multiple-range test, and error bars represent means ± SD. Scale bars, 2 cm (e). Source data are provided as a Source Data file.
To investigate the effect of S5779181 on the promoter activities of GNP2, we evaluated pGNP2(G) and pGNP2(A) promoter activities via transient expression in rice protoplasts. The promoter activity of pGNP2(G) was significantly higher than that of pGNP2(A) (Fig. 2b). Furthermore, GNP and GNP2 expression levels were significantly higher in accessions harboring pGNP2(G) vs pGNP2(A) (Fig. 2c, d). These results suggest that the differences in GNP2 expression among germplasm resources are attributable to the S5779181 variation. We introduced the pGNP2(G) or pGNP2(A) promoter and GNP2 genomic sequence into the gnp2-ko background, resulting in GNP2 complementation plants (GNP2-CMG and GNP2-CMA) (Supplementary Fig. 5a). The reduced GNP of gnp2-ko plants was fully or partially restored in the complementation plants. The number of secondary branches, grain number on secondary branches, and GNP were significantly higher in GNP2-CMG vs GNP2-CMA plants, whereas the panicle length, number of primary branches, grain number on primary branches, plant height, and tiller number showed no obvious differences between them (Fig. 2e, f and Supplementary Fig. 5b–i). These results demonstrate that S5779181 in the GNP2 promoter is responsible for the variation in GNP in natural germplasm.
GNP5 acts upstream of GNP2
bZIP TFs play important roles in plant growth and development33. GNP5 encodes a TF containing a bZIP domain. The bZIP domain preferentially binds to A-box (TACGTA), C-box (GACATC), and G-box (CACGTG) motifs in the promoters of their target genes34,35. Notably, the functional variation S5779181 of GNP2 is located in the G-box. We therefore investigated whether GNP5 directly binds to the G-box to regulate GNP2 expression. The expression of GNP2 in young panicles was significantly reduced in gnp5-ko, but increased in GNP5-OE, compared to Nip plants (Fig. 3a). In a yeast one-hybrid assay (Y1H), GNP5 interacted more strongly with the pGNP2(G) promoter than with the pGNP2(A) promoter in yeast cells (Fig. 3b). We then co-transfected rice protoplasts with the GNP5 effector and the pGNP2:LUC reporter constructs and performed dual-luciferase assays. GNP5 activated the transcription of GNP2, with stronger activation of the pGNP2(G) vs pGNP2(A) promoter (Fig. 3c, d). An electrophoretic mobility shift assay (EMSA) confirmed the binding of GNP5 to the P2 region in vitro (Fig. 3e). GNP5 bound more strongly to pGNP2(G) than to pGNP2(A) (Fig. 3f, g). Chromatin immunoprecipitation (ChIP) qRT-PCR assays revealed that GNP5 binds to three putative binding sites (P1–P3) in the GNP2 promoter, with the strongest binding occurring at P1/2 containing the S5779181 variation (Fig. 3h). These results indicate that GNP5 directly regulates the expression of GNP2 by binding to its promoter. To investigate the genetic relationship between GNP5 and GNP2, we generated the gnp5-ko gnp2-ko plants. The GNP of the gnp5-ko gnp2-ko plants was similar to that of gnp2-ko plants (Fig. 3i, j and Supplementary Fig. 6a, b). Therefore, we propose that GNP5 acts upstream of GNP2 to regulate GNP.
a Relative expression level of GNP2 in Nip, gnp5-ko, and GNP5-OE plants (n = 3). b Y1H assays showing GNP5 can bind to both pGNP2(G) and pGNP2(A). lacZ, β-galactosidase. c, d The dual-luciferase assays showing GNP5 activated the transcription of GNP2 by co-transfecting in rice protoplasts (n = 3). REN renilla luciferase. LUC luciferase. e, f EMSA assays showing the binding of GNP5 to the GNP2 promoter in vitro. g Quantification of the EMSA results in (f) using ImageJ software (n = 4). h ChIP qRT-PCR assays showing the binding of GNP5 to the GNP2 promoter in vivo (n = 3). i Panicle morphology of Nip, gnp5-ko, gnp2-ko, and gnp5-ko gnp2-ko plants. j Statistical results for GNP of Nip, gnp5-ko, gnp2-ko, and gnp5-ko gnp2-ko plants. In a, g asterisks indicate statistical significance by two-tailed Student’s t-tests (**P < 0.01,***P < 0.001, data represent means ± SD). In d, h, j, different lowercase letters indicate statistically significant differences at P = 0.05 by one-way ANOVA with Duncan’s multiple-range test, and error bars represent means ± SD. Scale bars, 2 cm (i). In b, e, f, a representative experiment from three independent experiments is shown. Source data are provided as a Source Data file.
GNP2 phosphorylates and stabilizes Gnp4/LAX2
GNP2 is mainly expressed in rice roots and panicles at different developmental stages, as revealed by qRT-PCR, which is consistent with the GUS activities present in these tissues in pGNP2:GUS transgenic plants (Supplementary Fig. 7a, b). In transiently transfected rice protoplasts, GNP2 mainly localized to the nucleus and cytoplasm (Supplementary Fig. 7c). GNP2 encodes GSK3/SHAGGY-like kinase, a multifunctional kinase that is highly conserved across eukaryotes36,37. To identify the substrates of GNP2, we constructed a cDNA expression library from young panicles to identify its interacting factors through a yeast two-hybrid (Y2H) assay and identified eight interacting proteins in yeast cells (Fig. 4a and Supplementary Table 3). We selected Gnp4/LAX2 for further study. Gnp4/LAX2 encodes a nuclear protein containing a plant-specific conserved domain, which regulates rice axillary meristem formation to influence plant growth and panicle development. Expression pattern analysis revealed that Gnp4/LAX2 is specifically expressed in secondary branches. The gnp4/lax2 mutants exhibited significantly reduced numbers of secondary branches and GNP38,39.
a, f Y2H assays showing GNP2 interacts with Gnp4/LAX2 in yeast cells. Yeast cells were cultured on SD minimal medium lacking Trp and Leu (SD/-T-L) or lacking Trp, Leu, His, and Ade (SD/-T-L-H-A). aa, amino acid. b, c RNA in situ hybridization analysis of GNP5 expression during rice inflorescence development. d, e RNA in situ hybridization analysis of GNP2 expression during rice inflorescence development. PBM primary branch meristem, SBM secondary branch meristem. g BiFC assays showing that GNP2 interacts with Gnp4/LAX2 in rice protoplasts. As a negative control, GNP2 and OsGSK3 did not show any interaction. h LCI assays showing that GNP2 interacts with Gnp4/LAX2 in epidermal cells of tobacco leaves. As a negative control, GNP2 and OsGSK3 did not show any interaction. i Co-IP assays showing the interaction between GNP2 and Gnp4/LAX2 in rice protoplasts. j In vitro pull-down assays showing the interaction between GNP2 and Gnp4/LAX2. Scale bars, 200 μm (b–e), 20 μm (g). In a–j, a representative experiment from three independent experiments is shown. Source data are provided as a Source Data file.
In situ hybridization experiments showed that GNP5 and GNP2 share similar expression patterns with Gnp4/LAX2— displaying weaker expression in primary branches but stronger signals in secondary branches (Fig. 4b–e and Supplementary Fig. 7d). Compared to Nip, the GNP of the CRISPR/Cas9 mutant gnp4/lax2-ko was significantly reduced, with similar phenotypes to gnp2-ko mutants (Supplementary Fig. 8a–d). These collective findings suggest that GNP5, GNP2, and Gnp4/LAX2 may coordinately regulate secondary branch development to influence rice panicle architecture. GNP2 interacted with Gnp4/LAX2 only in the presence of the segment comprising amino acids 139–394 of Gnp4/LAX2 in yeast cells (Fig. 4f). We validated the interaction between GNP2 and Gnp4/LAX2 in plant cells using bimolecular fluorescence complementation (BiFC) and firefly luminescence complementation imaging (LCI) assays (Fig. 4g, h). We then conducted a co-immunoprecipitation (Co-IP) assay by co-transfecting rice protoplasts with GNP2-GFP and HA, FLAG-tagged Gnp4/LAX2 (Gnp4/LAX2-HF). Gnp4/LAX2-HF was detected among the proteins that were precipitated with anti-HA antibody (Fig. 4i). Furthermore, a pull-down assay showed that the GNP2-GST fusion protein interacted with the Gnp4/LAX2-C-MBP (amino acids 139–394) fusion protein in vitro (Fig. 4j). These findings suggest that GNP2 directly interacts with Gnp4/LAX2 in vitro and in vivo.
Given that GNP2 is a kinase, we reasoned that Gnp4/LAX2 might be a substrate for its phosphorylation. Indeed, GNP2 phosphorylated Gnp4/LAX2 in in vitro phosphorylation assays (Fig. 5a). To confirm this phosphorylation in planta, we co-expressed Gnp4/LAX2-HF with either GNP2-GFP or GFP in rice protoplasts. The relative intensity of the phosphorylated band of Gnp4/LAX2-HF significantly increased after co-expression with GNP2-GFP, but not GFP alone (Fig. 5b). Subsequently, liquid chromatography–tandem mass spectrometry (LC-MS/MS) revealed the presence of phosphorylated Thr175 and Thr262 in Gnp4/LAX2-C-MBP when it was incubated with GNP2-GST, but not with GST (Supplementary Figs. 9 and 10). We then constructed the Gnp4/LAX2T175A-C-MBP, Gnp4/LAX2T262A-C-MBP, and Gnp4/LAX2T175,262A-C-MBP fusion proteins to mimic the non-phosphorylated form of Gnp4/LAX2. The phosphorylation levels of Gnp4/LAX2T175A-C-MBP, Gnp4/LAX2T262A-C-MBP, and Gnp4/LAX2T175,262A-C-MBP were reduced, especially Gnp4/LAX2T175,262A-C-MBP, compared to the Gnp4/LAX2-C-MBP control (Fig. 5c), suggesting that Thr175 and Thr262 of Gnp4/LAX2 are the phosphorylation sites targeted by GNP2. Taken together, these results indicate that GNP2 phosphorylates Gnp4/LAX2 in vivo and in vitro.
a, b GNP2 phosphorylates Gnp4/LAX2 in vitro (a) and in vivo (b). c GNP2 phosphorylates the Thr175 and Thr262 of Gnp4/LAX2 in vitro. d, e Analysis of the stability of Gnp4/LAX2 protein in cell-free assays. Gnp4/LAX2-C-GST was incubated with equal amounts of total proteins extracted from WT, gnp2-ko, or GNP2-OE plants in the presence of 10 mM ATP. Gnp4/LAX2-C-GST was detected with anti-GST antibody. WT wild type. f Analysis of the stability of Gnp4/LAX2-C-GST and Gnp4/LAX2T175,262A-C-GST proteins in a cell-free assay. Actin was used as an internal control. Protein levels in the untreated condition were set to 1.00. Immunoblot results were quantified using ImageJ software. g Panicle morphology of Nip, gnp2-ko, gnp4/lax2-ko, and gnp2-ko gnp4/lax2-ko plants. h Statistical results for GNP of Nip, gnp2-ko, gnp4/lax2-ko, and gnp2-ko gnp4/lax2-ko plants. In h, different lowercase letters indicate statistically significant differences at P = 0.05 by one-way ANOVA with Duncan’s multiple-range test, and error bars represent means ± SD. Scale bars, 2 cm (g). In a–f, a representative experiment from three independent experiments is shown. Source data are provided as a Source Data file.
The phosphorylation of substrate proteins is generally associated with protein stability40,41. To investigate the effect of GNP2 on Gnp4/LAX2 stability, we conducted a cell-free degradation assay. Gnp4/LAX2-C-GST was degraded more rapidly when incubated with total proteins from gnp2-ko compared to Nip, but more slowly when incubated with total proteins from GNP2-OE plants (Fig. 5d, e). When we incubated either Gnp4/LAX2T175,262A-C-GST or Gnp4/LAX2-C-GST with total proteins extracted from GNP2-OE plants, Gnp4/LAX2T175,262A-C-GST exhibited faster degradation than Gnp4/LAX2-C-GST (Fig. 5f). To investigate the genetic relationship between GNP2 and Gnp4/LAX2, we generated the gnp2-ko gnp4/lax2-ko plants using CRISPR/Cas9 (Supplementary Fig. 11a, b). The GNP of gnp2-ko gnp4/lax2-ko plants was similar to that of gnp4/lax2-ko plants (Fig. 5g, h), suggesting that Gnp4/LAX2 functions downstream of GNP2 to regulate GNP. Together, these results suggest that GNP2 regulates GNP in rice by phosphorylating Gnp4/LAX2 and promoting its stability.
The GNP2–Gnp4/LAX2 module regulates GNP-related TFs activity
To identify the downstream genes involved in the GNP2-Gnp4/LAX2 pathway, we conducted transcriptome deep sequencing (RNA sequencing [RNA-seq]) of young panicles of Nip, gnp2-ko, and gnp4/lax2-ko plants. We identified differentially expressed genes (DEGs) by applying the criteria of a significant difference (P < 0.05) and an absolute fold-change > 1. Compared to Nip, 1494 DEGs were identified in gnp2-ko plants, including 642 upregulated and 852 downregulated DEGs (Figs. 6a), and 1829 DEGs were identified in gnp4/lax2-ko plants, including 722 upregulated and 1107 downregulated DEGs (Fig. 6b). Among these, 814 DEGs were detected in both comparisons. We defined different gene clusters of DEGs from this analysis: Cluster 1, representing DEGs that were significantly upregulated in gnp2-ko and gnp4/lax2-ko compared to Nip; and Cluster 2, representing DEGs that were significantly downregulated in gnp2-ko and gnp4/lax2-ko compared to Nip (Fig. 6c). We conducted gene ontology (GO) analysis of each cluster to explore the biological processes linked to the GNP2-Gnp4/LAX2 pathway and identified ten significantly enriched terms. A larger proportion of genes in both clusters are associated with transcriptional regulation processes (Fig. 6d), supporting the finding that Gnp4/LAX2 regulates gene transcription and protein–protein interactions39,42.
a, b The volcano plot illustrated the number of DEGs in the young panicle of gnp2-ko (a) or gnp4/lax2-ko (b) compared to Nip. c Clustering analysis of DEGs co-regulated by GNP2 and Gnp4/LAX2. d GO analysis showing the significantly enriched biological pathways. e Gnp4/LAX2 interacted with TFs related to panicle development. Yeast cells were cultured on SD minimal medium lacking Trp and Leu (SD/-T-L) or lacking Trp, Leu, His, and Ade (SD/-T-L-H-A). f The dual-luciferase assays showing the effect of Gnp4/LAX2-HF and Gnp4/LAX2T175,262D-HF on the transcriptional activation activity of TFs in (e) (n = 6). The relative activity of firefly luciferase (LUC) under control of the GAL4-binding element UAS was measured. Renilla luciferase (REN) activity was used as a reference, and VP16 as the positive control. g Panicle morphology of Nip, gnp2-ko, gnp2-ko Gnp4/LAX2-OE, and gnp2-ko Gnp4/LAX2T175,262D-OE plants. h Statistical results for GNP of Nip, gnp2-ko, gnp2-ko Gnp4/LAX2-OE, and gnp2-ko Gnp4/LAX2T175,262D-OE plants. In a, b, d Significant DEGs were identified as those with a P-value (one-way ANOVA test) of differential expression above the threshold (P-value < 0.05, log2(|fold change|) > 1). In f, h, different lowercase letters indicate statistically significant differences at P = 0.05 by one-way ANOVA with Duncan’s multiple-range test, and error bars represent means ± SD. Scale bars, 2 cm (g). Source data are provided as a Source Data file.
We used Gnp4/LAX2-BD as the bait to screen a library comprising genes related to rice yield (generated by the Laboratory of Rice Germplasm, Genomics, and Molecular Breeding, China Agricultural University). Gnp4/LAX2 interacted with many TFs related to panicle development, including LAX1, MOG1, GL6, Ghd2, An-1, and OsMADS15, in yeast cells (Fig. 6e). In a transient transcriptional activity assay, the transcriptional activation activities of LAX1, MOG1, and Ghd2 were significantly enhanced when these TFs, which positively regulate GNP, bound to Gnp4/LAX2 and even more so when they interacted with the phosphorylated form of Gnp4/LAX2T175,262D. By contrast, when the TFs GL6, MADS15, and An-1, which negatively regulate GNP, bound to Gnp4/LAX2, their transcriptional activation activity significantly decreased, and it decreased further when they interacted with phosphorylated Gnp4/LAX2T175,262D. These results indicate that the interactions between Gnp4/LAX2 and different TFs play critical roles in either promoting or inhibiting their transcriptional activation activity, with the phosphorylated Gnp4/LAX2T175,262D generated by GNP2 phosphorylation showing a more pronounced regulatory effect (Fig. 6f).
To investigate whether Gnp4/LAX2 phosphorylation modulates GNP, we constructed transgenic rice plants overexpressing Gnp4/LAX2 or Gnp4/LAX2T175,262D driven by the 35S promoter in the gnp2-ko background (Supplementary Fig. 12a). The GNP was significantly higher in gnp2-ko Gnp4/LAX2-OE vs gnp2-ko plants and even higher in gnp2-ko Gnp4/LAX2T175,262D-OE plants. These results indicate that Gnp4/LAX2 phosphorylation plays an important role in regulating GNP (Fig. 6g, h and Supplementary Fig. 12b). Taking these results together, we propose that the GNP2–Gnp4/LAX2 module regulates the activities of TFs involved in panicle development.
Identification of an elite joint haplotype of GNP5 and GNP2
To investigate the natural variation of GNP5 in 496 rice accessions, we performed haplotype analysis using the variants of GNP5 identified by GWAS. We classified the gene allelic variations into two haplotypes, GNP5-Hap1 and GNP5-Hap2, based on one significant SNP in the promoter region and three significant non-synonymous SNPs in the coding region of this gene (Supplementary Fig. 13 and Supplementary Table 4). The accessions containing GNP5-Hap1 exhibited significantly higher GNP than the accessions containing GNP5-Hap2 in Nanning and Yuxi (Supplementary Table 5). Therefore, we conclude that GNP5-Hap1 is the superior haplotype of GNP5 in terms of GNP.
We performed joint haplotype analysis of GNP2 and GNP5 based on the GNP2-Hap-G (S5779181 locus in the GNP2 promoter is the haplotype with the G base), GNP2-Hap-A (S5779181 locus in the GNP2 promoter is the haplotype with the A base), GNP5-Hap1, and GNP5-Hap2 haplotypes among the 496 rice accessions. We detected four haplotypes in the indica and japonica subpopulations (Types I–IV) (Table 2). The accessions harboring Type I exhibited the highest GNP values in indica and japonica rice grown in Nanning and Yuxi (Table 2). To further confirm that haplotype Type I is the dominant combination of GNP2 and GNP5 in rice germplasm, we used the indica rice Type I accession CX122 and Type II accession CH1275 to construct near-isogenic lines (NILs). We screened the BC4 backcross progeny to obtain NIL-Type IICH1275 and NIL-Type ICX122 (Fig. 7a, b). NIL-Type ICX122 and NIL-Type IICH1275 showed no significant differences in grain length, grain width, and 1000-grain weight (Supplementary Fig. 14a–d), but NIL-Type ICX122 exhibited significantly higher GNP and grain yield per plant than NIL-Type IICH1275 (Fig. 7c–e). These results suggest that Type I is the superior combination of GNP2 and GNP5 in rice germplasm, suggesting that pyramiding these alleles could improve GNP and yield.
a, b The schematic diagram for the construction process and plant morphology of NIL-Type ICX122 and NIL-Type IICH1275. c–e Panicle morphology (c), statistical results for GNP (d), and grain yield per plant (e) of NIL-Type ICX122 and NIL-Type IICH1275. f Haplotype network analysis of GNP5 and GNP2. Or O. rufipogon, trj tropical japonica, tej temperate japonica, aro aromatic. g Geographical distribution of accessions containing different combinations of GNP5 and GNP2. The map was made using the maps package in R. h Frequencies of the GNP5, GNP2, and different combinations of GNP5-GNP2 haplotypes in landraces (LAN) and improved varieties (IMP). In d, e, different lowercase letters indicate statistically significant differences at P = 0.05 by one-way ANOVA with Duncan’s multiple-range test, and error bars represent means ± SD. Scale bars, 20 cm (b), 2 cm (c). Source data are provided as a Source Data file.
To explore the origin and dissemination of the elite Type I haplotype, we conducted haplotype network analysis of GNP2 and GNP5 among 66 Oryza rufipogon wild rice and 379 cultivated rice accessions worldwide, finding that Type I originated from wild rice and was widely present in indica rice (Fig. 7f). We then performed geographic distribution analysis of the four joint haplotypes using 1716 indica rice and 788 japonica rice accessions from the 3010 Rice Genomes (3k) database43 (Supplementary Data 4). Type I was prevalent in indica rice worldwide but relatively uncommon in japonica rice (Fig. 7g and Supplementary Table 6). To investigate whether the Type I haplotype has been utilized in rice breeding, we examined the proportions of the four haplotypes in landraces (LAN) and improved varieties (IMP) within the 3k database (Supplementary Data 5). The findings indicated that Type I has been thoroughly utilized in the breeding programs for aus, aromatic, and indica rice, whereas there remains considerable opportunity for its application in the breeding of temperate and tropical japonica rice varieties (Fig. 7h).
Enhanced expression of GNP5 and GNP2 increases rice yield in the field
To further evaluate the potential of GNP5 and GNP2 for utilization in molecular breeding, we tested the effects of the genotypes at GNP5 and GNP2 on other agronomic traits. Plant height, grain length, grain width, and 1000-grain weight showed no obvious differences between Nip and GNP2-OE plants. However, the number of primary and secondary branches, and tiller number were significantly higher in GNP2-OE plants compared to Nip, resulting in 18.4% and 19.2% increases in grain yield per plant in GNP2-OE1 and GNP2-OE2, respectively (Fig. 8a, b and Supplementary Fig. 15a–h). Similarly, plant height, tiller number, grain length, and 1000-grain weight showed no obvious differences between Nip and GNP5-OE plants, with a slight decrease in grain width, whereas the numbers of primary and secondary branches were significantly higher in GNP5-OE plants than in Nip, resulting in 17.0% and 20.3% increases in grain yield per plant in GNP5-OE1 and GNP5-OE2, respectively (Fig. 8c, d and Supplementary Fig. 16a–h). In field plot yield trials, the grain yield of GNP2-OE1 and GNP2-OE2 plants increased by 11.4% and 10.3%, respectively, compared to Nip (Fig. 8e–g). These results support the notion that increasing the expression of GNP5 and GNP2 can improve rice grain yield.
a, b Plant morphology (a) and statistical results (b) for grain yield per plant of Nip, GNP2-OE1, and GNP2-OE2 plants. c, d Plant morphology (c) and statistical results (d) for grain yield per plant of Nip, GNP5-OE1, and GNP5-OE2 plants. e–g Field trials of Nip, GNP2-OE1, and GNP2-OE2 plants. h A proposed model illustrating GNP5-GNP2-Gnp4/LAX2 module regulating rice GNP. In b, d asterisks indicate statistical significance by two-tailed Student’s t-tests (**P < 0.01, ***P < 0.001, data represent means ± SD). In g, different lowercase letters indicate statistically significant differences at P = 0.05 by one-way ANOVA with Duncan’s multiple-range test, and error bars represent means ± SD. Scale bars, 20 cm (a, c). Source data are provided as a Source Data file.
Discussion
In this study, we found that the GSK3-like kinase GNP2 improves rice yield by regulating GNP. Based on this finding, we propose the following model: During the evolution of rice, artificial selection has led to high and stable yield becoming an important criterion for evaluating rice varieties. Currently, some existing rice varieties have developed a well-established mechanism for high and stable yield. Specifically, when the functional variation site in the GNP2 promoter is a G base, the G base aligns with the conserved G-box motif, which can be recognized by bZIP TFs. This interaction facilitates the binding of GNP5 to the variation site, thereby enhancing the expression of GNP2. The increased expression of GNP2 promotes the phosphorylation and stability of Gnp4/LAX2. The phosphorylated form of Gnp4/LAX2T175,262D regulates the activity of TFs associated with panicle development, which increases GNP and ultimately improves yield. In contrast, when the functional variation site in the GNP2 promoter is an A base, the expression of GNP2 is reduced, leading to a decrease in GNP and consequently affecting the yield (Fig. 8h). The mechanism by which GSK3-like kinases regulate rice GNP at the transcriptional level breaks the limitations of previous research, which primarily focused on post-transcriptional regulation of GSK3-like kinases, and provides a breeding strategy for achieving high and stable rice yields.
Crop yield is influenced by multiple factors, including light, water, temperature, nutrients, and fertilizers. Although elevated GNP2 expression significantly enhances yield in the rice model variety Nip, its limited ecological adaptability in China may restrict the manifestation of its yield-enhancing effects in practical production. Future efforts should focus on localized genetic enhancement through molecular breeding in different local cultivars. In the combined haplotype, Type I of GNP5 and GNP2 significantly increases GNP in both indica and japonica rice, outperforming the other three types. (Table 2). Throughout the rice domestication process in diverse geographical regions, Type I has consistently represented a significant proportion of indica rice. These findings indicate that the favorable phenotype conferred by Type I has been widely leveraged in indica rice across different regions. By contrast, Type I currently constitutes just 20.2% of the japonica rice population (Fig. 7g). The superior GNP2 haplotype constitutes 30.8% of landraces of temperate japonica rice but just 7.5% of improved varieties. In tropical japonica rice, this haplotype represents 36.1% of landraces but 60.7% of improved varieties (Fig. 7h). The superior haplotype GNP5 is present in very high proportions of both landraces and improved varieties of temperate japonica rice. In tropical japonica rice, it accounts for 58.9% of landraces and 55.6% of improved varieties (Fig. 7h). We also examined the genotypes of GNP5 and GNP2 in high-yielding, high-quality rice varieties developed by the Rice Research Institute of the Guangxi Academy of Agricultural Sciences, China, and found that many of these varieties belong to Type I (Supplementary Data 6). These findings point to the potential of introducing Type I into modern rice varieties to improve yield.
BRs are one of the six major classes of plant hormones. These polyhydroxylated steroid compounds play a crucial role in regulating plant growth and development44,45,46,47,48,49,50,51. Recent studies have shown that moderately suppressing the BR signaling pathway can actually increase crop yield. For example, in semi-dwarf wheat (Triticum aestivum), the deletion of the r-e-z genotype block significantly enhances yield by weakening BR signal transduction52. Similarly, the tissue-specific expression of the BRD3 gene in rice can inhibit the BR signaling pathway, thereby promoting the formation of panicle branches and increasing yield13. OsGSK2 is one of the homologous genes of BIN2 in Arabidopsis and negatively regulates the expression of downstream BR-responsive genes in rice. OsGSK2 knockout plants show an increase in grain length, and RNAi plants have both longer and wider grains. In contrast, overexpressing plants typically exhibit shorter grains and reduced thousand-grain weight13,53,54. However, in FLAG-aGSK2 overexpressing plants, the ability to inhibit BR signaling is weaker, and FLAG-aGSK2 primarily accumulates in the pedicels, while significantly less is found in the spikelets. Therefore, compared to the ZH11 variety, there is no significant difference in grain size or weight. Nevertheless, due to the aggregation of multiple small panicles, FLAG-aGSK2 overexpressing plants show a more compact panicle shape. This specific enhancement in grain number leads to a 14.7% to 27% increase in grain yield per plant compared to ZH1113. These findings have revealed the important role of BRs and their associated genes in regulating rice yield. OsGSK1 is another homologous gene of BIN2 from Arabidopsis. GNP2, as an allele of OsGSK1, negatively regulates the BR signaling pathway (Supplementary Fig. 17a–c). Our research indicates that the phosphorylation mediated by GNP2 stabilizes Gnp4/LAX2, thereby regulating the formation of rice panicle architecture and yield. Compared to the Nip variety, GNP2 overexpressing plants exhibit the following characteristics: a significant increase in tiller number and GNP; changes in grain length, width, and 1000-grain weight, though these variations are not statistically significant (Fig. 1i–m; 8a, b and Supplementary Fig. 15a–h). These phenotypic changes are similar to those observed in FLAG-aGSK2 overexpressing plants. It is proposed that the yield increase mediated by GNP2 may primarily be achieved through an increase in grain number.
In rice, the GSK3-like kinase family consists of nine members, which can be divided into 4 subgroups based on their function. The second subgroup includes four members (GSK1 to GSK4), which play an important role in regulating BR signal transduction21. Compared to the ZH11 variety, single and double gene mutants (such as gsk2, gsk1,2, gsk2,3, gsk2,4) show a slight increase in plant height, while gsk2,3,4 and gsk1,2,3,4 mutants have significantly reduced plant height. Although most mutants do not show significant changes in tiller number, gsk1,2,3,4 mutants show a slight increase in tiller number. These results suggest functional redundancy between members of the GSK3-like kinase family24. In our study, compared to the Nip variety, there were no significant differences in plant height and tiller number in gnp2-ko plants (Supplementary Fig. 3e, h), which is consistent with the above conclusion and may be due to functional redundancy between members of the GSK3-like kinase family. In quantitative experiments, we found that GNP2 is highly expressed in the young panicles of rice, which may be related to the significant changes in the grain number of gnp2-ko plants (Fig. 1g, h and Supplementary Fig. 7a, b). bZIP TFs play a central role in plant growth, development, stress response, and metabolic regulation. A total of 89 genes encoding bZIP TFs have been identified and characterized in the rice genome. By aligning the sequences of these OsbZIP proteins, all OsbZIP TFs, except for OsbZIP80, can be classified into ten subfamilies33. Among them, GNP5 (OsbZIP35) belongs to subfamily B, which contains 11 OsbZIP TFs. These TFs not only share highly similar bZIP domains but also possess additional conserved motifs, suggesting they may have similar biological functions. Studies have shown that two members of the B subfamily, RF2a (OsbZIP75) and RF2b (OsbZIP30), when overexpressed in dominant negative mutants, exhibit symptoms similar to rice tungro virus disease; however, overexpressing these two genes enhances rice resistance to the virus55. These results suggest that the OsbZIP TFs in subfamily B may have functional redundancy. In our study, compared to the Nip variety, there were no significant differences in plant height and tiller number in gnp5-ko plants (Supplementary Fig. 4e, h), which may also be due to the functional redundancy among OsbZIP TFs.
Methods
Plant materials, sequencing, and growth conditions
A total of 496 cultivated rice accessions from around the world were analyzed for GNP, including 174 accessions from the core germplasm collection and 322 accessions from the International Rice Molecular Breeding Network56,57. All germplasm materials were planted under conventional cultivation conditions in Yuxi City, Yunnan Province, China in 2015, and in Nanning City, Guangxi Province, China in 2020. The sequence data were obtained from the Rice 3000 rice genomes project58. For haplotype network analysis, sequencing data from 66 wild rice accessions obtained from published datasets were used59.
CRISPR/Cas9 technology was used to generate gnp5-ko, gnp2-ko, gnp5-ko gnp2-ko, and gnp2-ko gnp4/lax2-ko plants. Two target site fragments were cloned into the binary plasmid pHUE411. The recombinant plasmids were then transformed into callus tissue of Nip and gnp2-ko via Agrobacterium-mediated transformation. To construct the GNP5 and GNP2 overexpression plasmids, the full-length coding sequences of GNP5 and GNP2 were respectively amplified from the cDNA of Nip and inserted into the binary vectors PCM1307 and pSuper1300-MYC. The recombinant plasmids were then transformed into Nip. For the construction of the pGNP2:GUS vector, a 2.5-kb promoter region of the GNP2 gene was amplified from Nip and cloned into the vector pMDC162, followed by transformation into Nip. To construct the GNP2-CMG and GNP2-CMA vectors, a 2.5-kb promoter region and the genomic sequence of the GNP2 gene were amplified from Nip and cloned into the vector pMDC163, which was then transformed into gnp2-ko plants. For the construction of the gnp2-ko Gnp4/LAX2-OE and gnp2-ko Gnp4/LAX2T175,262D-OE vectors, the full-length coding sequence of Gnp4/LAX2 was amplified from the cDNA of Nip and inserted into the binary vector pMDC32, and then transformed into gnp2-ko plants. CH1275 was used as the recurrent parent, and NIL-Type ICX122 and NIL-Type IICH1275 were constructed through backcrossing and selfing.
Genome-wide association study
In the R environment, a genome-wide association study was performed using the compressed mixed linear model method from the GAPIT package (version 3), with a total of 4,890,951 SNPs used60. The data underwent quality control, and the minimum allele frequency was set to greater than or equal to 0.05. The genome-wide significance threshold was determined by conducting 1000 permutation tests61. If a region contained more than two consecutive significant SNPs, that region was considered a single association signal, with the SNP having the smallest P-value within the signal considered as the lead SNP. Linkage disequilibrium was analyzed using the standardized disequilibrium coefficient (D′), and the allele frequency squared correlation coefficient (r²) for SNP pairs was determined using the TASSEL 5.0 program62. Subsequently, the Linkage disequilibrium decay distance around the lead SNPs identified in this study was also calculated.
RNA isolation and real-time qPCR assay
Total RNA was extracted from various tissues of rice at different developmental stages using an RNA extraction kit (Aidlab, Beijing, China). According to the instructions of the kit, reverse transcription was performed using HiScript II reverse transcriptase (Vazyme, Nanjing, China). qPCR experiments were conducted following the protocol provided by the manufacturer (TaKaRa, Kyoto, Japan), with three independent RNA samples prepared for biological replication. The rice OsActin (LOC_Os03g50885) was used as the internal reference gene.
Histochemical GUS staining
Different rice tissues were collected from the pGNP2:GUS transgenic lines and incubated in GUS staining buffer containing 5-bromo-4-chloro-3-indolyl-β-D-glucuronide (X-gluc) as the substrate. The rice tissues were stained for 10 h at 37 °C in the dark. After staining, the samples were washed with anhydrous ethanol to remove excess staining solution.
Subcellular localization assay
To investigate the subcellular localization of GNP2, the full-length cDNA of GNP2 was cloned from Nip and inserted into the pSuper1300-GFP vector. The constructed recombinant vector was then transformed into rice protoplasts. Fluorescent signals were observed and captured using an LSM880 confocal laser scanning microscope (Zeiss, Oberkochen, Germany).
Yeast one-hybrid assay
The promoter region of GNP2 was amplified from Nip and cloned into the pLacZi2μ vector to construct the pGNP2(G):lacZ and pGNP2(A):lacZ reporter gene plasmids. The full-length coding sequence of GNP5 was amplified from Nip and cloned into the pB42AD vector to obtain the pB42AD-GNP5 recombinant plasmid. Subsequently, pB42AD-GNP5 was co-transformed with pGNP2(G):lacZ and pGNP2(A):lacZ plasmids into the yeast strain EGY48. The transformed yeast cells were cultured on SD dropout medium containing X-Gal (5-bromo-4-chloro-3-indolyl-β-D-galactopyranoside), and the blue color development was performed according to the protocol provided in the manufacturer’s user guide (Clontech).
Luciferase activity assay in rice protoplasts
To investigate the activity of different types of GNP2 promoters, we amplified the promoter pGNP2(G) from Nip and generated the point mutation pGNP2(A). Both of them were cloned into the pGreenII 0800-LUC vector to generate the reporter plasmids. We amplified and cloned the full-length CDS of GNP5 from Nip into the pSuper1300-MYC vector, thereby constructing the effector plasmid. To evaluate the transcriptional regulatory activity, we amplified and cloned the full-length CDS of LAX1, MOG1, GL6, OsMADS15, An-1, and Ghd2 from Nip and fused them with GAL4BD to construct effector plasmids. Meanwhile, the empty vector LUC and REN were used as reporter genes. To investigate the effect of Gnp4/LAX2 on the transcriptional activity of the aforementioned TFs, the full-length CDS of Gnp4/LAX2 from Nip and Gnp4/LAX2T175,262D generated from point mutation were cloned into the PCM1307 vector as an effector plasmid. The analysis of promoter activity and transcriptional activity was performed in protoplasts extracted from two-week-old rice leaf sheaths of Nip. The transfected protoplasts were incubated at 28 °C in the dark for 16 h, followed by cell lysis. Luciferase activity was detected using the dual-luciferase reporter assay kit (Promega, E1960). The Renilla luciferase (REN) gene was used as an internal control for normalization. Each plasmid combination was tested in at least three independent transformation experiments.
Chromatin immunoprecipitation-quantitative PCR
Samples of inflorescences were collected from wild-type and GNP5-OE plants for chromatin immunoprecipitation analysis. The procedure is as follows: 2 grams of inflorescence samples were cross-linked with a 1% formaldehyde solution for 5 min. The cross-linking reaction was then terminated by adding 0.125 M glycine, and the reaction was allowed to continue for 5 min. The samples were ground to a fine powder in liquid nitrogen to isolate the chromatin, followed by sonication to fragment the chromatin to an average size of approximately 500 bp. The fragmented chromatin solution was pre-incubated with protein A/G agarose beads for 6 h, followed by overnight immunoprecipitation at 4 °C with Flag M2 affinity gel (Sigma, A2220). The precipitated DNA was then recovered and purified, and quantitative analysis was performed using qPCR to assess the enrichment of target genes.
Electrophoretic mobility shift assay
The full-length gene of GNP5 was amplified and cloned into the pGEX-4T-1 vector. The recombinant GNP5-GST protein was incubated with biotin-labeled probes, competing probes, or mutated probes at 25 °C for 20 min. DNA gel shift assays were performed according to the instructions provided in the LightShift Chemiluminescent EMSA Kit (Thermo Fisher Scientific, 20148).
RNA in situ hybridization
Digoxigenin (Roche)-labeled RNA probes, specifically targeting the GNP5 and GNP2 sequences, were synthesized. Rice young panicle tissues were fixed overnight at 4 °C under vacuum in in situ hybridization fixative (containing formaldehyde, glacial acetic acid, and 50% ethanol). The fixed tissues were dehydrated, embedded in Paraplast Plus wax (Sigma-Aldrich, St. Louis, MO, USA), sectioned using a microtome (RM2235; Leica, Wetzlar, Germany), and the sections were mounted onto Poly-Prep slides (Sigma-Aldrich, St. Louis, MO, USA).
Yeast two-hybrid assay
To screen for proteins that interact with GNP2 in rice, the CDS of GNP2 was first amplified and cloned into the pGBKT7 vector. Then, the GNP2-BD, which does not have self-activation activity, was used as a bait to screen the cDNA library from rice young panicles. The screening and plasmid isolation procedures were carried out according to the manufacturer’s manual (Clontech). This experiment was conducted using the yeast strain AH109.
In vitro pull-down assay
To verify the in vitro interaction between GNP2 and Gnp4/LAX2-C(amino acids 139–394), the GNP2 and Gnp4/LAX2-C(amino acids 139–394) were amplified and cloned into the pGEX-4T-1 and pMAL-c5x vectors, respectively. The recombinant plasmids were then transformed into Escherichia coli BL21 cells for expression. Next, GNP2-GST and GST proteins were incubated with glutathione beads (GE Healthcare, cat. no. 17-0756-01) at 4 °C for 2 h, followed by the addition of Gnp4/LAX2-C-MBP protein and continued incubation for 1 h. After elution, the protein samples obtained from the beads were subjected to western blot analysis, using an anti-MBP antibody (Sigma, M6295, dilution 1:4000) to detect Gnp4/LAX2-C-MBP.
Co-immunoprecipitation assay
To verify the in vivo interaction between GNP2 and Gnp4/LAX2, the plasmids Gnp4-HF and GNP2-GFP were co-transformed into rice protoplasts and incubated at 28 °C in the dark for 16 h. Total protein was extracted using 500 µL of IP buffer [containing 150 mM NaCl, 50 mM Tris-HCl (pH 7.5), 0.1% NP-40, 10 mM MgCl₂, 3 mM DTT, and 1 × protease inhibitor mixture], and the lysate was centrifuged at 13,000 × g for 20 min at 4 °C. Then 60 µL of the supernatant was taken as input, and the remaining supernatant was incubated with GFP agarose beads (Chromo Tek) at 4 °C for 2 h. Subsequently, western blot analysis was performed using an anti-HA antibody (Sigma, H3663, diluted 1:3,000) for detection.
Bimolecular fluorescence complementation assay
The coding sequences of GNP2, OsGSK3, and Gnp4/LAX2 were amplified from Nip and cloned into nEYFP or cEYFP fusion plasmids, respectively. After co-transformation of these plasmids into rice protoplasts, the samples were incubated at 28 °C in the dark for 16 h. Fluorescence signals were then detected using a confocal laser scanning microscope.
Split-luciferase complementation assay
The coding sequences of GNP2, OsGSK3, and Gnp4/LAX2 were fused to the C-terminus or N-terminus of luciferase, respectively. These constructs were transformed into Agrobacterium EHA105 and co-infiltrated into N. benthamiana leaves. After 48 h of incubation under normal growth conditions, the leaves were incubated with 1 mM luciferin.
In vitro phosphorylation and liquid chromatography–tandem mass spectrometry assay
The recombinant Gnp4/LAX2-C-MBP protein was incubated with GNP2-GST or GST protein in a reaction buffer (containing 25 mM Tris-HCl, pH 7.5, 10 mM MgCl₂, 1 mM DTT, and 25 mM ATP) at 30 °C for 1 h. GST protein was used as a negative control. The reaction was terminated by adding 5× SDS sample loading buffer. The phosphorylation products were then analyzed by phos-tag SDS-PAGE. Phosphorylation products and input samples were detected using anti-phosphothreonine (Cell Signaling, 9381, diluted 1:500), anti-GST (Sigma, SAB5300159, dilution 1:2000), and anti-MBP (Sigma, M6295, dilution 1:4000) antibodies.
To identify the phosphorylation sites of Gnp4/LAX2, the phosphorylated protein was first excised from the SDS-PAGE gel, then reduced with DTT and alkylated with iodoacetamide. The protein was subsequently digested with trypsin for 12 h at 37 °C, pH 8.5. The analysis was performed by liquid chromatography–tandem mass spectrometry63.
Cell-free degradation assay
The specific procedure for the in vitro protein degradation assay of Gnp4/LAX2 is as follows: First, total proteins were extracted from young panicles using the following buffer: 5 mM Tris-HCl (pH 7.5), 10 mM MgCl₂, 1 mM CaCl2, 3 mM DTT, and 1× protease inhibitor mixture. Then, equal amounts of total proteins from Nip, gnp2-ko and GNP2-OE plants were mixed with recombinant Gnp4/LAX2-C-GST protein and incubated at 30 °C for various time periods, with 10 mM ATP and 50 µM MG132. After the incubation, protein separation was performed using 10% (v/v) SDS-PAGE gel, and Gnp4/LAX2-C protein was detected by anti-GST antibody (Sigma, SAB5300159, dilution 1:2000).
RNA sequencing analysis
For the RNA-seq analysis, approximately 1 g of young panicles at the 3-mm stage from Nip, gnp2-ko and gnp4/lax2-ko varieties were collected, and total RNA was extracted from three biological replicates using TRIzol reagent. The RNA library construction and sequencing were performed by Novogene (Beijing), and sequencing was conducted on the Illumina NovaSeq platform64. The raw data (in fastq format) underwent quality control, processed using FastQC (v.0.11.9), and the cleaned reads were aligned to the Nip reference genome (IRGSP1.0). Differentially expressed genes were selected based on the criteria: P-value < 0.05 and log2(|fold change|) > 1. The expression level heatmap of differentially expressed genes was constructed using TBtools software65.
Haplotype and phylogenetic analysis of GNP2
Based on the GWAS results, we performed haplotype analysis of SNPs with P < 1.0 × 10−4 in the GNP5 and GNP2. In the joint haplotype analysis, we combined the functional loci of GNP5 and GNP2, and compared the phenotypic differences between different haplotypes using Duncan’s multiple range test. To investigate the origin of GNP5-GNP2, we selected 66 wild rice accessions of O. rufipogon and 379 cultivated rice varieties to construct a haplotype network. The definition of different haplotypes was performed using ARLEQUIN v.3.566, and the distance matrix results were displayed in HAPSTAR-0.667. The geographical distribution was mapped on a world map using R software.
Primers
The relevant primers are listed in Supplementary Data 7.
Statistical analyses
Data are presented as mean ± standard deviation with error bars. A two-tailed Student’s t-test implemented in Microsoft Excel 2016 was used to compare differences between two groups, with P < 0.05 considered statistically significant. More than two experimental groups were analyzed by one-way ANOVA combined with Tukey multiple range test in IBM SPSS software (version 21.0).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
RNA-Seq data generated in this work have been deposited in the Genome Sequence Archive database under accession CRA026304 in the National Genomics Data Center, China National Center for Bioinformation. The mass spectrometry proteomics data have been deposited in the ProteomeXchange Consortium under accession PXD068686. Gene sequences of GNP2 and GNP5 are provided in Supplementary Data 8, which are also available at the Rice Annotation Project database (https://rapdb.dna.affrc.go.jp/). Source data are provided with this paper.
References
Xing, Y. Z. & Zhang, Q. F. Genetic and molecular bases of rice yield. Annu. Rev. Plant Biol. 61, 421–442 (2010).
Sasaki, T. & Burr, B. International rice genome sequencing project: the effort to completely sequence the rice genome. Curr. Opin. Plant Biol. 3, 138–141 (2000).
Zhang, Z. Y. et al. Fine mapping and cloning of the grain number per-panicle gene (Gnp4) on chromosome 4 in rice (Oryza sativa L.). Agr. Sci. China 10, 1825–1833 (2011).
Guo, T. et al. GRAIN SIZE AND NUMBER1 negatively regulates the OsMKKK10-OsMKK4-OsMPK6 cascade to coordinate the trade-off between grain number per panicle and grain size in rice. Plant Cell 30, 871–888 (2018).
Ashikari, M. et al. Cytokinin oxidase regulates rice grain production. Science 309, 741–745 (2005).
Kurakawa, T. et al. Direct control of shoot meristem activity by a cytokinin-activating enzyme. Nature 445, 652–655 (2007).
Wu, B. et al. Suppressing a phosphohydrolase of cytokinin nucleotide enhances grain yield in rice. Nat. Genet. 55, 1381–1389 (2023).
Komatsu, K. et al. LAX and SPA: Major regulators of shoot branching in rice. Proc. Natl. Acad. Sci. USA 100, 11765–11770 (2003).
Komatsu, M. et al. The LAX1 and FRIZZY PANICLE 2 genes determine the inflorescence architecture of rice by controlling rachis-branch and spikelet development. Dev. Biol. 231, 364–373 (2001).
Chen, Q. et al. Translational repression of FZP mediated by CU-rich element/OsPTB interactions modulates panicle development in rice. Plant J. 110, 1319–1331 (2022).
Komatsu, M. et al. FRIZZY PANICLE is required to prevent the formation of axillary meristems and to establish floral meristem identity in rice spikelets. Dev 130, 3841–3850 (2003).
Taguchi-Shiobara, F. et al. A loss-of-function mutation of rice DENSE PANICLE 1 causes semi-dwarfness and slightly increased number of spikelets. Breed. Sci. 61, 17–25 (2011).
Zhang, X. X. et al. Enhancing rice panicle branching and grain yield through tissue-specific brassinosteroid inhibition. Science 383, adk8838 (2024).
Han, Y. C. et al. Natural variation in MORE GRAINS 1 regulates grain number and grain weight in rice. J. Integr. Plant Biol. 66, 1440–1458 (2024).
Liu, J. H. et al. Ghd2, a CONSTANS-like gene, confers drought sensitivity through regulation of senescence in rice. J. Exp. Bot. 67, 5785–5798 (2016).
Xue, W. Y. et al. Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat. Genet. 40, 761–767 (2008).
Lu, S. J. et al. Overexpression of a transcription factor OsMADS15 modifies plant architecture and flowering time in rice (Oryza sativa L.). Plant Mol. Biol. Rep. 30, 1461–1469 (2012).
Huang, X. H. et al. Genome-wide association study of flowering time and grain yield traits in a worldwide collection of rice germplasm. Nat. Genet. 44, 32–39 (2012).
Cui, D. et al. Genomic insights on the contribution of introgressions from Xian/Indica to the genetic improvement of Geng/Japonica rice cultivars. Plant Commun. 3, 100325 (2022).
Zhang, Q. et al. Natural variation of RGN1a regulates grain number per panicle in japonica rice. Front. Plant Sci. 13, 1097622 (2022).
Youn, J. H. & Kim, T. W. Functional insights of plant GSK3-like kinases: multi-taskers in diverse cellular signal transduction pathways. Mol. Plant 8, 552–565 (2015).
Li, C. X., Zhang, B. & Yu, H. GSK3s: nodes of multilayer regulation of plant development and stress responses. Trends Plant Sci. 26, 1286–1300 (2021).
Koh, S. et al. T-DNA tagged knockout mutation of rice OsGSK1, an orthologue of Arabidopsis BIN2, with enhanced tolerance to various abiotic stresses. Plant Mol. Biol. 65, 453–466 (2007).
Liu, D. P. et al. Diversification of plant agronomic traits by genome editing of brassinosteroid signaling family genes in rice. Plant Physiol. 187, 2563–2576 (2021).
Xiao, Y. H. et al. GSK2 stabilizes OFP3 to suppress brassinosteroid responses in rice. Plant J. 102, 1187–1201 (2020).
Huang, Y. S. et al. Ribonuclease H-like gene SMALL GRAIN2 regulates grain size in rice through brassinosteroid signaling pathway. J. Integr. Plant Biol. 64, 1883–1900 (2022).
Hu, J. L. et al. Rice stripe virus suppresses jasmonic acid-mediated resistance by hijacking brassinosteroid signaling pathway in rice. PLoS Pathog. 16, e1008801 (2020).
Sun, S. Y. et al. Natural selection of a GSK3 determines rice mesocotyl domestication by coordinating strigolactone and brassinosteroid signaling. Nat. Commun. 9, 2523 (2018).
Gao, X. Y. et al. Rice qGL3/OsPPKL1 functions with the GSK3/SHAGGY-like kinase OsGSK3 to modulate brassinosteroid signaling. Plant Cell 31, 1077–1093 (2019).
Ying, J. Z. et al. TGW3, a major QTL that negatively modulates grain length and weight in rice. Mol. Plant 11, 750–753 (2018).
Xia, D. et al. GL3.3, a novel QTL encoding a GSK3/SHAGGY-like kinase, epistatically interacts with GS3 to produce extra-long grains in rice. Mol. Plant 11, 754–756 (2018).
Hu, Z. J. et al. A novel QTL qTGW3 encodes the GSK3/SHAGGY-like kinase OsGSK5/OsSK41 that interacts with OsARF4 to negatively regulate grain size and weight in rice. Mol. Plant 11, 736–749 (2018).
Nijhawan, A. et al. Genomic survey and gene expression analysis of the basic leucine zipper transcription factor family in rice. Plant Physiol. 146, 323–324 (2008).
Choi, H. I. et al. ABFs, a family of ABA-responsive element binding factors. J. Biol. Chem. 275, 1723–1730 (2000).
Li, Z. Y. et al. The transcription factor bZIP68 negatively regulates cold tolerance in maize. Plant Cell 34, 2833–2851 (2022).
Yoo, M. J., Albert, V. A., Soltis, P. S. & Soltis, D. E. Phylogenetic diversification of glycogen synthase kinase 3/SHAGGY-like kinase genes in plants. Bmc. Plant Biol. 6, 3 (2006).
Ahmar, S., Zolkiewicz, K. & Gruszka, D. Analyses of genes encoding the Glycogen Synthase Kinases in rice and Arabidopsis reveal mechanisms which regulate their expression during development and responses to abiotic stresses. Plant Sci. 332, 111724 (2023).
Tabuchi, H. et al. LAX PANICLE2 of rice encodes a novel nuclear protein and regulates the formation of axillary meristems. Plant Cell 23, 3276–3287 (2011).
Zhang, Z. Y. et al. Gnp4/LAX2, a RAWUL protein, interferes with the OsIAA3–OsARF25 interaction to regulate grain length via the auxin signaling pathway in rice. J. Exp. Bot. 69, 4723–4737 (2018).
Zeng, R. et al. Natural variation in a type-A response regulator confers maize chilling tolerance. Nat. Commun. 12, 4713 (2021).
Li, J. F. et al. The GSK3-like kinase BIN2 is a molecular switch between the salt stress response and growth recovery in arabidopsis thaliana. Dev. Cell 55, 367–380 (2020).
Huang, Y. et al. Evolution and conservation of polycomb repressive complex 1 core components and putative associated factors in the green lineage. Bmc. Genomics 20, 533 (2019).
Wang, W. S. et al. Genomic variation in 3,010 diverse accessions of Asian cultivated rice. Nature 557, 43–49 (2018).
Clouse, S. D. Brassinosteroids. Arabidopsis Book 9, e0151 (2011).
Tong, H. N. & Chu, C. C. Functional specificities of brassinosteroid and potential utilization for crop improvement. Trends Plant Sci. 23, 1016–1028 (2018).
Fang, N. et al. SMALL GRAIN 11 controls grain size, grain number and grain yield in rice. Rice 9, 64 (2016).
Hong, Z. et al. A rice brassinosteroid-deficient mutant, ebisu dwarf (d2), is caused by a loss of function of a new member of cytochrome P450. Plant Cell 15, 2900–2910 (2003).
Zhou, Y. et al. GNS4, a novel allele of DWARF11, regulates grain number and grain size in a high-yield rice variety. Rice 10, 34 (2017).
Tanabe, S. et al. A novel cytochrome P450 is implicated in brassinosteroid biosynthesis via the characterization of a rice dwarf mutant, dwarf11, with reduced seed length. Plant Cell 17, 776–790 (2005).
Zhu, X. L. et al. Brassinosteroids promote development of rice pollen grains and seeds by triggering expression of carbon starved anther, a MYB domain protein. Plant J. 82, 570–581 (2015).
Bai, M. Y. et al. Functions of OsBZR1 and 14-3-3 proteins in brassinosteroid signaling in rice. Proc. Natl. Acad. Sci. USA 104, 13839–13844 (2007).
Song, L. et al. Reducing brassinosteroid signalling enhances grain yield in semi-dwarf wheat. Nature 617, 118–124 (2023).
Tong, H. N. et al. DWARF AND LOW-TILLERING acts as a direct downstream target of a GSK3/SHAGGY-like kinase to mediate brassinosteroid responses in rice. Plant Cell 24, 2562–2577 (2012).
Lyu, J. et al. Control of grain size and weight by the GSK2-LARGE1/OML4 pathway in rice. Plant Cell 32, 1905–1918 (2020).
Dai, S. H. et al. Transgenic rice plants that overexpress transcription factors RF2a and RF2b are tolerant to rice tungro virus replication and disease. Proc. Natl. Acad. Sci. USA 105, 21012–21016 (2008).
Zhang, H. L. et al. A core collection and mini core collection of Oryza sativa L. in China. Theor. Appl. Genet. 122, 49–61 (2011).
Yu, S. B. et al. Molecular diversity and multilocus organization of the parental lines used in the International Rice Molecular Breeding Program. Theor. Appl. Genet. 108, 131–140 (2003).
Alexandrov, N. et al. SNP-Seek database of SNPs derived from 3000 rice genomes. Nucleic Acids Res. 43, D1023–D1027 (2015).
Huang, X. H. et al. A map of rice genome variation reveals the origin of cultivated rice. Nature 490, 497–501 (2012).
Wang, J. B. & Zhang, Z. W. GAPIT version 3: boosting power and accuracy for genomic association and prediction. Genom. Proteom. Bioinform. 19, 629–640 (2021).
Guo, H. F. et al. Differentiation, evolution and utilization of natural alleles for cold adaptability at the reproductive stage in rice. Plant Biotechnol. J. 18, 2491–2503 (2020).
Bradbury, P. J. et al. TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23, 2633–2635 (2007).
Liu, Z. Y. et al. Plasma membrane CRPK1-mediated phosphorylation of 14-3-3 proteins induces their nuclear import to fine-tune CBF signaling during coldresponse. Mol. Cell. 66, 117–128. e5 (2017).
Li, Y. P. et al. The CRY2-COP1-HY5-BBX7/8 module regulates blue light-dependent cold acclimation in Arabidopsis. Plant Cell 33, 3555–3573 (2021).
Chen, C. J. et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol. Plant. 13, 1194–1202 (2020).
Excoffier, L. & Lischer, H. E. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 10, 564–567 (2010).
Teacher, A. & Griffiths, D. HapStar: automated haplotype network layout and visualization. Mol. Ecol. Resour. 11, 151–153 (2011).
Acknowledgements
This work was funded by the Biological Breeding-National Science and Technology Major Project (2023ZD0406803 to Zhanying Z.), National Natural Science Foundation of China (32572301 and 32272123 to Zhanying Z.), Chinese Universities Scientific Fund (2025TC142 to Zhanying Z.), and the 2115 Talent Development Program of China Agricultural University.
Author information
Authors and Affiliations
Contributions
Zhanying Z. conceived and designed the study. Q.H. carried out most of the experiments. Q.H., H.X., Zhikun Z., H.G., X.Z., and X.W. analysed the data. Yong Z., L.M., Z.M., P.X., and R.Z. performed part of the experiments. Q.H. and Zhanying Z. wrote the manuscript. Yawen Z., Y.P., D.L., X.S., J.L., H.Z,. and Z.L. provided critical guidance for the revision of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Chengqiang Ding, Damian Gruszka, Hongning Tong, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hu, Q., Zhao, Z., Ma, L. et al. Natural variation of GNP2 enhances grain number to benefit rice yield. Nat Commun 16, 8848 (2025). https://doi.org/10.1038/s41467-025-64564-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64564-y