Abstract
Chikungunya virus (CHIKV) is a mosquito-borne alphavirus that causes febrile illness and acute or chronic arthritis. Most therapeutics are still in the pre-clinical stage. In this study, we report the isolation of two neutralizing antibodies, C34 and C37, from a convalescent patient and investigate their mechanisms of action. Both C34 and C37 exhibit high neutralizing activities in vitro and demonstrate protective effects against CHIKV in a female mouse model. Our functional and structural studies reveal a mechanism that inhibits multiple stages of the virus infection cycle. Both antibodies bind with high affinity to an epitope spanning E2, E1, and the connecting β-strands, facilitating intra- and inter-virion crosslinking. Cryo-EM structures additionally identify a minor patch located beneath the E3 binding site on E2, which is allosterically exposed upon E3 dissociation during virus maturation. Functional and structural data further suggest that binding to the CHIKV receptor, Mxra8, is obstructed due to a clash between the antibodies and the stalk region of Mxra8. Our results highlight the potential of antibody-based therapeutics against CHIKV and elucidate the mechanisms of monoclonal antibody protection.
Similar content being viewed by others
Introduction
Chikungunya virus (CHIKV), a member of the Alphavirus genus within the Togaviridae family, is the most common mosquito-borne arthritogenic alphavirus infecting humans1,2. It is prevalent in tropical and subtropical regions worldwide3. CHIKV infection can lead to symptoms such as myalgia, high fever, severe and chronic polyarthritis, and even death4,5,6,7.
Like other alphaviruses, CHIKV is an enveloped virus of 65–70 nm in diameter. It possesses a single-stranded, positive-sense RNA genome approximately 11.8 kb in length, consisting of two open reading frames (ORFs)1,8,9. The N-terminal ORF encodes four nonstructural proteins, nsP1-4, which are essential for virus replication, immune evasion, and protein modification10,11. The C-terminal ORF encodes four structural proteins: capsid (C), envelope (p62–E1), initially synthesized as a polyprotein and cleaved into C, p62, 6K, and E1. The p62 protein is further cleaved into E3 and E2 by furin-like proteases in the trans-Golgi Network (TGN), generating the mature form of CHIKV1,8,9. The mature viral core consists of 240 copies of capsid proteins packaging a linear genome RNA molecule, surrounded by a lipid bilayer with 240 E1–E2 heterodimers embedded and assembled into 80 trimeric spikes with T = 4 icosahedral symmetry12. E3 acts as a chaperone for the proper folding of p62 and prevents E2-E1 conformational change during Golgi processing13,14,15. The E2 protein comprises three domains (DA, DB, and DC), with domain B located outermost on the virion surface, domain A at the center of each trimeric spike, and domain C close to the viral membrane. E2 is responsible for host receptor recognition, with domains DA and DB being primary targets of neutralizing antibodies16,17,18,19,20,21. The E2 protein prevents the premature exposure of the fusion loop, ensuring that the virus remains non-infectious until it encounters an acidic environment, such as that in the endosome, which triggers the conformational changes required for fusion. The E1 protein is a class II fusion protein composed of three ectodomains (Domain I, DI; Domain II, DII; and Domain III, DIII) with a fusion loop located at DII. E1 forms a heterodimer with E2 and lies proximal to the viral membrane, with its fusion loop hidden beneath the E2 domain B under neutral pH16,22,23,24.
Mxra8, a dual-immunoglobulin-like domain molecule, is the bona fide cell entry receptor for CHIKV, engaging both E2 and E1 proteins, as revealed by X-ray crystallography and cryo-EM studies17,25,26,27,28. Following receptor binding, the CHIKV virion is internalized into an endosome, where acidic conditions induce a conformational change in the E1–E2 heterodimer, exposing the E1 fusion loop29,30. The insertion of the fusion loop into the endosomal membrane facilitates membrane fusion, leading to nucleocapsid disassembly and viral RNA release. After translation and replication of the genome, newly synthesized virions bud at the plasma membrane31.
Currently, at least two vaccines, Ixchiq and Vimkunya, are approved by the FDA for human use against CHIKV infection32. No antiviral therapeutics have been approved thus far. While vaccines offer population-wide protection and are essential for preventing infection, mAbs have unique advantages in specific therapeutic contexts, such as post-exposure prophylaxis, treatment for immunocompromised individuals, and the potential for personalized medicine. Additionally, monoclonal antibodies (mAbs) can target highly conserved epitopes, offering protection against emerging variants. Protective mAbs are widely regarded as promising and feasible approaches for the prevention and treatment of viral infections, as evidenced by their efficacy against SARS-CoV-2 in recent years33. It is crucial to identify CHIKV-specific neutralizing antibodies and elucidate their functionalities, epitopes, and neutralizing mechanisms. Previous research has shown that effective neutralizing antibodies against CHIKV target various epitopes on E2 and E134,35,36,37,38,39,40. However, the structural basis for their neutralizing mechanisms, virus specificity, and how these antibodies affect CHIKV’s interaction with its receptor remains poorly understood.
In this study, we isolated and characterized two CHIKV-specific neutralizing mAbs, C34 and C37, which showed potent efficacy both in vitro and in vivo. We determined their structures in complex with CHIKV VLPs, revealing a unique virus-specific epitope on the E2 protein and a common mechanism for Mxra8 blockade. This highlights a critical site for antibody-mediated neutralization, offering valuable insights for developing therapeutic antibodies and vaccines against CHIKV.
Results
Isolation of CHIKV-E reactive human mAbs
We obtained a panel of mAbs from a convalescent patient of CHIKV infection. Peripheral blood mononuclear cells (PBMCs) were isolated from a blood sample and subjected to antigen-reactive single memory B cell sorting using a recombinant CHIKV E1–E2 fusion protein (denoted as EFL) as bait. With this strategy41, we sorted a total of 82 CHIKV E-reactive IgG+ B cells from approximately 1 million PBMCs. From these sorted B cells, we performed single B cell RT-PCR and successfully recovered 16 mAbs from 12 different B cell lineages based on the germline gene usage of VH.
Characterization of mAbs targeting CHIKV- EFL
We generated human IgG1 plasmid constructs from the V–D–J sequences for recombinant expression of the isolated mAbs and assessed their binding to EFL using ELISA. Nine of the sixteen mAbs bound tightly to the antigen (Fig. S1a). Each of these nine mAbs was genetically distinct from the others (Fig. S1b). We also tested their binding to the E1–E2 heterodimer as well as the constituent 6K-E1 and E2 proteins using flow cytometry. Two (C19 and C59) and four (C31, C37, C47 and C52) of the mAbs bound to soluble 6K-E1 and E2, respectively, while the remaining three mAbs (C18, C34, and C48) only bound to the E1–E2 heterodimer (Figs. 1a and S2). It is worth noting that although C37 shows weak binding to the E2 protein, its binding ability to the E1–E2 heterodimer is significantly stronger. This may be due to the more stable conformation of the E1–E2 heterodimer, which could result in differences in the epitopes between the E2 monomer and the E1–E2 dimer. We further quantitatively assessed the binding kinetics of the corresponding Fab molecules with the E1–E2 heterodimer using a surface plasmon resonance (SPR) assay. Four of the nine Fab molecules (C34, C37, C48, and C59) demonstrated sub-nanomolar KD values (Figs. 1c and S3).
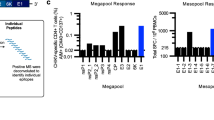
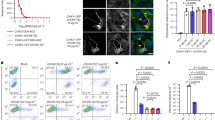
a Heat map displaying cell cytometry-based binding of sorted mAbs to full-length E protein (E-FL), p62–E1, and E2 of CHIKV, as well as full-length E proteins of BFV, VEEV, and ONNV. The intensity of the heatmap corresponds to the percentage of positive cells. b Dose-dependent neutralization curves and a table of IC50 values for the indicated mAbs. Data represent mean ± s.d. of three technical replicates of two biological replicates. c Representative binding kinetics of C34 (left) and C37 (right) Fab molecules to the CHIKV E protein, as derived from the BIAcore 8K system. E proteins were captured on chips. The KD values are calculated as geometric mean ± s.d. of three biological replicates. d, e Pre- and post-attachment assays for IgG and Fab forms of C34 (d) and C37 (e). PRNT50 values were determined by curve fitting of 3-fold serially diluted relative infection values using nonlinear regression. Data represent the mean ± s.d. from three technical replicates. f Egress inhibition assays of C34 and C37. Both antibodies have inhibiting activity for the viral egress. The significance of the identical concentration of C34 and C37 IgG are compared with isotype. p-values for C34 IgG: p = 0.0013, 0.0006 and 0.0004 for 10, 1 and 0.1 μg/ml. p-values for C37 IgG: p = 0017, 0.0008 and 0.0045 for 10, 1 and 0.1 μg/ml. g Fusion from without (FFWO) assays of C34 and C37 IgG. The significance of the identical concentration of C34 and C37 IgG are compared with isotype. p-values for C34 IgG: 0.0011, 0.0019, 0.0025, 0.0036 for 100, 10, 1, and 0.1 μg/ml. p-values for C37 IgG: 0.0013, 0.0043, 0.1199, 0.7240 for 100, 10, 1, and 0.1 μg/ml. f, g Data are the mean ± s.d. from three biological replicates. Significance was determined by one-way ANOVA with Dunnett’s posttest compared to the isotype control. (**p < 0.01, ***p < 0.001, ****p < 0.0001).
To evaluate cross-reactivity, we performed flow cytometry-based binding tests of the obtained mAbs with E proteins from other alphaviruses. Only five of the nine mAbs showed cross-reactivity with o’nyong-nyong virus (ONNV) E protein, which has the highest sequence similarity with CHIKV-E (Figs. 1a and S2). No cross-reactivity was detected with E proteins of Barmah Forest virus (BFV), Venezuelan equine encephalitis virus (VEEV), Mayaro virus (MAYV), or Ross River virus (RRV) (Fig. S2).
Potent in vitro neutralizing activity of mAbs C34 and C37
To quantitatively assess the neutralizing potency of the mAbs, we performed a FACS-based assay in Vero cells. Two of the mAbs, namely C34 and C37, displayed superior inhibitory activity with IC50 values lower than 100 ng/ml (Fig. 1b). We further conducted pre- and post-attachment neutralization assays to investigate the viral entry inhibition activities of these two antibodies. In brief, either the IgG or Fab forms of C34 or C37 were added before or after virion attachment to cells at 4 °C. In the pre-attachment assay, both C34 IgG and C37 IgG effectively neutralized the infection, with PRNT50 values of 16.61 and 77.56 ng/ml, respectively, comparable to previous in vitro neutralizing assays. In the post-attachment assay, C34 and C37 IgG also exhibited good neutralizing activities, though with moderately decreased PRNT50 values of 48.69 ng/ml and 163.5 ng/ml, respectively. Interestingly, the Fab forms of C34 and C37 also showed comparable neutralizing activities to their IgG counterparts. The PRNT50 values for the pre-attachment assay were 19.23 ng/ml and 79.55 ng/ml, respectively, and for the post-attachment assay, 97.03 ng/ml and 727.2 ng/ml, respectively (Fig. 1d, e). This phenotype indicates that the attachment-inhibiting activities are strongly associated with the Fab regions of the two mAbs.
Given the high neutralizing activity of C34 and C37 IgG observed in the post-attachment assay, we further explored the fusion inhibition and budding suppression capabilities of these two antibodies. We first employed the fusion-from-without (FFWO) assay to investigate whether these antibodies could inhibit the fusion of the virus with host cells or the ability of the virus to spread between cells. Our results showed that both C34 and C37 IgG exhibited notable membrane fusion inhibition activity. Notably, C37 demonstrated a strong dose-dependent effect, while C34’s fusion inhibition activity did not show a clear dose dependency. This could be attributed to the fact that, at all experimental concentrations tested, C34 maintained consistently high inhibition levels.
We also investigated the budding suppression activity of these two antibodies. For this, we incubated the virus with Vero cells (MOI = 1) at 37 °C for 2 h, followed by six washes with PBS. The antibodies, diluted in DMEM containing 2% serum and 25 mM NH4Cl, were then added and incubated at 37 °C for an additional 6 h. The virus RNA content in the supernatant was measured by qPCR. The results showed that, after 6 h, both C34 and C37 exhibited significant budding suppression activity, with the inhibition strength showing a clear dose-dependent trend.
Evaluation of in vivo protection efficacies of C34 and C37 mAbs
The protective effects of the mAbs C34 and C37 were evaluated using an immunodeficient mouse model. We also included C59 IgG as a negative control. For the therapeutic study, groups of naïve 8–12-week-old mice were injected with PBS or 10 mg/kg of anti-CHIKV mAbs C34 or C37 24 h before subcutaneous inoculation with 100 LD50 of CHIKV 181/25 strain (Fig. 2a). All mice treated with the neutralizing mAbs were completely protected against lethality, whereas those in the PBS group and 80% of mice in C59 group succumbed to CHIKV infection within five days (Fig. 2b). Both mAbs significantly reduced footpad swelling compared to the PBS-treated animals, although relative weight change was not significant after day 2 post-administration of mAbs or PBS (Fig. 2c, d). On day one in the therapeutic group, a dramatic increase in footpad swelling was observed, which was related to the inoculation of the virus through the footpad.
a–d In vivo therapeutic efficacy of C34, C37, and C59 (non-neutralizing control) mAbs against CHIKV infection in mice. a Mice were administered 10 mg/kg of the indicated mAbs by intraperitoneal injection 24 h after a lethal CHIKV challenge. Dpi days post-infection. b Survival rates of each group. The number of mice in each group are indicated in brackets. Survival rates of mAb-treated mice were compared to the PBS group. p = 0.039, 0.0039, and 0.5846 for C34, C37, and C59 groups against the PBS group, respectively. c Percentage of weight change in each group of mice. p = 0.0493 for day1 C34 vs PBS group. d Footpad swelling was measured as the percentage change in footpad size in each group of mice. For day 2, p = 0.0169 for C34. For day 3, p = 0.0002 and 0.0370 for C37 and C59, respectively. For day 4, p = 0.0002 and 0.0001 for C34 and C3,7, respectively. e–h In vivo prophylactic efficacy of C34, C37, and C59 mAbs against CHIKV infection in mice. e Mice were administered 10 mg/kg of the indicated mAbs by intraperitoneal injection 24 h before a lethal CHIKV challenge. f Survival rates of each group, with the number of mice indicated in brackets. Survival rates of mAb-treated mice were compared to the PBS group. p = 0.034, 0.0034, and 0.0528 for C34, C37, and C59 groups against the PBS group, respectively. g Percentage of weight change in each group of mice. For day 3, p = 0.0287 for C34. For day 4, p = 0.0013 and 0.0017 for C34 and C37, respectively. For day 5, p = 0.0215 and 0.0112 for C34 and C37, respectively. For day 6, p = 0.0233 and 0.0088 for C34 and C37, respectively. h Footpad swelling was measured as the percentage change in footpad size in each group of mice. For day 3, p = 0.0294 and 0.0190 for C34 and C37, respectively. For day 4, p = 0.0312 and 0.0478 for C34 and C59, respectively. For day 5, p = 0.0144 for C59. For c, d, g, h, data represent mean ± s.d. of n = 5 biological repeats. For b, f, statistical analysis was performed using the Mantel-Cox log-rank test: **p < 0.01, ***p < 0.001, ****p < 0.0001. For c, d, g, h, statistical analysis was performed using two-way ANOVA and Dunnett’s multiple comparisons test: **p < 0.01, ***p < 0.001, ****p < 0.0001.
For the prophylactic study, the experimental setup was identical to the therapeutic group, except that the administration of mAbs or PBS occurred one day before virus inoculation (Fig. 2e). Mice given PBS exhibited significant footpad swelling and body weight reduction 4 days post-inoculation (dpi) and showed a 100% mortality rate by 8 dpi. In contrast, mice administered with C34 or C37 demonstrated a 100% survival rate and minimal signs of body weight change or footpad swelling (Fig. 2f–h). These results clearly demonstrate that mAbs C34 and C37 have exceptional therapeutic and prophylactic efficacy against CHIKV infection.
Overall structures of CHIKV-specific neutralizing mAbs complexed with VLP or E heterodimer
To further investigate the structural basis of CHIKV neutralization by mAbs, we generated high-resolution cryo-EM reconstructions of CHIKV VLPs in isolation as well as complexed with C34 or C37 Fab molecules (Fig. 3a–d, and S4 and S5). We also generated the structure of an infectious CHIKV virion particle with E3 associated for comparison. All four above structures are first solved with T = 4 icosahedral symmetry imposed and determined to a moderate resolution ranging from 3.63 Å to 4.35 Å. To further improve the resolution by overcoming the heterogeneity of the sample and by correcting the defocus gradient from various parts of the VLP, we divided the icosahedral reconstructions into blocks of 2-fold (2f), 3-fold (3f), and 5-fold (5f) symmetry using a block-based algorithm42 and separately calculated their reconstructions with subparticles re-extracted, followed by localized refinement. Due to the resolution limitations of the local refinement process, where the density of the Fab and VLP complex remains unresolved in certain regions and a small portion of the side-chain densities cannot be fully reconstructed, we further analyzed the cryo-EM structure of the C34 Fab in complex with the E1–E2 fusion protein and the crystal structure of the C37 Fab in complex with the E1–E2 fusion protein as complementary structural analysis tools. The resolutions of these two structures were 3.05 Å and 3.20 Å, respectively, with side chain density better resolved. An overview of reconstructions and their calculated resolutions are provided in the Supplement Table 1.
a, c Cryo-EM density maps of C34 (a) and C37 (c) Fab are shown as equatorial cross-sections. Surface representations of the entire particle are depicted as insets at the bottom left of each graph. Symmetry axes are indicated by arrows. The E1, E2, and capsid proteins of CHIKV VLPs and C34/C37 Fabs are colored the same as those indicated in (b, d), and this applies to all figures unless otherwise noted. b, d Surface-rendered representations of the asymmetric unit reconstructed with Fab molecules. e, f Reconstructions of the 2f (e) and 5f (f) blocks of VLPs complexed with C34 Fab. g, h Distances in Ångströms between Cα atoms of terminal heavy chain constant domains (CH1) of neighboring Fab molecules are shown as dashed lines. E2 from q3 and i3 spikes are indicated in the 2f blocks (g). All spikes around the 5f (h) vertex are q3 spikes. The glycoproteins E1 and E2 are faded for clarity. i, j Reconstructions of the 2f (i) and 5f (j) blocks of VLPs complexed with C37 Fab. k, l Distances in Ångströms between Cα atoms of terminal heavy chain constant domains (CH1) of neighboring Fab molecules are shown as dashed lines. E2 from q3 and i3 spikes are indicated in the 2f blocks (k). All spikes around the 5f (l) vertex are q3 spikes. m Models illustrating contact interfaces between Fab constant domains of neighboring Fab molecules: two C34 Fab molecules around the 2f vertex (left) and C37 Fab molecules around the 5f vertex (right). The closest distance binding sites are indicated by red circles. The residues involved in crosslinking are indicated by black short arrows. n–p Dynamic light scattering (DLS) was used to assess the dispersity and particle diameter change of VLPs complexed with the indicated mAbs in Fab or IgG forms. o Diameter in nm showing C34, C37, or CHK-263 IgG at different mAb to Ag molar ratios. Data represent mean ± s.d. of three independent experiments. p Diameter in nm showing C34, C37, or CHK-263 Fab molecules at different molar ratios to Ag. Data represent mean ± s.d. of three independent experiments.
The reconstructed infectious virion structure is in good agreement with the VLP structures determined here and previous structures43,44,45,46,47,48. The densities of various domains of E1, E2, and E3 are overall well defined (Fig. S7). We identified additional blob densities above the E2 domain B and the β-ribbon connecting E2 domains A and B (namely the A–B strand) at both the 5f and 2f axes, revealing the definite shape of Fabs on both VLP structures complexed with C34 and C37 (Fig. 3b, d).
By comparing the overall VLP structures of various Fab molecules bound, upon binding of C34 and C37 Fab molecules, no significant change in the diameter of the VLP nor the structure of the E1–E2 heterodimer was observed compared to the VLP alone structure, indicating that the binding of these Fab molecules did not alter the overall virus assembly (Fig. S8). We constructed models for the CHIKV E1, E2, and E3 proteins and the VH/VL regions of the Fab molecules for each asymmetric unit within the corresponding block-based reconstruction, performing a rigid-body fit for the CH1/CL regions in locally filtered maps to account for resolution decay in the Fab region. On each reconstruction with Fab molecules bound, we could observe densities in all 60 equivalent binding positions. However, at the 5f vertex, Fab densities for the C37 Fab’s CH1/CL region were smeared, but well defined for the VH/VL region, likely due to the flexibility of the peripheral CH1/CL region. In contrast, C34 Fab densities were highly smeared around the 5f vertex. To determine whether this broken electron density was due to partial occupancy or high flexibility, we conducted localized 3D classification on subparticles around the 5f vertex for C34 Fab-VLP local reconstruction without imposing symmetry. By varying T values from 4 to 35, we consistently obtained a single class of particles with over 95% showing five Fab molecules bound simultaneously around the 5f vertex. A minority population showed 1–4 Fab molecules bound around the same vertex. The model of C34 Fab was fitted into the densities around the 5f vertex without apparent clashes between neighboring Fab molecules. We speculated that the congested space at the 5f vertex may cause the simultaneous binding of five C34 Fab molecules to be unstable, leading to a higher turnover rate, although full occupancy is theoretically possible.
Within an asymmetric unit, all four C34 Fab molecules bound to the E1–E2 heterodimer in similar conformations with an average RMSD value of each pairwise comparison of less than 0.6. For C37 Fab molecules, only the VH/VL region bound to the E1–E2 heterodimer in an identical manner, while the CH1/CL region exhibited varying degrees of swing relative to the VH/VL region, presumably to accommodate as many Fab molecules as possible at the 5f and 2f vertices. Notably, an identical spatial distribution of neighboring Fab molecules was observed across different symmetry counterparts. The S26 residue of the C34 light chain was in close proximity to the CH domain of a neighboring Fab, while the R218 residue of the C37 heavy chain was near the Q156 residue of a neighboring Fab’s light chain (Fig. 3m, n). This compact distribution of Fab molecules around both the 5f and 2f axes suggests a weak but widespread crosslinking effect among neighboring Fab molecules, potentially enhancing multivalent effects and neutralization potency.
Inter- and intra-virion crosslinking effects may contribute to the neutralization potency of C34 and C37
Previous research on EEEV antibodies has demonstrated that the intravirion and intervirion crosslinking capacity of IgG, determined by the spatial distribution of cysteine residues at the end of each CH1 domain of the corresponding Fab and the binding orientation of Fab molecules on the virus surface, plays a critical role in neutralizing potency49. We measured the distances between Cα atoms of the C224 residues at the end of CH1 domains from neighboring Fab fragments around the 2f and 5f vertices. The distances between neighboring Fab molecules around the 5f vertex were 47.9 Å (Fig. 3h) and 30.7 Å (Fig. 3l) for C34 and C37 Fab molecules, respectively. For C37 Fab molecules around the 2 f vertex, all measured distances were within the 50 Å theoretical radius50 for intravirion crosslinking, while slightly above 50 Å for C34 (Fig. 3g, k). Next, we tested the potential for intervirion crosslinking by measuring the hydrodynamic diameters of VLP particles using dynamic light scattering (DLS). A dramatic increase in VLP diameter from ~60 nm to ~800 nm was observed with the addition of an excess amount of the IgG form of C34 or C37, whereas no aggregation occurred when their Fab counterparts were added (Fig. 3o, p). We further investigated the aggregation of VLPs induced by C34 and C37 in both IgG and Fab forms with a titration of the molar ratio between antibody and VLP. In brief, we incubated different molecular weight forms of Fab or equivalent Fab (i.e., each IgG molecule containing two Fab molecules) with VLPs for the same period, and then measured the particle size by DLS. We found that the IgG forms of C34 and C37 caused VLP aggregation when the molar ratio with VLPs exceeded 25:1 and 120:1, respectively (i.e., the equivalent ratio of Fab to E protein dimer was approximately 0.3:1 and 1.5:1). In contrast, neither of the Fab forms of the antibodies induced VLP aggregation at a molar ratio of 400:1. We also observed that the IgG form of C34 and C37 behaved differently as CHK-263 did in inducing the aggregation of VLPs. The reason is that C34 and C37 bind to VLP with heavy chain adjacent to the virion surface, leaving C-terminal tails of CH pointing outwards, while in CHK-263, the heavy chain is positioned oppositely and pointing downwards. Together, these results indicate that IgG forms of C34 and C37 can cross-link virus particles, whereas the Fab forms of these antibodies cannot. Nonetheless, in pre-attachment neutralization assays, C34 Fab (PRNT50 = 19.23 ng/ml) and C37 Fab (PRNT50 = 79.55 ng/ml) exhibited nearly the same potency as their corresponding IgG forms, suggesting the existence of additional neutralization mechanisms beyond virion aggregation induction.
C34 and C37 target an immunodominant patch allosterically exposed by E3 dissociation on E2
To circumvent the complexity arising from the compact and crosslinked distribution of Fab molecules on VLP vertices, we determined the monovalent complex structures of C34 and C37 Fab molecules with the E1–E2 heterodimer. The cryo-EM structure of the C34 Fab-E1–E2 complex was resolved to a resolution of 3.05 Å, and the crystal structure of the C37 Fab-E1–E2 complex was determined at 3.20 Å. These high-resolution structures enabled precise model building of the epitopes (Fig. 4a, b). The C34 Fab binds to an epitope that spans E2 domain B, the β-ribbon connecting domain B and domain A (referred to as the A–B strand), and two residues, K69 and K61, from E1. The heavy chain of the C34 Fab engages the CHIKV E1–E2 heterodimer with a buried surface area of 573 Ų, while the light chain buries an area of 622 Ų, resulting in a total buried interface of 1195 Ų. C34 mainly utilized both heavy and light chain variable regions to interact with CHIKV E protein, including hydrogen bonds between D106CDRH3-E247E2, Y108CDRH3-D250E2, Y109CDRH3-L248E2, Y109CDRH3-G249E2, Y49CDRL2-D250E2, S53CDRL2-D250E2, L54CDRL2-K252E2, R28CDRL1-E208E2, S31CDRL1-R198E2, G68L-L210E2, D31CDRH1-K69E1, and N105CDRH3-K61E1. By contrast, the C37 Fab utilized mainly CDRH1, CDRH2, and CDRL3 to recognize an epitope with more interactions with E2 domain B, partially overlapping with the C34 epitope. The C37 Fab binds to E2 with a buried interface of 941 Å2, mainly achieved by abundant hydrogen bonds including G92CDRL3-T212E2, S32CDRL1-T212E2, N93CDRL3-K215E2, W94CDRL3-L210E2, R91CDRL3-T196E2, D50CDRL2-K233E2, Y60CDRH2-R198E2, Y35CDRH1-R198E2, Y34CDRH1-D250E2, and Y34CDRH1-K252E2 (Fig. 4a–d).
a, b Overall structures of the C34–E1–E2 cryo-EM structure (a) and the C37-E crystal structure (b). Fab molecules and CHIKV E proteins are shown in ribbon representation, with interactions having a buried surface area (BSA) larger than 10 Ų depicted as sticks. Hydrogen bonds are indicated by dashed lines. c, d Surface representations showing the electrostatic potential of epitopes on the E1–E2 dimer for C34 (c) and C37 (d). Positive, negative, and neutral charged residues are colored blue, red, and white, respectively. Non-interacting residues on E1–E2 are colored the same as the corresponding text labels, respectively. e Superposition of the E2 loop from various structures: infectious virions (green), VLP (red), C34:VLP complex (blue), C34:VLP complex (pink), C34–E1–E2 complex (magenta), and C37–E1–E2 crystal structure (tint). The side chain flip and displacement distances of D250 and K252 from the infectious virion and VLP structures are indicated by stick representation and arrows. The density of the VLP alone structure is shown in red surface representation. f Multiple sequence alignment of the E2 protein covering the epitopes of C34 and C37 mAbs together with representative mAbs targeting similar (CHK-263 and SKW24) or distinct (506.A08 and 506.C01) epitopes across seven alphaviruses: CHIKV (OQ567725.1), ONNV (OQ238859.1), RRV (MK028844.2), MAYV (PP505832.1), VEEV (U34999.1), WEEV (PP747354.1), and EEEV (MK028842.1). Antibody contact residues are delineated below the alignment. Antibodies discovered in this study are represented as squares, antibodies from previous publications are represented as circles.
The large footprint areas of the C34 and C37 Fab molecules on CHIKV correlate with their high binding affinity and potent neutralizing activity. Previous studies have identified two patches—spanning residues 180-199 on the A–B strand of E2 and residues 209–223 on the C-C’ strand—as dominant and broadly neutralizing hotspots across all alphaviruses due to their relatively low sequence variation and conformational stability. Antibodies including CHK-265, 506.A08 and 506.C01 all target these two patches35,36,51,52,53,54. C34 and C37 Fab molecules interact intensively with these two patches as well. Additionally, a third patch, spanning residues 245NAELGDRK252 on CHIKV E2, is involved in the recognition by C34, C37, and some other previously characterized antibodies, including CHK-26355 for CHIKV and SKW2456 for WEEV (Fig. 4f). Even though, the exact epitopes of these four antibodies are not fully overlap. This interaction is largely dependent on the residues N245 and D250, which are also the interacting residues with CHK-263 (Fig. S9). However, these residues are not conserved in sequence or conformation among alphaviruses, except for ONNV, which aligns with the limited cross-reactivities of the two mAbs. Notably, N245 is an insertion mutant and not a canonical glycosylation site, as no glycans were observed near N245. Docking simulations of C34 and C37 Fab molecules onto equivalent epitopes of E proteins from various alphaviruses revealed clashes primarily in this additional patch, with no significant conflicts observed on the A–B and C-C’ strand epitopes.
Previous research on alphavirus particle preparations has demonstrated the gradual release of cleaved E3 from budded viruses during alphavirus maturation43,57. By superimposing the C34 and C37 Fab molecules onto an E3-associated structure of CHIKV, we observed distinct clashes between the Fab molecules and E3. We incubated C34 Fab with freshly collected CHIKV infectious particles and prepared cryogenic samples to analyze the interaction between C34 Fab molecules and E3. As expected, C34 Fab minimally bound to E3-associated infectious particles, with definite densities of E3 associated with the A–B strand. Only at very high contour levels (σ = 0.015) did we observe weak densities of C34 Fab molecules, likely binding to a minority population of particles with E3 released (Fig. S10).
As illustrated by the structure of the infectious virion, the cleaved E3 associates with the A–B strand mainly through hydrophobic interactions, stabilizing E2 DB under acidic conditions during Golgi processing. Comparing the structures of infectious particles with fully matured VLP particles, we found that the strand composing the additional C34/C37 epitope undergoes structural rearrangement after E3 dissociation. Specifically, the loop comprising this epitope adopts a “squeezed” conformation when E3 is associated. Upon E3 dissociation, the side chain of the conserved residue K252 flips out to the opposite direction, pointing towards the solvent environment, along with a mainchain displacement of about 5.1 Å. This flip and shift twist the upstream residues to minimize the loop’s energy, causing a significant orientation change of the D250 side chain, making it exposed. We examined this loop’s conformational change under various local environments, including VLPs with or without antibody binding, as well as the C34/C37 Fab–E1–E2 structures (cryo-EM and crystal structure), and observed negligible backbone movements, indicating that the conformational change is induced by E3 dissociation rather than antibody recognition (Fig. 4e). The exposure of D250 is central to the recognition by C34 and C37 mAbs.
C34 and C37 block the Mxra8 attachment by interfering with the stalk region
Although multiple pathways contribute to the neutralizing potency of C34 and C37, the obvious differences in neutralizing efficacy observed in pre- and post-attachment experiments imply a possible role of virus attachment blockade for these two antibodies. We first conducted a VLP blocking test with flow cytometry and observed a complete blockade of VLP attachment to Mxra8-expressing cells (Fig. S11). However, docking Mxra8 into Fab:VLP structures showed no direct steric hindrance between either C34 or C37 Fab molecules and Mxra8 within a single E1–E2 trimeric spike, as Mxra8 and the Fab molecules approach the E2 domain B from opposite directions. Considering the T = 4 icosahedral symmetry of CHIKV, however, VH and VL regions of the Fab molecules are either in close proximity (within 3 Å) or overlapping with Mxra8 D1 on the adjacent spike when we tried to align the Fab-bound structure with the Mxra8-bound one. We then analyzed three distinct interfaces between adjacent spikes (Fig. 5a). These interfaces are originated from two types of interactions between trimers: (1) the interface between two neighboring quasi-three-fold trimers (interface 1 and 2) in which a small angle is formed between two spikes, so that the distance (between Mxra8 and Fab molecules) in interface 2 is larger than that in interface 1 as the two molecules are away from the five -fold axis. (2) The interface between quasi-three-fold and icosahedral-three-fold spike (interface 3 and 4) in which the edges of two spikes are strictly antiparallel to each other. As the density of the stalk region located in the C-terminal of hMxra8 D1 was not resolved in the crystal structure28, we used the mMxra8-bound VLP structure to analyze the possibility of direct steric hindrance between the stalk region of the receptor and the antibodies17. Docking Mxra8 and the corresponding Fab molecules onto these three interfaces revealed apparent steric hindrance at all these interfaces for C34 Fab (Fig. 5b). For C37 Fab, however, the overlap was to a lesser extent than that of C34. Considering the stalk region residues downstream being highly glycated and the hydrophobicity of these modeled residues, we speculate potential steric hindrance may exist, though no direct overlapping of C37 fab and Mxra8 was observed in this alignment (Fig. 5c).
a Diagrammatic representation of the organization of E1, E2 subunits, Mxra8, and Fab molecules according to T = 4 icosahedral symmetry. Dashed triangles indicate two types of spikes: i3 spikes in black and q3 spikes in red. Black capital letters indicate the asymmetric units around the 5f vertex, delineated as black solid pentagons. White numbers delineating independent binding subunits within the asymmetric unit (ASU) follow the sequence identical to Fig. 3b, d. Symmetry axes are designated by a pentagon (5f), a triangle 3f, and a diamond 2f. b, c Direct alignment of C34 (b), C37 (c) bound CHIKV-E spike against mMxra8 bound spike. Three types of contact interfaces from (a) are illustrated, respectively, for each Fab. The first nine residues in the stalk region of mouse Mxra8 are colored black in stick representation. Docking of C34 or C37 variable regions into corresponding interfaces, accommodated with Mxra8, to show direct overlapping. d Experimental setup for studying the blockage of Mxra8 binding by Fab molecules. Twin strep-tags were fused to the N- or C-termini of Mxra8 D1 or D2, replaced with an equal length of G4S linker or truncated as illustrated. The binding of different constructs of Mxra8 was determined via flow cytometry. α-strep antibodies were used for surface staining. e APC-labeled anti-Mxra8 were examined as an indicator of Mxra8 binding capacity on CHIKV-E trimer transiently expressed in 293 T cells preincubated with 3f serially diluted antibodies. The four set-ups correspond to the constructions in (d). mAbs of C34, C37, CHK-263, and SKT-20 were colored with red, blue, orange, and gray, respectively. The fluorescence intensities of 293 T cells without antibodies incubated were colored yellow. Representative results of three biological replicates were plotted. A dashed line indicates the threshold of fluorescence intensity of Mxra8-bound and non-bound cells. f quantification of the relative binding percentage of (e). Relative binding percentage is defined as the change in the ratio of Mxra8-bound cells over all cells under the corresponding mAbs concentration. Data are presented as mean ± s.d. from three independent experiments. Statistical analysis are performed using two-way ANOVA and Dunnett’s multiple comparisons test for 100 μg/ml of C34, C37, and CHK-263 against SKT-20. p-values are indicated correspondingly.
To test this hypothesis, we conducted the receptor blocking tests. The CHIKV E spike was expressed on the cell surface, and the twin-strep-tagged Mxra8 proteins (with an intact stalk or a GS-substituted stalk) were used for CHIKV E binding in the presence or absence of the indicated antibodies by flow-cytometry. The anti-strep signal serves as an indicator of Mxra8 binding to CHIKV E trimer on the cell surface (Fig. 5d). In order to examine the influence of the stalk region on antibody blocking efficiency, we interchangeably expressed E trimer on the surface of the cell and tested Mxra8 binding activity change upon antibody incubation. The reduction of bound Mxra8 serves as an indicator for blocking activities of tested antibodies. We placed a twin-strep tag on the N- or C-terminal of MXRA8 and tested its feasibility as an indicator of E protein binding. As the VLP-bound MXRA8 molecules on the cell surface always adopt an “inverted” orientation with a 48-a.a.-long stalk region protruding from the C-terminal of the D1 Ig domain, a simple deletion of the stalk region in the C-terminal tagged set-up may reduce the avidity of the tag, so we replace the stalk region with a GS-linker of the same length in the C-terminal tagged set-up. Without antibody incubation, all four configurations in Fig. 5d. shows definite binding between soluble MXRA8 and cell surface-expressed E trimers, indicating a feasible set-up (Fig. 5d). In the N-terminal tagged configuration, deleting the stalk region resulted in a significant loss of blocking by C34, C37 and CHK-263 mAbs while no significant Mxra8 binding ratio change by a fusion loop targeting neutralizing antibody SKT2056, indicating the existence of interaction between the above three mAbs with stalk region of CHIKV E, and this interaction contribute to blocking of Mxra8 binding.,. In the C-terminal tagged configuration, however, replacing the stalk region with a GS-linker can only partially weaken the blocking activities of the three antibodies rather than fully abolish the blocking effect (Fig. 5e). This observation is in accordance with the assumption that the stalk region interferes with the three antibodies as substituting the stalk region into smaller side chain residues may reduce the potential clash between antibodies and stalk region. In both setups with original stalk region and N- or C-terminal strep labeled, Mxra8 binding to E trimer was blocked by C34, C37, and CHK-263 in a dose-dependent manner (Fig. 5f), while SKT20 showed no blocking activity. For the constructions with a stalk region deleted or replaced by GS-linker, the blocking activities are markedly reduced. This finding aligns with our structural analysis, indicating that C34 and C37 Fab molecules block Mxra8 binding by clashing with the stalk region of CHIKV E.
In summary, three aspects of the neutralizing-protecting mechanism have been identified for antibodies targeting the novel epitope spanning CHIKV E2 and E1 proteins. Firstly, the bound antibodies facilitate the crosslinking of adjacent virions, leading to the aggregation of virus particles and inhibiting their dispersion. Secondly, these antibodies block the attachment of virions to the cell surface by interfering with the interaction between the virus and the stalk region of Mxra8. Thirdly, the antibodies inhibit the exposure of the fusion loop, which is essential for membrane fusion and viral entry. By stabilizing the E2 and E1 protein conformation, the antibodies prevent the conformational changes required for the fusion of the viral and cellular membranes. This combined mechanism effectively neutralizes and protects against CHIKV infection by targeting multiple stages of the viral life cycle.
Discussion
Neutralizing antibodies play a critical role in antiviral immunity, showcasing significant therapeutic potential. Several antibody treatments for viruses such as SARS-CoV-2, respiratory syncytial virus (RSV), and Ebola virus (EBOV) have received clinical approval58,59,60. CHIKV, a mosquito-borne pathogen, causes substantial morbidity through acute febrile illness and chronic arthralgia. The immune response, particularly neutralizing antibodies, is essential for controlling CHIKV infection. Identifying protective epitopes on arthritogenic alphaviruses and thoroughly investigating the mechanisms of protective mAbs can enhance our understanding of humoral immune responses and facilitate the iterative design of vaccines.
In this study, we isolated a panel of naturally occurring human mAbs from a CHIKV convalescent patient and identified two potent neutralizing antibodies, C34 and C37. These mAbs exhibited strong neutralizing activity in vitro and provided significant protection in an immunodeficient mouse model, highlighting their therapeutic potential against CHIKV.
Our structural analysis revealed that C34 and C37 are two antibodies targeting an overlapping epitope positioned on the E2 domain B and the β-ribbon connecting domain A and domain B. On alphaviruses, E2 domain B is located at the most membrane-distal region, and has been demonstrated as a main target of neutralizing antibodies. Most E2 domain B targeting antibodies are involved in interfering with multiple stages of viral infection, which is also the case for C34 and C37. Two epitopes spanning residues 180–199 and 209–223 are common among E2 domain B targeting antibodies, like CHK-265, 506.A08, and 506.C01. A less characterized epitope spanning residues 245-252 was also covered by some antibodies against various alphaviruses, like CHK-26355 against CHIKV, SKW2456 against WEEV, and in this study, C34 and C37.
Among the antibodies with existing structural analyses, the highest epitope overlap with human antibodies C34 and C37 is observed with the murine antibody CHK-263. Based on a 4–5 Å medium-resolution structure, it is inferred that the neutralization mechanism of CHK-263 is inhibition of fusion by locking E1 and E2. Despite the significant epitope overlap between C34, C37, and CHK-263, structural and functional assays reveal at least two key differences: (1) CHK-263’s binding is heavily dependent on the light chain, which is located on the virus membrane-facing side, whereas for C34 and C37, the heavy chain faces the membrane, with the light chain at the far end. A significant outcome of this is that CHK-263 cannot mediate cross-linking of different viral particles through bivalency effects, while C34 and C37 can, as their C-terminal end of the CH region points towards the solvent region, facilitating viral particle cross-linking. This finding is consistent with the results of DLS (Fig. 3o). (2) The post-attachment activity of C34 and C37 in their Fab form declines less dramatically compared to their IgG form, unlike most domain B-targeting antibodies, including CHK-263. This suggests that antibodies with similar binding epitopes may exhibit distinct neutralization mechanisms. Further investigation into the mechanisms of C34 and C37 reveals that these antibodies block receptor binding by forming a spatial hindrance with the stalk region of the receptor Mxra8, which aligns with the phenotypic observation that these antibodies act during the attachment phase in functional studies. In further analysis, we found that the neutralizing mechanisms of C34 and C37 mAbs are multifaceted. The precise positioning of Fab molecules on the virus surface ensures full occupancy and potential crosslinking between neighboring Fab molecules of adjacent trimeric spikes, contributing to their neutralizing mechanism. The neutralizing mechanisms employed by C34 and C37 do not act in isolation but cooperatively to enhance overall neutralization efficacy. We speculate this cooperative action ensures a robust and multi-layered defense against CHIKV.
Our structural analysis also revealed that C34 and C37 bind to a less-studied binding site on the E2 protein, a site normally covered by the E3 protein in immature virions. The binding of C34 and C37 is considered a “mimic” of E3. A direct observation is that C34 or C37 Fab has high binding affinity with E1–E2 heterodimer, while only a weak interaction was observed for binding with the constituent E1 or E2 protein (Figs. 1a and S2). However, structural studies revealed that the binding sites of these two Fab molecules are mainly located on the E2 protein. We speculate that the binding of C34 or C37 has an effect of stabilizing the β-ribbon connecting DB and DA of E2, which is believed to inhibit the exposure of the fusion loop.
Alphavirus envelope E1 is a class II fusion protein with homologs in many different enveloped viruses, such as flaviviruses, rubella viruses, and bunyaviruses. A shared feature of class II fusogens is the exposure of the fusion loop triggered by low pH in endosomes, leading to the insertion of the fusion loop into the target membrane. During virion maturation in the TGN, a mildly acidic pH environment requires a protective mechanism. In alphaviruses, E3 acts as a chaperone to prevent early exposure of the fusion loop by stabilizing E2-DB, while the counterpart in flaviviruses is the pr moiety. We revealed an allosteric effect of E3 dissociation, resulting in the exposure of a latent binding site for neutralizing antibodies on E2. This mechanism is similar to that observed in flaviviruses, where the 150-loop rearranges upon prM dissociation at neutral pH61. This suggests the potential for latent neutralizing hotspots in flaviviruses, warranting further study.
The blocking of Mxra8 attachment by clashing with the stalk region is another significant mechanism employed by C34 and C37. Similar to the mouse antibody CHK-263, these antibodies interfere with virus attachment. Our previous study demonstrated that the 48-amino-acid stalk region of Mxra8 is critical for CHIKV entry into cells. In this study, we showed that the stalk region could also serve as a neutralization hotspot for antibodies.
Taken together, these observations suggest that mAbs with outstanding neutralizing potency derived from convalescent CHIKV patients employ different working mechanisms compared to those derived from VLP vaccination. The latter typically neutralize viruses at specific stages of the viral replication cycle, including attachment, entry, fusion, or egress. In contrast, the former may block multiple pathways throughout the entire viral lifecycle. This observation is not only true for C34 and C37 mAbs but is also consistent with several prior studies.
In addition to elucidating the mechanisms of these two mAbs, we generated high-resolution reconstructions for CHIKV VLPs and virions with E3 attached, providing a more precise model of CHIKV particles. These high-resolution models, along with models of various alphaviruses, can guide further structural studies to identify critical determinants of antigenicity and inform the rational design of vaccines against alphaviruses.
A limitation of this study is related to structural determination. Despite similar sample preparation and data processing methodologies, we could determine the icosahedral structure of native virions to 3.6 Å but not better than 4 Å for VLP:Fab.
complex. The higher resolution structure obtained with block-based methodology comes at the expense of uncovering heterogeneities arising from Fab binding. Thorough analysis of “imperfect” subparticles using cryo-EM analysis proved laborious and unfruitful in this research. However, recent studies on the intrinsic heterogeneity of alphaviruses using subtomogram analysis may shed light on future studies about neutralizing antibody mechanisms at the VLP level.
In summary, our study clearly defines the function and working mechanisms of a group of fully mature virion-specific neutralizing mAbs, and reveals that the blockage targeting the stalk region of Mxra8 is the common characteristic of mAbs in this class. The findings can inform future vaccine and immunotherapy design against CHIKV and other related alphaviruses.
Methods
Ethics
Peripheral blood was obtained from a female subject in P.R.China in 2017 following written informed consent. The participant was compensated. The gender of participants was not considered in the study design. The protocols were approved by the Committee on Ethics of the Institute of Microbiology, Chinese Academy of Sciences (approval no. HP-APIMCAS2025230). The animal experiments in this study were approved by the Committee on the Ethics of Animal Experiments of the Institute of Microbiology, Chinese Academy of Sciences (approval no. APIMCAS2023067).
Cells, viruses, and animals
Vero cells (ATCC, CCL-81) and HEK293T cells (ATCC CRL-3216) were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM, Gibco) supplemented with 10% heat-inactivated fetal bovine serum (FBS, Gibco) and 1% penicillin–streptomycin (p/s) at 37 °C in a 5% CO2 atmosphere. Expi293F (A14527, Thermo Fisher Scientific) cells were cultured in SMM 293-TII-N medium (Sino Biological Inc.) with 1% p/s at 37 °C in a 5% CO2 atmosphere. Vero cells (#ATCC CCL-81) were purchased from ATCC. Sf9 cells (#11496015) were purchased from Invitrogen. High Five cells (#B85502) were purchased from Invitrogen. All cell lines routinely tested negative for mycoplasma contamination. All cell lines were authenticated with PCR assays.
CHIKV (strain CHIKV-181/25) was sourced from previous stocks and passaged in Vero cells. All procedures involving authentic CHIKV were conducted in a biosafety level 3 (BSL-3) laboratory at IMCAS. Virus titers were determined using the 50% tissue culture infectious dose (TCID50) method.
Eight- to twelve-week-old female Ifnar1-KO mice (Cat. NO. NM-KO-18038) were purchased from Shanghai Model Organisms Center, Inc. and maintained in an animal biosafety level 3 (ABSL3) facility at IMCAS, with use approved by the National Health Commission of China. All animals were allowed free access to water and a standard chow diet and provided with a 12-h light and dark cycle (temperature, 20–25 °C; humidity, 40–70%).
Isolation of CHIKV EFL-specific single memory B cells and determination of variable region sequences
PBMCs from a CHIKV convalescent patient were isolated as previously described60. Briefly, PBMCs were incubated with his-tagged CHIKV EFL at 400 nM and subsequently stained with anti-human CD3, CD16, CD235a, CD19, CD27, CD38, IgG, and anti-His. Antigen-specific memory B cells (CD3−, CD16−, CD235a−, CD38−, CD19+, CD27+, IgG+, and His+) were sorted into 96-well PCR plates using a BD FACSAria III flow cytometer, then subjected to gene amplification and sequencing. Variable regions were amplified by 5’ RACE and nested PCR, and the full-length mAbs were generated by linking variable region genes to a human IgG1 constant region.
Monoclonal antibody and Fab production and digestion
mAbs were expressed in HEK-293F cells following transient transfection for 96 h. Supernatants containing secreted antibodies were collected by centrifugation at 12,000×g for 1 h and applied to a HiTrap protein A HP column. Eluted antibodies were further purified using a Superdex 200 gel-filtration column.
Fab fragments were produced in two ways. For functional studies, including SPR, pre- and post-attachment assays, and flow cytometry-based blocking assays, Fab molecules were obtained by papain digestion of mAbs overnight and purified using a protein A column to remove digested Fc fragments. The flow-through containing Fab molecules was pooled and concentrated to working concentrations. For crystal screening and cryo-EM studies, Fab molecules were generated by recombinant expression in HEK-293F cells with a His-tag fused to the C-terminal of the heavy chain. Supernatants were applied to a HisTrap FF column and eluted with a linear gradient of imidazole, then further purified using a Superdex 200 column.
Protein expression and purification
The E1–E2 heterodimer was expressed as a fusion protein using the Bac-to-Bac baculovirus expression system with an identical construction method as described before28. Shortly saying, recombinant bacmids sequentially encoding E3 (1–64 a.a.), E2 (65–405 a.a., corresponding to 1–341 a.a. of E2), and E1 (1–412 a.a.) were transfected and passaged in Sf9 cells to produce baculovirus stock, which was used to express protein in High5 cells for 48 h. The 4× GGGGS linker is inserted between the E2 and the E1 domains of above mentioned construction. A twin-strep tag following a His tag was fused and expressed in the C-terminal of the above construction. Supernatants were collected and purified using a HisTrap FF column followed by Superdex 200 gel-filtration. The His-tag was not removed. For the formation of the E1–E2-C34 Fab complex. The elution from HisTrap was subjected to a tandem Q-SP column under 100 mM NaCl, 20 mM Tris, pH 8.0. The target protein was bound to the SP column and was eluted with a 100–1000 mM NaCl gradient. E1–E2 fusion protein was further purified by Superdex 200 gel-filtration under 150 mM NaCl, 20 mM Tris, pH 8.0.
For E1–E2–C37 Fab complex, the purified E1–E2 fusion protein was incubated with recombinant expressed C37 Fab at a molar ratio of Fab:E1–E2 complex = 1.5:1, under 20 mM Tris, pH 7.5, 100 mM NaCl on ice for 30 min and subjected to further purification by gel-filtration chromatography using a Superdex 200 column.
VLP production and purification
CHIKV VLPs were produced with modifications to previously described methods28. In brief, genes encoding CHIKV capsid and envelope proteins (strain 119067; GenBank: APA34057) were cloned into the pCAGGS vector. After 48 h of expression, supernatants were centrifuged at 11,000×g for 10 min to remove cell debris and concentrated approximately 200-fold using Amicon Ultra-6 100 kDa cutoff centrifugal concentrators (Millipore). Concentrated supernatants were subjected to discrete sucrose gradient (20–60% w/v) ultracentrifugation in PBS at 100,000×g, 4 °C for 4 h. Fractions containing CHIKV VLP were carefully collected and buffer-exchanged or diluted to appropriate concentrations.
Flow cytometry-based binding assay
For binding activity assays of selected mAbs with subunits of CHIKV E protein, HEK293T cells were transiently transfected with pEGFP-N1-CHIKV-E full length, pEGFP-N1-CHIKV-6k-E1, or pEGFP-N1-CHIKV-p62 plasmids. At 24 h post-transfection, cells were detached and aliquoted into 96-well plates. After washing with PBS, cells were stained with excess mAbs (5 μg/ml) at 4 °C for 30 min. After washing three times with PBS, anti-human IgG-Fc/APC was added and incubated for another 30 min. Cells were analyzed on a BD FACS Canto II after washing three times with PBS.
For cross-binding activity assays against E proteins of BFV, VEEV, RRV, MAYV, and ONNV, the same procedure was followed with corresponding E proteins cloned into a pCAGGS vector.
SPR
SPR measurements were performed at 25 °C in single-cycle mode on a BIAcore 8 K system with CM5 sensor chips. All proteins were exchanged to a running buffer of 1× PBS, pH 7.5, supplemented with 0.05% Tween-20 by gel filtration. A blank channel served as a negative control. CM5 biosensor chips were first immobilized with α-His antibodies at 10,000–20,000 response units. Serially diluted E proteins were then injected, followed by capture of 2f diluted mAbs. After each cycle, sensors were regenerated with Gly-HCl. Kd values for each mAb to E protein were calculated by fitting to a 1:1 binding model.
In vitro neutralization assay
CHIKV was passaged in Vero cells. Cells were seeded at 6 × 104 cells per well in 24-well culture plates and cultured overnight. Serially diluted mAbs were mixed with DMEM supplemented with 1% FBS and incubated with CHIKV (103 PFUs/ml) for 1 h at 37 °C. Virus and DMEM with 1% FBS served as blank controls. Cells were washed twice before infection with 350 μl per well of the virus-antibody mixture. After incubation for 1 h at 37 °C, DMEM with 1% FBS was added to a total volume of 1 ml/well. Cells were grown for another 24 h at 37 °C, then washed, fixed, and stained with C34 IgG followed by anti-human IgG/FITC. After three washes, fluorescence was measured on a BD FACSCanto II. All procedures before fixing and staining were conducted in a BSL-3 lab at IMCAS.
Pre- and post-attachment assay
Twenty-four-well tissue culture plates were seeded with 6 × 104 Vero cells per well overnight. CHIKV stock was defrosted at 4 °C for 2 h. For pre-attachment assays, three-fold dilutions of the IgG or Fab were prepared at 4 °C in DMEM with 2% FBS. CHIKV stock was diluted to 106 PFU/mL and mixed with an equal volume of antibody. After incubation at 4 °C for 1 h, 400 μL per well of the mixture was added to pre-chilled Vero cells and incubated at 4 °C for 1 h. Non-adsorbed virus was removed by washing twice with cold DMEM with 2% FBS, and adsorbed virus was allowed to internalize during a 37 °C incubation for 30 min. Supernatant was removed, and 400 μL per well of DMEM with 2% FBS was added and incubated at 37 °C.
For post-attachment assays, CHIKV stock was diluted to 106 PFU/mL and mixed with an equal volume of cold DMEM with 2% FBS. 400 μL per well of the mixture was added to pre-chilled Vero cells and incubated at 4 °C for 1 h. Non-adsorbed virus was removed by washing twice with cold DMEM with 2% FBS. Diluted IgG or Fab was mixed with an equal volume of cold DMEM with 2% FBS and added to Vero cells, incubated at 4 °C for another hour. Adsorbed virus was allowed to internalize during a 37 °C incubation for 30 min. Supernatant was removed, and 400 μL per well of DMEM with 2% FBS was added and incubated at 37 °C. After 24 h, infection rates reached about 65%, and the plates were harvested and analyzed for antigen-specific FACS as described above.
In vivo protection studies on mice
The protective efficacy of each mAb was assessed using an IFNAR1-KO mouse model. For determining the median lethal dose (LD50), five groups of ten female mice (8-12 weeks old) were challenged by footpad injection with 5 × 107, 5 × 106, 5 × 105, 5 × 104, or 0 PFU/ml of CHIKV (181/25 strain). Mice were monitored for 14 days post-infection for survival.
In the prophylactic experiment, groups of 8–12-week-old mice (five per group) were injected intraperitoneally with PBS or 10 mg/kg of mAbs. After 24 h, mice were subcutaneously infected with 100 LD50 of CHIKV. Weight and foot swelling were monitored daily for 15 days post-infection.
In the therapeutic experiment, groups of 8–12-week-old mice (five per group) were subcutaneously infected with 100 LD50 of CHIKV 24 h before intraperitoneal injection of PBS or 10 mg/kg of mAbs. Weight and foot swelling were monitored daily for 15 days post-infection.
FACS-based blocking assay
HEK293T cells were seeded into 6-well plates and transfected with pEGFP-N1-CHIKV E (4 μg/well). After 48 h, cells were harvested, and the surface-expressed E protein was confirmed. The cells were then harvested and aliquoted into 100 μl portions to 96-well plates, each well containing 3 × 105 cells. Cells were incubated with various mAbs in a 3f serial dilution starting from a concentration of 100 μg/ml for 30 min at 4 °C. For CHK-263, the Fc region was replaced by a human IgG Fc region for the eligibility of detection by the following antibodies. Following incubation, the cells were washed twice with PBS, and then incubated with different constructions of Mxra8 (100 μg/mL) for an additional 30 min. After two more washes, the cells were incubated with mouse anti-Strep II antibody (1:1000) for 30 min at 4 °C. Cells were washed twice again, followed by incubation with APC-conjugated goat anti-mouse IgG (1:200) for 30 min. After final washes, the cells were analyzed for APC fluorescence using a FACS Canto II flow cytometer.
VLP blocking assay with flow cytometry
VLPs (20 μg/mL) were mixed with antibodies at an equivalent molar ratio of E trimer: antibody = 1:10 and incubated for 30 min at room temperature. Meanwhile, Mxra8 was expressed on the surface of HEK293T cells transfected with pEGFP-N1-Mxra8 (expressed for 24 h). The cells were then harvested and aliquoted into 100 μl portions, each containing 3 × 105 cells. The cells were incubated with 200 μL of VLP-antibody mixture for 30 min at room temperature. After two washes, cells were stained with APC-conjugated anti-human IgG-Fc (5 μL for 1 × 106 cells, Biolegend Inc.) for 30 min. Following three washes, the samples were analyzed using a FACS Canto II flow cytometer.
DLS
VLPs were diluted to ~0.1 mg/ml in 20 mM Tris, pH 7.5, 100 mM NaCl, 1 mM EDTA buffer. Corresponding Fab molecules or IgGs of equal molar ratio to E1–E2 protein were mixed with VLPs at Fab:VLP molar ratios ranging from 5:1 to 400:1, and incubated on ice for 10 min. The size of complex particles was measured at 4 °C in 20 repeats, and the results were plotted. The fold changes of particle size compared to VLP alone were calculated. Three independent repeats were conducted and plotted with the mean and error.
Egress inhibition assay
Vero cells were cultured in 12-well plates and inoculated with the CHIKV 181/25 strain at MOI = 1. After incubating at 37 °C for 2 h, cells were washed 6 times with PBS. The antibodies were diluted with DMEM supplemented with 2% serum and 25 mM NH4Cl. Cells were incubated with diluted IgG at 37 °C for 6 h. The cell supernatant was collected and subjected to viral RNA amount assay with RT-qPCR. The copy number was calculated and plotted.
FFWO assay
Vero cells were seeded in 24-well plates, washed once with RPMI 1640 supplemented with 0.2% BSA, 10 mM HEPES pH 7.4, and 20 mM NH4Cl, and incubated for 15 min at 4 °C. CHIKV-181/25 (MOI of 15) was prepared in the same medium and added to cells for 1 h at 4 °C, and then the free virus was removed. Subsequently, DMEM containing 2% FBS with or without CHIKV-specific or isotype mAbs at 100 μg/ml, 10 μg/ml, 1 μg/ml, or 0.1 μg/ml was added to cells and incubated for 1 h at 4 °C. FFWO was induced by the addition of pre-warmed fusion media (RPMI 1640, 0.2% BSA, 10 mM HEPES, and 30 mM succinic acid at pH 5.5) for 2 min at 37 °C. In parallel wells, control media (RPMI 1640, 0.2% BSA, 10 mM HEPES at pH 7.4) were added for 2 min at 37 °C to ensure that infection occurred only through pH-dependent plasma membrane fusion. Medium was removed and cells were incubated in DMEM supplemented with 5% FBS, 10 mM HEPES, and 20 mM NH4Cl (pH 7.4); NH4Cl prevented secondary infection through endosomal fusion pathways. Cells were detached 24 h later, fixed, and permeabilized with BD Cytofix/Cytoperm fixation and permeabilization solution according to the manufacturer’s instructions. Cells were incubated sequentially with C48-APC antibody. Infection was evaluated on a Canto flow cytometer and analyzed using FlowJo software.
Crystallization
Eluates from gel-filtration of the C37 Fab-CHIKV E protein complex were collected and concentrated using Amicon Ultra-6 30 kDa cutoff centrifugal concentrators (Millipore). Initial crystals were obtained via sitting-drop vapor-diffusion at 18 °C, with 300 nL of protein solution (5 or 10 mg/mL) mixed with 300 nL of reservoir solution. The following crystallization kit was used for screening: Wizard 1-4, Structure Screen 1-2, PEG/Rx 1-2, PEG/Ion, PEG/Ion 2, SaltRx 1-2, Crystal Screen 1–2, Index, Natrix 1–2, and JCSG plus. Optimized crystallization conditions were achieved by sitting-drop (1 μL + 1 μL) vapor-diffusion, resulting in improved crystals under the condition of 100 mM HEPES/Sodium hydroxide pH 7.0, 15% (w/v) PEG 20000 at 18 °C.
X-ray diffraction data collection, structure determination, and refinement
Diffraction data were collected at the Shanghai Synchrotron Radiation Facility (SSRF) beamline BL19U1 using an RDI CMOS_8M detector with a wavelength of 0.97853 Å. The datasets were processed using the HKL2000 software package62.
The crystals diffracted to 3.20 Å resolution and belonged to the P212121 space group. The complex structure was determined by molecular replacement (MR) with PHASER63 using the crystal structures of the CHIKV prefusion E1–E2 heterodimer (PDB ID 6JO828) and Fab (Alphafold2 generated) as search models. The complex atomic model was refined through iterative rounds using Coot64 and phenix.refine65. The final model had Rwork and Rfree values of 0.222 and 0.277, respectively, with 95.10%, 4.90%, and 0% of residues in favored, allowed, and outlier regions of the Ramachandran plot, respectively. The final model was validated with MolProbity66. Detailed statistics for data collection and structure refinement are shown in Table S2.
Cryo-EM sample preparation and data acquisition
CHIKV VLPs were incubated with a 2-f excess amount (molar ratio of Fab:VLP = 720:1) of C34 Fab or C37 Fab at 4 °C in buffer containing 20 mM Tris, pH 7.5, 250 mM NaCl, 1 mM EDTA for 5 min, then prepared for VLP: Fab complex specimen preparation. VLP and infectious virus samples were prepared in the same buffer condition. A 3.5 μL sample at a concentration of ~1 mg/mL was applied to 200 mesh Quantifoil R2/1 gold grids that had been glow-discharged in oxygen and argon for 1 minute. The grids were flash-cooled in liquid ethane under BSL-2 containment using an FEI Vitrobot Mark IV with a blotting time of 4 s at 100% humidity, 8 °C. Cryo-EM movies were recorded with EPU (Thermo Fisher) using a K3 Summit electron detector (Gatan) attached to a Titan Krios microscope operating at 300 kV, or with SerialEM using a K2 Summit electron detector (Gatan) attached to a Titan Krios microscope operating at 300 kV. An electron dose of 50 or 60 e−/Å2 fractioned into 32 frames and pixel sizes of 1.32 Å, 1.36 Å, or 1.04 Å were used for different datasets (detailed in Table S1). All datasets were recorded under a defocus range between −1.5 μm and −2.2 μm.
Cryo-EM data processing
The movies were imported into cryoSPARC67 and corrected for beam-induced motion with patch-based motion correction. The contrast transfer function (CTF) parameters were determined for the aligned and dose-weighted micrographs with patch-CTF estimation in cryoSPARC. Micrographs with poor Thon rings and severe contamination were manually excluded. Particles were picked from micrographs using Topaz68 with a ResNet16 (32units) pretrained model and subjected to several rounds of 2D classification and extraction within cryoSPARC. Ab initio 3D models were generated using cryoSPARC without any symmetry imposed. For all the VLP- or virion-related datasets, particles were subjected to heterorefinement and NU-refinement with icosahedral (I1) symmetry imposed. For the VLP dataset, 36,959 particles were picked from 2860 movies, of which 23,118 particles generated a 3.82 Å refined map. For C34 Fab:VLP dataset, 61,787 particles were picked from 2136 movies, of which 39,766 particles generated a 4.35 Å refined map. For C37 Fab:VLP dataset, 101,173 particles were picked from 3535 movies, of which 74,737 particles generated a 4.01 Å refined map. And for infectious virion:C34 Fab dataset, 124,010 particles were picked from 3488 movies, of which 106,658 particles generated a 3.63 Å refined map.
To further improve resolution, a block-based algorithm was applied. Using the C34 Fab:VLP complex structure as an example, 39,766 refined particles were imported to Relion-3.0.869 using the csparc2star.py script in pyem70, and blocks around 5f, 3f, and 2f axes were generated. Subparticles were extracted with the calculated defocus and various box sizes (detailed in Fig. S5). The extracted subparticles underwent 3D classification without shift or rotation alignment. For the 5f block, 414,392 out of 2,384,724 subparticles were selected and imported back to cryoSPARC. Subsequently, local refinement and local CTF refinement were performed with a mask generated from the reconstructed volume around the 5 f axis, producing a final density map with a resolution of 3.90 Å. For the 2f block, 545,547 subparticles generated a 3.89 Å resolution map, and for the 3f block, 391,860 subparticles resulted in a 3.84 Å resolution map.
Similar block-based processing was conducted for all VLP and virion-related datasets, yielding density maps with various resolutions. All resolutions were estimated using the gold-standard Fourier shell correlation (FSC) criterion of 0.143. Local resolutions were estimated using cryoSPARC’s built-in program.
For the subparticles of C34 Fab:VLP complex around 5 f axis, duplicate particles were removed around 5f axis, duplicate particles were removed, and particles underwent no-alignment 3D classification with a denovo mask centered at the 5 Fab molecules around the 5f axis with a 95 Å radius. Subparticles were classified into 8 or 10 classes with T values of 4, 10, 20, and 35, respectively. Particles generating density maps with 1 to 5 C34 Fab molecules were imported into cryoSPARC for non-uniform local refinement, with resolutions estimated locally.
For the C34 Fab–E1–E2 (ectodomain) complex dataset, a total of 3883 movies were collected and imported into cryoSPARC. A similar pre-processing procedure was followed, including motion correction, CTF estimation, micrograph panning, and particle picking. Particles were extracted with a box size of 320 pixels × 1.32 Å/pixel, larger than the estimated particle dimensions (180–220 Å), to achieve robust 2D classification. A total of 1,310,873 particles were picked and extracted. After several rounds of 2D classification, 634,904 particles were kept for ab-initio reconstruction and heterorefinement. Classes of particles generating density maps with visible E protein subdomains were selected for non-uniform refinement, yielding a 3.05 Å resolution density map.
Model building and refinement for cryo-EM structures
The cryo-ET structure of CHIKV VLP (PDB ID: 8CFG), E1–E2 heterodimer or E3 subunit from CHIKV E trimer structure (PDB ID: 6JO8), the C34 Fab structure predicted by AlphaFold, and the C37 Fab crystal structure from this study were fitted into the corresponding density map as starting references. For all structures, the four different assemblies within each asymmetric unit were refined against different blocks: assembly 4 was refined against the 5 f blocks, and the remaining assemblies 1–3 were refined against the 2f blocks. For C34 and C37 Fab-bound structures, additional refinement against the 3 f blocks was conducted for the assemblies on the i3 spike. We conducted a local filter on the local refined blocks maps by the local filtering option in cryoSPARC in order to maintain the low-frequency densities for model fitting. For each model fitting, the coordinates were first refined against the local filtered maps of each block for several rounds and refined against sharpened maps in the last round of refinement. The sharpened maps were deposited in EMDB. Models were refined through iterative rounds of real-space refinement in PHENIX65, manual adjustments in Coot64, and EM-related tools-based refinement in Rosetta71. Low-resolution regions, including CH1/CL of C37 Fab molecules and C34 Fab molecules around the 5 f axis, were subjected to real-space refinement in PHENIX as rigid bodies with secondary structure and Ramachandran restraints. The final models were validated with MolProbity. Detailed statistics for structure refinement are shown in Table S1. Figures presenting the structures were prepared using UCSF Chimera72, UCSF ChimeraX73, and PyMOL (Schrödinger LLC, New York, NY).
Quantification and statistical analysis
All statistical analyses were performed using GraphPad Prism version 9.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Atomic coordinates and associated structure factors from crystallographic studies have been deposited in the Protein Data Bank (PDB) under accession code 9IW2. Cryo-EM structures have been deposited in the Electron Microscopy Data Bank (EMDB) and PDB with the following accession codes: C34 Fab:VLP complex: EMD-60997, 60999, 61002 and PDB 9IYI; C37 Fab:VLP complex: EMD-61025, 61026, 61030 and PDB 9IZ9; CHIKV VLP alone: EMD-60970, 60972, and PDB 9IXI, C34 Fab:infectious virion complex: EMD-61168, 61169 and PDB 9J6D [[https://www.rcsb.org/structure/9I6D]], C34 Fab-E1–E2 ectodomain complex: EMD-60966 and PDB 9IXA. CHK-263 structure is derived from PDB with accession code 7CVZ and 7CW2. Source data underlying Figs. 1, 2, 3n–p, 5f and Supplementary Figs 1a, 3, and 9b, c are provided as a Source Data file. Source data are provided with this paper.
References
Strauss, J. H. & Strauss, E. G. The alphaviruses: gene expression, replication, and evolution. Microbiol. Rev. 58, 491–562 (1994).
Weaver, S. C., Winegar, R., Manger, I. D. & Forrester, N. L. Alphaviruses: population genetics and determinants of emergence. Antivir. Res. 94, 242–257 (2012).
Silva, L. A. & Dermody, T. S. Chikungunya virus: epidemiology, replication, disease mechanisms, and prospective intervention strategies. J. Clin. Invest. 127, 737–749 (2017).
Reiter, P., Fontenille, D. & Paupy, C. Aedes albopictus as an epidemic vector of chikungunya virus: Another emerging problem?. Lancet Infect. Dis. 6, 463–464 (2006).
Tsetsarkin, K. A., Vanlandingham, D. L., McGee, C. E. & Higgs, S. A single mutation in chikungunya virus affects vector specificity and epidemic potential. PLoS Pathog. 3, e201 (2007).
Lumsden, W. H. An epidemic of virus disease in Southern Province, Tanganyika Territory, in 1952–53. II. General description and epidemiology. Trans. R. Soc. Trop. Med. Hyg. 49, 33–57 (1955).
Mason, P. J. & Haddow, A. J. An epidemic of virus disease in Southern Province, Tanganyika Territory, in 1952-53; an additional note on Chikungunya virus isolations and serum antibodies. Trans. R. Soc. Trop. Med. Hyg. 51, 238–240 (1957).
Simizu, B., Yamamoto, K., Hashimoto, K. & Ogata, T. Structural proteins of chikungunya virus. J. Virol. 51, 254–258 (1984).
Jose, J., Snyder, J. E. & Kuhn, R. J. A structural and functional perspective of alphavirus replication and assembly. Future Microbiol. 4, 837–856 (2009).
Schwartz, O. & Albert, M. L. Biology and pathogenesis of chikungunya virus. Nat. Rev. Microbiol. 8, 491–500 (2010).
Solignat, M., Gay, B., Higgs, S., Briant, L. & Devaux, C. Replication cycle of chikungunya: a re-emerging arbovirus. Virology 393, 183–197 (2009).
Sun, S. et al. Structural analyses at pseudo atomic resolution of Chikungunya virus and antibodies show mechanisms of neutralization. Elife 2, e00435 (2013).
Uchime, O., Fields, W. & Kielian, M. The role of E3 in pH protection during alphavirus assembly and exit. J. Virol. 87, 10255–10262 (2013).
Sjöberg, M., Lindqvist, B. & Garoff, H. Activation of the alphavirus spike protein is suppressed by bound E3. J. Virol. 85, 5644–5650 (2011).
Snyder, A. J. & Mukhopadhyay, S. The alphavirus E3 glycoprotein functions in a clade-specific manner. J. Virol. 86, 13609–13620 (2012).
Voss, J. E. et al. Glycoprotein organization of chikungunya virus particles revealed by X-ray crystallography. Nature 468, 709–712 (2010).
Basore, K. et al. Cryo-EM structure of chikungunya Virus in complex with the Mxra8 receptor. Cell 177, 1725–1737.e16 (2019).
Davis, N. L., Pence, D. F., Meyer, W. J., Schmaljohn, A. L. & Johnston, R. E. Alternative forms of a strain-specific neutralizing antigenic site on the Sindbis virus E2 glycoprotein. Virology 161, 101–108 (1987).
Mukhopadhyay, S. et al. Mapping the structure and function of the E1 and E2 glycoproteins in alphaviruses. Structure 14, 63–73 (2006).
Ubol, S. & Griffin, D. E. Identification of a putative alphavirus receptor on mouse neural cells. J. Virol. 65, 6913–6921 (1991).
Vrati, S., Kerr, P. J., Weir, R. C. & Dalgarno, L. Entry kinetics and mouse virulence of Ross River virus mutants altered in neutralization epitopes. J. Virol. 70, 1745–1750 (1996).
Cherrier, M. V. et al. Structural basis for the preferential recognition of immature flaviviruses by a fusion-loop antibody. EMBO J. 28, 3269–3276 (2009).
Lescar, J. et al. The Fusion glycoprotein shell of Semliki Forest virus: an icosahedral assembly primed for fusogenic activation at endosomal pH. Cell 105, 137–148 (2001).
Roussel, A. et al. Structure and interactions at the viral surface of the envelope protein E1 of Semliki Forest virus. Structure 14, 75–86 (2006).
Zhang, R. et al. Mxra8 is a receptor for multiple arthritogenic alphaviruses. Nature 557, 570–574 (2018).
Jung, Y.-K. et al. DICAM, a novel dual immunoglobulin domain containing cell adhesion molecule interacts with alphavbeta3 integrin. J. Cell Physiol. 216, 603–614 (2008).
Jung, Y.-K. et al. DICAM inhibits osteoclast differentiation through attenuation of the integrin αVβ3 pathway. J. Bone Min. Res. 27, 2024–2034 (2012).
Song, H. et al. Molecular basis of arthritogenic alphavirus receptor MXRA8 binding to chikungunya virus envelope protein. Cell 177, 1714–1724.e12 (2019).
van Duijl-Richter, M. K. S., Hoornweg, T. E., Rodenhuis-Zybert, I. A. & Smit, J. M. Early Events in chikungunya virus infection-from virus cell binding to membrane fusion. Viruses 7, 3647–3674 (2015).
Li, L., Jose, J., Xiang, Y., Kuhn, R. J. & Rossmann, M. G. Structural changes of envelope proteins during alphavirus fusion. Nature 468, 705–708 (2010).
Abdelnabi, R., Neyts, J. & Delang, L. Towards antivirals against chikungunya virus. Antivir. Res. 121, 59–68 (2015).
Ly, H. Ixchiq (VLA1553): The first FDA-approved vaccine to prevent disease caused by chikungunya virus infection. Virulence 15, 2301573 (2024).
Dörner, T. & Radbruch, A. Antibodies and B cell memory in viral immunity. Immunity 27, 384–392 (2007).
Warter, L. et al. Chikungunya virus envelope-specific human monoclonal antibodies with broad neutralization potency. J. Immunol. 186, 3258–3264 (2011).
Pal, P. et al. Development of a highly protective combination monoclonal antibody therapy against Chikungunya virus. PLoS Pathog. 9, e1003312 (2013).
Fox, J. M. et al. Broadly neutralizing alphavirus antibodies bind an epitope on E2 and inhibit entry and egress. Cell 163, 1095–1107 (2015).
Fong, R. H. et al. Exposure of epitope residues on the outer face of the chikungunya virus envelope trimer determines antibody neutralizing efficacy. J. Virol. 88, 14364–14379 (2014).
Smith, S. A. et al. Isolation and characterization of broad and ultrapotent human monoclonal antibodies with therapeutic activity against chikungunya virus. Cell Host Microbe 18, 86–95 (2015).
Quiroz, J. A. et al. Human monoclonal antibodies against chikungunya virus target multiple distinct epitopes in the E1 and E2 glycoproteins. PLoS Pathog. 15, e1008061 (2019).
Malonis, R. J. et al. Near-germline human monoclonal antibodies neutralize and protect against multiple arthritogenic alphaviruses. Proc. Natl. Acad. Sci. USA 118, e2100104118 (2021).
Tian, S. et al. Distinct BCR repertoires elicited by SARS-CoV-2 RBD and S vaccinations in mice. Cell Discov. 7, 91 (2021).
Zhu, D. et al. Pushing the resolution limit by correcting the Ewald sphere effect in single-particle Cryo-EM reconstructions. Nat. Commun. 9, 1552 (2018).
Zhang, R. et al. 4.4 Å cryo-EM structure of an enveloped alphavirus Venezuelan equine encephalitis virus. EMBO J. 30, 3854–3863 (2011).
Chen, L. et al. Implication for alphavirus host-cell entry and assembly indicated by a 3.5Å resolution cryo-EM structure. Nat. Commun. 9, 5326 (2018).
Wang, A. et al. Structure of infective Getah virus at 2.8 Å resolution determined by cryo-electron microscopy. Cell Discov. 8, 12 (2022).
Wang, M. et al. Structural insights into alphavirus assembly revealed by the cryo-EM structure of Getah virus. Viruses 14, 327 (2022).
Chmielewski, D. et al. Cryogenic electron microscopy and tomography reveal imperfect icosahedral symmetry in alphaviruses. PNAS Nexus 3, pgae102 (2024).
Ribeiro-Filho, H. V. et al. Cryo-EM structure of the mature and infective Mayaro virus at 4.4 Å resolution reveals features of arthritogenic alphaviruses. Nat. Commun. 12, 3038 (2021).
Williamson, L. E. et al. Structural constraints link differences in neutralization potency of human anti-Eastern equine encephalitis virus monoclonal antibodies. Proc. Natl. Acad. Sci. USA 120, e2213690120 (2023).
Sharma, A. et al. The epitope arrangement on flavivirus particles contributes to Mab C10’s extraordinary neutralization breadth across Zika and dengue viruses. Cell 184, 6052–6066.e18 (2021).
Fox, J. M. et al. A cross-reactive antibody protects against Ross River virus musculoskeletal disease despite rapid neutralization escape in mice. PLoS Pathog. 16, e1008743 (2020).
Powell, L. A. et al. Human mAbs broadly protect against arthritogenic alphaviruses by recognizing conserved elements of the Mxra8 receptor-binding site. Cell Host Microbe 28, 699–711.e7 (2020).
Goh, L. Y. H. et al. Neutralizing monoclonal antibodies to the E2 protein of chikungunya virus protects against disease in a mouse model. Clin. Immunol. 149, 487–497 (2013).
Earnest, J. T. et al. Neutralizing antibodies against Mayaro virus require Fc effector functions for protective activity. J. Exp. Med 216, 2282–2301 (2019).
Zhou, Q. F. et al. Structural basis of Chikungunya virus inhibition by monoclonal antibodies. Proc. Natl. Acad. Sci. USA 117, 27637–27645 (2020).
Sutton, M. S. et al. Vaccine elicitation and structural basis for antibody protection against alphaviruses. Cell 186, 2672–2689.e25 (2023).
Wu, S.-R., Haag, L., Sjöberg, M., Garoff, H. & Hammar, L. The dynamic envelope of a fusion class II virus. E3 domain of glycoprotein E2 precursor in Semliki Forest virus provides a unique contact with the fusion protein E1. J. Biol. Chem. 283, 26452–26460 (2008).
Pascal, K. E. et al. Development of clinical-stage human monoclonal antibodies that treat advanced ebola virus disease in nonhuman primates. J. Infect. Dis. 218, S612–S626 (2018).
Joyce, M. G. et al. Iterative structure-based improvement of a fusion-glycoprotein vaccine against RSV. Nat. Struct. Mol. Biol. 23, 811–820 (2016).
Shi, R. et al. A human neutralizing antibody targets the receptor-binding site of SARS-CoV-2. Nature 584, 120–124 (2020).
Vaney, M.-C. et al. Evolution and activation mechanism of the flavivirus class II membrane-fusion machinery. Nat. Commun. 13, 3718 (2022).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D. Biol. Crystallogr. 66, 12–21 (2010).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, e42166 (2018).
Schorb, M., Haberbosch, I., Hagen, W. J. H., Schwab, Y. & Mastronarde, D. N. Software tools for automated transmission electron microscopy. Nat. Methods 16, 471–477 (2019).
Lugmayr, W. et al. StarMap: a user-friendly workflow for Rosetta-driven molecular structure refinement. Nat. Protoc. 18, 239–264 (2023).
Pettersen, E. F. et al. UCSF chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Goddard, T. D. et al. UCSF ChimeraX: Meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Acknowledgements
We gratefully acknowledge the support of the Center for Biological Imaging, Core Facilities for Protein Science at the Institute of Biophysics (IBP), Chinese Academy of Sciences (CAS), and the Institute of Physics, CAS & Beijing Branch of Songshan Lake Materials Laboratory, for providing access to cryo-EM data collection. Special thanks to B. Zhu, X. Huang, X-J. Li, L. Chen, and other staff at IBP, CAS, and to D. Sun at the Institute of Physics, CAS, for their invaluable assistance in cryo-EM data collection. We also extend our gratitude to the staff at the BL19U1 beamline at the SSRF (Shanghai, People’s Republic of China) for their assistance during X-ray data collection. We thank Y. Bi and the staff of the Biosafety Level 3 (BSL-3) laboratories at the Institute of Microbiology, Chinese Academy of Sciences, for their support throughout this study. We appreciate the contributions of X. Jin, X. Bi, Z. Zhao, and H. Yang at IMCAS for their participation in the early stages of this research. This work is supported by the Major Research Plan of the National Natural Science Foundation of China, no. 92369202 (J.Y. and X.H.).
Author information
Authors and Affiliations
Contributions
Conceptualization: G.F.G., J.Y., and X.H. Methodology: X.H., C.J., S.T., F.W., G.F.G., and J.Y. Investigation: X.H., C.J., S.T., F.W., D.L., X.D., Q.W., Q.H., and J.Q. Resources: G.-P.C., Z.T., and B.-D.Z. Writing—original draft: X.H. and J.Y. Writing—review and editing: all authors. Supervision: G.F.G. and J.Y. Project administration: X.H., G.F.G., and J.Y. Funding acquisition: J.Y. and X.H.
Corresponding authors
Ethics declarations
Competing interests
J.Y., X.H., C.J., S.T., F.W., Z.T., and G.F.G. are listed as inventors on a patent application for the use of C34 and C37 antibodies. All other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Aileen Chang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Han, X., Ji, C., Tian, S. et al. Neutralizing antibodies against Chikungunya virus and structural elucidation of their mechanism of action. Nat Commun 16, 9682 (2025). https://doi.org/10.1038/s41467-025-64687-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64687-2