Abstract
Human T-cell leukemia virus type 1 (HTLV-1)-associated myelopathy (HAM) is a debilitating neuroinflammatory disease with no available effective treatments. A hallmark of HAM is the transformation of HTLV-1-infected cells into T helper type 1 (Th1)-like cells, characterized by excessive interferon (IFN)-γ production that drives chronic inflammation. However, the molecular mechanisms fuel this aberrant Th1-like transformation and sustained inflammation remain poorly understood. We hypothesized that HAM-characteristic chromatin remodeling plays a pivotal role in the overexpression of key genes driving inflammatory pathogenesis. Using transcriptomic analysis, chromatin accessibility profiling, and biomarker evaluations across HTLV-1-related diseases, we identify MAP3K8 as a key gene that defines the unique inflammatory profile of HAM. MAP3K8 overexpression promotes Th1-like differentiation and constitutively activates the MEK-ERK signaling pathway. Furthermore, we elucidate the mechanism by which HTLV-1 Tax, Fosl2, and c-Jun collaboratively induce HAM-characteristic chromatin remodeling at the enhancer region of the MAP3K8 locus. Crucially, we demonstrate that mitogen-activated protein kinase kinase (MEK) inhibitors effectively suppress the MAP3K8-MEK signaling cascade and significantly mitigated inflammatory pathogenesis in an ex vivo culture assay. Our findings provide critical insights into the virus-host interactions underpinning HAM and propose the MAP3K8-MEK-ERK axis as a promising therapeutic target for this challenging condition.
Similar content being viewed by others
Introduction
ATP-dependent chromatin remodeling alters the higher-order structure of chromatin and thereby regulates transcription by controlling the access of transcription factors to chromatin1,2,3,4,5,6. Chromatin remodeling, mediated by mechanisms such as histone modifications, histone variant incorporation, and nucleosome repositioning, dynamically regulates the formation of open chromatin regions (OCRs) in a cell type– and stimulus-specific manner. Disruption of this regulation leads to pathological gene expression programs, underscoring the critical role of chromatin dynamics in development and disease7,8.
As a model for studying abnormal chromatin remodeling in disease, we focus on the neuroinflammatory condition known as human T-cell leukemia virus type 1 (HTLV-1)-associated myelopathy (HAM), also termed tropical spastic paraparesis. HTLV-1 infects an estimated 5‒10 million people worldwide, leading to adult T-cell leukemia/lymphoma (ATL) in ~5% of infected individuals and HAM in 0.3‒3%9,10. HTLV-1 primarily infects CD4+ T cells, with infected cells in HAM patients exhibiting a distinct T helper type 1 cell (Th1)-like phenotype and polyclonal proliferation. In contrast, ATL cells demonstrate a regulatory T cell (Treg)-like phenotype and monoclonal proliferation. This divergence raises a critical question: how does a single viral infection result in such distinct pathological outcomes?
Differential gene expression profiles in the infected cells likely play a pivotal role in the divergent progression of HAM and ATL11,12,13: For example, infected cells in patients with HAM express higher levels of HTLV-1 tax, a viral regulatory gene, compared with ATL cells14. This drives the transformation of these infected cells into Th1-like cells that overproduce interferon (IFN)-γ, creating a chronic inflammatory state dominated by Th1 immune responses15. In the spinal cord, these infected cells enhance cellular immunity through IFN-γ overproduction, stimulating astrocytes to secrete CXCL10, which recruits additional these infected and inflammatory cells16. This self-perpetuating inflammatory loop underpins HAM pathogenesis, making these aberrant infected cells a critical target for therapeutic intervention. However, the molecular mechanisms underlying the transition of the infected cells to this pathogenic HAM phenotype remain poorly understood.
In various cancers, genetic mutations in chromatin remodelers have been shown to create abnormal chromatin structures that drive disease progression17. In contrast, the role of abnormal chromatin structures in inflammatory disease development remains less well understood. We previously reported that HTLV-1 infection leads to chromatin restructuring and epigenetic dysregulation, contributing to leukemogenesis18. Based on this finding, we hypothesized that disease-specific chromatin remodeling in infected T cells drives the overexpression of key genes involved in inflammatory pathogenesis.
To test this hypothesis, we conducted transcriptomic and chromatin accessibility analyses of HTLV-1-infected T cells across HTLV-1-related diseases and integrated these data with markers of disease activity in HAM. Our study revealed a mechanism by which virus-host interactions drive inflammatory pathogenesis in HAM and identified promising therapeutic strategies targeting these pathways.
Results
HAM-specific gene expression profiles among HTLV-1-infected CD4+ T cells in HTLV-1-associated diseases
HTLV-1-infected cells are characterized by CD4 and CADM1 surface markers. In infected cells in patients with HAM, polyclonality is observed, along with CD7 expression, whereas ATL cells exhibit monoclonal expansion and downregulation of CD719,20,21,22. To investigate the transcriptomic profile of the infected cells in patients with HAM, we sorted CD4+CADM1+CD7+ cells (designated as the “D” population) from the peripheral blood mononuclear cells (PBMCs) of patients with HAM (n = 13) and performed deep RNA sequencing (RNA-seq), achieving over 30 million reads per sample.
Previously, transcriptomic data were obtained using the same sorting strategy from HTLV-1 uninfected CD4+ cells (CD4+CADM1−CD7+ cells, “P” population), HTLV-1 infected CD4+ T cells (CD4+CADM1+CD7+, “D” population), and ATL cells (CD4+CADM1+CD7−, “N” population) in asymptomatic carriers (AC) and patients with indolent ATL (iATL) and acute ATL (aATL)18. By integrating these datasets with transcriptomic profiles of HTLV-1–infected cells from patients with HAM, we conducted a comparative analysis across the spectrum of HTLV-1-associated disease phenotypes (Fig. 1A).
A Workflow illustrating the collection and processing of CD4+CADM1+CD7+ cells from the PBMCs of patients with HAM (D population, n = 13). The transcriptome data from these cells were integrated with those of HTLV-1-infected CD4+ T cells from asymptomatic carriers (P, D, and N populations, n = 10, 8, and 9, respectively), patients with indolent ATL (D and N populations, n = 5 and 9, respectively), and patients with acute ATL (N population, n = 13). P, D, and N populations represent CD4+CADM1−CD7+ cells, CD4+CADM1+CD7+ cells, and CD4+CADM1+CD7− cells, respectively. B Non-hierarchical dimensional reduction using t-SNE based on the expression of all genes (22,074 genes). AC_P (black), AC_D (orange), and AC_N (green) represent HTLV-1-uninfected, CD7-positive HTLV-1-infected, and CD7-negative HTLV-1-infected CD4+ T cells in asymptomatic individuals, respectively. HAM_D (red) represents CD7-positive HTLV-1-infected CD4+ T cells from patients with HAM. iATL_D (gray) and iATL_N (blue) represent CD7-positive and CD7-negative ATL cells, respectively, in patients with indolent ATL. aATL_N (purple) represents CD7-negative ATL cells in patients with acute ATL (C) Hierarchical clustering based on the expression of all genes using HTLV-1-uninfected CD4+ T cells (AC_P: black), HTLV-1-infected CD4+ T cells from patients with HAM (HAM_D: red), and acute ATL cells (aATL_N: purple). D The heatmap depicts relative expression levels of significantly differentially expressed genes in HAM_D, AC_D, AC_N, iATL_N, and aATL_N compared to AC_P, respectively (log2 FC ≧ 1, adjusted P < 0.05, TPM ≧ 2). Statistical analysis was performed using a two-sided test with Bonferroni correction for multiple comparisons. Adjusted P values are reported. E Heatmap showing genes upregulated only in HAM_D and AC_D compared with AC_P (groups a + b); genes commonly upregulated in HAM_D, AC_D, AC_N, iATL_N, and aATL_N compared with AC_P (group c); and genes upregulated only in iATL_N and aATL_N compared with AC_P (d + e). Enriched gene ontology terms are depicted for the upregulated gene sets. P values were calculated using a one-sided Fisher’s exact test and are shown as −log10(P). Nominal P values are reported.
We applied non-hierarchical dimensionality reduction using t-distributed Stochastic Neighbor Embedding (t-SNE) to visualize transcriptomic differences. Distinct clusters were formed between HTLV-1-uninfected and HTLV-1-infected CD4+ T cells. Within the HTLV-1-infected cell cluster, cells from patients with HAM and those from aATL showed a tendency to segregate (Fig. 1B). Hierarchical clustering further confirmed that the transcriptomic profiles of HTLV-1−infected cells from patients with HAM and leukemic cells from patients with aATL formed distinct clusters (Fig. 1C).
To identify characteristic gene expression patterns in infected cells from patients with HAM, we compared the gene expression levels of HTLV-1-infected cells in different disease states with those of uninfected CD4+ T cells. This analysis identified 893 genes with significantly elevated expression in at least one infected cell type (log2 fold change (FC) ≧ 1, adjusted P < 0.05, TPM: transcripts per million ≧ 2). These genes were classified into several groups:
(a) genes highly expressed exclusively in infected cells from patients with HAM,
(b) genes highly expressed in both HAM and AC CD4+CADM1+CD7+ cells,
(c) genes commonly expressed in HTLV-1-infected CD4+ T cells across all HTLV-1-associated diseases,
(d) genes expressed in iATL and aATL CD4+CADM1+CD7− cells,
(e) genes exclusively expressed in aATL cells (Fig. 1D).
Group (c) included ectopically expressed genes such as CADM1, TIAM2, EVC, and EVC2, which were shared across HTLV-1-infected T cells in both disease and asymptomatic states23,24,25. Genes in groups (a) and (b) exhibited differential expression that accompanied the emergence of HTLV-1–infected cells from patients with HAM. Gene ontology analysis of these groups revealed enrichment for inflammatory immune response-related functions, including regulation of immune and inflammatory response, reflecting the inflammatory characteristics of HAM (Fig. 1E).
Conversely, group (c) genes were enriched for cell division- and proliferation-related functions, while genes strongly expressed in aATL cells (groups d and e) were enriched for gene expression regulation and DNA replication. These findings suggest that HTLV-1-infected cells from patients with HAM are characterized by a gene expression profile involved in inflammatory immunity, highlighting specific genes likely to play central roles in HAM inflammatory pathogenesis.
Exploration of inflammatory genes within HAM-characteristic OCRs
To test the hypothesis that disease-specific abnormal chromatin remodeling in HTLV-1−infected T cells from patients with HAM leads to the overexpression of critical genes driving inflammatory pathogenesis, we performed an assay for transposase-accessible chromatin (ATAC) with sequencing (ATAC-seq). Chromatin accessibility profiles were generated from CD4+CADM1+CD7+ cells sorted from PBMCs of HAM patients (Supplementary Fig. 1). These data were integrated with previously obtained ATAC-seq data from CD4+CADM1+CD7− cells isolated from PBMCs of patients with acute ATL (aATL) for comparative analysis18.
We investigated the relationship between chromatin accessibility and gene expression by focusing on 190 genes strongly expressed in infected cells from patients with HAM (groups a and b) and from ACs (group b) and 270 genes highly expressed in aATL cells (groups d and e). A positive correlation was observed between chromatin accessibility and gene expression, indicating that the chromatin accessibility plays a regulatory role in HAM- or ATL-specific gene expression (Fig. 2A). Furthermore, we focused on a set of 647 genes that were significantly differentially expressed in HTLV-1–infected T cells from HAM patients compared to those from ACs (fold change ≧ 1.5, adjusted P < 0.05), and examined their relationship with chromatin accessibility. The results showed a positive correlation between these two parameters, suggesting that chromatin accessibility contributes to the regulation of gene expression profiles characteristic of HAM and ACs (Fig. 2B). To more accurately identify HAM-characteristic OCRs, we compared ATAC-seq profiles of HTLV-1–infected cells from HAM patients with those of ACs and sorted normal CD4⁺ T cells, either unstimulated or activated with anti-CD3/CD28 antibodies (GSE116696). OCRs were identified using the MACS2 peak-calling algorithm. Comparative analysis revealed 132, 455, and 679 differentially accessible regions when infected cells from HAM patients were compared to ACs, unstimulated CD4⁺ T cells, and activated CD4⁺ T cells, respectively (Fig. 2C). Thirteen OCRs were commonly enriched across all comparisons, and we defined these as HAM-characteristic OCRs.
A The scatter plot shows differences in ATAC-seq peak values in promoter regions (x axis) and differences in gene expression levels from RNA-seq data (y-axis) in HTLV-1–infected cells from HAM patients compared with acute ATL cells. Red and purple dots represent 190 and 270 genes strongly expressed in HTLV-1–infected cells from HAM patients (groups a and b) and aATL cells (groups d and e), respectively. Spearman’s correlation coefficient was calculated using a two-sided test (R = 0.41, P < 0.001). B Similar scatter plot comparing HTLV-1–infected cells from HAM patients and ACs. Red and ocher dots represent 434 and 213 genes significantly upregulated in HAM and AC samples, respectively (fold change > 1.5, adjusted P < 0.05). Spearman’s correlation coefficient was calculated using a two-sided test (R = 0.52, P < 0.001). C Venn diagram showing the overlap of OCRs detected in HTLV-1–infected T cells from patients with HAM versus those from ACs (red), normal CD4+ T cells (green), and CD4+ T cells stimulated with anti-CD3/CD28 antibodies (blue). D Heatmap (left) depicting Pearson correlation coefficients between transcripts levels (TPM) of eight selected genes and the CSF concentrations of CXCL10 (pg/mL) or neopterin (pmol/mL). Scatter plots (middle and right) show the correlation between MAP3K8 expression and CSF levels of CXCL10 or neopterin (R = 0.823 and 0.753, respectively). E Boxplot showing normalized MAP3K8 expression levels in AC_P (n = 10), AC_D (n = 8), and AC_N (n = 9) in asymptomatic individuals; HAM _D (n = 13) in patients with HAM; iATL_D (n = 5) and iATL_N (n = 9) in patients with indolent ATL; and aATL_N (N = 13) in patients with acute ATL, based on RNA-seq data. Each data point represents an individual patient sample. Boxes indicate the interquartile rage (IQR), with the line representing the median. Whiskers extend to 1.5 * IQR, and outliers beyond this range are shown as individual points. Statistical significance was assessed using one-way ANOVA. Exact P value are as follows: AC_P vs HAM_D, P = 0.000078, AC_D vs HAM_D, P = 0.0196, HAM_D vs iATL_D, P = 0.0128, HAM_D vs AC_N, P = 0.00005, HAM_D vs iATL_N, P = 0.0000096, HAM_D vs aATL_N, P = 0.0000067. Asterisks indicate statistical significance (* P < 0.05, ** P < 0.01, *** P < 0.001). (F) Representative histogram images of MAP3K8 from ATAC-seq and RNA-seq in normal CD4+ T cells, AC_D, HAM_D, and aATL_N. The highlighted region shows HAM-characteristic OCR.
To pinpoint genes most relevant to HAM inflammatory pathogenesis, we calculated the correlation coefficients between the expression levels of the 8 protein-coding genes among the 13 candidates and two established biomarkers of HAM disease activity: CXCL10 and neopterin concentrations in cerebrospinal fluid (CSF) (Fig. 2D). Among these, MAP3K8 and ENSA exhibited the highest correlation coefficient for both biomarkers, strongly implicating its role in the inflammatory pathogenesis of infected cells from patients with HAM. Given that MAP3K8, but not ENSA, was included in group (a) in Fig. 1D, we subsequently focused our analysis on MAP3K8. Expression analysis further revealed that MAP3K8 expression levels in HTLV-1−infected cells from HAM patients were significantly higher than those in both HTLV-1-uninfected and HTLV-1-infected CD4+ T cells, as well as in ATL cells from HTLV-1-infected individuals. Notably, MAP3K8 expression in HTLV-1−infected cells from HAM patients was 4.3-fold higher than in aATL cells (Fig. 2E). In addition, ATAC-seq analysis identified a pronounced HAM-characteristic open chromatin peak within the intronic region of MAP3K8 (Fig. 2F), suggesting that the open chromatin state in this region may drive the high MAP3K8 expression in HAM.
MAP3K8 overexpression constitutively activates the MEK-ERK pathway, increases inflammatory gene expression and induces Th1 differentiation
MAP3K8 has been implicated in Th1 and Th17 differentiation in knockout mouse models26. To investigate whether MAP3K8 promotes Th1 differentiation in human CD4+ T cells, we transduced the MAP3K8 gene into CD4+ T cells from multiple human donors using lentiviral vectors. MAP3K8 overexpression significantly increased the proportion of cells expressing T-bet, the master transcription factor that governs Th1 differentiation (Fig. 3A).
A Induction of T-bet expression in CD4+ T cells by MAP3K8, using lentiviral vectors (right: MAP3K8-expressing construct, left: empty-construct). Statistical analysis was performed using a two-sided paired t-test. n = 3 biologically independent samples, each derived from a different healthy donor. The asterisk indicates statistical significance (*P = 0.036). B Constitutive activation of MEK, ERK, and p38 phosphorylation in Jurkat cells transduced with a lentiviral vector expressing MAP3K8. β-actin was used as a loading control. Representative results from two to three independent experiments using cells from multiple independent donor samples are shown. C Relative expression levels of IFNG (upper panel) and TBX21 (lower panel) in Jurkat cells overexpressing MAP3K8. Data are presented as the mean +SD. Statistical analysis was performed using a two-sided paired t-test. n = 3 biologically independent experiments, each performed on separate days using independently cultured Jurkat cells. The asterisk indicates statistical significance (**P = 0.0011). D Genes upregulated in Jurkat cells overexpressing MAP3K8 compared with mock-overexpressing cells. The plot displays fold change (MAP3K8/mock) and ranking order of upregulated genes. E Gene ontology (GO) analysis for the 906 genes upregulated in MAP3K8-overexpressing Jurkat cells compared to mock-overexpressing cells (fold change > 2). GO terms enriched in the upregulated genes are displayed in order of P-value (−log10). Statistical analysis was performed using a two-sided Fisher’s exact test. Nominal P values are reported.
Given the high MAP3K8 expression observed in HTLV-1–infected cells from patients with HAM, we established three stable T cell lines with elevated MAP3K8 expression (MAP3K8-Jurkat) to evaluate its functional effects. MAP3K8 overexpression led to marked increases in phosphorylation levels of MEK1/2, ERK1/2, and p38, indicating constitutive activation of the MEK-ERK and p38 signaling pathways (Fig. 3B). This activation was accompanied by significant upregulation of IFNG and TBX21, key genes involved in the inflammatory response and Th1 differentiation (Fig. 3C).
To further explore the transcriptomic changes induced by MAP3K8 overexpression, we compared the gene expression profiles of MAP3K8-Jurkat and mock-Jurkat cells using RNA-Seq. This analysis identified 906 genes that were upregulated by more than 2-fold in MAP3K8-Jurkat cells compared with mock controls. Gene ontology analysis of these upregulated genes revealed strong enrichment for pathways associated with immune response, inflammatory response, and the mitogen-activated protein kinase pathway (Fig. 3D and E).
MEK inhibitors suppress MAP3K8-derived MEK-ERK pathway activation and Th1 differentiation
Based on the hypothesis that MAP3K8-driven activation of the MEK-ERK pathway contributes to HAM pathogenesis, we evaluated the effects of blocking this pathway using MEK inhibitors clinically approved for other diseases. Administration of trametinib, a MEK inhibitor, to MAP3K8-Jurkat cells completely abolished ERK phosphorylation (Fig. 4A). Similarly, when trametinib was applied to HCT-4 and HCT-5 cell lines (HTLV-1-infected cell lines derived from HAM patients), it effectively inhibited ERK pathway activation (Fig. 4B and Supplementary Fig. 2A). This inhibition was accompanied by decreased IFNG expression and reduced IFN-γ production (Fig. 4C and Supplementary Fig. 2B). Moreover, trametinib suppressed the proliferative capacity of HTLV-1-infected cell lines derived from HAM patients (Fig. 4D and Supplementary Fig. 2C). In addition to trametinib, other MEK1/2 inhibitors (binimetinib, selumetinib, and cobimetinib) similarly inhibited ERK1/2 phosphorylation and reduced cell proliferation (Fig. 4E and Supplementary Fig. 2D).
A Inhibitory effect of trametinib (1000 nM) on ERK phosphorylation in MAP3K8-overexpressing Jurkat cells. β-actin was used as a loading control. B Dose-dependent inhibitory effect of trametinib on ERK phosphorylation in HCT-4 cells. β-actin was used as a loading control. C IFNG relative expression levels (upper panel) and IFN-γ production levels (lower panel) in HCT-4 cells treated with trametinib. Data are presented as the mean +SD. n = 3 biologically independent experiments, each performed on separate days using independently cultured HCT-4 cells. D Percentages of cell proliferation in HCT-4 cells treated with trametinib. Data are presented as the mean +SD. n = 3 biologically independent experiments, each performed on separate days using independently cultured HCT-4 cells. E Inhibitory effect of MEK inhibitors on ERK phosphorylation in HCT-4 cells. β-actin was used as a loading control. F Downregulated gene expression levels in HCT-4 cells treated with trametinib compared with DMSO-treated cells. The plot shows fold changes (Trametinib/DMSO) and their respective ranking order. G Gene ontology (GO) analysis for the 452 genes downregulated in HCT-4 cells treated with trametinib compared with those treated with DMSO. GO terms enriched in the trametinib-treated group are displayed in order of P value (−log10). Statistical analysis was performed using a two-sided Fisher’s exact test. Nominal P values are reported. H Flow cytometric plots of IFN-γ and T-bet in normal CD4+ T cells stimulated with α-CD3/28 antibodies and treated with trametinib (1000 nM). The numbers in the plots indicate the percentage of positive cells. I Bar graphs indicating the percentage of T-bet-positive cells in normal CD4+ cells stimulated with α-CD3/28 antibodies and treated with trametinib (1000 nM). Data are presented as mean values + SD. n = 3 biologically independent experiments, each performed using CD4+ T cells isolated from different healthy donors. Asterisks indicate statistical significance (* P < 0.05, ** P < 0.01, *** P < 0.001). Statistical analysis was performed using a two-sided unpaired t-test (Fig.4C) and a one-way ANOVA (Fig.4D and Fig.4I). Representative immunoblot results from two to three independent experiments using cultured cell lines are shown (Fig. 4A, B, and E).
To assess the broader transcriptomic effects of MEK1/2 inhibitors, RNA-seq analysis was performed on HCT-4 cells treated with trametinib. A total of 452 genes were identified with significantly reduced expression compared to the dimethyl sulfoxide (DMSO)-treated control group. Gene ontology analysis revealed that these downregulated genes were highly enriched for pathways related to immune response, protein phosphorylation, and inflammatory response (Fig. 4F and G). Trametinib also reduced IFNG, CD69, and CTLA4 expression levels, which are representative genes significantly downregulated by trametinib in HCT-4 cells, in both HCT-1 and HCT-5 cells (Supplementary Fig. 2E).
To further evaluate the effects of MEK inhibition on Th1 differentiation, we stimulated human CD4+ T cells with anti-CD3/28 antibodies, which to mimic T cell receptor (TCR) activation. This stimulation increased T-bet expression and IFN-γ production, indicative of Th1 differentiation. Importantly, trametinib markedly suppressed both T-bet expression and IFN-γ production, demonstrating potent inhibition of Th1 differentiation (Fig. 4H and I).
To accurately evaluate the function of MAP3K8, we performed knockdown of endogenous MAP3K8 via siRNA electroporation in HAM-derived cell lines, which resulted in reduced MAP3K8 expression and a concomitant decrease in IFNG and TBX21 expression levels (Supplementary Fig. 2F and 2G). In contrast, knockdown of MAP3K8 in ATL cell lines and primary ATL cells did not significantly affect IFNG expression. This is likely due to the closed chromatin configuration at the IFNG locus in ATL cells, as compared to the more accessible structure observed in HTLV-1–infected cells from HAM patients (Supplementary Fig. 12).
These findings demonstrate that MEK1/2 inhibitors effectively suppress the MEK-ERK pathway downstream of MAP3K8. This suppression leads to reduced expression of genes associated with inflammatory responses, cell proliferation, and Th1 differentiation, highlighting the therapeutic potential of MEK inhibition in HAM pathogenesis.
Single cell multiome analysis reveals MAP3K8highHTLV-1-infected CD4+ T cells with distinct transcriptional profiles
To investigate the heterogeneity of HTLV-1−infected T cells, we conducted single-cell (sc) multiome analyses, integrating scRNA-seq and scATAC-seq data from PBMC derived from patients with HAM and ACs. Transcriptome and chromatin accessibility profiles were obtained at the single-cell level from PBMCs of HAM patients (n = 2) and ACs (n = 3). This approach enabled precise mapping of gene expression and chromatin accessibility profiles within individual cells. A total of 49,645 cells from each library were analyzed.
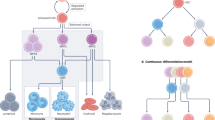
Uniform Manifold Approximation and Projection (UMAP) clustering of scRNA-seq and scATAC-seq data revealed multiple clusters. Based on marker gene expression, the clusters were classified into HTLV-1–infected CD4⁺ T cells, uninfected CD4⁺ T cells, HTLV-1–infected CD8⁺ T cells, uninfected CD8⁺ T cells, CD4⁺/CD8⁺ double-positive T cells, NK cells, B cells, and monocytes/dendritic cells. This analysis revealed distinct cluster distributions between patients with HAM and ACs (Fig. 5A). Clusters 0 and 16, representing specific subpopulations of HTLV-1–infected T cells, were predominantly observed in HAM #14 and HAM #15. In contrast, ACs (AC #1, AC #2, and AC #3) primarily exhibited clusters 1 and 17, corresponding to a different HTLV-1–infected subpopulation (Supplementary Fig. 3A and 3B). ATAC peaks at the MAP3K8 locus and MAP3K8 expression levels in HTLV-1–infected CD4⁺ T cell subpopulations more clearly demonstrated differences between HAM patients and ACs rather than Fig. 2F (Supplementary Fig. 3C and 3D). These results suggest that HTLV-1–infected T cells in patients with HAM and ACs comprise distinct subpopulations characterized by divergent gene expression and chromatin accessibility profiles.
A Weighted Nearest Neighbor (WNN) UMAP of single-cell multiome data (49,645 cells) derived from PBMCs of patients with HAM (n = 2) and ACs (n = 3). Each dot represents an individual cell, color-coded according to cell type annotations based on integrated clustering of transcriptomic and chromatin accessibility profiles. The upper panel displays the overall WNN UMAP (red, HTLV-1-infected CD4+ T cell; ocher, Monocyte; yellow-green, CD8+ T cell; green, CD4+ T cell; emerald green, NK cell; light blue, CD4/CD8 double positive cell; indigo, HTLV-1-infected CD8+ T cel; pink, B cell), while the lower panels present the WNN UMAPs segregated by individual donor (red, HAM #14; yellow-green, HAM #15; green, AC #1; blue. AC #2; purple, AC #3). B UMAP of scATAC-seq data (31,663 cells) showing PBMCs and CD4+CADM1+CD7+ cells from patients with HAM (n = 2). Cells are color-coded by annotated identity (brown, MAP3K8high HTLV-1+CD4+; purple, MAP3K8low HTLV-1+CD4+; red, uninfected CD4+T cell; muted yellow-green, Infected CD8+ T cell; pink, uninfected CD8+ T cell; blue, NK cell; orange, B cell; green, monocyte/DC). The upper right panel shows UMAPs for each sample library (upper left, CD4+CADM1+CD7+ cells in HAM #14; upper right, PBMCs in HAM #14; lower left, CD4+CADM1+CD7+ cells in HAM #15; lower right: PBMCs in HAM #15). The lower panels illustrate expression levels of cell-type marker genes used for annotation, displayed from upper left to lower right: NCAM1, CD19, CD14, CD8A, CD8B, CD4, CADM1, CCR4, MAP3K8, Provirus, and Virus RNA. C Hallmark gene set enrichment analysis (GSEA) of MAP3K8highHTLV-1+CD4+ compared with MAP3K8lowHTLV-1+CD4+. Enriched gene sets were evaluated by normalized enrichment score and nominal P value. Statistical analysis was performed using a one-sided permutation test (1,000 permutations), as implemented in the GSEA algorithm. Nominal P values are reported, and gene sets with P < 0.05 are considered significantly enriched. The hallmark inflammatory response gene set is presented graphically. D The hierarchical clustering of enrichment levels of transcriptional factor motifs across cell types using scATAC-seq data. Heatmap displays z-score-normalized enrichment values, focusing on differences between MAP3K8highHTLV-1+CD4+ and MAP3K8lowHTLV-1+CD4+ cells. E Flowchart for identifying transcription factors specific to MAP3K8highHTLV-1+CD4+ cells. F Correlation between FOSL2 TPM and MAP3K8 TPM in HTLV-1-infected CD4+ T cells from patients with HAM (n = 13). Pearson’s correlation coefficient R = 0.91. G Boxplot of normalized FOSL2 expression levels in AC_P (n = 10), AC_D (n = 8), HAM_D (n = 13), iATL_D (n = 5), AC_N (n = 9), iATL_N (n = 9), and aATL_N (N = 13) from RNA-seq data. Each data point represents an individual patient sample. Boxes indicate the interquartile rage (IQR), with the line representing the median. Whiskers extend to 1.5 * IQR, and outliers beyond this range are shown as individual points. Statistical analysis was performed using one-way ANOVA. Exact P value are as follows: AC_P vs HAM_D, P = 0.00000037, AC_D vs HAM_D, P = 0.0154, HAM_D vs iATL_D, P = 0.0047, HAM_D vs AC_N, P = 0.00238, HAM_D vs iATL_N, P = 0.000022, HAM_D vs aATL_N, P = 0.00000057. Asterisks indicate statistical significance (* P < 0.05, ** P < 0.01, *** P < 0.001). (H) Correlation of FOSL2 expression (TPM) in HTLV-1-infected T cells with CSF biomarkers in HAM (n = 13): CSF CXCL10 (left panel, R = 0.77) and neopterin (right panel, R = 0.74).
Furthermore, we performed pseudotime analysis using single-cell multiome data and the Monocle3 algorithm to reconstruct cellular trajectories within the CD4⁺ T cell compartment. This analysis revealed distinct pseudotemporal states between HTLV-1–infected cells from HAM patients and those from ACs. Specifically, the inferred trajectory suggested a progression from cluster 4, which is enriched for uninfected CD4⁺ T cells, to cluster 1 (dominated by infected cells from ACs), and further toward clusters 0 and 16 (dominated by infected cells from HAM patients) (Supplementary Fig. 3E). Along this trajectory, we observed a transition toward increased chromatin accessibility at the HAM-characteristic OCR within the MAP3K8 locus, which was accompanied by upregulation of MAP3K8 and IFNG expression (Supplementary Fig. 3C, 3D, and 3E).
To further investigate the heterogeneity of HTLV-1-infected T cells in patients with HAM, we conducted single-cell multiome reanalysis. Transcriptome and chromatin accessibility profiles were obtained at the single-cell level from PBMCs of two HAM patients (n = 2) after enrichment of HTLV-1−infected T cells via flow cytometry (CD4+CADM1+CD7+ cells from HAM #14 and HAM #15). A total of 31,633 cells from each library were reanalyzed. UMAP clustering revealed multiple cell clusters. Notably, the HTLV-1−infected CD4+ T cell population in HAM patients was further subdivided into two distinct subclusters based on MAP3K8 expression: MAP3K8 highHTLV-1+CD4+ and MAP3K8 lowHTLV-1+CD4+ (Fig. 5B). The MAP3K8 highHTLV-1+CD4+ cells exhibited elevated expression of IFNG and TBX21 (Supplementary Fig. 4A). Gene set enrichment analysis (GSEA) revealed that MAP3K8 highHTLV-1+CD4+ cells had increased expression of inflammatory response-related genes compared to MAP3K8 lowHTLV-1+CD4+ cells (Fig. 5C).
To further characterize MAP3K8 highHTLV-1+CD4+ cells, we examined differential genome accessibility of transcription factors across cell populations using scATAC-seq data. MAP3K8 highHTLV-1+CD4+ cells showed distinct chromatin accessibility profiles compared to MAP3K8lowHTLV-1+CD4+ and uninfected CD4+ T cells. Notably, these cells exhibited increased accessibility for transcription factors, such as FOSL2-JUN and FOS-JUND (Fig. 5D), suggesting their involvement in MAP3K8 expression. We employed a multi-step approach to identify transcription factors critical for MAP3K8 expression1: From RNA-seq data, we identified 7,917 genes expressed in HTLV-1–infected cells from HAM patients (TPM > 10)2. From these genes, we selected 2,550 genes that were more highly expressed in MAP3K8highHTLV-1+CD4+ cells than in MAP3K8lowHTLV-1+CD4+ or uninfected CD4+ T cells3. Differential accessibility analysis identified 14 transcription factors with increased chromatin accessibility in MAP3K8highHTLV-1+CD4+ cells compared to MAP3K8lowHTLV-1+CD4+ and uninfected CD4+ T-cells (Fig. 5E)4. Among these, correlation analysis revealed a strong association between FOSL2 (fos-like 2) expression and MAP3K8 expression levels (Fig. 5F and Supplementary Fig. 4C). FOSL2 expression levels in HTLV-1–infected cells from HAM patients were significantly higher than those in HTLV-1–infected cells from ACs and from patients with ATL (Fig. 5G). Furthermore, FOSL2 expression strongly correlated with CSF concentrations of CXCL10 and CSF neopterin, two established biomarkers of HAM disease activity (Fig. 5H).
Fosl2, c-Jun, and HTLV-1 Tax cooperatively enhance MAP3K8 transcription through HAM-characterisic OCR binding
To determine whether Fosl2 (FOSL2 protein) binds to the HAM-characteristic OCR within the MAP3K8 locus, we performed chromatin immunoprecipitation sequencing (ChIP-seq) with Fosl2 in HCT-4 cells. The analysis revealed strong binding of Fosl2 to the HAM- characteristic OCR (Fig. 6A). We also examined the genomic localization of HTLV-1 Tax, a viral cofactor known to amplify transcriptional activity by binding to transcription factors. ChIP-seq for HTLV-1 Tax confirmed its strong binding to both the MAP3K8 transcription start site and the HAM-characteristic OCR (Fig. 6A). Further analysis of Fosl2 ChIP-seq data revealed significant enrichment of c-Jun and Fosl2 binding motifs within the Fosl2-bound genomic region (Fig. 6B). Given that Fosl2 is a transcription factor containing a basic leucine zipper (bZIP) domain, it forms heterodimers with other bZIP domain-containing transcription factors, functioning as part of the AP-1 transcription factor complex. Therefore, we hypothesized that Fosl2 functions by forming heterodimers with c-Jun in HTLV-1–infected cells from patients with HAM.
A Fosl2 and HTLV-1 Tax enrichment (ChIP-seq) in HCT-4 cells and chromatin accessibility (ATAC-seq) in HTLV-1-infected CD4+ T cells in patients with HAM, highlighting the HAM-characteristic OCR. Genomic DNA from HCT-4 cells prior to immunoprecipitation served as input control. ChIP-seq and ATAC-seq peaks were visualized with IGV (data range: ChIP-seq, 0-30; ATAC-seq, 0-100). B Enrichment analysis of transcription factor motifs in ChIP-seq with Fosl2 using the HOMER algorithm. Motifs are ranked by -log10(P). Statistical analysis was performed using a one-sided hypergeometric test. Nominal P values are reported. C Enhancer assays using the MAP3K8 promoter and HAM-characteristic OCR at the MAP3K8 locus. Relative luciferase activity for Fosl2 with partner factors (c-Jun, JunB, BATF, and BATF3) was examined. Data are presented as mean values + SD. Statistical significance was assessed by one-way ANOVA followed by two-sided Tukey’s honestly significant difference (HSD) test for multiple comparisons. Asterisks indicate statistical significance (*** P < 0.001). n = 3, 4, or 5 biologically independent experiments, each performed on separate days using independently transfected HEK293 cells. D Enhancer assays using the MAP3K8 promoter and HAM-characteristic OCR, with Fosl2, c-Jun, and HTLV-1 Tax. Yellow arrows indicate OCR orientation (5’ → 3’ or 3’ → 5’). Data are presented as mean values + SD. Statistical significance was assessed by one-way ANOVA followed by two-sided Tukey’s HSD test for multiple comparisons. Exact P value are as follows: MAP3K8 promoter with Fosl2, c-Jun, and HTLV-1 Tax vs MAP3K8 promoter and HAM-characteristic OCR (5’ → 3’) with Fosl2, c-Jun, and HTLV-1 Tax, P = 0.000010, MAP3K8 promoter with Fosl2, c-Jun, and HTLV-1 Tax vs MAP3K8 promoter and HAM-characteristic OCR (3’ → 5’) with Fosl2, c-Jun, and HTLV-1 Tax, P = 0.000021. Asterisks indicate statistical significance (*** P < 0.001). n = 4 or 7 biologically independent experiments, each performed on separate days using independently transfected HEK293 cells. E Representative histograms depicting chromatin accessibility at the MAP3K8 locus, as determined by ATAC-seq analysis, across various CD4⁺ T cell populations: resting normal CD4⁺ T cells, HAM_D subset, CD4⁺ T cells transiently expressing HTLV-1 Tax for 7 days via a lentiviral system (Tax-cell), and CD4⁺ T cells activated with anti-CD3/CD28 antibodies (GSM5554060). The highlighted region shows HAM-characteristic OCR. ATAC-seq peaks were visualized with IGV (data range: 0-100).
To test whether the AP-1 complex, formed by Fosl2 and c-Jun, induces enhancer activity through binding to the HAM-characteristic OCR (located approximately +13,000 bp from the MAP3K8 transcription start site), we conducted a luciferase assay. The human MAP3K8 promoter is reported to be in the region from −918 to +178 bp relative to the transcription start site of MAP3K827. Insertion of the HAM-characteristic OCR (457 bp) upstream of the MAP3K8 promoter markedly increased MAP3K8 transcriptional activity induced by Fosl2 and c-Jun (Fig. 6C). Notably, neither JunB (a member of the c-Jun family) nor BATF and BATF3 (ATF family transcription factors with bZIP domains) induced enhancer activity, highlighting the unique role of the Fosl2-c-Jun combination for this enhancer activity. We also evaluated whether HTLV-1 Tax, which binds the HAM-characteristic OCR at the MAP3K8 locus, further amplifies the enhancer activity induced by Fosl2-c-Jun. Indeed, HTLV-1 Tax significantly augmented the enhancer activity (Fig. 6D). To evaluate the expression levels of the transgenes, we performed western blot analysis. The results confirmed robust expression of Fosl2, c-Jun, JunB, BATF, BATF3, and Tax proteins (Supplementary Fig. 5). Interestingly, overexpression of BATF and BATF3 appeared to suppress Fosl2 protein levels. This observation aligns with previous reports suggesting that BATF can negatively regulate Fosl2 expression by competing for AP-1 complex formation, thereby destabilizing Fos-containing complexes28,29.
To evaluate whether HTLV-1 Tax drives the formation of HAM-characteristic OCR within the MAP3K8 locus, we analyzed ATAC-seq data from CD4 T+ cells transiently expressing HTLV-1 Tax for 7 days using a lentiviral vector system18. The data demonstrated an open chromatin state in this region, indicating that HTLV-1 Tax triggered chromatin remodeling at this site (Fig. 6E). Moreover, our analysis confirmed that chromatin accessibility at the MAP3K8 locus, which was identified as a HAM-characteristic OCR, was significantly lower in anti-CD3/CD28–stimulated CD4⁺ T cells (GSM5554060) compared to HTLV-1 Tax-expressing cells (fold change = 6.82, adjusted P = 1.78E-8).
To explore the regulation of FOSL2 and JUN (c-Jun) expression by the MEK-ERK pathway, we treated HCT-4 cells with trametinib and compared their expression levels using RNA-Seq. Trametinib treatment significantly reduced FOSL2 expression, whereas JUN expression remained unchanged (Supplementary Fig. 6A). A cross-sectional comparison of JUN expression across HTLV-1-associated diseases revealed consistently high expression of JUN in both HTLV-1-infected and -uninfected CD4+ T cells (Supplementary Fig. 6B). These findings suggest that FOSL2 is regulated downstream of the MEK-ERK pathway, whereas JUN is constitutively upregulated in CD4+ T cells.
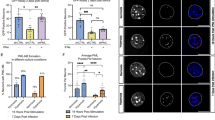
ChIP-seq analysis of HTLV-1 Tax revealed strong binding of HTLV-1 Tax to the FOSL2 promoter region in HCT-4 cells (Supplementary Fig. 6C), suggesting that HTLV-1 Tax promotes FOSL2 expression. Additionally, motif analysis by ChIP-seq with HTLV-1 Tax showed significant enrichment of Fosl2 and c-Jun binding motifs (Supplementary Fig. 6D). HTLV-1 Tax does not possess intrinsic DNA-binding activity. However, previous work by Vandermeulen et al. demonstrated that HTLV-1 Tax can interact with c-Jun via a yeast two-hybrid system and subsequent validation assays30. To further investigate this mechanism, we performed proximity ligation assays (PLA) to examine protein–protein interactions among HTLV-1 Tax, c-Jun, and Fosl2. The PLA results revealed that these proteins are in close proximity (within 40 nanometers), suggesting that they form a functional complex (Supplementary Fig. 7A and 7B). Based on these findings, we propose that HTLV-1 Tax is indirectly recruited to the MAP3K8 enhancer region and the FOSL2 promoter region through interactions with the AP-1 complex, particularly via c-Jun and Fosl2, thereby facilitating the transcriptional activation of MAP3K8.
To analyze the correlation between the expression of HTLV-1 Tax and those of MAP3K8, FOSL2, and JUN, we analyzed the correlation between the expression levels of these genes across six T-cell lines (HCT-4, HCT-5, MT-2, C91-PL, MT-1, and Jurkat), which exhibit varying levels of HTLV-1 Tax expression. Our analysis revealed a positive correlation among the expression levels of these genes, supporting the notion that HTLV-1 Tax promotes MAP3K8 transcription in concert with Fosl2 and c-Jun (Supplementary Fig. 6E).
Together, these findings indicate that Fosl2 and c-Jun colocalize with HTLV-1 Tax in the genome of HTLV-1−infected cells from HAM patients, cooperatively enhancing MAP3K8 transcriptional activity and contributing to HAM pathogenesis.
Inhibitory effects of MEK inhibitors on inflammatory responses in ex vivo PBMCs from patients with HAM
Given that constitutive activation of the MEK-ERK pathway by MAP3K8 amplifies inflammatory responses, we assessed the anti-inflammatory effects of MEK inhibitors using ex vivo PBMCs from patients with HAM (HAM-PBMCs). In ex vivo cultures, these PBMCs spontaneously proliferate in culture due to the expansion of HTLV-1-infected cells and immune responses against HTLV-1-infected T cells, especially CD8+ T cells (Supplementary Fig. 8A)31,32.
When HAM-PBMCs were cultured with various MEK inhibitors (binimetinib, cobimetinib, selumetinib, and trametinib), a significant reduction in 3H-thymidine incorporation was observed, indicating decreased cell proliferation, especially CD8+ T cells (Supplementary Fig. 8A and 8B). Prednisone (PSL) was used as a positive control, while vemurafenib, a B-Raf enzyme inhibitor targeting an upstream signaling factor of the MEK-ERK pathway unrelated to MAP3K8, served as a negative control and showed no effect, indicating that MEK-ERK pathway activation in HTLV-1−infected cells from HAM patients is independent of B-Raf.
Flow cytometry analysis demonstrated that MEK inhibitors, particularly trametinib, reduced the frequency of CCR4+Ki67+ cells, suggesting inhibition of mitotic entry in HTLV-1-infected cells (Supplementary Fig. 9A and Supplementary Fig. 10A). The MEK inhibitors also significantly reduced the number of Ki67+CD8+ T cells, which are critical contributors to the inflammatory pathogenesis of HAM (Supplementary Fig. 9B and Supplementary Fig. 10B). Importantly, in two PSL-refractory cases, MEK inhibitors reduced the frequency of both CCR4+Ki67+CD4+ and Ki67+CD8+ T cells (Supplementary Fig. 10C). The spontaneous proliferation of HAM-PBMCs results in an excessive immune activation, including the expansion of HTLV-1 Tax-specific cytotoxic T cells (CTLs), which play a pivotal role in HAM pathology33. MEK inhibitors suppressed both the number and proliferation of HTLV-1 Tax-specific CTLs (Supplementary Fig. 9C and Supplementary Fig. 10D and 10E).
Inflammatory cytokines such as IFN-γ, tumor necrosis factor (TNF)-α, and interleukin (IL)−6 are produced at high levels in the supernatant of cultured HAM-PBMCs, exacerbating inflammatory pathogenesis34. Treatment with MEK inhibitors, particularly trametinib and cobimetinib, significantly reduced the production levels of IFN-γ and TNF-α proteins, while IL-2 levels remained unaffected (Supplementary Fig. 9D).
Discussion
Although it is increasingly recognized that abnormal chromatin remodeling is involved in various disease pathogeneses, the detailed mechanisms by which abnormal chromatin remodeling leads to disease development remain unclear. In this study, we focused on the differences in HTLV-1-infected T cell characteristics between HAM and ATL, two distinct diseases caused by the same virus, HTLV-1. We hypothesized that HTLV-1 induces disease-specific chromatin remodeling in infected cells, thereby determining their fate and being responsible for disease development.
In this study, we identified MAP3K8 as a key gene regulating HTLV-1–infected cell characteristics from patients with HAM. We also found that abnormal HAM-characteristic chromatin remodeling occurred in the enhancer region of the MAP3K8 locus, where Fosl2, c-Jun, and HTLV-1 Tax bind to enhance MAP3K8 expression. Furthermore, we demonstrated that HTLV-1 Tax triggered chromatin remodeling in this region. The study also showed that MAP3K8 expression contributes to the differentiation and proliferation of HTLV-1-infected cells into Th1-like cells via MEK-ERK pathway activation and is closely linked to increased spinal cord inflammation in patients with HAM. HTLV-1 Tax and MEK-ERK pathway activation also contributed to FOSL2 expression. These results suggest that HTLV-1 Tax induces Fosl2 expression and that HTLV-1 Tax, Fosl2, and c-Jun induce abnormal chromatin remodeling in the enhancer region of the MAP3K8 gene locus, leading to MAP3K8 overexpression, which further induces Fosl2, resulting in a vicious cycle that contributes to HAM-characteristic pathogenesis (Supplementary Fig. 11). We further demonstrated that blocking this vicious cycle may be a promising strategy for treating HAM.
Our analysis revealed specific gene expression profiles unique to HTLV-1–infected cells from patients with HAM, distinguishing them from those of ATL and AC. Among the 328 genes commonly overexpressed in the infected cells, we identified well-known HTLV-1-infected cell markers, such as IL2RA, CCR4, and CADM1 (Supplementary Table 4), underscoring the accuracy of our gene profiles. In particular, MAP3K8 and FOSL2 were prominently expressed in HTLV-1–infected cells from patients with HAM, suggesting their pivotal roles in HAM pathogenesis. We detected differentially accessible 892 OCRs by comparing the chromatin accessibility profiles between HTLV-1–infected cells from patients with HAM and those with aATL (Supplementary Table 3). These OCRs included TBX21, a master transcription factor for Th1, and the Th1 cell markers IFNG and CXCR3, indicating that the Th1-like differentiation phenotype of HTLV-1–infected cells from HAM patients was epigenetically maintained (Supplementary Fig. 12). These results highlight the critical role of disease-characteristic chromatin remodeling in defining the distinct outcomes of HTLV-1 infection, such as HAM and ATL.
This study unveiled a mechanism for HAM-characteristic abnormal chromatin remodeling in the enhancer region of the MAP3K8 locus driven by the collaborative binding of Fosl2, c-Jun, and HTLV-1 Tax, which amplified the enhancer activity of MAP3K8 gene expression. HTLV-1–infected cells from patients with HAM express higher levels of HTLV-1 Tax compared to those from ACs and ATL patients14,35. HTLV-1 Tax lacks a DNA-binding domain and thus cannot bind directly to DNA but regulates the transcription of target genes by binding to transcription factors. HTLV-1 Tax also binds to BRG1, a representative ATPase subunit of the chromatin remodeling factor switch/sucrose nonfermenting (SWI/SNF) complex, and CBP/p300, a histone acetyltransferase, and transports these factors into the genome to induce chromatin remodeling and transcriptional activity amplification36,37. We demonstrated that HTLV-1 Tax interacts with both Fosl2 and c-Jun, and is recruited to HAM-characteristic OCRs containing the AP-1 binding sites (Fosl2 and c-Jun). Thus, HTLV-1 Tax can act directly to maintain HAM-characteristic OCR at the MAP3K8 locus. Furthermore, the AP-1 complex functions as a pioneer factor that transiently interacts with heterochromatin and directly recruits the SWI/SNF complex38,39,40. HTLV-1 Tax increased FOSL2 expression41, and JUN was highly expressed in CD4+ T cells, including the infected cells. Indeed, HTLV-1 Tax triggered chromatin remodeling and formed an open chromatin structure in this region of CD4+ T cells. Therefore, HTLV-1 Tax is suggested to be the HAM pathogenesis initiator, forming an AP-1 complex comprising Fosl2 and c-Jun in HTLV-1-infected cells, which, together with chromatin remodeling enhancement by HTLV-1 Tax, induces chromatin remodeling in the intronic region of MAP3K8 locus and forms the HAM-characteristic OCR.
HTLV-1 Tax expression is transient and limited to a small subset of infected cells. In contrast, the majority of infected cells exhibiting high MAP3K8 expression lack detectable levels of HTLV-1 Tax. We hypothesize that the initial recruitment of the HTLV-1 Tax–Fosl2–c-Jun complex to the MAP3K8 enhancer acts as a chromatin-opening event, thereby facilitating subsequent access by transcription factors such as Fosl2 even in the absence of continued HTLV-1 Tax expression. Notably, activation of the MAP3K8–MEK–ERK signaling axis has been shown to upregulate FOSL2 expression (Supplementary Fig. 6A), suggesting the establishment of a positive feedback loop between Fosl2 and the MAP3K8-MEK-ERK pathway (Supplementary Fig. 11). Thus, even after the decline of HTLV-1 Tax expression, sustained occupancy of the MAP3K8 enhancer by the Fosl2–c-Jun complex may maintain elevated MAP3K8 transcription in HTLV-1–infected T cells from patients with HAM.
This study showed that MAP3K8, which is overexpressed in HTLV-1–infected cells from patients with HAM owing to abnormal chromatin remodeling, enhances inflammatory responses by promoting Th1-like differentiation and constitutively activating the MEK-ERK pathway. The strong correlation between MAP3K8 expression in HTLV-1–infected cells from patients with HAM and CSF markers of disease activity (CXCL10 and neopterin) indicates a central role for MAP3K8 in HAM inflammatory pathology. Activation of the MAP3K8-MEK-ERK pathway drives IFN-γ production in HTLV-1–infected cells from HAM patients and further induces FOSL2 expression. Thus, activation of this pathway maintains the constant MAP3K8 expression and sustained IFN-γ production, leading to CXCL10 and neopterin overproduction, resulting in a chronic inflammatory loop. Therefore, the mechanism that initiates constant MAP3K8 overexpression may be pivotal in HAM pathogenesis.
Preventing this vicious cycle of chronic inflammation, centered on MAP3K8 overexpression, is crucial for developing effective HAM therapies. MEK inhibitors, particularly trametinib, suppressed the proliferation of HTLV-1–infected cells and CD8+ T cells, including Tax-specific CTLs, in an ex vivo inflammatory response model using HAM-PBMCs. Since MEK-ERK pathway activation is essential for the proliferation of TCR-stimulated CD8+ T cells42,43, MEK inhibitors could be effective against hyperactivated CD8+ T cells by viral antigens. Trametinib and cobimetinib also significantly suppressed the amount of IFN-γ produced by HTLV-1–infected and inflammatory cells from HAM patients, making these MEK inhibitors promising candidates for controlling inflammatory pathologies in HAM. Given the challenges in developing additional drugs for rare diseases like HAM, repurposing existing drugs such as MEK inhibitors could provide a practical and promising therapeutic approach.
In conclusion, we identified MAP3K8 as a molecule that defines the unique pathological characteristics of HAM. By integrating transcriptome, chromatin accessibility profiles, and disease activity biomarkers across HTLV-1-related diseases, we elucidated the mechanism of MAP3K8 overexpression induced by HAM-characteristic abnormal chromatin remodeling. Our findings highlight the central role of MAP3K8-MEK-ERK pathway in HAM pathogenesis and its potential as a therapeutic target. This study underscores the importance of viral factors driving abnormal chromatin remodeling and disease pathogenesis, providing crucial insights into pathogen-host interactions and offering a foundation for future therapeutic development.
Methods
Participants and sample preparation
PBMCs were collected from patients with HAM (N = 35) and ATL (N = 2), and ACs (N = 3) using Pancoll (PANBiotech GmbH, Aidenbach, Germany). CSF was obtained from the patients via lumbar puncture. HTLV-1 seropositivity was determined using a particle agglutination assay (Serodia-HTLV-1; FUJIREBIO. Inc., Tokyo, Japan) and confirmed by western blotting (SRL Inc., Tokyo, Japan). HAM was diagnosed according to the World Health Organization guidelines44. This study was approved by the Institutional Ethics Committee of St. Marianna University (approval number 1646) and the University of Tokyo (approval numbers 19-51 and 18-265), and all participants provided written informed consent.
HTLV-1-infected CD4+T cell sorting
HTLV-1-infected CD4+ T cells were obtained using the HAS-flow method, as previously described20,21. Purified PBMCs were stained with the following fluorescent-labeled antibodies: anti-CADM1-biotin (1:100; CM004-6, MBL), anti-CD7-APC (5:100; clone CD7-6B7), anti-CD3APC-Cy7 (5:100; clone SK7), anti-CD4-Pacific Blue (5:100; clone RPA-T4), and anti-CD14-Pacific Orange (5:100). After washing with phosphate-buffered saline (PBS), phycoerythrin-conjugated streptavidin (2:100; 554061; BD Biosciences) was added. Propidium iodide (Sigma-Aldrich) was added to the samples to exclude dead cells. CD3+CD4+CADM1+CD7+CD14−cells from HTLV-1-infected T cells of patients with HAM were sorted using FACSAria-II SORP (BD Biosciences).
RNA-sequence
Total RNA from each sample was extracted using the TRIzol reagent (Invitrogen), and the quantity and quality were assessed using an Agilent 2100 Bioanalyzer (Agilent Technologies). A total of 20 ng of RNA with a RIN value above seven was used for library preparation. Library preparation and sequencing were performed using GeneWiz (Azenta Life Sciences). Libraries with different indices were multiplexed and loaded on an Illumina HiSeq instrument per manufacturer instructions (Illumina). Sequencing was conducted using a 2 × 150-bp paired-end configuration, and image analysis and base calling were performed using the HiSeq Control Software (HCS v2.2.38 or later) along with OLB and GAPipeline-1.6 (Illumina). For quality control, technical sequences, including adapters, PCR primers, and low-quality bases (below Q20), were removed from the pass-filter data in fastq format using Trimmomatic (v0.30) to generate high-quality clean data. The reference genome was indexed using Hisat2 (v2.0.1), and the clean data were aligned with the reference genome using Hisat2.
ATAC-sequence
HTLV-1-infected CD4+ T cells (100 K) in patients with HAM were isolated using a FACSAria-II cell sorter (BD Biosciences). These cells were fixed with 1% formaldehyde, and further processing and ATAC experiments, including chromatin extraction, fragmentation, and library preparation, were performed using Active Motif (Carlsbad, CA, USA). The resulting materials were sequenced with a paired end of 42 bp on a NovaSeq6000 (Illumina). The output reads were aligned using the Burrows-Wheeler Aligner algorithm. Duplicate reads were removed and only reads mapped as matched pairs and uniquely mapped reads were used for further analysis. Peaks with significant differences were identified using the MACS algorithm. For the whole-genome region, the number of sequenced reads was converted to tag numbers, which were used as epigenomic data. In addition, bigwig format files that could draw read counts were drawn using Integrative Genomics Viewer (IGV). EaSeq software (v1.111) was used to calculate peak values.
Single-cell (sc) multiome analysis
The sc multiome libraries were constructed using the Chromium Controller and the 10x Genomics Chromium Next GEM Single Cell Multiome ATAC plus Gene Expression kit, per manufacturer protocols (CG000365 Rev C, CG000338 Rev F, 10x Genomics). Libraries were sequenced using the NovaSeq 6000 system (Illumina)per manufacturer instructions. For ATAC libraries, sequencing was performed with a 50 × 49-bp paired-end configuration, while RNA library sequencing used a 28 × 91-bp paired-end configuration. The sc multiome dataset was initially processed using Cell Ranger ARCv2.0.0 (Cell Ranger ARC, 10x Genomics). BCL files were converted to FASTQ files using the cellranger_ark mkfastq command with the default parameters. The FASTQ files were then processed using the cellranger_ark count and merged using cellranger-arc agr. To remove the batch effect, the sc Multiome RNA dataset was processed using Seurat (v4.3.0) reciprocal PCA (clustering parameters: PCA dimensions 1–30, resolution 0.5). The sc Multiome ATAC dataset was recounted using Signac (v1.9.0) using merged peak bed files and processed using Harmony (v0.1.1). Cell Ranger ATAC was used to perform differential accessibility analysis to identify transcription factor motifs whose accessibility was specific to each cluster. The output data were subjected to hierarchical clustering analysis using R (v4.3.0).
Trajectry analysis using Monocle3
Psudotime trajectory analysis was performed using the Monocle3 package (v1.3.7) in R (v4.4.1) to infer lineage relationship among CD4+ T cell populations. A subset of cells annotated as CD4+ T cells was extracted from the integrated Seurat object containing single-cell multiome data. The Seurat object was then converted to a cell_data_set object using as.cell_data_set() function. Dimensionality reduction was performed UMAP embeddings derived from WNN analysis. Clusters were learned with the cluster_cells() function, and a principal graph was constructed using learn_graph(). The root node was manually assigned based on the cluster enriched in uninfected CD4+ T cells. Pseudotime values were calculated using the order_cell() function, and the inferred trajectory was visualized over the WNN UMAP. All trajectory analyses were conducted following the recommended Monocle3 workflow, and cell state transitions were interpreted in the context of chromatin accessibility and gene expression changes along pseudotime.
Measurement of CXCL10 and neopterin in CSF
CSF neopterin and CXCL10 levels were measured using high-performance liquid chromatography at a commercial laboratory (SRL Inc.) and a cytometric bead array (BD Biosciences), respectively.
Cell culture
PBMCs from patients with HAM were cultured in RPMI 1640 medium (FUJIFILM Wako Chemicals) supplemented with 10% heat-inactivated fetal bovine serum (FBS; GIBCO) and 1% penicillin/streptomycin (FUJIFILM Wako Chemicals) in 96-well round-bottom plates. Binimetinib (ONO PHARMACEUTICAL Co., LTD), cobimetinib (Roche Applied Science), selumetinib (AstraZeneca), and trametinib (NOVARTIS) were used as the MEK inhibitors. Vemurafenib (Cayman Chemical), a B-Raf kinase inhibitor, was used as the negative control. PSL (LKT Laboratories, Inc.) was used as a positive control. Jurkat (a T-cell acute lymphoblastic leukemia cell line), HUT102, C91-PL, and MT-2 (HTLV-1-infected T cell lines), and MT-1 and TL-Om1 (ATL cell lines) were cultured in RPMI 1640 medium with 10% heat-inactivated FBS and 1% penicillin/streptomycin. HCT-4, HCT-5, and HCT-1, HTLV-1-infected T cell lines derived from patients with HAM, and KK-1, an ATL cell line, were cultured in RPMI 1640 medium supplemented with 10% or 20% heat-inactivated FBS, 1% penicillin/streptomycin, and 100 U/ml or 200 U/mL recombinant human IL-2 (PEPRO TECH, Thermo Fisher Scientific). HEK293FT, HEK293, and HeLa cells were cultured in Dulbecco’s modified eagle medium supplemented with 10% heat-inactivated FBS and 1% penicillin/streptomycin. Normal (HTLV-1-uninfected) CD4+ T cells from healthy donors were purchased from Lonza (Basel, Switzerland).
Measurement of inflammatory cytokines (IFN-γ, TNF-α, IL-2, and IL-6)
HAM-PBMCs were cultured in RPMI 1640 medium with 10% heat-inactivated FBS and 1% penicillin/streptomycin, with or without MEK inhibitors, for 72 h in a 96-well round-bottom plate. The supernatants were collected, and inflammatory cytokines (IFN-γ, TNF-α, IL-2, and IL-6) were measured using a cytometric beads array (BD Biosciences). All assays were performed per manufacturer instructions.
Cell proliferation assay
PBMCs from patients with HAM (n = 7) were seeded at a density of 1 × 105 cells/well in 96-well round-bottom plates. The cells were cultured for 7 days in the presence or absence of the MEK inhibitors, vemurafenib, or PSL. During the final 16 h, 1 μCi of 3H-thymidine was added to each well, after which the cells were harvested and counted using a 2450 MicroBeta2 Plate Counter (Perkin Elmer, Boston, MA). The assay was performed in triplicates. PBMCs from seven additional patients with HAM were used to verify the effects of MEK inhibitors. Using the average counts of 3H-thymidine incorporation in the DMSO-treated group as 100%, the average relative value in each drug-treated group was calculated and defined as the 3H-thymidine incorporation rate (%). All primary cultures were maintained at 37 °C with 5% CO2.
Lentiviral vector-mediated transfection
To overexpress MAP3K8, a replication-defective, self-inactivating lentivirus vector (CSII-EF-MCS-IRES-Venus) was used (Riken, BRC), and hemagglutinin (HA)-tagged MAP3K8 cDNA was subcloned into the lentivirus vector. The established viral vectors were co-transfected with the packaging plasmid (pCAG-HIVgp) and VSV-G- and Rev-expressing plasmids (pCMV-VSV-G-RSV-Rev) into 293FT cells. High-titer viral solutions were prepared by centrifugation and used for transduction into Jurkat cells. HA-tagged MAP3K8 expression was validated by western blotting using an anti-HA antibody (#561-5, MBL).
Primers used for the cloning of the MAP3K8 gene were as follows:
MAP3K8 Fw: GCGGCCGCatggatatctatccatatgacgttccagattatg
MAP3K8 Rv: GGATCCtcagccatattcaagcgttggtg
Gene knockdown by siRNA
To knock down MAP3K8, siRNA targeting the MAP3K8 gene was introduced into HAM-derived cell lines, ATL cell lines, and primary ATL cells (2 × 10⁵ cells each) by electroporation using the AMAXA Nucleofector II system in conjunction with either the Cell Line Nucleofector Kit V or the Human T Cell Nucleofector Kit according to the manufacturer’s instructions (both from Lonza Bioscience, Switzerland). Electroporation was performed using Nucleofector programs X-001 and U-014. Transfected cells were harvested after 24 hours, and their RNA was extracted using TRIzol reagent (Thermo Fisher Scientific) followed by purification with the RNeasy Mini Kit (Qiagen). The following siRNA were used in this study:
MAP3K8 siRNA: J-003511-17-0005 (Dharmacon, Lafayette, CO)
Non-targeting control siRNA: D-001810-10-0 (Dharmacon)
Western blot
Total cellular proteins were isolated using cell lysis buffer (10 mM Tris-HCl, pH 7.4, 0.1% sodium dodecyl-sulfate (SDS), 1% NP-40, 0.1% Sodium Deoxycholate, 0.15 M NaCl, and 1 mM EDTA) including Protease inhibitor cocktail (NACALAI TESQUE, Kyoto, Japan). Whole-cell lysates were subjected to SDS-polyacrylamide gel electrophoresis and transferred to a polyvinylidene difluoride membrane. Membranes were probed with primary antibodies, followed by alkaline phosphatase-conjugated secondary antibodies. The antibodies and detection reagents used were as follows:
Anti-MAP3K8 antibody (N3C3, GeneTex, Irvine, CA)
Anti-HA antibody (#561-5, MBL, Tokyo, Japan)
Anti-phospho-MEK1/2 (Ser217/221) antibody (#9121, Cell Signaling Technology)
Anti-MEK1/2 antibody (#9154S, Cell Signaling Technology)
Anti-phospho-ERK1/2 (Thr202/Tyr204) antibody (#4370, Cell Signaling Technology)
Anti-ERK1/2 antibody (#4695S, Cell Signaling TECHNOLOGY)
Anti-phospho-p38 (Thr180/Tyr182) antibody (#9211, Cell Signaling Technology)
Anti-p38 antibody (#53520, BD Transduction Laboratories)
Anti-β-actin antibody (sc-69879, Santa Cruz Biotechnology)
Anti-mouse IgG (H + L) AP Conjugate secondary antibody (S3728; Promega).
Anti-rabbit IgG (H + L) AP Conjugate secondary antibody (S373B; Promega).
Immunoreactive proteins were visualized using the BCIP/NBT Color Development Substrate (5-bromo-4-chloro3-indolyl-phosphate/nitro blue tetrazolium, Promega).
Anti-c-Jun antibody (9165S, Cell Signaling Technology)
Anti-JunB antibody (sc-8051, Santa Cruz Biotechnology)
Anti-BATF antibody (8638 T, Cell Signaling Technology)
Anti-BATF3 antibody (GTX130125, Gene Tex)
Anti-Tax antibody (LT-4, kindly gifted by Dr. Yuetsu Tanaka)
Anti-Rabbit IgG (whole molecule)-Peroxidase antibody (A9169, Sigma-Aldrich)
Anti-Mouse IgG (whoke molecule)-Peroxidase antibody (A5278, Sima-Aldrich)
ECL Prime Western Blotting Detection Reagent (Cytiva, Little Chalfront, England)
Proximity ligation assay (PLA)
Proximity ligation assays were performed using the Duolink In Situ Detection Reagents Red (DUO92008, Sigma-Aldrich, St. Louis, MO) according to the manufacturer’s instructions. Briefly, HeLa cells (1 × 10⁵) were transfected using Lipofectamine 2000 (Thermo Fisher Scientific) with 0.5 μg of plasmids encoding FOSL2, JUN, JUNB, and HTLV-1 Tax. After culture for 24 hours, cells were fixed by 4% paraformaldehyde in PBS for 15 minutes at room temperature, followed by permeabilization with 0.1 % Triton X in PBS for 10 minutes. After blocking with Duolink Blocking solution (Sigma-Aldrich) for 1 hour at 37°C, cells were incubated overnight at 4°C with a pair of primary antibodies raised in different spices (e.g., rabbit anti-FOSL2 and mouse anti-HTLV-1 Tax). Following washes, secondary antibodies conjugated with PLA probes (PLUS and MINUS) were added and incubated for 1 hour at 37°C. Ligation and amplification reactions were carried out sequentially according to the kit instructions. Fluorescent signals indicating protein-protein proximity (<40 nm) were visualized using a fluorescence microscope (BZ-X810, KEYENCE, Osaka, Japan). DAPI was used for nuclear counterstaining. Image analysis was performed using ImageJ (version: 2.16.0/1.54p). PBMCs isolated from patients with HAM and cultured for 48 hours were subjected to proximity ligation assays to assess protein-protein interactions among Fosl2, c-Jun, and HTLV-1 Tax. PBMCs from a healthy donor served as a negative control.
Enhancer luciferase assay
HEK293 cells (1 × 10⁵) were transfected using Lipofectamine 2000 (Thermo Fisher Scientific) with 0.5 μg each of plasmids encoding FOSL2, JUN, JUNB, BATF, BATF3, and HTLV-1 Tax. Empty plasmids were used as negative control. Additionally, cells were co-transfected with 0.5 μg of a pGL3 vector containing the MAP3K8 promoter, with or without the HAM-characteristic OCR inserted in the forward (5’ to 3’) or reverse (3’ to 5’) orientation upstream of a firefly luciferase reporter gene (Promega), and 0.05 μg of a RSV renilla luciferase reporter plasmid (Promega) for normalization purposes. Transfections were performed in 24-well plates. After 48 hours, cells were harvested and lysed using Passive Lysis Buffer (Promega). Firefly and renilla luciferase activities were measured using the Dual-Luciferase Reporter Assay System (Promega) and Varioskan LUX multimode microplate reader (Thermo Fisher Scientific) according to the manufacturer’s instructions.
Real-time quantitative polymerase chain reaction (qPCR) analysis
MAP3K8-overexpressing Jurkat cells were lysed, and total RNA was extracted and reverse-transcribed using ReverTra Ace qPCR RT Master Mix with gDNA Remover (TOYOBO). Relative mRNA quantification was performed using a thermal cycler dice real-time PCR system (Takara Bio Inc., Shiga, Japan) and the delta-delta Ct method was used to calculate relative gene expression levels. The RPL19 gene was used as an internal control. The primers used were as follows:
RPL19 Fw: accaaggaagcacgcaagc
RPL19 Rv: cagacaaagtgggaggttttatttc
IFNG Fw: acgagatgacttcgaaagctga
IFNG Rv: gacagttcagcatcacttgga
TBX21 Fw: aggattccgggagaactttg
TBX21 Rv: ccccaaggaattgacagttgg
MAP3K8 Fw: gcaagaggctgctgagtagga
MAP3K8 Rv: attcctcggtgcttcctgtg
CD69 Fw: acaactggttcaacgttacagg
CD69 Rv: agcagcatccactgacacag
CTLA4 Fw: ctacctgggcataggcaacg
CTLA4 Rv: ccccgaactaactgctgcaa
Flow cytometric analysis
To analyze the effects of MEK inhibitors on PBMCs from patients with HAM, PBMCs cultured with MEK inhibitors were stained with fluorochrome-conjugated antibodies and analyzed using FACSCanto-II and FACSCelesta (BD Biosciences). To detect Ki-67 expression, PBMCs were fixed using the Foxp3/Transcription Factor Staining Buffer Set (00-5523-00; eBioscience). HTLV-1 tax-specific CTLs in patients with HAM with HLA-A*24:02 were detected using the HLA-A*24:02-restricted Tax tetramer (HTLV-1 Tax 301-309 Peptide, TS-M018-1, MBL).
The antibodies used were as follows:
Anti-CD3-BV510 (clone: UCHT1, BioLegend)
Anti-CD4-APC (clone: RPA-T4, TONBO; Cytek Biosciences, Fremont, CA, USA)
Anti-CD8-PE-cy7 (clone: RPA-T8, TONBO)
Anti-CD45RO-BV785 (clone: UCHL1, BioLegend)
Anti-CCR4-BV421 (clone: 1G1, BD Biosciences)
Anti-Ki67-FITC (clone: MOPC-21, BD Biosciences)
Anti-CD19-PEcy7 (clone: HIB19, eBioscience, Thermo Fisher Scientific)
Anti-CD56-PE (clone: MEM-188, Biolegend)
Anti-CD8-APC (clone: RPA-T8, TONBO)
Anti-CD4-PEcy7 (clone: OKT4, Biolegend)
Anti-CD3-APCcy7 (UCHT1, TONBO)
MAP3K8 induction to normal CD4-positive T cells
High-titer lentiviral solutions for MAP3K8 induction were prepared by centrifugation and used to transduce normal CD4+ T-cells (1 × 106 cells). After 72 h of cell culture in RPMI 1640 medium with 10% heat-inactivated FBS and 1% penicillin/streptomycin, the cells were treated with GolgiStop (BD Biosciences) for an additional 5 h. The cells were then fixed using the Human FoxP3 Buffer Set (BD Biosciences) and stained with anti-T-bet-BV421 (clone: 4B10, Cell Signaling Technology) and anti-IFN-γ-APC (clone: 4SB3, BioLegend). Since Venus-positive cells indicate MAP3K8-expressing cells, T-bet and IFN-γ expression in Venus-positive cells was analyzed using FACSCanto-II flow cytometry (BD Biosciences).
Activation of CD4 T cells using anti-CD3 and anti-CD28 antibodies
Anti-CD3 antibodies (final concentration: 1 μg/mL, clone: OKT3, Miltenyi, Bergisch Gladbach, Germany) and anti-CD28 antibodies (final concentration: 5 μg/mL, clone: 15E8, Miltenyi) were coated on 24-well plates. Normal CD4+ T cells were incubated and stimulated for 48 h in RPMI 1640 medium supplemented with 10% heat-inactivated FBS and 1% penicillin/streptomycin in the presence of DMSO or 1 μM trametinib. The cells were fixed using the Human FoxP3 Buffer Set (BD Biosciences) and then stained with anti-T-bet-BV421 (Cell Signaling Technology) and anti-IFN-γ-APC (BioLegend). The cells were analyzed using an FACSCanto-II flow cytometer (BD Biosciences).
ChIP-seq
HCT-4 cells (4 × 106) were fixed by adding 1/37 volume of freshly prepared formaldehyde solution (final conc. 1%, FUJIFILM Wako Chemicals) to the medium and incubated for 10 min at room temperature. Fixation was stopped by adding a glycine solution and incubating for 5 min at room temperature. ChIP experiments, including chromatin extraction, fragmentation, and antibody precipitation, were performed using the SimpleChIP Enzymatic Chromatin IP Kit (#9003, Cell Signaling Technology) per manufacturer instructions. For chromatin digestion, 0.5 μL of micrococcal nuclease was added and incubated for 20 min at 37°C, followed by sonication to extract chromatin from the nucleus using Bioruptor UCD-250 (Cosmo Bio LTD., Japan). For immunoprecipitation (IP), 400 ng of anti-Fosl2 antibody (Cell Signaling Technology, #19967), anti-Tax (Lt-4) antibody (kindly provided by Dr. Yuetsu Tanaka), and Normal Rabbit IgG (#2729 Cell Signaling Technology) were used. IP fragments were washed and purified, and the purified DNA fragments were used for Illumina sequencing library preparation using GENEWIZ (Azenta Life Sciences). Illumina sequencing libraries were prepared from ChIP and input DNAs using standard enzymatic steps: end-polishing, dA-addition, and adaptor ligation. After the final PCR amplification, the resulting DNA libraries were quantified and sequenced on an Illumina Novaseq (150 nt reads, paired-end). The reads were aligned to the human genome (hg38) using the Bowtie2 algorithm (v2.5.2). The output SAM files were converted to BAM files using Samtools (v1.19), and peak calling for Fosl2 and Tax was performed using the MACS2 algorithm (v2.2.7.1) with a Q-value cutoff of 10-2. Motif analysis of Fosl2 and Tax ChIP-seq data was performed using the HOMER algorithm (v4.11).
Data analysis
The IGV tool was used to visualize RNA-seq, ATAC-seq, and ChIP-seq results. GSEA was performed using GSEA software (v4.1.0) (https://www.gsea-msigdb.org/gsea) with 1000 permutations. Gene sets were selected from the MSigDB hallmark gene sets (https://www.gsea-msigdb.org/gsea/msigdb/human/collections.jsp). DAVID was used for Gene Ontology (GO) analysis.
Data visualization
Box, beeswarm, and bar plots were generated and visualized using R (v4.3).
Statistical and reproducibility
All bar and line graphs that summarizing multiple datasets represent mean values, and box plots indicate median values as the center lines. Significant differences in gene expression and other biological assays between the two groups were analyzed using a two-sided Student’s t-test or paired t-test, as appropriate. A two-sided one-way analysis of variance or a two-sided the Kruskal‒Wallis test was used for comparisons among three or more groups. Correlations between two variables were assessed using two-sided Pearson’s or Spearman’s correlation coefficients. Bonferroni correction was employed to identify genes with statistically significant differences in RNA-seq data comparing HTLV-1–infected cells from patients with HAM and ATL, and those of ACs.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Sequencing data (FASTQ format), including RNA-seq, ATAC-seq, scRNA-seq, and scATAC-seq data for patients with HAM samples were deposited in the National Bioscience Database Center Human Database (JGAS000835, https://humandbs.dbcls.jp/en/hum0225-v2). Previously published RNA-seq and ATAC-seq datasets (JGAS000301 and JGAS000553, https://humandbs.dbcls.jp/en/hum0252-v3) were compared with the HAM patient dataset. The reference human genome, hg38, was downloaded from the UCSC Genome Browser. The publicly available chromatin accessibility data used in this study are available in the Gene Expression Omnibus (GEO) database under accession code GSE116696 and GSM5554060. Source data are provided with this paper.
References
Knezetic, J. A. & Luse, D. S. The presence of nucleosomes on a DNA template prevents initiation by RNA polymerase II in vitro. Cell 45, 95–104 (1986).
Han, M. & Grunstein, M. Nucleosome loss activates yeast downstream promoters in vivo. Cell 55, 1137–1145 (1988).
Li, B., Carey, M. & Workman, J. L. The role of chromatin during transcription. Cell 128, 707–719 (2007).
Klemm, S. L., Shipony, Z. & Greenleaf, W. J. Chromatin accessibility and the regulatory epigenome. Nat. Rev. Genet 20, 207–220 (2019).
Struhl, K. Histone acetylation and transcriptional regulatory mechanisms. Genes Dev. 12, 599–606 (1998).
Clapier, C. R. & Cairns, B. R. The biology of chromatin remodeling complexes. Annu Rev. Biochem 78, 273–304 (2009).
Santos-Rosa, H. & Caldas, C. Chromatin modifier enzymes, the histone code and cancer. Eur. J. Cancer 41, 2381–2402 (2005).
Berson, A., Nativio, R., Berger, S. L. & Bonini, N. M. Epigenetic regulation in neurodegenerative diseases. Trends Neurosci. 41, 587–598 (2018).
Einsiedel, L. et al. Human T-Lymphotropic Virus type 1 infection in an Indigenous Australian population: epidemiological insights from a hospital-based cohort study. BMC Public Health 16, 787 (2016).
Bangham, C. R., Araujo, A., Yamano, Y. & Taylor, G. P. HTLV-1-associated myelopathy/tropical spastic paraparesis. Nat. Rev. Dis. Prim. 1, 15012 (2015).
Yamagishi, M., Fujikawa, D., Watanabe, T. & Uchimaru, K. HTLV-1-mediated epigenetic pathway to adult T-cell leukemia-lymphoma. Front Microbiol 9, 1686 (2018).
Bangham, C. R. M. Human T cell leukemia virus type 1: persistence and pathogenesis. Annu Rev. Immunol. 36, 43–71 (2018).
Olière, S. et al. HTLV-1 evades type I interferon antiviral signaling by inducing the suppressor of cytokine signaling 1 (SOCS1). PLoS Pathog. 6, e1001177 (2010).
Yamano, Y. et al. Correlation of human T-cell lymphotropic virus type 1 (HTLV-1) mRNA with proviral DNA load, virus-specific CD8(+) T cells, and disease severity in HTLV-1-associated myelopathy (HAM/TSP). Blood 99, 88–94 (2002).
Araya, N. et al. HTLV-1 induces a Th1-like state in CD4+CCR4+ T cells. J. Clin. Invest 124, 3431–3442 (2014).
Ando, H. et al. Positive feedback loop via astrocytes causes chronic inflammation in virus-associated myelopathy. Brain 136, 2876–2887 (2013).
Kadoch, C. & Crabtree, G. R. Mammalian SWI/SNF chromatin remodeling complexes and cancer: Mechanistic insights gained from human genomics. Sci. Adv. 1, e1500447 (2015).
Mizuike, J. et al. Rewired chromatin structure and epigenetic gene dysregulation during HTLV-1 infection to leukemogenesis. Cancer Sci. https://doi.org/10.1111/cas.16388 (2024).
Yamagishi, M. et al. Chronological genome and single-cell transcriptome integration characterizes the evolutionary process of adult T cell leukemia-lymphoma. Nat. Commun. 12, 4821 (2021).
Kobayashi, S. et al. CADM1 expression and stepwise downregulation of CD7 are closely associated with clonal expansion of HTLV-I-infected cells in adult T-cell leukemia/lymphoma. Clin. Cancer Res 20, 2851–2861 (2014).
Sato, T. et al. Mogamulizumab (Anti-CCR4) in HTLV-1-associated myelopathy. N. Engl. J. Med 378, 529–538 (2018).
Nagasaka, M. et al. Mortality and risk of progression to adult T cell leukemia/lymphoma in HTLV-1-associated myelopathy/tropical spastic paraparesis. Proc. Natl. Acad. Sci. USA 117, 11685–11691 (2020).
Sasaki, H. et al. Overexpression of a cell adhesion molecule, TSLC1, as a possible molecular marker for acute-type adult T-cell leukemia. Blood 105, 1204–1213 (2005).
Wong, R. W. J. et al. Enhancer profiling identifies critical cancer genes and characterizes cell identity in adult T-cell leukemia. Blood 130, 2326–2338 (2017).
Takahashi, R. et al. Epigenetic deregulation of Ellis Van Creveld confers robust Hedgehog signaling in adult T-cell leukemia. Cancer Sci. 105, 1160–1169 (2014).
Acuff, N. V., Li, X., Kirkland, R., Nagy, T. & Watford, W. T. Tumor progression locus 2 differentially regulates IFNγ and IL-17 production by effector CD4+ T cells in a T cell transfer model of colitis. PLoS One 10, e0119885 (2015).
Schmid, S., Sachs, D. & tenOever, B. R. Mitogen-activated protein kinase-mediated licensing of interferon regulatory factor 3/7 reinforces the cell response to virus. J. Biol. Chem. 289, 299–311 (2014).
Rauscher, F. J. 3rd, Voulalas, P. J., Franza, B. R. Jr & Curran, T. Fos and Jun bind cooperatively to the AP-1 site: reconstitution in vitro. Genes Dev. 2, 1687–1699 (1988).
Shetty, A. et al. A systematic comparison of FOSL1, FOSL2 and BATF-mediated transcriptional regulation during early human Th17 differentiation. Nucleic Acids Res 50, 4938–4958 (2022).
Vandermeulen, C. et al. The HTLV-1 viral oncoproteins Tax and HBZ reprogram the cellular mRNA splicing landscape. PLoS Pathog. 17, e1009919 (2021).
Itoyama, Y. et al. Spontaneous proliferation of peripheral blood lymphocytes increased in patients with HTLV-I-associated myelopathy. Neurology 38, 1302–1307 (1988).
Ijichi, S. et al. In vitro modulation of lymphocyte proliferation by prednisolone and interferon-alpha in patients with HTLV-I-associated myelopathy (HAM). J. Neuroimmunol. 23, 175–178 (1989).
Tanaka, Y. et al. Potential role of HTLV-1 Tax-specific cytotoxic t lymphocytes expressing a unique t-cell receptor to promote inflammation of the central nervous system in myelopathy associated with HTLV-1. Front Immunol. 13, 993025 (2022).
Yamauchi, J. et al. Mogamulizumab, an anti-CCR4 antibody, targets human T-lymphotropic virus type 1-infected CD8+ and CD4+ T cells to treat associated myelopathy. J. Infect. Dis. 211, 238–248 (2015).
Araya, N. et al. Human T-lymphotropic virus type 1 (HTLV-1) and regulatory T cells in HTLV-1-associated neuroinflammatory disease. Viruses 3, 1532–1548 (2011).
Kwok, R. P. et al. Control of cAMP-regulated enhancers by the viral transactivator Tax through CREB and the co-activator CBP. Nature 380, 642–646 (1996).
Wu, K. et al. Protein profile of tax-associated complexes. J. Biol. Chem. 279, 495–508 (2004).
Renoux, F. et al. The AP1 transcription factor Fosl2 promotes systemic autoimmunity and inflammation by repressing treg development. Cell Rep. 31, 107826 (2020).
Wolf, B. K. et al. Cooperation of chromatin remodeling SWI/SNF complex and pioneer factor AP-1 shapes 3D enhancer landscapes. Nat. Struct. Mol. Biol. 30, 10–21 (2023).
Barral, A. & Zaret, K. S. Pioneer factors: roles and their regulation in development. Trends Genet 40, 134–148 (2024).
Arainga, M., Murakami, H. & Aida, Y. Visualizing spatiotemporal dynamics of apoptosis after G1 arrest by human T cell leukemia virus type 1 Tax and insights into gene expression changes using microarray-based gene expression analysis. BMC Genomics 13, 275 (2012).
DeSilva, D. R. et al. Inhibition of mitogen-activated protein kinase kinase blocks T cell proliferation but does not induce or prevent anergy. J. Immunol. 160, 4175–4181 (1998).
D’Souza, W. N., Chang, C. F., Fischer, A. M., Li, M. & Hedrick, S. M. The Erk2 MAPK regulates CD8 T cell proliferation and survival. J. Immunol. 181, 7617–7629 (2008).
Osame, M. Review of WHO Kagoshima meeting and diagnostic guidelines for HAM/TSP. Human retrovirology, (ed. Blattner W.A.) 191–197 (New York, NY: Raven Press; 1990).
Acknowledgements
We thank H. Miyoshi and A. Miyawaki for providing venus-encoding lentiviral vectors. The authors acknowledge the excellent technical assistance provided by Katsunori Takahashi, Yasuo Kunitomo, Yumiko Hasegawa, Mikako Koike, Yumi Saito, and Miho Ishikawa. The Platform for Advanced Genome Science project provided sequencing technologies. The super-computing resource was provided by Human Genome Center, the Institute of Medical Science, the University of Tokyo (http://sc.hgc.jp/shirokane.html). This study was supported by AMED (grant numbers: JP25ek0109827, JP25fk0108733, JP24ek0109735, JP23fk0108672, JP23wm0325056, JP24ama221534, and JP23ck0106860), JSPS KAKENHI (grant numbers: JP25K02584, JP24K10148, JP22H02987, JP23K24248, JP24K02317, JP22K07507, JP22H04925 [PAGS], and JP22H04923 [CoBiA]), and a grant from the Japan Research Foundation for Clinical Pharmacology and International Joint Usage/Research Center, Institute of Medical Science, University of Tokyo.
Author information
Authors and Affiliations
Contributions
M.N. performed cell culture assays, RNA-seq analysis, epigenomic profiling, ChIP-seq analyses, single-cell gene expression and chromatin accessibility profiling, bioinformatics analyses, enhancer assays, and experimental and clinical data analyses, and wrote the paper. K.N. performed cell culture assays, single-cell gene expression and chromatin accessibility profiling, bioinformatics analyses, and experimental and clinical data analyses. N.T. curated the clinical data, provided advice, and co-wrote the paper. A.N. performed cell culture assays and experimental and clinical data analyses. K.Y. curated the single-cell data and performed bioinformatics analyses. J.M. prepared chromatin accessibility profiles. S.T. and S.A. prepared the plasmids for enhancer assays and provided advice. N.Y., E.H., T.W., T.S., and Y.N. curated clinical data and provided advice. Y.S. performed bioinformatics analyses and provided advice. K.U., M.Y., and Y.Y. conceived and supervised the project and co-wrote the paper. All the authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Charles Bangham and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nakashima, M., Nagai, K., Takao, N. et al. Chromatin remodeling enhances MAP3K8 expression in HAM: a key pathogenesis for therapeutic intervention. Nat Commun 16, 9874 (2025). https://doi.org/10.1038/s41467-025-64836-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64836-7