Abstract
The formation of foamy macrophages is often considered a pathological hallmark of atherosclerosis, but the underlying mechanisms remain elusive. Herein, we demonstrated that the transcription factor TFEC could upregulate the expression of Glycoprotein non-metastatic melanoma protein B (GPNMB) in atherosclerotic plaque foamy macrophages. Circulating levels of soluble GPNMB correlated positively with atherosclerotic severity. Moreover, mice with systemic Gpnmb-mutation or myeloid-specific Gpnmb knockout exhibited a reduced atherosclerotic burden. Live-cell imaging revealed that GPNMB-positive vesicles were involved in lipoprotein internalization and transport within macrophages and facilitated lipid droplet formation. In Gpnmb-mutant macrophages, impaired lipid droplet formation from internalized lipoproteins, combined with enhanced lipid β-oxidation and lysosomal lipolysis, led to reduced macrophage foaming. Notably, mice treated with siRNA-loaded lipid nanoparticles targeting Gpnmb in lesional foamy macrophages showed alleviation of atherosclerotic burden. Overall, our findings elucidate the intracellular lipoprotein processing in macrophages and suggest GPNMB as a potential therapeutic target for atherosclerosis.
Similar content being viewed by others
Introduction
Atherosclerosis, a leading global cause of death1, remains a significant area of research focusing on its pathogenesis, prevention, and treatment. Foamy macrophage formation is often considered a pathological hallmark of atherosclerosis2. How to reverse this foaming process has yet to be explored, which is one of the keys to the regression and resolution of atherosclerosis3. Considering vascular macrophages physiologically uptake and process atherogenic low-density lipoproteins (LDLs) and their modified forms (e.g., oxidized low-density lipoproteins, OxLDL), lipid storage in macrophages can also be viewed as a protective mechanism against a lipid-rich environment and associated elevated lipid uptake4. Preserving the lipoprotein-processing function of vascular macrophages has become a pressing issue.
Macrophages within the vascular wall directly interact with and engulf subendothelial LDL and its modified forms (e.g., OxLDL). After endocytosis mediated by caveolae and clathrin and subsequent fusion with endosomes, internalized lipoproteins are subsequently degraded within lysosomes and undergo various metabolic fates5. Primarily, the released free cholesterol is esterified by acyl-CoA cholesterol acyltransferase 1 (ACAT1) and stored in lipid droplets or effluxed via ATP-binding cassette subfamily members-mediated transport6. Understanding how macrophages process lipoprotein particles intracellularly, particularly transport and storage, is a significant area of research. Only cells expressing scavenger receptors, such as atherosclerotic plaque macrophages, can take up excess OxLDL in an atherogenic lipid-rich environment7,8. Therefore, storage of cholesterol esters in lipid droplets (LDs) undoubtedly prevents intracellular free cholesterol accumulation and associated lipotoxicity. However, unrestricted lipoprotein influx and the subsequent breakdown of the intracellular lipid degradation system can lead to the accumulation of lipid droplets7, resulting in macrophages appearing as foam cells and exacerbating atherosclerotic plaque development. Further research is warranted to elucidate how macrophages process lipoproteins while resisting lipotoxicity, which may facilitate the regulation of lipid droplet accumulation and potentially lead to the regression of foam cells.
Glycoprotein non-metastatic melanoma protein B (GPNMB) has emerged as a prominent signature of plaque foam cells in recent years9,10,11,12,13,14. While GPNMB has been reported to be located at the endosomal system in mature macrophages15,16,17 and identified as a membrane protein of lipid droplets in foamy THP-1-derived macrophages18, its role in the intracellular processing of lipoproteins and macrophage foaming remains unclear. The role of GPNMB in atherosclerosis progression is controversial19,20,21, potentially due to its varying functions in different cell types22. Clarifying the role of highly expressed GPNMB in foamy macrophages may help resolve these contradictions. Although targeted elimination of GPNMB-positive endothelial cells has been shown to reduce atherosclerotic plaque burden and ameliorate metabolic dysfunction19, the high expression of GPNMB in foamy macrophages also suggests it as a potential therapeutic target for atherosclerosis through regulating foam cells. Additionally, soluble GPNMB (sGPNMB) has been reported as a potential plasma or serum biomarker for several metabolic diseases, including non-alcoholic steatohepatitis (NASH)23, obesity24, Gaucher disease25,26,27, and Niemann–Pick disease type C28,29. Comprehensive research is needed to evaluate the potential of GPNMB as a therapeutic target and sGPNMB as a biomarker for atherosclerosis.
Herein, we observed that GPNMB expression was significantly upregulated in macrophages in response to OxLDL stimulation. GPNMB mediated lipoprotein internalization, transport, and lipid droplet formation to resist intracellular lipotoxicity, thereby promoting macrophage foaming in an atherosclerotic environment. Both systemic mutation and myeloid-specific knockout of Gpnmb markedly reduced OxLDL-induced foamy macrophage formation. The underlying mechanism involves inhibition of intracellular lipid droplet production and a significant increase in lipid metabolism via lysosomes and lipid oxidation, which leads to reduced atherosclerotic plaque burden in Gpnmb-deficient mice. Furthermore, delivering small interfering RNAs (siRNAs) targeting Gpnmb to macrophages and foam cells within plaques using lipid nanoparticles (LNPs) offers a promising therapeutic approach for atherosclerosis. Additionally, the plasma concentration of sGPNMB was identified as an independent risk factor for the severity of atherosclerosis.
Results
High expression of GPNMB correlated with atherosclerosis progression
Consistent with the omics data above, we verified that Gpnmb expression was significantly higher in human atheroma plaques than in adjacent carotid tissue based on the Gene Expression Omnibus database (GEO database, GDS5083)30 (Fig. 1A). To investigate the role of GPNMB in atherosclerosis progression and considering the potential of sGPNMB as a disease severity indicator, we collected plasma samples from patients with stable and unstable coronary atherosclerosis and controls. As expected, plasma sGPNMB levels were significantly elevated in patients with coronary atherosclerosis compared to control participants (Fig. 1B). Although there was no difference in plasma sGPNMB between patients with stable and unstable coronary atherosclerosis (Fig. S3a), plasma sGPNMB levels correlated moderately positively with the severity of atherosclerosis as assessed by Gensini scores, SCORE risk score, and ASCVD risk score (Fig. 1C, D, and S3b, c). Additionally, plasma sGPNMB levels were higher in patients with a history of coronary artery disease (CAD) or hypertension and those taking lipid-lowering or antihypertensive drugs (Fig. 1E). Furthermore, we analyzed the correlations between plasma sGPNMB and other atherosclerosis-related variables. Interestingly, plasma sGPNMB correlated positively with total cholesterol (TCHO), HDL-cholesterol (HDL-C), and Hemoglobin A1C (HbA1c) but negatively with Hemoglobin when single-factor correlation analysis was performed (Figs. 1F and S3d–f). The multivariable linear regression model further indicated an independent correlation between plasma sGPNMB and total cholesterol (β = 0.151, P = 0.021, Fig. 1G and Supplementary Table 2). When multivariable linear regression analysis was performed to predict carotid plaque severity (Gensini score) at baseline, sGPNMB was identified as an independent risk factor, improving the accuracy of predicting atherosclerosis severity (β = 0.218, P = 0.031, Fig. 1H and Supplementary Tables 3). Considering the limited correlation between circulating sGPNMB levels and AS severity, it is necessary to expand the clinical samples and incorporate additional biomarker parameters (e.g., inflammatory cytokines, advanced lipid profiling) for further analysis. We prefer the integration of plasma sGPNMB within a multi-modal biomarker panel for atherosclerosis risk stratification.
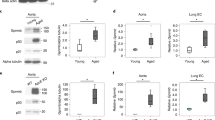
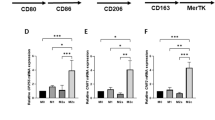
A Relative mRNA expression levels of GPNMB in human carotid artery atheromatous plaques and control intact tissue from hypertensive patients (mean ± SEM, n = 32, based on dataset GDS5083). B Plasma sGPNMB levels in patients with coronary atherosclerosis and the controls (Control, n = 31; Patients, n = 89; mean ± SEM). C The linear relationship of sGPNMB with the Gensini score (left, n = 79) and plasma sGPNMB levels of CAD patients with varying severity based on the Gensini score (right, High/Intermediate/Low). D The linear relationship of sGPNMB with the SCORE risk score (left, n = 154) and plasma sGPNMB levels of CAD patients with varying severity based on the SCORE risk score (right, High/Intermediate/Low). E Panels illustrated the comparisons of sGPNMB across categories of coronary artery disease (upper left; NC, n = 61; CAD, n = 25), hypertension (upper right; NC, n = 67; Hypertension, n = 47), lipid-lowering therapy (lower left; Without, n = 70; With, n = 41), and antihypertensive therapy (lower right; Without, n = 93; With, n = 19). F The linear relationship of sGPNMB with plasma total cholesterol (n = 148). G Forest plot of the correlation between sGPNMB and other AS-associated independent variables at baseline (HR (95% CI), n = 182, derived from the multivariable linear regression model reported in Supplemental Table 2). H Forest plot of predicted multivariable linear regression model for carotid plaque severity (Gensini score) at baseline (HR (95% CI), n = 182, derived from the multivariable linear regression models reported in Supplemental Table 3). I Western blot showing the expression of GPNMB in the aortas of ApoE−/− mice fed with a CD or WD for different weeks (left) and the quantitative analysis results (right) (mean ± SEM, n = 3). P values were calculated by unpaired Student t-test (A, B, E, and I) or one-way ANOVA (C, D). Spearman rank correlation test, P < 0.05 was regarded as the statistical criterion to set thresholds (C, D, and F).
Next, we simulated atherosclerosis of varying severity in ApoE−/− mice fed a Western Diet (WD). As the feeding time increased, the percentage of BODIPYhi foam cells in aortic macrophages, as determined by flow cytometry analysis, increased gradually (Fig. S3g). Additionally, GPNMB expression in the aortas and aortic macrophages of ApoE−/− mice fed a WD was significantly higher compared to control mice (ApoE−/− mice fed a chow diet) (Fig. 1I).
These observations aligned with previous omics data regarding GPNMB abundance in human and mouse atherosclerosis plaques, suggesting a progressive increase in GPNMB expression during atherosclerosis progression. Moreover, plasma sGPNMB levels emerged as a potential independent predictor of atherosclerosis.
GPNMB localized within macrophages in atherosclerotic plaques
Single-cell RNA-seq analysis of human and mouse atherosclerotic lesions revealed that GPNMB was primarily expressed in macrophage-derived foam cells. We subsequently analyzed the immune cell composition of human atherosclerotic plaques from the Biobank of Karolinska Endarterectomies (BiKE, GSE21545) submitted by Orecchioni et al.31. GPNMB abundance was even higher than TREM2 and other markers in Trem2hi macrophages, the widely recognized foam macrophage population in both human and mouse plaques (Fig. 2A). Plaque compositions were extracted using Cibersort for representative GPNMBhigh (left) and GPNMBlow (right) endarterectomy plaques, and the percentages of macrophages were higher in GPNMBhigh plaques (Figs. 2B and S4e). Additionally, markedly higher GPNMB expression was observed in plaques with increased macrophage infiltration (Fig. 2C). Further analysis showed that GPNMB expression correlated positively with M0 macrophage infiltration but negatively with M2 macrophage infiltration in plaques (Fig. 2D).
A mRNA levels of GPNMB and TREM2hi macrophage marker genes measured by Affymetrix gene array (BiKE database, GSE21545) expressed as RMA (robust multi-array average; log scale, mean ± SEM, n = 126). B Plaque composition extracted with Cibersort for representative GPNMBhigh (left) and GPNMBlow (right) endarterectomy plaques. C GPNMB expression as a function of macrophage content ratio (low, <0.5, and high, >0.7; 61 subjects total) as determined by Cibersort. D The relationships between GPNMB expression and the content ratio of total macrophages (left), M0 macrophages (middle), and M2 macrophages (right). E Representative immunohistochemistry staining of α-SMA+ smooth muscle cells, CD31+ endothelial cells, CD68+ macrophages, and GPNMB+ cells in human carotid artery atheromatous plaques. N = 3 independent experiments. Scale bar, 200 μm. F Representative images from immunofluorescence of GPNMB+ cells (Red) with CD68+ macrophages (Green, upper) and BODIPYhi foam cells (Green, lower) in human plaques. N = 3 independent experiments. G–H Representative images from immunofluorescence of GPNMB+ cells (Red) with F4/80+ macrophages (Green) and BODIPYhi foam cells (Green) in atherosclerotic plaque from ApoE−/− mice and the quantification of the degree of colocalization (H mean ± SEM, n = 3). I–J Representative western blot images showing the expression of GPNMB in BMDMs from ApoE−/− mice treated with OxLDL (50 μg/mL) for different hours (I) and corresponding quantitative analysis (J, mean ± SEM, n = 3). The statistically significant P values for comparisons against the 0 h group in Fig. 2J were 0.0485, 0.0162, 0.0368, 0.0045, 0.0118, 0.0046, 0.0214, 0.0003, and 0.0113 (from left to right). K Representative western blot images showing the expression of GPNMB in THP-1 derived macrophages treated with OxLDL (50 μg/mL) for 48 h and corresponding quantitative analysis (mean ± SEM, n = 3). P values were calculated by unpaired Student t-test (C and K) or one-way ANOVA (J). Spearman rank correlation test, P < 0.05 was regarded as the statistical criterion to set thresholds (D).
In parallel with these dataset findings, we observed that GPNMB was specifically expressed on macrophages (Mφ, marked with CD68), distinct from endothelial cells (EC, marked with CD31) and smooth muscle cells (SMC, marked with α-SMA) in human carotid plaque tissue (Fig. 2E). Further immunofluorescence analysis confirmed the co-localization of GPNMB+ cells with CD68-labeled macrophages and BODIPY-labeled foam cells in human carotid plaque tissue (Fig. 2F). Similarly, GPNMB+ cells co-localized with F4/80-labeled macrophages and BODIPY-labeled foam cells in aortic root plaque from ApoE−/− mice (Fig. 2G–H).
Based on the aforementioned results, we hypothesized that GPNMB plays a crucial role in foam cell formation. To investigate this, macrophage foaming was induced in bone marrow-derived macrophages (BMDMs) from C57BL/6 J mice (WT BMDMs) or ApoE−/− mice by OxLDL treatment. GPNMB expression was increased after prolonged treatment with OxLDL (50 μg/mL) as well as increased with increasing OxLDL concentrations (Figs. 2I–J and S3h–i). Consequently, the macrophage foaming model was established by treating BMDMs with 50 μg/mL OxLDL for 48 h for further mechanistic studies. Similarly, treatment with 50 μg/mL OxLDL for 48 h induced GPNMB expression in THP-1 cells (Fig. 2K).
These findings indicated that GPNMB was specifically localized to macrophages and foam cells in human and mouse atherosclerotic plaques, and GPNMB expression was significantly elevated with macrophage foaming.
TFEC induced Gpnmb expression in macrophages during atherosclerosis progression
The factors upregulating Gpnmb expression during atherosclerosis progression and macrophage foaming remain to be investigated. Previous studies have suggested that MITF is required for Gpnmb induction in adipose tissue macrophages (ATMs)32 and that Gpnmb is a direct transcriptional target of TFE3 in Renal Cell Carcinoma33. Given that MITF, TFE3, and TFEC are members of the MiT/TFE family of transcription factors, and TFEC is macrophage-restricted34, we explored whether this family controlled Gpnmb induction during macrophage foaming in atherosclerosis. We first compared the mRNA expression of all four MiT/TFE family members in human atheroma plaques and adjacent carotid tissue using the GDS5083 database30 and analyzed their correlation with GPNMB expression. MITF, TFEC, and TFEB mRNA expression were significantly higher in plaques than in control tissues, while TFE3 expression showed little difference (Figs. 3A and S4a, b). Notably, MITF or TFEC mRNA expression exhibited strong positive correlations with GPNMB expression (MITF: R = 0.8485, P < 0.0001; TFEC: R = 0.8306, P < 0.0001; Fig. 3B), whereas TFEB or TFE3 mRNA expression showed weak correlations (Fig. S4c, d). Similarly, a strong correlation between GPNMB expression and MITF or TFEC gene expression and a weak correlation between GPNMB expression and TFEB or TFE3 gene expression were observed in plaque macrophages (based on BiKE, GSE21545, Figs. 3C and S4e–h). Given that MITF is reported to induce Gpnmb expression in ATMs directly, we focused on the role of TFEC.
A mRNA levels of MITF (left) and TFEC (right) in human carotid artery atheromatous plaques and control intact tissue from hypertensive patients (mean ± SEM, n = 32, based on dataset GDS5083). B The relationships between Gpnmb expression with Mitf expression (left) or Tfec expression (right) in human carotid artery atheromatous plaques (based on dataset GDS5083). C The relationships between Gpnmb expression with Mitf expression (left) or Tfec expression (right) in macrophages from human carotid plaque (based on dataset GSE21545, n = 126). Representative western blot images showing increased expression of GPNMB and TFEC in the aortas of ApoE−/− mice fed with a WD (D) and WT BMDMs treated with OxLDL (E) (mean ± SEM, n = 3). F Representative images from immunofluorescence of TFEC (magenta) with Hoechst-labeled cell nucleus (cyan) in WT BMDMs treated with or without OxLDL and corresponding models built by Imaris (left). The quantitative analysis of total fluorescence intensity (right) and the ratio of nucleus fluorescence intensity to cytoplasm fluorescence intensity (middle) of TFEC, N = 3 independent experiments, mean ± SEM, n = 26 in NC group, n = 34 in OxLDL group. G, H 293T cells were transfected with pEGFP-TFEC plasmid, pEGFP-MITF plasmid, pEGFP-TFEB plasmid, or pEGFP-TFE3 plasmid, or co-transfected with pEGFP-TFEC plasmid and one plasmid of the other three plasmids. Representative western blot images showing the expression of GPNMB and the overexpression efficacy of these plasmids with an antibody against GFP (G) and the quantitative analysis (H, mean ± SEM, n = 3). I Activation of Gpnmb promoter in HEK293T cells transiently expressing pEGFP-TFEC plasmid, pEGFP-MITF plasmid, pEGFP-TFEB plasmid, pEGFP-TFE3 plasmid, or negative control plasmid was measured using a dual luciferase reporter assay (mean ± SEM, n = 3). J Representative images from immunofluorescence of TFEC (magenta) with Hoechst-labeled cell nucleus (cyan) in WT BMDMs transfected with siRNA against Tfec (siTfec) or nontarget control siRNA (siNC) and the quantitative analysis of total fluorescence intensity of TFEC, N = 3 independent experiments (mean ± SEM, n = 23 in siNC group, n = 20 in siTfec group). K Quantitative PCR analysis of Tfec and Gpnmb expression in WT BMDMs transfected with siTfec or siNC (mean ± SEM, n = 4). L Representative western blot images showing the expression of GPNMB and TFEC in WT BMDMs transfected with siTfec or siNC (left) and the quantitative analysis (right) (mean ± SEM, n = 3). P values were calculated by unpaired Student t-test (A, D, E, F, and J–L) or one-way ANOVA (H–I). Differences are significant for *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Spearman rank correlation test, P < 0.05 was regarded as the statistical criterion to set thresholds (B, C).
In atherosclerotic aortas from ApoE−/− mice, TFEC protein levels were dramatically increased (Figs. 3D and 4i). Treatment with OxLDL also upregulated TFEC protein expression in WT BMDMs, consistent with fluorescence staining results (Fig. 3E, F). Additionally, clear translocation of TFEC to the nucleus was observed in BMDMs treated with OxLDL (Fig. 3F). Furthermore, the promoter of human GPNMB, a 2-kb region upstream of the transcription start site (TSS), was inserted into a luciferase reporter plasmid. When reporter plasmids or control plasmids and overexpression plasmids of MiT/TFE family transcription factors were co-transfected in HEK-293T cells, remarkable upregulation of Gpnmb promoter activity and GPNMB expression upon the overexpression of TFEC, MITF, TFEB, or TFE3 was observed (Fig. 3G–I). Additionally, TFEC/MITF, TFEC/TFEB, and TFEC/TFE3 co-overexpression also upregulated GPNMB expression and enhanced Gpnmb promoter activity (Figs. 3G, H and S4j). These findings indicated that GPNMB was a direct transcriptional target of MiT/TFE family transcription factors. To confirm the regulatory role of TFEC in controlling Gpnmb expression in macrophages, we performed siRNA-mediated knockdown targeting TFEC in BMDMs. Knockdown efficiency was determined by immunofluorescence (Fig. 3J) and quantitative PCR analysis (Fig. 3K). As expected, TFEC knockdown suppressed GPNMB expression at both the mRNA and protein levels (Fig. 3K, L).
In summary, these experimental data demonstrated that upregulated TFEC expression significantly induced GPNMB expression in macrophages during atherosclerosis progression. Given the non-negligible role of MITF, future studies should comprehensively investigate how the MiT/TFE family of transcription factors co-regulates Gpnmb expression.
Gpnmb-mutation or myeloid-specific Gpnmb knockout alleviated atherogenesis in ApoE −/− mice
Next, we investigated the impact of the Gpnmb-mutation on atherogenesis. The parental DBA/2J strain of mice carries a nonsense variant in Gpnmb (GpnmbR150X)35. C57BL/6J × DBA/2J (BXD) mice and GpnmbR150XApoE−/− mice were generated. To minimize background effects, we backcrossed the generated mice with two parental strains, creating BXD-D and BXD-C backgrounds, respectively. The Gpnmb variant was confirmed (Fig. S1g–i). We next examined the physiological and biochemical characteristics of GpnmbR150X mice and GpnmbR150XApoE−/− mice on the BXD-D background fed with a control diet (CD) or a WD, as well as wild-type (WT), heterozygous (HET), GpnmbR150X, WT ApoE−/−, HET ApoE−/−, and GpnmbR150XApoE−/− mice on the BXD-C background fed with a WD (concluded in Supplementary Tables 4–7).
The aortas of WT, HET, and GpnmbR150X mice on the BXD-C crossed background appeared to be in good condition (Fig. S5b). En-face Oil-Red-O (ORO) staining of aortas and photographs of aortas and aortic arches revealed a smaller plaque burden in GpnmbR150XApoE−/− mice compared to WT ApoE−/− mice on the BXD-C crossed background fed with a Western diet (Figs. 4A and S5f). Similar results were obtained with female mice (Fig. S5g, h). Flow cytometry analysis further verified fewer CD45+ CD11b+ macrophages, fewer CD36hi BODIPYhi foam cells, and lower median fluorescence intensity (MFI) of aortic macrophages in atherosclerotic aortas from male GpnmbR150XApoE−/− mice (Figs. 4B and S5i). Similar results were observed with female mice (Fig. S5j). Consistently, hematoxylin and eosin (H&E) staining and Modified Russell-Movat (Movat) staining of aortic root sections revealed lower atherosclerosis burden in GpnmbR150XApoE−/− mice with smaller total plaque size, smaller necrotic cores, more collagen, and thicker fibrous caps, especially in the advanced atherosclerosis stage (Figs. 4C−E and S5k). Similar results were obtained with ORO staining (Fig. S5l). A tendency towards lower percentages of F4/80-labeled macrophages and BODIPY-labeled foam cells and thicker α-SMA-labeled fibrous caps was observed in cross sections of the aortic root from GpnmbR150XApoE−/− mice compared to WT ApoE−/− mice (Fig. S5m, n).
WT ApoE−/−, HET ApoE−/−, and GpnmbR150X ApoE−/− (all on hybrid DBA/2-C57BL/6 J background) mice were fed a WD to induce lesion development. A Representative photomicrographs and the quantification of en face lesion area in the whole aortas of WT ApoE−/−, HET ApoE−/−, and GpnmbR150X ApoE−/− mice (male) fed a WD for 20, 24, and 28 weeks (mean ± SEM, n = 3). B Representative flow cytometric expression of CD45 and CD11b in atherosclerotic aortas (upper dot plots) and CD36 and BODIPY levels in aortic macrophages (lower dot plots and histogram) (male, n = 3/group). The percentages of flow cytometric BODIPYhi CD36hi foam cells in aortic macrophages and the flow cytometry median fluorescence intensity (MFI) of aortic macrophages (mean ± SEM, male, N = 5 independent experiments, n = 3/group for each experiment). C Representative histological analysis of cross sections of the male aortic sinus stained with hematoxylin and eosin (H&E), and dashed lines delimited the plaque area. The quantification of plaque area percentages (mean ± SEM, male, n = 17 in WT ApoE−/− mice group, n = 13 in GpnmbR150X ApoE−/− mice group). Representative histological analysis of cross sections of the aortic sinus stained with Movat pentachrome (E) and the quantification of the percentage of total plaque, collagen, necrotic core, and fiber cap area (D) (mean ± SEM, male, n = 5 in WT ApoE−/− mice group, n = 4 in GpnmbR150X ApoE−/− mice group). The statistically significant P values for comparisons in D were 0.0236, 0.0003, 0.0029, and 0.0002 (from left to right). F Female WT, HET, and GpnmbR150X (all on hybrid DBA/2-C57BL/6J background) mice were fed a WD for 20 weeks. The quantification of increased total cholesterol (TCHO, left) and HDL-C (right) was shown in the graph (mean ± SEM, n = 4 in WT mice group, n = 6 in HET and GpnmbR150X mice group). G WT ApoE−/− and GpnmbR150X ApoE−/− (all on hybrid DBA/2-C57BL/6J background) mice were fed a WD for 20 weeks. The quantification of total cholesterol (TCHO, male, left) and HDL-C (male, right) was shown in the graph (mean ± SEM, n = 8−18/group). Representative photomicrographs (H) and the quantification (I) of en face lesion area in the whole aortas of Gpnmbfl/fl ApoE−/− and Gpnmbfl/fl Lyz2cre/cre ApoE−/− mice (male) fed a WD for 12, 16, and 20 weeks (mean ± SEM, n = 3). J Representative images of cross sections of the aortic sinus from immunofluorescence of F4/80+ macrophages (Green) and Nile Red+ foam cells (Red) in atherosclerotic plaque from Gpnmbfl/fl ApoE−/− and Gpnmbfl/fl Lyz2cre/cre ApoE−/− mice (right, female) and the quantification of plaque area percentages (left, mean ± SEM, female, n = 5) and intensity of Nile Red-labeled plaque (middle, mean ± SEM, female, n = 5). K, Male Gpnmbfl/fl ApoE−/− and Gpnmbfl/fl Lyz2cre/cre ApoE−/− mice were fed a WD for 16 weeks. The quantification of increased total cholesterol (TCHO, upper) and LDL-C (lower) was shown in the graph (mean ± SEM, n = 6). P values were calculated by unpaired Student t-test (A, C, D, G, and I−K) or one-way ANOVA (B, F). Differences are significant for *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Obesity appeared to be more easily induced by a Western diet in Gpnmb-mutant mice on a BXD-C crossed background, which was accompanied by a higher level of serum triglyceride in GpnmbR150XApoE−/− mice (Fig. S5a, c−e and Supplementary Tables 5). Conversely, female GpnmbR150X mice and male GpnmbR150XApoE−/− mice showed lower serum total cholesterol and HDL-C levels after feeding with a WD than respective control mice (Fig. 4F, G). Interestingly, this finding corresponded to the positive correlation between the serum levels of sGPNMB and HDL-C in patients with atherosclerosis (Fig. S3d).
To avoid genetic background interference, we also examined the characteristics of atherogenesis in GpnmbR150XApoE−/− mice on a BXD-D crossed background. WD feeding resulted in increased body weight, fat pad weight, serum total triglyceride levels, and aortic macrophage percentages in GpnmbR150X mice on BXD-D (Fig. S6a−g). Surprisingly, we found no significant differences in atherosclerotic plaques between GpnmbR150XApoE−/− mice on a BXD-D crossed background fed with a CD or a WD within 20 weeks (Fig. S6h−l). These findings suggest that the Gpnmb-mutation is a likely reason for the low susceptibility to developing atherosclerotic aortic lesions in DBA/2J strain mice. A similar atherogenesis difference was observed in GpnmbR150XApoE−/− mice fed with a WD on either a BXD-C-crossed background or a BXD-D-crossed background.
To further investigate whether the atheroprotective phenotypes in Gpnmb-mutation mice were attributed to the role of GPNMB in macrophages, we constructed a myeloid-specific Gpnmb knockout mouse model (Gpnmbfl/flLyz2cre/cre, GpnmbΔMAC). Gpnmb knockout efficiency was determined by both Western blot and flow cytometry (Fig. S1j−m). As expected, both en-face ORO staining of aortas and aortic root sections indicated a smaller plaque burden in GpnmbΔMAC ApoE−/− mice compared to Gpnmbfl/flApoE−/− mice fed with a Western diet (Figs. 4H−I and S7a, b, e, f). The decreased plaque area and lipid contents were also confirmed by H&E staining analysis (Fig. S7c, d) and immunofluorescence assays (Fig. 4J). Female GpnmbΔMAC ApoE−/− mice exhibited lower serum total cholesterol and LDL-C levels after feeding with a WD than Gpnmbfl/flApoE−/− mice (Figs. 4Kand S7g).
Collectively, the genetic mutation and myeloid-specific knockout of Gpnmb mitigated atherogenesis in ApoE−/− mice, accompanied by decreased serum total cholesterol levels. Given the reduced atherosclerosis in myeloid-specific Gpnmb knockout mice, inhibiting the function of GPNMB in macrophages represents a potential atheroprotective strategy.
GPNMB mediated the internalization, trafficking, and degradation of lipoprotein in macrophages
Considering the high expression and specific location of GPNMB in atherosclerotic plaques, we hypothesized that GPNMB may exert pro-atherosclerotic effects by promoting macrophage foaming. Strikingly, GPNMB was identified as one of the membrane proteins of LDs in THP-1-derived macrophage foam cells18. Indeed, we observed the colocalization of GPNMB with BODIPY-labeled LDs in BMDMs treated with OxLDL, as well as the colocalization of GPNMB with Dil-OxLDL (Fig. 5A). Live cell imaging of BMDMs transfected with a pEGFP-GPNMB expressing plasmid and then treated with Dil-OxLDL was monitored for 48 h. As shown in Fig. 5B, Dil-OxLDL surrounded BMDMs at 2 h, and part of the added Dil-OxLDL was taken up by BMDMs at 24 h, with high colocalization with pEGFP-GPNMB. Imaris was used to model the shapes and locations of GPNMB (irregular green circles) and Dil-OxLDL (blue balls represented the shortest distance to GPNMB+ circles more than 0; purple balls represented the distance less than 0), respectively. Supplementary Movies 1 and 2 visualized GPNMB+ circles contacting with Dil-OxLDL (upper line) and mediating the uptake of Dil-OxLDL (lower line) in BMDMs.
A Representative images from immunofluorescence of GPNMB expression (Red) and BODIPY levels (Green) in BMDMs treated with OxLDL (50 μg/mL) for 48 h (upper), representative living cell images of Dil-OxLDL in BMDMs transfected with pEGFP-GPNMB plasmid (lower), and the quantification of the degree of colocalization (mean ± SEM, N = 5 independent experiments). B Representative living cell images of BMDMs transfected with pEGFP-GPNMB plasmid after treatment with Dil-OxLDL for 2 h or 24 h (mean ± SEM, N = 5 independent experiments). C Representative electron micrograph images of GPNMB expression (immuno-gold staining, indicated with red triangles) in BMDMs (mean ± SEM, N = 3 independent experiments). Scale bars, 500 nm. D The fluorescence peak diagram of pEGFP and Nile Red in BMDMs transfected with pEGFP-GPNMB plasmid or empty vector (left) and the quantitative analysis of intensity (right, mean ± SEM, N = 4 independent experiments). E, Representative images of Dil-OxLDL (Red) in BMDMs transfected with pEGFP-GPNMB plasmid or empty vector (left, Scale bar, 7 μm) and the quantitative analysis of Dil-OxLDL intensity (right, mean ± SEM, N = 3 independent experiments, n = 49 cells each group). F-G, Representative images from immunofluorescence of GPNMB (Red), EEA1 (Green), RAB7 (Green), COPB (Green), and ERGIC-53 (Green) expression in BMDMs and corresponding models built by Imaris (G) and the shortest distance of GPNMB+ spots to different organelles calculated by Imaris (F, mean ± SEM, N = 5 independent experiments, n = 375/787/464/700 cells each group). H Representative images from immunofluorescence of GPNMB (Green) and RAB5 (Red) expression in THP-1 derived macrophages (upper two lines) and BMDMs (lower two lines) (mean ± SEM, N = 3 independent experiments). I Representative living cell images of NEEP21, FLCN, or MANNⅡ expression in BMDMs transfected with pEGFP-GPNMB plasmid and treated with CF647-stained OxLDL (CF647-OxLDL) for 36 h (mean ± SEM, N = 3 independent experiments). J Pathway enrichment analysis assessing KEGG pathways or GO terms for biological processes based on proteins coprecipitated by anti-GPNMB antibody. Selected pathways are shown, ranked by Counts. Differentially expressed proteins were determined with the Limma program at p-adj (adjusted p value) <0.05 and logFC (log fold change > 1. K Heatmap of enriched proteins from BMDM lysates using an antibody targeting GPNMB. L Representative living cell images of MOESIN or KIF5B expression in BMDMs transfected with pEGFP-GPNMB plasmid and treated with CF647-OxLDL for 36 h (mean ± SEM, N = 3 independent experiments). M Representative images from immunofluorescence of GPNMB (Green) and SEC22B (Red) expression in BMDMs (mean ± SEM, N = 3 independent experiments). N Representative living cell images of AP-2 or RAB11A expression in BMDMs transfected with pEGFP-GPNMB plasmid and treated with CF647-OxLDL for 36 h (mean ± SEM, N = 3 independent experiments). O Representative living cell images of SEC61β expression in BMDMs transfected with pEGFP-GPNMB plasmid and treated with CF647-OxLDL for 36 h (mean ± SEM, N = 3 independent experiments). P values were calculated by unpaired Student t-test (D, E) or one-way ANOVA (F). Differences are significant for *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
To elucidate the role of GPNMB in the intracellular processes of OxLDL, we employed electron microscopy on BMDMs and identified GPNMB expression on the membrane of organelles, including the endoplasmic reticulum (ER), Golgi apparatus, lysosomes, autophagolysosome, and other vesicle-like structures (immuno-gold staining, indicated with red triangles, Figs. 5C and S8k). When exploring the potential connection between GPNMB and LDs, we conducted an overexpression of GPNMB in BMDMs and assessed the efficiency of this overexpression using flow cytometry (Figs. 5D and S8c) and high-content images (Fig. S8d). Notably, we observed that the intracellular lipid content determined by Nile Red staining and Dil-OxLDL content in BMDMs transfected with the pEGFP-GPNMB plasmid were significantly higher than those transfected with the empty control plasmid (Figs. 5D, E and S8b−f).
We further investigated how GPNMB affects the intracellular physiological processes of LDs in macrophages. We first validated the colocalization observed above using immunofluorescence, including early endosomes (labeled by EEA1), late endosomes (labeled by RAB7), Golgi apparatus and vesicles (labeled by COP-B), endoplasmic reticulum (ERGIC-53/KDEL/SEC61β/ER-Tracker Blue-labeled), Lyso-Tracker Red/LAMP1-labeled lysosomes, and Mito-Tracker-labeled mitochondria (Figs. 5G, O and S8g−i, l−n). Two distinct colocalization models were identified through 3D reconstruction and calculation of the shortest distance from GPNMB proteins to labeled organelles using Imaris. These included overlapping colocalization between GPNMB proteins with LDs, lipoproteins, and endosomal-lysosomal system-related vesicles, and partially binding colocalization between GPNMB proteins with the Golgi apparatus and the ER (Fig. 5F, G). The endosomal-lysosomal system is essential for the internalization, trafficking, and degradation of lipoproteins in macrophages. To elucidate the role of GPNMB in these processes, we monitored the dynamic changes of GPNMB localization with organelles in BMDMs treated with CF647-stained OxLDL (CF647-OxLDL) using live cell imaging (Movie S3-10). GPNMB-labeled vesicle-like structures highly co-localized with endosomes and lysosomes, exhibiting three distinct statuses: preliminary contact, partial fusion, and complete merge, which differed from the contacts of GPNMB with the ER (Fig. 5H, I and Movies S,3−6). These results suggest the existence of sole GPNMB+ vesicles that participate in the internalization, trafficking, and degradation of lipoproteins in macrophages.
To better understand the role of sole GPNMB+ vesicles in intracellular lipoprotein processes, we used monoclonal antibodies against mouse GPNMB to coprecipitate potentially interacting proteins in BMDMs. Subsequent immunoprecipitation-mass spectrometry (IP-MS) analysis revealed that the enriched potential GPNMB-interaction proteins were closely related to the vesicle-mediated transport pathway, consistent with the optical imaging results described above (Fig. 5J). The enriched proteins related to COPI-dependent Golgi-to-ER retrograde traffic were depicted in Fig. S8j. Representative enriched differential proteins, as shown in Fig. 5K, included the endoplasmic reticulum protein SEC61 Translocon Subunit Beta (SEC61B). Since GPNMB was found to colocalize with EEA1 and RAB5, markers of early endosomes, suggesting its role in the internalization of OxLDL, we then observed the colocalization of GPNMB and other marker protein-labeled vesicles. We selected AP-2 (σ2, representing classic clathrin-coated vesicles), Ras-related protein Rab-11A (RAB11A, representing recycling endosomes), and three enriched proteins associated with vesicle-mediated transport, including Moesin that modulates phagolysosomal biogenesis in macrophages36, Kinesin 5B (KIF5B) that regulates lysosomal distribution37, and SEC22b that is involved in vesicle-trafficking between the Golgi complex and the ER. We observed extensive colocalization of GPNMB with the endosomal system and an identical phenomenon where sole GPNMB+ vesicles interacted with endosomes, lysosomes, and other vesicles carrying OxLDL in BMDMs (indicated with gray arrows, Fig. 5I, L−N and Movie S7−10). Moreover, reconstruction using Imaris simulated this phenomenon occurring with Folliculin-labeled lysosomes (Movie S4). In addition, the interactions between GPNMB+ vesicles carrying OxLDL and the ER system created conditions for LDs budding from the ER (Figs. 5O and S8g).
We propose that sole GPNMB+ vesicles could interact with the endosomal system carrying OxLDL, transferring the latter to GPNMB+ endosomes/lysosomes through membrane fusion to coordinate intracellular lipoprotein processing in macrophages.
Gpnmb-mutation inhibited macrophage foaming
Considering the putative function of GPNMB in lipoprotein processing, we compared the uptake of Dil-OxLDL and lipid storage levels in BMDMs from GpnmbR150X mice (GpnmbR150X BMDMs) and WT mice (WT BMDMs) (both BXD-C crossed background) and rechecked these parameters in BMDMs from GpnmbR150XApoE−/− mice and WT ApoE−/− mice (both BXD-C crossed background) to align with our findings in mouse models of atherosclerosis. As a consequence, Gpnmb-mutation led to decreased BODIPY-labeled lipid contents after treatment with Ox-LDL and decreased uptake of Dil-OxLDL in ApoE−/− BMDMs, as confirmed using confocal and high-content microscopy (Figs. 6A and S9a, b). Flow cytometry further quantified these changes, demonstrating that the MFI of BODIPY-labeled lipid contents and Dil-OxLDL indeed declined in GpnmbR150X BMDMs compared to WT BMDMs, similar to the observations in BMDMs from GpnmbR150XApoE−/− mice (Figs. 6B−D and S9c, d). Neutral lipid content measured by Nile Red staining was consistently reduced in response to Gpnmb-mutation or myeloid-specific Gpnmb knockout (Fig. S9e, k). Dil-OxLDL uptake was also restricted in BMDMs from GpnmbΔMAC mice compared to those from Gpnmbflox/flox mice (Fig. S9l). Ultimately, both Gpnmb-mutation and GPNMB knockdown mitigated but did not eliminate lipid content in BMDMs.
A BMDMs from WT ApoE−/−, HET ApoE−/−, and GpnmbR150X ApoE−/− mice were treated with OxLDL for 24 h, followed by BODIPY staining (left upper) or Dil-OxLDL for 12 h (left lower), and images were taken by immunofluorescence microscopy. N = 3 independent experiments. The quantitative analysis of total fluorescence intensity of BODIPYTM (middle, n = 32/25/24) and Dil-OxLDL (right, n = 42/16/9). B The quantitative analysis of total fluorescence intensity of Dil-OxLDL in four groups of BMDMs from WT and GpnmbR150X mice treated with Dil-OxLDL for 12 h (mean ± SEM, N = 3 independent experiments, n = 3/group for each experiment). Representative FACS plots for the changes of GPNMB expression (C), the histogram of MFI of BODIPY in BMDMs from WT and GpnmbR150X mice treated with or without OxLDL for 48 h (D), and quantitative analysis of total fluorescence intensity of BODIPY in four groups of BMDMs in D (C, mean ± SEM, N = 4 independent experiments). E Representative images of WT BMDMs and GpnmbR150X BMDMs treated with Dil-OxLDL for 12 h (Red), followed by BODIPYTM staining (Green) (left) and the localization of Dil-OxLDL with LDs quantified with Mandar overlap R-value analysis (right) (mean ± SEM, N = 3 independent experiments, n = 13 images each group). F Representative images of expression of SEC61β (Blue) in WT BMDMs and GpnmbR150X BMDMs treated with Dil-OxLDL for 12 h (Red) and stained with followed by BODIPYTM staining (Green), N = 3 independent experiments. G Representative images of expression of MANNⅡ and KDEL (Red) in WT BMDMs and GpnmbR150X BMDMs treated with OxLDL for 24 h, followed by BODIPYTM staining (Green), N = 3 independent experiments. H The localization of BODIPYTM-labeled LDs (upper) or Dil-OxLDL (lower) with ER quantified with PCC (mean ± SEM, n = 8/4/5/4 in the upper figure, n = 5/5/9/10/4/4 in the lower figure). I Representative images of expression of RAB7, RAB11A, or SEC22B (Red) in WT BMDMs and GpnmbR150X BMDMs treated with OxLDL for 24 h, followed by BODIPYTM staining (Green), N = 3 independent experiments. J The localization of BODIPYTM-labeled LDs with Vesicles quantified with PCC (mean ± SEM, N = 3 independent experiments, n = 4/5/10/10/6/6/8/7/7/8). K Representative images of WT BMDMs and GpnmbR150X BMDMs treated with Dil-OxLDL for 12 h (Red) and stained with Mito-Tracker (Magenta), followed by BODIPYTM staining (Green) (left) and the localization of BODIPYTM-labeled LDs (middle, n = 51/44) or Dil-OxLDL (right, mean ± SEM, n = 9/10) with mitochondria quantified with Pearson’s Correlation Coefficient (PCC), N = 3 independent experiments. L Pathway enrichment analysis assessing GO terms for biological processes based on up-regulated proteins in GpnmbR150X BMDMs compared to WT BMDM, identified by mass spectrometry analysis. Selected pathways are shown, ranked by Counts. Differentially expressed proteins were determined with the Limma program at p-adj (adjusted p value) <0.05 and logFC (log fold change) > 1. M Sankey diagram of pathway enrichment analysis assessing GO terms for biological processes based on up-regulated proteins related to vesicle-trafficking in GpnmbR150X BMDMs compared to WT BMDM, identified by mass spectrometry analysis. Selected pathways are shown, ranked by Counts. Differentially expressed proteins were determined with the Limma program at p-adj (adjusted p value) <0.05 and logFC (log fold change) > 1. Data are expressed as mean ± SEM. P values were calculated by unpaired Student t-test (B, C, E, H, J, K) or one-way ANOVA (A). Differences are significant for *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
More interestingly, twelve hours after administration of Dil-OxLDL, GpnmbR150X BMDMs displayed markedly reduced OxLDL-LDs colocalization (Fig. 6E), indicating defective LD biogenesis derived from OxLDL, which was also demonstrated by the decreased colocalization of Dil-OxLDL with the ER (Figs. 6F, H and S9i). Nevertheless, the smaller size of LDs along with the enhanced colocalization of LDs with the ER likely symbolized a higher proportion of unbudded immature LDs in GpnmbR150X BMDMs compared to WT BMDMs (Figs. 6F−H and S9i). Meanwhile, in GpnmbR150X BMDMs, LDs showed significantly more contact with RAB7+ vesicles, RAB11A+ vesicles, and SEC22B+ vesicles, but not with RAB5+ endosomes or AP-2+ vesicles (Fig. 6I, J).
To elucidate the underlying mechanism, we applied bulk RNA-seq and mass spectrometry analysis to identify differentially expressed genes and proteins between GpnmbR150X BMDMs and WT BMDMs (Fig. S10a, b). Based on RNA-seq data, Gpnmb-mutation significantly affected the gene expression pattern of BMDMs when exposed to OxLDL stimulation, particularly for upregulated genes (Fig. S10c). Compared to WT BMDMs, pathway enrichment analysis revealed that many upregulated genes and proteins in GpnmbR150X BMDMs were related to the fatty acid metabolic process (Figs. 6L and S10d). Notably, compared to WT BMDMs, we observed upregulation of Pparγ and Nr1h3 at the mRNA level along with CD36 and LPL at both the mRNA and protein levels in GpnmbR150X BMDMs, suggesting activation of the pathway involved in uptake and processing of OxLDL and cholesterol removal38 (Fig. S9f, g). We validated the increased expression of CD36 caused by the Gpnmb-mutation (Fig. S9h). Based on the upregulated expression of Plin1 and Abhd5 related to lipolysis in mitochondria39 and the upregulated expression of multiple proteins related to fatty acid beta-oxidation in GpnmbR150X BMDMs (Fig. S9f, g), we further observed the colocalization of BODIPY-labeled LDs with Mito-Tracker-labeled mitochondria. The enhanced colocalization of LDs with mitochondria indicated enhanced fatty acid oxidation in GpnmbR150X BMDMs compared to WT BMDMs (Fig. 6K). However, the colocalization of Dil-OxLDL with mitochondria was unexpectedly decreased (Fig. 6K). In addition, the expression heatmap of previously reported signature genes of foam cells12,14,40 was summarized in Fig. S10f. A more comprehensive Sankey diagram summarized upregulated proteins that regulated lipid metabolism in GpnmbR150X BMDMs compared to WT BMDMs (Fig. S10g). Under transmission electron microscopy (TEM), the electron density of mitochondria in GpnmbR150X BMDMs appeared to be lower compared to WT BMDMs, suggesting lower levels of lipid peroxidation (Fig. S9j).
These findings substantiated that Gpnmb-mutation significantly suppressed macrophage foaming by restricting OxLDL-derived lipid droplet biogenesis and accelerating lipid oxidation. While the underlying mechanisms for the varying degrees of lipid droplet colocalization with different vesicles in GpnmbR150X BMDMs remain enigmatic, the vesicle transport-related proteins identified through mass spectrometry analysis offer promising avenues for further exploration (Figs. 6M and S10h).
Gpnmb-mutation facilitated lysosome-mediated lipolysis in macrophages
As depicted in Fig. S9f, g, elevated expression of Plin2 and Plin3 facilitated the formation of lipid droplet-lysosome contacts, leading to enhanced lipophagy39. Concurrently, the increased expression of lysosomal acid lipase/cholesteryl ester hydrolase (LICH, encoded by Lipa) suggested a potential augmentation of lipid hydrolysis within lysosomes of GpnmbR150X BMDMs compared to wild-type BMDMs. Indeed, the colocalization of lysosomes with either LDs or Dil-OxLDL was significantly increased in GpnmbR150X BMDMs (Fig. 7A−C). Given that GPNMB is localized on lysosomes and autophagolysosomes, these findings prompted the use of Lyso-Tracker Red and Acridine Orange (AO) staining to assess lysosomal degradation capacity and membrane integrity. We observed enhanced degradation ability and increased membrane integrity in lysosomes of GpnmbR150X BMDMs (Fig. 7D, E), mirroring similar observations in ApoE−/− BMDMs (Fig. 7H). Furthermore, the use of LysoSensor Yellow/Blue DND-160 (PDMPO) and R2pH-LAMP1-3×GLAG, additional sensitive lysosomal pH probes, demonstrated decreased lysosomal pH in GpnmbR150X BMDMs (Fig. 7F, G). Intriguingly, overexpression of GPNMB in wild-type BMDMs also led to increased lysosomal degradation (Fig. 7I, J), potentially due to higher intracellular lipid stress. This finding suggests that GPNMB itself may play a role in maintaining lysosomal function.
A Representative images of WT BMDMs and GpnmbR150X BMDMs treated with OxLDL for 24 h, followed by the staining of Lyso-Tracker Red (LTR, Red) and BODIPY staining (Green), N = 5 independent experiments, Scale bar, 5 μm. B, Representative living cell images of FLCN expression (Green) in WT BMDMs and GpnmbR150X BMDMs treated with CF647-OxLDL (Magenta) for 36 h, N = 5 independent experiments, Scale bar, 2 μm. C The localization of BODIPY-labeled LDs (left) or Dil-OxLDL (right) with lysosomes quantified with PCC (mean ± SEM, n = 5/4/7/10/8/10 in the left figure, n = 5/5/9/9 in the right figure). D Representative images of WT BMDMs and GpnmbR150X BMDMs stained with LTR-labeled lysosomes (Red) (Scale bar, 10 μm) and the quantitative analysis of LTR intensity (mean ± SEM, N = 5 independent experiments, n = 42 images each group). E Representative images of WT BMDMs and GpnmbR150X BMDMs stained with acridine orange (AO) (Scale bar, 5 μm) and the quantitative analysis of intensity per cell (mean ± SEM, N = 3 independent experiments, n = 9 images each group). F Representative images of WT BMDMs and GpnmbR150X BMDMs stained with LysoSensor Yellow/Blue DND-160 (PDMPO) (Scale bar, 10μm) and the quantitative analysis of probe intensity ratio (mean ± SEM, N = 3 independent experiments, n = 16 images each group). G Representative images of WT BMDMs and GpnmbR150X BMDMs transfected with R2pH-LAMP1-3×GLAG plasmid (Scale bar, 10 μm) and the quantitative analysis of intensity ratio (mean ± SEM, N = 3 independent experiments, n = 13 or 11 images). H Representative images of AO-stained BMDMs from WT ApoE−/−, HET ApoE−/−, and GpnmbR150X ApoE−/− mice (captured with a high-content microscope, Scale bar, 500 μm) and corresponding quantitative analysis of intensity ratio (mean ± SEM, N = 3 independent experiments, n = 9 images each group). I Representative images of LTR-labeled lysosomes (Red) in WT BMDMs transfected with pEGFP-GPNMB plasmid or empty vector (Scale bar, 10 μm) and the quantitative analysis of LTR intensity (mean ± SEM, N = 3 independent experiments, n = 8 images each group). J Representative images of LysoSensor Yellow/Blue DND-160 (PDMPO) staining in WT BMDMs transfected with pEGFP-GPNMB plasmid or empty vector (Scale bar, 10 μm) and the quantitative analysis of probe intensity ratio (mean ± SEM, N = 3 independent experiments, n = 11 or 7 images). K Heatmap of the expression of lysosome-associated genes and ATP6V family genes from RNA-seq data of WT BMDMs and GpnmbR150X BMDMs treated with or without OxLDL. L Representative images from immunofluorescence of GPNMB expression (Red) and ATP1A1 + ATP1A2 + ATP1A3 + ATP1A4 expression (Green, upper) or ATP6V0E2 expression (Green, lower) in WT BMDMs, N = 3 independent experiments. M Representative western blot images showing the expression of GPNMB and LAMP1 in BMDMs from WT, HET, and GpnmbR150X mice treated with or without OxLDL (upper) and the quantitative analysis (lower) (mean ± SEM, n = 3). N Force-directed network visualization of the STRING links between enriched proteins related to the V-ATPase (c) from whole WT BMDM lysates using an anti-GPNMB antibody. Data are expressed as mean ± SEM. P values were calculated by unpaired Student t-test (C–G, I, J) or one-way ANOVA (H, M). Differences are significant for *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Autophagosomes and autolysosomes play a pivotal role in lipophagy. Previous studies have demonstrated that endogenous GPNMB facilitates LC3 endosomal fusion and lysosomal acidification of phagosomes in BMDMs17. In contrast, the number of GFP-LC3-labeled autophagosomes was increased in GpnmbR150X ApoE−/− BMDMs (Fig. S11a, b). Moreover, Gpnmb-mutation led to a significant increase in mRFP-eGFP-LC3-indicated autolysosomes in ApoE−/− BMDMs (Fig. S11d−f), a finding supported by electron microscopy observations (Fig. S11g). When labeling autolysosomes with GFP-LC3 and LAMP1, similar results were obtained; however, the variability in LAMP1 expression between GpnmbR150X ApoE−/− BMDMs and WT ApoE−/− BMDMs precluded quantitative analysis (Fig. S11c). Interestingly, Gpnmb-mutation substantially increased LAMP1 expression in BMDMs (Fig. 7M).
To elucidate the underlying mechanisms of enhanced lysosomal function, we compared the expression of additional lysosome-associated genes in RNA-seq data from WT BMDMs and GpnmbR150X BMDMs treated with or without OxLDL (Fig. 7K). Notably, Gpnmb-mutation significantly upregulated the expression of ATP6V family (H+-ATPase, V-ATPase) genes in BMDMs (Fig. 7K). Furthermore, five members of the ATP6V family were identified as potential binding proteins of GPNMB (Fig. 7N), consistent with previous findings in HUVECs20. Subsequent experiments confirmed the colocalization of GPNMB with ATP1A1, a known binding partner, and ATP6V0E2, the ATP6V family member exhibiting the highest fold change (Fig. 7L).
The findings presented here suggest that Gpnmb-mutation enhances lysosomal degradation and lipophagy in BMDMs, likely through the upregulation of V-ATPases.
siRNA-lipid nanoparticles targeting Gpnmb in lesional macrophages alleviated atherosclerosis in ApoE −/− mice
Based on the aforementioned results, we hypothesized that inhibiting GPNMB expression and function in macrophages could be a promising strategy for atherosclerosis prevention. Targeted delivery of siRNA using lipid nanoparticle (LNP) technology has emerged as a potential approach for targeted therapy41,42. We synthesized siRNA targeting Gpnmb (siGpnmb) and confirmed its silencing efficiency (Fig. S12a). Consistent with previous findings, lipid content (ORO staining) and uptake of Dil-OxLDL were reduced when GPNMB expression was knocked down using siGpnmb in ApoE−/− BMDMs (Fig. S12b−d). To target lesional macrophages, we designed a protocol for synthesizing LNPs-siGpnmb with surfaces coupled with anti-CD36 and anti-TREM2 antibodies (Fig. 8A). We characterized the physicochemical properties of both targeted and nontargeted LNPs-siRNA (Fig. S12e and Supplementary Table 8). Subsequently, we evaluated their uptake efficiency in BMDMs at various concentrations (Fig. S12f) and found that Gpnmb expression was significantly inhibited at a siRNA concentration of 50 nM (Fig. 8B). Importantly, cell viability remained comparable between BMDMs treated with LNPs-siRNA at 50 nM or 100 nM and control groups (Fig. S12g), leading us to select 50 nM for in vitro experiments. BMDMs treated with LNPs-siGpnmb exhibited significantly lower lipid content, as assessed by BODIPY and Nile Red staining, and decreased uptake of Dil-OxLDL (Figs. 8C, D and S12h−j). Furthermore, we observed high colocalization of internalized LNPs-siRNA (red) with lysosomes and late endosomes (green) 1-hour post-incubation. However, after 12 hours, most LNPs-siRNA were located outside of endosomes, suggesting their escape from lysosomal and late endosomal degradation and release into the cytoplasm (Fig. 8E). In summary, treatment with LNPs-siGpnmb effectively inhibited macrophage foaming, aligning with observations from Gpnmb-mutated and myeloid-specific Gpnmb knockout macrophages.
A Schematic diagram of targeted siRNA-LNPs design. Created in BioRender. Wang, M. (2025) https://BioRender.com/39a6n06. B Quantitative PCR analysis of Gpnmb expression in BMDMs treated with the indicated concentration of siGpnmb-LNPs or siNC-LNPs, N = 3 independent experiments. Representative images of ApoE−/− BMDMs treated with siGpnmb-LNPs or siNC-LNPs followed by Dil-OxLDL-treatment for 12 h (right, C) or OxLDL-treatment for 24 h and Nile Red staining (left, D). N = 3 independent experiments. The quantitative analysis of total fluorescence intensity of Dil-OxLDL (C, left, n = 30 images, Scale bar, 10 μm) and Nile Red (C, right, n = 50 images, Scale bar, 15 μm), mean ± SEM, N = 3 independent experiments. E Representative confocal microscopic images of ApoE−/− BMDMs after incubation for 1, 3, or 12 h with Cy5.5-siRNA-LNPs (red). Nuclei were stained with Hoechst 33342 (blue), and late endosomes were stained with LysoTracker Red (green), N = 3 independent experiments. Scale bar, 2 or 3 μm. F At the fifth day after the whole eight injections of Cy5.5-siRNA-LNPs, in vivo MSOT imaging of Cy5.5-siRNA-LNPs in the aortic arch, descending thoracic aorta, thoracic abdominal aorta, and abdominal aorta from 16-week WD-fed ApoE−/− mice (Late Group, male). Aortic arch: White dotted lines frame, N = 3 independent experiments. G At 12 h after whole eight injections of Control-LNPs or CD36 + TREM2-LNPs, In Vivo Animal Live-Cell Imaging of Cy5.5-LNPs in 16-week WD-fed ApoE−/− mice (male), mean ± SEM, N = 3 independent experiments, n = 5/12/6 images for plaque/ liver/ kidney). H, At 24 h after whole eight injections of Control-LNPs or CD36 + TREM2-LNPs, ex vivo fluorescence imaging of isolated organs from 12-week WD-fed ApoE−/− mice (Early Group, male), mean ± SEM, N = 4 independent experiments). I Representative images of cross sections of the aortic sinus from immunofluorescence of F4/80+ macrophages (Green), Nile Red+ foam cells (Red), and Cy5.5-siRNA-LNPs (Magenta) in atherosclerotic plaque from 16-week WD-fed ApoE−/− mice (Late Group, male), N = 3 independent experiments. J, At the seventh day after the whole eight injections of Cy5.5-siRNA-LNPs, ex vivo fluorescence imaging of isolated aortas from 12-week WD-fed ApoE−/− mice (Early Group) and 16-week WD-fed ApoE−/− mice (Late Group), mean ± SEM, n = 3. K The increased body weight (= final weight – WD-starting weight, n = 5/7/3/4) and the levels of total triglyceride, TCHO, and LDL-C in serum from 12-week WD-fed ApoE−/− mice (Early Group, male, n = 7) and 16-week WD-fed ApoE−/− mice (Late Group, male, n = 6), mean ± SEM. L, On the sixth day after the whole eight injections of Cy5.5-siRNA-LNPs, the representative images of ultrasound examinations on the aortic arch and abdominal aorta from 12-week WD-fed ApoE−/− mice (male, n = 3). M, Representative photomicrographs and the quantification of en face lesion area in the whole aortas from 12-week WD-fed ApoE−/− mice (Early Group; left upper, male, n = 7; left lower, female, n = 3) and 16-week WD-fed ApoE−/− mice (Late Group, male, n = 5), mean ± SEM. N Representative flow cytometric expression of CD64 and BODIPY levels in aortic macrophages (left) and the quantitative percentages of flow cytometric BODIPYhi CD64hi foam cells in aortic macrophages (right, mean ± SEM, male, n = 3/group). O The quantification of plaque area percentages of Oil-Red-O-stained cross sections of aortic sinus from 12-week WD-fed ApoE−/− mice (Early Group; n = 5; female, n = 3) and 16-week WD-fed ApoE−/− mice (Late Group, male, n = 5), mean ± SEM. For each mouse, the average of the staining results from three consecutive sections was calculated for statistical analysis. P, Representative images of cross sections of the aortic sinus from immunofluorescence of F4/80+ macrophages (Green) and Nile Red+ foam cells (Red) in atherosclerotic plaque from 12-week WD-fed ApoE−/− mice (Early Group; n = 5; female, n = 3) and 16-week WD-fed ApoE−/− mice (Late Group, male, n = 5), mean ± SEM. For each mouse, the average of the staining results from three consecutive sections was calculated for statistical analysis. P values were calculated by unpaired Student t-test (B, C, G, H, J, K, M–P). Differences are significant for *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
To increase the targeting efficacy of Cy5.5-modified LNPs-siRNA, we utilized antibodies against CD36 and TREM2, two well-established surface markers of macrophage-derived foam cells in atherosclerotic lesions14,43,44. Antibodies against these proteins facilitated the targeting of LNPs-siRNA to foam cells (CD36 + TREM2-LNPs), compared to their respective isotype control antibodies (Control-LNPs). Employing in vivo fluorescence imaging, we observed a putative distribution of CD36 + TREM2-LNPs in mouse aortas and a higher abundance of Control-LNPs in liver and kidney compared to CD36 + TREM2-LNPs (Fig. S12k). According to the results of photoacoustic imaging, the highest CD36 + TREM2-LNPs signal was detected at the aortic arch (white dotted lines in Fig. 8F and red circle in Fig. S12l)45, colocalizing with HbO2 signal and distinct from the abdominal aorta (blue circle in Fig. S12l) on x,y-cross section45. In contrast, the signals from Control-LNPs-treated mice were predominantly distributed in the liver and kidneys, moderately in the spleen, rarely at the aortic arch (Fig. S12l). Further confirmation by in vivo small animal real-time cell imaging revealed a significantly increased distribution of CD36 + TREM2-LNPs in atherosclerotic plaques and kidneys, accompanied by a marked reduction in hepatic accumulation (Fig. 8G), while no significant differences were observed in other major organs (Fig. S12m). Ex vivo fluorescence images of isolated aortas (Fig. S12n) and other major organs (Fig. 8H) showed consistent results. Notably, immunofluorescence staining further verified the targeting of CD36 + TREM2-LNPs to plaque macrophages and foam cells (Fig. 8I).
The therapeutic effect of LNPs-siGpnmb in atherosclerotic ApoE−/− mice was further evaluated. Given that Gpnmb-mutation showed enhanced atheroprotective effects in ApoE−/− mice fed with a WD for a longer time (Fig. 5A), we built atherosclerotic ApoE−/− mice fed with a WD for 12 weeks (Early-stage group, marked as Early group in figures) and atherosclerotic ApoE−/− mice fed with a WD for 16 weeks (Late-stage group, marked as Late group in figures) to analyze the therapeutic effect of LNPs-siGpnmb in atherosclerosis of varying degrees of severity. Mice were injected intravenously with LNPs-siNC or LNPs-siGpnmb (1 nmol siRNA per injection, 200 μL) once every three days for a total of eight injections. Ex vivo fluorescence images showed that the signal of Cy5.5-LNPs corresponded to the location of observed plaques, but the area and intensity of the fluorescent signal did not appear to fully characterize the whole location and area of the plaque (Fig. 8J). Overall, physiological and biochemical analyses revealed lower weight gain and reduced serum levels of TG, TCHO, and LDL-C in ApoE−/− mice treated with LNPs-siGpnmb compared to those treated with LNPs-siNC (Figs. 8K and S12o). Importantly, hematological analysis and histopathological examination revealed that neither LNPs-siGpnmb nor LNPs-siNC exhibited significant toxicity in ApoE−/− mice (Supplementary Tables 9 and Fig. S12p). Using ultrasound examinations, we detected atherosclerotic plaques in the early-stage group and found that LNPs-siGpnmb treatment mitigated plaque burden in both the aortic arch and abdominal aorta compared to LNPs-siNC (Figs. 8L and S12q). Similarly, ORO staining of en face lesions in the entire aorta and cross sections of the aortic sinus confirmed the beneficial effects of LNPs-siGpnmb, which were observed in both male and female mice (Figs. 8M, N and S12r). Additionally, flow cytometry analysis confirmed the presence of Cy5.5-LNP+ cells in plaque-containing aortas (Fig. S12s). The percentage of macrophages was slightly decreased, and the percentage of foam cells within macrophages was notably reduced in ApoE−/− mice injected with LNPs-siGpnmb compared to LNPs-siNC (Figs. 8O and S12s). The mitigated plaque burden was further validated by immunofluorescence assays (Fig. 8P).
Overall, the LNPs-siGpnmb treatment effectively targeted macrophages and macrophage-derived foam cells in atherosclerotic plaques. This targeted therapy significantly alleviated the atherosclerotic burden in ApoE−/− mice with both early-stage and advanced-stage atherosclerosis, as demonstrated by the comprehensive staining results obtained from the entire aorta and cross-sections.
Discussion
Under normal physiological conditions, vascular macrophages phagocytose subendothelial OxLDL and regulate intracellular lipid turnover by forming inert lipid droplets to prevent lipotoxicity caused by excessive intracellular free cholesterol. The underlying mechanisms of this process remain to be fully elucidated6. We discovered that GPNMB+ vesicles are involved in the internalization and transport of lipoproteins in macrophages. In Gpnmb-mutant BMDMs, while the internalization of lipoproteins by early endosomes was not significantly altered, a combination of reduced lipoprotein uptake, impaired lipid droplet generation from internalized lipoproteins, increased localization of lipid droplets or lipoproteins to mitochondria and lysosomes, and enhanced lipid β-oxidation in the mitochondrial pathway and lipolysis in the lysosomal pathway resulted in decreased intracellular lipid droplet accumulation. Collectively, these findings suggest that GPNMB promotes the formation of lipid droplets from internalized lipoproteins in macrophages while simultaneously inhibiting lipid droplet degradation. This dual role contributes to the storage of intracellular lipoproteins as lipid droplets, thereby regulating lipid turnover. This process is crucial for macrophage decision-making regarding cholesterol storage and metabolism under both physiological and pathological conditions.
In the pathological context of atherosclerosis, characterized by excessive subendotheliadl OxLDL, lipid stress, and subsequent lysosomal deficits, led to compensatory upregulation of GPNMB expression. This upregulation promoted increased OxLDL uptake by macrophages and the formation of more lipid droplets to counter intracellular lipotoxicity, inadvertently accelerating macrophage foaming and the atherosclerotic process. These findings align with the high expression of GPNMB in various types of lipid-laden macrophages26,28,32,46,47 and partially explain the upregulation of GPNMB expression in response to multiple lysosome-stimulating factors32,48. Therefore, GPNMB may function as a protective agent against lipotoxicity and lysosomal dysfunction induced by lipid stress in both macrophages and foam cells. Unfortunately, the physiological function of GPNMB in macrophage processing of OxLDL has been shown to exacerbate plaque progression. Nevertheless, this research offers novel insights into facilitating plaque regression and remission by restoring foam cells to their proper function in lipoprotein processing by modulating the lipotoxicity-coping system. GPNMB emerges as a key target for future investigations within this area.
Current research on atherosclerosis treatment has primarily focused on inhibiting lipoprotein uptake and enhancing lipid efflux in foam cells49,50. However, regulating the intracellular processing of lipoproteins represents another promising therapeutic avenue. Our studies demonstrated that Gpnmb-mutation or myeloid-specific Gpnmb knockout mitigated atherogenesis in ApoE−/− mice. Building upon these findings, we employed LNPs specifically designed to target foam cells for tdhe delivery of siRNA aimed at silencing Gpnmb in plaque macrophages. This intervention led to a significant reduction in plaque burden and blood lipid levels in treated mice. While our results support GPNMB as a potential therapeutic target for atherosclerosis, further optimization of this treatment approach is necessary. Specifically, it is crucial to closely monitor additional toxicity indicators to ensure patient safety. Regarding the apparent distribution of CD36 + TREM2-LNPs in the kidneys, future research should focus on enhancing the targeting efficacy and extending the duration of action of LNPs-siGpnmb through modifications to the delivery strategy. In the kidney, CD36 is prominently expressed in proximal and distal tubular epithelial cells, podocytes, mesangial cells, microvascular endothelial cells, and interstitial macrophages51,52,53,54,55,56,57. Its expression is significantly elevated in chronic kidney disease (CKD), particularly in diabetic nephropathy (DN) contexts, where it correlates with dysregulated renal lipid accumulation51,52,57,58,59. This upregulation is further amplified in diabetic kidney disease (DKD) patients with concomitant hyperlipidaemia60. Moreover, renal biopsies from DKD patients demonstrate increased infiltration of TREM2+ macrophages, specifically localized to glomerular and tubulointerstitial compartments61. Based on the aforementioned literature, although renal macrophages are not our primary target cells, it is inevitable and undeniable that the CD36 + TREM2-LNPs accumulate and persist within these cells. This retention may partly account for the significant signal enhancement in the kidney observed in our study (Fig. 8G−H). GPNMB can serve as a biomarker of renal injury in both experimental models of kidney disease and CKD62,63. Uremic macrophages demonstrate elevated GPNMB expression, concomitant with heightened production of proinflammatory cytokines and suppressed expression of anti-inflammatory cytokines64. Although only minimal morphological abnormalities were detected in renal tissues of mice treated with CD36 + TREM2-LNPs in our study (Fig. S12p), further studies are warranted to determine the functional consequences of macrophage-specific Gpnmb knockdown on overall renal function. Furthermore, the enhanced signal in the kidneys may also be explained by accelerated metabolic clearance of the LNPs, which requires further investigation.
Notably, our observations highlight a remarkable reduction in circulating total cholesterol levels when considering GPNMB as a therapeutic target for atherosclerosis. Our clinical analyses revealed a significant positive correlation between circulating GPNMB levels and serum total cholesterol levels. Experimental validation across multiple models demonstrated consistent cholesterol-lowering effects upon GPNMB knockdown, specifically including systemic GPNMB-mutant mice, myeloid-specific Gpnmb knockout mice, and foamy macrophage-targeted Gpnmb-silencing models. We speculate two coordinated mechanisms1: Gpnmb-deficiency upregulated the expression of phagocytosis-associated proteins (e.g., CD36), enhancing macrophage uptake of OxLDL and other cholesterol components2; Concurrent activation of lysosomal lipid degradation pathways and ABC transporters (e.g., ABCA1 and ABCG1) potentiated reverse cholesterol transport. These synergistic adaptations collectively enhanced clearance of atherogenic lipoproteins (particularly LDL and modified forms) in preclinical models. However, despite the capacity of macrophages to internalize LDL and OxLDL, their impact on circulating total cholesterol and LDL-C concentrations is limited compared to the liver, which serves as the central hub for cholesterol catabolism and excretion65,66. Gong et al. demonstrated that hepatocyte-derived GPNMB activates the CD44–AKT–SREBP1c signaling axis in adipocytes, thereby promoting de novo lipogenesis, both in vivo and in vitro24. Betaine attenuates lipid accumulation and gluconeogenic activity in the liver, partially mediated by its suppression of GPNMB transcriptional expression22. This study primarily investigates the role of GPNMB during foam cell formation. However, further research is required to delineate the functional contribution of GPNMB to hepatic LDL metabolism, which may provide a mechanistic basis for the observed reductions in TCHO and LDL-C levels in both systemic Gpnmb-mutant models and macrophage-specific Gpnmb-knockout models.
In summary, we demonstrated that GPNMB plays a critical role in intracellular lipoprotein processing, maintaining uptaken OxLDL as stored lipid droplets in macrophages under physiological conditions. In the pathological state of atherosclerosis, these mechanisms sharpen the transformation of macrophages into foam cells. However, Gpnmb-mutation, myeloid-specific knockout of Gpnmb, and silence targeting Gpnmb in foam cells using siRNA effectively reversed this process, suggesting the potential of GPNMB as a therapeutic target for atherosclerosis. Additionally, sGPNMB showed promise as a biomarker for atherosclerosis, providing a valuable clinical indicator for monitoring disease severity. A graphical summary was provided in Fig. 9. These findings elucidate the role of GPNMB in intracellular lipoprotein processing within macrophages and highlight its significant potential as a therapeutic target for atherosclerosis.
GPNMB plays a critical role in intracellular lipoprotein processing, maintaining uptaken OxLDL as stored lipid droplets in macrophages under physiological conditions. In the pathological state of atherosclerosis, these mechanisms sharpen the transformation of macrophages into foam cells. However, Gpnmb-mutation, myeloid-specific knockout of Gpnmb, and silence targeting Gpnmb in foam cells using siRNA effectively reversed this process, suggesting the potential of GPNMB as a therapeutic target for atherosclerosis. Created in BioRender. Wang, M. (2025) https://BioRender.com/39a6n06.
Methods and materials
Human serum and carotid plaque sample
All samples were obtained from patients after receipt of written informed consent in accordance with the Declaration of Helsinki. The study protocol was approved by the Xiangya Hospital of Central South University, Hunan, China (202005379). The basic demographic information of all participants was given in Supplementary Table 1. Diagnoses of both stable and unstable patients were verified independently by the responsible clinicians. Treating physicians listed in Supplementary Table 1 were advised to apply guideline-based therapy regimens.
In all participants, blood samples were collected at the time of presentation before any intervention (Supplementary Table 1). The serum was extracted by centrifugation at 1000 × g for 10 min at 4 °C and then transferred to fresh tubes and stored at −80 °C. sGPNMB concentration was measured using a human ELISA kit (DY2550, R&D). Statistical analyses were conducted using the R statistical language (version 4.3.2), RStudio (Version 2023.9.1.494), and the following packages: mice, tableone, and forestploter. Calculation and visualization of differences and correlation were performed with GraphPad Prism (9.0.0) and IBM SPSS (28.0).
Human carotid plaque samples were fixed in 4% PFA for 24 h, and then dehydrated with a gradient of ethanol and embedded in paraffin. Sections (8 μm) were cut on a Finesse 325 (Thermo Scientific). After dewaxing and hydrating with xylene and gradient ethanol, antigen retrieval was performed on paraffin-embedded sections of human plaque sample by citric acid (pH 6.0, Sigma-Aldrich) antigen retrieval buffer, and endogenous catalase was blocked by treatment with 3% H2O2. After permeabilized with 0.15% Triton X-100 in PBS, paraffin-embedded sections were blocked with PBS containing 10% horse serum for 1 h at room temperature, and then incubated with Avidin for 15 min, followed by incubation with Biotin for 15 min. Afterward, sections were incubated with diluted primary antibodies overnight at 4 °C and subsequently with secondary antibody for one hour at room temperature. ABC reagent and NovaRed peroxidase substrate of the Vector-kit (Vector Laboratories) were used for color development according to the specifications. The slices were imaged with a Panoramic SCAN (3D Histech, Budapest, Hungary) and viewed in CaseViewer (3D Histech, software version 2.3.0). The following antibodies were used: Anti-CD68 antibody (ab213363, Abcam), Anti-CD31 antibody (ab9498, Abcam), Anti-αSMA antibody (ab7817, Abcam), and Anti-GPNMB antibody (ab222109, Abcam).
Mice and atherosclerosis induction
C57BL/6J mice and GpnmbR150X (DBA/2) mice were purchased from Huafukang (China), and the GpnmbR150X mutation was identified through sequencing. GpnmbR150X mice were crossed with C57BL/6J mice or ApoE−/− mice (on C57BL/6 background) and then backcrossed with C57BL/6J mice, DBA/2 mice, or ApoE−/− mice for 6 generations. The seventh filial generation mice were used to obtain the littermates of WT, HET, GpnmbR150X, WT ApoE−/−, HET ApoE−/−, and GpnmbR150X ApoE−/− mice on C57BL/6 background (BXD-C background) as well as GpnmbR150X and GpnmbR150X ApoE−/− mice on DBA/2 background (BXD-D background). During breeding, mutant Gpnmb was confirmed by generating PCR as previously described17,23. Mice were fed a western diet (Huafukang, China) for 16–28 weeks to induce atherosclerosis.
Gpnmb-flox mice (Strain NO. T008779) were purchased from GemPharmatech (Nanjing, China). These mice were crossed with C57BL/6 Lyz2cre/cre mice, and the resulting offspring were crossed to generate Gpnmbfl/fl and Gpnmbfl/flLyz2cre/cre (GpnmbΔMAC) littermates. Crossed with ApoE−/− mice, the resulting offspring were crossed to create Gpnmbfl/flApoE−/− and Gpnmbfl/flLyz2cre/creApoE−/− (GpnmbΔMAC ApoE−/−) littermates. Gpnmbfl/flApoE−/− and Gpnmbfl/flLyz2cre/creApoE−/− (GpnmbΔMAC ApoE−/−) littermates were fed with the WD (XT108C, Xietong Bio) for 12, 16, or 20 weeks to develop atherosclerotic plaques.
To assess the in vivo therapeutic effect of siRNA-LNPs for the treatment of atherosclerosis, we first fed six- to eight-week-old ApoE−/− mice with the WD (XT108C, Xietong Bio) for 7 or 11 weeks to establish the early or late atherosclerotic mouse models, respectively. The male mice were randomly divided into 4 groups (with 3–5 mice per group), and the female mice were randomly divided into 2 groups (with 3 mice per group). After establishing the mouse model of atherosclerosis, these mice were injected intravenously with LNPs-siNC (1 nmol siRNA per injection, 200 μL) or LNPs-siGpnmb (1 nmol siRNA per injection, 200 μL) once every three days (total eight times). During the treatment period, the mice were maintained on the WD. After treatment, the mice were fed with the WD for another week and then were euthanized to collect samples for follow-up experiments.
Our study examined male and female animals, and similar findings are reported for both sexes. Mice were euthanized, and then serum samples and tissue samples were collected. All mice were maintained at the SPF facilities of the Beijing Institute of Lifeomics. All experimental procedures in mice were approved by the Institutional Animal Care and Use Committee at the Beijing Institute of Lifeomics (IACUC-20220707-47MB).
Biochemical and physiological assays
After deep anesthetization, blood samples were collected from mouse hearts via cardiac puncture. For blood routine examination, 50 μL of fresh blood was collected in EDTA-coated collection tubes immediately. Routine blood analysis was performed using an Auto Hematology Analyzer (BC-5000 Vet, Mindray). For biochemistry examination, the whole blood was allowed to stand at room temperature and followed by centrifugation at 12000 rpm for 10 min to obtain serum samples. Serum biochemistry analysis was performed on a fully automatic biochemical analyzer (7020, Hitachi High-Technologies, Japan). All indicators were summarized in Supplementary Tables 4–7.
Staining of lesions in the aorta and aortic sinus
For en face analysis, after overnight fixation in 4% PFA, the thoracic aorta was opened longitudinally.
The opened aorta was incubated with propylene glycol for 5 min, and stained with 60% Oil Red O (Sigma-Aldrich) and imaged with a Digital Camera (EOS 70D, Canon).
Mouse heart samples were dehydrated and then embedded in paraffin (Sakura Finetek), followed by cryosectioning (10 μm) on a Finesse 325 (Thermo Scientific) to obtain cross sections of the aortic sinus. Paraffin-embedded sections were dewaxed and rehydrated using standard histological procedures. For hematoxylin and eosin (HE) staining, the sample sections were stained with hematoxylin, followed by differentiation with hydrochloric acid and alcohol, and blue return with loose water. The sample sections were then stained with eosin for 5 min and rinsed with water. For modified Russell-Movat staining, the sample sections were stained with Alixin blue (DG0041, LEAGENE) and differentiated with alkaline ethanol solution. The sample sections were then stained with Weigert hematoxylin for 30 min, followed by differentiation and counterstaining. Afterward, sections were successively stained with Safranin O (S8020, Solarbio)/Acid Fuchsin (71019360, Sinopharm Chemical Reagent) mixed solution (4:1), EVG dye solution(G1042, Servicebio), andSirius scarlet (PH1099, PHYGENE) according to the instructions. The stained sections were dehydrated with alcohol and xylene and then coverslip-mounted. The images of stained sections were captured using an Eclipse Ti-E Inverted Research Microscope (Nikon, Japan).
Flow cytometry and antibodies
Mice were perfused with at least 10 mL of PBS containing sodium heparin to eliminate blood contamination before isolating the aorta. The whole aortas were carefully dissected and cut into 2–5 mm pieces and incubated at 37 °C for 75 min with gentle shaking in a PBS solution with calcium and magnesium containing DNase I (90 U/mL), collagenase I (675 U/mL), collagenase XI (187.5 U/mL), hyaluronidase (90 U/mL)12. After enzyme digestion, the samples were disaggregated by passing through a 70 µm nylon sieve (BD Bioscience) and then centrifuged at 500 g for 5 min at 4 °C to collect the cell pellet. Cells were first blocked by anti-CD16/32 antibody and then stained with other antibodies against surface markers at 4 °C for 30 min. After centrifugation and cleaning, cells were stained with Ghost DyesTM (TONBO) and then permeabilized with the FoxP3 transcription factor-staining buffer set (eBioscience). After permeabilization and cleaning, foam cells were labeled with 40 nmol/L BODIPY493/503 (D3922, Invitrogen) in PBS at 4 °C for 30 min. Flow cytometric analyses were performed using a fluorescence-associated cell sorter (FACS) Fortessa or LSRII (BD Biosciences) instrument and FlowJo software (version 10.3; Tree Star Inc., Ashland, OR).
The following antibodies against mouse surface proteins were used: anti-CD16/CD32 (2.4G2), anti-CD45 (30-F11), anti-CD11b (XMG1.2), anti-Ly6C (HK1.4), anti-Ly6G (1A8), anti-CD64 (x54-5/7.1), anti-F4/80 (BM8), anti-CD36 (HM36). These antibodies were purchased from eBioscience, BioLegend, TONBO Bioscience, and Invitrogen. The GPNMB Monoclonal Antibody (CTSREVL, eFluor™ 660, 50-5708-82) was purchased from eBioscience. The gating strategies were shown in Fig. S2.
Antibodies and plasmids
To generate a pEGFP-GPNMB fusion plasmid and a pEGFP-TFEC fusion plasmid, the ORF of mouse Gpnmb and human Tfec was subcloned into pEGFP-N3 (purchased from TSINGKE, China). mEmerald–RAB5A and mCherry–KDEL plasmids were gifts from Prof. Li Dong (National Laboratory of Biomacromolecules). σ2-mCherry was a gift from Stephen Royle (Addgene plasmid #186579; http://n2t.net/addgene:186579; RRID:Addgene_186579)67. BFP-Sec61 beta was a gift from Gia Voeltz (Addgene plasmid #49154; http://n2t.net/addgene:49154; RRID:Addgene_49154)68. mApple-Moesin-N-14 was a gift from Michael Davidson (Addgene plasmid #54928; http://n2t.net/addgene:54928; RRID:Addgene_54928). mPlum-Rab11a-7 was a gift from Michael Davidson (Addgene plasmid #55998; http://n2t.net/addgene:55998; RRID:Addgene_55998). mEos2-NEEP21-N-18 was a gift from Michael Davidson (Addgene plasmid #57407; http://n2t.net/addgene:57407; RRID:Addgene_57407). mCherry-MannII-N-10 was a gift from Michael Davidson (Addgene plasmid #55074; http://n2t.net/addgene:55074; RRID:Addgene_55074). ptdTomato-N1-FLCN was a gift from Shawn Ferguson (Addgene plasmid #49174; http://n2t.net/addgene:49174; RRID:Addgene_49174)69. mCh-KIF5B*-strep was a gift from Juan Bonifacino (Addgene plasmid #120164; http://n2t.net/addgene:120164; RRID:Addgene_120164)70. pCAG R2pH-LAMP1-3xFLAG was a gift from Massimiliano Stagi (Addgene plasmid #157940; http://n2t.net/addgene:157940; RRID:Addgene_157940)71. pEGFP-N1-MITF-M (Addgene plasmid #38131; http://n2t.net/addgene:38131; RRID:Addgene_38131), pEGFP-N1-TFEB (Addgene plasmid #38119; http://n2t.net/addgene:38119; RRID:Addgene_38119), and pEGFP-N1-TFE3 (Addgene plasmid #38120; http://n2t.net/addgene:38120; RRID:Addgene_38120) were gifts from Shawn Ferguson72.
Monoclonal antibodies against mouse GPNMB (ab188222), GFP (ab183734), TFEC (ab185226), SEC22B (ab181076), ATP1A1 + ATP1A2 + ATP1A3 + ATP1A4 (ab300507), Rab5 (ab218624), LAMP1 (ab289548), and F4/80 (ab300421, Abcam) were purchased from Abcam. Monoclonal antibodies against ERGIC-53 (sc-365158), EEA1 (sc-137130), COPB (sc-393615), MITF (sc-515925), and TFEB (sc-166736) were purchased from Santa Cruz. Monoclonal antibody against Histone-H3 (68345-1) was purchased from Proteintech. Anti-β-Actin antibody (#4970) was purchased from Cell Signaling. Anti-GAPDH antibody (AC033) was purchased from ABclonal. Polyclonal antibody against ATP6V0E2 (NBP1-55100) was purchased from Novus.
Cell culture, treatment, and transfection
Bone marrow–derived macrophages (BMDMs) were isolated and cultured in DMEM medium (11965092, Gibco) containing 10% fetal bovine serum (FBS, 16000044, Gibco), 1% penicillin-streptomycin (C0222, Beyotime), and 10 ng/mL M-CSF (AF-315-02, PeproTech) at 37 °C in a humidified incubator with 5% CO2. Experiments were done on differentiated BMDMs 7 days after plating. Human monocyte line THP1 cells were cultured in RPMI 1640 medium (11875093, Gibco) containing 10% FBS and 1% penicillin-streptomycin and were induced to be macrophages with 162 nM PMA (19-144, Sigma-Aldrich). HEK-293T cells were cultured in the same environment as BMDMs.
Unless specifically stated in the text, treatments with 50 μg/mL OxLDL (YB-002, Yiyuan biotechnology), 30 μg/mL DiI-OxLDL (YB-0010, Yiyuan biotechnology), or 30 μg/mL CF647-stained OxLDL were used to simulate macrophage foaming. CF647-stained OxLDL was prepared with Mix-n-Stain CF647 Antibody Labeling Kits (MX647S100, Sigma-Aldrich) and OxLDL.
For transfection, siRNA and plasmids were transfected into cells by Lipofectamine 3000 (L3000001, Invitrogen) or INVI DNA RNA Transfection Reagent (IV1216025, Invigentech) according to instructions, and the efficiency was detected. Small interfering RNAs were purchased from GenePharma: control siRNA (siNC, sense 5′-UUCUCCGAACGUGUCACGUTT-3′ and antisense 5′-ACGUGACAC GUUCGGAGAATT-3′), Gpnmb siRNA (siGpnmb,sense 5′-GCCUGUUUGUCUCCAACAATT-3′ and antisense 5′-UUGUUGGAGACAAACAGGCTT-3′), and Tfec siRNA (siTfec, sense 5′-UGUUGCUUACAACUGACUTT-3′ and antisense 5′-AGUCAGUUGUAAGCAACAGTT-3′).
Mass spectrometry analysis and RNA sequencing (RNA-seq) analysis
For Immunoprecipitation-Mass Spectrometry (IP-MS), BMDMs were lysed on ice in NETN lysis buffer. The lysates were subjected to immunoprecipitation with anti-GPNMB antibody or control IgG antibody. For proteomic analysis, WT BMDMs and GpnmbR150X BMDMs were lysed on ice in NETN lysis buffer. Subsequent procedures were entrusted to the Mass Spectrometry Platform of the State Key Laboratory of Medical Proteomics (China). The parameters of the database search based on PD2.5IP were the following: database, UniprotSwissProt; taxonomy, mouse; search method, thorough. The mass spectrometry proteomics data have been deposited in the ProteomeXchange Consortium (https://proteomecentral.proteomexchange.org) via the iProX partner repository73,74 with the dataset identifier PXD053686. Differentially expressed proteins were determined with the Limma75 program at p-adj (adjusted p value) <0.05 and logFC (log fold change) > 1.
For RNA sequencing analysis, the RNA of WT BMDMs and GpnmbR150X BMDMs, and two kinds of BMDMs treated with 50 μg/mL OxLDL for 48 h, were extracted with TRIzol™ (15596, Invitrogen). Subsequent procedures were entrusted to Novogene (China). Each group was performed with three independent biological replicates. All samples passed quality control based on the results of FastQC. Differentially expressed genes were determined with the DESeq276 program at p-adj (adjusted p value) <0.05 and logFC (log fold change) > 1. Gene expression data obtained in these studies were deposited in the Gene Expression Omnibus database (GSE274954).
Bioinformatic analysis
For the analysis based on dataset GDS508330, the RMA (robust multi-array average; log scale) of specified genes was extracted to analyze the differences or correlations. For the analysis based on the BiKE database (GSE21545)77, the expression of Gpnmb and other genes was measured by Affymetrix gene array expressed as RMA. Plaque composition extracted with Cibersort (https://cibersortx.stanford.edu/) and further analysis were based on previously published literature31. Mass spectrometry analysis and RNA sequencing analysis were performed in R statistical language (version 4.3.2) and RStudio (Version 2023.9.1.494). An online platform (https://www.bioinformatics.com.cn) was used for data analysis and visualization78.
Immunofluorescence
Mouse heart samples were embedded in OCT compound (Sakura Finetek) and cryosectioned (10 μm) on a Cryotome FSE (Thermo Scientific) to obtain cross sections of the aortic sinus. For permeabilization, paraffin-embedded sections and rewarming frozen sections were applied with 0.15% Triton X-100 in PBS for 30 min. Subsequently, sections were blocked for 2 h with 10% donkey serum in PBS, followed by staining with antibodies diluted in PBS with 1% donkey serum for 10 h at 4 °C. The samples were incubated with secondary antibodies or BODIPY for 2 h and 1 μg/μl DAPI (Invitrogen) for 10 minutes at room temperature. The following antibodies were used: Alexa Fluor™ 488 Secondary Antibody (A21202, Invitrogen) and Alexa Fluor™ 568 Secondary Antibody (A10042, Invitrogen). Sections were mounted with Fluoromount G (YEASEN) and were acquired on a Nikon A1R plus N-STORM confocal microscope with a 10x/0.45 PLANAPO, 20X/0.75 PLANAPO, or 40X/0.75 PLANFLUOR objective lens.
For immunocytochemistry, cells were grown on a 35 mm Glass bottom Confocal dish (NEST, 801002) and were fixed with 4% paraformaldehyde (pH 7.2). Cells were permeablized with 50 mM NH4Cl (pH 7.2) and then blocked with 10% donkey serum in PBS. Subsequent primary and secondary antibody incubations were similar to section immunofluorescence staining. Nuclei were stained with DAPI or Hoechst 33342 (62249, Thermo Scientific). Alexa488 and Alexa594 conjugated secondary antibodies were obtained from Invitrogen. Covered the dish with PBS. Images were acquired with a Zeiss LSM 710 laser scanning confocal microscope using a 63X Plan Apo (NA = 1.4) oil immersion objective and analyzed using Zeiss Efficient Navigation (ZEN) software. Laser excitation wavelengths of 405 nm, 488 nm, 561 nm, and 647 nm detectors were used for fluorescence detection.
Live cell imaging
BMDMS were transfected with pEGFP-N3-GPNMB plasmid for 24 h following the instructions of the transfection reagent, and then the medium was replaced with fresh DMEM full culture medium. After a 24-h culture, the BMDMs were stained with Hoechst 33342, and the medium was then replaced with fresh DMEM full culture medium containing 30 μg/mL DiI-OxLDL. After a 2-h culture, the treated BMDMs were imaged with the High Resolution Live Cell Imaging System (Delta Vision) for 24 h.
BMDMS were co-transfected with pEGFP-N3-GPNMB plasmid and one of the other organelle marker plasmids (see “Antibodies and Plasmids” section) for 24 h following the instructions of the transfection reagent, and then the medium was replaced with fresh DMEM full culture medium containing 30 μg/mL CF647-OxLDL. After a 36-hour culture, the BMDMs were stained with Hoechst 33342 and then imaged with the High Resolution Live Cell Imaging System (Delta Vision) for 2 h. To improve the resolution and contrast in reconstructed images, sparse deconvolution was used by following the previous report79.
Immunoelectron microscope and transmission electron microscope
For immunoelectron microscopy studies, WT BMDMs were fixed with 3% glutaraldehyde and dehydrated with acetone, followed by embedding and polymerization with epoxy resin (02660, SPI).
Ultrathin 60–80 nm sections were prepared, placed on nickel grids, and incubated with the monoclonal antibody directed against GPNMB. Bound antibody was detected using a gold-labeled goat anti-rabbit antibody (BIOSS, China). Sections were counterstained with uranyl acetate-lead citrate and examined using a JEM-1400Flash transmission electron microscope (JEOL; Tokyo, Japan).
For transmission electron microscopy studies, BMDMs were fixed, dehydrated, embedded, and cut into sections, similar to the “Immunoelectron Microscope” section. Sections were examined using a JEM-1200EX transmission electron microscope (JEOL; Tokyo, Japan) at 100 kV.
Cellular staining
For Oil Red O (ORO) staining, BMDMs were fixed using 10% formalin and stained with ORO (Sigma-Aldrich) for 30 min at room temperature. Stained cells were washed with water and rinsed with 60% isopropanol. Multiple fluorescent dyes were used to stain BMDMs following respective instructions, including BODIPY493/503 (D3922, Invitrogen), Lyso-Tracker Red (LTR, C1046, Beyotime), Mito-Tracker Deep Red FM (C1032, Beyotime), Nile red Staining Solution (C0009, APPLYGEN), ER-Tracker Blue (E2380, Solarbio), LysoSensor™ Yellow/Blue DND-160(PDMPO, 40768ES50, YEASEN), and Acridine Orange Hydrochloride (AO, MX4217, MAOKANG). For imaging, the whole scan of 96-well plates was applied with a high-content microscope (Operetta High-Content Imaging System), and detailed images were acquired on a Zeiss LSM 710 laser scanning confocal microscope using a 63X Plan Apo (NA = 1.4) oil immersion objective. Laser excitation wavelengths of 405 nm, 488 nm, 561 nm, and 647 nm detectors were used for fluorescence detection.
Luciferase reporter gene assay
A Gpnmb-firefly luciferase reporter plasmid and negative control plasmids were obtained from GENECHEM (China). The reporter plasmid was constructed by inserting the promoter region of human Gpnmb, a 2-kb region upstream of the transcription start site (TSS), into a luciferase reporter plasmid. HEK-293T cells were seeded in a 48-well plate and co-transfected with the Gpnmb-firefly luciferase reporter plasmid or control plasmid and overexpression plasmid of MiT/TFE family transcription factors using PEI Transfection Reagent. After 12 h of transfection, the medium was replaced with fresh DMEM full culture medium for another 36 h of culture. Then, the Luciferase activity was detected with a Dual Luciferase Reporter Gene Assay Kit (Beyotime, RG027) using the dual-luciferase reporter assay system (Promega Corporation, E1910) according to the manufacturer’s instructions.
Western blotting
Total lysates of tissue or cultured cells were prepared in RIPA buffer (R1091, LABLEAD), including a cocktail of protease (C0101, LABLEAD) and phosphatase inhibitors (C0104, LABLEAD). Lysed samples were separated via SDS-polyacrylamide gel electrophoresis (PAGE) and following transferred to nitrocellulose membranes. Blots were incubated in blocking buffer (5%, w/v, dried skimmed milk in TBST) for one hour and followed by overnight incubation with appropriate antibodies (see “Antibodies and Plasmids” section) diluted in blocking buffer. Western blot images were acquired by Amersham ImageQuant 800 (Cytiva) and quantified using ImageJ (version 1.53c).
Quantitative real-time PCR
Total RNA was extracted (Trizol, Thermo) and reverse transcribed with PrimeScript RT kit (Takara). Quantitative real-time PCR was performed with a SYBR Green PCR kit (Toyobo, Japan) on the CFX96 Touch Real-Time Detection System (Bio-Rad). Relative gene expression was determined using the ΔΔCT method. The mRNA levels of the genes were expressed as a fold change over WT at baseline. The following primers were used: mouse Tfec-forward (5’-GGTCTCACGGATGCTCCTTG-3’) and mouse Tfec-reverse (5’-TCCAGCGCATATCAGGATCATTA-3’); mouse Gpnmb-forward (5’-CGCCTCCACCTTCAACTC-3’) and mouse Gpnmb-reverse (5’-CAATTGTGATGGTGGCTCTG-3’).
Synthesis of siRNA-loaded NPs
DSPE-PEG-Maleimide (80031306, Tanshtech), ((4-Hydroxybutyl)azanediyl)bis(hexane-6,1-diyl)bis(2-hexyldecanoate) (ALC-0315, 81000226, Tanshtech), Methoxypolyethyleneglycoloxy(2000)-N (ALC-0159, 81000227, Tanshtech), 1,2-distearoyl-sn-glycero-3-phosphocholine (DSPC, 81000047, Tanshtech) and Cholesterol (Plant derived) (CHOL, 80031306, Tanshtech) were dissolved in ethanol (molar ratio 0.5:50:1:10:38.5) with 1 mg of Cy5.5-COOH (80031801, Tanshtech). siRNA (siNC or siGpnmb) was dissolved in citrate buffer (100 mM, pH=4–6), and the aqueous and ethanol solutions were quickly mixed at a 3:1 volume ratio. Then, the mixed solution was incubated at room temperature to allow the lipid nanoparticle (LNP) to fully self-assemble, which was next purified in sterile PBS with a dialysis tube for 2 h to remove ethanol and buffer.
Monoclonal antibodies against TREM2 (ab305103, Abcam) and CD36 (sc-7309, Santa Cruz), or their respective isotype control antibodies (ab172730, Abcam; sc-3881, Santa Cruz), were separately dispersed in borate buffer salt (with 0.1% EDTA, 0.1 M, pH 8.0) added with an amount of Traut’s Reagent. The solution was incubated at room temperature for 1 hour, and the solution was then replaced through a 10 KDa ultrafiltration tube and then dispersed in PBS (with 0.1% EDTA, 10 mM, pH 7.4). The above solution was mixed with the liposome solution and incubated overnight at 4 °C. The coupled antibody was then added to the ultrafiltration tube and washed three times with water at 4 °C. Finally, the above solution was collected, filtered, and fixed, and relevant tests about the physicochemical characterization of siRNA-loaded LNPs were performed by Tanshtech.
Assessment of endosomal escape of siRNA-loaded LNPs
BMDMs were seeded in glass-bottom cell culture dishes (801002, NEST) and cultured for 7 days. Cy5.5-siRNA–loaded LNPs were then added and incubated with the cells for 1, 3, or 12 h. The cells were then rinsed three times with PBS and covered with fresh medium. Hoechst 33342 (H1399, Thermo Fisher Scientific) was used to label the nuclei, and Lyso-Tracker Red (LTR, C1046, Beyotime) was used to label lysosomes and late endosomes. The images were acquired on a Zeiss LSM 710 laser scanning confocal microscope using a 63X Plan Apo (NA = 1.4) oil immersion objective. Laser excitation wavelengths of 405 nm, 561 nm, and 647 nm detectors were used for fluorescence detection. The overlap of the red and green fluorescence signals in BMDMs suggests that siRNA-loaded LNPs are trapped inside the endosomes, while the separation of the red and green fluorescence signals suggests the successful endosomal escape of siRNA-loaded LNPs.
Macrophage uptake of LNPs and in vitro assessment of gene silencing by siRNA-loaded LNPs
BMDMs were seeded in 96-well plates and cultured for 7 days. Different concentrations of Cy5.5–siRNA–loaded LNPs were then added. After 24-h incubation, the medium containing LNPs was replaced with fresh medium. After a 24-h incubation, BMDMs were rinsed twice with PBS and then covered with PBS. Fluorescence images were captured with an inverted fluorescence microscope (Eclipse Ti-E, NIKON) to observe the macrophage uptake of LNPs. BMDMs were collected to extract total RNA using Trizol (Invitrogen). Quantitative Real-Time PCR was applied to detect gene expression levels of Gpnmb, indicating the gene silencing by siRNA-loaded LNPs.
In vitro toxicity study
BMDMs were seeded in 96-well plates and cultured for 7 days. Cy5.5-siRNA–loaded LNPs at the indicated concentrations were then added. After 24-hour incubation, the medium containing LNPs was replaced with fresh medium. After an additional 24, 36, or 48-h incubation, cytotoxicity was assayed using the alamarBlue cell viability reagent (40202ES60, YEASEN) according to the manufacturer’s protocol.
In vitro toxicity study
BMDMs were seeded in 96-well plates and cultured for 7 days. Cy5.5-siRNA–loaded LNPs at the indicated concentrations were then added. After 24-h incubation, the medium containing LNPs was replaced with fresh medium. After an additional 24, 36, or 48-h incubation, cytotoxicity was assayed using the alamarBlue cell viability reagent (40202ES60, YEASEN) according to the manufacturer’s protocol.
In vivo fluorescence imaging, photoacoustic imaging, and In vivo microscopy
In vivo fluorescence images were captured using the IVIS® SpectrumCT In Vivo Imaging System (PerkinElmer). At 24 h after the second, fourth, sixth, and eighth injection of Cy5.5-siRNA-LNP, mice were sedated with 3% (v/v) isoflurane, and anaesthetization was maintained with 1.5% (v/v) isoflurane within the IVIS® SpectrumCT In Vivo Imaging System chamber. The belly hair was removed by a trimmer before the fluorescence signal from mice (maximum 5 mice in the visualization chamber) was detected. Representative mice were placed, enabling both CT imaging and fluorescence imaging to further determine the spatial distribution of fluorescent signals between mice. Fluorescence imaging of various tissues and organs isolated from mice was captured after administration. The 2D planar fluorescence images were presented as radiance in photons/sec/cm2/sr.
At 5 days after the eighth injection of Cy5.5-siRNA-LNP, the photoacoustic imaging (PA) studies of mice were performed on a preclinical photoacoustic computerized tomography scanner (MSOT inSight/ inVision 256, iThera Medical). The illumination wavelength of 680 nm (corresponding to the absorption peak of Cy5.5-siRNA-LNP) was adopted. The mice (n = 4) were anesthetized using 3% isoflurane and had their body hair removed with hair removal cream before imaging. The signals of oxyhemoglobin (HbO2), hemoglobin (Hb), and Cy5.5-siRNA-LNP were collected using multispectral optoacoustic tomography software. In Vivo imaging quantification were performed using Living Image® software (version 4.4) as the following procesures: (i) Regions of interest (ROI) selection (identical-sized regions over kidneys/liver/aortas, avoiding background artifacts); (ii) Normalization protocol (fluorescence intensity normalized with automated background subtraction applied); and (iii) Replicates (three independent measurements per organ/mouse).
In Vivo Real-Time Cellular Imaging was performed using an intravital confocal and two-photon microscopy (IVM-CMS, IVIM Technology Inc., Daejeon, Korea). At 12 h after the whole eight injections, anesthetized animals received intravenous injection of fluorescently-labeled probes (488 nm), and sequential images were acquired using appropriate excitation/emission wavelengths (640 nm). Signals in the liver, kidneys, and spleen were acquired in live mice, whereas signals from the heart, lungs, and atherosclerotic plaques were collected post-euthanasia. For plaque imaging, the aorta was first exposed, and visually identifiable plaque tissues were selected. Imaging was performed after locating the regions of interest under low magnification (4×). Cy5.5 signals were quantified using the spot module in Imaris software for 3D reconstruction and particle counting.
Ultrasound examinations
Before euthanization of mice treated with siRNA-LNP, ultrasound examinations were conducted during this study using a portable digital ultrasound unit (MyLab OMEGA eXP VET, Esaote). The system was equipped with a high-frequency linear probe (10–22 MHz, SL3116) and used the MyLab One Vet default software for visualization and measurement of images.
Statistics
Data analysis was performed using GraphPad Prism (Version 9.0.0) to ensure the accuracy and reliability of the results. Continuous variables were expressed as mean ± Standard Error of the Mean (SEM). For comparison between two independent groups, Student’s t-test was applied for normally distributed continuous variables, while the Mann–Whitney U test was used for non-normally distributed data. When comparing differences among multiple groups, one-way ANOVA followed by Tukey’s HSD was used for normally distributed data, and the Kruskal-Wallis test followed by Dunn’s test was used for non-normally distributed data. Correlation analysis was performed using scatter plots and Pearson’s correlation analysis (GraphPad Prism 9.0.0). Linear regression models were fit using IBM SPSS (version 28.0). Two-sided tests were employed for all statistical analyses throughout this paper. Fiji-ImageJ software (1.53c) was used for area quantification of Oil red O staining and Movat staining. Imaris 10 (Oxford Instruments) was used for calculating the intensity, numbers, area, and other detail parameters of fluorescent labels via appropriate reconstruction from the original image.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The microarray datasets described in the paper have been deposited in the Gene Expression Omnibus database with accession numbers GSE274954, GDS5083, and GSE21545 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE274954). The mass spectrometry proteomics data have been deposited in the ProteomeXchange Consortium (https://proteomecentral.proteomexchange.org) via the iProX partner repository with the dataset identifier PXD053686 (https://www.iprox.cn/page/PSV023.html;?url=1726666205444yVEs, password: 14NC. Source data are provided with this paper.
References
Benjamin, E. J. et al. Heart disease and stroke statistics-2018 update: a report from the American Heart Association. Circulation 137, e67–e492 (2018).
Guerrini, V. & Gennaro, M. L. Foam cells: one size doesn’t fit all. Trends Immunol. 40, 1163–1179 (2019).
Susser, L. I. & Rayner, K. J. Through the layers: how macrophages drive atherosclerosis across the vessel wall. J. Clin. Investig. 132, e157011 (2022).
Deng, L., Kersten, S. & Stienstra, R. Triacylglycerol uptake and handling by macrophages: from fatty acids to lipoproteins. Prog. Lipid Res. 92, 101250 (2023).
Luo, J., Yang, H. & Song, B. L. Mechanisms and regulation of cholesterol homeostasis. Nat. Rev. Mol. Cell Biol. 21, 225–245 (2020).
Moore, K. J., Sheedy, F. J. & Fisher, E. A. Macrophages in atherosclerosis: a dynamic balance. Nat. Rev. Immunol. 13, 709–721 (2013).
Borén, J. et al. Low-density lipoproteins cause atherosclerotic cardiovascular disease: pathophysiological, genetic, and therapeutic insights: a consensus statement from the European Atherosclerosis Society Consensus Panel. Eur. Heart J. 41, 2313–2330 (2020).
von Eckardstein, A. LDL contributes to reverse cholesterol transport. Circ. Res. 127, 793–795 (2020).
Zernecke, A. et al. Integrated single-cell analysis-based classification of vascular mononuclear phagocytes in mouse and human atherosclerosis. Cardiovasc. Res. 119, 1676–1689 (2023).
Härdtner, C. et al. A comparative gene expression matrix in Apoe-deficient mice identifies unique and atherosclerotic disease stage-specific gene regulation patterns in monocytes and macrophages. Atherosclerosis 371, 1–13 (2023).
Depuydt, M. A. C. et al. Microanatomy of the human atherosclerotic plaque by single-cell transcriptomics. Circ. Res. 127, 1437–1455 (2020).
Kim, K. et al. Transcriptome analysis reveals nonfoamy rather than foamy plaque macrophages are proinflammatory in atherosclerotic murine models. Circ. Res. 123, 1127–1142 (2018).
Williams, J. W. et al. Limited proliferation capacity of aortic intima resident macrophages requires monocyte recruitment for atherosclerotic plaque progression. Nat. Immunol. 21, 1194–1204 (2020).
Zernecke, A. et al. Meta-analysis of leukocyte diversity in atherosclerotic mouse aortas. Circ. Res 127, 402–426 (2020).
Tomihari, M., Hwang, S. H., Chung, J. S., Cruz, P. D. Jr. & Ariizumi, K. Gpnmb is a melanosome-associated glycoprotein that contributes to melanocyte/keratinocyte adhesion in a RGD-dependent fashion. Exp. Dermatol. 18, 586–595 (2009).
Ripoll, V. M. et al. Microphthalmia transcription factor regulates the expression of the novel osteoclast factor GPNMB. Gene 413, 32–41 (2008).
Li, B. et al. The melanoma-associated transmembrane glycoprotein Gpnmb controls trafficking of cellular debris for degradation and is essential for tissue repair. Faseb J. 24, 4767–4781 (2010).
Robichaud, S. et al. Identification of novel lipid droplet factors that regulate lipophagy and cholesterol efflux in macrophage foam cells. Autophagy. 17, 3671–3689 (2021)
Suda, M. et al. Senolytic vaccination improves normal and pathological age-related phenotypes and increases lifespan in progeroid mice. Nat. Aging 1, 1117–1126 (2021).
Suda, M. et al. Glycoprotein nonmetastatic melanoma protein B regulates lysosomal integrity and lifespan of senescent cells. Sci. Rep. 12, 6522 (2022).
Nickl, B., Qadri, F. & Bader, M. Role of Gpnmb in atherosclerosis of female mice. Biochem. Biophys. Res. Commun. 621, 20–24 (2022).
Yu, X., Li, M., Wang, C. & Guan, X. Glycoprotein non-metastatic melanoma protein B (GPNMB): An attractive target in atherosclerosis. Biochem. Biophys. Res. Commun. 732, 150386 (2024).
Katayama, A. et al. Beneficial impact of Gpnmb and its significance as a biomarker in nonalcoholic steatohepatitis. Sci. Rep. 5, 16920 (2015).
Gong, X. M. et al. Gpnmb secreted from liver promotes lipogenesis in white adipose tissue and aggravates obesity and insulin resistance. Nat. Metab. 1, 570–583 (2019).
Zigdon, H. et al. Identification of a biomarker in cerebrospinal fluid for neuronopathic forms of Gaucher disease. PLoS ONE 10, e0120194 (2015).
Kramer, G. et al. Elevation of glycoprotein nonmetastatic melanoma protein B in type 1 Gaucher disease patients and mouse models. FEBS Open Bio. 6, 902–913 (2016).
Murugesan, V. et al. Validating glycoprotein non-metastatic melanoma B (gpNMB, osteoactivin), a new biomarker of Gaucher disease. Blood Cells Mol. Dis. 68, 47–53 (2018).
Marques, A. R. et al. GPNMB is a potential marker for the visceral pathology in Niemann-Pick type C disease. PLoS ONE 11, e0147208 (2016).
Fukaura, M. et al. Intracerebroventricular treatment with 2-hydroxypropyl-β-cyclodextrin decreased cerebellar and hepatic glycoprotein nonmetastatic melanoma protein B (GPNMB) expression in Niemann-Pick disease type C model mice. Int. J. Mol. Sci. 22, 452 (2021).
Ayari, H. & Bricca, G. Identification of two genes potentially associated in iron-heme homeostasis in human carotid plaque using microarray analysis. J. Biosci. 38, 311–315 (2013).
Orecchioni, M. et al. Olfactory receptor 2 in vascular macrophages drives atherosclerosis by NLRP3-dependent IL-1 production. Science 375, 214–221 (2022).
Gabriel, T. L. et al. Lysosomal stress in obese adipose tissue macrophages contributes to MITF-dependent Gpnmb induction. Diabetes 63, 3310–3323 (2014).
Baba, M. et al. TFE3 Xp11.2 translocation renal cell carcinoma mouse model reveals novel therapeutic targets and identifies GPNMB as a diagnostic marker for human disease. Mol. Cancer Res. 17, 1613–1626 (2019).
Rehli, M., Lichanska, A., Cassady, A. I., Ostrowski, M. C. & Hume, D. A. TFEC is a macrophage-restricted member of the microphthalmia-TFE subfamily of basic helix-loop-helix leucine zipper transcription factors. J. Immunol. 162, 1559–1565 (1999).
Lyons, M. A. et al. Quantitative trait loci that determine lipoprotein cholesterol levels in DBA/2J and CAST/Ei inbred mice. J. Lipid Res. 44, 953–967 (2003).
Gomez, C. P. & Descoteaux, A. Moesin and myosin IIA modulate phagolysosomal biogenesis in macrophages. Biochem. Biophys. Res. Commun. 495, 1964–1971 (2018).
Guardia, C. M. et al. Reversible association with motor proteins (RAMP): a streptavidin-based method to manipulate organelle positioning. PLoS Biol. 17, e3000279 (2019).
Chawla, A. et al. A PPAR gamma-LXR-ABCA1 pathway in macrophages is involved in cholesterol efflux and atherogenesis. Mol. Cell 7, 161–171 (2001).
Olzmann, J. A. & Carvalho, P. Dynamics and functions of lipid droplets. Nat. Rev. Mol. Cell Biol. 20, 137–155 (2019).
Cochain, C. et al. Single-cell RNA-seq reveals the transcriptional landscape and heterogeneity of aortic macrophages in murine atherosclerosis. Circ. Res. 122, 1661–1674 (2018).
Huang, X. et al. Synthesis of siRNA nanoparticles to silence plaque-destabilizing gene in atherosclerotic lesional macrophages. Nat. Protoc. 17, 748–780 (2022).
Tao, W. et al. siRNA nanoparticles targeting CaMKIIγ in lesional macrophages improve atherosclerotic plaque stability in mice. Sci. Translat. Med. 12, eaay1063 (2020).
Collot-Teixeira, S., Martin, J., McDermott-Roe, C., Poston, R. & McGregor, J. L. CD36 and macrophages in atherosclerosis. Cardiovasc. Res. 75, 468–477 (2007).
Willemsen, L. & de Winther, M. P. Macrophage subsets in atherosclerosis as defined by single-cell technologies. J. Pathol. 250, 705–714 (2020).
Cai, J. et al. Au/FeNiPO4-based multiple spectra optoacoustic tomography/CT dual-mode nanoprobe for systemic screening of atherosclerotic vulnerable plaque. Adv. Functional Mater. 34, 2406192 (2024).
Hendrickx, D. A. E. et al. Gene expression profiling of multiple sclerosis pathology identifies early patterns of demyelination surrounding chronic active lesions. Front. Immunol. 8, 1810 (2017).
van Eijk, M. & Aerts, J. The unique phenotype of lipid-laden macrophages. Int. J. Mol. Sci. 22, 4039 (2021).
van der Lienden, M. J. C., Gaspar, P., Boot, R., Aerts, J., & van Eijk, M. Glycoprotein non-metastatic protein b: an emerging biomarker for lysosomal dysfunction in macrophages. Int. J. Mol. Sci. 20, 16 (2018).
Vergallo, R. & Crea, F. Atherosclerotic plaque healing. N. Engl. J. Med. 383, 846–857 (2020).
Lu, Y. et al. The functional role of lipoproteins in atherosclerosis: novel directions for diagnosis and targeting therapy. Aging Dis. 13, 491–520 (2022).
Yang, X. et al. CD36 in chronic kidney disease: novel insights and therapeutic opportunities. Nat. Rev. Nephrol. 13, 769–781 (2017).
Kennedy, D. J. et al. CD36 and Na/K-ATPase-α1 form a proinflammatory signaling loop in kidney. Hypertension 61, 216–224 (2013).
Okamura, D. M., López-Guisa, J. M., Koelsch, K., Collins, S. & Eddy, A. A. Atherogenic scavenger receptor modulation in the tubulointerstitium in response to chronic renal injury. Am. J. Physiol. Ren. Physiol. 293, F575–F585 (2007).
Hua, W. et al. CD36 mediated fatty acid-induced podocyte apoptosis via oxidative stress. PLoS ONE 10, e0127507 (2015).
Ruan, X. Z., Varghese, Z., Powis, S. H. & Moorhead, J. F. Human mesangial cells express inducible macrophage scavenger receptor. Kidney Int. 56, 440–451 (1999).
Yang, P. et al. Inflammatory stress promotes the development of obesity-related chronic kidney disease via CD36 in mice. J. Lipid Res. 58, 1417–1427 (2017).
Gai, Z. et al. Lipid accumulation and chronic kidney disease. Nutrients. 11, 722 (2019).
Susztak, K., Ciccone, E., McCue, P., Sharma, K. & Böttinger, E. P. Multiple metabolic hits converge on CD36 as novel mediator of tubular epithelial apoptosis in diabetic nephropathy. PLoS Med. 2, e45 (2005).
Herman-Edelstein, M., Scherzer, P., Tobar, A., Levi, M. & Gafter, U. Altered renal lipid metabolism and renal lipid accumulation in human diabetic nephropathy. J. Lipid Res. 55, 561–572 (2014).
Fang, Z., Liu, R., Xie, J. & He, J. C. Molecular mechanism of renal lipid accumulation in diabetic kidney disease. J. Cell Mol. Med. 28, e18364 (2024).
Fu, J. et al. The single-cell landscape of kidney immune cells reveals transcriptional heterogeneity in early diabetic kidney disease. Kidney Int. 102, 1291–1304 (2022).
Satirapoj, B., Nast, C. C. & Adler, S. G. Novel insights into the relationship between glomerular pathology and progressive kidney disease. Adv. Chronic Kidney Dis. 19, 93–100 (2012).
Patel-Chamberlin, M. et al. Hematopoietic growth factor inducible neurokinin-1 (Gpnmb/Osteoactivin) is a biomarker of progressive renal injury across species. Kidney Int. 79, 1138–1148 (2011).
Pahl, M. V., Vaziri, N. D., Yuan, J. & Adler, S. G. Upregulation of monocyte/macrophage HGFIN (Gpnmb/Osteoactivin) expression in end-stage renal disease. Clin. J. Am. Soc. Nephrology 5, 56–61 (2010).
Feingold, K. R. Introduction to lipids and lipoproteins. in (Feingold K. R., et al., eds) (Endotext. South Dartmouth (MA): MDText.com, Inc. Copyright © 2000-2025, MDText.com, Inc., 2000).
Norum, K. R., Berg, T., Helgerud, P. & Drevon, C. A. Transport of cholesterol. Physiol. Rev. 63, 1343–1419 (1983).
Küey, C., Sittewelle, M., Larocque, G., Hernández-González, M. & Royle, S. J. Recruitment of clathrin to intracellular membranes is sufficient for vesicle formation. eLife. 11, e78929 (2022).
Zurek, N., Sparks, L. & Voeltz, G. Reticulon short hairpin transmembrane domains are used to shape ER tubules. Traffic. 12, 28–41 (2011).
Petit, C. S., Roczniak-Ferguson, A., Ferguson, S. M. Recruitment of folliculin to lysosomes supports the amino acid-dependent activation of Rag GTPases. J. Cell Biol. 202, 1107–1122 (2013).
Guardia, C. M. et al. Reversible association with motor proteins (RAMP): a streptavidin-based method to manipulate organelle positioning. PLoS Biol. 17, e3000279 (2019)
Ponsford, A. H. et al. Live imaging of intra-lysosome pH in cell lines and primary neuronal culture using a novel genetically encoded biosensor. Autophagy. 17, 1500-1518 (2021)
Roczniak-Ferguson, A. et al. The transcription factor TFEB links mTORC1 signaling to transcriptional control of lysosome homeostasis. Sci. Signal. 5, ra42 (2012).
Ma, J. et al. iProX: an integrated proteome resource. Nucleic Acids Res. 47, D1211-d7 (2019).
Chen, T. et al. iProX in 2021: connecting proteomics data sharing with big data. Nucleic Acids Res. 50, D1522-d7 (2022).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Folkersen, L. et al. Prediction of ischemic events on the basis of transcriptomic and genomic profiling in patients undergoing carotid endarterectomy. Mol. Med. 18, 669–675 (2012).
Tang, D. et al. SRplot: a free online platform for data visualization and graphing. PLoS ONE. 18, e0294236 (2023).
Zhao, W. et al. Sparse deconvolution improves the resolution of live-cell super-resolution fluorescence microscopy. Nat. Biotechnol. 40, 606–617 (2022).
Acknowledgements
This work was funded by the National Key Research and Development Program of China (No. 2020YFA0803903), the National Science Fund for the Distinguished Young Scholars of China (No. 82225009), the National Key R&D Program of China (No. 2023YFA1800203), the State Key Laboratory of Proteomics Foundation (No. SKLP-O201903), the Scientific Research Program of FuRong Laboratory (No. 2024PT5103), and the National Natural Science Foundation of China (No. 82173907). We thank the Imaging Platform (Shaoqiong Yi, Jun Chen), Animal Facility (Chen Qiu, Jiaang Zhao), Flow Cytometry Platform, Mass Spectrometry Platform, and Bioinformatics Platform of State Key Laboratory of Medical Proteomics (China) for providing instrumentation support. We thank Director Hongyue Wang for providing data analysis on MOVAT staining. We thank Qiao Chen and Linying Yue for collecting clinical plasma samples. We thank Quan Cheng and Wei Hou for collecting clinical plaque samples. We thank Yinxiao Du for providing clinical data analysis. We also thank Jianjian Zhou of Beijing Loham Trading Co., Ltd. for SPR assistance. We thank the Home for Researchers editorial team (www.home-for-researchers.com) for language editing services.
Author information
Authors and Affiliations
Contributions
Li Tang, Xiao-Ping Chen, and Junqi Wang conceived the project, designed the experiments, analyzed the data, and wrote the paper. Junqi Wang performed the experiments. Liming Peng provided the human serum samples and the corresponding pathology information. Huan Wang provided expertise in omics data analysis. Chunhua Zhang and Ziliang He provided imaging technology support. Di Liu, Wenting Yang, Fang Yan, and Dianyuan Zhao provided input and assistance in conducting the experiments. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ruibing Wang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, J., Peng, L., Wang, H. et al. Regulation of lipoprotein processing by GPNMB in foamy macrophages: potential therapeutic targets for atherosclerosis. Nat Commun 16, 10030 (2025). https://doi.org/10.1038/s41467-025-65224-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65224-x