Abstract
Palmitoylation is the only fully reversible post-translational lipid modification that impacts 10-20% of the human proteome, but its role during spermatogenesis remains enigmatic. In this study, through generating HA-tagged Abhd10 knock-in mice, Abhd10-null mice, and combining super-resolution fluorescence imaging and electron microscopy, we identify that the S-depalmitoylase ABHD10 (abhydrolase domain containing 10) is a mitochondrial matrix protein, specifically expressed in testis and is essential for male fertility. Abhd10 knockout mice manifest severe sperm motility defects accompanied by malformed mitochondrial sheaths of sperm. Mitochondrial proteomic analysis reveals that ABHD10 deficiency downregulates respiratory chain complex proteins and mitochondrial sheath formation factors SPATA19 and GK2. Using mass spectrometry-based mitochondrial acyl-biotin exchange assays, we systematically identify that loss of ABHD10 leads to the hyper-palmitoylation of multiple functionally critical proteins, including mitochondrial sheath formation factors (SPATA19 and GK2) and aerobic respiration regulators (PDHX, NDUFV1 and SDHB). Co-immunoprecipitation and proximity labeling assays reveal the physical interactions between ABHD10 and its substrates (SPATA19, GK2, PDHX). Collectively, ABHD10 may bind to and mediate the S-depalmitoylation of SPATA19, GK2, and PDHX, thereby regulating the formation of the sperm mitochondrial sheath and mitochondrial function. This work not only identifies S-depalmitoylase ABHD10 as a key determinant of male fertility but also advances our understanding of post-translational regulation during spermatogenesis.
Similar content being viewed by others
Introduction
Infertility affects ~17.5% of the global population, with male factors contributing to half of all cases1. Male factor accounts for 50% of all infertility cases. Among various causes, impaired sperm function represents the predominant etiology, highlighting an urgent need to elucidate molecular regulators of spermatogenesis. Spermatogenesis consists of three main stages: mitosis of spermatogonia, meiosis of spermatocytes, and spermiogenesis of spermatids2. Spermiogenesis is a complex process that involves the condensation of the nucleus, expansion of mitochondria, and formation of the flagellum. The mature sperm tail contains several accessory structures, including the mitochondrial sheath, outer dense fibers, and fibrous sheath, which divide the tail into three distinct parts: the midpiece (containing the mitochondrial sheath, outer dense fibers, and fibrous sheath), the principal piece (containing the outer dense fibers and fibrous sheath), and the end piece (containing the fibrous sheath). Sperm, as a specialized type of cell capable of rapid movement, consumes large amounts of adenosine triphosphate (ATP) during its motility3. The primary source of ATP for sperm motility is derived from glycolysis and oxidative phosphorylation (OXPHOS). Glycolysis takes place in the fibrous sheaths of the sperm head and principal piece, while OXPHOS occurs within the mitochondrial sheath of the sperm midpiece4,5. Oxidative phosphorylation is a complex process that relies on two key components of the inner mitochondrial membrane: the respiratory chain and ATP synthase3. The respiratory chain consists of four multimeric complexes (I, II, III, and IV) and two mobile carriers, coenzyme Q and cytochrome C.
Mitochondrial dysfunction is implicated in a wide array of diseases across various medical fields. Notably, over 40% of mitochondrial proteins are associated with human diseases, underscoring the critical role of mitochondria in maintaining human health6. In sperm, mitochondria are arranged in a distinctive helical pattern in the mid-piece of the sperm flagellum, forming a specialized structure referred to as the mitochondrial sheath7. The formation of mitochondrial sheath occurs during the stage of spermiogenesis. During this stage, spherical mitochondria in the cytoplasm are recruited and aligned around the sperm flagellum, a process known as the alignment phase. Subsequently, these spherical mitochondria will elongate to form crescent-like shape (interlocking phase). Following this, these crescent-shaped mitochondria will further lengthen and tightly wrap around the flagellum to form a helical structure (compact phase)8,9. As a result of these dynamic changes in distribution and morphology, mitochondria are ultimately confined to the mid-piece of the sperm, tightly encircling the flagellum, forming a structure known as the mitochondrial sheath. Armadillo repeat-containing protein 12 (ARMC12), glycerol kinase (GYKL1), Glycerol kinase 2 (GK2) and mitochondrial cardiolipin hydrolase (PLD6) have been reported to regulate the formation of mitochondrial sheath through regulating the aggregation of mitochondria, however, the mechanisms underlying mitochondrial sheath formation remain largely unknown. Our previous studies demonstrate that the structures of human and mouse mitochondrial sheaths share common characteristics10, thus mice are an ideal animal model to study the process of mitochondrial sheath formation.
S-palmitoylation is a widespread post-translational modification that affects 10-20% of the human proteome11,12. More importantly, palmitoylation is the only fully reversible protein-lipid modification, which can be modulated by “writer” enzymes (protein acyltransferases, PATs) and “eraser” enzymes (acyl protein thioesterases, APTs). This modulation regulates various aspects of protein behavior, including stability, subcellular localization, trafficking, activity, protein-protein interactions, and signaling transduction12,13,14. Although palmitoylation was identified several decades ago15, our understanding of this modification has progressed more slowly compared to other types. Abhydrolase domain containing 10 (ABHD10) has recently been identified as a S-depalmitoylase and regulates redox homeostasis via Peroxiredoxin-5 (PRDX5) in vitro16, however, its physiological function in vivo remains unknown.
In this study, we inserted an HA tag into ABHD10 gene, confirming that ABHD10 is a testis-specific protein expressed in the mitochondrial matrix in vivo. To investigate its functional role, we generated Abhd10 knockout mice to study its functions and molecular mechanisms in vivo. The genomic knockout of Abhd10 results in mitochondrial swelling, loss of the mitochondrial inner membrane, and abnormal mitochondrial sheath formation in sperm, ultimately leading to asthenospermia and male infertility. Molecularly, loss of ABHD10 leads to a significant decrease of proteins in mitochondria, particularly those involved in oxidative phosphorylation and mitochondrial sheath formation. Using acyl-biotin exchange (ABE) assays, we further revealed that ABHD10 deficiency leads to a significantly increased palmitoylation of proteins associated with sperm mitochondrial sheath formation (SPATA19 and GK2) and oxidative phosphorylation (PDHX, NDUFV1 and SDHB). Importantly, co-immunoprecipitation and proximity labeling assays identified SPATA19, GK2, and PDHX as interaction partners of ABHD10, establishing these proteins as its primary molecular targets. These results unveil that ABHD10 catalyzed mitochondrial S-depalmitoylase and is essential for male fertility.
Results
Tissue-specific-mitochondrial proteins tend to be localized in testis
To evaluate the heterogeneity of mitochondrial protein expression across mammalian tissues, we systematically analyzed 1140 mouse mitochondrial proteins across 14 major tissues using proteomic data from MitoCarta 3.0. Through calculation of the Specificity Measure (SPM) for each protein in a certain tissue, we identified that 20.70% (236/1140) of mouse mitochondrial proteins exhibited tissue-biased expression (SPM > 0.5) and 4.65% (53/1140) were strictly tissue specific (SPM = 1) (Fig. 1a, b and Supplementary Fig. 1a, b). Strikingly, among tissue-biased and tissue-specific mitochondrial proteins, 22.46% (53/236) and 33.96% (18/53), respectively, showed the highest specificity for the testis (Fig. 1a, c, d). This pronounced testis-specific enrichment suggests that mitochondrial proteins may evolve specialized functional adaptations to support unique physiological demands in male reproduction. Based on these findings, we subsequently focused our investigation on testis-specific mitochondrial protein expression profiles and their potential biological significance.
a Schematic workflow for analyzing mitochondrial proteins (MPs) across 14 murine tissues. Tissue specificity was determined by calculating the specificity measure (SPM) score for each protein, with SPM > 0.5 defining tissue-biased MPs and SPM = 1 indicating tissue-specific MPs. This panel was partly generated by Servier Medical Art and SciDraw (credit Milica Nedeljković, John Chilton and Macauley Smith Breault), licensed under CC BY 4.0 (https://creativecommons.org/licenses/by/4.0/). The drawing of testis is modified from an original provided by Servier Medical Art. b Proportional representation of mitochondrial protein populations: total MPs (cyan), tissue-biased MPs (blue), and tissue-specific MPs (purple). Tissue distribution of testis-enriched MPs among all tissue-biased (c) and tissue-specific (d) mitochondrial proteins, shown as percentage contributions. e The GO term of testis-enriched MPs. The most significantly enriched pathway was spermatogenesis. The names of the genes detected in each pathway are listed above the corresponding bars. Statistical significance (p-values) was determined based on the cumulative hypergeometric distribution. f, g RT-qPCR analyses of Abhd10 mRNA levels in tissues of mouse and human, and cell lines. n = 3 biologically independent mice. Data are presented as mean ± SEM.
In order to conduct a more comprehensive detection of testis-specific mitochondrial proteins, we isolated mitochondria from various tissues and conducted quantitative protein mass spectrometry analysis (Supplementary Fig. 1c). A total of 32 testis-enriched mitochondrial proteins were identified (Supplementary Fig. 1d and Data. 1). Gene Ontology (GO) enrichment analysis revealed that the functions of these proteins are primarily associated with mitochondrial transmembrane transport, piRNA processing (a male germ cell-specific pathway), pyruvate metabolism, spermatogenesis, aerobic respiration, and monocarboxylic acid metabolic processes (Fig. 1e). Among these identified testis-enriched mitochondrial proteins, Abhydrolase domain containing 10 (ABHD10) garnered our attention due to it is a recently discovered S-depalmitoylase and the function of it under physiological conditions in vivo has not been reported in any literature (Supplementary Fig. 1d). Quantitative PCR analysis of various mouse organs and human tissues or cell lines revealed that the mRNA of Abhd10 is predominantly expressed in the testes (Fig. 1f, g). Moreover, sequence alignment analysis across different species revealed that ABHD10 is highly conserved in mammals (Supplementary Fig. 1e), indicating that research conducted on mice has reference significance for human diseases as well.
Loss of Abhd10 leads to male infertility
To elucidate the physiological role of Abhd10 in vivo, we generated a mouse line with a deletion of exon 2 of the Abhd10 gene (Abhd10Del/Del, hereafter referred to as KO) (Fig. 2a). PCR genotyping and mRNA analysis of testicular tissue confirmed the successful generation of Abhd10 knockout mice (KO) (Fig. 2b–d). The adult Abhd10 knockout mice were viable and exhibited comparable body weights to the control mice (Fig. 2e and Supplementary Fig. 2a). Additionally, we assessed the fertility of adult Abhd10 knockout mice. Notably, Abhd10 KO males were completely sterile, while Abhd10 KO females were fertile (Fig. 2f and Supplementary Fig. 2b). Although Abhd10 KO males were infertile, there were no significant differences in the size and weight of their testes and epididymides compared to age-matched control littermates (Fig. 2g, h).
a Schematic diagram of the WT and KO alleles. Scissor: the in vitro transcribed gRNA was designed to target the Exon 2 of Abhd10. Images of scissors were generated by figdraw.com. b The location of the primers used for genotyping are shown. F1 and F2, forward primers, R1 and R2, reverse primers. c Representative PCR genotyping of WT (+/+), Abhd10 Het (+/Del), Abhd10 KO (Del/Del) alleles. d RT-qPCR analysis of Abhd10 expression in testes from WT and KO mice. e The body sizes of wild-type (WT) mice and knockout (KO) mice were similar. Scale bar: 2 cm. f The number of pups per plug in female mice after mating with WT and KO male mice aged 8–12 weeks. g Testes and epididymides from WT and KO mice (8-week-old). Scale bar: 5 mm. h Testis weight of WT and KO mice (8-week-old). i The I-XII stages of seminiferous tubules in WT and Abhd10 KO testes are shown by PAS staining. Scale bar: 50 μm. j PAS staining analyses of the stepwise development of spermatids in testes. Scale bar: 5 μm. k TUNEL staining of WT and KO testes indicated that ABHD10 does not affect the apoptosis of testicular cells. Scale bar: 50 μm. l PAS staining of caput, corpus, cauda of epididymis sections from WT and KO mice (8-week-old). Scale bar: 50 μm. m The representative phase-contrast micrograph of sperm released from WT and KO epididymides. Normal and abnormal spermatozoa are shown. The black arrow indicates discontinuity in the mitochondrial sheath of the sperm. Scale bar: 5 μm. n Sperm count was evaluated in WT and KO mice. o Statistics of motility (%), rapid motility (%) and progressive motility (%) of WT and KO mice, with at least 1000 sperm were examined for per individual. p In vitro fertilization (IVF) indicates that the deletion of ABHD10 in mice leads to the failure of in vitro fertilization. Scale bar: 50 μm (left), 100 μm (right). d, f, h, n, o display quantitative data presented as mean ± SEM, with n = 3–5 biologically independent mice per group. P values were determined using a two-sided unpaired Student’s t-test. e, g, i, j, k, l, m, p show representative images from at least three independent experiments with similar results.
To further investigate the causes of male infertility, we examined the histology of wild-type (WT) and Abhd10 knockout (KO) testes. PAS staining of the testes revealed that the seminiferous tubules of KO mice were similar to those of control mice, containing all types of germ cells. The percentage of each stage of seminiferous tubules in the testes of KO mice was comparable to that of the controls (Fig. 2i and Supplementary Fig. 2c). Additionally, stepwise analyses of round and elongating spermatids demonstrated normal head condensation and sperm acrosome biogenesis in KO mice (Fig. 2j), indicating that ABHD10 is nonessential for sperm head deformation. Furthermore, TUNEL staining revealed no abnormal apoptosis of germ cells in KO seminiferous tubule (Fig. 2k), suggesting that ABHD10 is not required for germ cell survival. Moreover, PAS staining of the epididymides indicated that the caput, corpus, and cauda are comparable in both WT and KO mice (Fig. 2l).
Then we released sperm from epididymides and performed CASA analyses, which revealed that the number of sperms was comparable, but the motility of sperm was significantly decreased (Fig. 2m-o and Supplementary Movie 1, 2). Interestingly, although the head and flagellar curls of the sperm appeared normal, an abnormal mid-piece was observed in Abhd10 knockout (KO) mice; this finding was further substantiated by HE staining analyses (Fig. 2m and Supplementary Fig. 2d, e). Subsequently, we collected sperm from WT and KO mice for IVF (In Vitro Fertilization). At the 6-h mark of IVF, WT sperm were able to swim to the oocytes and bind to the zona pellucida, whereas only a few KO sperm reached the periphery of the oocytes. After 1, 2, 3, and 4 days of fertilization, oocytes bound by WT sperm developed into 2-cell, 4-cell, morula, and blastocyst stages, respectively, while oocytes bound by KO sperm failed to develop (Fig. 2p), suggesting that Abhd10-deficient sperm failed to accomplish fertilization.
To rescue the male infertility phenotype and exclude potential off-target effects in Abhd10 KO mice, we generated a knock-in mouse strain expressing Abhd10 cDNA with its upstream promoter (Supplementary Fig. 2f). Transgenic knock in of Abhd10 fully restored fertility in Abhd10 KO males (Supplementary Fig. 2g), indicating that the observed phenotypes are attributable to Abhd10 loss. Collectively, these findings indicate that ABHD10 is not essential for the survival of germ cells, but is essential for sperm motility and male fertility.
ABHD10 deficiency leads to abnormal mitochondrial morphology and MIM defects
To unveil the cause of reduced sperm motility, we performed scanning electron microscopy (SEM) to examined the morphology of mature spermatozoa released from the cauda epididymides. In Abhd10 KO sperm, the mitochondrial sheath of midpiece was disrupted, while the neck and annulus structures located at the ends of the mitochondrial sheath are normal (Fig. 3a). In contrast to the intact mitochondrial sheaths observed in WT sperm, 100% of Abhd10 KO sperm exhibited severe mitochondrial sheath defects (1022 sperm counted from 3 knockout mice) (Fig. 3b, c). To better visualize the mitochondrial sheaths, we use a freeze-fracture method to process SEM samples. In the control group, ordered spiral mitochondria were observed in the midpiece of the sperm, while the knockout sperm manifested swollen mitochondria and disjointed mitochondrial sheaths (Fig. 3d). These results suggest that ABHD10 is essential for sperm mitochondrial sheath formation.
a Scanning electron microscopy (SEM) showing the ultrastructure of the sperm neck, mitochondrial sheath, and annulus. MS, mitochondria sheath. The red arrow indicates that the mitochondrial sheath is absent in the KO sperm. The white arrow indicates that the annulus structures of both WT and Abhd10 KO sperm are normal. Scale bar: 10 μm (left), 1 μm (right). b Immunofluorescent staining of WT and KO sperm by MitoTracker and DAPI. Scale bar: 10 μm. c Percentage of sperm with abnormal mitochondria sheath (MS) in the epididymides of WT and KO mice. d Freeze-fracture scanning electron microscopy (SEM) analysis reveals that in wild-type (WT) sperm, the mitochondria are arranged in a compressed spiral pattern. In contrast, within the mid-piece of knockout (KO) sperm, the mitochondria appear swollen. Scale bar: 10 μm (left), 1 μm (right). e Longitudinal section of sperm mitochondrial sheath. The mitochondria consist of the inner and outer membrane in WT sperm, while partial of inner membrane was loss in KO sperm. Red arrows indicate the absence of mitochondrial inner membrane. Scale bar: 500 nm. f Percentage of sperm with intact mitochondrial inner membrane. g Transverse sections showing the ultrastructure of the midpiece, principal piece, and end piece of sperm flagella from WT and KO testes and epididymides. The midpiece of KO spermatozoa exhibits structural defects, including malformations or swelling. Note that the 9 + 2 structure was intact in testicular sperm while was destructed in epididymal sperm. Scale bar: 200 nm. c, f display quantitative data presented as mean ± SEM, with n = 3–5 biologically independent mice per group. P values were determined using a two-sided unpaired Student’s t-test. a, b, d, e, g show representative images from at least three independent experiments with similar results.
To identify proteins that function synergistically with ABHD10 during mitochondrial sheath formation, we conducted a series of co-immunoprecipitation (co-IP) experiments by co-expressing ABHD10 with eight known mitochondrial sheath-associated proteins: ARMC12, DDHD1, GK2, KLC3, MFF, SPATA19, TBC1D21, and VDAC3 (Supplementary Fig. 3a). Interestingly, we did not detect the interaction between ABHD10 and these candidate proteins under our experimental conditions (Supplementary Fig. 3b–i). These results indicate that ABHD10 cannot form stable protein complexes with these known sperm mitochondrial sheath formation-associated proteins.
To examine the ultrastructure of mitochondria, we performed transmission electron microscopy (TEM) of sperm from epididymides. In longitudinal sections of WT sperm, uniformly sized mitochondria with two membranes—the inner and outer membranes—encapsulate the sperm axoneme, while the mitochondria of mutant sperm exhibited significant variation, characterized by severely destructed membrane organization or even devoid of the inner mitochondrial membrane and mitochondrial cristae (Fig. 3e, f). TEM analysis of transverse sections of Abhd10 KO sperm also revealed mitochondria with swelling and absence of inner mitochondrial membrane (Fig. 3g). Interestingly, we observed that typical 9 + 2 microtubule axoneme was disrupted in the principal piece, midpiece, and end piece of mutant sperm, whereas it remained intact in mutant testes (Fig. 3g). These results suggest that the formation of 9 + 2 microtubule axoneme is normal during spermatogenesis, while it was disrupted during the process of being released from the testis to the epididymis, hints that mitochondria serves as a physical protective structure for sperm, tightly wrapping around the axoneme to maintain its stability.
OXPHOS was disrupted in Abhd10 KO testes
To unveil the molecular regulatory network of ABHD10 protein in testis, WT and KO testes were isolated for quantitative proteomics analysis (Supplementary Fig. 4a). A total of 301 differentially expressed proteins were identified (Fig. 4a and Supplementary Fig. 4b and Data 2) and the function of these proteins, especially down-regulated proteins, were primarily enriched in mitochondrial respiratory chain complex assembly (Fig. 4b and Supplementary Fig. 4c). Given that ABHD10 is specifically localized to mitochondria, the deregulated proteins encoding the mitochondrial proteome were selected for further analysis17. A total of 48 dysregulated mitochondrial proteins were screened, of which 31 were downregulated and 17 were upregulated (Supplementary Fig. 4d). Strikingly, 70.97% (22/31) down-regulated mitochondrial proteins are localized in the inner mitochondrial membrane (IMM), and 77.27% (17/22) down-regulated IMM proteins are associated with oxidative phosphorylation (OXPHOS) (Fig. 4c).
a Volcano plot showing up- and down-regulated proteins in KO relative to WT testis. Two-sided unpaired Student’s t-test was used to calculate p-value. Only proteins meeting p < 0.05 and |FC| ≥ 2 were defined significantly differentially expressed. b GO enrichment of up- and down-regulated proteins. c Doughnut diagram showing down-regulated proteins mainly IMM proteins, especially OXPHOS-related proteins. d The model for isolating testicular mitochondria from WT and Abhd10 KO mice. Images of mitochondria were created by figdraw.com. Images of testis, equipment and mouse were generated by Servier Medical Art and SciDraw (credit Ethan Tyler and Lex Kravitz), licensed under CC BY 4.0 (https://creativecommons.org/licenses/by/4.0/). The drawing of testis is modified from an original provided by Servier Medical Art. e The volcano plot illustrates the up-regulated and down-regulated proteins in mitochondrial proteomics. Two-sided unpaired Student’s t-test was used to calculate p-value. Only proteins meeting p < 0.05 and |FC| ≥ 1.5 were defined significantly differentially expressed. f Pathways enriched analyses of downregulated proteins through MitoCarta3.0_MitoPathways. g Expression patterns of representative proteins participating in OXPHOS and sperm mitochondrial sheath formation. Each protein is color-coded based on their expression. Heatmaps and bubble plots are utilized to display the fold change and statistical significance. Of these downregulated mitochondrial OXPHOS proteins, four proteins (ND1, ND3, ND4, and ND5) are encoded by the mitochondrial genome, while the remaining proteins are nuclear-encoded. h Western blot was used to detect the expression of COXI (NDUFS4 and NDUFB7), COXIV (COXIV, COX5A, COX6B2 and COX7C), SPATA19 in WT control and Abhd10 KO testes. GAPDH was served as a loading control. Western blot analysis in this panel was performed using the same set of biological samples and this experiment was repeated at least 3 times with similar results. i The ATP levels were significantly decreased in KO testes and sperm compared to control group. j, k Baseline mitochondrial oxygen consumption rate (OCR) and extracellular acidification rate (ECAR) of WT and KO sperm were measured using a SeahorseXF96 analyzer. i, k display quantitative data presented as mean ± SEM, with n = 5 biologically independent mice per group. P values were determined using a two-sided unpaired Student’s t-test.
We then analyzed the function of mitochondrial proteins using MitoCarta3.0_MitoPathways. The downregulated proteins predominantly enriched at OXPHOS and are the components of OXPHOS Complex (Supplementary Fig. 4e). This finding further underscores the critical role of ABHD10 in the mitochondrial respiratory chain complexes and oxidative phosphorylation. In addition to the significant impact on oxidative phosphorylation, the marked downregulation of Spermatogenesis-associated protein 19 (SPATA19) and glycerol kinase (GYKL1) is particularly noteworthy, as these proteins are known to regulate mitochondrial sheath formation during spermiogenesis18,19 (Supplementary Fig. 4e). Therefore, we speculate that these two proteins are important downstream targets of ABHD10.
To further confirm the reliability of our proteomics observation that ABHD10 primarily regulates oxidative phosphorylation-related proteins, we performed mitochondrial proteomics using WT and ABHD10 KO testes (Fig. 4d). This approach utilized isolated mitochondrial fractions that enriching for mitochondrial proteins, while effectively eliminating potential interference from cytoplasmic and nuclear components, thereby substantially improving both the sensitivity and specificity of mitochondrial protein detection. Analysis of mitochondrial proteomics revealed 76 significantly downregulated and 5 upregulated proteins (fold-change > 2 and p < 0.05) in Abhd10 KO mitochondria compared to controls (Fig. 4e). The MitoPathways analysis revealed that MitoPathways analysis revealed that among these downregulated proteins, 63.16% of them were enriched in OXPHOS and functioned as OXPHOS subunits or OXPHOS assembly factors (Fig. 4f, g). As previously described, in mitochondrial proteomics, we also detected the downregulation of SPATA19, a protein associated with the formation of the sperm mitochondrial sheath (Fig. 4g). Consistent with our proteomic analyses, western blot results demonstrate that the proteins of complex I (NDUFS4, NDUFB7), complex IV (COXIV, COX5A, COX6B2, and COX7C), and SPATA19 were significantly downregulated in KO testes. Notably, some proteins, such as COX6B2, were nearly undetectable in KO testes (Fig. 4h).
Interestingly, the signal intensity of COXIV in WT and KO spermatocytes was found to be comparable by immunofluorescence, whereas the signal intensity was significantly decreased in round spermatids (Supplementary Fig. 5a), suggesting an abnormality in the mitochondria of round spermatids. Subsequently, we examined the mitochondrial morphology during the early stages of spermiogenesis, specifically in round spermatids, using transmission electron microscopy. Interestingly, in WT round spermatids, nearly all mitochondria exhibited a condensed morphology, while in Abhd10 KO round spermatids, the mitochondria predominantly displayed intermediate and elongated forms (Supplementary Fig. 5b–d). These findings indicate that the defects in the mitochondrial sheath of mature sperm may be attributed to the poor quality of mitochondria at earlier developmental stages.
Given that ATP is produced during oxidative phosphorylation and reflect oxidative phosphorylation function in mitochondria, we detected the ATP levels in both testes and sperm and observed that the ATP concentration was significantly decreased in both testes and sperm (Fig. 4i). To further confirmed the reason of decreased ATP concentration, we measured the extracellular acidification rate (ECAR) and oxygen consumption rate (OCR) using Seahorse XF analyzers. Notably, Abhd10 KO sperm exhibited significantly decreased OCR alongside increased ECAR compared to WT controls (Fig. 4j, k), indicating a shift from OXPHOS to glycolytic metabolism. This metabolic reprogramming was further supported by elevated lactate levels, a glycolytic byproduct, in KO group (Supplementary Fig. 5e), suggesting that the reduction in mitochondrial ATP production is partially compensated by increased glycolytic flux. Together, these findings provide compelling evidence that the disruption of oxidative phosphorylation during spermatogenesis severely impairs ATP production in sperm.
ABHD10 is a mitochondrial matrix protein specifically expressed in testes
To investigate the molecular mechanism of ABHD10 during spermatogenesis, we first studied its localization in mammals. Due to the absence of effective antibodies against the mouse ABHD10 protein, we engineered HA-tagged Abhd10 knock-in (KI) mice. The “HA tag-linker” cassette was inserted at the N-terminus of ABHD10 to ensure that the HA tag labels all transcripts of ABHD10. Given that the gene contains a transit peptide at the N-terminus, which is cleaved post-translation, the “HA tag-linker” cassette was inserted following the mouse transit peptide (amino acids 1-43) (Fig. 5a–c). Western blot analysis of multiple mouse organs revealed that the protein ABHD10 was exclusively expressed in the testes (Supplementary Fig. 5f). Spermatogenesis progressed to spermatogonia, early spermatocytes, round spermatocytes, elongated spermatocytes, and spermatids at 1, 2, 3, 4, 5, and 8 weeks postnatal, respectively. Protein expression analysis indicated that ABHD10 was not detected in neonatal P0 mouse testes, but expression significantly increased 1 week after birth in the testes (Supplementary Fig. 5g). We next determined the subcellular localization of ABHD10 during spermatogenesis by immunofluorescence. ABHD10 was highly expressed in spermatocytes, and round and elongating spermatids, weakly expressed in spermatogonia, but not in sertoli cells in adult mouse testes (Supplementary Fig. 5h), which was further validated by immunohistochemistry (IHC) staining of ABHD10 (Fig. 5d). These results suggest that ABHD10 is primarily localized in the mitochondria of spermatocytes and round/elongating spermatids.
a–c Schematic illustration of the targeting strategy for constructing ABHD10 protein HA-tagged mice. Due to the transit peptide is cleaved off during protein processing, HA tag was inserted after the transit peptide (b). Briefly, HA tag were inserted into the exon 2 of ABHD10 gene (a). After transcription and translation, ABHD10 HA tagged protein was produced (c). Images of mouse were generated by SciDraw (credit Ethan Tyler and Lex Kravitz), licensed under CC BY 4.0 (https://creativecommons.org/licenses/by/4.0/). d IHC staining using testicular tissue showed that ABHD10 was expressed in the testis and localized in the cytoplasm of germ cells and flagella of sperm. Scale bar: 20 μm. e WB analyses revealed that ABHD10 was expressed only in mitochondria of testes. Nc nuclei, Mito mitochondria, Cp cytoplasm (mitochondria has been removed). f Schematic for isolating mitochondrial subcellular component proteins. Mitochondria contains four sub-compartments: outer membrane (MOM), inner membrane (MIM) intermembrane space (IMS) and matrix. After treatment with sodium carbonate, mitochondria can be separated as pellet (PEL) containing integral IMM and integral OMM and supernatant (SN) containing Matrix, IMS, peripheral lMM and peripheral OMM. Image of mitochondria were generated by figdraw.com. g WB analyses of proteins in different sub-compartments of mitochondria. Mito Mitochondria, SN supernatant, PEL pellet. h Subcellular localization of ABHD10 in mouse testes using immunogold staining, followed by TEM. Representative image of mitochondria in testis showed that the specific gold particles localized in the matrix of mitochondria. Scale bar: 1 μm. i Super-resolution images of mitochondria in HEK293T cells. Green signal indicates GFP-ABHD10 fusion protein. IMM was labeled with PK mito (Purple signal). The white arrowheads indicate swollen mitochondria. Scale bar: 10 μm. j Subcellular localization of ABHD10 in HEK293T cells using immunogold staining, followed by TEM. The FLAG antibody was utilized to detect the expression of ABHD10 proteins. Representative image of mitochondria in HEK293T cells demonstrated that ABHD10 was localized in the matrix of mitochondria. Scale bar: 1 μm. k Diagrams showing the protein region organization of ABHD10 fused with GFP, including full length, the N-terminus deletion (ΔN), AB hydrolase-1 domain deletion (ΔD), and the C-terminus deletion (ΔC). l Representative images of 293T cells transfected with GFP vector or GFP-tagged ABHD10 full length or different deletions (ΔN, ΔD, and ΔC). The white arrowhead indicates that loss of mitochondrial co-localization of ABHD10 upon deletion of its N-terminus. Scale bar: 5 μm. d, e, g, h, i, j, l show representative images from at least three independent experiments with similar results.
To more precisely define the subcellular localization of ABHD10 protein, we purified nucleus, mitochondria and cytoplasm (with mitochondria removed) from mouse testes for western blot analyses. Our data indicated that ABHD10 was specifically localized in the mitochondria of the testes (Fig. 5e). Mitochondria contain four sub-compartments: the outer membrane (MOM), the inner membrane (MIM), and two soluble compartments, the intermembrane space (IMS) and the matrix (Fig. 5f), We are curious what part of ABHD10 is present. To further characterize the localization of ABHD10, we treated mitochondria with sodium carbonate, which can break mitochondrial membrane, thereby separating the integral proteins in the pellet (PEL) and peripheral soluble proteins in the supernatant (SN) (Fig. 5f, g). Following this fractionation, ABHD10 was detected in the soluble phases (Fig. 5g), indicating that ABHD10 is a soluble protein rather than an integral protein of either the mitochondrial inner or outer membrane. Furthermore, we applied immunoelectron microscopy and confirmed that ABHD10 is localized in the mitochondrial matrix of testes (Fig. 5h).
To further validate the localization of ABHD10, we performed structured illumination microscopy (SIM) live cell imaging analysis to obtain super-resolution images. GFP-ABHD10 fusion proteins were transfected into COS-7 cells and ABHD10 protein was observed to be embedded within the gap of MIM (Fig. 5i). In addition, immunoelectron microscopy confirmed the mitochondrial matrix localization of ABHD10 (Fig. 5j). Interestingly, the overexpression of ABHD10 protein led to mitochondrial swelling, indicating that the expression levels of ABHD10 is critical for mitochondrial morphology.
Additionally, The formation of the sperm mitochondrial sheath relies on the precise control of mitochondrial aggregation19,20,21. Therefore, we overexpressed GFP-ABHD10 protein in 293T cells (Fig. 5k, l), but unfortunately, ABHD10 did not induce mitochondrial aggregation. This indicates that the abnormal morphology of the mitochondrial sheath in sperm is not caused by mitochondrial aggregation defects in ABHD10 knockout mice. Given that the functions of each domain of ABHD10 remain unknown, we generated ABHD10 mutants lacking the N-terminus, AB hydrolase-1 domain, and C-terminus separately and transiently expressed them in 293T cells (Fig. 5k). We then performed live cell staining to detect the colocalization of different truncated ABHD10 proteins with MitoTracker (Fig. 5l). The results showed that deletion of the N-terminus of ABHD10 prevents its localization to mitochondria, results in ABHD10 protein aggregate in the cytoplasm. Collectively, these data suggest that ABHD10 is a testis-specific mitochondrial matrix protein and its N-terminus is essential for its recruitment to mitochondria.
ABHD10 regulates the palmitoylation of mitochondrial sheath formation-associated proteins
To elucidate the regulatory role of ABHD10 in mitochondrial protein depalmitoylation, we conducted an acyl-biotin exchange (ABE) assay followed by mass spectrometry (Fig. 6a). As anticipated, palmitoylation of mitochondrial proteins was found to be dysregulated in ABHD10-deficient testes, with the majority of the affected proteins exhibiting increased levels of palmitoylation (Fig. 6b and Supplementary Data 3). The functions of these proteins, which showed elevated palmitoylation levels, are predominantly enriched in aerobic respiration, respiratory electron transport, mitochondrial gene expression, and nucleoside phosphate metabolic processes (Fig. 6c). To validate our proteomic findings, we selected proteins (SPATA19, GK2, PDHX, NDUFV1, SDHB, SUCLG1, and PMPCB) that exhibited significantly increased palmitoylation levels in KO mice for western blot analysis and this analysis confirmed elevated palmitoylation levels for all targets (Fig. 6d).
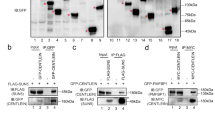
a Diagram of ABE assay used for the detection of protein acylation in WT and Abhd10 KO testes. NEM, N-ethylmaleimide. Images of mitochondria were generated by figdraw.com. Images of testis, equipment and mouse were generated by Servier Medical Art and SciDraw (credit Ethan Tyler and Lex Kravitz), licensed under CC BY 4.0 (https://creativecommons.org/licenses/by/4.0/). The drawing of testis is modified from an original provided by Servier Medical Art. b Volcano plots showing the differentially palmitoylated mitochondrial proteins in WT and KO testes. Two-sided unpaired Student’s t-test was used to calculate p-value. Only proteins meeting p < 0.05 and |FC| ≥ 2 were defined significantly differentially expressed. c GO enrichment of up-palmitoylated mitochondrial proteins in KO testes. d ABE assay of WT and KO testes followed by immunoblot to detect the palmitoylation levels of PDHX, PMPCB, SUCLG1, SDHB, SPATA19 and GK2. Western blot analysis in this panel was performed using the same set of biological samples. e, f Sperm from WT and KO mice are stained with anti-SPATA19 and anti-GK2 antibodies. The white arrow indicates the aggregation of SPATA19 and GK2 proteins. Scale bar: 5 μm. g The activity of pyruvate dehydrogenase (PDH) was significantly decreased in KO testes compared to controls. h, i Increased pyruvate levels and decreased acetyl-CoA levels were observed in KO testes. j The functional enrichment analyses of ABHD10 interacting proteins. Statistical significance (p-values) was determined based on the cumulative hypergeometric distribution. k Venn diagrams show the overlaps of up-regulated palmitoylated proteins and ABHD10-interacted proteins detected by IP-MS. The box on the right sides displays the overlapped 29 proteins. l Immunoprecipitation (IP) coupled with western blot were used to detect ABHD10 interacted proteins (SPATA19, GK2 and PDHX) in testes. m, n Western blot analysis of streptavidin pull-down samples, with or without biotin treatment. TurboID-ABHD10, GFP-GK2 and GFP-SPATA19 were transfected into 293T cells. GAPDH served as a negative control. g–i display quantitative data presented as mean ± SEM, with n = 3 biologically independent mice per group. P values were determined using a two-sided unpaired Student’s t-test. d, e, l–n) show representative images from at least three independent experiments with similar results.
Given the established role of palmitoylation in regulating protein stability, we conducted an integrative analysis of palmitoylation proteomics and mitochondrial proteomics datasets. This approach revealed mitochondrial sheath protein SPATA19 and electron transport chain components (NDUFA5, NDUFA8, and COX6B2) exhibited increased palmitoylation (FC < 0.5, P < 0.05) alongside decreased protein abundance (FC < 0.5, P < 0.05) (Supplementary Fig. 6a and Supplementary Data 2–3). These molecular alterations closely correlate with the observed mitochondrial sheath disorganization and OXPHOS impairment in Abhd10-deficient mice. To investigate whether the stability of other proteins with increased palmitoylation levels was affected, we analyzed proteomics sequencing data and performed western blotting. The results demonstrated that the protein levels of SPATA19, GK2, NDUFV1, and SUCLG1 were significantly reduced in KO testes (FC < 0.67, P < 0.05), PDHX and SDHB exhibited a moderate but statistically significant decrease (FC < 0.8, P < 0.05), whereas PMPCB expression remained unchanged (P > 0.05) (Supplementary Fig. 6b, c).
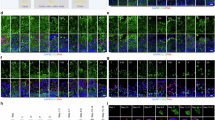
Notably, SPATA19 and GK2 have been reported to regulate the formation of mitochondrial sheaths and genetic knockout of either protein leads to a phenotype similar to that observed in Abhd10 knockout mice18,19. Therefore, SPATA19 and GK2 may serve as important downstream proteins of ABHD10 that regulate the formation of sperm mitochondrial sheaths. To further investigate the impact of Abhd10 knockout on SPATA19 and GK2, we performed immunofluorescence assays. The absence of ABHD10 resulted in a significant reduction in their signal intensity accompanied by the appearance of distinct punctate fluorescent aggregates within the mitochondrial sheath (Fig. 6e, f). To rule out the possibility that the aggregates was due to changes in mitochondrial arrangement, we stained a commonly used mitochondrial marker, TOMM20, and as expected, TOMM20 did not show significant aberrant aggregated signal in KO sperm (Supplementary Fig. 6d). These findings suggest that excessive hydrophobic palmitoylation of SPATA19 and GK2 may drive the formation of insoluble protein aggregates, which can subsequently induce protein degradation. This observation provides a plausible explanation for the observed morphological abnormalities of mitochondrial sheath.
Pyruvate dehydrogenase complex component X (PDHX) is the component of the pyruvate dehydrogenase complex (PDHc)22. To investigate the impact of elevated PDHX palmitoylation on mitochondria, we assessed the activity of PDHc and observed a decrease in its enzymatic activity (Fig. 6g). Additionally, we measured the levels of pyruvate and acetyl-CoA, which are the substrate and product of pyruvate dehydrogenase, respectively23. The results revealed an increase in pyruvate levels and a decrease in acetyl-CoA levels (Fig. 6h, i). Acetyl-CoA is the critical initial substrate for the tricarboxylic acid (TCA) cycle, its reduced levels could significantly impair the oxidative phosphorylation (OXPHOS) process and the knockdown of PDHX have been reported could significantly impair the oxidative phosphorylation process23,24. Thus, we speculate that ABHD10 may catalyze the S-depalmitoylation of PDHX, thereby stabilizing PDHX-mediated OXPHOS machinery. Given that SUCLG1 mutations are known to be associated with mtDNA depletion syndromes25,26, we assessed whether knockout of Abhd10 affected mtDNA levels in testes. Surprisingly, the mtDNA copy number in KO testes did not differ significantly from WT controls (Supplementary Fig. 6e), suggesting that SUCLG1 may function as a dispensable downstream effector of ABHD10.
To identify the direct targets of ABHD10 protein, we performed immunoprecipitation (IP) using testicular mitochondria from ABHD10-HA-tagged transgenic mice to identify candidates that interact with ABHD10 endogenously (Supplementary Data 4). We totally identified 136 ABHD10 interacted proteins and functional enriched analysis showed that these interacted proteins were enriched at organic acid catabolic process, acyl-CoA metabolic process, mitochondrion organization, mitochondrial protein degradation and protein localization to mitochondrion (Fig. 6j). Then, we compared the up-regulated palmitoylated proteins with the proteins that interact with ABHD10. A total of 29 proteins exist in both datasets was detected (Fig. 6k). Among these, pyruvate dehydrogenase complex component X (PDHX), mitochondrial sheath formation associated proteins SPATA19 and GK2 drawn our attention (Fig. 6k). Western blot validates a robust protein-protein interaction between ABHD10 and PDHX (Fig. 6l). Interestingly, we detected GK2 and SPATA19 in ABHD10 IP-MS, while these interactions were not detected in western blot (Fig. 6k, l and Supplementary Fig. 6f). Given that western blot detection is less sensitive than mass spectrometry, we hypothesized that ABHD10 might form weak/transient interactions with SPATA19 and GK2. To test this, we employed proximity labeling technology—a powerful enzymatic approach that enables covalent tagging of neighboring proteins, particularly effective for capturing transient and weak molecular interactions. As anticipated, both SPATA19 and GK2 were successfully biotinylated in these assays, confirming their physical association with ABHD10 (Fig. 6m, n). This strongly suggests that ABHD10 binds to and catalyzes the depalmitoylation of GK2 and SPATA19, thereby regulating their protein stability.
Additionally, since NDUFV1, SDHB, SUCLG1, and PMPCB also exhibited elevated palmitoylation levels, we employed both immunoprecipitation and proximity labeling technology to examine its potential interaction with ABHD10. The results showed that ABHD10 failed to bind NDUFV1, SDHB, SUCLG1 and PMPCB (Supplementary Fig. 6g, h), indicating that they are not a direct catalytic target of ABHD10-mediated depalmitoylation. Collectively, our findings demonstrate that SPATA19, GK2 and PDHX are essential ABHD10 substrates during spermatogenesis (Fig. 7).
Loss of ABHD10 impairs sperm motility that phenocopies human asthenozoospermia. The potential mechanism is that ABHD10 deficiency induces hyper-palmitoylation of key mitochondrial proteins (including SPATA19, GK2 and PDHX), leading to malformation of the mitochondrial sheath and disruption of oxidative phosphorylation (OXPHOS). Image of WT mitochondria were generated by figdraw.com.
Discussion
The role of palmitoylation during spermatogenesis
Among the three most prevalent lipid modifications, myristoylation, prenylation, and palmitoylation, only palmitoylation is reversible, regulated by “writers” (protein acyltransferases, PATs) and “erasers” (acyl protein thioesterases, APTs)12,13. In humans, there are 23 known PATs characterized by DHHC structural domains. In contrast, APTs have been studied even less extensively, with only eight identified, including PPT1/2 localized in the lysosome, APT1/2 in the cytoplasm, ABHD17A/B/C, and the mitochondrially-located ABHD1012,16. In recent years, palmitoylation has been demonstrated as an important protein modification that regulates stability, trafficking, localization and activity of proteins and is crucial for various biological processes12,13, garnering considerable attention from researchers. However, there is currently limited research on the role of palmitoylation during mammalian spermatogenesis compared to other fields. Although ZDHHC19 have been reported to be specifically expressed in testis and is essential for male fertility in mice27, the function of depalmitoylation during spermatogenesis has not been investigated. ABHD10 recently was identified as a S-depalmitoylase and regulates redox homeostasis via PRDX5 in vitro28, however, its physiological function in vivo remains unknown. In this study, we demonstrate S-depalmitoylase ABHD10 as a critical regulator of sperm mitochondrial sheath formation and male fertility. Although studies concerning palmitoylation modifications during spermatogenesis were rare, we believe that in the future palmitoylation modifications will ushered an explosive development.
Comparison with reported proteins associated with sperm mitochondrial sheath formation
The genes reported to be associated with mitochondrial sheath integrity were summarized in Supplementary Fig. 3a. Given the similar phenotypes, we sought to investigate the relationship between these proteins and ABHD10. We considered whether any of these proteins are downstream of ABHD10. Proteomic analysis unveiled that the protein expression levels of Spata19, Gk2 and Gykl1 were downregulated in Abhd10 knockout testes. Interestingly, SPATA19 was a mitochondrial protein and 100% of Spata19 knockout sperm exhibited disorganized mitochondrial structure18. The integrity of the mitochondrial respiratory chain complex and ATP production were similarly disrupted, mirroring the observations in Abhd10 knockout testes. More importantly, the palmitoylation levels of SPATA19 and GK2 were significantly increased. Therefore, we hypothesized that SPATA19 and GK2 are key downstream proteins of ABHD10.
In addition, recently Mise et al. reported that the absence of mitochondrial sheath in Vdac3 KO sperm tends to occur in mitochondria located near the neck and annulus28. In contrast to the phenotype observed in knockout mice of the Vdac3 gene, we did not observe a positional bias in the loss of mitochondrial sheaths in the spermatozoa of Abhd10 knockout mice. This finding further confirms that the function of ABHD10 in mitochondria is independent of VDAC3. Interestingly, overexpression of ARMC12 has been reported to lead to the aggregation of mitochondria20. However, this phenomenon was not observed in cells overexpressing ABHD10, suggesting that ABHD10 functions independently of ARMC12.
The mechanism of deformed mitochondrial sheath
Our findings demonstrate that ABHD10 deficiency leads to hyperpalmitoylation and destabilization of mitochondrial sheath proteins SPATA19 and GK2 - two crucial regulators of mitochondrial sheath formation whose deficiency phenocopies our ABHD10 knockout observations19,21. This molecular dysregulation provides a direct explanation for the deformed mitochondrial sheath observed in Abhd10 knockout sperm. While SPATA19/GK2 dysregulation may be the primary defect, we cannot exclude potential secondary contributions from OXPHOS impairment that may further compromise mitochondrial membrane integrity and morphology. Together, these findings establish ABHD10-mediated depalmitoylation as a critical regulator of mitochondrial sheath formation through its effects on key structural proteins.
The cause of damaged OXPHOS
In this study, we observed significant mitochondrial dysfunction characterized by reduced oxygen consumption, decreased OXPHOS protein levels, and impaired ATP production. This metabolic disorder may through three distinct yet interconnected mechanisms. First, ABHD10-mediated depalmitoylation of PDHX is essential for maintaining pyruvate dehydrogenase activity, and its impairment leads to reduced acetyl-CoA production, thereby limiting TCA cycle flux and subsequent oxidative phosphorylation. Second, the deformed mitochondrial sheath causes structural damage and electron transport chain component loss, thus compromising OXPHOS capacity. Third, elevated palmitoylation of ETC proteins affects OXPHOS. However, the absence of detectable physical interaction between ABHD10 and ETC components suggests this regulation may occur through indirect mechanisms that warrant further investigation. Collectively, these findings position ABHD10 as a master regulator that orchestrates both metabolic flux and structural integrity to sustain mitochondrial energy production.
The substrates of ABHD10
Comparative analysis of ABHD10-interaction proteins and differential palmitoylation proteins revealed that 29 of interacting proteins exhibited significant alterations in palmitoylation levels. Among these candidates, three proteins—SPATA19, GK2, and PDHX—emerged as particularly noteworthy: SPATA19 and GK2 based on their established roles in mitochondrial sheath formation18,21, and PDHX due to its critical function in OXPHOS24. Based on these findings, we propose that SPATA19, GK2, and PDHX represent key catalytic substrates of ABHD10’s depalmitoylase activity. While we consider these three proteins to be critical targets based on their functional relevance and palmitoylation dynamics, we acknowledge the possibility that additional substrates may exist beyond our current detection limits due to technical constraints in protein-protein interaction assays and palmitoylation proteomics coverage. Regarding the impact of excessive palmitoylation on proteins, we observed that the absence of ABHD10 led to abnormal protein aggregation of SPATA19 and GK2, thus we hypothesize that excessive palmitoylation increases protein hydrophobicity, leading to abnormal aggregation and reduced stability. In addition, given the well-established role of SUCLG1 in maintaining mitochondrial integrity25,26, we evaluate mtDNA levels in our experimental model. Surprisingly, our results revealed that ABHD10 knockout did not significantly alter mtDNA copy number. One possible explanation for this discrepancy is that the ABHD10-mediated depalmitoylation of SUCLG1 may not be sufficiently robust to perturb its functional capacity. Another explanation could be that compensatory pathways in germ cells may effectively buffer against this disruption.
Together, these findings provide a possible mechanism for ABHD10 as a critical regulator of both mitochondrial structure (via SPATA19/GK2) and function (via PDHX).
The decreased expression of mitochondrial proteins may result from various possibilities, such as changes in mitochondrial import, protein translation, protein degradation, or mitochondrial biogenesis. Since we observed reduced protein levels of SPATA19, GK2, PDHX alongside elevated palmitoylation levels, we hypothesize that excessive palmitoylation of SPATA19, GK2, and PDHX may lead to their protein degradation. These abnormalities can lead to structural and functional abnormities of mitochondria, thereby causing dysregulation of proteins throughout the mitochondria. However, whether additional mechanisms are involved in the dysregulation of mitochondrial protein expression warrants further investigation.
Mitochondrial sheath maintains the integrity of 9 + 2 structure
Mitochondrial sheaths have multi-faceted function. Sperm have a strong ability to continuously move and this activity consumes substantial amounts of adenosine triphosphate (ATP)3. The sperm mitochondrial sheath generates ATP through oxidative phosphorylation, providing the necessary energy for flagellar movement, which is vital for normal sperm activity and effective swimming. In addition, the mitochondrial sheath surrounds the axonemal complex and the nine outer dense fibers at the midpiece to maintain its proper alignment and structural integrity. This structural stability is essential for proper sperm function and motility29. Interestingly, our study observed that the sperm mitochondrial sheath contributes to the structural stability of 9 + 2 microtubular arrangement. In Abhd10 KO mice the “9 + 2” axonemal structure is intact in the immobile sperm of testes, whereas a disarrangement of “9 + 2” axonemal structure is observed in motile epididymal sperm, suggesting that the formation of “9 + 2” axonemal structure during spermatogenesis is normal, but it is disrupted during the movement of sperm.
Methods
Ethics statement
All mice were housed in a pathogen-free facility at the Laboratory Animal Center of Sir Run Run Shaw Hospital, Zhejiang University School of Medicine. All animal procedures were conducted in accordance with the guidelines established by the Animal Care and Research Committee of Sir Run Run Shaw Hospital, Zhejiang University School of Medicine (approval number: SRRSH2025-0028). All experiments involving human participants were approved by the Ethics Committees of Sir Run Run Saw Hospital, Zhejiang University School of Medicine (approval number 2022-726-01).
Mice
The Abhd10 knockout (KO) mice were generated using CRISPR/Cas9-mediated genome editing technology. The T7 promoter along with a guide sequence designed to target exon 2 of the Abhd10 gene was employed, using a pair of single-guide RNAs (sgRNAs). The sgRNAs were amplified via PCR using the following primers: Abhd10-sgRNA-Forward: 5′-GTTGGGAGGTAATGTCTGAA-3′ and Abhd10-sgRNA-Reverse: 5′-TTCAGCCACTGGGCTTTTAC-3′. Cas9 mRNA (20 ng) and sgRNA (10 ng) were mixed and injected into C57BL/6 zygotes, which were subsequently implanted into pseudo-pregnant C57BL/6 females. Genomic DNA was extracted from founder mice, followed by Sanger sequencing confirmation and PCR analysis. Founder heterozygous mice were backcrossed to wild-type (WT) C57BL/6 mice to obtain F1 heterozygous mice, and F2 generation mice were genotyped and intercrossed to produce Abhd10 KO mice for experiments.
The Abhd10 transgenic knock in mice were generated using our previously established methods30. Briefly, the endogenous Abhd10 promoter was amplified from mouse genomic DNA and used to replace the CMV promoter in the pmCherry-N1 vector (Clontech, 632523). Subsequently, Abhd10 cDNA, was cloned from total RNA isolated from adult mouse testes and cloned into the modified pmCherry-N1 vector. DNA fragment containing Abhd10 promoter and cDNA was microinjected into the pronuclei of one-cell zygotes to generate founder mice. Genotyping of the mice was conducted using PCR amplification of genomic DNA extracted from mouse tails.
To generate Abhd10-HA-tagged transgenic mice, the “HA tag-linker” cassette was inserted downstream of the eighth amino acid following the mouse transit peptide (aa.1-43) of ABHD10. To engineer the donor vector, homology arms were generated by PCR using BAC clone RP23-89A17 as a template. Ribonucleoprotein (RNP) was injected into fertilized C57BL/6 J mouse oocytes along with the donor vector for mouse production. F0 founder mice were identified through PCR followed by Sanger sequencing analysis, and were subsequently mated with wild-type C57BL/6J mice to produce the F1 generation. The primers utilized for genotyping are listed in Supplementary Table 1.
All mice used in this study were C57BL/6J strains. Male mice aged 2–3 months were used in this study, unless otherwise specified in figure legends or methods. All animals were housed in a pathogen-free facility under a 12-h light/dark cycle, with ambient temperature maintained at 23 ± 1 °C and relative humidity at 55 ± 5%. They were provided with food and water ad libitum and kept on clean bedding, which was replaced weekly.
Mitochondrial proteome data analysis
Mitochondrial protein expression profiles across 14 tissues were obtained from mouse MitoCarta3.0 (https://personal.broadinstitute.org/scalvo/MitoCarta3.0/mouse.mitocarta3.0.html). To identify tissue-biased and tissue-specific mitochondrial proteins, we calculated the Specificity Measure (SPM) score for each protein based on its expression across 14 major tissues. Proteins with SPM > 0.5 were classified as exhibiting highly biased expression in a single tissue, while those with SPM = 1 were considered exclusively tissue-specific. The resulting high-confidence tissue-biased and tissue-specific mitochondrial proteomes were then subjected to hierarchical clustering analysis (Fig. 1a) and further statistical analysis (Fig. 1b–d).
Total RNA extraction and RT-qPCR
Total testicular RNA was isolated using Trizol (Invitrogen, 15596-018) according to the manufacturer’s instructions. Complementary DNA (cDNA) was transcribed from total testicular RNA (800 ng) using PrimeScript RT Master Mix (TaKaRa, RR047A). Real-time quantitative PCR was performed using a two-step PCR program on CFX96 Real-Time PCR Detection System (Bio-Rad, CFX96). The primers employed in the experiment are listed in Supplementary Table 1.
Histological analysis
Testes and epididymides were dissected and fixed overnight at 4 °C in Bouin’s solution (Sigma, HT10132). Following fixation, samples were dehydrated through a graded ethanol series (70%, 80%, 90%, and 100%), cleared in xylene, and embedded in paraffin. Serial sections (5 μm thick) were prepared using a microtome. For periodic acid-Schiff (PAS) staining, sections were deparaffinized in xylene and rehydrated through descending ethanol concentrations. The staining procedure was performed using the PAS Staining Kit (Solarbio, G1281) according to manufacturer’s protocol. Briefly, sections were oxidized with 0.5% periodic acid solution for 10 min at room temperature in the dark, then thoroughly rinsed with distilled water. Schiff’s reagent was applied for 30–60 min at 37 °C in a humidified chamber. After washing, sections were counterstained with hematoxylin for 30 s. Following dehydration through graded alcohols and xylene clearing, sections were mounted with neutral balsam (Solarbio, G8590). Hematoxylin and Eosin (H&E) Staining was performed using a Modified Hematoxylin-Eosin Staining Kit (Solarbio, G1121) according to the manufacturer’s protocol. Following deparaffinization in xylene and hydration through a graded ethanol series, sections were stained with Hematoxylin Solution, briefly differentiated, and blued. Cytoplasmic counterstaining was carried out using Eosin Solution. After staining, sections were dehydrated through graded alcohols, cleared in xylene, and mounted with neutral balsam. All slides were examined under a light microscope (Olympus BX43) and images were captured using Olympus imaging software.
TUNEL analyses
Testicular sections were deparaffinized in xylene and rehydrated through descending ethanol concentrations. Subsequently, tissue sections were treated with 20 μg/mL proteinase K (Beyotime, ST533) for 20 min at 37 °C to expose DNA breaks. Endogenous peroxidases were blocked with 3% H₂O₂ for 20 min. The TUNEL reaction mix (Beyotime, C1091) was prepared by combining TdT enzyme and Biotin-dUTP (1:9 ratio) and applied to sections for 60 min at 37 °C in a humidified chamber. Streptavidin-HRP (1:50 dilution) was then added for 30 min, followed by DAB substrate incubation (5–30 min). Nuclei were counterstained with hematoxylin. Apoptotic cells (brown DAB signal) were imaged using an Olympus BX43 microscope. Negative controls omitted TdT enzyme, while DNase I-treated sections served as positive controls.
CASA
Epididymides were harvested and subsequently dissected into several smaller pieces. The tissue samples were then incubated in pre-warmed TYH medium (Nanjing Aibei Biotechnology Co., Ltd, M2035) at 37 °C for 30 min to facilitate the release of spermatozoa. Sperm motility and concentration were analyzed using computer-assisted sperm analysis (CASA) (Version 6.3, Microptic, Barcelona, Spain), with a minimum of 500 spermatozoa counted per mouse. Dual-sided chamber slides (with a depth of 20 μm, Goldcyto, SCA 20-04-01) were employed for dispensing sperm suspension in computer-assisted sperm analysis (CASA).
In vitro fertilization
Twenty-one-day-old female mice were injected with 5 IU of PMSG. After 44 h post-PMSG injection, the mice received a second injection of 5 IU of hCG. Sixteen hours later, the mice were euthanized, and their oviducts were collected. Oocyte–cumulus complexes were released from the oviducts using needles and subsequently transferred to HTF medium. Next, 10 μL of capacitated sperm in TYH medium (Nanjing Aibei Biotechnology, M2035) was added to the HTF drops (Nanjing Aibei Biotechnology, M1135) containing the oocyte–cumulus complexes. After 6 h of incubation, the zygotes were washed with HTF medium to remove sperm and granule cells. The zygotes were then transferred to KSOM mini-drops (Nanjing Aibei Biotechnology, M1435). Embryos were photographed at 6, 24, 48, 72, and 96 h post in vitro fertilization.
Scanning electron microscopy
Spermatozoa were released from the epididymides of mice and smeared onto coverslips that had been pre-coated with 3-Aminopropyltriethoxysilane (Solarbio, A7440). Then sperm-attached coverslips were fixed in 2.5% glutaraldehyde solution at room temperature for 2 h, followed by overnight fixation at 4 °C. The samples were rinsed three times with 0.1 M PB (pH 7.4) for 10 min each. Subsequently, the samples were fixed in 1% osmium tetroxide solution for 1.5 h and rinsed again three times with 0.1 M PB (pH 7.4) for 10 min each. Dehydration of the samples was performed using graded ethanol solutions (50%, 70%, 90%), each for 15 min, followed by two treatments with 100% ethanol for 20 min each. The samples were then subjected to critical point drying, coated with a conductive layer and imaged with an S-3400N scanning electron microscope (Hitachi, Tokyo, Japan).
Transmission electron microscopy
Spermatozoa and testes from mice were initially preserved in a 2.5% glutaraldehyde solution at 4 °C overnight. The samples were then washed three times with 0.1 M PB (pH 7.4) for 15 min each. The specimens were then treated with 1% osmium tetroxide for 1.5 h and subsequently washed three times with 0.1 M PB (pH 7.4) for 15 min each. Dehydration was performed using a graded series of ethanol solutions (50%, 70% and 90%) for 30 min each, followed by two 30-min immersions in 100% ethanol, a 30-min treatment with a 1:1 ethanol-acetone mixture, and two 30-min immersions in pure acetone. The specimens were then infiltrated with mixtures of epon resin and acetone (1:1 for 2 h, 3:1 overnight), embedded, and polymerized at 65 °C for 24–48 h. Ultrathin sections (60 nm) were prepared and stained with uranyl acetate and lead citrate. Ultrastructural analysis was carried out using a Tecnai 10 or 12 microscope (Philips) at 120 kV.
Immuno-electron microscopy
In the process of immunogold-based electron microscopy (Ig-EM) for mouse testis and 293T cells, the following steps were performed. First, the samples were fixed in a solution containing 2.5% glutaraldehyde and 3% paraformaldehyde within a 0.1 M cacodylate buffer (pH 7.4) at 4 °C for 8 h. Following this, the samples were washed in 0.1 M PB overnight. Subsequently, the samples underwent dehydration through a series of graded ethanol solutions. They were then embedded in acrylic resin and polymerized at 53 °C for 24 h. Ultrathin sections with a thickness ranging from 60 to 70 nm were prepared. These sections were immunostained overnight at 4 °C using either rabbit anti-HA or anti-FLAG antibodies. This was followed by a 1-h incubation with 10 nm colloidal gold-conjugated goat anti-rabbit or goat anti-mouse IgG. Finally, the immunostained sections were imaged using a JEM-1400 transmission electron microscope (manufactured by JEOL).
Cell culture and plasmids construction
HEK-293T (Procell, CL-0005) and COS-7 (Procell, CM-0069) cells were cultured in DMEM mediums supplemented with 10% (v/v) fetal bovine serum and 1% penicillin–streptomycin at 37 °C with 5% CO2. For cell passaging, aspirate the spent medium from the cell culture dish (Bioland, CCD06) and gently rinse the adherent cell layer with PBS. Add pre-warmed trypsin-EDTA to dissociate the cells and incubate briefly at 37 °C. Once detached, neutralize trypsin with Fetal bovine serum. Transfer the cell suspension to a 15 mL centrifuge tube (Bioland, ATS05-15) and centrifuge at 1000 rpm for 3 min. After centrifugation, carefully aspirate the supernatant and resuspend the cell pellet in fresh complete medium. For Cryopreservation, resuspend the cell pellet in Serum-free Cell Freezing Medium (Solarbio, C2950). Aliquot the cell suspension into cryovials (Bioland, AC05-20) and transfer them to a −80 °C freezer for storage.
Full-length mouse Abhd10, Armc12, Ddhd1, Gk2, Klc3, Mff, Spata19, Tbc1d21, and Vdac3 cDNA were obtained from mouse testis cDNA, which were then cloned into the vectors pCMV-C-Flag, pEGFP-N1 or modified C1(1-29)-TurboID-V5 (Addgene, 107173). All construction of plasmids was performed using Clon Express Ultra One Step Cloning Kit (Vazyme, C115). Each required tagged protein expression construct was transfected in HEK-293T or COS-7 cells with QuickShuttle-293 (Biodragon, KX0110044) according to the manufacturer’s protocol. Cells were used for experiments after 48 h transfection.
Immuno-precipitation
Testicular mitochondria or HEK293T cells were homogenized in IP buffer (Beyotime, P0013) supplemented with complete protease inhibitor cocktail (Roche, 11697498001). The lysates were incubated for 30 min at 4 °C, followed by centrifugation at 15,000 × g for 15 min to clarify the supernatant. The cleared supernatant was then incubated with 50 µl of pre-cross-linked antibody-conjugated magnetic beads (MCE, HY-K0202) overnight at 4 °C with gentle rotation. The bead-protein complexes were washed five times with cold IP buffer. For immunoblotting analysis, bound proteins were eluted using 1× SDS loading buffer. For mass spectrometry (MS) analysis, proteins were eluted with SDT buffer (4% SDS, 100 mM DTT, 100 mM Tris-HCl). MS samples were processed by tryptic digestion as previously described, and the resulting peptides were analyzed using a Q-Exactive Plus mass spectrometer (Thermo Fisher Scientific).
Immunohistochemistry
Paraffin-embedded testicular sections (5 μm) were deparaffinized in xylene and rehydrated through graded ethanol. Antigen retrieval was performed in citrate buffer (pH 6.0) at 95 °C for 20 min. Endogenous peroxidase activity was blocked with 3% H₂O₂ for 15 min, followed by 5% BSA blocking for 30 min at RT. Sections were incubated overnight at 4 °C with primary antibodies, then with HRP-conjugated secondary antibodies (1:500) for 1 h at RT. DAB substrate was applied for 1–5 min for signal development, and nuclei were counterstained with hematoxylin. Negative controls omitted primary antibodies. Slides were dehydrated, cleared in xylene, and mounted. Images were captured using an Olympus BX43 microscope. For antibody information, see Supplementary Table 2.
Immunofluorescence
Mouse testes were fixed in 4% paraformaldehyde (PFA) overnight, embedded in paraffin, and sectioned to a thickness of 8 μm. The sections were rehydrated through graded ethanol, subjected to antigen retrieval, and permeabilized in PBS containing 0.1% Triton X-100 for 30 min. Following a 1-h blocking step with 10% donkey serum (GeneTex, GTX30972) at room temperature, sections were incubated with the primary antibody overnight at 4 °C in a humidified chamber. After three washes with PBST, sections were incubated with an Alexa Fluor-conjugated secondary antibody for 1 h at room temperature. Following another three PBST washes, the sections were incubated with Hoechst 33342 (Invitrogen, H3570) for 5 min at room temperature, washed with PBST again, and then mounted with antifade mounting medium (Invitrogen, P36970). For the immunostaining of MitoTracker, 293T cells or sperm were incubated with MitoTracker Red CMXRos (Beyotime, C1049B) for 15 min at 37 °C. After incubation, the samples were washed three times with Hanks’ Balanced Salt Solution (with Ca2+ & Mg2+) (Beyotime, C0219). Images were captured using a Zeiss LSM900. For antibody information, see Supplementary Table 2.
HIS-SIM imaging
Super-resolution imaging of mitochondria was conducted using HIS-SIM (High Intelligent and Sensitive SIM) provided by Guangzhou CSR Biotech Co. Ltd. Images were acquired using a 100×/1.5 NA oil immersion objective (Olympus). Cells were placed in a 35 mm glass bottom dish (Cellvis, D35-20-1.5-N) and maintained at 37 °C with 5% CO2 in a humidified chamber for live SIM imaging. Image data were collected and analyzed following the procedures described previously31. To enhance image quality, sparse deconvolution was applied as outlined in prior studies32.
Mitochondrial isolation and purification
Mitochondria were isolated from the mouse brain, heart, liver, and testis using the Mitochondria Isolation and Protein Extraction Kit (Proteintech, PK10016). The tissues were harvested from the mice and homogenized on ice in Isolation Reagent A at a 1:10 (w/v) ratio, utilizing a Dounce homogenizer. The homogenization process involved five strokes at a moderate speed. Subsequently, the homogenate was carefully layered onto an equal volume of Isolation Reagent B and centrifuged at 700 × g for 10 min at 4 °C. The resulting supernatant was transferred to a new Eppendorf tube and subjected to further centrifugation at 10,000 × g for 10 min at 4 °C, with the aim of collecting the crude mitochondrial pellet. To further purify the mitochondria, the pellet was resuspended in wash buffer. A separation solution was prepared by mixing Mitochondria Isolation Buffers C and D in a 17:3 ratio. The resuspended crude mitochondrial fraction was carefully layered on top of the separation solution, followed by centrifugation at 22,000 × g for 10 min at 4 °C to isolate the mitochondria. The isolated mitochondria were then resuspended in 1 ml of wash buffer and centrifuged at 16,000 × g for 5 min at 4 °C to eliminate any remaining impurities. The final pellet obtained represents the highly purified mitochondria.
The isolation of mitochondrial subcompartment protein
The mitochondrial pellet was resuspended in 30 µL of Na₂CO₃ (100 mM) solution and incubated on ice for 30 min. After adding 1 µL of PMSF (100 mM), the suspension was transferred to specialized centrifuge tubes of the ultra-speed TLA-110 rotor and centrifuged at 75,000 × g for 40 min at 4 °C in a Beckman MAX-XP. After centrifugation, the supernatant (S) was carefully collected and transferred to a new tube. Subsequently, the pellet was washed with 100 µL of 1X TD buffer (49.99 mM Trizma base, 274.13 mM NaCl, 20.12 mM KCl, 13.95 mM Na2HPO4), followed by centrifugation at 20,000 × g for 5 min at 4 °C. This washing step was repeated twice to ensure the desired levels of purity. Subsequently, the pellet (P) was resuspended in 20 µL of 1X TD buffer and stored on ice until further analysis.
BCA assay
All procedures were performed using an enhanced BCA (bicinchoninic acid) protein assay kit (Beyotime, P0012). A stock solution with a concentration of 0.5 mg/mL was prepared and diluted with PBS to prepare standard solutions with final concentrations of 0.025, 0.05, 0.1, 0.2, 0.3, and 0.4 mg/mL. The working reagent was prepared by mixing Reagent A and Reagent B in a 50:1 ratio. Then, 200 μL of this working reagent was added to each sample. Protein samples were diluted with lysis buffer to a final volume of 20 μL, using one-tenth of the original sample volume. The samples and working reagent were incubated together at 37 °C for 30 min. After incubation, absorbance was measured at 562 nm using a SpectraMax ABS Microplate Reader (Molecular Devices). A standard curve was plotted based on the absorbance readings of the standard solutions, and the concentrations of the protein samples were calculated accordingly.
Protein extract and western blot
Mouse tissues were lysed in RIPA Lysis Buffer (Beyotime, P0013B) supplemented with a protease inhibitor cocktail (Roche) at a ratio of 1:10 (w/v). The tissues were homogenized using a tissue grinder and then sonicated for 15 cycles, each consisting of 2 s on and 2.5 s off. The homogenates were centrifuged at 12,000 × g for 15 min at 4 °C, and the supernatants were collected. The supernatants were mixed with 5× loading buffer (X-Blot, Xblot-0025) and denatured at 100 °C for 10 min. Protein samples were separated by 12.5% SDS-PAGE and transferred to PVDF membranes using a Bio-Rad blotting system. The membranes were blocked with 5% nonfat milk in TBS containing 0.1% Tween 20 (TBST) for 1 h at room temperature. After blocking, the membranes were incubated with primary antibodies overnight at 4 °C. The membranes were then washed three times with TBST and incubated with HRP-conjugated secondary antibodies for 1 h at room temperature. Western blots were developed using the ChemiDoc MP imaging system (Bio-Rad).
Protein preparation and mass spectrometry analysis
Protein samples were homogenized using SDT lysis buffer containing 4% SDS, 100 mM DTT, and 100 mM Tris-HCl (pH 8.0). Samples were boiled for 3 min and further ultra-sonicated. To inhibit reduced cysteine residues, detergent, DTT, and IAA in UA buffer were added. The protein suspension was then subjected to overnight digestion with trypsin (Promega) at 37 °C. Following digestion, the peptide mixtures were desalted using C18 StageTip prior to LC-MS analysis.
LC-MS/MS analysis was performed with an Orbitrap Astral mass spectrometer coupled with a Vanquish Neo UHPLC system (Thermo Fisher Scientific). The eluted peptides were analyzed on Orbitrap Astral mass spectrometer. DIA approach involved a survey scan in the range of 380–980 m/z with a resolution of 240,000, an AGC target of 500%, and an injection time of 5 ms. DIA MS/MS scans were acquired within the 150–2000 m/z range, utilizing a 2 m/z isolation window, with an AGC target of 500% and an injection time of 3 ms. The normalized collision energy was set to 25, and the cycle time was 0.6 seconds. Full MS and DIA scan spectra were recorded in profile and centroid modes, respectively.
DIA MS data were analyzed using DIA-NN 1.8.133,34. Database searches were conducted with results filtered to achieve a false discovery rate (FDR) of less than 1% at both peptide-spectrum-matched (PSM) and protein levels. Bioinformatics analyses were performed using Microsoft Excel and R statistical software, with hierarchical clustering and volcano plot generation executed in R. Differentially expressed proteins were identified based on both statistical significance and magnitude of change. A two-sided unpaired Student’s t-test was used to assess differences between experimental groups, and proteins with a p < 0.05 were considered statistically significant. Additionally, a fold change (FC) threshold was applied. Only proteins meeting both criteria were deemed significantly differentially expressed.
Acyl-biotin exchange (ABE) assays
First, WT and KO mitochondria were isolated from 40 WT and 40 KO testes. The isolated mitochondria were suspended in 1 ml of lysis buffer containing 50 mM Tris-HCl (pH 7.4), 150 mM NaCl, 5 mM EDTA, 1 % Triton X-100, and protease inhibitors, and subject to end over-end rotation 1 h at 4 °C. Centrifugation was performed at 15,000 × g for 15 min at 4 °C to pellet the cell debris, and subsequently, the supernatant was collected and precipitated by chloroform-methanol treatment. The resulting pellet was dissolved by 4% SDS buffer (4% SDS, 50 mM Tris-HCl, 5 mM EDTA, pH 7.4) containing 10 mM NEM at 37 °C for 10 min.
Then, Block buffer (150 mM NaCl, 50 mM Tris-HCl, 5 mM EDTA, 20 mM NEM, 0.2% Triton X-100, pH 7.4) containing protease inhibitors was added to each tube, and incubated with end-over-end rotation at 4 °C overnight. After incubation, the samples were subjected to three sequential CM precipitations to remove NEM. For the third time, the resulting pellet was dissolved with 4% SDS buffer and divided equally into two parts, which were then added to two tubes labeled as the +HAM tube and the -HAM tube, respectively. +HAM buffer (0.7 M hydroxylamine, 1 mM HPDP-biotin, 0.2% Triton X-100, pH 7.4) was added to the +HAM tube and -HAM buffer (50 mM Tris-HCl, 1 mM HPDP-biotin, 0.2% Triton X-100, pH 7.4) was added to the -HAM tube. The tubes were incubated for 1 h at RT. After this incubation, samples were precipitated by CM, dissolved and incubated with low-HPDP-biotin buffer (150 mM NaCl, 50 mM Tris-HCl, 0.2 mM HPDP-biotin, 0.2% Triton X-100, pH 7.4) for 1 h at RT.
For WB analysis, after this incubation, the samples were subjected to three sequential CM precipitations, and the pellet was dissolved with 2% SDS buffer (2% SDS, 50 mM Tris-HCl, 5 mM EDTA, pH 7.4) for the third time. The solution was diluted with 0.2% Triton X-100 solution to 20-fold its original volume and subjected to end-over-end rotation for 30 min at RT. Centrifugation was carried out at 15,000 × g for 1 min, and the supernatant was collected and incubated with the washed Pierce Streptavidin Magnetic Beads (Thermo Fisher, 88816) overnight at 4 °C. After being washed by washing buffer (150 mM NaCl, 50 mM Tris-HCl, 5 mM EDTA, 0.1% SDS, 0.2% Triton X-100, pH 7.4) four times, beads were heated with 1X loading buffer and analyzed by WB.
For mass spectrometry, after HPDP-biotin incubation, the peptides were re-dissolved in 8 M UA solution, and trypsin was digested overnight. The digested peptides were desalted using a C18 Cartridge, and an equal amount of biotin-labeled peptides was enriched using Streptavidin Magnetic Beads, and the Streptavidin Magnetic Beads were eluted several times to remove non-specific enriched peptides, and finally palmitoylated peptides were enriched using TCEP elution; IAA (20 mM) was added and incubated at room temperature for 1 h. After drying, the sample was desalted, concentrated in vacuum, and dissolved by adding 10 µL of 0.1% FA for mass spectrometry.
ATP detection
Mouse testes were lysed in ten volumes of lysis buffer according to the protocol provided by the ATP Assay Kit (Beyotime, S0027). The homogenized tissues were then centrifuged at 12,000 rpm for 15 min at 4 °C to obtain the supernatant. ATP standard solutions were prepared by diluting the ATP stock solution with ATP assay lysis buffer to achieve final concentrations of 0.01, 0.03, 0.1, 0.3, 1, 3, and 10 μM. The working reagents were allowed to equilibrate at room temperature for 3 min to eliminate background ATP. Following this, the supernatants were added to the working reagent. Chemiluminescence was recorded at 2-s intervals using a SpectraMax M5 microplate reader. Protein concentration was measured using a BCA Protein Assay Kit (Beyotime, P0012S). The ATP concentration was converted to nanomoles per milligram of protein.
OCR and ECAR assessment
Spermatozoa oxygen consumption rate (OCR) and extracellular acidification rate (ECAR) were measured using a Seahorse XFe96 Extracellular Flux Analyzer (Agilent) with minor modifications to previously described protocols35,36. The XF96 plastic microplate was pre-coated with 0.5 mg/mL concanavalin A (Sigma, C5275) in PBS for 1 h at 37 °C, followed by washing with ddH₂O and air-drying overnight at room temperature (RT). Spermatozoa were collected from the caudal epididymides and released into Seahorse XF DMEM Medium (Agilent, 103575-100) for 10 min at 37 °C. The sperm suspension was then centrifuged at 2000 rpm for 3 min, and the pellet was resuspended in pre-warmed Seahorse assay medium (Seahorse XF DMEM Medium supplemented with 10 mM glucose, 1 mM pyruvate, and 2 mM L-glutamine) to achieve a final concentration of 10⁶ sperm per 50 µL. 50 µL sperm suspension was loaded into each well of the pre-coated XF96 microplate. To enhance sperm adhesion, the plate was subjected to two sequential centrifugation steps (2000 rpm, 2 min each) in forward and reverse orientations. Subsequently, 130 µL of pre-warmed assay medium was gently added along the sidewall of each well to avoid disturbing the adhered spermatozoa. OCR and ECAR were measured using the Seahorse XFe96 Analyzer (Agilent) with 1 min-mixing and 2 min-measurement cycles.
Measurement of lactate concentration
Lactate levels were quantified using the CheKine™ Lactate Assay Kit (Abbkine, KTB1100) following the manufacturer’s protocol. Briefly, WT and KO testis homogenates were prepared in Lactate Assay Buffer and centrifuged at 12,000 × g for 5 min at 4 °C. A working reagent containing lactate dehydrogenase, cofactor, WST-8, and enhancer was mixed with supernatants or standards (0.0313–2 mM L(+)-lactate). After 30 min incubation at 37 °C in the dark, absorbance was measured at 450 nm using a microplate reader. Lactate concentrations were calculated based on a standard curve and normalized to sample fresh weight.
Genomic DNA extraction and mtDNA copy number analysis
Total genomic DNA was extracted from WT and KO testes using the Genomic DNA Purification Mini Spin Kit (Beyotime, D0063) following the manufacturer’s protocol. Briefly, 5 mg testes were lysed in 180 μL Lysis Buffer A containing 20 μL Proteinase K at 55 °C overnight. 4 μL RNase A (100 mg/mL) was added. After complete lysis, 200 μL Lysis Buffer B was added, mixed thoroughly, and incubated at 70 °C for 10 min, followed by precipitation with 200 μL ethanol. The lysate was loaded onto a DNA purification column and centrifuged (6000 × g, 1 min). The column was washed sequentially with 500 μL Wash Buffer I and 600 μL Wash Buffer II (18,000 × g, 1 min each), followed by a final centrifugation to remove residual ethanol. DNA was eluted in 30 μL Elution Buffer (room temperature, 3 min) and stored at −20 °C. For mtDNA copy number quantification, qPCR was performed using primers targeting mitochondrial mt-Nd1 and nuclear Hk2 genes. The relative mtDNA copy number was calculated using the comparative ΔΔCt method: 2^- (ΔCt_KO - ΔCt_WT), where ΔCt = Ct_mtDNA - Ct_nuclear. All reactions were performed in triplicate to ensure reproducibility.
Assessment of pyruvate dehydrogenase (PDH) activity
The pyruvate dehydrogenase (PDH) activity was measured using the CheKine™ PDH Activity Assay Kit (Abbkine, KTB1270) with a microplate reader capable of detecting absorbance at 605 nm. testes were homogenized in Extraction Buffer I (1 mL per 0.1 g tissue) supplemented with Extraction Buffer II (10 μL per 0.1 g tissue), followed by centrifugation (11,000 × g, 10 min, 4 °C) to obtain supernatants, which were kept on ice for immediate analysis. The reaction was initiated by mixing 10 μL of sample with 190 μL of freshly prepared working reagent (Reagent I, II, and III) in a 96-well plate, and absorbance readings were taken at 20 s (A₁) and 5 min 20 s (A₂) to calculate ΔA (A₁–A₂). PDH activity was normalized to sample weight (U/g tissue) using the formula 384.76 × ΔA ÷ sample weight (g).
Pyruvate concentration assay
The pyruvate (PA) Concentration was determined using the CheKine™ Pyruvate (PA) Assay Kit (Abbkine, KTB1121) following the manufacturer’s protocol. For sample preparation, testes were homogenized in Extraction Buffer (1 mL per 0.1 g tissue) and centrifuged (4000 × g, 10 min) to obtain supernatants. Standards (1.094–70 µg/mL) and samples (75 µL) were mixed with Chromogen A (25 µL) in microplates. After 2 min incubation at RT, Chromogen B (125 µL) was added and absorbance was immediately measured at 520 nm. PA concentrations were calculated using a standard curve and normalized to sample mass (µg/g).
Evaluation of Acetyl-CoA
The Acetyl-CoA concentration was measured using the Acetyl-CoA ELISA Kit (BBI, D751001) following the manufacturer’s protocol. The assay employs a sandwich ELISA technique, where samples or standards are added to anti-A-CoA antibody-coated wells, followed by incubation with biotin-conjugated detection antibody and HRP-streptavidin. The reaction produces a colorimetric signal (TMB substrate) measured at 450 nm, with Acetyl-CoA concentration proportional to absorbance. For sample preparation, testes were homogenized in PBS (1:9 w/v) and centrifuged (5000 × g, 10 min). Standards (0–20 ng/mL) were serially diluted, and all reagents were equilibrated to room temperature before use. The procedure involved: (1) adding 100 µL samples/standards to wells (90 min, 37 °C); (2) incubating with biotin-antibody (60 min, 37 °C) and HRP-streptavidin (30 min, 37 °C); (3) washing four times with 300 µL/well washing buffer; (4) adding TMB (15 min, 37 °C); and (5) stopping with 50 µL acidic solution. Absorbance was read at 450 nm within 5 min. Data were analyzed using a 4-parameter logistic standard curve, with sample values falling outside the standard range being appropriately diluted and reanalyzed. The relative acetyl-CoA concentration was calculated by dividing the acetyl-CoA concentration in the KO group by that in the WT group, with experiments conducted in biological triplicates to ensure reproducibility.
Proximity labeling
Proximity labeling were performed as previously described with slight modifications37,38,39. In brief, 293T cells expressing ABHD10-TurboID constructs were incubated in complete DMEM medium supplemented with 50 μM biotin (Sigma, B4639) for 16 h at 37 °C under 5% CO₂. Following biotinylation, cells were washed twice with ice-cold PBS (pH 7.4) and lysed in NP-40 buffer (50 mM HEPES, 150 mM NaCl, 1% NP-40, PH 7.4) freshly supplemented with protease inhibitor cocktail and PMSF. Cell lysates were gently rotated at 4 °C for 30 min to ensure complete solubilization, then clarified by centrifugation at 15,000 × g for 15 min at 4 °C. The resulting supernatants were subjected to affinity purification using pre-equilibrated Pierce Streptavidin Magnetic Beads (Thermo Fisher Scientific, 88816), with end-over-end rotation overnight at 4 °C. Subsequently, the beads were washed twice with NP-40 buffer, once with NP-40 buffer containing 1 M NaCl, twice with 2 M urea in 10 mM Tris-HCl (pH 8.0), and finally twice again with NP-40 buffer. Biotinylated proteins were eluted by heating the beads in 1× protein loading buffer containing 2 mM biotin and 20 mM DTT at 95 °C for 10 min. Both input lysates and streptavidin-enriched fractions were analyzed by western blot.
Statistics and reproducibility
Statistical analyses were conducted using GraphPad Prism 9 and RStudio software. The quantitative data are presented as mean ± SEM. For comparisons between two groups, a two-sided unpaired Student’s t-test was employed. The specific statistical tests utilized for each individual experiment are detailed in the corresponding figure legends. Statistical significance was determined at a threshold of P < 0.05. All experimental procedures including histology, electron microscope, immunofluorescence, western blot, immunoprecipitation, and qPCR were performed at least three independent biological replicates. All uncropped and unprocessed scans of the western blots were supplied in Source Data and Supplementary Information.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw protein spectrum data uploaded to the ProteomeXchange (https://www.ebi.ac.uk/pride/archive) database can be accessed by the code PXD067158. Mitochondrial protein expression profiles across 14 tissues were obtained from mouse MitoCarta3.0 (https://personal.broadinstitute.org/scalvo/MitoCarta3.0/mouse.mitocarta3.0.html). Source data for Figs. 1–7 and Supplementary Figs. 1–6 are provided in Supplementary Fig. or Source Data files. Source data are provided with this paper.
References
1 in 6 people globally affected by infertility: WHO. https://www.who.int/news/item/04-04-2023-1-in-6-people-globally-affected-by-infertility.
Sang, Q., Ray, P. F. & Wang, L. Understanding the genetics of human infertility. Science 380, 158–163 (2023).
Piomboni, P., Focarelli, R., Stendardi, A., Ferramosca, A. & Zara, V. The role of mitochondria in energy production for human sperm motility. Int. J. Androl. 35, 109–124 (2012).
du Plessis, S. S., Agarwal, A., Mohanty, G. & van der Linde, M. Oxidative phosphorylation versus glycolysis: what fuel do spermatozoa use? Asian J. Androl. 17, 230–235 (2015).
Ferramosca, A. & Zara, V. Bioenergetics of mammalian sperm capacitation. Biomed. Res. Int. 2014, 902953 (2014).
Morgenstern, M. et al. Quantitative high-confidence human mitochondrial proteome and its dynamics in cellular context. Cell Metab. 33, 2464–2483.e18 (2021).
Teves, M. E. & Roldan, E. R. S. Sperm bauplan and function and underlying processes of sperm formation and selection. Physiol. Rev. 102, 7–60 (2022).
Ho, H.-C. & Wey, S. Three dimensional rendering of the mitochondrial sheath morphogenesis during mouse spermiogenesis. Microsc. Res. Tech. 70, 719–723 (2007).
Otani, H., Tanaka, O., Kasai, K.-I. & Yoshioka, T. Development of mitochondrial helical sheath in the middle piece of the mouse spermatid tail: Regular dispositions and synchronized changes. Anat. Rec. 222, 26–33 (1988).
Gu, N.-H., Zhao, W.-L., Wang, G.-S. & Sun, F. Comparative analysis of mammalian sperm ultrastructure reveals relationships between sperm morphology, mitochondrial functions and motility. Reprod. Biol. Endocrinol. 17, 66 (2019).
Blanc, M., David, F. P. A. & van der Goot, F. G. SwissPalm 2: protein S-palmitoylation database. Methods Mol. Biol. 2009, 203–214 (2019).
Mesquita, S. et al. Mechanisms and functions of protein S-acylation. Nat. Rev. Mol. Cell Biol. 25, 488–509 (2024).
Du, G. et al. ROS-dependent S-palmitoylation activates cleaved and intact gasdermin D. Nature 630, 437–446 (2024).
Linder, M. E. & Deschenes, R. J. Palmitoylation: policing protein stability and traffic. Nat. Rev. Mol. Cell Biol. 8, 74–84 (2007).
Schmidt, M. F. & Schlesinger, M. J. Fatty acid binding to vesicular stomatitis virus glycoprotein: a new type of post-translational modification of the viral glycoprotein. Cell 17, 813–819 (1979).
Cao, Y. et al. ABHD10 is an S-depalmitoylase affecting redox homeostasis through peroxiredoxin-5. Nat. Chem. Biol. 15, 1232–1240 (2019).
Rath, S. et al. MitoCarta3.0: an updated mitochondrial proteome now with sub-organelle localization and pathway annotations. Nucleic Acids Res 49, D1541–D1547 (2021).
Mi, Y., Shi, Z. & Li, J. Spata19 is critical for sperm mitochondrial function and male fertility. Mol. Reprod. Dev. 82, 907–913 (2015).
Chen, Y. et al. Glycerol kinase-like proteins cooperate with Pld6 in regulating sperm mitochondrial sheath formation and male fertility. Cell Discov. 3, 1–18 (2017).
Shimada, K. et al. ARMC12 regulates spatiotemporal mitochondrial dynamics during spermiogenesis and is required for male fertility. Proc. Natl. Acad. Sci. USA 118, e2018355118 (2021).
Shimada, K., Kato, H., Miyata, H. & Ikawa, M. Glycerol kinase 2 is essential for proper arrangement of crescent-like mitochondria to form the mitochondrial sheath during mouse spermatogenesis. J. Reprod. Dev. 65, 155–162 (2019).
Jiang, Z. et al. PDHX acetylation facilitates tumor progression by disrupting PDC assembly and activating lactylation-mediated gene expression. Protein Cell 16, 49–63 (2025).
Sutendra, G. et al. A nuclear pyruvate dehydrogenase complex is important for the generation of acetyl-CoA and histone acetylation. Cell 158, 84–97 (2014).
Chen, C. et al. Oxidative phosphorylation enhances the leukemogenic capacity and resistance to chemotherapy of B cell acute lymphoblastic leukemia. Sci. Adv. 7, eabd6280 (2021).
Maalej, M. et al. Clinical, Molecular, and Computational Analysis in two cases with mitochondrial encephalomyopathy associated with SUCLG1 mutation in a consanguineous family. Biochem Biophys. Res Commun. 495, 1730–1737 (2018).
Randolph, L. M. et al. Fatal infantile lactic acidosis and a novel homozygous mutation in the SUCLG1 gene: a mitochondrial DNA depletion disorder. Mol. Genet. Metab. 102, 149–152 (2011).
Wu, Y. et al. ZDHHC19 localizes to the cell membrane of spermatids and is involved in spermatogenesis. Biol. Reprod. 106, 477–486 (2022).
Mise, S. et al. Kastor and Polluks polypeptides encoded by a single gene locus cooperatively regulate VDAC and spermatogenesis. Nat. Commun. 13, 1071 (2022).
Boguenet, M., Bouet, P.-E., Spiers, A., Reynier, P. & May-Panloup, P. Mitochondria: their role in spermatozoa and in male infertility. Hum. Reprod. Update 27, 697–719 (2021).
Xiong, M. et al. ADAD2 interacts with RNF17 in P-bodies to repress the Ping-pong cycle in pachytene piRNA biogenesis. J. Cell Biol. 222, e202206067 (2023).
Huang, X. et al. Fast, long-term, super-resolution imaging with Hessian structured illumination microscopy. Nat. Biotechnol. 36, 451–459 (2018).
Zhao, W. et al. Sparse deconvolution improves the resolution of live-cell super-resolution fluorescence microscopy. Nat. Biotechnol. 40, 606–617 (2022).
Barkovits, K. et al. Reproducibility, specificity and accuracy of relative quantification using spectral library-based data-independent acquisition. Mol. Cell. Proteom.: MCP 19, 181–197 (2020).
Demichev, V., Messner, C. B., Vernardis, S. I., Lilley, K. S. & Ralser, M. DIA-NN: neural networks and interference correction enable deep proteome coverage in high throughput. Nat. Methods 17, 41–44 (2020).
Kuang, W. et al. SLC22A14 is a mitochondrial riboflavin transporter required for sperm oxidative phosphorylation and male fertility. Cell Rep. 35, 109025 (2021).
Moon, N. et al. Stress increases sperm respiration and motility in mice and men. Nat. Commun. 15, 7900 (2024).
Makar, A. N. et al. The V-ATPase complex component RNAseK is required for lysosomal hydrolase delivery and autophagosome degradation. Nat. Commun. 15, 7743 (2024).
Kim, K.-E. et al. Dynamic tracking and identification of tissue-specific secretory proteins in the circulation of live mice. Nat. Commun. 12, 5204 (2021).
Ryu, K. W. et al. Cellular ATP demand creates metabolically distinct subpopulations of mitochondria. Nature 635, 746–754 (2024).
Acknowledgements
We thank the funding from the National Key Research and Development Program of China (No. 2024YFC2706800 to F.S.), National Natural Science Foundation of China (No. U24A20657, 82371613 to F.S., and 82401865 to B.Z.) and the Key Research and Development Program of Zhejiang Province (No. 2023C03035 to F.S.) in support of our work. We appreciate Li Wang, Chenyu Yang and Dandan Song at the Center of Electron Microscopy at Zhejiang University for their assistance with Electron Microscopy. Thanks to Bioprofile for providing the proteomics sequencing services. Special thanks to Milica Nedeljković, Ethan Tyler, Lex Kravitz, John Chilton and Macauley Smith Breault for graphic plotting. All images provided by Servier Medical Art and SciDraw were licensed under CC BY 4.0 (https://creativecommons.org/licenses/by/4.0/).
Author information
Authors and Affiliations
Contributions
S.Z., H.Z., H.X., M.X., S.G., D.L., and Y.Z. performed benched experiments; S.Z., M.X., S.G., Z.Y., Y.Z., B.Z., C.L. provided technical support and conceptual advice. S. Z. and H.X. wrote the paper. F.S. revised the paper. F.S. supervised this study. All authors discussed the results and critically reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Sara Cogliati, Tsan Xiao and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhou, S., Zhou, H., Xu, H. et al. The S-depalmitoylase ABHD10 is essential for sperm mitochondrial sheath formation and male fertility. Nat Commun 16, 10334 (2025). https://doi.org/10.1038/s41467-025-65300-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65300-2