Abstract
Wide variation of responses to identical stimuli presented to genetically inbred mice suggests the hypothesis that stochastic non-genetic variation, such as in chromatin state or enhancer activity during neurodevelopment, can mediate such phenotypic differences. However, this hypothesis is largely untested since capturing pre-existing molecular states requires non-destructive, longitudinal recording. Therefore, we tested the potential of Calling Cards (CC) to record transient neuronal enhancer activity during postnatal development in mice, and thereby associate such non-genetic variation with a subsequent phenotypic presentation – degree of seizure response to the pro-convulsant pentylenetetrazol. We show that recorded differences in enhancer activity at 243 loci predict a severe vs. mild response, and that these are enriched near genes associated with human epilepsy. We also validated pharmacologically a seizure-modifying role for two previously unassociated genes, Htr1f and Let7c. This proof-of-principle supports using CC broadly to discover predisposition loci for other neuropsychiatric traits and behaviors. Finally, as human disease is also influenced by non-genetic factors, similar epigenetic predispositions are possible in humans.
Similar content being viewed by others
Introduction
Neuropsychiatric disease risk, while heavily impacted by genetic makeup, is also influenced by non-genetic factors during development. For example, unique epigenetic states may be established early in life in response to factors such as environmental exposures (e.g., infections, head traumas) or psychosocial background, causing some cells in the body to activate specific sets of enhancers. Thus, inter-individual differences could be attributed partially to naturally-occurring variation in enhancer states across the population. This variation could contribute to brain disorders like epilepsy, which can have variable presentation even between identical twins1,2,3. However, a substantial challenge in studying any kind of epigenetic risk factors is disentangling pre-existing molecular states from those occurring as a consequence of an experimental manipulation or the disease itself. Overcoming this challenge would represent an important milestone, as connecting non-genetic variation to stochastic differences in pre-existing enhancer states could help identify novel mediators of seizure propensity or other naturally variable neuropsychiatric disease traits.
Epilepsy affects 46 million individuals worldwide and is characterized by unprovoked, recurrent seizures that can be highly variable in onset, frequency, duration, or intensity4,5. Human genome-wide association studies (GWAS) have identified many inherited risk factors for epilepsy. Many monogenic forms are causally linked to loss- or gain-of-function mutations in proteins influencing membrane excitability, such as SCN1A or GABRA16. Furthermore, other GWAS implicated genes like BCL11A or RORB are transcription factors that influence neuron development and identity, rather than membrane excitability7,8, and many gene variants that cause neurodevelopmental disorders or intellectual disability (ID) broadly also exhibit comorbid epilepsy9. Nevertheless, even among individuals with the same mutation, seizure frequency and intensity can vary substantially, suggesting other processes influence risk. Indeed, it is well-established that non-genetic factors such as traumatic brain injury or stroke also increase risk for developing seizures, and in some at-risk individuals, stress, alcohol consumption, or sleep deprivation can trigger an episode10. These lines of evidence argue that non-genetic risk factors arising stochastically during typical neurodevelopment, or that are uniquely acquired, may also influence seizure risk.
Variability in neuropsychiatric phenotypes influenced by non-genetic factors also exists in animals. For example, mouse models of acute seizure (e.g., induction with GABA-A receptor antagonist pentylenetetrazol; PTZ), result in a range of seizure responses even in effectively isogenic mice11. Specifically, while most mice with identical genetic backgrounds experience generalized tonic-clonic activity, some only exhibit mild freezing while others proceed to death. We hypothesize that this natural variability in presentation might be due to stochastic variation in pre-existing epigenetic states established during brain development. We therefore posited that PTZ induction in naive mice might represent a model system to examine 1) whether such pre-existing molecular states can influence subsequent phenotype or presentation, and 2) if so, to identify naturally-occurring susceptibility factors from such molecular states.
However, most methods of assessing molecular state, such as RNA-seq or ChIP-seq, are destructive, and can therefore only provide a snapshot of the molecular state at the time of sacrifice. They cannot, therefore, be used to connect pre-existing cellular states to behavioral outcomes12. Indeed, several studies using such methods have investigated global epigenetic changes such as genome-wide DNA methylation13,14,15,16,17,18, transcription factor19,20 and modified histone21 binding, chromatin accessibility and conformation19, following seizure induction. However, the prior states that influenced these changes remain unknown. Calling Cards (CC) is a transposon-transposase based technology that can create a permanent record of one such molecular state: transient transcription factor-DNA interactions22,23,24. The crux of CC is a self-reporting transposon that is directed to a genomic locus by the transposase hyperPiggyBac, which has a natural affinity for BRD4 binding sites in the genome. BRD4 is a histone acetylation reader that preferentially binds at active enhancers and promoters; increased levels of BRD4 binding often predict increased gene expression25,26. The transposase-DNA interaction results in a donor transposon insertion at a TTAA site at that locus, serving as a permanent record (a “calling card”) of nearby BRD4 binding that can be read out through targeted sequencing after sacrifice. Unlike epigenetic profiling tools like ChIP-seq or ATAC-seq, CC can record cumulative enhancer activity across extended time windows (i.e., across development prior to PTZ injection). Because CC insertion and subsequent transposon reporter expression requires relatively long time windows (24–48 h), minimal recording of molecular state occurs in the few minutes between PTZ injection and seizure (in contrast to rapid RNA induction of immediate early genes like c-fos). Therefore, CC could circumvent the chief challenge of disentangling pre-existing molecular states from those occurring as a consequence of seizure induction. If so, then CC could be applied broadly to understanding variability in many other neurological diseases or behaviors beyond acute seizure induction (Fig. 1).
A Frontflip hyperPiggyBac (hyPB) transposase (top) and barcoded self-reporting transposon (SRT) (bottom) constructs. ITR, inverted-terminal repeat. SD splice donor, SA splice acceptor, LTR long-terminal repeat, bc barcode. B Unfused hyPB transposase naturally interacts with BRD4 and directs donor transposons to enhancer sites, as marked by histone 3 (H3) acetylation (Ac). C A self-reporting transposon (red box) is inserted into the genome that reports its genomic locus via transcription. D Over days to weeks, SRTs are inserted across the genome. E Approach to use Calling Cards to connect pre-existing epigenetic states with behavioral variability in order to identify novel disease mechanisms, therapeutic targets, and epigenetic risk loci. Insertion of CC transposons are denoted by the presence of red transposon boxes into the genome. A–E Created in BioRender. Dougherty, J. (2025). https://biorender.com/brst9u3. Schoch, K. (2025). https://biorender.com/tw2kqal. Schoch, K. (2025). https://biorender.com/69p9l0u.
Here, we examined whether such a pre-recorded epigenetic status can predict future seizure susceptibility. We implemented neuron-specific CC in inbred, isogenic mice immediately after birth to record cumulative BRD4 binding events through one month of age. We then induced acute seizures and sacrificed animals after 15 min of treatment. Next, we stratified mice based on seizure responses and tested for differential CC transposon insertions between severe vs. mild seizure responders. We identified numerous loci with pre-existing enhancer activity differences between these groups. Further, these putative enhancer sites mapped near known epilepsy risk genes, and susceptibility loci were enriched for genes influencing neurodevelopmental maturity. Finally, we implemented pharmacological methods to support the identification of two previously understudied seizure-modifying genes from our dataset. This work supports the hypothesis that pre-existing molecular states that affect novel or established epilepsy pathways can influence seizure susceptibility. Our study thus provides a template for examining antecedent epigenetic and molecular states for various other neuropsychiatric diseases and traits.
Results
Recording pre-seizure Calling Cards profiles in a pharmacological model of acute seizures
Studies in mice have shown that there is substantial variability in seizure susceptibility across genetically identical individuals27,28,29,30. Therefore, we sought to record pre-existing variation in enhancer activity and correlate inter-individual difference in activity with seizure severity in a PTZ-induced acute seizure model in isogenic mice. We delivered conditionally expressed (FLEX, Front Flip), AAV Calling Cards viral vectors (Fig. 1A) to the cortices of a cohort of P0-1 mouse pups of the Syn1::Cre genotype. These CC reagents record the binding of BRD4, an epigenetic factor which marks active enhancer regions22,31,32. The FLEX system with neuronal (Syn1) Cre ensures recording occurs in neurons and limits the background contribution from other cells. At P28, after recording pre-seizure CC occupancy profiles in each animal, mice were administered acute seizures through intraperitoneal injection of the GABA-A receptor antagonist PTZ at 65 mg/kg. Mice were observed for 15 min and were then immediately euthanized, reasoning 15 min would allow insufficient time for any seizure-related CC insertions and expression (especially relative to the 4 weeks of recording that have already occurred), while allowing long enough to stratify seizure responses. Cortical tissues were then collected for CC analysis (Fig. 2A). Seizures were scored based on a modified Racine scale, ranging from 1 (immobility) to 5 (generalized tonic-clonic seizure) to 8 (death). We therefore trichotomized animals by seizure scores into mild (score 1–4), moderate (score 5), or severe (score 6–8) (Table 1). Of the 32 mice tested, 12 were mild responders, 9 were moderate, and 11 were severe (Fig. 2B, Supplementary Data 1). Multiple regression analysis revealed that seizure scores were not related to factors such as sex or weight, likely due to our controlling for weight with dosing (Supplementary Fig. S1A, B, Supplementary Data 1).
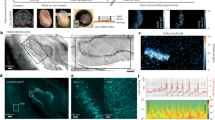
A Experimental paradigm of recording molecular states from postnatal day (P)0 to P28 prior to seizure with Calling Cards. Created in BioRender. Schoch, K. (2025) https://biorender.com/pmdb5nq. B Frequencies of seizure score by modified Racine scale after pentylenetetrazol (PTZ) injection. Categorization for seizure severity into mild, moderate, or severe responses are indicated by teal, gray, or purple respectively.
To confirm that our CC recordings were not influenced by the 15-min seizure, we investigated the dynamics of transposon hopping (transposition) in a simplified in vitro system. For this, we used the BrokenHeart (BH) transposon construct, in which the DsRed coding sequence is interrupted by a donor transposon. PiggyBac (PB) transposase activity removes the donor transposon from the construct and inserts it into the genome, leaving an intact DsRed sequence (Supplementary Fig. S2A). An N2A cell line, engineered to be constitutively expressing PB, was electroporated with BH plasmid and collected for lysis on a time course starting at 15 min following electroporation (Supplementary Fig. S2B). Transposition was detectable beginning 2–5 h following electroporation with BH (Supplementary Fig. S2C, D). Therefore, a 15 min seizure would not allow sufficient time for hopping. Furthermore, this simplified system represents the fastest scenario for transposition, because the PB enzyme is already present. Furthermore, unlike with SRTs, new transcription is not required for measurement of transposition in this system, as we are only measuring excision of the transposon from the donor construct. For the in vivo CC using self-reporting transposons, this step is followed by insertion of the transposon into the genome and transcription of the transposon. The additional time required for these steps would further preclude the detection of new CC insertions deposited during seizure that could influence the recording profiles. Therefore, we are confident that our in vivo CC will measure only pre-existing enhancer profiles.
Calling Cards identifies enhancer loci related to later seizure responses
To test whether epigenetic enhancer activity (indicated by BRD4-directed CC insertions) might predict seizure severity, we sought to compare CC profiles between mice that experienced mild or severe responses. We hypothesized that the stochastic distribution in seizure presentation would be matched by stochastic differences at the molecular level. Therefore, we expected that mice with moderate responses would appear as an intermediate state that incorporates both susceptibility and resistance molecular phenotypes. Therefore, a primary comparison between the mild (~1/3) and the severe (~1/3) groups would optimize our chance of detecting susceptibility/resistance states. To do this, we prepared and sequenced CC libraries from all mild (12) and severe (11) responders to recover CC insertions. To increase our sensitivity to detect unique insertions, libraries were prepared from separate tissue fragments of each cortex (up to six per animal). We selected the cortex for analysis because the cortex is quickly activated following intraperitoneal PTZ administration in rodents33. CC enhancer profiles were generally very similar across animals (Supplementary Fig. S3A, B). Animals passing QC metrics (methods) were included for downstream analysis, leaving 10 mice per group. As expected, we found that insertions were distributed across the genome (Supplementary Fig. S3C) and could not be attributed to any one mouse (Supplementary Fig. S3D). We used peak calling to define regions of enriched insertion density, for a total of 16,872 peaks (Supplementary Data 2). We expected that these peaks should include known enhancer sites. Indeed, peaks were located predominantly within intergenic regions and introns, rather than promoters, as would be expected (Supplementary Fig. S3E). Furthermore, overlaying CC peaks with ChIP-seq datasets of histone modifications from developing mouse forebrain34 revealed strong enrichment of active enhancer-associated marks H3K27ac (Fig. 3A, B, Supplementary Fig. S3F, I) and H3K4me1 (Supplementary Fig. S3G, I), but not of repressive mark H3K27me3 (Supplementary Fig. S3H, I), as expected from prior work establishing the AAV CC system22. CC peak centers had 2.6-fold enrichment over flanking regions for H3K27ac (p = 0), 1.5-fold enrichment for H3K4me1 (p = 0), and no enrichment for H3K27me3 (p = 0.13) (Supplementary Fig. S3J). This enrichment was specific to CC peaks, as shifting the peak coordinates by 20 kb resulted in loss of this enrichment (Supplementary Fig. S3F–H).
A Representative genome browser track depicting number of insertions, insertion density, peaks annotated by CC pipeline, and H3K27ac ChIP-seq density from P0 mouse forebrain (ENCODE) around the Snx10 and Skap2 loci (chr6: 51500K–51900K). Insertions enriched in mild or severe responses are separated, and annotated peaks represent genomic locations with strong density across responders. The H3K27ac track demonstrates a presence of established enhancer sites near peaks. B Enrichment plot (top) and heatmap (bottom) of H3K27ac ChIP-seq (same as A) relative enrichment, quantified as fold change over ChIP-seq control, centered on CC peaks. C Volcano plot depicting log2 fold enrichment (x-axis) and adjusted p-values (y-axis) of differential peaks between mild (teal) and severe responders (purple) using Fisher’s exact test with Benjamini–Hochberg correction. Peaks are annotated with nearby genes. D Genome browser track for representative peak around Ramac/Homer2 (chr7:81775773-81779508) depicting differential enrichment of mild insertions. E Genome browser track for multiple representative peaks around Zic1/Zic4 (chr9:91355K-91450K) depicting differential enrichment of severe insertions. F–I Two-sided Spearman correlations between individual animal seizure scores and insertion counts at loci near (F) Gabrb1/Gabra4 (Spearman ρ = −0.73, p = 0.02), G Ramac/Homer2 (ρ = −0.71, p = 0.02), H Man1a (ρ = 0.81, p = 0.004), and I Kcnc1 (ρ = 0.72, p = 0.02).
To test the hypothesis that there might be loci whose developmental epigenetic activity predicts later seizure presentation, we compared the number of insertions falling within each peak between mild and severe responders using Fisher’s exact test. We expected this categorical distinction between mild and severe to provide high sensitivity to detect differences related to seizure score. After Benjamini-Hocherberg correction, we observed 110 regions enriched for CC binding in mild-responding animals and 133 for severe responders (Fig. 3C and Supplementary Data 2).
We next annotated peaks with nearby genes and plotted the directionality of peaks toward mild or severe responders. As an example, we found relative enrichment in CC insertions around the Ramac/Homer2 gene in mild animals (Fig. 3D). Similarly, we observed heavy enrichment of CC insertions in peaks around Zic1/Zic4 in the severe-response mice, with seven peaks reaching significance (Fig. 3E). Among the strongest enrichment in mild responders occurred around Htr1f and Heatr3, and Sall3 and Bche for severe. We found enrichment near both coding and non-coding genes, such as the microRNA MirLet7c-1. Next, we conducted a posthoc analysis on these peaks showing categorical differences to examine whether the magnitude of change in enhancer utilization is also sometimes linearly correlated with seizure severity across animals. This complementary analysis would allow us to explore the quantitative relationship between seizure score and enhancer usage. Overall, within the set of mild or severe-related peaks, we found 15 sites that were significantly correlated with seizure score (Supplementary Data 2). Insertions near Gabrb1/Gabra4, Ahcyl1, Ramac/Homer2, Ptprk/Themis, Ano10, Kctd16/Pabpc2, and Mfap5/Aicda were negatively correlated with seizure score (Gabrb1/Gabra4 and Ramac/Homer2 shown in Fig. 3F, G). Insertions near Kcnc1, Man1a, Ifitm7, Nr2f2, Nyap2/Dock10, Sncaip/Zfp474, Lnp1/Tmem45a, and Snhg14/Snrpn were positively correlated with seizure score (Kcnc1 and Man1a shown in Fig. 3H, I). Notably, insertion numbers from strongly enriched genes like Ramac/Homer2 negatively correlated with numerical seizure score (ρ = −0.71, p = 0.02), while others like Zic1/Zic4 did not show strong correlation (ρ = 0.14, p > 0.3). These results provide evidence for a subset of peaks having a dose-dependent relationship between enhancer occupancy and seizure outcomes.
To ensure that the differential peaks we discovered represented robust epigenetic differences related to seizure vulnerability and not mere individual differences, we shuffled the comparison groups 50 times to comprise animals with mixed seizure severity scores and performed the same peak calling and differential analysis. Indeed, the number and proportion of differential peaks we observed in our dataset (1.44%) were significantly higher than those observed in 49/50 shuffled arrangements (mean 0.44%, st. dev. 0.35%, p = 0.02), indicating there is a biological coherence to these group assignments that was better than chance (Supplementary Fig. S4A). This suggests a substantial proportion of the 243 differentially bound regions we identified represented pre-existing variation in BRD4 binding that was predictive of later seizure severity. Furthermore, to determine the sensitivity of our CC approach to discovering differential peaks, we downsampled our dataset of insertion profiles to various percentages to reflect reduced sample sizes and quantified the number of differential peaks retained from the full dataset. Interestingly, while the number of differential peaks showed a relatively linear relationship with downsampling percentage, we did observe a plateau in the number of differential peaks significantly correlated to seizure score when downsampled to 80% (Supplementary Fig. S4B). Taken together, these results suggest that our analysis has high specificity (so that most of the differentially bound peaks we identified are likely bona fide) and reasonable sensitivity at this power.
Genes associated with seizure responses are associated with neurodevelopment and epilepsy
If pre-existing variation in neuronal enhancer activity predicts seizure severity, then genes near differentially active enhancers should be enriched for neurodevelopmental pathways and genes previously implicated in seizure disorders. We performed Gene Ontology (GO) for mild-response genes or severe-response genes against a background list consisting of all genes annotated near CC peaks. (We note that because only Syn1-Cre positive cells (neurons) had transposase activity, all loci should be from neurons). Surprisingly, even against this background of neuron loci, we found enrichment of multiple neurodevelopmental pathways in the severe-response genes, but not in mild-response genes. For example, enriched processes associated with severe genes included “generation of neurons” and “neuron differentiation” (Fig. 4A). Interestingly, severe-associated genes were also enriched for “L-glutamate transmembrane transporter activity,” such as transporters (Slc1a7, Slc7a11) and receptors (Grm1) (Fig. 4A). These suggest severe responders might carry a greater proportion of neurons in a more neurodevelopmentally immature state and/or might have cell populations more polarized towards excitatory neurons.
A Gene Ontology enrichment for severe genes. Representative genes are annotated for “L-glutamate transmembrane transport” and “neuron migration” processes. Graph shows log10(p-values) after Benjamini–Hochberg false discovery rate (FDR) correction. Distribution of expected gene overlaps between CC genes identified for mild (B), severe (C), or both (D) and comprehensive epilepsy/disease genes list from Prevention Genetics (PG) (list = 1481 genes) using permutation testing. Observed overlap between significant gene sets indicated by black dashed line. E Distribution of expected gene overlaps between CC genes identified for both response groups (mild and severe) and comprehensive epilepsy/disease genes list from Zhang et al. 2024 (list = 2945 genes). F Distribution of expected gene overlaps between CC genes identified for both response groups and gene list associated with cancer (list = 1164 genes). G Heatmap summary of z-scores (left, red-blue spectrum) and p-values (right, magenta-green spectrum) for overlap analysis between seizure response-associated genes with epilepsy, neuropsychiatric disease, and other gene lists (not adjusted for multiple comparisons). The gray dotted line marks p < 0.05.
We were specifically interested in whether genes from either list were enriched with established epilepsy genes, either related to monogenic forms of epilepsy or identified as a risk factor in common variant genome-wide association studies. This would be consistent with a model where slight epigenetic variations in activities across numerous established epilepsy genes could summate to alter risk. To measure enrichment for known epilepsy genes, we tested our significant mild and severe gene sets for the number of overlaps with known epilepsy gene lists, and compared this to an expected distribution of overlaps. We used permutation testing to generate the expected distributions of overlap using sets of all genes identified by CC peaks to control for the neuronal bias inherent in genes drawn from neuronal CC peaks. As a negative control, we also included lists of genes implicated in cancer, hypertension, and rheumatoid arthritis. Prevention Genetics clinical panels include a list of seizure and epilepsy genes for which seizures are a major clinical feature (Supplementary Fig. S5F–H), as well as a comprehensive list of seizure-and epilepsy-associated genes (Fig. 4B–D). We found that of the 326 mild genes, 8 were associated with monogenic epilepsy (z-score 0.42, p = 0.3888) (Supplementary Fig. S5F), and 37 with broader epilepsy associations (z-score 2.26, p = 0.0187) (Fig. 4B). Among the 381 severe genes, 15 were associated with monogenic epilepsy (z-score 2.48, p = 0.0152) (Supplementary Fig. S5G), and 47 with broader epilepsy associations (z-score 3.17, p = 0.0023) (Fig. 4C). A recently updated list of seizure genes6 includes and categorizes genes related to epilepsy with varying levels of clinical evidence, ranging from monogenic forms of epilepsy to potentially-associated genes (Supplementary Fig. S5A–E, Fig. 4E). Of the mild genes, 5 overlapped with monogenic epilepsy genes (z-score 1.17, p = 0.1828) (Supplementary Fig. S5C). Of the severe genes, 6 overlapped with monogenic epilepsy genes (z-score 1.35, p = 0.1404) (Supplementary Fig. S5D). In total, 129 genes from both mild and severe groups (686) overlapped with the list of all epilepsy genes (z-score 2.85, p = 0.003) (Fig. 4E). Owing to a similar etiology, we expected to observe overlap with other neurodevelopmental disorders. Indeed, mild and severe genes each showed strong overlap with autism genes35 (35 for mild, z-score 3.2, p = 0.0017; 44 for severe, z-score 4.08, p = 0.0001) (Supplementary Fig. S5I–K) and intellectual disability genes36 (44 for mild, z-score 2.71, p = 0.0057; 51 for severe, z-score 2.89, p = 0.003) (Supplementary Fig. S5L–N). As expected, the overlap with genes for cancer37,38 (22 mild, z-score 0.62, p = 0.2958; 20 severe, z-score −0.59, p = 0.754) and other less related diseases (hypertension39, rheumatoid arthritis40) across both response groups was not significantly different from other genes nearby to CC peaks (Fig. 4F, Supplementary Fig. S6). Even the overlap with Parkinson’s disease genes41 (5 mild, z-score 0.97, p = 0.2329; 6 severe, z-score 1.13, p = 0.1899) did not significantly differ from that of other CC genes (Supplementary Fig. S5O–Q), despite its being a neurological disease, potentially due to its neurodegenerative rather than neurodevelopmental etiology. Overall, our analysis indicates there is significant overlap between the epigenetic risk loci defined here and previously defined genetic risk loci.
Manipulation of seizure-response genes modifies seizure severity or latency
While there was significant overlap with known epilepsy genes, most CC implicated loci did not contain known epilepsy genes. Therefore, we predicted that these seizure response-related genes might represent previously uncharacterized pathways influencing seizure activity, and further, targetable pathways for future therapeutic leads. We screened the list of target genes for those that had small molecule antagonists or agonists using the Gene Drug Interactive Database. We found 56 such genes that had an interaction score of at least 10, representing likely target engagement (Supplementary Data 3). Owing to the availability of highly-selective, blood-brain barrier penetrant agonists, we selected Htr1f, which encodes the serotonin receptor 1f, for further study. Additionally, we had previously developed an antisense oligonucleotide (ASO) inhibitor for another potential target non-coding microRNA Mirlet7c-1 that can be delivered safely to the central nervous system. A peak at Htr1f was enriched in mild responders (mild/severe log2FC of 1.96), and two peaks at Mirlet7c-1 were modestly enriched in severe responders (log2FC 0.59 and 0.16; Fig. 3C, Supplementary Fig. S7, and Supplementary Data 2). Notably, neither HTR1F nor MIRLET7C has any existing ties to epilepsy disorders.
Htr1f encodes the serotonin receptor 1F, which is an inhibitory G-protein coupled receptor, and its most abundant expression in the central nervous system is in the retina and cortex42. To evaluate whether HTR1F activity can influence seizure responses, we delivered to P28 wildtype mice either vehicle or 8 mg/kg HTR1F agonist LY344864 by intraperitoneal injection for 3 days, then administered PTZ (Fig. 5A). LY344864 is a highly selective serotonin receptor 1F agonist that can cross the blood-brain barrier43,44. We evaluated both the maximal seizure score and the latencies for each mouse to achieve a given score. LY344864 did not influence seizure score, but significantly extended time until mice experienced seizure responses (Fig. 5B). The first response, usually represented by freezing or immobility behavior, was delayed by 25 s (median) (Fig. 5C). Score 3, which is marked by a “tail pop”, trended to a delay of 30 s (Fig. 5D). Score 5, which is the onset of a generalized tonic-clonic seizure, was delayed by 180 s (Fig. 5E).
A Paradigm for treatment of mice with HTR1F agonist LY344864 or Let-7 antagonist (antisense oligonucleotide [ASO]) and subsequent administration of PTZ to characterize seizure responses. Created in BioRender. Schoch, K. (2025) https://biorender.com/y53fyfl. B Mice were treated with HTR1F agonist or dimethyl sulfoxide (DMSO) vehicle for 3 days at P28 by daily intraperitoneal injection and responses to seizure induction were characterized. Seizure scores were plotted between control treatment versus LY344864 (n = 12,12; two-tailed Mann–Whitney p = 0.5466). Cumulative distribution demonstrating time in seconds between administration of PTZ and onset of C first response (n = 12, 12; 2-way ANOVA; p = 0.06), D score 3 (n = 11,12), E or score 5 (n = 10, 11; p = 0.02). Source data are provided as a Source Data file. F Female mice were treated with Let-7 ASO or scramble ASO control for 4 weeks from 3 months of age by intracerebroventricular injection, and responses to seizure induction were characterized. Seizure scores were plotted between scramble ASO sequence and Let-7 ASO (n = 12,12; two-tailed Mann–Whitney p = 0.0474). G–I Survival curves demonstrating time in seconds between administration of PTZ and onset of G first response (n = 12, 12; 2-way ANOVA, p = 0.003), H score 3 (n = 11, 12; p = 0.06), I or score 5 (n = 6,10). Error bars in (B, F) represent SEM.
Let-7c is a member of the Let-7 miRNA family, which share an identical seed sequence for target binding. Let-7 miRNAs are widely-expressed throughout the brain and suppress hundreds of target mRNAs, including those influencing neuronal synaptic development45. To evaluate whether Let-7 inhibition could influence seizures, we delivered an ASO to neutralize Let-7 miRNAs by intracerebroventricular injection to 3 month old mice, an age at which ASOs can be administered ICV easily (Fig. 5A). After one month, we administered PTZ and recorded seizure responses. Let-7 inhibition worsened seizure score and hastened the first response by 22.5 s with a trend to delaying score 3 by 17.5 s (Fig. 5F–I).
Discussion
CC provides a unique opportunity to record prior enhancer status for future readout. Using CC, we sought to evaluate if PTZ-seizure severity can be predicted by antecedent molecular states. We found activity at 243 genomic loci that distinguished mild from seizure responders, and upon mapping these sites to nearby genes, we observed an enrichment with neurodevelopmental pathways and epilepsy risk genes. For a subset of 15 of these, antecedent enhancer activity was linearly related to later seizure severity. Finally, we demonstrated the utility of this strategy to identify seizure-related targets by pharmacologically manipulating activity of two genes in these loci, Htr1f and Mirlet7c-1. Our results support the hypothesis that epigenetic risk factors during neurodevelopment can influence seizure vulnerability, and broadly that CC may be used to probe antecedent events related to non-genetic risk factors for a variety of outcomes. While our strategy focused on using BRD4 binding as a proxy for enhancer usage events, the non-genetic factors that contribute to seizure susceptibility likely encompass a myriad of other epigenetic changes such as DNA methylation and histone modifications. By fusing the transposase to transcription factors or chromatin binding proteins, the CC method could be used in future studies to correlate other types of pre-existing epigenetic changes with behavioral outcomes.
Our data suggest that natural variation in epilepsy risk can arise from stochastic trajectories that are enacted postnatally and that are tuned across early development. We note that the term ‘epigenetics’ classically encompasses a range of non-genetic influences46. For example, one could hypothesize differences in either cell proportions or in cell states could explain some of the observed CC differences related to seizure vulnerability. Consistent with the first hypothesis, we found enrichment with specific neuronal markers, such as Bcl11b (Citp2) and Sox2 in severe and Pou3f3 (Brn1) and Pou3f2 (Brn2) in mild responders47. This observation hints that differences in proportions of cells, such as in numbers of Brn1/Brn2 + (Layer 2–4 excitatory neurons) or Ctip2 + (Layer 5 excitatory) neurons, arising during development could be a contributing factor to seizure responses. Another example was the enrichment of “glutamate signaling” in GO for severe-response genes. This might suggest greater abundance of excitatory neurons, whereas the enrichment of insertions near GABA receptor subunits (Gabra4, Gabrb1, Kcdt16) and transporters (Slc6a1 and Slc6a11) in the mild group might suggest enrichment with inhibitory neurons. On the other hand, epigenetic differences could also reflect cellular states, circuit wiring, or of course, the more recent definition of epigenetics meaning chromatin states. Indeed, the presence of GABA receptor subunits could reflect differences of excitability in each cell, rather than differences in cell proportions. Future studies implementing a single-cell Calling Cards approach may help distinguish these possibilities. Nonetheless, our current analysis did still identify key pathways with seizure modifying activity.
The precise timing of when the differential enhancer usage occurred also remains elusive. Because CC records continuously from delivery (P0-P1)22, epigenetic differences could either be attributed to trajectory over development or the matured state at sacrifice. Even within development, our recording window from P0 to P28 covers the periods of synaptogenesis, synaptic pruning, gliogenesis, programmed cell death, and myelination48,49,50–any of which could have influenced seizure vulnerability. These questions motivate the need for inducible CC systems to enable temporal control of recording during desired windows. Such systems would also allow for examination of non-genetic factors contributing to a range of phenotypes presenting on longer time scales.
Nonetheless, we found interesting loci that may act epigenetically to influence seizure propensity. Most known genetic forms of epilepsy are related to gain or loss of function mutations in a single gene. We predicted that slight differences in activity of these genes might be associated with increased epilepsy risk. Indeed, mild responses were linked with CC activity near epilepsy genes like Gabrb1 and Slc6a1, for example. Severe responses had greater CC activity near genes such as Chd2 and Kcnc1. Compared to established, clinically-used panels of epilepsy genes (Prevention Genetics and6), we found strong enrichment across both mild and severe genes. This suggests the epigenetic signatures detected by CC might act in either direction; indeed, BRD4 can act as either a repressor or activator, depending on the context32,51,52.
Polygenic risk for epilepsy has been explored in GWAS mega-analyses from The International League Against Epilepsy Consortium on Complex Epilepsies, and we might expect the CC-tagged risk might also overlap with such common genetic risk. Genome-wide mega-analyses in 2018 and 2023 have respectively identified 16 and 26 risk loci related to various epilepsy subtypes7,8. Of the five loci associated with all forms of epilepsy in their analyses, three were linked with seizure response in our analysis: VRK2/FANCL (mapped to BCL11A, severe), RORB (mild), and HEATR3 (mild). Of those linked with generalized epilepsy, six sites associated with seizure response: POU3F3, PTPRK, PCDH7, COX7B2/GABRA4 (mapped to GABRA2), GRIK1, and ACVRL1/ACVRL1B (mapped to SCN8A). The locus including VRK2/FANCL/BCL11A has been strongly associated with epilepsy risk, and its mapping to VRK2 or BCL11A has not been clear8,53. An interesting observation in our results is the identification of their homologs Vrk1 and Bcl11b as both in loci associated with seizure response, despite being in distinct loci. If both can influence severity, this suggests the human locus’s association may be driven by both genes. Collectively, our data argue that slight variations in activity at both established monogenic epilepsy genes and common variant associated loci occur stochastically, and these variations during neurodevelopment might be predictive of future risk.
We also hypothesized that previously unassociated genes could suggest unexplored mechanisms contributing to epilepsy, or targets for therapeutic intervention. One intriguing example in our dataset with strong enrichment in severe-responders was the ZIC family (Zic1, Zic2, Zic4, and Zic5). The genomic locus that contains Zic1 and Zic4 had seven significant peaks, and the region containing Zic2 and Zic5 had five peaks. Although the ZIC loci did not show a quantitative relationship with seizure score across the full set of mice, the increased CC activity of ZIC loci seen in a subset of severe mice might represent loci with a critical point in which even small changes in enhancer activity can elicit larger effects on seizure vulnerability, or where a locus has potential to have a strong effect in individual animals, but is only perturbed infrequently across the population of animals. ZICs are zinc-finger proteins and transcriptional activators that direct neural progenitor proliferation in neurodevelopment54,55. Deficiency of ZIC1/ZIC4 proteins result in Dandy-Walker formation, which involves cerebellar hypoplasia and delayed motor and cognitive development. Loss of ZIC2 results in holoprosencephaly in humans and disrupted cortical lamination in mouse models. In later development, ZIC proteins influence neuronal migration and axon refinement, both of which could modify seizure propensities. For example, ZIC2 may regulate Cajal-Retzius cell migration55 through control of tyrosine kinase receptor EphB1 expression. The ZIC family may influence seizure susceptibility through its transcriptional regulation of genes involved in synaptic control, such as Camk2a56, the serotonin transporter Slc6a4 (SERT)57, and the D1 dopamine receptor DRD158. Given the presence of multiple peaks in our dataset, we suspect ZIC proteins might themselves represent epilepsy genes and anti-seizure targets, but as yet, no pharmacological agent for this family is available.
Most current anti-epileptic drugs operate by subduing neuronal excitability, for example by inhibiting sodium channels or augmenting GABAergic signaling59, so we sought to evaluate other mechanisms contributing to seizure activity. One gene target identified from our dataset that has established pharmacological tools was the serotonin receptor 1F (Htr1f). HTR1F is an inhibitory G-protein-coupled receptor that responds to serotonin but has relatively unknown functions. While agonism of HTR1F with LY344864 produced modest seizure-modifying activity in our assay, we predict that further development may yield a promising anti-epileptic. The non-coding microRNA Let-7 was also an intriguing target, owing to the presence of two severe-associated peaks near the Mirlet7c-1 genomic locus. We found that inhibiting Let-7 activity worsened seizure responses, validating its contribution to seizure activity. While Let-7 is not an established epilepsy target, this microRNA regulates many genes controlling neuronal excitability, including MECP245, BDNF60, and multiple autophagy related genes61, which may contribute to effects seen. We do note that the effect size of these two manipulations was not nearly as strong as current treatments for epilepsy like diazepam: when tested in a similar paradigm, pretreatment of diazepam completely blocked PTZ-induced convulsions in 7/7 mice, rather than simply modifying onset time62. Notably, our Calling Cards analysis did identify the GABA receptor locus (i.e., the targets of diazepam) as associated with seizure severity.
Therefore, we anticipate that further exploring previously understudied pathways could also inspire additional therapies for epilepsy once tools to manipulate these pathways have been developed. Overall, epigenetic risk factors such as those suggested here might yield novel drug targets or adjuvants for existing therapies providing untapped strategies to affect disease. However, these need to be first thoroughly tested in preclinical models of chronic epilepsy before advancing towards the clinic.
Beyond epilepsy, an individual’s outcomes or risk for neuropsychiatric diseases like Alzheimer’s disease or major depression hinge on natural variation. In these circumstances, epigenetic variability might manifest in an individual’s unique presentation, disease course, or response to therapy. In behavioral neuroscience, this variability might be modeled in “learners” or “non-learners” in social operant tasks that might stratify mice based on social motivation63, or distinguish mice that respond to social defeat stress with longer term depressive like behavior (vulnerable vs. resilient mice)64. CC provides the opportunity to isolate the epigenetic factors in balance during prodromal states, to reveal what protective pathways support successful therapeutic response, or to identify factors that make genetically identical individual animals respond differently to the same event. This work should serve as a template for applying this strategy broadly to reveal mechanisms of disease and targets for therapeutic intervention.
Methods
In vitro Calling Cards
To investigate the dynamics of transposon hopping, we engineered a cell line that could constitutively express the PB transposase. To generate the transposase construct, PB-ERT2 and T2A-GFP inserts were cloned into the pSMAL backbone. After confirmation of the construct by sequencing, HEK293T cells were transfected with the PB-ERT2-T2A-GFP construct (available upon request) to produce lentivirus. Neuro-2a (N2A) cells were transduced with PB-ERT2-T2A-GFP lentivirus. After visual confirmation of GFP expression, GFP-positive cells were bulk sorted to establish a polyclonal stable line.
The N2A parental cell line was obtained from ATCC (cat. no. CCL-131) and authenticated using ATCC standard procedures including karyotyping and testing for fungal, bacterial, ectromelia viral, and mycoplasma contamination. The N2A parental line tested negative for mycoplasma contamination. The engineered N2A-PB-ERT2-GFP line was not tested for mycoplasma contamination.
The N2A-PB-ERT2-T2A-GFP cells were cultured in EMEM (Corning #10-009-CV) + 10% fetal bovine serum (Gibco #A56708-01) + 1 uM 4-hydroxytamoxifen (4-OHT) (Sigma catalog: H6278-50MG) before and during the experiment for constitutive expression of PiggyBac transposase. Three replicate electroporations were performed. For each, a T-75 flask (6–7 × 106 cells) was split and prepared for electroporation using the Neon NxT Electroporation System 100uL Kit (ThermoFisher) following the manufacturer’s instructions. Six μg of BrokenHeart (BH) transposon plasmid (Addgene: 203394) was electroporated using the following parameters: 1 pulse at 1400 V for 20 ms. The cells were immediately plated in culture medium into 24-well plates and collected in technical triplicate at the following timepoints after electroporation: 15 min, 1 h, 2 h, 5 h, 9 h, 24 h, and 48 h. Cell collection occurred by aspirating the culture medium into a 1.5 ml tube, lifting cells remaining in the well with 0.25% Trypsin-EDTA (Gibco #25200056), and centrifuging the mixture for 3 min at 500 × g. This process ensured capture of both adhered and not yet adhered cells to account for the adherence status at a wide range of timepoints. Following aspiration of the supernatant, cell pellets were frozen until completion of the time course. Thawed cell pellets were resuspended and lysed in 25 mM NaOH with 0.2 mM EDTA, pH = 12, at 99 °C for 30 min. Following lysis, the lysate was neutralized with 40 mM Tris-HCl, pH = 5.2. Lysates were used as input for PCR amplification of BH using NEBNext Ultra II Q5 Master Mix (New England Biolabs #M0544X) with the following thermal cycling conditions: 98 °C for 30 sec; 35 cycles of 98 °C for 10 sec, 67 °C for 15 sec, and 72 °C for 30 sec; 72 °C for 2 min. Primer sequences were as follows: DsRed Fwd: 5′-CGACTACAAGAAGCTGTCCTTCC-3′, DsRed Rev: 5′-GCCCAGAGTCTTCTTCTGCAT-3′. PCR products were run on a 1.5% agarose gel stained with ethidium bromide and imaged on a UVP GelSolo (Analytik Jena). Fiji was used to quantify band intensities following background subtraction.
Animals
All practices and procedures using mice were approved by the Washington University in St. Louis Institutional Animal Care and Use Committee (protocol numbers 19-0875 and 22-0253) in accordance with National Institutes of Health (NIH) guidelines. All mice used in this study were bred and maintained in the Washington University in St. Louis vivarium, where they were kept on a 12-h light/dark cycle, constant temperature and humidity, and had unlimited access to food and water. All mice were on a C57BL/6J background from Jackson Laboratory (catalog: 000664), and experiments were balanced for sex, except the Let7 ASO study (see Drug treatment and preparation). Syn1-Cre mice used for implementation of Calling Cards were ordered from Jackson Laboratory: B6.Cg-Tg(Syn1-cre)671Jxm/J65 (catalog: 003966) and were inbred with C57BL/6J. Non-transgenic mice were used in drug studies and were euthanized in their home cage using carbon dioxide.
In vivo Calling Cards constructs, virus production, and injections
Transposase and donor transposon plasmids were packaged into AAV9 capsids by the Hope Center Viral Vectors Core at Washington University School of Medicine. For the donor transposons, we used a barcoded pool of Self-Reporting Transposons (SRTs) (Addgene: 1000000213). SRTs report their location via RNA transcription and enable about an order of magnitude more insertions to be recovered per animal, compared to non-SRT transposons22. Insertions are recovered following reverse transcription and PCR amplification of the SRTs in the RNA - and each inserted locus can produce multiple RNA copies, whereas non-SRTs begin with fewer copies as they are DNA-based transposons66. The barcoded SRT is a further innovation that allows us to distinguish insertions at identical genomic locations by their different barcodes, increasing resolution by a factor of the number of unique barcodes67. SRTs were co-injected with the ‘Front Flip’ Cre-dependent Transposase, which cleanly drives activity only in Cre-expressing cells22. Titers for each virus ranged between 6.8 × 1013 viral genomes/ml. A 1:1 mixture by volume of transposase and donor transposon viruses was intracranially injected into the cortex of P0-1 Syn1-Cre male and female mice (two sites per hemisphere; 1 μL of viral mix per site). This injection paradigm results in widespread delivery throughout the brain22,24, though not necessarily evenly within and between animals. Therefore, QC steps described in the analysis sections below are performed to ensure sufficient expression of CC reagents for downstream analysis. Prior to injections, pregnant females were monitored daily, and newborn pups were injected within 24 h. CC injected pups were not weaned prior to sacrifice at P28.
Seizure induction for Calling Cards and evaluation
Intraperitoneal (i.p.) injection of pentylenetetrazol (PTZ) is the most well-established method for PTZ-induced seizure induction30. PTZ delivered by i.p. rapidly enters the brain, typically resulting in epileptiform signs within 1–5 min. Dosing for i.p. PTZ delivery was selected because a literature-wide review of this approach found that 50% of mice exhibited generalized tonic-clonic activity (score 5) with a 65 mg/kg injection30. This dose was selected to achieve sizable groups for mild (score <5) and severe (score >5).
Mice were injected i.p. with 65 mg/kg PTZ (Sigma catalog: P6500-25G) and placed in a standard home cage, where they were immediately evaluated for seizure activity for 15 min while also being recorded with a video camera. Immediately afterwards, mice were anesthetized with isoflurane and decapitated. Cortices were extracted, and each hemi-cortex was dissected into three parts and flash frozen in liquid nitrogen. For CC libraries, Syn1-Cre animals were used at P28; for validation drug studies, non-transgenic mice were used at either P28 or at 3 months of age.
A previously defined ‘modified Racine scale’ was used to score seizure responses during the 15 min period of observation11,29,68 (Table 1.) For the purposes of this study, we created pre-determined severity categories to assign scores of 1–4 as “mild” responses; score 5 as “moderate”; and scores 6–8 as “severe.” The time at which each mouse achieved a score threshold was marked, and the latency from PTZ injection was calculated for time to “first response”, “score 3” (tail pop), or “score 5” (generalized tonic clonic activity). Comparisons of seizure latencies between groups were performed using only mice that achieved at least that score. Seizures were observed and recorded. Seizure scoring was performed by a blinded examiner based exclusively on the video recordings.
Calling Cards library preparation and sequencing
Calling Cards protocol proceeded largely as described24. Briefly, RNA was extracted from cortical tissues according to the manufacturer’s instructions (Zymo RNA Clean & Concentrator-5 Kit and QIAGEN RNEasy Plus Mini Kit). SRT sequencing libraries were prepared from these RNA samples as previously described24 and sequenced on the NovaSeq X Plus platform (Illumina). Multiple cortical pieces (up to six per animal) were processed separately to further increase our ability to detect unique insertions even beyond the number of available barcodes. Data were then pooled by animals for subsequent analysis. Libraries were prepared from all mild and severe animals, and successfully sequenced for 11 mild and 11 severe animals.
Raw reads were processed into QBED files containing transposon insertion sites (at TTAA sites) using the nf-core/callingcards69 Nextflow70,71 pipeline with the default mammalian parameters for mouse as described24. Briefly, read quality control (QC) was performed with FastQC72 and reads were split for parallel processing with SeqKit73. Barcodes were extracted by UMItools74 and reads were trimmed with Trimmomatic75. Reads were aligned to mm10 using bwamem276. Insertions were counted using callingCardsTools77 and alignment QC was performed using samtools78, Picard Tools79, and RSeQC80. QC data was presented with MultiQC81. QBEDs from each cortical piece were merged per animal with the ‘cat’ shell command and sorted using BEDOPS82. Reads aligning to the same TTAA site with separate barcodes, or from spatially separate cortical pieces, were considered unique insertions. After filtering for insertions with read depth >1, all unique insertions were considered equally, regardless of read depth.
Software and package versions are listed in Supplementary Data 4.
Calling Cards peak calling and annotation
Significantly enriched insertion peak calling was performed with the pycallingcards module83. Insertion profiles for all animals within each severity category were merged to create mild and severe profiles. Peaks were called on the mild and severe profiles separately using the MACCs method, then merged using pybedtools for a final set of peaks. The following parameters were used for the peak calling: reference = “mm10”, pvalue_adj_cutoff = 0, window_size = 2000, step_size = 800, lam_win_size = 1000000, pseudocounts = 1.
For quality control, pairwise Pearson correlation coefficients were calculated on insertion counts per million within each peak between animals. The average pairwise correlation coefficient was then calculated for each animal. Animals with average correlation coefficients above >0.91 were used for downstream analysis, including re-calling the peaks, resulting in 10 mild and 10 severe animals. Principal components analysis was also performed on the insertion counts per million within each peak between animals to assess similarity among samples. The two excluded animals with the lowest correlation coefficients, which were also outliers on the PCA, reflected the animals with the lowest total insertion counts.
Peak annotation was performed using bedtools84 and HOMER85. Because enhancers usually exert their effects on nearby genes86,87, genes were assigned to peaks based on proximity. We used bedtools to find up to five of the closest genes within one megabase of the peak, filtering out “Gm” and “Rik” genes that have not been assigned names. Genomic features of the peaks were identified by HOMER.
Histone modification enrichment analysis
Qualitative enrichment analysis of histone modifications at CC peaks and non-peak loci was performed using deepTools88. The following P0 mouse forebrain ChIP-Seq34 datasets from ENCODE89 were used: H3K27ac (ENCSR094TTT), H3K4me1 (ENCSR465PLB), and H3K27me3 (ENCSR070MOK). Enrichment was quantified by comparing ChIP-Seq signal at CC peak centers, defined as ±0.5 kb from center coordinates, versus at flanking regions, defined as two standard deviations of peak length away from peak centers (±6.4–7.4 kb), using paired two-tailed t-tests.
Differential peak analysis
For the main differential peak analysis, we compared the number of insertions falling within each peak between mild and severe groups (or between shuffled groups) using Fisher’s exact test implemented in pycallingcards83. Peaks with adjusted p-values < 0.05 after Benjamini–Hochberg FDR correction were considered differential peaks. In addition, as a post-hoc we confirmed the correlation between insertion counts per million at each of the significant peaks and seizure score using the Spearman test, with Benjamini–Hochberg correction.
For the label shuffling analysis, animal profiles were shuffled into two groups in 50 random arrangements such that each group had 4–6 mild and 4–6 severe animals and roughly equal numbers of insertions. Downstream peak calling and differential analysis proceeded as described here. An empirical p-value was determined by dividing the number of arrangements with equal or greater significant peaks than the observed value by the total number of arrangements.
For the downsampling analysis, insertion profiles for the mild and severe groups were randomly downsampled three times per percentage to 20–90% of the full dataset using pycallingcards83. Differential peak analysis was performed as described above using the peaks called on the full dataset.
Gene ontology enrichment analysis
Gene ontology was performed using Panther classification system to identify enrichment in “biological” processes. Genes associated with mild responders or with severe responders were evaluated against a background of all genes annotated with CC peaks (16,240 genes). Processes achieving p < 0.05 after Benjamini–Hochberg FDR correction are displayed.
Gene overlap significance testing
The significance of overlap between gene sets was statistically evaluated. The observed overlap between significant gene sets (mild, severe, both) and disease-related genes was compared to an expected distribution. This distribution was generated through 10,000 random permutations. In each permutation, genes were randomly sampled without replacement from the list of all genes annotated to CC peaks, with the sample size matched to each significant gene set being tested. The overlap between these random gene sets and the disease gene list was calculated. An empirical p-value was determined by dividing the number of permutations with equal or greater overlap than the observed value by the total number of permutations. Note the random gene lists and the significant sets are both drawn from peaks found in neurons, and thus this controls for any general neuron-specific bias to any disease.
The following disease gene datasets were used for the overlap analysis (Supplementary Data 5): monogenic epilepsy (Prevention Genetics Epilepsy and Seizure Panel, test code 7347; https://www.preventiongenetics.com/testInfo?val=Epilepsy-and-Seizure-Panel), epilepsy-related (Prevention Genetics Comprehensive Epilepsy and Seizure Panel, test code 16005; https://www.preventiongenetics.com/tests/pgmax-comprehensive-epilepsy-and-seizure-panel), monogenic and all epilepsy-related6, autism (SFARI genes; https://gene.sfari.org/database/human-gene/)35, ID (SysNDD phenotype search “intellectual disability”, category: definitive; https://sysndd.dbmr.unibe.ch/Phenotypes)36, Parkinson’s disease (Gene4PD, rare genes; http://genemed.tech/gene4pd/download)41, cancer (OncoKB; https://www.oncokb.org/cancer-genes)37,38, hypertension (CVD Atlas, PedAM Disease-variant association; https://ngdc.cncb.ac.cn/cvd/disease/CVDD000096)39, rheumatoid arthritis (RADB: http://www.bioapp.org/RADB/index.php/Index/index)40. Human disease gene IDs were converted to mouse gene IDs prior to overlap testing based on the Mouse Genome Informatics nomenclature.
Drug treatment and preparation
Experimental drug testing was performed in non-transgenic mice. Mice were randomly allocated into each group with balancing for sex, except for the Let-7 ASO study. This experiment only used female mice because there were differences in (expected) weight gain between male/female mice when using this ASO, which would interfere with PTZ dosing. Investigators were blinded to treatment groups in drug studies and during seizure scoring.
The HTR1F agonist LY344864 was ordered from Med Chem Express (Cat. No.: HY-13788). LY344864 was reconstituted in DMSO to stock concentration of 60 mg/mL, aliquoted, and kept at −80 °C. On the day of use, LY344864 was resuspended as 0.5% DMSO/drug in corn oil. LY344864 was delivered by intraperitoneal injection at 8 mg/kg. Corn oil containing 0.5% DMSO was used as vehicle control. Non-transgenic mice were injected once-daily for 3 days from P28-P30. One hour after the final injection, PTZ was administered, and seizure responses were recorded.
The Let-7 inhibitor was ordered from Integrated DNA Technologies with the sequence AACTATACAACCTACTACCTCA. A non-targeting scrambled sequence ATAACACTCTAACCACTATCAC was used as a control. Each oligonucleotide was fully modified with 2-methoxyethyl bases and a phosphorothioate backbone. 20 nanomoles of Let7 inhibitor, corresponding to 173.4 micrograms, was delivered by intracerebroventricular injection at age 3 months as previously described90. A total volume of 10 microliters was injected at a rate of 1 microliter per second, with the needle left in place for 2 min both before and after injection. Coordinates from bregma were the following: X = −1.0 mm lateral (right), Y = +0.3 mm anterior, Z = −3.0 mm ventral. Let7 inhibition in the CNS using the indicated sequence has previously been validated91,92. Four weeks after injection, mice were administered PTZ, and seizure responses were recorded.
Behavioral statistics
Multiple linear regressions were used to examine association between seizure score with mouse sex or weight. Seizure scores were non-normally distributed ordinal values, therefore non-parametric two-tailed Mann–Whitney tests were used to compare between treatment groups. Seizure latencies were compared between treatment groups using log-rank (Mantel–Cox) test.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw and processed CC data generated in this study have been deposited in the Gene Expression Omnibus (GEO) under accession no. GSE305773. Source data are provided with this paper.
Code availability
Code is publicly available on GitHub: https://github.com/Dougherty-Lab/seizure-calling-cards.
References
Berkovic, S. F., Howell, R. A., Hay, D. A. & Hopper, J. L. Epilepsies in twins: genetics of the major epilepsy syndromes. Ann. Neurol. 43, 435–445 (1998).
Kjeldsen, M. J. et al. Genetic factors in seizures: A population-based study of 47,626 US, Norwegian and Danish twin pairs. Twin Res. Hum. Genet. J. Int. Soc. Twin Stud. 8, 138–147 (2005).
Krenn, M. et al. Phenotypic variability of GABRA1-related epilepsy in monozygotic twins. Ann. Clin. Transl. Neurol. 6, 2317–2322 (2019).
Beghi, E. et al. Global, regional, and national burden of epilepsy, 1990–2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol. 18, 357–375 (2019).
Devinsky, O. et al. Epilepsy. Nat. Rev. Dis. Prim. 4, 1–24 (2018).
Zhang, M.-W. et al. Epilepsy-associated genes: an update. Seizure Eur. J. Epilepsy 116, 4–13 (2024).
Abou-Khalil, B. et al. Genome-wide mega-analysis identifies 16 loci and highlights diverse biological mechanisms in the common epilepsies. Nat. Commun. 9, 5269 (2018).
Stevelink, R. et al. GWAS meta-analysis of over 29,000 people with epilepsy identifies 26 risk loci and subtype-specific genetic architecture. Nat. Genet. 55, 1471–1482 (2023).
Zacher, P. et al. The genetic landscape of intellectual disability and epilepsy in adults and the elderly: A systematic genetic work-up of 150 individuals. Genet. Med. 23, 1492–1497 (2021).
Wassenaar, M., Kasteleijn-Nolst Trenité, D. G. A., Haan, G.-J., Carpay, J. A. & Leijten, F. S. S. Seizure precipitants in a community-based epilepsy cohort. J. Neurol. 261, 717–724 (2014).
Van Erum, J., Van Dam, D. & De Deyn, P. P. PTZ-induced seizures in mice require a revised Racine scale. Epilepsy Behav. 95, 51–55 (2019).
Johnson, D. S., Mortazavi, A., Myers, R. M. & Wold, B. Genome-wide mapping of in vivo protein-DNA interactions. Science 316, 1497–1502 (2007).
Bahabry, R. et al. Alterations in DNA 5-hydroxymethylation patterns in the hippocampus of an experimental model of chronic epilepsy. Neurobiol. Dis. 200, 106638 (2024).
Berger, T. C. et al. Neuronal and glial DNA methylation and gene expression changes in early epileptogenesis. PLOS ONE 14, e0226575 (2019).
Dębski, K. J. et al. Etiology matters – genomic DNA methylation patterns in three rat models of acquired epilepsy. Sci. Rep. 6, 25668 (2016).
Kobow, K. et al. Deep sequencing reveals increased DNA methylation in chronic rat epilepsy. Acta Neuropathol. 126, 741–756 (2013).
Miller-Delaney, S. F. C. et al. Differential DNA methylation patterns define status epilepticus and epileptic tolerance. J. Neurosci. 32, 1577–1588 (2012).
Williams-Karnesky, R. L. et al. Epigenetic changes induced by adenosine augmentation therapy prevent epileptogenesis. J. Clin. Invest. 123, 3552–3563 (2013).
Fernandez-Albert, J. et al. Immediate and deferred epigenomic signatures of in vivo neuronal activation in mouse hippocampus. Nat. Neurosci. 22, 1718–1730 (2019).
Stephens, G. S. et al. Genes bound by ΔFosB in different conditions with recurrent seizures regulate similar neuronal functions. Front. Neurosci. 14, 472 (2020).
Liu, Z. et al. Epigenomic tomography for probing spatially defined chromatin state in the brain. Cell Rep. Methods 4, 100738 (2024).
Cammack, A. J. et al. A viral toolkit for recording transcription factor–DNA interactions in live mouse tissues. Proc. Natl Acad. Sci. 117, 10003–10014 (2020).
Wang, H., Mayhew, D., Chen, X., Johnston, M. & Mitra, R. D. Calling Cards” for DNA-binding proteins in mammalian cells. Genetics 190, 941–949 (2012).
Yen, A. et al. Calling Cards: a customizable platform to longitudinally record protein-DNA interactions over time in cells and tissues. Curr. Protoc. 3, e883 (2023).
Devaiah, B. N. et al. BRD4 is a histone acetyltransferase that evicts nucleosomes from chromatin. Nat. Struct. Mol. Biol. 23, 540–548 (2016).
Kanno, T. et al. BRD4 assists elongation of both coding and enhancer RNAs guided by histone acetylation. Nat. Struct. Mol. Biol. 21, 1047–1057 (2014).
Aydin-Abidin, S., Yildirim, M., Abidin, İ, Akca, M. & Cansu, A. Comparison of focally induced epileptiform activities in C57BL/6 and BALB/c mice by using in vivo EEG recording. Neurosci. Lett. 504, 165–169 (2011).
Copping, N., Adhikari, A., Petkova, S. & Silverman, J. Genetic backgrounds have unique seizure response profiles and behavioral outcomes following convulsant administration. Epilepsy Behav. EB 101, 106547 (2019).
DeVos, S. L. et al. Antisense reduction of tau in adult mice protects against seizures. J. Neurosci. 33, 12887–12897 (2013).
Yuskaitis, C. J. et al. Factors influencing the acute pentylenetetrazole-induced seizure paradigm and a literature review. Ann. Clin. Transl. Neurol. 8, 1388–1397 (2021).
Gogol-Döring, A. et al. Genome-wide profiling reveals remarkable parallels between insertion site selection properties of the MLV retrovirus and the piggyBac transposon in primary human CD4+ T cells. Mol. Ther. 24, 592–606 (2016).
Latif, A.-L. et al. BRD4-mediated repression of p53 is a target for combination therapy in AML. Nat. Commun. 12, 241 (2021).
Keogh, B. P. et al. BOLD-fMRI of PTZ-induced seizures in rats. Epilepsy Res. 66, 75–90 (2005).
He, Y. et al. Spatiotemporal DNA methylome dynamics of the developing mouse fetus. Nature 583, 752–759 (2020).
Abrahams, B. S. et al. SFARI Gene 2.0: a community-driven knowledgebase for the autism spectrum disorders (ASDs). Mol. Autism 4, 36 (2013).
Kochinke, K. et al. Systematic phenomics analysis deconvolutes genes mutated in intellectual disability into biologically coherent modules. Am. J. Hum. Genet. 98, 149–164 (2016).
Chakravarty, D. et al. OncoKB: A Precision Oncology Knowledge Base. JCO Precis. Oncol. 1–16 https://doi.org/10.1200/PO.17.00011 (2017).
Suehnholz, S. P. et al. Quantifying the expanding landscape of clinical actionability for patients with cancer. Cancer Discov. 14, 49–65 (2024).
Qian, Q. et al. CVD Atlas: a multi-omics database of cardiovascular disease. Nucleic Acids Res. 848, https://doi.org/10.1093/nar/gkae848 (2024).
Zhang, R. et al. RADB: a database of rheumatoid arthritis-related polymorphisms. Database 2014, bau090 (2014).
Li, B. et al. Gene4PD: a comprehensive genetic database of Parkinson’s disease. Front. Neurosci. 15, 679568 (2021).
Acevedo-Triana, C. A., León, L. A. & Cardenas, F. P. Comparing the expression of genes related to serotonin (5-HT) in C57BL/6J mice and humans based on Data Available at the Allen Mouse Brain Atlas and Allen Human Brain Atlas. Neurol. Res. Int. 7138926 https://doi.org/10.1155/2017/7138926 (2017).
Phebus, L. A. et al. Characterization of LY344864 as a pharmacological tool to study 5-HT1F receptors: Binding affinities, brain penetration and activity in the neurogenic dural inflammation model of migraine. Life Sci. 61, 2117–2126 (1997).
Scholpa, N. E., Lynn, M. K., Corum, D., Boger, H. A. & Schnellmann, R. G. 5-HT1F receptor-mediated mitochondrial biogenesis for the treatment of Parkinson’s disease. Br. J. Pharmacol. 175, 348–358 (2018).
McGowan, H. et al. Hsa-let-7c miRNA regulates synaptic and neuronal function in human neurons. Front. Synaptic Neurosci. 10, 19 (2018).
Haig, D. The (dual) origin of epigenetics. Cold Spring Harb. Symp. Quant. Biol. 69, 67–70 (2004).
Loo, L. et al. Single-cell transcriptomic analysis of mouse neocortical development. Nat. Commun. 10, 134 (2019).
Chini, M. & Hanganu-Opatz, I. L. Prefrontal cortex development in health and disease: lessons from rodents and humans. Trends Neurosci. 44, 227–240 (2021).
Thion, M. S. & Garel, S. On place and time: microglia in embryonic and perinatal brain development. Curr. Opin. Neurobiol. 47, 121–130 (2017).
Zeiss, C. J. Comparative milestones in rodent and human postnatal central nervous system development. Toxicol. Pathol. 49, 1368–1373 (2021).
Liu, B. et al. BRD4-directed super-enhancer organization of transcription repression programs links to chemotherapeutic efficacy in breast cancer. Proc. Natl Acad. Sci. USA 119, 2109133119 (2022).
Sakamaki, J. et al. Bromodomain protein BRD4 is a transcriptional repressor of autophagy and lysosomal function. Mol. Cell 66, 517–532 9 (2017).
International League Against Epilepsy Consortium on Complex Epilepsies Genetic determinants of common epilepsies: a meta-analysis of genome-wide association studies. Lancet Neurol. 13, 893–903 (2014).
Aruga, J., Tohmonda, T., Homma, S. & Mikoshiba, K. Zic1 promotes the expansion of dorsal neural progenitors in spinal cord by inhibiting neuronal differentiation. Dev. Biol. 244, 329–341 (2002).
Murillo, B., Ruiz-Reig, N., Herrera, M., Fairén, A. & Herrera, E. Zic2 controls the migration of specific neuronal populations in the developing forebrain. J. Neurosci. 35, 11266–11280 (2015).
Sakurada, T., Mima, K., Kurisaki, A., Sugino, H. & Yamauchi, T. Neuronal cell type-specific promoter of the α CaM kinase II gene is activated by Zic2, a Zic family zinc finger protein. Neurosci. Res. 53, 323–330 (2005).
García-Frigola, C. & Herrera, E. Zic2 regulates the expression of Sert to modulate eye-specific refinement at the visual targets. EMBO J. 29, 3170–3183 (2010).
Yang, Y., Hwang, C. K., Junn, E., Lee, G. & Mouradian, M. M. ZIC2 and Sp3 repress Sp1-induced activation of the humanD1A dopamine receptor gene. J. Biol. Chem. 275, 38863–38869 (2000).
Sills, G. J. & Rogawski, M. A. Mechanisms of action of currently used antiseizure drugs. Neuropharmacology 168, 107966 (2020).
Nguyen, T., Su, C. & Singh, M. Let-7i inhibition enhances progesterone-induced functional recovery in a mouse model of ischemia. Proc. Natl Acad. Sci. 115, E9668–E9677 (2018).
Petri, R. et al. let-7 regulates radial migration of new-born neurons through positive regulation of autophagy. EMBO J. 36, 1379–1391 (2017).
File, S. E. Tolerance to the anti-pentylenetetrazole effects of diazepam in the mouse. Psychopharmacology 79, 284–286 (1983).
Maloney, S. E. et al. A comprehensive assay of social motivation reveals sex-specific roles of autism-associated genes and oxytocin. Cell Rep. Methods 3, 100504 (2023).
Russo, S. J., Murrough, J. W., Han, M.-H., Charney, D. S. & Nestler, E. J. Neurobiology of resilience. Nat. Neurosci. 15, 1475–1484 (2012).
Zhu, Y. et al. Ablation of NF1 function in neurons induces abnormal development of cerebral cortex and reactive gliosis in the brain. Genes Dev. 15, 859–876 (2001).
Moudgil, A. et al. Self-reporting transposons enable simultaneous readout of gene expression and transcription factor binding in single cells. Cell 182, 992–1008.e21 (2020).
Lalli, M. et al. Measuring transcription factor binding and gene expression using barcoded self-reporting transposon calling cards and transcriptomes. NAR Genomics Bioinforma. 4, lqac061 (2022).
Lüttjohann, A., Fabene, P. F. & van Luijtelaar, G. A revised Racine’s scale for PTZ-induced seizures in rats. Physiol. Behav. 98, 579–586 (2009).
Mateusiak, C. & Hanssen, F. Caling Cards v1.0.0 - Initial Release. Zenodo https://doi.org/10.5281/zenodo.11281598 (2024).
Di Tommaso, P. et al. Nextflow enables reproducible computational workflows. Nat. Biotechnol. 35, 316–319 (2017).
Ewels, P. A. et al. The nf-core framework for community-curated bioinformatics pipelines. Nat. Biotechnol. 38, 276–278 (2020).
Andrews, S. Babraham Bioinformatics - FastQC A Quality Control tool for High Throughput Sequence Data. https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2010).
Shen, W., Le, S., Li, Y. & Hu, F. SeqKit: a cross-platform and ultrafast toolkit for FASTA/Q file manipulation. PLOS ONE 11, e0163962 (2016).
Smith, T., Heger, A. & Sudbery, I. UMI-tools: modeling sequencing errors in Unique Molecular Identifiers to improve quantification accuracy. Genome Res. 27, 491–499 (2017).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Vasimuddin, M. D., Misra, S., Li, H. & Aluru, S. Efficient architecture-aware acceleration of BWA-MEM for multicore systems. In 2019 IEEE International Parallel and Distributed Processing Symposium (IPDPS) 314–324, https://doi.org/10.1109/IPDPS.2019.00041 (2019).
Mateusiak, C. cmatKhan/callingCardsTools: v1.2.0. Zenodo https://doi.org/10.5281/zenodo.10042067 (2023).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Broad Institute. Picard Tools - By Broad Institute. https://broadinstitute.github.io/picard/ (2019).
Wang, L., Wang, S. & Li, W. RSeQC: quality control of RNA-seq experiments. Bioinformatics 28, 2184–2185 (2012).
Ewels, P., Magnusson, M., Lundin, S. & Käller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinforma. Oxf. Engl. 32, 3047–3048 (2016).
Neph, S. et al. BEDOPS: high-performance genomic feature operations. Bioinformatics 28, 1919–1920 (2012).
Guo, J. et al. Pycallingcards: an integrated environment for visualizing, analyzing, and interpreting Calling Cards data. Bioinforma. Oxf. Engl. 40, 070 (2024).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Sun, F. et al. Promoter-enhancer communication occurs primarily within insulated neighborhoods. Mol. Cell 73, 250–263.e5 (2019).
Symmons, O. et al. Functional and topological characteristics of mammalian regulatory domains. Genome Res. 24, 390–400 (2014).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Dunham, I. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
DeVos, S. L. & Miller, T. M. Direct intraventricular delivery of drugs to the rodent central nervous system. J. Vis. Exp. JoVE 75, e50326 (2013).
Dubinsky, A. N. et al. Let-7 coordinately suppresses components of the amino acid sensing pathway to repress mTORC1 and induce autophagy. Cell Metab. 20, 626–638 (2014).
Koval, E. D. et al. Method for widespread microRNA-155 inhibition prolongs survival in ALS-model mice. Hum. Mol. Genet. 22, 4127–4135 (2013).
Acknowledgements
The authors thank Harrison Gabel for helpful comments on the manuscript, Rebecca Chase and Emma Jones for assistance with data and code accessibility, the DNA Sequencing Innovation Lab, Genome Technology Access Center at the McDonnell Genome Institute (GTAC@MGI), and the High Throughput Computing Facility at the Center for Genome Sciences and Systems Biology. This work was supported by the Hope Center Viral Vectors Core at Washington University School of Medicine. This work was supported by grants from the National Institute of Mental Health (RF1MH117070, RF1MH126723 to R.D.M., J.D.D., and F30MH136688-01A1 to M.A.G.); National Institute of Aging (F30AG082394-01A1 to B.D.B.).
Author information
Authors and Affiliations
Contributions
Project conceptualization: B.D.B., M.A.G., A.J.C., R.D.M., J.D.D., T.M.M. Method development, experiments, and data collection: B.D.B., M.A.G., D.A.G., S.M.M., M.S., A.Y., X.C., T.S. Formal analysis: B.D.B., M.A.G., J.G. Figures and data visualization: B.D.B., M.A.G., J.G. Writing-original draft: B.D.B., M.A.G., R.D.M., J.D.D., T.M.M. Writing-review and editing: B.D.B., M.A.G., D.A.G., A.J.C., R.D.M., J.D.D., T.M.M. Project coordination: B.D.B., M.A.G., R.D.M., J.D.D., T.M.M. Funding acquisition: B.D.B., M.A.G., R.D.M., J.D.D., T.M.M.
Corresponding authors
Ethics declarations
Competing interests
T.M.M. is a consultant for Ionis Pharmaceuticals, Biogen, and Arbor Biosciences. T.M.M. has licensing agreements with Ionis Pharmaceuticals and C2N Diagnostics. R.D.M. has submitted patents related to Calling Cards technology. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Elizabeth Heller and Wei-Ping Liao for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Boros, B.D., Gachechiladze, M.A., Guo, J. et al. Antecedent enhancer activity predicts future susceptibility to seizures in mice. Nat Commun 17, 300 (2026). https://doi.org/10.1038/s41467-025-65346-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65346-2