Abstract
Liquid biopsies hold promise to improve the diagnosis, assessment of response to therapy, and ultimately guide the management of cancer patients. However, implementation of this approach in brain tumors has proven challenging due to the limited passage of molecules across the blood-brain barrier (BBB). We recently reported results from a phase I clinical trial in which the BBB was transiently opened in glioblastoma (GBM) patients using skull-implantable low-intensity pulsed ultrasound combined with microbubbles (LIPU/MB). In this study, treatment and BBB opening was performed every 3 weeks with paclitaxel administration until disease progression or up to 6 cycles (NCT04528680). As an exploratory objective of this trial, here we investigate extracellular vesicles and particles (EVPs/EPs) released into circulation in the context of tumor cell death as a potential biomarker for response to treatment. We develop and validate a microfluidic device designed to capture tumor-derived EVPs in glioblastoma patients (GlioExoChip). This approach leverages GBM-based expression of phosphatidylserine and Annexin-V chemistry that is traditionally used to measure apoptosis. EVPs are characterized using nanoparticle tracking analysis, proteomics, western blot, and scanning electron microscopy. Proteomic analysis of circulating EVPs isolated from GBM patients reveals distinct expression patterns to that of healthy individuals, and scRNA-seq analysis of these genes supported their tumoral origin within the GBM microenvironment. In vitro, paclitaxel-susceptible glioma cells treated with this drug exhibit apoptosis and dose-dependent EVP release. In concordance, we find changes in EVP release following the initiation of paclitaxel with LIPU/MB correlated with overall survival in GBM patients. Thus, our study introduces an efficient microfluidic platform for the capture of circulating GBM EVPs and demonstrates that release upon BBB opening is predictive of outcomes following paclitaxel treatment. This approach represents a real-time surrogate biomarker for treatment response for a disease where imaging-based assessment of response has not been shown to be reliable. Future prospective validation is warranted.
Similar content being viewed by others
Introduction
Glioblastoma (GBM) is the most common and malignant primary brain tumor in adults, for which effective therapies are lacking. Patients typically undergo maximal safe resection followed by a standardized regimen of chemotherapy and radiation1, yet even with full treatment, the expected median survival is less than two years2,3,4. Decades of research have focused on improving outcomes for GBM patients, however, efforts are largely hindered by the blood-brain barrier (BBB), which limits the passage of drugs to the brain. In parallel, these tumors have variable susceptibility to drugs, and given that these are in the brain, the BBB limits the use of circulating biomarkers to quantify tumor burden or response to treatment for a given therapy. Innovative approaches in these areas are essential to spur progress in clinical management and improve outcomes for patients with GBM.
Clinicians rely on histological and molecular analyses performed at the time of surgery to predict outcomes and make decisions on future treatments, as ongoing changes in GBM biology or therapeutic response cannot be repeatedly measured. Though tumor progression may be suggested based on increases in MRI enhancement, treatments such as radiation and immunotherapy can result in similar radiographic patterns, a phenomenon termed “pseudo-progression”. This complicates decisions on clinical management, as the precise timing of tumor recurrence, and thus the need for additional surgical resection or changes in adjuvant therapies, is often unclear5,6. Accurate monitoring of progression is also a key component of clinical trials, as timely and precise measurement of susceptibility to treatment is needed for evaluation of new therapies.
Identifying longitudinal changes in a patient’s GBM, including tumor progression, may be improved using liquid biopsy, which provides a non-invasive alternative for serial sampling of tumors7. This approach relies on the presence of circulating biomarkers, such as released tumor cells, cell-free nucleic acids, or extracellular vesicles and particles (EVPs), that can be detected in various biological fluids (e.g., plasma, saliva, cerebrospinal fluid). EVPs are released by all eukaryotic cells8, typically as a result of stress. They constitute multiple subpopulations (exomeres, small and large exosomes), and range in size from 30 to 150 nm9. Small EVPs are promising biomarkers in GBM, as they carry pathological materials from tumor cells, exist at diagnostically meaningful concentrations in peripheral blood, and can be tied to neoplastic tissue based on the expression of surface markers10,11. However, the use of EVP profiling and other liquid biopsy approaches have thus far proven challenging in GBM, likely owing to an impenetrable BBB that limits the diffusion of these tumor-specific biomarkers into peripheral circulation for detection.
We recently reported results of a phase I clinical trial in which recurrent GBM patients underwent opening of the BBB for delivery of albumin-bound paclitaxel (PTX) (Abraxane) chemotherapy, using a skull-implantable low-intensity pulsed ultrasound device combined with injections of microbubbles (LIPU/MB, “sonication”)12. We validated that this device, termed the “SonoCloud-9” (SC-9), successfully opens the BBB in patients and increases brain parenchymal drug concentrations relative to non-sonicated brain12,13,14,15, consistent with results that we’ve observed in carefully controlled animal models16. Moreover, we observed that with optimal timing for LIPU/MB and drug infusion, the transient BBB opening can lead to drug trapping in the sonicated brain17. After implantation, patients underwent periodic outpatient sonication sessions in which PTX was administered during the opening of the BBB, as well as interval MR imaging every 3 cycles (or as clinically needed). Plasma samples were systematically collected during these infusion visits, both before and after sonication, providing a unique resource to understand changes in circulating tumor material after BBB opening and its relationship to clinical outcome and tumor evolution.
Due to the sensitivity and specificity that microfluidic technologies offer, our team has engineered a system for isolation and characterization of EVPs in cancer and other diseases that relies on this approach18,19,20,21,22. In this study we optimize a sensitive, rapid, and cost-effective microfluidic device (“GlioExoChip”) to harvest circulating tumor-specific EVPs in GBM patients. Using GBM cell lines and patient plasma specimens, we demonstrate that GlioExoChip exhibits high capture yields of vesicles and particles derived from tumor cells, which can be readily released for downstream characterization such as profiling of EVP size and concentration, and protein measurement. Indeed, proteomic profiling of EVPs captured from the plasma of GBM patients exhibited distinct patterns relative to normal control plasma, consistent with gene expression signatures observed in malignant glioma cells. Using GlioExoChip, we characterized longitudinal EVP profiles from GBM patients undergoing PTX with concomitant opening of the BBB. Using GBM cell lines, we observe that PTX treatment results in EVP release in drug-susceptible, but not resistant GBM cell lines. Consequently, stratification of patients based on changes in EVP release after PTX administration in the first two cycles was predictive of overall survival in our clinical trial patients. Collectively, our work describes an efficient and minimally invasive method to longitudinally monitor chemotherapy susceptibility in GBM patients.
Results
Development and validation of GlioExoChip for capture of GBM-derived EVPs
We optimized a microfluidic platform capable of harvesting tumor-specific EVPs from GBM cell lines and patient plasma specimens (termed GlioExoChip). We reasoned that capture of these vesicles and particles specifically from GBM tumor cells would increase the detection of clinically relevant signals (as opposed to all EVPs present in the plasma), and that longitudinal collection across multiple data time points could identify temporal patterns relative to therapy and patient outcomes. EVPs, predominantly those in the exosomal size range, are captured when fluid (e.g., plasma) is passed through channels enclosed between PDMS and a glass slide, while vesicle-depleted fluid exits at the distal end of each device (Fig. 1A). Cancer-specific EVPs can be isolated by depositing antibodies along the base of each channel that harbor an affinity for select proteins or phospholipids expressed on EVPs of interest.
A Microfluidic devices were designed to isolate EVPs from cell media and patient plasma. Each device is fabricated from a PDMS top and glass slide bottom. Channels contain interconnected 500-micron diameter wells to enhance EVP capture. In total, each chip has 30 × 60 microwells. Scale bar in SEM image: 500 µm. B Western blot was used to optimize and test antibodies with cell lysate and cell line derived ultracentrifuged (UC) EVPs in two replicates. GBM6 cells had strong GFAP and calnexin bands (cellular markers) that were not found in UC EVPs. While EGFR and TSG101 was detectable in cells, no bands were seen for UC EVPs. On the contrary, Flotillin-1 (a marker for vesicles) was stronger in EVPs compared to cells. C Schematic of EVP capture and release on the GlioExoChip device. Annexin-V is coated on each slide, which binds to PS on the surface of GBM EVPs in the presence of Ca2+ ions. Exosomes are released for downstream applications by chelating Ca2+ ions using EDTA. D EVPs from two GBM cell lines (U87 and GBM6) were captured with high efficiency on Annexin-V functionalized devices (93.03 ± 3.31% and 96.77 ± 0.72% for U87 and GBM6, respectively, n = 3 technical replicates) when compared with anti-EGFR and anti-CD63 based techniques. Non-neoplastic astrocytes (IM-HA) were not well-captured using Annexin-V. Individual recordings are shown as small blue dots, values less than zero are marked in red and plotted on the x-axis. E Fluorescent images show DiO stained EVPs (bright green dots) under a light microscope for different device surface functionalization conditions. The coverage of EVPs in the wells qualitatively follows the trend observed in (D), with the Annexin-V functionalized device showing maximum capture, followed by CD63 and EGFR. Negligible amount of EVPs were observed on an unfunctionalized (No antibody) device. F Nanoparticle tracking analysis (NTA) plot of GBM6 EVPs after release from GlioExoChip using EDTA. The strongest concentration peaks are in the exosome size range, marked with green dotted box. G SEM image of GBM6 cell line-derived EVPs captured in wells of GlioExoChip. Inset shows representative EVPs in a magnified view, which fall in the expected 30–15 nm size range. For each panel, source data are provided as a Source Data file.
We collected ultracentrifuged vesicles and particles from the supernatant of cultured GBM cells to investigate baseline levels of protein expression and explore possible targets for the capture of GBM EVPs using the ExoChip. Western blot revealed the expected expression of the cell markers GFAP23 and calnexin24 in GBM cells, with intensity that varied based on the amount of protein loaded (Fig. 1B). By contrast, the vesicular marker flotillin-125,26 was mostly detected in EVPs, with a weaker signal observed in cell-derived protein isolate. Notably, the exosome marker TSG10127 was attenuated in EVPs isolated using ultra-centrifugation, suggesting a relatively low yield of exosomes with this method. To improve capture efficiency and isolation of disease-specific EVPs, we next explored three distinct antibodies (Annexin-V, anti-EGFR, and anti-CD63) for targeting on our microfluidic platform. Annexin-V is a protein that binds to phosphatidylserine (PS) with high affinity in the presence of Ca2+ ions, as shown in Fig. 1C28. We explored several antigens expressed by glioma cells as means of capturing tumor EVPs, including PS, Epidermal Growth Factor Receptor (EGFR), and CD63. PS is found on the inner leaflet of normal cell membranes but is externalized during periods of cell stress and thereby located on the outer leaflet of EVPs from cancer cells29,30,31,32. EGFR is an extracellular protein frequently overexpressed in GBM and has previously been correlated with poor prognosis33. Finally, CD63 is a universally expressed tetraspanin found on the surface of exosomes, with varying levels of expression dependent on the cell type34, and is previously reported for isolation of exosomes.
We observed that devices coated (functionalized) with Annexin-V exhibited higher capture efficiency when compared with devices coated with anti-EGFR or anti-CD63, with the capture of 93.03 ± 3.31% and 96.77 ± 0.72% for the U87 and GBM6 cell lines, respectively; Fig. 1D. Notably, this approach failed to adequately capture EVPs from immortalized non-neoplastic astrocytes. This result suggested elevated and consistent expression of PS on the surface of GBM-derived EVPs from these two cell lines that was not found in non-tumor astrocytes, and low or non-uniform expression of EGFR and anti-CD63. Consistent with this, fluorescent staining demonstrated enhanced capture of EVs in Annexin-V-based devices compared to CD63 or EGFR (Fig. 1E). EVPs were released by chelating calcium using EDTA, and the discharged vesicles and particles were subsequently quantified with nanoparticle tracking analysis (NTA). This analysis revealed 89% of EVPs to be in the exosome size range among the U87 and GBM6 cell lines (Fig. 1F), which was further confirmed by scanning electron microscopy (SEM) imaging (Fig. 1G). Given these results, we conclude that Annexin-V conjugated ExoChip, referred as GlioExoChip, has been optimized for GBM EVP capture, with a dominance of the small EVP population.
Isolation of patient-derived GBM EVPs from plasma by GlioExoChip
We next tested GlioExoChip for EVP isolation using plasma samples from GBM patients. We processed four specimens using two different plasma volumes (100 µL or 30 µL; diluted to a total volume of 300 µL) to determine if additional information was gained by using a higher amount of input material. We obtained similar or higher concentrations of EVPs from 30 µL of plasma compared to 100 µL (Fig. S1). Thus, we concluded that 30 µL of plasma is sufficient to provide EVPs in the detectable range (order of 109 EVPs/mL) using GlioExoChip.
To assess the reproducibility of EVP quantification across different GlioExoChip devices, plasma collected from one patient (P112) and one healthy individual (HYC) were divided and processed separately through 3 identical devices. The concentration profile of released EVPs from each device was independently measured and compared. The NTA plot in Fig. 2A demonstrates that distinct concentration profiles were observed for diseased and healthy plasma (average concentration of 5.28 × 1010 versus 1.31 × 109 EVPs/mL in GBM vs. healthy, respectively, Student’s t-test p = 0.027), with the peak concentration values for both samples between 30 and 150 nm (small EVP) size range. Notably, reproducibility within each sample type among three replicates was high, as demonstrated by the linear patterns observed when the concentration profile on each device is shown in logarithmic scale (Fig. 2B; correlation of 95% or greater between each pairwise device). Similar to GBM cell lines, we also compared the capture performance of Annexin-V and anti-CD63 using patient and healthy plasma. We did not use anti-EGFR for comparisons with plasma because this antibody had the least capture efficiency among the three with cell-derived EVPs. In line with our earlier results, fluorescent staining confirmed enriched capture of GBM patient-derived EVPs when Annexin-V was utilized, compared to devices that relied on anti-CD63 antibodies (Fig. 2C). By contrast, fluorescent staining of healthy plasma demonstrated relatively higher capture with anti-CD63 (Fig. S2), suggesting the minimal presence of PS-expressing EVPs (associated with GBM) in healthy subjects. Examination of captured products by SEM yielded similar findings, whereby Annexin-V-based EVP capture resulted in increased EVPs in patients over healthy controls (Fig. 2D), while no differences were observed in EVP density on anti-CD63 devices (Fig. S3). These results suggest that GlioExoChip can reliably and specifically capture GBM EVPs from patient plasma when using Annexin-V.
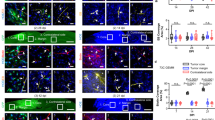
A Reproducibility was tested by flowing plasma of one patient (P112) and one healthy control (HYC) through three devices each. A nanoparticle tracking analysis (NTA) plot is shown of EVP concentration and size profile for both samples across replicates. The average EVP concentrations are 5.28 × 1010 and 1.31 × 109 EVPs/mL for GBM and healthy, respectively (two-sided Students’s t-test, p = 0.027). B EVP concentrations are plotted in logarithmic scale with each axis denoting measurements from a different microfluidic device (used to run the same sample). The position of every dot denotes the composite values from three devices. Colors red and blue are used for patient and healthy samples, respectively, while the size of dots denotes the size of EVP. The clustering of dots along a straight line indicates high reproducibility of GlioExoChip (paired correlation values are 0.96, 0.97, and 0.95 between devices). C DiO staining after capturing of EVPs from patient plasma (P112, cycle 2, pre-son) revealed high fluorescence intensity in the device functionalized with Annexin-V, with relatively less in anti-CD63 and negative control (No Antibody) devices. Inset shows magnified sections of each well. Scale bar: 100 µm. D SEM images of GlioExoChip-isolated EVPs on glass slide from GBM patient (P112, cycle 2, pre-son) and healthy (HDAK) plasma. E The protein amount extracted from GBM patients using GlioExoChip was significantly higher when compared with healthy control plasma for Annexin-V functionalized devices (p = 0.0013; two-sided Mann–Whitney test). A high protein amount was also obtained from Annexin-V (n = 9, n = 4, n = 2 biological replicates for each condition, respectively) devices when compared with CD63 (p = 0.036 for patient group) (n = 5, n = 4, n = 2 biological replicates for each condition, respectively). Each individual dot corresponds to one plasma sample. No Ab control corresponds to devices without plasma. F Protein characterization of EVPs from GBM patients and healthy plasma isolated using Annexin-V (AV) and CD63 functionalization. Strong flotillin-1 and GFAP bands were seen only in patient EVPs isolated using AV. Plasma from GBM patients included P112 (C2, Pre) and P114 (C4, Post), and healthy controls included HAM and HH. For each panel, source data are provided as a Source Data file.
We next quantified protein content from the EVPs among these samples. From healthy donors, we collected an average of 5.85 µg of protein per subject (± 4.75 µg), indicating that EVPs harbored sufficient material to perform common types of downstream characterization such as Western blot or proteomics (Table S1). Similar to our NTA results, we found that chips relying on Annexin-V isolated significantly more protein than CD63-based capture per volume of plasma (Welch’s t-test p-value for patients in the two groups, p = 0.036; Fig. 2E and Table S2). Furthermore, EVPs isolated from patient samples harbored a large increase in protein compared to healthy controls (p = 0.0013 from Mann–Whitney test and p = 0.0039 from Wilcoxon Signed Rank test; average of 85.1 µg), concordant with the increased total EVP concentration reported in Fig. 2A.
To confirm the source and purity of isolated EVPs, we performed Western blot of Glial Fibrillary Acidic Protein (GFAP; a marker for glial cell origin23), flotillin-1 (a small-EV marker25,26), TSG101 (an exosome marker), and calnexin (a negative marker that is present only in cells24). Patient EVPs isolated using Annexin-V exhibited strong flotillin-1 bands and were positive for GFAP, while EVPs from a healthy control had only flotillin-1 expression, indicating a lack of glial cell origin (Fig. 2F). By contrast, CD63-isolated EVs showed only trace bands for flotillin-1, consistent with weaker EVP capture using this approach. All samples tested had minimal signal for calnexin, confirming high purity of the isolated EVPs. The protein TSG10135 was strongly identified among GBM EVPs as compared to healthy controls (Fig. 3A), confirming an exosome-like subpopulation in these cases that was poorly captured using ultracentrifugation approaches (see also Fig. 1B).
A Protein characterization shows the presence of GFAP (glial marker) and TSG101 (exosome marker) in patient but not in healthy control plasma. All samples are devoid of cellular contamination (negative calnexin). B Proteomic profiling of EVPs derived from GBM patients (n = 7) and healthy controls (n = 3) identified disease-specific expression patterns. Shown is a circos plot of all detected genes across two groups. C The first principal component of proteomics data accounted for 49% of the total variability, and was sufficient to separate GBM patient EVPs (red) from healthy control EVPs (blue). D A volcano plot of differentially expressed genes identified proteins with significant fold changes compared to healthy controls. Significant up or downregulated genes are highlighted in red (identified by thresholding at |log2FC| > 1 and adjusted p-value < 0.05) with the top three genes labeled. Statistical analyses were performed with DESeq2 using the two-sided Wald test. E GBM patient-derived EVPs exhibited significant down-regulation of APOC1 and DMKN, and upregulation of SERPINA3, which most closely matched expression patterns seen in malignant cells of the GBM microenvironment based on scRNA-seq data41. Expression values for each column are normalized to show the range of each gene across cell types. For each panel, source data are provided as a Source Data file.
To further characterize EVPs captured from GBM patients, we performed proteomic profiling and compared this data to a cohort of healthy individuals. A total of 218 proteins were identified among EVPs harvested from plasma (Fig. 3B), and principal component analysis (PCA) of the proteomic data revealed that GBM patients exhibited distinctive protein expression patterns compared to healthy individuals. Indeed, nearly half of the observed variability in our dataset (48.4%) was explained by the first principal component, which reflected the source of each profiled sample (i.e., GBM patient vs. healthy individual; Fig. 3C). Analysis of differential abundance in tumor and control EVPs identified several proteins that were significantly elevated in GBM patients (Fig. 3D). Of note, several of these are known to play important roles in GBM biology and have been proposed as possible prognostic biomarkers, such as SERPINA336,37,38 and APOC139,40. Using single-cell RNA-seq data of GBM samples41 we investigated the potential cell origin of these proteins within the GBM microenvironment, and found that the top differentially expressed proteins exhibited patterns most consistent with malignant cells in GBM tissue (Fig. 3E). These included SERPINA3 (upregulated in GBM patient EVes) and APOC1 (downregulated in GBM patient EVs). Collectively, our results demonstrate that GlioExoChip successfully captures GBM-derived EVPs), validated using NTA, proteomics, and SEM imaging.
Concordance of circulating EVPs with radiographic enhancement during therapy
Having validated the yield and specificity of GlioExoChip for capture of GBM EVPs, we next used this device to characterize plasma specimens obtained during our recent phase 1 clinical trial, in which patients underwent longitudinal administration of PTX during LIPU/MB-based BBB disruption using a skull implantable ultrasound12. Plasma specimens were obtained at every therapeutic cycle before and after LIPU/MB and drug delivery (Fig. 4A, B), providing an opportunity to study EVP dynamics associated with BBB opening, as well as local disease control and overall survival for this patient cohort. The population characteristics of this cohort are provided in Table 1 of our previously published clinical trial12.
A GBM patients underwent implantation of the SonoCloud-9 device for ultrasound-based opening of blood-brain barrier (LIPU/MB: Low-intensity pulsed ultrasound assisted by microbubbles). The tight junctions of endothelial cells in the BBB are loosened by microbubbles pulsation induced by ultrasound, which alters release of extracellular vesicles and particles into circulation. B Plasma samples from 18 patients were collected to characterize EVP (30–150 nm size range) concentrations during longitudinal paclitaxel (PTX) treatment, both before and after BBB opening. The number of total cycles for each patient ranged from 2 to 6. A total of 66 paired plasma samples (pre and post) underwent analysis with GlioExoChip. C The timing of blood sample collection is shown for each longitudinal clinical visit, relative to sonication time (t = 0), stratified by patient. Each visit has pre and post-sonication collection points connected by a line. Blue lines represent visits in which both pre and post sonication blood samples were collected within 75 min of sonication (indicated by red dashed lines; 46 of 66 pre/post paired EVP samples). Patients with at least three such visits are shaded in dark grey, and analyzed in (F). D Longitudinal measurements of EVP concentration are plotted for P113 and P117, including specimens before (pre; teal) and after (post; purple) sonication. The blue and red dashed lines represent the timing of tumor recurrence and death, respectively. E Changes in EVP concentrations after sonication were associated with the degree of tumor enhancement at each corresponding timepoint (p < 0.01; Pearson correlation). Each dot represents an EVP ratio measurement (post/pre-sonication), colored according to the post-op day. The standard error associated with the regression line generated by all points is shown in grey. F Changes in EVP concentration after sonication closely mirrored the amount of MRI enhancement in each patient over longitudinal clinic visits. Each datapoint represents an MRI (orange) or exosome (grey) measurement, and has been normalized to the highest value for that modality. Amount of enhancement is linearly interpolated at each longitudinal clinic visit (open circles). The right panels show longitudinal MRI images from a representative patient (P114) and demonstrate progressive growth of the enhancing region. For each panel, source data are provided as a Source Data file.
We processed 132 longitudinal plasma specimens (66 pre and post sonication pairs) using GlioExoChip, and calculated EVP concentration and purity (ratio of 30–150 nm EVPs to total EVPs) for each sample. We focused specifically on the small EVP population (30–150 nm) among captured EVPs, due to its enrichment on GlioExoChip and efficient storage of intra-cellular cargo (discussed above) and referred them as EVPs in the following sections. Nearly all EVP concentrations fell on the order of 109/mL (Fig. S4A). To understand the effects of BBB opening on EVP release, we focused initial analyses on EVP concentration changes occurring before and after sonication (Fig. S4B). Fourteen patients had at least three therapeutic cycles available for paired analysis (both pre- and post-sonication available) (Fig. 4C). We observed clear associations between pre- and post-sonication measurements within these patients (Figs. 4D and S5). Notably, however, paired comparison of samples before/after BBB opening did not identify consistent changes in EVP concentration when considered across patients and cycles (p = 0.82; Fig. S4B). As the amount of MRI enhancement can approximate overall tumor burden, we examined if this variable might correlate with longitudinal EVP release after BBB opening (ratio of post/pre). Indeed, analysis of our aggregate dataset revealed an association of these variables, whereby EVP concentration changes were related to enhancement level in each patient at the time of sonication (p = 0.005, r = 0.52; Pearson correlation; using samples acquired within 75 min of BBB opening) (Fig. 4C and E).
We next performed a longitudinal analysis of EVPs from plasma samples collected during outpatient clinic visits in which chemotherapy was administered during BBB opening, aiming to understand if GBM EVPs were reflective of radiographic changes (MRI enhancement) during this period. Though a blood draw was performed before and after the opening of the BBB with each sonication cycle, the timing of specimen collection could be variable (Fig. 4C). To optimize the detection of clinically relevant signals, we limited our dataset to pre/post sample pairs in which both specimens were collected within 75 min of sonication.
During each patient’s treatment course, periodic MR imaging was performed to track tumor progression. Volumetric quantification of enhancement was performed as previously reported12,15, providing an opportunity to compare longitudinal tumor enhancement to circulating EVP concentration in individual patients. We performed linear interpolation to synchronize tumor enhancement levels to each EVP collection date. We focused on patients that had at least three longitudinal cycles of LIPU/MB for tracking (n = 6 patients). Among the resulting data, the change in EVPs after sonication (post/pre ratio) closely resembled enhancement on MRI over time (Fig. 4F), though repeated measures correlation (an estimate of within-individual association between two variables measured on multiple occasions42) did not reach statistical significance (Fig. S6A; p = 0.08; r = 0.38). Notably, when this analysis was repeated among the same patients using the pre-sonication EVP concentrations (rather than post/pre ratio), the association was much weaker (Fig. S6B, C; p = 0.59; r = −0.12). This result suggests that opening of the BBB enhances the release of EVPs into the blood circulation, whereby consideration of only non-sonicated values partially obscures the relationship of EVP concentration with radiographic progression of enhancing disease.
EVP release following LIPU/MB as a biomarker for glioblastoma susceptibility and response to paclitaxel
As GlioExoChip is designed to capture EVPs shed from apoptotic GBM cells (PS+), we hypothesized that our longitudinal data could provide insights into each patient’s response to PTX treatment. To investigate a possible association, we first analyzed in vitro the effects of PTX treatment on EVP release in GBM cells with variable susceptibility to this drug. Specifically, we used patient-derived xenograft (PDX) cell lines to compare the effects of PTX treatment on GBM cells with relative susceptibility to this drug to those with resistance (GBM43C and GBM43T, respectively; Fig. 5A). Exposure to increasing doses of PTX resulted in higher levels of cell apoptosis in the PTX-susceptible cell line GBM43C, indicated by increasing Annexin-V positivity (Figs. 5B and S7). Indeed, when we co-administered an inhibitor of apoptosis for this cell line we observed that the efficacy of PTX was reduced, confirming an essential role of this cell death pathway (Fig. S8). This result is consistent with a previous CRISPR screen in which we identified genes associated with PTX susceptibility in GBM43, which included a number of markers associated with apoptosis (Table S3). However, minimal changes were observed in Annexin-V for the PTX-resistant cell line GBM43T. Notably, elevated apoptosis also resulted in a dose-dependent increase in EVP release among PTX-sensitive (but not resistant) cells (Fig. 5C). We did not observe changes in the distribution of EVP sizes (Fig. 5D). These results suggest that GBM cells that are susceptible to PTX release more EVPs in response to treatment with this drug, while those that are resistant do not.
A The cell titer glow (CTG) dose response curve to paclitaxel (PTX) is shown for susceptible (GBM43C, purple) and resistant (GBM43T, green) GBM cell lines. At each concentration, n = 3 technical replicates are measured for each sample. Error bars indicate the standard error. B Measurement of Annexin-V (+) cells at increasing doses of PTX (0, 50, and 500 nM) revealed a progressive increase of apoptosis only in the susceptible cell line GBM43C. C NTA quantification of EVPs after treatment of GBM cell lines with PTX indicates that GBM43C has increasing EVP release associated with increased dose of PTX. D The distribution of EVP sizes did not change significantly with administration of increasing PTX, and remained in the EVP size range. E The slope direction of EVP ratios between cycles #1 and #2 was related to overall survival across patients (Cox hazards ratio = 0.203; Wald test p = 0.05, two-sided). A slope of zero is marked by the dashed line, and patients with positive and negative slopes are colored light blue and red, respectively. The grey ribbon represents the standard error of the fitted regression line. F Patients with negative EVP slopes had worse overall survival, shown in this Kaplan–Meier curve (log-rank p = 0.035). G Our data supports a model whereby patients that harbor paclitaxel-susceptible tumors have increased cell death with treatment, resulting in elevated EVP release and longer overall survival. Schema created using BioRender. For each panel, source data are provided as a Source Data file.
Based on this insight, we next investigated the relationship of EVP release and GBM susceptibility to PTX (determined by overall survival) across GBM patients that received the maximum dose in our clinical trial. We reasoned that changes in EVP concentration after BBB opening (ratio of post/pre-sonication) between therapeutic cycles one and two might reflect how patient GBM cells were responding to this treatment, as susceptible cells would release more EVPs in response to drug. We therefore defined a new variable, termed “EVP slope”, which was calculated as difference in post/pre ratios between two cycles, divided by the number of days between those cycles (see “Methods” section). This metric gives a measure of how tumor material release is changing over time with respect to BBB opening. During the first cycle, patients had only been exposed to paclitaxel for a few minutes at the time of plasma collection, while at cycle two, the tumors had had three weeks to undergo apoptosis and release EVPs following exposure to the drug. We observed that a subset of patients had negative slopes during this period (Fig. 5E; red squares), suggesting that less EVPs were being released in response to sonication over time. The remaining patients had positive slopes (blue squares), and notably, we found that on average these patients exhibited a doubling of overall survival compared to the first group (median OS of 610 vs 223 days, Students T-test p = 0.05). Indeed, stratification of patients into negative and positive slope groups at this timepoint was predictive of patient overall survival (Fig. 5E; Cox Hazard Ratio = 0.203, Wald test p = 0.05), and resulted in distinctive Kaplan–Meier curves with a significant difference in log-rank test (Fig. 5F; p = 0.035). These results suggest that patients who were responding to treatment with PTX released more EVPs during BBB opening after initiation of this chemotherapy (Fig. 5G).
In similar fashion, we examined the correlation of EVP slope with radiographic variables, including progression-free survival (number of days between surgery and radiographic growth of enhancement) and the amount of tumor-related MRI enhancement change during the period of EVP collection. In contrast to associations identified with overall survival, both radiographic variables missed significance (p = 0.53 and 0.27, respectively; Fig. S9). Collectively, our results suggest that EVP release dynamics may provide valuable insights towards predicting GBM response to paclitaxel after BBB opening.
Discussion
Liquid biopsy is emerging as a diagnostic and tumor burden biomarker in cancer, yet its application for GBM remains a challenge. In this study, we designed and validated a microfluidic device for capture of GBM-specific EVPs based on PS expression, and leveraged this technology to explore longitudinal changes in circulating EVPs after opening of the BBB. Our results distinguished patients with PTX-susceptible versus resistant GBMs based on changes in EVP concentrations after one cycle of this drug, suggesting that apoptotic release of PS+ EVPs may represent a surrogate marker for tumor response to cytotoxic therapy.
Several emerging liquid biopsy technologies are under investigation in GBM, including the detection of mutations and methylation patterns of cell-free DNA and exosomes, among others44,45. Our clinical analyses focused on small EVP quantification (the dominant population isolated by our device), as these vesicles and particles are enriched during periods of environmental stress, have diagnostically meaningful concentrations in peripheral blood46, can be tied to neoplastic tissue based on surface markers10, and use a biogenesis pathway that results in enhanced packaging of intracellular cargos for harvest and characterization8. Compared to conventional methods of isolating EVPs from plasma using size and density (e.g., ultracentrifugation), or charge-based approaches (e.g., dielectrophoresis)47, more recent studies in GBM have utilized immunoaffinity to improve specificity48,49,50,51. We optimized a microfluidic platform that was previously developed for lung-cancer exosomes for the capture of GBM-derived vesicles and particles from plasma. This approach leverages preferential expression of PS on the membrane of cancer and damaged cells31,52,53, which we observed to improve yield compared to EGFR and CD63. Captured EVPs exhibited proteomic patterns similar to GBM cells, which distinguished them from those harvested from normal control plasma. Our work thereby establishes an approach to capture GBM EVPs that can be utilized for characterization of proteins, lipids, DNA, and RNA from minimal plasma input.
GBM is a highly aggressive brain tumor in which current treatment approaches result in rapid tumor evolution and heterogeneous responses.15,43,54,55,56,57 Patient susceptibility to chemotherapies such as temozolomide (the current standard of care)57 and PTX12,15,43 is often unpredictable and may change over time. Due to pseudoprogression58,59, the use of MRI is limited in determining tumor resistance to therapy and whether a given treatment might be beneficial for individual patients. It is critical to determine early whether a given therapy will lead to prolongation of survival, as prompt modification of ineffective treatment plans can save valuable time and maximize patient benefit. Similar to what has been described for other cancers60,61,62, we’ve previously reported variable susceptibility of GBM to PTX, which translates into survival differences15,43. In this study, we found that PTX-susceptible GBM cell lines release PS+ EVPs during therapy-related induction of apoptosis, but this phenomenon is not observed in PTX-resistant GBM cell lines. Consequently, GlioExoChip, which isolates EVPs based on PS expression, detected changes in PTX-responsive patients after a single cycle of therapy. This was demonstrated as a change in release of tumor material after BBB opening between subsequent treatment cycles, we which measured as EVP slope. Our data thereby suggests that patients with PTX-susceptible GBM shed EVPs in during treatment-induced apoptosis of tumor cells, which can serve as a real-time assessment of individual patient response. Our data also indicated that autophagy and pyroptosis may play a role in PTX response in GBM cells, and future work might focus on investigating markers of these pathways. As there may be other potential unknown factors that could explain these results, prospective validation in a larger cohort will be needed.
Prior studies have established the importance of vesicles in GBM biology and their potential as a prognostic biomarker49,50,51,63,64,65,66,67,68,69,70. GBM patients are known to exhibit elevated levels of circulating EVs with unique cargos compared to healthy controls68,71, and EV quantity and material has been shown to correlate with lower overall survival63,64,65. Comparison of multiple EV time points in GBM patients has been more limited, though recent studies have investigated MGMT status using EGFR-based immuno-affinity50 and IDH mutations based on EV RNA70. Importantly, most previous EV studies have been performed in the setting of an intact BBB. The BBB acts as an impediment to the release of tumor material into circulation and can thereby obscure the precision and accuracy of blood-based liquid biopsy attempts for GBM72,73. Opening of the BBB using LIPU/MB and other techniques has been used to successfully deliver circulating drugs into the GBM microenvironment12,13,74 and is recently being explored to enhance the sensitivity of liquid biopsy for these tumors75,76. Interestingly, we found that the total amount of EVPs isolated after sonication was at times less than the amount isolated before sonication in the same patient. Despite this, we observed that changes in the release of EVPs after BBB opening were predictive of GBM patient survival during PTX treatment in a prospective clinical trial, while measurements before BBB opening (“pre-son”) were not. This result underscores the complexity of measuring circulating tumor material such as EVPs, which can be affected by numerous factors such as location of the tumor relative to sonication, effectiveness of BBB opening, time between post-sonication and pre-sonication blood draws, and interpatient heterogeneity. Further studies that focus on BBB dynamics and release of tumor EVPs could provide additional insights into this phenomenon.
Our study has several limitations and raises additional questions that remain to be answered. The clinical trial cohort analyzed is relatively small, and independent validation of our findings in a larger prospective cohort is warranted before adoption of this technology as readout for GBM therapy response. Moreover, given that the biomarker required BBB opening using skull implantable ultrasound, this approach might have limited applicability for patients who do not have these implants. A granular molecular characterization of EVPs is challenging owing to the limited cargo present in them and would require advanced technologies for proteomics and nucleic acid studies that might not be available at all clinical institutions. Our results captured on the order of hundreds of peptides from circulating EVPs (similar to other recent studies77,78,79,80,81,82,83), which likely represents only a fraction of the diversity that can be measured from circulating tumor material. Though proteomic data from captured EVPs was consistent with gene expression profiles of GBM cells, analysis of a larger dataset would be important to confirm the origin of harvested material from within the tumor microenvironment. Future studies are warranted to understand how protein and nucleic acid content of these EVPs might change over time during therapy for GBM, and also expand the depth of material captured using our platform. Finally, the use of PS to isolate GBM EVPs bears some caveats. It is probable that the isolated material is not strictly limited to tumor-derived EVPs, as inflammation or stress response can also trigger expression of PS. Specific tumor-associated markers would need to be incorporated in those situations. Nonetheless, we still observe a clear biological signal in our data, suggesting clinical utility in the isolated species. Lastly, it is possible that PS+ EVPs might provide a response biomarker that applies to other therapies associated with glioma cell apoptosis beyond paclitaxel. Dedicated studies and analyses are warranted to investigate this.
In summary, we present an efficient microfluidic device for isolation of disease-specific EVPs in GBM patients and demonstrate that EVPs are predictive of PTX response after opening of the BBB. Application of this device in additional settings, such as different cytotoxic drugs and cancer types), could yield new insights into EVP biology and biomarkers of patient treatment response. With standardized collection protocols and collection of disease-specific material for characterization, EVP-based liquid biopsy holds promise to optimize the management and outcomes of patients with malignant glioma.
Methods
Patient enrollment and collection of samples
Specimens used in this project were obtained from the SC-9 phase I clinical trial performed at Northwestern University (NCT#04528680)12. Patients underwent informed consent for molecular analyses, and all procedures were approved by the Institutional Review Board of Northwestern University. Healthy control specimens were also recruited through an IRB protocol at the University of Michigan. An honest broker was used to de-identify subjects prior to EVP quantification (performed at University of Michigan) and analysis to protect patient privacy. Longitudinal specimens were collected during post-operative clinic visits for 18 patients. These typically included both a pre- and post-sonication plasma sample. For each specimen used in this study, 10–15 milliliters of blood were drawn in a Streck tube and then centrifuged at 1600 × g with slow deceleration. Subsequently, an additional spin was performed at 16,000 × g. Plasma was then harvested, split into aliquots, and stored at −80 °C for future EVP isolation.
Device fabrication and functionalization
A device mold was designed to optimally extract circulating EVPs from plasma and fabricated on a silicon wafer in the Lurie Nanofabrication facility at the University of Michigan, as discussed previously84. This mold was subsequently used to produce microfluidic devices for characterization of GBM. Our design includes 30 × 60 intermittent wells of diameter 500 µm each along the microfluidic channel. Higher cross-sectional area of wells reduces the flow velocity of medium to enhance the capture of EVPs in those regions. Each device top was prepared using polydimethylsiloxane (PDMS). Liquid PDMS (SYLGARD 184 Silicone Elastomer Base) was poured into the mold curing agent (SYLGARD 184) in the ratio of 10:1, degassed for 30 min, and cured in a 65 C oven for a minimum of 4 h. After solidification, holes were punched in the inlet and outlet (Uni-core-0.75, Pat #7093508). Glass slides (Fisher Scientific Cat #12-544-1) of size 75 × 25 mm were used for the device bottoms. PDMS and glass were treated with O2 plasma (Femto Science Covance). The two were bonded and placed on a 95 °C hot plate for 10 min to enhance adhesion. Functionalization steps were performed on the same day, including ethanol treatment, followed by 3-mercaptopropylmethyl-dimethoxysilane (Geleste, Inc. Cat #SIM6474.0-100GM) (1:25 in ethanol). Silane was incubated for an hour, followed by N-γ-maleimidobutyryl-oxysuccinimide ester (Thermo Fisher Scientific, Cat #22309) (3 μL per mL of ethanol) and incubated for 30 min. After washing the excess with ethanol, devices were incubated overnight with neutravidin (Thermo Fisher Scientific Cat # A2666) (1:10 in PBS) in a 4 °C fridge. Devices were then used the day after or stored for up to a month.
Harvesting EVs from GBM cell lines
The GBM6 and U87 cell lines were used to optimize GlioExoChip devices for harvesting of GBM-derived EVPs (Mayo Clinic and ATCC, respectively), and underwent authentication via Short Tandem Repeats. Both cells were cultured in complete media-90% DMEM media (Sigma Aldrich; Cat# 11995065) supplemented with 10% fetal bovine serum (Cat #F2442-500ML) and 1% antibiotic/antimycotic (Thermo Fisher Scientific; Cat #15240062) in a 37 °C incubator with 5% CO2. Three million cells were seeded in a 100 mm petri dish (Sarstedt Inc Cat# 50465226) for 24 h in complete media. The media was replaced by serum-free media after 3 PBS washes. Media with EVPs was collected after 50–72 h of incubation and centrifuged at 2000 × g for 15 min. Next, the supernatant was collected and centrifuged at 12,000 × g for 20 min. Finally, the supernatant was ultracentrifuged in two consecutive steps using a Thermo Fisher Sorvall WX+ Ultraseries ultracentrifuge. In the first step, 38 mL of liquid was centrifuged at 100,000 × g for 90 min. In the second step, 33 mL of liquid was replaced by the same volume of PBS and centrifugation was done under the same conditions as in the previous step. Once done, all liquid was gradually poured out, and 200 μL of PBS was used to collect pure EVPs in a vial. Samples were stored in −20 °C for short-term use (~ 2 weeks), or in −80 °C for long-term usage.
Sample preparation and experimental protocol for microfluidic capture using GlioExoChip
On the day of experiments, GlioExoChip devices were washed with PBS, followed by washing with a binding buffer. The binding buffer was prepared by diluting 10× binding buffer (BB, Cat #51-66121E) with DI water. After washing, 100 µL of 12 µg/mL of Annexin-V (Cat #556417) was injected into the devices and incubated for 40 min. A needle (Bstean 30-Gauge Cat #X0012ZMF33) was used to insert 3-inch tubings (Cole-Parmer Cat #AAD02091-CP) into the inlet and outlet sides. Syringe pumps were used to flow samples through each device (Harvard Apparatus PHD 2000). Plasma sample, originally stored in −80 °C was thawed on ice, then centrifuged at 3000 RPM (1811 × g) for 5 min and diluted in BB (30:270 µL). Samples were then pumped through each device for EVP capture, at a rate of 600 µL/h and 0.3 mL total volume. EVPs are captured when medium is passed through channels running between PDMS and a glass slide, while EV-depleted fluid exits at the distal end of each device. Washing was subsequently performed with 0.2 mL BB at 1 mL/h. Captured EVPs were released using 20 mM Ethylenediaminetetraacetic acid (EDTA) (prepared in DI water), at a flow rate of 1 mL/h and 0.3 mL total volume (0.15 mL injection, followed by 15 min incubation and 0.15 mL injection). Final washing was performed at 1.2 mL/h and EVPs were collected for further processing.
Nanoparticle tracking analysis
EVP quantification was performed using Particle Metrix Zetaview ×30 with the NTA module. Concentration versus size measurements were taken in triplicates and an average of the three readings has been reported. For each reading, 1 mL of diluted sample was prepared, based on ratios determined from sensitivity tests (typically in the range of 10–100). All measurements were performed using standardized settings: sensitivity = 100, shutter = 80, min brightness = 20, min size = 5, max size = 1000, positions = 11. Focus calibrations were completed using a standard suspension (1:250,000 diluted in DI water) of 100 nm polystyrene beads at the beginning of each set of runs. Videos were taken at 30 frames per second and analyzed using the Zetaview analysis software (version 8.05.12 SP1). Microsoft Excel was used for further data analysis to extract EVPs in the 30–150 nm size range. Two outlier measurement (values > 75 times the standard error) were removed as technical failures.
Capture efficiency, purity, and recovery calculations
Measurements of device performance and sample content were calculated according to the following formulas:
Fluorescent staining of extracellular vesicles and particles
On-chip staining of EVPs was performed using a lipophilic dye. After EVPs were captured and washed, a dye mixture was prepared using PBS and DiO (Cat #V22886, Invitrogen VybrantTM DiO cell-labeling solution) in the ratio of 100:1. The mixture was flown through the device at 20 µL/min (200 µL total) and then incubated at room temperature for 20 min. Excess dye was washed away with 0.3 mL PBS at a rate of 1 mL/h. Stained wells were imaged under a light microscope (Nikon Ti2 Eclipse) using a fluorescence channel (FITC, excitation wavelength of 488 nm). Images were post-processed using NIS-Elements AR Analysis software (version 4.00.03).
Western blot analysis
Protein analysis using Western blot was performed on relevant markers, including calnexin (a marker of cellular content), GFAP (a marker for glial cell origin), TSG101 (a marker of exosomes), and flotillin-1 (a marker of extracellular vesicles). For cell culture experiments, protein was extracted from 1 million pelleted cells using 100 µL of radio-immunoprecipitation assay (RIPA) buffer (Cat# 89900) and protease inhibitor cocktail (PIC, Cat# 1862209). Cells were resuspended and incubated in the buffer for 90 min on ice, followed by centrifugation at 8500 rpm (7674 × g) for 15 min. Supernatant was collected and stored in −80 °C for further use. For EVPs captured from GlioExoChip, 150 µL of RIPA and PIC was flown through the device at 20 µL/min. The first 30 µL was not collected, followed by a 50 µL injection that was incubated on ice for 20 min, and finally, the remaining 70 µL was flown. A total of 120 µL of protein extract was thereby collected for further processing. Samples underwent immediate protein quantification by Bichinchoninic acid (BCA) assay. A minimum of 5–10 µg of protein or a maximum of 37.5 µL of protein extract, whichever was more, was used for western blot (the minimum protein amount therefore, varied across samples). After protein quantification, 37.5 µL of original protein sample was combined with 12.5 µL of Laemmli mixture (90% of 4× Laemmli sample buffer (Cat# 1610747) and 10% of beta-mercaptoethanol (Cat# M3148-25ML)). Proteins were denatured at 100 °C for 7 min and immediately cooled on ice, followed by spin down and gentle vortex. We loaded 50 µL of sample and 7 µL of Precision-plus Protein Kaleidoscope onto a mini-PROTEAN TGX stain-free gel (Cat# 4568094; BioRad). Gel electrophoresis was run at 250 V for 27 min using PowerPac high-current electrophoresis equipment. The protein was then transferred to an Immuno-Blot PVDF membrane (Cat# 162-0261; BioRad) using the Trans-Blot Turbo Transfer system for 7 min. A set of two stacks (Cat# BR20221017) were used for the transfer arrangement and transfer buffer was prepared using Trans-Blot Turbo 5× transfer buffer (Cat# 10026938). The blot was blocked using EveryBlot blocking buffer (Cat# 12010020; BioRad) for 5 min at 4 °C, then replaced by primary antibody for overnight incubation (12 to 16 h) at 4 °C on a rocker (concentrations- 1:2000 GFAP rabbit polyAb (Cat# 16825-1-AP), 1:1500 Calnexin rabbit mAb (Cat# C5C9), and 1:1000 Flotillin-1 rabbit pAb (Cat# A6220), 1:500 TSG101 mouse mAb (Cat#SC-7964)). The following day, washing was done with 1× TBST (10× TBST buffer; Cat# 1706435, BioRad) diluted in DI water with 0.1% Tween-20 (Cat# 9005-64-5). This included 3 quick washes for 40–50 s each and 3 long washes for 10 min each in a rocking platform (speed 4) at room temperature. The blot was incubated in secondary antibody (1:20000 of anti-rabbit HRP conjugated (Cat# 7074S) for 90 min at 4 °C on a rocking platform (speed 3) or 1:2000 of anti-mouse HRP conjugated (Cat# HAF018), followed by washing as described above. After the last wash, the blot was incubated in SuperSignalTM West-Pico PLUS chemiluminescence substrate (Cat# 34577) for 5 min or SuperSignalTM West Femto (Cat# 34094) for 2–3 min. Imaging was done immediately after using the ChemiDoc Imaging system under colorimetric and chemiluminescent modes (auto rapid). Image files were post-processed on Sciugo portal. Typically, one blot was used for GFAP and Calnexin detection first, then reused for Flotillin-1, followed by TSG101 after stripping of antibodies. Stripping was performed by incubating the blot in Restore PLUS western blot stripping buffer (Cat# 46430) for 20 min on a rocking platform (speed 2.5) at room temperature. The blot was washed (as described above) and was ready to be reused.
BCA for total protein quantification
The ThermoScientific microBCATM Protein assay kit (Cat# 23235) was used for total protein quantification. Standard solutions of 200, 40, 20, 10, 5, 2.5, and 1 µg/mL were prepared as described in the manufacturer’s manual. Protein samples extracted from patient, healthy, and cell samples using GlioExoChip were diluted 1:4 to make 200 µL of final solution. Each sample was loaded in duplicate, with 100 µL of the diluted sample and 100 µL of working reagent (as prepared using manufacturer’s instructions) added to each well of a 96-well plate. After incubation for 2 h at 37 °C in darkness (no CO2), readings were taken using the BioTek Syngergy Neo2 Gen 5.3.1 plate reader. A protocol was set to measure absorbance at 532 nm wavelength and non-linear regression was used to generate a calibration curve. Total protein concentration is reported as the average of duplicates.
Scanning electron microscopy (SEM)
SEM imaging was performed on EVPs that were captured using GlioExoChip. SEM preparation steps included fixation of EVPs in 2% glutaraldehyde for 1 h on ice, followed by 1% Tannic acid for the 20 min at room temperature. Immediately after fixation, the PDMS and glass layers of GlioExoChip were separated and glass sections of 1 × 1 cm were cut out. Glass sections were submerged in PBS for 10 min. Next, dehydration steps were performed for 10 min each, including exposure to 50, 70, 95, 100, and 100% ethanol. The final steps of dehydration involved dipping glass slides in 50% hexamethyldisilazane (HMDS, Cat# 120581000) for 10 min, followed by 100% HMDS overnight. Samples were subsequently coated with 5 nm gold nanoparticles by sputtering at 18 mA for 90 s. SEM images were taken using TFS Nova 200 at 10 kV accelerating voltage and 0.4 nA current.
Proteomic profiling of plasma EVPs from GBM patients and healthy controls
Protein extraction was performed using on chip lysis (Pierce RIPA buffer; cat#89900) from pre-sonication GBM samples in treatment cycle #1 (P102, P106, P107, P113, P117, P118, P119) and healthy controls (HAM, HDH, HRA). Incubation on chip lasted 30 min and was performed without protease inhibitors. Samples (5.9 μg/condition) were submitted to the Proteomics Resource Facility at the University of Michigan for processing and mass spectrometry data acquisition. Briefly, upon reduction (5 mM DTT, for 30 min at 45 °C) and alkylation (15 mM 2-chloroacetamide, for 30 min at room temperature) of cysteines in samples, the proteins were precipitated by adding 6 volumes of ice-cold acetone followed by overnight incubation at −20 °C. The precipitate was spun down, and the pellet was allowed to air dry. The pellet was resuspended in 0.1 M TEAB and overnight (~ 16 h) digestion with trypsin/Lys-C mix (1:25 protease: protein; Promega) at 37 °C was performed with constant mixing using a thermomixer. The TMT 10-plex reagents were dissolved in 41 μl of anhydrous acetonitrile and labeling was performed by transferring the entire digest to TMT reagent vial and incubating at room temperature for 1 h. Reaction was quenched by adding 8 μl of 5% hydroxylamine and further 15 min incubation. Labeled samples were mixed, and dried using a vacufuge. An offline fractionation of the combined sample (~ 200 μg) into 8 fractions was performed using high pH reversed-phase peptide fractionation kit according to the manufacturer’s protocol (Pierce; Cat #84868). followed by an additional clean-up step with ZipTip. Samples were then run on the Orbitrap Ascend with FAIMSpro, including relative quantitation (TMT 10-plex) with 4 fractions. Fractions were dried and reconstituted in 9 μl of 0.1% formic acid/2% acetonitrile in preparation for LC-MS/MS analysis.
To obtain superior quantitation accuracy, we employed multinotch-MS385, which minimizes the reporter ion ratio distortion resulting from fragmentation of co-isolated peptides during MS analysis. Orbitrap Ascend Tribrid equipped with FAIMS source (Thermo Fisher Scientific) and Vanquish Neo UHPLC was used to acquire the data. Two μl of the sample was resolved on an Easy-Spray PepMap Neo column (75 μm i.d. × 50 cm; Thermo Scientific) at the flow-rate of 300 nl/min using 0.1% formic acid/acetonitrile gradient system (3–19% acetonitrile in 72 min; 19−29% acetonitrile in 28 min; 29–41% in 20 min followed by 10 min column wash at 95% acetonitrile and re-equilibration) and directly spray onto the mass spectrometer using EasySpray source (Thermo Fisher Scientific). FAIMS source was operated in standard resolution mode, with a nitrogen gas flow of 4.2 L/min, and inner and outer electrode temperature of 100 °C and dispersion voltage of −5000 V. Two compensation voltages (CVs) of −45 and −65 V, 1.5 s per CV, were employed to select ions that enter the mass spectrometer for MS1 scan and MS/MS cycles. Mass spectrometer was set to collect MS1 scan (Orbitrap; 400–1600 m/z; 120 K resolution; AGC target of 100%; max IT in Auto) following which precursor ions with charge states of 2–6 were isolated by quadrupole mass filter at 0.7 m/z width and fragmented by collision induced dissociation in ion trap (NCE 30%; normalized AGC target of 100%; max IT 35 ms). For multinotch-MS3, the top 10 precursors from each MS2 were fragmented by HCD followed by Orbitrap analysis (NCE 55; 45 K resolution; normalized AGC target of 200%; max IT 200 ms, 100–500 m/z scan range).
Proteome Discoverer (v3.0; Thermo Fisher) was used for data analysis. MS2 spectra were searched against SwissProt human protein database (20359 entries; Homo sapiens (sp_canonical TaxID = 9606) (v2024-03-27)) using the following search parameters: MS1 and MS2 tolerance were set to 10 ppm and 0.6 Da, respectively; carbamidomethylation of cysteines (57.02146 Da) and TMT labeling of lysine and N-termini of peptides (229.16293 Da) were considered static modifications; oxidation of methionine (15.9949 Da) and deamidation of asparagine and glutamine (0.98401 Da) were considered variable. Identified proteins and peptides were filtered to retain only those that passed ≤1% FDR threshold. Quantitation was performed using high-quality MS3 spectra (Average signal-to-noise ratio of 10 and <50% isolation interference). A total of 218 unique proteins were identified. PCA was performed on normalized protein abundances using RStudio (v2023.12.1 + 402), and ExpressAnalyst was used to perform differential gene expression. Differentially expressed genes were plotted using GraphPad Prism. The threshold for log2(fold change) was set at 1 or −1. Pathway analysis was performed using SRplot86. A heatmap of all genes was generated in the same tool using scaled protein abundances. KEGG pathway enrichment analysis was used to generate a bubble plot.
Correlation of EVP concentrations with outcome variables
The ratio in small EVP (30–150 nm size) concentration measured before (pre) and after (post) sonication was calculated for each longitudinal clinic visit for each patient (post divided by pre), using all paired (pre/post) datapoints. The timing of each blood draw was also noted relative to the initiation of sonication (Fig. 4C). To compare EVP ratios to MRI results, we quantified the level of MRI enhancement within the region of sonication as previously reported15. As MRI dates were typically not synchronous with EVP acquisition dates, we performed linear interpolation of enhancement values to calculate data on overlapping timepoints. To decipher meaningful associations between enhancement and EVP ratios, we only considered patients who had at least three longitudinal datapoints (EVP collections) for comparison. Separate analyses were performed using all remaining EVP data, vs. filtering to include only EVP samples that were collected within 75 min of sonication. The latter group contained data from six total patients after filtering, all of which are shown in Fig. 4F. An additional analysis was performed to correlate pre-sonication EVP concentrations (rather than post/pre ratios) with enhancement. To aggregate data across patients, we used the repeated measures correlation approach, implemented in the R package rmcorr42. To detect associations between EVP and patient survival, we calculated the change (or “slope”) of EVP release associated with sonication in treatment cycles #1 and #2. Specifically, the post/pre ratio from cycle #2 was subtracted from the post/pre ratio from cycle #1, and the resulting value was divided by the number of elapsed days between cycles #1 and #2, as shown below:
Analysis of GBM single cell RNA-seq data
Publicly available scRNA-seq data from GBM studies was analyzed on the GBMap platform41, which includes harmonized samples comprised of 1.1 M cells across 15 tissues. The dataset was filtered to include relevant biological cell populations that each had > 100 k cells available for analysis (malignant cells, glial cells, neural cells, macrophages, lymphocytes, microglial cells). The data plotted in Fig. 3E is normalized within each gene to show the range across cell types. The top three most differentially expressed genes (SERPINA3, APOC1, DMKN) between GBM and control EVPs were selected for plotting across cell types.
Measurement of EVP release after treating glioma cell lines with paclitaxel
The PDX cell line GBM43 was purchased from Mayo Clinic by independent investigators (denoted GBM43T and GBM43C), and after passage under distinct media conditions (serum-free media and complete media, respectively), was found to exhibit unique paclitaxel response curves upon testing. Cell lines were seeded on 6-well plates (four replicates per treatment group), with 120 k cells for each well. The next day, cells were washed three times with PBS, then incubated in their respective media with paclitaxel (0, 50, or 500 nM) for 50–72 h. The media was then collected and centrifuged at 2000 × g for 15 min, followed by supernatant collection (the cells were preserved for flow cytometry, discussed below). Supernatant was subsequently centrifuged at 12000 × g for 20 min, then frozen for NTA processing at the University of Michigan (same procedure as above).
Flow cytometry of GBM cells after paclitaxel treatment
The six-well plates used for EVP media collection were treated with 0.25% Trypsin-EDTA to detach cells. Apoptotic cells were then stained using APC-conjugated Annexin-V (Biolegend Cat. #640920) and Propidium Iodide (PI, Invitrogen Cat. # 00-6990-50) following the manufacturer’s protocol. Cell apoptosis was quantified using an LSRFortessa cell analyser (BD Biosciences) with the BD FACSDIVA software (BD Biosciences v 6.0), and analysis was carried out using FlowJo v10 to determine percentages of early apoptotic, late apoptotic, and total apoptotic cells.
Cell titer glow after paclitaxel treatment
Opaque-walled 96-well plates were used to grow the GBM43T and GBM43C cell lines in parallel with the 6-well plates used for EVP quantification and flow cytometry. All cell lines were plated on 96-well plates (three replicates per treatment group), and 4 k cells were seeded in each well. To assess cell line dose-response to PTX, cells were grown in 100 μL of media and treated with multiple PTX concentrations starting at 12.5 nM and increasing two-fold up to 1 μM (including the 500 nM treatment used for flow cytometry and exosome quantification). Cells were treated in the same conditions and timeline as for EVP release quantification. The Promega CellTiter-Glo (R) protocol was used to perform cell viability assays 72 h after treatment. CellTiter-Glo (R) reagent (3 mL) was diluted in DPBS (9 mL). 100 μL of the reagent was added to each of the wells and incubated on an orbital shaker for 2 min, followed by room temperature incubation for 10 min. Luminescence was then recorded by Gen5 on a Synergy 2 Microplate Reader (BioTek) for each well and values were normalized to untreated columns to determine cell viability after Paclitaxel treatment.
Paclitaxel cell death pathway inhibition
The role of specific cell death pathways in paclitaxel-induced cytotoxicity was assessed using pharmacologic inhibitors. GBM43C cells were seeded in triplicate at a density of 4000 cells per well and incubated for 24 h. Cells were pre-treated with pathway-specific inhibitors for 6 h prior to PTX administration. The following inhibitors were used: Z-VAD-FMK (50 µM; Promega) for apoptosis (pan-caspase inhibition), Necrostatin-1 (10 µM; Selleckchem) for necroptosis, Ferrostatin-1 (5 µM; MedChemExpress) for ferroptosis, Disulfiram (10 µM; Selleckchem) for pyroptosis, and 3-Methyladenine (2 mM; MedChemExpress) for autophagic cell death. PTX was then added at a final concentration of 12.5 nM to both inhibitor-treated and control wells. Cell viability was assessed 24 and 48 h after PTX treatment using the CellTiter-Glo® luminescent assay (Promega). Luminescence values were normalized to untreated control wells, and statistical comparisons between PTX-only and PTX + inhibitor conditions were performed using unpaired t-tests.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data are provided with this paper. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE87 partner repository with the dataset identifier PXD063771. Source data are provided with this paper.
References
Stupp, R. et al. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N. Engl. J. Med. 352, 987–996 (2005).
Brandes, A. A. et al. MGMT promoter methylation status can predict the incidence and outcome of pseudoprogression after concomitant radiochemotherapy in newly diagnosed glioblastoma patients. J. Clin. Oncol. 26, 2192–2197 (2008).
Mirimanoff, R. O. et al. Radiotherapy and temozolomide for newly diagnosed glioblastoma: recursive partitioning analysis of the EORTC 26981/22981-NCIC CE3 phase III randomized trial. J. Clin. Oncol. 24, 2563–2569 (2006).
Stupp, R. et al. Effect of tumor-treating fields plus maintenance temozolomide vs maintenance temozolomide alone on survival in patients with glioblastoma: a randomized clinical trial. JAMA 318, 2306–2316 (2017).
Mullins, M. E. et al. Radiation necrosis versus glioma recurrence: conventional MR imaging clues to diagnosis. AJNR Am. J. Neuroradiol. 26, 1967–1972 (2005).
Zeng, Q. S. et al. Multivoxel 3D proton MR spectroscopy in the distinction of recurrent glioma from radiation injury. J. Neurooncol. 84, 63–69 (2007).
Abouali, H., Hosseini, S. A., Purcell, E., Nagrath, S. & Poudineh, M. Recent advances in device engineering and computational analysis for characterization of cell-released cancer biomarkers. Cancers (Basel) 14, 288 (2022).
van Niel, G., D’Angelo, G. & Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 19, 213–228 (2018).
Onukwugha, N. E., Kang, Y. T. & Nagrath, S. Emerging micro-nanotechnologies for extracellular vesicles in immuno-oncology: from target specific isolations to immunomodulation. Lab Chip 22, 3314–3339 (2022).
Kok, V. C. & Yu, C. C. Cancer-derived exosomes: their role in cancer biology and biomarker development. Int. J. Nanomed. 15, 8019–8036 (2020).
Hoshino, A. et al. Extracellular vesicle and particle biomarkers define multiple human cancers. Cell 182, 1044–1061.e18 (2020).
Sonabend, A. M. et al. Repeated blood–brain barrier opening with an implantable ultrasound device for delivery of albumin-bound paclitaxel in patients with recurrent glioblastoma: a phase 1 trial. Lancet Oncol. 24, 509–522 (2023).
Carpentier, A. et al. Repeated blood-brain barrier opening with a nine-emitter implantable ultrasound device in combination with carboplatin in recurrent glioblastoma: a phase I/II clinical trial. Nat. Commun. 15, 1650 (2024).
Arrieta, V. A. et al. Ultrasound-mediated delivery of doxorubicin to the brain results in immune modulation and improved responses to PD-1 blockade in gliomas. Nat. Commun. 15, 4698 (2024).
Habashy, K. J. et al. Paclitaxel and carboplatin in combination with low-intensity pulsed ultrasound for glioblastoma. Clin. Cancer Res. 30, 1619–1629 (2024).
Zhang, D. Y. et al. Ultrasound-mediated delivery of paclitaxel for glioma: a comparative study of distribution, toxicity, and efficacy of albumin-bound versus cremophor formulations. Clin. Cancer Res. 26, 477–486 (2020).
Habashy, K. J. et al. Pharmacokinetic analysis of carboplatin and fluorescein brain retention following ultrasound-based blood-brain barrier opening. Clin. Cancer Res. 31, 3562–3570 (2025).
Kang, Y. T. et al. Dual-isolation and profiling of circulating tumor cells and cancer exosomes from blood samples with melanoma using immunoaffinity-based microfluidic interfaces. Adv. Sci. 7, 2001581 (2020).
Kang, Y. T., Hadlock, T., Jolly, S. & Nagrath, S. Extracellular vesicles on demand (EVOD) chip for screening and quantification of cancer-associated extracellular vesicles. Biosens. Bioelectron. 168, 112535 (2020).
Kang, Y. T. et al. On-chip biogenesis of circulating NK cell-derived exosomes in non-small cell lung cancer exhibits antitumoral activity. Adv. Sci. 8, 2003747 (2021).
Kang, Y. T. et al. Multiplex isolation and profiling of extracellular vesicles using a microfluidic DICE device. Analyst 144, 5785–5793 (2019).
Kim, B. et al. Palmitate and glucose increase amyloid precursor protein in extracellular vesicles: missing link between metabolic syndrome and Alzheimer’s disease. J. Extracell. Vesicles 12, e12340 (2023).
Hol, E. M. & Pekny, M. Glial fibrillary acidic protein (GFAP) and the astrocyte intermediate filament system in diseases of the central nervous system. Curr. Opin. Cell Biol. 32, 121–130 (2015).
Bonsergent, E. et al. Quantitative characterization of extracellular vesicle uptake and content delivery within mammalian cells. Nat. Commun. 12, 1864 (2021).
Kowal, J. et al. Proteomic comparison defines novel markers to characterize heterogeneous populations of extracellular vesicle subtypes. Proc. Natl. Acad. Sci. USA 113, E968–E977 (2016).
Meister, M. & Tikkanen, R. Endocytic trafficking of membrane-bound cargo: a flotillin point of view. Membranes 4, 356–371 (2014).
Raiborg, C. & Stenmark, H. The ESCRT machinery in endosomal sorting of ubiquitylated membrane proteins. Nature 458, 445–452 (2009).
Swairjo, M. A., Concha, N. O., Kaetzel, M. A., Dedman, J. R. & Seaton, B. A. Ca(2+)-bridging mechanism and phospholipid head group recognition in the membrane-binding protein annexin V. Nat. Struct. Biol. 2, 968–974 (1995).
Desai, T. J., Toombs, J. E., Minna, J. D., Brekken, R. A. & Udugamasooriya, D. G. Identification of lipid-phosphatidylserine (PS) as the target of unbiasedly selected cancer specific peptide-peptoid hybrid PPS1. Oncotarget 7, 30678–30690 (2016).
Lea, J. et al. Detection of phosphatidylserine-positive exosomes as a diagnostic marker for ovarian malignancies: a proof of concept study. Oncotarget 8, 14395–14407 (2017).
Kang, Y. T. et al. Isolation and profiling of circulating tumor-associated exosomes using extracellular vesicular lipid-protein binding affinity based microfluidic device. Small 15, e1903600 (2019).
Matsumura, S. et al. Subtypes of tumour cell-derived small extracellular vesicles having differently externalized phosphatidylserine. J. Extracell. Vesicles 8, 1579541 (2019).
Saadeh, F. S., Mahfouz, R. & Assi, H. I. EGFR as a clinical marker in glioblastomas and other gliomas. Int. J. Biol. Markers 33, 22–32 (2018).
Khushman, M. et al. Exosomal markers (CD63 and CD9) expression pattern using immunohistochemistry in resected malignant and nonmalignant pancreatic specimens. Pancreas 46, 782–788 (2017).
Willms, E. et al. Cells release subpopulations of exosomes with distinct molecular and biological properties. Sci. Rep. 6, 22519 (2016).
Yuan, Q. et al. Highly expressed of SERPINA3 indicated poor prognosis and involved in immune suppression in glioma. Immun. Inflamm. Dis. 9, 1618–1630 (2021).
Norton, E. S., Da Mesquita, S. & Guerrero-Cazares, H. SERPINA3 in glioblastoma and Alzheimer’s disease. Aging (Albany NY) 13, 21812–21813 (2021).
Luo, D. et al. Serpin peptidase inhibitor, clade A member 3 (SERPINA3), is overexpressed in glioma and associated with poor prognosis in glioma patients. Onco Targets Ther. 10, 2173–2181 (2017).
Hilbert, M., Kuzman, P., Mueller, W. C., Meixensberger, J. & Nestler, U. Feasibility of ApoC1 serum levels as tumor biomarker in glioblastoma patients: a pilot study. Sci. Rep. 12, 16981 (2022).
Zheng, X. J. et al. Apolipoprotein C1 promotes glioblastoma tumorigenesis by reducing KEAP1/NRF2 and CBS-regulated ferroptosis. Acta Pharm. Sin. 43, 2977–2992 (2022).
Ruiz-Moreno, C. et al. Charting the single-cell and spatial landscape of IDH-wild-type glioblastoma with GBmap. Neuro Oncol. 27, 2281–2295 (2025).
Bakdash, J. Z. & Marusich, L. R. Repeated measures correlation. Front. Psychol. 8, 456 (2017).
Dmello, C. et al. Translocon-associated protein subunit SSR3 determines and predicts susceptibility to paclitaxel in breast cancer and glioblastoma. Clin. Cancer Res. 28, 3156–3169 (2022).
Mair, R. & Mouliere, F. Cell-free DNA technologies for the analysis of brain cancer. Br. J. Cancer 126, 371–378 (2022).
Müller Bark, J., Kulasinghe, A., Chua, B., Day, B. W. & Punyadeera, C. Circulating biomarkers in patients with glioblastoma. Br. J. Cancer 122, 295–305 (2020).
Xu, M. et al. The biogenesis and secretion of exosomes and multivesicular bodies (MVBs): intercellular shuttles and implications in human diseases. Genes Dis. 10, 1894–1907 (2023).
Liangsupree, T., Multia, E. & Riekkola, M. L. Modern isolation and separation techniques for extracellular vesicles. J. Chromatogr. A 1636, 461773 (2021).
Reátegui, E. et al. Engineered nanointerfaces for microfluidic isolation and molecular profiling of tumor-specific extracellular vesicles. Nat. Commun. 9, 175 (2018).
Shao, H. et al. Protein typing of circulating microvesicles allows real-time monitoring of glioblastoma therapy. Nat. Med. 18, 1835–1840 (2012).
Shao, H. et al. Chip-based analysis of exosomal mRNA mediating drug resistance in glioblastoma. Nat. Commun. 6, 6999 (2015).
Chen, C. et al. Microfluidic isolation and transcriptome analysis of serum microvesicles. Lab Chip 10, 505–511 (2010).
Maccarrone, M., Melino, G. & Finazzi-Agrò, A. Lipoxygenases and their involvement in programmed cell death. Cell Death Differ. 8, 776–784 (2001).
Kenis, H. & Reutelingsperger, C. Targeting phosphatidylserine in anti-cancer therapy. Curr. Pharm. Des. 15, 2719–2723 (2009).
Nicholson, J. G. & Fine, H. A. Diffuse glioma heterogeneity and its therapeutic implications. Cancer Discov. 11, 575–590 (2021).
Qazi, M. A. et al. Intratumoral heterogeneity: pathways to treatment resistance and relapse in human glioblastoma. Ann. Oncol. 28, 1448–1456 (2017).
Arrieta, V. A. et al. ERK1/2 phosphorylation predicts survival following anti-PD-1 immunotherapy in recurrent glioblastoma. Nat. Cancer 2, 1372–1386 (2021).
Hegi, M. E. et al. MGMT gene silencing and benefit from temozolomide in glioblastoma. N. Engl. J. Med. 352, 997–1003 (2005).
Billan, S., Kaidar-Person, O. & Gil, Z. Treatment after progression in the era of immunotherapy. Lancet Oncol. 21, e463–e476 (2020).
Li, Y. et al. Advanced imaging techniques for differentiating pseudoprogression and tumor recurrence after immunotherapy for glioblastoma. Front. Immunol. 12, 790674 (2021).
Poelman, S. M. et al. Human breast cancer susceptibility to paclitaxel therapy is independent of Bcl-2 expression. Clin. Cancer Res. 6, 4043–4048 (2000).
Mimori, K. et al. Reduced tau expression in gastric cancer can identify candidates for successful Paclitaxel treatment. Br. J. Cancer 94, 1894–1897 (2006).
Li, Y. et al. Resistance to paclitaxel increases the sensitivity to other microenvironmental stresses in prostate cancer cells. J. Cell Biochem. 112, 2125–2137 (2011).
Manda, S. V. et al. Exosomes as a biomarker platform for detecting epidermal growth factor receptor-positive high-grade gliomas. J. Neurosurg. 128, 1091–1101 (2018).
Skog, J. et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat. Cell Biol. 10, 1470–1476 (2008).
Figueroa, J. M. et al. Detection of wild-type EGFR amplification and EGFRvIII mutation in CSF-derived extracellular vesicles of glioblastoma patients. Neuro Oncol. 19, 1494–1502 (2017).
Graner, M. W. et al. Proteomic and immunologic analyses of brain tumor exosomes. Faseb J. 23, 1541–1557 (2009).
Balaj, L. et al. Tumour microvesicles contain retrotransposon elements and amplified oncogene sequences. Nat. Commun. 2, 180 (2011).
Noerholm, M. et al. RNA expression patterns in serum microvesicles from patients with glioblastoma multiforme and controls. BMC Cancer 12, 22 (2012).
Peng, J. et al. Current understanding of exosomal microRNAs in glioma immune regulation and therapeutic responses. Front. Immunol. 12, 813747 (2021).
Batool, S. M. et al. Clinical utility of a blood based assay for the detection of IDH1.R132H-mutant gliomas. Nat. Commun. 15, 7074 (2024).
Osti, D. et al. Clinical significance of extracellular vesicles in plasma from glioblastoma patients. Clin. Cancer Res. 25, 266–276 (2019).
De Mattos-Arruda, L. et al. Cerebrospinal fluid-derived circulating tumour DNA better represents the genomic alterations of brain tumours than plasma. Nat. Commun. 6, 8839 (2015).
Miller, A. M. et al. Tracking tumour evolution in glioma through liquid biopsies of cerebrospinal fluid. Nature 565, 654–658 (2019).
Idbaih, A. et al. Safety and feasibility of repeated and transient blood–brain barrier disruption by pulsed ultrasound in patients with recurrent glioblastoma. Clin. Cancer Res. 25, 3793–3801 (2019).
Meng, Y. et al. MR-guided focused ultrasound liquid biopsy enriches circulating biomarkers in patients with brain tumors. Neuro Oncol. 23, 1789–1797 (2021).
Zhang, D. Y. et al. Ultrasound-mediated blood-brain barrier opening increases cell-free DNA in a time-dependent manner. Neurooncol. Adv. 3, vdab165 (2021).
Tamarindo, G. H. et al. Distinct proteomic profiles of plasma-derived extracellular vesicles in healthy, benign, and triple-negative breast cancer: candidate biomarkers for liquid biopsy. Sci. Rep. 15, 12122 (2025).
Muraoka, S. et al. Proteomic profiling of extracellular vesicles separated from plasma of former National Football League players at risk for chronic traumatic encephalopathy. Aging Dis. 12, 1363–1375 (2021).
Torres, A. et al. Comparing the proteomic profiles of extracellular vesicles isolated using different methods from long-term stored plasma samples. Biol. Proced. Online 26, 18 (2024).
Sundar, I. K., Li, D. & Rahman, I. Proteomic analysis of plasma-derived extracellular vesicles in smokers and patients with chronic obstructive pulmonary disease. ACS Omega 4, 10649–10661 (2019).
Su, H. et al. Multiomics analysis to evaluate the enrichment of extracellular vesicles from human plasma. J. Lipid. Res. 66, 100877 (2025).
Veerman, R. E. et al. Molecular evaluation of five different isolation methods for extracellular vesicles reveals different clinical applicability and subcellular origin. J. Extracell. Vesicles 10, e12128 (2021).
Suresh, P. S. & Zhang, Q. Comprehensive comparison of methods for isolation of extracellular vesicles from human plasma. J. Proteome Res. 24, 2956–2967 (2025).
Kanwar, S. S., Dunlay, C. J., Simeone, D. M. & Nagrath, S. Microfluidic device (ExoChip) for on-chip isolation, quantification and characterization of circulating exosomes. Lab Chip 14, 1891–1900 (2014).
McAlister, G. C. et al. MultiNotch MS3 enables accurate, sensitive, and multiplexed detection of differential expression across cancer cell line proteomes. Anal. Chem. 86, 7150–7158 (2014).
Tang, D. et al. SRplot: a free online platform for data visualization and graphing. PLoS ONE 18, e0294236 (2023).
Perez-Riverol, Y. et al. The PRIDE database at 20 years: 2025 update. Nucleic Acids Res. 53, D543–d553 (2025).
Acknowledgements
This work was supported by National Institutes of Health 1R01CA245969-01A1 (A.M.S. & R.S.), P50CA221747 (A.M.S. & R.S.), 5-R33-CA-202867 (S.N.), 1-R01-CA-208335-01-A1 (S.N.), and 1U19CA264338-01 (A.M.S. & R.S.). AMS also received funding support from the Lou and Jean Malnati Brain Tumor Institute, philanthropic support from the Moceri Family Foundation in memory of Sharon Moceri, as well as Tina and Vic Kedaitis. The authors acknowledge the financial support from Forbes Institute for Cancer Discovery at University of Michigan. This work was also supported by the United States Department of Defense (M.W.Y.; CA230856), an American Brain Tumor Association Basic Research Fellowship (M.W.Y.; fully supported by Tap Cancer Out), and the Focused Ultrasound Foundation (A.S.). The Michigan Center for Materials Characterization (MC)2 is acknowledged for their assistance with electron microscopy. The authors thank Venky Basrur from the Proteomics Core Facility (Michigan Medicine) for his help with planning and design of proteomics experiments, Sarah Spanninga from the Biointerfaces Institute (Nanotechnicum) for her help with NTA measurements, and the Deepak Nagrath lab members for their support with protein analysis. Sponsors had no role in study design, data collection and analysis, or manuscript writing.
Author information
Authors and Affiliations
Contributions
A.K. and M.W.Y. contributed equally to the work. A.K. conceptualized, optimized, and conducted all experiments with plasma. M.W.Y. conceptualized and conducted all analyzes, and in-vitro experiments. Y.T.K. led preliminary experiments in this study. A.G., K.H., M.G., H.L., T.M., T.K.S., and C.D. helped with wet-lab experiments. A.G., K.H., L.C., R.W., K.M., K.C.P., E.M.T.P., A.Z., M.N., S.V., D.P., K.D., R.V.L., P.K., G.B., M.C., C.A., and R.S. assisted with clinical specimen collection, processing, and analysis. H.C. and H.Z. helped with statistical analyses. A.M.S. and S.N. supervised the work.
Corresponding authors
Ethics declarations
Competing interests
A.M.S. is a paid consultant for Carthera and EnClear Therapies. A.M.S. and R.S have received funding support from Carthera and Agenus Bio, and in-kind support by Agenus, Carthera, and BMS. R.S. serves as a member of the scientific advisory board of Alpheus Medical, Carthera, and has provided consultancy to AstraZeneca, Boston Scientific, Novartis and Novocure. RS has equity in Alpheus Medical and Carthera. M.C. and G.B. are full-time employees of Carthera. M.C. has patents and equity interests in Carthera related to this technology. All other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Agnieszka Bronisz, Peter Canoll and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Youngblood, M.W., Kumari, A., Kang, YT. et al. Dynamic release of extracellular particles after opening of the blood-brain barrier predicts glioblastoma susceptibility to paclitaxel. Nat Commun 16, 11045 (2025). https://doi.org/10.1038/s41467-025-65681-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65681-4