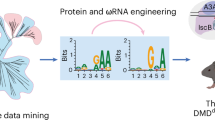
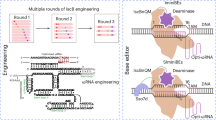
Abstract
IscB, as the putative ancestor of Cas9, possesses a compact size, making it suitable for in vivo delivery. OgeuIscB is the first IscB protein known to function in eukaryotic cells but requires a complex TAM (NWRRNA). Here, we characterize a CRISPR-associated IscB system, named DelIscB, which recognizes a flexible TAM (NAC). Through systematically engineering its protein and sgRNA, we obtain enDelIscB with an average 48.9-fold increase in activity. By fusing enDelIscB with T5 exonuclease (T5E), we find that enDelIscB-T5E displays robust efficiency comparable to that of enIscB-T5E in human cells. Moreover, by fusing cytosine or adenosine deaminase with enDelIscB nickase, we establish efficient miniature base editors (ICBE and IABE). Finally, we efficiently generate mouse models by microinjecting mRNA/sgRNA of enDelIscB and enDelIscB-T5E into mouse embryos. Collectively, our work presents a set of enDelIscB-based miniature genome-editing tools with great potential for diverse applications in vivo.
Similar content being viewed by others
Introduction
The clustered regularly interspaced short palindromic repeats (CRISPR) and CRISPR-associated proteins (Cas) systems as prokaryotes’ adaptive immune systems against invading phages or plasmids have been harnessed for diverse genome manipulation in eukaryotes1,2,3. The CRISPR system is mainly divided into class1 and class2 according to whether the Cas proteins are multiple subunits. The class1 includes three subtypes (type I, III, and IV) and class2 includes another three subtypes (type II, V, and VI)4. Among them, Cas9 from type II and Cas12a from type V are the most widely used gene editors in eukaryotic cells due to their simple composition, which includes a single Cas effector and a single guide RNA (sgRNA) to introduce site-specific DNA double-strand breaks (DSB)1,5,6. The cells then repair the DSB through either an error-prone non-homology end joining repair (NHEJ) pathway to disrupt the target gene or a precise homology-directed repair (HDR) pathway for precise gene editing7. In addition to DSB-dependent gene editing tools, DSB-independent genome engineering tools have been developed based on Cas9 nickase or dead Cas fusion with other elements, including base editors (BEs), primer editors (PEs), and epigenetic editors for safer therapeutic purposes8,9. However, the large size of the initially identified Cas9 or Cas12a proteins (generally more than 1000 amino acids [aa]) makes them or their derived editors difficult to deliver with a single adeno-associated virus (AAV). AAV is one of the most commonly utilized nucleic acid drug delivery vehicles approved by the FDA, but it has a loading ceiling of only 4.7 kb10. Recently, a series of compact RNA-guided nucleases (RGNs) suitable for AAV delivery have been developed, including the homologs of Cas12a and Cas9, such as Cas12j (700–800 aa)11, Cas12f (400–700 aa)12,13,14,15, Cas12n (400–700 aa)16, Cas9d (450–1050 aa)17. In addition, their ancestral protein TnpB (~400 aa) and IscB (~400 aa)18,19,20, and the TnpB’s eukaryotic homolog-Fanzor (400–700 aa) have been also described21,22.
IscB (insertion sequences Cas9-like OrfB) proteins as predicted ancestors of Cas9, consisting of ~400 aa and exploits a single noncoding RNA named ωRNA for RNA-guided cleavage of double-stranded DNA with appropriate target-adjacent motif (TAM)18. IscB protein shared a Cas9-liked domain composition (Fig. 1a), which contains both RuvC and HNH endonuclease domain to cleave the non-target strand (NTS) and the target strand (TS) respectively23. Therefore, compared with Cas12-like miniature proteins, which only have one RuvC endonuclease domain to cleave two DNA strands, IscB is more feasibly converted into a nickase to establish compact BEs or PEs. OgeuIscB is the first identified IscB protein that can induce insertions/deletions (indels) in human cells, although its editing efficiency is rather low18. Han et al. engineered OgeuIscB protein and its ωRNA in human cells and named the engineered system enIscB. When enIscB fused with T5 exonuclease (T5E), its resulting efficiency is further increased, which is comparable to that of SpG Cas9 and reduces chromosome translocation effects24. Moreover, efficient miniature base editors (miBEs) were generated through the fusion of cytosine or adenosine deaminase with enIscB nickase24. Subsequently, several other research teams reported similar efforts in engineering the OgeuIscB system and developing base editors25,26,27,28. However, it remains unclear whether other IscB systems that recognize different TAMs can be engineered into effective gene editors.
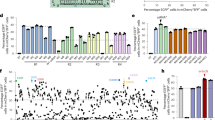
a Comparison of size and domain architecture of different Cas9s and IscBs. b Schematic illustration of the plasmid interference assay. This schematic diagram is adapted from Fig. 1B of the paper by Song et al.48, which is published under a CC BY 4.0 Attribution 4.0 International license (https://creativecommons.org/licenses/by/4.0/). c, d Representative E. coli clone images from plasmid interference assays. T targeting, NT non-targeting; 16 nt/20 nt/24 nt (guide lengths). NNLS N-terminal nuclear localization signal, CNLS C-terminal nuclear localization signal. Each experiment was repeated independently two times with similar results. e Procedure for detecting IscB cleavage activity in HEK293T cells using a fluorescent reporter system, which contains an editor plasmid with IscB-T5 exonuclease(T5E), sgRNA, and mCherry, and a reporter plasmid with BFP and EGFxxFP. NLS, nuclear localization signal. T2A, a self-cleaving 2A peptide from Thosea asigna virus. f Fluorescent images of HEK293T cells co-transfected with editor plasmids and reporter plasmids. The fluorescent channels are shown at the top of the figure. BF, bright field. The scale bars represent 100 μm. g The representative FACS analysis of IscB-T5E with non-target (NT) or target sgRNA. h Flow cytometry calculated the percentage of EGFP-activated (EGFP+) cells in both mCherry and BFP-activated (mCherry+BFP+) cells. Data represent mean ± s.d. of three independent biological replicates. Source data are provided as a Source data file.
In this study, we focus on a CRISPR-associated IscB system identified from the Delaware Bay aquatic sample metagenome18, so we abbreviate it as DelIscB. Altae-Tran et al. have determined the TAM and the noncoding RNA (ncRNA) for DelIscB and confirmed its enzymatic activity with an in vitro cleavage assay18, but whether it functions in mammalian cells remains unknown. DelIscB recognizes a more flexible TAM (NAC) and is more compact in size, with only 457 residues. In contrast, OgeuIscB requires a complicated TAM (NWRRNA) and is composed of 496 residues18. In our study, we initially discovered that DelIscB could efficiently cleave the plasmids in E. coli through a plasmid interference assay. We then truncated its ncRNA and removed the polyT motif, expecting it to be able to function in eukaryotic cells. We then determined that DelIscB was able to cleave fluorescent reporter plasmids albeit with limited efficiency in HEK293T cells. Subsequently, we engineered both the protein and sgRNA of DelIscB and named the engineered system enDelIscB. The enDelIscB remarkably increased the cleavage activity across 24 endogenous gene loci by an average of about 48.9-fold compared to the initial version. When enDelIscB fused T5E, its activity is comparable to that of enIscB-T5E. At the same time, we evaluated the specificity of both editors. Moreover, we developed enDelIscB nickase-based adenine and cytosine base editors (named IABE and ICBE, respectively) which exhibit remarkable base editing efficiency. Finally, we investigated the activity of enDelIscB and enDelIscB-T5E in mouse cells and embryos, and efficiently generated mouse models. In summary, our research not only expands the compact gene-editing toolbox for fundamental research and therapeutic applications but also offers valuable perspectives for efficiently engineering RGNs.
Results
DelIscB enables programmed plasmid DNA cleavage in E. coli and human cells
To assess the activity of IscB proteins, we developed a plasmid interference assay in E. coli. This assay contains two compatible plasmids, each conferring a distinct antibiotic resistance (Fig. 1b). Specifically, one plasmid (P1) expresses Cas9 or IscB protein, while the other plasmid (P2) expresses sgRNA or ωRNA and simultaneously provides targeted sites or non-targeted sites as control. After co-transfection of the two plasmids to E. coli, E. coli clones cannot grow on agar plates containing both antibiotics once the P2 plasmid is cleaved. Initially, we examined the activity of AwaIscB and OgeuIscB in cleaving plasmids in E. coli at their optimal length of guide sequences, with SpCas9 as a positive control. We found that AwaIscB and OgeuIscB did not efficiently cleave the plasmids in E. coli, whereas SpCas9 could (Fig. 1c). Inspired by the ability of DelIscB to cleave the PAM library plasmids in E. coli to determine TAM18, we hypothesized that DelIscB may exhibit more efficient plasmid-cleaving activity. We then performed the same plasmid interference experiment on DelIscB and found that it was able to cleave plasmids using a 16-nt guide length, and its efficiency is similar to that of SpCas9 (Fig. 1c).
Before validating the activity of DelIscB in human cells, we performed an in-depth analysis of its ncRNA. Unlike the ωRNA of OgeuIscB, the ncRNA of DelIscB contains spacers and repeats similar to those in CRISPR arrays, along with an anti-repeat region (Supplementary Fig. 1a). Combining the secondary structures predicted by RNAfold and Altae-Tran et al.18, we hypothesized that the hairpin formed by the direct repeat (DR)/anti-repeat duplex could be truncated to facilitate its transcription in human cells. Therefore, we established a series of truncations of varying lengths around the DR/anti-repeat region and found that certain truncates were able to retain the function of the plasmid cleavage (Supplementary Fig. 1b). We selected a relatively shorter yet functional truncation (Δ20’−121’ + GAAA) to serve as the template for subsequent engineering and named it sgRNA-V0.1. Considering that Pol III pauses at the poly-T signal when using the U6 promoter to transcribe sgRNA in mammalian cells29, we mutated poly-T motif by U136’A in sgRNA-V0.1, and named this mutated sgRNA as sgRNA-V1, which retained its cleavage activity (Supplementary Fig. 1b). Furthermore, to investigate whether the stem-loop structures in other regions are functionally redundant, a series of truncations were performed based on either sgRNA-V0.1 or sgRNA-V1 (Supplementary Fig. 1b). However, these truncated variants exhibited a loss of functionality, which implies that either these regions are essential for the function or require more precise truncations based on the actual three-dimensional structure (Supplementary Fig. 1b).
Next, we expressed wild type (WT)-DelIscB fused with nuclear localization signals (NLS) at both terminals and co-expressed sgRNA-V1 targeting different gene loci in HEK293T cells, and with SpCas9 serving as a positive control. The T7 endonuclease I (T7EI) assay is commonly used for the rapid detection of indels introduced by gene editing in the genome. Its core principle relies on the specific recognition and cleavage of mismatched regions in double-stranded DNA by T7EI. By analyzing the proportion of cleavage products through gel electrophoresis, the editing efficiency can be evaluated. However, the results of T7EI assay showed that WT-DelIscB+sgRNA-V1 did not exhibit significant genome cleavage activity at six genomic loci (Supplementary Fig. 1c). We hypothesized that the fusion of NLS at both termini of WT-DelIscB might impair its activity and the activity of WT-DelIscB is too weak to effectively cleave the genome DNA. Therefore, we conducted plasmid interference experiments to verify whether the NLS fused at the N-terminus or C-terminus of DelIscB impacts its activity to cleave the target plasmid. Our data indicated that fusion with NLS at the N-terminus of DelIscB inhibits it from effectively cleaving the plasmid, suggesting that the exposed N-terminus of DelIscB may be essential for its cleavage activity (Fig. 1d).
By using a fluorescent reporter system, which is sensitive in detecting the cleavage activity of the IscB system in mammalian cells24, we found that WT-DelIscB-CNLS+sgRNA-V1 was able to cleave plasmids in human cells with an efficiency of merely approximately 5%. After fusion with T5E, the relevant fluorescence signal was enhanced to 15% (Fig. 1e-h, Supplementary Fig. 1d). Using this fluorescence reporter system, we further explored the optimal guide length of the sgRNA of DelIscB. Our data showed that the optimal length is 18-nt at the first reporter sequence (EGFxxFP-traget1) and 16-nt at the other two reporter sequences (EGFxxFP-traget2/3.1) (Supplementary Fig. 1e).
Furthermore, we used this fluorescence reporter system to reconfirm that fusion with the NLS at the N-terminus of DelIscB results in activity reduction, even in different forms of fusion (Supplementary Fig. 1f). Notably, the reduction in activity after removing NLS from the N-terminal of enIscB-T5E was relatively limited (Supplementary Fig. 1f). We further re-quantified the previous sgRNA modifications using this fluorescent reporter system. Our findings indicated that both the truncation in the DR/anti-repeat region and the mutation of U136’A rendered the sgRNA of DelIscB more amenable to function in mammalian cells (Supplementary Fig. 1g). However, additional detailed truncations did not display activity superior to that of sgRNA-V1 (Supplementary Fig. 1g). In summary, our data indicated that fusing NLS at the N-terminus of DelIscB diminishes its activity, whereas fusing T5E at its C-terminus increases its activity. The currently obtained sgRNA-V1 version was used for subsequent experiments.
Engineering the DelIscB protein to enhance its activity in mammalian cells
Given that DelIscB-T5E+sgRNA-V1 demonstrated only 15% plasmid cleavage activity in HEK293T cells, its activity remains notably low when compared to enIscB-T5E. As a result, we began to explore strategies for engineering the protein of DelIscB to boost its activity. Firstly, considering enIscB as the first highly active IscB protein24, we wondered whether it would be feasible to map the seven highly active substitutions in enIscB (K39R, E85R, D97R, H369R, S387R, A402R, and S457R) to DelIscB. Therefore, we employed AlphaFold2 (AF2)30,31 to predict the protein structure of DelIscB and aligned it with the structure of OgeuIscB (7UTN)23. We found five corresponding amino acid residues (Y37, Q85, S97, K350 and A404) in DelIscB (Fig. 2a). It is worth noting that the predicted structure of DelIscB revealed the absence of the P1 stem-loop interaction domain (P1D) compared to OgeuIscB (Supplementary Fig. 2a-c). Consequently, the amino acids corresponding to S387 and A402 of OgeuIscB could not be located within the structure of DelIscB (Fig. 2a). We then replaced the five corresponding residues (Y37, Q85, S97, K350, and A404) with arginine. The results showed that only one mutation, S97R, led to a 2-fold change in activity, while the other four substitutions (Y37R, Q85R, K350R, and A404R) all resulted in reduced activity (Fig. 2b).
a An alignment was made between the predicted protein structure of DelIscB and the cryo-electron microscopy (cryo-EM) structure (7UTN) of OgeuIscB-ωRNA bound to the dsDNA target, to identify the amino acids on DelIscB that correspond to the highly active mutant residues of enIscB. b Arginine was used to replace the five amino acids of DelIscB, which structurally correspond to the highly active mutations of enIscB. Data represent mean ± s.d. of three independent biological replicates. c Multiple sequence alignment of DelIscB with its 10 homologous proteins. Representative regions are displayed, and candidates for mutagenesis are highlighted in red or cyan boxes. The type of mutant is shown at the top of the box, with red indicating a mutation to arginine or lysine and cyan indicating a mutation to other amino acid types. d Substitutions of amino acid residues of DelIscB protein. Each dot represents activity for a single variant. The dashed line represents a 1.2-fold change in activity relative to the wild type. Data represent mean ± s.d. of two independent biological replicates. e The activity of a series of mutant combinations was detected with reporter sequence EGFxxFP-target1. Data represent mean ± s.d. of three independent biological replicates. f The activity of a series of mutant combinations was confirmed with another reporter sequence (EGFxxFP-target2). Data represent mean ± s.d. of three independent biological replicates. Source data are provided as a Source data file.
Moreover, acknowledging that recent approaches for engineering AsCas12f1 or OgeuIscB involve enhancing the protein’s affinity to DNA or sgRNA through arginine or lysine substitution, we decided to adopt a similar strategy24,32. Given the lack of actual structural information, to identify the appropriate positions of DelIscB for arginine substitution, we first performed a phylogenetic tree analysis of the known IscB proteins18 (Supplementary Fig. 3). Subsequently, we selected 10 IscB proteins closely related to DelIscB for multiple sequence alignment (MSA) (Supplementary Fig. 4). Based on the result of MSA, we focused on selecting neutral or negatively charged residues in DelIscB. If the corresponding aligned positions in DelIscB homologs harbored positively charged residues, we mutated the residues to arginine and generated a total of 122 single mutations. Among these, eight mutants (C12R, Q192R, E317R, Q340R, A343R, A364R, V396R, and S436R) exhibited activity changes of more than 1.2-fold (Fig. 2c, d). During the process of observing the MSA result, we noticed that at certain alignment positions, DelIscB had unique residues distinct from those of its homologous proteins. We then attempted to mutate these unique residues into the ones that were more prevalent at the alignment positions. By using this strategy, we constructed 66 mutants and obtained 5 mutants (L90E, S100A, G104S, W117F, and A142P) with activity changes exceeding 1.2-fold (Fig. 2c, d).
Notably, among the 14 variants screened above, the S97R variant exhibited the highest activity (Fig. 2d). We also noticed that a lysine residue was present at the position aligned with S97 in its homologs (Fig. 2c). Therefore, we examined the activity of S97K and found that its activity was slightly higher than that of S97R (Supplementary Fig. 5a). We further examined the activity of the corresponding lysine replacement mutants (C12K, Q192K, E317K, Q340K, A343K, A364K, and V396K) of the other 7 arginine replacement mutants (Supplementary Fig. 5a). Additionally, considering the obtained highly active variants may be sequence-dependent, we employed two additional reporter sequences to further verify the activity of the aforementioned mutants. The results showed that, under the concurrent validation of the three reporter sequences, the activity change of all mutants except C12K, Q192K, E317K, and L90E was more than 1-fold (Supplementary Fig. 5a–c). After calculating the average fold change in activity of the above mutants across the three reporter sequences, we ultimately selected 14 mutants (S97R/K, V396R/K, E317R, C12R, A364R, Q340R/K, W117F, A142P, S100A, G104S, S436R) with an average increase of over 1.2-fold for subsequent combinations (Supplementary Figs. 5d and 2e). The outcomes of the mutant combinations revealed that a combination of 10 variants (S97K, V396K, E317R, C12R, Q340K, W117F, A142P, S100A, G104S, S436R) displayed the highest activity (Fig. 2e). We named this combination DelIscB-V10.1, which is approximately 5-fold more active than wild-type and even more active than enIscB-T5E under characterization using the reporter sequence EGFxxFP-target1 (Fig. 2e). We also validated the activity of the combined mutants under another reporter sequence (EGFxxFP-target2) and found that DelIscB-V10.1 still performed optimally (Fig. 2f). In summary, through screening a vast number of mutants and combining highly active mutants, we have successfully obtained the currently best-performing protein version, DelIscB-V10.1.
Engineering the DelIscB sgRNA scaffold to further increase activity
Previous reports have indicated that truncating the stem-loop structures in different regions of the sgRNA in AsCas12f1 and introducing mutations to enhance the stability of the ωRNA in OgeuIscB can contribute to enhancing their activity15,24,33. For further optimizing the DelIscB sgRNA scaffold, WT-DelIscB-T5E paired with sgRNA-V1 served as the baseline control. Based on the predicted secondary structure of sgRNA-V1 (Fig. 3a), we first truncated its stem-loop structures to different extents (Fig. 3b). Among those, the truncations at stem1.3 and stem2 led to an increase in activity, i.e., V1 + Δ35-41 + GAAA and V1 + Δ67-71 + GAAA, which we named sgRNA-V2 and sgRNA-V2.1, respectively (Fig. 3b). Although the truncated body V1 + Δ221-240 + GAAA at stem6 did not significantly alter its activity, considering its ability to substantially reduce the total length of sgRNA, we still incorporated it into the subsequent study and named it sgRNA-V2.2 (Fig. 3b).
a The predicted secondary structure of DelIscB sgRNA-V1 contains a total of 267 nucleotides (nt). Among them, the secondary structure of 1 to 207 nt is plotted according to the ncRNA (CRISPR-associated IscB) predicted structure of Altae-Tran et al.18, and the secondary structure of 208 to 267 nt is predicted according to Mfold (Supplementary Fig. 6). b Several predicted stem-loops of sgRNA-V1 were truncated to enhance its activity. The dashed line represents the cleavage activity of WT-DelIscB-T5E+sgRNA-V1, i.e., 1-fold change. c To improve the stability of sgRNA-V1, the A-U or G-U weak base pairs in stem regions were replaced with G-C stronger base pairs. d To improve flexibility at the sgRNA-V1, the G-U base pair, or several other types of base pairs in the pseudoknot region were replaced with unpaired bases. e Combinations of effective truncated and mutant versions of sgRNA-V1. f Combinations of engineered protein and sgRNA versions of DelIscB with or without T5E. All data represent mean ± s.d. of three independent biological replicates. Source data are provided as a Source data file.
Next, aiming to enhance the stability of sgRNA-V1, we replaced the relatively weak base pairs A-U or G-U with the strong base pair G-C or paired several bases that were unpaired at the pseudoknot. We found that the activities of the two mutants, V1 + U15C + A28G and V1 + U65C + A73G, increased by more than one-fold. Therefore, we named them sgRNA-V3 and sgRNA-V3.1 respectively for further investigation (Fig. 3c).
During the process of stabilizing sgRNA, we found that stabilization mutations at the pseudoknot, such as U205C and A206G, actually decreased the activity. Additionally, inspired by the finding that increased flexibility of CasX sgRNA can enhance its activity34, we hypothesized that introducing a series of mismatched mutations at the pseudoknot of sgRNA-V1 might increase its flexibility. Our experimental data revealed that the U205A mutation resulted in an activity increase of more than 1.5 times and the resulting variant was named sgRNA-V4 (Fig. 3d).
Subsequently, we combined the different versions of the sgRNA aforementioned and found that the combination of sgRNA-V4/V2/V2.2 maximally increases its activity, which we named sgRNA-V5 (Fig. 3e). Finally, we combined sgRNA-V5 with DelIscB-V10.1-T5E and DelIscB-V10.1, respectively. Remarkably, the activity of each combination was further augmented (Fig. 3f). We named these two final combinations as enDelIscB-T5E and enDelIscB. Under the reporter sequence EGFxxFP-target1, both enDelIscB-T5E and enDelIscB exhibited higher activity than enIscB-T5E (Fig. 3f).
In addition, we utilized two additional reporter sequences (EGFXXFP-target2 and EGFXXFP-target3) to compare the activities of the different optimized sgRNA versions. The results indicated that, consistent with the findings validated by the reporter sequence EGFXXFP-target1 (Fig. 3e), all the different optimized versions (except sgRNA-V2.2) demonstrated varying degrees of activity improvement compared to sgRNA-V1 (Supplementary Fig. 7a, b). Among them, sgRNA-V5 maintained the highest performance (Supplementary Fig. 7a, b). This suggests that the series of optimized sgRNAs we screened are not dependent on the target context.
Furthermore, we compared the activity of sgRNA-V1 and sgRNA-V5 in combination with DelIscB-V10.1-T5E using two additional report sequences (EGFXXFP-target2 and EGFXXFP-target3) (Supplementary Fig. 7a, b). We found that when combined with DelIscB-V10.1-T5E, sgRNA-V5 showed only a slight increase in activity compared to sgRNA-V1 (Supplementary Fig. 7a, b), which is consistent with the performance observed under the reporting sequence EGFXXFP-target1 (Fig. 3f). Meanwhile, we also compared the activities of DelIscB-V10.1 combined with sgRNA-V1 and sgRNA-V5 under three reporter sequences (Supplementary Fig. 7c–e). The results showed that the activity of sgRNA-V5 was slightly higher than that of sgRNA-V1 under all three reporter sequences (Supplementary Fig. 7c–e). Collectively, sgRNA-V5 demonstrates marginally enhanced efficacy compared to sgRNA-V1 when co-expressed with either DelIscB-V10.1-T5E or DelIscB-V10.1. However, this enhancement is not as prominent as when combined with WT-DelIscB-T5E. We speculate that DelIscB-V10.1 has enhanced its activity to a level approaching saturation.
Evaluating the genome editing efficiency and precision of enDelIscB and enDelIscB-T5E
To evaluate the genome editing activity of enDelIscB and enDelIscB-T5E at endogenous gene loci, all-in-one plasmids expressing enDelIscB/enDelIscB-T5E, sgRNA, and mCherry were constructed, and then transfected into HEK293T cells. Forty-eight hours post-transfection, cells were harvested and the top 25% of mCherry-positive cells were sorted via flow cytometry. These sorted cells were then subjected to lysis and amplicon sequencing to determine indel efficiency (Fig. 4a). We selected 24 loci from six genes for validation, with WT-DelIscB+sgRNA-V1 and enIscB-T5E included as controls (Fig. 4b). Given that the OgeuIscB’s TAM (NWRRNA) differs from the DelIscB’s TAM (NAC), we selected target sequences with TAM positions as close as possible to each other for activity comparison. Across these 24 loci, enDelIscB demonstrated an average increase in gene editing activity by about 48.9-fold on average compared to WT-DelIscB+sgRNA-V1 (Fig. 4b, c). Moreover, when enDelIscB was fused with T5E, its activity was on par with that of enIscB-T5E, with values of 54.27 ± 28.07% versus 53.70 ± 29.84% (Fig. 4b, c).
a The experimental workflow for evaluating indel formation efficiency of IscB editors at endogenous loci. b Comparison of indel frequency by WT-DelIscB+sgRNA-V1, enDelIscB, enDelIscB-T5E and enIscB-T5E across 24 endogenous genomic loci in HEK293T cells. Data represent mean ± s.d. of three independent biological replicates. c The summary dot plot shows a comparison of the activity of WT-DelIscB+sgRNA-V1, enDelIscB, enDelIscB-T5E and enIscB-T5E in HEK293T cells. Each dot represents the mean indel level of three independent biological replicates measured for each endogenous locus mentioned in (b). P values were determined by a two-tailed unpaired t test. All data are presented as mean ± s.d., n = 24. d The indel patterns generated by WT-DelIscB+sgRNA-V1, enDelIscB and enDelIscB-T5E in HEK293T cells at the EMX1-sg2 site. Data are shown as one independent biological replicate. e Analysis of enDelIscB and enDelIscB-T5E off-target effect at three targets (EMX1-sg1, VEGFA-sg2, and CXCR4-sg2). The top line of each panel is an on-target sequence, with 16 lines below representing the in silico predicted off-target sites. The TAM is displayed in blue, and the mismatched bases of the off-target sites relative to the on-target sites are marked in lowercase and red. The numbers and color intensity within each box of the heatmap on the right represent the indel levels of an editor at that site. Data are presented as the mean of three independent biological replicates. Source data are provided as a Source data file.
Concurrently, we examined the indel patterns generated by WT-DelIscB+sgRNA-V1, enDelIscB and enDelIscB-T5E in the genome (Fig. 4d). At the EMX1-sg2 locus, the insertion and deletion produced by WT-DelIscB+sgRNA-V1 were not only inefficient (reaching up to about 1.5%) but also narrow (around 10 bp) (Fig. 4d). In contrast, enDelIscB generated a wide deletion range (about 90 bp) and a certain proportion of insertions (about 17%) (Fig. 4d). The enDelIscB-T5E had a narrower deletion range than enDelIscB (about 60 bp), yet its deletion efficiency was higher than that of enDelIscB (Fig. 4d). Meanwhile, enDelIscB-T5E caused almost no insertion events, similar to the cleavage pattern of enIscB-T5E (Fig. 4d)24.
Furthermore, we utilized Cas-OFFinder to analyze the off-target effects of enDelIscB and enDelIscB-T5E at three targets, respectively. The results showed that enDelIscB and enDelIscB-T5E had almost no indels at 16 predicted off-target sites corresponding to two targets (EMX1-sg4 and CXCR4-sg2), while there were four significant off-target events at VEGFA-sg2 (Fig. 4e).
To evaluate genome-wide off-target effects of enDelIscB and enDelIscB-T5E, we employed GUIDE-seq35 at four targeted loci (Fig. 5). Given the short guide sequence (16 nt) and NAC TAM sequence (3 nt) of DelIscB, we allowed up to six mismatches within the 19-nt bait in GUIDE-seq analysis. We selected off-target sites with more than 1% sequencing reads for presentation (Fig. 5), and all off-target site sequences are listed in Supplementary Data 3. Our data showed that the top-ranked sequences captured by GUIDE-seq at four loci for enDelIscB or enDelIscB-T5E are all target sequences (Fig. 5a–d).
a–d Off-target editing events induced by enDelIscB and enDelIscB-T5E in HEK293T cells at AAVS1-sg3 (a), EMX1-sg4 (b), AAVS1-sg2 (c), and VEGFA-sg2 (d). A 19-nt sequence that includes TAM (3 nt) and the guide sequence (16 nt) is shown (percentiles of reads >1% are listed). e Summary of off-target site hits and the proportion of off-target editing events versus total edits.
We counted the number of off-target sites and the proportion of off-target editing events versus total editing events (Fig. 5e). Our analysis revealed that enDelIscB-T5E displays a higher proportion of off-target reads compared to enDelIscB (Fig. 5e). This observation indicates that fusion with T5E not only enhances activity at on-target sites but also elevates activity at off-target sites. In addition, we assessed the insertion rates of dsODN and found that enDelIscB-T5E exhibits a lower insertion rate than enDelIscB (Supplementary Fig. 14). We hypothesize that the degradation of the 5’ end of DNA at the break site by the T5 exonuclease leads to the formation of sticky ends, thereby reducing the insertion efficiency of dsODN. This further implies that fusion with T5E may diminish the detection sensitivity of GUIDE-seq.
We utilized WebLogo336 to analyze the profiles of the off-target sequences for enDelIscB at four different loci (Supplementary Fig. 15). We observed that the shortest sequence effectively recognized by DelIscB should be the 15 nucleotides upstream of the TAM sequence (Supplementary Fig. 15), which is consistent with the findings presented in Supplementary Fig. 1e. This suggests DelIscB’s off-target effects arise from both its intrinsic mismatch tolerance and relatively short effective guide length. Kannan et al. extended the effective guide length of OrufIscB to 20-nt by inserting the REC domain of II-D Cas9, which not only increased its activity but also improved its specificity37. This strategy thus offers a valuable approach for further enhancing the specificity of DelIscB.
Taken together, enDelIscB and enDelIscB-T5E exhibited robust genome editing activity in mammalian cells, while off-target events still need to be considered in future applications.
Developing base editors based on enDelIscB nickase
Given that DelIscB possesses the features of both RuvC and HNH nuclease domains, we consider developing base editors based on enDelIscB nickase. Through full-sequence alignment between DelIscB and OgeuIscB, we identified presumed active residues (D60, E171, H226, H324, and D327) in DelIscB, which correspond to the RuvC and HNH active residues of OgeuIscB23 (Supplementary Fig. 8a). We then mutated these putative active residues to alanine and validated its nickase activity using a pair of fluorescent reporter plasmids carrying a single or dual target sequence (Supplementary Fig. 8b). The results showed that the corresponding single mutations in the RuvC domain (D60A, E171A, H324A, and D327A) and the H226A mutation in the HNH domain all lost their double-strand cleavage activity while retaining nickase activity (Supplementary Fig. 8c). When validating activity with a single target reporter, we noticed that these five single mutations yielded approximately 10% more EGFP+ cells than the NT control. We believe this may be attributed to the background that nicked plasmids are more prone to spontaneous double-strand breaks. In addition, enDelIscB with D60A + H226A exhibited no cleavage activity and was referred to as dead enDelIscB (Supplementary Fig. 8c).
To construct C-to-T DNA base editors, we selected enDelIscBD60A nickase and fused the cytosine deaminase hAPOBEC3AW98Y/W104A/Y130F (the high-fidelity version of hAPOBEC3A, referred as hA3A*)38 at either its N-terminus or C-terminus, along with two copies of uracil glycosylase inhibitor (2×UGI) (Fig. 6a). At two endogenous loci (CXCR4-sg2 and PCSK9-sg3), the base editor with hA3A* fused at the C-terminus (enDelIscBD60A-hA3A*−2×UGI) exhibited a significantly higher C-to-T conversion efficiency than the one with hA3A* fused at the N-terminus (hA3A*-enDelIscBD60A-2×UGI) (Fig. 6b). We named the former as ICBE (i.e., IscB based cytosine base editor) (Fig. 6b). This outcome is consistent with the previous finding that fusing NLS at the N-terminus of DelIscB diminishes its activity. Meanwhile, we constructed adenine base editors by fusing TadA8eV106W (the high-fidelity version of TadA8e, referred as TadA*)39 with enDelIscBD60A in different forms (Fig. 6c). We found that fusing the TadA* dimer to the C-terminus of enDelIscBD60A displays the most optimal activity of A-to-G conversion at both target sites, which we named as IABE (Fig. 6d).
a Schematic for constructing a miniature cytosine base editor by fusing cytosine deaminase at the N-terminus or C-terminus of enDelIscB nickase. hA3A* represents human APOBEC3AW98Y/W104A/Y130F. UGI, uracil glycosylase inhibitor. b Comparison of C-to-T conversion efficiency of hA3A*-enDelIscBD60A-2×UGI and enDelIscBD60A-hA3A*−2×UGI at 2 endogenous loci (CXCR4-sg2 and PCSK9-sg3). Data represent mean ± s.d. of three independent biological replicates. c Schematic for constructing a miniature adenosine base editor by fusing adenosine deaminase with enDelIscB nickase in a different pattern. TadA* represents TadA8eV106W. d Comparison of A-to-G conversion efficiency of TadA*-enDelIscBD60A-TadA*, enDelIscBD60A-TadA*, and enDelIscBD60A-2×TadA* at 2 endogenous loci (EMX1-sg2 and EMX1-sg5). Data represent mean ± s.d. of three independent biological replicates. e Comparison of C-to-T editing efficiency for ICBE and miCBE across 9 endogenous loci. Data represent mean ± s.d. of three independent biological replicates. f Comparison of A-to-G editing efficiency for IABE and miABE across 9 endogenous loci. Data represent mean ± s.d. of three independent biological replicates. Source data are provided as a Source data file. The schematic diagrams of the base editors in (a, c) were adapted from a licensed CRISPR/Cas9 (Pink) icon. This icon is available at https://togotv.dbcls.jp/en/togopic.2014.75.html, sourced from TogoTV50 (© 2016 DBCLS TogoTV) and distributed under the CC-BY-4.0 license (https://creativecommons.org/licenses/by/4.0/).
Next, we compared the editing windows and efficiencies of the base editors based on enIscBD61A and enDelIscBD60A across multiple genomic loci (Fig. 6e, f). miCBE and miABE were derived from enIscBD61A, which was the engineered version of OgeuIscB24. By analyzing the conversion efficiencies of each C-to-T or A-to-G at all targeted sites, we found that for ICBE, its editing window ranges approximately from position −1 to 12 (counting the TAM as positions 17–19), while the editing window for miCBE ranges from position 2 to 14 (Supplementary Fig. 9a). For IABE, its editing window is between target positions −4 and 12, while the editing window for miABE is between positions 2 and 12 (Supplementary Fig. 9b). The average highest base editing efficiency at each locus was statistically analyzed, and the highest efficiency of ICBE was 49.26 ± 23.22% compared to 40.93 ± 33.85% for miCBE (Supplementary Fig. 9c). For IABE, the highest efficiency is 60.22 ± 22.47%, while that of miABE is 38.89 ± 31.93% (Supplementary Fig. 9c). The results of the differential analysis indicated that there is no significant difference in the editing efficiency of base editors based on enIscBD61A and enDelIscBD60A across these endogenous loci (Supplementary Fig. 9c).
We also examined the indels generated by these four base editors at the target sites. Our findings revealed that all four base editors induced a low frequency of indels (Supplementary Fig. 9d, e), which is consistent with previous reports24. Collectively, these results suggest that enDelIscBD60A-based base editors (ICBE and IABE) offer an alternative and efficient option for expanding the miniature base editing toolbox.
Efficient generation of mouse models with enDelIscB and enDelIscB-T5E
To further explore the enDelIscB system’s potential for in vivo applications, we microinjected the enDelIscB system into mouse embryos. We chose the tyrosinase (Tyr) gene as a model target based on the following scientific rationale: The Tyr gene encodes the black coat color phenotype in C57BL/6 mice, and its mutation causes albinism. By targeting Tyr, we could intuitively assess the efficiency of genome editors through observable coat color changes in mice. Furthermore, this target represents a widely adopted benchmark for validating the efficiency of gene editing tools in mouse models27,40,41. We wish to explicitly clarify that the selection of this gene in no way implies or endorses its potential use in human gene editing for the purpose of modifying skin color.
To identify suitable loci for enDelIscB or enDelIscB-T5E to target efficiently, we screened a dozen sgRNAs with different guide sequences targeting the exon 1 or exon 4 of Tyr gene in mouse neuro2a (N2a) cells (Fig. 7a). The results of the indel detection showed that 9 out of the 18 sgRNAs exhibited detectable activity, suggesting that enDelIscB-T5E can also function in mouse cells (Fig. 7b). Among these sgRNAs, Tyr-sg12, which targets exon 1 of the Tyr gene, demonstrated the highest editing efficiency (Fig. 7b).
a Schematic representation of the mouse tyrosinase (Tyr) gene, which contains five exons. We choose exon1 and exon4 for screening target sequences. Each target sequence is annotated below its corresponding exon. b enDelIscB-T5E-mediated indel formation efficiency at different targets of Tyr gene in mouse N2a cells. Data represent mean ± s.d. of three independent biological replicates. c Phenotypic photographs of F0-generation mice with Tyr gene disruption generated by microinjecting enDelIscB or enDelIscB-T5E mRNA and sgRNA (Tyr-sg12). Photos were taken when the mice were 21 days old. d The indel rates of the Tyr gene in F0 mice were edited by enDelIscB (n = 39) and enDelIscB-T5E (n = 47). e Summary of the condition in constructing a mouse model of albinism through microinjection. Source data are provided as a Source data file.
Subsequently, Tyr-sg12 was in vitro transcribed and mixed with the mRNA of enDelIscB or enDelIscB-T5E and co-injected into the zygotes of C57BL/6 J mice. The phenotypes of the newborn mice were as follows: among the 39 F0 generation mice injected with enDelIscB, 35 mice had completely white fur, 2 mice had a mix of black and white fur, only 1mouse had black fur, 1 mouse died prematurely and was not photographed (Fig. 7c, e). Regarding the 47 F0 generation mice injected with enDelIscB-T5E, 44 mice had pure white fur, while 3 mice had black-and-white fur (Fig. 7c, e). The results of the genotype analysis indicate that among the 39 F0 mice injected with enDelIscB, only 4 mice were not fully edited, with an average indel level of 96.77% (Fig. 7d). Among the 47 F0 mice injected with enDelIscB-T5E, only 3 mice were not fully edited, with an average indel level of 98.67% (Fig. 7d). The phenotype of each mouse corresponded to its genotype (Fig. 7c and Supplementary Fig. 10). In summary, these findings demonstrate that both enDelIscB and enDelIscB-T5E exhibit robust gene editing activity in mouse embryos, which have great potential for in vivo applications.
Discussion
In this study, through the characterization and engineering of a CRISPR-associated IscB system, DelIscB (which has a relatively flexible TAM, NAC), we obtained two potent genome editors, enDelIscB and enDelIscB-T5E. Furthermore, based on the enDelIscB nickase, we developed two efficient miniature base editors, ICBE and IABE. Finally, we efficiently constructed mouse models using enDelIscB and enDelIscB-T5E.
Unlike OgeuIscB, which is guided by a single ωRNA, DelIscB is guided by an ncRNA located upstream of its ORF and containing the CRISPR array. This ncRNA contains repeat and anti-repeat regions, which are similar to the CRISPR RNA (crRNA) and trans-activating crRNA (tracrRNA) of the Cas9 systems5. This implies that DelIscB may represent an intermediate in the evolutionary progression from the ancestral IscB system to the CRISPR-Cas9 system. However, further investigation is required to determine whether and how this ncRNA is processed into two components similar to crRNA and tracrRNA, which could enable targeting different genomic loci using diverse spacer sequences.
For engineering the DelIscB protein, we initially employed structural comparison to map the high-activity amino acid mutation of enIscB to DelIscB. However, only one effective corresponding mutation, S97R, was obtained. Subsequent protein homology analysis revealed that DelIscB and OgeuIscB share only 28.14% sequence identity, indicating significant structural divergence between the two proteins. This low sequence similarity suggests that a direct structural comparison between DelIscB and OgeuIscB is not feasible for guiding further engineering efforts.
Next, we employed the MSA strategy to introduce the naturally occurring mutations from its homologous proteins into DelIscB. We focused on the substitution of arginine, while the substitution of other residues that frequently appear at aligned positions also demonstrated effective results. This approach provides an operational insight for engineering RGNs. Although this strategy does not exhaustively screen all potential beneficial mutations, it offers a more rational alternative to the unbiased arginine scanning substitution strategy24, as it is obvious that mutations in conserved regions of the protein are highly likely to inactivate IscB.
The increase in activity of the effective DelIscB mutants was relatively modest, with most mutants exhibiting a 1.2-fold to 1.6-fold change in activity. This is in contrast with the more significant activity enhancements seen in engineered OgeuIscB mutant (>1.8-fold), likely due to the inherently low activity of DelIscB. Compared to OgeuIscB, DelIscB lacks the P1 domain, which is crucial for sgRNA interaction, and has a smaller protein size. These features reduce the contact area for nucleic acid interactions, potentially explaining its weaker baseline activity. However, through our systematic engineering of the DelIscB protein and sgRNA, and by maximizing the accumulation of beneficial mutations, the activity of DelIscB has been increased by an average of approximately 48.9-fold across 24 endogenous sites. Given that nearly all wild-type IscB systems exhibit inherently low cleavage activity, our findings provide a framework for employing diverse strategies to screen and accumulate beneficial mutations, thereby facilitating the development of efficient and compact gene editing tools based on these systems.
For the truncation of the DelIscB sgRNA scaffold, we have truncated the initial length from 365-nt (ncRNA) to 267-nt (sgRNA-V1), and further to 248-nt (sgRNA-V5). We anticipate that if the cryo-EM structure of the DelIscB and sgRNA complex bound to target DNA is available, the sgRNA can be further truncated based on structure information to improve its performance and enable chemical synthesis. Such structural insights would also provide a reference basis for structure-guided protein engineering.
For rationalizing the effects of beneficial protein mutations, we have employed AlphaFold342 to predict the structure of the DelIscB-sgRNA-V1-DNA complex, with all beneficial mutations highlighted in the structure (Supplementary Fig. 11). Among these, C12R localizes to the PLMP domain and is likely involved in stabilizing the DelIscB-sgRNA-V1 complex. The mutations S97K, S100A, G104S, W117F, and A142P are anticipated to facilitate recognition of the guide:target heteroduplex by the BH and Linker (Rec) domains. E317R and Q340K reside in the RuvC III domain and are predicted to enhance the affinity for the NT strand during allosteric cleavage. Structurally, V396K and S436R appear to strengthen affinity for the sgRNA scaffold. Additionally, we have marked the positions of effective truncations and mutations in the predicted structure of sgRNA-V1 (Supplementary Fig. 12). Notably, the effective truncated stem-loop regions exhibit no interaction with the protein interface (Supplementary Fig. 12).
To explain the potential mechanism by which the N-terminal fusion of NLS to DelIscB may reduce its activity, we utilized AlphaFold342 to predict the structures of the DelIscB-sgRNA-V1-target DNA complex and the protein structure of NNLS-DelIscB alone (Supplementary Fig. 13a, d). Structural alignment reveals strong correspondence between the predicted DelIscB-sgRNA-V1-target DNA complex and the cryo-EM structure of the OgeuIscB–ωRNA–target DNA complex (PDB: 7XHT) (Supplementary Fig. 13b, c), validating the prediction accuracy of the model. After aligning the predicted structure of NNLS-DelIscB with the predicted structure of the DelIscB-sgRNA-V1-target DNA complex, we found that the spatial positioning of the NNLS conflicts with the 3’ stem-loop of sgRNA-V1 (Supplementary Fig. 13d). Considering that the PLMP domain at the N-terminus of OgeuIscB interacts with the RuvC domain and ωRNA, it plays a role in stabilizing the IscB–ωRNA complex43. Therefore, we hypothesize that the NNLS affects the natural interaction between the PLMP domain and the 3’ stem-loop of sgRNA-V1, thereby disrupting the stability of the entire ribonucleoprotein (RNP) complex and leading to a decrease in activity.
In addition, the N-terminal fusion of hA3A* in DelIscB exhibits lower base editing activity, whereas this is not observed for the C-terminal fusion (Fig. 6a, b). We employed AlphaFold342 to predict the individual protein structures of hA3A*-enDelIscBD60A-2×UGI and enDelIscBD60A-hA3A*−2×UGI (ICBE). Structural alignment of the predicted models with the predicted structure of DelIscB-sgRNA-V1-target DNA complex revealed that the N-terminally fused hA3A* exhibits steric clash with the 3’ stem-loop of sgRNA-V1, whereas the C-terminally fused hA3A* shows no such spatial conflict. (Supplementary Fig. 13e, f). This suggests that the N-terminal fusion of hA3A* may hinder the formation of a stable RNP complex between DelIscB protein and sgRNA-V1, leading to a decrease in activity.
However, fusing TadA* at both the N-terminus and C-terminus of DelIscB does not lead to a significant decrease in activity compared to enDelIscBD60A-2×TadA* (IABE) (Fig. 6c, d). In contrast, fusing a single TadA* only at the C-terminus shows lower activity than the former two (Fig. 6c, d). We used AlphaFold342 to predict the protein structures of TadA*-enDelIscBD60A-TadA*, enDelIscBD60A-2×TadA* (IABE), and enDelIscBD60A-TadA*, and aligned them with the predicted structure of the DelIscB-sgRNA-V1-target DNA complex individually (Supplementary Fig. 13g–i). Predictive structural analysis revealed that the two TadA* domains within the TadA*-enDelIscBD60A-TadA* form a homodimer and engage DNA without steric clash with the 3’ stem-loop (Supplementary Fig. 13g). Similarly, the TadA* domains in both enDelIscBD60A-2×TadA* (IABE) and enDelIscBD60A-TadA* configurations are predicted to exhibit DNA binding without steric hindrance to the 3’ stem-loop (Supplementary Fig. 13h, i). This predicted structural compatibility likely underlies their enhanced activity relative to hA3A*-enDelIscBD60A-2×UGI. However, the predicted structure may not fully represent the true structure. To clarify the underlying mechanisms, a combination of biochemical and structural experimental approaches may be required for in-depth investigation. Furthermore, C-terminal mono-TadA* fusion exhibits reduced catalytic efficiency relative to di-TadA* configurations (Fig. 6c, d), likely because TadA natively operates as a homodimer44.
Compared to miCBE and miABE, the base editors ICBE and IABE that we developed exhibit a slightly wider editing window, extending even beyond the scope of the guide sequence (Supplementary Fig. 9a, b). This may result from the inherent properties of enDelIscB or the different fusion patterns between the deaminase and IscB nickase. However, the editing windows of these four IscB nickase-based base editors appear to be quite broad. It is particularly important to narrow their editing windows for more precise base editing. Inspired by the idea that inserting TadA inside Cas9 can narrow the ABE editing window45, fusing deaminases inside IscB may be a feasible strategy to reduce the editing window.
Given that gene editing typically targets the coding sequence (CDS) of genes, we systematically calculated the distribution density of various nucleases’ PAM/TAM across the entire human CDS using a sliding window algorithm46. This density is defined as the total number of PAM or TAM sites divided by the total length of cDNA sequences (Supplementary Fig. 22). Statistical analysis revealed that DelIscB exhibits a higher TAM density than OgeuIscB (10.10% vs. 7.39%) (Supplementary Fig. 22), indicating that DelIscB has a broader targeting scope within the CDS region compared to OgeuIscB. Furthermore, the targeting scope of SpCas9 is broader than that of both DelIscB and OgeuIscB (Supplementary Fig. 22).
We also employed the enDelIscB and enDelIscB-T5E editors to efficiently establish albinism models by microinjecting mouse embryos. This indicates that enDelIscB and enDelIscB-T5E have great potential for in vivo applications. Also considering the advantages of DelIscB’s compact size, gene therapy research can be carried out in the future by delivering enDelIscB, enDelIscB-T5E, ICBE, and IABE through AAV vectors, further expanding their utility in precision medicine. However, we strongly urge that any potential applications of genome editing in humans be approached with extreme caution. We also emphasize the imperative of establishing robust ethical oversight, implementing rigorous regulations, and building broad societal consensus.
It is worth noting that several mice (#14, #31, #56, #66, #86) exhibited both black and white fur patches, suggesting that these animals may have mosaic genomes. For detecting the frequency of mosaicism, we performed sampling and sequencing of different skin patches with varying fur colors from three mice of the F0 generation with black and white fur (#14, #31, #56). This included skin samples from the tail, ears, and the left, middle, and right regions of the back (Supplementary Figs. 16–18). Sequencing statistics reveal that different tissues within each mouse exhibit distinct frequencies of mosaic genomes (Supplementary Figs. 16–18).
After photographing the F0 generation mice at 21 days postnatally, we retained 12 mice for subsequent breeding, and all of them have been living healthily for 19 weeks (Supplementary Fig. 19a, b). We measured their body weight and body length, and observed no growth differences compared with wild-type (WT) mice. (Supplementary Fig. 19c). Furthermore, we mated some F0 generation mice with WT mice, while F0 mice #74 and #59 were bred with each other. Both breeding schemes successfully yielded offspring carrying Try mutations (Supplementary Figs. 19d, 20, and 21). These results demonstrate that mice edited with the enDelIscB or enDelIscB-T5E genes are healthy and possess normal reproductive capabilities.
Methods
All experiments in this article comply with all relevant ethical regulations and are approved by the Committee of Biosafety, Ethics and Laboratory Animal Management at the Institute of Biophysics, Chinese Academy of Sciences, under approval number SYXK2024235. The animal experimental protocols were approved by the Institutional Animal Care and Use Committee of the Institute of Biophysics, Chinese Academy of Sciences.
Plasmid constructions
For prokaryotic expression plasmid construction, the ORFs of SpCas9, OgeuIscB, AwaIscB, and DelIscB from addgene plasmids (#44758, #176540, #176538 and #176588) were cloned into the downstream of the lac operon on the pACYC184 (p15A replicon, chloramphenicol resistance) vector by Gibson Assembly, respectively. The corresponding sgRNA/ωRNA/ncRNA with target or non-target guide sequences were cloned into the downstream of the J23119 promoter on the pRSFduet-1 (RSF replicon, kanamycin resistance, and lacI expression) vector by Gibson Assembly. All fragments used in Gibson Assembly were obtained by PCR using Q5 Hot start High-Fidelity DNA Polymerase (NEB). The assembly reagents used in Gibson Assembly are NEBuilder HiFi DNA Assembly Cloning Kit (NEB) or ClonExpress Ultra One Step Cloning Kit (Vazyme).
For eukaryotic expression plasmid construction, we cloned the human codon-optimized DelIscB sequence (synthesized by Tsingke) and its sgRNA under U6 promoter into the pCAG vector obtained from enIscB (Addgene, #205410). For the construction of the fluorescent reporter plasmid, we cloned the BFP-T2A-EGFxxFP sequence into the pCDNA3.1 backbone (Addgene, #176540) and under a CMV promoter. For the engineering of DelIscB and sgRNA, DNA fragments were amplified using PCR primers with mutations and overlaps, and assembled with the backbone digested by AgeI and HindIII or NotI. For targeting different genome loci, pairs of oligonucleotides were annealed and ligated into BsaI or BbsI-linearized plasmid vector by T4 DNA ligase (GenStar). For the construction of DelIscB-T5E, ICBE and IABE plasmids, the elements of T5E, hAPOBEC3AW98Y/W104A/Y130F, UGI, TadA8eV106W and linkers were cloned from the plasmids of enIscB-T5E, miCBE and miABE (Addgene, #205411, #205413 and #205412), which were gifts from Hui Yang and Yingsi Zhou. All DNA, protein, and primer sequences are listed in Supplementary Data 1. All primer sequences were synthesized by Tsingke.
Plasmid interference assay in E. coli
The plasmid interference assay was based on our previous reports47,48. Briefly, 50 ng of plasmid 1 and 50 ng of plasmid 2 were co-transformed into 25 μl of Tsuro chemically competent cells (Tsingke) and then plated on LB agar plates containing chloramphenicol (33 μg/ml), kanamycin (50 μg/ml), and IPTG (1 mM). Incubate the plate overnight at 37 °C and take pictures using a gel scanner (Tanon 3500).
Mammalian cell culture
HEK293T cells were purchased from the American Type Culture Collection (ATCC) (CRL-11268) and were cultured in Dulbecco’s modified Eagle medium (Millipore) supplemented with 10% (v/v) fetal bovine serum (Gibco), 1% (v/v) non-essential amino acids, 1% (v/v) glutamine and 1% (v/v) β-mercaptoethanol at 37 °C with 5% CO2. Cells are passaged at a 1:10 ratio every 3 days.
Neuro-2a cells were purchased from the ATCC (CCL-131) were cultured in Minimum Essential Medium (Gibco) with 10% (v/v) fetal bovine serum (Gibco) and 1% (v/v) non-essential amino acids at 37 °C with 5% CO2. Cells are passaged at a 1:4 ratio every 2 days.
Mammalian cell transfection and fluorescence-activated cell sorting (FACS) analysis
The HEK293T cells were seeded in 24-well plates at 2 × 105 per well and transfected after 16 h. For engineering the protein and sgRNA, cells were transfected with 1 μg of mixed plasmids containing the reporter and editor plasmids in a molar ratio of 1:1 using Lipofectamine 2000 (GenStar). After 48 h, mCherry, BFP, and EGFP fluorescence were analyzed by BD FACSAria IIIu flow cytometer. FACS data were analyzed by FlowJo (vX.0.7). For detection of indel and base editing at endogenous sites, 1 μg of editor plasmids were transfected with Lipofectamine 2000. After 48 h, the top roughly 20% of mCherry-positive cells were sorted using BD FACSAria IIIu flow cytometer.
T7 endonuclease I assay
To preliminarily evaluate the genome editing efficiency in mammalian cells, the harvested cells were suspended with QuickExtract DNA Extraction Solution (LGC Biosearch Technologies) and cycled at 65 °C for 15 min, 68 °C for 15 min then 95 °C for 10 min to lyse cells. Genome loci containing the target were amplified using Q5 High-Fidelity 2x Master Mix (NEB). Primer sequence was listed in Supplementary Data 1. Then T7 endonuclease I reaction was performed according to the manufacturer’s instructions. Subsequently, the reaction products were digested by proteinase K to eliminate the enzymes that interacted with DNA and finally were fractionated in a 2.5 % agarose gel and visualized by Gel Imaging System (Tanon 3500).
Next-generation sequencing (NGS) and data analysis
About 20,000 sorted cells were lysed in 10 μl of QuickExtract DNA Extraction Solution (LGC Biosearch Technologies), and 1 μl of lysed cells were used as input into each PCR reaction. The target genomic regions were amplified with the first round of PCR using 2×Phanta Flash Master Mix (Vazyme) to add Illumina adapters, and followed by a second round of PCR to add index1/2 and P5/P7. The libraries were gel extracted and sequenced by 150-bp paired-end reads Illumina NovaSeq Xplus platform (Majorbio Bio-Pharm Technology Co. Ltd.). The Indel or base conversion frequency was analyzed using CRISPResso2 as described49. All primers used are provided in Supplementary Data 1.
Off-target analysis using Cas-OFFinder
To evaluate the specificity of enDelIscB and enDelIscB-T5E, we used Cas-OFFinder to predict potential off-target sites. Briefly, 5’-NNN-3’ PAM type was chosen due to the lack of equivalent PAM for DelIscB. The query sequence should include 16-nt on-target guide sequence and ‘NAC’. The target genome was selected as vertebrate and human (GRCh38/hg38) with the number of mismatches set to 3. In the sequence exported from the website (Supplementary Data 2), we manually selected 16 off-target sites with NAC TAM and 1 or 2 mismatches distributed at different positions for NGS analysis. The primers for PCR were provided in Supplementary Data 1.
GUIDE-seq for identifying genome-wide off-target editing
To comprehensively evaluate and compare the genome-wide off-target editing effects of enDelIscB and enDelIscB-T5E, GUIDE-seq assays were carried out as previously described with modifications35. Briefly, the targeting plasmids and dsODN (double-stranded oligodeoxynucleotide) were transfected into HEK293T cells by using Lipofectamine 2000 (GenStar) according to the manufacturer’s instructions. The cells were harvested 72-h post transfection for evaluating the genome-wide off-target effects. The genomic DNA was extracted and sheared to an average length of 500 bp by using NEBNext dsDNA Fragmentase (NEB) according to the manufacturer’s instruction. The fragmented gDNA was end-repaired, dA-tailed and ligated to an adapter harboring an 8-nt random molecular index by using VAHTS Universal DNA Library Prep Kit for Illumina V3 (Vazyme) according to the manufacturer’s instruction. The dsODN-containing fragments were enriched with two rounds of nested anchored PCR. The library was subjected to Illumina NovaSeq 6000 sequencing. Output sequence data was analyzed using GUIDE-seq pipeline (https://github.com/aryeelab/guideseq).
In vitro transcription of RNA
For the transcription of enDelIscB and enDelIscB-T5E mRNA, mMESSAGE mMACHINE® T7 Ultra Kit (Invitrogen) was used according to the manufacturer’s instructions. The in vitro transcription template is derived from a linearized plasmid containing the ORF of enDelIscB and enDelIscB-T5E under the T7 promoter. The sgRNA was transcribed from the PCR products using the MEGAshortscript™ (Invitrogen) following the manufacturer’s instructions. The primers with the T7 promoter sequence were provided in Supplementary Data 1.
Microinjection of mouse zygotes
Mouse embryos were obtained by superovulation of C57BL/6 J female mice and in vitro fertilization (IVF) with male mice of the same genetic background. Zygotes were injected into the cytoplasm with solutions that contained enDelIscB or enDelIscB-T5E mRNA (100 ng/μl) and the sgRNA of Tyr-sg12 (200 ng/μl). The injected zygotes were cultured in KSOM embryo culture medium overnight. The embryos developed to the 2-cell stage were transferred into the ampulla of the fallopian tube of pseudo-pregnant ICR female mice. All mice were housed under a specific pathogen-free environment, with 12 h dark/light cycle, 22 ± 2 °C, and 40% ~ 60% humidity conditions. A total of 10 C57BL/6 J female mice aged 4 weeks, one C57BL/6 J male mouse aged 12 weeks, and 8 ICR female mice aged 10 weeks were used in this experiment.
Extraction and sequencing of DNA from mouse tissues
Small tissue samples are first collected from the mouse’s tail or toe, from which DNA is extracted. Next, amplicon sequencing is performed on the DNA sequences flanking the target site of the Tyr gene, and the indel proportion is analyzed using CRISPResso2. When statistically analyzing the genotype and phenotype of each F0 generation mouse, we did not take gender into account, as the target gene Tyr is not related to the sex chromosomes.
Analyzing the targeting scope of RGNs on the entire human CDS
The GRCh38 human CDS (https://ftp.ensembl.org/pub/release-114/fasta/homo_sapiens/cds/) was used for analysis. For a given RGN, all PAM/TAM sequences or their complementary sequences on CDS were counted by a sliding window algorithm46. The targeting scope on CDS was calculated by using the number of targetable TAM/PAM to divide the total length of the CDS.
Statistics and reproducibility
GraphPad Prism (Version 10.4.0) was used for statistics analysis. All values are shown as mean ± standard deviations (SD). A two-tailed unpaired t test was used for comparisons and P < 0.05 was considered to be statistically significant. Each experiment was repeated independently at least two or three times with similar results. No statistical method was used to predetermine sample size; sample sizes were chosen based on established standards in the field. No data were excluded from the analyses. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Web site used in this study
Cas-OFFinder (http://www.rgenome.net/cas-offinder/)
RNAfold (http://rna.tbi.univie.ac.at/cgi-bin/RNAWebSuite/RNAfold.cgi)
Mfold (https://www.unafold.org/mfold/applications/rna-folding-form.php)
MUSCLE (https://www.ebi.ac.uk/jdispatcher/msa/muscle?stype=protein)
AlphaFold2 (https://colab.research.google.com/github/sokrypton/ColabFold/blob/main/AlphaFold2.ipynb)
AlphaFold3 (https://alphafoldserver.com/)
WebLogo3 (https://weblogo.threeplusone.com/create.cgi)
TogoTV (https://togotv.dbcls.jp/en/pics.html)
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All deep sequencing data are available at the National Center for Biotechnology Information (NCBI) Sequence Read Archive database under the BioProject accession code PRJNA1292198. Plasmids are available on Addgene. Source data are provided with this paper.
References
Pacesa, M., Pelea, O. & Jinek, M. Past, present, and future of CRISPR genome editing technologies. Cell 187, 1076–1100 (2024).
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Knott, G. J. & Doudna, J. A. CRISPR-Cas guides the future of genetic engineering. Science 361, 866–869 (2018).
Makarova, K. S. et al. Evolutionary classification of CRISPR–Cas systems: a burst of class 2 and derived variants. Nat. Rev. Microbiol. 18, 67–83 (2019).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163, 759–771 (2015).
Adli, M. The CRISPR tool kit for genome editing and beyond. Nat. Commun. 9, 1911 (2018).
Anzalone, A. V., Koblan, L. W. & Liu, D. R. Genome editing with CRISPR-Cas nucleases, base editors, transposases and prime editors. Nat. Biotechnol. 38, 824–844 (2020).
McCutcheon, S. R., Rohm, D., Iglesias, N. & Gersbach, C. A. Epigenome editing technologies for discovery and medicine. Nat. Biotechnol. 42, 1199–1217 (2024).
Madigan, V., Zhang, F. & Dahlman, J. E. Drug delivery systems for CRISPR-based genome editors. Nat. Rev. Drug Discov. 22, 875–894 (2023).
Pausch, P. et al. CRISPR-CasΦ from huge phages is a hypercompact genome editor. Science 369, 333–337 (2020).
Wu, Z. W. et al. Programmed genome editing by a miniature CRISPR-Cas12f nuclease. Nat. Chem. Biol. 17, 1132–1138 (2021).
Xu, X. S. et al. Engineered miniature CRISPR-Cas system for mammalian genome regulation and editing. Mol. Cell 81, 4333 (2021).
Kim, D. Y. et al. Efficient CRISPR editing with a hypercompact Cas12f1 and engineered guide RNAs delivered by adeno-associated virus. Nat. Biotechnol. 40, 94 (2022).
Wu, Z. W. et al. Structure and engineering of miniature Cas12f1. Nat. Catal. 6, 695–709 (2023).
Chen, W. Z. et al. Cas12n nucleases, early evolutionary intermediates of type V CRISPR, comprise a distinct family of miniature genome editors. Mol. Cell 83, 2768 (2023).
Aliaga Goltsman, D. S. et al. Compact Cas9d and HEARO enzymes for genome editing discovered from uncultivated microbes. Nat. Commun. 13, 7602 (2022).
Altae-Tran, H. et al. The widespread IS200/IS605 transposon family encodes diverse programmable RNA-guided endonucleases. Science 374, 57 (2021).
Karvelis, T. et al. Transposon-associated TnpB is a programmable RNA-guided DNA endonuclease. Nature 599, 692 (2021).
Xiang, G. H. et al. Evolutionary mining and functional characterization of TnpB nucleases identify efficient miniature genome editors. Nat. Biotechnol. 42, 745 (2024).
Saito, M. et al. Fanzor is a eukaryotic programmable RNA-guided endonuclease. Nature 620, 660–668 (2023).
Jiang, K. Y. et al. Programmable RNA-guided DNA endonucleases are widespread in eukaryotes and their viruses. Sci. Adv. 9, eadk0171 (2023).
Schuler, G., Hu, C. Y. & Ke, A. L. Structural basis for RNA-guided DNA cleavage by IscB-ωRNA and mechanistic comparison with Cas9. Science 376, 1476 (2022).
Han, D. Y. et al. Development of miniature base editors using engineered IscB nickase. Nat. Methods 20, 1029 (2023).
Han, L. X. et al. Engineering miniature IscB nickase for robust base editing with broad targeting range. Nat. Chem. Biol. 20, 1629–1639 (2024).
Yan, H. et al. Assessing and engineering the IscB-ωRNA system for programmed genome editing. Nat. Chem. Biol. 20, 1617–1628 (2024).
Xue, N. N. et al. Engineering IscB to develop highly efficient miniature editing tools in mammalian cells and embryos. Mol. Cell 84, 3128–3140.e3124 (2024).
Guo, R. et al. Engineered IscB-ωRNA system with improved base editing efficiency for disease correction via single AAV delivery in mice. Cell Rep. 43, 114973 (2024).
Nielsen, S., Yuzenkova, Y. & Zenkin, N. Mechanism of eukaryotic RNA polymerase III transcription termination. Science 340, 1577–1580 (2013).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Wu, T. et al. An engineered hypercompact CRISPR-Cas12f system with boosted gene-editing activity. Nat. Chem. Biol. 19, 1384 (2023).
Hino, T. et al. An AsCas12f-based compact genome-editing tool derived by deep mutational scanning and structural analysis. Cell 186, 4920 (2023).
Tsuchida, C. A. et al. Chimeric CRISPR-CasX enzymes and guide RNAs for improved genome editing activity. Mol. Cell 82, 1199 (2022).
Tsai, S. Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33, 187–197 (2014).
Crooks, G. E., Hon, G., Chandonia, J.-M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Kannan, S. et al. Evolution-guided protein design of IscB for persistent epigenome editing in vivo. Nat. Biotechnol. https://doi.org/10.1038/s41587-025-02655-3 (2025).
Wang, X. et al. Cas12a base editors induce efficient and specific editing with low DNA damage response. Cell Rep. 31, 107723 (2020).
Richter, M. F. et al. Phage-assisted evolution of an adenine base editor with improved Cas domain compatibility and activity. Nat. Biotechnol. 38, 883–891 (2020).
Li, Z. F. et al. Engineering a transposon-associated TnpB-ωRNA system for efficient gene editing and phenotypic correction of a tyrosinaemia mouse model. Nat. Commun. 15, 831 (2024).
Zuo, E. W. et al. One-step generation of complete gene knockout mice and monkeys by CRISPR/Cas9-mediated gene editing with multiple sgRNAs. Cell Res. 27, 933–945 (2017).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Kato, K. et al. Structure of the IscB–ωRNA ribonucleoprotein complex, the likely ancestor of CRISPR-Cas9. Nat. Commun. 13, 6719 (2022).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Li, S. et al. Docking sites inside Cas9 for adenine base editing diversification and RNA off-target elimination. Nat. Commun. 11, 5827 (2020).
Chen, Y. et al. Synergistic engineering of CRISPR-Cas nucleases enables robust mammalian genome editing. Innovation 3, 100264 (2022).
Song, G. X. et al. AcrIIA5 inhibits a broad range of Cas9 orthologs by preventing DNA target cleavage. Cell Rep. 29, 2579 (2019).
Song, G. X. et al. Discovery of potent and versatile CRISPR-Cas9 inhibitors engineered for chemically controllable genome editing. Nucleic Acids Res. 50, 2836–2853 (2022).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Kawano, S., Ono, H., Takagi, T. & Bono, H. Tutorial videos of bioinformatics resources: online distribution trial in Japan named TogoTV. Brief. Bioinform. 13, 258–268 (2012).
Acknowledgements
This work was supported by grants from the National Key R&D Program of China (2022YFF0710700、2021YFF0702800、2020YFA0803501), the National Natural Science Foundation of China (32270567, 32070533), and Biological Resource Program of Chinese Academy of Sciences (KFJ-BRP-005). All grants are obtained by Y.T. We express our gratitude to Hui Yang and Yingsi Zhou from HUIDAGENE Therapeutics Inc. for their generous gift of plasmids. We are grateful to Junying Jia and Shu Meng (Protein Science Core Facility, Institute of Biophysics, Chinese Academy of Sciences) for their technical support in flow cytometry. We also appreciate Xiaoxiao Zhu and Zhuanzhuan Xing for their technical assistance with mice. We are thankful to Quan Meng for assisting with the local configuration of CRISPResso2 on the computer. The image of Fig. 6a, c are from TogoTV (© 2016 DBCLS TogoTV, CC-BY-4.0 https://creativecommons.org/licenses/by/4.0/) and have been modified.
Author information
Authors and Affiliations
Contributions
F.Z. conceived and designed the research. F.Z. and Y.P. performed most of the experiments. F.Z. analyzed the data. D.F. performed the microinjection experiment on the mouse embryo. G.S. provided technical suggestions and assistance. X.G. performed in vitro transcription of RNA. F.Z. wrote the paper. Y.T. directed the project and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
F.Z., Y.T., Y.P., D.F., G.S., and X.G. have filed a patent application (CN 120173914A) related to this work through the Institute of Biophysics, Chinese Academy of Sciences.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, F., Peng, Y., Fan, D. et al. Engineering a CRISPR-associated IscB system for developing miniature genome-editing tools in human cells and mouse embryos. Nat Commun 16, 10693 (2025). https://doi.org/10.1038/s41467-025-65724-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65724-w