Abstract
Acute myeloid leukemia (AML) is primarily driven by leukemic stem cells (LSCs), the main cause of relapse and therapy resistance. Here, we discover that LSCs are predominantly small and mechanically soft. These mechanical properties enable their selective isolation using microfluidic chips. Single-cell RNA-sequencing of primary human AML bone marrow identifies enrichment of LSCs within the FSClow ALDH1A1+ subpopulation, which exhibits long-term stemness in functional assays. Notably, inhibiting ALDH1A1 in these cells promotes F-actin polymerization and increases cellular stiffness, reducing their stemness while enhancing their susceptibility to natural killer (NK) cell-mediated cytotoxicity. In AML patient-derived xenograft models, the combination of ALDH1A1 inhibition with NK cell therapy markedly suppresses leukemia progression. These findings suggest that targeting the mechanical properties of LSC offers a promising strategy to overcome AML treatment resistance, providing insights into stem cell mechanobiology and paving the way for combining targeted therapies with immunotherapy to improve clinical outcomes.
Similar content being viewed by others
Introduction
Acute myeloid leukemia (AML) is an aggressive hematologic malignancy characterized by the uncontrolled accumulation of immature myeloid cells in the bone marrow, peripheral blood and other tissues1,2. Despite significant advances in treatment, the prognosis for AML remains poor, primarily due to the persistence of chemotherapy-resistant LSCs3,4,5. These cells not only initiate leukemia in immunocompromised mice, a standard model for studying LSCs, but also exhibit unique interactions with the immune system, often evading anti-tumor immune responses6,7. Thus, understanding the mechanisms underlying LSC maintenance and developing effective strategies to target these cells are critical for improving AML treatment outcomes.
Over the past decade, considerable progress has been made in unraveling the complex biology of LSCs, with efforts focusing on molecular8,9, cellular4,10, genetic11,12, epigenetic13,14, and metabolic aspects15, in both human and mouse models. Despite extensive investigations, the unique properties of LSCs are not fully understood. Emerging evidence suggests that the mechanical properties of cells may play a pivotal role in their functional behaviors, influencing cell fate and responses to therapy16,17. These properties, governed by the cytoskeleton—a dynamic network of polymers and associated proteins—are modulated by both intrinsic and extrinsic cues18,19,20. In stem cell biology, extrinsic mechanical cues, such as the rigidity of the extracellular matrix (ECM), provide structural support that affects stem cell behaviors, including self-renewal and differentiation20,21. Our previous work demonstrated that the intrinsic mechanical properties of hematopoietic stem cells (HSCs) are crucial for niche retention and regeneration22, and that corticotropin-releasing hormone enhances the homing and engraftment of hemopoietic stem and progenitor cells (HSPCs) by modulating cytoskeletal mechanics23. However, the mechanical properties of LSCs in AML, and their contribution to treatment resistance remain poorly understood. There is a need for further investigation into how changes in these mechanical properties might affect LSC behavior and potentially improve patient outcomes.
In this study, we explore the relationship between the mechanical properties of LSCs and their roles in AML progression and treatment resistance. Utilizing cutting-edge techniques such as microfluidic sorting, single-cell RNA sequencing (scRNA-seq), and mechanical phenotype characterization, we identify a specific LSC subpopulation, characterized by low forward scatter (FSClow) and high aldehyde dehydrogenase 1 family member A1 (ALDH1A1+) expression, which displays reduced cellular stiffness and enhanced stemness. Notably, inhibiting ALDH1A1, an enzyme previously known for its role in cancer stem cell maintenance, restores cellular stiffness and improves the efficacy of NK cell-based immunotherapy. These findings suggest that targeting the mechanical properties of LSCs could improve AML treatment outcomes, offering promising avenues for combination therapies that integrate mechanical modulation with immunotherapy.
Results
Small AML cells exhibit dominant stem cell potential and increased leukemia burden
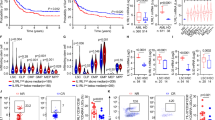
AML is characterized by substantial cellular heterogeneity, yet the impact of cell size on leukemia cell function remains largely unexplored. To investigate this, we performed Wright-Giemsa staining on the human leukemia cell lines KG-1a and U937, revealing a marked difference in cell size among the leukemia cells (Fig. 1a and Supplementary Fig. 1a). To assess whether these size differences are related to leukemia cell function, we isolated small and large cells from KG-1a and U937 by FACS sorting (Fig. 1b), and confirmed the separation purity by measuring the cell size (Fig. 1c). Interestingly, small leukemia cells displayed a significantly enhanced capacity for colony formation compared to their larger counterparts in both KG-1a and U937 cells (Fig. 1d and Supplementary Fig. 1b), suggesting a potential link between cell size and LSC potential.
a Wright-Giemsa staining of KG-1a cells showing small (red arrows) and large (blue arrows) cells. b Flow cytometry-based sorting of small and large KG-1a cell subsets by forward scatter (FSC-A). c Quantification cell size (n = 30 cells per group). d Colony formation by sorted small and large KG-1a cells (n = 3 biologically independent experiments). e–h Experimental design of KG-1a-Luc transplantation into M-NSG mice (3 × 105 cells per mouse) (e). Representative bioluminescence imaging (f) and total flux (p/s) quantification (g) in KG-1a-Luc engrafted mice. Kaplan-Meier survival curves of engrafted mice (h) (n = 6 mice per group). i Wright-Giemsa staining of primary AML BM Lin- cells. j–r In vitro and in vivo analysis of small and large primary BM Lin- cells from AML patients. Schematic overview (j). Representative sorting strategy of small and large primary BM Lin- cells by FSC-A (k). Quantification of cell size (l) (n = 50 cells per group from five AML samples). G0 phase distribution (m, n = 6), CD11b expression (n, left, n = 8), and CD14 expression (n, right, n = 5) in small and large primary AML BM Lin- cells. Colony formation by sorted small and large BM Lin- cells (o) (n = 5 biologically independent experiments). Flow cytometry of human CD45+ cell engraftment in mouse BM post-transplantation (p) (n = 6 mice per group). Hematoxylin eosin (HE) staining of BM from recipient mice (q) (n = 3 mice per group). Limiting dilution assay assessing leukemic initiating potential in M-NSG mice (r) (n = 4 mice per group). For (c, d, g, l, o, p, r), data were analyzed using two-tailed unpaired Student’s t test. For (h), data were analyzed using log-rank test (No adjustment for multiple comparisons was made). For (m and n), data were analyzed using two-tailed paired t test. Data are mean ± SEM and representative of at least three independent experiments. Source data are provided as a Source Data file.
To further investigate the stemness potential of small leukemia cells in vivo, we established a leukemia xenograft model by injecting small and large KG-1a-Luc cells into immunodeficient NOD-PrkdcscidIl2rgem1/Smoc (M-NSG) mice via the tail vein (Fig. 1e). Bioluminescence imaging showed a notably higher leukemic burden in mice injected with small cells (Fig. 1f, g). Consistently, the small-cell group exhibited significantly lower survival rates compared to the large-cell group (Fig. 1h). Similar results in U937 cells confirmed that small cells exhibited higher colony formation capacity and leukemic burden compared to large cells (Supplementary Fig. 1b, c).
To explore whether these findings extend to human primary AML cells, we performed Wright-Giemsa staining on primary bone marrow (BM) lineage-negative (Lin-) cells from AML patients, identifying both small and large leukemia cell populations (Fig. 1i). Using a combination of in vitro and in vivo assays, we isolated these populations for functional testing (Fig. 1j-l). Small cells were notably enriched in the G0 phase, exhibited a less differentiated phenotype, and displayed increased colony-formation capacity, indicative of LSC-like properties (Fig. 1m-o). In a patient-derived xenograft (PDX) model, mice engrafted with small primary BM Lin- cells exhibited a greater leukemic burden in the BM, accompanied by extensive leukemic cell infiltration and expansion (Fig. 1p, q). Remarkably, small leukemia cells significantly increased the frequency of LSCs in the PDX model, as demonstrated by limiting dilution assays (LDA) (Fig. 1r).
Given the treatment challenges associated with LSCs, we examined the sensitivity of small versus large AML cells to Ara-C, a commonly used cell-cycle-specific chemotherapy agent in AML therapy. While Ara-C effectively suppressed large AML cells, small leukemia cells exhibited marked resistance (Supplementary Fig. 1d), underscoring their potential role in disease persistence and relapse. Together, our findings suggest that smaller cell size is associated with enhanced LSC potential, which may contribute to increased leukemogenesis and poorer prognosis in AML.
Cytoskeletal dynamics and mechanical properties of small leukemia cells modulate LSC Potential in AML
Cytoskeletal remodeling, especially actin dynamics, plays a critical role in regulating cell size24,25. Changes in filamentous actin (F-actin) organization can lead to localized alterations in cell mechanics26,27. To investigate this, we assessed F-actin levels and cortical stiffness in small and large cells from KG-1a, U937, and primary BM Lin- cells from AML patients, using confocal microscopy and atomic force microscopy (AFM) (Fig. 2a-d and Supplementary Fig. 2a-d). Notably, small leukemia cells exhibited substantially lower F-actin levels compared to their larger counterparts (Fig.2b and Supplementary Fig. 2a, b). Correspondingly, small cells also exhibited markedly lower cortical stiffness (Fig. 2c and Supplementary Fig. 2c). Additionally, a significant correlation between cell stiffness and size was observed (Fig. 2d and Supplementary Fig. 2d), supporting the hypothesis that LSCs are characterized by small size and soft cortical properties. To test whether this relationship is conserved in normal hematopoiesis, we examined CD34⁺ HSPCs and CD34⁻ non-HSPCs from healthy bone marrow using AFM. Small CD34⁺ HSPCs displayed significantly lower stiffness than their larger counterparts and CD34⁻ non-HSPCs, and stiffness inversely correlated with size across the population (Supplementary Fig. 2e, f).
a Schematic of the AFM setting for cell cortical stiffness measurement. b Representative immunofluorescence images of F-actin (red) and nuclei (blue) in small and large AML BM Lin- cells, with quantification of F-actin intensity (n = 113 cells for small, and n = 191 cells for large; from four AML samples). c AFM-based stiffness measurement (n = 23 cells per group, from four AML samples). d Correlation between cell size and stiffness (n = 46 cells, from four AML samples). Imaging flow cytometry of F-actin (e) and the cell area (f) for Actinlow and Actinhigh cells from six AML samples. g–j F-actin staining (g, h, n = 5 AML samples) and stiffness quantification (i, n = 19 individual cells per group, from five AML samples) in small and large primary BM Lin- cells treated with 1 μM Jas or 1 μM Lat A for 6 h. Colony formation assay (j, n = 3 AML samples) of treated and untreated AML BM Lin⁻ cells (k, Overview of RNA-seq, in vivo xenografts, and drug sensitivity assays using size-sorted AML BM Lin⁻ cells treated with Jas or Lat A (n = 5 AML samples). l DEGs between small and large cells. m GSEA of DEGs between small and large cell. n DEGs between untreated and Jas-treated small cells. o GSEA of DEGs between untreated and Jas-treated small cells. p DEGs between untreated and Lat A-treated large cells. q GSEA of DEGs between untreated and Lat A-treated large cells. r Flow cytometry analysis of BM engraftment following transplantation of treated AML BM Lin⁻ cells (n = 4 mice per group, from three AML samples). s Cell viability assay following Ara-C treatment (48 h) in size-sorted AML BM Lin⁻ cells pretreated with Jas or Lat A (n = 5 AML samples). For (b, c, s), data were analyzed using two-tailed unpaired Student’s t test. For (d), data were analyzed using Pearson correlation. For (f, l–q), data were analyzed using two-tailed paired t test. For (h–j, r), data were analyzed using one-way ANOVA with Tukey correction (F = 38.13, 41.74, 24.37 and 56.32, respectively). Data are mean ± SEM and representative of at least three independent experiments. Source data are provided as a Source Data file.
Given the relationships among cell size, stemness, and cellular mechanics, we next examined whether cytoskeletal dynamics are crucial for maintaining LSC potential. We employed two cytoskeletal modulating agents: latrunculin A (Lat A, 1 μM) to disrupt actin filaments and soften leukemia cells, and Jasplakinolide (Jas, 1 μM) to stabilize actin filaments and stiffen cells. To assess the effects of cytoskeletal modulation, we isolated Actinlow and Actinhigh cells from KG-1a and U937 cells by FACS sorting (Supplementary Fig. 2g), and confirmed the separation by measuring F-actin content and cortical stiffness (Supplementary Fig. 2h-j). Jas treatment increased F-actin levels and stiffness in Actinlow cells, and also led to a significant increase in cell size (Supplementary Fig. 2h–k). In contrast, Lat A treatment decreased F-actin levels and stiffness in Actinhigh cells and reduced their cell size (Supplementary Fig. 2h–k). Functionally, these biomechanical changes were accompanied by altered clonogenicity. Jas-treated Actinlow cells exhibited markedly reduced colony-forming ability compared to untreated controls, whereas Lat A-treated Actinhigh cells showed increased clonogenicity (Supplementary Fig. 2l, m). Together, these findings demonstrate that F-actin dynamics regulate not only cell stiffness and size, but also functional stemness, highlighting the mechanobiological control of LSC potential in AML.
In primary human BM Lin- cells from AML patients, Actinlow cells were significantly smaller than Actinhigh cells (Fig. 2e, f), indicating that smaller, Actinlow leukemia cells may be enriched for LSCs. To test whether actin modulation directly affects LSC stemness, we treated small and large primary BM Lin- cells with Jas and Lat A. Consistent with previous findings, Jas treatment of small primary BM Lin- cells increased F-actin levels and cortical stiffness (Fig. 2g-i), but decreased colony formation (Fig. 2j). Conversely, Lat A treatment of larger cells decreased F-actin levels, reduced cortical stiffness, and enhanced colony formation (Fig. 2g-j). These findings suggest that in AML, a smaller cell size is essential for LSC maintenance, primarily through changes in the F-actin cytoskeleton.
To further investigate whether F-actin dynamics reprogram LSC-like properties at the molecular and functional levels, we conducted bulk RNA sequencing (RNA-seq), in vivo transplantation, and Ara-C sensitivity assays using size-sorted primary AML Lin- cells following pharmacologic modulation of actin polymerization (Fig. 2k). Bulk RNA-seq data revealed that small leukemic cells were significantly enriched for LSC and stemness-related signatures, while large cells displayed a more differentiated transcriptional profile (Fig. 2l–m). Treatment with Jas, which stabilizes F-actin, downregulated LSC-associated genes (e.g., FLT328, DNMT3B29,30, NOTCH428,31, CD3428,29,32 and CD13332) in small cells and suppressed LSC-related pathways (Fig. 2n–o). Conversely, disrupting F-actin with Lat A in large cells induced a stem-like transcriptional state, including upregulation of key LSC-related genes (e.g., NOTCH1/228, RUNX328, WNT9A33 and BCL2A/BCL2L134) and enrichment of LSC gene sets (Fig. 2p–q). Functionally, Lat A–treated large cells exhibited enhanced leukemogenicity in xenograft models, whereas Jas-treated small cells showed markedly reduced engraftment (Fig. 2r). Moreover, F-actin stabilization by Jas sensitized small leukemic cells to Ara-C treatment, while actin disruption by Lat A conferred chemoresistance in large cells (Fig. 2s). Together, these findings demonstrate that actin cytoskeletal dynamics regulate not only mechanical properties but also LSC-associated transcriptional programs, leukemogenic potential, and chemotherapeutic response.
Microfluidic isolation of SSLCs with enhanced stem cell potential in AML
Given that smaller leukemia cells exhibit reduced F-actin levels and lower cortical stiffness (Fig. 2 and Supplementary Fig. 2), and that cell deformability is primarily influenced by softness35, we next sought to investigate the mechanical properties and functional characteristics of these cells by isolating soft-small leukemic cells (SSLCs) and stiff-large leukemic cells (SLLCs). To achieve this, we developed a microfluidic sorting device that differentiates cells based solely on their mechanical properties, specifically stiffness and deformability. The device consists of two main components: a microweir and a channel chamber (Supplementary Fig. 3a, b). The microweir with height restrictions allows smaller, softer cells to pass more easily under the same hydraulic pressure, whereas larger, stiffer cells take longer to pass or may fail to do so. The microweir was calibrated to a height of 12 μm for sorting U937 cells and 5 μm for sorting primary AML cells. The channel chamber, designed with a height of 40 μm, accommodates cells that fail to pass through the microweir, preventing blockage. Additionally, partitions within the chamber prevent interference from incomplete cell clusters during sorting. Utilizing this device, we successfully isolated SSLCs and SLLCs from primary BM Lin- cells from AML patients and U937 cells (Fig. 3a and Supplementary Fig. 3c). The separation purity was confirmed by in vitro assays, where SSLCs exhibited lower F-actin levels, reduced cell size, and decreased cortical stiffness compared to SLLCs (Fig. 3b-d and Supplementary Fig. 3d-g). Furthermore, a significant correlation was observed between cell size and stiffness (Fig. 3e and Supplementary Fig. 3h).
a Schematic overview of soft-small leukemic cells (SSLCs) and stiff-large leukemic cells (SLLCs) in primary BM Lin- cells isolated by microfluidic devices. b Representative immunofluorescence images showing F-actin (red) and nuclei (blue) in SSLCs and SLLCs from five independent AML patient samples; each dot represents an individual cell (n = 297 cells per group). c Quantification of cell size in SSLCs and SLLCs (n = 21 cells per group, from five AML samples). d Cortical stiffness measurements by AFM for SSLCs and SLLCs from primary BM Lin- cells (n = 21 cells per group, from five AML samples). e Correlation between cell size and cortical stiffness in SSLCs and SLLCs from primary BM Lin- cells (n = 42 cells, from five AML samples). Flow cytometry analysis showing quantification of G0 phase (f) (n = 4 AML samples), CD11b expression (g) (n = 7 AML samples), and CD14 expression (h) (n = 4 AML samples) in SSLCs and SLLCs from primary BM Lin- cells. i Colony formation assay showing the number of colonies formed by SSLCs and SLLCs of BM Lin- cells from seven primary AML samples. Cells were seeded at 1 × 105 cells per well, with colony counts enumerated (n = 7 biologically independent experiments). j, k Flow cytometry analysis illustrating BM engraftment of SSLCs and SLLCs in primary BM Lin- cells from four independent AML patients (5 × 106 cells per mouse). Mice were sacrificed 2-3 months post- engraftment, and their BM was analyzed by flow cytometry (j) (n = 5 mice per group) or HE staining (k) (n = 3 mice per group). For (b–d, j), data were analyzed using by two-tailed unpaired Student’s t test. For (e), data were analyzed using Pearson correlation. For (f–h, i), data were analyzed using two-tailed paired t test. Data are mean ± SEM and representative of at least three independent experiments. Source data are provided as a Source Data file.
Further analysis of primary BM Lin- cells from AML patients showed that SSLCs were predominantly in the G0 phase of the cell cycle, exhibited lower differentiation levels as indicated by reduced CD11b and CD14 expression (Fig. 3f-h), and demonstrated significantly higher colony-forming ability compared to SLLCs (Fig. 3i and Supplementary Fig. 3i). These findings underscore the enhanced stem cell-like potential of SLLCs in AML.
To evaluate the role of SSLCs in AML maintenance and progression in vivo, we established two leukemia xenograft models by injecting SSLCs and SLLCs from primary BM Lin- cells from AML patients and U937 cells into M-NSG mice via the tail vein. Consistent with the in vitro findings, mice injected with SSLCs exhibited a significantly higher leukemic burden compared to those injected with SLLCs (Fig. 3j and Supplementary Fig. 3j). Hematoxylin-eosin (HE) staining confirmed a marked expansion and infiltration of leukemic cells in the BM of the SSLC-injected mice (Fig. 3k and Supplementary Fig. 3k). Collectively, our results indicate that SSLCs, isolated via microfluidic sorting, are enriched for LSCs in AML, with small cell size and cortical softness emerging as key physical characteristics of these cells.
The FSClow ALDH1A1+ subpopulation effectively enriches LSCs in primary AML patients
To explore the mechanisms underlying the enhanced LSC potential in smaller leukemia cells, we performed single-cell RNA sequencing (scRNA-seq) on FACS-sorted small (FSClow) and large (FSChigh) BM Lin- cells from primary AML patients (Fig. 4a). This analysis yielded 9943 high-quality single cells, including 5608 small cells and 4335 large cells (Fig. 4b and Supplementary Fig. 4a, b). We identified six distinct cell clusters, each expressing unique marker genes (Supplementary Data 1), representing the primary types of leukemia cells within the dataset (Fig. 4b, c).
a Schematic diagram showing scRNA-seq analysis of size-sorted small and large primary BM Lin- cells from four AML patients. b UMAP plots showing six cell clusters among a total of 9943 cells, colored by cluster. c Violin plots showing the expression of canonical gene markers across each cluster. d Box plots displaying the LSC 6 score in each cluster from scRNA-seq data (n = 282 for LSC, 582 for EPC, 3134 for GMP, 137 for DC-like, 2269 for HSPC, and 3539 for Blast). Each dot represents a single cell. The boxes extend from the 25th to the 75th percentiles, with a distinct line marking the median. Whiskers extend to the maximum and minimum data points. e Heatmap showing the average expression of the top five gene markers in each subset. f Violin plot illustrating ALDH1A1 expression across each cluster. g Representative immunofluorescence images of ALDH1A1 (red) in small and large primary BM Lin- cells of three AML samples. Nuclei are stained with DAPI (blue). Each dot represents an individual cell (n = 158 cells per group). Flow cytometry analysis quantifying the percentages of ALDH1A1+ (h) (n = 7 AML samples) and ALDEFLUOR+ (i) (n = 8 AML samples) cells in small and large primary BM Lin- cells. j Pie charts showing the percentages of LSCs in the small (FSClow) and large (FSChigh) (j), and in FSClow ALDH1A1+ and FSClow ALDH1A1- subpopulations (k) within primary BM Lin- cells from AML patients, based on scRNA-seq data. For (g), data were analyzed using two-tailed unpaired Student’s t test. For (h, i), data were analyzed using two-tailed paired t test. For (g–i), data are mean ± SEM and representative of at least three independent experiments. Source data are provided as a Source Data file.
To characterize LSCs, we analyzed LSC-specific gene signatures across these clusters (Supplementary Data 2). Both the LSC630 and LSC4836 scores were notably elevated in the LSC cluster, confirming their enrichment (Fig. 4d and Supplementary Fig. 4c, d). Developmental pseudotime analysis showed an advanced differentiation trajectory for the LSC cluster (Supplementary Fig. 4e-g), and cell cycle analysis indicated that LSCs were predominantly in the G0/G1 phase, suggesting a quiescent state (Supplementary Fig. 4h).
By examining genes highly expressed in LSCs, we identified ALDH1A1, SPINK2, and LGALS3BP as key candidates (Fig. 4e). This finding was further supported by public bulk RNA-seq data (GSE199452), which revealed significantly higher expression levels of SPINK2 and LGALS3BP in LSCs with poorer prognosis than in non-LSCs (Supplementary Fig. 4i-k). Notably, elevated ALDH1A1 expression in LSCs correlated with poor overall survival in AML patients (Fig. 4f and Supplementary Fig. 4j, k). ALDH1A1, an enzyme within the aldehyde dehydrogenase (ALDH) family, plays a critical role in cellular detoxification and is linked to stem cell characteristics such as self-renewal and resistance to oxidative stress37. The activity of this enzyme is frequently increased in cancer stem cells (CSCs), contributing to enhanced stemness and therapy resistance38,39.
To assess ALDH1A1 expression in different AML cell subpopulations, we conducted immunofluorescence and FACS analyses of human primary AML BM Lin- cells. Both analyses confirmed that smaller AML subpopulations exhibited significantly higher levels of ALDH1A1 than did larger cells (Fig. 4g, h). Furthermore, using the ALDEFLUOR+ assay—a standard method for identifying cells with high ALDH activity40—we observed a higher proportion of ALDEFLUOR+ cells in the smaller AML subpopulations than in the larger ones (Fig.4i). These findings suggest that ALDH1A1 may play a crucial role in LSC-enriched, small leukemia populations in AML.
Consistent with their stemness characteristics, LSCs were significantly enriched in the small (FSClow, 5.2%) group compared with the large (FSChigh, 1.4%) group (Fig. 4j), aligning with the observed increase in leukemic burden in small AML cells (Fig. 1p-r). To further investigate the role of ALDH1A1 in LSC maintenance, we analyzed the FSClow ALDH1A1+ subset in our scRNA-seq data. This subset exhibited remarkable enrichment, with 80.3% of FSClow ALDH1A1+ cells displaying LSC characteristics (Fig. 4k)—approximately a 16-fold increase over the general FSClow group (5.2%) (Fig. 4j). Moreover, the proportion of LSC in FSClow ALDH1A1+ cells was 5.3 times higher than that in ALDH1A1+ cells alone, supporting previous observations in AML patients41. Overall, these findings highlight the FSClow ALDH1A1+ subpopulation as a significantly enriched source of LSCs, offering a potential strategy for more effective LSC identification in AML.
FSClow ALDH1A1+ cells display unique cytoskeletal features and enhanced LSC potential in AML
To identify the molecular characteristics of FSClow ALDH1A1+ cells that contribute to LSC enrichment in AML, we performed differential expression gene (DEG) analysis (Supplementary Data 3) of downregulated genes (down-DEGs) in the FSClow ALDH1A1+ subpopulation compared with other populations within primary AML BM Lin- cells in our scRNA-seq data. This analysis identified 156 protein-coding genes (Supplementary Data 4) that were significantly downregulated in FSClow ALDH1A1+ cells (Fig. 5a). Gene Ontology (GO) analysis (Supplementary Data 5) of these down-DEGs revealed that they were mainly enriched in actin cytoskeleton-related pathways (Fig. 5b). Further F-actin scoring confirmed that FSClow ALDH1A1+ cells exhibited markedly lower F-actin-related mechanics compared to other groups (Fig. 5c).
a Venn diagram showing the overlap of downregulated DEGs between FSClow ALDH1A1+ and FSClow ALDH1A1-, FSChigh ALDH1A1-, and FSChigh ALDH1A1+ BM Lin- cells from four AML samples, based on scRNA-seq data. b GO term analysis of overlapping down-DEGs in (a), illustrating cytoskeletal- and actin-related pathways. c F-actin gene scores in the indicated cell populations based on scRNA-seq. d Representative immunofluorescence images of F-actin (red) and nuclei (blue) in FSClow, ALDH1A1+ and FSClow ALDH1A1+ AML BM Lin- cells, with quantification of F-actin intensity (n = 60, 68 and 64 cells for the respective groups, from five AML samples). e AFM-based stiffness measurements for FSClow, ALDH1A1+ and FSClow ALDH1A1+ cells (n = 11, 23, 25 cells for the respective groups, from five AML samples). f Colony formation assay of FSClow, ALDH1A1+ and FSClow ALDH1A1+ AML BM Lin- cells (n = 3 biologically independent experiments). g Representative immunofluorescence images and quantification of F-actin (red) in FSClow ALDH1A1+ AML BM Lin- cells (n = 60 cells per group, from five AML samples). h AFM-based stiffness measurements for FSClow ALDH1A1+ cells after Jas treatment (n = 42 cells per group, from five AML samples). i G0 phase proportions by flow cytometry (n = 6 AML samples). j Colony formation assay for FSClow ALDH1A1+ cells after Jas treatment (n = 5 biologically independent experiments). Flow cytometry plots (k) and statistical analyses (l–n) of human CD45+ cell chimerism in the BM of M-NSG recipients after primary transplantation with sorted FSClow, ALDH1A1+ and FSClow ALDH1A1+ cells (1 × 10⁵ cells per mouse) (l, n = 4 mice), secondary (m, n = 6 mice) and tertiary recipients (n, n = 5 mice) were transplanted with 1 × 106 BM cells. For (b), data were analyzed using Wilcoxon Rank Sum Test. For (d–f, l–n), data were analyzed using one-way ANOVA with Tukey correction (F = 58.03, 22.71, 40.04, 115.05, 34.12 and 82.09, respectively). For (g, h and j), data were analyzed using two-tailed unpaired Student’s t test. For (i), data were analyzed using two-tailed paired Student’s t test. Data are mean ± SEM and representative of at least three independent experiments. Source data are provided as a Source Data file.
Given that ALDH1A1 is a key marker for LSCs (Fig. 4e, f), we evaluated its expression in small leukemia cells, finding a range from 3% to 33% (Fig. 4h, i). We then examined the LSC potential of ALDH1A+ and FSClow ALDH1A1+ subpopulations in vitro. Compared with FSClow or ALDH1A1+ cells, FSClow ALDH1A1+ cells from AML patients displayed significantly lower F-actin levels and cortical stiffness, but greater colony formation ability (Fig. 5d-f), indicating that ALDH1A1 may contribute to the unique mechanical properties and enhanced stem-like potential observed in LSC-enriched FSClow ALDH1A1+ cells. Given the distinct F-actin and stiffness profiles of FSClow ALDH1A1+ cells, we next examined whether modulation of the cytoskeleton could impact these LSC-enriched characteristics. To explore this, we isolated FSClow ALDH1A1+ cells from primary BM Lin- AML samples and treated them with Jas to promote F-actin polymerization and increase cortical stiffness (Fig. 5g, h). Jas treatment effectively enhanced F-actin polymerization, increased cortical stiffness, reduced the proportion of cells in the G0 phase, and decreased colony formation in FSClow ALDH1A1+ cells (Fig. 5g–j). These results emphasize the role of the F-actin cytoskeleton in regulating the mechanical properties and stem-like potential of FSClow ALDH1A1+ cells.
To further validate the LSC properties of FSClow ALDH1A1+ cells in vivo, we performed long-term serial transplantation experiments. Notably, FSClow ALDH1A1+ cells exhibited significantly enhanced engraftment not only in primary and secondary recipients but also in tertiary recipients, compared with other subpopulations (Fig. 5k–n), confirming their robust functional stemness. These in vivo results provide compelling evidence that FSClow ALDH1A1+ cells represent a highly enriched LSC subpopulation with sustained repopulating capacity. Together, these findings support the notion that FSClow ALDH1A1+ cells represent a mechanically and functionally distinct LSC-enriched subpopulation in AML.
ALDH1A1 inhibition enhances F-actin polymerization and stiffness while reducing stemness in LSC-enriched AML cells
To investigate the role of ALDH1A1 in cell mechanics and its contribution to LSC potential, we treated FSClow ALDH1A1+ cells isolated from primary BM Lin- cells of AML patients with the ALDH1 inhibitor Disulfiram in vitro (Fig. 6a). Compared with FSClow ALDH1A1- cells, FSClow ALDH1A1+ cells displayed lower F-actin levels, reduced cellular stiffness, and higher colony-forming ability (Fig. 6b-d), which was consistent with previous findings (Fig. 5d-f). Disulfiram treatment induced a marked increase in F-actin polymerization and cortical stiffness in FSClow ALDH1A1+ cells, along with a significant reduction in colony formation, suggesting that ALDH1A1 regulates F-actin organization and LSC-like properties (Fig. 6b-d).
a Schematic of in vitro analysis of FSClow ALDH1A1+ and FSClow ALDH1A1- primary BM Lin- cells isolated from AML patients by FACS sorting. b Representative immunofluorescence images of F-actin (red) and nuclei (blue) in FSClow ALDH1A1- and FSClow ALDH1A1+ AML BM Lin- cells treated with 1 μM Disulfiram for 48 h (n = 71, 68, and 71 cells for the respective groups, from three AML samples). c Cortical stiffness measurements by AFM in the same cell groups (n = 20 cells per group, from three AML samples). d Colony formation assay of the indicated cells (n = 5 biologically independent experiments). e Schematic of in vitro and in vivo assays of FSClow ALDH1A1+ KG-1a cells with ALDH1A1 knockdown. f Representative immunofluorescence images of F-actin (red) and nuclei (blue) in FSClow ALDH1A1+ KG-1a cells with ALDH1A1 knockdown. with quantification of F-actin intensity (n = 21 cells for sh-NC, n = 16 cells for shALDH1A1#1, and n = 15 cells for shALDH1A1#2). g AFM-based stiffness measurements for FSClow ALDH1A1+ KG-1a cells with ALDH1A1 knockdown (n = 10 cells for sh-NC, n = 30 cells for shALDH1A1#1, and n = 16 cells for shALDH1A1#2). h Colony formation assays for FSClow ALDH1A1+ KG-1a cells with ALDH1A1 knockdown (n = 3 biologically independent experiments). i, j Flow cytometry analysis illustrating BM engraftment of FSClow ALDH1A1+ KG-1a cells with ALDH1A1 knockdown (1 × 105 cells per mouse). Mice were sacrificed 20-30 days post-engraftment, and the BM was analyzed by HE staining (i) and flow cytometry (j) (n = 6 mice per group). For (b–d, f–h and j), data were analyzed using one-way ANOVA with Tukey correction (F = 106.90, 47.08, 21.45, 577.1, 47.45, 58.71 and 28.51, respectively). Data are mean ± SEM and representative of at least three independent experiments. Source data are provided as a Source Data file.
To further explore this regulatory role, we used shRNA to knock down ALDH1A1 in FSClow ALDH1A1+ KG-1a cells and examined their LSC potential both in vitro and in vivo (Fig. 6e). Knockdown of ALDH1A1 was confirmed by a substantial reduction in both the mRNA and protein levels (Supplementary Fig. 5a, b). In FSClow ALDH1A1+ KG-1a cells, ALDH1A1 knockdown led to enhanced F-actin polymerization, increased cortical stiffness (Fig. 6f, g), and reduced colony formation (Fig. 6h), indicating that ALDH1A1 is essential for maintaining LSC potential.
To assess the impact of ALDH1A1 inhibition on leukemia progression in vivo, we established a xenograft model by transplanting ALDH1A1-knockdown FSClow ALDH1A1+ KG-1a cells into M-NSG mice. Consistent with in vitro findings, ALDH1A1 knockdown significantly reduced the leukemic burden in the BM (Fig. 6i, j) and spleen (Supplementary Fig. 5c) compared with that in the controls. Overall, these data support the hypothesis that ALDH1A1 inhibition reduces stemness while promoting F-actin polymerization and increasing cellular stiffness, thereby reducing LSC potential in FSClow ALDH1A1+ leukemic cells.
Leukemia cell stiffening enhances the efficacy of NK cell immunotherapy by ALDH1A1 inhibition
Our recent studies have demonstrated that enhancing immunological synapse (IS) formation can improve NK cell cytotoxicity against leukemia cells42,43. IS formation relies on F-actin remodeling, which facilitates the polarization of lytic granules such as perforin and granzymes toward the synapse and the microtubule-organizing center (MTOC)44,45. This cytoskeletal remodeling is essential for effective NK cell-mediated killing46,47. Since ALDH1A1 inhibition promotes F-actin polymerization and increases cellular stiffness in LSC-enriched AML cells (Fig. 6), we hypothesized that ALDH1A1 inhibition might improve NK cell-mediated cytotoxicity by increasing leukemia cell stiffness.
To test this hypothesis, we treated FSClow ALDH1A1+ KG-1a cells with the ALDH1 inhibitor Disulfiram to increase cellular stiffness and examined the effect on NK cell-mediated killing (Fig. 7a). Compared with FSClow ALDH1A1+ cells, FSClow ALDH1A1- cells exhibited enhanced F-actin polymerization, increased cortical stiffness, and reduced colony-forming ability (Supplementary Fig. 6a-c). Disulfiram treatment further enhanced F-actin polymerization and stiffness in FSClow ALDH1A1+ cells (Supplementary Fig. 6d, e). To assess the effect of increased stiffness on NK cell interactions, we co-cultured Disulfiram-treated FSClow ALDH1A1+ cells with NK cells and stained for perforin and F-actin to evaluate IS formation (Fig. 7b). We quantified the polarization index by measuring the distance of perforin from the IS (Fig. 7c). Disulfiram treatment significantly promoted perforin polarization toward the IS (Fig. 7b-d) and enhanced F-actin levels and stiffness (Supplementary Fig. 6d, e), thereby strengthening IS formation and improving NK cell cytotoxicity (Fig. 7e). Additionally, co-culture of Disulfiram-treated FSClow ALDH1A1+ cells with NK cells resulted in a significant reduction in colony formation, indicating increased NK cell-mediated lysis of LSC-enriched leukemic cells (Fig. 7f). Similarly, ALDH1A1 knockdown in FSClow ALDH1A1+ KG-1a cells increased their susceptibility to NK cell-mediated lysis (Supplementary Fig. 6f). Compared with control cells, NK cells co-cultured with ALDH1A1-knockdown FSClow ALDH1A1+ KG-1a cells presented a significant reduction in colony number (Supplementary Fig. 6g). To further visualize the dynamic interactions between NK cells and LSCs, we performed live-cell time-lapse imaging. Disulfiram-treated FSClow ALDH1A1+ KG-1a cells exhibited markedly increased susceptibility to NK cell–mediated cytotoxicity compared to untreated cells (Supplementary Movie 1, Fig. 7g-i). These real-time observations provide direct evidence that Disulfiram disrupts LSC-mediated immune evasion and enhances NK cell killing efficiency.
a Schematic of in vitro analysis of FSClow ALDH1A1+ KG-1a cells treated with 1 μM Disulfiram for 48 h. b Confocal images showing F-actin (red) and perforin (green) localization at the IS in target-NK cell conjugates at different time points. c Diagram illustrating the calculation of polarization index, based on the distance from perforin to the IS center. d Quantification of the polarization index from six donors. Each dot represents an individual conjugate. For the Control group, n = 16 (30 min), 27 (1 h), 36 (2 h), and 31 (4 h); for the Disulfiram group, n = 16 (30 min), 42 (1 h), 21 (2 h), and 22 (4 h). e Flow cytometric analysis of target cell lysis after co-culture with CB-NK cells (E:T = 5:1, 6 h; n = 6 biologically independent experiments). f Colony formation assay of target cells post NK cell exposure (n = 6 biologically independent experiments). Live-cell imaging of NK cell cytotoxicity: representative killing snapshots (g), NK cell killing time (minutes) (h), and cytotoxicity rate (%) (i) in control vs. Disulfiram-treated target cells (n = 3 biologically independent experiments). j Schematic of in vivo AML PDX model: NSG mice were transplanted with primary AML blasts (7 × 10⁶ cells/mouse), treated with Disulfiram (15 mg/kg, i.p., days 1–5) ± CB-NK cell infusion (5 × 10⁶ cells, i.v., day 6). k Flow cytometry of hCD45⁺ leukemic cells in mouse BM at endpoint (n = 4 mice for Control, 5 for Disulfiram, 5 for NK, and 5 for Disulfiram + NK). l Representative HE staining of BM sections. m Diagram visually illustrating the enhanced efficacy of NK cell immunotherapy through ALDH1A1 inhibition-mediated stiffening of leukemic cells. For (e), data were analyzed using two-tailed paired t test. For (d, f, h, i), data were analyzed using two-tailed unpaired Student’s t test. For (k), data were analyzed using one-way ANOVA with Tukey correction (F = 42.17). Data are mean ± SEM and representative of at least three independent experiments. Source data are provided as a Source Data file.
Notably, NK cells co-cultured with Disulfiram-treated leukemic cells did not exhibit enhanced effector molecule expression of IFN-γ, TNF-α, granzyme B, perforin, or CD107a (Supplementary Fig. 6h). Next, we investigated whether Disulfiram directly affects NK cell function. Treatment of cord blood (CB)-derived NK (CB-NK) cells with Disulfiram alone did not alter their proliferation or apoptosis, nor did it impair their cytotoxicity against KG-1a target cells (Supplementary Fig. 6i–l). Furthermore, direct exposure to Disulfiram had no significant impact on the expression of IFN-γ, TNF-α, granzyme B, perforin, or CD107a in NK cells following target cell stimulation (Supplementary Fig. 6m). We next examined NK ligand expression on leukemic cells. Baseline levels of MICA/MICB and HLA-A/B/C were comparable between FSClow ALDH1A1⁺ cells and the broader FSClow population (Supplementary Fig. 6n, o). Upon Disulfiram treatment, MICA/MICB, but not HLA-A/B/C, were significantly upregulated (Supplementary Fig. 6p, q), consistent with stress-induced sensitization to NK cells. Collectively, these findings indicate that the increased NK cell-mediated killing is not a result of enhanced NK cell activation, but is more likely due to Disulfiram-induced changes in the mechanical properties of the leukemic target cells.
To evaluate the therapeutic potential of targeting the mechanical properties of leukemic cells in vivo, we combined Disulfiram treatment with NK cell infusion in an AML PDX model (Fig. 7j). Compared with Disulfiram or NK cell treatment alone, the combined treatment significantly reduced leukemogenesis (Fig. 7k). Both Disulfiram and NK cell treatments individually decreased leukemia burden, but the combination therapy eradicated the majority of leukemia cells in the BM (Fig. 7k, l). HE staining confirmed the reduced leukemic burden in mice treated with the combination therapy (Fig. 7l). To investigate the safety of ALDH1A1 inhibition in normal hematopoiesis, we assessed its expression and functional impact in human CB-derived CD34+ HSPCs. ALDH1A1 was highly expressed in CD34+ HSPCs (Supplementary Fig. 6r), but treatment with Disulfiram, a pharmacologic ALDH1A1 inhibitor, did not impair HSPC viability or clonogenic potential in colony-forming assays (Supplementary Fig. 6s–t). These results are consistent with a previous study in murine models showing that genetic deletion of ALDH1A1 does not compromise HSC self-renewal or hematopoietic recovery under homeostatic or stress conditions48. Together, our data indicate that normal HSPCs are functionally preserved upon ALDH1A1 inhibition, supporting the therapeutic window for selectively targeting ALDH1A1+ LSCs.
Taken together, these findings support the hypothesis that NK cell immunotherapy, when combined with ALDH1A1 inhibition to increase cytoskeletal stiffness in LSCs, enhances anti-leukemic efficacy. This strategy could improve treatment outcomes and reduce the risk of AML recurrence by increasing the susceptibility of LSCs to immune-mediated elimination (Fig. 7m).
Discussion
Despite substantial progress in understanding the biology of human LSCs, developing effective therapies targeting these cells remains challenging. Traditional approaches have largely focused on the biochemical characteristics, such as the molecular and genetic features, of LSCs, often overlooking the potential role of their mechanical properties8,9,10,11,12,13,49,50,51,52,53,54,55. This study addresses this gap by uncovering a critical relationship between the mechanical properties of LSCs and their contribution to AML progression and therapy resistance. Our results reveal that human LSCs are predominantly small, soft, and enriched in the FSClow ALDH1A1+ subpopulation, validated for long-term stemness potential. Notably, targeting ALDH1A1, a key regulator in this subpopulation, alters the mechanical properties of LSCs, leading to increased cellular stiffness, decreased stemness, and enhanced susceptibility to NK cell-mediated cytotoxicity. These findings highlight LSC mechanics as a promising therapeutic target in AML, offering alternative avenues for intervention.
While stem cells are generally characterized by their small size, whether this feature is essential for their function remains unclear. Recent research has suggested that small cell size is important for HSC function in vivo56. However, the relationship between cell size and LSCs is not well understood. In this study, we demonstrate that smaller leukemic cells exhibit enhanced stem cell potential, characterized by increased colony formation, leukemogenicity, and drug-resistance. We also establish a link between cell size and mechanical properties. These smaller leukemic cells show reduced F-actin cytoskeleton organization and decreased stiffness, which corresponds to their enhanced LSC characteristics. Notably, increasing F-actin levels and cellular stiffness in these cells suppressed their malignant phenotype, as evidenced by reduced in vivo leukemogenicity, diminished stemness, and decreased chemoresistance. These findings provide direct evidence that the cytoskeletal architecture and mechanical properties of LSCs are functionally coupled to their stem-like behavior. Our findings align with growing evidence that CSCs often display distinct mechanical properties compared with differentiated cells, with reduced stiffness associated with increased stem cell potential18,57,58,59. Importantly, we show that cellular softness is not merely a passive attribute but a modifiable property that directly contributes to LSC function and therapy resistance.
ALDH1A1, an NADP-dependent enzyme in the ALDH family, plays a critical role in both physiological and pathological processes60,61. It has been identified as a marker for tumor-initiating cells, playing a key role in maintaining stemness, angiogenesis, metastasis, and drug resistance38,39,62,63. In AML, ALDH1A1 supports the survival of leukemia-initiating cells and is essential for establishing human AML xenografts in mice63,64. In the present study, using scRNA-seq and functional assays, we identified an LSC-enriched FSClow ALDH1A1+ subpopulation with enhanced stemness. We also uncovered a role for ALDH1A1 beyond its identity as a stem cell marker. It acts as a key regulator of the mechanical properties of LSCs, specifically by modulating F-actin dynamics to influence cell stiffness. This newly identified role of ALDH1A1 links its activity to both stem cell maintenance and the biophysical characteristics that enable LSCs to resist therapeutic interventions. These findings provide a rationale for targeting cytoskeletal regulators, such as F-actin, to modify the physical properties of LSCs and reduce their therapeutic resistance. Future research should further elucidate the molecular mechanisms by which ALDH1A1 regulates F-actin dynamics and stiffness in LSCs, and explore whether other mechanical regulators contribute to LSC maintenance in AML, as well as in the progression of other types of cancer.
Another important finding in this work is that stiffening leukemic cells via ALDH1A1 inhibition enhances NK-cell cytotoxicity and improves the efficacy of adoptive NK-cell therapy against leukemia in AML PDX models. NK cell cytotoxicity relies on the formation of lytic IS with target cells, a process that requires F-actin-dependent plasma membrane remodeling44,45. This remodeling drives the migration and polarization of lytic granules towards the synapse, enabling cytotoxic exocytosis46,47. Our recent research has demonstrated that venetoclax activates NK cells, enhancing their cytotoxicity against leukemia through NF-κB-mediated metabolic and cytoskeletal changes42,43. While extensive studies have focused on the biochemical signals that boost NK cell function, the role of LSC mechanical properties in modulating immune responses remains underexplored. Here, we demonstrate that ALDH1A1 inhibition increases F-actin polymerization and cellular stiffness in LSC-enriched FSClow ALDH1A1+ leukemic cells, enhancing their susceptibility to NK cell-mediated cytotoxicity. This alteration in LSC mechanics improves NK cell cytotoxicity and presents a promising strategy for enhancing AML therapy. The combination of ALDH1A1 inhibition with NK cell therapy significantly suppressed leukemia progression in PDX models. We further demonstrate that ALDH1A1 targeting has no detectable toxicity toward HSPCs or NK cells, consistent with previous findings in both murine and human systems48,65. This favorable safety profile strongly underscores the therapeutic promise of ALDH1A1 inhibition in AML. Of note, a recent study demonstrated that increasing cancer cell stiffness through cholesterol depletion enhances CD8+ T-cell cytotoxicity, a process associated with enhanced F-actin dynamics19. Our findings suggest that mechanical modulation of LSCs through ALDH1A1 inhibition could enhance both NK and CD8+ T-cell immunotherapy, broadening the applicability of this approach in targeting CSCs.
In conclusion, our work underscores the importance of mechanical properties in LSC biology and proposes these properties as viable therapeutic targets in AML. By combining the modulation of LSC stiffness with immunotherapy, we propose a potential strategy to overcome treatment resistance and improve patient outcomes. Future research should focus on refining this approach, exploring its application across diverse AML subtypes, and evaluating its potential for integration into existing treatment protocols. Moreover, this strategy may have broader implications in other stem cell-driven malignancies, advancing the field of cancer treatment and tissue regeneration.
Methods
Cell lines
The leukemia cell lines (KG-1a, KG-1a-LUC, and U937) and HEK293T cells were generously provided by Professor Changcheng Zheng (Department of Hematology, The First Affiliated Hospital of University of Science and Technology of China). Cells were maintained in either IMDM (Biosharp, Cat# BL312A) or RPMI 1640 (Biosharp, Cat# BL303A) media, supplemented with 10% FBS (Sigma-Aldrich, Cat# F7524), 100 U/mL Penicillin Streptomycin Amphotericin (Biosharp, Cat# BL142A). All cell lines were cultured in humidified incubator at 37 °C with 5% CO2 and were negative for Mycoplasma contamination. Cell line identity was verified by Short Tandem Repeat (STR) genotyping prior to use in this study.
Animals
Female 6–8-week-old M-NSG mice were obtained from Shanghai Model Organisms Center, Inc., and were used for xenograft transplantation studies involving KG-1a, U937, and primary human AML cells. Female mice were utilized to avoid aggressive interactions inherent to males of this immunodeficient strain. All the mice were housed in a specific pathogen-free animal care center at the University of Science and Technology of China (USTC) with ad libitum access to food and water, a 12/12 h light/dark cycle, and controlled temperature (22–24 °C) and humidity (40–60%). Each experimental group typically comprised a minimum of four mice, with individual data points presented for each sample. Processed data from these experiments are provided in the Source Data file.
Human samples
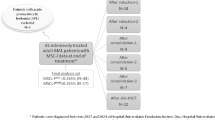
Primary human AML BM cells and umbilical CB samples were obtained from the First Affiliated Hospital of USTC. This study included a total of 81 participants (34 females and 47 males). No financial compensation was provided to the participants. AML patient samples were collected at the time of initial diagnosis, and detailed patient characteristics, including sex, age group, molecular and cytogenetic features as well as their corresponding usage in figures, are provided in Supplementary Data 6. No sub-analyses were performed based on sex or gender. A sex-based analysis was not conducted as the study was not designed or powered for this objective. The study was primarily focused on validating the LSCs mechanobiological properties and their response to immunotherapy and the analysis was focused on this goal.
Wright-Giemsa staining
AML cells were suspended in PBS and smeared onto adhesion microscope slides. Briefly, cells were stained by Wright-Giemsa stain (Biosharp, Cat# BL800A), rinsed in deionized water, and mounted with coverslips for observation under a SLIDEVIEW VS200 Research Slide Scanner microscopy (Olympus). Images were captured using OlyVIA V3.3 software (Olympus).
Cell sorting and flow cytometry analysis
AML bone marrow mononuclear cells (BMMNCs) were isolated by density gradient centrifugation via Ficoll (TBDScience, Cat# LTS1077). For magnetic-activated cell sorting (MACS), primary AML BM Lin- cells were purified from human AML BMMNCs using the Lineage Cell Depletion Kit (Miltenyi Biotec, Cat# 130-092-211), following the manufacturer’s instructions. For sorting human CB CD34+ HSPCs, CD34+ cells were enriched using human CD34 microbead kits (Miltenyi Biotec, Cat# 130-046-702) following manufacturer instructions. For fluorescence-activated cell sorting (FACS), AML Lin- cells were purified using a FACSAria III cytometer (BD Biosciences) or CytoFLEX SRT sorter (Beckman Coulter Life Sciences) based on the human CD45+CD3−CD19- phenotype. All antibodies used throughout this study were commercially sourced from Biolegend and BD and validated by the manufacturers. Anti-human-CD45-FITC (1:100, Biolegend, Cat# 304038, Lot# B380023), Anti-human-CD45-BV510 (1:100, Biolegend, Cat# 304036, Lot# B426008), Anti-human-CD45-APC/Cy7 (1:100, Biolegend, Cat# 304014, Lot# B402225), Anti-human-CD3-PE/Cy7 (1:100, Biolegend, Cat# 300420, Lot# 403211), Anti-human CD3-APC/Cy7 (1:100, Biolegend, Cat# 344818, Lot# B403971), Anti-human CD3-PerCP/Cy5.5 (1:100, Biolegend, Cat# 317336, Lot#B396188); Anti-human CD19-FITC (1:100, Biolegend, Cat# 302206, Lot# B350703) and Anti-human CD19-APC/Cy7 (1:100, Biolegend, Cat# 302218, Lot# B448668), antibodies are used for the sorting and analysis of the leukemic Lin- population. The enriched resting Lin- cells (purity > 95%) were then cultured in StemSpan SFEM (STEMCELL Technologies, Cat# 09600) supplemented with rhFlt3L (100 ng/mL, Novoprotein, Cat# CA82), rhTPO (50 ng/mL, Novoprotein, Cat# CJ95), rhSCF (100 ng/mL, Novoprotein, Cat# C034), and 100× Penicillin Streptomycin Amphotericin (Biosharp, Cat# BL142A).
For sorting small and large cells, AML cell lines (U937, KG-1a-LUC, KG-1a) and human AML BM Lin- cells were separated by FACS on a BD FACSAria III (BD Biosciences) or CytoFLEX SRT sorter (Beckman Coulter Life Sciences) on the basis of forward and side scatter (FSC-A, SSC-A). Small and large cells were gated from the bottom and top 10–15% of the FSC-A and SSC-A distribution, respectively, and then used as inputs for scRNA-seq. Similarly, cells with low and high actin levels (actinlow and actinhigh) were sorted from the bottom and top 10–15% of SIR-Actin (1:1000, Cytoskeleton, Inc., Cat# CY-SC001) distribution (Supplementary Fig. 7).
For analysis and sorting on the basis of ALDH activity (Supplementary Fig. 7), an ALDEFLUOR kit (STEMCELL Technologies, Cat# 01700) was used to identify AML BM Lin- cells, KG-1a cells, and human CB CD34+ HSPCs and CD34- non-HSPCs with high ALDH enzymatic activity. Briefly, cells (1 × 106/well) were incubated with the ALDH substrate BAAA for 30 min at 37 °C, following the manufacturer’s instructions. Cells treated with the ALDH inhibitor diethylaminob enzaldehyde (DEAB) established baseline fluorescence to define the cut-off for ALDEFLUOR-positive cells. Small (FSClow, left 10–15%) and large (FSChigh, right 10-15%) cells were then sorted by FSC-A and SSC-A, with FSClow ALDH1A1+ cells further verified. For flow cytometry analysis of ALDH1A1 expression, human AML BM Lin- cells were stained with an anti-ALDH1A1 antibody (1:100, Cell Signaling Technology, Cat# 36671, Lot# 1), followed by an Alexa-Fluor-488-labeled secondary antibody (1:400, Biolegend, Cat# 406416, Lot# B323038).
For general flow cytometry analysis, AML BMMNCs were stained with hCD45, hCD3, and hCD19 to identify BM Lin- cells. For the cell cycle analysis, cells were incubated with Hoechst 33342 (10 µg/ml, Beyotime, Cat# C1022) for 15–20 min at 37 °C, followed by Pyronin Y (1 µg/ml, Polysciences, Inc., Cat# 18614-5) staining for 20 min. For cell differentiation analysis, the purified cells were stained with CD11b-FITC or PE (1:200, Biolegend, Cat# 101206 or 301306, Lot# B260639 or B434405) and CD14-PE (1:200, Biolegend, Cat# 325606, Lot# B368423). All flow cytometry data were analyzed using FlowJo software (10.8.1).
Colony formation assay
To evaluate the clonogenic potential of AML cells, U937, KG-1a, primary human AML BM Lin- cells, human CB CD34+ HSPCs and CD34⁻ non-HSPCs by MACs using human CD34 microbead kits (Miltenyi Biotec, Cat# 130-046-702) were resuspended in MethoCultTM H4434 medium (STEMCELL Technologies, Cat# H4434), and assessed in colony-forming unit (CFU) methylcellulose assays according to the manufacturer’s instructions. For assessing the effect of NK cells on clonogenic growth, AML cells (500 cells) were co-cultured with CB-NK cells at a ratio of 5:1 for 6 h. Subsequently, the cells were resuspended in MethoCultTM H4434 medium and scored in CFU methylcellulose assays. Colony formation, defined as a cluster of ≥ 50 leukemia cells, was quantified using a Thunder Leica DM6B microscope (Leica).
Xenotransplantation of human AML
For the AML xenograft models, small KG-1a-Luc cells (3 × 105 cells/mouse), ALDH1A1-knockdown FSClow ALDH1A1+ KG-1a cells (1 × 105 cells/mouse), U937 cells (2.5 × 104 cells/mouse), primary BM Lin- cells from AML patients (5 × 106 cells/mouse for SSLCs, 7×106 cells/mouse for small AML cells), small or large primary BM Lin- cells from AML patients treated with Jas or Lat A (4 × 105 cells/mouse for small, small+Jas, large, large+Lat A group, respectively), and primary BMMNCs from AML patients (7 × 106 cells/mouse) were transferred to female 6-8-week-old M-NSG mice. Eight hours before the leukemia cells were injected via the tail vein to establish a xenograft model, the mice were sublethally irradiated (250 cGy). Leukemia progression was monitored through bioluminescence imaging or flow cytometry at various timepoints. Leukemic burden in flow cytometry analyses was determined by the presence of human CD45+ (1:100, Biolegend, Cat# 304038, Lot# B380023) cells.
Bioluminescence imaging
Mice engrafted with KG-1a-LUC cells were injected intraperitoneally with D-luciferin substrate (15 mg/ml, YEASEN, Cat# 40902ES03) and imaged using an IVIS Spectrum Imaging System (PerkinElmer). The bioluminescent signals were analyzed by Living Image Software v 4.7.3 (PerkinElmer Inc.) to enable consistent comparison of images taken at the same exposure and time points after substrate administration. Signal intensity was quantified as radiance (photons/sec/cm2/steradian, p/s/cm2/sr) with pseudocolor images illustrating intensity levels. To normalize for background luminescence, an additional ROI was applied to each image during analysis.
Limiting dilution assay
For in vivo limiting dilution assays (LDAs), small and large primary BM Lin- cells from AML patients were sorted by FACS and injected into the tail vein of M-NSG mice at the specified cell doses (10000 and 100000 cells/mouse). Eight hours before engraftment, the mice were sublethally irradiated (250 cGy). Two to three months later, leukemia burden was evaluated by FACS analysis of human CD45+ cells in the BM.
Immunofluorescence and image acquisition
Cells were fixed with 4% paraformaldehyde for 15 min at room temperature, followed by washing with PBS. To block nonspecific binding, cells were incubated with QuickBlock™ Blocking Buffer (Beyotime, Cat# P0260) at 4 °C for 1–2 h.
For F-actin staining, cells were incubated with Actin-Tracker Red-555 (1:100, Beyotime, Cat# C2203S) or Actin-Tracker Green-488 (1:100, Beyotime, Cat# C2201S) for 1 h at 4 °C. For ALDH1A1 staining, cells were incubated overnight with an anti-ALDH1A1 antibody (1:100, Cell Signaling Technology, Cat# 36671, Lot# 1) followed by an incubation with an Alexa-Fluor-488-labeled secondary antibody (1:400, Biolegend, Cat# 406416, Lot# B323038). The cells were then washed three times with PBS, and nuclear staining was conducted using DAPI (1 mg/mL, Biosharp, Cat# BL105A) for 5 min. Images were acquired using a confocal laser scanning microscope (Leica), and F-actin intensity was quantified using ImageJ software (National Institutes of Health).
Measurement of cell cortical stiffness by AFM
The stiffness of human AML cells was measured by AFM (Bruker Resolve 8). After undergoing plasma treatment for 15 min, the glass-bottom Petri dishes were then coated with laminin (2 μg/ml, Nuwacell, Cat# RP01025) for 2–4 h. AML cell samples (1 × 105 cells in 500 μl culture medium) were stained with Hoechst 33342 (10 µg/ml, Beyotime, Cat# C1022) for 30 min and then cultured in the prepared dishes for 8 h at 37 °C. The sample stage of the AFM was replaced with the motorized displacement stage of the confocal microscope, allowing for high-magnification observation. Microspheres with a radius of 10 µm were modified on the tip of AFM probe (NPO-10) to facilitate indentation measurements. The cantilever spring constant was 0.06 N/m. To align the AFM probe accurately with the nuclei labeled with Hoechst, the center of the microsphere at the tip of the AFM cantilever was visualized via confocal microscopy. Cell stiffness was assessed by analyzing force curves and calculating the Young’s modulus. This was done by fitting the approach curve to an indentation depth of less than 500 nm using NanoScope Analysis software (Bruker). Briefly, force-indentation curves were analyzed using NanoScope Analysis 1.9 software and fitted with the Hertz contact model for a spherical indenter:
where E is the Young’s modulus, ν the Poisson’s ratio (assumed to be 0.5 based on prior studies66, R the radius of the cantilever tip, δ the cell deformation distance, and F the applied force. The maximal force applied to each cell is 0.7 nN. Cell stiffness was quantified by the average Young’s modulus.
Imaging flow cytometry
Leukemia cells were stained with Hoechst 33342 (10 µg/ml, Beyotime, Cat# C1022) and SiR-Actin (1:1000, Cytoskeleton, Inc., Cat# CY-SC001) at 37 °C for 30 min. Following staining, the cells were washed. Imaging was performed at 60x magnification under low-speed settings using an Amnis ImageStreamX Mark II Imaging Flow Cytometer (Merck Millipore). A minimum of 50 cells were captured per sample. Data analysis and cell labeling were conducted with IDEAS software (EMD Millipore).
Bulk RNA-seq and data analysis
Small and large primary BM Lin- cells were isolated by FACS and treated with 1 μM Jas or 1 μM Lat A for 6 h. Following treatment, total RNA was extracted using TRIzol (Invitrogen, Cat# 15596018), and libraries were generated using the VAHTS Universal V8 RNA-seq Library Prep Kit. Sequencing was performed in single-read mode on an Illumina Novaseq X Plus platform.
Raw RNA-seq reads were mapped to the human GRCh38 assembly reference genome using the STAR alignment tool (v2.7.11b). Heatmaps were produced using unsupervised clustering, and differential gene expression analysis was conducted using DESeq2 (v1.34.0), with a fold-change threshold of 1.5 and a p-value < 0.05 considered statistically significant. GSEA was conducted using the clusterProfiler package (v4.2.2), with gene sets obtained from the MSigDB collection (v7.5.1). The list of the DEGs and enriched signaling pathways is provided in Supplementary Data 7.
Microfluidic chip preparation and sorting
The design was inspired by and modified from prior work58, with substantial optimization for improved resolution in separating small, deformable cells from larger, stiffer ones. Specifically, our microfluidic device comprises two core components: a microweir structure and a collection chamber. The microweir imposes size- and deformability-dependent thresholds, allowing softer and smaller cells to pass through under constant hydraulic pressure, while larger or stiffer cells are delayed or blocked. The channel chamber with a 40 μm ceiling, designed to accommodate retained cells and prevent clogging. Custom thresholds were set (12 μm for U937 cells, 5 μm for primary AML cells) based on the size range of the populations of interest. For microfluidic sorting, soft-small and stiff-large BM Lin- (CD45+CD3-CD19-) cells were sorted and functionally assessed on a per-sample basis. The detailed protocol comprised the following sequential steps:
The 4-inch monocrystalline wafers used to invert the microfluidic piping were first ultrasonically cleaned to remove surface impurity particles. Dewatering baking was performed to eliminate surface moisture and enhance photoresist adhesion. Since the photoresist contains solvent molecules, the coated wafers were further dried at about 80 °C. Microchem SU-8 negative photoresist was then applied to create a mask based on the pipeline drawing designed by Klayout software, and thus, the pipeline structure on the substrate can be obtained through a single exposure. For further etching, a dry etch process using plasma was employed to remove unprotected areas of the wafer surface. Finally, to prevent the inverted mold material from adhering to the wafer surface during the curing process, a layer of Au or Cr nanoparticles was coated on the finished wafer surface, forming an anti-adhesive layer.
Microfluidic chips are prepared using silicon wafers processed by PDMS (Polydimethylsiloxane) inversion lithography. The device was connected to a pressure pump via 15 ml centrifuge tubes for sample introduction. Before sample loading, the chip was sequentially washed with 75% ethanol and 1 × PBS. Cell suspensions (density ranging from 0.5 × 104 /ml) were connected to the inlet of the chip, driven by a pump operating at pressures ranging from 100 to 400 mbar. After the cells passed through the chips of 12 μm height (for U937 cells) or 5 μm height (for human primary AML samples), SSLCs were collected from the chip outlets. The residual cells that did not pass through the chip (with a height of 12 or 5 μm) and SSLCs were collected from the inlet with a reverse flow of fluids.
To drive microfluidic flow, a positive-pressure air compressor (TW3801-4C, Shanghai Tuowen Machinery Co., Ltd.) and a vacuum pump (VP18 PLUS, Beijing Laibotaike Instrument Co., Ltd.) were used. Pressure and vacuum were precisely regulated using an OB1 mk3+ multi-channel microfluidic controller (Elveflow, France), which enabled stable and tunable control of flow rates within the chip. The air compressor delivered a maximum flow rate of 20 L/min at 0.7 MPa, while the vacuum pump provided an ultimate vacuum of 20 mbar at a pumping rate of 18 L/min. A schematic of the fluidic setup and tubing configuration is shown in Supplementary Fig. 3a-b.
ScRNA-seq data analysis
For scRNA-Seq library construction, small primary AML BM Lin- cells from four individuals were sorted by a BD FACSAriaTM III sorter (BD Biosciences, United States). Then, scRNA-Seq was performed on both large and small cell populations. According to the instructions for the single-cell 3′ solution v3 reagent kit (10X Genomics), pooled cells were counted and resuspended at a concentration of 1000 cells/μl, with an estimated yield of 6000 cells per library. Single-cell libraries were constructed following the manufacturer’s protocols and sequenced on the Illumina NovaSeq 6000 system.
To acquire the sequencing data, the FASTQ files of AML samples were aligned at the hg19 reference genome, with UMI counts quantified using the default parameters of 10X Genomics Cell Ranger pipeline (v.3.1.0)67. Data analysis was conducted in R (v4.1.2) using the Seurat package (v4.1.2)68. Quality control (QC) was performed to remove cells expressing fewer than 500 genes or more than 4000 genes or having over 10% mitochondrial gene content. Additionally, genes detected in fewer than five cells were excluded. Seurat objects for the large and small cell groups were created, and data integration was conducted using the “IntegrateData” function with the top 15 canonical correlation analysis (CCA) dimensions for anchor weighting. To identify cell clusters, we performed PCA on the integrated object, setting the resolution to Z and selecting the top Y dimensions for PCA. The top 15 principal components were then used for uniform manifold approximation and projection (UMAP) analysis. Six cell subpopulations were identified, and the top five genes with cluster-specific expression were visualized using Seurat’s “FindAllMarkers” function.
To confirm the signature of the LSC cluster, the LSC score, cell cycle score and F-actin score were determined via the “AddModuleScore” function. Gene sets for these analyses were sourced from published signatures and the MsigDB website (https://www.gsea-msigdb.org/gsea/msigdb /human/geneset), with details provided in Supplementary Data 2. For pseudotime analysis, the Monocle R package (v2.18.0) was used on selected cell subsets (HSPC, LSC, EPC, GMP, DC-like and Blast). The trajectories were mapped using the DDRTree dimensionality reduction algorithm, with cell ordering established by the orderCells function. Gene Ontology (GO) analysis was conducted using Metascape (http://metascape.org/gp/index.html).
Public RNA-seq and survival analysis
To verify the expression of the top markers (ALDH1A1, SPINK2, and LGALS3BP), we analyzed their expression in LSCs and non-LSCs of AML patients from public RNA-seq dataset. This bulk RNA-seq dataset, generated by ref.69, is archived at NCBI GEO with accession code GSE199452. To confirm whether the top markers (ALDH1A1, SPINK2, and LGALS3BP) were associated with clinical survival, we conducted survival analysis of AML patients. We downloaded processed RPKM expression levels of 179 samples from theTCGA-LAML study70 (https://tcga-data.nci.nih.gov/docs/publications/laml_2012) and survival data from the cBioPortal (http://www.cbioportal.org/). In building Kaplan-Meier curves, the AML patients were split into high and low expression groups according to the optimal cut-off point calculated by the surv_cut point function in the R package survminer v. 0.4.9 (https://github.com/kassambara/survminer) with minprop = 0.3. Finally, survival analysis was performed with the survfit function in the R package survival v.3.4.0 (https://github.com/therneau/survival).
Serial transplantation assays
For primary transplantation, sub-lethally irradiated M-NSG mice were intravenously injected via the tail vein with 1 × 105 FACS-sorted FSClow, ALDH1A1+, or FSClow ALDH1A1+ subpopulations from AML patient-derived Lin- cells, suspended in 0.2 mL PBS. Mice were sacrificed 5–6 weeks post-transplantation, and BM mononuclear cells were collected from one femur per mouse. Human CD45+ cell engraftment was assessed by flow cytometry. For secondary transplantation, 1 × 10⁶ BM cells from primary recipients were injected into sub-lethally irradiated secondary M-NSG mice via tail vein. Mice were euthanized 4 weeks later, and BM engraftment was evaluated as above. For tertiary transplantation, 1×10⁶ BM cells from secondary recipients were injected into tertiary M-NSG recipients. Human CD45⁺ cell chimerism in the BM was analyzed by flow cytometry 3 weeks post-transplantation.
Knockdown assay in KG-1a cells
The RNAi target sequences and knockdown plasmids containing shALDH1A1 were designed and generated in our laboratory. All plasmids were constructed using restriction enzyme digestion and ligation. The shRNA sequences targeting human ALDH1A1 were cloned into the pLKO.1 vector (Addgene). All constructs were verified by Sanger sequencing. The primers used were: ALDH1A1 (shRNA-1), 5’-ccgggctgatttaatc gaaagagatctcgagatctctttcgattaaatcagctttttg-3’ (forward), 5’-aattcaaaaagctgatttaatcgaaagagatctc gagatctctttcgattaaatcagc-3’ (reverse); and ALDH1A1 (shRNA-2), 5’-ccggcaccgatttgaagattcaatac tcgagtattgaatcttcaaatcggtgtttttg-3’ (forward), 5’-aattcaaaaacaccgatttgaagattcaatactcgagtattgaat cttcaaatcggtg-3’ (reverse). Lentiviruses were produced in HEK293T cells transfected using Polyethylenimine (PEI, Polysciences, Cat# 23966) with pMD2.G and pSPAX2 packaging constructs. Viral supernatants were collected at 48 h and 72 h after transfection, filtered, concentrated and stored at −80 °C. KG-1a cells were infected with the virus and polybrene (8 µg/ml, Sigma-Aldrich, Cat# H9268). ShRNA-transduced cells were sorted by FACS on day 4 after infection.
Real-time quantitative PCR
Total RNA was isolated using TRIzol Reagent (Invitrogen, Cat# 15596018), and then reverse transcribed using the HiScript III RT SuperMix (Vazyme, Cat# R323-01) for RT-PCR. The reverse transcription products were subjected to real-time quantitative PCR with ChamQ Universal SYBR qPCR Master Mix (Vazyme, Cat# Q711-02). All primers were designed and synthesized by Sangon Biotech (Shanghai). GAPDH mRNA levels served as an internal normalization control. The primers used were: ALDH1A1, 5’-acgccagacttacctgtcctactc-3’ (forward), 5’-tcttgccactcactgaatcatgcc-3’ (reverse); and GAPDH, 5’-ccagcaagagcacaagaggaa-3’ (forward), 5’-caaggggtctacatggcaact −3’ (reverse).
Immunoblotting
Cells were lysed in cell lysis buffer containing 50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 1 mM EDTA, and 1% Nonidet P-40 (Beyotime, Cat# P0013F) at 4 °C for 40 min. For immunoblotting, the protein samples were separated on 4–12% SDS-PAGE gels (Epizyme, Cat# LK307) and then transferred onto PVDF membranes. After blocking for 30 min at room temperature, the PVDF membranes were incubated with primary antibodies overnight at 4 °C. Following washes, the membranes were incubated with secondary antibodies at room temperature for 2 h and then subjected to imaging. The targeted proteins were visualized using the enhanced chemiluminescence (ECL) method, and the immunoblots were detected by exposure to an SH-Compact 523 imaging system (SHSTBio).
Human NK cell isolation and culture
CB-NK cells were purified from CB mononuclear cells (CBMCs) with the NK Cell Isolation Kit (Miltenyi Biotec, Cat # 130-092-657) following the manufacturer’s instructions. Each purification typically yielded between 5 × 106 and 1 × 107 NK cells, with a purity of over 95%. The enriched resting NK cells were then cultured in IMDM supplemented with 10% FBS (Sigma-Aldrich, Cat# F7524) and 100× Penicillin Streptomycin Amphotericin (Biosharp, Cat# BL142A). NK cell activation was induced with rhIL-12 (10 ng/mL, PeproTech, Cat # 210-12) and rhIL-15 (50 ng/mL, PeproTech, Cat # 210-15).
Lytic granule polarization assay
FSClow ALDH1A1+ KG-1a cells were isolated by FACS sorting (Supplementary Fig. 7f), pretreated with or without 1 μM Disulfiram for 48 h, and then co-cultured with CB-NK cells (1.5 × 105 cells per well) from six individual donors at an effector-to-target ratio of 1.5:1 for different durations (30 min, 1 h, 2 h and 4 h). The cell mixtures were seeded on poly-L-lysine-coated coverslips (Sigma-Aldrich, Cat# P4707). Following co-culture, the cells were fixed with 4% paraformaldehyde for 15 min and blocked with QuickBlock Blocking Buffer (Beyotime, Cat# P0260) for 2 h.
The cells were then incubated in blocking buffer containing anti-perforin-FITC antibody (1:100, Biolegend, Cat# 353310, Lot# B405039) at 4 °C overnight. Subsequently, the cells were incubated with Alexa Fluor-488-conjugated anti-mouse antibody (1:400, Biolegend, Cat# 4408, Lot# 24) for 30 min, and with Actin-Tracker Red-555 (1:150, Beyotime, Cat# C2203S) for 2 h at room temperature. After washing, the cells were mounted on slides and imaged by a confocal laser-scanning microscope with a 63× objective lens (Leica). Z-stacks were acquired at 0.5 μm increments (covering a 10 μm total range) for both the Alexa Fluor 555 and the Alexa Fluor 488 channels for each NK cell-target cell conjugate. The intensities of Alexa 555 and Alexa 488 for each NK cell-target cell conjugate were quantified for perforin via projected images exported from the z stack data.
The perforin distribution was quantified via projected images from the z-stack data. Based on the NK cell area opposite the immune synapse, distances were segmented into five polarization indices (0.2, 0.4, 0.6, 0.8 and 1.0), representing varying distances from the synapse. The polarization index was used to quantify the average perforin ratio within each NK cell relative to the immune synapse across these indices.
NK cell functional assays
For NK cell cytotoxicity assay, FSClow ALDH1A1+ KG-1a cells were pretreated with or without 1 μM Disulfiram for 48 h, followed by staining with the CellTrace Violet Cell Proliferation Kit (0.5 μM, Thermo Fisher Scientific, Cat# C34557) for 15–20 min at 37 °C. After washing, these FSClow ALDH1A1+ KG-1a cells (2 × 104) were co-cultured with CB-derived NK cells (from six individual donors) at an E: T ratio of 5:1 for 6 h in 96-well plates at 37 °C. Following co-culture, the cells were stained with Annexin V/PI (Elabscience, Cat# E-CK-A217) and analyzed by flow cytometry. Specific killing (%) was calculated using the following formula: % specific killing = (% Annexin-V+ with NK cells - % Annexin-V+ without NK cells).
For NK cell degranulation assay, CB-NK cells from three individual donors were first stained with surface markers Anti-human CD3-APC/Cy7 (1:100, Biolegend, Cat# 344818, Lot# B403971) and Anti-human CD56-BV421 (1:100, BD, Cat# 562751, Lot# 4333719) (CD3-CD56+) for 30–60 min at 4 °C, then co-cultured with target cells at a 5:1 ratio for 6 h. Anti-CD107a-PE or FITC antibody (1:100, Biolegend, Cat# 328608 or 328606, Lot# B376271 or B278334) was added at the beginning of co-culture to monitor degranulation.
For intracellular cytokine and cytolytic granule staining, IL-12 and IL-15 pre-activated NK cells were surface-stained and then fixed and permeabilized for intracellular staining with antibodies against IFN-γ (1:100, Biolegend, Cat# 506528, Lot# B373056), TNF-α (1:100, Biolegend, Cat# 502912, Lot# B419917), granzyme B (1:100, Biolegend, Cat# 372222, Lot# B402218), and perforin (1:100, Biolegend, Cat# 353310, Lot# B405039). All samples were analyzed using a Beckman Coulter CytoFLEX flow cytometer, and data were processed with FlowJo software (v10.8.1).
For NK cell proliferation assay, CB-NK cells treated with or without 1 µM Disulfiram were seeded (2 × 10⁴ cells/well) in 96-well plates. At days 0, 1, 2, and 3, proliferation was assessed using the Cell Counting Kit-8 (CCK-8, MCE, Cat# HY-K0301). Fresh IMDM medium (100 μL; 10% FBS, 10 ng/mL IL-12, and 10 ng/mL IL-15) supplemented with 10 μL CCK-8 reagent was added to each well and incubated for 2 h at 37 °C in the dark. Absorbance was measured at 450 nm.
For NK cell viability assay, resting or IL-12 (10 ng/mL) + IL-15 (50 ng/mL)-activated CB-NK cells were treated with increasing concentrations of Disulfiram (0, 500, 1000, 2000 nM) for 24 h, 48 h, or 72 h. Cell viability was assessed by Annexin V/PI staining followed by flow cytometry. In parallel, Disulfiram-treated CB-NK cells from five donors were co-cultured with KG-1a cells (2 × 10⁴ cells/well) at a 5:1 E:T ratio for 6 h at 37 °C, and cytotoxicity was evaluated by flow cytometry.
Live-cell imaging
For live-cell imaging, 5 × 105 FSClow ALDH1A1+ KG-1a cells pretreated with Disulfiram (1 μM) for 48 h were seeded onto dishes precoated with poly-lysine (1 mg/ml, Beyotime, Cat# ST508). Cells were allowed to adhere to the poly-L-lysine substrate for 3–5 h at 37 °C. Meanwhile, 5 × 105 CB-NK cells were stained with Hoechst 33342 (10 µg/ml, Beyotime, Cat# C1022) for 30 min at 37 °C, followed by washing and addition to the adherent KG-1a cells. Co-cultures were placed in a live-cell imaging chamber maintained at 37 °C and 5% CO2. Time-lapse imaging and three-dimensional scanning were performed using a Leica confocal laser scanning microscope (TCS SP8) to monitor NK cell cytotoxicity behavior. The timing of cytolytic contact between NK and KG-1a cells was manually identified based on bright-field time-lapse recordings using LAS X software. The percentage of target cell killing by same-donor CB-NK cells against control and Disulfiram-treated KG-1a cells were quantified under identical effector-to-target (E:T) ratios.
Treatment regimens for AML xenografts
For the AML PDX model, primary AML BMMNCs (7 × 106 cells/mouse) from four AML patient samples were transplanted into sublethally irradiated (250 cGy) M-NSG mice via tail vein injection. Successful engraftment was observed 6–10 weeks post-inoculation, with ~0.1–10% human CD45+ cells detected in the peripheral blood. The mice were then randomly divided into four groups and assigned the following treatments: untreated control (PBS), Disulfiram (15 mg/kg, administered intraperitoneally, daily for 5 days), NK cells (5 × 106 NK cells /mouse), and a combination of Disulfiram and NK cells.
In a separate experiment to investigate leukemia cell stiffening with Disulfiram, engrafted mice received Disulfiram from day 0 to day 5 post-leukemia establishment. Disulfiram was administered intraperitoneally for 5 consecutive days in both the Disulfiram and combination groups. On day 6, activated NK cells (5 × 106 NK cells/mouse) from six individual CB donors were infused intravenously via the tail vein in the NK and combination groups. To support NK cell activity, rhIL-15 (1 μg/mouse, PeproTech, Cat# 210-15) was administered intraperitoneally every 3 days following NK cell infusion until the study endpoint. At the end of treatment period, the mice were sacrificed and the BM were collected for further analysis. Untreated mice served as controls, and their leukemia cells were collected on the same day as those from the treatment groups.
Statistics and reproducibility
The sample size was determined based on resource availability and experimental feasibility, and were sufficient to demonstrate statistically significant differences between experimental groups. No statistical methods were used to predetermine sample sizes. All data meeting quality control criteria were included in the analyses, and no data were excluded. All mice were randomly assigned to experimental groups. The investigators were not blinded to allocation during experiments and outcome assessment. GraphPad Prism 8 (GraphPad Software) was used for statistical analysis. For each dataset, data were presented as mean ± standard error of the mean (SEM) unless otherwise indicated in the figure legends. The statistical tests used for each figure are detailed in the corresponding figure legends. When two groups were compared, Student’s t-tests were used. When multiple groups were compared, one-way ANOVA was used. Correlation analyses were conducted using Pearson’s test, and survival analyses were performed with the log-rank test. A p-value of <0.05 was considered statistically significant. All FACS data and microscopy images are representative of at least three independent biological experiments. Numbers of biologically independent experiments are indicated in the figure legends.
Ethics statement
All animal experiments were conducted following the guidelines of the USTC Laboratory Animal Care and Use Committee (USTCACUC22240122004 and USTCACUC25120124093). Animals that met predefined humane endpoints—including weight loss (>20% post-irradiation or >15% post-treatment), or persistent signs of distress such as lethargy, piloerection, or hunched posture for over 6 h—underwent mandated euthanasia. No animals in this study exceeded these ethical limits, ensuring compliance with welfare standards. All experimental protocols involving human samples were reviewed and approved by the Medical Ethics Committees of the First Affiliated Hospital of USTC (2021 KY no.150 and 2022 KY no.022), and written informed consent was obtained from all participants prior to sample collection.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequencing data generated in this study, including both scRNA seq and bulk RNA-seq data, have been deposited in the Genome Sequence Archive for Human (GSA-Human) at the National Genomics Data Center71, China National Center for Bioinformation / Beijing Institute of Genomics, Chinese Academy of Sciences, under accession numbers HRA010191 and HRA011960, respectively. These data are active and available under controlled access due to the sensitive nature of human genomic data and to protect participant privacy in accordance with relevant laws. Access can be obtained by submitting a request to the corresponding author. The processed expression data generated in this study are provided in the Supplementary Information. The scRNA-seq raw data and the processed data have also been deposited in the Gene Expression Omnibus (GEO) under accession number GSE267998 and are publicly accessible. Source data are provided with this paper.
Code availability
The tools supporting this paper are available on GitHub, including Seurat (v4.1.2), Cell Ranger (v3.1.0), R (v4.1.2), Survival (v.3.4.0), Survminer (v. 0.4.9). The original code is publicly available as of the date of publication through https://github.com/XiXi1994bb/LSC72 (https://doi.org/10.5281/zenodo.17250099).
References
DiNardo, C. D., Erba, H. P., Freeman, S. D. & Wei, A. H. Acute myeloid leukaemia. Lancet 401, 2073–2086 (2023).
Subedi, A. et al. Nicotinamide phosphoribosyltransferase inhibitors selectively induce apoptosis of AML stem cells by disrupting lipid homeostasis. Cell Stem Cell 28, 1851–1867.e1858 (2021).
Shlush, L. I. et al. Tracing the origins of relapse in acute myeloid leukaemia to stem cells. Nature 547, 104–108 (2017).
Haubner, S. et al. Cooperative CAR targeting to selectively eliminate AML and minimize escape. Cancer Cell 41, 1871–1891.e1876 (2023).
Hou, S., Liu, J. & Zhu, Y. Multi-omics advances for molecular characterization, precision medicine, and prognostic implications in leukemia. Cell Investig. 1, 100007 (2025).
Su, R. et al. Targeting FTO suppresses cancer stem cell maintenance and immune evasion. Cancer Cell 38, 79–96.e11 (2020).
Saw, P. E., Liu, Q., Wong, P. P. & Song, E. Cancer stem cell mimicry for immune evasion and therapeutic resistance. Cell Stem Cell 31, 1101–1112 (2024).
Li, Z. et al. ILC1s control leukemia stem cell fate and limit development of AML. Nat. Immunol. 23, 718–730 (2022).
Park, S. M. et al. Dual IKZF2 and CK1α degrader targets acute myeloid leukemia cells. Cancer Cell 41, 726–739.e711 (2023).
Leick, M. B. et al. Non-cleavable hinge enhances avidity and expansion of CAR-T cells for acute myeloid leukemia. Cancer Cell 40, 494–508.e495 (2022).
Wang, J. et al. Leukemogenic chromatin alterations promote AML leukemia stem cells via a KDM4C-ALKBH5-AXL signaling axis. Cell Stem Cell 27, 81–97.e88 (2020).
Nicosia, L. et al. Therapeutic targeting of EP300/CBP by bromodomain inhibition in hematologic malignancies. Cancer Cell 41, 2136–2153.e2113 (2023).
Li, Y. et al. TET2-mediated mRNA demethylation regulates leukemia stem cell homing and self-renewal. Cell Stem Cell 30, 1072–1090.e1010 (2023).
Liang, W. W. et al. Integrative multi-omic cancer profiling reveals DNA methylation patterns associated with therapeutic vulnerability and cell-of-origin. Cancer Cell 41, 1567–1585.e1567 (2023).
Jones, C. L. et al. Inhibition of amino acid metabolism selectively targets human leukemia stem cells. Cancer Cell 34, 724–740.e724 (2018).
Uroz, M. et al. Differential stiffness between brain vasculature and parenchyma promotes metastatic infiltration through vessel co-option. Nat. Cell Biol. 26, 2144–2153 (2024).
Chaudhuri, O. et al. Extracellular matrix stiffness and composition jointly regulate the induction of malignant phenotypes in mammary epithelium. Nat. Mater. 13, 970–978 (2014).
Northey, J. J., Przybyla, L. & Weaver, V. M. Tissue force programs cell fate and tumor aggression. Cancer Discov. 7, 1224–1237 (2017).
Lei, K. et al. Cancer-cell stiffening via cholesterol depletion enhances adoptive T-cell immunotherapy. Nat. Biomed. Eng. 5, 1411–1425 (2021).
Vining, K. H. & Mooney, D. J. Mechanical forces direct stem cell behaviour in development and regeneration. Nat. Rev. Mol. Cell Biol. 18, 728–742 (2017).
Yang, C., Tibbitt, M. W., Basta, L. & Anseth, K. S. Mechanical memory and dosing influence stem cell fate. Nat. Mater. 13, 645–652 (2014).
Ni, F. et al. Ptpn21 controls hematopoietic stem cell homeostasis and biomechanics. Cell Stem Cell 24, 608–620.e606 (2019).
Wu, M. et al. Enhanced engraftment of human haematopoietic stem cells via mechanical remodelling mediated by the corticotropin-releasing hormone. Nat. Biomed. Eng. 9, 754–771 (2025).
Daeden, A. et al. Polarized branched Actin modulates cortical mechanics to produce unequal-size daughters during asymmetric division. Nat. Cell Biol. 25, 235–245 (2023).
Cao, M. et al. An actin filament branching surveillance system regulates cell cycle progression, cytokinesis and primary ciliogenesis. Nat. Commun. 14, 1687 (2023).
Pernier, J. et al. Myosin 1b is an actin depolymerase. Nat. Commun. 10, 5200 (2019).
Gardel, M. L., Schneider, I. C., Aratyn-Schaus, Y. & Waterman, C. M. Mechanical integration of actin and adhesion dynamics in cell migration. Annu. Rev. Cell Dev. Biol. 26, 315–333 (2010).
Goardon, N. et al. Coexistence of LMPP-like and GMP-like leukemia stem cells in acute myeloid leukemia. Cancer Cell 19, 138–152 (2011).
Ng, S. W. et al. A 17-gene stemness score for rapid determination of risk in acute leukaemia. Nature 540, 433–437 (2016).
Elsayed, A. H. et al. A six-gene leukemic stem cell score identifies high risk pediatric acute myeloid leukemia. Leukemia 34, 735–745 (2020).
Zhu, R. et al. Inhibition of NOTCH4 sensitizes FLT3/ITD acute myeloid leukemia cells to FLT3 tyrosine kinase inhibition. Leukemia 38, 1581–1591 (2024).
Zeng, Z. et al. Regulation and signaling pathways in cancer stem cells: implications for targeted therapy for cancer. Mol. Cancer 22, 172 (2023).
Nusse, R. & Clevers, H. Wnt/β-catenin signaling, disease, and emerging therapeutic modalities. Cell 169, 985–999 (2017).
Pei, S. et al. A novel type of monocytic leukemia stem cell revealed by the clinical use of venetoclax-based therapy. Cancer Discov. 13, 2032–2049 (2023).
Zhu, J. et al. Custom microfluidic chip design enables cost-effective three-dimensional spatiotemporal transcriptomics with a wide field of view. Nat. Genet. 56, 2259–2270 (2024).
Eppert, K. et al. Stem cell gene expression programs influence clinical outcome in human leukemia. Nat. Med. 17, 1086–1093 (2011).
Zhao, Z. et al. Activity-based sensing reveals elevated labile copper promotes liver aging via hepatic ALDH1A1 depletion. Nat. Commun. 16, 1794 (2025).
Calvanese, V. et al. Mapping human haematopoietic stem cells from haemogenic endothelium to birth. Nature 604, 534–540 (2022).
Piao, S. et al. ALDH1A1 and HLTF modulate the activity of lysosomal autophagy inhibitors in cancer cells. Autophagy 13, 2056–2071 (2017).
Storms, R. W. et al. Distinct hematopoietic progenitor compartments are delineated by the expression of aldehyde dehydrogenase and CD34. Blood 106, 95–102 (2005).
Cheung, A. M. et al. Aldehyde dehydrogenase activity in leukemic blasts defines a subgroup of acute myeloid leukemia with adverse prognosis and superior NOD/SCID engrafting potential. Leukemia 21, 1423–1430 (2007).
Wang, Y. et al. Venetoclax acts as an immunometabolic modulator to potentiate adoptive NK cell immunotherapy against leukemia. Cell Rep. Med. 5, 101580 (2024).
Wang, Y. et al. Protocol to measure cell avidity between cord blood-derived NK cells and leukemia cell line KG-1a. STAR Protoc. 5, 103387 (2024).
Zheng, X. et al. Tumors evade immune cytotoxicity by altering the surface topology of NK cells. Nat. Immunol. 24, 802–813 (2023).
Mukherjee, M., Mace, E. M., Carisey, A. F., Ahmed, N. & Orange, J. S. Quantitative imaging approaches to study the CAR immunological synapse. Mol. Ther. 25, 1757–1768 (2017).
Phatarpekar, P. V. et al. The septin cytoskeleton regulates natural killer cell lytic granule release. J. Cell Biol. 219, e202002145 (2020).
Lagrue, K., Carisey, A., Morgan, D. J., Chopra, R. & Davis, D. M. Lenalidomide augments actin remodeling and lowers NK-cell activation thresholds. Blood 126, 50–60 (2015).
Levi, B. P., Yilmaz, O. H., Duester, G. & Morrison, S. J. Aldehyde dehydrogenase 1a1 is dispensable for stem cell function in the mouse hematopoietic and nervous systems. Blood 113, 1670–1680 (2009).
Majeti, R. et al. CD47 is an adverse prognostic factor and therapeutic antibody target on human acute myeloid leukemia stem cells. Cell 138, 286–299 (2009).
Bai, L. et al. A POtent and Selective Small-molecule Degrader of STAT3 achieves complete tumor regression in vivo. Cancer Cell 36, 498–511.e417 (2019).
Ding, L. et al. Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing. Nature 481, 506–510 (2012).
Chen, S. et al. Arsenic trioxide rescues structural p53 mutations through a cryptic allosteric site. Cancer Cell 39, 225–239.e228 (2021).
Cheng, Y. et al. Decoding m(6)A RNA methylome identifies PRMT6-regulated lipid transport promoting AML stem cell maintenance. Cell Stem Cell 30, 69–85.e67 (2023).
Qiu, K. et al. Targeting USP10–FAK pathway sensitizes BCR-ABL+ leukemia cells to tyrosine kinase inhibitors. Cell Investig. 1, 100017 (2025).
Zhang, L. et al. RNA m6A modification: a key regulator in normal and malignant processes. Cell Investig. 1, 100023 (2025).
Lengefeld, J. et al. Cell size is a determinant of stem cell potential during aging. Sci. Adv. 7, eabk0271 (2021).
Debaugnies, M. et al. RHOJ controls EMT-associated resistance to chemotherapy. Nature 616, 168–175 (2023).
Lv, J. et al. Cell softness regulates tumorigenicity and stemness of cancer cells. EMBO J. 40, e106123 (2020).
Hayward, M. K., Muncie, J. M. & Weaver, V. M. Tissue mechanics in stem cell fate, development, and cancer. Dev. Cell 56, 1833–1847 (2021).
Dinavahi, S. S., Bazewicz, C. G., Gowda, R. & Robertson, G. P. Aldehyde dehydrogenase inhibitors for cancer therapeutics. Trends Pharmacol. Sci. 40, 774–789 (2019).
Chen, Y., Thompson, D. C., Koppaka, V., Jester, J. V. & Vasiliou, V. Ocular aldehyde dehydrogenases: protection against ultraviolet damage and maintenance of transparency for vision. Prog. Retin. Eye Res. 33, 28–39 (2013).
Liu, C. et al. ALDH1A1 activity in tumor-initiating cells remodels myeloid-derived suppressor cells to promote breast cancer progression. Cancer Res 81, 5919–5934 (2021).
Elcheva, I. A. et al. RNA-binding protein IGF2BP1 maintains leukemia stem cell properties by regulating HOXB4, MYB, and ALDH1A1. Leukemia 34, 1354–1363 (2020).
Venton, G. et al. Aldehyde dehydrogenases inhibition eradicates leukemia stem cells while sparing normal progenitors. Blood Cancer J. 6, e469 (2016).
Chute, J. P. et al. Inhibition of aldehyde dehydrogenase and retinoid signaling induces the expansion of human hematopoietic stem cells. Proc. Natl. Acad. Sci. USA 103, 11707–11712 (2006).
Rheinlaender, J. et al. Cortical cell stiffness is independent of substrate mechanics. Nat. Mater. 19, 1019–1025 (2020).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Zeng, A. G. X. et al. A cellular hierarchy framework for understanding heterogeneity and predicting drug response in acute myeloid leukemia. Nat. Med 28, 1212–1223 (2022).
Ley, T. J. et al. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 368, 2059–2074 (2013).
Database Resources of the National Genomics Data Center, China National Center for Bioinformation in 2022. Nucleic Acids Res. 50, D27-d38 (2022).
Huang B.LSC.GitHub https://github.com/XiXi1994bb/LSC, https://doi.org/10.5281/zenodo.17250099 (2024).
Acknowledgements
This work was supported by the Strategic Priority Research Program of the Chinese Academy of Sciences, No. XDB0940000 (F.N.), the Natural Science Foundation of China, Nos. 32070916 (F.N.), 82370159 (F.N.),12025207 (H.J.), and 12532015 (H.J.), USTC Research Funds of the Double First-Class Initiative, Nos. YD9100002505 (F.N.), YD2090002024 (H.J.), and YD2090002502 (H.Y.), the Extension Grant of the Anhui Provincial Distinguished Young Scholars Science Foundation, No. 2408085JX012 (F.N.), Research Funds of Center for Advanced Interdisciplinary Science and Biomedicine of IHM, No. QYPY20220007 (F.N.), and the Fundamental Research Funds for the Central Universities (H.Y.). We thank Ping Wang for helpful suggestions on our experiments. We also thank the Experimental Center for Life Science at USTC, particularly Zhen Yang, Qian Wang, and Zhenbang Liu, for their technical assistance in the experiments, and the Center for Micro and Nanoscale Research and Fabrication at USTC for providing technical support for this work.
Author information
Authors and Affiliations
Contributions
F.N., and H.J. conceived and conducted the project; F.N., H.J., M.Z., and H.Y. designed the experiments; M.Z., H.Y., and K.Q. conducted the research and summarized the data; B.H., T.L., and M.Z. analyzed the scRNA-seq data and conducted GSEA analysis; M.Z. contributed to mouse model development and cell culture; H.L. and M.Z. performed molecular knock-down experiments; M.W., Y.W., M.G., and X.L. provided critical advice on the experimental design and interpretation of the data; N.Z., X.S., X.Z. (Xuxu Zhao), M.G., Y.Z., X.H., Q.L., C.Z., X.Z. (Xiaoyu Zhu), and M.Z. collected human samples and clinical information; R.Z. and L.Q. provided comments and suggestions; M.Z., H. Y., K.Q., H.J., and F.N. wrote the manuscript. F.N. supervised the project. All authors commented and contributed to the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks John Pimanda, who co-reviewed with Henry Hampton, Todd Sulchek and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhu, M., Yang, H., Qiu, K. et al. Targeting leukemic stem cell biomechanics suppresses stemness and enhances NK cell-mediated immunotherapy. Nat Commun 16, 10800 (2025). https://doi.org/10.1038/s41467-025-65842-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65842-5