Abstract
KP.3.1.1 became a dominant successor to JN.1 by the second half of 2024 but the intrinsic pathogenicity and virological feature of KP.3.1.1 remain incompletely understood. Here, we comprehensively evaluated the pathogenesis and characteristics of KP.3.1.1 in comparison to JN.1 and other JN.1-derived variants including JN.1.7, KP.2, and KP.3. The unique S31del mutation on KP.3.1.1 spike confers further evasion to the clinically authorized mAb Pemivibart and reduces convalescent serum neutralization efficiency. Structural analysis indicates that S31del induces novel glycosylation sites that facilitates evasion of neutralizing antibodies. We further reveal that S31del significantly enhances pseudovirus entry efficiency in all evaluated cell types including the human primary nasal epithelial cells. Nevertheless, the intrinsic pathogenicity of KP.3.1.1 is similar to JN.1 and KP.3, and higher than that of JN.1.7 and KP.2 in a male hamster model. Interestingly, the increased virus infectivity conferred by S31del in KP.3.1.1 spike is counterbalanced by the NSP10 S33C mutation. Overall, our study indicates that a single spike mutation can confer both enhanced immune escape and increased viral infectivity. The opposing effects of spike and non-spike mutations highlight the complex interplay of viral genomic elements in shaping their overall fitness, and reveal the high plasticity of coronavirus evolution.
Similar content being viewed by others
Introduction
SARS-CoV-2 Omicron first identified in late 2021 has demonstrated substantially elevated infectability, pronounced immune evasion, and attenuated pathogenicity when compared to ancestral SARS-CoV-2 and earlier SARS-CoV-2 variants of concern (VOCs)1,2,3,4,5,6,7,8. Over the past three years, Omicron subvariants continue to adapt and circulate in the human population9,10,11,12,13,14. Omicron JN.1 emerged in 2023 and subsequently became the predominant strain, in part due to its spike L455S mutation that confers further increased immune evasion, infectivity, and transmissibility15,16,17,18,19,20. Among the JN.1 sublineages, KP.3.1.1 was classified by the World Health Organization as a variant under monitoring (VUMs), which became a dominant successor to JN.1 by the second half of 202421. Recent studies have characterized the immune escape and infectivity mediated by the KP.3.1.1 spike protein22,23,24. Nevertheless, the intrinsic pathogenicity and virological characteristics of KP.3.1.1 remain incompletely understood. In this study, we investigated the in vitro and in vivo virological features of KP.3.1.1 and included JN.1.7, KP.2, and KP.3 in comparative studies. We found that the unique S31del change in KP.3.1.1 spike significantly enhanced both immune evasion and viral infection fitness in the human nasal epithelial cells (hNECs), thereby conferring KP.3.1.1 a distinct competitive advantage over other JN.1-derived subvariants. Using a reverse genetics approach, we further revealed that the unique NSP10 S33C mutation in KP.3.1.1 negatively regulates virus replication, which equilibrates the overall pathogenicity of KP.3.1.1. Together, our results indicate that emerging Omicron subvariants may evolve spike mutations with dual advantages on immune evasion and intrinsic infectivity, highlighting the importance of continuous surveillance and characterization of emerging subvariants.
Result
Evolution of JN.1-derived subvariants JN.1.7, KP.2, KP.3, and KP.3.1.1
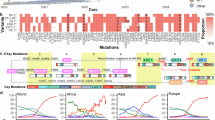
Following the global dominance of JN.1 in early 2024, multiple JN.1-derived subvariants have emerged, including JN.1.7, JN.1.11.1.2 (KP.2), JN.1.11.1.3 (KP.3), and JN.1.11.1.3.1.1 (KP.3.1.1) (Fig. 1a and Fig. S1a). All JN.1-derived subvariants maintain the L455S mutation in their spike proteins, a feature that has been demonstrated by us and others to contribute to immune evasion and enhanced infectivity25,26. The JN.1.7 subvariant contains two additional spike mutations (T572I and E1150D) relative to its parental JN.1 strain (Fig. 1b). KP.2 spike differs from that of JN.1 by three amino acid substitutions (R346T, F456L, and V1104L), while the KP.3 spike contains a distinct set of mutations (F456L, Q493E, and V1104L) relative to JN.1. Notably, the KP.3.1.1 spike protein carries an additional deletion at residue 31 (S31del) compared to KP.3 (Fig. 1b). These variants demonstrate distinct fitness advantages relative to JN.1. Globally, JN.1.7 exhibits a 21% fitness advantage, KP.2 shows a 42% advantage, and KP.3 demonstrates a 52% advantage. The relative fitness advantage of KP.3.1.1 reaches 80% compared to JN.1 (Supplementary Fig. 1b, c). Due to this substantial competitive advantage, KP.3.1.1 has become one of the most globally prevalent SARS-CoV-2 variants worldwide (Fig. 1c).
a Representative phylogenetic tree highlighting SARS-CoV-2 evolution. b Summary of the mutations of SARS-CoV-2 lineages JN.1, JN.1.7, KP.2, KP.3, and KP.3.1.1 compared to ancestral spike proteins with >10% frequency. The color represents the proportion of each mutation in each variant. c Variant frequency of SARS-CoV-2 lineages JN.1, JN.1.7, KP.2, KP.3, and KP.3.1.1 worldwide and in Asia, Europe and North America.
Antibody evasion of JN.1.7, KP.2, KP.3, and KP.3.1.1
To assess the recognition of the JN.1-derived variants by monoclonal antibodies (mAbs) either approved for COVID-19 therapy or currently in development, we performed pseudovirus neutralization assays evaluating both naturally circulating strains (JN.1, JN.1.7, KP.2, KP.3, and KP.3.1.1) and engineered pseudoviruses containing specific mutant combinations (JN.1 + R346T, JN.1 + F456L, JN.1 + R346T/F456L, JN.1 + T572I, and JN.1 + Q493E)24. These mAbs include clinically approved mAbs (Pemivibart27, Sipavibart28, and S30929) and the pre-clinical mAb SA5530, chosen for their demonstrated efficacy against the JN.1 variant (Fig. 2a). Consistent with recent studies27,31, the clinically authorized mAb Pemivibart demonstrated activity against JN.1, JN.1.7, and KP.2. However, its potency was reduced against JN.1 + R346T/F456L and KP.3, with an IC50 of 0.36 µg/mL and 0.633 µg/mL, respectively. Notably, Pemivibart exhibited further reduced neutralization potency against KP.3.1.1 (IC50 = 4.716 µg/mL). While Sipavibart maintained efficacy against JN.1 subvariants lacking the F456L mutation, its antiviral activity was dramatically diminished against JN.1 + F456L, JN.1 + R346T/F456L, KP.2, KP.3, and KP.3.1.1. These findings demonstrated that the F456L substitution in the spike protein is primarily responsible for the loss of neutralizing activity by Sipavibart. In contrast, mAb S309 displayed weak neutralization activity against all tested JN.1 subvariants. Interestingly, mAb SA55 remained highly effective against all evaluated JN.1 subvariants, likely because the mutated residues in these variants do not directly interact with SA55.
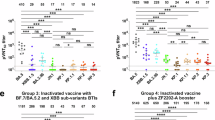
a Neutralization curve and IC50 heatmap of mAbs against Omicron-spike pseudoviruses (n = 3). b Neutralization of the indicated Omicron-spike pseudoviruses by sera collected from individuals 14–28 days after XBB or JN.1 reinfection, following prior BA.5 infection and two to three doses of inactivated vaccine (n = 16). The horizontal dashed line represents the detection limit (20-fold dilution). c Antigenic map and principal component analysis (PCA) map based on breakthrough infection serum neutralization data from (b). Virus positions are indicated by closed circles, while serum positions are represented by open squares. d Antigenic map and PCA map based on all infected hamster serum neutralization data from (e). Virus positions are represented by closed circles. In contrast, serum positions are shown as open squares. e Omicron-spike pseudoviruses were neutralized with sera from infected hamsters (n = 8 for BA.2, JN.1, and KP.3.1.1, n = 9 for JN.1.7, and KP.2, n = 7 for KP.3). The horizontal dashed line indicates the detection limit (20-fold). The n number represents biological repeats. Data represents mean ± SEM. Statistical significance in (b) was determined with one-way ANOVA with post hoc Tukey’s multiple comparisons test. *p < 0.05, ** p < 0.01, *** p < 0.001, ****p < 0.0001. ns represents not statistically significant. Data were obtained from three independent experiments. Source data are provided as a Source Data file.
Immune evasion of JN.1.7, KP.2, KP.3, and KP.3.1.1 with convalescent serum neutralization
To systematically characterize humoral immune evasion profiles, we evaluated both circulating JN.1-derived subvariants and engineered JN.1 variants containing single or combined mutations using convalescent plasma from two distinct cohorts of inactivated vaccine recipients who experienced breakthrough infections. The first cohort (n = 16) consisted of individuals who received inactivated vaccines, subsequently experienced BA.5 breakthrough infection, and later contracted XBB reinfection. The second cohort (n = 16) comprised inactivated vaccine recipients who experienced BA.5 breakthrough infection followed by JN.1 reinfection (Fig. 2b). Consistent with recent studies32,33, our neutralization analyses revealed that sera from XBB-reinfected individuals demonstrated weak neutralizing activity against all JN.1-derived subvariants, showing a maximum geometric mean titer (GMT) of 724 against JN.1. These sera exhibited significantly reduced neutralization titers against KP.2, KP.3, and KP.3.1.1 pseudoviruses, while maintaining relatively stable activity against JN.1.7. Importantly, we identified that both R346T and F456L mutations independently conferred immune escape capabilities, with their combination demonstrating further increased impact on evasion from neutralizing antibodies. The KP.2 pseudovirus, incorporating R346T, F456L, and V1104L spike mutations, exhibited significantly enhanced immune evasion relative to JN.1, demonstrating a 2.3-fold reduction in GMT using sera from XBB-reinfected individuals with prior BA.5 infection. Similarly, the KP.3 pseudovirus containing F456L, Q493E, and V1104L spike mutations showed a 2.4-fold GMT decrease. Most notably, the KP.3.1.1 pseudovirus displayed additional immune evasion capacity through its unique S31del mutation, evidenced by a further 1.8-fold GMT reduction compared to KP.3 (Fig. 2b and Supplementary Fig. 2a). Antigenic cartography of serum neutralization data revealed distinct clustering patterns among the variants. While the JN.1.7 pseudovirus maintained close antigenic similarity to its parental JN.1 strain, the emerging subvariants KP.3 and KP.3.1.1 formed a separate cluster, with KP.2 occupying an intermediate antigenic position distinct from both groups (Fig. 2c). These findings demonstrate that KP.3, KP.3.1.1, and KP.2 have undergone antigenic evolution from their parental JN.1 strain, highlighting both their shared antigenic properties and enhanced resistance to neutralizing antibodies compared to earlier variants34. In the JN.1 breakthrough infection cohort, we observed consistently higher neutralizing titers against all JN.1-derived subvariants relative to the XBB-reinfected group, with KP.3.1.1 showing the lowest GMT of 1124 (Fig. 2b and Supplementary Fig. 2b). Notably, neutralizing titers against KP.2, KP.3, and KP.3.1.1 pseudoviruses showed substantial reductions, displaying 2.4-fold, 2.4-fold, and 2.6-fold decreases in GMT, respectively, when compared to the parental JN.1 variant. These results demonstrated that the acquired mutations in these variants contribute to immune evasion. Specifically, our results indicated the F456L mutation as a major determinant of immune escape for KP.2, KP.3, and KP.3.1.1, establishing the foundation for their evasion capacity. Additionally, S31del conferred a modest but measurable enhancement of immune evasion, ultimately rendering KP.3.1.1 the overall most resistant variant. Antigenic cartography confirmed these observations, with JN.1 and JN.1.7 maintaining in close proximity while KP.3 and KP.3.1.1 formed a distinct antigenic cluster (Fig. 2c). Intriguingly, while KP.2 shares the F456L mutation with KP.3 and KP.3.1.1, its distinct antigenic positioning reveals substantial evolutionary divergence among these JN.1-derived variants, suggesting differential impacts of their respective spike mutations on immune recognition. Since the spike N-terminal domain (NTD) S31del in KP.3.1.1 conferred increased immune evasion, we additionally assessed the immune evasion capability of XEC, which similarly contains NTD mutations. While KP.3.1.1 spike differs from KP.3 spike at NTD S31del, XEC spike differs from KP.3 spike at NTD T22N and F59S. We found that when compared with the KP.3.1.1 pseudovirus, the XEC pseudovirus demonstrated a subtle 1.1-fold and 1.3-fold increase in evasion against BA.5 + XBB and BA.5 + JN.1 breakthrough infection sera, respectively (Supplementary Fig. 3a). Overall, these findings indicate that the NTD mutations on the spike of KP.3.1.1 and XEC confer increased immune evasion, which may facilitate the circulation of these variants.
Antigenicity of JN.1.7, KP.2, KP.3, and KP.3.1.1
To systematically compare immunogenicity across JN.1-derived variants, we established a hamster infection model using authentic viruses of JN.1, JN.1.7, KP.2, KP.3, and KP.3.1.1, with BA.2 included as an ancestral comparator. Serum samples were collected at 16 days post-infection (dpi) for subsequent immunological analyses (Fig. 2d, e). Neutralization assays revealed that convalescent sera from BA.2-infected hamsters demonstrated low neutralizing activity against all tested JN.1-derived subvariants (Fig. 2e). Strikingly, pseudoviruses bearing KP.3 and KP.3.1.1 spike proteins exhibited pronounced resistance to neutralization by convalescent sera from hamsters infected with BA.2, JN.1, JN.1.7, and KP.2, an effect primarily attributed to the Q493E mutation. Comparative analysis of neutralization profiles revealed that S31del conferred additional immune evasion properties to KP.3.1.1 relative to KP.3. Antigenic cartography through both distance mapping and principal component analysis (PCA) consistently grouped KP.3 and KP.3.1.1 into a shared cluster, while distinctly separating JN.1, JN.1.7, and KP.2 into independent antigenic groups35 (Fig. 2d). Collectively, these findings demonstrate that neutralization cross-reactivity patterns among JN.1 subvariants directly reflect their phylogenetic relationships. Importantly, our data establish that KP.3 and KP.3.1.1 have evolved substantial antigenic divergence from other JN.1-derived variants, including KP.2, suggesting these emerging strains may possess enhanced capacity to evade population immunity established through prior infection with earlier JN.1 subvariants.
Receptor interaction of KP.2, KP.3, and KP.3.1.1 spike
The binding affinity between SARS-CoV-2 spike protein and human angiotensin-converting enzyme 2 (hACE2) represents a critical determinant of viral fitness through modulating virus entry. To systematically evaluate these interactions, we quantified the binding kinetics of spike trimers from both naturally circulating JN.1 subvariants and engineered JN.1 mutants (JN.1 + R346T, JN.1 + F456L, JN.1 + R346T/F456L, JN.1 + Q493E, and JN.1 + T572I) with hACE2 using surface plasmon resonance (SPR) (Supplementary Fig. 4). In keeping with previous findings36,37, the R346T (KD = 6.97 nM), F456L (KD = 7.43 nM), R346T/F456L (KD = 5.62 nM), and T572I (KD = 5.89 nM) mutations did not substantially alter the hACE2-binding affinity of JN.1 (KD = 6.3 nM). Similarly, JN.1.7 (KD = 6.6 nM), which carries the T572I mutation, displayed a comparable hACE2 binding affinity to JN.1 + T572I (KD = 5.89 nM). KP.2 (KD = 3.7 nM), which harbors both the R346T and F456L mutations, exhibited slightly enhanced receptor binding affinity compared to the JN.1 (KD = 6.3 nM). Interestingly, introduction of the Q493E mutation in the JN.1 background (KD = 47.8 nM) resulted in substantially reduced receptor binding capacity. Notably, while the Q493E mutation alone substantially impaired receptor binding (JN.1 + Q493E, KD = 47.8 nM), its combination with F456L in KP.3 (KD = 4.76 nM) and KP.3.1.1 (KD = 5.76 nM) restored high-affinity hACE2 interaction. This observation suggests an epistatic interaction between L456 and E493 that counteracts the affinity loss induced by Q493E mutation.
To further elucidate the differences in spike-receptor binding caused by different amino acid mutations, we first determined the structure of ACE2 in complex with the stabilized prefusion ectodomains of KP.2 S(6 P) trimers using cryo-electron microscopy (cryo-EM) (Fig. 3). The 3D reconstruction revealed a single conformational state of the KP.2 spike-ACE2 complexes with three up-RBDs bound to three ACE2 molecules (Fig. 3a), resolved at a resolution of 2.27 Å (Supplementary Fig. 5). To improve the density around the spike-ACE2 interface, the NTD_RBD_ACE2 regions of KP.2 were locally refined to 2.90 Å, enabling the unambiguous model building for the RBD domain and ACE2 (Fig. 3b and Supplementary Fig. 5). The interfaces of the JN.1 spike-ACE2 and KP.2 spike-ACE2 complexes display a high degree of structural similarity, with the notable exception of a local variation induced by the F456L mutation (Fig. 3c, d). In the KP.2 spike-ACE2 complex, the side chain of ACE2K31 is positioned between L456 and Q493 of RBD, forming a salt bridge with Q493 and a hydrogen bond with the backbone of F490 (Fig. 3d). In contrast, in the JN.1 spike-ACE2 complex, the side chain of F456 is oriented towards the ACE2K31, preventing its side chain from inserting into the RBD. This hindrance disrupts both the hydrogen bonds and the salt bridge between ACE2K31 and RBD, resulting JN.1 with the weakest binding affinity for ACE2 when compared to its L456-containing descendants (KP.2, KP.3, KP.3.1.1) (Fig. 3c). Compared to the KP.2 spike, the KP.3 spike RBD incorporates a Q493E mutation, which introduces additional salt bridges between E493 and H34, further enhancing its binding affinity for ACE238 (Fig. 3e). KP.3.1.1 contains the same F456L and Q493E RBD mutations as KP.3 but possesses the S31del mutation at its NTD. We determined the structure of KP.3.1.1 spike at a global resolution of 3.06 Å. Local refinement improved the density map of NTD and RBD domains, resulting in a local resolution of 3.37 Å (Supplementary Fig. 6). Structural analysis uncovered an interesting consequence of the S31 deletion on KP.3.1.1 spike, which converted the adjacent N30 residue into a novel N-glycosylation site. The S31del change also induced significant conformational rearrangement of the F32 side chain compared to its orientation in both JN.1 and KP.2 spike structures (Fig. 3f, g). The conformational change of F32 resulted in the formation of a π−π interaction region among Y28-F32-Y91-F216-Y266, which might stabilize the local conformation of KP.3.1.1 (Fig. 3h). Interestingly, another glycosylation site was observed at N61 in KP.3.1.1 due to the S31 deletion (Fig. 3f).
a Cryo-EM structure of the KP.2 spike in complex with ACE2. Two perpendicular views of KP.2 spike-ACE2 are depicted as surface, with ACE2 in dodger blue and the trimeric spike in pink, medium purple, and purple. b Structure of KP.2 RBD-ACE2. c Detailed interactions between the JN.1 spikeF456, Q493-ACE2 (PDB: 8YZD) and the local cryo-EM maps. d Detailed interactions between the KP.2 spikeL456, Q493-ACE2, and the local cryo-EM maps. e Detailed interactions between the KP.3 spikeL456, E493-ACE2 (PDB: 9IUP), and the local cryo-EM maps. f Structural superposition of JN.1 (PDB:8Y5J) and KP.3.1.1 spike trimer. JN.1 and KP.3.1.1 are colored red orange and purple. N30, F32, N61 and glycosylation sites are shown as sticks. The red arrow indicates the conformational change of F32 in the structure of KP.3.1.1 spike trimer compared to JN.1. g The local cryo-EM maps of N30, F32, N61 and glycosylation sites in the structure of KP.3.1.1 spike trimer. h Structural superposition of JN.1 (PDB:8Y5J) and KP.3.1.1 spike trimer. JN.1 and KP.3.1.1 are colored red orange and purple. Red dashed circle indicates the interaction region comprising of Y28, F32, Y91, F216 and Y266, which were shown as sticks. i Structural prediction of the binding between NTD specific antibody C1717 or DH1052 and N30 site of SARS-CoV-2 WT or KP.3.1.1 spike protein. j, k Neutralization curve and IC50 heatmap of selected mAbs against KP.3 and KP.3.1.1 pseudoviruses, as well as their N30A mutant variants (n = 3). The n number represents biological repeats. Data represent mean ± SEM. Data were obtained from three independent experiments. Source data are provided as a Source Data file.
We next investigated the structural basis of NTD-directed antibody recognition against KP.3.1.1 spike. The S31 deletion facilitates a novel glycosylation event at the N30 site and induces a conformational rearrangement of residue F32, both of which are situated within the previously characterized antigenic site vi24,39. This region is known to be targeted by multiple antibodies, including C171740, DH105241, S2M2439, and polyclonal Fabs COV5742. Since high-resolution structural data are currently limited to C1717 and DH1052, we performed structural superposition of the KP.3.1.1 NTD onto these known antibody-complex structures to evaluate potential steric or electrostatic conflicts (Fig. 3i). A close-up structural analysis of the spike-Fab interface revealed that the newly acquired N30 glycan would sterically clash with residue I57 of the C1717 heavy chain, likely disrupting antibody binding. Similarly, for DH1052, key residues including S31 and Y32 in the light chain, as well as Y100H and H100I in the heavy chain, would experience steric hindrance from the N30 glycosylation (Fig. 3i). These structural observations suggest that the S31del change in KP.3.1.1 spike facilitates immune evasion by introducing glycosylation-mediated steric interference, thereby impairing the recognition and binding of NTD-targeting antibodies such as C1717 and DH1052.
To experimentally validate the impact of N30 glycosylation on antibody neutralization, we performed pseudovirus-based neutralization assays using KP.3 and KP.3.1.1 spike pseudoviruses against four spike-NTD-targeting antibodies, including 4–18, 4–19, C1717, and DH105240,41,43,44,45. Among these antibodies, C1717 and DH1052 recognize epitopes that encompass the N30 site, and therefore their binding is likely influenced by glycosylation at this position. Our results showed that antibodies 4-18 and 4-19 retained activity against KP.3, with IC₅₀ values of 2.315 μg/mL and 4.050 μg/mL, respectively, and showed comparable neutralization against KP.3.1.1, although 4-19 exhibited a moderate reduction in potency. In contrast, DH1052 and C1717 showed markedly reduced neutralization against KP.3.1.1 (IC₅₀ = 6.516 μg/mL and 3.876 μg/mL) compared with KP.3 (IC₅₀ = 1.221 μg/mL and 0.035 μg/mL, respectively), indicating that the S31 deletion in the spike protein confers substantial immune evasion (Figs. 3j and 3k). Next, we introduce the N30A mutation in the background of KP.3.1.1 spike to abolish the N30 glycosylation. Similarly, N30A was introduced in the background of KP.3 spike as a control. We evaluated the neutralization activity of C1717 against KP.3-N30A and KP.3.1.1-N30A pseudoviruses. In the KP.3 background, C1717 maintained a robust activity against the KP.3-N30A pseudovirus (IC₅₀ = 0.033 μg/mL), with IC₅₀ value comparable to that against the wild-type KP.3 pseudovirus (IC₅₀ = 0.035 μg/mL). Interestingly, in the KP.3.1.1 background, the N30A mutation renders the KP.3.1.1-N30A pseudovirus highly susceptible to C1717 neutralization (IC₅₀ = 0.054 μg/mL), which is an over 70-fold difference when compared to the wild-type KP.3.1.1 pseudovirus (IC₅₀ = 3.876 μg/mL) (Fig. 3j, k). Overall, these validation experiments confirm that the S31 deletion and the consequent N30 glycosylation significantly enhance immune evasion by interfering with antibody binding, thereby supporting the structural and functional role of this modification in facilitating immune escape.
Virological features of JN.1.7, KP.2, KP.3, and KP.3.1.1 in vitro
To systematically compare fusogenic activity across variants, we first employed a GFP reporter-based cell-cell fusion assay in 293T cells expressing spike proteins from JN.1.7, KP.2, KP.3, and KP.3.1.1. Quantitative analysis revealed all tested subvariant spikes mediated significantly less membrane fusion than their parental JN.1 spike protein (Fig. 4a). Notably, we observed comparable membrane fusion activity between KP.3 and KP.3.1.1 spike-expressing cells, indicating that the S31del mutation did not substantially alter the fusogenic potential of spike protein (Fig. 4a). We next evaluated the plaque morphology of JN.1 subvariants. To this end, we infected VeroE6-TMPRSS2 cells with authentic viruses (JN.1.7, KP.2, KP.3, and KP.3.1.1) with BA.2 and JN.1 included as controls. Plaque size analysis at 96 and 120 hpi demonstrated that KP.2 produced the smallest plaque size among all tested sublineages, which hinted its less robust replication fitness when compared with JN.1, JN.1.7, KP.3, and KP.3.1.1 (Fig. 4b). Additionally, we monitored the replication of JN.1-derived subvariants in Calu-3 cells and found that KP.2, KP.3, and KP.3.1.1 all replicated less efficiently compared to JN.1 (Figs. 4c and 4d). Area under the curve (AUC) analysis of virus replication in infected Calu-3 cells from 2 to 72 hpi showed that the infectious titer of KP.2, KP.3, and KP.3.1.1 in Calu-3 supernatants were 0.1- (P = 0.0002), 0.2- (P = 0.0005), and 0.5-fold (P = 0.0369) than that of JN.1, respectively (Fig. 4d).
a SARS-CoV-2 spike protein-mediated cell-cell fusion in 293T cells. The effector cells were co-transfected with GFP1-10 and spike protein expression plasmids and co-cultured with target cells which were co-transfected with GFP11 and human ACE2 protein expression plasmids for 24 h (n = 6). For analysis, images were split into RGB channels, and the green channel was selected for integrated density measurement of the target signal. A uniform threshold was applied across all images to minimize background noise and ensure consistent quantification. Scale bar, 200 μm. b Comparison of plaque size among Omicron subvariants. VeroE6-TMPRSS2 cells were infected with authentic viruses for 2 h before the viral inoculum was removed and cells were overlaid with agarose. At 96 and 120 hpi, cells were fixed with 4% paraformaldehyde and stained with crystal violet. Plaque sizes were measured with Adobe Photoshop (n = 12). c, d The replication of Omicron subvariants in Calu-3 cells. Calu-3 cells in 96-well plates were challenged with Omicron subvariants at 2 MOI. Cells lysates were extracted for sgE gene copy detection at the indicated time points (n = 6) (c). Infectious titers of supernatant samples were determined with TCID50 assays (n = 4 for BA.2, JN.1, KP.2, KP.3, and KP.3.1.1, n = 3 for JN.1.7) (d). The area under the curve was calculated by Graphpad Prism 10.0. e Entry of VSV-based Omicron-spike pseudoviruses in cell lines. 293T, Calu-3, Caco-2, and VeroE6-TMPRSS2 cells were transduced with pseudoviruses carrying spike proteins of Omicron subvariants. After 24 h, pseudovirus entry was quantified by measuring luciferase activity (n = 6). The fold change in luciferase signal was normalized to the mean signal of the JN.1-spike pseudovirus. The n number represents biological repeats. Data represent mean ± SEM. Statistical significance in (a–e) was determined with one-way ANOVA with post hoc Tukey’s multiple comparisons test. *p < 0.05, ** p < 0.01, *** p < 0.001, ****p < 0.0001. NS represents not statistically significant. Data were obtained from three independent experiments. Source data are provided as a Source Data file.
Pseudovirus entry of JN.1.7, KP.2, KP.3, and KP.3.1.1
To systematically evaluate spike-mediated viral entry efficiency, we constructed VSV-based pseudoviruses expressing spike proteins from JN.1.7, KP.2, KP.3, and KP.3.1.1 subvariants or engineered spike proteins, and quantified pseudovirus entry in 293T, Calu-3, Caco-2, and VeroE6-TMPRSS2 cells. We found that while KP.2 and KP.3 pseudoviruses demonstrated reduced entry efficiency, the KP.3.1.1 pseudovirus displayed markedly enhanced infectivity when compared to JN.1 or other JN.1-derived subvariants across all tested cell lines. Since KP.3.1.1 and KP.3 spike only differs at S31del, these results indicate that the S31del change in KP.3.1.1 spike confers increased infectivity21 (Fig. 4e). To delineate the entry pathway of these JN.1-derived subvariants, we employed Camostat (serine protease inhibitor) and E64D (cathepsin inhibitor) to selectively block the cell surface and endosomal entry pathway, respectively. Our inhibitor profiling revealed that JN.1.7, KP.2, and KP.3 pseudoviruses utilized largely similar entry pathways when compared to the JN.1 pseudovirus (Supplementary Fig. 7). Interestingly, the KP.3.1.1 pseudovirus displayed reduced sensitivity to both Camostat and E64D when compared to KP.3, suggesting that the S31del change might modulate the entry pathway of KP.3.1.1 (Supplementary Fig. 7). We also evaluated the role of S31del in using other proteases including members of the matrix metalloproteinase (MMP) family and the a disintegrin and metalloproteinase (ADAM) family, as they were found to facilitate Omicron entry46. Our results revealed no significant difference in KP.3.1.1 pseudovirus entry relative to the KP.3 pseudovirus across any of the tested MMPs and ADAMs (Supplementary Fig. 8). Since the NTD S31del change in KP.3.1.1 increased infectivity, we additionally evaluated the entry of XEC pseudovirus, which carries the NTD T22N and F59S mutations in spike when compared to KP.3. Our results demonstrated that the XEC pseudovirus is more robust than the JN.1 pseudovirus in virus entry in all evaluated cells. Nevertheless, the level of entry enhancement gained by T22N and F59S was in general less pronounced than that observed with S31del (Supplementary Fig. 3b)
Virological features of JN.1.7, KP.2, KP.3, and KP.3.1.1 in human nasal epithelial cells
Next, we evaluated the replication fitness of the JN.1-derived subvariants in differentiated primary hNECs. We found that while JN.1.7 and KP.2 replicated less efficiently than JN.1, KP.3 and KP.3.1.1 replicated to similar level when compared to JN.1 (Fig. 5a, b). Interestingly, the replication of KP.3.1.1 was 1.5-fold higher than that of KP.3 albeit not reaching statistical difference, hinting a potential role of S31del in infecting hNECs (Fig. 5b). In addition, we analyzed spike cleavage of JN.1-derived subvariants in infected hNECs. Western blot of spike processing in hNECs indicated comparable cleavage efficiency among JN.1-derived subvariants and JN.1 with the exception of KP.2, which demonstrated reduced S1/S2 cleavage (Fig. 5c). To further assess spike-mediated virus entry in hNECs, we transduced hNECs with BA.2, JN.1, JN.1.7, KP.2, KP.3, and KP.3.1.1 pseudoviruses. We found that while the JN.1 pseudovirus entered hNECs more efficiently when compared to the BA.2 pseudovirus, JN.1.7, KP.2, and KP.3 pseudovirus entry in hNEC were largely comparable to that of the JN.1 pseudovirus (Fig. 5d). Remarkably, the KP.3.1.1 pseudovirus demonstrated significantly higher infectivity in the hNECs, which was 7.3-fold (P < 0.0001) and 8.1-fold (P < 0.0001) higher when compared to the JN.1 pseudovirus and the KP.3 pseudovirus, respectively (Fig. 5d). Interestingly, this significantly elevated infectivity of KP.3.1.1 was not observed in immunofluorescence assays, which detected nucleocapsid protein expression at 24 hpi in hNECs (Fig. 5e).
a The infectious titer of Omicron subvariants in differentiated hNECs. The infectious titer in the supernatant was determined by TCID50 assays (n = 3). The area under the curve was calculated by Graphpad Prism 10.0. b The replication of Omicron subvariants in differentiated hNECs. Differentiated hNECs were challenged with Omicron subvariants at 2 MOI for 2 h. At 96 hpi, cell lysates were harvested and RNA was extracted for sgE gene detection (n = 3). c Spike cleavage of Omicron subvariants in hNECs. Differentiated hNECs were challenged with Omicron subvariants at 2 MOI and protein samples were collected at 96 hpi for Western blot analysis (n = 3). The intensity of the bands was quantified using ImageJ 1.53. d Pseudovirus entry in hNECs. Differentiated hNECs were transduced with pseudoviruses carrying Omicron subvariant spike proteins for 24 h. Pseudovirus entry was quantified and normalized to the mean luciferase signal of the JN.1-spike pseudovirus (n = 6). e Immunofluorescence staining of authentic viruses infected hNECs. Infected hNECs were fixed at 48 hpi to visualize ciliated cell marker beta-tubulin (red) and SARS-CoV-2 nucleocapsid protein (green). The DAPI channel was included in the merged images. Scale bar, 20 μm. The signal intensity was quantified with ImageJ 1.53 (n = 3). f Pseudovirus entry with S31del mutation in cell lines and differentiated hNECs. 293T, Calu-3, Caco-2, VeroE6-TMPRSS2, and differentiated hNECs were transduced with pseudoviruses carrying Omicron subvariants spike proteins with or without the S31del mutation (n = 8 for 293T, Calu-3, Caco-2, and VeroE6-TMPRSS2, n = 6 for hNECs). The fold change of signal readout was normalized to the mean signal readout of pseudoviruses carrying their wildtype counterparts. The n number represents biological repeats. Data represent mean ± SEM. Statistical significance was determined with one-way ANOVA with post hoc Tukey’s multiple comparisons test (a–e) or two-sided Student’s t tests (f). *p < 0.05, ** p < 0.01, *** p < 0.001, ****p < 0.0001. ns represents not statistically significant. Data were obtained from three independent experiments. Source data are provided as a Source Data file.
Since pseudovirus experiments performed in cell lines (Fig. 4e) and hNECs (Fig. 5d) both suggested a role of S31del in enhancing pseudovirus entry, we further evaluated the impact of S31del on virus entry. To this end, we constructed pseudoviruses expressing BA.2-, XBB.1-, and JN.1-spike proteins with or without the S31del mutation and compared entry efficiency of these pseudoviruses. Our data demonstrated that the S31del-containing pseudoviruses entered hNECs and cell lines significantly more robustly when compared to their control counterparts, confirming that S31del is capable of independently enhancing spike-mediated infectivity (Fig. 5f). To investigate the mechanism of how S31del enhances KP.3.1.1 infectivity, we first examined whether the S31del-induced N30 glycosylation contributes to the increased infectivity. We introduced the N30A substitution in the background of KP.3.1.1 spike to abolish the N30 glycosylation and produced the corresponding pseudoviruses. Similarly, N30A was introduced in the background of KP.3 as a control. We compared the entry efficiency of KP.3, KP.3 + N30A, KP.3.1.1, and KP.3.1.1 + N30A pseudoviruses in 293T, Calu-3, and Caco-2 cells. Our result demonstrated that the entry efficiency of the KP.3 + N30A pseudovirus was comparable to that of the KP.3 pseudovirus in 293T and Caco-2 cells. Importantly, KP.3.1.1 + N30A pseudovirus entry was also similar to that of the KP.3.1.1 pseudovirus in 293T and Caco-2 cells, which remained significantly higher than that of the KP.3 + N30A pseudovirus. This result indicates that the enhanced infectivity of KP.3.1.1 spike is not primarily mediated by N30 glycosylation (Supplementary Fig. 9). Interestingly, the N30A mutation in the background of both KP.3 and KP.3.1.1 substantially reduced virus infectivity in Calu-3 cells, suggesting that this mutation is not tolerated by the two viral spike proteins for Calu-3 cell entry. Next, we evaluated whether the spike-S31del change might enhance spike protein expression, thereby increasing the infectivity of both authentic KP.3.1.1 virus and corresponding pseudoviruses. To test this hypothesis, we transfected 293T cells with four pairs of spike plasmids, including the spike from BA.2, BA.2+S31del, XBB.1, XBB.1+S31del, JN.1, JN.1+S31del, KP.3, and KP.3+S31del (KP.3.1.1). We quantified the total and cell surface expression level of the spike proteins with flow cytometry at 24 h post transfection. Interestingly, we found that the S31del change increased both the total and cell surface expression level of spike when it is introduced in the background of all four spike proteins (Supplementary Fig. 10 and Supplementary Fig. 11). Together, these results suggest that the S31 deletion in KP.3.1.1 spike likely promoted infectivity through increasing the spike expression level.
Intrinsic pathogenicity of JN.1.7, KP.2, KP.3, and KP.3.1.1 in hamsters
We next assessed the intrinsic pathogenicity of JN.1.7, KP.2, KP.3, and KP.3.1.1 in golden hamsters with their ancestral JN.1 strain included as a control. At 2 days post-infection (dpi), lung tissues were collected for comprehensive analysis including quantification of subgenomic RNA copies, determination of infectious virus titers, viral antigen expression profiling, and histopathological evaluation (Fig. 6a). Quantitative analysis of virological parameters revealed that the JN.1-derived subvariants in general replicated less efficiently in hamster lungs when compared to their ancestral JN.1 strain, which is in keeping with findings obtained in Calu-3 cells (Fig. 6b–d). In hamster nasal turbinates, the viral burden of JN.1, JN.1.7, KP.3, and KP.3.1.1 were largely comparable with the exception of KP.2, which demonstrated markedly reduced virus replication (Supplementary Fig. 12). Immunofluorescence analysis of infected hamster lungs revealed reduced viral antigen expression for JN.1.7 (0.1-fold) and KP.2 (0.2-fold) relative to JN.1, while antigen expression of KP.3 and KP.3.1.1 was more comparable to that of JN.1 (Fig. 6d, f). In parallel, our findings from histology studies were largely in keeping with the virus replication results, which identified attenuated lung pathology in JN.1.7- and KP.2-infected hamsters, characterized by minimal alveolar exudates, inflammatory infiltration, and epithelial damage. In contrast, KP.3 and KP.3.1.1 induced similar histopathological changes in hamster lungs when compared with JN.1 (Fig. 6e, g). Omicron replication in the lung tissues is attenuated when compared to ancestral SARS-CoV-2 or earlier SARS-CoV-2 variants such as Delta6. We additionally compared the replication of JN.1 and KP.3.1.1 with that of Delta and found that the infectious titer of both JN.1 and KP.3.1.1 in hamster lung was more than 100-fold lower than that of Delta, which is keeping with our recent reports on the continuous attenuation of emerging Omicron subvariants10,11,12,20 (Supplementary Fig. 13).
a Schematic of hamster infection and sample harvest. Golden hamsters were intranasally infected with 1 × 105 PFU JN.1, JN.1.7, KP.2, KP.3, or KP.3.1.1 (n = 5). Hamster lungs were harvested at 2 dpi for viral load detection or histological analysis. Created in BioRender. Zhu, R. (2025) https://BioRender.com/syp9nuf. b Viral gene copy detection in hamster lungs by RT-qPCR. RNA was extracted from homogenized hamster lungs, and RT-qPCR was conducted to detect the sgE and β-actin gene copies (n = 5). c Detection of infectious titer in infected hamster lungs by TCID50 assay. Homogenized hamster lung samples were 10-fold serial diluted to infect VeroE6-TMPRSS2 cells in 96-well plates (n = 5). Cellular cytopathic effect was recorded after 4 days of incubation to calculate the infectious titer. d, f Immunofluorescence staining of infected hamster lungs. Infected hamster lungs were embedded and sectioned, followed by immunostaining with SARS-CoV-2 nucleocapsid (N) protein antibody. Cell nuclei were labeled with DAPI stain (n = 5) (f). The fluorescence signal intensity was measured using ImageJ 1.53 (d). Scale bar, 200 μm. e, g Infected hamster lungs H&E staining and histological scoring. Sectioned hamster lungs were stained with hematoxylin and eosin. Images were taken under a microscope (g) and histological scoring was conducted based on the degree of tissue damage (n = 5) (e). Scale bar, 70 μm. The n number represents biological repeats. Data represent mean ± SEM. Statistical significance in (b–e) was determined with one-way ANOVA with post hoc Tukey’s multiple comparisons test. *p < 0.05, *** p < 0.001. ns represents not statistically significant. Source data are provided as a Source Data file.
The role of NSP10-S33C in virus replication in vivo and vitro
Peculiarly, while our results clearly indicated a positive role of S31del in KP.3.1.1 spike in augmenting virus entry, we did not detect a significant increase in KP.3.1.1 replication in vitro or in vitro. With sequence alignment, we found that KP.3.1.1 contains an additional ORF1a S4286C (NSP10 S33C) mutation that is absent in previous VOCs and other Omicron-derived variants including KP.3 (Fig. 7a). We speculated that this mutation might play a role in offsetting the replication fitness gained by the spike S31del change in KP.3.1.1. To evaluate this hypothesis, we introduced the NSP10-S33C mutation into the recombinant BA.1 (rBA.1) backbone (Fig. 7b). The modified plasmid was sequence-verified and transfected into VeroE6-TMPRSS2 cells for virus rescue (Fig. 7c). Comparative analysis of viral replication in VeroE6, VeroE6-TMPRSS2, and Calu-3 cells revealed that rBA.1-NSP10-S33C replicated significantly less efficiently than wild-type rBA.1 across all three evaluated cell lines (Fig. 7d, e). We further introduced the spike-S31del change into rBA.1-NSP10-S33C. Our results demonstrated that S31del rescued the attenuated replication fitness caused by NSP10-S33C in all evaluated cell types. For example, the infectious titer of rBA.1-NSP10-S33C was 16-fold (P = 0.0061) lower than that of rBA.1 in Calu-3 cells at 72 h post infection. Introducing the spike-S31del change substantially restored virus replication. The infectious titer of rBA.1-NSP10-S33C-spike-S31del was largely comparable with that of rBA.1 in Calu-3 cells at 72 h post infection of (P = ns) (Figs. 7d and 7e). Furthermore, we challenged hamsters with rBA.1 and rBA.1-NSP10-S33C, and detected virus replication in lungs and nasal turbinates (NT) at 2 dpi (Fig. 7f). Our results indicated that the NSP10-S33C mutation reduced the replication of rBA.1 in hamster lung and NT tissues (Fig. 7g, h). In keeping with these observations, immunofluorescence staining of infected hamster lungs similarly indicated a reduction of antigen expression contributed by the introduction of the NSP10-S33C mutation (Fig. 7i, j). Collectively, these findings demonstrate that the NSP10-S33C mutation attenuates SARS-CoV-2 replication both in vitro and in vivo. Coronavirus NSP10 is known to interact with NSP14 and the interaction is important for the exoribonuclease activity of NSP1447,48. Structural modeling revealed that residue S33 in NSP10 forms a hydrogen bond with N67 in NSP14 (Fig. 7k, l). The S33C substitution introduces a cysteine residue that may disrupt native hydrogen bonding networks and induces conformational rearrangements in the α2’ helix of NSP10 within its N-terminal domain, likely impairing NSP10-NSP14 complex formation, which provides a mechanistic basis for the attenuated replication observed in KP.3.1.1.
a Sequence comparison at ORF1a residue 4286 among SARS-CoV-2 sublineages. b Schematic of reverse genetic modification. Created in BioRender. Zhu, R. (2025) https://BioRender.com/8k3tw9y. c Sequence comparison of the constructed rBA.1-NSP10-S33C mutant with rBA.1. d, e Replication of rBA.1-NSP10-S33C and rBA.1-NSP10-S33C-spike-S31del in cell lines compared to rBA.1. VeroE6 and VeroE6-TMPRSS2 cells were challenged with viruses at 0.01 MOI and Calu-3 cell was challenged at 0.5 MOI. Cell lysate and supernatant were harvested for viral sgE gene qPCR (n = 6) (d) and infectious titer detection (n = 4) (e). f Schematic of the hamster infection experiment. Golden hamsters were intranasally infected with 1 × 105 PFU rBA.1 or rBA.1-NSP10-S33C (n = 4). Created in BioRender. Zhu, R. (2025) https://BioRender.com/msxbmo1. g, h Viral load of rBA.1-NSP10-S33C in hamster lungs and NTs. Hamster lungs and NTs were harvested for sgE gene qPCR detection (g) or infectious titer detection (h) (n = 4). i, j Antigen expression in hamster lungs. Infected hamster lungs were stained with SARS-CoV-2 N specific antibody (j). The fluorescent intensity was quantified with ImageJ 1.53 (i) (n = 4). Scale bar, 200 μm. k Structural superposition of NSP10 (PDB code: 6ZCT) and the NSP10-NSP14 complex (PDB code: 7DIY). Unbound NSP10 is depicted in gray cartoon. In the complex, NSP10 is rendered in salmon cartoon while NSP14 is shown in green surface presentation. l Close-up view of the interaction between NSP10 and NSP14. Residue S33 of NSP10 and residues Q65-N67 of NSP14 are shown in stick mode. The red arrow indicates the movement of α2’ helix in NSP10 upon binding to NSP14. The dashed yellow line indicates the hydrogen bond. The n number represents biological repeats. Data represent mean ± SEM. Statistical significance was determined with two-way ANOVA with post hoc Tukey’s multiple comparisons test (d, e) or two-sided Student’s t tests (g–i). *p < 0.05, ** p < 0.01, *** p < 0.001, ****p < 0.0001. ns represents not statistically significant. Data were obtained from three independent experiments. Source data are provided as a Source Data file.
Discussion
JN.1 is a descendent variant of BA.2.86 characterized by its acquisition of the L455S mutation in spike. L455S conferred enhanced immune evasion capacity, which contributes to the global dominance of JN.1 in late 2023 and early 202420,49,50,51,52. As JN.1 continues to circulate, JN.1-derived variants retain the L455S change while have evolved additional mutations for further gained immune escape, including the R346T, F456L, Q493E, and T572I mutations in spike. These accumulated mutations progressively enhanced the competitive fitness of emerging sublineages, ultimately driving the replacement of the ancestral JN.1 variant37,53. In this study, we comprehensively investigated the virological features of KP.3.1.1, which replaced JN.1 as the dominant circulating variant by late 2024. Through comparative analyses of KP.3.1.1 against JN.1 and additional JN.1-derived variants including JN.1.7, KP.2, and KP.3, we identified mechanistic advantages that underlie KP.3.1.1’s global predominance.
KP.3.1.1 differs from KP.3 by a spike S31del mutation and a ORF1a S4286C (NSP10 S33C) mutation. In antibody evasion assays, we demonstrated that the clinically approved monoclonal antibody Pemivibart exhibited further reduced neutralization potency against KP.3.1.1 when compared to KP.3, indicating that the spike S31del mutation confers substantial immune evasion. In convalescent sera neutralization assays, we showed that while R346T and F456L mutations both conferred immune escape, additional immune evasion capacity is acquired by KP.3.1.1 through its unique spike S31del mutation. The reduced neutralization titers against KP.3.1.1 is in keeping with results from others22,23,24. By using NTD-directed antibodies including C1717 as well as mutagenesis at the N30 position, we experimentally validated the impact of N30 glycosylation on antibody neutralization. These findings in part explained how KP.3.1.1 became a globally dominant variant compared to its ancestral JN.1 variant.
Interestingly, we found that the S31del mutation significantly enhances viral infectivity. Compared with that of the KP.3 pseudovirus, the KP.3.1.1 pseudovirus is more efficient in virus entry in 293T (7.2-fold), Calu-3 (16.7-fold), Caco-2 (4.2-fold), VeroE6-TMPRSS2 (3.4-fold), and hNEC (8.1-fold). The increased virus entry does not appear to be due to better usage of a particular entry pathway, as the enhanced entry was observed in cells that predominantly support endosomal entry (293T) and cells that predominantly support plasma membrane entry (Calu-3 and hNEC). Importantly, when we introduced S31del into BA.2-, XBB.1-, and JN.1-spike pseudoviruses, we found that S31del consistently promoted the entry of these pseudoviruses when compared to their wildtype counterparts. Further investigation revealed that S31del enhances the expression level of the modified spike protein compared to their wild-type counterparts. The elevated expression provides a mechanistic explanation for the increased infectivity observed in KP.3.1.1 pseudovirus carrying this mutation.
Despite the elevated viral infectivity, we did not observe a significant increase in KP.3.1.1 replication in infected cells or hamsters when compared to KP.3. This naturally led to the postulation that the only other mutation between KP.3.1.1 and KP.3, the non-spike NSP10 S33C mutation, may play a role in offsetting the increased virus entry capacity conferred by S31del in KP.3.1.1. To evaluate this hypothesis, we engineered recombinant BA.1 carrying NSP10 S33C and demonstrated that the NSP10 S33C mutation significantly reduced virus replication in both in vitro and in vivo settings. Structural modeling suggests that the S33C mutation induces a conformational change in NSP10 that weaken its interaction with NSP14, which may explain the observed replication deficiency54,55,56. The identification of the role of the NSP10 S33C mutation on virus replication is highly interesting and we cannot explain why KP.3.1.1 retains a non-spike mutation that negatively impacts virus replication, since it will result in lower virus fitness when compared to viruses with a wildtype NSP10. At the current stage, we speculate that NSP10 S33C may play an unknown role important for KP.3.1.1’s survival or that unidentified epistasis may exist between spike and NSP10 proteins, which will warrant further investigations.
Overall, our study identified the S31del mutation in KP.3.1.1 spike NTD as a pivotal determinant of enhanced immune evasion and cellular infectivity, which collectively drove its global dominance (Supplementary Fig. 14). Immune escape is often achieved at the cost of virus fitness57,58,59. Here, our results indicate that the spike NTD S31del mutation confers enhanced immune escape while at the same time increases viral infectivity. Future variants may develop similar mutations in the spike protein that provide dual advantages to virus fitness, highlighting the importance of continued surveillance and characterization of emerging variants of SARS-CoV-2 to optimize pandemic control strategies, including vaccination.
Methods
Virus infection in animal models
All our animal experiments were approved by the Committee on the Use of Live Animals in Teaching and Research (CULATR) of The University of Hong Kong (HKU) under CULATR 22-139. Golden Syrian hamsters were sourced from the Center for Comparative Medicine Research (CCMR) at HKU and housed in individually ventilated cages under controlled conditions (65% humidity, 21–23 °C, 12 h/12 h light/dark cycle). For hamster infection, 4- to 6-week-old male hamsters were randomly allocated to experimental groups and anesthetized with ketamine and xylazine, followed by intranasal inoculation of 50 µL pre-diluted 1ⅹ105 PFU Omicron JN.1, JN.1.7, KP.2, KP.3 and KP.3.1.1 virus in DMEM. At 2 days post-infection (dpi), hamsters were euthanized and sacrificed for lung tissue collection to perform virological and histopathological examinations as we previously described60.
Viruses and biosafety
SARS-CoV-2 Omicron BA.2 (GISAID: EPI_ISL_9845731), JN.1 (GISAID: EPI_ISL_18841631), JN.1.7 (GISAID: EPI_ISL_19351034), KP.2 (GISAID: EPI_ISL_19351035), KP.3 (GISAID: EPI_ISL_19535068), and KP.3.1.1 (GISAID: EPI_ISL_19534723) were isolated from laboratory-confirmed COVID-19 patients in Hong Kong. Virus isolation was performed on archived clinical specimens obtained during routine clinical practice from laboratory-confirmed COVID-19 patients in Hong Kong. This study was approved by the Institutional Review Board of the University of Hong Kong/Hospital Authority Hong Kong West Cluster (HKU/HA HKW IRB) (Reference No.: UW 21-313; Protocol No.: 202104). The requirement for informed consent was waived by the IRB as the research involved no additional interventions and utilized only anonymized archival specimens. All SARS-CoV-2 subvariants were propagated in VeroE6-TMPRSS2 cells and titrated using plaque assays. All in vivo and in vitro experiments involving authentic viruses were conducted in the Biosafety Level 3 (BSL-3) facility at the Department of Microbiology, The University of Hong Kong, following approved standard operating procedures61.
Human serum collection
Peripheral blood samples were collected from vaccinated individuals (2–3 doses of inactivated vaccines) with prior BA.5 infection followed by XBB or JN.1 reinfection at Nanjing Hospital of Chinese Medicine. Serum was isolated by centrifugation and stored at −80 °C. All collections complied with Declaration of Helsinki guidelines under approved protocols (Ethics Committee of Nanjing Hospital of Chinese Medicine Affiliated to Nanjing University of Chinese Medicine, KY2023073). All participants provided written informed consent. The background information of the convalescent donors is summarized in Supplementary Table 1.
Phylogenetic, mutation, and epidemic analysis
A total of 3863 SARS-CoV-2 genome sequences and pair-wised phylogenetic tree were downloaded from nextstrain (https://nextstrain.org/)62 with open data63 from January 2022 to October 2024 (Supplementary Table 2). We accessed recent 6-month global data on November 05, 2024, which highlight the Omicron variants. We re-colored the phylogenetic tree by labeling all variants as “BA.2,“ “JN.1,” “JN.1.7,” “KP.2,” “KP.3,” and “KP.3.1.1”. We analyzed the mutation and epidemic by using tools from covSPECTRUM (https://cov-spectrum.org/)64 based on the GISAID data (https://www.gisaid.org/)63 on November 4, 2024. The mutation proportion of each variant was based on all genome sequences obtained on June 27, 2025. Here, only mutations with >10% frequency were included. The variants frequency of each variant was calculated based on genome sequences from November 1, 2023 to November 1, 2024 obtained on June 27, 2025.
Fitness analysis
Relative fitness advantage was analyzed based on covSPECTRUM (https://cov-spectrum.org/)64. The analysis employed a logistic growth model to estimate the relative fitness advantage of SARS-CoV-2 variants compared to the reference variant JN.1 across different geographic regions65. Input data consisted of daily variant counts stratified by date, variant designation, observed counts, and geographic region information from November 1, 2023 to November 1, 2024 obtained on June 27, 2025. Temporal dynamics of fitness advantage for major SARS-CoV-2 lineages was analyzed following previous publications65,66. The analysis assumed a viral generation time of 3.5 days derived from past research67. A sliding window of 90 days was used to ensure temporal resolution while maintaining statistical stability. For each variant, a logistic growth model was applied within each 90-day window advancing weekly. The model compared the variant’s frequency against JN.1, using a binomial generalized linear model with a logit link. The response variable was binary (1 for the variant, 0 for JN.1), and the predictor was time (in days). Only windows with ≥5 data points and non-zero total samples were included. The growth rate was derived from the slope coefficient, and was converted to relative fitness advantage, which is expressed as a percentage change per generation (3.5 days). Confidence intervals (95%) were calculated using standard errors from the model covariance matrix. All analyses were implemented in R version 4.3.0 utilizing glm for model fitting, purrr for iterative processing across variants, and ggplot2 for visualization.
Protein expression and purification
We generated mammalian expression plasmids encoding stabilized prefusion spike trimers (S6P) for JN.1, JN.1.7, KP.2, KP.3, and KP.3.1.1 variants, along with JN.1 derivatives containing individual or combined mutations. Each construct was synthesized by cloning the full-length spike sequence into pCMV3, incorporating C-terminal 8×His and 2×Strep-tag II affinity tags. Soluble dimeric ACE2 expression plasmids were prepared using established protocols68. The Expi293 cells were transiently transfected with the corresponding expression plasmids using 1 mg/mL polyethylenimine (PEI, Polysciences) for protein expression. Five days post-transfection, culture supernatants were collected and subjected to affinity purification using Ni-Smart resin (Smart Lifesciences, Changzhou, China; catalog #SA036100, Ni Smart Beads 6FF) according to the manufacturer’s protocol. Purified proteins were quality-controlled by SDS-PAGE to verify molecular weight and purity before experimental use.
Antibody expression and purification
For this study, the monoclonal antibody Sipavibart was commercially sourced (AntibodySystem, Catalog #DVV00353), whereas Pemivibart, S309, SA55, 4-18, 4-19, C1717, and DH152 monoclonal antibodies were obtained from established in-house collections as described in our previous publications20,45. Briefly, the variable region genes were codon-optimized for human cell expression and synthesized by HuaGene™ (Shanghai, China) into gWiz plasmids encoding the constant regions of human IgG1 heavy and light chains. Antibodies were expressed in Expi293F cells (ThermoFisher, A14527) by co-transfection of plasmids encoding the heavy and light chains using polyethylenimine (Polysciences). The cells were cultured at 37 °C with shaking at 125 rpm under 8% CO₂. Five days post-transfection, the supernatants were harvested for antibody purification using MabSelect™ PrismA affinity chromatography (Cytiva, 17549801).
Surface plasmon resonance (SPR)
SPR experiments were conducted using the Biacore 8 K system (Cytiva) following previously described protocols69. Spike trimer proteins were immobilized on CM5 sensor chips (Cytiva, 29149603) via amine coupling (Cytiva, BR100050) to achieve a target density of 2000 response units (RU). Serial dilutions of purified human ACE2 (6.25, 12.5, 25, 50, 100, and 200 nM) were injected across the functionalized sensor surface. Binding interactions were monitored at room temperature using Biacore 8 K Evaluation Software 3.0 (Cytiva), with sensorgrams analyzed by fitting the raw data to a 1:1 Langmuir binding model.
Hamster serum collection
Four-week-old male Syrian hamsters were intranasally inoculated with 2 × 105 PFU of Omicron BA.2, JN.1, JN.1.7, KP.2, KP.3, or KP.3.1.1 under anesthesia. Sixteen days post-inoculation, serum samples were harvested, centrifuged, and heat-inactivated at 56 °C for 30 min.
Production of pseudoviruses
Pseudoviruses were prepared as we previously described70,71. Plasmids encoding spike proteins from Omicron variants JN.1, JN.1.7, KP.2, KP.3, and KP.3.1.1, along with engineered JN.1 variants containing single or combined mutations, were synthesized. Expi293F cells were grown to 3 × 106 cells/mL before transfection with the indicated spike gene using polyethylenimine (PEI, Polysciences). Cells were cultured overnight at 37 °C with 8% CO2 and VSV-G pseudo-typed ΔG-luciferase (G*ΔG-luciferase, Kerafast) was used to infect the cells in DMEM at a multiplicity of infection (MOI) of 5 for 4 h. Subsequently, cells were washed three times with PBS containing 2% FBS (PBS/2%). After 24 h, the transfection supernatant was collected and clarified by centrifugation at 300 × g for 10 min. Each viral stock was then incubated with 20% I1 hybridoma (anti-VSV-G; CRL-2700, ATCC) supernatant for 1 h at 37 °C to neutralize residual VSV-G pseudotyped ΔG-luciferase virus prior to titer determination and aliquot storage at −80 °C.
Pseudovirus neutralization
Neutralization assays were performed by incubating pseudoviruses with serially diluted monoclonal antibodies or serum samples, with neutralizing activity quantified through luciferase reporter gene expression reduction as previously established72,73. In brief, VeroE6 cells were seeded in a 96-well plate at a density of 2 × 104 cells per well. On the following day, pseudoviruses were incubated with serial dilutions of the test samples in triplicate for 30 min at 37 °C. The mixture was then added to cultured cells and incubated for 24 h, after which luminescence was measured using the Luciferase Assay System (Beyotime). IC50 was defined as the dilution at which the relative light units (RLUs) were reduced by 50% compared to the virus control wells (virus + cells) after subtracting the background signal in the control groups with cells only. The IC50 values were calculated using nonlinear regression analysis in GraphPad Prism 10.0.
Antigenic cartography
The antigenic maps were generated as we described previously20. The antigenic map was constructed through antigenic cartography analysis of serum neutralization data using the Racmacs package (https://acorg.github.io/Racmacs/). The map was generated in R with 10,000 optimization steps and default two-dimensional parameters. Map distances between sublineages and serum positions were optimized to reflect fold-changes in neutralizing ID50 titers relative to each serum’s maximum titer, with one map unit representing a two-fold titer difference.
Cryo-EM data collection and processing
The stabilized prefusion ectodomains of the KP.3.1.1 S(6P) trimer, the KP.2 S(6P) trimer and ACE2 were expressed in Expi293F cells and purified following previously described protocols20. The purified KP.2 spike trimer was concentrated to 0.6 mg/mL, mixed with ACE2 ectodomain at a 1:1.2 molar ratio, and incubated at 4 °C for 12 min. The KP.3.1.1 spike trimer was concentrated to 0.5 mg/mL, and 3 μL of the protein was applied to a novel light-emitting porous Ni-Ti amorphous alloy film supported by 400-mesh gold grid. Samples were vitrified using a Vitrobot Mark IV (Thermo Fisher Scientific) with the following parameters: 2 s blot time, blot force of -3, and 10 s wait time. Cryo-EM data were acquired on a 300 kV Titan Krios G4 microscope (Thermo Fisher Scientific) equipped with a Falcon 4i direct electron detector and Selectris X energy filter (20 eV slit width), using EPU software for automated data collection in AFIS mode. EER movie stacks were collected in super-resolution mode, KP.2 S-ACE2 magnification of 130,000×, corresponding to a physical pixel size of 0.932 Å, dose fractioned to 1080 frames. For KP.3.1.1-spike trimer, the magnification was 165,000×, corresponding to a physical pixel size of 0.760 Å. The defocusing range is −1.0 to −3.0 μm, and the total dose is about 50e−/Å274,.
For the KP.2 spike-ACE2 complex dataset, all dose-fractionated images underwent motion correction and dose weighting in RELION, followed by contrast transfer function estimation using cryoSPARC patch CTF analysis75. Particle selection was performed through automated blob picking and template matching, followed by extraction at 320-pixel box size with Fourier-space cropping to 160 pixels. All subsequent 2D/3D classification and refinement procedures were conducted in cryoSPARC75 (Supplementary Fig. 5a). 958,918 particles were selected after 2D classification, and these particles were used to do ab initio reconstruction heterogeneous refinement. This process yielded 393,648 converged particles, corresponding to a “3-up” conformation with three ACE2 bound to each RBD. Next, these particle sets were re-extracted with a box size of 400 pixels and subjected to non-uniform refinement, achieving a final a resolution of 2.27 Å. To analyze the receptor interaction interface, a focused mask covering the NTD, RBD, and ACE2 regions was applied for alignment-free 3D classification and local refinement after achieving global particle convergence and applying C3 symmetry expansion. The provided particles depict a local reconstruction of the NTD-RBD-ACE2 region, resulting in a 2.90 Å map generated using 393,648 particles (Supplementary Fig. 5b).
For the KP.3.1.1 spike dataset, motion correction, CTF estimation, particle picking, and 2D classification were performed in cryoSPARC as previously described. From this processing, 83,990 high-quality particles were selected for ab initio reconstruction and heterogeneous refinement (4 classes). The optimal class served as input for Topaz training. The trained particle set was then merged with particles from the best refinement class, followed by removal of duplicate particles. Upon non-uniform refinement for these 75,262 particles, we yielded a map of 3.06 Å resolution. For further improving the density of NTD and RBD regions, 3D classification and local refinement with mask were performed and resulted in a 3.37 Å map using 36,563 particles (Supplementary Fig. 6).
Model building and refinement
The sharpened maps were generated by EMReady76. For model building, the initial model was generated by SWISS-MODEL77 and fitted to the map using UCSF Chimera78. The model was then manually adjusted in COOT and further refined using PHENIX79. Figures were produced by UCSF Chimera and UCSF ChimeraX80. Data collection and model refinement statistics are shown in Supplementary Table 3.
Cell culture and virus infection
293T, Caco-2, and VeroE6 cell lines (ATCC) were cultured in DMEM (11965-092, Gibco) supplemented with 10% FBS, 100 U/mL penicillin, and 100 μg/mL streptomycin. VeroE6-TMPRSS2 cells (JCRB Cell Bank) were maintained in DMEM containing 10% FBS, 100 U/mL penicillin, 100 μg/mL streptomycin, and 2% G418 (ant-gn-5, InvivoGen), while Calu-3 cells were cultured in DMEM/F12 (10565042, Gibco) with 10% FBS and antibiotics. All cells were incubated at 37 °C with 5% CO2 and routinely tested for mycoplasma contamination. For viral infection assays, Calu-3 cells were seeded in 96-well plates and infected with SARS-CoV-2 Omicron subvariants at MOI 2 for replication studies.
Human nasal epithelial cell air-liquid interface culture and virus infection
hNECs cultured at air-liquid interface (ALI) were obtained from Epithelix (EP02MP, Switzerland) and maintained in MucilAir medium (EP04MM) before infection. For viral challenges, hNECs were washed apically three times with medium to remove mucus and then infected with SARS-CoV-2 Omicron subvariants (MOI = 2) at the apical surface, followed by 2-h incubation at 37 °C for viral entry before inoculum removal. Viral load was assessed at 2, 24, 48, 72, and 96 h post-infection through apical collection of supernatants/cell lysates for infectious titer detection and RT-qPCR quantification of viral RNA. For immunofluorescence, cells were fixed with neutral-buffered formalin, permeabilized with 0.1% Triton X-100 (Sigma-Aldrich, 11332481001), and stained with rabbit anti-SARS-CoV-2 nucleocapsid serum (in-house) and mouse anti-β-tubulin (Sigma-Aldrich, T7941) to detect viral protein and ciliated cells, respectively, followed by incubation with Alexa Fluor-conjugated secondary antibodies (goat anti-mouse 594 [Thermo Fisher, A-11005] and goat anti-rabbit 488 [A-11034]). Samples were mounted with ProLong Diamond Antifade Mountant with DAPI (Thermo Fisher, P36962) and imaged on an LSM980 confocal microscope (Zeiss) using Z-stack acquisition and maximum intensity projection processing in ZEN Blue software.
The efficiency of Omicron-spike pseudovirus entry in cell lines and hNECs
To compare the entry efficiency of Omicron-spike pseudoviruses (BA.2, JN.1, JN.1.7, KP.2, KP.3, KP.3.1.1, and engineered JN.1 variants with single/combined mutations), 293T, Calu-3, Caco-2, and VeroE6-TMPRSS2 cells were seeded in 96-well plates for transduction. Parallel experiments using differentiated hNECs assessed entry of BA.2, JN.1, JN.1.7, KP.2, KP.3, and KP.3.1.1 spike pseudoviruses. After 24-h incubation, cells were PBS-washed and lysed (RG129M) for luminescence detection (Luciferase Assay System, RG066M, Beyotime). For inhibitor studies, cells were pretreated with 10 μM Camostat (HY-13512) or E64D (HY-100229) (MedChemExpress) for 2 h prior to transduction with the pseudovirus panel, with luminescence measured post-24h incubation.
Hamster lung histological and immunofluorescence staining
For SARS-CoV-2 antigen detection and histopathological evaluation, infected hamster lungs were immersion-fixed in 4% paraformaldehyde (24 h), ethanol-washed, and paraffin-embedded using a Leica TP1020 processor. Tissue sections (5 μm) were microtome-cut (Thermo Fisher), slide-mounted, and oven-dried (37 °C overnight). For immunofluorescence, dewaxed slides underwent antigen retrieval (Vector Labs H-3300, 90 s boiling), Sudan black B quenching, and 5% FBS blocking before incubation with primary (rabbit anti-SARS-CoV-2-N) and secondary antibodies (goat anti-rabbit IgG-FITC, Thermo 65-6111, 1:500 dilution), followed by DAPI mounting (Vector H-1200). For H&E staining, slides were processed with Gill’s hematoxylin/eosin. All imaging used an Olympus BX53 microscope, with nucleocapsid signal intensity quantified in ImageJ 1.53 and histopathology scored semi-quantitatively (n = 5 hamsters/group, 3 slides/hamster)46.
RNA extraction and quantitative reverse transcription polymerase chain reaction (qRT-PCR)
RNA from infected cell lysates or homogenized animal tissues was extracted using the RNeasy Mini Kit (74106, Qiagen), while RNA from infected cell supernatants was isolated with the Viral RNA Mini Kit (52906, Qiagen) or QIAsymphony RNA Kit (931636, Qiagen). To quantify gene expression levels in the extracted RNA samples, RT-qPCR was performed using the QuantiNova Probe RT-PCR Kit (208354, Qiagen) or QuantiNova SYBR Green RT-PCR Kit (208154, Qiagen) on a LightCycler 480 Real-Time PCR System (Roche, Basel, Switzerland). Primer and probe sequences are shown in Supplementary Table 4.
Flow cytometry
293T cells were transfected with spike expressing plasmids for 24 h before fixing. The transfected cells were detached with 10 mM EDTA in PBS, fixed in 4% paraformaldehyde, followed by immunolabelling with a rabbit anti-SARS-CoV-2-spike-S1 specific antibody (ab283942, abcam) (1:500). Flow cytometry was performed using a BD FACSCanto II flow cytometer (BD) and data was analyzed using FlowJo (version v10.7) (BD).
Construction of reverse genetic modified BA.1 expression plasmid
To engineer the recombinant BA.1 plasmid with targeted mutations, we employed the Lambda-Red recombination system using the Counter-Selection BAC Modification Kit (Gene Bridges K002). Following manufacturer protocols, we co-transformed the pSmartBac-BA.1 plasmid (previously constructed via homologous recombination) and pRed/ET plasmid into DH10b competent cells (Thermo Fisher EC0113) prior to modification. Two distinct DNA fragments were generated: one containing the rpsL counter-selection marker and another bearing the viral sequence with desired mutations. These fragments were sequentially introduced into cells for recombination. Positive clones were selected using appropriate antibiotics, and plasmid integrity was verified by sequencing after maxiprep purification.
Infectious titer quantification of infected hamster lung and nasal turbinate tissues
To assess the infectious titer in infected hamster tissues, a 50% tissue culture infectious dose (TCID50) assay was conducted. Briefly, VeroE6-TMPRSS2 cells were seeded into 96-well plates and inoculated with 10-fold serial dilutions of lung homogenates. After a 2-h incubation, residual viral particles were removed, and cells were washed three times with serum-free DMEM. At four days post-infection, cytopathic effects (CPE) were evaluated to determine the median tissue culture infectious dose (TCID50).
293T cell-cell fusion assay
A population of 293T cells was co-transfected with GFP1-10 (cat#68715, Addgene) along with SARS-CoV-2 spike expression plasmids to generate effector cells. Similarly, a separate set of 293T cells was co-transfected with GFP11 (cat#68716, Addgene) and human ACE2 expression plasmids to establish target cells. Twenty-four hours post-transfection, cells were harvested using EDTA-Trypsin (25200072, Gibco) and subsequently co-cultured at a 1:1 ratio in 8-well chamber slides for an additional 24 h. After incubation, cells were fixed with 10% formalin and permeabilized with 0.1% Triton X-100 (11332481001, Sigma, USA) at room temperature. Finally, slides were mounted using antifade mounting medium containing DAPI (H-1200, Vector Laboratories) and visualized using an Olympus BX73 fluorescence microscope (Olympus Life Science, Tokyo, Japan). The images were processed using ImageJ 1.53 software by separating the RGB channels, after which the green channel was selected for integrated density measurements. A uniform threshold was applied to all cell-cell fusion images to reduce background interference and ensure consistent quantification.
The analysis of spike cleavage in hNECs
To investigate spike cleavage of Omicron subvariants in hNECs, differentiated hNECs were infected with BA.2, JN.1, JN.1.7, KP.2, KP.3, and KP.3.1.1 at a MOI of 2 for 96 h prior to sample collection. Cells were lysed in RIPA buffer (89901, Thermo Fisher Scientific), and proteins were separated by 8% SDS-PAGE. For Western blot analysis, membranes were blocked with 5% non-fat milk for 2 h and incubated overnight at 4 °C with either a rabbit anti-SARS-CoV-2 spike S2 antibody (40590-T62, Sino Biological, China) or a β-actin antibody (clone AC-74, A5316, Sigma-Aldrich). Subsequently, horseradish peroxidase (HRP)-conjugated secondary antibodies (31460, Thermo Fisher Scientific) were applied, and protein bands were visualized using SuperSignal West Pico PLUS Chemiluminescent Substrate (34580, Thermo Scientific) with an Alliance Imager apparatus (Uvitec, Cambridge, UK).
Statistical analysis
Data in the figures are presented as means ± standard error of the mean (SEM). Comparisons between two groups were performed using two-sided Student’s t tests, while comparisons among three or more experimental groups were conducted using one-way or two-way ANOVA. Differences were considered statistically significant at p < 0.05. All data analyses were performed using GraphPad Prism 10.0.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Coordinates and maps associated with data reported in this manuscript were deposited to the Electron Microscopy Data Bank (EMDB) and Protein Data Bank (PDB) with accession numbers EMD-64078 and PDB 9UE7 (KP.2 spike-ACE2), EMD-64077 and PDB 9UE6 (KP.2 spike NTD-RBD-ACE2 local refinement), EMD-64086 (KP.3.1.1 spike) and EMD-64085, PDB 9UEL (KP.3.1.1 spike NTD-RBD local refinement). Source data are provided with this paper.
References
Liu, L. et al. Striking antibody evasion manifested by the Omicron variant of SARS-CoV-2. Nature 602, 676–681 (2022).
Meng, B. et al. Altered TMPRSS2 usage by SARS-CoV-2 Omicron impacts infectivity and fusogenicity. Nature 603, 706–714 (2022).
Cao, Y. et al. Omicron escapes the majority of existing SARS-CoV-2 neutralizing antibodies. Nature 602, 657–663 (2022).
Planas, D. et al. Considerable escape of SARS-CoV-2 Omicron to antibody neutralization. Nature 602, 671–675 (2022).
Suzuki, R. et al. Attenuated fusogenicity and pathogenicity of SARS-CoV-2 Omicron variant. Nature 603, 700–705 (2022).
Shuai, H. et al. Attenuated replication and pathogenicity of SARS-CoV-2 B.1.1.529 Omicron. Nature 603, 693–699 (2022).
Halfmann, P. J. et al. SARS-CoV-2 Omicron virus causes attenuated disease in mice and hamsters. Nature 603, 687–692 (2022).
McMahan, K. et al. Reduced pathogenicity of the SARS-CoV-2 omicron variant in hamsters. Med 3, 262–268 e264 (2022).
Yamasoba, D. et al. Virological characteristics of the SARS-CoV-2 Omicron BA.2 spike. Cell 185, 2103–2115 e2119 (2022).
Chan, J. F. et al. Virological features and pathogenicity of SARS-CoV-2 Omicron BA.2. Cell Rep. Med. 3, 100743 (2022).
Shuai, H. et al. The viral fitness and intrinsic pathogenicity of dominant SARS-CoV-2 Omicron sublineages BA.1, BA.2, and BA.5. EBioMedicine 95, 104753 (2023).
Hu, B. et al. Divergent trajectory of replication and intrinsic pathogenicity of SARS-CoV-2 Omicron post-BA.2/5 subvariants in the upper and lower respiratory tract. EBioMedicine 99, 104916 (2024).
Qu, P. et al. Immune evasion, infectivity, and fusogenicity of SARS-CoV-2 BA.2.86 and FLip variants. Cell 187, 585–595.e586 (2024).
Zhang, L. et al. SARS-CoV-2 BA.2.86 enters lung cells and evades neutralizing antibodies with high efficiency. Cell 187, 596–608 e517 (2024).
Kaku, Y. et al. Virological characteristics of the SARS-CoV-2 JN.1 variant. Lancet Infect. Dis. 24, e82 (2024).
Yang, S. et al. Fast evolution of SARS-CoV-2 BA.2.86 to JN.1 under heavy immune pressure. Lancet Infect. Dis. 24, e70–e72 (2024).
Planas, D. et al. Distinct evolution of SARS-CoV-2 Omicron XBB and BA.2.86/JN.1 lineages combining increased fitness and antibody evasion. Nat. Commun. 15, 2254 (2024).
Wang, Q. et al. XBB.1.5 monovalent mRNA vaccine booster elicits robust neutralizing antibodies against XBB subvariants and JN.1. Cell Host Microbe 32, 315–321 e313 (2024).
Li, P. et al. Distinct patterns of SARS-CoV-2 BA.2.87.1 and JN.1 variants in immune evasion, antigenicity, and cell-cell fusion. mBio 15, e0075124 (2024).
Liu, Y. et al. Lineage-specific pathogenicity, immune evasion, and virological features of SARS-CoV-2 BA.2.86/JN.1 and EG.5.1/HK.3. Nat. Commun. 15, 8728 (2024).
Kaku, Y. et al. Virological characteristics of the SARS-CoV-2 KP.3.1.1 variant. Lancet Infect. Dis. 24, e609 (2024).
Li, P. et al. Neutralization and spike stability of JN.1-derived LB.1, KP.2.3, KP.3, and KP.3.1.1 subvariants. mBio, e0046425 https://doi.org/10.1128/mbio.00464-25 (2025).
Uriu, K. et al. Antiviral humoral immunity induced by JN.1 monovalent mRNA vaccines against SARS-CoV-2 omicron subvariants including JN.1, KP.3.1.1, and XEC. Lancet Infect. Dis. 25, e61 (2025).
Liu, J. et al. Enhanced immune evasion of SARS-CoV-2 variants KP.3.1.1 and XEC through N-terminal domain mutations. Lancet Infect. Dis. 25, e6–e7 (2025).
Focosi, D., Quiroga, R., McConnell, S., Johnson, M. C. & Casadevall, A. Convergent evolution in SARS-CoV-2 spike creates a variant soup from which new COVID-19 waves emerge. Int. J. Mol. Sci. 24, https://doi.org/10.3390/ijms24032264 (2023).
Cao, Y. et al. Imprinted SARS-CoV-2 humoral immunity in duces convergent Omicron RBD evolution. Nature 614, 521–529 (2023).
Wang, Q., Guo, Y., Ho, J. & Ho, D. D. Activity of research-grade pemivibart against recent SARS-CoV-2 JN.1 sublineages. N. Engl. J. Med 391, 1863–1864 (2024).
Cai, Y. et al. AZD3152 neutralizes SARS-CoV-2 historical and contemporary variants and is protective in hamsters and well tolerated in adults. Sci. Transl. Med. 16, eado2817 (2024).
Pinto, D. et al. Cross-neutralization of SARS-CoV-2 by a human monoclonal SARS-CoV antibody. Nature 583, 290–295 (2020).
Cao, Y. et al. Rational identification of potent and broad sarbecovirus-neutralizing antibody cocktails from SARS convalescents. Cell Rep. 41, 111845 (2022).
Planas, D. et al. Escape of SARS-CoV-2 variants KP.1.1, LB.1, and KP3.3 from approved monoclonal antibodies. Pathog. Immun. 10, 1–11 (2024).
Kaku, Y. et al. Virological characteristics of the SARS-CoV-2 KP.3, LB.1, and KP.2.3 variants. Lancet Infect. Dis. 24, e482–e483 (2024).
Xu, K. et al. Neutralization of SARS-CoV-2 KP.1, KP.1.1, KP.2 and KP.3 by human and murine sera. NPJ Vaccines 9, 215 (2024).
Wilks, S. H. et al. Mapping SARS-CoV-2 antigenic relationships and serological responses. Science 382, eadj0070 (2023).
Tamura, T. et al. Virological characteristics of the SARS-CoV-2 XBB variant derived from recombination of two Omicron subvariants. Nat. Commun. 14, 2800 (2023).
Jian, F. et al. Evolving antibody response to SARS-CoV-2 antigenic shift from XBB to JN.1. Nature 637, 921–929 (2025).
Wang, Q. et al. Recurrent SARS-CoV-2 spike mutations confer growth advantages to select JN.1 sublineages. Emerg. Microbes Infect. 13, 2402880 (2024).
Feng, L. et al. Structural and molecular basis of the epistasis effect in enhanced affinity between SARS-CoV-2 KP.3 and ACE2. Cell Discov. 10, 123 (2024).
McCallum, M. et al. N-terminal domain antigenic mapping reveals a site of vulnerability for SARS-CoV-2. Cell 184, 2332–2347 e2316 (2021).
Feng, Z. et al. Structural and functional insights into the evolution of SARS-CoV-2 KP.3.1.1 spike protein. Cell Rep. 44, 115941 (2025).
Li, D. et al. In vitro and in vivo functions of SARS-CoV-2 infection-enhancing and neutralizing antibodies. Cell 184, 4203–4219 e4232 (2021).
Barnes, C. O. et al. Structures of human antibodies bound to SARS-CoV-2 spike reveal common epitopes and recurrent features of antibodies. Cell 182, 828–842 e816 (2020).
Liu, L. et al. Antibodies targeting a quaternary site on SARS-CoV-2 spike glycoprotein prevent viral receptor engagement by conformational locking. Immunity 56, 2442–2455 e2448 (2023).
Wang, Z. et al. Analysis of memory B cells identifies conserved neutralizing epitopes on the N-terminal domain of variant SARS-Cov-2 spike proteins. Immunity 55, 998–1012 e1018 (2022).
Liu, L. et al. Potent neutralizing antibodies against multiple epitopes on SARS-CoV-2 spike. Nature 584, 450–456 (2020).
Chan, J. F. et al. Altered host protease determinants for SARS-CoV-2 Omicron. Sci. Adv. 9, eadd3867 (2023).
Bouvet, M. et al. Coronavirus Nsp10, a critical co-factor for activation of multiple replicative enzymes. J. Biol. Chem. 289, 25783–25796 (2014).
Lin, S. et al. Crystal structure of SARS-CoV-2 nsp10 bound to nsp14-ExoN domain reveals an exoribonuclease with both structural and functional integrity. Nucleic Acids Res 49, 5382–5392 (2021).
Zhang, L. et al. Rapid spread of the SARS-CoV-2 JN.1 lineage is associated with increased neutralization evasion. iScience 27, 109904 (2024).
Lewnard, J. A. et al. Immune escape and attenuated severity associated with the SARS-CoV-2 BA.2.86/JN.1 lineage. Nat. Commun. 15, 8550 (2024).
Li, L. et al. Spike structures, receptor binding, and immune escape of recently circulating SARS-CoV-2 Omicron BA.2.86, JN.1, EG.5, EG.5.1, and HV.1 sub-variants. Structure 32, 1055–1067.e1056 (2024).
Yang, H. et al. Structural basis for the evolution and antibody evasion of SARS-CoV-2 BA.2.86 and JN.1 subvariants. Nat. Commun. 15, 7715 (2024).
Li, P. et al. Neutralization escape, infectivity, and membrane fusion of JN.1-derived SARS-CoV-2 SLip, FLiRT, and KP.2 variants. Cell Rep. 43, 114520 (2024).
Czarna, A. et al. Refolding of lid subdomain of SARS-CoV-2 nsp14 upon nsp10 interaction releases exonuclease activity. Structure 30, 1050–1054.e1052 (2022).
Matsuda, A. et al. Despite the odds: formation of the SARS-CoV-2 methylation complex. Nucleic Acids Res. 52, 6441–6458 (2024).
Imprachim, N., Yosaatmadja, Y. & Newman, J. A. Crystal structures and fragment screening of SARS-CoV-2 NSP14 reveal details of exoribonuclease activation and mRNA capping and provide starting points for antiviral drug development. Nucleic Acids Res. 51, 475–487 (2023).
Dadonaite, B. et al. Spike deep mutational scanning helps predict success of SARS-CoV-2 clades. Nature 631, 617–626 (2024).
Moulana, A. et al. Compensatory epistasis maintains ACE2 affinity in SARS-CoV-2 Omicron BA.1. Nat. Commun. 13, 7011 (2022).
Li, W. et al. Key mechanistic features of the trade-off between antibody escape and host cell binding in the SARS-CoV-2 Omicron variant spike proteins. EMBO J. 43, 1484–1498 (2024).
Chen, Y. et al. COVID-19 mRNA vaccine protects against SARS-CoV-2 Omicron BA.1 infection in diet-induced obese mice through boosting host innate antiviral responses. EBioMedicine 89, 104485 (2023).
Chu, H. et al. Coronaviruses exploit a host cysteine-aspartic protease for replication. Nature 609, 785–792 (2022).
Hadfield, J. et al. Nextstrain: real-time tracking of pathogen evolution. Bioinformatics 34, 4121–4123 (2018).
Khare, S. et al. GISAID’s role in pandemic response. China CDC Wkly 3, 1049–1051 (2021).
Chen, C. et al. CoV-Spectrum: analysis of globally shared SARS-CoV-2 data to identify and characterize new variants. Bioinformatics 38, 1735–1737 (2022).
Chen, C. et al. Quantification of the spread of SARS-CoV-2 variant B.1.1.7 in Switzerland. Epidemics 37, 100480 (2021).
Meijers, M., Ruchnewitz, D., Eberhardt, J., Luksza, M. & Lassig, M. Population immunity predicts evolutionary trajectories of SARS-CoV-2. Cell 186, 5151–5164 e5113 (2023).
Chan, L. Y. H. et al. Estimating the generation time for SARS-CoV-2 transmission using household data in the United States, December 2021–May 2023. medRxiv, 2024.2010.2010.24315246 https://doi.org/10.1101/2024.10.10.24315246 (2024).
Zhao, X. et al. Potent and broadly neutralizing antibodies against sarbecoviruses induced by sequential COVID-19 vaccination. Cell Discov. 10, 14 (2024).
Chu, H. et al. Host and viral determinants for efficient SARS-CoV-2 infection of the human lung. Nat. Commun. 12, 134 (2021).
Wang, X. et al. Neutralization of Omicron BA.4/BA.5 and BA.2.75 by booster vaccination or BA.2 breakthrough infection sera. Cell Discov. 8, 110 (2022).
Wang, X. et al. Enhanced neutralization of SARS-CoV-2 variant BA.2.86 and XBB sub-lineages by a tetravalent COVID-19 vaccine booster. Cell Host Microbe 32, 25–34.e25 (2024).
Wang, X. et al. Neutralization of distinct Omicron sublineages by longitudinal vaccination sera. J. Med. Virol. 94, 5090–5092 (2022).
Hu, B. et al. Spike mutations contributing to the altered entry preference of SARS-CoV-2 omicron BA.1 and BA.2. Emerg. Microbes Infect. 11, 2275–2287 (2022).
Hsieh, C. L. et al. Structure-based design of prefusion-stabilized SARS-CoV-2 spikes. Science 369, 1501–1505 (2020).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
He, J., Li, T. & Huang, S. Y. Improvement of cryo-EM maps by simultaneous local and non-local deep learning. Nat. Commun. 14, 3217 (2023).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–w303 (2018).
Pettersen, E. F. et al. UCSF Chimera-a visualization system for exploratory research and analysis. J. Comput Chem. 25, 1605–1612 (2004).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr D. Struct. Biol. 74, 531–544 (2018).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Acknowledgements
We gratefully acknowledge all data contributors, i.e., the Authors and their Originating laboratories responsible for obtaining the specimens, and their Submitting laboratories for generating the genetic sequence and metadata and sharing via the GenBank, on which this research is based. Supplementary table is attached. We also thank the Center of Cryo-Electron Microscopy, Core Facility of Shanghai Medical College, Fudan University for the support on cryo-EM data collection. Hin Chu acknowledge supports by funding from National Natural Science Foundation of China Excellent Young Scientists Fund (Hong Kong and Macau) (32122001); the Health and Medical Research Fund (COVID1903010-14, 23220522, CID-HKU1-5), the Food and Health Bureau, The Government of the Hong Kong Special Administrative Region; the General Research Fund (17118621, 17119122), Collaborative Research Fund (C7103-22G, C7060-21G), and Theme-Based Research Scheme (T11-709/21-N), the Research Grants Council of the Hong Kong Special Administrative Region; InnoHK initiative of the Innovation and Technology Commission of the Hong Kong Special Administrative Region Government; National Key Research and Development Program of China (projects 2021YFC0866100 and 2023YFC3041600); General Programme, Guangdong Provincial National Science Foundation, China (2023A1515012325, 2023A1515011891); the University of Hong Kong Outstanding Young Researcher Award; the University of Hong Kong Research Output Prize (Li Ka Shing Faculty of Medicine); and the support from Dr. Gallant Ho on the Gallant Ho Outstanding Young Professorship. This study was also supported by funding from Research Fellow Scheme (HKU RFS2425-7S06 to J.F.-W.C.), the Research Grants Council of the Hong Kong Special Administrative Region, China, the National Key Research and Development Program of China (2023YFC2605400 and 2023YFC3404000 to X.Z.; 2021YFC2302500 to L.S.), National Natural Science Foundation of China (32270142 to P.W.; 32300121 to X.Z; 92469108 to L.S.), Shanghai Rising-Star Program (22QA1408800 to P.W.), Shanghai Pujiang Programme (23PJD007 to X.Z.), the Program of Science and Technology Cooperation with Hong Kong, Macao and Taiwan (23410760500 to P.W.), and Shanghai Pilot Program for Basic Research—Fudan University 21TQ1400100 (25TQ011 to P.W.). P.W. acknowledges support from Xiaomi Young Talents Program. Lei Sun acknowledges support from the R&D Program of Guangzhou Laboratory (SRPG22-003). Maozhou He acknowledges support from National Science Foundation of China (32401012); Zhejiang Provincial Natural Science Foundation of China (LQ24C050004); Research Funds from Hangzhou Institute for Advanced Study (2024HIAS-Y011). L.Z. acknowledges support from the National Natural Science Foundation of China (32273099). B.-Z.Z., H.C., and Y.-F.H. also thank support from Major Program of the National Natural Science Foundation of China (92369201). All donors and funding sources had no role in the study design, data collection, analysis, interpretation, or writing of the manuscript.
Author information
Authors and Affiliations
Contributions
J.S., Y.L., H.L., Z.C., L.W., Y.H., Y.C., B.H., H.S., Y.W., X.Z., W.-M.C., L.-L.C., J.-P.C., B.S., H.Z., D.Y., L.Z., S.Y., J.Z., K.-Y.Y., K.K.-W.T., J.F.-W.C., and H.C. performed experiments, analyzed the data, or provided key resources for the in vitro and in vivo pseudovirus and authentic virus assays. X.Z., J.L., X.L., S.J., Y.Z., Q.W., and P.W. performed experiments, analyzed the data, or provided key resources for the neutralization assays, generated spike trimers, and constructed the pseudoviruses. X.J., Y.X., Y.W., M.H., and L.S. performed experiments, analyzed the data, or provided key resources for the Cryo-EM determination of spike-hACE2 interaction. Y.-F.H., J.-D.H., and B.-Z.Z. performed experiments, analyzed the data, or provided key resources for bioinformatic analysis. J.S., X.Z., M.H., L.S., P.W., and H.C. wrote the manuscript. H.C. conceptualized and supervised the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Vincent Munster and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shi, J., Zhao, X., Jin, X. et al. Pathogenicity, virological features, and immune evasion of SARS-CoV-2 JN.1-derived variants including JN.1.7, KP.2, KP.3, and KP.3.1.1. Nat Commun 16, 11002 (2025). https://doi.org/10.1038/s41467-025-66018-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66018-x