Abstract
Zika virus (ZIKV) is typically mild in humans but can cause severe congenital defects when contracted during pregnancy. Chimeric live-attenuated vaccine candidate YF-ZIK previously showed protective efficacy against lethal infection and developmental abnormalities in mice after a single dose. Here we demonstrate that YF-ZIK is safe, induces antiviral immunity, and protects rhesus macaques against high-dose experimental challenge. A single subcutaneous dose elicits neutralizing antibodies within 7–14 days, boosted by a second dose at 4 weeks. Passive serum transfer protects AG129 mice, supporting antibodies as a correlate of protection. YF-ZIK triggers balanced Th1/Th2 responses and a transcriptional profile resembling the licensed YF17D vaccine, involving multiple pathways favoring polyvalent immunity. Upon challenge, vaccinated macaques show no detectable viral RNA nor seroconversion to anti-ZIKV NS1 antibodies, suggesting sterilizing immunity. Systems analysis identifies TNFRSF17 as predictor of antibody responses to YF-ZIK, and GNAS and CD207 (Langerin) as potentially linked to clinical outcomes. The favorable preclinical safety, immunogenicity, and efficacy of YF-ZIK justify its future evaluation in humans.
Similar content being viewed by others
Introduction
Zika virus (ZIKV) is a mosquito-borne Orthoflavivirus, belonging to a genus that also contains other important human pathogens like the yellow fever virus (YFV), dengue virus (DENV), Japanese encephalitis virus (JEV), West Nile virus (WNV), and tick-borne encephalitis virus. Alike other orthoflaviviruses, ZIKV has a ~ 11 kb-long, single-stranded RNA genome containing one open-reading frame, encoding for three structural proteins, namely capsid, precursor membrane (prM), the envelope (E), and several non-structural (NS1-5) proteins1. Until 2007, ZIKV circulated only in Africa and Asia, sporadically causing self-limited asymptomatic infections in humans. However, from 2007 to 2016, ZIKV explosively spread to the Asian-Pacific and Americas, causing more than 700.000 documented infections in humans. Unexpectedly, during the last epidemic, ZIKV infections were associated with severe complications including a high rate of fetal microcephaly, congenital malformations in infants born to mothers infected during pregnancy, and neuromuscular disorders, namely Guillain-Barré syndrome, in adults2,3. Despite the significant decline in reported cases after 2016, the high chance of recurring outbreaks and transmission to previously spared regions by global travel and expansion of mosquito vectors urges the development of safe and effective ZIKV vaccines3,4.
In response to the ZIKV emergency, multiple vaccine candidates have been developed employing different technologies, including DNA, messenger RNA, recombinant protein, inactivated whole virus, virus-like particle, non-replicating, and live-attenuated viral vector vaccines. Several that proved safe and effective in mice and non-human primates (NHPs) have entered clinical trials, but none have been approved yet for use in humans3,4. Full clinical development has been impeded by a rapid decline in Zika cases after the 2015/2016 pandemic. Though ZIKV transmission continues across multiple regions5, setting up field studies for formal vaccine efficacy evaluation remain challenging due to limited surveillance and the impossibility to predict local outbreaks6. Future vaccine development may therefore need to consider alterative pathways towards licensure based on immunological correlates of protection, alike in the past for YFV and JEV vaccines7. All the more, understanding ZIKV pathogenesis and immunology in step-up models such as NHP is crucial to define relevant parameters that need to be considered for future clinical trial design.
The mechanisms conferring protective immunity against ZIKV are not fully understood. Virus-specific neutralizing antibodies (NAbs) targeting the structural proteins, especially E, are considered the major correlate of protection against ZIKV and other Orthoflavivirus infections7,8. Seminal studies testing several vaccine platforms, including purified inactivated, DNA, and adenovirus vector-based vaccine candidates have established that ZIKV-specific serum NAbs, either induced by active immunization or instilled by passive transfer of purified immunoglobulin, can protect from ZIKV infection in both mouse and NHP models9,10,11,12, in line with the widely accepted concept of seroprotection. Thereby, specific NAbs above a certain threshold protected from viremia, scored as suppression of viral RNA below RT-qPCR detection limits9,10,11. However, depending on the respective animal model employed, on the viral dose and route of inoculation used for challenge, as well as on the methodology used for antibody detection, thus determined critical NAb titres varied widely; ranging from about 1:100 to 1:1000 (ref. 13). Notably, lower NAb levels were required to prevent viremia in semi-permissive BALB/c mice challenged with a low (102 plaque forming units, PFU) virus dose of ZIKV9,10 than to protect readily susceptible rhesus macaques challenged with a moderately higher dose (103 PFU)10,11. In line, in these models also a potent monoclonal NAb could protect from viremia in a dose-dependent manner14. By contrast, highly susceptible IFN-α/β and γ receptor-deficient (AG129) mice15,16 could only partially be protected from lethal infection by NAbs, even when treated with high amounts of purified, fully functional IgG and despite use of a very low challenge dose12. Careful inspection further showed that abolition of viremia is not more than a proxy for a certain degree of protection. In fact, also macaques that stayed non-viremic following challenge showed later a raise in NAbs to saturating levels, whereby titres increased markedly by up to two orders of magnitude compared to post-vaccination levels10. Such a massive recall response suggests that humoral immunity was incomplete and not sterilizing.
Due to technical challenges and lack of reagents, the role of T cell responses has rarely been studied in NHPs, yet more extensively in mice. Intriguingly, while generally robust in mice, vaccine-induced T cell responses remained relatively weak in NHPs, hardly exceeding 100 Spot Forming Units/106 PBMC (∼0.01% of ZIKV E-specific T cells) by IFN-γ ELISpot9,10. Nevertheless, depletion and adoptive transfer experiments in mice suggest a complementary role of T cells in mediating vigorous and durable protection against ZIKV infection and disease17,18,19,20,21. Therefore, an ideal ZIKV vaccine should induce both strong humoral and cell-mediated immune (CMI) responses3,4,22,23.
Live-attenuated vaccines, such as the yellow fever vaccine YF17D, are known for their outstanding immunogenicity profile, durable protective immunity, and efficacy after single vaccination without the need for an adjuvant. With more than 1 billion administered doses, YF17D itself is recognized as one of the most effective human vaccines ever developed24,25,26. Consequently, there is a long history of developing live-attenuated chimeric YF17D-based Orthoflavivirus vaccines, exemplified by the licensed JEV (Imojev®) and DENV (Dengvaxia®) vaccines27,28. Generally, single-dose YF17D vaccination is efficient to induce vigorous innate, long-lasting humoral and balanced Th1/Th2 cellular responses that possibly warrant life-long protection in most vaccinees. Mechanistically, the early, broad, and unique transcriptional profile elicited by YF17D is considered critical to orchestrating a markedly polyvalent and durable immunity. Therefore, new vaccines recapitulating the favourable immunogenicity profile of YF17D are more likely to succeed in human clinical trials25,26,29,30,31,32,33.
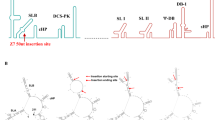
In response to the ZIKV outbreak of 2015/16, we developed a YF17D-based live-attenuated Zika vaccine candidate (YF-ZIK) by replacing the antigenic surface glycoproteins prM/E and the C anchor domain of YF17D with a prototypic Asian ZIKV isolate (Fig. 1a). YF-ZIK was shown to be highly attenuated and safe as demonstrated by smaller plaque phenotype, poor replication in mosquito cells, and no neurovirulence in suckling mice. Single-dose YF-ZIK immunization rapidly elicited protective immunity and conferred within 7–12 days vigorous protection against both lethal ZIKV infection and congenital malformations in mice22.
a Schematic of YF17D and YF17D-based Zika vaccine candidate YF-ZIK. The antigenic surface glycoproteins and the capsid anchor of YF17D were replaced with a prototypic Asian lineage Zika virus (ZIKV). C core, NS non-structural protein, UTR untranslated region. b Schematic of non-human primates (NHPs) vaccination and challenge study. Rhesus macaques (Macaca mulatta) were immunized s.c. twice (at day 0 and day 28) with 105 PFU of YF-ZIK (n = 6) or medium (sham; n = 4). At day 56, all macaques were i.d. infected with 105 PFU of ZIKV DR/2016/PD1 strain. Weight and temperature changes were monitored daily until day 70. Blood was sampled weekly for hematology and serology (NAb) analyses. Whole blood was collected for transcriptional analysis after prime (at day 0, 2, 7), boost (at day 28, 30, 35), and challenge (at day 56, 57, 60); and PBMCs to assess CMI responses by ICS (at day 42). Viremia and ZIKV NS1-specific antibody levels after challenge were quantified until day 70. Created in BioRender. Ma, J. (2025) https://BioRender.com/0jgftwr.
In this study, we demonstrate the safety, immunogenicity, and protective efficacy of YF-ZIK in a stringent non-human primate ZIKV challenge model. A single dose induces rapid seroconversion in macaques, generating high-titer ZIKV-specific NAbs that may confer sterilizing immunity, with responses further enhanced by a booster dose. Transcriptomic profiling reveals a balanced Th1/Th2 response and a signature closely resembling that of the parental YF17D vaccine, involving multiple pathways linked to broad antiviral immunity. Systems-level analysis identifies TNFRSF17 as a correlate of antibody responses, and GNAS and CD207 (Langerin) as predictive biomarkers of vaccine efficacy against high-dose viral challenge.
Results
Immunization and safety profile of YF-ZIK in macaques
NHPs exhibit key characteristics of ZIKV infection in humans, including rapid viremia, the presence of viral RNA in multiple organs and body fluids, and the development of NAbs and CMI responses; hence, NHPs represent a relevant model for the study of ZIKV pathogenesis and preclinical evaluation of potential interventions34,35. In a first step, we assessed the vaccine safety of our previously described chimeric yellow fever-Zika virus vaccine candidate YF-ZIK (Fig. 1a)22. To that end, six Orthoflavivirus-seronegative rhesus macaques (sex ratio 1:1) were immunized subcutaneously (s.c.) twice (at day 0 and day 28) with each 105 PFU of YF-ZIK, while four macaques received medium as sham controls (Fig. 1b). The vaccine was well tolerated as no signs of discomfort (normal behaviour and food intake), local reactogenicity (no swelling at the injection site or of draining lymph nodes), nor significant changes in weight and body temperature (Supplementary Fig. 1a, b) were observed in both groups. In addition, all hematological parameters tested during the immunization phase (blood cell counts and ratios, platelet counts, hemoglobin levels) remained within physiological ranges in both YF-ZIK-immunized and in sham-vaccinated animals (Supplementary Fig. 1c–l). Altogether, no adverse reactions were observed in rhesus macaques, confirming the previously observed favourable preclinical safety profile of YF-ZIK22,36.
YF-ZIK rapidly activates broad innate and adaptive transcriptional programs
As a benchmark of successful human viral vaccines, YF17D-induced immunity has been extensively studied. Data collectively suggest that YF17D induces a unique transcriptional master profile integrating multiple arms of the innate and adaptive immune system to develop broad, polyfunctional, and persistent immunity in humans24,26,30,31,32,33. However, despite serving as well-established viral vector, there is limited information about the molecular mechanisms by which YF17D-derived vaccines act and protect29,37. To fill this gap, we used nCounter digital transcriptomics for the profiling of whole blood samples collected both after prime (at day 2 and 7, as compared to day 0) and boost (at day 30 and 35, as compared to day 28) (Fig. 1b). More specifically, we analyzed the expression profile of >750 genes associated with vaccination (especially YF17D immunization in humans), immune response and viral infection26,30,31,32,33.
Our analysis revealed that YF-ZIK immunization rapidly induces a gene expression profile associated with activation of multiple pathways as early as 2 days after prime and then further pronounced and shaped after vaccine boost (Fig. 2a–e, see Source Data file for raw values and descriptive statistics).
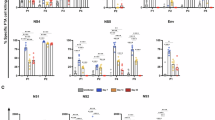
a, c Volcano plot of differentially expressed genes (DEGs) at day 2 post-prime (as compared with the baseline of day 0 pre-prime vaccination) (a) and day 2 post-boost vaccination (as compared with the baseline of day 28 pre-boost vaccination) (c). Log P-values are shown in y-axis; dotted lines represent cut-off values based on the adjusted P-value. P-values were calculated using a negative binomial generalized linear model (NanoString’s method) with Benjamini–Hochberg false discovery rate correction. b, d Heatmap of different immune pathways scores and hierarchical clustering (Euclidean distance metric and complete linkage agglomeration method) of different immune pathways in YF-ZIK immunized macaques over time after prime (b) and boost vaccination (d). e Representative immune pathways score in individual macaques after prime (n = 3) and after boost (n = 6). Symbols represent individual animals. Horizontal bars in post-prime subsets indicate the median; boxes in post-boost subsets represent the interquartile range (25th–75th percentile) with the median. Whiskers indicate the minimum and maximum values. Data were analyzed by one-way ANOVA with Tukey’s multiple comparisons test. f Pathway enrichment analysis at day 2 after prime (n = 3) and boost (n = 6). Line thickness represents the number of interacting genes between two nodes; node diameter indicates the number of genes involved.
More specifically, prime vaccination induced an early upregulation of CXCL10 (or IP-10; pro-inflammatory response to viral infection), EIF2AK2 (or PKR), OAS1, OAS2, MX1, MX2, IFIT2, STAT1, STAT2, IFIH1, IFI16, IFI44, and DDX58 (antiviral and IFN-related genes), CSF1R, CD163, CD68 and SIGLEC1 (related to monocytes/macrophages immunity), and CYBB and CD38 (related to adaptive immunity) within 2 days after immunization (Fig. 2a). Overall, YF-ZIK immunization in macaques appeared to induce an earlier peak in DEG (at day 2) than reported for YF17D immunization in humans, which peaks around day 7 after vaccination31. Upregulation of CXCL10 and OAS1 transcripts by YF-ZIK at day 2, genes commonly associated with successful immunization, was particularly fast, especially when contrasted with the delayed response observed with YF17D vaccination, where similar changes were not seen until day 21 (ref. 32). Downregulated DEG in the whole blood transcriptome included TLR9 (TLR signaling), TRAF1 and TRAF6 (related to antiviral immune responses), and CD59 (related to complement system) amongst others. A downregulation of TLR pathways and simultaneous upregulation of antiviral interferon signaling (Fig. 2b, e, Supplementary Fig. 2c, d, and Source Data) would align with RIG-I/DDX58 and MDA5/IFIH1 possibly acting as major cytosolic RNA sensors during early Orthoflavivirus infections. Likewise, a reduced enrichment of gene activation to adaptive immunity and T cell activation (LCP2 and KLRB1), FADD and CASP8 (related to cell apoptosis), GATA3 (related to Th2 immune responses) (Fig. 2b) may result from cell recruitment and migration from peripheral blood into draining lymph nodes in early response to YF-ZIK. The pronounced early transcriptional response was transient, with transcript levels and pathway signatures returning to baseline by day 7 post-vaccination, indicating rapid contraction of the immune response following primary immunization (Supplementary Fig. 2a–c).
A booster dose of YF-ZIK administered at day 28 activated a significantly stronger and broader transcriptional response compared to the primary immunization. This included enhanced activation of DEG clusters that were already upregulated by prime vaccination, as well as an additional gene set associated with further maturation and shaping of the immune response (Fig. 2c–e and Supplementary Fig. 2c). DEG included, among others, the upregulation of canonical antiviral and IFN-related genes (OAS1-3, PML, IRF5, MX1, MX2, IFITM1, ISG20, EIF2AK2, IFI16, STAT2, and SOCS1), genes related to monocytes/macrophages immunity (SIGLEC1 and CD68), to interleukin signaling (IL1RN), to TLR signaling (TLR7, ELK1, and MEF2A), and complement activation (SERPING1). Besides a strong innate antiviral signal, thus identified DEG included key genes involved in humoral responses and upregulation of T cell activity (MR1 and CD4), cell proliferation (TNFSF13B and BST2), and apoptosis (TNFSF10) (Fig. 2c), indicative for a strong adaptive recall response.
In summary, though often measured at low amplitude and in many cases with limited statistical power due to the small number of samples suitable for analysis, our whole blood transcriptomic analyses suggest a rapid, canonical IFN-centered innate antiviral response and an early, concerted induction of adaptive immunity after prime vaccination, which is further enhanced by a booster dose (Fig. 2f).
YF-ZIK induces polyfunctional immunity, including a strong NAbs response as major correlate of protection
Next, we assessed the immunogenicity of YF-ZIK, focusing on the induction of polyfunctional immunity and the generation of NAbs, a key correlate of protection. The immunogenicity was evaluated following a two-dose regimen (Fig. 1b) analogous to the schedule recommended for the chimeric YF17D-based JEV vaccine (Imojev®). Primary YF17D vaccination may result in transient, low-level viremia (typically 3–9 days; <104 genome RNA copies/mL) in both humans and NHP38,39,40,41. After YF-ZIK prime immunization, viremia remained very low, exceeding quantification limits occasionally in only one macaque (R10128) (Supplementary Fig. 3a, b). Though rare, the detection of viral RNA in the bloodstream proves that live-attenuated YF-ZIK is capable of active replication, amplification, and systemic spread. Nevertheless, 3 out of 6 YF-ZIK-immunized macaques (50%) developed ZIKV-specific NAbs as early as 7 days after prime vaccination, with a log10 geometric mean titre (GMT) of 2.0, including previously mentioned individual R10128 (NAb titre: 2.5 log10).
At day 14, all animals (6/6) had seroconverted with a peak in NAbs at a log10 GMT of 3.2 (Fig. 3a and Source Data file). NAbs were absent in the sham group. NAbs titres were determined by an in-house developed fluorescence-based reporter virus seroneutralization assay, which has been validated and shown to yield a comparable reliability and accuracy as a classical PRNT assay used as benchmark (Supplementary Fig. 3c). Notably, by day 28 post-prime vaccination, ZIKV-specific NAb titres (log10 GMT of 2.8) still exceeded established correlates of protection for several licensed Orthoflavivirus vaccines in humans (NAb titre > 1:10 or 1:40), as well as the more stringent threshold proposed for ZIKV (NAb titres > 1:25 to 1:100) derived from challenge studies in macaques9,10,11,42. This suggests that single-dose YF-ZIK vaccination might be sufficient for protection. As expected, a second YF-ZIK dose given at day 28 significantly boosted ZIKV-specific NAb levels, resulting in a marked 25-fold increase by day 35 (log10 GMT of 4.2) as compared to day 28 (prior to boost; log10 GMT of 2.8), persisting until day 56 (log10 GMT of 3.8) immediately before animals were challenged (Fig. 3a and Source Data file).
a Kinetics of ZIKV-specific SNT90 NAbs in macaques. LLOD, the lower limit of detection. b Total ZIKV-specific IgG, NAbs (left y-axis) and the respective ratio (right y-axis) at different timepoints after vaccination. c Kinetics of Activation-induced cytidine deaminase (AICDA) transcript levels in circulating PBMCs of YF-ZIK-immunized macaques. d, e Passive serum transfer study scheme (d) and survival rate after ZIKV MR766 challenge (e). AG129 mice received a single dose of 450 µl of NHPs serum pool from either YF-ZIK-immunized macaques or sham-immunized macaques by intraperitoneal (i.p) injection. Mice were challenged 24 h later by i.p. injection with 10 PFU of ZIKV MR766 (n = 9–10 from two independent experiments). Differences in Kaplan–Meier survival curves (e) were assessed using the two-sided Log-rank (Mantel-Cox) test. d was created in BioRender. Ma, J. (2025) https://BioRender.com/jebjnt0.
The early production of NAbs, which was further boosted in all animals within 7 days after the second dose, coincided with a rapid contraction of the immediate transcriptional response associated with IFN signaling and innate immune pathways (Fig. 2b, e and Supplementary Fig. 2a–d). This sequence of events following booster vaccination suggests a functional relationship between the resolution of an early innate immune activation in response to the live vaccine virus and a rapid onset of a previously primed adaptive humoral response shortly thereafter. Notably, the raise in ZIKV-specific IgG levels measured in parallel to NAbs displayed only slightly delayed kinetics, indicating an early antibody class switch (Fig. 3b). In support for this, we found that transcript levels of activation-induced cytidine deaminase (AICDA), encoding for a key enzyme required for antibody diversification via somatic hypermutation and class switching from IgM to IgG, increased markedly, peaking at day 7 after prime vaccination and again 2 days after the booster dose (Fig. 3c).
To determine whether vaccine-elicited NAbs are sufficient for protection, we performed adoptive serum transfer in AG129 mice, a very sensitive and stringent mouse model for ZIKV challenge and infection studies15,22,43. AG129 mice are highly susceptible to infection, with as little as 1 PFU of ZIKV (MR766 strain) being sufficient to induce disease and full mortality (5/5) (Supplementary Fig. 3d, e). For this purpose, serum pools from YF-ZIK-immunized macaques (from day 21, 35, and 42 with log10 GMT of 3.7), or sham-vaccinated macaques were used. Antiserum transfer was well tolerated (Fig. 3e and Supplementary Fig. 3f, g) and resulted in uniformly high serum concentrations of ZIKV-specific NAbs (log10 GMT of 3.2) immediately after dosing. Titres declined rapidly to undetectable levels by day 14, consistent with the short half-life of NHP antibodies in mice2,3,4,23 (Supplementary Fig. 3h). Upon challenge with 10 PFU (i.e., a more than tenfold lethal dose) of ZIKV one day after serum transfer, all mice that received the serum from sham-vaccinated macaques developed severe disease and succumbed to infection (Fig. 3d, e). In contrast, 7 out of 9 mice that received serum from YF-ZIK-immunized macaques remained healthy and were fully protected, as confirmed by the absence of weight loss, clinical signs of disease, or mortality following MR766 challenge (Fig. 3e and Supplementary Fig. 3g). These data confirm the critical role of NAbs in conferring protection against lethal ZIKV infection. However, additional immune mechanisms triggered by YF-ZIK vaccination are be expected to contribute to vaccine efficacy2,3,4,23.
Systems vaccinology analyses suggest mechanistic similarities with YF17D vaccine
Several clinical trials have demonstrated the great complexity of molecular and cellular events that are triggered by the prototypic YF17D vaccine to induce long-lasting polyfunctional immunity. Thereby, the mounting of vigorous adaptive responses and immunological memory has been linked to a highly predictive transcriptional pattern early after vaccination within specific cell compartments26,30,31,32,33. To validate and translate our NHP data to the human model, we compared the gene expression profile triggered by YF-ZIK vaccination of macaques with what can be expected for YF17D vaccination following meta-analysis of available human data. Using UMAP analysis, we first plotted the YF-ZIK and YF17D specific expression profiles to the human blood transcriptome44 to demonstrate that both vaccines affected multiple leukocyte populations (amongst others, dendritic cells, macrophages and neutrophils; naive and memory B cell, plasmablasts and plasma cells; CD4+ T, and CD8+ T and NK T cells), following a comparable pattern (Fig. 4a and Supplementary Fig. 4a).
a Uniform Manifold Approximation and Projection (UMAP) representation of blood cells extracted from the Tabula Sapiens database. Left panel, cells colored by cell type; middle and right panels, the color gradient represents the normalized score of the gene set comprising upregulated genes 2 days after prime vaccination with YF-ZIK and YF17D. b Correlation of differentially expressed genes at day 2 after YF17D vaccination, and 30 days after vaccination with YF-ZIK. c Venn diagram showing overlap between the number of differentially expressed genes by YF17D vaccination in humans and YF-ZIK vaccination in macaques, at day 2 (upper panel) and day 30 (lower panel). Enrichment P-values were calculated using a one-sided hypergeometric test.
For quantitative comparison, publicly available early expressions data for YF17D (prime only: day 3 and day 7) were plotted over those gathered in this study for early time points after YF-ZIK administration (i.e., day 2 post-prime and day 2 post-boost) (Fig. 4b). The thus aligned DEGs revealed a strong overlap for both vaccines after prime (at day 2; 39 shared genes, enrichment p = 0.0013) and boost (at day 30; 48 shared genes, enrichment p < 10−6) (Fig. 4b, c). This strong correlation does not appear to be markedly affected by differences in vaccine doses and experimental schedules, or their possible impact on DEG expression kinetics. By contrast, YF-ZIK and YF17D shared an upregulation of numerous canonical antiviral and IFN-related genes (e.g., EIF2AK2, OAS1, MX1, STAT1/2, ISG20, DDX58) (Fig. 4c and Source Data), genes related to T cell immunity, to TNF and chemokine signaling and to the involvement of macrophages and complement. However, a most striking overlap was observed for genes associated with humoral immunity, BCR signaling, and B cell proliferation (see also Fig. 7d–f below)26,31. An intriguingly similar yet enhanced transcriptional response triggered by a second dose of YF-ZIK shows that booster immunization remains feasible and may be beneficial to further increase ZIKV-specific NAb levels (Fig. 3a). Overall, our systems vaccinology analyses suggest a high similarity in the molecular mechanism of both vaccines, as well as the validity and translational value of data obtained in the macaque model.
Besides obvious commonalities, the comparison of YF-ZIK and YF17D also unveiled unique DEG sets. At day 2, YF-ZIK exclusively upregulated 26 genes that were not altered by YF17D at any time point. These included several IFN-stimulated genes such as SIGLEC1, IRF5, TMEM173 (the gene encoding IFN activator STING), HLA-DRA, and HLA-DQA1. Likewise, YF17D exclusively upregulated 45 genes, including several IFN-stimulated genes. Of interest, these genes were also upregulated by infection with unrelated viral pathogens such as SARS-CoV-2 in rhesus macaques and humans, as well as in multiple other animal species (ferrets, hamsters, and mice; Fig. 4c and Source Data). It is hence tempting to speculate whether the favorable safety profile of YF-ZIK (Supplementary Fig. 1), observed as a reduced neurovirulence in suckling mice and no lethality for AG129 mice in contrast to parental YF17D22,36,43, might be linked to the absence of any of these transcripts. The 16 genes exclusively downregulated by YF17D were not significantly enriched in any particular pathway.
Finally, for four genes, the two vaccines induced opposing expression patterns. SOCS1 (Suppressor Of Cytokine Signaling 1, an inhibitor of type I and II IFN signaling) and TICAM1 (TIR Domain Containing Adaptor Molecule 1, a TLR signaling adapter protein) (highlighted in Fig. 4b), and anti-apoptotic BLC2L1 (Bcl-2-Like Protein 1) were upregulated by YF17D and downregulated by YF-ZIK, while PTPRC (encoding pan-leukocyte marker CD45) was downregulated by YF17D and upregulated by YF-ZIK. An elevated CD45 and diminished BLC2L1 expression 2 days after YF-ZIK vaccination are in agreement with a simultaneous increase in monocytes/macrophages and a decrease in apoptosis (Fig. 2 and Supplementary Fig. 2), features not shared with YF17D26,32. Overall, however, transcriptomics analyses unveiled marked similarities rather than differences between chimeric YF-ZIK and its parent YF17D.
YF-ZIK immunization protects macaques against ZIKV infection
The magnitude and quality of YF-ZIK-induced humoral and cellular responses supported the hypothesis that vaccinated animals would be protected against ZIKV infection. To test this, we performed an experimental challenge, whereby all animals were intradermally (i.d.) injected with a high dose (105 PFU) of ZIKV DR/2016/PD1 strain eight weeks after priming (day 56, Fig. 1b). Consistent with previous reports34,35, ZIKV infection resulted in a high, protracted blood plasma viremia in the sham-immunized macaques (Fig. 5a and Supplementary Fig. 5a). The viremia rapidly increased with viral RNA detectable in 3/4 animals within 24 h after inoculation and peaked around 4 days post-challenge (log10 GMT of 5.7 viral genome copies, 95% CI 3.9–7.5). As expected for rhesus macaques, infection was mild (no signs of weight loss, nor fever) and self-limiting (Supplementary Fig. 5). Following induction of an original antiviral IFN-related and adaptive response (Supplementary Fig. 5a, b), viral clearance from plasma occurred within 10 days post-infection (Fig. 5a and Supplementary Fig. 5a), concomitant with seroconversion to ZIKV-specific NAb and anti-NS1 antibodies (Fig. 5b, c).
a Blood plasma ZIKV RNA kinetics after challenge of Sham (n = 4) or YF-ZIK (n = 6) immunized macaques. Circles represent individual animals; solid lines with shaded area, mean ± SEM. b ZIKV NS1-specific antibody levels in the blood plasma after ZIKV challenge. Data presented as median ± IQR. Data analysed by two-way variance (ANOVA) with Tukey multi-comparison tests. c ZIKV-specific SNT50 NAbs in Sham and YF-ZIK immunized macaques after ZIKV challenge. Colour-coded lines represent individual animals. LLOD, lower limit of detection.
In contrast, YF-ZIK immunized macaques were protected against vigorous ZIKV challenge, as demonstrated by the absence of any detectable viral RNA in the blood plasma after challenge, except for a singular transient event in one individual (R13010), yet markedly lower and delayed in onset by about 5 days compared to sham-vaccinated animals (Fig. 5a and Supplementary Fig. 5a). Induction of ZIKV-specific anti-NS1 antibodies was assessed as a sensitive biomarker of ZIKV exposure. Compared to the high levels observed in sham-vaccinated macaques at day 70 (14 days after challenge), only a minimal increase of ZIKV NS1-specific IgG was detectable in YF-ZIK-immunized animals (Fig. 5b). Notably, no substantial increase in ZIKV-specific NAb could be observed after challenge in YF-ZIK-immunized macaques, suggesting that vaccination had already elicited saturating antibody levels, resulting in almost sterilizing immunity (Fig. 5c). Of note, the vaccine candidate proved safe as no enhancement of disease nor virus replication by pre-existing immunity was observed in vaccinated animals (Supplementary Fig. 5b, c). In summary, YF-ZIK immunization protected macaques against vigorous high-dose i.d. ZIKV infection.
Rapid recall of polyfunctional immune responses in YF-ZIK immunized macaques contribute to protection
To explore the molecular and cellular underpinnings of such a high degree of protection, we extend our transcriptional analysis to days 1 and 4 after challenge, compared to pre-challenge (day 56 after priming). The initial response to ZIKV challenge in sham-vaccinated macaques remained limited, both regarding the number and amplitude of affected DEGs at day 57 (1 day after infection) without statistical significance (Supplementary Fig. 6; see Source Data file for raw values and descriptive statistics). Likewise, a slight yet marked upregulation of genes related to IFNs (including OASL, OAS1, OAS2, OAS3, MX1, MX2, DDX58, IRF7, IFIT1, IFIT2, IFIT3, IFI44, IFN 3), metabolism (SLC2A1, CHIT1, ALOX15), host defense against infections and adaptive immunity (LTF, DEFA1, ICAM4) appeared delayed at day 60 (4 days after challenge) (Fig. 6a and Supplementary Fig. 6), in line with the capacity of ZIKV to antagonize antiviral host responses2,3,4,23.
Volcano plot of DEGs at day four post-challenge (as compared with the baseline of day 0 before challenge) in sham (n = 4) (a) and YF-ZIK (n = 6) (b) immunized macaques. Log p-values are shown in y-axis; dotted lines represent cut-off values based on the adjusted p-value. P-values were calculated using a negative binomial generalized linear model (NanoString’s method) with Benjamini–Hochberg false discovery rate correction. c Heatmap of different immune pathways score and hierarchical clustering (Euclidean distance metric and complete linkage agglomeration method) of different immune pathways after challenge. d Pathway enrichment analysis at day 4 post-challenge. Line thickness represents the number of interacting genes between two nodes; node diameter indicates the number of genes involved.
In contrast, YF-ZIK immunized macaques showed a stronger, earlier, and broader transcriptional signature compared to the sham group; especially pronounced at day 60 (4 days after challenge; Fig. 6b and Supplementary Fig. 6; see Source Data file for raw values and descriptive statistics). While largely limited to a primary IFN response in sham animals, YF-ZIK immunized macaques showed a strong upregulation of a large number of genes related to the (re)activation of both the innate and adaptive arms of the immune system, suggestive for early involvement in vaccine-induced protection. These included the activation and modulation of TLR-mediated, interleukin-, Fc Receptor-, MAPK-, VEGF-, NF-kB-induced immune responses (such as IRAK1, MYD88, TRAF6, MAVS, TNFRSF14, IKBKE, etc.), macrophage involvement (SIGLEC1, CD68, and CSF3R), complement (SERPING1 and C1R), antigen presentation (HLA-DMB), T cell function (CXCR3, CXCR,4 and CD4), and metabolism (CHIT1, PLA2G2A, and ALOX15) (Fig. 6b, c and Supplementary Fig. 6b, c). Collectively, this fast switch in the expression of multiple DEGs from several interlinked antiviral pathways upon virus exposure (Fig. 6d and Supplementary Fig. 6d) lays a molecular basis for vaccine-induced protection and rapid control of ZIKV infection in YF-ZIK immunized macaques.
Vaccine-induced transcriptional patterns predict immunity and clinical outcome
Finally, we used linear and logistic regression models and machine learning algorithms to identify gene signatures that could serve as predictors for successful vaccination and protective immunity against ZIKV infection in macaques. Herein, several biomarkers routinely measured in human cohort studies were considered, namely peak viral load (at day 60), overall viral load (AUC, area under the curve across all time points), and anti-NS1 antibody levels (as a measure of exposure to replicating ZIKV). Due to the limited number of samples available at day 0 and day 3, predictions were performed from digital transcriptomic profiles obtained at day 7 and day 28 (after prime), as well as day 30 and day 35 (after boost). After multiple uni- and multivariate regression models were tested, best-fitting univariate predictors for NAb levels and clinical outcomes were identified by linear regression. Among all time points, gene expression profiles from day 30 (2 days after boost) generally provided the best predictive models. Demographic variables (age, sex, and weight of animals) were not further considered as they did not lead to any better predictions despite correlation to some serological and transcriptional parameters (Fig. 7 and Source Data).
a, b Top 50 most predictive genes (quantified by digital transcriptomics at day 30) for neutralizing antibody levels at day 56 (before challenge) form a densely linked network centered around B cell activation (a). Color codes show significantly enriched pathways and protein networks identified by STRING analysis (b). c Kinetics of B Cell Activating Factor (BAFF, also known as BLyS, encoded by TNFSF13B), and its receptor (B Cell Maturation Antigen, BCMA, encoded by TNFRSF17) after vaccination. Transcript quantification was performed by digital transcriptomics. Multiple comparisons were analyzed using a mixed-effects model with Restricted Maximum Likelihood (REML) estimation; all tests were two-sided. d, e Univariate linear regression and ranking (attribute selection by machine learning) of top 20 most predictive genes for NAbs at day 56 after vaccination (n = 6, d) and for peak viral load in plasma at day 60 (n = 10, e). Individual data points represent independent animals. The solid line indicates the mean of the simple linear regression; the shaded area representing the 95% confidence interval. The goodness of fit (R) and corresponding p-values are shown in each panel. f Heatmap of Spearman’s correlations and hierarchical clustering (Euclidean distance metric and complete linkage agglomeration method) of selected parameters, including immune pathway scores, transcript levels, viral load, and frequency of cytokine-producing T cells. The bar on the right indicates the scale of the correlation values.
The top 50 most predictive DEGs (quantified at day 30) for NAb levels at day 56 (prior to challenge) form a densely linked protein-protein interaction network centered around B cell activation including the highly connected canonical B cell marker CD19, B cell receptor-associated CD79A/CD79B and MS4A1 (CD20), B cell activation markers CD40 and CD86, as well as several B cell specific members from the TNF receptor superfamily, such as the two bona fide receptors for TNFSF13B/BAFF (B cell activating factor) called TNFRSF13C (BAFFR) and TNFRSF17 (B cell maturation antigen, BCMA) (Fig. 7a, b).
As shown in Fig. 7c, TNFSF13B/BAFF transcript levels were significantly increased after both prime and boost, which paralleled the previously observed immune pathway activation (Fig. 7a, b) and NAb kinetics (Fig. 3a), as well as an increased expression of its receptor TNFRSF17, reaching a plateau starting from day 7 after vaccination, like human YF17D data. In addition, TNFRSF17, together with FASLG (Fas Ligand), appeared to be the most predictive DEG for NAb production after YF-ZIK immunization in macaques (Fig. 7d). Notably, TNFRSF17 turned similarly out as the most accurate predictor for the YF17D-induced NAb response in humans with up to 100% accuracy26.
ZIKV-specific NAb alone may not be sufficient to confer full protection. Our transcriptional analysis revealed another two transcripts in peripheral blood to most accurately predict viral load at day 60, the peak of ZIKV infection in macaques, and overall viremia (AUC), respectively, being CD207 (Langerin), present on circulating conventional dendritic cells, and GNAS, a pleiotropic gene involved in anti-inflammatory cytokines production and GPCR signaling. Thereby, low expression of either DEG prior to ZIKV infection correlated with protection from virus replication (Fig. 7e and Supplementary Fig. 7a). High viral loads were similarly associated with an exhausted CD8+ T cell profile and high LAIR1 (inhibitory leukocyte-associated Ig-like receptor, CD305) expression (Fig. 7e and Supplementary Fig. 7a), further implicating CD8+ T cells as relevant mediators of protective immunity against ZIKV infection3,4,17,23,42,45.
ZIKV-specific NS1 antibody levels (at day 70) correlated strongly with peak viral loads (at day 60) (Fig. 5a, b). Seroconversion to anti-NS1 may hence serve as a sensitive marker for exposure to replicating ZIKV. Consequently, low transcript levels for CCL5/RANTES (leucocyte migration) and PDCD1 (encoding PD-1 indicative for inactive lymphocytes) in the blood prior to challenge translated into a barely detectable anti-ZIKV NS1 IgG response (Supplementary Fig. 7b) and, thus, protection in macaques imprinted by YF-ZIK vaccination. In contrast, extensive virus replication observed in sham-vaccinated animals may reflect delayed tissue recruitment and slow responsiveness of T cells in naïve subjects during an acute ZIKV infection.
Multiple studies highlight the importance of ZIKV-specific CMI responses for protective immunity2,3,4,17,23,42. Induction of a balanced (Th1/Th2/Treg) and polyfunctional (CD4+/CD8+) ZIKV-specific antiviral CMI response by YF-ZIK is corroborated by a comprehensive DEG and correlation analysis of transcriptional, virological, serological and cellular parameters comparing YF-ZIK and sham-vaccinated animals (summarized in Fig. 7f). For additional experimental evidence, intracellular cytokine staining (ICS) was performed on PBMCs collected at day 42; i.e., 14 days after administration of the booster dose (Fig. 1b). Upon stimulation with ZIKV E protein-derived peptides, ZIKV E-specific CD4+ T cells producing IFN-γ, IL-2 (promoting CMI responses; Th1) or IL-4 (promoting humoral responses; Th2), as well as CD8+ T cells producing IFN-γ or IL-2 could be identified in YF-ZIK immunized macaques (Supplementary Fig. 7d). Restricted in scope and depth by technical issues and by a limited number of specimens, our ICS analysis remains preliminary and merely suggestive (no statistical significance). Nevertheless, based on our experimental evidence, a balanced Th1/Th2 cellular response can be expected to complement humoral immunity through the antiviral activity of cytotoxic T cells and the provision of T cell help, thereby enhancing the quality and longevity of YF-ZIK-induced protection.
Taken together, protective immunity triggered by YF-ZIK appears to extend beyond a certain NAb level serving as proxy and serological correlate for successful immunization. We further demonstrate the translational relevance of our NHP data to humans and identified TNFRSF17 as a common predictor for NAbs for both YF17D and YF-ZIK. Several factors in peripheral blood such as CD207, GNAS, LIAR1, PDCD1, and CCL5 are inversely correlated with a protective cellular immune response and forecast an adverse infection outcome.
Discussion
In this study, we assessed the safety, immunogenicity, and efficacy of our chimeric live-attenuated Zika vaccine candidate, YF-ZIK, in rhesus macaques. Safety of YF-ZIK had been demonstrated before, amongst others, by lack of neurovirulence in suckling mice22,43. Here, we show that YF-ZIK is also well tolerated in NHPs, further supporting its excellent preclinical safety profile (Supplementary Fig. 1).
In mice, single-dose YF-ZIK immunization elicited strong antibody and polyfunctional T cell responses, which conferred full protection against ZIKV infection and congenital malformations22. In macaques, YF-ZIK immunization also induces rapid (detectable as early as 7 days after prime) and strong antibody responses (Fig. 3a), and ZIKV E-specific CD4+ (with balanced Th1/Th2 cytokines production) and CD8+ T cell responses (Supplementary Fig. 7d). Importantly, in YF-ZIK immunized macaques, ZIKV-specific NAbs (log10 GMT of 2.8) are at day 28 already excessively higher than what is considered protective against ZIKV infection in macaques and humans (log10 GMT of 1.4–2.0)3,4,9,46,47, even without a booster, suggesting that a single YF-ZIK vaccination might be sufficient for protection (Fig. 3a). Though tempting, our 2-dose study design (Fig. 1) does not allow to prove this claim. Given that ZIKV is mainly endemic in remote areas in resource-limited and less-developed countries, implementation of such a Zika vaccine with single-dose efficacy would facilitate population-wide immunization.
A recent study defines a markedly higher log10 GMT of 3.8 as seroprotective antibody level48. However, in that study, inactivated whole ZIKV was used as vaccine antigen, that may lack key features that contribute to the overall immunogenicity and efficacy of live-attenuated vaccines. While we confirmed by passive serum transfer that ZIKV-specific NAb can indeed serve as major immunological correlate of protection, other arms of the immune system might complement NAb to confer broad and lasting protection (Fig. 3d, e)2,3,4,23. Previous investigations have thus shown that CD4+ T cells are critical in mediating the magnitude, quality, and longevity of ZIKV-specific humoral responses. Further, both CD4+ and CD8+ T cells can directly be involved in controlling and eliminating active ZIKV replication17,18,20,23,42,49. Therefore, the balanced T cell responses induced by YF-ZIK (Supplementary Fig. 7d) can be expected to form a second line of defence against ZIKV infection.
YF17D has been widely used as benchmark and model of a successful human vaccine to understand human immune responses to vaccinations24,26,30,31,32,33, as well as a vector for the development of novel live-attenuated vaccines22,29,37,43,50,51,52. Nevertheless, despite remarkable safety, immunogenicity, and efficacy in both animal models and humans, the underlying immune mechanisms of YF17D-based vaccines remain only partially understood. However, a unique and early YF17D-induced DEG pattern is believed to be critical in orchestrating its polyfunctional and long-term immunogenicity in humans26,30,31,32,33. Here, we sought to understand the strong immunogenicity and high degree of protection induced by our YF17D-based live ZIKV vaccine by analysing vaccine-induced early transcriptional profiles in NHPs.
Our results demonstrated that, like YF17D, YF-ZIK triggered an immediate IFN-centered antiviral gene expression response that favors and activates multiple downstream pathways (Fig. 2). We found that YF-ZIK and YF17D share highly overlapping expression profiles, particularly pronounced after boost (Fig. 4), certainly shaping the YF-ZIK-induced immunity. Interestingly, though high titres of ZIKV-specific NAbs have been rapidly induced in macaques, we found limited gene expression and immune pathways activation after prime (Fig. 2a, c and Supplementary Fig. 2a, b). This is consistent with recent work showing how YF17D distinctly downregulates most immune pathways one day after immunization, whereas they are commonly upregulated in responses to other vaccines32. Furthermore, we observed richer and stronger DEGs expression, and stronger and broader immune pathway activation after YF-ZIK boost as compared to prime, suggesting benefits from YF-ZIK boost (Fig. 2a–d and Supplementary Fig. 2a, b).
Some unexplained quantitative differences remain when comparing the somewhat delayed antiviral and IFN responses that are observed in humans by day 7 after YF17D immunization. We observed transient activation of IFN signaling as early as 2 days after YF-ZIK prime and boost, which is earlier than in YF17D immunization in humans, peaking at day 7 (Fig. 2b, d). YFV and other orthoflaviviruses antagonize host IFN signaling to establish infection. This may cause a delay in the activation of the innate antiviral program to live YF17D immunization53,54. Next to possible technical issues (different doses and immunization schedules used), a relatively early YF-ZIK-induced IFN activation might be explained by (1) host-specific differences between NHPs and humans, (2) a change in the tropism of the recombinant YF17D-derived vaccine vector, whereby (3) attenuated YF-ZIK may have lost certain antagonism for IFN signalling, possibly favouring vaccine safety55,56.
YF-ZIK prime and boost showed the same overall DEG patterns in all macaques, yet different in their magnitude. Transient type I IFN-centered responses dominated both YF-ZIK prime and boost (Fig. 2a–d and Supplementary Fig. 2c, d). Such early IFN responses have been seen with several human vaccines, including YF17D, vesicular stomatitis virus-based Ebola and influenza vaccines; essentially linked to the induction of early antibody and T cell activations3,25,31,32,33. Though initial immune activation after YF-ZIK prime is substantially weaker than after homologous boost (Fig. 2a–d), multiple human studies have consistently demonstrated that also YF17D boost does not provide additional benefits if original vaccination, confirmed by the presence of NAbs remains effective57,58.
ZIKV is naturally transmitted by infected mosquitos that inoculate an estimated 250 PFU59. In our study, we use an about 40-fold higher dose of 105 PFU for vigorous intradermal challenge. YF-ZIK immunization hence protects against stringent, high-dose experimental ZIKV infection, as confirmed by the absence of viremia and ZIKV NS1-specific antibodies in plasma of YF-ZIK immunized macaques (Fig. 5a and Supplementary Fig. 5a). In addition, YF-ZIK elicited almost sterilizing immunity with saturated antibody titres. This is remarkable considering that earlier seminal NHP studies used a 100-fold lower dose (103 PFU) for challenge9,10,11,12. Yet even under these relaxed conditions, the thus tested vaccine candidates failed to fully contain ZIKV replication as obvious from a marked increase in NAb levels post-challenge by up two orders of magnitude10. Notably, full protection is thought to be important to prevent viremia and congenital abnormalities during pregnancy60,61, as supported by evidence from NHP models of gestational ZIKV infection62,63.
Further transcriptional analysis of serial blood samples after ZIKV challenge demonstrated that, different from the limited and delayed primary IFN response in the sham group, a rapid and markedly stronger and richer immune (re-) activation contributes to rapid control of infection in YF-ZIK immunized macaques (Fig. 6a–d and Supplementary Fig. 6a, b). Our transcriptional analysis largely expands our understanding of the superior immunogenicity and efficacy of YF17D-vectored vaccines beyond protective NAbs. This can be informative to develop next-generation YF17D-based vaccines29,37.
Finally, we found that the DEG patterns triggered by YF-ZIK in macaques significantly overlapped with YF17D immunization in humans (Fig. 4a–c). TNFRSF17 essentially involved in B cell development, was identified as the most reliable predictor for good NAb responses for both YF-ZIK and YF17D (Fig. 7c)26,64. It can hence be speculated to what extent TNFRSF17 might serve as a common marker for other YF17D-based vaccines as well. In addition, we identified several new transcriptional biomarkers defining clinical outcomes, such CD207 (the gene encoding Langerin), a marker of skin Langerhans cells but to our knowledge not previously described as a vaccine response marker in blood leukocytes, GNAS (Guanine Nucleotide-Binding Protein G(S) Subunit Alpha Isoforms Short), and LAIR1 (Leukocyte Associated Immunoglobulin Like Receptor 1). Of note, LAIR1 expression was identified as the best predictor of type I IFN-NAbs in severe/critical COVID-1965,66 (Fig. 7d, e).
Limitations of the study
We present a comprehensive mechanistic study assessing the molecular and cellular basis of the immunogenicity and protective efficacy of a live-attenuated ZIKV vaccine candidate in macaques as step-up translational animal model. Despite clear virological and serological evidence for protection against vigorous experimental ZIKV infection, several open questions remain. Firstly, our attempts to demonstrate ZIKV-specific antiviral T cell immunity of marked magnitude and broad functionality (Supplementary Fig. 7d) remains limited and may demand confirmation in larger cohorts (n > 3) with refined methodologies (e.g., multi-parameter cytometry; in vivo depletion of specific T cell subsets, etc.). In any case, our longitudinal transcriptomics analysis of NHP over the entire 10-week course of vaccination (prime and boost) and ZIKV infection is unique and in full support of our preliminary evidence for the YF-ZIK induced CMI and its contribution to ZIKV protection (Fig. 7f). Second, the extraordinarily high NAb titre induced by YF-ZIK9,10,11,12 and their potency to protect from lethal challenge in a stringent mouse model15,22,43 (Fig. 3e) may further obscure any dependency on CMI. At least our methodology to quantify NAb was validated by comparison with gold standard PRNT (Supplementary Fig. 3c). In general, the contribution of CMI for Orthoflavivirus immunity remains underappreciated2,3,4,17,23,42. Third, though NS1 has also been discussed as protective Orthoflavivirus antigen67, current YF-ZIK construct expresses YFV NS122, which is unlikely to confer cross-protection for ZIKV. However, the antigenic distance between NS1 from different virus species allowed to employ seroconvertion to ZIKV NS1-specific IgG antibodies as sensitive diagnostic marker for protection. Fourth, despite sharing many commonalities with parental live YF17D (Fig. 4), vaccine viremia, which has been linked to YF17D potency38 remained mostly below detection limits (Supplementary Fig. 3b). On the other hand, elevated and prolonged viremia is frequently seen as a safety concern68. In line, YF-ZIK was well tolerated in AG129 mice22,36 and NHP (Supplementary Fig. 1). Nevertheless, direct quantitative comparison of YF-ZIK responses in our non-human primate study with data from human trials of the parental YF17D vaccine should be interpreted with caution. We employed a fixed, relatively high dose of 10⁵ PFU for vaccination and intermittent blood sampling, which provides only a preliminary view of pharmacokinetic and pharmacodynamic parameters such as the magnitude and duration of viremia, or transcriptional kinetics. Intriguingly, other YF17D-based vaccines such as licensed JEV vaccine Imojev®69,70 and the clinically validated WNV candidate ChimeriVaxWN0271,72,73,74 have demonstrated consistent safety, immunogenicity, and efficacy in both NHPs and humans across a wide dose range (10²–10⁵ PFU). Similarly, recent trials assessing fractional dosing of the four WHO-prequalified YF17D vaccines reported no significant differences in seroprotection efficacy, despite spanning a dose range of approximately 102.7–104.6 IU75,76 (i.e., ~103.7–105.6 PFU)77,78. Thus, while caution is warranted, a qualitative comparison with YF17D human trial data may nonetheless be justified. Finally, prior ZIKV infection, like prior DENV infection, is associated with an increased risk of disease with certain DENV serotypes79,80. In principle, cross-reactivity among orthoflaviviruses should be considered when assessing vaccine safety and efficacy. Recently, new promising DENV vaccines have been introduced81,82 with no evidence yet for any negative impact on DENV or ZIKV morbidity83. Implementation of vaccination strategies that deploy a combination of both DENV and ZIKV vaccines into future routine immunization may mitigate the remaining risk by conferring global seroprotection against the entire virus complex84.
Several Zika vaccine candidates have been developed and entered human clinical trials, but none has acquired market authorization yet2,3,4,22,23. The promising preclinical safety profile, recently established means of high-yield manufacturing36,85, rapid induction of immunity, and possibility to protect as single-dose vaccine, makes YF-ZIK comparable with, if not superior to, other vaccine candidates. Our systems vaccinology analysis combines comprehensive serological, cellular, and transcriptional data from a vaccine-challenge study in NHPs to provide in-depth insight into the performance of YF-ZIK. Moreover, it provides a publicly available multiparametric data set to explore the molecular and cellular mechanism of protection conferred by YF17D-based vaccines, identifying several candidate biomarkers that predict vaccine immunogenicity and efficacy. It is tempting to speculate that these markers could be applicable to other live-attenuated viral vaccines or be implemented in future human clinical trials.
Methods
Cell culture and virus
Vero E6 (African green monkey kidney) and BHK-21J (baby hamster kidney) cells were provided by Dr. P. Bredenbeek and maintained in minimum essential medium (Gibco). Cells were maintained in Dulbecco’s modified Eagle medium (Gibco). All media were supplemented with 10% fetal bovine serum (FBS, Hyclone), 2 mM L-glutamine (Gibco), and 1% sodium bicarbonate (Gibco).
YF-ZIK vaccine stock, earlier called YF-ZIKprM/E, was generated as supernatant of infected Vero E6 cells22. ZIKV PD1 strain (ZIKV DR/2016/PD1) was kindly provided by Dr. L. Barzon, University of Padua, Italy, which was isolated from a traveler returning sick from the Dominican Republic86. ZIKV MR766 strain was obtained from the European Virus Archive (EVA) (Ref-SKU: 001v-EVA143). Virus titres expressed as plaque-forming units (PFU/mL) were determined as described before22. In brief, Vero E6 cells (1 × 10⁶/well) were seeded in 6-well plates overnight. After PBS washes, cells were incubated with serial virus dilutions for 1 h at 37 °C. Post-infection, cells were washed and overlaid with 0.5% low melting agarose (Invitrogen) in Minimal Essential medium (MEM) (Gibco) with 2% FBS. After agar solidification, plates were incubated for 7 days at 37 °C, then fixed with 4% formaldehyde and stained with 1% crystal violet. Plaques were counted manually and expressed as PFU/mL.
NHP study
All housing and animal procedures involving NHP took place at the BPRC, upon positive advice by the independent ethics committee (DEC-BPRC), under project license AVD502002016496 issued by the Dutch Central Committee for Animal Experiments and following approval of the detailed study protocol by the institutional animal welfare body. The NHP study was conducted on rhesus macaques (Macaca mulatta; 7 males, 3 females, approximately 4 years old with a body weight of at least 6 kg). All animals were pre-screened as negative for IgG antibodies against other related orthoflaviviruses (including ZIKV, DENV, JEV, and YFV) by ELISA. A group of six macaques was immunized twice in BSL3 facilities at Biomedical Primate Research Centre (BPRC, Netherlands) by subcutaneous injection of 105 PFU of YF-ZIK at day 0 and day 28, respectively. Another group of four macaques received MEM containing 2% FBS (referred to as sham-vaccinated) subcutaneously at the matched time point. Animals were randomly allocated to the experimental groups. All animals were monitored daily for clinical signs, appearance of injection site, and swelling of local draining lymph nodes by palpation. Blood draws, body weight, and temperature measurements were performed every 2 to 3 days during the first week after every vaccination and then once a week up to day 56. At day 56, all macaques were intradermally challenged with 105 PFU of ZIKV PD1 strain. After the challenge, macaques were followed daily for clinical signs including body temperature, body weight, food intake, dehydration, and abnormal behaviors. Blood draws were performed daily until day 70.
Hematology test
Hematology parameters were measured by the clinical laboratory at the BPRC with a Sysmex XT-2000iV Automated Hematology Analyzer (Sysmex Nederland B.V., Etten-Leur, The Netherlands).
Serum neutralizing test (SNT)
Quantification of NAb by SNT using ZIKV-mCherry reporter viruses has been described previously in detail52,87. In brief, serum dilutions were incubated with 10 TCID50 ZIKV–mCherry for 1 h at 37 °C, after which serum–virus complexes were transferred for 72 h to 104 BHK-21J cells. The percentage of mCherry-expressing cells was quantified on a Cell Insight CX5/7 High Content Screening platform (Thermo Fischer Scientific) to determine neutralization titres, including half-maximal (SNT50) and 90% (SNT90) inhibitory concentration. Neutralization values were calculated by normalizing to virus-only (100%) and cell-only (0%) controls, followed by curve fitting in GraphPad Prism. Our SNT assay has been validated and showed strong correlation with the gold standard plaque reduction neutralization test (PRNT) (Supplementary Fig. 3c).
Flow cytometry and intracellular staining (ICS)
Cryopreserved peripheral blood mononuclear cells (PBMCs) were thawed at 37 °C and 1 × 106 cells were cultured in RPMI 1640 with L-glutamine (Lonza), supplemented with 10% fetal calf serum (FCS), penicillin (100 units/ml; Continental Pharma), and streptomycin (100 μg/ml; Continental Pharma). Cells were stimulated with interleukin-2 (IL-2) (50 ng/ml; PeproTech), anti-human CD28 (Clone CD28.2, 1 μg/ml; BD Biosciences), and anti-human CD49d (Clone R1-2, 1 μg/ml; BD Biosciences) for 72 h at 37 °C/5% CO2. To measure vaccine-specific T cell responses, cells were either or not stimulated with PepMix ZIKV E protein peptide (PM-ZIKV-E, 5 μM; JPT Peptide Technologies). As a positive control, cells were stimulated with PMA (50 ng/ml; Sigma-Aldrich) and ionomycin (500 ng/ml; Sigma-Aldrich). Four hours prior to staining, cells were incubated with Brefeldin A (GolgiPlug, 1:1000; BD Biosciences) and Monensin (GolgiStop, 1:1500; BD Biosciences). Subsequently, cells were incubated with FcR-blocking reagent (0.5 μl/well; Miltenyi Biotec) and labelled for extracellular markers, including with CD45 BV786 (Clone D058-1283), CD3 BUV 395 (Clone SP34-2), CD4 BB515 (Clone L200) and CD8 PE (Clone SK1, BioLegend) antibody in FACS buffer (Ca2+/Mg2+-free PBS, supplemented with 2% FCS and 2 mM ethylenediaminetetraacetic acid (EDTA)). Dead cells were excluded using Zombie Aqua 516 viability dye (BioLegend). Intracellular staining with IFN-γ BV711 (Clone 4S.B3), IL-2 BV421 (Clone MQ1-17H12), and IL-4 APC (Clone 8D4-8) was performed with the Fixation and Permeabilization kit (BD Biosciences) according to the manufacturer’s protocol. Unless stated otherwise, antibodies were purchased from BD Biosciences. Flow cytometry was performed on the BD LSR Fortessa X20, and data were analyzed with FlowJo (LLC, V10). The gating strategy is shown in Supplementary Fig. 7c. Frequencies of ZIKV-specific T cells were calculated by subtracting non-peptide stimulated samples from corresponding stimulated samples. Due to cryopreservation-related viability loss, PBMC samples from three YF-ZIK-immunized individuals showed insufficient viable cell counts for ICS and were therefore excluded from analysis.
AG129 mice
All mouse experiments were approved by the Ethical Committee of the Animal Research Center of KU Leuven (project number P100/2019) and conducted in full accordance with the Belgian guidelines for animal experimentation and the guidelines of the Federation of European Laboratory Animal Science Associations. 129/SvEv Ifnar1tm1Agt Ifngr1tm1Agt mice deficient in interferon-α/β and γ receptors (AG129 mice; B&K Universal, Marshall Bio resources, UK) were bred in-house at the Experimental Animal Facilities of KU Leuven and randomly assigned to study groups. Sex was not considered in the study design. Animals were housed in groups of up to n = 5 in individually ventilated cages (IsoCage N Biocontainment System, Tecniplast) at a temperature of 21 °C, humidity of 55%, and 12:12 dark/light cycles, with ad libitum access to food and water and cage enrichment (cardboard houses and wood blocks).
ZIKV MR766 titration and passive serum transfer experiment in mice
For ZIKV MR766 titration, 8–10 weeks old AG129 mice were injected intraperitoneally (i.p.) with 1 PFU, 10 PFU, 102 PFU, and 103 PFU of MR766 virus, or medium as control. Mice were monitored daily for changes in body weight and clinical signs of disease. Sick mice were euthanized based on morbidity criteria (hind limb paralysis, hunch posture, ruffled fur, and weakness) or if weight loss exceeded 25%.
For the serum transfer experiment, 8–10 weeks old AG129 mice received a single-dose of 450 µl of immune serum pool from samples collected on days 21, 35, and 42 from YF-ZIK-immunized macaques (mean neutralizing titre of 5492), or from sham-vaccinated macaques by i.p. injection. Twenty-four hours after passive serum transfer, mice were infected with 10 PFU of ZIKV MR766 by i.p. injection (Fig. 3d). For survival analysis (Fig. 3e), results from two independent experiments were pooled from in total n = 9 mice (4 + 5) receiving antiserum and n = 10 mice (5 + 5) receiving sham serum. A separate group of AG129 mice (n = 3) that received the serum pool from YF-ZIK-immunized macaques was not infected and bled weekly to determine NAbs kinetics after serum transfer (Supplementary Fig. 3h). Mice were monitored daily for weight change and bled weekly to determine the antibody titres. Sick mice were euthanized based on morbidity criteria (hind limb paralysis, hunch posture, ruffled fur, and weakness) or if weight loss exceeded 25%.
Quantitative ZIKV RNA analysis in plasma
ZIKV RNA in plasma samples from EDTA-treated blood was quantified using real-time PCR as described before86. Briefly, viral RNA was extracted by using the Qiagen QIAamp® Viral RNA Mini Kit. The cDNA synthesis and PCR amplification were performed using the Brilliant II QRT-PCR Core Reagent Kit, 1-Step kit (Agilent), and RNA quantification was performed on a Bio-Rad CFX Connect real-time system.
ZIKV NS1-specific antibody ELISA
The ZIKV (Suriname) NS1 recombinant protein (The Native antigen company, cat. no. ZIKVSU-NS1-100) was used at a concentration of 1 µg/ml in Dulbecco’s phosphate-buffered saline (DPBS, Gibco) to coat 96-well microplates overnight at room temperature (RT). The antigen-coated plates were washed three times with PBST-20 (0.05% Tween-20 in DPBS), blocked with 3% Bovine Serum Albumin (BSA) for 1 h and incubated with NHP serum (1:500 dilution in DPBS containing 0.5% BSA and 0.2% Tween-20) for another 30 min. Subsequently, the plates were incubated with a goat anti-monkey IgG, IgA, IgM (Fc Specific), secondary antibody conjugated with horseradish peroxidase (1:1000, Nordic-MUBio) for 30 min. Plates were washed (3–5 times) between incubations steps with PBST-20. The reaction was developed with tetramethylbenzidine liquid substrate system for ELISA (Sigma-Aldrich) for 10 min and stopped with 1 M hydrochloric acid (HCl). The absorbance at 450 nm was measured using SPARK Magellan plate reader (Tecan Life Technologies). All measurements were conducted in triplicates.
Transcriptional and systems immunology analyses
Whole blood samples were collected in PAXgene tubes (Qiagen) for RNA preservation. Digital transcriptomics (nCounter platform, Nanostring Ltd.) was performed as previously described for other viral pathologies and their vaccine outcomes65,66,88,89. In brief, purified RNA samples were hybridized to unique capture/reporter pairs (50 bp each) targeting 770 transcripts (754 immune transcripts and 16 housekeeping genes present in the “Non-human Primate Immunology” Panel V2, as well as 6 positive and 8 negative control probes (all from NanoString). Results were sequentially corrected for background (negative control probes), technical variation (positive control probes) and RNA content (housekeeping genes), followed by differential expression, cell type deconvolution, pathway enrichment and network analysis using nSolver 4.0 (NanoString), MSigDb90, and STRING, in conjunction with publicly available bulk and single-cell transcriptomic datasets GEO and CZ CELLxGENE Discover (https://cellxgene.cziscience.com/)91,92. Resulting signature and cell type scores are the combined normalized total of all z-scores for all genes in a certain pathway or cell type. Gene sets used to define individual pathways are listed in the section of the Source Data corresponding to Fig. 2. Data mining of demographic, serological, and transcriptomic data with machine learning algorithms for prediction (attribute selection, decision trees) was performed using Weka (version 3.8.4). Uniform Manifold Approximation and Projection (UMAP) and dot plot representations were performed in RStudio (version 2024.04.2 + 764), using the Seurat package (v5)93 and the Tabula Sapiens - Blood database44.
Statistics and reproducibility
GraphPad Prism Version 9 (GraphPad Software, Inc.) was used for all statistical analyses. One-way or two-way analysis of variance (ANOVA) with multiple comparisons was applied to analyze the time-series observations for independent groups. The nonparametric two-tailed Mann–Whitney U-test was used for the comparison between paired groups. Kruskal–Wallis test was used to analyze differences among more than two groups with a non-normal distribution and corrected for multiple comparisons using Dunn’s method. Univariate and multivariate linear and logistic regression models were tested in GraphPad Prism Version 9 and XL-STAT. No statistical method was used to predetermine sample size. Details of statistical analyses are specified in each figure legend. Values were considered significantly different at P < 0.05. All raw, individual-level data are presented in the Source Data file.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data associated with this study are provided in the paper, the Supplementary materials or the Source Data file. Transcriptomic data from this study are deposited and accessible at https://www.ebi.ac.uk/biostudies/arrayexpress/studies/E-MTAB-15887. Unique materials used in this study are available from the corresponding author upon reasonable request. Source data are provided with this paper.
Code availability
No specific code was generated for this project.
References
Barrows, N. J. et al. Biochemistry and molecular biology of flaviviruses. Chem. Rev. 118, 4448–4482 (2018).
Musso, D., Ko, A. I. & Baud, D. Zika virus infection - after the pandemic. N. Engl. J. Med. 381, 1444–1457 (2019).
Shan, C., Xie, X. & Shi, P. Y. Zika virus vaccine: progress and challenges. Cell Host Microbe 24, 12–17 (2018).
Barouch, D. H., Thomas, S. J. & Michael, N. L. Prospects for a Zika virus vaccine. Immunity 46, 176–182 (2017).
Rabe, I. B. et al. A review of the recent epidemiology of Zika virus infection. Am. J. Trop. Med. Hyg. 112, 1026–1035 (2025).
Ostrowsky, J. T. et al. Zika virus vaccines and monoclonal antibodies: a priority agenda for research and development. Lancet Infect Dis. 25, e402–e415 (2025).
Hombach, J., Cardosa, M. J., Sabchareon, A., Vaughn, D. W. & Barrett, A. D. Scientific consultation on immunological correlates of protection induced by dengue vaccines report from a meeting held at the World Health Organization 17-18 November 2005. Vaccine 25, 4130–4139 (2007).
Barrett, A. D. T. Yellow fever vaccine: the conundrum of 2 doses, one dose, or one-fifth dose to induce and maintain protective immunity. J. Infect. Dis. 221, 1922–1924 (2020).
Abbink, P. et al. Protective efficacy of multiple vaccine platforms against Zika virus challenge in rhesus monkeys. Science 353, 1129–1132 (2016).
Abbink, P. et al. Durability and correlates of vaccine protection against Zika virus in rhesus monkeys. Sci. Transl. Med. 9, eaao4163 (2017).
Dowd, K. A. et al. Rapid development of a DNA vaccine for Zika virus. Science 354, 237–240 (2016).
Maciejewski, S. et al. Distinct neutralizing antibody correlates of protection among related Zika virus vaccines identify a role for antibody quality. Sci. Transl. Med. 12, eaaw9066 (2020).
Larocca, R. A. et al. Vaccine protection against Zika virus from Brazil. Nature 536, 474–478 (2016).
Nkolola, J. P. et al. Protective threshold of a potent neutralizing Zika virus monoclonal antibody in rhesus macaques. J. Virol. 98, e0142924 (2024).
Aliota, M. T. et al. Characterization of lethal Zika virus infection in AG129 mice. PLoS Negl. Trop. Dis. 10, e0004682 (2016).
Zmurko, J. et al. The viral polymerase inhibitor 7-deaza-2’-C-methyladenosine is a potent inhibitor of in vitro Zika virus replication and delays disease progression in a robust mouse infection model. PLoS Negl. Trop. Dis. 10, e0004695 (2016).
Lucas, C. G. O. et al. Critical role of CD4(+) T cells and IFNgamma signaling in antibody-mediated resistance to Zika virus infection. Nat. Commun. 9, 3136 (2018).
Hassert, M. et al. CD4+T cells mediate protection against Zika associated severe disease in a mouse model of infection. PLoS Pathog. 14, e1007237 (2018).
Elong Ngono, A. et al. CD8(+) T cells mediate protection against Zika virus induced by an NS3-based vaccine. Sci. Adv. 6, eabb2154 (2020).
Elong Ngono, A. et al. CD4+ T cells promote humoral immunity and viral control during Zika virus infection. PLoS Pathog. 15, e1007474 (2019).
Grubor-Bauk, B. et al. NS1 DNA vaccination protects against Zika infection through T cell-mediated immunity in immunocompetent mice. Sci. Adv. 5, eaax2388 (2019).
Kum, D. B. et al. A yellow fever-Zika chimeric virus vaccine candidate protects against Zika infection and congenital malformations in mice. NPJ Vaccines 3, 56 (2018).
Lima, N. S., Rolland, M., Modjarrad, K. & Trautmann, L. T cell immunity and Zika virus vaccine development. Trends Immunol. 38, 594–605 (2017).
Pulendran, B. Learning immunology from the yellow fever vaccine: innate immunity to systems vaccinology. Nat. Rev. Immunol. 9, 741–747 (2009).
Pulendran, B., Oh, J. Z., Nakaya, H. I., Ravindran, R. & Kazmin, D. A. Immunity to viruses: learning from successful human vaccines. Immunol. Rev. 255, 243–255 (2013).
Querec, T. D. et al. Systems biology approach predicts immunogenicity of the yellow fever vaccine in humans. Nat. Immunol. 10, 116–125 (2009).
Appaiahgari, M. B. & Vrati, S. Clinical development of IMOJEV (R)-a recombinant Japanese encephalitis chimeric vaccine (JE-CV). Expert Opin. Biol. Ther. 12, 1251–1263 (2012).
Guy, B. et al. A recombinant live attenuated tetravalent vaccine for the prevention of dengue. Expert Rev. Vaccines 16, 1–13 (2017).
Bonaldo, M. C., Sequeira, P. C. & Galler, R. The yellow fever 17D virus as a platform for new live attenuated vaccines. Hum. Vaccin Immunother. 10, 1256–1265 (2014).
Fourati, S. et al. Pan-vaccine analysis reveals innate immune endotypes predictive of antibody responses to vaccination. Nat. Immunol. 23, 1777–1787 (2022).
Gaucher, D. et al. Yellow fever vaccine induces integrated multilineage and polyfunctional immune responses. J. Exp. Med. 205, 3119–3131 (2008).
Hagan, T. et al. Transcriptional atlas of the human immune response to 13 vaccines reveals a common predictor of vaccine-induced antibody responses. Nat. Immunol. 23, 1788–1798 (2022).
Querec, T. et al. Yellow fever vaccine YF-17D activates multiple dendritic cell subsets via TLR2, 7, 8, and 9 to stimulate polyvalent immunity. J. Exp. Med. 203, 413–424 (2006).
Dudley, D. M. et al. A rhesus macaque model of Asian-lineage Zika virus infection. Nat. Commun. 7, 12204 (2016).
Osuna, C. E. et al. Zika viral dynamics and shedding in rhesus and cynomolgus macaques. Nat. Med. 22, 1448–1455 (2016).
Gobel, S. et al. Parallel multifactorial process optimization and intensification for high-yield production of live YF17D-vectored Zika vaccine. Vaccines (Basel) 12, 755 (2024).
Sanchez-Felipe, L., Alpizar, Y. A., Ma, J., Coelmont, L. & Dallmeier, K. YF17D-based vaccines - standing on the shoulders of a giant. Eur. J. Immunol. 54, e2250133 (2024).
Akondy, R. S. et al. Initial viral load determines the magnitude of the human CD8 T cell response to yellow fever vaccination. Proc. Natl. Acad. Sci. USA 112, 3050–3055 (2015).
Kalimuddin, S. et al. Vaccine-induced T cell responses control Orthoflavivirus challenge infection without neutralizing antibodies in humans. Nat. Microbiol. 10, 374–387 (2025).
Mantel, N. et al. Cynomolgus macaques as a translational model of human immune responses to yellow fever 17D vaccination. J. Virol. 98, e0151623 (2024).
Mudd, P. A. et al. The live-attenuated yellow fever vaccine 17D induces broad and potent T cell responses against several viral proteins in Indian rhesus macaques-implications for recombinant vaccine design. Immunogenetics 62, 593–600 (2010).
Elong Ngono, A. et al. Mapping and role of the CD8(+) T cell response during primary Zika virus infection in mice. Cell Host Microbe 21, 35–46 (2017).
Kum, D. B. et al. A chimeric yellow fever-Zika virus vaccine candidate fully protects against yellow fever virus infection in mice. Emerg. Microbes Infect. 9, 520–533 (2020).
Tabula Sapiens, C. et al. The tabula sapiens: a multiple-organ, single-cell transcriptomic atlas of humans. Science 376, eabl4896 (2022).
Meyaard, L. The inhibitory collagen receptor LAIR-1 (CD305). J. Leukoc. Biol. 83, 799–803 (2008).
Kwek, S. S. et al. A systematic approach to the development of a safe live attenuated Zika vaccine. Nat. Commun. 9, 1031 (2018).
Lecouturier, V. et al. An optimized purified inactivated Zika vaccine provides sustained immunogenicity and protection in cynomolgus macaques. NPJ Vaccines 5, 19 (2020).
Acosta, C. J. et al. Predicting efficacy of a purified inactivated Zika virus vaccine in flavivirus-naive humans using an immunological correlate of protection in non-human primates. Microorganisms 12, 1177 (2024).
Scott, J. M. et al. Cellular and humoral immunity protect against vaginal Zika virus infection in mice. J. Virol. 92, e00038–00018 (2018).
Lemmens, V. et al. YF17D-vectored Ebola vaccine candidate protects mice against lethal surrogate Ebola and yellow fever virus challenge. NPJ Vaccines 8, 99 (2023).
Ma, J. et al. Live-attenuated YF17D-vectored COVID-19 vaccine protects from lethal yellow fever virus infection in mouse and hamster models. EBioMedicine 83, 104240 (2022).
Sanchez-Felipe, L. et al. A single-dose live-attenuated YF17D-vectored SARS-CoV-2 vaccine candidate. Nature 590, 320–325 (2021).
Best, S. M. The many faces of the flavivirus NS5 protein in antagonism of type I interferon signaling. J. Virol. 91, e01970–01916 (2017).
Laurent-Rolle, M. et al. The interferon signaling antagonist function of yellow fever virus NS5 protein is activated by type I interferon. Cell Host Microbe 16, 314–327 (2014).
Bastard, P. et al. Auto-antibodies to type I IFNs can underlie adverse reactions to yellow fever live attenuated vaccine. J. Exp. Med. 218, e20202486 (2021).
Hernandez, N. et al. Inherited IFNAR1 deficiency in otherwise healthy patients with adverse reaction to measles and yellow fever live vaccines. J. Exp. Med. 216, 2057–2070 (2019).
Bovay, A. et al. Minimal immune response to booster vaccination against Yellow Fever associated with pre-existing antibodies. Vaccine 38, 2172–2182 (2020).
Kongsgaard, M. et al. Adaptive immune responses to booster vaccination against yellow fever virus are much reduced compared to those after primary vaccination. Sci. Rep. 7, 662 (2017).
Dudley, D. M. et al. Infection via mosquito bite alters Zika virus tissue tropism and replication kinetics in rhesus macaques. Nat. Commun. 8, 2096 (2017).
Sapparapu, G. et al. Neutralizing human antibodies prevent Zika virus replication and fetal disease in mice. Nature 540, 443–447 (2016).
Shan, C. et al. A single-dose live-attenuated vaccine prevents Zika virus pregnancy transmission and testis damage. Nat. Commun. 8, 676 (2017).
Li, A. et al. Role of non-human primate models in accelerating research and developing countermeasures against Zika virus infection. Lancet Microbe. 6, 101030 (2025).
Van Rompay, K. K. A. et al. DNA vaccination before conception protects Zika virus-exposed pregnant macaques against prolonged viremia and improves fetal outcomes. Sci. Transl. Med. 11, eaay2736 (2019).
Mackay, F., Schneider, P., Rennert, P. & Browning, J. BAFF AND APRIL: a tutorial on B cell survival. Annu. Rev. Immunol. 21, 231–264 (2003).
van der Wijst, M. G. P. et al. Type I interferon autoantibodies are associated with systemic immune alterations in patients with COVID-19. Sci. Transl. Med. 13, eabh2624 (2021).
Cuypers, L. et al. Immunovirological and environmental screening reveals actionable risk factors for fatal COVID-19 during post-vaccination nursing home outbreaks. Nat. Aging 3, 722–733 (2023).
Carpio, K. L. & Barrett, A. D. T. Flavivirus NS1 and its potential in vaccine development. Vaccines (Basel) 9, 622 (2021).
Monath, T. P. et al. Live virus vaccines based on a yellow fever vaccine backbone: standardized template with key considerations for a risk/benefit assessment. Vaccine 33, 62–72 (2015).
Monath, T. P. et al. Chimeric live, attenuated vaccine against Japanese encephalitis (ChimeriVax-JE): phase 2 clinical trials for safety and immunogenicity, effect of vaccine dose and schedule, and memory response to challenge with inactivated Japanese encephalitis antigen. J. Infect. Dis. 188, 1213–1230 (2003).
Monath, T. P. et al. Chimeric yellow fever virus 17D-Japanese encephalitis virus vaccine: dose-response effectiveness and extended safety testing in rhesus monkeys. J. Virol. 74, 1742–1751 (2000).
Biedenbender, R., Bevilacqua, J., Gregg, A. M., Watson, M. & Dayan, G. Phase II, randomized, double-blind, placebo-controlled, multicenter study to investigate the immunogenicity and safety of a West Nile virus vaccine in healthy adults. J. Infect. Dis. 203, 75–84 (2011).
Campi-Azevedo, A. C. et al. Subdoses of 17DD yellow fever vaccine elicit equivalent virological/immunological kinetics timeline. BMC Infect. Dis. 14, 391 (2014).
Dayan, G. H., Bevilacqua, J., Coleman, D., Buldo, A. & Risi, G. Phase II, dose ranging study of the safety and immunogenicity of single dose West Nile vaccine in healthy adults >/= 50 years of age. Vaccine 30, 6656–6664 (2012).
Monath, T. P. et al. A live, attenuated recombinant West Nile virus vaccine. Proc. Natl. Acad. Sci. USA 103, 6694–6699 (2006).
Juan-Giner, A. et al. Immunogenicity and safety of fractional doses of yellow fever vaccines: a randomised, double-blind, non-inferiority trial. Lancet 397, 119–127 (2021).
Kimathi, D. et al. Low-dose yellow fever vaccine in adults in Africa. N. Engl. J. Med. 392, 788–797 (2025).
Kum, D. B. et al. Limited evolution of the yellow fever virus 17d in a mouse infection model. Emerg. Microbes Infect. 8, 1734–1746 (2019).
Verhoeff, N. P. et al. Focus localization in patients with partial epilepsy with 99Tcm-HMPAO SPECT under continuous surface EEG monitoring. Nucl. Med. Commun. 13, 127–136 (1992).
Katzelnick, L. C. et al. Zika virus infection enhances future risk of severe dengue disease. Science 369, 1123–1128 (2020).
Zambrana, J. V. et al. Primary exposure to Zika virus is linked with increased risk of symptomatic dengue virus infection with serotypes 2, 3, and 4, but not 1. Sci. Transl. Med. 16, eadn2199 (2024).
Thomas, S. J. Is new dengue vaccine efficacy data a relief or cause for concern? NPJ Vaccines 8, 55 (2023).
Wilder-Smith, A. B., Freedman, D. O. & Wilder-Smith, A. Edging towards a third dengue vaccine. Lancet Infect. Dis. 24, 1182–1184 (2024).
Zambrano, B. et al. Zika and dengue interactions in the context of a large dengue vaccine clinical trial in Latin America. Am. J. Trop. Med. Hyg. 104, 136–144 (2021).
Wilder-Smith, A., Cherian, T. & Hombach, J. Dengue vaccine development and deployment into routine immunization. Vaccines (Basel) 13, 483 (2025).
Gobel, S. et al. Optimization of YF17D-vectored Zika vaccine production by employing small-molecule viral sensitizers to enhance yields. Vaccines (Basel) 13, 757 (2025).
Barzon, L. et al. Isolation of infectious Zika virus from saliva and prolonged viral RNA shedding in a traveller returning from the Dominican Republic to Italy, January 2016. Eur. Surveill. 21, 30159 (2016).
Rasulova, M. et al. A high-throughput yellow fever neutralization assay. Microbiol. Spectr. 10, e0254821 (2022).
Feys, S. et al. Lung epithelial and myeloid innate immunity in influenza-associated or COVID-19-associated pulmonary aspergillosis: an observational study. Lancet Respir. Med. 10, 1147–1159 (2022).
Menezes, S. M. et al. Blood transcriptomic analyses reveal persistent SARS-CoV-2 RNA and candidate biomarkers in post-COVID-19 condition. Lancet Microbe 5, 100849 (2024).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Megill, C. et al. cellxgene: a performant, scalable exploration platform for high dimensional sparse matrices. Preprint at https://www.biorxiv.org/content/10.1101/2021.04.05.438318v1 (2021).
Program, C. Z. I. C. S. et al. CZ CELLxGENE Discover: a single-cell data platform for scalable exploration, analysis and modeling of aggregated data. Nucleic Acids Res. 53, D886–D900 (2025).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e3529 (2021).
Acknowledgements
We thank Prof. Luisa Barzon (University of Padua, Italy) for providing the ZIKV challenge strain. Dr. Dominique Van Looveren, Dr. Madina Rasulova, Jasmine Paulissen, Dr. Winnie Kerstens and Nathalie Thys (TPVC) contributed to serological assessments; Dr. Dieudonné B. Kum assisted with virus stock generation; Dr. Niraj Mishra, Dr. Sapna Sharma, Carolien De Keyzer and Lindsey Bervoets (MVVD) supported in vivo studies; and Katrien Geerts and Sarah Debaveye (MVVD) contributed to in vitro cell and virus culture. We also acknowledge the staff of the animal facilities at BPRC and the KU Leuven Rega Institute for scientific-technical support and Dr. Gregory Fanning and Dr. Frederik Stevenaert for critical discussion of the manuscript. Current work was supported by the EU-TRANSVAC2 (project number TRANSVAC-TNA-1802-2) to L.C., J.N. and K.D. and the European Union’s Horizon 2020 Research and Innovation Program under Grant Agreements No. 733176 to L.C., J.N., E.J.V. and K.D. (RABYD-VAX consortium), No. 734584 (ZikaPLAN) and No. 734548 (ZIKAlliance) to J.N. and No. 101137459 to K.D. (Yellow4Flavi). B.M.D., Y.A.A. and K.D. received support from the joined KU Leuven/University of Edinburgh Funds Global Seed Fund (GSF/25/074; EarlyViroPath) and K.D. from KU Leuven Internal Funds (C3/19/057; Lab of Excellence). J.V.W. was supported by KU Leuven (“Vaast Leysen Leerstoel”) and Flanders Research Foundation (FWO) Grants G0A0621N and G065421N.
Author information
Authors and Affiliations
Contributions
Conceptualization: K.D., L.C., J.V.W., H.J.T. Methodology: J.M., B.M.D, J.V.W., B.E.V., G.K., E.J.V., T.V., H.J.T., L.S.F., M.P.A.J., N.D.G., P.M., K.D. Investigation: J.M., B.E.V., B.M.D, Y.A.A., J.V.W., H.J.T. Visualization: J.M., B.M.D., J.V.W., Y.A.A., H.J.T. Funding acquisition: L.C., J.N., E.J.V., B.M.D., Y.A.A., K.D. Project administration: L.C., K.D. Supervision: K.D. Writing–original draft: J.M., J.V.W., H.J.T. Writing–review & editing: all authors.
Corresponding authors
Ethics declarations
Competing interests
J.N. and K.D. are mentioned as inventors on a patent application claiming the discovery and use of chimeric yellow fever—Zika virus vaccines. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks John J. Suschak, Kuan Rong Chan, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ma, J., Malengier-Devlies, B., Verstrepen, B.E. et al. Immunological and transcriptomic profile of chimeric live-attenuated Zika vaccine linked to protection in non-human primates. Nat Commun 16, 11221 (2025). https://doi.org/10.1038/s41467-025-66073-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66073-4