Abstract
Sensory information is fundamental for navigation. Visual motion is used by animals to estimate their traveling distance and direction, and landmarks allow animals to tether their location and orientation to their environment. How such signals are integrated in the vertebrate brain is poorly understood. Here we investigate the representation of directional whole field visual motion and landmark position in the larval zebrafish head direction circuit. Using calcium imaging we show that these stimuli are represented in the habenula, interpeduncular nucleus and anterior hindbrain. In the dorsal interpeduncular nucleus, both stimuli are topographically arranged and align with the representation of the heading signal. Neuronal ablations show that the landmark responses, but not the whole field motion responses, require intact habenula input. Our findings suggest the interpeduncular nucleus as a site for integration of the heading signal with visual information, shedding light on how navigational signals are processed in the vertebrate brain.
Similar content being viewed by others
Introduction
To navigate the environment, an animal must know where it is heading. Neural networks that represent the animal’s orientation within its environment, called heading direction (HD) networks, have been found in mammals, birds, fish and insects1,2,3,4,5. In the absence of landmarks, animals can path integrate self-motion cues such as vestibular input, self-generated optic flow and motor commands to maintain an updated internal representation of heading6,7,8,9,10,11,12. This path integration is nevertheless prone to the accumulation of errors, making HD networks less stable in the absence of visual landmarks13,14. Anchoring the HD network therefore requires the incorporation of sensory input into this internal representation, but the mechanisms by which this is achieved in vertebrates remain largely unknown.
In mammals, HD neurons (HDNs) have been found in several brain regions, including the dorsal tegmental nucleus (DTN), lateral mammillary nucleus (LMN), anterodorsal thalamic nucleus (ADN), postsubiculum (PoSub) and entorhinal and retrosplenial cortices2,6,12,15,16,17. The heading signal is internally generated by a network that integrates vestibular signals and motor commands, most likely in the DTN and LMN16,18,19,20. From there, this signal is transmitted to the thalamus and PoSub. Previous studies have shown that activity of HDNs in the ADN, PoSub and retrosplenial cortex, is sensitive to visual information14,21,22. While some studies suggest that sensory information is most likely integrated in the PoSub23 or the retrosplenial cortex22,24, it is still not known whether such information affects HDNs in other parts of the network.
In recent years, the topography and connectivity of the insect heading direction network has been discovered and described in great detail1,25,26,27,28,29. The insect HDNs form a network in a structure called the central complex in the center of the insect brain1. The neuropil of the insect HDNs is organized as wedges in the ellipsoid body, a circular structure located within the central complex. Heading direction representation in this network can be shifted by motor information (if the animal turns) or by sensory information, such as visual or somatosensory cues (e.g., if a landmark in the environment moves to a new location)1,30,31,32,33,34. This rotation is performed by shift neurons, which have their axons tiling the ellipsoid body in a similar manner to the HDNs but with a small offset. Surprisingly, the neurons that provide sensory information to this system do not form localized arborizations but rather connect to all HDNs29,35. The fact that many other types of neurons that innervate the ellipsoid body, or other parts of the central complex, tile a specific wedge, row or column in this region29,36 allows for local modulation of these global inputs. This principle of organization appears to allow for very flexible routes of information flow in the insect navigation system.
In addition to being where the HD signal is generated, the insect central complex is also a site where this signal is integrated with other information that is relevant for navigation, such as the animal’s traveling direction. While HDNs themselves are not sensitive to translational visual motion, another type of cell in this region responds to visual motion in a particular direction37,38. It is currently unclear where such integration exists in the vertebrate brain.
Recently, we discovered a heading direction network in the zebrafish anterior hindbrain (aHB) and suggested this as the homologous structure of the mammalian DTN5. This is due to three reasons: (1) the DTN develops from rhombomere 1, where the zebrafish HD network is located, (2) both regions contain a group of GABAergic HD neurons and (3) the aHB and DTN show similar connectivity patterns with the habenula and interpeduncular nucleus (IPN)12,16,39,40,41.
Here, we investigate the representation of different types of visual information in the aHB-IPN-habenula network. We show that all three structures contain visually tuned neurons. We further show that both direction of optic flow and azimuthal landmark position are topographically represented in the dorsal IPN (dIPN). Finally, we use ablations to show that habenula input is required for representation of landmark position in the dIPN, but not for representation of visual motion or basic functioning of the HD network.
Results
Whole brain tuning to directional visual motion
Our first goal was to detect neurons, throughout the brain, tuned to directional visual motion, a visual signal that indicates an animal’s traveling direction. We used lightsheet microscopy to obtain whole brain imaging datasets recorded at 2 Hz from Tg(elavl3:H2B-GCaMP6s) zebrafish larvae presented with whole field translational visual motion (Fig. 1a, see Methods). Fish were head restrained in agarose, with their tail free to allow tracking of tail movements. Fish were presented with a natural pattern on their bottom field of view. In each trial, the projected pattern moved in one of the eight cardinal or inter-cardinal directions with respect to the fish, chosen in a random order. Neurons responding reliably to visual motion in at least one direction were detected in the tectum, pretectum, left habenula, aHB and IPN, in line with previous studies42,43,44 (Fig. 1b–g, Supplementary Figs. 1–3). Out of the regions containing reliably responding neurons (see Methods for reliability index calculation), the representation of different directions in the pretectum and aHB was found to be lateralized such that neurons responding to leftward motion are on the left hemisphere of the brain, and neurons responding to rightward motion are on the right hemisphere. In contrast, neurons in the left habenula and tectum do not exhibit this lateralization and visually tuned neurons in these regions do not have a particular anatomical distribution (Fig. 1c, d).
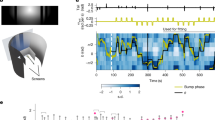
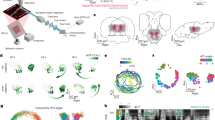
a Illustration of experimental design. Left, side view of an embedded zebrafish larva under the microscope. Center, top view of embedded zebrafish larva in a lightsheet imaging chamber. Agarose is removed from the tail to allow swimming and tail tracking. Agarose is also removed from the side and front to prevent the laser from scattering. Right, a zebrafish on top of a pink noise pattern moving in 8 possible directions. b horizontal and sagittal views of the zebrafish brain (MapZeBrain atlas reference brain). Lines point to specific regions that were found to respond to whole field visual motion. PT: pretectum, OT: optic tectum, aHB: anterior hindbrain, IPN: interpeduncular nucleus. c Three views of all ROIs extracted from a whole brain dataset (one example fish). Neurons are colored according to their correlation value with rightward motion. d Three views of the same fish, only reliably responding neurons are shown (top 5% of reliability index). Each neuron is colored according to the direction it is tuned to. e Same as d but for all fish in the dataset, registered to a reference fish (n = 15). f Example neurons that are tuned to different directions. Left, tuning curves of 8 neurons. Right, the full traces from the entire experiment for the same neurons. The colorful shadings indicate the direction that is being presented. g Distributions of correlation values with each direction for neurons in the left habenula (LHab), IPN, aHB and pretectum (n = 15).
Heading direction neurons in the aHB are not tuned to visual motion
The aHB, the region containing the zebrafish HDNs, also contains many neurons that are tuned to visual motion (Fig. 1). We next wanted to check if HDNs in the aHB respond to visual motion. We used a lightsheet microscope to image GABAergic neurons in the aHB in both darkness and while presenting directional whole field motion (Fig. 2a). HDNs were selected based on their activity in the darkness phase as previously described in5. Our results showed that HDNs and visually tuned neurons form two non-overlapping populations (Fig. 2). In all of our fish, neurons classified as HDNs did not reliably respond to translational visual motion (Fig. 2c, d) and did not show high correlation with any of the sensory regressors (Fig. 2e, f). Similarly, neurons that reliably respond to the visual stimulus were never identified as HDNs, i.e. they were less negatively correlated with other neurons during the darkness phase (Fig. 2c–f).
a Left, anatomy of the aHB of a Tg(gad1b:Gal4; UAS:GCaMP6s) fish. HDNs are indicated in green. Right, traces of heading direction neurons, sorted according to their phase. The tail trace and colorful shading indicating presentation of visual motion are shown on top of the traces. b Left, anatomy of the aHB in the same fish. Visually tuned neurons are labeled in purple. Right, traces of visually tuned neurons. c Scatter plot showing the HDNs and the visually tuned neurons are functionally two separate populations (same example fish as in (a) and (b). HDNs do not show reliable responses to visual motion (x axis) and show more negative correlation values with other neurons during the darkness phase of the experiment (y axis). Heading direction neurons have more negative correlation values with other neurons during darkness and visually tuned neurons have higher values of the reliability index. d Same as c but for all fish (n(fish)=3, n(HDNs)=[74, 172, 137], n(visually tuned)=[56, 367, 236]). e Scatter plot showing the HDNs and the visually tuned neurons are functionally two separate populations (same example fish as in (a) and (b)). HDNs do not show high correlation values with visual motion, unlike visually tuned neurons. f Same as e but for all fish.
Visual motion is topographically arranged in the dIPN
Our whole brain imaging data revealed neurons tuned to directional whole field motion in the IPN, where HDNs from the aHB project their dendrites and axons to5. We next imaged different transgenic lines that label different components of the IPN, and characterized their responses to directional visual motion. To characterize the tuning of IPN neuropil to visual motion, we generated tuning maps in which each pixel is colored according to the direction it is tuned to (Fig. 3, Supplementary Fig. 4). We first imaged Tg(elavl3:GCaMP6s) fish, in which GCaMP6s is expressed in the cytosol of all neurons, thus labeling both somata and neuropil. In this panneuronal line, we noticed a strong tuning to visual motion in the dIPN. This activity is topographically organized, with each direction represented in a single parasagittal stripe on the rostro-caudal axis. The leftmost stripe is tuned to backward-left motion and the rightmost stripe is tuned to backward-right motion (Fig. 3c, d). This topographic organization is highly consistent across different planes in the dIPN (Supplementary Fig. 4) and across fish (Fig. 3). We further quantified the average correlation with each direction as a function of position, and found that the topographic organization exists in the lateral-medial, but not in the rostro-caudal direction (Supplementary Figs. 5 and 6).
a Illustration of the zebrafish brain, horizontal (left) and sagittal (right) views. The IPN and its afferent structures are highlighted (habenula is shown in purple, IPN is shown in light blue, aHB is shown in gold) dashed square and line indicate the dIPN (field of view in following plots). b Illustration of heading direction representation in the dIPN (based on5). Left, illustration of the morphology of three HDNs and their projections in the dIPN: each HDN has a single neurite that goes ventrally and splits into an ipsilateral dendrite and a contralateral axon. Right, illustration of the activity of HDN neuropil in the IPN when the fish is facing different directions. When an HDN is active, both its axon and dendrite are active in the dIPN (c) Illustration of the IPN and its different components as expected in the pan-neuronal line: local IPN cells, neuropil from the aHB and habenula axons. d Left, median tuning map of the dIPN in the Tg(elavl3:GCaMP6s) line labeling the cytosol pan-neuronally (n = 4). Each pixel is colored according to the direction it is tuned to. Right, tuning maps of three example fish. e Illustration of habenula input to the dIPN. Axons from the left habenula enter the IPN and then wrap around it. f Left, median tuning map of the dIPN in the Tg(16715:Gal4) line labeling only habenula axons (n = 12). Right, tuning maps of three example fish. g Illustration of IPN cells and their neuropil. h Left, median tuning map of the dIPN in the Tg(s1168t:Gal4) line labeling IPN cells and neuropil (n = 15). Right, tuning maps of three example fish. i Illustration of GABAergic elements in the dIPN, which consists mostly from sparse expression in the IPN itself with additional neuropil that mostly originates in the aHB. j Left, median tuning map of the dIPN in the Tg(gad1b:Gal4) fish (n = 21). Right, tuning maps of three example fish. Arrows indicate neuropil stripes that are tuned to the same direction on both sides of the dIPN.
To better understand how this striped pattern is formed, we imaged fish lines with more restricted expression patterns. Next, we imaged the Tg(16715:Gal4; UAS:GCaMP6s) line, labeling only neurons in the habenula, a major source of excitatory input to the IPN in mammals and fish45,46,47,48. In zebrafish, the left habenula projects to both dorsal and ventral parts of the IPN while the right habenula projects mostly to the ventral IPN49,50. In both cases, individual habenular axons wrap around large portions of the IPN51. Despite this unspecific axonal innervation, the activity of habenular axons in the dIPN shows a topographical tuning to the visual stimulus. This pattern is like the one we observed in the Tg(elavl3:GCaMP6s) line (Fig. 3c, d), with habenula axons showing tuning that is organized in stripes in the rostro-caudal axis (Fig. 3e, f, Supplementary Figs. 4–6). This organization is surprising given the afore-mentioned fact that habenular axons wrap around the dIPN and do not form localized projections.
We next imaged the Tg(s1168t:Gal4; UAS:GCaMP6s) fish line that labels IPN neurons and their neuropil, but none of the structures that project to the IPN (Supplementary Fig. 7). The cell bodies of these neurons are aligned from left to right along the rostral boundary of the dIPN, and their neuropil extends caudally. This neuropil is tuned to directional motion in a very similar manner to that found in the habenula axons (Fig. 3g, h). This tuning is organized in a striped pattern, such that left stripes are tuned to leftward visual motion and right stripes are tuned to rightward visual motion. The overall organization is very similar to that observed in the pan-neuronal and habenula lines, with the exception of lack of tuning to backward motion by IPN cells (Supplementary Figs. 4–6).
We next imaged Tg(gad1b:Gal4; UAS:GCaMP6s) fish, in which only GABAergic neurons are labeled (Fig. 3i, j). When imaging the GABAergic neuropil in the dIPN, we imaged dendrites and axons of IPN neurons as well as those of aHB and potentially other GABAergic neurons projecting to the dIPN. These experiments revealed that the GABAergic neuropil in the dIPN is strongly tuned to visual motion in different directions. The tuning appears again to be organized in a striped pattern, such that different directions are represented by different neuropil stripes (Fig. 3i). However, the organization of the GABAergic neuropil is different from that observed for IPN cells and habenula axons. While in the previous datasets tuning to each direction appeared in a single stripe, when imaging the GABAergic neuropil, we found that each direction appears on both sides of the IPN. This is very similar to the structure of the heading direction signal in the dIPN (Fig. 3b5). As our data shows that HDNs themselves do not respond to visual motion, in the likely case that this tuning arises from aHB neurons, this would suggest that visually tuned neurons in the aHB have a similar morphology to that of HDNs.
Landmark position is topographically arranged in the dIPN
Many animals use visual landmarks to anchor their heading direction to their sensory environment. We next wanted to check if landmark position is at all represented in the dIPN. Neurons that have local receptive fields could represent this type of information. To detect such neurons, we used a two-photon microscope to record activity of IPN neurons in zebrafish larvae that were presented with a light bar at different azimuth angles in the range ±60° in their front visual field (Fig. 4a, see Methods).
a Illustration of experimental setup of presentation of frontal visual stimulus to embedded fish. b Illustration of somata distribution in the dIPN as seen in the Tg(elavl3:H2BGCaMP6s) line. c Tuning of dIPN cells to light position. Only reliably responding ROIs are shown, each ROI is colored according to the position it is tuned to. Left, data from multiple fish registered to one another (n = 7). Right, same as in the left panel but for four individual fish. The dIPN in each dataset is marked with a dashed circle. d Illustration of dIPN somata and neuropil. e Tuning of dIPN neuropil to landmark position. Left, average tuning map of Tg(s1168t:Gal4; UAS:GCaMP6s) fish, showing the average response pattern, each pixel is colored according to the direction that it is tuned to (n = 14). Right, example fish tuning maps of the dIPN in three example fish. f Illustration of the zebrafish brain, the habenula is highlighted in pink. g Tuning of habenula cells to landmark position as found in Tg(elavl3:H2B-GCaMP6s) and Tg(elavl3:GCaMP6s) fish. Only reliably responding ROIs are shown, each ROI is colored according to the position it is tuned to. Left, data from multiple fish morphed to one another (n = 16). Right, same as in the left panel but for four individual fish. The left habenula in each dataset is marked with a dashed circle. h Responses of eight neurons from the left habenula to the appearance of a light bar in the eight possible locations. For each neuron the mean ± sem response is shown. Orange shading indicates stimulus presentation.
When imaging cell bodies in the IPN, we found neurons, in all fish, that responded to light in a particular position of the visual field. In all fish, these light-responsive neurons tiled the IPN retinotopically. In addition, for most fish, this retinotopic map was aligned: neurons on the right side of the IPN responded to light on the right side of the visual field, whereas neurons on the left responded to stimuli on the left (Fig. 4b, c). For some fish though, (see first individual fish in Fig. 4c), this map was “phase shifted”. i.e. there was a constant offset between the left-right alignment of the visual field and the left-right alignment in the IPN.
We next wanted to better characterize the representation of landmark position in the dIPN. To do this we imaged the IPN neuropil while showing the same light bar stimulus. In these experiments, we observed a more graded pattern than we observed in the somata, with the azimuthal angle in the visual field similarly encoded in the left-right dIPN axis in stripes that extended rostro-caudally (Fig. 4d-e), similar to that observed in response to visual motion (Fig. 3g) and to the neuropil of HDNs (Fig. 3b and5). The striped pattern is consistent across different planes within the dIPN (Supplementary Fig. 8). Just as we had observed for the cell bodies, the representation of landmark position by IPN neuropil was not always consistent across fish. While in all fish we detected a consistent retinotopic map (within and across planes), this also appeared phase shifted by a constant offset in some fish (Fig. 4e).
Representation of landmark position in the habenula
Given that the habenula is the most prominent input to the IPN45,46,47,48, and that the left habenula neurons are known to respond to whole field changes in luminance levels52,53,54, we wanted to investigate the left habenula as a potential source for light responses in the IPN.
The visual receptive field properties of left habenula neurons have not been characterized, so we next imaged the habenula while showing the fish the same light bar stimuli as before. We found a population of neurons in the habenula that reliably responded to this stimulus (Fig. 4f, g). In line with previous studies, a large fraction of left habenula neurons (30 ± 10%, mean ± sd) reliably responded to the light stimulus, and a smaller number in the right habenula (10 ± 7%, mean ± sd). In the left habenula, we could detect neurons that had a receptive field localized in the azimuthal direction (Fig. 4h, Supplementary Fig. 9). Unlike the responses in the IPN, we could not detect any organization of this azimuthal position representation in the habenula (Fig. 4g). It appears that, as for visual motion (Fig. 1c, d), habenula neurons that respond to light in a particular position are not topographically organized.
Habenula axons wrap around the IPN and do not form localized contacts. However, as our data shows that habenula axons in the dIPN show topographically organized responses to directional visual motion (Fig. 3f), we wanted to investigate if a similar pattern exists for landmark representation. We next imaged habenula axons in the dIPN in fish presented with the light bar stimulus (Supplementary Fig. 10). Our data shows that habenula axons are tuned to landmark position, though we could not observe a fine structure as we did for the neuropil of dIPN cells. We find, in all fish (n = 17), that habenula axons on the left side of the dIPN are tuned to landmarks on the left side of the visual field, while habenula axons on the right side of the dIPN are tuned to landmarks on the right side of the visual field. In the center of the dIPN, we could not identify any specific tuning (these pixels appear white in the image), likely because axons from multiple neurons which are tuned to different positions are anatomically mixed.
The habenula is not the source of visual motion information
Our data shows that habenula neurons respond to visual motion and that their axons in the IPN show topographically organized tuning to visual motion direction (Figs. 1, 3). Given that the habenula projects to both the IPN and aHB, we next wanted to check if it is the left habenula that relays visual motion information to these structures. We used lightsheet microscopy to image a group of fish before and after chemogenetic habenula ablations. In these experiments we used triple transgenic fish; expressing Tg(elavl3:H2B-GCaMP6s), Tg(16715:Gal4) and Tg(UAS:Ntr-mCherry). In these fish nitroreductase (Ntr) is expressed exclusively in habenula neurons, such that in the presence of nifurpirinol (NFP) habenula cells are ablated (Supplementary Fig. 11 shows the expression pattern of the Tg(16715:Gal4) line). As controls, we used siblings of ablated fish that were found not to express Ntr at all (from now on referred to as treatment controls). Figure 5a shows a whole brain stack of a fish brain before and after NFP treatment. Following overnight treatment with NFP, habenula neurons are gone, as well as their axon terminals in the IPN (Fig. 5b).
a Z projection of a confocal stack of a Tg(elavl3:H2B-GCaMP6s; 16715:Gal4; UAS:Ntr-mCherry) fish before (left) and after (right) NFP treatment. b Same as a but zoomed in view of the IPN. c Pooled ROIs from 11 fish, each neuron is colored according to its correlation value with rightward motion before (left) and after (right) habenula ablation. Only reliably responding neurons are shown (top 5% of reliability index). d Same as c but for control fish not expressing Ntr in the habenula (n(fish)=11). e Number of reliably responding cells in the IPN and aHB before and after habenula ablation (distributions are not significantly different, one-sided Wilcoxon signed-rank test, p(IPN) = 0.09, p(aHB)=0.23, n(fish)=11). Cell count was done by counting all reliably responding neurons in the IPN/aHB (neurons with a reliability threshold > 0.25). f Same as e but for control fish not expressing Ntr (distributions are not significantly different, one-sided Wilcoxon signed-rank test, p(IPN) = 0.84, p(aHB)=0.48, n = 11). g Distribution of correlation values with all regressors before and after ablation for the IPN and aHB for Ntr+ fish (mean ± sd, n(fish)=11). The variances of the distributions before ablation are not significantly larger than those of the distributions post ablation (Wilcoxon signed-rank test, p(IPN) = 0.25, p(aHB)=0.12). h Distribution of correlation values with all regressors before and after ablation for the IPN and aHB for the control group (mean ± sd, n(fish)=11). The variances of the distributions before ablation are not significantly larger than those of the distributions post ablation (Wilcoxon signed-rank test, p(IPN) = 0.91, p(aHB)=0.28).
Figure 5 shows data from ablated and treatment control fish. Fish from both groups show directional tuning in the tectum, pretectum, aHB, IPN and left habenula before the treatment, as expected. Following NFP treatment, only Ntr+ fish lose visual tuning in the left habenula, indicating that the ablation was successful (Fig. 5c). To visualize changes in responses to visual motion we first voxelized the responses in all fish and computed the difference in response amplitude before and after ablation (Supplementary Fig. 12, see Methods). When comparing whole brain response patterns, it appears that habenula ablations do not affect representation of whole field motion in the brain. This can be seen not only in the voxelized data, but also when comparing the single cell correlation values before and after ablations (Supplementary Figs. 13 and 14). Specifically, visual tuning in the aHB and IPN remains intact (Fig. 5c–f, Supplementary Fig. 15), suggesting that the habenula is not required for representation of visual motion in these regions.
The habenula provides landmark information to the IPN
Our results show that both habenula and IPN neurons respond to light in localized parts of the visual field (Fig. 4). We wanted to check if the habenula is the source of visual landmark information to the dIPN. To do so, we performed chemogenetic ablations of the habenula and recorded activity in the IPN before and after ablation (Fig. 6). Before ablation, we could identify IPN neurons that responded to light in a particular position of the visual field in all fish. Following NFP treatment, IPN neurons in Ntr+ fish with ablated habenula showed no response to the presented stimuli (Fig. 6, Supplementary Fig. 16). IPN neurons in treatment control fish, in which the habenula was left intact, showed similar responses before and after NFP treatment (Fig. 6, Supplementary Fig. 16). Combined with our previous results, this data shows that while both the habenula and IPN contain neurons that respond to visual motion and neurons that respond to light position, only the representation in the IPN of the latter requires an intact habenula.
a An example plane showing correlation values of IPN cells with light in a particular position before and after NFP treatment for an Ntr+ (top) and a control (bottom) fish. b Maximum correlation maps for all fish in the dataset. In each panel, each pixel shows the maximum absolute correlation value across all fish, planes and landmark positions (n(Ntr + )=8, n(Ntr-)=8). c Left, distribution of correlation values of IPN neurons with the RF regressors before (blue) and after (purple) NFP treatment for Tg(16715:Gal4; UAS:Ntr-mCherry) fish (habenula ablated group, mean ± sd, n(fish)=8). The variance of the pre-ablation distribution is significantly larger than the variance of the post ablation distribution (Wilcoxon signed-rank test, p = 0.004). Right, distribution of correlation values of IPN neurons with the sensory regressors before (blue) and after (purple) NFP treatment for Ntr-fish (control group, mean ± sd, n(fish)=8). The variance of the pre-ablation distribution is not significantly larger than the post ablation (one-sided Wilcoxon signed-rank test, p = 0.9). d Number of reliably tuned neurons in the IPN before and after NFP treatment for Ntr+ (left) and control (right) fish. NFP significantly reduces the number of responsive cells in the IPN in the Ntr+ group (Wilcoxon signed-rank test, p = 0.004, n(fish)=8) but not in the control group (one-sided Wilcoxon signed-rank test, p = 0.87, n(fish)=8). Cell count was done by counting all reliably responding neurons in the IPN (neurons with a reliability threshold > 0.25).
The habenula is not required for the heading direction network
Recently, the habenula was found to directly contact HDNs in the zebrafish aHB55. In addition, our data shows that the habenula provides visual information that could be used for landmark based navigation to the IPN (Fig. 6). In rodents, a small part of habenular neurons respond to changes in angular heading velocity and could contribute to the construction of the heading signal39,45,56.
We next wanted to check if the habenula is necessary for the heading direction network in the aHB to function in darkness. We used two-photon ablations in order to sever the fasciculus retroflexus (FR), the bundles of axons coming from the habenula to the IPN (Fig. 7a). The day after ablation we imaged the fish and checked that the habenular innervation was abolished. We next imaged the aHB of ablated fish and were able to detect the heading direction network (Fig. 7b–d). Figure 7c shows the activity of the network in three different fish with ablated habenula. All these fish show an activity bump that moves around the aHB in a way that is correlated with the fish’s estimated heading direction (mean ± sem: −0.88 ± 0.02, n = 3), as previously described5. This data shows that habenular input is not necessary for the basic functioning of the ring attractor network or for this network to integrate motor commands and estimate the heading direction of the fish. Combined, our results show that while habenula neurons represent different types of signals, their input to the zebrafish heading direction network is specific, in the case of the stimuli we tested, to visual landmark stimuli.
a Left, z projection of the brain of a Tg(gad1b:Gal4; 16715:Gal4; UAS:GCaMP6s) fish expressing GCaMP6s both in habenular neurons and GABAergic neurons before two-photon ablation. Right, z projection of the brain of a different fish following a two-photon ablation of the FR. Red arrows indicate the two sites of laser ablation. Note habenular axons are gone from the IPN. b aHB anatomy overlaid with HDNs colored according to their phase. c Neural dynamics following two-photon ablation of habenular input in three fish. For each fish: top, tail trace showing the motor activity of the fish during the experiment. Bottom, traces of heading direction neurons sorted according to their phase showing the activity bump changing as the fish turns. Black scale bar for each fish indicated 100 seconds of recording.
Discussion
In this study we investigated the neural representation of visual information in the habenula - IPN - aHB circuit. We focus on two types of visual information that are relevant to navigation: directional whole field motion and landmark position. We find that both signals are represented in the dIPN, the region containing the neuropil of HDNs in zebrafish. These representations are topographically organized in parasagittal stripes along the dIPN, in a similar manner to the organization of the HD neuropil in this region. We further show that the habenula is necessary for the representation of landmark position in the dIPN, but not for the representation of visual motion or the basic function of the heading direction neurons in the aHB, namely, the integration of motor commands to represent the heading direction of the fish. Our findings suggest that the dIPN is a site of integration of visual information with the heading signal.
We show that in the aHB, visual motion and heading direction are represented by two separate populations. We further show that representation of visual motion in GABAergic neuropil in the dIPN, most of which comes from the aHB, has a similar structure to the representation of heading direction. In both cases, tuning appears in a striped pattern. Furthermore, the tuning appears in both sides of the dIPN and could be the result of aHB neurons that have a dendrite on one side and an axon on the other. Recent papers have shown different populations of cells in the zebrafish aHB that differ by expression of biochemical markers or morphology55,57. Interestingly, cells that express different markers were found to target different regions of the IPN and show different activity patterns57. Future studies are needed to conclude if this is also the case for the HDNs and the visually tuned neurons in this region.
Our data shows that different types of visual information are represented in the dIPN in stripes. We show that the anatomical organization of directional visual motion in the dIPN is consistent across fish, unlike the HD signal. When imaging habenular axons in the dIPN or the neuropil of dIPN neurons we see that each direction is represented on a single stripe in the rostro-caudal axis. The topographic organization of tuning in habenular axons is surprising given their morphology. Habenular axons wrap around the IPN and do not form localized projections within the IPN51. Previous studies have shown that the activity of habenula axons in the IPN is shaped by axoaxonic synapses58,59. Specifically, habenula axons have been shown to be modulated by GABAergic input in the IPN59. It is possible that similar modulation exists in the context of visual information representation in the IPN. Such GABAergic modulation could stem from local IPN neurons, aHB neurons that arborize in the IPN55 or a different population that targets the IPN such as the nucleus incertus57.
The representation of landmark position in the dIPN neuropil is also striped, such that each stripe represents a different position. In most fish we find that the topographic organization is consistent: the leftmost stripe is tuned to landmarks on the left while the rightmost stripe is tuned to landmarks on the right. However, in some fish we noticed a phase shift of this organization. Interestingly, when we imaged habenula axons in the dIPN, we find that landmark representation is consistent, and never identified a phase shift. These findings suggest that IPN cells and their neuropil could encode different types of landmark related information via different subpopulations of cells. It appears that some cells represent landmark position in egocentric coordinates and with respect to the fish orientation. Other cells are tied to the heading direction signal, which is allocentric, and thus represent landmark position with respect to the animal heading. As light position information is transmitted from the retina through the eminentia thalami, to the habenula and finally to the IPN53, it undergoes several transformations from a simple receptive field response to a landmark that could be used to tether the heading direction system to the visual scene. It remains unclear how this transformation occurs and it should be investigated in future studies.
The IPN does not contain HDNs, yet it is a core part of the vertebrate heading direction system. In mammals it is bidirectionally connected with the DTN39 and in zebrafish it is the region in which HDNs are connected to one another5. Furthermore, IPN lesions in rats lead to impairment of both landmark based navigation and path integration, indicating the importance of this region in these processes60. IPN lesions also affect the stability of HDNs in the thalamus, suggesting that its effects are not restricted to the DTN but encompass the broader HD network61. Here, we show that different types of spatial signals are similarly organized in the dIPN: heading direction5, visual motion direction, and landmark position all share a striped pattern. The similar organization suggests the IPN functions as a site of spatial information integration for navigation. However, the mechanisms underlying this integration remain unclear. Interestingly, while representation of heading direction is allocentric and varies between fish, the representation of visual motion direction is consistent and the representation of landmark position is semi-consistent. In most fish, the representation of visual motion direction and landmark position is aligned: the leftmost stripe of the dIPN is tuned to leftward motion and landmarks positioned on the left side of the visual field. Several possible explanations exist for this alignment. First, the two signals may be aligned with the heading signal to allow for integration of heading with either visual motion (for estimation of traveling direction) or landmark position (for anchoring of the heading signal to the visual scene). Alternatively, the two signals may be directly aligned to one another to allow for integration of different aspects of the visual scene. Future experiments should be done in order to reveal what signals are integrated and the underlying mechanisms.
In our experiments, we characterized the representation of landmark position on the front visual field of fish, in the range of 120°. Surprisingly, this range seems to activate regions across the entire dIPN, even though zebrafish larvae have a much larger visual field62. This could suggest that the representation of landmarks in the dIPN is not uniform, with larger areas devoted to representation of frontal landmarks than to landmarks appearing on the back visual field. Indeed, such non-uniformity in spatial representation was identified in other brain regions, such as the pretectum63. Alternatively, there could be overlap between the representation of different receptive fields, or that the back visual field is represented elsewhere in the brain. Finally, as these recordings were obtained from developing animals in the larval stage, it is possible that the representation of visual information is still developing and would be different in adults.
The habenula is a major source of input to the IPN45,46,47,48, yet our findings reveal a very specific role for the habenula in the heading direction system. While the habenula does contain neurons responding to whole field visual motion, its absence does not affect the representation of that information in either the aHB or IPN. In addition, the habenula is not necessary for the generation of the heading signal, and in the absence of habenula input, fish can still integrate motor command information and use that to update their heading representation (as they do in the dark)5. However, the habenula does provide landmark information to the circuit, allowing it to tether the heading direction signal to the visual scene. Our data shows that ablation of the habenula reduces both the negative and positive correlation values of the activity of dIPN neurons with landmark position. As habenula input is mostly excitatory45,46,47,48, this suggests that this input is being modulated in the IPN, likely by GABAergic IPN or aHB neurons, though this should be investigated in further studies.
The striped pattern in which IPN responses are organized is very reminiscent of the insect central complex. The insect central complex is also a site of integration of different signals that are relevant for navigation, such as heading direction and the animal’s traveling direction37,38. While the insect HDNs themselves are not sensitive to translational visual motion, another type of cells in this region responds to visual motion in a particular direction37,38. Both types of cells tile regions of the central complex, their neuropil forming columns, each tuned to a different heading/ traveling direction. Our data shows that a similar organization exists also in the vertebrate brain.
In many animals, the heading direction signal is tethered to the outside world and can be updated by shifting a landmark to a new position3,21. This is crucial for navigation, as, while most animals are capable of using path integration, this mechanism is prone to suffer from accumulating errors. The strong effect of visual landmarks on spatial sensation is evident not only in the heading direction network, but also in heading-direction-modulated place cells in the rodent hippocampus64. The directional tuning of place cells is preserved in virtual reality, demonstrating its dependence on visual cues rather than vestibular information64. Furthermore, the visual tuning in the hippocampus is also present in immobile rats presented with a moving bar of light65. The existence of visual tuning, independent of the rat motor actions, demonstrates that the control of visual landmarks on spatial orientation extends beyond the classic heading direction network65.
In insects, the tethering of the heading direction signal to the external visual environment is done by ring neurons: neurons that show receptive field responses and carry this information to the ellipsoid body28,30. Unlike many other neurons in the insect heading direction system, ring neurons do not tile a specific wedge of the ellipsoid body, but rather form synapses throughout the structure. Through yet unknown mechanisms of plasticity, these neurons manage to convey information about landmark position to specific HDNs in different environments28.
The morphology of the larval zebrafish habenula neurons is strikingly similar to the morphology of the insect ring neurons. They both have a single axon that wraps around the target structure and forms synapses in a non-localized manner36,51. Interestingly, habenula ablation in rats impairs several cognitive functions, one of which is landmark based navigation66. Here we show that a group of habenula neurons, similarly to the drosophila ring neurons, respond to light in a particular part of the visual field. We further show that ablating the habenula results in absence of landmark responses in the IPN, showing that habenular input is necessary for this representation. These results imply that a group of habenula neurons provide landmark related information to the dIPN, where it is poised to be integrated into the zebrafish heading direction system. The similarities between larval zebrafish habenula neurons and drosophila ring neurons suggests that this information could be integrated via similar mechanisms, revealing further potential analogies between the insect and vertebrate heading direction systems67.
Methods
Zebrafish husbandry
All procedures related to animal handling were conducted following protocols approved by the Technische Universität München and the Regierung von Oberbayern (TVA # 55-2-1-54-2532-101-12 and TVA ROB-55.22532.Vet_02-24-5). Adult zebrafish (Danio rerio) from Tüpfel long fin (TL) strain were kept at 27.5-28 °C on a 14/10 light cycle, and hosted in a fish facility that provided full recirculation of water with carbon-, bio-and UV filtering and a daily exchange of 12% of water. Water pH was kept at 7.0-7.5 with a 20 g/liter buffer and conductivity maintained at 750-800 µS using 100 g/liter. Fish were hosted in 3.5 liter tanks in groups of 10 to 17 animals. Adults were fed with Gemma micron 300 (Skretting) and live food (Artemia salina) twice per day and the larvae were fed with Sera micron Nature (Sera) and ST-1 (Aquaschwarz) three times a day.
All experiments were conducted on 5-10 dpf larvae of yet undetermined sex (the sex of the fish cannot be determined at this developmental stage). The week before the experiment, one male and one female or three male and three female animals were left breeding overnight in a breeding tank (Tecniplast). The day after, eggs were collected in the morning, rinsed with water from the facility water system, and then kept in groups of 20-40 in 90 mm Petri dishes filled with 0.3x Danieau’s solution (17.4 mM NaCl, 0.21 mM KCl, 0.12 mM MgSO4, 0.18 mM Ca(NO3)2, 1.5 mM HEPES, reagents from Sigma-Aldrich) until hatching and in groups of 20 larvae in water from the fish facility afterwards. Larvae were kept in an incubator that maintained temperature at 28.5 °C and a 14/10 hour light/dark cycle, and their solution was changed daily. At 4 or 5 dpf, animals were lightly anesthetized with Tricaine mesylate (Sigma-Aldrich) and screened for fluorescence under an epifluorescence microscope. Animals positive for GCaMP6s/ mCherry fluorescence were selected for the imaging experiments. Animals older than 5 dpf were kept in small breeding tanks and fed daily with Sera micron Nature and Rotifers.
Transgenic animals
Tg(elavl3:H2B-GCaMP6s) fish were used for whole brain imaging experiments68. Imaging of GABAergic neurons was done using double transgenic animals expressing Tg(gad1b/GAD67:Gal4-VP16)mpn155 (referred to as Tg(gad1b:Gal4)) which drives expression in a subpopulation of GABAergic cells under gad1b regulatory elements69 and Tg(UAS:GCaMP6s)mpn10170. For chemogenetic ablation experiments, we used a transgenic line expressing three different elements: Tg(elavl3:H2B-GCaMP6s), Tg(16715:Gal4) and Tg(UAS:Ntr-mCherry)71. For imaging IPN neurons and their neuropil we used the double transgenic animals expressing Tg(S1168t:Gal4)72 and Tg(UAS:GCaMP6s)mpn101. For imaging the pan-neuronal neuropil in the IPN we imaged Tg(elavl3:GCaMP6s) fish73. All the transgenic animals were also mitfa-/-and thus lacked melanophores74.
Lightsheet imaging
Lightsheet experiments were done as previously described5,75. Briefly, animals were embedded in 2-2.5% low-melting point agarose (Thermofisher) in a custom lightsheet chamber with a glass coverslip sealed on the sides in the position where the beams of the lightsheet enters the chamber, and a square of transparent acrylic on the bottom, for behavioral tracking. The chamber was filled with water from the fish facility system and agarose was removed along the optic path of the lateral laser beam (to prevent scattering), and around the tail of the animal, to enable movements of the tail. After embedding, fish were left recovering 1 to 6 hours before the imaging session.
Imaging experiments were performed using a custom-built lightsheet microscope76 as previously described5,75. In whole brain experiments, 20 planes were acquired over a range of approximately 250 µm, slightly adjusted for every fish. The resulting imaging data had a resolution of 10 × 0.6 × 0.6 µm/voxel, and a temporal resolution of 2 Hz. In experiments imaging only the aHB, 8 planes were acquired over a range of approximately 100 µm, slightly adjusted for every fish. The resulting imaging data had a resolution of 10 × 0.6 × 0.6 µm/voxel, and a temporal resolution of 5 Hz.
Two-photon microscopy
Two-photon experiments were done as previously described5,43. Briefly, animals were embedded in 2-2.5% low-melting point agarose (Thermofisher) in 30 mm petri dishes. The agarose around the tail, caudal to the pectoral fins, was cut away with a fine scalpel to allow for tail movement. The dish was placed onto an acrylic support with a light-diffusing screen and imaged on a custom-built two-photon microscope. In experiments in which visual stimuli were projected in the front visual field of the fish, the front half of the petri dish was covered with a light diffusing paper. The costume Python package brunoise was used to control the microscope hardware77.
Full frames were acquired at 3 Hz in four, 0.83 μm spaced interlaced scans, which resulted in x and y pixel dimensions of 0.3 −0.6 μm (varying resolutions depending on field of view covered). After acquisition from one plane was done, the objective was moved downward by 2 −8 μm and the process was repeated.
Visual stimuli were generated using a custom written Python script with the Stytra package78, and were projected at 60 frames per second using an Asus P2E microprojector and a red long-pass filter (Kodak Wratten No.25) to allow for simultaneous imaging and visual stimulation. Fish were illuminated using infrared light-emitting diodes (850 nm wavelength) and imaged from below at up to 200 frames per second using an infrared-sensitive charge-coupled device camera (Pike F032B, Allied Vision Technologies). Tail movements were tracked online using Stytra.
Tail tracking and stimulus presentation
To monitor tail movements during the imaging session, an infrared LED source (RS Components, UK) was used to illuminate the larvae from above. A camera (Ximea, Germany) with a macro objective (Navitar, USA) was aimed at the animal through the transparent bottom of the lightsheet chamber with the help of a mirror placed at 45° below the imaging stage. A longpass filter (Thorlabs, USA) was placed in front of the camera. A projector (Optoma, Taiwan) was used to display visual stimuli; light from the projector was conveyed to the stage through a cold mirror that reflected the projected image on the 45°-mirror placed below the stage. The stimuli were projected on a white paper screen positioned below the fish, with a triangular hole that kept the fish visible from the camera. The behavior tracking part of the rig was very similar to the setup for restrained fish tracking described in78.
Frames from the behavioral camera were acquired at 400 Hz and tail movements were tracked online using Stytra78 with Stytra’s default algorithm to fit to the tail 9 linear segments. The “tail angle” quantity used for controlling the closed-loop was computed online during the experiment in the Stytra program as the difference between the average angle of the first two and last two segments of the tail and saved with the rest of the log from Stytra. The stimulus presentation and the behavior tracking were synchronized with the imaging acquisition with a ZMQ-based trigger signal supported natively by Stytra.
Visual stimulation
To study neural responses to visual motion we presented fish with a pink noise pattern from below. The pattern could move in 8 different directions with even 45 degrees spacing. In each trial, the pattern moved in one direction (chosen randomly) for 10 seconds and then paused for 5/10 seconds before the next trial.
To study neural responses to landmark position, we presented fish with a bar of light in its front visual field (in the range ±60°). The fish was embedded in a plastic dish and the front half of the dish was covered with filter paper. A projector was placed in front of the dish as illustrated in Fig. 5. A red rectangle appeared in one of 8 possible locations in the fish’s front visual field in a random order. The red rectangle appeared for 5/10 seconds and then disappeared for 5/10 seconds before appearing again in a new location. For these experiments we chose to use two-photon microscopy, as in a lightsheet microscope the fish can see the blue laser which could be perceived as an additional landmark.
Chemogenetic ablations
For chemical ablations of habenular neurons we used fish expressing three transgenic elements: Tg(elavl3:H2B-GCaMP6s), Tg(16715:Gal4), and Tg(UAS:Ntr-mCherry). In these fish, Ntr is only expressed in the habenula. Ntr+ and Ntr- fish were imaged before ablation at 5-6 dpf. Following the first imaging session, fish were left in a 5 µM NFP solution in a light protected box for 16 hours as previously described71,79. Fish were washed several times and left to recover for 24 hours before they were imaged again at 7-8 dpf.
To check for the completion of chemical ablations of habenular neurons we used a confocal microscope to image fish before and after habenula ablations. For confocal experiments, larvae were embedded in 2% agarose and anesthetized with Tricaine mesylate (Sigma-Aldrich). Whole brain stacks of 5 dpf fish expressing Tg(elavl3:H2B-GCaMP6s), Tg(16715:Gal4), and Tg(UAS:Ntr-mCherry) transgenes were acquired using a 10x water immersion objective (NA = 0.45) with a voxel resolution of 1 × 0.83 × 0.83 µm (LSM 880, Carl Zeiss, Germany). Stacks of the IPN were acquired using a 20x water immersion objective (NA = 1) with a voxel resolution of 1 × 0.28 × 0.28 µm. The fish were freed from the agarose, treated with NFP for 16 hours and imaged again with identical parameters on 7 dpf.
The confocal experiments detailed above show that our ablation protocol ablated all habenula cells expressing Ntr. To ensure a large fraction of habenula neurons are labeled by our transgenic line (Tg(16715:Gal4)), we next imaged fish expressing three transgenic elements: Tg(elavl3:H2B-mCherry), Tg(16715:Gal4), and Tg(UAS:GCaMP6s). In these fish GCaMP6s is only expressed in the habenula and mCherry is expressed in all neurons. We used a confocal microscope (as described above) to image the habenula of 6 fish at 6 dpf, using a 20x water immersion objective (NA = 1) with a voxel resolution of 1 × 0.21 × 0.21 µm.
Our confocal stacks show that the Tg(16715:Gal4) line labels most habenula neurons. We would like to point out that even though this line does not label all neurons in the habenula, our data shows that the relevant neurons in the habenula were ablated in each one of our experiments, for the following reasons:
-
1.
In our experiments investigating ablation effects on representation of visual motion (Fig. 5), we performed whole brain imaging, allowing us to see the effects of the ablation not only on the IPN and aHB, but also on the habenula itself. The figure clearly shows that following habenula ablation, the representation of visual motion in this region is gone. Thus, even if some cells survived the ablation, cells tuned to visual motion are no longer present in the habenula, showing that this information in this region is not the source of visual motion information in the aHB or IPN.
-
2.
In our experiments investigating ablation effects on representation of landmark information (Fig. 6), we only imaged the IPN. In each fish we checked that the habenula ablation was successful, yet we did not take stacks to quantify the completeness of the ablation. However, as the effect of habenula ablation on light responses in the IPN is so strong, it is not likely that visually tuned cells remained in the habenula of ablated fish.
Two photon laser ablations
For two-photon laser ablations of the fasciculus retroflexus (FR) tracts of habenular axons going in the IPN we used fish expressing three transgenic elements: Tg(gad1b:Gal4), Tg(16715:Gal4), and Tg(UAS:GCaMP6s). In these fish GCaMP6s is expressed in the habenula and in GABAergic neurons in the brain. The labeling of the FR tracts allowed us to target the two-photon laser to a thin section (5×1 µm) of each of the two tracts. This section was targeted with the two-photon laser at 130 mW for 200 ms. Each tract was cut twice, about 20 and 40 µm rostral to the IPN. Following laser ablation, the fish were freed from agarose and left to recover overnight at 28 degrees with available food. The following day, fish health was assessed and only fish which were active and fed were chosen for the following experiments. Fish were embedded again and a whole brain stack was acquired to ensure that habenular axons were ablated and missing from the IPN. Next the GABAergic neurons in the aHB were imaged for detection of the heading direction network in ablated fish.
Imaging data preprocessing
The imaging stacks were saved in hdf5 files and then directly fed into suite2p, a Python package for calcium imaging data registration and ROI extraction80. We did not use suite2p algorithms for spike deconvolution. Parameters used for registration and source extraction in suite2p can be found in the shared analysis code. The parameters used by the suite2p algorithm were different based on the microscope used and the transgenic line. From the raw F traces saved from suite2p (F.npy file), ∆F/F_baseline was calculated taking F_baseline as the average fluorescence in a rolling window of 900 s, to compensate for some small amount of bleaching that was observed in some acquisition. The signal then was smoothed with a median filter from scipy (medfilt from scipy.signal), and Z-scored so that all traces were centered on 0 and normalized to a standard deviation of 1. The coordinate of each ROI was taken as the centroid of its voxels.
Reliability index
Reliability index was calculated as previously described81. Briefly, we calculated for each ROI the average correlation of the responses across all individual presentations of the presented stimuli. To use an objective criterion to select responsive cells, we used Otsu’s method from the SciPy package to set a threshold on the obtained histogram.
Regressor analysis
Regressor analysis was done as previously described82. Briefly, regressors were generated from stimulus related variables. In the whole field visual motion experiments, we constructed 8 regressors, one for each direction of motion. In the landmark experiments we constructed 8 regressors, one for each landmark position. The regressors were convolved with an exponential decay kernel that was found to fit our data. The constructed regressors were correlated with traces extracted from segmented ROIs or individual pixels (depending on the analyzed data).
Tuning maps
Tuning maps were generated using the results from the regressor analysis described above. For each analyzed pixel/ ROI we had 8 different correlation values with the 8 regressors. For generation of tuning maps, each pixel/ ROI was assigned two values: angle and amplitude. The angle represents the preferred direction of each pixel by indicating which direction elicits the strongest response, and is indicated in the maps by the pixel’s hue. The amplitude represents the magnitude of directional tuning and is indicated in the maps by the pixel’s saturation.
Quantification of neural tuning to landmark position
To characterize the representation of landmark position in the habenula, tuning curves were generated for each left habenula neuron based on the fluorescence measured when the landmark was presented in different positions in the range ±60 degrees. Each tuning curve was fit with a von Mises function using least squares optimization.
Voxel-wise differences
For the voxel-wise quantification of neuronal tuning differences as a result of habenula ablations (Supplementary Fig. 12), the MapZeBrain83 reference brain was first split in cubic voxels of 5 μm per side. Then for each one of the four imaging sessions (Ntr+ and control fish, for both pre-and post-ablation imaging sessions) all detected ROIs were assigned to their corresponding voxel containing them, and the final tuning amplitude for each voxel was computed as the average amplitude from all ROIs contained in it (top and middle rows in Supplementary Fig. 12). The difference in tuning amplitude between pre-and post-ablation imaging sessions were computed as the voxel-wise difference of this value across the two experimental sessions (bottom rows in Supplementary Fig. 12).
Anatomical registrations
For whole brain lightsheet experiments, all individual brains were registered to the MapZeBrain atlas reference brain83. Brain registration was performed on the anatomical stacks obtained from averaging selected frames in the corresponding dataset along the temporal dimension. First, a manual registration was performed using a custom napari-based GUI84. Results from this initial alignment were then used as the initial registration and fed into an ANTsPy registration pipeline that included both affine and diffeomorphic transformations85. In order to manipulate the different anatomical spaces, the brainglobe-space package from Brain-Globe86 was used. Coordinates for each ROI were computed as the centroid of all of its encompassing voxels. Registration of Two-photon was done using a similar pipeline. Two-photon datasets were not registered to the MapZeBrain atlas but to one of the fish from the specific dataset. Selection of cells belonging to a particular brain region in Fig. 1g was done using manually drawn masks drawn using a napari-based GUI.
Data analysis and statistics
All parts of the data analysis were performed using Python 3.7, and Python libraries for scientific computing, in particular Numpy, Scipy and Scikit-learn. The Python environment required to replicate the analysis in the paper can be found in the paper code repository. All figures were produced using Matplotlib. All the analysis code will be available upon publication.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All source data used in the functional imaging analysis (dF/F traces, ROI coordinates, behavioral tracking traces, and stimulus logs from Stytra) and for the anatomical observations (confocal stacks) are available at: https://doi.org/10.5281/zenodo.16180143.
Code availability
All scripts for stimuli generation, data pre-processing, analysis and plots generation are available at: https://github.com/portugueslab/Lavian_et_al_2025.
References
Seelig, J. D. et al. Neural dynamics for landmark orientation and angular path integration. Nature 521, 186–191 (2015).
Taube, J. S. et al. Head-Direction Cells Recorded from the Postsubiculum in Freely Moving Rats. I. Description and Quantitative Analysis. The Journal of Neuroscience (1990).
Vinepinsky, E. et al. Representation of edges, head direction, and swimming kinematics in the brain of freely-navigating fish. Sci. Rep. 10, 1–16 (2020).
Yartsev, M. M. et al. Grid cells without theta oscillations in the entorhinal cortex of bats. Nature 479, 103–107 (2011).
Petrucco, L. et al. Neural dynamics and architecture of the heading direction circuit in zebrafish. Nat. Neurosci. 26, 765–773 (2023).
Cullen, K. E. et al. Our sense of direction: Progress, controversies and challenges. Nat. Neurosci. 20, 1465–1473 (2017).
Hulse, B. K. et al. Annual Review of Neuroscience Mechanisms Underlying the Neural Computation of Head Direction. https://doi.org/10.1146/annurev-neuro-072116 (2019)
Beugnon, G. et al. Homing in the field cricket, Gryllus campestris. J. Insect Behav. 2, 187–198 (1989).
Blair, H. T. et al. Visual and vestibular influences on head-direction cells in the anterior thalamus of the rat. Behav. Neurosci. 110, 643–660 (1996).
Mittelstaedt, H. et al. Homing by path integration. Proc. Life Sci. 119–156 https://doi.org/10.1007/978-3-642-68616-0_29 (1982).
Stackman, R. W. et al. Firing properties of head direction cells in the rat anterior thalamic nucleus: Dependence on vestibular input. J. Neurosci. 17, 4349–4358 (1997).
Taube, J. S. The head direction signal: Origins and sensory-motor integration. Annu. Rev. Neurosci. 30, 181–207 (2007).
Goodridge, J. P. et al. Cue control and head direction cells. Behav. Neurosci. 112, 749–761 (1998).
Goodridge, J. P. et al. Preferential Use of the Landmark Navigational System by Head Direction Cells in Rats. Behav. Neurosci. 109, 49–61 (1995).
Chen, L. L. et al. Head-direction cells in the rat posterior cortex - I. anatomical distribution and behavioral modulation. Exp. Brain Res. 101, 8–23 (1994).
Sharp, P. E. et al. Angular velocity and head direction signals recorded from the dorsal tegmental nucleus of gudden in the rat: Implications for path integration in the head direction cell circuit. Behav. Neurosci. 115, 571–588 (2001).
Stackman, R. W. et al. Firing properties of rat lateral mammillary single units: Head direction, head pitch, and angular head velocity. J. Neurosci. 18, 9020–9037 (1998).
Brown, J. E. et al. Polysynaptic pathways from the vestibular nuclei to the lateral mammillary nucleus of the rat: Substrates for vestibular input to head direction cells. Exp. Brain Res. 161, 47–61 (2005).
Senzai, Y. et al. The brain simulates actions and their consequences during REM sleep. https://doi.org/10.1101/2024.08.13.607810 (2024).
Peyrache, A. et al. Internally organized mechanisms of the head direction sense. Nat. Neurosci. 18, 569–575 (2015).
Taube, J. S. et al. Head-direction cells recorded from the postsubiculum in freely moving rats. II. Effects of environmental manipulations. J. Neurosci. 10, 436–447 (1990).
Sit, K. K. et al. Coregistration of heading to visual cues in retrosplenial cortex. Nat. Commun. 14, (2023).
Yoder, R. M. et al. Origins of landmark encoding in the brain. Trends Neurosci. 34, 561–571 (2011).
Jacob, P. Y. et al. An independent, landmark-dominated head-direction signal in dysgranular retrosplenial cortex. Nat. Neurosci. 20, 173–175 (2017).
Green, J. et al. A neural circuit architecture for angular integration in Drosophila. Nature 546, 101–106 (2017).
Kim, S. S. et al. Ring attractor dynamics in the Drosophila central brain. Sci. (80-.) 356, 849–853 (2017).
Green, J. et al. A neural heading estimate is compared with an internal goal to guide oriented navigation. Nat. Neurosci. 22, 1460–1468 (2019).
Fisher, Y. E. et al. Sensorimotor experience remaps visual input to a heading-direction network. Nature 576, 121–125 (2019).
Hulse, B. K. et al. A connectome of the drosophila central complex reveals network motifs suitable for flexible navigation and context-dependent action selection. Elife 10, 1–180 (2021).
Seelig, J. D. et al. Feature detection and orientation tuning in the Drosophila central complex. Nature 503, 262–266 (2013).
Okubo, T. S. et al. A Neural Network for Wind-Guided Compass Navigation. Neuron 107, 924–940.e18 (2020).
Sun, X. et al. How the insect central complex could coordinate multimodal navigation. Elife 10, 1–21 (2021).
Turner-Evans, D. et al. Angular velocity integration in a fly heading circuit. https://doi.org/10.7554/eLife.23496.001 (2017).
Hulse, B. K. et al. A rotational velocity estimate constructed through visuomotor competition updates the fly’s neural compass. bioRxiv 2023.09.25.559373 (2023).
Hanesch, U. et al. Neuronal architecture of the central complex in Drosophila melanogaster. Cell Tissue Res 257, 343–366 (1989).
Wolff, T. et al. Neuroarchitecture and neuroanatomy of the Drosophila central complex: A GAL4-based dissection of protocerebral bridge neurons and circuits. J. Comp. Neurol. 523, 997–1037 (2015).
Lyu, C. et al. Building an allocentric travelling direction signal via vector computation. Nature 601, 92–97 (2022).
Lu, J. et al. Transforming representations of movement from body- to world-centric space. Nature 601, 98–104 (2022).
Groenewegen, H. J. et al. Cytoarchitecture, fiber connections, and some histochemical aspects of the interpeduncular nucleus in the rat. J. Comp. Neurol. 249, 65–102 (1986).
Liu, R. et al. The dorsal tegmental nucleus: an axoplasmic transport study. Brain Res 310, 123–132 (1984).
Shibata, H. et al. Efferent projections of the interpeduncular complex in the rat, with special reference to its subnuclei: a retrograde horseradish peroxidase study. Brain Res 296, 345–349 (1984).
Chen, X. et al. Brain-wide Organization of Neuronal Activity and Convergent Sensorimotor Transformations in Larval Zebrafish. Neuron 100, 876–890.e5 (2018).
Dragomir, E. I. et al. Evidence accumulation during a sensorimotor decision task revealed by whole-brain imaging. Nat. Neurosci. 23, 85–93 (2020).
Naumann, E. A. et al. From Whole-Brain Data to Functional Circuit Models: The Zebrafish Optomotor Response. Cell 167, 947–960.e20 (2016).
Contestabile, R. A. et al. Afferent connections of the interpeduncular nucleus and the topographic organization of the habenulo-interpeduncular pathway: An HRP study in the rat. J. Comp. Neurol. 196, 253–270 (1981).
Gottesfeld, Z. et al. Cholinergic projection of the diagonal band to the interpeduncular nucleus of the rat brain. Brain Res 156, 329–332 (1978).
Kataoka, K. et al. Habenulo-interpeduncular tract: a possible cholinergic neuron in rat brain. Brain Res 62, 264–267 (1973).
Villani, L. et al. Cholinergic projections in the telencephalo-habenulo-interpeduncular system of the goldfish. Neurosci. Lett. 76, 263–268 (1987).
Aizawa, H. et al. Laterotopic Representation of Left-Right Information onto the Dorso-Ventral Axis of a Zebrafish Midbrain Target Nucleus. Curr. Biol. 15, 238–243 (2005).
Gamse, J. T. et al. Directional asymmetry of the zebrafish epithalamus guides dorsoventral innervation of the midbrain target. Development 132, 4869–4880 (2005).
Bianco, I. H. et al. Brain asymmetry is encoded at the level of axon terminal morphology. Neural Dev. 3, (2008).
Dreosti, E. et al. Left-Right Asymmetry Is Required for the Habenulae to Respond to Both Visual and Olfactory Stimuli. Curr. Biol. 24, 440–445 (2014).
Zhang, B. et al. Left Habenula Mediates Light-Preference Behavior in Zebrafish via an Asymmetrical Visual Pathway. Neuron 93, 914–928.e4 (2017).
Zhao, H. et al. Circadian firing-rate rhythms and light responses of rat habenular nucleus neurons in vivo and in vitro. Neuroscience 132, 519–528 (2005).
Wu, Y. K. et al. Anatomical and functional organization of the interpeduncular nucleus in larval zebrafish. bioRxiv https://doi.org/10.1101/2024.10.09.617353 (2024).
Sharp, P. E. et al. Movement-related correlates of single cell activity in the interpeduncular nucleus and habenula of the rat during a pellet-chasing task. Behav. Brain Res. 166, 55–70 (2006).
Spikol, E. D. et al. Genetically defined nucleus incertus neurons differ in connectivity and function. Elife 12, 1–33 (2024).
Zaupa, M. et al. Trans-inhibition of axon terminals underlies competition in the habenulo-interpeduncular pathway. Curr. Biol. 31, 4762–4772.e5 (2021).
Paoli, E. et al. Modulation of habenula axon terminals supports action-outcome associations in larval zebrafish. (2025).
Clark, B. J. et al. Deficits in Landmark Navigation and Path Integration After Lesions of the Interpeduncular Nucleus. Behav. Neurosci. 123, 490–503 (2009).
Clark, B. J. et al. Head direction cell instability in the anterior dorsal thalamus after lesions of the interpeduncular nucleus. J. Neurosci. 29, 493–507 (2009).
Easter, S. S. Jr. et al. The Development of Vision in the Zebrafish. Dev. Biol. 180, 646–663 (1996).
Wang, K. et al. Parallel Channels for Motion Feature Extraction in the Pretectum and Tectum of Larval Zebrafish. Cell Rep. 30, 442–453.e6 (2020).
Acharya, L. et al. Causal Influence of Visual Cues on Hippocampal Directional Selectivity. Cell 164, 197–207 (2016).
Purandare, C. S. et al. Moving bar of light evokes vectorial spatial selectivity in the immobile rat hippocampus. Nature 602, 461–467 (2022).
Lecourtier, L. et al. Habenula lesions cause impaired cognitive performance in rats: Implications for schizophrenia. Eur. J. Neurosci. 19, 2551–2560 (2004).
Heinze, S. Neuroscience: Fish and fly headed in the same direction. Curr. Biol. 33, R677–R679 (2023).
Freeman, J. et al. Mapping brain activity at scale with cluster computing. Nat. Methods 11, 941–950 (2014).
Förster, D. et al. Genetic targeting and anatomical registration of neuronal populations in the zebrafish brain with a new set of BAC transgenic tools. Sci. Rep. 7, 1–11 (2017).
Thiele, T. R. et al. Descending Control of Swim Posture by a Midbrain Nucleus in Zebrafish. Neuron 83, 679–691 (2014).
Palieri, V. et al. The preoptic area and dorsal habenula jointly support homeostatic navigation in larval zebrafish. Curr. Biol. 34, 489–504.e7 (2024).
Scott, E. K. et al. The cellular architecture of the larval zebrafish tectum, as revealed by Gal4 enhancer trap lines. Front. Neural Circuits 3, (2009).
Kim, D. H. et al. Pan-neuronal calcium imaging with cellular resolution in freely swimming zebrafish. Nat. Methods 14, 1107–1114 (2017).
Lister, J. A. et al. Nacre encodes a zebrafish microphthalmia-related protein that regulates neural-crest-derived pigment cell fate. Development 126, 3757–3767 (1999).
Markov, D. A. et al. A cerebellar internal model calibrates a feedback controller involved in sensorimotor control. Nat. Commun. 12, 1–21 (2021).
Štih, V. et al. Sashimi (v0.2.1). (2020). https://doi.org/10.5281/zenodo.5932227
Štih, V. et al. portugueslab/brunoise: Alpha (0.1). (2020). https://doi.org/10.5281/zenodo.4122063
Štih, V. et al. Stytra: An open-source, integrated system for stimulation, tracking and closed-loop behavioral experiments. PLoS Comput. Biol. 15, 1–19 (2019).
Bergemann, D. et al. Nifurpirinol: A more potent and reliable substrate compared to metronidazole for nitroreductase-mediated cell ablations. Wound Repair Regen. 26, 238–244 (2018).
Pachitariu, M. et al. Suite2p: beyond 10,000 neurons with standard two-photon microscopy. bioRxiv. Bioarxiv 20, 2017 (2017).
Prat, O. et al. Comparing the Representation of a Simple Visual Stimulus across the Cerebellar Network. eNeuro 11, 1–15 (2024).
Portugues, R. et al. Whole-brain activity maps reveal stereotyped, distributed networks for visuomotor behavior. Neuron 81, 1328–1343 (2014).
Kunst, M. et al. A Cellular-Resolution Atlas of the Larval Zebrafish Brain. Neuron 103, 21–38.e5 (2019).
Sofroniew, N. et al. napari: a multi-dimensional image viewer for Python. https://doi.org/10.5281/zenodo.3555620 (2019)
Tustison, N. J. et al. The ANTsX ecosystem for quantitative biological and medical imaging. Sci. Rep. 11, 9068 (2021).
Claudi, F. et al. BrainGlobe Atlas API: a common interface for neuroanatomical atlases. J. Open Source Softw. 5, 2668 (2020).
Acknowledgements
We thank all the members of the Portugues lab members for their input. H.L. would like to thank Inbal Shainer and Barak Shalom for helpful discussions. We thank Herwig Baier for sharing the Tg(gad1b/GAD67:Gal4-VP16)mpn155 and Tg(UAS:GCaMP6s)mpn101 transgenic lines. This research was funded by the German Research Foundation (DFG) under Germany’s Excellence Strategy within the framework of the Munich Cluster for Systems Neurology (EXC 2145 SyNergy, identifier 390857198) and through the “Enhanced resolution microscopy” project DFG – Projektnummer 518284373, by the Volkswagen Stiftung via a Life? grant and by the Max Planck Foundation.
Author information
Authors and Affiliations
Contributions
The conception and design of the study was done by H.L. and R.P. Experiments were done by H.L. and O.P with help from V.S. and L.P. Data analysis was done by H.L. with help from O.P., V.S., and L.P. The paper was written by H.L. and R.P. with help from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interest.
Peer review
Peer review information
Nature Communications thanks Yu Mu and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lavian, H., Prat, O., Petrucco, L. et al. Visual motion and landmark position align with heading direction in the zebrafish interpeduncular nucleus. Nat Commun 16, 9924 (2025). https://doi.org/10.1038/s41467-025-66084-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66084-1
This article is cited by
-
Plastic landmark anchoring in zebrafish compass neurons
Nature (2026)