Abstract
Coagulase-negative staphylococci are dominant human skin colonizers, producing natural products that shape the community and prevent pathogen colonization. The molecular mechanisms by which these natural products mediate interbacterial competition are not fully understood. Here, we identify a plasmid-borne daptide bacteriocin (hominicin) from a human skin isolate of Staphylococcus hominis, which features an unusual N2-N2-dimethyl-1,2-propanediamine C-terminus. Heterologous expression of the reconstituted biosynthetic loci yields a daptide product of the same molecular mass that exhibits antimicrobial activity against the skin pathogen Staphylococcus aureus, with amino-modified termini being essential for activity. Membrane permeability and voltage-clamp lipid bilayer experiments support a mechanism by which the daptide rapidly dissipates the transmembrane potential by forming peptidic channels. Additionally, we identify a cognate homI gene that confers resistance against membrane damage. Finally, the purified daptide effectively protects mouse skin from S. aureus-induced epicutaneous injury, as evidenced by reduced bacterial burden, inflammation, and transepithelial water loss, highlighting its therapeutic potential for treating bacterial skin infections. Our findings elucidate a mechanism of action, biosynthesis, and resistance for a staphylococcal bacteriocin belonging to a class of natural products called daptides.
Similar content being viewed by others
Introduction
Human skin serves as both a physical and immunologic barrier between the body and the environment; a hostile landscape characterized by dryness, low pH, and high levels of salt1. Yet, the skin microbiota stably colonizes in this niche and plays an integral role in shaping the diversity of the community, preventing pathogen colonization, fortifying the epithelial barrier, and priming immune responses2. Coagulase-negative staphylococci (CoNS) are prominent members of the skin microbiome, comprising a heterogenous group of 38 known species3. CoNS maintain skin homeostasis by secreting natural products that antagonize microbial competitors and pathogens4. While the roles of CoNS in the microbiome have primarily focused on Staphylococcus epidermidis, emerging evidence underscores the beneficial roles of Staphylococcus hominis, a common CoNS on human skin, in defending against bacterial pathogens. S. hominis secretes autoinducing peptides that inhibit the accessory gene regulator (agr) system of the opportunistic skin pathogen Staphylococcus aureus, thereby preventing S. aureus-mediated inflammation and cutaneous damage5,6. Furthermore, S. hominis produces an arsenal of bacteriocins, including lantibiotics and other peptidic antibiotics, which inhibit S. aureus in vitro and in vivo7,8,9.
Bacteriocins are often ribosomally synthesized and post-translationally modified peptides (RiPPs) with targeted antibacterial activity against specific genera and species10. These RiPPs are generally encoded by complex and variable biosynthetic gene clusters (BGCs) that include the precursor bacteriocin, post-translationally modifying enzymes, and accessory proteins involved in export and producer immunity. Bacteriocin production is traditionally regarded as a probiotic trait, as it facilitates the producer strain in colonizing a niche, eliminating competing microbes, and potentially modulating the immune responses11. Contextually, the loss of bacteriocin-producing CoNS is associated with increased burden of S. aureus on the skin of subjects with atopic dermatitis (AD), underscoring the importance of commensal bacteria in fortifying the skin’s antimicrobial barrier7. In a post-antibiotic era, bacteriocins present a promising source of potent antimicrobials that have the potential to inhibit pathogens. Human clinical trials have demonstrated the safe delivery and effectiveness of lantibiotic-producing S. hominis as a bacteriotherapy treatment for S. aureus-colonized AD skin, highlighting the effect of a bacteriocin producer in reducing S. aureus colonization and disease severity12.
(Meta)genome-based and activity-based screening approaches continue to advance the discovery of RiPPs, which are categorized by class-defining chemical features13. Daptides emerged as a new class of RiPPs, characterized by an unusual (S)-N2-N2-dimethyl-1,2-propanediamine (Dmp)-modified C-terminus and defined in Microbacterium paraoxydans14,15,16. Daptide BGCs are widely distributed across the Actinomycetota, Bacillota, and Pseudomonadota phyla14, implying a potential biological role for these small peptides. Characterization of M. paraoxydans (Mpa) daptides revealed hemolytic activity against bovine erythrocytes, but no antibacterial properties were observed14. Since then, no other daptides have been isolated; however, an earlier report described hominicin17, a Dmp-carrying peptide, as antimicrobial, although its genetic and biosynthetic origin as a RiPP remains undetermined. Thus, the target specificity, mode of action, and mechanism of resistance for daptides have not been fully elucidated. Such knowledge is critical to the field of drug discovery for developing novel and rational interventions that reduce morbidity and mortality from bacterial infections.
Herein, we identify a daptide bacteriocin produced by a human skin isolate of S. hominis. The expression of the reconstituted biosynthesis genes sufficiently confers antimicrobial activity. We isolate the daptide from culture supernatant and solve its structure using mass spectrometry and nuclear magnetic resonance (NMR). Mechanistically, the daptide induces membrane damage by dissipating the transmembrane potential of susceptible bacteria through the formation of peptidic channels. Genetic studies identify a cognate protein, HomI, as the determinant for resistance against daptide-induced membrane damage. Finally, we demonstrate that topical treatment with the purified daptide reduces S. aureus colonization and cutaneous injury in a murine model of epicutaneous infection. This study elucidates a mechanism of action, biosynthesis, and resistance for a staphylococcal bacteriocin belonging to an understudied class of natural products called daptides.
Results
Antimicrobial activity of a skin commensal S. hominis strain AH5011
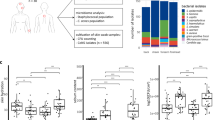
We screened the cell-free, conditioned media (CM) from a human skin isolate collection of S. hominis for antimicrobial activity and determined that the strain AH5011 (D11), an isolate from non-lesional skin of a subject with atopic dermatitis, suppressed the growth of S. aureus strain AH6350 (Fig. 1a, c). To determine the antimicrobial specificity of AH5011, we tested the susceptibility of skin commensals and pathogenic strains of Staphylococcus, including S. aureus, S. hominis, S. epidermidis, S. capitis, S. haemolyticus, S. lugdunensis, S. simulans, S. warneri, and S. pseudintermedius, as well as Micrococcus luteus and Streptococcus pyogenes (Fig. 1b, Supplementary Data 3). Streptococcus agalactiae and Enterococcus faecalis were included as non-skin-associated bacteria. Antimicrobial susceptibility was assessed by calculating the ratio of optical density of the test condition (40% v/v CM) to untreated cells grown in tryptic soy broth (TSB). We defined inhibition as a ratio of <0.5. This analysis revealed that AH5011 is inhibitory towards staphylococci, M. luteus, and S. pyogenes, but not S. agalactiae or E. faecalis. However, the inhibitory activity was strain dependent, as evident by a bimodal distribution of susceptibility for S. aureus, S. hominis, S. epidermidis, S. lugdunensis, and S. warneri. Intriguingly, this observation was not seen for S. agalactiae and E. faecalis. Therefore, we hypothesize that S. hominis AH5011 targets specific members of the skin community, largely the Staphylococcus genus. We observed that the antimicrobial function inhibits the S. hominis reference strain ATCC 27844 (AH4553) from the American Type Culture Collection in a concentration-dependent manner (Fig. 1c–e). This strain was adopted as a sensitive control strain for subsequent experiments. The antimicrobial activity is thermally stable and can be inactivated by the serine protease Proteinase K (Supplementary Fig. 1). Additionally, AH5011 is resistant to the inhibitory effect from its own CM (Fig. 1f, g), which indicates a self-protection mechanism.
a, d, f S. aureus strain AH6350 (a), S. hominis strains AH4553 (d), and AH5011 (f) were incubated with 40% CM from AH5011 or TSB for 8 h and monitored by cell density (OD600) and colony-forming units (CFUs). Data from n = 3 biological replicates are reported as the mean and standard deviation. Statistical significance in comparison to the TSB control group was determined using repeated measures two-way analysis of variance (ANOVA). Exact p values from left to right are as follows: a (CFU) 0.0320, 0.0488, a (OD600) < 0.0001, <0.0001, d (CFU) 0.0201, 0.0007, d (OD600) 0.0092, <0.0001, f (CFU) 0.3912, >0.9999, f (OD600) > 0.9999, >0.9999. b Antimicrobial susceptibility of Staphylococcus sp., Micrococcus luteus, Streptococcus pyogenes, Streptococcus agalactiae, and Enterococcus faecalis. “N” values indicate the number of bacterial strains tested which are listed in Supplementary Data 3. The data represents the mean of n = 2 biological replicates for each strain tested. c Representative images of spot-on-lawn bioactivity of AH5011. Scale bar = 10 mm. e, g S. hominis strains AH4553 (e) and AH5011 (g) were incubated in 40%, 20%, 10% CM from AH5011 or TSB for 8 h. Data from n = 3 biological replicates are reported as the mean and standard deviation. Statistical differences were compared to TSB control group at 8 h using ordinary one-way ANOVA followed by Dunnett’s multiple comparison test. Exact p values from top to bottom are as follows: e < 0.0001, <0.0001, <0.0001, g 0.9345, 0.8103, 0.7501. Source data are provided as a Source Data file.
S. hominis AH5011 bacteriocin is a member of the daptide family of RiPPs
The whole-genome sequence of AH5011 revealed two plasmids, p1 and p2, that are 19.8 kilo bases (kb) and 22 kb, respectively. Bacteriocin biosynthesis machineries are often found on genetically mobile plasmids18. To assess whether the inhibitory function was plasmid-encoded, we generated plasmid-cured mutants of AH5011 upon exposure to acriflavine and tested for loss of activity. The absence of p1, but not p2, led to a complete loss of activity, as evidenced by the lack of an inhibitory zone and detectable activity in the CM from the ∆p1 and ∆p1∆p2 mutants against the sensitive control strain AH4553 (Fig. 2a, Supplementary Fig. 2a). Furthermore, the growth of ∆p1 and ∆p1∆p2 mutants was attenuated in AH5011’s CM, but not that of the ∆p2 mutant (Supplementary Fig. 2b–d). Thus, we inferred that the p1 plasmid is the main contributor to the antimicrobial activity and resistance.
a Spot-on-lawn bioactivity of AH5011 and plasmid-cured mutants against a lawn of AH4553, with inhibition zone diameters represented by black dashed lines. Red dashed line indicates the limit of detection. Scale bar = 10 mm. Data represents the mean of n = 5 biological replicates and error bars show standard deviation. Statistical significance was determined using one-way ANOVA followed by Dunnett’s multiple comparison test. Exact p values from left to right: a < 0.0001, 0.4946, <0.0001. Source data are provided as a Source Data file. b Gene arrangement and annotation of the hom BGC that spans 9596 bp in length. c HomA encodes a 65-amino acid precursor peptide, consisting of an N-terminal leader and a C-terminal core sequences. d The proposed post-translational modifications of the core peptide, with the modified sites shown and the modifying enzymes indicated. Image was generated in BioRender. Orange dehydrobutyrine (Dhb), green dehydroalanine (Dha), blue d-alanines, red glutamate.
We analyzed the AH5011 plasmids using the BAGEL4 database19 and identified a gene cluster containing a lanthipeptide dehydratase and serine peptidase in the p1 plasmid. The 9.6-kb gene cluster consists of ten genes encoding the precursor peptide, seven biosynthesis enzymes, a peptidase, and a hypothetical protein (Fig. 2b, Supplementary Table 5). Based on sequence similarity with proteins of known functions and/or domains, homB encodes a domain of unknown function (DUF)-fused RiPP recognition element (RRE) protein; homC encodes an enoyl-(acyl carrier protein) reductase family member; homD encodes a class-III pyridoxal phosphate-dependent aminotransferase; homJ encodes a NAD(P)H-dependent oxidoreductase, belonging to the flavoprotein-like superfamily; homM1 and homM2 both encode S-adenosyl-l-methionine-dependent methyltransferases; homP encodes a serine peptidase, belonging to the subtilisin family; homMD encodes a DUF4135 domain-containing protein that represents only the dehydratase domain of class II lanthipeptide synthetases (LanM)20. Therefore, we proposed to designate this single domain enzyme as LanMD to indicate its proposed dehydration activity21,22.
RiPP precursor peptides consist of an N-terminal leader sequence, which serves as the binding element for modifying enzymes, and a C-terminal core sequence, which receives the post-translational modifications. The homA gene encodes a 65-residue precursor bacteriocin (Fig. 2c), with the core peptide mapping to the reported final structure of hominicin, a peptidic natural product that features the Dmp-modified C-terminus17. We detected differences at the 10th and 17th residues between the core peptide of the AH5011 bacteriocin and the final structure of hominicin (Supplementary Fig. 14d). Although the genetic and biosynthetic information of the hominicin-producing strain MBBL 2-917 are not publicly available, we infer that the AH5011 bacteriocin is likely hominicin. The Dmp-modified C-terminus is a class-defining feature of daptides, which has been characterized in M. paraoxydans14 (Supplementary Fig. 14a, b). The Dmp moiety arises from enzymatic modifications of a C-terminal threonine residue, beginning with the oxidative decarboxylation of the hydroxyl group to a ketone, catalyzed by the DUF-RRE and alcohol dehydrogenase DapBC14. This is followed by transamination via transaminase DapD and a dimethylation step by the methyltransferase DapM14. We propose that the hom BGC contains the necessary tailoring enzymes, HomBCDM2, that modify the C-terminal Thr into Dmp (Fig. 2d, Supplementary Fig. 14a). Additionally, the LanMD likely functions as a dehydratase to convert the core threonines into dehydrobutyrines and serines into dehydroalanines20,21,22. We anticipate that the flavin-dependent oxidoreductase HomJ likely catalyzes the reduction of dehydroalanines to d-alanines, as previously reported in the biosynthesis of lanthipeptides23,24,25. The modified core is proteolytically cleaved from the leader peptide by the serine peptidase HomP26 while the second methyltransferase HomM1 is likely responsible for the dimethylation of the free N-terminal isoleucine, which results in a fully modified and bioactive daptide.
Detection and structural characterization of AH5011 daptide
Using ultraperformance liquid chromatography-mass spectrometry (UPLC-MS), we analyzed the CM from daptide-positive AH5011 and daptide-negative ∆p1∆p2 strains. We detected a distinct chromatographic peak at retention time 5.32 min with a measured m/z value of 1020.1114 in the CM from AH5011 but absent from ∆p1∆p2. The measured mass was within 2 ppm of the predicted m/z (1020.1103) for a doubly-charged protonated ion of the daptide (Fig. 3a, Supplementary Fig. 3a). The observed MS-MS fragmentation pattern of this m/z in combination with the genetic data allowed us to propose the bacteriocin sequence as DmIle–Dhb–Pro–Ala–Dhb–Pro–Phe–Dhb–Pro–Ala–Ile–Thr–Glu–Ile–Dhb–Ala–Ala–Val–Ile–Ala–Dmp (Fig. 3c, d, Supplementary Fig. 3b). The initial hominicin investigation lacked genetic information for S. hominis strain MBBL 2-917, so the precursor peptide sequence was not proposed. However, the final modified bacteriocin structure remains consistent with the previously proposed structure of hominicin. The determination of the accurate mass of hominicin from S. hominis AH5011 confirmed its molecular weight to be 2038.5110 Da. 1D and 2D NMR experiments were carried out to confirm the proposed structure of AH5011 hominicin (Fig. 3e, Supplementary Fig. 6 and Supplementary Data 1).
a The base peak chromatograms of conditioned media from AH5011, ∆p1∆p2, and the TSB control. A peak matching the predicted m/z of the [M + 2H]2+ of the daptide bacteriocin (1020.1103) within 2 ppm of the measured mass (1020.1114) elutes at a retention time of 5.32 min, as indicated by the pink trace for AH5011. b Base peak chromatograms of conditioned media from S. aureus strains expressing homABCDIJM1M2PlanMD and empty vector. c MS-MS spectrum showing major fragments for the precursor ion calculated at m/z 1020.1103 identified as the bacteriocin [M + 2H]2+ ion in conditioned media from AH5011. d Molecular structure and annotated MS/MS fragments of the AH5011 hominicin in conditioned media with precursor ion m/z 1020.110. e Key HMBC, NOESY and COSY/TOCSY NMR correlations of AH5011 hominicin.
To characterize the configurations of hominicin’s amino acids, we performed a full hydrolysis of hominicin followed by Marfey’s derivatization27. The results (Supplementary Fig. 7) indicate that all hominicin amino acids are of l-configuration, with the exception of alanine, which was detected in both the l- and d-configurations with a ratio of 1.2:1 L:D (calculated from UPLC-MS peak area). These findings are consistent with the biosynthetic model that indicates a product containing three ribosomally derived l-alanines and two d-alanines derived from serines (L:D ratio of 1.5:1). The configuration of the stereocenter of the Dmp group on the C-terminus was not confirmed for hominicin, although previous work has shown this functional group to have an (S) configuration in other daptides14.
To determine whether the hom BGC from AH5011 is sufficient to produce the daptide, we cloned the 9.6-kb BGC into the plasmid pCM28 and heterologously expressed the construct in a restriction-deficient cloning host, S. aureus strain RN4220. From the CM of the hom BGC expressing S. aureus, but not the empty vector, an ion was detected with the same retention time (5.33 mins), accurate mass (1020.1115 m/z), and MS-MS fragmentation pattern as the ion detected in the AH5011 strain (Fig. 3b, Supplementary Fig. 3c, d). Thus, we inferred that the expression of the hom BGC was sufficient for the biosynthesis and secretion of hominicin.
Given the antimicrobial potency from CM and optimal aerobic growth conditions, we enriched hominicin through chloroform extraction, then subjected the material to flash chromatography, followed by preparative high-performance liquid chromatography (HPLC) (Supplementary Fig. 4). The peptide product (6.4 mg) was isolated from 892 mg of the chloroform extract with >95% purity, corresponding to a yield of 0.8%. UPLC-MS analysis of the product yielded an ion with mass and fragmentation pattern consistent with hominicin (Supplementary Fig. 4), and this product was further subjected to NMR and Marfey’s analysis (Supplementary Fig. 6 and 7) confirming the hominicin structure. A simpler extraction by n-butanol also provided a partially purified and enriched antimicrobial fraction from conditioned media (Supplementary Fig. 5). Therefore, the AH5011 extract obtained via n-butanol extraction was adopted for further experiments.
Activity of AH5011 hominicin against skin commensals and pathogens
Based on the inhibitory effect of the CM against a wide range of Gram-positive bacteria, as shown in Fig. 1, we next measured the minimum inhibitory concentration (MIC) of the purified daptide against a selection of skin commensal and pathogenic bacteria (Table 1) using a broth microdilution assay adapted from the Clinical Laboratory Standards Institute (CLSI) guidelines28. Hominicin demonstrated antimicrobial potency comparable to that of the clinically relevant antibiotic vancomycin across all tested strains (Table 1). Full MIC curves are included as supporting information (Supplementary Fig. 8).
The amino-modified termini are essential to hominicin’s activity
Given that the hom BGC represents a fully assembled gene cluster for a staphylococcal daptide, we next investigated the minimal genes required for antimicrobial activity. We generated plasmid constructs lacking homBDJM2 (i.e., predicted involvement in the Dmp biosynthesis and oxidoreduction of dehydroalanines), homD (i.e., predicted transamination in the Dmp biosynthesis), homM1 (i.e., predicted dimethylation of the N-terminus), lanMD (i.e., predicted dehydratase function), and homI, a gene of unknown function (Fig. 4a). Each construct was heterologously expressed into RN4220 and spot plated against AH4553. The expression of the full-length hom BGC confers antimicrobial activity, as evidenced by an inhibitory zone with a mean diameter of 17.3 ± 0.6 mm, compared to the empty vector (10.0 ± 0.2 mm), whereas the omission of homBDJM2, homD, homM1, and lanMD genes abolished antimicrobial activity (Fig. 4b). Interestingly, the omission of homI gene had no apparent impact on the antimicrobial activity, suggesting that HomI is not essential for hominicin biosynthesis.
a Gene organization of the minimal hom BGC (no flanking genes included), with the constructed gene omissions depicted. b Representative images of spot-on-lawn bioactivity of S. aureus RN4220 wild-type (wt) and expression mutants on a bacterial lawn of AH4553, with inhibition zone diameters indicated by black dashed lines. Red dashed line indicates the limit of detection. Scale bar = 10 mm. Data represents the mean of n = 5 biological replicates and error bars show standard deviation. c S. hominis AH4553 was cultured in 40% v/v CM from engineered S. aureus strains for 8 h. Data represents the mean of n = 3 biological replicates and error bars show standard deviation. Statistical significance in comparison to the S. aureus strain expressing empty vector was determined by ordinary one-way ANOVA followed by Dunnett’s multiple comparison test. Exact p values from left to right are as follows: b < 0.0001, <0.0001, 0.9591, 0.9990, >0.9999, 0.3197, c 0.1457, 0.9983, 0.1486, 0.1599, <0.0001, <0.0001. Source data are provided as a Source Data file.
We next tested whether the activity, or lack thereof, could be detected in the CM of each mutant. The CM from strains expressing the full-length hom gene cluster and HomABCDJM1M2PLanMD (i.e., omission of homI) were growth suppressive; however, CM from strains lacking homBDJM2, homD, homM1, and lanMD were not inhibitory (Fig. 4c). To corroborate the activity results, we performed mass spectrometric analysis of each CM to confirm the presence or absence of hominicin. The omission of homI resulted in a 21-residue product with an m/z value for the [M + 2H]2+ ion of 1020.1095 [∆ = 1 ppm] (Supplementary Fig. 9), corresponding to hominicin. However, the calculated daptide mass was not detected in the CM from mutants lacking homBDJM2; instead, multiple peptide species of similar mass to hominicin were detected with the most abundant ion displayed an [M + 2H]2+ of 1026.5679 (Supplementary Fig. 10a). The MS-MS spectra displayed a fragmentation pattern that is distinct from hominicin (Supplementary Fig. 10b). This structure likely differs from hominicin in the lack of C-terminal modification and oxidoreduction of dehydroalanine residues, matching the predicted m/z of 1026.5659 [∆ = 2 ppm] (Supplementary Fig. 10c). No peptide product was detected in the expression strain lacking the lanMD gene, suggesting this biosynthetic step is needed for stability of the peptide (Supplementary Fig. 11a). For the homM1 omission, mass spectrometric analysis indicated the production of a peptide lacking the N-terminal methylations while retaining formation of d-Ala from l-Ser (Supplementary Fig. 11a, b). Surprisingly, the observed product also lacked the Dmp modification on the C-terminus. The biosynthetic rationale for this is unclear as the installation of the C-terminal modifications should precede removal of the leader peptide needed for N-terminal methylation14. Whether this observation is due to polar effects on transcription or a requirement for these N-methylations to form the Dmp motif remains to be elucidated. For the homD omission, mass spectrometric analysis indicated the formation of a hominicin analogue with the expected N-terminal methylations and d-Alas, but a C-terminal ketone instead of the fully modified Dmp moiety (Supplementary Fig. 11c). This indicates that the N-terminal methylations can be installed even without formation of the Dmp motif.
AH5011 hominicin is a membrane-disruptive, pore-forming antimicrobial
The structural characteristics of daptides include a hydrophobic core and positively charged amino termini14. These features are found in many membrane-active, pore-forming antimicrobial peptides29. Whether daptides also form membrane pores has not been demonstrated although previous work observed that daptides from M. paraoxydans exhibit hemolytic activity14. We first asked whether hominicin could permeabilize the cytoplasmic membranes of S. aureus by measuring the uptake of the impermeant SYTOX Green fluorescent probe into bacterial cells. Upon cell entry, the fluorescence intensity of SYTOX Green is enhanced after binding to nucleic acids30. S. aureus cells treated with 2x or 4x conditioned media from AH5011, prepared from n-butanol extraction, which contained approximately 8.2 µg and 16.3 µg of hominicin, respectively, were significantly permeabilized compared to ∆p1∆p2-treated cells (Fig. 5a). We next examined the effects of hominicin on the membrane potential by measuring the release of potentiometric, cationic 3,3’-dipropylthiadicarbocyanine iodide [DiSC3(5)] dye from bacterial cells. DiSC3(5) accumulates on polarized membranes and translocates into lipid bilayers, resulting in fluorescence self-quenching31. S. aureus cells treated with AH5011 extract were depolarized in a dose-dependent manner, as indicated by the increased fluorescence intensity in the culture medium due to the release of DiSC3(5) (Fig. 5b).
a AH6350 cells were stained with SYTOX Green and treated with conditioned media prepared by n-butanol extraction (AH5011 or ∆p1∆p2), 35% vol/vol ethanol, or left untreated. The concentrations of hominicin in the AH5011 extract were 4.1 µg (1x), 8.2 µg (2x), and 16.3 (4x) µg. Fluorescence intensity was measured 3 h post-treatment. Data represents the mean of n = 3 biological replicates, and error bars show standard deviation. Statistical significance was determined using two-tailed unpaired t-test. Exact p values from left to right are as follows: a 0.0163, 0.0155, 0.5177. b AH6350 cells were stained with DiSC3(5) and treated with either AH5011 or ∆p1∆p2 extract, 1% Triton X-100, or left untreated. Fluorescence intensity was measured every 5 min. Data represents the mean of n = 3 biological replicates, and error bars show standard deviation. Statistical significance was compared to the untreated control group at 65 min using one-way ANOVA analysis followed by Dunnett’s multiple comparison test. Exact p values from left to right are as follows: b 0.9729, 0.8197, 0.0014, <0.0001. c Current/voltage (I/V) relationship of hominicin, unmodified core peptide, alpha-toxin and vehicle under symmetrical conditions of 1 M KCl and 10 mM HEPES, pH 7.2. Data represents the mean of n = 5 independent membrane recordings, and error bars show the standard error of the mean. d, e Representative current traces showing single channel openings and closings recorded at +100 mV (d) and –100 mV (e) for hominicin, unmodified core peptide, and vehicle. Source data are provided as a Source Data file.
To test whether hominicin exhibits ion channel activity, we reconstituted the pure peptide in artificial lipid bilayers bathed in a solution of 1 M KCl and 10 mM HEPES and measured ion flux across the membrane using voltage-clamp electrophysiology (Supplementary Fig. 12a). The current/voltage relationship of hominicin displayed an ohmic relationship with a calculated channel conductance of 692.4 ± 70.4 pS while the synthetic unmodified core peptide and buffer alone exhibited no channel activity (Fig. 5c). As a control, we added alpha-toxin32, a known pore-forming protein produced by S. aureus, and observed a channel conductance of 489.9 ± 82.8 pS. At both positive and negative holding potentials, hominicin exhibited ion-conducting activity, as evidenced by observable openings and closings of single channels (Fig. 5d, e). While hominicin exhibited antimicrobial activity, its unmodified form was inactive against S. aureus (Supplementary Fig. 12b). This suggests that the post-translational modifications of hominicin are crucial for its membrane insertion and channel activity. The unmodified peptide did not alter the current trace from baseline, whereas hominicin caused stepwise increases in the current trace at +50 mV and decreases at −50 mV and −100 mV holding potentials (Supplementary Fig. 12c–e). The changes in current traces are indicative of ion flux across the bilayer through the opening of membrane pores.
To test whether hominicin disrupts host membranes, we investigated the hemolytic activity induced by AH5011 using whole human blood. Alpha-toxin-producing S. aureus strains AH1263 and AH6350 were included, and AH1263 ∆agr mutant served as a non-hemolytic control33. AH1263 and AH6350 exhibited significant hemolysis when compared to uninfected (PBS) control while AH5011 did not induced hemolysis (Supplementary Fig. 13a). We did not observe differences in hemolysis of the AH5011 extract when compared to treatment with the ∆p1∆p2 extract (Supplementary Fig. 13b). In confirmation, we did not observe zones of hemolysis for AH5011, ∆p1∆p2 mutant, and AH1263 ∆agr mutant when plated on sheep’s blood agar (Supplementary Fig. 13c). We next measured cytotoxicity in human N/TERT keratinocytes using lactate dehydrogenase (LDH) release assays. AH1263 and AH6350 induced significant cytotoxicity on host cells whereas cytotoxicity caused by AH5011 and ∆p1∆p2 mutant was undetectable compared to untreated cells (Supplementary Fig. 13d). Altogether, these data indicate the mode of action of AH5011 hominicin involves impairing the membrane integrity by depolarizing the bacterial transmembrane potential through the formation of pores.
The genetic diversity and conservation of daptide BGC in Staphylococcus
To assess the diversity and conservation of the hom BGC in Staphylococcus, we performed a gene cluster comparison using the CAGECAT pipeline34, which enables detailed homology search, filtering, gene neighborhood estimation, and visualization of resulting variant BGCs. Fourteen assemblies were identified, carrying homologs of the precursor peptide along with at least one or two associated biosynthesis genes (Supplementary Table 3). These assemblies were sourced from shotgun sequencing data of various staphylococci, including S. aureus, S. pseudintermedius, S. felis, S. epidermidis, S. warneri, S. agnetis, and S. cohnii. Some of the BGCs exhibited variations in the composition of tailoring enzymes and the number of precursor peptides (Supplementary Fig. 14c). Seven assemblies lack the genes encoding for the dehydratase LanMD and oxidoreductase HomJ, suggesting that the installation of dehydroamino acids and the conversion of serines to d-alanines may have evolved independently from the Dmp biosynthesis. Additionally, we identified accessory genes likely involved in bacteriocin export and immunity. These were predicted to encode ATP-binding cassette (ABC) transporters, previously reported in the export of bacteriocins across the cytoplasmic membrane as a resistance mechanism35. Intriguingly, these were not found in S. hominis AH5011; however, the gene of unknown function, homI, was present in seven BGCs, including AH5011 and seems to correlate with hominicin-like molecules. We also observed that these daptide BGCs are located on or near mobile genetic elements, such as plasmids, recombinases, and transposases. This suggests the acquisition of daptides via horizontal gene transfer. Alignment of the precursor peptides revealed diversity in sequence and length while conservation of the C-terminal threonine was observed (Supplementary Fig. 14d).
HomI is the immunity protein that provides resistance to AH5011 hominicin
Producer strains are often protected from the activity of their bacteriocin by self-immunity systems36,37,38. The mechanisms of producer immunity to daptides have never been investigated. We ascertained the role of homI given that 6 out of the 14 daptide BGCs encode this gene (Supplementary Fig. 14c) and previous work observed the co-occurrence of HomI in staphylococcal daptide BGCs14. We hypothesized that HomI is the immunity protein that provides cognate resistance to hominicin. Using a genetically tractable and hominicin-susceptible S. aureus strain Newman, we cloned homI on a multicopy plasmid and observed no growth defects between the wild-type and homI-expressing mutant in TSB (Fig. 6a). Under growth conditions with AH5011 CM, the mutant exhibited significant resistance while the growth of wild-type and empty vector were attenuated (Fig. 6b). HomI-mediated resistance was confirmed by the lack of inhibitory zones in the presence of hominicin-producing AH5011 and S. aureus strain RN4220 expressing hom BGC (Fig. 6c).
a, b Bacterial growth curves of S. aureus strains Newman, homI expression mutant, and empty vector grown in TSB (a) or 40% v/v CM from AH5011 (b). Data represents the mean of n = 3 biological replicates and error bars show standard deviation. Statistical significance was compared to the growth of the wild-type strain Newman at 8 h using one-way ANOVA followed by Dunnett’s multiple comparison test. Exact p values from left to right are as follows: a 0.2729, 0.9983, b < 0.0001, 0.1459. c Representative images of spot-on-lawn bioactivity of AH5011 and RN4220 expressing hom BGC against lawns of S. aureus, with inhibition zone diameters represented by black dashed lines. Red dashed line indicates the limit of detection. Scale bar = 10 mm. Data represents the mean of n = 5 biological replicates, and error bars show standard deviation. Statistical significance was determined using one-way ANOVA followed by Dunnett’s multiple comparison test. Exact p values from left to right: c (diameter of AH5011 inhibition) <0.0001, 0.9743, c (diameter of AH6425 inhibition) <0.0001, 0.9921. d The HomI-hominicin complex model superimposed onto the AlphaFold model of HomI. The models are presented in a side-profile view (top) and a top-down view (bottom). Hominicin is represented in purple as sticks. HomI modeled without hominicin is represented in yellow, and HomI modeled in complex with hominicin is represented in teal. e–g The release of DiSC3(5) from Newman wild-type (e), homI-expressing mutant (f), and empty vector (g) in the culture medium. The 8x AH5011 extract contains 32.7 µg of hominicin. Data represents the mean of n = 3 biological replicates and error bars show standard deviation. Statistical significance was compared to the untreated control group at 65 min using one-way ANOVA analysis followed by Dunnett’s multiple comparison test. Exact p values from left to right are as follows: e 0.7120, 0.0159, f (8x AH5011) 0.7705, f (8x ∆p1∆p2) 0.8201, g 0.6444, 0.0269. Source data are provided as a Source Data file.
HomI is predicted to be a 32.8 kDa membrane-associated protein with five transmembrane helices39 (Supplementary Fig. 15a). To identify structural homologs, we predicted the structure of S. hominis HomI using AlphaFold40 and submitted the structure as a query in Foldseek41. This analysis revealed that HomI shares structural homology with the YidC family of membrane protein insertases (Supplementary Fig. 15b, c), suggesting that HomI may function at the membrane interface. Alignments of the HomI model with an AlphaFold-predicted structure of S. aureus YidC and a crystal structure of Bacillus halodurans YidC (PDB ID 3WO7_A) yielded RMSD values of 1.41 Å and 1.76 Å, respectively, indicating high structural similarity. To investigate the potential mechanism behind HomI-mediated resistance against hominicin, we used Boltz-1x to predict a HomI-hominicin complex structure40,42. For comparison, the AlphaFold-predicted HomI structure was superimposed onto the HomI-hominicin complex. Hominicin docked within the elongated positively charged pocket in the HomI core, with minimal structural changes between HomI modeled with and without hominicin. Intriguingly, the N-terminal α1 helix exhibited a conformational shift, suggesting a potential binding interaction between HomI and hominicin (Fig. 6d, Supplementary Fig. 15d, e).
We next examined the potential function of HomI in protecting against daptide-induced membrane damage by measuring the release of DiSC3(5) in the culture medium. Under ∆p1∆p2-treated and untreated conditions, the wild-type strain Newman, homI-expressing mutant, and empty vector strains exhibited fluorescence self-quenching, which reflects the uptake of DiSC3(5) and the energized and/or polarized state of the cells. When exposed to AH5011 extract, the wild-type (Fig. 6e) and empty vector (Fig. 6f) strains displayed membrane depolarization. However, the homI-expressing mutant retained its membrane potential, comparable to the untreated control (Fig. 6g). Altogether, we identified homI as the genetic determinant that confers resistance against membrane damage caused by hominicin.
AH5011 hominicin restricts S. aureus skin infection in vivo
The translational potential of daptides to protect against invading pathogens has never been tested. Therefore, we investigated hominicin-mediated skin protection from S. aureus infection. S. aureus is the most common cause of skin and soft tissue infections in the United States and frequently associated with high morbidity, morality, and healthcare costs43,44. As an opportunistic pathogen, S. aureus can exacerbate diseases, such as atopic dermatitis, an inflammatory skin disease with a complex pathogenesis marked by impaired skin barrier function and structure, immunological abnormalities, and dysbiosis45. Nasal carriage is a known risk factor for invasive S. aureus infections46. Therefore, we selected the strain AH6350, a high-toxin producing isolate from an acute lesion of pediatric AD, as a clinical organism to study in vivo (Fig. 7a, Supplementary Fig. 13). AH6350 is highly susceptible to hominicin, with a minimum inhibitory concentration of 1 µg/mL (Table 1). Genomic analysis revealed that it is closely related to strains from sequence type (ST) 15 lineage with an accessory gene regulator (agr) quorum-sensing system of type II (Fig. 7b). ST15 is commonly associated as nasal carriage lineage47 and often expresses surface proteins with skin-binding capacity48,49 and has the potential pathogenicity to cause skin-related diseases50,51.
a Representative image of hemolysis from S. aureus strain AH6350 on 5% sheep blood agar. Scale bar = 10 mm. b Multiple sequence alignment of AgrD, the peptide precursor of thiolactone auto-inducing peptide, from representative S. aureus (SAUR) strains. Colors indicate residue side-chain properties as standard for Clustal X alignments. c–f Five male and five female C57BL6/J mice were epicutaneously infected with S. aureus AH6350 or left uninfected (phosphate saline buffer) and given either hominicin, vehicle (methanol), or left untreated (n = 10 biological replicates per group). c Representative images of skin lesions for indicated groups. d Total bacterial burden in epicutaneous lesions at 72 h post-infection (hpi). Data represents the mean and standard deviation. ND indicates no detection. Statistical significance in comparison to the S. aureus-infected untreated group was determined using two-tailed Mann-Whitney U-test. Exact p values from top to bottom are as follows: d < 0.0001, <0.0001. e Skin barrier integrity was assessed by measuring transepidermal water loss (TEWL) prior to infection and at 72 hpi. Data represents the mean and standard deviation of n = 5 female C57BL6/J mice per group. f The severity of skin inflammation was quantified using a total disease score, which is the sum of the individual grades for erythema, edema, erosion and scaling. Data represent the mean and standard deviation of n = 10 biological replicates per group. Statistical significance in comparison to the uninfected group was determined using ordinary one-way ANOVA followed by Dunnett’s multiple comparison test. Exact p values from top to bottom are as follows: e (0 hpi) 0.9897, 0.5398, 0.9744, e (72 hpi) 0.9989, 0.0025, 0.0069, f 0.2013, <0.0001, <0.001. Source data are provided as a Source Data file.
Epicutaneous infection of female and male C57BL6/J mice with AH6350 alone resulted in severe skin scaling, erosion, and erythema (Fig. 7c). Infection with S. aureus was attenuated by the topical addition of hominicin to the skin surface, as evidenced by a reduced bacterial burden compared to the untreated group (Fig. 7d). Additionally, the infected mice treated with hominicin retained skin barrier function and integrity, as measured by transepithelial water loss (Fig. 7e). The severity of local inflammation on mouse skin at 72 h post-infection was assessed using a skin disease score, which represents the sum of individual grades for erythema, edema, erosion and scaling52. We observed minimal skin damage with hominicin alone in the absence of infection (Fig. 7f, Supplementary Fig. 16). Topical treatment with hominicin on infected skin resulted in significantly less skin inflammation and cutaneous injury, which correlates to the reduction of S. aureus on the skin.
Discussion
In a polymicrobial setting, skin microbes must contend with neighboring competitors and have evolved mechanisms for interbacterial competition, including the production of antimicrobial natural products53. Daptides are an understudied class of RiPPs. Here, we describe a member of this class produced by S. hominis and explore its biosynthesis and mechanism of action using bioinformatic, microbiological, biochemical, and biophysical techniques and propose a comprehensive model for hominicin (Supplementary Fig. 17). Bacteriocins inhibit similar or closely related bacteria that the producer strain encounters in its niche54. This study investigates the breadth of hominicin’s activity against members of the skin community. While we observed varying levels of growth inhibition against related skin staphylococci, the lack of inhibition against S. agalactiae and E. faecalis aligns with this principle, as these species are not commonly found in the human skin microbiome55. Moreover, inhibition against M. luteus and S. pyogenes–the former is a commensal species56 and the latter a known etiological agent of skin and soft infections57–points to the microbiome-shaping role of daptides. The disparate sensitivities between strains could be attributed to the presence of HomI homologs, canonical transporter systems for bacteriocin (e.g., ATP-binding cassette transporters), or the insufficient amount of hominicin in the CM to elicit observable sensitivity. While this study examined inhibition against a subset of commensal and pathogenic species, a combinatorial approach that leverages in vitro activity and metagenomic analysis could provide insights into how daptides coalesce in a polymicrobial setting and affect the ecological structure and function of the community.
Structural characterization of AH5011 hominicin revealed several post-translational modifications, which include dehydrobutyrines, d-alanines, a dimethylated N-terminus, and a Dmp-modified C-terminus (Fig. 3). Ren et al. identified the biosynthetic machinery that coordinates the conversion of a C-terminal threonine to (S)-N2-N2-dimethyl-1,2-propanediamine14. By exploiting the conservation of the DapBCDM enzymes, co-occurrence analysis of the neighboring genes led to the discovery of several putative subgroups of daptide BGCs that encode an array of additional biosynthetic enzymes such as lanthipeptide dehydratases, glycosyltransferases, and thiopeptide pyridine synthases14. These ancillary enzymes highlight the potential structural diversity of daptides that remains to be characterised. Indeed, the hom BGC encodes the dehydratase LanMD that modifies Thr2, Thr5, Thr8, Ser10, Thr15, and Ser17 into dehydroamino acids. We observed that Thr12 remains unmodified, which could be explained by structural constraints, enzyme specificity, or interference by other modifications that may influence its accessibility. Furthermore, we identified a tertiary modification mediated by the HomJ oxidoreductase that converts dehydroalanines, derived from Ser10 and Ser17, into d-alanines. Incorporation of d-amino acids into RiPPs imparts several advantageous properties, such as stabilization of peptide structures, reduced susceptibility to proteolysis, and enhancement of bioactivity25,58. Lastly, amino-modified termini are common features present in many defense peptides. Notably, the non-ribosomal aquimarins from a marine bacterial genus Aquimarina feature a C-terminal amine, which was demonstrated to be indispensable for antibacterial activity59. N-terminal dimethylation has been reported in other RiPPs, such as linaridins (e.g., cypemycin60) and thiazole/oxazole-modified microcins (e.g., plantazolicin61,62), and has been shown to be essential for the peptides’ activity. Other bodies of work corroborate the importance of modified C-termini for peptide-membrane interaction and structural stabilization63,64,65. We observed that omission of the genes involved in the N-terminal dimethylation and C-terminal Dmp biosynthesis resulted in loss of antimicrobial activity, implying that amino-modified termini are essential to hominicin’s mode of action (Fig. 4). Future studies will be needed to define the efficacy of interaction, specificity, and stability of these moieties for membranes.
Collectively, the MS, genetic, and NMR data reveal that AH5011 daptide is structurally identical to hominicin. Hominicin was originally classified as a lanthipeptide lacking the lanthionine thioether bridge17, but it is more correctly assigned as a daptide14. Based on gene sequence data and expression studies, our work provides insight into the biosynthesis and antimicrobial function of hominicin. Here, we have chosen to adopt the hominicin name for the peptide to be consistent with the daptide nomenclature. Furthermore, our report proposes that the core peptide contains serines at the 10th and 17th residues, which are post-translationally converted into d-Ala to form the hominicin peptide. This finding is significant, as it resolves the biosynthetic gap between the primary sequence of the precursor peptide and the final structure of hominicin, a connection not identified in the original report.
Unlike hominicin, Mpa daptides do not exhibit antibacterial activity14. Intriguingly, Mpa daptides are hemolytic against bovine erythrocytes, suggesting their capability to interact with cell membranes. Worth mentioning is that the Mpa daptides do not harbor dehydroamino acids and an N-terminally dimethylated residue. Using in vivo membrane potential measurements coupled with electrophysiology, we provide evidence that hominicin permeabilizes the cytoplasmic membrane of S. aureus, likely driven by its capacity to penetrate and self-organize into ion-conducting channels in the membrane (Fig. 5). The interaction at the membrane interface is likely facilitated by the charged amino termini via electrostatic interactions. With its hydrophobic moieties, hominicin efficiently partitions into the hydrophobic core of the membrane and spontaneously self-assembles individual monomers to form a transmembrane pore. The lipid bilayer experiment supports this working model given that hominicin associates with the phospholipid membrane without the mediation of other surface receptors or proteins. Our results provide a molecular picture of hominicin as a membrane-depolarizing, pore-forming agent that consequently leads to the loss of membrane integrity and the collapse of transmembrane potential in susceptible bacteria. However, the observed channel activity is not sufficient to fully resolve hominicin’s structure within the membrane. An in-depth knowledge of the three-dimensional structure of the hominicin monomer would be valuable in the study of its peptidic channel.
Additionally, we identified a daptide resistance protein, HomI. To our knowledge, there has been no previous report of an immunity mechanism for daptides. We observed that HomI sufficiently protects against hominicin-induced membrane depolarization (Fig. 6). HomI shares structural homology with the YidC family of membrane protein insertases (Supplementary Fig. 15). YidC is involved in facilitating membrane insertion of small proteins or assisting the protein-conducting channel SecYEG, which mediates the translocation of secretory proteins across the membrane. As such, the Enterocin P bacteriocin from Enterococcus faecium has been shown to require the sec-dependent pathway for its secretion66, and the sec pathway can be alternatively used to secrete dehydrated lanthipeptides in the absence of their lantibiotic transporter, NisT67. Our structural modeling shows hominicin docks within HomI’s charged core with high predictability (Fig. 6d). Glu13 of hominicin is predicted to form a salt bridge with Arg66 residue of HomI (Supplementary Fig. 15c). It has been demonstrated that this Arg residue, conserved within YidC proteins, is essential for its function68. We hypothesize that HomI operates in a similar manner to mediate resistance against hominicin. While YidC facilitates protein insertion into the membrane, HomI may sequester hominicin within its core or participate in the transport of hominicin. Other studies suggest that YidC may interact with immunity protein substrates involved in bacteriocin transport and resistance69,70. Interestingly, we detected homologs of ABC transporters in a number of daptide BGCs (Supplementary Fig. 14), which may represent alternative resistance mechanisms. This warrants future investigations to define the role of these putative transporters, as well as further study into the molecular mechanism of HomI-mediated daptide resistance.
In this study, we evaluated the effectiveness of treating S. aureus-infected skin with hominicin in vivo (Fig. 7). Our discovery of a commensal bacterium producing a potent daptide is a strain-specific feature that is often overlooked in large genomic datasets. Nevertheless, the presence of small-molecule biosynthetic gene clusters is prevalent in human-associated metagenomes71, highlighting the commensal bacteria as an untapped reservoir of bioactive molecules. Our results add to many examples of S. hominis as a protective skin commensal, showcasing its beneficial role in the production of bacteriocins. The translatability of characterizing these natural products and evaluating their utility as antimicrobial therapies presents an avenue for therapeutic development. Utilizing hominicin or daptide-producing staphylococci could be an effective strategy for preventing S. aureus colonization and infection, as has been previously done with a lantibiotic-producing strain of S. hominis12. The introduction of hom genes into an exclusively commensal species could pave the way towards a precision-based bacteriotherapy to target specific colonizing pathogens or provide colonization resistance against future infections. Collectively, our discoveries provide unprecedented insights into the molecular mechanism of hominicin activity and highlight its antibacterial property as an anti-staphylococcal agent.
Methods
Collection of bacteria from human subjects
Bacterial isolates from human skin swabs were used from a previously published collection5,72. Swabs from a 5-cm2 area of the antecubital fossa skin were collected from both the left and right arms from healthy subjects and patients with AD. Skin swab samples were suspended in 1 mL TSB containing 15% v/v glycerol and plated on mannitol salt agar with egg yolk for the selective growth of Staphylococcus species. Participants included individuals of both sexes who had not used an antibiotic in the preceding month. Age and sex were self-reported. Skin swab collection was done in accordance with protocols approved by the University of California, San Diego (UCSD) Institutional Review Board under project number 140144. Bacterial isolates from skin swabs of healthy subjects were collected in accordance with protocols approved by COMIRB under protocol number 19-2218 and UCSD Institutional Review Board under project number 071032. Informed consent was obtained from all subjects. Isolates used in this study are listed in Supplementary Data 3.
Growth conditions and reagents
The bacterial strains and plasmids used in this study are listed in Supplementary Table 1. S. hominis isolates were confirmed to be S. hominis by matrix-assisted laser desorption/ionization (MALDI)-time of flight (TOF) mass spectrometry prior to experimentation. All staphylococcal strains and Micrococcus luteus were grown in TSB growth medium at 37 °C with shaking at 250 rpm. Escherichia coli was grown in Luria broth (LB) at 37 °C with shaking. Enterococcus faecalis was grown in Brain Heart Infusion at 37 °C with shaking. Streptococcus agalactiae was grown statically in Todd Hewitt broth at 37 °C. Streptococcus pyogenes was grown statically in Todd Hewitt broth supplemented with yeast extract at 37 °C + 5% CO2. For strains with pCM28, chloramphenicol was added to a final concentration of 10 µg/mL. The unmodified core peptide was custom synthesized by AnaSpec (Fremont, CA).
Bacteriocin antimicrobial activity assays
For conditioned medium (CM) assays, overnight bacterial cultures of S. hominis or S. aureus strains were pelleted, and the spent media was filtered through a 0.22-µm cellulose acetate Spin-X filter (Costar). 100 µL of 80% (vol/vol) CM was added to a 96-well culture plate unless otherwise indicated. The indicator strains were prepared by diluting overnight cultures 1:50 in fresh TSB. 100 µL of the indicator strain was added to the culture plate to achieve a final volume of 200 µL per well. Cultures were grown in a Stuart humidified incubator at 37 °C with shaking at 800 rpm. Cell density (OD600) was measured on a Tecan Group Ltd. Infinite Pro plate reader. At specified time-points, 20 µL of culture was removed, serially diluted, and plated on TSA for colony counting.
For the spot-on-lawn assays, S. hominis AH4553 was used as the sensitive strain for all experiments, unless otherwise stated. 100 µL of the 1:10 diluted suspension of AH4553 was spread on TSA using sterile glass beads and air dried. 10 µL of the daptide-producing or non-producing strains were inoculated onto the bacterial lawn. Plates were incubated at 37 °C for 16–20 h, and images of the zones of inhibition were collected and analyzed using ImageJ software (version 1.53).
Isolation of plasmid-cured mutants
S. hominis AH5011 was subcultured 1:100 in fresh TSB and allowed to reach an OD600 of 0.50. Acriflavine was added to the bacterial culture at a final concentration of 25 µg/mL and incubated for 1 h. The culture was diluted and plated on TSA to obtain single colonies, which were then picked and patched on fresh TSA. Bacterial strains cured of p1 failed to produce inhibition zones when spotted on a lawn of AH4553. Strains cured of p2 failed to grow on agar containing 1 µg/mL mupirocin. To confirm the loss of plasmids, polymerase chain reaction (PCR) was performed using primers for the p1- and p2- specific genes, homA and mupA, respectively.
Cloning of the hom biosynthetic gene cluster
All primers used in this study are listed in Supplementary Table 2 and purchased from Integrated DNA Technologies (Coralville, Iowa). The hom gene cluster was cloned into pCM28 by a Gibson assembly strategy. The construction of the S. aureus shuttle vector pCM28 has been previously described73. The pCM28 vector was linearized by restriction digestion using BamHI-HF and EcoRI-HF (New England Biolabs). The hom gene cluster was amplified in four parts from the genomic DNA of S. hominis AH5011 using primers homBGC_p1_Frw/_Rev, homBGC_p2_Frw/_Rev, homBGC_p3_Frw/_Rev, and homBGC_p4_Frw/_Rev. The resulting 15-kb construct, pAN1, was introduced into chemically competent E. coli DH5-alpha cells and selected on LB agar with 100 µg/mL ampicillin. The plasmid was verified by DNA sequencing. The pAN1 and empty vector pCM28 were electroporated into competent S. aureus RN4220, as previously described74. Transformants were selected on TSA with 10 µg/mL chloramphenicol.
For gene omission studies, expression constructs were generated as mentioned above. To construct pCM28-homABCDJM1M2PlanMD (i.e., omission of homI), the gene cluster was amplified in three parts using primers homBGC_p3_Fwd/_Rev, homBGC_p4_Fwd/_Rev, and homBGC_p7_Fwd/_Rev. The resulting 14-kb construct was designated as pAN2. To construct pCM28-homACIM1PlanMD (i.e., omission of BDJM2), the gene cluster was amplified in three parts using primers homBGC_p4_Fwd/_Rev, homBGC_p5_Fwd/_Rev, and homBGC_p6_Fwd/_Rev. The resulting 11-kb construct was designated as pAN3. To construct pCM28-homABCDIJM2PlanMD (i.e., omission of homM1), the gene cluster was amplified in four parts using primers homBGC_p1_Fwd/_Rev, homBGC_p2_Fwd/_Rev, homBGC_p14_Fwd/_Rev, and homBGC_p15_Fwd/_Rev. The resulting 14-kb construct was designated as pAN4. To construct pCM28-homABCIJM1M2PlanMD (i.e., omission of homD), the gene cluster was amplified in four parts using primers homBGC_p3_Fwd/_Rev, homBGC_p4_Fwd/_Rev, homBGC_p8_Fwd/_Rev, and homBGC_p9_Fwd/_Rev. The resulting 14-kb construct was designated as pAN6. To construct pCM28-homABCDIJM1M2P (i.e., omission of lanMD), the gene cluster was amplified in four parts using primers homBGC_p1_Fwd/_Rev, homBGC_p2_Fwd/_Rev, homBGC_p10_Fwd/_Rev, and homBGC_p11_Fwd/_Rev. The resulting 13-kb construct was designated as pAN7.
Cloning of the immunity gene, homI, was performed as follows: The pCM28 vector was linearized by restriction digestion using BamHI-HF and PstI-HF. The insert was PCR-amplified using primers homI_Frw/_Rev. Purified vector and insert were then ligated using T4 ligase. The resulting 6.6-kb construct, designated as pAN5, was introduced into DH5-alpha cells and verified by DNA sequencing. pAN4 was passaged through RN4220 and subsequently introduced into S. aureus strain Newman by electroporation.
Mass spectrometric identification of the purified daptide and in conditioned media
Samples were analyzed on a Q Exactive Plus mass spectrometer (Thermo Fisher Scientific, Waltham, MA) with a heated electrospray ionization source coupled to an Acquity ultrahigh-performance liquid chromatography (UPLC) system (Waters Corp., Milford, MA). Conditioned media samples, obtained from three independent biological replicates, were centrifuged to pellet bacterial cells and the resulting supernatants were passed through 0.22 µm PVDF syringe filters prior to analysis. A culture medium blank processed under identical conditions was also included. Purified daptide samples were dissolved in LC-MS grade methanol and injected in triplicate as technical replicates. All samples were injected with a 3-7 µL volume and eluted from an Acquity UPLC BEH C18 1.7 μm 2.1 × 50 mm column (Waters Corporation) at a 0.3 mL/min flow rate using a binary solvent system consisting of 0.1% formic acid (A) and acetonitrile (CH3CN) with 0.1% formic acid (B). Mass spectra were collected using positive ion mode electrospray ionization with two scan events, a full-scan event over a mass range of 300 to 2000 at a resolving power of 35,000, and a data-dependent tandem mass spectrometry (MS/MS) scan event selecting the calculated m/z or a data-independent scan event using all ion fragmentation. The solvent gradient began with a 1.5 min isocratic hold at 20% B which was diverted to waste. Subsequently, the flow was diverted to the mass spectrometer and a linear increase to 60% B was performed over 5 min. The gradient was held isocratic from 6.5 min to 7.0 min and then increased to 100% B at 8.0 min. The column was washed at 100% B for 1 min and then returned to the starting conditions to allow equilibration for 1.0 min prior to the next injection.
Marfey’s analysis
Marfey’s analysis was conducted to determine the stereochemistry of amino acids in the sample, employing a modified version of the procedure27. Each amino acid standard was prepared in water at approximately 4 mg/mL. Each prepared standard (50 µL) was then combined with 1 M NaHCO₃ (20 µL), and 1% Marfey’s reagent (Nα-(2,4-dinitro-5-fluorophenyl)-l-alaninamide) in acetone (100 µL), and the mixture was incubated at 40 °C for 1 h. The reaction was quenched by adding 10 µL of 2 N HCl, and the resulting mixture was dried under a stream of nitrogen. The residue was reconstituted in approximately 200 µL of methanol. Additionally, approximately 0.1 mg of purified hominicin was hydrolyzed with 0.5 mL of 6 N HCl at 90 °C for 24 h. After hydrolysis, the sample was evaporated under a stream of nitrogen. The residue was reconstituted in 25 µL of water, followed by the addition of 10 µL of 1 M NaHCO₃ and 50 µL of 1% Marfey’s reagent in acetone. The mixture was agitated at 40 °C for 1 h, then quenched with 5 µL of 2 N HCl. The derivatized sample was dried under a stream of nitrogen and reconstituted in approximately 125 µL of methanol.
Each derivatized standard was injected individually (1 µL) onto the UPLC-MS, and a mixed standard containing aliquots of all derivatized standards was also prepared and injected. Samples were analyzed on a Q Exactive Plus mass spectrometer (Thermo Fisher Scientific, Waltham, MA) with a heated electrospray ionization source coupled to an Acquity ultrahigh-performance liquid chromatography (UPLC) system (Waters Corp., Milford, MA). Chromatographic separation was performed using a BEH C18 column (150 mm × 2.1 mm, 1.7 µm particle size) at a flow rate of 0.3 mL/min. The mobile phase consisted of 0.1% formic acid in water (solvent A) and methanol (solvent B). A gradient of 10–70% solvent B over 10 min was employed, followed by re-equilibration to the initial conditions (90% A, 10% B) over 1.5 min with a total runtime of 11.5 min.
Mass spectrometric analysis was performed in positive with spray voltage 3000 V, capillary temperature 256 °C, probe heater temperature 350 °C, sheath gas flow 48 (arbitrary units), auxiliary gas flow 11 (arbitrary units), and S-Lens RF level 50. Data acquisition was performed in full-scan MS mode over a mass range of m/z 150–1000. Stereochemical assignments were made by comparing retention times and m/z values for the amino acids detected from hominicin to those of d- and l-amino acid standards (Supplementary Fig. 7).
Butanol extraction of AH5011 hominicin
n-Butanol (100 mL) was added to 300 mL of cell-free, conditioned media and thoroughly mixed before setting aside for several min until the phases settled. After centrifugation at 2000 x g for 5 min, the upper butanol phase was collected and evaporated in a rotatory vacuum evaporator (Buchi Rotavapor R-3000) at 45 °C. The dried extract was resuspended and concentrated in nuclease-free water or PBS, which was stored at −80 °C until use.
Isolation of the AH5011 hominicin
The bacteriocin was concentrated by liquid-liquid partitioning of culture media with chloroform (1:1, v/v). The chloroform was removed by evaporation under nitrogen gas, and the material was subjected to reversed-phase flash chromatography using an automated CombiFlash RF system (Teledyne-Isco). A 34.7-minute method was implemented with a gradient of water (A) and methanol (B). The gradient was performed at a flow rate of 75 mL/min on a 130 g C18 RediSep column beginning at 50% B and linearly increasing to 100% B, followed by an isocratic hold at 100% B for approximately 8.5 min. Subsequently, preparative scale reversed phase high performance liquid chromatography (HPLC) was performed for further purification on a Varian HPLC system (Agilent, Santa Clara, CA) equipped with ProStar 210 pumps, a ProStar 710 fraction collector, a ProStar 335 photodiode array detector with and Galaxie Chromatography Workstation software (version 1.9.3.2). A 40-minute method was implemented with a gradient of water (A) and acetonitrile (B) on a Phenomenex Gemini – NX C18 column (5 μm; 250 × 21.20 mm). The gradient separation was performed at a flow rate of 21.2 mL/min and began at 30% B and linearly increased to 50% B.
NMR data collection
NMR experiments were performed on an Agilent 700 MHz NMR spectrometer equipped with a cryoprobe, operating at 700 MHz for 1H and 175 MHz for 13C. Spectra were collected using a water suppression enhanced through T1 effects (WET) pulse sequence. The sample was dissolved in methanol-d3, and the residual solvent signals (δH = 3.31 ppm and δC = 49.0 ppm) were used as reference peaks.
Determination of minimum inhibitory concentration (MIC)
The MICs of the purified hominicin were determined for a panel of skin microbes using a broth microdilution assay modified from the Clinical Laboratory Standards Institute (CLSI) guidelines. Serial dilutions of hominicin were prepared in 96-well microtiter plates containing bacterial cultures adjusted to 5 × 105 CFU/mL in either TSB media (staphylococci) or in cation-adjusted Mueller-Hinton Broth with laked horse blood (streptococci). As a control, the MICs of vancomycin were also obtained for the same panel of skin microbes. Plates were incubated overnight at 37 °C at 250 rpm, and bacterial growth was assessed by measuring optical density at 600 nm using a BioTek Synergy H1 microplate reader. The MIC was defined as the lowest concentration of hominicin that completely inhibited visible bacterial growth.
SYTOX Green influx to detect membrane permeabilization
Mid-log S. aureus cells were washed and then suspended in a final concentration of 2 µM SYTOX Green (Invitrogen) in nuclease-free water. 100 µL of SYTOX-bacterial suspension was dispensed into a 96-well black culture plate (Corning). Plates were incubated for 15 min in the dark. Then, 100 µL of extract from AH5011 or ∆p1∆p2, 70% v/v ethanol, or water were added to achieve a final volume of 200 µL per well. Fluorescence intensity was measured with a Tecan plate reader at an excitation/ emission wavelength of 480 nm/ 522 nm and normalized to “no bacteria” controls.
DiSC3(5) efflux to detect membrane depolarization
To measure DiSC3(5) efflux, mid-log S. aureus cells were washed and suspended in a final concentration of 4 µM DiSC3(5) (Invitrogen) in buffer (5 mM HEPES, 20 mM glucose, pH 7.2). The labeled bacteria were dispensed into a 96-well black culture plate and allowed to reach equilibrium. Next, 100 µL of extract from AH5011 or ∆p1∆p2, 1% Triton X-100, or HEPES/glucose buffer were added to achieve a final volume of 200 µL per well. Fluorescence measurements were taken every 5 min at an excitation/ emission wavelength of 620 nm/670 nm and normalized to “no bacteria” controls.
Voltage-clamp planar lipid bilayer
Electrophysiology experiments were performed using a 4-channel micro-electrode cavity array (100 µm MECA, Ionera Technologies) on an Orbit Mini (Nanion Technologies) system. Recordings were collected using the Elements Data Reader software, with parameters set to 200 pA gain and 1.25 kHz sampling rate and subsequently analyzed using the Elements Data Analyzer (v. 1.4.6). For preparation of the horizontal lipid bilayer system, 150 µL of the bathing solution (1 M KCl, 10 mM HEPES, pH 7.2) was added to the measurement chamber. 1,2-diphytanoyl-sn-glycerol-3-phosphocholine (DphpC) from Avanti Polar Lipids was dissolved in n-octane to a concentration of 10 mg/mL (Sigma, electronics grade). Membranes were painted over the microcavities using the air bubble technique75 until the capacitance reached 15–30 pF, as recommended by the manufacturer. A transmembrane voltage of +50 mV was applied, unless otherwise stated. Approximately 40 ng of hominicin, 40 ng of synthetic unmodified core peptide, 0.125% v/v methanol, or 8 ng of alpha-toxin was added per aperture, and each membrane was monitored for fusion spikes to indicate insertion of peptides into the lipid bilayer. Current traces were monitored until single-channel conductance events were observed at the indicated voltage. For current-voltage analysis, the current was recorded for 10 seconds at 20 mV increments with a return to 0 mV between each step, starting from −100 mV to +100 mV. Single-channel conductance was determined from the current divided by the transmembrane voltage.
Hemolysis assay
Whole human blood samples were collected separately from three healthy adult donors in sodium heparin blood collection tubes (BD Vacutainer). Overnight cultures of bacterial strains were pelleted, washed with PBS, and normalized to 5 × 107 CFU/mL. 200 µL of bacterial suspension, conditioned media extract, or water was incubated with 200 µL of whole blood at 37 °C for 4 h. Following incubation, samples were centrifuged at 5500 x g, and 100 µL of the supernatant was transferred to a 96-well plate. Erythrocyte lysis was determined by measuring the absorbance at 543 nm. Percent hemolysis was determined by normalizing absorbance values to those of water-treated samples (i.e., 100% total lysis).
Lactate dehydrogenase leakage assay with N/TERT-2G keratinocytes
Immortalized human N/TERT-2G keratinocytes were obtained from Dr. James Rheinwald laboratory and were grown in serum-free keratinocyte growth medium (Gibco) supplemented with 25 µg/mL bovine pituitary extract, 0.2 ng/mL epidermal growth factor, and 0.3 mM calcium chloride as previously described76. Low passage ( < 15), undifferentiated N/TERT-2G cells were seeded to 1 × 105 cells/mL into 24-well tissue culture plates (Corning) and cultured at 37 °C with 5% CO2 until reaching ~90% confluency. Cell monolayers were washed with PBS and media was replaced. Overnight cultures of bacteria were pelleted, washed, and then added to cell monolayers at 1 × 106 CFU and allowed to incubate for 4 h. LDH release was measured using the CyQUANT LDH Cytotoxicity Kit (Invitrogen), following the manufacturer’s instructions.
Murine epicutaneous infection with S. aureus
Age- and gender-matched five male and five female 8-week-old C57BL6/J mice were used for epicutaneous exposure to S. aureus5,52,77. 24 h prior to bacterial inoculation, the dorsal skin of anesthetized C57BL6/J mice (2% isoflurane) was shaved and depilated with Nair (Church & Dwight Co., Inc.). S. aureus strain AH6350 was subcultured 1:50 in TSB and allowed to reach an OD600 of 1. Cells were pelleted, washed, and then resuspended in PBS to achieve an inoculum of 1 × 108 CFU in 100 µL volume. S. aureus was inoculated on a sterile 2-cm2 gauze pad and affixed to the back skin with a Tegaderm dressing and secured with adhesive bandages (BAND-AID, Johnson and Johnson) for 72 h. For mice receiving treatment, hominicin (100 µg suspended in methanol) or vehicle (2% v/v methanol) were combined with S. aureus immediately prior to application on the gauze pad. Inoculum concentration was verified by serial dilution, plating, and colony counting. Transepithelial water loss was measured using a Tewameter TM300 device (Courage & Khazaka Electronic GmbH) before infection and at 72 h post-infection. Two sites per lesion were analyzed to minimize error in measurements. The disease score as a measure of skin inflammation severity was evaluated by a blinded observer from digital photographs and quantified as a sum of three individual grades for erythema, edema (each graded: 0, 1, 2, 3) and scaling/erosion (scaling graded: 0, 1, 2, 3 while erosion graded as 4, 5, 6, 7)52. To enumerate bacterial burden, the full-thickness 2-cm2 atopic lesions were excised, suspended in 0.50 mL PBS with 1-mm zirconia-silica homogenization beads (Biospec), and subsequently homogenized for three 1-minute intervals. The tissue homogenates were serially diluted and plated on nonselective (TSA) and selective (mannitol salt agar [MSA]) media. Plates were incubated overnight before counting colonies.
Whole-genome assembly and annotation
Genomic DNA from S. hominis AH5011 and S. aureus AH6350 were isolated by phenol-chloroform extraction for whole-genome sequencing, genome assembly, and annotation at the SeqCenter facility (Pittsburgh, USA). SeqCenter-prepared DNA libraries were sequenced on the NextSeq 2000 Illumina and the ONT MinION sequencer. Quality control and adapter trimming were performed with blc2fastq (version 2.20.0.445) and porechop (version 0.2.3_seqan2.1.1) for Illumina and Oxford Nanopore sequencing, respectively. Unicycler (version 0.5.0) was used to assemble Illumina and ONT reads78. Bandage (version 0.8.1)79 and BUSCO (version 5.2.2)80 were used to assess assembly completeness. Genomes were annotated using Prokka (version 1.14.5)81. BAGEL4 was used to detect biosynthetic gene clusters19. Multi-locus sequence typing of S. aureus genomes was performed using PubMLST database82. Unless otherwise stated, default parameters were used for all software. The hom BGC and its homologs were identified using BLASTp. Genomes of various staphylococci containing the hom BGC were downloaded from NCBI and are listed in Supplementary Table 3. Comparative gene cluster analysis and visualization were performed using CAGECAT, a web-based platform that integrates Cblaster83 and Clinker84 modules. Multi-sequence alignment was performed using Clustal Omega85 and visualized using Geneious Prime (version 2024.0.7). Structural protein homology was analyzed and visualized using Foldseek (version 10) and PyMOL (version 3.0.3).
Quantification and statistical analysis
Statistical details of experiments, including the methods used and defined significance values, can be found in the figure legends. All statistical analyses were performed with GraphPad Prism software (version 10.4.0). To determine the statistical significance of differences between two groups, we used two-tailed Student’s t-tests. For multiple comparisons involving more than two groups, we chose a one-way analysis of variance (ANOVA) followed by Dunnett’s multiple comparison test. For bacterial growth curve experiments, we selected a repeated measures two-way ANOVA analysis.
Ethics statement
All vertebrate animal experiments were approved and conducted in accordance with the Institutional Animal Care and Use Committee of the University of Colorado Anschutz Medical Campus under protocol number 00941. Eight-week-old male and female C57BL6/J mice (strain #000664) were obtained from the Jackson Laboratory (Bar Harbor, ME) and housed in specific pathogen-free facilities at the University of Colorado Anschutz Medical Campus Animal Care Facility under controlled conditions (12-h light/dark cycle; temperature: 20-22 °C; humidity: 30-70%). Mice were provided ad libitum access to food and water and allowed to acclimate for 1 week prior to experimentation. At experimental endpoints, mice were euthanized via CO2 inhalation followed by cervical dislocation. This method is consistent with the recommendations of the American Veterinary Medical Association (AVMA) Guidelines for the Euthanasia of Animals. All procedures involving human blood samples were conducted in accordance with guidelines approved by the Colorado Multiple Institutional Review Board (COMIRB) under protocol number 17-1926 and the Institutional Biosafety Committee under protocol number IBC-00001264. Informed consent was obtained from all subjects prior to participation.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings of this study are presented in the article, supplementary information, and source data file. Reagents and strains created in this study are available upon request to the corresponding author. The raw sequencing data have been deposited in the NCBI Sequence Read Archive under SRS24496481 and SRS24496482. The nucleotide sequencing data for S. hominis AH5011 and S. aureus AH6350 have been deposited in the National Center for Biotechnology Information (NCBI) GenBank under BioProject PRJNA1222585 and accession codes JBLOOY000000000 and CP183426. The genomes used in this study are listed in Supplementary Table 3. The mass spectrometry data have been deposited in the Zenodo database. Links to the deposited data can be found here: Hominicin LC-MS Quantification [https://zenodo.org/records/14834706], S. hominis spent media LC-MS and MS/MS [https://zenodo.org/records/14833876], Staphylococci Knockout LC-MS Screening [https://zenodo.org/records/14834696], and Staphylococci Knockout Screening 2 LC-MS [https://zenodo.org/records/16877503]. The NMR data have been deposited in the Natural Products Magnetic Resonance Database (NP-MRD) under project code NP0350779 [https://doi.org/10.57994/3763]. Source data are provided with this paper.
References
Chen, Y. E., Fischbach, M. A. & Belkaid, Y. Skin microbiota-host interactions. Nature 553, 427–436 (2018).
Harris-Tryon, T. A. & Grice, E. A. Microbiota and maintenance of skin barrier function. Science 376, 940–945 (2022).
Becker, K., Heilmann, C. & Peters, G. Coagulase-negative staphylococci. Clin. Microbiol Rev. 27, 870–926 (2014).
Parlet, C. P., Brown, M. M. & Horswill, A. R. Commensal Staphylococci influence staphylococcus aureus skin colonization and disease. Trends Microbiol 27, 497–507 (2019).
Severn, M. M. et al. The ubiquitous human skin commensal staphylococcus hominis protects against opportunistic pathogens. mBio 13, e0093022 (2022).
Williams, M. R. et al. Quorum sensing between bacterial species on the skin protects against epidermal injury in atopic dermatitis. Sci. Transl. Med 11, eaat8329 (2019).
Nakatsuji, T. et al. Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Sci. Transl. Med 9, eaah4680 (2017).
Liu, Y. et al. Skin microbiota analysis-inspired development of novel anti-infectives. Microbiome 8, 85 (2020).
Aftab Uddin, M. et al. A plant endophyte Staphylococcus hominis strain MBL_AB63 produces a novel lantibiotic, homicorcin and a position one variant. Sci. Rep. 11, 11211 (2021).
Arnison, P. G. et al. Ribosomally synthesized and post-translationally modified peptide natural products: overview and recommendations for a universal nomenclature. Nat. Prod. Rep. 30, 108–160 (2013).
Dobson, A., Cotter, P. D., Ross, R. P. & Hill, C. Bacteriocin production: a probiotic trait?. Appl Environ. Microbiol 78, 1–6 (2012).
Nakatsuji, T. et al. Development of a human skin commensal microbe for bacteriotherapy of atopic dermatitis and use in a phase 1 randomized clinical trial. Nat. Med 27, 700–709 (2021).
Montalban-Lopez, M. et al. New developments in RiPP discovery, enzymology and engineering. Nat. Prod. Rep. 38, 130–239 (2021).
Ren, H. et al. Genome mining unveils a class of ribosomal peptides with two amino termini. Nat. Commun. 14, 1624 (2023).
Nguyen, D. T., Mitchell, D. A. & van der Donk, W. A. Genome Mining for New Enzyme Chemistry. ACS Catal. 14, 4536–4553 (2024).
Nguyen, N. A., Forstater, J. H. & McIntosh, J. A. Decarboxylation in natural products biosynthesis. JACS Au 4, 2715–2745 (2024).
Kim, P. I. et al. Characterization and structure identification of an antimicrobial peptide, hominicin, produced by Staphylococcus hominis MBBL 2-9. Biochem Biophys. Res Commun. 399, 133–138 (2010).
Heilbronner, S., Krismer, B., Brotz-Oesterhelt, H. & Peschel, A. The microbiome-shaping roles of bacteriocins. Nat. Rev. Microbiol 19, 726–739 (2021).
van Heel, A. J. et al. BAGEL4: a user-friendly web server to thoroughly mine RiPPs and bacteriocins. Nucleic Acids Res 46, W278–W281 (2018).
Knerr, P. J. & van der Donk, W. A. Discovery, biosynthesis, and engineering of lantipeptides. Annu Rev. Biochem 81, 479–505 (2012).
Pei, Z. F., Zhu, L. & Nair, S. K. Core-dependent post-translational modifications guide the biosynthesis of a new class of hypermodified peptides. Nat. Commun. 14, 7734 (2023).
Freeman, M. F. et al. Metagenome mining reveals polytheonamides as posttranslationally modified ribosomal peptides. Science 338, 387–390 (2012).
Yang, X. & van der Donk, W. A. Post-translational introduction of D-Alanine into ribosomally synthesized peptides by the dehydroalanine reductase NpnJ. J. Am. Chem. Soc. 137, 12426–12429 (2015).
Lohans, C. T., Li, J. L. & Vederas, J. C. Structure and biosynthesis of carnolysin, a homologue of enterococcal cytolysin with D-amino acids. J. Am. Chem. Soc. 136, 13150–13153 (2014).
Cotter, P. D. et al. Posttranslational conversion of L-serines to D-alanines is vital for optimal production and activity of the lantibiotic lacticin 3147. Proc. Natl Acad. Sci. USA 102, 18584–18589 (2005).
Eslami, S. M. & van der Donk, W. A. Proteases involved in leader peptide removal during RiPP biosynthesis. ACS Bio Med Chem. Au 4, 20–36 (2024).
Rivera-Chavez, J. et al. Biosynthesis of fluorinated peptaibols using a site-directed building block incorporation approach. J. Nat. Prod. 80, 1883–1892 (2017).
Weinstein, M. P. & Patel, J. B. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically: M07-A11. 11. edition edn, 92 (Committee for Clinical Laboratory Standards, 2018).
Yamasaki, K. & Gallo, R. L. Antimicrobial peptides in human skin disease. Eur. J. Dermatol 18, 11–21 (2008).
Yasir, M., Dutta, D. & Willcox, M. D. P. Mode of action of the antimicrobial peptide Mel4 is independent of Staphylococcus aureus cell membrane permeability. PLoS One 14, e0215703 (2019).
Farha, M. A., Verschoor, C. P., Bowdish, D. & Brown, E. D. Collapsing the proton motive force to identify synergistic combinations against Staphylococcus aureus. Chem. Biol. 20, 1168–1178 (2013).
Belmonte, G. et al. Pore formation by Staphylococcus aureus alpha-toxin in lipid bilayers. Dependence upon temperature and toxin concentration. Eur. Biophys. J. 14, 349–358 (1987).
Salam, A. M. et al. Castaneroxy a from the leaves of castanea sativa inhibits vIRULENCe in Staphylococcus aureus. Front Pharm. 12, 640179 (2021).
van den Belt, M. et al. CAGECAT: the comparative gene cluster analysis toolbox for rapid search and visualisation of homologous gene clusters. BMC Bioinforma. 24, 181 (2023).
Smits, S. H. J., Schmitt, L. & Beis, K. Self-immunity to antibacterial peptides by ABC transporters. FEBS Lett. 594, 3920–3942 (2020).
Bastos Mdo, C., Coelho, M. L. & Santos, O. C. Resistance to bacteriocins produced by Gram-positive bacteria. Microbiol. (Read.) 161, 683–700 (2015).
Stein, T., Heinzmann, S., Solovieva, I. & Entian, K. D. Function of Lactococcus lactis nisin immunity genes nisI and nisFEG after coordinated expression in the surrogate host Bacillus subtilis. J. Biol. Chem. 278, 89–94 (2003).
Diep, D. B., Skaugen, M., Salehian, Z., Holo, H. & Nes, I. F. Common mechanisms of target cell recognition and immunity for class II bacteriocins. Proc. Natl Acad. Sci. USA 104, 2384–2389 (2007).
Omasits, U., Ahrens, C. H., Muller, S. & Wollscheid, B. Protter: interactive protein feature visualization and integration with experimental proteomic data. Bioinformatics 30, 884–886 (2014).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
van Kempen, M. et al. Fast and accurate protein structure search with Foldseek. Nat. Biotechnol. 42, 243–246 (2024).
Wohlwend, J. et al. Boltz-1 Democratizing Biomolecular Interaction Modeling. Preprint at bioRxiv 10.1101/2024.11.19.624167 (2024).
Ray, G. T., Suaya, J. A. & Baxter, R. Incidence, microbiology, and patient characteristics of skin and soft-tissue infections in a U.S. population: a retrospective population-based study. BMC Infect. Dis. 13, 252 (2013).
Vella, V. et al. The incidence of skin and soft tissue infections in the united states and associated healthcare utilization between 2010 and 2020. Open Forum Infect. Dis. 11, ofae267 (2024).
Nutten, S. Atopic dermatitis: global epidemiology and risk factors. Ann. Nutr. Metab. 66 Suppl 1, 8–16 (2015).
Wertheim, H. F. et al. The role of nasal carriage in Staphylococcus aureus infections. Lancet Infect. Dis. 5, 751–762 (2005).
Monecke, S., Luedicke, C., Slickers, P. & Ehricht, R. Molecular epidemiology of Staphylococcus aureus in asymptomatic carriers. Eur. J. Clin. Microbiol Infect. Dis. 28, 1159–1165 (2009).
Crosby, H. A. et al. Host-derived protease promotes aggregation of Staphylococcus aureus by cleaving the surface protein SasG. mBio 15, e0348323 (2024).
Mills, K. B. et al. Staphylococcus aureus skin colonization is mediated by SasG lectin variation. Cell Rep. 43, 114022 (2024).
Yang, C. et al. Genomic epidemiology and phenotypic characterization of Staphylococcus aureus isolated from atopic dermatitis patients in South China. Sci. Rep. 15, 4773 (2025).
Saheb Kashaf, S. et al. Staphylococcal diversity in atopic dermatitis from an individual to a global scale. Cell Host Microbe 31, 578–592 e576 (2023).
Liu, H. et al. Staphylococcus aureus Epicutaneous Exposure Drives Skin Inflammation via IL-36-Mediated T Cell Responses. Cell Host Microbe 22, 653–666.e655 (2017).
O’Sullivan, J. N., Rea, M. C., O’Connor, P. M., Hill, C. & Ross, R. P. Human skin microbiota is a rich source of bacteriocin-producing staphylococci that kill human pathogens. FEMS Microbiol Ecol. 95, fiy241 (2019).
Riley, M. A. & Wertz, J. E. Bacteriocins: evolution, ecology, and application. Annu Rev. Microbiol 56, 117–137 (2002).
Grice, E. A. & Segre, J. A. The skin microbiome. Nat. Rev. Microbiol 9, 244–253 (2011).
Byrd, A. L., Belkaid, Y. & Segre, J. A. The human skin microbiome. Nat. Rev. Microbiol 16, 143–155 (2018).
Johansson, L., Thulin, P., Low, D. E. & Norrby-Teglund, A. Getting under the skin: the immunopathogenesis of Streptococcus pyogenes deep tissue infections. Clin. Infect. Dis. 51, 58–65 (2010).
Fernandez-Lopez, S. et al. Antibacterial agents based on the cyclic D,L-alpha-peptide architecture. Nature 412, 452–455 (2001).
Dieterich, C. L. et al. Aquimarins, peptide antibiotics with amino-modified c-termini from a sponge-derived aquimarina sP. BACTerium. Angew. Chem. Int Ed. Engl. 61, e202115802 (2022).
Claesen, J. & Bibb, M. Genome mining and genetic analysis of cypemycin biosynthesis reveal an unusual class of posttranslationally modified peptides. Proc. Natl Acad. Sci. USA 107, 16297–16302 (2010).
Molohon, K. J. et al. Structure determination and interception of biosynthetic intermediates for the plantazolicin class of highly discriminating antibiotics. ACS Chem. Biol. 6, 1307–1313 (2011).
Kalyon, B. et al. Plantazolicin A and B: structure elucidation of ribosomally synthesized thiazole/oxazole peptides from Bacillus amyloliquefaciens FZB42. Org. Lett. 13, 2996–2999 (2011).
Shahmiri, M. & Mechler, A. The role of C-terminal amidation in the mechanism of action of the antimicrobial peptide aurein 1.2. EuroBiotech J. 4, 25–31 (2020).
Zhu, S., Li, W., O’Brien-Simpson, N., Separovic, F. & Sani, M. A. C-terminus amidation influences biological activity and membrane interaction of maculatin 1.1. Amino Acids 53, 769–777 (2021).
Dennison, S. R. & Phoenix, D. A. Influence of C-terminal amidation on the efficacy of modelin-5. Biochemistry 50, 1514–1523 (2011).
Cintas, L. M., Casaus, P., Havarstein, L. S., Hernandez, P. E. & Nes, I. F. Biochemical and genetic characterization of enterocin P, a novel sec-dependent bacteriocin from Enterococcus faecium P13 with a broad antimicrobial spectrum. Appl Environ. Microbiol 63, 4321–4330 (1997).
Kuipers, A. et al. Sec-mediated transport of posttranslationally dehydrated peptides in Lactococcus lactis. Appl Environ. Microbiol 72, 7626–7633 (2006).
Kumazaki, K. et al. Structural basis of Sec-independent membrane protein insertion by YidC. Nature 509, 516–520 (2014).
Shiota, N., Shimokawa-Chiba, N., Fujiwara, K. & Chiba, S. Identification of Bacillus subtilis YidC Substrates Using a MifM-instructed Translation Arrest-based Reporter. J. Mol. Biol. 435, 168172 (2023).
Froderberg, L., Houben, E. N., Baars, L., Luirink, J. & de Gier, J. W. Targeting and translocation of two lipoproteins in Escherichia coli via the SRP/Sec/YidC pathway. J. Biol. Chem. 279, 31026–31032 (2004).
Donia, M. S. et al. A systematic analysis of biosynthetic gene clusters in the human microbiome reveals a common family of antibiotics. Cell 158, 1402–1414 (2014).
Cau, L. et al. Staphylococcus epidermidis protease EcpA can be a deleterious component of the skin microbiome in atopic dermatitis. J. Allergy Clin. Immunol. 147, 955–966.e916 (2021).
Boles, B. R., Thoendel, M., Roth, A. J. & Horswill, A. R. Identification of genes involved in polysaccharide-independent Staphylococcus aureus biofilm formation. PLoS One 5, e10146 (2010).
Schenk, S. & Laddaga, R. A. Improved method for electroporation of Staphylococcus aureus. FEMS Microbiol Lett. 73, 133–138 (1992).
Zaitseva, E. et al. Electrophysiology on channel-forming proteins in artificial lipid bilayers: next-generation instrumentation for multiple recordings in parallel. Methods Mol. Biol. 2188, 67–92 (2021).
Smits, J. P. H. et al. Immortalized N/TERT keratinocytes as an alternative cell source in 3D human epidermal models. Sci. Rep. 7, 11838 (2017).
Patrick, G. J. et al. Epicutaneous Staphylococcus aureus induces IL-36 to enhance IgE production and ensuing allergic disease. J. Clin. Invest 131, e143334 (2021).
Wick, R. R., Judd, L. M., Gorrie, C. L. & Holt, K. E. Unicycler: Resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput Biol. 13, e1005595 (2017).
Wick, R. R., Schultz, M. B., Zobel, J. & Holt, K. E. Bandage: interactive visualization of de novo genome assemblies. Bioinformatics 31, 3350–3352 (2015).
Manni, M., Berkeley, M. R., Seppey, M. & Zdobnov, E. M. BUSCO: Assessing Genomic Data Quality and Beyond. Curr. Protoc. 1, e323 (2021).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Jolley, K. A., Bray, J. E. & Maiden, M. C. J. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 3, 124 (2018).
Gilchrist, C. L. M. et al. cblaster: a remote search tool for rapid identification and visualization of homologous gene clusters. Bioinform Adv. 1, vbab016 (2021).
Gilchrist, C. L. M. & Chooi, Y. H. clinker & clustermap.js: automatic generation of gene cluster comparison figures. Bioinformatics 37, 2473–2475 (2021).
Madeira, F. et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 47, W636–W641 (2019).
Acknowledgements
A.R.H., N.B.C., and R.L.G. are supported by the National Institute for Allergy and Infectious Diseases (NIAID) grant R01AI153185. Z.L.B. received support from T32AT008938 from the National Center for Complementary and Integrative Health (NCCIH). J.R.C. is supported by the National Institute of General Medical Sciences of the National Institutes of Health (R35GM147439). We thank Dr. Emily Gurnee at Children’s Hospital Colorado for providing clinical isolates and SeqCenter (Pittsburgh, USA) for their technical assistance with whole-genome sequencing. Mass spectrometric data were collected in the Triad Mass Spectrometry Facility, and we thank Dr. Warren Vidar for his help with data collection. This work was performed in part at the Joint School of Nanoscience and Nanoengineering, a member of the National Nanotechnology Coordinated Infrastructure (NNCI), which is supported by the National Science Foundation (Grant ECCS-2025462).
Author information
Authors and Affiliations
Contributions
A.N. and Z.L.B. contributed equally to this work. A.N. performed direct cloning, heterologous expression, bioactivity assays, bioinformatics, membrane-based assays, and animal experiments. Z.L.B. performed compound purifications, UPLC-MS analysis, MIC assays, and stereochemical and functional group determination with oversight from N.B.C. I.Y.M., M.R., and T.N.G. designed, performed, and analyzed NMR experiments with oversight from N.H.O. U.T. assisted the design, collection, and analysis of electrophysiological recordings. J.C.T. performed and analyzed the AlphaFold and ligand modeling. M.M.S. designed and performed the bioactivity assays. T.N. designed and collected bacterial isolates from skin swabs with oversight from R.L.G. J.R.C. analyzed the bioinformatics data and stereochemical and functional group determination. A.N. and Z.L.B wrote the manuscript with editorial oversight from A.R.H. and N.B.C., along with input from all other authors. N.B.C. and A.R.H. conceived of and supervised the overall project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Daniel Wozniak, who co-reviewed with Pranav Rana, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nguyen, A., Bunch, Z.L., Martinez-Aldino, I.Y. et al. An antimicrobial daptide from human skin commensal Staphylococcus hominis protects against skin pathogens. Nat Commun 16, 11459 (2025). https://doi.org/10.1038/s41467-025-66259-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66259-w