Abstract
The WEE1 kinase negatively regulates CDK1/2 to control DNA replication and mitotic entry. Genetic factors that determine sensitivity to WEE1 inhibitors (WEE1i) are largely unknown. A genome-wide insertional mutagenesis screen revealed that mutation of EIF2A, a translation regulator, sensitized to WEE1i. Additionally, a genome-wide CRISPR-Cas9 screen revealed that inactivation of integrated stress response (ISR) kinase GCN2 or its co-factor GCN1 rescued WEE1i-mediated cytotoxicity. Conversely, loss of the collided ribosome sensor ZNF598 increased sensitivity to WEE1i. Mechanistically, WEE1i induced paradoxical GCN2 activation, ATF4 upregulation, and altered ribosome dynamics. ISR activation was independent of WEE1 presence, pointing at off-target GCN2 engagement by multiple chemically distinct WEE1i. ISR activation was observed in cancer cells as well as non-transformed cells, and required GCN1 and ongoing translation. Consequently, WEE1i induce multiple independent cellular effects: DNA damage, premature mitotic entry and sensitization to DNA-damaging chemotherapeutics in an ISR-independent fashion, as well as ISR activation independently of CDK1/2 activation. Importantly, low-dose WEE1 inhibition did not induce ISR activation, while it still synergized with PKMYT1 inhibition. Taken together, WEE1i trigger toxic ISR activation and translational shutdown, which can be prevented by low-dose or combination treatments, while retaining the cell cycle checkpoint-perturbing effects.
Similar content being viewed by others
Introduction
Cell cycle progression is driven by the temporally controlled activation of specific cyclin-dependent kinases (CDKs) in complex with their cognate cyclin partners1. Although the roles and regulation of cyclin-CDKs in mammalian cells are complex, distinct roles of specific cyclin-CDKs have been defined; entry into the cell cycle is largely driven by CDK4/6 in complex with cyclin D, initiation and progression of S-phase is promoted by CDK2 in complex with cyclin E and cyclin A, whereas cyclin B1-CDK1 drives entry into mitosis1.
An important regulatory mechanism controlling the activity of CDK1 and CDK2 involves inhibitory phosphorylation. Both CDK1 and CDK2 are phosphorylated on Tyr15 (Y15) by the WEE1 kinase2,3,4, which limits their activity and safeguards DNA replication and prevents premature entry into mitosis, respectively5. In addition, CDK1 is phosphorylated on Thr14 (T14) by the PKMYT1 kinase (also known as Myt1) to prevent unscheduled CDK1 activation6,7. To activate CDK1/2, the inhibitory phosphorylation at Y15 and T14 is removed by CDC25 phosphatases8. In situations of DNA damage, the kinases ATR and ATM trigger activation of the downstream checkpoint kinases CHK1 and CHK29, which subsequently phosphorylate and inhibit CDC258,10. In parallel, CHK1 activates WEE111. Consequently, DNA lesions and incomplete DNA replication lead to maintenance of CDK1/2 inhibitory phosphorylation, which arrests cell cycle progression and allows cells to repair DNA or complete replication.
Inactivation of cell cycle control is cytotoxic in situations of cell-intrinsic or therapy-induced DNA damage12,13. This concept is therapeutically exploited by development of a range of cell cycle checkpoint inhibitors, including inhibitors of WEE114. WEE1 inhibitors (WEE1i) were demonstrated to be preferentially cytotoxic in TP53 mutant cells14,15, as well as in cells with elevated levels of replication stress, due to oncogene overexpression16,17 or nucleotide deficiency18. Moreover, WEE1i treatment has been demonstrated to sensitize tumor cells to a range of genotoxic chemotherapeutic agents19,20,21,22.
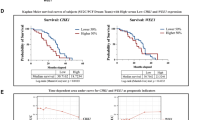
Initial clinical studies focused on combination treatment of the WEE1i adavosertib (AZD1775) with platinum-based chemotherapeutics in patients with high-grade serous ovarian cancer. This combination treatment showed promising results, although dose-limiting toxicities occurred23,24,25,26. Although all included patients harbored TP53 mutation in their tumors, considerable variation in response to WEE1i treatment was observed. Clearly, other tumor-associated features beyond TP53 mutations influence WEE1i response. In this study, we explore genetic determinants of WEE1i treatment response.
Results
A haploid genetic screen identifies genes determining sensitivity to the WEE1 inhibitor AZD1775
To identify gene mutations that sensitize cells to inhibition of WEE1, an insertional mutagenesis screen was performed in near-haploid HAP1 cells. To this end, HAP1 cells were mutagenized using a gene trap virus encoding a strong splice acceptor (Supplementary Fig. 1a)27, and subsequently treated with the WEE1i AZD1775 (Fig. 1a). Several genes were identified that conferred sensitivity to AZD1775 treatment upon inactivation (Fig. 1b), including multiple genes involved in the one-carbon metabolic pathway, which uses folate to catabolize serine into purines and pyrimidines28. Specifically, MTHFD1, MTHFD1L, MTHFD2, and SHMT2 of this pathway were identified (Fig. 1b). Of note, pharmacological targeting of MTHFD2 was previously shown to induce replication stress and increase sensitivity to cell cycle checkpoint inhibitors, including WEE1i29. Furthermore, we identified ubiquitin ligase subunits, including the F-box factor FBXW7 that regulates cyclin E1/CDK2 activity, which was previously described to determine WEE1i sensitivity16,17,30. Additionally, inactivation of the PP2A phosphatase activator PTPA and PPP2R5E (encoding the PP2A subunit B56ε) conferred sensitivity to AZD1775, in line with a previously described role for PP2A in regulating the WEE1/CDC25C pathway31. Identification of these genes underscored the validity of our screening approach.
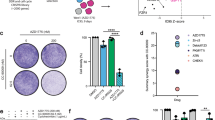
a Experimental setup of the viral insertional mutagenesis screen in (b). b Sense/total insertion ratios from mutagenesis screens performed in HAP1 cells treated with AZD1775 or DMSO. Genes with significantly less sense insertions in AZD1775-treated cells are indicated. c Schematic representation of high-ranking gene mutations causing sensitization to WEE1i. d Pathway enrichment analysis, depicting the most highly enriched pathways involved in sensitization to WEE1i. e, f Dose-response curve of AZD1775 in control RPE1 TP53KO and RPE1 TP53KO EIF2AKO cells (e) and OVCAR3, OVCAR3 EIF2AKO #1 and OVCAR3 EIF2AKO #2 cells (f). Cells were treated for 5 days and cellular viability was measured by MTT conversion. Data represent mean ± standard deviation (SD) (n = 3). g Immunofluorescence microscopy analysis of γH2AX in control RPE1 TP53KO and EIF2AKO cells treated with AZD1775 (500 nM) for 24 h. Mean foci intensity per cell is plotted. Medians of n = 218, 212, 228 or 206 cells, respectively, measured across 3 independent experiments. Paired t-test (two-sided) of medians, p ≤ 0.05 was considered significant. h scEdU-seq maximum normalized log counts are plotted for control RPE1 TP53KO and RPE1 TP53KO EIF2AKO cells, ranked according to S-phase progression (y-axis) and binned per 400 kb (x-axis). A 40 megabase region of chromosome 2 is shown. All replicates are biological replicates unless indicated otherwise. Source data are provided as a Source data file.
Remarkably, we identified multiple genes involved in the regulation of mRNA translation, including many aminoacyl-tRNA synthetases, such as NARS2, RARS2, CARS2, LARS2, SARS2, and WARS1 (Fig. 1b–d). Inactivation of these genes mildly affected the viability of control cells but further reduced the viability of WEE1-inhibited cells. In contrast, mutation of EIF2A did not affect HAP1 cell viability but caused a significant loss of viability upon WEE1 inhibition. eIF2A was initially implicated in mRNA translation using alternative initiation sites upon stress32, with a reported interaction of eIF2A along with eIF5B33 to 40S ribosome subunits34. Recent reports, however, demonstrated that eIF2A in human cells has a limited role in alternative translation35,36. Instead, eIF2A was suggested to regulate 40S ribosome subunit recycling36. Nevertheless, the exact role of eIF2A in translation remains unclear35,37,38. To confirm its sensitization to WEE1i, EIF2A was inactivated using CRISPR/Cas9 in HAP1, RPE1 TP53KO, and OVCAR3 cells (Supplementary Figure 1b–f). In a panel of HAP1-EIF2A knockout (KO) clones, decreased clonogenic survival was observed when cells were treated with WEE1i (Supplementary Fig. 1b). Likewise, inactivation of eIF2A in RPE1 TP53KO and OVCAR3 cells conferred WEE1i sensitivity in short-term viability assays (Fig. 1e, f) and clonogenic survival assays (Supplementary Fig. 1c–f). Combined, these results indicate that loss of eIF2A leads to cellular sensitivity to the WEE1i AZD1775.
WEE1i sensitivity is not explained by increased replication stress in cells lacking eIF2A
We hypothesized that loss of EIF2A may sensitize cells to WEE1 inhibition by increasing DNA replication stress and DNA damage. Baseline levels of mean γH2AX foci intensity and pan-nuclear γH2AX in EIF2AKO cells were similar to those of control RPE1 TP53KO cells (Fig. 1g and Supplementary Fig. 1i, j). Upon AZD1775 treatment, both control cells and EIF2AKO cells showed increased levels of γH2AX foci intensity (Fig. 1g), with EIF2AKO cells showing a slightly stronger increase in pan-nuclear γH2AX (Supplementary Fig. 1i, j).
In-depth analysis of replication dynamics using single-cell EdU sequencing (scEdUseq) did not reveal obvious differences between control-treated RPE1 TP53KO and EIF2AKO cells (Fig. 1h and Supplementary Fig. 2a). Also, upon AZD1775 treatment, control and EIF2AKO RPE1 TP53KO cells showed similar replication dynamics (Supplementary Fig. 2b, c). In addition, both the overall number of DNA replication forks per cell and the distribution of active replication forks over the various stages of S-phase were similar between control RPE1 TP53KO and EIF2AKO cells, and did not show obvious changes upon AZD1775 treatment (Supplementary Fig. 2d–f). DNA fiber analysis confirmed these findings (Supplementary Fig. 3a), while expected WEE1i-mediated perturbation of cell cycle dynamics was observed (Supplementary Fig. 3b). Only at high-dose AZD1775 treatment (1 µM), DNA replication was dramatically perturbed (Supplementary Fig. 3c), which coincided with a strong decrease in active replication forks per cell (Supplementary Figure 3d, e). Importantly, the observation that replication dynamics were not perturbed in EIF2AKO cells after low-dose AZD1775 treatment (Supplementary Fig. 2b, c), at which dose EIF2AKO cells already showed increased AZD1775 sensitivity (Fig. 1e, f) suggests that alterations in the DNA replication program are not driving the increased sensitivity of EIF2AKO cells to WEE1 inhibition.
WEE1 inhibition induces the integrated stress response through GCN2 activation
Comparison of baseline mRNA expression levels between control RPE1 TP53KO and EIF2AKO cells revealed significant downregulation of gene sets related to mRNA translation (Fig. 2a and Supplementary Fig. 4a and Supplementary Data 1), in line with a proposed regulatory role for eIF2A in translation upon integrated stress response activation39,40. Proteomics analysis using stable isotope labeling by amino acids in cell culture (SILAC) confirmed loss of eIF2A protein in EIF2AKO cells, while none of the other identified hit from the genetic screen showed altered abundance, indicating that effects of EIF2A inactivation are not likely explained by indirect inactivation of other identified genes (Fig. 2b). Treatment with AZD1775 induced altered expression of translation regulators, including LARP141,42,43, specifically in EIF2AKO cells (Fig. 2c). Using puromycin or L-azidohomoalanine (AHA) incorporation assays to measure ongoing mRNA translation, we observed that two chemically distinct WEE1i, AZD1775 and Debio 0123, decreased ongoing mRNA translation in RPE1 TP53KO cells (Fig. 2d, e). Importantly, mRNA translation attenuation upon WEE1i-treatment was even more pronounced in EIF2AKO cells (Fig. 2d). Downregulation of translation upon WEE1i treatment was confirmed in HCC38 triple-negative breast cancer cells (Supplementary Fig. 4b). WEE1i-mediated attenuation of translation was rescued by inhibition of the integrated stress response (ISR) using ISRIB44 (Fig. 2e). To provide insights into how WEE1 inhibition affects mRNA translation, we performed a CRISPR-based screen to identify gene mutations that confer resistance to AZD1775 in RPE1 hTERT TP53KO cells. (Fig. 2f). The strongest hits included the eIF2α kinase GCN2 (EIF2AK4) and its coactivator GCN1 (GCN1L1), which function in the integrated stress response, as well as negative mTORC1 regulators TSC1 and TSC2 (Fig. 2g)45. To corroborate these results, RPE1 GCN2KO and GCN1KO cells were generated, and WEE1i sensitivity was analyzed. Indeed, both GCN2KO and GCN1KO cells were less sensitive to AZD1775 treatment compared to RPE1 control cells in short-term proliferation assays (Fig. 2h), clonogenic survival assays (Supplementary Fig. 4c, d), and cell competition assays (Supplementary Fig. 4e, f).
a Gene set enrichment analysis of RNA-seq data of RPE1 TP53KO control and EIF2AKO cells (more detailed analysis in Supplementary Fig. 3a). Circle size represents gene set size (control: n = 3, EIF2AKO: n = 1). b, c SILAC-based proteomics analysis of RPE1 TP53KO control and EIF2AKO cells, showing differential protein expression at baseline (b) and upon AZD1775 treatment for 6, 24 and 48 h (c) (n = 2). Red datapoint indicates eIF2A, green datapoints indicate other screen hits. d RPE1 TP53KO control and EIF2AKO cells were treated with AZD1775 or Debio 0123 (1 µM) for 24 h, followed by a 10’ puromycin pulse. Puromycin incorporation was visualized by immunoblotting (left) and quantified (right). Data represent mean ± SD; RPE1 TP53KO EIF2AKO treated with Debio 0123 (n = 4); control RPE1 TP53KO treated with DMSO or AZD1775 (n = 7); Other conditions (n = 5). e RPE1 TP53KO PACKO cells were treated with AZD1775 (250 nM) and ISRIB for 24 h and subsequently labeled with L-azidohomoalanine (AHA). Immunoblot of AHA-labeled proteins and quantification is shown. Data represent mean and individual datapoints (n = 2). f Schematic overview of the CRISPR/Cas9 genome-wide AZD1775 resistance screen. g MAGeCK scores for individual genes in the RPE1 TP53KO AZD1775 positive selection screen (day 12 vs day 0). h Cell confluency analysis of parental RPE1 TP53KO PACKO or GCN2KO cells. Mean ± range, n = 3 for control and GCN2KO #1, n = 2 for GCN2KO #2. i Quantification of clonogenic survival assays of parental RPE1 TP53KO PACKO and ATF4KO cells (corresponding images provided in Supplementary Fig. 4h). Mean ± SD, n = 3 for all conditions. j, k Representative histograms (j) and quantification (k) of ATF4-mScarlet flow cytometry measurements in RPE1 TP53KO ATF4-mScarlet reporter cells treated with thapsigargin (1 µM), AZD1775 (1 µM), Debio 0123 (1 µM) and/or A92 (1 µM) for 24 h. Data represent mean ± SD (n = 3). Statistical analysis of (d, k): unpaired t-test (two-sided) with p ≤ 0.05 considered significant. All replicates are biological replicates unless indicated otherwise. Source data are provided as a Source data file.
Subsequently, we analyzed whether signaling downstream of GCN2 also determined AZD1775-induced cytotoxicity. Time course analysis of AZD1775-induced activation of GCN2 and the downstream transcription factor ATF446 showed GCN2 phosphorylation and ATF4 upregulation already at 30 min after AZD1775 treatment (Supplementary Fig. 4g). Inactivation of ATF4 led to reduced sensitivity of RPE1 TP53KO cells to AZD1775 (Fig. 2i and Supplementary Fig. 4h). These findings were confirmed in RPE1 TP53KO cells harboring a fluorescent ATF4 reporter, consisting of two upstream open reading frames (uORFs) and the first 84 base pairs of the ATF4 gene (ORF3) fused to an mScarlet cassette47. Under unstressed conditions, only the two uORFs are translated, whereas upon ISR activation, delayed translation reinitiation leads to ribosome trailing past the initiation site of uORF2 and translation of the ATF4-mScarlet fusion protein47. Thapsigargin, a known activator of the ISR via the PERK kinase, confirmed the ability of these reporter cell lines to read out ATF4 upregulation (Supplementary Fig. 4i, j). Treatment with AZD1775 or the chemically distinct WEE1i Debio 0123 led to ATF4-mScarlet upregulation, which was fully rescued by pharmacological GCN2 inhibition (Fig. 2j, k and Supplementary Fig. 4i, j). Supporting this finding, pharmacological inhibition of GCN2 also rescued AZD1775-mediated reduction of mRNA translation in the breast cancer cell line HCC38 (Supplementary Fig. 4b). Taken together, these data show that WEE1i trigger activation of the ISR and decrease global mRNA translation in a GCN2-dependent manner.
ISR induction and cell cycle deregulation are independent consequences of WEE1i
We next analyzed whether GCN2 activation by WEE1 inhibition is dependent on cell cycle status. Quantitative image-based cytometry (QIBC) showed that ATF4 expression was upregulated upon AZD1775 treatment in RPE1 TP53KO cells throughout the cell cycle, which was prevented by ISRIB co-treatment (Fig. 3a). To further investigate whether ATF4 upregulation by AZD1775 requires ongoing cell cycle progression, RPE1 TP53KO were synchronized in mitosis using the microtubule-poison nocodazole and released into a round of cell division. Alternatively, cells were arrested in G1 phase after release from mitosis, using the CDK4/6 inhibitor palbociclib (Fig. 3b). Subsequently, cells were treated with AZD1775 and/or the GCN2 inhibitor A92. A palbociclib-mediated arrest in G1 was confirmed by low abundance of cyclin B (Supplementary Fig. 5a). AZD1775 treatment resulted in upregulation of ATF4, both in proliferating cells as well as G1-arrested cells (Fig. 3c and Supplementary Fig. 5b). Conversely, GCN2 inhibition prevented ATF4 upregulation in both cycling and non-cycling cells, indicating that the GCN2-mediated ATF4 upregulation in response to WEE1 inhibition is independent of cell cycle status (Fig. 3c).
a RPE1 TP53KO PACKO cells were treated with AZD1775 (250 nM) for 6 h in the presence or absence of ISRIB (1 μM), pulse labeled with EdU, and processed for quantitative image-based cytometry. Cell cycle stage was defined by DNA content and EdU positivity. Each point represents a cell. Two independent replicates are defined by two different colors. Bars represent means of each replicate. Experimental set-up (b) and flow cytometry analysis (c) of RPE1 TP53KO ATF4-mScarlet-NLS reporter cells. Cells were treated overnight with nocodazole. Mitotic cells were isolated and replated in the presence or absence of palbociclib. After 2 h, AZD1775 (1 µM) and/or A92 (1 µM) was added for 20 h. Data represent mean ± SD (n = 3). Flow cytometry gating strategy (d) and analysis of γH2AX in RPE1 TP53KO cells after treatment with AZD1775 (1 µM) and/or A92 (1 µM) in the absence (e, left) or presence (e, right) of nocodazole. Data represent mean ± SD (n = 4). f Flow cytometry analysis of MPM2-positivity in RPE1 TP53KO cells after treatment with nocodazole, doxorubicin, AZD1775 (1 µM) and/or A92 (1 µM). Data represent mean ± SD (n = 3). g Flow cytometry analysis of MPM2-positivity in control RPE1 TP53KO PACKO and ATF4KO #1 or GCN2KO #1 cells after treatment with nocodazole, doxorubicin and AZD1775 (1 µM). Data represent mean ± SD (parental conditions n = 4, KO conditions n = 3). h Immunoblot of resting PBMCs or PBMCs stimulated with anti-CD3/CD28 beads. PBMCs were treated with DMSO, AZD1775 (500 nM) or thapsigargin (500 nM) for 24 h. Representative blot of n = 3 independent experiments. i RPE1 TP53KO PACKO and GCN2KO cells were treated with AZD1775 and/or cisplatin for 5 days. Cell survival was analyzed by MTT conversion. Data represent mean ± SD (n = 4). Statistical analysis for c, e, f, g was performed using unpaired t-tests (two-sided), with p ≤ 0.05 considered significant. All replicates are biological replicates unless indicated otherwise. Source data are provided as a Source data file.
WEE1i are well described to cause DNA damage and premature entry into mitosis12,48. Since loss of GCN2 reduced WEE1i-induced cytotoxicity, we analyzed whether GCN2 was required for WEE1i-induced DNA damage and premature mitotic entry. As expected, WEE1 inhibition significantly increased the percentage of γH2AX-positive cells, which is caused by unscheduled cleavage of replication forks49, a phenotype that was not rescued by GCN2i (Fig. 3d, e, left panel). WEE1 inhibition induced entry of γH2AX-positive cells in mitosis, which also occurred in GCN2-inhibited cells (Fig. 3d, e, right panel). Of note, GCN2i partially suppressed the entry of γH2AX-positive cells into mitosis, which may have been caused by slower proliferation of GCN2i-treated cells, a possibility supported by the observation that the total number of mitotic cells was lower upon pharmacological GCN2 inhibition (Supplementary Fig. 5c).
WEE1i can also force entry into mitosis in cells treated with DNA-damaging chemotherapeutic agents20,50. Indeed, WEE1 inhibition resulted in an override of the G2 checkpoint arrest induced by the DNA topoisomerase II inhibitor doxorubicin, which was not prevented by GCN2i (Fig. 3f and Supplementary Fig. 5d). To confirm these findings, we analyzed the proportion of mitotic cells upon AZD1775 treatment in the RPE1 TP53KO GCN2KO or ATF4KO cell lines. Neither inactivation of GCN2 nor ATF4 affected the ability of AZD1775 to override the doxorubicin-induced G2 cell cycle arrest (Fig. 3g and Supplementary Fig. 5e, f). Taken together, these results show that ISR induction and cell cycle deregulation are independent effects of WEE1i.
To investigate whether WEE1 inhibition also leads to ISR activation in non-transformed cells, peripheral blood mononuclear cells (PBMCs) were treated with AZD1775 or thapsigargin as a positive control. Non-stimulated PBMCs were compared with PBMCs that were activated using anti-CD3/CD28 T-cell activator beads (Fig. 3h). Ongoing mRNA translation levels were below detection level in unstimulated PBMCs, along with very low levels of WEE1 protein expression (Fig. 3h). In contrast, activated PBMCs showed an overall increase in protein abundance and ongoing mRNA translation (Fig. 3h and Supplementary Fig. 5g). WEE1i treatment in activated PBMCs robustly induced ATF4 expression, and suppressed active mRNA translation (Fig. 3h and Supplementary Fig. 5g). In line with the observed ATF4 induction, GCN2-T899 phosphorylation was increased upon WEE1i treatment (Fig. 3h), without altering total GCN2 levels (Supplementary Fig. 5h). These data show that ISR activation by WEE1i is not limited to immortalized or transformed cells and can occur in human primary cells.
AZD1775 has been evaluated in combination with platinum-based chemotherapeutic agents23,24,25,26. We analyzed whether GCN2 activation is required for the potentiating effects of AZD1775 on cisplatin sensitivity. Loss of GCN2 partially rescued the cytotoxicity induced by AZD1775 as a single agent, but not the additive cytotoxicity of combined WEE1i and cisplatin treatment (Fig. 3i) in RPE1 TP53KO cells. These findings are consistent with the observation that GCN2 inactivation did not prevent γH2AX induction and cell cycle checkpoint override following AZD1775 treatment (Fig. 3e–g). Similar findings were observed in triple-negative breast cancer cell lines HCC1806 (Supplementary Fig. 6a) and MDA-MB-231 (Supplementary Fig. 6b). Combined, these data show that activation of GCN2 by WEE1i is cell cycle-independent, and that GCN2 loss does not prevent WEE1i-induced DNA damage, premature mitotic entry and sensitization to genotoxic chemotherapeutics.
WEE1i alter the response to translation perturbations
Since WEE1i activate the ISR via GCN2, independently of cell cycle status or DNA damage induction, we tested if WEE1i directly affected ribosome dynamics. A CRISPR-based negative selection screen identified predicted synergistic WEE1i interactors, including PKMYT1, suggesting redundancy in the regulation of CDK inhibition, and confirming previous findings51,52 (Fig. 4a, b). Moreover, loss of the DNA polymerase epsilon subunit POLE3 and the ribonucleotide reductase catalytic subunit RRM1 sensitized cells to WEE1i (Fig. 4b). Loss of these genes was previously described to sensitize cells to ATR inhibition, emphasizing the shared roles of WEE1 and ATR in guarding faithful S-phase progression53. We also identified WEE1, pointing to drug-induced genetic insufficiency, possibly involving inactivation of one WEE1 allele, rendering cells more sensitive to WEE1i treatment54. Finally, we identified a subset of genes that function in ribosome quality control (RQC), including ZNF598, PELO, HBS1L, USP9X, RNF25, and components of the ASC-1 complex (ASCC2 and ASCC3)55,56,57,58,59,60 (Fig. 4b, c). These observations suggest that mechanisms that resolve ribosome collisions are important to determine WEE1i sensitivity, with GCN2 and GCN1 loss appearing as drivers of WEE1i resistance in this screen (Fig. 4b). Sensitization to WEE1i upon inactivation of ZNF598 was confirmed in Incucyte cell proliferation assays (Supplementary Fig. 7a).
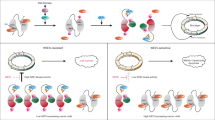
a Schematic overview of the CRISPR screen in (b). b NormZ scores of individual genes in an RPE1 TP53KO PACKO Cas9 negative selection AZD1775 CRISPR screen (day 15 vs. day 0). Positive normZ scores indicate suppression, and negative normZ scores indicate synergy. Genes are color-coded by category; Orange: cell cycle checkpoint kinases, Green: ribosome quality control, Blue: ISR kinases GCN1L1 (GCN1) and EIF2AK4 (GCN2). c Schematic representation of ribosome quality control pathway members with high-ranking screen hits sensitizing to WEE1i. d Schematic representation of single ribosome mRNA translation kinetics measured by socRNA-mediated GFP translation. e Representative image of socRNA-mediated GFP translation as analyzed in (f). f GFP foci intensity over time (n = 1, tracking of 67 (no drug) or 73 (AZD1775) socRNAs). Lines and shaded areas indicate mean ± SD. g Elongation speed of a single ribosome on a socRNA in control or AZD1775-treated cells. Data represent mean ± SD (no drug: n = 58, AZD1775: n = 54, from 2 experiments), unpaired t-test (two-sided) with p ≤ 0.05 considered significant. h Schematic representation of a socRNA containing an Xbp1 pause sequence. Puromycin addition triggers the release of ribosomes and nascent peptide chains from a socRNA, allowing quantification of ribosome copies per socRNA. i Visual representation of ribosome release upon puromycin addition to socRNA, as quantified in (j). j Fraction of analyzed socRNAs containing indicated numbers of translating ribosomes per socRNA after AZD1775 treatment (300 nM, 24 h) with or without GCN2i A92 (1 µM). Mean ± range (No drug n = 3; AZD1775 n = 4, AZD1775 + A92 n = 2). k Immunoblot of RPE1 TP53KO cells after 30 min treatment with emetine (0.5 µg/ml) and puromycin pulse (n = 1). l Immunoblot of RPE1 TP53KO cells co-treated with AZD1775 (250 nM) and indicated doses of emetine according to the indicated timeline. Representative experiment of n = 2. m Immunoblot of RPE1 TP53KO cells treated with AZD1775 (250 nM) for 4.5 h, and during the last 30 min co-treatment with emetine (0.5 µg/ml). Representative experiment of n = 2. All replicates are biological replicates unless indicated otherwise. Source data are provided as a Source data file.
These observations led us to investigate whether WEE1i might influence ribosome dynamics. To investigate translation dynamics in more detail, we used the Stopless-ORF circular (soc)RNA assay, a recently-developed single-molecule imaging approach to visualize translation dynamics with high precision61. socRNAs encode SunTag peptide epitopes, which are co-translationally labeled by stably expressed STAb-GFP, resulting in local accumulation of GFP at socRNAs upon translation, allowing real-time measurements of translation dynamics (Fig. 4d, e). No difference in ribosome elongation speed was observed between WEE1i-treated cells and untreated cells (Fig. 4f, g). In addition, the number of translating ribosomes per socRNA was not different in cells treated with WEE1i (Supplementary Fig. 7b), indicating that ribosome processivity was also unaffected61. To specifically examine ribosome quality control in WEE1i-treated cells, we next analyzed a socRNA harboring the well-established Xbp1(S255A) ribosome pausing sequence (Fig. 4h). Ribosomes will stall at this sequence, causing ribosome collisions, which will eventually be recycled in a ZNF598-dependent manner59,62. In this assay, collision-induced ribosome recycling efficiency can be assessed by measuring the number of ribosomes per Xbp1(S255A) socRNA, which can be determined by inducing ribosome release from the socRNA through puromycin treatment and counting the number of GFP foci (each representing a single ribosome) that split from a single socRNA61 (Fig. 4h, i). When compared to control-treated cells, WEE1i-treated cells contained more ribosomes per Xbp1(S255A) socRNA, suggesting that collision-induced ribosome recycling was suppressed in WEE1i-treated cells, with an effect size similar to that reported for ZNF598 depletion (Fig. 4j)61. These effects on ribosome recycling were not due to a change in pause duration upon WEE1i treatment (Supplementary Fig. 7c). Polysome profiling revealed elevated ribosome occupancy on mRNAs upon WEE1i treatment, supporting the socRNA data showing higher ribosome occupancy upon AZD1775 treatment (Supplementary Fig. 7d). Interestingly, combined treatment with AZD1775 and the GCN2i A92 completely rescued the increase in ribosome occupancy, demonstrating that the ribosome recycling defect in WEE1i-treated cells was dependent on GCN2. Ribosome occupancy in cells treated with both inhibitors was even slightly below the occupancy in untreated cells (Fig. 4j). These effects are reminiscent of the decrease in ATF4 levels below baseline upon combined treatment with AZD1775 and A92 (Fig. 2j, k), suggesting baseline GCN2 activity in our cell line models.
Defective ribosome recycling after WEE1 inhibition may cause long-lived ribosome collisions in these cells, which results in increased GCN2 activation, explaining the genetic interaction of WEE1 inhibition with proteins involved in ribosome quality control.
The observed defective ribosome recycling suggested that GCN2 responds to translation perturbations caused by the WEE1i. To test this concept more directly, we asked whether ongoing mRNA translation was necessary for the activation of GCN2 by WEE1i. To do so, we treated RPE1 TP53KO cells with the translation inhibitor emetine63 at a dose that caused ribosome stalling (Fig. 4k). Notably, we observed that co-treatment of cells with emetine and WEE1i completely blocked GCN2 activation (Fig. 4l). We also observed that emetine addition to cells with WEE1i-induced ISR activation resulted in suppressed GCN2 activation within as little as 30 min (Fig. 4m), indicating that ongoing translation is required for sustained GCN2 activation by WEE1i treatment.
To further dissect how WEE1i alter ribosome processivity, mRNA-seq and Ribo-seq of AZD1775-treated RPE1 TP53KO cells were performed. Differential expression analysis of bulk mRNA-seq data revealed a robust ISR and hallmarks of GCN2 activation, characterized by increased expression of amino acid biosynthesis genes (e.g. ASNS, CBS, and PSAT1), along with aminoacyl-tRNA synthetases (Supplementary Fig. 8a). Moreover, gene set enrichment analysis identified robust upregulation of pathways related to mRNA translation at late time-points of AZD1775 treatment, specifically involving eukaryotic translation initiation and elongation, peptide chain elongation, cap-dependent translation initiation, and seleno-amino acid metabolism in response to 48 h AZD1775 treatment (Supplementary Fig. 8b).
Ribosome profiling (Ribo-seq) analysis showed that high-dose WEE1i treatment for 24 h resulted in a global decrease in translation, independently of mRNA abundance (Supplementary Fig. 9a, b). Pathway enrichment analysis showed that high-dose WEE1i led to significant downregulation of translation-related pathways, including “eukaryotic translation elongation” and “seleno-amino acid metabolism” (Supplementary Fig. 9c). Upregulated transcription of a panel of ATF4 target genes (i.e., DDIT3, DDIT4) (Supplementary Fig. 9d), as well as amino acid biosynthesis factors such as ASNS and PSAT1 (Supplementary Fig. 8a, b), confirmed ISR activation. Although not statistically significant, likely due to limited sample size, the levels of the ATF4 target genes ATF3 and PPP1R15A also appeared to increase (Supplementary Fig. 9d). Ribosome occupancy was only mildly shifted after 24 h of WEE1i treatment (Supplementary Fig. 9e), possibly due to a shift in expressed genes due to the induction of an alternative translation program by ISR activation, and overall lower coding DNA sequence (CDS) reads (Supplementary Fig. 9e–g). However, overall, translation extension kinetics were similar between conditions (Supplementary Fig. 9e), in line with our observations of socRNA ribosome kinetics (Fig. 4f). Taken together, these data suggest that WEE1i alters the response to translation perturbations and affects ribosome processivity, triggering cellular adaptation.
The WEE1i AZD1775 activates GCN2 as an off-target activity
To analyze whether inhibitors of other cell cycle checkpoint kinases also activate the ISR, we measured ATF4-mScarlet upregulation upon treatment of RPE1 TP53KO ATF4 mScarlet reporter cells with inhibitors of PKMYT1 (RP-6306), ATR (VE-822), and CHK1 (AZD7762). From this panel, only AZD1775 treatment resulted in clear ATF4-mScarlet upregulation (Fig. 5a).
a Relative ATF4-mScarlet MFI in RPE1 ATF4-mScarlet reporter cells after treatment with 125, 250, 500, or 1000 nM of AZD1775, RP-6306, VE-822 or AZD7762. Data represent mean ± SD (DMSO: n = 4; other conditions: n = 3). Statistical analysis was performed using a one-way ANOVA with Dunnett’s multiple comparisons with p ≤ 0.05 considered significant. b Immunoblot of RPE1 TP53KO cells treated with siRNA targeting WEE1 for 72 h and AZD1775 (250 nM) for 2.5 or 5 h. Representative blot of n = 3 experiments. WEE1 and GCN2 engagement as measured by NanoBRET kinase target engagement assay. HEK293 cells were transfected with WEE1-NanoLuc (c) and NanoLuc-GCN2 (d) and incubated with K-10 tracer (0.5 μM), and indicated doses of CC1, GCN2iB, AZD1775, Debio 0123, ZNL-02-096, RP-6306 or neratinib for 2 h prior to substrate addition and BRET signal detection. Data represent mean ± SD (n = 3). Representative histograms (e) and quantification (f) of ATF4-mScarlet flow cytometry measurements in RPE1 TP53KO ATF4-mScarlet reporter cells treated with siRNAs targeting GCN1 and AZD1775 or neratinib (1 µM) for 5 h. Data represent mean ± SD (n = 3), statistical analysis: two-way ANOVA with Tukey’s multiple comparisons test, with p ≤ 0.05 considered significant. g Model of WEE1 inhibitor mode of action and consequences. All replicates are biological replicates unless indicated otherwise. Source data are provided as a Source data file.
Small molecule inhibitors of eIF2α kinases, PERK and PKR, as well as tyrosine kinase inhibitors including neratinib and dovitinib have previously been shown to paradoxically activate GCN264,65. The proposed model for GCN2 activation by these drugs involves binding to the ATP binding pocket of one of the two GCN2 monomers within a GCN2 dimer, thereby enhancing the affinity of the other binding pocket for ATP64,65. To test whether the presence of WEE1 is required for ISR activation, RPE1 TP53KO cells were depleted of WEE1, and subsequently treated with AZD1775. WEE1 depletion did not prevent ATF4 upregulation or GCN2 phosphorylation, indicating that ISR activation by AZD1775 does not require WEE1 presence (Fig. 5b and Supplementary Fig. 10a, b). To determine whether WEE1 inhibitors could similarly activate GCN2 kinase as previously described kinase inhibitors, a NanoBRET assay66 was used to address the previously reported observation that AZD1775 could bind the recombinant kinase domain67. Specifically, we assessed the ability of the WEE1i AZD1775, Debio 0123 and ZNL 02-096, a WEE1 PROTAC consisting of AZD1775 linked to an E3 ubiquitin ligase, as well as the GCN2 inhibitor GCN2iB, the PKMYT1 inhibitor RP-6306 and the tyrosine kinase inhibitor neratinib to displace a fluorescent tracer compound at the active site of luciferase-tagged WEE1 or GCN2 kinase domain in a cellular environment. In these assays, all three WEE1 inhibiting compounds potently displaced the fluorescent tracer from the WEE1 kinase domain at biologically relevant doses and resulted in its inhibition, while PKMYT1i only interacted at high doses (Fig. 5c). Neither GCN2iB nor neratinib were able to engage with the WEE1 kinase domain (Fig. 5c). In accordance with these findings, AZD1775 and Debio 0123, but not GCN2iB or neratinib, engaged with full-length recombinant WEE1 in kinase profiling assays (Supplementary Fig. 10c). As expected, both the non-specific pan-kinase inhibitor (CC1) and GCN2iB engaged with the GCN2 kinase domain in NanoBRET analysis (Fig. 5d). AZD1775, Debio 0123 and ZNL 02-096 were found to engage with the GCN2 kinase domain, albeit at relatively high doses (Fig. 5d). However, these doses were similar to the dose at which neratinib engaged with the GCN2 kinase in this assay (Fig. 5d). RP-6306 did not show any engagement with the GCN2 kinase domain (Fig. 5d). Kinase profiling showed efficient engagement of both AZD1775 and neratinib with full-length recombinant GCN2, with IC50 values of 140 and 507 nM respectively (Supplementary Fig. 10d). Taken together, these data suggest that ISR activation by chemical WEE1i AZD1775, Debio 0123 or ZNL 02-096 is likely caused by direct engagement with the GCN2 kinase.
Similar to the WEE1i, treatment of RPE1 TP53KO cells with the WEE1 PROTAC ZNL 02-096 resulted in upregulation of ATF4 expression and GCN2 phosphorylation (Supplementary Fig. 10e). Intriguingly, whereas AZD1775 treatment resulted in sustained ISR activity, we observed a decrease in ATF4 upregulation at 12 h after ZNL 02-096 treatment. Previous studies have shown that GCN2 requires GCN1 for its activation68,69. To study whether GCN1 is also required for WEE1i-mediated GCN2 activation, GCN1 was depleted in RPE1 TP53KO ATF4-mScarlet-NLS cells, prior to treatment with AZD1775. Analysis of ATF4 and pGCN2 levels showed that both AZD1775 and neratinib treatment led to increased ATF4-mScarlet levels in control-depleted cells but were significantly reduced in GCN1-depleted cells (Fig. 5e, f and Supplementary Fig. 10f). Interestingly, residual phosphorylation of GCN2 was observed upon treatment of GCN1-depleted cells (Supplementary Fig. 10g), suggesting that WEE1i can prime GCN2, resulting in autophosphorylation, but that GCN1 is required for downstream ISR signaling. Of note, at untreated conditions, ATF4 levels were slightly decreased in GCN1-depleted cells (Fig. 5e, f), indicating that these cells have baseline ISR signaling.
Combined, these data show that ISR activation by chemically distinct WEE1i is independent of WEE1 and involves an off-target interaction with GCN2. This leads to paradoxical GCN2 activation that requires GCN1 and ongoing translation for potent ISR activation. Our data suggest a model in which WEE1i causes WEE1-dependent loss of cell cycle checkpoint control, leading to replication stress, DNA damage accumulation, and premature mitotic entry, leading to cell death. Simultaneously, WEE1i also triggers activation of GCN2 in cells with ongoing mRNA translation, leading to ISR activation and translation attenuation (Fig. 5g).
WEE1i and PKMYT1i synergistically kill TP53 mutant cells without WEE1i-mediated induction of the ISR
In line with WEE1 and PKMYT1, both regulating CDK1 activity, their combined inhibition showed synergy even at individual sub-cytotoxic doses51,70. Combined WEE1 and PKMYT1 inhibition in solid tumors is currently being evaluated in a phase 1/1b clinical trial (NCT04855656). To investigate whether WEE1i combined with RP6306, a PKMYT1i71,72, can synergistically induce cytotoxicity without WEE1i-mediated ISR activation, ATF4 protein expression was measured following treatment with a dose range of Debio 0123 or AZD1775 in RPE1 TP53KO cells (Fig. 6a and Supplementary Fig. 11a, b) and in triple-negative breast cancer cell lines HCC1806 and MDA-MB-231 (Supplementary Fig. 11c, d). Debio 0123 treatment only resulted in ATF4 upregulation from 250 nM onwards in HCC1806 cells (Supplementary Fig. 11c upper panel), and from 400 to 500 nM onwards in MDA-MB-231 and RPE1 TP53KO cells respectively (Fig. 6a and Supplementary Fig. 11c lower panel), whereas AZD1775-mediated ATF4 expression was observed already at 5 nM in HCC1806 (Supplementary Fig. 11d upper panel), and at 20 nM onwards in RPE1 TP53KO and MDA-MB-231 cells (Supplementary Fig. 11b, d lower panel).
a Relative ATF4-mScarlet MFI in RPE1 TP53KO ATF4-mScarlet reporter cells after treatment with the indicated doses of Debio 0123. Data represent mean ± SD (n = 3). b Cell viability in parental RPE1 TP53KO PACKO, GCN2KO #1 and GCN2KO #2 cells after treatment with Debio 0123 and RP-6306 at the indicated doses (n = 5). c ZIP synergy scores from cell survival matrices shown in (b, d). Data represent mean ± SD (n = 5). d Synergy plots with ZIP synergy scores of heatmaps shown in (b). e Relative ATF4-mScarlet MFI in RPE1 TP53KO ATF4-mScarlet reporter cells treated with the indicated doses of AZD1775, Debio 0123 and RP-6306. Data represent mean ± SD (n = 4). Statistical analysis (a, c, e) was performed using paired t-tests (two-sided), with p ≤ 0.05 considered significant. All replicates are biological replicates unless indicated otherwise. Source data are provided as a Source data file.
We observed synergy between WEE1i and PKMYT1i, with a larger reduction in viability in the combination treatment compared to the highest doses of each single treatment in all cell lines (Fig. 6b and Supplementary Fig. 11e–m). Specifically, RPE1 TP53KO cells treated with Debio 0123 and PKMYT1i showed a mean synergy (ZIP) score of 33.56, ranging from 17.4 to 40.9 between experiments (Fig. 6c). GCN2KO cells required slightly higher doses of the combination treatment to reach a similar viability reduction compared to parental cells (Fig. 6b, d), although the synergy between Debio 0123 and PKMYT1i treatment was maintained in GCN2KO cells, with mean ZIP scores of 27.5 and 40.08 for GCN2KO #1 and #2, respectively (Fig. 6b–d). Importantly, synergy between Debio 0123 (250 nM) and RP-6306 (60 nM) was observed at doses that did not induce ATF4 expression, either in single agent treatment or in combination treatments (Fig. 6a–e). Similar results were obtained with AZD1775 treatment in RPE1 TP53KO cells, with mean synergy scores of 22.4, ranging from approximately 9 to 33 (Supplementary Fig. 11c–e). Again, GCN2KO cells required higher doses of both inhibitors to achieve the same level of toxicity as parental RPE1 TP53KO cells, although GCN2KO #1 and #2 cell lines still exhibited synergy with mean synergy scores of 14.3 and 18.4, respectively (Supplementary Figs. 11e and 12a, b).
Synergy analysis in HCC1806 and MDA-MB-231 cells resulted in similar findings when compared to RPE1 TP53KO cells (Supplementary Fig. 12c–n). Synergy upon combined treatment with Debio 0123 and RP-6306 was retained upon GCN2 inhibition in both HCC1806 (mean synergy score of 44.44 without GCN2i and 32.75 with GCN2i) and MDA-MB-231 (mean synergy score of 36.73 without GCN2i and 34.58 with GCN2i) (Supplementary Fig. 12c–h). Importantly, synergistic reduction in viability was observed at doses of Debio 0123 below those needed to induce ATF4 expression in MDA-MB-231 cells. For AZD1775 and RP-6306, we observed a consistent decrease in synergy score upon GCN2i treatment in HCC1806 and MDA-MB-231 cells, although synergy was retained as indicated by the mean synergy score over 10 (Supplementary Fig. 12i–n), and in line with AZD1775 inducing ATF4 activation at virtually all tested doses. Taken together, these findings show that combined treatment of low-dose Debio 0123 WEE1i with PKMYT1i synergistically induces cytotoxicity, independently of ISR activation.
Discussion
In this study, we report that the sensitivity of cells to clinically used WEE1i is determined by the integrated stress response (ISR) and ribosome quality control (RQC) pathways, in addition to its known effects on cell cycle perturbation. The WEE1i AZD1775 and Debio 0123 trigger activation of GCN2 in transformed and untransformed cells, in a cell cycle-independent manner. Specifically, loss of ISR-kinase GCN2, its binding partner GCN1, or its downstream effector ATF4 decrease WEE1i cytotoxicity. Yet, cells in which the ISR is inactivated remain susceptible to WEE1i-mediated cell cycle checkpoint override and synergistic effects of WEE1i with genotoxic chemotherapeutics and with PKMYT1 inhibition. Clearly, current WEE1i trigger two mechanistically distinct cytotoxic responses.
Remarkably, we found that activation of the ISR by the WEE1i used in this study was most likely independent of WEE1. Although siRNA-mediated depletion may leave residual target protein, WEE1 depletion did not prevent ISR activation, pointing at off-target effects of WEE1i. A potential explanation for this observation is that the AZD1775 and Debio 0123 molecules directly bind and paradoxically activate GCN2, analogous to other tyrosine kinase inhibitors64,65,73 that are known to bind the ATP-binding pocket of one of the monomers of the GCN2 dimer. In line with this hypothesis, previously reported kinome profiling studies employing phage display reported binding of AZD1775 to recombinant GCN2 kinase domain67,74. A more recent study also suggested this mode of action for AZD1775, when tested at high concentrations75. In this study, we found evidence for direct engagement of WEE1 with GCN2 by WEE1i, using both cellular NanoBRET assays and in vitro GCN2 kinase profiling. It should be mentioned that the in vitro kinase profiling resulted in kinase engagement at lower doses than observed with the NanoBRET assay, which may be due to lower sensitivity of this assay. Importantly, we demonstrate that paradoxical activation of GCN2 by WEE1i not only requires the presence of GCN2. Rather, we show that GCN1 presence, ongoing translation, and translational stress are required for paradoxical GCN2 activation (Fig. 5g). Specifically, GCN1 inactivation conferred resistance to AZD1775 treatment, and an emetine-mediated block in translation prevented AZD1775-mediated ISR activation, while socRNAs without pausing sequence did not lead to altered ribosome kinetics in AZD1775-treated cells. Thus, our findings support a model in which paradoxical activation of GCN2 by WEE1i requires a conformation change induced by GCN1. Of note, the observation that GCN1 depletion also prevents ISR activation by the tyrosine kinase inhibitor neratinib suggests that these requirements also apply to other paradoxical activators of GCN2.
The WEE1i-induced ISR activation resulted in an attenuation of global mRNA translation and upregulation of stress response factors. In the absence of eIF2A, GCN2 can still be activated, leading to eIF2α phosphorylation, which subsequently results in reduced cap-dependent translation and presumably increased translation of stress-related ISR target proteins. In a context where eIF2A is not available, translation of these stress-related ISR target proteins is likely reduced, possibly due to a requirement of eIF2A for their translation32, likely leading to critically low translation levels, which could explain the increased sensitivity of EIF2AKO cells to WEE1i. More recently, a role in human cells for eIF2A in alternative initiation of translation has been challenged35,36, with a demonstrated role for eIF2A in RQC36. This study showed that eIF2A is involved in regulating 40S recycling or degradation, and suggested that loss of eIF2A may cause premature or faulty translation reinitiation36. This recently described role for eIF2A aligns well with our identification of other ribosome rescue pathway factors, of which inactivation resulted in increased sensitivity to WEE1i treatment. These genetic contexts of impaired RQC lead to increased ribosomal stress and trigger ISR activation. Specifically, previous studies have shown that loss of ZNF598 results in the readthrough of ribosomal stalling sites59,76. In yeast, loss of the ZNF598-ortholog Hel2 was demonstrated to increase GCN2 activation77. While in conditions of infrequent ribosome collisions, ZNF598/Hel2 is activated, in conditions of excessive ribosome collisions, GCN2 will be activated as well77. Possibly, GCN2 activation in response to loss of RQC factors may become cytotoxic when GCN2 is additionally activated by WEE1i. In the setting where WEE1i activates GCN2, and RQC factor expression is lost, GCN2 might be occupied and therefore unable to respond to baseline collisions, which are normally resolved by RQC factors. Therefore, WEE1i may lead to increased levels of stalled or collided ribosomes, which are not resolved or recycled, subsequently resulting in an increased number of ribosomes per RNA containing a stall site. This model aligns with our socRNA data. In accordance with this, we also observed an increase in 80S ribosomal subunits after AZD1775 treatment, which could be due to reduced 80S splitting by ABCE178. Of note, we observed that pharmacological inhibition of GCN2 rescued ribosome recycling, suggesting that ISR activation is not only a downstream effector of ribosome stress but also signals to the RQC complex to regulate ribosome dynamics.
In previous studies, WEE1i was studied as a single agent or in combination regimens with genotoxic agents to induce uncontrolled cell cycle progression in the presence of DNA lesions to trigger mitotic catastrophe30. Clinical evaluation of WEE1i demonstrated favorable responses along with considerable bone marrow toxicity23,25,26,79. Although ISR activation is a mechanism to restore homeostasis upon stress, chronic stimulation of ISR kinases can induce apoptosis80. Therefore, ISR activation may contribute to these adverse side effects. Indeed, we observed that ISR activation by WEE1i also occurred in non-transformed blood cells. However, since unstimulated blood cells have low levels of mRNA translation, we anticipate that ISR activation will only manifest itself in subpopulations that are metabolically active.
The notion that WEE1i in tumor cells exerts multiple mechanistically distinct effects—loss of cell cycle control and ISR activation—may challenge biomarker identification for WEE1i-eligible patients. Indeed, biomarker studies have predominantly focused on tumor characteristics related to defective cell cycle control (i.e., TP53 inactivation) or replication stress (i.e., SETD2 loss, CCNE1 amplification). Of note, the WEE1i AZD1775 was previously reported to also have off-target activity against PLK167. Although our screening results did not identify genes involved in PLK1-related mechanisms, it is clear that next-generation WEE1i should be developed with increased specificity, including an inability to activate the ISR, to ultimately improve patient benefit.
Alternatively, combination strategies based on combining low doses of currently used WEE1i with other agents could also be considered. Indeed, our data show that the previously described synergistic combination of the WEE1i Debio 0123 with PKMYT1i70 is effective at doses that do not trigger ISR activation. Combination of Debio 0123 with RP-6306 at low doses may therefore represent a therapeutic strategy to limit off-target cytotoxicity through ISR activation, while maintaining efficacy against cancer cells. Similarly, the anticipated development of next-generation dual PKMYT1/WEE1 inhibitors may provide therapeutic advantages of CDK1/2 modulation without the potential liability of ISR induction. However, it is also possible that ISR induction may be therapeutically desirable under some circumstances. Indeed, based on our genetic screening results, tumors with defective RQC pathways or loss of eIF2A are expected to be increasingly sensitive to current WEE1i. Overall, our study describes how on-target and off-target effects of WEE1 inhibitors influence their therapeutic potential.
Methods
Cell line models
RPE1 hTERT TP53KO (RPE1 TP53KO) and derived cell lines, as well as MDA-MB-231 cells, were cultured in Dulbecco′s Modified Eagle′s Medium (Gibco) with low glucose (1 g/L; DMEM-L), supplemented with 10% fetal calf serum (FCS, Lonza) and 50 units/mL penicillin and 50 μg/mL streptomycin (Gibco, 15140122). RPE1 hTERT TP53KO (RPE1 TP53KO) PACKO cells, RPE1-FLAG-Cas, HEK293T cells, and derivative lines were maintained in DMEM with 4.5 g/L glucose (DMEM-H), 1 mM sodium pyruvate, supplemented with 10% fetal calf serum (FCS). Media for RPE1 TP53KO ZNF598KO, ATF4KO, and GCN2KO cells were supplemented with MEM non-essential amino acids, or DMEM with 1 g/L glucose supplemented with 10% FCS and 50 units/mL penicillin and 50 μg/mL streptomycin. For CRISPR/Cas9 screens, RPE1 cells were grown in DMEM-H, 1 mM sodium pyruvate, 1x penicillin-streptomycin (Wisent), and 10% FCS. HAP1 cells were cultured in Iscove’s Modified Dulbecco’s Medium (IMDM, Gibco), supplemented with 2 mM L-glutamine (Gibco), 50 units/mL penicillin, 50 μg/mL streptomycin (Gibco, 15140122), and 10% FCS. OVCAR3 cells were cultured in Roswell Park Memorial Institute (RPMI)-1640 media supplemented with 50 units/mL penicillin, 50 μg/mL streptomycin, and 20% FCS. Peripheral Blood Mononuclear Cells (PBMCs) were cultured in RPMI-1640 medium supplemented with 10% FCS and 2 mM L-glutamine. HCC38 and HCC1806 cells were maintained in RPMI-1640 medium supplemented with 50 units/mL penicillin, 50 μg/mL streptomycin, and 10% FCS. U2OS cells stably expressing tetR, ALFA-Nb-CAAX, and STAb-GFP were cultured in DMEM-H with 50 units/mL penicillin, 50 μg/mL streptomycin, and 5% FCS. All cells were cultured at 37 °C and 5% CO2.
Inhibitors and fine chemicals
The chemicals used in this study were AZD1775 (#1494, Axon Medchem or S1525, Selleck Chemicals), Debio 0123 (S9778, Selleckchem), A92 (#2720, Axon Medchem), RP-6306 (gift from Repare Therapeutics), AZD7762 (#1399, Axon Medchem), VE-822 (#2452, Axon Medchem), ZNL 02-096 (7240, Tocris), thapsigargin (T9033, Merck), doxorubicin (sc-280681, Santa Cruz Biotechnology), nocodazole (M1404, Sigma), palbociclib (1505, Axon Medchem), integrated stress response inhibitor (ISRIB; SML0843, Sigma or S7400, Selleckchem), salubrinal (S2923, Selleckchem), GCN2iB (S8929, Selleckchem), neratinib (#1526, Axon Medchem) and Emetine (E521535, Toronto Research Chemicals).
Haploid genetic screen
HAP1 cells were mutagenized as described previously30,81,82. In summary, ∼6 × 107 cells were transduced with pGT gene-trap retrovirus. Mutagenized cells were treated for 9 days with AZD1775 (125 nM) prior to fixation in ‘Fixation Buffer 1’ (Becton Dickinson). Genomic DNA was isolated from ∼4 × 107 cells. DNA was subjected to a linear amplification (LAM)-PCR protocol, and sequenced using the Genome Analyzer platform (Illumina). Gene trap insertion mapping and data analysis were performed as described previously83. In brief, the dataset was normalized to combined insertion counts of four HAP1 wildtype datasets81. A binomial test was performed for the distribution of sense and antisense orientation insertions in one AZD1775-treated HAP1 dataset. Loss of a gene was considered sensitizing to AZD1775 when the sense-over-antisense insertion ratio was decreased in the AZD1775-treated HAP1 dataset compared to the four HAP1 wildtype datasets81,84 (SRA accession no. SRP058962). To this end, two-sided Fisher’s exact tests were performed for each gene against all wildtype datasets, and the gene was considered a sensitizer when it passed all Fisher’s tests (p < 0.05; effect size 20%). Sequencing data have been deposited at the BioProject Archive of the NCBI under accession PRJNA1152540.
For pathway analysis, sense insertion ratios and insertion frequency of WEE1i-treated cells were compared with 4 DMSO-treated control datasets. P values for each gene were log-10 transformed, and genes were ranked according to average P value. The top 250 genes in which WEE1i-treated cells had more antisense insertions were analyzed by DAVID84, using the KEGG pathway gene sets. Only gene sets with a false-discovery rate <0.05 are indicated.
Genome-wide CRISPR screens with Toronto KnockOut sgRNA libraries
For CRISPR/Cas9 screens, RPE1 TP53KO Flag-Cas9 cells were transduced at low multiplicity of infection (MOI 0.3-0.5) with Toronto KnockOut (TKO) v1 90k sgRNA or v2 71k sgRNA libraries for the sensitizer and resistance screen, respectively85,86,87. The next day, infected cells were selected with 20 µg/ml puromycin with trypsinization 24 h after selection and then expanded in the presence of puromycin and blasticidin. Cells were then split into technical duplicates maintaining the appropriate library coverage on day T0. Cells were split once more (day T3) before addition of AZD1775 on day T6. For the negative selection screen, cells were split every three days in the presence of 80 nM AZD1775. Screens were stopped at an estimated 10 population doublings (T21 for AZD1775 comparisons shown in this study). For the positive selection screen with AZD1775, RPE1 TP53KO Flag-Cas9 cells were split once dishes were 90% confluent (6-9 days after first addition of 200 nM AZD1775) and then collected a second time (T18-T21). Genomic DNA was extracted from samples taken at each time point (T0, T6, T18-21) using the QIAamp DNA Blood kit (Qiagen). Genomic DNA was normalized by measurement on a Nanodrop spectrophotometer. TKO sgRNA cassettes were isolated by PCR using KAPA HiFi HotStart ReadyMix (Roche). Illumina i5 and i7 sequencing multiplex barcodes were added in a second round of PCR and libraries were purified by agarose gel electrophoresis. Sequencing libraries were processed on an Illumina NextSeq500 platform at the Lunenfeld-Tanenbaum Research Institute NBCC facility (https://nbcc.lunenfeld.ca). sgRNA representation was determined by aligning sequencing reads to reference libraries and then using read counts as input for MAGeCK or DrugZ analysis to compute gene-level sgRNA enrichment and depletion, respectively. DrugZ values are provided in Supplementary Data 2. Negative log10-transformed positive MAGeCK scores for individual genes were plotted. Raw MAGeCK scores are provided in Supplementary Data 3.
CRISPR/Cas9-based gene editing
A sgRNA targeting exon 4 of EIF2A (Supplementary Data 4) was cloned into the PX458 vector, which was a gift from Fengh Zhang (Addgene #48138). Plasmids were introduced in RPE1 TP53KO and OVCAR3 cell lines using Fugene HD and were flow-sorted based on GFP expression.
Genomic DNA was isolated using QuickExtract DNA solution (Lucigen), followed by PCR using Taq polymerase (New England Biolabs) according to manufacturer’s instructions. Samples were sequenced by Eurofins Genomics using a Mix2Seq kit (Eurofins). TIDE was used to identify the mutational pattern after CRISPR-Cas9 mediated editing of EIF2A88. RPE1 TP53KO EIF2AKO cells had 93.8% out-of-frame mutations. OVCAR3 EIF2AKO #1 cells had 36% out-of-frame mutations, and OVCAR3 EIF2AKO #1 cells had 32.3% out-of-frame mutations and 13.4% in-frame indels.
RPE1 TP53KO PACKO ATF4KO and GCN2KO CRISPR/Cas9 edited cells were generated by transfection of parental cells with Cas9 ribonucleoprotein complexes (RNP) using Lipofectamine CRISPRMAX reagent (Life Technologies), and single clones were selected. sgRNAs were generated by in vitro transcription using the TranscriptAid T7 High Yield kit (ThermoFisher) (Supplementary Data 4). RPE1 TP53KO PACKO ZNF598KO cells were generated by transfecting parental cells with plentiCRISPRv2 derived sgRNA expression vector plentiGuide (Supplementary Data) and selecting for single clones. To assess sgRNA editing efficiency in polyclonal populations, transfected cells were passaged at least once before collection for genomic DNA extraction, PCR, and Sanger sequencing. Cas9 editing was assessed using the tracking of indels by decomposition (TIDE, Netherlands Cancer Institute) or inference of CRISPR edits (ICE) algorithm (Synthego). CRISPR editing in clonal cell lines was confirmed by Sanger sequencing and immunoblotting.
DNA cloning and retroviral transduction
The ATF4-mScarlet-NLS insert of pLHCX-ATF4 mScarlet NLS, which was a gift from David Andrews (Addgene, #115970), was obtained by PCR (Supplementary Data 4). Restriction digest was applied on pMSCV-blast (Addgene, #75085) using SalI (ThermoFisher, FD0644) and BamHI (ThermoFisher, FD0054) according to the manufacturer’s instructions. The ATF4-mScarlet NLS insert was cloned into the digested pMSCV-blast using NEBuilder HIFI DNA assembly Master Mix (New England Biolabs) according to the manufacturer’s instructions. Selected colonies were sequenced by Eurofins Genomics using a Mix2Seq kit (Eurofins). TIDE analysis was used to check the integration of the insert88.
The pMSCV-ATF4-mScarlet-NLS construct was transfected into HEK293T cells along with pRetro-VSV-G and pRetro-gag/pol, and subsequently, RPE1 TP53KO cells were transduced with retrovirus. Successfully transduced cells were selected with 7.5 µg/ml blasticidin S HCl (ThermoFisher, R21001) and sorted monoclonally using the Sony SH800S cell sorter.
siRNA transfection
For knockdown of WEE1 or GCN1, RPE1 TP53KO or RPE1 TP53KO ATF4-mScarlet-NLS cells, respectively, were transfected with siRNA targeting WEE1 (ThermoFisher, HSS111337) or GCN1 (ThermoFisher, 4392420 s21624 and s21626) using oligofectamine (Invitrogen, 12252-011) according to the manufacturer’s instructions. After 72 h of transfection, cells were treated with AZD1775 or neratinib for the indicated time points. Cells were harvested and further analyzed by Western blot or flow cytometry.
Viability assays
MTT assays: To assess the sensitivity of EIF2AKO cells to AZD1775, RPE1 TP53KO cells were seeded at a density of 200 cells/well of a 96-well plate, while OVCAR3 cells were seeded at 500 cells/well. For AZD1775 and cisplatin synergy assays, 100 cells/well were seeded for RPE1 TP53KO PACKO control cells, and GCN2KO cells were seeded at a 200 cells/well density. For AZD1775 and cisplatin synergy assays, treated with A92, HCC1806, and MDA-MB-231 cells were seeded at 600 cells/well. For PKMYT1i and WEE1i synergy assays, RPE1 TP53KO PACKO control and GCN2KO cells were plated at a density of 100–400 cells/well. For PKMYT1i and WEE1i synergy assays treated with A92, HCC1806 cells were seeded at 600–800 cells/well and MDA-MB-231 cells were seeded at 600 cells/well. 24 h after seeding, drugs were added at the indicated doses for 5 days. 5 mg/mL of methyl-thiazol tetrazolium (MTT; Sigma, M2128) was added for 4 h, after which medium was removed and formazan crystals were dissolved in DMSO. Absorbance values were determined using a spectrophotometer at a wavelength of 520 nm. For synergy analysis, SynergyFinder version 3.0 (synergyfinder.fimm.fi) was used to generate heatmaps and calculate ZIP synergy scores89.
Clonogenic assays: RPE1 TP53KO cells were seeded at a density of 200 cells/well of a 6-well plate, while OVCAR3 cells were seeded at 500 cells/well. 24 h after seeding, drugs were added at the indicated doses for 10 days. Culture medium was removed, and cells were fixed and stained using 0.2% Coomassie Brilliant Blue solution containing 50% methanol (Merck) and 14% acetic acid (Merck) in PBS. Plates were washed in tap water and air-dried overnight. Images of the plates were loaded into Adobe Photoshop, and a concentric circle was drawn to isolate the same area for each well. The magic wand tool was used to select colonies, and data per colony was exported. Subsequently, data for each colony were loaded into RStudio, and all areas with a circularity measure below 0.1 and an area below 12, which represent technical artifacts, were removed. The experiment was summarized to calculate area covered per well.
For experiments performed with RPE1 TP53KO PACKO ATF4KO and GCN2KO clones, cells were seeded at a density of 100 cells/well in 6-well plates or at 300–1000 cells/dish in 10 cm dishes containing dilutions of AZD1775 and vehicle. After 12–20 days, surviving colonies were fixed and stained with crystal violet (0.4% w/v crystal violet (Sigma), 20% methanol). Quantification of experiments was performed using a GelCount scanner (Oxford Optronix), and surviving fractions were calculated by normalization to vehicle treatments.
For experiments with the IncuCyte S3 Live-Cell Analysis Instrument (Essen Bioscience, Sartorius), RPE1 TP53KO PACKO cells were seeded in 96-well plates (100-300 cells per well) in the presence of serial dilutions of the indicated concentrations of AZD1775 and vehicle controls. Cells were imaged with a 10X objective once the vehicle controls reached confluency (approximately 6–9 days). Cell confluency was calculated automatically using IncuCyte segmentation algorithms based on phase contrast images and normalized to vehicle treatments.
For fluorescence-based cell competition assays, RPE1 TP53KO FLAG-Cas9 expressing cells were transduced at high MOI (MOI: 1) with plentiCRISPRv2 derived sgRNA expression vectors (plentiGuide) encoding EGFP-NLS or mCherry-NLS fluorescent proteins. For mCherry control expression vectors, β-galactosidase (LacZ) targeting sequences were used. Two days after infection (T−1), cells were seeded at low density in a 1:1 ratio in 12-well or 24-well plates to begin imaging the next day (T0) on a GE InCell Analyzer 6000 confocal microscope using a 4x objective. Where indicated, drug treatments were initiated on T1, and cultures were split in 1:20 fractions each time cells reached 90% confluency (approximately 3–4 days of growth). Cell fitness was calculated in each experimental condition by charting the ratio of GFP-positive to mCherry-positive cells at each imaging timepoint and normalizing the GFP:RFP ratio on day T0 to 1. Therefore, defects in cell fitness are presented as values < 1, and gain of fitness is represented by values > 1. The total number of GFP and mCherry-positive nuclei was segmented and counted in each well of an assay plate using a custom Acapella script (PerkinElmer).
Western blotting
Cells were trypsinized and lysed in Mammalian Protein Extraction Reagent (M-PER™, ThermoFisher) supplemented with Halt phosphatase inhibitor and Halt protease inhibitor (Thermo Scientific). Protein concentration was measured using Pierce BCA Protein Assay Kit (Thermo Scientific). Proteins were separated by SDS-PAGE gel electrophoresis and transferred to methanol-activated PVDF membranes (Immobilon) using the Trans-Blot Turbo Transfer System. Subsequently, membranes were blocked in 3% Bovine Serum Albumin Fraction V (Sigma-Aldrich) or 5% skim milk (Sigma-Aldrich) in 0.05% TBS-Tween (TBST). Immunodetection was performed using antibodies directed against puromycin (MABE343, Merck, 1:10000), GCN2 (3302, Cell Signaling, 1:1000), pGCN2 (ab75836, Abcam, 1:500), ATF4 (11815, Cell Signaling, 1:1000), Vinculin (ab129002, Abcam, 1:2000), HSP90 (sc-13119, Santa Cruz, 1:5000), pCDK1/2/3/5 (ab133463, Abcam, 1:1000), eIF2A (ab169528, Abcam, 1:1000), WEE1 (4936S, Cell signaling, 1:1000), rabbit anti-cyclin B1 (Santa Cruz, sc-752, 1:1000), and GCN1L1 (A301-843A, Bethyl laboratories, 1:1000) followed by staining with secondary antibodies Goat Anti-Rabbit Immunoglobulins/HRP (Dako, P0448) or Rabbit Anti-Mouse Immunoglobulins/HRP (Dako, P0260). Chemiluminescence detection was performed using Lumi-Light (Lumi-Light, Roche Diagnostics) or SuperSignal West Femto (SuperSignal, ThermoFisher) using ChemiDoc XRS+ System or ChemiDoc Imaging System (Bio-Rad).
For the RPE1 TP53KO PACKO, GCN2KO, ATF4KO, ZNL 02-096, and emetine immunoblot experiments, whole cell lysates were prepared by scraping cells in warm 2x Laemmli sample buffer (2% SDS, 167 mM Tris HCl pH 6.8, 20% glycerol, 0.02% bromophenol blue, 2 mM DTT). Proteins were separated by SDS-PAGE and the gel was stained with Ponceau S. Afterwards, proteins were transferred onto nitrocellulose membranes and subsequently blocked with 5% milk or in 0.1% TBST and blotted with the following primary antibodies for 1 h at room temperature (RT) or overnight at 4 °C: 1:1000 rabbit anti-phospho-eIF2a Ser51 (Cell Signaling, #3398), 1:1000 rabbit anti-phospho-eIF2a Ser51 (Cell Signaling, #9721), 1:1000 rabbit anti-eIF2a (Cell Signaling, #9722), 1:1000 rabbit anti-phospho-GCN2 Thr899 (Abcam, ab75836), 1:1000 rabbit anti-GCN2 (Abcam, ab134053), 1:1000 rabbit anti-ZNF598 (Sigma, HPA041760), 1:1000 rabbit anti-PERK (Cell Signaling, #5683), 1:1000 rabbit anti-phospho-cdc2 (CDK1) Tyr15 (Cell Signaling, #4539), 1:500 rabbit anti-ATF4 (Cell Signaling, #11815), 1:500 mouse anti-ATF4 (Cell Signaling, #97038), 1:500 rabbit anti-WEE1 (Cell Signaling, #13084). Blots were stained with the following secondary antibodies (1:10000 in 5% milk in 0.1% TBST) for 1 h at RT, and chemiluminescent signal (SuperSignal West Dura, Thermo Fisher) was captured on film: horseradish peroxidase (HRP) conjugated AffiniPure goat anti-rabbit IgG (Jackson ImmunoResearch 111-035-144), HRP AffiniPure goat anti-mouse IgG (Jackson 115-035-003), HRP sheep anti-mouse IgG (Amersham NA931V).
Puromycin assays
To assess protein synthesis rates, 20 μg/mL of puromycin was added to cells for 10 min before harvesting and lysis to label nascent proteins with puromycin. Further sample processing was the same as the general western blotting procedure as mentioned above.
Western blot quantification
Quantification of puromycin incorporation and WEE1, GCN2, and pGCN2 protein levels was performed by measuring grey values of puromycin lanes, loading control bands, and background in Adobe Photoshop. Grey values were reverted, background signal values for puromycin or the protein of interest, and loading controls were subtracted from their respective sample values. Corrected puromycin or protein signal values were corrected using loading control values. pGCN2 values were divided by total GCN2 values to obtain the plotted ratios. Treated conditions were normalized to the control condition.
Metabolic labeling with L-azidohomoalanine
To assess bulk mRNA translation, RPE1 TP53KO PACKO cells were seeded in 15 cm dishes and then treated or not with AZD1775 and ISRIB for 24 h. Cells were washed once in PBS and then grown in DMEM high glucose without glutamine, methionine, or cysteine (Wisent) with 10% FBS containing 25 μM Click-iT L-azidohomoalanine (AHA, Thermo Fisher) for 2 h. Cells were collected by trypsinization and washed twice with PBS before lysis in buffer containing 1% SDS, 50 mM Tris pH 8.0, 1x complete protease inhibitor cocktail (Roche), and 150 U/ml benzonase nuclease (Sigma). Whole cell lysates were incubated on ice for 30 min and pelleted by centrifugation in a refrigerated microcentrifuge for 10 min at 4 °C. Supernatants were normalized using a Nanodrop spectrophotometer, and 1 μl of 10 mM IRDye800CW-DBCO (Li-Cor Biosciences) was added to each sample and incubated for 2 h at RT in the dark. Excess dye was removed by centrifugation of samples through Zeba spin desalting columns (7 K MWCO, 0.5 ml, Thermo Fisher) according to the manufacturer’s instructions. Reactions were boiled in 1X Laemmli sample buffer, and SDS-PAGE was performed with Novex 4–12% Tris-Glycine gels (Invitrogen). IRDye800CW infrared fluorescence was detected on an Odyssey 9120 imager (Li-Cor), and total proteins were stained with Coomassie staining solution (0.1% w/v Coomassie Brilliant Blue, 20% methanol, 10% acetic acid). Total AHA labeled proteins were quantified by densitometry in Fiji (ImageJ) with background IRDye800CW fluorescence subtracted from measurements and normalized to AZD1775 and ISRIB vehicle (DMSO).
Flow cytometry
For cell cycle analysis, RPE1 TP53KO PACKO control, ATF4KO #1, GCN2KO #1, and GCN2KO #2 cells were seeded at a density of 1.20-1.75*105 cells/T25 and treated with 500 nM doxorubicin for 1 h before the addition of 62.5 ng/ml nocodazole to arrest cells in mitosis. Additionally, in RPE1 TP53KO-only experiments, cells were treated with A92 and AZD1775 simultaneously with 500 nM doxorubicin. After 16 h of treatment, cells were harvested and fixed in ice-cold 70% ethanol. Cells were stained with primary antibodies against phospho-Ser/Thr-Pro MPM-2 (05-368, Millipore, 1:100) and γH2AX (#9718, Cell Signaling, 1:100), and Alexa-Fluor 488 and 647 conjugated secondary antibodies (A32728 and A21206, ThermoFisher). DNA was stained using Propidium Iodide (PI; Sigma, P4864) while lysing RNA using RNAse A (Sigma, R6513). Samples were measured on a Quanteon NovoCyte (NovoCyte Quanteon, Agilent) and analyzed using FlowJo VX.
For ATF4-mScarlet reporter experiments, RPE1 TP53KO ATF4-mScarlet-NLS cells were seeded at a density of 30,000 cells/well in a 6-well plate. Cells were treated with indicated doses of thapsigargin, AZD1775, Debio 0123, A92, RP-6306, AZD7762 or VE-822. After 24 h, or for GCN1 siRNA experiments after 5 h, cells were harvested and measured on a Quanteon flow cytometer (NovoCyte Quanteon, Agilent) and analyzed using FlowJo VX. For the G1 arrest experiments, cells were plated at 90,000 cells/T25 density and treated with 62.5 ng/ml nocodazole or vehicle for 16 h before performing mitotic shakeoff, or in case of nocodazole-free controls, harvested by trypsinization. Subsequently, nocodazole was washed out and the cells were replated in the presence of 1.25 µM palbociclib or vehicle. After the cells had adhered, 1 µM AZD1775 and 1 µM A92 were added for approximately 20 h. After treatment, cells were harvested, measured on a Quanteon NovoCyte (NovoCyte Quanteon, Agilent), and analyzed using FlowJo VX.
PBMC activation
PBMCs were seeded at 1.25 M cells/ml density in 12-well plates. CD3+ cells were activated with a ratio of 1:8 Dynabeads Human T-Activator CD3/CD28 beads (Gibco, 11131) per cell for 4 days before the addition of inhibitors. 24 h after the addition of AZD1775 and thapsigargin, the PBMCs were pelleted and subsequently prepared for western blot as described.
DNA fiber assays
RPE1 TP53KO and OVCAR3 cells were seeded at 50,000 cells/well in a 6-well plate and incubated for 24 h. Cells were pulse-labeled with 25 μM 5-iodo-2′-deoxyuridine (IdU; Sigma-Aldrich) for 20 min, washed three times with pre-warmed culture media, and subsequently pulse-labeled with 250 μM 5-chloro-2′-deoxyuridine (CldU; Sigma-Aldrich) for 20 min. Cells were trypsinized and diluted to a concentration of 50,000 cells/mL. Cells were lysed in lysis buffer (0.5% sodium dodecyl sulfate, 200 mM Tris pH 7.4, 50 mM EDTA) and allowed to spread by gravity flow at an angle of approximately 20 degrees. Slides were air-dried, fixed in methanol/acetic acid (3:1 ratio) for 15 min, and denatured in 2.5 M HCl for 1.5 h. A primary antibody directed against BrdU (BD Biosciences, 347580, 1:250, mouse) was used to detect IdU, while CldU was detected using a primary antibody directed against BrdU (Abcam, ab6326, 1:1000, rat). Slides were incubated with primary antibodies for 2 h at RT, followed by incubation with Alexa Fluor 488- or 647-conjugated secondary antibodies (1:500) for 1.5 h. Images were acquired on a Leica DM-6000 fluorescence microscope, equipped with Leica Application Suite software. The lengths of IdU and CldU tracks were measured using ImageJ.
Immunofluorescence microscopy
Eighteen millimeter square coverslips were transferred into 6-well plates and sterilized in a microwave at 900 W for 2 min. RPE1 TP53KO and EIF2AKO cells were seeded onto the coverslips at a 30,000 cells/well density and incubated for 24 h. Soluble proteins were extracted by incubation with CSK buffer pH 7 (10 mM PIPES pH 7, 100 mM NaCl, 300 mM sucrose, 3 mM MgCl2, 0.5% Triton X-100) for 2 min on ice before fixation using 2% paraformaldehyde, permeabilization using PBS-0.1% Triton X-100 and blocking with 3% BSA V. Cover slips were incubated overnight at 4 °C in primary antibodies directed against γH2AX (05-636, Millipore, 1:300). Cells were then incubated with an Alexa Fluor 647-conjugated secondary antibody (A32728, ThermoFisher) for 2 h at RT (1:500) and stained with DAPI. Finally, cells were mounted with Vectashield anti-fade mounting medium (Vectorlabs). Imaging was performed using the Zeiss Axio Imager Z2, and analysis was performed in FIJI. Experimental details for quantitative image-based cytometry (QIBC) of ATF4 levels are provided in the Supplementary methods.
SILAC labeling
Labeled isotopes (10Arg and 8Lys), SILAC DMEM (5 g/L glucose), and SILAC FBS were obtained from Silantes. Medium supplemented with labeled isotopes is called “heavy”, while medium without labeled isotopes is called “light”. RPE1 TP53KO and RPE1 TP53KO EIF2AKO cells were cultured in either heavy or light medium for 5 passages (19 days) to saturate all proteins with isotope-labeled or unlabeled amino acids. Cells were treated with DMSO or 125 nM of AZD1775 to avoid excessive cell death in the more sensitive EIF2AKO cell line. For all samples, a replicate was performed, swapping the heavy and light labels to obtain n = 2 within the experiment. At 0, 6, 24, and 48 h, samples were taken by trypsinization, washing with PBS, and the dry cell pellet was snap-frozen and stored at −20 °C. SILAC samples were lysed in MPER supplemented with protease and phosphatase inhibitors, and protein concentration was determined using a BCA assay. Heavy and light samples were mixed to obtain a 1 to 1 ratio of proteins containing 50 μg of protein in 50 μL of MPER. Per sample, 6X loading buffer with 10% β-mercaptoethanol was added, and samples were boiled for 5 min at 95 °C. Samples were loaded onto an SDS-PAGE gel, and a single fraction was isolated for in-gel protein digestion. Slices were cut into 1 mm pieces and destained with 100 mM ammonium bicarbonate (ABC) in 50–70% acetonitrile. Reduction (10 mM dithiothreitol in 100 mM ABC) and alkylation (55 mM iodoacetamide in 100 mM ABC) steps were performed to block cysteines. Gel pieces were dehydrated and incubated overnight with 10 ng per μL trypsin (Promega), diluted in 100 mM ABC at 37 °C. Peptides were subsequently extracted with 5% formic acid for 20 min. Mass spectrometry (MS) instrument details are included in the Supplementary methods.
The MS data were analyzed with PEAKS Studio 10.6 build 20201221, and mapped on the UniProt human proteome build with accession number UP000005640. Missed cleavages were set to 3; for SILAC ratios, condition 1 was set to light(lys0/arg0), condition 2 was set to heavy (lys6/arg10); proteins were filtered for protein ID (FDR 1%) and for common contaminants. Protein ratios were calculated from normalized protein counts and converted to a log fold change.
WEE1 and GCN2 kinase engagement assays
NanoBRET assays: To assess AZD1775, Debio 0123, ZNL 02-096, GCN2iB, RP-6306, CC1 and neratinib binding to WEE1 or the GCN2 kinase domain in cells, HEK293 cells were reverse transfected with NanoLuc-EIF2AK4 (Domain 2) (Promega, NV3051) or WEE1-NanoLuc (Promega, NV2231) and transfection carrier DNA in opaque 96-well plates (Corning, #3917). 24 h after transfection, fluorescent tracer K-10 (for GCN2; Promega, N2640) or K-5 (for WEE1; Promega, N282) was added to cells at a final concentration of 0.5 μM or 0.125 µM, respectively, and plates were mixed for 15 s on an orbital shaker at 900 rpm. Serial dilutions of each compound were added to cells, and plates were incubated for 2 h. NanoBRET signal was measured by incubating cells with diluted Nano-Glo substrate according to the manufacturer’s instructions and detecting donor and acceptor emissions with an EnVision microplate reader (PerkinElmer). BRET ratios corresponding to tracer occupancy were determined by dividing acceptor emission by donor emission and multiplying by 1000 to compute mBu units. Background mBu signal was measured in wells where tracer compound was absent. Fractional occupancy of the kinase domain for each compound was calculated using the following formula: [1 – (X-Z)/(Y-Z)] * 100, where X = mBu in the presence of test compound and tracer, Y = mBu in the presence of vehicle (100% BRET signal), and Z = mBu in the absence of tracer compound (no BRET control).
To assess full-length GCN2 and WEE1 kinase activity upon treatment with WEE1i in vitro, AZD1775, Debio 0123, GCN2iB, and neratinib were sent to Eurofins for analysis with KinaseProfiler (items 14-934KP and 14-925KP, Eurofins).
Single cell EdU sequencing (scEdU-seq)
To assess replication dynamics, scEdU-seq was performed as described previously90. Briefly, RPE1 TP53KO control and EIF2AKO cells were treated as indicated and pulsed with 10 µM EdU for 15 min. The cells were subsequently harvested and fixed. Subsequently, pelleted cells were subjected to an EdU click reaction using to EdU-Click 647 imaging kit (Invitrogen) using the manufacturer’s instructions with minor modifications. Instead of Azide-647, 2 mM of azide-PEG3-biotin conjugate (Sigma) was used and supplemented with 6 mM tris((1-hydroxy-propyl-1H-1,2,3-triazol-4-yl)methyl)amine (Jena Bioscience). After this, cells were sorted to achieve single cells per well and lysed with Proteinase K. To digest the genome, NIaIII was added, and the resulting overhangs were processed to blunt ends using Klenow large fragment. End-repaired DNA fragments were A-tailed, and these fragments were ligated to T7 promoter-containing adapters with cell barcodes and UMI. Further details of cDNA preparation are provided in the Supplementary Methods. cDNA libraries were sequenced using v2.5 chemistry on a NextSeq500 or NextSeq2000 (Illumina; NextSeq control software v.2.2.0.4; RTA v.2.4.11) with 100 cycles for read 1 (cell index and tables were filtered for the presence of a NlaIII restriction site, a mapping quality >30, the molecule has a pair of reads assigned, the molecule is unique and should not have alternative alignment positions in the genome. The code for analysis and plotting is available on GitHub: https://github.com/jervdberg/WEE1.
Polysome profiling
To assess bulk mRNA translation, RPE1 TP53KO PACKO cells were treated with 250 nM AZD1775 or DMSO for 6 h. Cells from six 15 cm dishes per condition were pre-treated with 100 μg/ml cycloheximide (CHX) for 5 min and washed once with cold PBS containing 100 μg/ml CHX. Cells were scraped in PBS-CHX on ice and pelleted before resuspension in hypotonic buffer (5 mM Tris HCl, pH 7.5, 2.5 mM MgCl2, 1.5 mM KCl, 1 mM DTT, 100 U/ml RNasin recombinant RNase inhibitor (Promega), 100 μg/ml CHX). Cells were lysed by addition of 10% Triton X-100 (final concentration 0.5%) and 10% sodium deoxycholate (0.5% final) and vortexing for 5 s. Whole cell lysates were cleared by centrifugation for 10 min at maximum speed in a cooled tabletop microcentrifuge. Lysates were normalized by measuring absorbance at 260 nm (A260) on a Nanodrop spectrophotometer. For each sample, 15 A260 optical units were loaded on 5–50% sucrose gradients prepared in polysome buffer and centrifuged in an Optima XPN-80 ultracentrifuge with SW41Ti rotor (Beckman Coulter) at 39,000 rpm for 2 h at 4 °C. Polysome profiles were collected with a density gradient fractionation system (Brandel) measuring absorbance at 254 nm.
Stopless ORF circular (soc)RNA assays
To investigate changes in ribosome kinetics, assays using socRNAs were performed as described previously61. Briefly, U2OS cells expressing STAb-GFP, tet repressor, and ALFA nanobody-CAAX were seeded at 25% confluency in 96-well glass-bottom plates (Matriplates, Brooks Life Science Systems) and after 24 h, were transfected with the plasmid encoding the socRNA using Fugene (Promega). Subsequently, cells were treated with 300 nM AZD1775 and/or 1 µM A92 for 24 h. Imaging was done the following day by replacing the medium with pre-warmed imaging medium (CO2-independent Leibovitz’s-15 medium (Gibco) containing 5% FCS (Sigma-Aldrich) and 1% penicillin/streptomycin (Gibco)). Ninety minutes prior to the start of imaging, doxycycline (Dox, 1 µg/mL) was added to the cells to induce socRNA expression. All live-cell imaging experiments were performed at 37 °C using the Nikon TI inverted microscope with NIS Element Software equipped with a perfect focus system, a Yokagawa CSU-X1 spinning disc, an iXon Ultra 897 EM-CCD camera (Andor), and a motorized piezo stage (Nanocan SP400, Prior). The microscope was equipped with a temperature-controlled box. A 100 × 1.49 NA oil immersion objective was used for all imaging experiments. To precisely determine the number of translating ribosomes on socRNAs, the translation inhibitor puromycin (0.1 mg/mL; ThermoFischer Scientific) was added to cells 1–2 h after the start of imaging to induce nascent chain release. For live-cell imaging of socRNAs, the x, y positions for imaging were chosen based on the presence of translating socRNAs in cells. Images were acquired every 90 s for 2–3 h, with an exposure time of 100 ms for the 488 nm laser. Single z-plane images were acquired with focus on SunTag-GFP foci on the plasma membrane. For experiments in which the GFP fluorescence intensity of individual 24xSunTag arrays was measured, the cells were transfected with a plasmid encoding the 24xSunTag-CAAX protein.
mRNA sequencing (mRNA-seq) and ribosome profiling (Ribo-seq)
For mRNA sequencing in Fig. 2a and Supplementary Figure 4a, RNA was isolated from RPE1 TP53KO and RPE1 TP53KO EIF2AKO cells during log-phase growth using an RNeasy Mini kit (Qiagen) according to manufacturer’s instructions. 25 * 106 paired-end reads of 75 bp per sample from a NovaSeq 6000 were aligned to the Hg19 genome build using STAR, to obtain gene-level counts. Genes with low read counts were removed based on the optimal Jaccard similarity index computed using the R package HTSFilter91, and differentially expressed genes were identified using edgeR92.
For mRNA sequencing in Supplementary Fig. 8a, b, total RNA for mRNA-seq was prepared from RPE1 TP53KO PACKO cells harvested after 24 or 48 h of treatment with 250 nM AZD1775 or DMSO in biological duplicate using DirectZol RNA MiniPrep kits (Zymo Research) according to the manufacturer’s instructions. cDNA libraries were generated using TruSeq Stranded Total RNA library kits with Ribo-Zero Gold rRNA depletion (Illumina) and sequenced as 300-nucleotide paired-end reads on a NextSeq500 platform (LTRI NBCC). mRNA-seq reads were processed by alignment with Salmon to the reference transcriptome (human genome assembly GRCh38.p13 cDNA)93. Differential gene expression changes were calculated with DESeq2 with an FDR cutoff <0.1 for gene significance94. Gene set enrichment analyses were first performed with ClusterProfiler95. Gene set enrichment analysis (GSEA) software was used to identify the enrichment of genes transcriptionally regulated by AZD1775 (mRNA-seq) in genes identified by ribosome profiling. For Ribo-Seq profiling in Supplementary Fig. 9, RPE1 TP53KO cells were treated with 1 µM of AZD1775 for 24 h. Ribo-seq was performed as detailed in the Supplemental Methods.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The mass spectrometry proteomics data generated in this study have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository, under dataset identifier PXD054741. The single-cell EdU-Seq data generated in this study have been deposited to the NCBI-GEO repository, under identifier GSE287601. Sequencing data of the HAP1 screen generated in this study have been deposited in the NCBI Sequence Read Archive (SRA), under BioProject number PRJNA1152540. The CRISPR screen data generated in this study have been deposited at the NCBI SRA, with BioProject number PRJNA1233308. RNA sequencing data of RPE1 TP53KO control and EIF2AKO cells generated in this study have been deposited in the Gene Expression Omnibus under accession numbers GSE286143 and GSE291445. Raw sequencing Riboseq data generated in this study, along with metadata, and count tables, have been made available in the Gene Expression Omnibus under the accession number GSE291029. All other data are available in the article and its supplementary files or from the corresponding author upon request. Source data are provided with this paper.
Code availability
Scripts for RNA-seq and Ribo-seq library QC evaluation and to produce figures are available on GitHub (https://github.com/mvanins/WEE1_manuscript). The code for analysis and plotting of scEdU-Seq data is available on GitHub (https://github.com/jervdberg/WEE1)96.
References
Malumbres, M. & Barbacid, M. Cell cycle, CDKs and cancer: a changing paradigm. Nat. Rev. Cancer 9, 153–166 (2009).
Featherstone, C. & Russell, P. Fission yeast p107wee1 mitotic inhibitor is a tyrosine/serine kinase. Nature 349, 808–811 (1991).
Igarashi, M., Nagata, A., Jinno, S., Suto, K. & Okayama, H. Wee1(+)-like gene in human cells. Nature 353, 80–83 (1991).
Parker, L. L. & Piwnica-Worms, H. Inactivation of the p34cdc2-cyclin B complex by the human WEE1 tyrosine kinase. Science 257, 1955–1957 (1992).
Krek, W. & Nigg, E. A. Mutations of p34cdc2 phosphorylation sites induce premature mitotic events in HeLa cells: evidence for a double block to p34cdc2 kinase activation in vertebrates. EMBO J. 10, 3331–3341 (1991).
Booher, R. N., Holman, P. S. & Fattaey, A. Human Myt1 is a cell cycle-regulated kinase that inhibits Cdc2 but not Cdk2 activity. J. Biol. Chem. 272, 22300–22306 (1997).
Mueller, P. R., Coleman, T. R., Kumagai, A. & Dunphy, W. G. Myt1: a membrane-associated inhibitory kinase that phosphorylates Cdc2 on both threonine-14 and tyrosine-15. Science 270, 86–90 (1995).
Donzelli, M. & Draetta, G. F. Regulating mammalian checkpoints through Cdc25 inactivation. EMBO Rep. 4, 671–677 (2003).
Sanchez, Y. et al. Conservation of the Chk1 checkpoint pathway in mammals: linkage of DNA damage to Cdk regulation through Cdc25. Science 277, 1497–1501 (1997).
Matsuoka, S., Huang, M. & Elledge, S. J. Linkage of ATM to cell cycle regulation by the Chk2 protein kinase. Science 282, 1893–1897 (1998).
O’Connell, M. J., Raleigh, J. M., Verkade, H. M. & Nurse, P. Chk1 is a wee1 kinase in the G2 DNA damage checkpoint inhibiting cdc2 by Y15 phosphorylation. EMBO J. 16, 545–554 (1997).
Aarts, M. et al. Forced mitotic entry of S-phase cells as a therapeutic strategy induced by inhibition of WEE1. Cancer Discov. 2, 524–539 (2012).
O’Connor, M. J. Targeting the DNA damage response in cancer. Mol. Cell 60, 547–560 (2015).
Hirai, H. et al. Small-molecule inhibition of Wee1 kinase by MK-1775 selectively sensitizes p53-deficient tumor cells to DNA-damaging agents. Mol. Cancer Ther. 8, 2992–3000 (2009).
Leijen, S., Beijnen, J. H. & Schellens, J. H. M. Abrogation of the G2 checkpoint by inhibition of Wee-1 kinase results in sensitization of p53-deficient tumor cells to DNA-damaging agents. Curr. Clin. Pharmacol. 5, 186–191 (2010).
Chen, X. et al. Cyclin E overexpression sensitizes triple-negative breast cancer to wee1 kinase inhibition. Clin. Cancer Res. 24, 6594–6610 (2018).
Kok, Y. P. et al. Overexpression of Cyclin E1 or Cdc25A leads to replication stress, mitotic aberrancies, and increased sensitivity to replication checkpoint inhibitors. Oncogenesis 9, 88 (2020).
Pfister, S. X. et al. Inhibiting WEE1 selectively kills histone h3k36me3-deficient cancers by dNTP starvation. Cancer Cell 28, 557–568 (2015).
Bridges, K. A. et al. MK-1775, a novel Wee1 kinase inhibitor, radiosensitizes p53-defective human tumor cells. Clin. Cancer Res. 17, 5638–5648 (2011).
Hirai, H. et al. MK-1775, a small molecule Wee1 inhibitor, enhances anti-tumor efficacy of various DNA-damaging agents, including 5-fluorouracil. Cancer Biol. Ther. 9, 514–522 (2010).
Rajeshkumar, N. V. et al. MK-1775, a potent Wee1 inhibitor, synergizes with gemcitabine to achieve tumor regressions, selectively in p53-deficient pancreatic cancer xenografts. Clin. Cancer Res. 17, 2799–2806 (2011).
Mir, S. E. et al. In silico analysis of kinase expression identifies WEE1 as a gatekeeper against mitotic catastrophe in glioblastoma. Cancer Cell 18, 244–257 (2010).
Embaby, A. et al. WEE1 inhibitor adavosertib in combination with carboplatin in advanced TP53 mutated ovarian cancer: a biomarker-enriched phase II study. Gynecol. Oncol. 174, 239–246 (2023).
Fu, S. et al. Multicenter phase II trial of the WEE1 inhibitor adavosertib in refractory solid tumors harboring CCNE1 amplification. J. Clin. Oncol. 41, 1725–1734 (2023).
Leijen, S. et al. Phase I study evaluating WEE1 Inhibitor AZD1775 as monotherapy and in combination with gemcitabine, cisplatin, or carboplatin in patients with advanced solid tumors. J. Clin. Oncol. 34, 4371–4380 (2016).
Leijen, S. et al. Phase II study of WEE1 inhibitor AZD1775 plus carboplatin in patients with TP53-mutated ovarian cancer refractory or resistant to first-line therapy within 3 months. J. Clin. Oncol. 34, 4354–4361 (2016).
Carette, J. E. et al. Global gene disruption in human cells to assign genes to phenotypes by deep sequencing. Nat. Biotechnol. 29, 542–546 (2011).
Ducker, G. S. & Rabinowitz, J. D. One-carbon metabolism in health and disease. Cell Metab. 25, 27–42 (2017).
Bonagas, N. et al. Pharmacological targeting of MTHFD2 suppresses acute myeloid leukemia by inducing thymidine depletion and replication stress. Nat. Cancer 3, 156–172 (2022).
Heijink, A. M. et al. A haploid genetic screen identifies the G1/S regulatory machinery as a determinant of Wee1 inhibitor sensitivity. Proc. Natl. Acad. Sci. USA 112, 15160–15165 (2015).
Margolis, S. S. et al. Role for the PP2A/B56delta phosphatase in regulating 14-3-3 release from Cdc25 to control mitosis. Cell 127, 759–773 (2006).
Komar, A. A. & Merrick, W. C. A retrospective on eIF2A and not the alpha subunit of eIF2. Int. J. Mol. Sci. 21, 2054 (2020).
Kim, E. et al. eIF2A, an initiator tRNA carrier refractory to eIF2α kinases, functions synergistically with eIF5B. Cell. Mol. Life 75, 4287–4300 (2018).
Grove, D. J., Levine, D. J. & Kearse, M. G. Increased levels of eIF2A inhibit translation by sequestering 40S ribosomal subunits. Nucleic Acids Res. 51, 9983–10000 (2023).
Roiuk, M., Neff, M. & Teleman, A. A. Human eIF2A has a minimal role in translation initiation and in uORF-mediated translational control in HeLa cells. eLife 14, RP105311 (2025).
Yigit, M. et al. The alternative initiation factor eIF2A regulates 40S subunit turnover in ribosome-associated quality control. Preprint at https://doi.org/10.1101/2025.05.22.655611 (2025).
Gaikwad, S., Ghobakhlou, F., Zhang, H. & Hinnebusch, A. G. Yeast eIF2A has a minimal role in translation initiation and uORF-mediated translational control in vivo. eLife 12, RP92916 (2024).
Ichihara, K. et al. Combinatorial analysis of translation dynamics reveals eIF2 dependence of translation initiation at near-cognate codons. Nucleic Acids Res. 49, 7298–7317 (2021).
Merrick, W. C. & Anderson, W. F. Purification and characterization of homogeneous protein synthesis initiation factor M1 from rabbit reticulocytes. J. Biol. Chem. 250, 1197–1206 (1975).
Zoll, W. L., Horton, L. E., Komar, A. A., Hensold, J. O. & Merrick, W. C. Characterization of mammalian eIF2A and identification of the yeast homolog. J. Biol. Chem. 277, 37079–37087 (2002).
Levy, S., Avni, D., Hariharan, N., Perry, R. P. & Meyuhas, O. Oligopyrimidine tract at the 5’ end of mammalian ribosomal protein mRNAs is required for their translational control. Proc. Natl. Acad. Sci. USA 88, 3319–3323 (1991).
Philippe, L., van den Elzen, A. M. G., Watson, M. J. & Thoreen, C. C. Global analysis of LARP1 translation targets reveals tunable and dynamic features of 5’ TOP motifs. Proc. Natl. Acad. Sci. USA 117, 5319–5328 (2020).
Yamashita, R. et al. Comprehensive detection of human terminal oligo-pyrimidine (TOP) genes and analysis of their characteristics. Nucleic Acids Res. 36, 3707–3715 (2008).
Zyryanova, A. F. et al. ISRIB blunts the integrated stress response by allosterically antagonising the inhibitory effect of phosphorylated eIF2 on eIF2B. Mol. Cell 81, 88–103.e6 (2021).
Costa-Mattioli, M. & Walter, P. The integrated stress response: from mechanism to disease. Science 368, eaat5314 (2020).
Harding, H. P. et al. Regulated translation initiation controls stress-induced gene expression in mammalian cells. Mol. Cell 6, 1099–1108 (2000).
Nougarède, A. et al. Improved IRE1 and PERK pathway sensors for multiplex endoplasmic reticulum stress assay reveal stress response to nuclear dyes used for image segmentation. Assay. Drug Dev. Technol. 16, 350–360 (2018).
De Witt Hamer, P. C., Mir, S. E., Noske, D., Van Noorden, C. J. F. & Würdinger, T. WEE1 kinase targeting combined with DNA-damaging cancer therapy catalyzes mitotic catastrophe. Clin. Cancer Res. 17, 4200–4207 (2011).
Duda, H. et al. A mechanism for controlled breakage of under-replicated chromosomes during mitosis. Dev. Cell 39, 740–755 (2016).
van Vugt, M. A. T. M., Brás, A. & Medema, R. H. Polo-like kinase-1 controls recovery from a G2 DNA damage-induced arrest in mammalian cells. Mol. Cell 15, 799–811 (2004).
Aarts, M. et al. Functional genetic screen identifies increased sensitivity to WEE1 inhibition in cells with defects in Fanconi anemia and HR pathways. Mol. Cancer Ther. 14, 865–876 (2015).
Toledo, C. M. et al. Genome-wide CRISPR-Cas9 screens reveal loss of redundancy between PKMYT1 and WEE1 in glioblastoma stem-like cells. Cell Rep. 13, 2425–2439 (2015).
Hustedt, N. et al. A consensus set of genetic vulnerabilities to ATR inhibition. Open Biol. 9, 190156 (2019).
Giaever, G. et al. Genomic profiling of drug sensitivities via induced haploinsufficiency. Nat. Genet. 21, 278–283 (1999).
Clancy, A. et al. The deubiquitylase USP9X controls ribosomal stalling. J. Cell Biol. 220, e202004211 (2021).
Juszkiewicz, S., Speldewinde, S. H., Wan, L., Svejstrup, J. Q. & Hegde, R. S. The ASC-1 complex disassembles collided ribosomes. Mol. Cell 79, 603–614.e8 (2020).
Oltion, K. et al. An E3 ligase network engages GCN1 to promote the degradation of translation factors on stalled ribosomes. Cell 186, 346–362.e17 (2023).
Pisareva, V. P., Skabkin, M. A., Hellen, C. U. T., Pestova, T. V. & Pisarev, A. V. Dissociation by Pelota, Hbs1 and ABCE1 of mammalian vacant 80S ribosomes and stalled elongation complexes. EMBO J. 30, 1804–1817 (2011).
Sundaramoorthy, E. et al. ZNF598 and RACK1 regulate mammalian ribosome-associated quality control function by mediating regulatory 40s ribosomal ubiquitylation. Mol. Cell 65, 751–760.e4 (2017).
Joazeiro, C. A. P. Mechanisms and functions of ribosome-associated protein quality control. Nat. Rev. Mol. Cell Biol. 20, 368–383 (2019).
Madern, M. F. et al. Long-term imaging of individual ribosomes reveals ribosome cooperativity in mRNA translation. Cell 188, 1896–1911 (2025).
Han, P. et al. Genome-wide survey of ribosome collision. Cell Rep. 31, 107610 (2020).
Dmitriev, S. E., Vladimirov, D. O. & Lashkevich, K. A. A quick guide to small-molecule inhibitors of eukaryotic protein synthesis. Biochemistry 85, 1389–1421 (2020).
Szaruga, M. et al. Activation of the integrated stress response by inhibitors of its kinases. Nat. Commun. 14, 5535 (2023).
Tang, C. P. et al. GCN2 kinase activation by ATP-competitive kinase inhibitors. Nat. Chem. Biol. 18, 207–215 (2022).
Machleidt, T. et al. NanoBRET-A Novel BRET platform for the analysis of protein-protein interactions. ACS Chem. Biol. 10, 1797–1804 (2015).
Zhu, J.-Y. et al. Structural basis of wee kinases functionality and inactivation by diverse small molecule inhibitors. J. Med. Chem. 60, 7863–7875 (2017).
Garcia-Barrio, M., Dong, J., Ufano, S. & Hinnebusch, A. G. Association of GCN1-GCN20 regulatory complex with the N-terminus of eIF2alpha kinase GCN2 is required for GCN2 activation. EMBO J. 19, 1887–1899 (2000).
Anda, S., Zach, R. & Grallert, B. Activation of Gcn2 in response to different stresses. PLoS ONE 12, e0182143 (2017).
Benada, J. et al. Synthetic lethal interaction between WEE1 and PKMYT1 is a target for multiple low-dose treatment of high-grade serous ovarian carcinoma. NAR Cancer 5, zcad029 (2023).
Gallo, D. et al. CCNE1 amplification is synthetic lethal with PKMYT1 kinase inhibition. Nature 604, 749–756 (2022).
Szychowski, J. et al. Discovery of an orally bioavailable and selective PKMYT1 inhibitor, RP-6306. J. Med. Chem. 65, 10251–10284 (2022).
Carlson, K. R., Georgiadis, M. M., Tameire, F., Staschke, K. A. & Wek, R. C. Activation of Gcn2 by small molecules designed to be inhibitors. J. Biol. Chem. 299, 104595 (2023).
Li, Z. et al. Development and characterization of a Wee1 kinase degrader. Cell Chem. Biol. 27, 57–65.e9 (2020).
Drainas, A. P. et al. GCN2 is a determinant of the response to WEE1 kinase inhibition in small-cell lung cancer. Cell Rep. 43, 114606 (2024).
Juszkiewicz, S. et al. ZNF598 is a quality control sensor of collided ribosomes. Mol. Cell 72, 469–481.e7 (2018).
Yan, L. L. & Zaher, H. S. Ribosome quality control antagonizes the activation of the integrated stress response on colliding ribosomes. Mol. Cell 81, 614–628.e4 (2021).
Pisarev, A. V. et al. The role of ABCE1 in eukaryotic posttermination ribosomal recycling. Mol. Cell 37, 196–210 (2010).
Schutte, T. et al. Clinical development of WEE1 inhibitors in gynecological cancers: a systematic review. Cancer Treat. Rev. 115, 102531 (2023).
Zappa, F., Muniozguren, N. L., Conrad, J. E. & Acosta-Alvear, D. The integrated stress response engages a cell-autonomous, ligand-independent, DR5-driven apoptosis switch. Cell Death Dis. 16, 101 (2025).
Blomen, V. A. et al. Gene essentiality and synthetic lethality in haploid human cells. Science 350, 1092–1096 (2015).
Stok, C. et al. FIRRM/C1orf112 is synthetic lethal with PICH and mediates RAD51 dynamics. Cell Rep. 42, 112668 (2023).
Raaijmakers, J. A. et al. BUB1 is essential for the viability of human cells in which the spindle assembly checkpoint is compromised. Cell Rep. 22, 1424–1438 (2018).
Sherman, B. T. et al. DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 50, W216–W221 (2022).
Hart, T. et al. High-resolution CRISPR screens reveal fitness genes and genotype-specific cancer liabilities. Cell 163, 1515–1526 (2015).
Hart, T. et al. Evaluation and design of genome-wide CRISPR/SpCas9 knockout screens. G3 7, 2719–2727 (2017).
Noordermeer, S. M. et al. The shieldin complex mediates 53BP1-dependent DNA repair. Nature 560, 117–121 (2018).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Ianevski, A., Giri, A. K. & Aittokallio, T. SynergyFinder 3.0: an interactive analysis and consensus interpretation of multi-drug synergies across multiple samples. Nucleic Acids Res. 50, W739–W743 (2022).
van den Berg, J. et al. Quantifying DNA replication speeds in single cells by scEdU-seq. Nat. Methods 21, 1175–1184 (2024).
Rau, A., Gallopin, M., Celeux, G. & Jaffrézic, F. Data-based filtering for replicated high-throughput transcriptome sequencing experiments. Bioinforma. Oxf. Engl. 29, 2146–2152 (2013).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinforma. Oxf. Engl. 26, 139–140 (2010).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. Omics J. Integr. Biol. 16, 284–287 (2012).
van den, J. berg. jervdberg/WEE1: WEE1 scEdU-seq v1 (Version v1). Zenodo https://doi.org/10.5281/zenodo.17298047 (2025).
Acknowledgements
We thank members of the Durocher and van Vugt lab for critical input. We thank Repare Therapeutics for providing the PKMYT1 inhibitor RP-6306, and Prof. Sir Stephen Jackson for sharing unpublished data and coordination of publication. We thank Henrique Melo for initial CRISPR screen analyses, as well as Krzysztof Szkop, Shannon Mclaughlan, Ivan Topisirovic, Ola Larsson for advice and help with polysome profiling and a RiboSeq experiment that was not included in this final manuscript; Kin Chan, and Monica Hasegan of the NBCC at the LTRI for expert assistance with sequencing and high content microscopy, respectively. We thank Maarten Hekkelman of the NKI for help with bioinformatics analysis of HAP1 screens. We thank Lisa Koob for assistance with the AZD1775 negative selection CRISPR screen. This work was supported by the Dutch Cancer Society (KWF-11352 to M.A.T.M.v.V., S.d.J., T.R.B., and KWF-12911 to M.A.T.M.v.V.), the Netherlands Organization for Scientific Research (NWO-VICI #09150182110019 to M.A.T.M.v.V) and the Canadian Institutes for Health Research (CIHR, grant PJT 180438 to DD). A.v.O. is supported by European Research Council Advanced grant (ERC-AdG grant no. 101053581-scTranslatomics). J.v.B. and A.v.O. are supported by Novo Nordisk Fonden Synergy Program (#0091873-Lost Memories).
Author information
Authors and Affiliations
Contributions
S.d.J., T.R.B., D.D., M.A.T.M.v.V. conceived the project. R.B.T., T.F.N., M.R., C.B., M.E., F.J.B., and M.L. performed wet lab experiments. N.H. and T.F.N. carried out the CRISPR screens that were analyzed by L.H. D.B. and T.R.B. performed and analyzed the HAP1 insertional mutagenesis screen. N.M. carried out the NanoBret experiments. J.Y. prepared purified GCN2 under supervision of F.S. S.Y. and M.E.T. performed and analyzed the socRNA experiments. M.V., A.v.O., and J.v.d.B. performed and analyzed scEdU-seq and RiboSeq experiments. H.R.d.B. analyzed the SILAC proteomics data. R.B.T., T.N., M.R., D.D., and M.A.T.M.v.V. wrote the manuscript. All authors provided input on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
D.D. and F.S. are shareholders and advisors for Repare Therapeutics. M.A.T.M.v.V. acted on the Scientific Advisory Board of Repare Therapeutics. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tjeerdsma, R.B., Ng, T.F., Roorda, M. et al. WEE1 inhibitors trigger GCN2-mediated activation of the integrated stress response. Nat Commun 16, 11598 (2025). https://doi.org/10.1038/s41467-025-66514-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66514-0