Abstract
Methane, a predominant component of human intestinal gas, has been reported to be associated with a reduction in intestinal transit speed, as well as correlations with elevated body mass index. While the gut methanogenic archaea that produce this gas have been studied, the countervailing role of methane-consuming bacteria (methanotrophs) within the human gut ecosystem remains a critical, under-explored area. The potential for these bacteria to act as a built-in sink for intestinal methane and thereby mitigate its negative physiological effects is unknown. Here, we isolate an unreported methanotroph from human fecal samples, classified as Methylocystis intestini. Using a mouse model, we observe that methane challenge is associated with gastrointestinal motility and fat metabolism. We then demonstrate that the administration of Methylocystis intestini effectively reverses these dysfunctional processes, restoring motility and metabolic parameters. Additional analysis of methane-oxidation genes abundance in 1207 public metagenomic sequences from individuals with varying health statuses, including obesity and constipation, provides consistent correlative support for our experimental conclusions. Expanding this view to a global scale, we conducted a metagenomic survey of 550 human fecal samples from populations across five continents. This broader analysis reveals that methane-oxidizing genes are not a rarity but a common feature of the human gut microbiome, being detectable in over 91% of samples. This ubiquity underscores their fundamental role in human biology. Collectively, our findings establish gut methanotrophs as key mediators of intestinal methane level. Their presence is widespread across global populations, and their functional capacity can balance the effects of methane on host physiology. This work elucidates a crucial component of gut homeostasis and opens a promising avenue for developing microbiome-based therapeutic strategies aimed at managing methane-related gastrointestinal disorders by harnessing the power of these native methane-consuming bacteria.
Similar content being viewed by others
Introduction
The human gut microbiome is a complex and dynamic ecosystem that significantly influences various physiological processes, including digestion, metabolism, and immune function. One critical aspect of gut health that has garnered increasing attention is the role of gut-derived gases, particularly hydrogen (H₂), carbon dioxide (CO₂), and methane (CH₄). These gases are produced as by-products of microbial fermentation and can have profound effects on gastrointestinal motility, nutrient absorption, and overall gut function1,2. Among these gases, more and more studies have proved that methane plays an important role in modulating gastrointestinal health, and its presence in the human gut is closely linked to several gastrointestinal disorders, including constipation, bloating, and irritable bowel syndrome3,4.
Methane in the gut is primarily produced by methanogenic archaea, which metabolize hydrogen gas and carbon dioxide to produce methane. This microbial process, although beneficial in reducing hydrogen levels, can have deleterious effects when methane accumulates excessively5. Elevated methane levels have been associated with delayed gastrointestinal motility. Studies have demonstrated that increased methane production in the gut leads to reduced intestinal peristalsis, slower gastric emptying, and decreased colonic transit time, all of which contribute to symptoms of constipation and discomfort6. Moreover, methane accumulation has been implicated in altered metabolic processes, including fat metabolism, suggesting that it could play a role in obesity and metabolic syndrome7,8,9.
However, much less attention has been paid to the microbial counterparts that could mitigate the effects of excess methane in the gut. Methanotrophs, a group of bacteria capable of oxidizing methane, represent an important functional group active in many environmental niches. However, they remain an underexplored component of the gut microbiota. These bacteria utilize methane as their sole carbon and energy source, converting it into biomass and carbon dioxide. This metabolic ability could offer a natural mechanism to reduce methane levels in the gut and potentially alleviate methane-induced gastrointestinal dysfunction. To date, the presence, diversity, functional roles of methanotrophs and their potential impact on gastrointestinal motility and metabolic regulation remain poorly understood.
The lack of detailed studies on gut methanotrophs stands in contrast to the growing body of research on methane-producing archaea, highlighting a significant gap in our understanding of the microbial dynamics in the human gut. While methanogens have been studied extensively due to their role in methane production, the functional contribution of methanotrophic bacteria in maintaining gut homeostasis and mitigating methane-induced disorders has been largely overlooked. Given the potential therapeutic benefits of methane reduction, it is essential to investigate the role of methanotrophs in the gut ecosystem and their ability to modulate gastrointestinal function and metabolic health.
In this study, we isolate and characterize a methanotrophic bacterium from human fecal samples, assess its methane-oxidizing capabilities, and evaluate its impact on methane-induced gastrointestinal dysfunction and metabolic disturbances. Our work provides some insights into the ecological role of gut methanotrophs and explores their therapeutic potential in alleviating conditions such as constipation and obesity, both of which are linked to altered methane dynamics in the gut, using mice as an experimental model. By filling the knowledge gap on the prevalence and functional significance of gut methanotrophs, this study may pave the way for microbiome-based therapies for methane-associated gastrointestinal and metabolic disorders.
Results
Isolation and characterization of a gut methanotroph
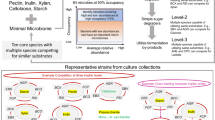
To isolate methanotrophs from the human intestine, we inoculated NMS1 medium with human fecal material. Growth of bacteria was observed in medium with methane as the sole carbon source (Fig. 1A). Using primers designed for the methane-oxidation gene (MOG) pmoA, we confirmed its presence in the fifth generation of the subcultures (Fig. 1B). And fluorescence in situ hybridization (FISH) further demonstrated the enrichment of methanotrophic bacteria (Fig. 1C). The metagenomic sequencing revealed a progressive increase in the relative abundance of Methylocystis after continuous subculturing (Fig. 1D). Analysis of draft genomes identified a complete pmoCAB methane oxidation gene cluster in only a subset of Methylocystis genomes (Fig. S1).
A The pipeline of isolation of gut methane-oxidizing bacteria (MOB). B Detection of the amplified pmoA-gene obtained with the primer pair Adj-pmoA. S1-S4 means four 5th cultures, and NK means non-treatment blank group. Single experiment. C Representative Fluorescence micrograph of the bacteria from gut contents. From left to right means micrograph using prokaryote staining DAPI, the MOB labeled with the red fluorescent, and an overlay of the blue DAPI stained cells with red MOB results in pink cells. Two biological replicates showed the similar results. D The altered relative abundance of gut microbes with culturing. Only the top three microbes with the most relative abundance in different samples are shown. E Neighbor-joining phylogenetic tree of the MOB based on the 16S rRNA genes, with gut MOB in red font. Type strains are identified with ‘T’. Bootstrap 1000 times. Bar, 0.01 substitutions per nucleotide position. F Representative electron micrographs of ultrathin sections of gut MOB cells. Bar, 0.2 µm. ICM: Intracytoplasmic membranes. Four biological replicates showed similar results. G The whole genome circle plot of gut MOB, and the region of methane-oxidizing genes are shown under the circle plot. Source data are provided as a Source Data file.
Following liquid enrichment and strain purification, we isolated a methanotrophic bacterial colony that was identified as a member of the genus Methylocystis within the class Alphaproteobacteria (Fig. 1E). Based on average nucleotide identity (ANI) analysis, we confirmed that the isolated single colony and the enriched culture containing methane oxidation genes represented the same strain (ANI > 99%; Fig. S2). However, genome comparison with known Methylocystis species revealed an ANI of less than 85.6%, suggesting that this strain represents a potential unreported species (Fig. S3). We tentatively designated it Methylocystis intestini. Transmission electron microscopy further revealed a distinct intracytoplasmic membrane (ICM) system characteristic of type II methanotrophs (Fig. 1F). Genomic analysis further indicated the presence of three copies of the pmo gene clusters (pmoCAB) and the complete set of genes for the downstream methylotrophy (Figs. 1G and S4). More than 15% of the genes (573 out of 3776) in M. intestini shared less than 70% amino acid sequence identity with those of other Methylocystis species (Fig. S5). In addition, M. intestini exhibited optimal growth at 37 °C compared with 30 °C (Fig. S6). And we found that the methane concentration in the headspace gas of the culture bottles inoculated with M. intestini decreased significantly, indicating active methane oxidation (Fig. S7). These findings highlight unique genomic and physiological features of this strain, which may underlie its adaptation to the gut environment.
Gut methanotrophs alleviate methane-associated constipation
Methane has been reported to be related to slow intestinal peristalsis10. To investigate whether M. intestini could mitigate this effect, we conducted a series of mice experiments (Fig. 2A). Using metagenomic sequencing and RT-qPCR, we found the M. intestini can successfully colonize in the intestine within five days after gavage (Figs. 2B and S8). Exhaled gas analysis after one week of intervention demonstrated that oral methane administration significantly increased the methane concentration in mice, whereas colonization with M. intestini markedly reduced the elevated methane levels (Fig. S9). Consistent with previous studies, methane exposure significantly reduced fecal water content (Fig. 2C, D) and increased intestinal transit time (Fig. 2E–G). And these effects can be alleviated by the intervention of M. intestini but not heat-killed M. intestini or other bacteria (Fig. S10). Additionally, histological examination revealed no obvious differences in epithelial morphology between the control and CH₄ groups (Fig. S11). Shotgun metagenomic sequencing showed that CH₄ significantly reduced the α-diversity and disrupted the microbial community structure in the colon, whereas supplementation with M. intestini restored diversity to control levels (Fig. S12A–C). And we did not observe enrichment of methanotrophs after CH₄ administration, nor suppression of methanogens following M. intestini gavage (Fig. S12D).
A The experimental design. B The relative abundance of M. intestini in different regions of the intestinal tract (n = 3). P-value: 1.05 × 10−7 (CH4+MOB vs. CH4 in ileum), 1.44 × 10−5 (CH4+MOB vs. Control in ileum), 3.03 × 10−5 (CH4+MOB vs. CH4 in cecum), 1.15 × 10−5 (CH4+MOB vs. Control in cecum). C The representative feces samples in different groups, and the water content of feces are shown in (D) (n = 5). E The whole gut transit time (n = 5), F, G small intestine transit rate in different groups (n = 5). H Spontaneous contractile activity of intestinal segment from different groups, and the frequency spectrums of them are shown in the (I). The frequency with the highest average amplitude is marked with a red box. Data are presented as means ± standard of mean in (B, G), except for boxplots in (D, E) (center line or point, median; box limits, 25th and 75th percentile; whiskers, Tukey; points, outliers). Statistical analysis was performed with 1-way ANOVA with Dunnett’s corrections in (B), two-tailed Mann-Whitney-Wilcoxon test with Benjamini/Hochberg correction in D and two-tailed unpaired t-test with Benjamini/Hochberg correction (E, G). Individual data points are independent biological replicates unless otherwise stated. Source data are provided as a Source Data file.
To further rule out other methane-induced factors, we performed an in vitro organ bath experiment (Fig. 2H). Methane exposure reduced peristaltic frequency to less than 0.2 Hz, while the addition of methane-oxidizing bacteria restored it to levels greater than 0.5 Hz, comparable to the control group. Similar results were also observed in rats (Fig. S13).
Association of gut methanotrophs with fat metabolism
It is reported that excessive methane is related to the accumulation of fat in overweight patients8,9. To explore this, we established three groups of mice and administered interventions by oral gavage for four weeks (Fig. 3A). While body weight remained comparable across groups (Fig. S14), methane exposure significantly increased fat deposition in some types of white adipose tissue (Fig. 3B). These effects were reversed by methane-oxidizing bacteria intervention. However, this effect was not altered by the intervention with heat-killed methane-oxidizing bacteria or by other bacterial treatments (Fig. S15). Moreover, a similar pattern was observed in an additional mouse strain (Fig. S16). In addition, we observed a reduction in blood cholesterol levels following methane exposure, whereas the group treated with M. intestini exhibited a decreasing trend (Fig. 3C).
A The experimental design for four-week intervention of methane and gut MOB. B Comparisons among different groups in adipose tissue weight from different depots (n = 4). C Blood indexes of mice. TC: total cholesterol. HDL: high-density lipoprotein. TG: triglyceride. LDL: low-density lipoprotein (n = 4). D GSEA of the ‘Fat digestion and absorption’ pathway based on ileum transcriptomes collected after 1 week in different groups. NES: normalized enrichment score. P-value estimation is based on an adaptive multi-level split Monte-Carlo scheme (FGSEA method) with Benjamini/Hochberg correction. E Volcano plot highlighting different genes belonging to the ‘Fat digestion and absorption’ pathway. Only genes in this pathway with |log2 fold change| > 2 and P-value < 0.05 are marked with color, DESeq2 method. F Venn diagram showing the number of unique and overlapping genes between the different gene sets belonging to the ‘Fat digestion and absorption’ pathway. The left circle represents CH4 vs. Control. The right circle represents CH4+MOB vs. CH4. And the qPCR result of the overlapping gene (Fabp1) is shown (n = 3). G Experimental design for the two-week diet intervention. Mice were subjected to a high-fat and high-sugar treatment by oral gavage of 0.4 ml lard per mouse per day and ad libitum access to high-concentration sucrose water for two weeks. H Representative eWAT in different groups. I The adipose tissue weight from different depots (n = 4). J Cross sections of various adipose tissue depots stained with hematoxylin and eosin (n = 3). (WAT: white adipose tissue; BAT, brown adipose tissue; eWAT: epididymal WAT; ingWAT: inguinal WAT; mWAT: mesenteric WAT). Data are presented as means ± standard of mean in F and J, except for boxplots in B, C and E (center line or point, median; box limits, 25th and 75th percentile; whiskers, Tukey; points, outliers). Statistical analysis was performed with two-tailed Mann–Whitney–Wilcoxon test (B, C, I), and the two-tailed unpaired t-test with Benjamini/Hochberg correction (F, J). Individual data points are independent biological replicates unless otherwise stated. Source data are provided as a Source Data file.
To further investigate the molecular responses to methane and M. intestini intervention, we performed RNA sequencing of ileal tissue collected after one week of treatment. Principal component analysis revealed that CH₄-treated samples were clearly separated from the control group, whereas the CH₄ + MOB group shifted closer to the control group (Fig. S17). Gene set enrichment analysis revealed that methane upregulated pathways involved in intestinal fat metabolism, which were restored by the methanotrophs (Fig. 3D). Pla2g3 was the only gene that exhibited increased expression in the CH₄ + MOB group compared with the CH₄ group. In contrast, four genes that were significantly upregulated following methane exposure (Fabp1, Apoa4, Clps, and Cd36) showed a consistent trend of downregulation after additional MOB intervention, among which Fabp1 displayed a statistically significant decrease (Fig. 3E, F). Quantitative PCR confirmed the downregulation of Fabp1, a gene associated with obesity (Fig. 3F)11. Similar blood indexes alteration was observed in a high-sugar, high-fat diet experiment (Fig. S18), where methane exposure increased the weight of epididymal white adipose tissue (eWAT) and mesenteric white adipose tissue (mWAT) (Fig. 3H, I). Methanotroph intervention reduced adipocyte size and mitigated methane-induced whitening of brown adipose tissue (Fig. 3J).
The widespread occurrence of gut methanotrophs
To better understand the occurrence of gut-associated methane-oxidizing bacteria, we curated a dataset of 73,125 methane-oxidizing genes and constructed a representative database, complemented by a Non-MOGs database to minimize false positives (Fig. 4A). Analysis of 550 metagenomic fecal samples from five continents revealed methane-oxidizing genes in 91.64% of samples, with over 60% showing more than 10 reads (Fig. 4B). A significant positive correlation was observed between methane-oxidizing gene reads and sequencing depth (Spearman’s Rho = 0.6569, P = 1.51 × 10−63, Fig. 4C), suggesting that adequate sequencing depth is crucial for detecting these bacteria (Fig. 4D). This may explain why intestinal methanotrophs have not been previously identified, as their detection requires higher sequencing depth.
A The establishment of Methane-oxidation genes (MOGs) and Non-MOGs databases. B Geographic distribution of samples with the fecal samples detected MOG. Each point indicates one sampling location. To visualization, overlapping points randomly increase latitude and longitude. C The correlation between the sequencing depth and detected MOG read count (mapped read). D The difference of the sequencing depth between detected MOG samples and Not-detected MOG samples (Detected n = 504, Not-detected n = 46). E The different gut MOB relative abundance among different cohorts (Health n = 500, Obesity n = 475, Constipation n = 232). F The correlation between the BMI and relative abundance of gut MOB. G The correlation between the fecal consistency and relative abundance of gut MOB. H ROC analysis of gut MOB relative abundance in cohorts. Solid lines represent ordinary least-squares linear regressions, and the error bands indicate the 95% confidence intervals (C, F, H). Statistical analysis was performed with two-tailed Spearman’s correlation analysis (C, F, G) and two-tailed Mann–Whitney–Wilcoxon test with Benjamini/Hochberg correction (D, E). Individual data points are independent biological replicates unless otherwise stated. Source data are provided as a Source Data file.
Validation of the roles of gut methanotroph in multi-cohorts
We analyzed fecal metagenomic data from healthy individuals (N = 500), obese patients (N = 475), and constipation disorder patients (N = 232). Methane-oxidizing bacteria were significantly less abundant in obese and constipated patients compared to healthy individuals (Fig. 4E). A negative correlation was observed between methanotroph abundance and BMI (Spearman’s Rho = −0.312, P = 6.98 × 10−20, Fig. 4F), while a positive correlation was found with stool consistency in constipated patients (Spearman’s Rho = 0.2659, P = 6.83 × 10−14, Fig. 4G). A classification model based on methanotroph abundance achieved high accuracy in distinguishing obesity and constipation from healthy controls (ROC curve areas of 84.71% and 83.61%, respectively, Fig. 4H). These results suggest that methanotroph abundance could serve as a diagnostic biomarker for methane-related disorders. The widespread presence of methane-oxidizing genes in human gut microbiomes across diverse populations highlights their potential global significance in maintaining gut health and metabolic balance.
Discussion
In this study, we isolated a human-derived methanotroph, M. intestini. Our results demonstrate that M. intestini harbors a complete set of methane-oxidizing genes and is capable of consuming methane both in vitro and in vivo. Administration of CH₄ in mice was associated with delayed intestinal transit and increased adiposity, whereas colonization with viable M. intestini partially reversed these phenotypes.
Methane has been implicated in slowing gastrointestinal motility, with prior studies linking its accumulation in the gut to decreased intestinal peristalsis and the onset of constipation10. Experimental evidence further indicates that methane can disrupt smooth muscle contractility and impair peristaltic activity both in vitro and in vivo12,13. Consistent with these findings, our results showed that introducing methane-oxidizing bacteria into mice significantly reduced intestinal methane levels and improved gut motility. Although a type of inactivated methylotroph has been reported to exert anti-inflammatory effects that may alleviate intestinal inflammation14, our supplemental experiments demonstrated that heat-killed methanotrophs failed to reverse methane-induced motility abnormalities. Moreover, no substantial intestinal tissue damage was observed following methane exposure, leading us to hypothesize that M. intestini primarily restores normal gastrointestinal function by counteracting the inhibitory effects of methane through its metabolic consumption.
Previous studies have suggested a link between reduced intestinal motility and obesity15,16. Excessive methane production in the gut has likewise been associated with metabolic disturbances and increased adiposity8,9. However, whether methane-induced alterations in intestinal motility contribute to these metabolic effects or whether methane exerts a direct influence remains unclear. In our study, methane exposure was accompanied by marked changes in fat metabolism, consistent with previous findings17. These changes were reversed upon the intervention of M. intestini. Although there are no reports on the direct mechanism of methane regulation of fat metabolism, our study found that after methane excess, genes related to the fat digestion and absorption pathway in the intestinal tissue were upregulated, including Fabp111, Cd3618, Apoa419 and Clps20. These genes have established roles in enterocyte lipid uptake, intracellular trafficking and lipoprotein assembly, and are regulated in part by PPAR-family nuclear receptors. Moreover, after intestinal colonization with M. intestini, methane levels in the body decreased and were accompanied by the downregulation of these genes. This reversal effect, to a certain extent, further emphasizes the potential function of methane. Here, M. intestini colonization also increased expression of Pla2g3, an enzyme capable of generating lysophospholipid signaling molecules that may influence intracellular trafficking responses21,22. We speculate that when fat absorption related pathways are perturbed, the intestinal epithelium upregulates Pla2g3 to hydrolyze membrane phospholipids, thereby releasing fatty acids or lysophospholipids as a means to maintain intracellular lipid supply or remodel membrane components in the short term.
In addition, a similar reversal effect after M. intestini intervention was also reflected in the gut microbial community structure in the lower part of the intestine. Gut microbiota dysbiosis has been linked to metabolic disorders23; however, it is unclear whether this methane-induced dysbiosis is an indirect or direct result of methane-induced constipation. A possible explanation for this might be that prolonged retention likely alters substrate availability within the gut, leading to the depletion of simple carbohydrates and favoring the expansion of Bacteroidota, a phylum that has a very broad metabolic potential24. This shift toward a Bacteroidota-enriched and altered diverse microbial community mirrors the compositional changes observed in individuals with chronic constipation25. Therefore, methane-induced slowing of intestinal transit may indirectly shape the microbial ecosystem, establishing a feedback loop between impaired motility and microbial imbalance. Collectively, these findings provide a potential mechanistic explanation for methane-associated metabolic alterations. M. intestini may mitigate methane-induced dysbiosis by regulating intestinal methane concentrations, thereby restoring microbial diversity in the distal gut. At the same time, it weakens the overactivation of host fat absorption-related PPAR signaling pathways, ultimately leading to the downregulation of fat metabolism genes, thereby achieving the regulation of host fat metabolism.
Methanotrophs were found to thrive in widely ecological niches where methane serves as a dedicated carbon source26, therefore, their presence in the human gut, which is colonized by methanogenic microbes27, is not unexpected. Our data indicate that metagenomic samples in which methane oxidation genes were undetectable often had lower sequencing depth, and that the relative abundance of gut methanotrophs is likely low, which helps explain why these organisms have long been overlooked within the complex gut microbial community. A major contributing factor is the insufficient sequencing depth to capture rare microbial populations28,29. Moreover, the genome of the human-derived methanotroph in our study showed only about 85% similarity with currently characterized members of the genus, suggesting a relatively unique genomic background that has not yet been represented in existing microbial genome databases. This genomic distinctiveness may further account for the historical underestimation of methanotrophs in the human gut.
In addition, methane oxidation by methanotrophs requires the presence of oxygen. Although the intestinal lumen is predominantly anaerobic, previous studies have demonstrated the existence of oxygen gradients along the mucosal surface and the intestinal axis, giving rise to localized microaerophilic niches, particularly in the proximal small intestine30. Our observation that methanotroph abundance was highest in the small intestine following gavage supports this possibility. An alternative explanation is that methanotrophs may acquire oxygen directly from host epithelial cells. Nevertheless, the precise source of oxygen that supports methane oxidation in the gut remains to be clarified.
While our findings suggest a potential link between gut methanotrophs, intestinal motility, and fat metabolism, the current study should be regarded as exploratory. The causal relationship between methanotrophic activity and the observed physiological outcomes remains to be established, and future work employing more direct experimental strategies will be necessary to confirm these mechanisms. In addition, the abundance of methanotrophs in the human gut appears to be relatively low, which may be as a limiting factor of their functional impact within the complex microbial community. Thus, further studies are warranted to clarify the extent to which gut methanotrophs contribute to host physiology and to determine the conditions under which their effects become biologically significant.
In conclusion, our study suggests the role of gut methanotrophs in mitigating methane-induced gastrointestinal dysfunction and regulating fat metabolism. These bacteria offer a natural mechanism for reducing methane levels in the gut, which in turn may improve motility and metabolic outcomes. Given the widespread presence of methanotrophs in the human gut, these microbes may represent an underexplored but crucial component of gut health. Future research will be necessary to fully understand the mechanisms by which methanotrophs influence gut function and metabolism and to explore their potential as therapeutic agents for methane-related disorders.
Methods
All research conducted for this manuscript complies with ethical regulations including approval by the Institutional Review Board (IRB) of School of Xiangya Basic Medical Sciences and Institutional Ethics Committee, Central South University (2021-KT75, CSU-2024-0315).
Sample collection and gut methanotroph culturing
Fecal samples were collected from 10 adult healthy volunteers, who signed informed consent forms and did not take any antibiotics for the preceding six months.
To isolate the gut microbes that can metabolize methane, the Nitrate Mineral Salts Medium (NMS1) medium (Table S1)31 was utilized and methane was used as the only carbon source. Briefly, we replaced 25% (v/v) of air with sterile methane in the headspace of serum bottles (with an interior volume of 200 mL). For further details regarding the culturing device, we refer to He et al.32. In detail, 20 g of each fecal sample were dissolved and mixed in the base substrate liquid to make a sample master mix, and 5 ml of sample master mix was inoculated into each culture bottle. Subsequently, these bottles were incubated at 30 °C at 180 rpm/min for two to three weeks. During this period, the culture was transferred about five times (approximately every 5–7 days by inoculating fresh NMS1 medium) to gradually eliminate non-methanotrophic microbes and enrich for intestinal methanotrophs.
After five subsequent serial dilutions in liquid medium, the enriched culture was then streaked onto NMS1 plates with 1% Agar (Sangon Biotech Co., Ltd) which were incubated at 30 °C in an atmosphere containing 25% methane in gas-tight cylinders for isolation of individual colonies. At this stage, we observed colonies with different morphologies, including potential methanotrophs and other methylotrophs such as Methylorubrum rhodesianum (M. rhodesianum which formed pink colonies capable of growing on methanol but not on methane as the sole carbon source). These organisms were initially difficult to separate purely based on colony morphology. To ensure successful isolation of true methanotrophs, a rigorous purification protocol was followed. Specifically, nearly all distinct colony types were picked and tested by re-inoculation into fresh NMS1 liquid medium under a methane atmosphere. Growth was monitored over 1–2 weeks. Only true methanotrophs could be further enriched under these strict conditions. After confirming growth in liquid medium, cultures derived from methanotroph colonies were subsequently re-streaked onto fresh NMS1 agar plates to obtain single colonies. This cycle of liquid enrichment followed by solid-phase isolation was repeated. After ~20 rounds of streaking and enrichment, we obtained a stable axenic culture. The purity of the isolate was confirmed by phase-contrast microscopy, by the inability to grow on complex LB medium (10 g/L tryptone, 5 g/L yeast extract, 10 g/L NaCl)33, and by sequencing of the 16S rRNA gene using 27F (5′-AGAGTTTGATCCTGGCTCAG-3′) and 1492R (5′-GGTTACCTTGTTACGACTT-3′) bacterial universal primers and the Sanger technology, which yielded a single sequence. The final isolation shared a uniform morphology, appearing as small, white, pinpoint colonies on NMS1 agar plates after extended incubation. All the consumables used were sterile, and all the above operations were carried out in the biosafety cabinet.
Whole genome sequencing
For short-read sequencing, 3 μg of genomic DNA was used for library construction. A paired-end library with insert sizes of 400 bp was prepared following the Illumina standard protocol. The library was sequenced using the Illumina NovaSeq 6000 platform (Shanghai Biozeron Biotechnology Co., Ltd, Shanghai, China).
The long-read sequencing was performed using the Pacific Biosciences Sequel IIe technology (PacBio). SMRTbell libraries were prepared according to the PacBio standard protocol. Samples were pooled into a single multiplexed library and the size was selected using Sage Sciences’ BluePippin, which uses the 0.75% DF Marker S1 High-Pass 6 kb–10 kb v3 run protocol and S1 marker. A size selection cutoff of 8000 (BPstart value) was used. The size-selected SMRTbell library was annealed and bound according to the SMRT Link Set Up and sequenced on a Sequel IIe.
All raw reads were trimmed and quality controlled by Fastp (version 0.23.4)34. Clean data obtained by the above quality control were used for further analysis. The Illumina data were used to evaluate the complexity of the genome and to correct the PacBio long reads. We used unicycler (version 0.4.8) (https://github.com/rrwick/Unicycler) to perform genome assembly with default parameters and received the optimal results of the assembly. The genome was circularized with Circlator (http://sanger-pathogens.github.io/circlator/).
Phylogenetic analysis
Taxonomic classification was determined by sequencing the 16S rRNA gene that was isolated using a bacterial genomic DNA extraction kit (Omega Bio-Tek, USA) according to the protocols recommended by the manufacturer. PCR amplification of 16S rRNA gene was performed using primers 27F and 1492R (same as above). The cycling conditions were 5 min at 95 °C, followed by 35 cycles of 45 s at 94 °C, 60 s at 56 °C, 90 s at 72 °C, and a final 10 min at 72 °C in a GeneAmp PCR system 9700 instrument (Perkin Elmer, Norwalk, CT, USA). The purification and sequencing of amplified products were conducted by Sangon Biotech Co., Ltd (Shanghai, China). The type strain sequence data were downloaded from BACDIVE35. Molecular evolutionary analysis and phylogenetic tree construction of the 16S rRNA gene sequence data were performed using MEGA 6.0 software (https://www.megasoftware.net).
In addition, we used the complete strain whole genome sequences after uploading into TYGS database (https://tygs.dsmz.de/)36. ANI analysis was carried out using FastANI (version 1.33)37.
Metagenomic sequencing and analyses
To study the microbial composition in the subculture samples and mice feces, the shotgun metagenomic sequencing was carried out. Total microbial genomic DNA was extracted using the QIAGEN DNeasy PowerWater Kit (14900-100-NF). The extracted microbial DNA was processed to construct metagenome shotgun sequencing libraries with insert sizes of 400 bp by using Illumina TruSeq Nano DNA LT Library Preparation Kit. Each library was sequenced by Illumina HiSeq X-ten platform (Illumina, USA) with PE150 strategy.
To ensure the data accuracy and obtain the clean reads, raw sequences were processed to remove low-quality (average quality<35 or contains more than 16 unclear bases) reads using Fastp34 (version 0.23.4) and FastUniq38 (version 1.1.0) to eliminate duplicates in paired short DNA sequence reads in a FASTQ format. The sequences from human were filtered out by mapping to the mouse reference genome (mm39) and human reference genome (hg38) using Bowtie239 (version 2.3.5) with sensitive mode. For identification of potential methanotrophs, the clean reads were assembled to contigs using Megahit40 (version 1.2.9) with default settings. Metabat241 was used to construct metagenome assembled genomes (MAGs) with checkm2 (version 1.0.1)42 to check quality, and only medium- or high- quality MAGs (completeness ≥ 70%; contamination ≤ 10%) were further considered. After annotating them using gtdb-tk (version 1.7.0)43, we extracted MAGs classified as Methylocystis. To check methane-oxidizing genes presence, these were annotated using bakta (version 1.9.2)44. For the metagenomic sequencing data of mice feces, we used CoverM (version 0.6.1) (https://github.com/wwood/CoverM) to estimate the relative abundance of gut methanotrophs by mapping reads to the complete genome of M. intestini.
Detection of pmoA gene and fluorescence in situ hybridization
According to metagenomic sequencing data, we adjusted the A189F/mb661r primer pair45 to Adj-pmoAF (5′-GGNGACTGGGANTTCTGG-3′) and Adj-pmoAR (5′-CCGGMRCAACGTCYTTACC-3′). Using this pair of adjusted primer, the pmoA gene was PCR amplified with conditions as follows: 5 min at 95 °C, followed by 30 cycles of denaturation at 95 °C for 15 s, 51.6 °C for 30 s, 72 °C for 60 s and 72 °C for 10 min. The products were visualized in 1% agarose gels and sequenced using Sanger technology.
FISH was applied to the mixed fecal samples. The samples were transferred to 50 ml NMS1 with 25% sterile methane for three days and pelleted at 10,000 rpm. Three types of pmoA probes (AAGGCGGAAGTTCACCCACCAGAA; GCCGTGATCACATAGGAGCCCGAC′; GCGCTCGACCATGCGGATGTATTC) were designed according to the sequenced pmoA and mixed with 58 μl hybridization buffer (20 mM Tris, 900 mM NaCl, 0.01% SDS, 10% dextran sulfate). Each probe was labeled with Cy3 at the 5′ site. The cells were washed with 0.9% (w/v) NaCl, transferred to microscope slides (5 μl per well). Cells were fixed on the slides using an in-situ hybridization fixative (Servicebio, G1113), and washed in PBS (pH 7.4) on a decolorization shaker for 3 times. Protease K (20 μg/ml) was added (40 °C) for 5 min. After this, the pre-hybridization solution was removed and slides moved to the probe-containing hybridization solution and incubated overnight. DAPI staining solution was added to the slides followed by incubation in the dark for 8 min, after which the anti-fluorescence quenching sealant agent was added to the slides, after washing. Sections were observed under a Nikon upright fluorescence microscope and images were collected.
Electron microscopy
The 2.5% glutaraldehyde fixative (Biosharp Cat. No BL911A) was added to the tube, and the precipitation was re-suspended in the fixative, then fixed at 4 °C for preservation and transportation. Cell preparation and freeze-etching were performed following the protocol described in detail in Rachel et al.46. The cuprum grids were observed under Transmission Electron Microscope (HITACHI, Ltd HT7800), and images were taken.
Description of Methylocystis intestini sp. nov
Methylocystis intestini (in.tes.ti'ni. N.L. gen. n. intestini, of the intestine, referring to the isolation source from human feces). Cells are gram-negative and short rods coccoid with a diameter of ~0.7 μm. Strain in this study (S13) is the type strain which deposited in Agricultural Culture Collection Center (Jiangsu, China) under accession No. of JSACC32502. Methane can be used as catabolic substrates. The strain was isolated from the healthy people feces in China. The complete circular genome size is 3.8 Mb, has a DNA G+C content of 65.26 mol%, two copy of the 16S rRNA gene, 23S rRNA gene and 5S rRNA gene, and 56 tRNAs.
Animals
All animal experiments were approved by the Institutional Animal Care and Use Committee. 5-week-old male Kunming, C57BL/6 mice and 8-month-old Sprague-Dawley rats (specific-pathogen-free, SPF) were used for the experimental procedures in this study. These animals were obtained from Hunan Sleek Jingda (Changsha, China) and were housed in cages with metal grid covers under sterile conditions, with a controlled environment (temperature: 25 ± 5 °C, humidity: 60–70%, and a 12-h light/dark cycle). They had ad libitum access to potable water and standard rodent chow (AIN-93G). Only male mice were used in this study. Sex consideration was based on practical feasibility issues of epididymal fat. All animal experiments were carried out in compliance with the National Institutes of Health Guide for the Care and Use of Laboratory Animals and approved by the Institutional Ethics Committee of Central South University (CSU-2024-0315).
Assessment of intestinal motility
To estimate fecal water content, mice were housed individually with ad libitum access to food and water. Feces were collected within 2 h, and the wet weight was recorded. After drying at 100 °C for 1 h, the dry weight was measured. Water content was then calculated as: (wet weight–dry weight)/wet weight of feces.
For the preparation of methylcellulose red dye (MRD), 0.2 g Methylcellulose and 0.6 g carmine were dissolved in 10 mL PBS. Afterwards, mice were placed in individual cages and gavaged with 0.2 mL of prepared MRD. Feces were collected and tested for the presence of red carmine dye by rubbing on white paper. The whole gut transit time was determined as the time elapsed from the initiation of gavage to the first excretion of red-colored feces, with a maximum time limit of 8 h47.
To estimate the small intestine transit rate, mice were provided with Indian ink (Biosharp Cat. No BL1540A) by gavage and sacrificed thirty minutes later. The gastrointestinal tract (from stomach to rectum) was taken out, straightened on a white waterproof board, and the length of the small intestine and the distance of Indian ink progression were measured. Small intestine transit rate was calculated by dividing the distance of Indian ink progression within the small intestine by the total length of the small intestine. All mice were fasted for 24 h prior to the experiment, with free access to water48.
In vitro ileal mobility
Animals were killed by cervical dislocation. The ileum was put into the Krebs solution at 37 °C49. For the details of this experiment we refer to Jahng et al.50. Changes in isometric tension were measured using MD3000-C force transducers (Zhenghua Biology Instrument Equipment Co., Ltd).
Briefly, we examined the effects of different gases on intestinal peristalsis rate by replacing the types of gases introduced. The bath was gassed with Control (20% oxygen and 80% nitrogen), CH4 (99.999% methane), or CH4+MOB (the top air collected from the 1 L sealed bottle which inoculating M. intestini with 200 ml NMS1 and 40 ml methane for one month) gases using peristaltic pump with a similar flow. The pH of the bath was constantly monitored and maintained. To ensure the stability of the results, the tension signals were collected after 20 min of equilibrium under different gas conditions.
Sample collection
Fecal samples were collected via the stress defecation method one day prior to mice sacrifice. Sterile cotton swabs were used to gently stimulate defecation by wiping the anal area, and the feces were collected in sterile tubes for storage at −80 °C. Blood samples were obtained during sacrifice after isoflurane anesthesia, via orbital vein puncture, followed by immediate euthanasia through cervical dislocation. Mice were fasted for 12 h before measuring blood indexes. The blood was immediately collected to estimate blood indexes using blood glucose meter (Voice+, Sinocare lnc, China) and blood lipid meter (PFS-30, URIT Co. Ltd, China). Animals were perfused with formaldehyde, and various adipose tissue depots (epididymal, inguinal, mesenteric and interscapular BAT) weighed, and stored in PBS. Samples were sent to the Servicebio Inc. paraffin embedding, slicing and hematoxylin and eosin (H&E) staining. The size of fat droplet was measured by ImageJ software.
Establishment of animal model for methane intervention
For the construction of abnormal intestinal peristalsis model caused by methane, we refer to Naiying Shen et al.51 methods, with slight modifications. To prepare Methane-rich saline (MRS), methane gas was dissolved in sterile PBS (pH 7.4) under a pressure of 0.4–0.6 MPa for 8 h. For CH4 group, MRS (0.2 ml) and methane gas (0.2 ml) were both administered to animals by oral gavage using large-caliber syringes to prevent gas from escaping. For CH4+MOB and CH4+M. rhodesianum group, 0.5 ml of M. intestini or M. rhodesianum suspension (OD600 > 1, washed with PBS) was additionally administered by oral gavage 4–5 h after methane intervention. For CH4+ heat-killed MOB group, the live 0.5 ml M. intestini was replaced by the same bacteria after PBS wash but sterilized by autoclaving at 121 °C for 20 min. For Control group, we also administered 0.2 ml PBS and 0.2 ml air by oral gavage. These interventions were administered twice daily until the end of the experiment. After one week of adaptive feeding (at 6-week-old), the KM and C57BL/6 mice began the experiment intervention.
For diet intervention, the high-level fat was provided with 0.4 ml lard (Yuanye Bio-Technology Co., Ltd, S26610) by gavage per mouse once daily, and the high-level sugar was provided with 60% (w/v) sucrose (Macklin Ltd, S818946) by ad libitum drinking.
Bacterial quantification by Real-time PCR
To quantify the abundance of M. intestini, its 16S rRNA gene fragment was PCR-amplified and inserted into the pET-28a (+) vector to generate a standard template. Genomic DNA from samples, together with a series of diluted plasmid standards, served as templates for qPCR analysis. The absolute copy number was then determined and expressed as 16S rRNA gene copies per 1 μg of total DNA or per gram of fecal material.
Measurement of methane concentration
Methane concentration was measured using a mid-infrared methane analyzer (HA300-CH4, HORASEN Co., Ltd), which is based on the principle that methane exhibits strong absorption of infrared light at specific wavelengths. The analyzer determines methane concentration by measuring the attenuation of laser intensity after transmission through the gas sample. For in vitro measurements of headspace methane in bacterial cultures, 50 ml of headspace gas was withdrawn from the culture bottle using a 50 ml gas-tight syringe and immediately injected into the analyzer for methane quantification. For in vivo measurements of exhaled methane in mice, individual mice were placed in a sealed 200 ml chamber connected to a circulation system with a gas pump. To prevent CO₂ accumulation and potential CO₂ toxicity, the circulation system was equipped with a sufficient amount of soda lime to absorb CO₂. Thirty minutes after methane gavage, mice were transferred into the chamber, and exhaled gas was collected continuously for 45 min for methane concentration determination.
RNA-sequencing and Real-time PCR
Total RNA was extracted from the ileum tissue using TRIzol Reagent according to the manufacturer’s instructions (Invitrogen) and genomic DNA was removed using DNase I (TaKara). RNA quality was determined using 2100 Bioanalyser (Agilent) and quantified using the ND-2000 (NanoDrop Technologies). RNA-seq transcriptome libraries were prepared following TruSeqTM RNA sample preparation Kit from Illumina (San Diego, CA), using 1 μg of total RNA. Briefly, messenger RNA was isolated with polyA selection by oligo(dT) beads and fragmented using fragmentation buffer. cDNA synthesis, end repair, A-base addition and ligation of the NGS-indexed adaptors were performed. Libraries were then size-selected for cDNA target fragments of 200–300 bp in 2% Low Range Ultra Agarose followed by PCR amplification using Phusion DNA polymerase (NEB) for 15 PCR cycles. After quantification using the TBS380, Paired-end libraries were sequenced (BIOZERON Co., Ltd).
The raw sequence was quality-controlled by Fastp34 with default parameters. Then, clean reads were separately aligned to reference genome (GRCm39) with orientation mode using Hisat252 with default parameters. Gene Set Enrichment Analysis (GSEA) was performed using R package FGSEA53, and differential gene analysis was performed by DESeq254. For real-time PCR, the gut tissues frozen at −80 °C were mixed with Trizol reagent (SK1312; Sango, Shanghai, China) to extract the total RNA according to the manufacturer’s instructions. RNA concentration was calculated by measuring absorbance at 260 nm using a microplate reader and NanoDrop 2000 software. The RNA was reverse transcribed into cDNA by the AMV First Strand cDNA Synthesis kit (SK2445; Sango, Shanghai, China). The Real-time PCR was conducted using Takara SybrGreen PCR Master Mix (RR820A; Takara, Kyoto, Japan) on SLAN-96S Real-time PCR System (Hongshi Co., Ltd). Primers: (Fabp1, F: AGGAGTGCGAACTGGAGACCAT, R: GTCTCCATTGAGTTCAGTCACGG). The quantitative expression level of Fabp1-gene was analyzed by the ΔΔCT method and normalized to that of ACTB gene.
Construction of methane oxidation gene database and relative gene abundance
By searching methane keywords in NCBI, UNIPROT, IMG/M (https://img.jgi.doe.gov/) and KEGG, we downloaded all the gene sequences encoding subunits of the two known methane monooxygenases. The sequences included the pmo genes (pmoA, pmoB, pmoC) and mmo genes (mmoX, mmoY, mmoZ, mmoB, mmoC, mmoD). Both particulate methane monooxygenase (pMMO, encoded by pmoABC) and soluble methane monooxygenase (sMMO, encoded by mmoXYZBCD) catalyze the initial and rate-limiting step of methane oxidation, namely the conversion of methane (CH₄) to methanol (CH₃OH). The pmo complex is membrane-bound and more commonly found in type II methanotrophs, whereas mmo is cytoplasmic and expressed in some methanotrophs under copper-limited conditions. Next, we used mmseq255 (version 13.45111) based on clustering threshold >90% to cluster and select representative gene sequences, so as to construct the MOG database. Because the database only contains methane oxidation gene sequences, to reduce the false positive of the recognition sequence, we additionally constructed a pseudo-MOG database, which is based on Eggnogs (version 5.0, http://eggnog5.embl.de) to remove the sequences of methane oxidation-related genes.
For the raw data, we performed the same quality control pipeline (Method: Metagenomic sequencing and analyses; above). Next, we mapped clean reads to the databases by diamond (version 2.0.2)56, and only the record with e-value < 0.001 was reserved. For the pair-read samples, if the forward or reverse read can map to MOG database, but not to the pseudo-MOGs database, we considered that the read truly belonged to a methane oxidation gene. The relative abundance of methane-oxidizing bacteria in a sample equal to mapped read count per sample sequencing depth.
Analysis of public metagenomic datasets
To understand the distribution of methane-oxidizing bacteria in the intestinal tract in the global population, we collected the geographical locations of different metagenomic samples from the metadata in the UHGG database (http://ftp.ebi.ac.uk/pub/databases/metagenomics/mgnify_genomes/human-gut/) and randomly selected metagenomic samples belonging to different regions to download.
For the datasets with known health-related condition, 500 samples were randomly selected for analysis of obesity-related microbiomes, from three projects: PRJEB8094, PRJNA388263, PRJNA422434. As the data of Obesity group, 475 samples were retained after downloading quality controls (Project: PRJEB12357, PRJEB4336, PRJEB12123, PRJNA290729, PRJEB14215, PRJEB6997, PRJDB3601, PRJEB15371), and 232 samples were selected for the Constipation condition group (Project: PRJNA779475, PRJEB3792 and PRJNA612367).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequence data reported in this paper have been deposited in the Genome Sequence Archive in National Genomics Data Center, China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences, under accession number CRA025930 (RNA sequencing data), CRA025960 (Subculture sample metagenomic sequencing data), CRA025961 (Mice fecal sample metagenomic sequencing data) that are publicly accessible at https://bigd.big.ac.cn/gsa. The methane oxidation gene database generated in this study have been deposited in the zenodo (https://doi.org/10.5281/zenodo.15124190). The public metagenomic sequencing dataset in this study are stored in Source data file. Source data are provided with this paper.
References
Sobko, T. et al. Generation of NO by probiotic bacteria in the gastrointestinal tract. Free Radic. Biol. Med. 41, 985–991 (2006).
Kalantar-Zadeh, K., Berean, K. J., Burgell, R. E., Muir, J. G. & Gibson, P. R. Intestinal gases: influence on gut disorders and the role of dietary manipulations. Nat. Rev. Gastroenterol. Hepatol. 16, 733–747 (2019).
Triantafyllou, K., Chang, C. & Pimentel, M. Methanogens, methane and gastrointestinal motility. J. Neurogastroenterol. Motil. 20, 31 (2014).
Roccarina, D. et al. The role of methane in intestinal diseases. Off. J. Am. Coll. Gastroenterol. 105, 1250–1256 (2010).
Waqar, S. H. B. & Rehan, A. Methane and constipation-predominant irritable bowel syndrome: entwining pillars of emerging neurogastroenterology. Cureus 11, e4764 (2019).
Pimentel, M., Gunsalus, R. P., Rao, S. S. & Zhang, H. Methanogens in human health and disease. Am. J. Gastroenterol. 1, 28 (2012).
Mathur, R. et al. Intestinal Methanobrevibacter smithii but not total bacteria is related to diet-induced weight gain in rats. Obesity 21, 748–754 (2013).
Mathur, R. et al. Intestinal methane production is associated with decreased weight loss following bariatric surgery. Obes. Res. Clin. Pract. 10, 728–733 (2016).
Basseri, R. J. et al. Intestinal methane production in obese individuals is associated with a higher body mass index. Gastroenterol. Hepatol. 8, 22 (2012).
Kunkel, D. et al. Methane on breath testing is associated with constipation: a systematic review and meta-analysis. Dig. Dis. Sci. 56, 1612–1618 (2011).
You, H. et al. Serum FABP1 levels correlate positively with obesity in Chinese patients after laparoscopic sleeve gastrectomy: a 12-month follow-up study. Obes. Surg. 30, 931–940 (2020).
Pimentel, M. et al. Methane, a gas produced by enteric bacteria, slows intestinal transit and augments small intestinal contractile activity. Am. J. Physiol.-Gastrointest. Liver Physiol. 290, G1089–G1095 (2006).
Park, Y., Lee, Y., Hussain, Z., Lee, Y. & Park, H. The effects and mechanism of action of methane on ileal motor function. Neurogastroenterol. Motil. 29, e13077 (2017).
Jensen, B. A. et al. Lysates of Methylococcus capsulatus Bath induce a lean-like microbiota, intestinal FoxP3+ RORγt+ IL-17+ Tregs and improve metabolism. Nat. Commun. 12, 1093 (2021).
Pecora, P., Suraci, C., Antonelli, M., De Maria, S. & Marrocco, W. Constipation and obesity: a statistical analysis. J. Biol. Res.-Boll. Soc. Biol. Sper. 57, 2384–2388 (1981).
Xiang, N., Xu, L., Qian, H. & Zhang, D. Multiple obesity indices suggest a close relationship between obesity and constipation: evidence from NHANES. BMC Public Health 24, 1273 (2024).
Mathur, R. et al. Metabolic effects of eradicating breath methane using antibiotics in prediabetic subjects with obesity. Obesity 24, 576–582 (2016).
Love-Gregory, L. & Abumrad, N. A. CD36 genetics and the metabolic complications of obesity. Curr. Opin. Clin. Nutr. Metab. Care 14, 527–534 (2011).
Qu, J. et al. Low-density lipoprotein receptor-related protein 1 (LRP1) is a v receptor for apolipoprotein A4 (APOA4) in adipose tissue. Sci Rep. 11, 13289 (2021).
Bae, Y. J. et al. Time-course microarray analysis for identifying candidate genes involved in obesity-associated pathological changes in the mouse colon. Genes Nutr. 11, 30 (2016).
Starkl, P., Marichal, T. & Galli, S. J. PLA2G3 promotes mast cell maturation and function. Nat. Immunol. 14, 527–529 (2013).
Gijs, H. L. et al. Primary cilium suppression by SREBP1c involves distortion of vesicular trafficking by PLA2G3. Mol. Biol. Cell 26, 2321–2332 (2015).
Dao, M. C. & Clément, K. Gut microbiota and obesity: Concepts relevant to clinical care. Eur. J. Intern. Med. 48, 18–24 (2018).
Lapébie, P., Lombard, V., Drula, E., Terrapon, N. & Henrissat, B. Bacteroidetes use thousands of enzyme combinations to break down glycans. Nat. Commun. 10, 2043 (2019).
Wang, J. et al. Characteristics of the gut microbiome and serum metabolome in patients with functional constipation. Nutrients 15, 1779 (2023).
Nazaries, L. et al. Environmental drivers of the geographical distribution of methanotrophs: Insights from a national survey. Soil Biol. Biochem. 127, 264–279 (2018).
Hoegenauer, C., Hammer, H. F., Mahnert, A. & Moissl-Eichinger, C. Methanogenic archaea in the human gastrointestinal tract. Nat. Rev. Gastroenterol. Hepatol. 19, 805–813 (2022).
Pereira-Marques, J. et al. Impact of host DNA and sequencing depth on the taxonomic resolution of whole metagenome sequencing for microbiome analysis. Front. Microbiol. 10, 1277 (2019).
Rodriguez-R, L. M. & Konstantinidis, K. T. Estimating coverage in metagenomic data sets and why it matters. ISME J. 8, 2349–2351 (2014).
Albenberg, L. et al. Correlation between intraluminal oxygen gradient and radial partitioning of intestinal microbiota. Gastroenterology 147, 1055–1063.e1058 (2014).
Dedysh, S. N. & Dunfield, P. F. Cultivation of methanotrophs. in Hydrocarbon and Lipid Microbiology Protocols: Isolation and Cultivation, 231–247 (Springer, 2014).
He, L. et al. A methanotrophic bacterium to enable methane removal for climate mitigation. Proc. Natl. Acad. Sci. USA 120, e2310046120 (2023).
Reuß, J. et al. Isolation of methanotrophic bacteria from termite gut. Microbiol. Res. 179, 29–37 (2015).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Reimer, L. C. et al. Bac Dive in 2022: the knowledge base for standardized bacterial and archaeal data. Nucleic Acids Res. 50, D741–D746 (2022).
Meier-Kolthoff, J. P. & Göker, M. TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat. Commun. 10, 2182 (2019).
Jain, C., Rodriguez-R, L. M., Phillippy, A. M., Konstantinidis, K. T. & Aluru, S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat. Commun. 9, 5114 (2018).
Xu, H. et al. FastUniq: a fast de novo duplicates removal tool for paired short reads. PLoS ONE 7, e52249 (2012).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, D., Liu, C. M., Luo, R., Sadakane, K. & Lam, T. W. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31, 1674–1676 (2015).
Kang, D. D. et al. MetaBAT 2: an adaptive binning algorithm for robust and efficient genome reconstruction from metagenome assemblies. PeerJ 7, e7359 (2019).
Chklovski, A., Parks, D. H., Woodcroft, B. J. & Tyson, G. W. CheckM2: a rapid, scalable and accurate tool for assessing microbial genome quality using machine learning. Nat. Methods 20, 1203–1212 (2023).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk: a toolkit to classify genomes with the Genome Taxonomy Database. Bioinformatics 36, 1925–1927 (2020).
Schwengers, O. et al. Bakta: rapid and standardized annotation of bacterial genomes via alignment-free sequence identification. Microb. Genomics 7, 000685 (2021).
Knief, C. Diversity and habitat preferences of cultivated and uncultivated aerobic methanotrophic bacteria evaluated based on pmoA as molecular marker. Front. Microbiol. 6, 1346 (2015).
Rachel, R. et al. Methods in Cell Biology, Vol. 96, 47–69 (Elsevier, 2010).
Dey, N. et al. Regulators of gut motility revealed by a gnotobiotic model of diet-microbiome interactions related to travel. Cell 163, 95–107 (2015).
Zhang, C. C. et al. A key genetic factor governing arabinan utilization in the gut microbiome alleviates constipation. Cell Host Microbe 31, https://doi.org/10.1016/j.chom.2023.10.011 (2023).
Bailey, L. E. & Ong, S. D. Krebs-Henseleit solution as a physiological buffer in perfused and superfused preparations. J. Pharmacol. Methods 1, 171–175 (1978).
Jahng, J., Jung, I., Choi, E., Conklin, J. & Park, H. The effects of methane and hydrogen gases produced by enteric bacteria on ileal motility and colonic transit time. Neurogastroenterol. Motil. 24, 185–e192 (2012).
Shen, N., Wang, Z., Wang, C., Zhang, J. & Liu, C. Methane alleviates inflammation and apoptosis of dextran sulfate sodium-induced inflammatory bowel diseases by inhibiting toll-like receptor 4 (TLR4)/Myeloid differentiation factor 88 (MyD88)/Nuclear translocation of nuclear factor-κb (NF-κB) and endoplasmic reticulum stress pathways in mice. Med. Sci. Monit. 26, e922248–922241 (2020).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Korotkevich, G. et al. Fast gene set enrichment analysis. Preprint at biorxiv https://www.biorxiv.org/content/10.1101/060012v3 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 1–21 (2014).
Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
Acknowledgements
This work was funded by the National Natural Science Foundation of China (32170071 to Z.Y. and 32300051 to J.H.), Natural Science Foundation of Hunan Province (2025JJ50101 to Z.Y. and 2023JJ30651 to J.H.) and the Innovation-Driven Research Programme of Central South University (No. 2023CXQD059 to Z.Y.). We express our gratitude to Dr. Andong Zha for the valuable assistance in the help of H&E staining. We thank Prof. Xin Yan for his valuable comments on the article. And thank Ms. Zhen Wang for the help in molecular experiments.
Author information
Authors and Affiliations
Contributions
Conceptualization and Supervision: Z.Y., J.H., L.C. Funding acquisition: Z.Y., J.H. Formal analysis and Visualization: Y.M.Z. Writing - original draft: Y.M.Z., H.C. Writing - review & editing: All authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ali Rezaie, and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhao, Y., Chen, H., Huang, J. et al. The gut methanotroph Methylocystis intestini modulates intestinal peristalsis and fat metabolism via reducing methane levels. Nat Commun 17, 2 (2026). https://doi.org/10.1038/s41467-025-66596-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66596-w