Abstract
Light, a ubiquitous environmental stimulus, shapes behavior and physiology across all domains of life. While photoreceptors are widespread in bacterial genomes, their functional roles and signal transduction mechanisms in non-photosynthetic bacteria remain understudied. Light represses biofilms and virulence factors through a bacteriophytochrome photoreceptor BphP and response regulator AlgB in the human pathogen Pseudomonas aeruginosa. Here, we used transposon mutagenesis screening to identify a conserved hypothetical microprotein, DimA, as the master activator of the photo-sensing cascade. Transcriptomics, luciferase reporter assays and physiological assays revealed that deletion of dimA abolishes light-dependent suppression of virulence factors and biofilms. Mechanistically, we demonstrated that DimA activates the site-I protease AlgW, triggering regulated intramembrane proteolysis of the anti-sigma factor MucA, liberating sigma factor AlgU (σ22), which promotes algB expression. We discovered a positive feedback loop where light-activated AlgB upregulates dimA expression, thereby amplifying the photosensory response. This work establishes DimA as a crucial activator of photo sensing and expands our understanding of bacterial adaptation to changing light environments.
Similar content being viewed by others
Introduction
Bacteria employ sophisticated molecular mechanisms to perceive and respond to environmental stimuli through specialized sensory apparatuses that are crucial for survival, pathogenesis, and ecological interactions, allowing bacteria to thrive in diverse and dynamic environments. Amongst primary sensing mechanisms are two-component signal transduction systems (TCS), comprising histidine kinases and response regulators, which detect various physicochemical parameters and initiate adaptive responses1,2,3. Environmental perception encompasses chemoreception via membrane-bound receptors that recognize specific molecules, mechanosensation, cell-to-cell chemical communication via quorum sensing and photoreception via photoreceptor proteins such as bacteriophytochromes4,5,6,7,8. Signal transduction cascades convert these environmental inputs into cellular responses through complex molecular networks. These include phosphorylation cascades, second messenger systems (particularly cyclic nucleotides like cAMP and c-di-GMP), and modulation of sigma factors and transcriptional regulators9,10,11,12,13. These pathways culminate in the activation of specific genetic programs, enabling bacteria to alter metabolism, virulence factor production, biofilm formation, and motility accordingly.
Light, a ubiquitous environmental stimulus, is detected by photoreceptors in all domains of life14. Particular photoreceptor photosensory domains are activated by specific wavelengths of light15, with bacteriophytochromes being the most abundant photoreceptors16,17. Although photoreceptors are abundant in bacterial genomes, the molecular underpinnings of photo-sensing signal transduction in non-photosynthetic bacteria remain poorly understood, often due to the lack of a known phenotypic output. We previously reported that the human pathogen Pseudomonas aeruginosa senses and responds to light to inhibit biofilm formation and virulence factor production18. The photo-sensing signaling cascade in P. aeruginosa is initiated by the activation of the biliverdin-containing bacteriophytochrome photoreceptor BphP by light. BphP is a histidine kinase that, in response to light, phosphorylates and activates the response regulator AlgB. Phospho-AlgB, in turn, represses the expression of biofilm and virulence genes. While this BphP-AlgB photo-sensing two-component system is conserved in diverse bacterial phyla, relatively little is known about the factors that control photo sensing. One negative regulator of photo sensing is the phosphatase KinB that dephosphorylates AlgB to dampen the photo-sensing response18,19. Previous studies on transcriptional regulation of bphP have implicated the stationary phase sigma factor RpoS20. The expression of algB is promoted by the alternative sigma factor σ22 (also known as AlgT/AlgU) and in turn, by regulated intramembrane proteolysis (RIP)21,22,23,24,25,26,27. RIP is a widely conserved signal transduction mechanism involving sequential protein cleavage by Site-1 (S1P) and Site-2 (S2P) proteases28,29,30,31,32. In P. aeruginosa, the anti-sigma factor MucA, the homolog of Escherichia coli RseA, is a signaling protein with a periplasmic domain, a single transmembrane helix and an N-terminal domain that binds and antagonizes AlgU33,34. MucA is sequentially cleaved via RIP by AlgW (S1P) and MucP (S2P). Following RIP, MucA is degraded by cytoplasmic ClpXP protease, and AlgU is released, leading to the activation of algB expression.
In this work, we performed genome-wide transposon mutagenesis screening to identify positive regulators of photo-sensing signaling in P. aeruginosa. We discovered that a previously uncharacterized microprotein, PA14_20480, that we name DimA (Diaphotos Initiator Microprotein of AlgB), positively regulated photo-sensing driven repression of biofilm formation and virulence factor production. DimA, a periplasmic protein, activated the site-I protease AlgW, triggering proteolysis of the anti-sigma factor MucA, and releasing the alternate sigma factor AlgU in turn to promote algB expression. We further identified a positive feedback loop whereby AlgB transcription factor drives dimA expression under far-red light, reinforcing the photo-sensing response.
Results
DimA promotes photo-sensing response
We previously discovered that P. aeruginosa responds to light via the BphP-AlgB two-component system that represses biofilm formation and virulence factor production in the presence of light (Fig. 1a, b)18. Specifically, far-red light inhibits the expression of the major biofilm matrix exopolysaccharide Pel in wildtype (WT) P. aeruginosa UCBPP-PA14 (hereafter called PA14) and virulence factors such as pyocyanin, hydrogen cyanide and lectin A18. To discover regulators of the BphP-AlgB photo-sensing pathway, we first generated a light responsive reporter system by engineering a chromosomal fusion of the promoter of the gene encoding lectin A to a promoter-less luciferase cassette (attB::PlecA’-‘luxCDABE, hereafter referred to as lecA-lux) (Supplementary Fig. 1). As expected, upon shining far-red light this reporter strain exhibits approximately ten-fold lower luminescence compared to dark condition (Fig. 1c). Next, we screened ~35,000 transposon insertion mutants for colonies exhibiting a “bright” phenotype, i.e., increased luminescence over the parent strain (Supplementary Fig. 1) and identified 17 genes that had multiple independent insertions (Supplementary Table 1). These can be broadly classified into three categories: (1) known photo-sensing components (bphP and algB), (2) known factors that control the transcription of algB (algU, clpX, clpP, algW, and mucP), and (3) hypothetical proteins. Category 1 hits serve as a direct photoreceptor-response regulator (BphP-AlgB) TCS that can detect and respond to specific wavelengths of light, while Category 2 factors (AlgU, ClpX, ClpP, AlgW, and MucP) form the RIP signaling cascade that controls algB expression (Fig. 1d and Supplementary Figs. 1, 2). Accordingly, Western blot analysis using custom raised antibodies against AlgB showed that AlgB protein levels were significantly reduced in the Category 2 transposon mutants compared to the single mutant of ΔkinB as well as WT PA14 (Supplementary Fig. 2b).
a Schematic of BphP-AlgB photo-sensing signaling cascade in P. aeruginosa. Light (denoted by “sun” symbol) stimulates the BphP kinase (green) to auto-phosphorylate and subsequently transfer the phosphoryl group to AlgB (brown) to activate AlgB. AlgB~P activates transcription of genes required for repression of group behaviors such as biofilm formation and virulence factor production. KinB (gray) acts as a phosphatase to antagonize AlgB. A “P” in a circle denotes addition or removal of a phosphate moiety. Arrow indicates activation, and T-bar indicates inhibition. b Colony biofilms under dark (denoted by “black crescent” symbol) and far-red (denoted by “red bulb” symbol) for WT, bphPSTOP, ΔalgB. Scale bar, 5 mm. Representative images shown of 3 independent experiments. c Quantification of lecA-lux activity in WT and mutant backgrounds under dark (black bars) and far-red light (red bars) conditions. d Quantification of lecA-lux activity for Tn5-candidates. e Quantification of lecA-lux activity in WT and mutant backgrounds under ambient light conditions. c–e Error bars represent SEM of three biological replicates (n = 3). Statistical significance was determined using two-way ANOVA for 1c and unpaired, two-tailed Welch’s t tests for pairwise comparisons for 1d-e in GraphPad Prism software (version 10.6.0). **** P < 0.0001, ** P < 0.005, * P < 0.05, ns not significant. Significant P-values were calculated as (left to right): For 1c 0.0010, for 1 d 0.0064, 0.0017, 0.0058, 0.0161, 0.0284, 0.0415, 0.0367, for 1e 0.0025, 0.0022, 0.0025, 0.0022.
Here, we focus on one transposon hit in a hypothetical gene, PA14_20840, that exhibited increased luminescence similar to those exhibited by the bphP::Tn and algB::Tn insertion mutants (Fig. 1d and Supplementary Fig. 1). The gene PA14_20480 encodes for a 92 amino acid microprotein (defined as proteins between 50 and 100 amino acids)35, that we refer to as DimA. To verify that DimA plays a role in lecA expression, we generated an in-frame marker-less deletion of dimA in the WT and ΔkinB strains bearing the lecA-lux reporter. Under ambient light conditions, the WT PA14, bphPSTOP and ΔkinB bphPSTOP strains exhibited a strong luminescence signal, while the absence of the negative regulator of photo-sensing KinB reduced lecA-lux reporter activity about 130-folds compared to WT (Fig. 1e). Consistent with the transposon mutant result, the ΔkinB ΔdimA double mutant was unable to abolish lecA expression, similar to the ΔkinB bphPSTOP double mutant (Fig. 1e), suggesting that DimA is an inhibitor of Lectin A production. Next, we compared lecA-lux reporter activity in WT and the ΔdimA mutant under far-red light. Whereas WT cells exhibited a four-fold reduction in lecA-lux reporter activity under far-red light, this repression was lost in the ΔdimA mutant (Fig. 2a). We conclude that DimA is required for far-red light–dependent suppression of lecA expression.
a Quantification of lecA-lux activity in WT and mutant ΔdimA under dark and far-red light conditions. Error bars represent SEM of five biological replicates (n = 5). b Pyocyanin production (OD695) was measured and normalized to growth (OD600) in WT PA14 and the designated mutants grown under ambient light conditions. Pyocyanin levels in WT PA14 were set to 100%. Error bars represent SEM of three biological replicates (n = 3). a, b Only comparisons that had a P-value < 0.05 are denoted. Statistical significance was determined using two-way ANOVA for 1a and unpaired, two-tailed t tests in GraphPad Prism software (version 10.6.0). * P < 0.05, *** P < 0.001, **** P < 0.0001. Significant P-values were calculated as (left to right): For 2a 0.0010, for 2b < 0.0001, 0.0003, 0.0004. c Colony biofilms under dark (denoted by “black crescent” symbol), ambient (denoted by “sun” symbol) and far-red (denoted by “red bulb” symbol) for WT and mutant strains. Scale bar, 5 mm. Representative images shown of three independent experiments. d Volcano plots of RNA-seq data for WT and ΔdimA in the presence or absence of far-red light. Dark gray solid circles represent genes that are significantly repressed by far-red light by > 2-fold while mustard yellow solid circles represent genes that are significantly activated by far-red light > 2-fold. Light gray solid circles represent genes with gene expression fold changes ≥ − 2 or ≤ 2 or Padj ≥ 0.05 (P-value adjusted for multiple comparisons using the Benjamini-Hochberg procedure). e Venn diagrams showing overlaps in genes that are differentially regulated in the presence and absence of light between WT and ΔdimA mutant. Numbers indicate the number of genes differentially regulated in each intersection.
DimA could function as a specific inhibitor of lecA expression or be a general photo-sensing regulator. To distinguish between these possibilities, we reasoned that a positive regulator of photo-sensing must repress other behaviors inhibited by light, such as pyocyanin production and biofilm formation. Accordingly, we measured the production of the phenazine pyocyanin36. Similar to the lecA-lux reporter, pyocyanin production was not repressed in WT or the ΔdimA single mutant under ambient light conditions (Fig. 2b). However, the ΔkinB mutant had 75% lower pyocyanin levels compared to WT, while the absence of DimA restored pyocyanin production (Fig. 2b). Complementation of the ΔkinB ΔdimA double mutant by introducing dimA under its native promoter in the attB chromosomal location reduced pyocyanin production by 72% (Fig. 2b), thereby demonstrating that DimA is a negative regulator of pyocyanin biosynthesis.
Along with repression of virulence factors, the AlgB-BphP photo-sensing system regulates biofilm formation (Fig. 1b)18. To determine whether DimA regulates light-mediated repression of biofilms, we assayed our strains for colony biofilm formation on Congo Red plates under light and dark conditions. In the absence of light, all the strains formed a rugose colony pattern indicative of biofilm formation (Fig. 2c). Exposure to ambient light, resulted in smooth colony pattern and repression of biofilm formation for the single mutant of ΔkinB (Fig. 2c). Absence of the photoreceptor BphP or DimA in the ΔkinB background showed de-repression of biofilms formation i.e., rugose colony phenotypes (Fig. 2c). The complemented strain ΔkinB ΔdimA attB::PdimA-dimA restored its ability to repress biofilms similar to the ΔkinB strain (Fig. 2c). Exposure to far-red light reduced colony rugosity in the WT when compared to dark condition but had little effect in the ΔdimA mutant. We conclude that DimA is a repressor of biofilm formation.
To define the extent of DimA’s role in photo sensing, we used RNA sequencing (RNA-seq) to compare the genome-wide transcriptional profiles of the WT and ΔdimA strains. We performed the experiment under dark and far-red light conditions. Principal component analysis (PCA) of normalized read counts showed that samples of dark conditions clustered separately from those of far-red light (Supplementary Fig. 3). Comparative transcriptomic analysis revealed a total of 553 differentially expressed genes (DEGs; genes with expression fold-changes ≤ − 2 and ≥ 2, and the P-values (Padj), adjusted using the Benjamini-Hochberg procedure, < 0.05) in the WT under far-red light compared to dark (Fig. 2d, e and Supplementary Data 6). These results demonstrate that far-red light drives large-scale transcriptional reprogramming in P. aeruginosa, controlling ~10% of the genome. In the ΔdimA mutant, we find a total of 133 DEGs, of which 101 transcripts overlapped with the light regulon in WT. Thus, the ΔdimA mutant showed a 76% reduction in the number of DEGs compared to WT (Fig. 2d, e). To further validate the RNA-seq data, transcriptional analyses of selected genes were performed using Quantitative Reverse Transcription PCR (qRT-PCR) (Supplementary Fig. 4). Specifically, PA14_08150, PA14_62680 and dimA were upregulated in WT under far-red light when compared to dark (Supplementary Fig. 4a–c), and mucE, PA14_20890, and PA14_60860 were downregulated in WT under far-red light compared to dark conditions (Supplementary Fig. 4d–f). Consistent with DimA’s role as an activator of photo-sensing response, regulation of these genes by far-red light was abolished in the absence of DimA (Supplementary Fig. 4). Thus, we conclude that DimA is a master activator of the photo-sensing response in P. aeruginosa.
DimA positively regulates AlgB transcription
One way in which DimA might promote photo sensing is by activating the expression of the photoreceptor BphP and/or the response regulator AlgB. Therefore, we determined the protein levels of BphP and AlgB in the WT and ΔdimA mutant strains. Western blot analysis using anti-FLAG antibody to probe for chromosomally encoded BphP-3xFLAG revealed that the protein levels of the photoreceptor BphP were not altered by the presence or absence of DimA (Fig. 3a, b). In contrast, Western blot analysis using custom raised antibodies against AlgB showed that the levels of the response regulator AlgB were significantly reduced in the ΔkinB ΔdimA double mutant compared to the single mutant of ΔkinB, as well as WT PA14 (Fig. 3a, c). Furthermore, ectopic complementation of dimA in the ΔkinB ΔdimA double mutant restored AlgB levels comparable to the ΔkinB mutant, indicating that DimA functions as an activator of AlgB protein (Fig. 3a, c). We conclude that while DimA did not alter BphP production, the absence of DimA significantly reduced AlgB protein levels.
a Western blot analysis of BphP-3xFLAG and AlgB protein levels. Overnight cultures of P. aeruginosa wildtype and the designated mutant strains were grown in triplicates in the presence of ambient light. Representative blot of one of the three biological replicates is shown, RpoB was used as a loading control. Molecular weight markers (kDa) are shown on the left. b, c Quantification of Western blots probing for BphP and AlgB. The values have been normalized to the RpoB protein as a loading control. n = 3 biological replicates. d Western blot analysis of AlgB protein levels. Overnight cultures of P. aeruginosa wildtype and the designated mutant strains were grown in three biological replicates under dark in the presence of 0.2% glucose (Glu) or 1% arabinose (Ara). Representative blot of one of the replicates is shown, RpoB was used as a loading control. Molecular weight markers (kDa) are shown on the left. (top panel). Quantification of Western blots probing for AlgB (bottom panel). The values have been normalized to the RpoB protein as a loading control and expressed relative to the WT sample, which was set to 1.0. n = 3 biological replicates. e Relative expression levels of algB measured by RT-qPCR in WT, ΔdimA, and WT harboring PBAD-driven dimA expression strain (attB::PBAD-dimA). WT and ΔdimA strains were grown in plain LB, while the complemented strain was grown under either repressing (0.2% glucose) or inducing (1% arabinose) conditions. All cultures were grown in the dark. Data were normalized to the geometric mean of rpsO and rpoD as internal reference genes and expressed relative to the glucose-grown sample of the same strain (set to 1.0). n = 3 biological replicates. f Pyocyanin production (OD695) was measured and normalized to growth (OD600) in WT PA14, ∆dimA, and the overexpression mutant grown overnight under ambient light conditions in the presence of 0.2% glucose or 1% arabinose, where indicated. Pyocyanin levels in WT PA14 were set to 100%. n = 3 biological replicates. b–f Error bars represent SEM of three biological replicates. Statistical significance was determined using unpaired, two-tailed Welch’s t tests for pairwise comparisons in GraphPad Prism software (version 10.6.0). * P < 0.05, *** P < 0.001. Significant P-values were calculated as (left to right): For 3c 0.0229, 0.0462, 3 d 0.0418, 0.0478, 0.0449, 3e 0.048, 0.0197, 3 f 0.0002. g Colony biofilm phenotypes of WT PA14 and the designated mutants on Congo red agar medium supplemented with carbenicillin and grown for 72 h. Strains were grown under dark (Top row, denoted by “black crescent” symbol) and far-red (Bottom row, denoted by “red bulb” symbol). Strains are harboring an empty vector of the pUCP18 plasmid (pEV) or are harboring a vector overexpressing AlgB protein (palgB). Scale bar is 5 mm for all images. Representative images shown of 3 independent experiments.
To determine whether the presence of DimA is sufficient to promote AlgB levels, we overexpressed DimA from an arabinose inducible promoter inserted at the attB locus in the PA14 genome (attB::PBAD-dimA), confirmed an increase in dimA transcript levels upon addition of arabinose via qRT-PCR (Supplementary Fig. 5) and examined AlgB protein abundance under dark conditions (Fig. 3d). Indeed, dimA overexpression increased AlgB protein levels (Fig. 3d). Next, to determine how AlgB protein levels were regulated by DimA, algB gene expression was measured using qRT-PCR. While deletion of dimA resulted in a two-fold decrease in algB transcript levels, overexpression of dimA led to a four-fold increase in the expression of algB under dark conditions and a concomitant decrease in pyocyanin production when the cells were exposed to light (Fig. 3e, f). We note that a similar result of dimA overexpression was obtained in a ΔdimA ΔkinB bphPSTOP triple mutant background, in which AlgB cannot be phosphorylated, indicating that DimA enhances AlgB accumulation independently of light and AlgB phosphorylation (Supplementary Fig. 5). We conclude that in the absence of DimA, AlgB protein levels were low due to a reduction in transcription of the algB gene.
A direct prediction of our model is that overexpression of algB should compensate for the absence of DimA. Accordingly, we overexpressed algB on a multicopy plasmid pUCP18 and tested biofilm formation in the presence and absence of light. Wildtype PA14 transformed with a vector containing algB under a constitutively active Plac promoter exhibited a reduction in biofilms when exposed to ambient light (Fig. 3g). Conversely, the biofilms were not repressed in the absence of the photoreceptor BphP even upon algB overexpression, as AlgB cannot get phosphorylated and therefore activated (Fig. 3g). However, overexpression of algB was sufficient to repress biofilms under light in the ΔdimA background, demonstrating that AlgB’s function is epistatic to DimA (Fig. 3g). Taken together, we conclude that DimA is an upstream positive regulator of AlgB.
The WTF C-terminal motif of DimA is essential for its function
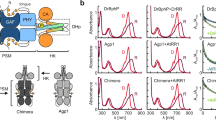
To gain insight into DimA’s mode of action, we questioned whether DimA is present in other bacteria. Examination of over ninety complete P. aeruginosa genomes identified DimA orthologues in both clinical and environmental isolates (Fig. 4a and Supplementary Fig. 6). In addition, reciprocal BLAST identified DimA homologs in other Pseudomonads such as Pseudomonas syringae and Pseudomonas chlororaphis (Fig. 4a and Supplementary Fig. 6). The amino acid sequences of DimA orthologues are highly conserved (Supplementary Fig. 7a), suggesting that DimA has a conserved function. Next, we examined the predicted AlphaFold structure of the protein (Fig. 4b and Supplementary Fig. 7b). Although the structure showed a very low average of per-residue model confidence score (pLDDT = 49.03), it was predicted to have an N-terminal α-helix that could function as a signal peptide for the Sec secretion system (Fig. 4c).
a Phylogenetic distribution and presence/absence of select genes across Pseudomonas spp. Rings surrounding the tree depict metadata and gene presence/absence across genomes − 4 innermost rings: presence (colored) or absence (gray) of four regulatory genes: dimA (purple), algB (brown), bphP (teal), and algW (orange), as determined by reciprocal BLAST against the P. aeruginosa PA14 genome. Outer ring: classification of P. aeruginosa strains as clinical (red) or environmental (gray). Colored branches in the tree represent P. aeruginosa (black), other Pseudomonas spp. (blue), and outgroup genomes (gray). b AlphaFold2 structure prediction model for DimA (PA14_20480) protein. The N- and C- termini are shown by their respective arrows. The structure is colored according to the per-residue model confidence score (pLDDT). c Protter visualization plot of DimA amino acid sequence. The predicted signal peptide is colored red, and the rest of the sequence is outlined in purple. d Confocal microscopy analysis of DimA and its N-terminal deletion mutant in P. aeruginosa. The cells were grown overnight in LB medium supplemented with 100 μg/ml tetracycline and 1 mM IPTG. Top row: PA14 ΔdimA strain harboring the plasmid pME6032-dimA-mNeonGreen, bottom row: PA14 ΔdimA strain harboring the plasmid pME6032-‘dimA -mNeonGreen fusion under induced conditions. Left to right: Membrane stained with FMTM4-64 (red), DimA-mNeonGreen fusion protein (green), and an overlay of both channels. Scale bar is 3 μm for all images. Representative images shown of three independent experiments. e Measurement of alkaline phosphatase activity. Overnight cultures of P. aeruginosa WT and ΔkinB strains bearing the dimA-phoA translational fusion were normalized for their growth at OD600 = 1. The phosphatase activity was measured at OD405 by the production of a yellow-colored product of the pNPP substrate. f Different truncation mutants of DimA were assayed for pyocyanin production. g Alanine substitutions in the WTF motif of DimA were tested for pyocyanin production. f, g Pyocyanin production (OD695) was measured and normalized to growth (OD600) in WT PA14 and the designated mutants grown overnight under ambient light conditions. Pyocyanin levels in WT PA14 were set to 100%. e–g Error bars represent SEM of three biological replicates (n = 3). Only pairwise comparisons that had P-value < 0.05 are denoted. Statistical significance was determined using unpaired, two-tailed t tests for pairwise comparisons in GraphPad Prism software (version 10.6.0). **** P < 0.0001, *** P < 0.001, ** P < 0.005. Significant P-values were calculated as (left to right): For 4e 0.0066, < 0.0001, for 4 f 0.0007, < 0.0001, < 0.0001, for 4 g 0.0003, 0.0004, 0.0044, 0.0004, 0.0004.
To define the localization of DimA, we engineered a C-terminal translational fusion of DimA with mNeonGreen fluorescent protein under IPTG-inducible promoter using the plasmid pME6032. Upon IPTG induction, the mNeonGreen signal colocalized with FM™ 4-64, a red fluorescent membrane dye, indicative of membrane localization for the fluorescently tagged DimA protein (Fig. 4d). Deletion of the N-terminal predicted signal peptide (M1-D28) resulted in uniform cytoplasmic localization of mNeonGreen which highlights the importance of the N-terminal domain for the localization of DimA (Fig. 4d). Expression of the DimAΔM1-D28, hereafter called ‘DimA variant, from an ectopic locus (attB::PdimA-dimAΔM1-D28) in the dimA mutant failed to complement DimA function (Supplementary Fig. 8). To further verify that DimA’s functional domain is localized in the periplasm, we generated a C-terminal fusion with alkaline phosphatase (PhoA) that is active only when it is in the periplasm37. Both the WT and ΔkinB strains containing the DimA’-‘PhoA fusion were able to degrade p-nitrophenyl phosphate disodium salt hexahydrate (pNPP) substrate, showing activity of the PhoA enzyme and localization of the fusion in the periplasm (Fig. 4e). Combining experimental evidence with the prediction from SignalP-6.0, which identifies a signal peptide cleavage site between amino acid positions 27 and 28 with 97.2% probability, we infer that DimA localizes and functions in the periplasmic space38.
Next, to investigate how a periplasmic microprotein could affect levels of the cytoplasmic response regulator AlgB, we generated several truncated versions of dimA and expressed them from its native promoter at an ectopic locus (attB::PdimA-dimAM1-W51/dimAM1-Q61/dimAM1-T71/dimAM1-L81/dimAM1-A89/dimAM1-F92). We performed pyocyanin production assay for these DimA variants in the sensitized ΔkinB mutant background and found that only the full-length DimA protein was able to repress pyocyanin production and even deletion of the last three amino acids (DimA1-89 or ΔWTF) completely abolished DimA’s function (Fig. 4f). To define the amino acid specificity of the WTF motif, alanine substitutions were made in the WTF motif and pyocyanin production was measured again as a proxy for the protein’s functionality. Interestingly, the alanine substitutions showed that pyocyanin production was repressed only when Trp90 and Phe92 were kept intact, while Thr91 was dispensable for the function of DimA (Fig. 4g). Conclusively, these data demonstrate that the C-terminal aromatic amino acids Trp90 and Phe92 are necessary for DimA function.
DimA activates the RIP site-I protease AlgW
One way in which DimA might promote AlgB protein levels is via the RIP cascade that governs the availability of the alternate sigma factor AlgU. Furthermore, algW was hit in the same transposon screen that identified DimA (Fig. 1d and Supplementary Figs. 1, 2). To test the idea that DimA could trigger AlgW-mediated proteolysis of MucA, we first generated a markerless deletion of algW in the WT and ΔkinB mutant backgrounds and examined whether the absence of AlgW showed similar physiological responses as the deletion of dimA. Indeed, repression of pyocyanin production (Fig. 5a) and biofilm formation (Fig. 5b) under ambient light were abolished in the ΔkinBΔalgW mutant, mirroring the effects observed in the ΔkinBΔdimA mutant. These findings are consistent with AlgW being a critical interacting partner of DimA, mediating light-dependent repression of pyocyanin production and biofilm formation in P. aeruginosa. Next, to test our hypothesis further, we employed the AlphaFold3 model prediction and found that DimA is predicted to bind through the WTF motif to the PDZ domain of AlgW. The C-terminal WTF motif of DimA closely resembles the WVF motif found at the C-terminus of the well-characterized envelope-stress activator MucE, which binds to the PDZ domain of AlgW to trigger proteolysis of MucA39 (Supplementary Fig. 9). This structural similarity suggests that DimA exploits a similar recognition mechanism: its exposed C-terminal tryptophan–phenylalanine (W90 and F92) residues engage the PDZ pocket of AlgW, likely leading to catalytic activation of the protease domain (Fig. 5c and Supplementary Figs. 9–11).
a AlgW mutant was assayed for pyocyanin production. Pyocyanin production (OD695) was measured and normalized to growth (OD600) in WT PA14 and the designated mutants grown overnight under ambient light conditions. Pyocyanin levels in WT PA14 were set to 100%. Error bars represent SEM of three biological replicates. Only pairwise comparisons that had P-value < 0.05 are denoted. Statistical significance was determined using unpaired, two-tailed Welch’s t tests for pairwise comparisons in GraphPad Prism software (version 10.6.0). ** P < 0.005, * P < 0.01. Significant P-values were calculated as (left to right): 0.0012, 0.0379, 0.0049, 0.0047. b Colony biofilm phenotypes of WT PA14 and the designated mutants on Congo red agar medium. The strains were grown for 72 h under dark (top row - denoted by the “black crescent” symbol) or ambient light conditions (bottom row – denoted by the “sun” symbol). Top-down view of stereoscope images. Scale bar is 5 mm for all images. Representative image from one of the three biological replicates is shown. c Alphafold3 multimer prediction model of AlgW periplasmic domain (colored in yellow) and DimA (colored in purple) structures. The interaction between the DimA WTF motif and AlgW PDZ domain has been boxed, and the key residues involved in the interaction have been enlarged and displayed as sticks (left panel). V286 and M342 residues of AlgW are colored orange and have been mutated to Ala in the ‘AlgWbind variant. W90 and F92 residues of DimA are colored magenta and have been mutated to Ala in the SUMO-‘DimAATA variant. The catalytic center of AlgW has been boxed, and the key residues involved in catalysis have been enlarged and displayed as sticks (right panel). H123, D153, and S227 residues of AlgW are colored red and have been mutated to Ala in the ‘AlgWcat mutant. The model was generated using the full-length AlgW sequence, but only its periplasmic domain is shown here for clarity (see full structure in Supplementary Fig. 9a). The PDB file generated by AlphaFold3 is provided as Supplementary Data 1. d In vitro analysis of MBP-‘MucA proteolysis in the presence of ‘AlgW protease WT or its designated variants and the ‘DimA activator in WT or its ATA mutant variant. SDS-PAGE gel stained with Coomassie Brilliant Blue (top panel) and corresponding anti-MBP immunoblot (bottom panel). Representative image shown of 3 independent experiments. Molecular weight markers (kDa) are shown on the left. e Levels of ‘MucA proteolysis were determined by quantifying the intensity of cleaved and full-length MBP-‘MucA bands in the Western blots. The % full-length ‘MucA was calculated by the formula (full-length/(cleaved + full-length) x 100). Error bars represent SEM of three independent experiments (n = 3).
To determine if DimA activates AlgW directly, an in vitro proteolysis assay was conducted. Specifically, we generated an N-terminal fusion of maltose binding protein (MBP) with the periplasmic domain of MucA (MBP-‘MucA), an N-terminal fusion of 6xHis-SUMO tag to the periplasmic part (amino acids 28–92) of DimA (SUMO-‘DimA) and an N-terminal fusion of 6xHis tag to the periplasmic domain of AlgW (‘AlgW). Purified ‘AlgW, ‘MucA and ‘DimA proteins were incubated together over a period of time and proteolysis of MucA was monitored via Coomassie Brilliant Blue staining as well as Western blot using MBP-specific antibodies. Incubation of ‘MucA with only ‘AlgW did not result in any significant proteolysis of ‘MucA (Fig. 5d, e). In contrast, the addition of ‘DimA induced proteolysis of ‘MucA as the levels of uncleaved MBP-‘MucA decreased over time while the levels of cleaved ‘MucA increased concomitantly (Fig. 5d, e). To test the importance of the C-terminal WTF motif for activation of AlgW-mediated proteolysis, we purified a ‘DimA variant where the WTF motif was changed to ATA (SUMO-‘DimAATA). The purified ‘DimAATA variant failed to activate AlgW-dependent degradation of ‘MucA (Fig. 5d, e). Notably, in addition to the degradation of ‘MucA, we also observed degradation of ‘DimA over time in vitro. However, heat-inactivated ‘AlgW failed to degrade both ‘MucA and ‘DimA, showing that catalytically active AlgW is required for MucA and DimA proteolysis (Supplementary Fig. 12). To confirm that AlgW is a specific protease, we incubated ‘AlgW with bovine serum albumin (BSA) protein in the presence of ‘DimA and did not observe any degradation of BSA over time (Supplementary Fig. 12). Furthermore, MBP alone was not cleaved by AlgW, demonstrating that the cleavage observed was specific to the MucA moiety of MBP-‘MucA (Supplementary Fig. 13a). A Coomassie-stained gel of all purified proteins at 5 μM confirmed the high purity of the preparations used in these assays (Supplementary Fig. 13b).
To further substantiate these findings, we performed proteolysis assays using catalytic and binding-site mutants of AlgW. A catalytic triple mutant of ‘AlgW (H123A/D153A/S227A), denoted as ‘AlgWcat, was completely defective in MucA cleavage, in the presence of wild-type ‘DimA (Fig. 5d, e). Similarly, an AlgW variant carrying V286A and M342A substitutions, denoted as ‘AlgWbind, where residues predicted by structural modeling to interact with the DimA C-terminal WTF motif, also failed to cleave MucA (Fig. 5d, e). Titration experiments further revealed that increasing concentrations of wildtype ‘DimA but not the ‘DimAATA variant (5 μM, 10 μM, 20 μM) accelerated MucA cleavage by wildtype ‘AlgW in a dose-dependent manner (Supplementary Fig. 14). In contrast, increasing concentrations of wildtype ‘DimA failed to elicit any ‘MucA cleavage when ‘AlgW catalytic or binding mutant was used (Supplementary Fig. 15). The concentrations of ‘AlgW and ‘MucA were kept constant at 1 μM and 5 μM, respectively in all of these reactions. Taken together, these data demonstrate that DimA directly activates AlgW protease function through its C-terminal WTF motif.
DimA is a light-induced microprotein
The nucleotide sequences of the upstream promoter regions of dimA in P. aeruginosa strains are highly conserved (Supplementary Fig. 16), suggesting that the regulation of its expression is universal across P. aeruginosa isolates. We found a putative AlgB binding motif 40 and hypothesized that dimA expression is induced by light via AlgB (Fig. 6a and Supplementary Fig. 16). To monitor the expression of dimA, we generated a chromosomal transcriptional reporter by integrating the β-galactosidase gene, lacZ, downstream of dimA in the PA14 genome. By measuring β-galactosidase activity, we had an estimate for transcriptional levels of dimA. Exposure to far-red light achieved striking upregulation of dimA in the WT PA14 strain compared to the dark condition (Fig. 6b). Deletion of the response regulator algB or the photoreceptor bphP abrogated this induction, demonstrating their essential roles in dimA expression. In addition, the absence of the stress-induced alternative sigma factor algU did not affect the upregulation of dimA (Fig. 6a).
a Alignment of PdimAproximal wildtype and mutant version. The region scrambled in the mutant is highlighted within an orange box. b β-galactosidase assays of PdimA-lacZ transcriptional fusion for background genotypes indicated on the X-axis and grown in dark and far-red light conditions. Error bars represent SEM of three biological replicates (n = 3). Statistical significance was determined using unpaired, two-tailed Welch’s t tests for pairwise comparisons in GraphPad Prism software (version 10.6.0). *** P < 0.001, **** P < 0.0001. Only pairwise comparisons that had P-value < 0.05 are denoted; Significant P-values were calculated as (left to right): 0.000085, 0.001074. c Representative images of three independent experiments of electromobility shift assay of PdimA variants, with the probe used in each panel indicated on the left. Lane 1 (Ø) represents the no AlgB control. The wedge indicates increasing concentrations of AlgB. Concentrations of AlgB added to lanes 2-10 are 0.03125, 0.0625, 0.125, 0.25, 0.5, 1, 2, 4, and 6 µM, respectively. Open caret represents unbound probe, gray caret represents shift, and black caret represents super-shift. d Fraction of bound DNA plotted against AlgB concentration. Error bars represent SEM of three independent experiments (n = 3). e Model for DimA-dependent activation of photo-sensing. (Left) In the absence of light, BphP (green) is inactive, AlgB (brown) is not phosphorylated and photo-sensing signaling is in an “OFF” state. (Right) Light activates BphP-AlgB TCS, P~ AlgB in turn activates the expression of DimA (purple). DimA triggers regulated intramembrane proteolysis via the site-I protease AlgW (orange) to release the sigma factor AlgU (tan) that leads to increased AlgB levels, which in turn, upon phosphorylation by BphP, represses biofilms and virulence factors in the photo-sensing “ON” state. Arrow indicates activation, and T-bar indicates inhibition.
To test whether AlgB directly bound the promoter region of dimA, an electrophoretic mobility shift assay (EMSA) was performed. Two ~ 250 bp probes were generated: one spanning the gene-proximal (PdimA proximal) and the other the gene-distal (PdimA distal) promoter region of dimA. Each probe was incubated with purified AlgB protein across a concentration gradient. An initial shift in mobility was observed in the proximal probe in the presence of 0.5 µM AlgB, which was further enhanced with increasing AlgB concertation, while no shift was observed in the distal probe even at the highest concentration of AlgB (Fig. 6c, d). These data indicate that AlgB specifically bound to the proximal region, but not to the distal region. Sequence analysis of the promoter-proximal region revealed two sets of inverted repeat sequences (TGCANNNTGCA) separated by 20 base pairs, resembling the putative AlgB binding motif40 (Fig. 6a). To assess the importance of these repeats for AlgB binding, both repeats were scrambled (PdimAproximalmut) (Fig. 6a). Binding was completely abolished when both repeats were scrambled, indicating that this region is important for AlgB association with the PdimA promoter (Fig. 6c, d). We conclude that dimA is a direct target of AlgB, and the inverted repeats are crucial for binding of AlgB to PdimA in vitro. Taken together, these data establish that DimA is a photo-induced microprotein whose expression is upregulated under far-red light directly by the light-activated AlgB response regulator.
Discussion
In this study, we have identified and characterized DimA, a novel 92-amino acid microprotein that functions as a master activator of the photo-sensing response in P. aeruginosa. Our findings reveal a sophisticated regulatory mechanism through which P. aeruginosa senses and responds to light via a complex signaling cascade involving DimA, the BphP-AlgB two-component system, and the regulated intramembrane proteolysis (RIP) pathway. DimA serves as a critical intermediary in this photo-sensing pathway, amplifying the light-induced signal through a positive feedback mechanism. Positive feedback in transcriptional regulation creates efficiency by limiting production of inactive regulatory proteins that require activation (such as phosphorylation) to function41,42. We propose that the DimA-mediated regulatory mechanism prevents wasteful synthesis of AlgB protein in the absence of light (Fig. 6e) and reduces the risk of inappropriate activation by non-physiological partners, as reported in other bacterial systems like sporulation in Bacillus subtilis and virulence pathways in Salmonella enterica43,44.
The molecular mechanism by which DimA regulates AlgB involves the RIP pathway. Our structural and functional analyses revealed that DimA is a periplasmic microprotein with a C-terminal domain containing a critical WTF motif. This motif bears striking similarity to the WVF motif in MucE, a known activator of the site-I protease AlgW, which initiates the cleavage of MucA in the regulated intramembrane proteolysis pathway of P. aeruginosa45,46. Indeed, our AlphaFold3 modeling and experimental data strongly suggest that DimA interacts with AlgW through its WTF motif, activating the protease and triggering the RIP cascade that ultimately leads to the release of the alternative sigma factor AlgU and subsequent upregulation of algB. Therefore, our discovery that DimA activates the same proteolytic cascade but in response to light stimuli represents a novel integration of environmental sensing with this established RIP signaling pathway. Furthermore, our phylogenetic analyses identified DimA homologs in other Pseudomonads such as P. chlororaphis, P. syringae, and P. arsenicoxydans, and AlphaFold predictions suggest that these DimA homologs interact with the corresponding AlgW proteases (Fig. 4 and Supplementary Figs. 6, 11). We speculate that DimA plays a similar role in these bacteria that contain functional BphP-AlgB photo-sensing cascades8,18.
The identification of DimA adds to our growing understanding of the critical role of microproteins in bacterial signaling networks. Microproteins, typically encoded by small open reading frames (sORFs), have emerged as important regulatory elements in diverse cellular processes, including ion transport, oxidative phosphorylation, and stress signaling47,48. These small proteins, often comprising the “dark matter” of bacterial proteomes, remain largely uncharacterized despite being widely distributed across all domains of life49,50,51,52,53. Of the characterized microproteins, some function as allosteric regulators of larger proteins or protein complexes, modifying or fine-tuning their activities, while others serve as independent signaling molecules. Small proteins involved in the regulation of two-component systems include SafA and MgrB, that function as negative regulators of the histidine kinase PhoQ in the PhoQ/PhoP two-component system in Escherichia coli and Salmonella enterica, respectively54,55,56,57. DimA exemplifies the paradigm of microproteins as critical regulatory elements in several ways. First, like many functional microproteins, DimA can effectively interface between different cellular compartments - in this case, between the periplasmic and cytoplasmic environments. Second, DimA functions not through enzymatic activity of its own, but as an allosteric regulator by modulating the activity of a larger protein complex (the AlgW protease) through a specific protein-protein interaction via its C-terminal WTF motif. Third, DimA operates as a signal amplifier as it enhances an initial environmental signal (light) into a robust cellular response (changes in virulence and biofilm formation). In essence, DimA represents a perfect example of how microproteins, despite their small size, can serve as sophisticated molecular switches in complex bacterial signaling networks, connecting environmental sensing to coordinated cellular responses.
The high conservation of DimA across both clinical and environmental P. aeruginosa isolates underscores its fundamental role in P. aeruginosa physiology and environmental adaptation. As an environmental cue, light provides bacteria with critical information about their location, and recent studies have shown that growth in darkness promotes virulence-associated traits and enhanced pathogenic potential in P. aeruginosa58,59. From a clinical perspective, these findings open promising therapeutic avenues against P. aeruginosa, a significant opportunistic pathogen associated with serious infections in immunocompromised individuals and cystic fibrosis patients. Indeed, artificial overexpression of DimA homolog in the P. aeruginosa clinical isolate IHMA879472 led to increased resistance to human plasma60. Specifically, targeting the DimA-mediated photo-sensing pathway could offer novel strategies to disrupt biofilm formation and virulence factor production, two critical determinants of P. aeruginosa pathogenicity in clinical settings. This approach aligns with emerging interest in developing anti-virulence therapies that do not directly kill bacteria but rather interfere with their pathogenic capabilities, potentially reducing selective pressure for resistance development.
In sum, our study identifies DimA as a critical regulator of the photo-sensing response in P. aeruginosa and elucidates its mechanism of action through activation of the RIP pathway. This work opens several promising avenues for future research in this important pathogen. First, kinetic and structural studies to elucidate the precise molecular interactions between DimA and the periplasmic domain of AlgW should provide insights into the proteolysis of DimA, potentially guiding the development of peptide inhibitors or small molecules that could modulate this interaction. Second, exploring the potential crosstalk between the photo-sensing pathway and other environmental stress responses, such as oxidative stress or antibiotic exposure, could reveal integrated regulatory networks. Finally, since P. aeruginosa infections often involve polymicrobial communities, examining how DimA-mediated photo-sensing affects interspecies interactions could yield insights into complex microbial behaviors in clinical and environmental settings.
Methods
Strains and growth conditions
P. aeruginosa UCBPP-PA14 strain was grown in lysogeny broth (LB) (10 g tryptone, 5 g yeast extract, 5 g NaCl per L) and on LB plates fortified with 1.5% agar at 37 ˚C. When appropriate, antimicrobials were included at the following concentrations: 400 µg/mL carbenicillin, 50 µg/mL gentamycin, 100 μg/mL irgasan, 500 μg/mL trimethoprim, 50 µg/mL kanamycin, and 100 μg/mL tetracycline.
Strain construction
Strains and primers used in this study are listed in Supplementary Tables 2–4. To construct marker-less in-frame chromosomal deletions in P. aeruginosa, DNA fragments flanking the gene of interest were amplified, assembled by the Gibson method, and cloned into pEXG261. The resulting plasmids were used to transform Escherichia coli SM10λpir, and subsequently, mobilized into P. aeruginosa PA14 via biparental mating. Exconjugants were selected on LB containing gentamicin and irgasan, followed by recovery of deletion mutants on LB medium containing 5% sucrose. Candidate mutants were confirmed by PCR. The complementation strain for PA14_20480 and its variants were cloned into pUC18T-mini-Tn7T-Tp plasmid62. The resulting plasmids were transformed to Escherichia coli DH5α, and mobilized into P. aeruginosa PA14 via quad-parental mating using as helper strains DH5α/pTNS3 and DH5α/pRK2013 to facilitate Tn7 transposition at the attB site as described here62. Insertion of the Tn7 transposon region was confirmed by PCR followed by Sanger sequencing for verifying the truncated versions and point mutations of PA14_20480. Similarly, the construction of the luminescent lecA reporter strains was performed by quad-parental mating between PA14 strains, DH5α/pUC18T-mini-Tn7T-Tp-PlecA-luxCDABE, and the helper strains DH5α/pTNS3 and DH5α/pRK2013. For the construction of the arabinose-inducible overexpression strains, PA14_20480 was cloned into the pTJ1 plasmid63. The resulting plasmid was transformed to Escherichia coli DH5α, and mobilized into P. aeruginosa PA14 via quad-parental mating using as helper strains DH5α/pTNS3 and DH5α/pRK2013.
To construct the transcriptional reporter fusions of P20480ΩlacZ, ~ 1000 bp of DNA upstream of the stop codon of PA14_20480 and the DNA encoding the lacZ open-reading frame were amplified using P. aeruginosa PA14 genomic DNA and the plasmid pIT2 as templates, respectively. Next, a DNA fragment of ~1000 bp downstream of PA14_20480 was amplified from P. aeruginosa PA14 genomic DNA. The three DNA fragments were assembled by the Gibson method and cloned into pEXG2. The resulting plasmid was used to transform E. coli SM10λpir, and subsequently mobilized into P. aeruginosa PA14 WT and the ΔalgB, ΔalgU, bphPSTOP mutants via biparental mating as described above.
To construct the translational fusion of PA14_20480-PhoA, ~1000 bp of DNA upstream and downstream of PA14_20480, the DNA encoding the phoA open-reading frame, were amplified using P. aeruginosa PA14 genomic DNA and the plasmid pCM639 as templates, respectively64. The three DNA fragments were assembled by the Gibson method and cloned into pEXG2. The resulting plasmid was used to transform E. coli SM10λpir, and subsequently mobilized into P. aeruginosa PA14 WT.
For the construction of PA14_20480-mNeonGreen or PA14_20480Cterm-mNeonGreen the open-reading frame of PA14_20480 or PA14_20480Δaa2−28, and the mNeonGreen were amplified using P. aeruginosa PA14 genomic DNA and the plasmid pmNeonGreen-N1 as templates, respectively65. The two DNA fragments were assembled by the Gibson method and cloned into the pME6032 plasmid66.
Protein production constructs were generated by amplifying the algWΔaa1-32, mucAΔaa1-105, dimAΔaa1−28 coding regions and cloning them in pET28b (Addgene, Watertown, MA, USA), pMAL-c6T (New England Biolabs, Ipswich, MA, USA), and pTB14667 expression vectors to obtain pET28b-His6-‘AlgW, pMAL-MBP-‘MucA, and pTB146-SUMO-‘DimA, respectively. The pET28b-His6-‘AlgWcat and pET28b-His6-‘AlgWbind were constructed by incorporating specific point mutations at H123A, D153A, S227A for the ‘AlgWcat and V286A,M342A mutations for the ‘AlgWbind variant. The ‘AlgW variant plasmids were obtained by GenScript Biotech (NJ, USA).
Transposon mutagenesis screen
We generated a sensitized reporter strain by deleting the genes encoding the known negative regulator of photo-sensing, KinB and the biliverdin-producing heme oxygenases BphO and HemO68,69. This sensitized ΔkinBΔbphOΔhemO attB::PlecA’-‘luxCDABE strain was subjected to Mariner-based transposon mutagenesis using pBT20 vector70. Our rationale was that inactivation of a gene(s) encoding a component that promotes photo-sensing would sever the connection between light and repression of the lecA-lux reporter. Insertion mutants were selected on LB agar containing 100 µg/mL gentamycin, and 100 μg/mL irgasan was included in the agar to counter select against the E. coli donor. The plates were incubated at 25 °C for 48 hours in the presence of light and screened for colonies expressing increased luminescence as detected by an Amersham ImageQuant 800 (Cytiva) imager. Transposon insertion locations were determined by arbitrary PCR and Sanger sequencing71.
Luminescence reporter assay
PA14 strains harboring the chromosomally encoded attB::PlecA-luxCDABE were grown overnight at 37 °C in LB growth medium under either dark or far-red light (730 nm, 1 Watt/m2). The next day, 5 μL of the overnight cultures were used to inoculate 1 mL cultures and grew them at 37 °C in LB medium until they reached OD600 = 0.5. 10-fold serial dilutions were carried out and plated on LB agar plates. The plates were incubated in different light conditions for 48 h and then measured the chemiluminescence with 3 min exposure time using an Amersham ImageQuant 800 (Cytiva) imager. Quantification of the luminescent values was performed with Image Lab 6.1 software (Bio-Rad). The luminescent values were normalized to the culture growth from the CFU measurement of the serial dilutions. Source data are provided as a Source Data file.
Colony biofilm assay
One microliter of P. aeruginosa cultures (OD600 = 0.5) grown at 37 °C in LB broth was spotted onto 60 × 15 mm Petri plates containing 10 mL 1% Tryptone medium supplemented with 40 mg/L Congo red, 20 mg/L Coomassie brilliant blue dyes, and solidified with 1% agar. Colonies were grown at 25 °C under dark, ambient light (1 Watt/m2, 14 W LED Grow Light Lamp - Full Spectrum), or far-red light (730 nm, 25 Watt/m2, Single Color LED-Starboard SST-10) conditions, and images were acquired after 120 h using a Zeiss AxioZoom v16 microscope. Note that a higher intensity of far-red light was used, as we have empirically found that biofilm cells are less sensitive to light.
Pyocyanin assay
PA14 strains were grown overnight in LB liquid medium at 37 °C with shaking at 220 rotations per minute (rpm). The cells were pelleted by centrifugation at 15,000 × g for 2 min, and the clarified supernatants were passed through 0.22 μm filters (Millipore, Burlington, MA, USA) into clear plastic cuvettes. The OD695 of each sample was measured on a spectrophotometer (Genesys 20, ThermoScientific) and normalized to the culture cell density, which was determined by OD600. Source data are provided as a Source Data file.
Protein overexpression and purification
His6-AlgB
The pET28b-His6-AlgB vector was transformed into BL21(DE3), and the cultures were grown approximately to 0.8 OD600 in 1 L of LB supplemented with 50 μg/mL kanamycin at 37 °C with shaking at 220 rpm. Protein production was induced by the addition of 1 mM IPTG, and incubation of the culture for 3 h at 25 °C was followed. The cells were pelleted by centrifugation at 2200 x g for 20 min and resuspended in AlgB-lysis buffer (50 mM NaH2PO4 [pH 8.0], 300 mM NaCl, 1 mM MgCl2, 1 mM DTT, 5% glycerol, 0.1% Triton X-100, 10 mM imidazole, and 1 tablet of cOmplete™, Mini Protease Inhibitor Cocktail – Roche). The cells were lysed by sonication (20 s pulses for 30 min). The lysate was centrifuged at 20100 x g for 30 min at 4 °C. The resulting clarified supernatant was combined with Ni-NTA resin (Novagen) and incubated for 3 h at 4 °C. The bead/lysate mixture was loaded onto a 1.5 cm Econo-Column® (Bio-Rad, Hercules, CA, USA), the resin was allowed to pack, and then it was washed with AlgB-wash buffer (50 mM NaH2PO4 [pH 8.0], 300 mM NaCl, 1 mM MgCl2, 1 mM DTT, 5% glycerol, 0.1% Triton X-100, 30 mM imidazole, and protease inhibitor cocktail tablet) Resin-bound His6-AlgB was eluted twice with 2 column volumes (CV) AlgW-lysis buffer containing 250 mM imidazole. Fractions were analyzed by SDS-PAGE, and the gel was stained with Coomassie brilliant blue to assess His6-AlgB purity. Purified protein was dialyzed in AlgB-storage buffer (50 mM NaH2PO4 [pH 8.0], 300 mM NaCl, 1 mM MgCl2, 1 mM DTT, 5% glycerol, and 0.1% Triton X-100) and stored at − 80 °C18.
His6-‘AlgW and its variants
The pET28b-His6-AlgW, pET28b-His6-AlgWcat, pET28b-His6-AlgWbind vectors were transformed into BL21(DE3) and the cultures were grown approximately to 0.8 OD600 in 1 L of LB supplemented with 50 μg/mL kanamycin at 37 °C with shaking at 220 rpm. Protein production was induced by the addition of 1 mM IPTG, and incubation of the culture overnight at 37 °C was followed. The cells were pelleted by centrifugation at 2200 x g for 20 min and resuspended in AlgW-lysis buffer (25 mM Tris-HCl (pH 7.5), 150 mM NaCl, and 5% glycerol). The cells were lysed by sonication (20 s pulses for 30 min). The lysate was centrifuged at 20100 x g for 30 min at 4 °C. The resulting clarified supernatant was combined with Ni-NTA resin (Novagen) and incubated for 1 h at 4 °C. The bead/lysate mixture was loaded onto a 1.5 cm Econo-Column® (Bio-Rad, Hercules, CA, USA), the resin was allowed to pack, and then it was washed with AlgW-wash buffer (25 mM Tris-HCl (pH 7.5), 150 mM NaCl, and 5% glycerol, 25 mM imidazole) Resin-bound His6-AlgW was eluted twice with 3 column volumes (CV) AlgW-lysis buffer containing 300 mM imidazole. Fractions were analyzed by SDS-PAGE, and the gel was stained with Coomassie brilliant blue to assess His6-AlgW purity. Purified protein was dialyzed in degradation buffer (50 mM sodium phosphate (pH 7.4), 200 mM KCl, and 10% glycerol) and stored at − 80 °C.
His6-SUMO-‘DimA and His6-SUMO-‘DimAATA
The pTB146-SUMO-‘DimA and pTB146-SUMO-‘DimAATA vector were transformed into BL21(DE3), and the culture was grown approximately to 0.6 OD600 in 1 L of LB supplemented with 200 μg/mL carbenicillin at 37 °C with shaking at 220 rpm. Protein production was induced by the addition of 1 mM IPTG, and incubation of the culture overnight at 37 °C was followed. The cells were pelleted by centrifugation at 2200 x g for 20 min and resuspended in DimA-lysis buffer (50 mM Na2HPO4, 300 mM NaCl and 10 mM imidazole, 1 tablet of cOmplete™, Mini Protease Inhibitor Cocktail - Roche). The cells were lysed by sonication (20 s pulses for 30 min). The lysate was centrifuged at 20100 x g for 30 min at 4 °C. The resulting clarified supernatant was combined with Ni-NTA resin (Novagen) and incubated for 1 h at 4 °C. The bead/lysate mixture was loaded onto a 1.5cm Econo-Column® (Bio-Rad, Hercules, CA, USA), the resin was allowed to pack, and then it was washed with DimA-wash buffer (50 mM Na2HPO4, 300 mM NaCl and 30 mM imidazole). Resin-bound His6-SUMO-‘DimA was eluted with 2 CV fractions of DimA-lysis buffer containing 50 mM, 100 mM, 250 mM, and 500 mM imidazole. Fractions were analyzed by SDS-PAGE, and the gel was stained with Coomassie brilliant blue to assess SUMO-DimA purity. Purified protein was dialyzed in degradation buffer and stored at − 80 °C.
MBP-‘MucA and MBP alone
The pMAL-MBP and pMAL-MBP-‘MucA vectors were transformed into BL21(DE3), and the cultures were grown approximately to 0.5 OD600 in 1 L of LB supplemented with 200 μg/mL carbenicillin and 0.2% glucose at 37 °C with shaking at 220 rpm. Protein production was induced by the addition of 1 mM IPTG, and incubation of the culture overnight at 16 °C was followed. The cells were pelleted by centrifugation at 2200 x g for 20 min and resuspended in MBP-‘MucA lysis buffer (20 mM Tris-HCl, 200 mM NaCl, 1 mM EDTA, and 1 tablet of cOmplete™, Mini Protease Inhibitor Cocktail). The cells were lysed by sonication (20 s pulses for 30 min). The lysate was centrifuged at 20100 x g for 30 min at 4 °C. The resulting clarified supernatant was combined with amylose resin (New England Biolabs Inc.) and incubated for 1 h at 4 °C. The bead/lysate mixture was diluted 1:5 in MBP-‘MucA lysis buffer loaded onto a 1.5 cm Econo-Column® (Bio-Rad, Hercules, CA, USA), the resin was allowed to pack. Then, it was washed with 12 CV of MBP-‘MucA lysis buffer and eluted in 1 CV of MBP-‘MucA lysis buffer containing 10 mM maltose. Fractions were analyzed by SDS-PAGE, and the gel was stained with Coomassie brilliant blue to assess MBP-MucA purity. Purified protein was dialyzed in degradation buffer and stored at − 80 oC.
Western blot assays
PA14 strains were grown overnight at 37 °C in LB medium. 1 mL culture of cells was harvested and resuspended in 200 μL of 2 × SDS-PAGE Laemmli (Bio-Rad Laboratories, Inc., Hercules, CA, USA). The samples were heated for 10 min at 95 °C. 10 μL samples were loaded onto 10% SDS-PAGE gels and were subjected to electrophoresis. Proteins were transferred to nitrocellulose membranes and afterwards blocked with 5% skim milk in TBS at room temperature for 1 h. Following, an incubation with primary antibodies anti-FLAG (Sigma-Aldrich, REF: A8592, LOT: 5LCF0816), anti-AlgB (house-made by Cocalico Biologicals Inc., Denver, PA, USA), anti-RNAP (Abcam, Inc., Cambridge, UK, REF: AB191598, LOT: 1115804-4), and anti-MBP (Sigma-Aldrich, REF: A4213-1VL, LOT: 0000133447) at 1:7500 dilution in 1% skim milk in TBS was performed overnight at 4 °C on a rocking platform. Membranes were washed 3 times with TBS-Tween 20 at room temperature for 10 min on a rocking platform. Incubation with the secondary antibody anti-rabbit IgG-Peroxidase produced in goat (Sigma-Aldrich, REF: A0545-1ML) at a 1:7500 dilution in TBS was followed for 1 h at room temperature. The membranes were washed again for 3 times with TBS-Tween 20 for 10 min and subsequently developed with a SuperSignal West Femto Kit (Thermo Scientific) and captured with an Amersham ImageQuant 800 (Cytiva) imager. Quantification of the western blots was performed with Image Lab 6.1 software (Bio-Rad). Source data are provided as a Source Data file.
Fluorescence microscopy
PA14 strains bearing the pME6032-PA14_20480-mNeonGreen or the pME6032-PA14_20480Cterm-mNeonGreen plasmid were grown overnight at 37 °C in LB medium containing 100 μg/mL tetracycline and 1 mM IPTG. 1 mL of culture was pelleted and washed once with 1 mL PBS. The pellets were resuspended in 50 μL solution of FMTM4-64 dye (Invitrogen) dissolved in PBS at 10 μg/mL and incubated in the dark for 1 min. The samples were washed once with 1 mL PBS and then resuspended in 100 μL PBS final volume. The samples were loaded onto a microscope slide and covered with 1 mm poly-L-Lysine coated coverslip. Micrographs were acquired at 60x magnification with immersion oil in a Stellaris 5 (Leica) microscope. Image processing was performed in the Fiji platform (Fiji−2012).
Alkaline phosphatase Assay
PA14 strains with the PA14_20480-phoA translation fusion were grown overnight at 37 °C in LB medium. The cultures were diluted to OD600 = 1 and 1 mL of culture was pelleted by centrifugation at 15,000 x g for 1 min. The pellets were washed once with 1 ml Tris pH = 8 and then resuspended in 200 μL of substrate p-nitrophenyl phosphate disodium salt hexahydrate (pNPP) at 1 mg/mL (Sigma). The substrate was resuspended in 0.1 M glycine buffer (pH 10.4), with 1 mM MgCl2 and 1 mM ZnCl2. The samples were incubated in the dark at room temperature for 10 min. To stop the reaction 50 μL of 3 M NaOH were added. The optical density at 405 nm was measured using a Synergy Neo2 microplate reader (Agilent BioTek). Source data are provided as a Source Data file.
β-galactosidase assay
PA14 strains with the PA14_20480ΩlacZ transcriptional fusion were grown overnight at 37 °C in LB medium under either dark or far-red light (730 nm, 1 Watt/m2) conditions and 1 mL cultures were collected and pelleted. The pellets were resuspended in 1 mL Z-buffer with the addition of 200 μg lysozyme for permeabilization. Samples were then incubated at 30 °C for 15 min and diluted 1:2 in a total volume of 500 μL. The colorimetric reaction initiated with the addition of 100 μL of 4 mg/mL ONPG (ortho-Nitrophenyl-β-galactoside) dissolved in Z-Buffer to the samples, and time was recorded. The samples were incubated at 30 °C until sufficient color change was observed. The reaction was quenched by 250 μL 1 M Na2CO3. The optical density of each sample was then measured at 420 nm and 550 nm. Standard activity was calculated in Miller units by this formula: ((OD420 – (OD550*1.75))*1000)/ (time * volume lysate * OD600). Source data are provided as a Source Data file.
MucA proteolysis assay
All the added proteins had been dialyzed in the same degradation buffer (50 mM sodium phosphate (pH 7.4), 200 mM KCl, and 10% glycerol). 5 μM of MBP-‘MucA, 1 μM of ‘AlgW or ‘AlgWcat or ‘AlgWbind, and 5 μM of SUMO-‘DimA or SUMO-‘DimAATA were mixed, and the reaction was incubated at 37 °C. Samples were collected after stopping the reactions by adding 4 x Laemmli Sample Buffer (Bio-Rad Inc.) containing 100 mM DTT and heating the samples at 95 °C for 10 min. For heat inactivation of ‘AlgW and SUMO-‘DimA, the proteins were heated at 95 °C for 10 min and then added to the mixture.
The samples were run in SDS-PAGE, and the gel was stained with Coomassie brilliant blue to assess MBP-‘MucA and SUMO-‘DimA cleavage. MBP-‘MucA cleavage was also examined by performing Western blot analysis using α-MBP Peroxidase conjugated antibody (Sigma-Aldrich, REF: A4213-1VL, LOT: 0000133447) at a 1:7500 dilution and following the Western blot procedure described above. Quantification of the western blots was performed with Image Lab 6.1 software (Bio-Rad). Source data are provided as a Source Data file.
Electrophoretic mobility shift assay
Two DNA probes of approximately 250 bp regions spanning the dimA promoter regions were generated by PCR amplification using PA14 chromosomal DNA and the following primer pairs, PdimA distal (DM208/DM209) and PdimA proximal (DM210/DM211). Each of these was further amplified with primers JW71/DM209 and JW71/DM211, respectively, to generate fluorescent-tagged probes. For PdimA proximal mutant probe, a gBlocks™ (Integrated DNA Technologies, Inc.) was generated, AYgb3 in which both inverted repeats were scrambled. AYgb3 was further amplified using primers JW71/DM211 to generate a fluorescent tagged version. Fluorescently labeled DNA probes were stored in the dark at 4 °C until further use. AlgB was diluted to a series of concentrations (0.03125, 0.0625, 0.125, 0.25, 0.5, 1, 2, 4, 6 µM) in binding buffer (50 mM NaH2PO4 pH 8.0, 300 mM NaCl, 1 mM MgCl2, 1 mM DTT, 5% glycerol and 0.1% Triton X-100). A total of 10μL binding reactions were prepared, which included AlgB at various concentrations and fluorescent probes at a final concentration of 10 nM. Binding reactions were incubated at room temperature in the dark for 30 min. A 5.5% native polyacrylamide gel was prepared using 29:1 acrylamide/bisacrylamide (Bio-Rad), 1 × Tris-Glycine-EDTA buffer (25 mM Tris base, 250 mM glycine, 1 mM EDTA pH 8.0 and 5% glycerol). Glycerol was added to the binding reactions at a final concentration of 10% to facilitate loading, and 12 μL of each reaction mixture was fractionated on the 5.5% native gel in 1 × TGE running buffer at room temperature for 70 min at 100 V. Gels were imaged directly using a Typhoon FLA 9500 scanner with the Cy5 filter, Auto PMT, and a pixel size of 100 µM. Image Lab 6.1 (Biorad) was used to quantify the unbound DNA in each lane from at least three replicates. The amount of unbound DNA was normalized to the first lane of each gel, and to calculate the amount of bound DNA, unbound DNA was subtracted from 1. The fraction bound was plotted against AlgB concentration using GraphPad Prism software (version 10.6.0). Source data are provided as a Source Data file.
Quantitative real-time PCR (qPCR) analysis
For the RNA-Seq validation experiments, WT and ΔdimA strains were grown under the same conditions described in the RNA-Seq Sample Collection and Sequencing section. For dimA overexpression experiments, cultures were supplemented with 0.2% (w/v) glucose for repression or 1% (w/v) arabinose for induction of the PBAD-dimA promoter under dark conditions. Cultures were incubated for 16 h at 37 °C in triplicate, and 500 µL of cell culture was pelleted for RNA extraction. Pellets were resuspended in TRIzol, and total RNA was isolated using the Direct-zol RNA Miniprep Kit (Zymo Research) according to the manufacturer’s instructions. RNA (0.5 µg) was reverse-transcribed into cDNA using the PrimeScript RT Reagent Kit with gDNA Eraser (Takara Bio, San Jose, CA, USA). Quantitative real-time PCR (qPCR) was performed using the Applied Biosystems PowerTrack SYBR Green Master Mix on a Bio-Rad C1000 Touch Thermal Cycler.
qPCR data were analyzed using the comparative Cq (ΔΔCq) method as described by Livak and Schmittgen72. Cq values of target genes were normalized to the geometric mean of the reference genes rpoD and rpsO to obtain ΔCq values73. Relative expression levels were calculated as 2^−ΔΔCq, with expression in the indicated control condition (e.g., wildtype grown with glucose) set to 1.0. Source data are provided as a Source Data file.
Phylogenetic tree generation and annotation
Genome assemblies were downloaded from the NCBI Datasets database. A phylogenetic species tree was inferred following the general approach outlined by Parks et al.74. Specifically, 120 ubiquitous single-copy marker genes (the bac120 marker set) were identified based on profiles from the Pfam and TIGRFAMs databases. Protein sequences matching these profiles were extracted and aligned to their corresponding HMMs using HMMER v3.4. The resulting alignments were concatenated into a single multiple sequence alignment (MSA), and an unrooted maximum-likelihood tree was inferred using FastTree v2.1.11 using the LG amino acid substitution model. The resulting tree was manually rooted using the known outgroup Burkholderia ubonensis. Putative orthologs were identified by reciprocal BLAST. A protein was considered a putative ortholog if the best forward BLAST hit in a target genome reciprocally identified the original Pseudomonas aeruginosa UCBPP-PA14 query protein as its top hit in a reverse BLAST. Query genes used in the reciprocal BLAST included dimA (PA14_20480), bphP (PA14_10700), algB (PA14_72380), and algW (PA14_57760). Resulting data was used to annotate the tree using the interactive tree of life (iTOL). Code for generating the species tree is available at: https://github.com/JonWinkelman/bac120_tree_inference75.
RNA-Seq Sample collection and sequencing
P. aeruginosa WT and ΔdimA strains were grown in triplicate at 37 °C overnight in LB medium. The overnight cultures were subcultured into 1 mL of fresh LB medium and incubated for 12 h at room temperature under either dark conditions or far-red light (730 nm, 1 Watt/m2). Cells pellets were collected by centrifugation and flash-frozen in liquid nitrogen. The cells pellets were processed for library preparation and sequencing by the DFI Microbial Metagenomics Facility.
RNA-Seq Processing and differential expression analysis
Paired-end RNA sequencing reads were processed using a custom nf-core-compatible pipeline for prokaryotic transcriptomes, available at https://github.com/trestle-biosciences/rnaseq_prokaryotes75. All reads were aligned to the Pseudomonas aeruginosa UCBPP-PA14 reference genome (NCBI assembly accession: GCA_000014625.1). Briefly, the pipeline performed quality control using FastQC and summarized results using MultiQC. Adapter sequences and low-quality bases were trimmed using Trim Galore (which integrates Cutadapt and FastQC). Filtered reads were aligned to the reference genome with Bowtie2, and aligned reads were assigned to annotated gene features using featureCounts from the Subread package. Gene-level counts were then analyzed for differential expression using DESeq276. To compare the relative sizes of light-responsive regulons between the wildtype and ΔdimA backgrounds, differential expression analysis was conducted to identify genes whose transcript abundance changed in response to far-red light compared to dark conditions. Genes were included in downstream analysis if they exhibited a nominal P-value < 0.05 (adjusted using the Benjamini-Hochberg procedure) and exceeded a minimum fold-change threshold (as visualized in volcano plots). Plots were generated with Python’s plotly library and edited in adobe illustrator. A table of differentially expressed genes (DEGs) in the WT and ΔdimA under far-red light compared to dark is provided as Supplementary Data 6.
Protein structure prediction modeling
For the monomeric DimA protein structure, the PDB structure file was retrieved by AlphaFold Protein Structure Database (AF-A0A0H2ZE95-F1-model_v6, accessed April 2023). Protein complex structures were predicted using AlphaFold3, via the AlphaFold server77. The amino acid sequences of each protein were provided as input, and multimeric assemblies were generated using default parameters. Predicted structures were visualized and analyzed in PyMOL (Schrödinger, LLC). No further manual refinement of the models was performed.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All relevant data are provided within the manuscript and the supplementary information. The raw RNA sequencing data generated in this study has been deposited to the NCBI SRA under accession PRJNA1345097: Pseudomonas aeruginosa PA14 response to Far-red light exposure (https://www.ncbi.nlm.nih.gov/sra/PRJNA1345097). The AlphaFold3-predicted structures generated in this study are provided as Supplementary Data 1–5. Source data are provided in this paper.
Code availability
The code for generating the species tree is available at: https://github.com/JonWinkelman/bac120_tree_inference75. The code and the customized nf-core-compatible pipeline for prokaryotic transcriptomes, for processing the paired-end RNA sequencing reads is available at https://github.com/trestle-biosciences/rnaseq_prokaryotes75.
References
Groisman, E. A. & Mouslim, C. Sensing by bacterial regulatory systems in host and non-host environments. Nat. Rev. Microbiol. 4, 705–709 (2006).
Capra, E. J. & Laub, M. T. Evolution of two-component signal transduction systems. Annu. Rev. Microbiol. 66, 325–347 (2012).
Papon, N. & Stock, A. M. Two-component systems. Curr. Biol. 29, R724–R725 (2019).
Muok, A. R., Olsthoorn, F. A. & Briegel, A. Unpacking alternative features of the bacterial chemotaxis system. Annu. Rev. Microbiol 78, 169–189 (2024).
Persat, A., Inclan, Y. F., Engel, J. N., Stone, H. A. & Gitai, Z. Type IV pili mechanochemically regulate virulence factors in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 112, 7563–7568 (2015).
Kühn, M. J. et al. Mechanotaxis directs Pseudomonas aeruginosa twitching motility. Proc. Natl. Acad. Sci. USA 118, https://doi.org/10.1073/pnas.2101759118 (2021).
Mukherjee, S. & Bassler, B. L. Bacterial quorum sensing in complex and dynamically changing environments. Nat. Rev. Microbiol. 17, 371–382 (2019).
Burgie, E. S. et al. Signaling by a bacterial phytochrome histidine kinase involves a conformational cascade reorganizing the dimeric photoreceptor. Nat. Commun. 15, 6853 (2024).
Isenberg, R. Y. & Mandel, M. J. Cyclic diguanylate in the wild: roles during plant and animal colonization. Annu. Rev. Microbiol. 78, 533–551 (2024).
Dahlstrom, K. M. & O’Toole, G. A. A symphony of cyclases: specificity in diguanylate cyclase signaling. Annu Rev. Microbiol. 71, 179–195 (2017).
Barchinger, S. E. & Ades, S. E. Regulated proteolysis: control of the Escherichia coli σ(E)-dependent cell envelope stress response. Subcell. Biochem. 66, 129–160 (2013).
Österberg, S., del Peso-Santos, T. & Shingler, V. Regulation of alternative sigma factor use. Annu. Rev. Microbiol. 65, 37–55 (2011).
Kazmierczak, M. J., Wiedmann, M. & Boor, K. J. Alternative sigma factors and their roles in bacterial virulence. Microbiol. Mol. Biol. Rev. 69, 527–543 (2005).
Kottke, T., Xie, A., Larsen, D. S. & Hoff, W. D. Photoreceptors take charge: emerging principles for light sensing. Annu. Rev. Biophys. 47, 291–313 (2018).
Shcherbakova, D. M., Shemetov, A. A., Kaberniuk, A. A. & Verkhusha, V. V. Natural photoreceptors as a source of fluorescent proteins, biosensors, and optogenetic tools. Annu. Rev. Biochem. 84, 519–550 (2015).
Gourinchas, G., Etzl, S. & Winkler, A. Bacteriophytochromes - from informative model systems of phytochrome function to powerful tools in cell biology. Curr. Opin. Struct. Biol. 57, 72–83 (2019).
Gomelsky, M. & Hoff, W. D. Light helps bacteria make important lifestyle decisions. Trends Microbiol. 19, 441–448 (2011).
Mukherjee, S., Jemielita, M., Stergioula, V., Tikhonov, M. & Bassler, B. L. Photosensing and quorum sensing are integrated to control Pseudomonas aeruginosa collective behaviors. PLoS Biol. 17, e3000579 (2019).
Chand, N. S., Clatworthy, A. E. & Hung, D. T. The two-component sensor KinB acts as a phosphatase to regulate Pseudomonas aeruginosa Virulence. J. Bacteriol. 194, 6537–6547 (2012).
Barkovits, K., Harms, A., Benkartek, C., Smart, J. L. & Frankenberg-Dinkel, N. Expression of the phytochrome operon in Pseudomonas aeruginosa is dependent on the alternative sigma factor RpoS. FEMS Microbiol. Lett. 280, 160–168 (2008).
Wozniak, D. J. & Ohman, D. E. Transcriptional analysis of the Pseudomonas aeruginosa genes algR, algB, and algD reveals a hierarchy of alginate gene expression which is modulated by algT. J. Bacteriol. 176, 6007–6014 (1994).
Hershberger, C. D., Ye, R. W., Parsek, M. R., Xie, Z. D. & Chakrabarty, A. M. The algT (algU) gene of Pseudomonas aeruginosa, a key regulator involved in alginate biosynthesis, encodes an alternative sigma factor (sigma E). Proc. Natl. Acad. Sci. USA 92, 7941–7945 (1995).
Boucher, J. C. et al. Two distinct loci affecting conversion to mucoidy in Pseudomonas aeruginosa in cystic fibrosis encode homologs of the serine protease HtrA. J. Bacteriol. 178, 511–523 (1996).
Xie, Z. D., Hershberger, C. D., Shankar, S., Ye, R. W. & Chakrabarty, A. M. Sigma factor-anti-sigma factor interaction in alginate synthesis: inhibition of AlgT by MucA. J. Bacteriol. 178, 4990–4996 (1996).
Schurr, M. J., Yu, H., Martinez-Salazar, J. M., Boucher, J. C. & Deretic, V. Control of AlgU, a member of the sigma E-like family of stress sigma factors, by the negative regulators MucA and MucB and Pseudomonas aeruginosa conversion to mucoidy in cystic fibrosis. J. Bacteriol. 178, 4997–5004 (1996).
Mathee, K., McPherson, C. J. & Ohman, D. E. Posttranslational control of the algT (algU)-encoded sigma22 for expression of the alginate regulon in Pseudomonas aeruginosa and localization of its antagonist proteins MucA and MucB (AlgN). J. Bacteriol. 179, 3711–3720 (1997).
Sommerfield, A. G., Wang, M., Mamana, J. & Darwin, A. J. In vivo and in vitro analyses of the role of the Prc protease in inducing mucoidy in Pseudomonas aeruginosa. J. Bacteriol. 206, e0022224 (2024).
Yu, H., Schurr, M. J. & Deretic, V. Functional equivalence of Escherichia coli sigma E and Pseudomonas aeruginosa AlgU: E. coli rpoE restores mucoidy and reduces sensitivity to reactive oxygen intermediates in algU mutants of P. aeruginosa. J. Bacteriol. 177, 3259–3268 (1995).
Damron, F. H. & Goldberg, J. B. Proteolytic regulation of alginate overproduction in Pseudomonas aeruginosa. Mol. Microbiol. 84, 595–607 (2012).
Chevalier, S. et al. Cell Envelope Stress Response in Pseudomonas aeruginosa. Adv. Exp. Med. Biol. 1386, 147–184 (2022).
Rhodius, V. A., Suh, W. C., Nonaka, G., West, J. & Gross, C. A. Conserved and variable functions of the sigmaE stress response in related genomes. PLoS Biol. 4, e2 (2006).
Lou, X. et al. Structural insights into regulated intramembrane proteolysis by the positive alginate regulator MucP from Pseudomonas aeruginosa. Biochem. Biophys. Res. Commun. 740, 150999 (2024).
Schofield, M. C. et al. The anti-sigma factor MucA is required for viability in Pseudomonas aeruginosa. Mol. Microbiol. 116, 550–563 (2021).
Cross, A. R., Raghuram, V., Wang, Z., Dey, D. & Goldberg, J. B. Overproduction of the AlgT Sigma Factor Is Lethal to Mucoid Pseudomonas aeruginosa. J. Bacteriol. 202, https://doi.org/10.1128/JB.00445-20 (2020).
Hassel, K. R., Brito-Estrada, O. & Makarewich, C. A. Microproteins: Overlooked regulators of physiology and disease. iScience 26, 106781 (2023).
Abdelaziz, A. A., Kamer, A. M. A., Al-Monofy, K. B. & Al-Madboly, L. A. Pseudomonas aeruginosa’s greenish-blue pigment pyocyanin: its production and biological activities. Micro. Cell Fact. 22, 110 (2023).
Hoffman, C. S. & Wright, A. Fusions of secreted proteins to alkaline phosphatase: an approach for studying protein secretion. Proc. Natl. Acad. Sci. USA 82, 5107–5111 (1985).
Teufel, F. et al. SignalP 6.0 predicts all five types of signal peptides using protein language models. Nat. Biotechnol. 40, 1023–1025 (2022).
Li, T. et al. Molecular basis of the versatile regulatory mechanism of HtrA-type protease AlgW from Pseudomonas aeruginosa. mBio 12, https://doi.org/10.1128/mbio.03299-20 (2021).
Trouillon, J. et al. Determination of the two-component systems regulatory network reveals core and accessory regulations across Pseudomonas aeruginosa lineages. Nucleic Acids Res. 49, 11476–11490 (2021).
Brandman, O., Ferrell, J. E., Li, R. & Meyer, T. Interlinked fast and slow positive feedback loops drive reliable cell decisions. Science 310, 496–498 (2005).
Mitrophanov, A. Y. & Groisman, E. A. Positive feedback in cellular control systems. Bioessays 30, 542–555 (2008).
Strauch, M. A., Wu, J. J., Jonas, R. H. & Hoch, J. A. A positive feedback loop controls transcription of the spoOF gene, a component of the sporulation phosphorelay in Bacillus subtilis. Mol. Microbiol. 7, 967–974 (1993).
Shin, D., Lee, E.-J., Huang, H. & Groisman, E. A. A positive feedback loop promotes transcription surge that jump-starts Salmonella virulence circuit. Science 314, 1607–1609 (2006).
Qiu, D., Eisinger, V. M., Rowen, D. W. & Yu, H. D. Regulated proteolysis controls mucoid conversion in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 104, 8107–8112 (2007).
Cezairliyan, B. O. & Sauer, R. T. Control of Pseudomonas aeruginosa AlgW protease cleavage of MucA by peptide signals and MucB. Mol. Microbiol. 72, 368–379 (2009).
Hobbs, E. C., Yin, X., Paul, B. J., Astarita, J. L. & Storz, G. Conserved small protein associates with the multidrug efflux pump AcrB and differentially affects antibiotic resistance. Proc. Natl. Acad. Sci. USA 109, 16696–16701 (2012).
Burton, A. T., Zeinert, R. & Storz, G. Large Roles of Small Proteins. Annu. Rev. Microbiol. 78, 1–22 (2024).
Simoens, L., Fijalkowski, I. & Van Damme, P. Exposing the small protein load of bacterial life. FEMS Microbiol. Rev. 47, https://doi.org/10.1093/femsre/fuad063 (2023).
Aoyama, J. J. & Storz, G. Two for one: regulatory RNAs that encode small proteins. Trends Biochem. Sci. 48, 1035–1043 (2023).
Fijalkowski, I., Willems, P., Jonckheere, V., Simoens, L. & Van Damme, P. Hidden in plain sight: challenges in proteomics detection of small ORF-encoded polypeptides. microLife 3, uqac005 (2022).
Gray, T., Storz, G. & Papenfort, K. Small proteins; big questions. J. Bacteriol. 204, e0034121 (2022).
Fesenko, I. et al. The hidden bacterial microproteome. Mol. Cell 85, 1024–1041 (2025).
Lippa, A. M. & Goulian, M. Feedback inhibition in the PhoQ/PhoP signaling system by a membrane peptide. PLoS Genet. 5, e1000788 (2009).
Yadavalli, S. S. et al. Functional determinants of a small protein controlling a broadly conserved bacterial sensor kinase. J. Bacteriol. 202, https://doi.org/10.1128/jb.00305-20 (2020).
Lippa, A. M. & Goulian, M. Perturbation of the oxidizing environment of the periplasm stimulates the PhoQ/PhoP system in Escherichia coli. J. Bacteriol. 194, 1457–1463 (2012).
Eguchi, Y., Ishii, E., Yamane, M. & Utsumi, R. The connector SafA interacts with the multi-sensing domain of PhoQ in Escherichia coli. Mol. Microbiol. 85, 299–313 (2012).
Arana, N. et al. Light regulation in critical human pathogens of clinical relevance such as Acinetobacter baumannii, Staphylococcus aureus and Pseudomonas aeruginosa. Photochem. Photobio. Sci. 22, 2019–2036 (2023).
Mesquita, C. S., Ribeiro, A., Gomes, A. C. & Santos, P. M. Absence of light exposure increases pathogenicity of Pseudomonas aeruginosa pneumonia-associated clinical isolates. Biology 10, https://doi.org/10.3390/biology10090837 (2021).
Janet-Maitre, M. et al. Genome-wide screen in human plasma identifies multifaceted complement evasion of Pseudomonas aeruginosa. PLoS Pathog. 19, e1011023 (2023).
Hmelo, L. R. et al. Precision-engineering the Pseudomonas aeruginosa genome with two-step allelic exchange. Nat. Protoc. 10, 1820–1841 (2015).
Choi, K. H. & Schweizer, H. P. mini-Tn7 insertion in bacteria with single attTn7 sites: Example Pseudomonas aeruginosa. Nat. Protoc. 1, 153–161 (2006).
Damron, F. H., McKenney, E. S., Schweizer, H. P. & Goldberg, J. B. Construction of a broad-host-range Tn 7 -based vector for single-copy P BAD -controlled gene expression in gram-negative bacteria. Appl. Environ. Microbiol. 79, 718–721 (2013).
Jacobs, M. A. et al. Comprehensive transposon mutant library of Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 100, 14339–14344 (2003).
Shaner, N. C. et al. A bright monomeric green fluorescent protein derived from Branchiostoma lanceolatum. Nat. Methods 10, 407–409 (2013).
Laventie, B.-J. et al. A surface-induced asymmetric program promotes tissue colonization by Pseudomonas aeruginosa. Cell Host Microbe 25, 140–152 (2019).
Cozy, L. M. et al. SlrA/SinR/SlrR inhibits motility gene expression upstream of a hypersensitive and hysteretic switch at the level of σ D in Bacillus subtilis. Mol. Microbiol. 83, 1210–1228 (2012).
Wegele, R., Tasler, R., Zeng, Y., Rivera, M. & Frankenberg-Dinkel, N. The heme oxygenase(s)-phytochrome system of Pseudomonas aeruginosa. J. Biol. Chem. 279, 45791–45802 (2004).
Barker, K. D., Barkovits, K. & Wilks, A. Metabolic flux of extracellular heme uptake in Pseudomonas aeruginosa is driven by the iron-regulated heme oxygenase (HemO). J. Biol. Chem. 287, 18342–18350 (2012).
Kulasekara, H. D. et al. A novel two-component system controls the expression of Pseudomonas aeruginosa fimbrial cup genes. Mol. Microbiol. 55, 368–380 (2005).
Kulasekara, H. D. Transposon mutagenesis. Methods Mol. Biol. 1149, 501–519 (2014).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 3, research0034.1 (2002).
Parks, D. H. et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat. Biotechnol. 36, 996–1004 (2018).
Winkelman, J. JonWinkelman/Manias_et_al_2025: v0.0.1. Zenodo https://doi.org/10.5281/zenodo.17536010 (2025).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Acknowledgements
We thank the DFI Microbial Metagenomics Facility for help with RNA sequencing. We thank all members of the Mukherjee lab for thoughtful discussions. Research reported in this publication was supported by the National Institute of General Medical Sciences of the National Institutes of Health (NIH) under Grants R35GM150803 and R00GM129424 and the Searle Scholars Program Grant SSP−2022-104 to S.M. We are grateful to Margot and Robert Haselkorn and Virginia Aronson for their generous gift of equipment. The content of this study is solely the responsibility of the authors and does not necessarily represent the official views of the funding agencies. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
D.M. and S.M. were responsible for conceptualization, experimentation, data curation, investigation, methodology, validation, visualization, and writing the manuscript. A.M. and G.C. were responsible for experimentation, data curation, investigation, methodology, and visualization. J.D.W. was responsible for formal analysis, visualization and software. S.M. was responsible for funding acquisition, project administration, and supervision. All authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Mingxu Fang and the other anonymous reviewer for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Manias, D., Mishra, A., Winkelman, J.D. et al. A light-induced microprotein triggers regulated intramembrane proteolysis to promote photo-sensing in a pathogenic bacterium. Nat Commun 17, 182 (2026). https://doi.org/10.1038/s41467-025-66862-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66862-x