Abstract
Induction of ferroptosis is a potential strategy to eliminate chemotherapy-resistant acute myeloid leukemia (AML) cells. Here, we investigate the role and mechanism of thyroid hormone receptor-associated protein 3 (THRAP3) in ferroptosis of AML cells. We show that high expression of THRAP3 is correlated with a poor prognosis in AML patients. THRAP3 knockdown suppresses AML cell proliferation, and delays orthotopic and subcutaneous tumor growth in male mice; however, THRAP3 overexpression exerts the opposite roles. THRAP3 overexpression promotes resistance of AML cells to RSL3/erastin-induced ferroptosis via inhibiting iron accumulation and promoting GSH synthesis. Mechanistically, THRAP3 recruits SLU7 homolog, splicing factor (SLU7) to facilitate GIT ArfGAP 2 (GIT2) Exon14 skipping. Inhibition of GIT2 Exon14 skipping reverses THRAP3-induced ferroptosis resistance in vitro and in vivo. Altogether, THRAP3 contributes to ferroptosis resistance of AML cells via interaction with SLU7 to trigger GIT2 Exon14 skipping, which suggests THRAP3 to be a therapeutic target for AML.
Similar content being viewed by others
Introduction
Acute myeloid leukemia (AML) is a malignancy caused by blocked myeloid differentiation and consequent uncontrolled deposition of myeloid progenitor cells, which may result in bone marrow failure and death1. Age and molecular subtypes influence the prognosis and therapeutic effect of AML patients2. Although great improvements have been made in the identification of the molecular and phenotypes of AML, poor response to therapies or relapse remains an enormous challenge for AML patients3,4. Therefore, it is urgent to uncover the underlying pathogenesis of AML to develop effective therapies.
Ferroptosis is a type of cell death, resulted from excessive lipid peroxidation and subsequent membrane injury5. Ferroptosis induction possesses great potential in the treatment of cancers and other disorders6. Currently, triggering ferroptosis has been recognized as a therapeutic strategy for hematological malignancies, including AML7. Erastin, a ferroptosis inducer, has been demonstrated to enhance the anticancer efficiency of venetoclax in AML cells8. Notably, AML cells showed lower sensitivity to erastin as compared with other hematological tumor cells, such as diffuse large B cell lymphomas9. So far, the possible mechanisms of ferroptosis resistance of AML cells remain largely unknown.
Alternative splicing (AS) is a key post-transcriptional process that modulates splicing of precursor messenger RNAs (pre-mRNAs) at specific sites10. Abnormal AS is associated with the development of multiple hematologic tumors, including AML11. As reported, aberrant AS events were found in about 30% genes in AML cells in comparison with normal CD34+ cells12. Abnormal AS events have been considered as potential biomarkers for AML13,14. Recent studies have identified that AS events are closely associated with ferroptosis. For example, circRAPGEF5 reduced ferroptosis sensitivity of endometrial cancer cells via modulating AS of TFRC15. Another study showed that abnormal AS of PCLAF contributed to hepatitis B virus-induced sorafenib resistance of hepatocellular carcinoma cells through repressing ferroptosis16. However, the involvement of aberrant AS in ferroptosis resistance of AML cells has not been explored.
Thyroid hormone receptor-associated protein 3 (THRAP3), also known as TRAP150, is a vital constituent of the spliceosome17. THRAP3 expression has been recognized as a prognostic marker for many cancers18. Mounting evidence has proved the regulation of THRAP3 in pre-mRNA spicing. For example, Lee et al. found that THRAP3 played a crucial role in pre-mRNA splicing and nuclear mRNA degradation19. Our bioinformation analysis data showed that THRAP3 was abnormally up-regulated in recurrent AML, which was closely correlated with a lower survival of AML patients. In addition, BioGRID, MINT, and STING databases predicted a potential interaction between THRAP3 and splicing factors. A previous study documented that THRAP3 could interact with splicing factor PSF to block the binding of target genes to RNA recognition motifs of PSF, thereby inhibiting pre-mRNA splicing in T cells20. Therefore, we predicted that THRAP3 might regulate pre-mRNA splicing to drive AML progression, which deserves to be investigated.
Here, we show that up-regulation of THRAP3 facilitates growth of AML cells via reducing ferroptosis sensitivity by recruiting SLU7 homolog, splicing factor (SLU7) to trigger GIT ArfGAP 2 (GIT2) Exon14 skipping. Our findings offer mechanistic insight into the influence of THRAP3 on AS event of AML cells, and uncover its contribution to ferroptosis resistance in AML cells, suggesting THRAP3 as a potential therapeutic target for AML.
Results
Up-regulation of THRAP3 indicates a poor prognosis of AML patients
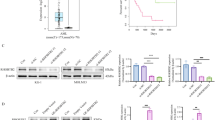
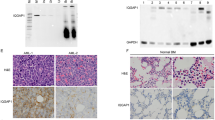
To validate the differential expression of THRAP3 in AML, expression signatures of 2128 primary AML patients and 416 recurrent AML patients from TARGET database were collected. We found that the expression of THRAP3 was significantly increased in relapsed AML patients compared with primary AML patients (Fig. 1A). In addition, we integrated the databases of TARGET, BEAT, and LEUCEGENE to analyze the differential expression of THRAP3 in AML. According to the results, THRAP3 was highly expressed in AML patients (Supplementary Fig. 1A, B). GEPIA database revealed that THRAP3 exhibited the highest level among 33 types of tumors (Supplementary Fig. 1C). As analyzed by TCGA database, high expression of THRAP3 was correlated with a lower survival rate of AML patients (Fig. 1B). Furthermore, RT-qPCR analysis indicated that THRAP3 level was higher in the BM samples of AML patients as compared with that of healthy volunteers (Fig. 1C). Notably, THRAP3 level was lower in complete remission (CR) group, but higher in relapse group in comparison with new diagnosis group of AML patients (Fig. 1D). Besides, high expression of THRAP3 was associated with a lower overall survival of AML patients (Fig. 1E). We also observed an elevated THRAP3 protein abundance in the BM samples of AML patients (Fig. 1F). Consistently, THRAP3 mRNA and protein levels were remarkably higher in AML cells than that in normal BMSCs, BMMNCs, and HS-5 cells (Fig. 1G, H). Taken together, THRAP3 was highly expressed in AML, which suggested a poor prognosis of AML patients.
A Up-regulation of THRAP3 was identified in relapsed AML patients (n = 416) and primary AML patients (n = 2128) from TARGET database. The box plots show the median (centre line), the 25th and 75th percentiles (box limits), and the minima and maxima. B TCGA database analyzed the correlation between THRAP3 expression and survival of AML patients (n = 132). C RT-qPCR analysis of THRAP3 expression in bone marrow (BM) of healthy volunteers and AML patients (normal group, n = 12; AML group, n = 82). D THRAP3 expression in AML patients from newly diagnosed (n = 82), complete remission (CR) (n = 42) and relapse groups (n = 16) was assessed by RT-qPCR. E Kaplan-Meier evaluated the correlation between THRAP3 expression and overall survival of AML patients (THRAP3 high expression, n = 41; THRAP3 low expression, n = 41). F Western blotting determined THRAP3 protein level in normal and AML samples (n = 12). N, normal BM samples; A, BM samples of AML patients. G, H RT-qPCR and Western blotting analysis of THRAP3 expression in normal BMSCs, BMMNCs, HS-5 and multiple AML cell lines (n = 3, biological replicates). The data are represented as mean ± standard deviation (SD). Experiments were repeated independently for 3 times with similar results, and representative image from one experiment is shown in (F, H). One-way ANOVA followed by Tukey’s post hoc test (for D, G, H), two-tailed Student’s t test (for A, C, F) and Log-rank test (for B, E) were performed for statistical analysis. *p < 0.05, **p < 0.01, ***p < 0.001. Source data are provided as a Source Data file.
THRAP3 overexpression accelerates AML tumorigenesis and reduces survival of mice
To explore the biological function of THRAP3 in AML tumorigenesis, THRAP3 was knocked out or overexpressed in THP-1 and HL-60 cells via transfection with sgTHRAP3 or THRAP3 overexpression plasmid, respectively. The silencing or overexpression efficiency of THRAP3 was validated by RT-qPCR and Western blotting in THP-1 or HL-60 cells, respectively (Supplementary Fig. 2A, B). Functional experiments revealed that THRAP3 knockdown restrained AML cell proliferation, whereas THRAP3 overexpression resulted in the opposite results (Fig. 2A). However, THRAP3 inhibition did not affect the proliferation and apoptosis of normal lymphocytes HS-5 cells, and THRAP3 silencing promoted THP-1 cell apoptosis (Supplementary Fig. 3). Moreover, we established an orthotopic AML xenograft model in mice. We found a reduction in mouse survival after THRAP3 overexpression (Fig. 2B). The liver and spleen weights of mice were increased by THRAP3 overexpression (Fig. 2C, D). Flow cytometry showed that THRAP3 overexpression obviously enhanced the percentage of CD45+Ki67+ in the liver and spleen tissues, suggesting the infiltration of AML cells into other organs (Fig. 2E). Collectively, these data supported that THRAP3 facilitated AML progression and reduced survival of mice.
THP-1 cells were transfected with sgTHRAP3 or sgNC; HL-60 cells were transfected with THRAP3 expression plasmid or vector. A CCK-8 evaluated AML cell proliferation. Orthotopic AML xenografts were generated by injection with HL-60 cells into tail veins of NOD/SCID mice. B Survival of mice was monitored. C, D livers and spleens were collected from mice and weighed. E The percentage of CD45+Ki67+ in livers and spleens was detected by flow cytometry. n = 3, biological replicates for A; n = 6 mice/group for (B–E). The data are represented as mean ± standard deviation (SD). Experiments were repeated independently for 6 times with similar results, and representative images from one experiment are shown in (E). Two-tailed Student’s t test (for A, D, E) and Log-rank test (for B) was performed for statistical analysis. **p < 0.01, ***p < 0.001. Source data are provided as a Source Data file.
THRAP3 affects the sensitivity of AML cells to ferroptosis
Ferroptosis induction has been identified as an effective therapeutic strategy to delay AML development21. To evaluate ferroptosis sensitivity, several AML cell lines were treated with ferroptosis inducer Erastin or RSL3. A highest ferroptosis sensitivity was found in HL-60 cells, followed by NB-4 and KG-1 cells (Fig. 3A). However, ferroptosis resistance was found in THP-1 cells (Fig. 3A). Erastin and RSL3-mediated inhibition in AML cell viability was weakened by ferroptosis inhibitor Fer-1 or DFO, which was not affected by apoptosis inhibitor (ZVAD-FMK), necrosis inhibitor (Nec-1) or autophagy inhibitor (3-MA) (Supplementary Fig. 4). Interestingly, THRAP3 silencing enhanced the sensitivity of THP-1 cells to Erastin and RSL3, whereas THRAP3 overexpression reduced ferroptosis sensitivity in HL-60 cells (Fig. 3B). THRAP3 overexpression remarkably reduced MDA and iron levels, but not significantly affected proliferation, lipid ROS level, GSH/GSSG and NADPH/NADP+ ratios in THP-1 cells (Supplementary Fig. 5A–F). Furthermore, Erastin or RSL3-induced increase in lipid peroxidation and MDA level was intensified by THRAP3 depletion, but weakened by THRAP3 overexpression in THP-1 and HL-60 cells (Fig. 3C, D). In subcutaneous xenograft model, THRAP3 deficiency delayed tumor growth and reinforced RSL3-mediated inhibition in tumor growth. On the contrary, THRAP3 overexpression accelerated tumor growth and counteracted the anti-tumor effect of RSL3 (Fig. 3E, F). Besides, Ki-67 expression was reduced, but 4-HNE expression was raised by THRAP3 knockdown or RSL3 treatment, which was more obvious when THRAP3 inhibition was combined with RSL3 (Supplementary Fig. 6). Whereas THRAP3 overexpression resulted in contrary results and reversed RSL3-induced down-regulation of Ki-67 and up-regulation of 4-HNE (Supplementary Fig. 6). Furthermore, THRAP3 depletion-mediated elevation in peroxidation and MDA levels, inhibition in tumor growth, down-regulation of Ki-67 and up-regulation of 4-HNE were counteracted by Fer-1 (Fig. 3C–F and Supplementary Fig. 6). To sum up, THRAP3 facilitated AML development via inducing ferroptosis resistance.
A AML cells were treated with various concentrations of Erastin/RSL3, and cell viability was assessed by CCK-8. B AML cells were transfected with sgTHRAP3 or THRAP3 expression plasmid, followed by exposure to various concentrations of Erastin/RSL3, and cell viability was detected by CCK-8. sgTHRAP3 or THRAP3 expression plasmid-transfected AML cells were treated with or without Erastin/RSL3. C Lipid peroxidation was measured by C11-BODIPY 581/591 probe. D MDA level was assessed by commercial kit. Subcutaneous AML xenografts were generated by subcutaneous injection with HL-60 cells into BALB/c nude mice. E, F Tumor volume and weight were monitored. n = 3, biological replicates for (A–D); n = 6 mice/group for E&F. The data are represented as mean ± standard deviation (SD). Experiments were repeated independently for 3 times with similar results, and representative images from one experiment are shown in (C). One-way ANOVA followed by Tukey’s post hoc test (for A–F) was performed for statistical analysis. *p < 0.05, **p < 0.01, ***p < 0.001. Source data are provided as a Source Data file.
THRAP3 regulates iron metabolism and GSH synthesis in AML cells
To investigate the downstream targets of THRAP3, LC-MS/MS was carried out in THRAP3-knockout THP-1 cells. The heatmap showed that 1033 proteins were up-regulated, while 693 proteins were down-regulated by THRAP3 knockout in THP-1 cells (Supplementary Fig. 7A). As analyzed by KEGG pathway, these differentially expressed proteins were mainly associated with ferroptosis and metabolic pathways (Supplementary Fig. 7B). Figure 4A, B illustrates differential expression of ferroptosis-related proteins, including FTL, GSS, GCLM, GCLC, TFRC, and ACSL4 in THRAP3-knockout THP-1 cells. Western blotting further validated that THRAP3 depletion reduced FTL, GSS, GCLM, GCLC expression, and increased TFRC expression. However, THRAP3 overexpression led to opposite results in HL-60 cells (Fig. 4C). Of note, ACSL4 expression was not affected by THRAP3 (Fig. 4C). Additionally, iron and LIP levels were elevated by THRAP3 depletion, but reduced in THRAP3 overexpression (Fig. 4D, E). The anabolism of GSH is illustrated in Fig. 4F. The GSH/GSSG and NADPH/NADP+ ratios were decrease by THRAP3 knockdown, whereas THRAP3 overexpression exhibited the opposite roles (Fig. 4G, H). THRAP3 overexpression remarkably reduced ROS level in HL-60 cells, and THRAP3 silencing significantly increased ROS level in THP-1 and KG-1 cells (Fig. 4I). The silencing efficiency of THRAP3 was validated by RT-qPCR and Western blotting in KG-1 cells (Supplementary Fig. 2C). Moreover, treatment with antioxidant NAC, ferroptosis inhibitor Fer-1 or DFO recovered the reduced viability of THRAP3-deficient cells (Fig. 4J). In addition, Eratin or RSL3 treatment increased iron and ROS levels, and lowered GSH/GSSG and NADPH/NADP+ ratios, which could be enhanced by THRAP3 deficiency in THP-1 cells, but weakened by THRAP3 overexpression in HL-60 cells (Supplementary Fig. 7C–F). These findings suggested that THRAP3 inhibited iron accumulation and GSH synthesis to suppress ferroptosis in AML cells.
A, B THP-1 cells and THRAP3-knockout THP-1 cells were analyzed by LC-MS/MS. Heatmap showed differentially expressed proteins between control and THRAP3-knockout groups. THP-1 cells were transfected with sgTHRAP3 or sgNC; HL-60 cells were transfected with THRAP3 expression plasmid or vector. C The protein levels of FTL, GSS, GCLM, GCLC, TFRC, and ACSL4 were determined by Western blotting. D iron and (E) labile iron pool (LIP) levels were detected. F The anabolism of GSH was illustrated. G GSH/GSSG and (H) NADPH/NADP+ ratios were evaluated. I THP-1 cells were transfected with sgNC or sgTHRAP3; KG-1 cells were transfected with siNC or siTHRAP3. ROS level was detected by DCFH-DA probe. J siTHRAP3 or sgTHRAP3-transfected cells were pre-treated with antioxidant NAC, Fer-1 or DFO. Cell viability was measured by CCK-8. n = 3, biological replicates for (A–E, G–J). The data are represented as mean ± standard deviation (SD). Experiments were repeated independently for 3 times with similar results, and representative images from one experiment are shown in C&I. Two-tailed Student’s t test (for C–E, G–I) and one-way ANOVA followed by Tukey’s post hoc test (for J) was performed for statistical analysis. *p < 0.05, **p < 0.01, ***p < 0.001. Source data are provided as a Source Data file.
THRAP3 facilitates GIT2 Exon14 skipping in AML cells
Given that abnormal AS events occurred in AML patients22, we further evaluated the changes in AS caused by THRAP3 knockout in THP-1 cells via RNA-seq. Of note, THRAP3 knockout resulted in 67.63% skipping exon (SE) among all AS (Fig. 5A). Additionally, the GEO database was adopted to analyze AS events across three datasets: 17 healthy samples from GSE183252, 6 AML patients from GSE151594, and 54 AML patients from GSE49642. GIT2 and BPTF were screened out by overlapping the data of SE from AML patients and THRAP3-knockout THP-1 cells (Fig. 5B). Because the sequences of skipping exon for BPTF were different in AML patients and THRAP3-knockout THP-1 cells, this study focused on exploring the mechanism of GIT2 exon skipping. Subsequently, gene ontology (GO) enrichment analysis showed that the differentially expressed alternative splicing genes after THRAP3 knockout were mainly associated with metal ion homeostasis (Fig. 5C). Figure 5D illustrates GIT2 Exo14 skipping. We have performed RACE assay in THP-1 cells and found that there were two transcripts for GIT2: GIT2-S (NM_001330153.2) and GIT2-L (NM_057169.5), respectively (Supplementary Data 1). We detected the levels of GIT2 isoforms in various cell lines. GIT2-S/GIT2-L ratio was higher in THP-1 cells with high THRAP3 expression, while GIT2-S/GIT2-L ratio was lower in HL-60 cells with low THRAP3 expression (Fig. 5E). We further verified the influence of THRAP3 on GIT2 exon skipping in AML cells by RT-PCR. The results showed that GIT2 Exon14 skipping was facilitated by THRAP3 overexpression, while restrained by THRAP3 silencing (Fig. 5F). THRAP3 overexpression reversed THRAP3 silencing-mediated inhibition in GIT2 Exon14 skipping (Fig. 5F). RIP assay further validated that THRAP3 could directly bind to GIT2 pre-mRNA in AML cells (Fig. 5G). Furthermore, GIT2 Exon14 skipping was elevated in AML patients by RT-PCR (Fig. 5H). Pearson correlation analysis showed that THRAP3 mRNA level was positively correlated with GIT2 Exon14 skipping level in AML patients (Fig. 5I). Moreover, GIT2 Exon14 skipping level was lower in CR group, but higher in relapse group in comparison with newly diagnosed AML patients (Fig. 5J). The above observations suggested that THRAP3 contributed to GIT2 Exon14 skipping in AML cells.
A Alternative splicing (AS) events were analyzed in THP-1 cells with and without THRAP3 knockout. The distribution of AS events was illustrated in the pie chart. B Overlap of genes with exon skipping from RNA-Seq data of AML patients (GEO database) (normal, n = 17; AML, n = 60) and THRAP3 knockout-THP-1 cells (n = 3). C Gene ontology (GO) analysis of differentially expressed genes responsive to THRAP3 absence in THP-1 cells. D Schematic diagram of GIT2 Exo14 skipping. E The levels of GIT2 isoforms in various cell lines were analyzed by RT-PCR. F GIT2 Exo14 skipping in THRAP3-silenced or overexpressed AML cells was detected by RT-PCR. G The binding or THRAP3 to GIT2 pre-mRNA was validated by RIP assay. H GIT2 Exo14 skipping in control and AML samples was analyzed by RT-PCR (n = 12). N, normal BM samples; A, AML patients’ BM samples. I Pearson correlation analysis of THRAP3 and GIT2 Exon14 skipping (n = 12). (J) RT-PCR evaluated GIT2 Exo14 skipping in newly diagnosed (n = 82), complete remission (CR) (n = 42) and relapse AML patients (n = 16). n = 3, biological replicates for (E–G). The data are represented as mean ± standard deviation (SD). Experiments were repeated independently for 3 times with similar results, and representative images from one experiment are shown in (E, F, H). Hypergeometric test with Benjamini-Hochberg FDR correction (for C), two-tailed Student’s t test (for F, G, H) and one-way ANOVA followed by Tukey’s post hoc test (for E, J) was performed for statistical analysis. *p < 0.05, **p < 0.01, ***p < 0.001. Source data are provided as a Source Data file.
GIT2 Exon14 skipping is associated with AML cell ferroptosis
Given that GIT2 Exon14 skipping was promoted by THRAP3, we further explored the influence of GIT2 Exon14 skipping on ferroptosis sensitivity of AML cells. Figure 6A presents the schematic of designing shGIT2-L (before skipping) and shGIT2-S (after skipping) sequences. RT-qPCR demonstrated that shGIT2-L remarkably decreased GIT2-L level, but not affected GIT2-S level (Fig. 6B). Besides, shGIT2-S reduced GIT2-S level, but not affected GIT2-L level (Fig. 6B). Transfection with shGIT2-L reduced the sensitivity of THP-1 cells to erastin or RSL3. By contrast, shGIT2-S enhanced ferroptosis sensitivity of AML cells (Fig. 6C). Similarly, lipid peroxidation, MDA, iron, LIP levels were decreased, and GSH/GSSG and NADPH/NADP+ ratios were increased after GIT2-L knockdown or Fer-1 treatment. Whereas, these results were the opposite in shGIT2-S-transfected group in the presence or absence of RSL3 (Fig. 6D–I). In addition, FTL, GSS, GCLM, and GCLC were up-regulated, and TFRC was down-regulated by GIT2-L deficiency. On the contrary, GITT2-S knockout down-regulated FTL, GSS, GCLM, and GCLC, and up-regulated TFRC (Fig. 6J). To further elucidate the regulation of GIT2 exon 14 skipping in ferroptosis, AML cells were transfected with shGIT2-L or shGIT2-S, followed by treatment with buthionine sulfoximine (BSO) or glutathione ethyl ester (GSH-EE), respectively. We found that GIT2-L knockdown-induced down-regulation of lipid ROS, MDA, iron, and LIP levels, and up-regulation of GSH/GSSG and NADPH/NADP+ ratios were reversed by BSO co-treatment (Supplementary Fig. 8A–F). Accordingly, GIT2-S knockdown-induced up-regulation of lipid ROS, MDA, iron, and LIP levels, and down-regulation of GSH/GSSG and NADPH/NADP+ ratios were reversed by GSH-EE co-treatment (Supplementary Fig. 8A-F). Moreover, we used antisense oligonucleotide1-3 (ASO1-3) (targeting the binding sites of SLU7 to GIT2 pre-mRNA) to repress GIT2 exon14 skipping (Fig. 7A). ASO2 with high interference efficient for GIT2 exon14 skipping was selected (Fig. 7B). In addition, ASO2 enhanced ferroptosis sensitivity of AML cells, elevated lipid ROS, MDA, iron, LIP levels, and reduced GSH/GSSG and NADPH/NADP+ ratios (Fig. 7C-I). Furthermore, FTL, GSS, GCLM, and GCLC were down-regulated, and TFRC was up-regulated by ASO2 (Fig. 7J). For the in vivo xenograft model, tumor growth was accelerated by shGIT2-L or Fer-1 treatment, while delayed by shGIT2-S in the presence or absence of RSL3 (Fig. 6K, L). Moreover, GIT2-L depletion enhanced Ki-67 expression, but restrained 4-HNE expression in the tumor tissues treated with or without RSL3, and contrary results were caused by GIT2-S silencing (Supplementary Fig. 9). The above data proved that GIT2 Exon14 skipping facilitated AML cell ferroptosis resistance, thus favoring AML progression.
A Schematic outline of shRNA constructs targeting specific GIT2 exon splice junction (shGIT2-L and shGIT2-S). B The silencing efficiency of shGIT2-L and shGIT2-S was verified by RT-qPCR. C AML cells were transfected with shGIT2-L or shGIT2-S, followed by treatment with various concentrations of Erastin/RSL3, and cell viability was assessed by CCK-8. shGIT2-L or shGIT2-S-transfected AML cells were treated with or without RSL3. D Lipid peroxidation was determined by C11-BODIPY 581/591 probe in the presence or absence of RSL3. (E–G) MDA, iron, and LIP levels were measured in the presence or absence of RSL3. H GSH/GSSG and (I) NADPH/NADP+ ratios were detected. J The protein abundance of FTL, GSS, GCLM, GCLC, and TFRC was analyzed by Western blotting. K, L Subcutaneous AML xenograft growth was monitored. n = 3, biological replicates for (B–J); n = 6 mice/group for K&L. The data are represented as mean ± standard deviation (SD). Experiments were repeated independently for 3 times with similar results, and representative images from one experiment are shown in D & J. Two-tailed Student’s t test (for B) and one-way ANOVA followed by Tukey’s post hoc test (for C–L) was performed for statistical analysis. *p < 0.05, **p < 0.01, ***p < 0.001. Source data are provided as a Source Data file.
A Schematic outline of antisense oligonucleotide1-3 (ASO1-3). B The interference efficient for GIT2 exon14 skipping of ASO1-3 was evaluated by RT-PCR. C AML cells were transfected with NC or ASO2, followed by treatment with various concentrations of Erastin/RSL3, and cell viability was assessed by CCK-8. D Lipid peroxidation was determined by C11-BODIPY 581/591 probe in the presence or absence of RSL3. E–G MDA, iron, and LIP levels were assessed in the presence or absence of RSL3. H GSH/GSSG and (I) NADPH/NADP+ ratios were detected. J The protein levels of FTL, GSS, GCLM, GCLC, and TFRC were measured by Western blotting. n = 3, biological replicates for (B–J). The data are represented as mean ± standard deviation (SD). Experiments were repeated independently for 3 times with similar results, and representative images from one experiment are shown in (B, D, J). Two-tailed Student’s t test (for F–J) and One-way ANOVA followed by Tukey’s post hoc test (for B–D) was performed for statistical analysis. *p < 0.05, **p < 0.01, ***p < 0.001. Source data are provided as a Source Data file.
THRAP3 recruits SLU7 to promote GIT2 Exon14 skipping
We next uncovered the exact mechanism through which THRAP3 facilitated GIT2 Exon14 skipping. We identified 371 binding proteins for THRAP3 by Co-IP/MS (Supplementary Data 2). In addition, SLU7, SMU1, CPSF1, and SRPK1 were selected based on high prot scores and association with alternative splicing. As verified by Co-IP, SLU7 protein possessed highest binding efficiency after immunoprecipitation with THRAP3 antibody (Supplementary Fig. 10A). Thus, SLU7 was focused on in the subsequent experiments. Next, we measured SLU7 protein level in multiple AML cells and found that there was no significant change in SLU7 protein level among multiple AML cells (Supplementary Fig. 10B), suggesting that SLU7 is more a cell type-independent general splicing co-factor most likely ubiquitous expressed. Silver staining MS analysis validated the direct interaction between THRAP3 and SLU7 proteins (Fig. 8A). Consistently, Co-IP and immunofluorescent staining assays demonstrated that THRAP3 directly interplayed with SLU7 protein in AML cells (Fig. 8B, C). To identify the exact binding domain of THRAP3, we constructed multiple short-truncated plasmids for THRAP3 (Fig. 8D). Co-IP further verified that SLU7 could bind to THRAP3 when C359 domain was absent, but not when △N190 or △NC domain was absent (Fig. 8E), indicating that SLU7 bound to N-terminal 1-190aa of THRAP3. Similarly, Fig. 8F illustrated the construction of short-truncated plasmids for SLU7. Co-IP assay demonstrated that the binding of THRAP3 to SLU7 was almost abolished by truncated NLS or zinc knuckle regions, but not affected by truncated Leu or Lys region, indicating that THRAP3 could bind to zinc knuckle region of SLU7, rather than Leu or Lys region (Fig. 8G). Yeast two-hybrid assay confirmed the interaction between THRAP3 N-terminal and SLU7 protein (Supplementary Fig. 10C). Consistently, proximity ligation assay validated that THRAP3 N-terminal could directly bind to SLU7 protein (Fig. 8H). The silencing or overexpression efficiency of SLU7 was validated by RT-qPCR and Western blotting in THP-1 and HL-60 cells (Supplementary Fig. 2D, E). Furthermore, GIT2 Exon14 skipping was repressed by SLU7 depletion, but promoted by overexpression of SLU7 with absence of Leu or Lys. However, overexpression of SLU7 with absence of NLS or Zinc-knk did not affect GIT2 Exon14 skipping (Fig. 8I). Moreover, RIP assay revealed that the binding of SLU7 protein to GIT2 pre-mRNA was weakened by THRAP3 knockdown, but enhanced by THRAP3 overexpression (Fig. 8J, K and Supplementary Fig. 10D). However, SLU7 knockout did not affect the binding between THRAP3 and GIT2 pre-mRNA (Fig. 8K). Notably, knockout of THRAP3 abolished the promoting effect of SLU7 overexpression on the skipping of exon 14 of GIT2, and THRAP3 overexpression-mediated GIT2 Exon14 skipping was abolished by SLU7 depletion (Fig. 8L). Taken together, THRAP3 directly interacted with SLU7 to promote GIT2 Exon14 skipping.
A Immunoprecipitation of THRAP3-interacting protein SLU7 in AML cells was visualized by silver staining. B The direct interaction between THRAP3 and SLU7 proteins was validated by Co-IP. C Co-localization of THRAP3 and SLU7 proteins in AML cells was observed by immunofluorescent staining (scale bar = 20 μm). D Schematic of truncated plasmids for THRAP3. E 293 T cells were transfected with full-length or truncated plasmids of THRAP3. The binding of SLU7 to exact domains of THRAP3 protein was evaluated by Co-IP. F Schematic of truncated plasmids for SLU7. G 293 T cells were transfected with full-length or truncated plasmids of SLU7. The binding of THRAP3 to exact domains of SLU7 protein was analyzed by Co-IP. H Proximity ligation assay validated the interaction between THRAP3 N-terminal and SLU7 protein. (scale bar = 5 μm) (I) The regulation of various domains of SLU7 protein in GIT2 Exon14 skipping was evaluated by RT-PCR. J The interplay between SLU7 protein and GIT2 pre-mRNA was verified by RIP. K The binding of SLU7/THRAP3 protein to GIT2 pre-mRNA in sgTHRAP3/THRAP3 expression plasmid/sgSLU7-transfected AML cells was determined by RIP. L RT-PCR analysis of GIT2 Exon14 skipping in AML cells transfected with sgSLU7, THRAP3 expression plasmid, sgTHRAP3 or SLU7 expression plasmid. n = 3, biological replicates for (A–C, E, G–L). The data are represented as mean ± standard deviation (SD). Experiments were repeated independently for 3 times with similar results, and representative images from one experiment are shown in (A–C, E, G–I, L). One-way ANOVA followed by Tukey’s post hoc test was performed for statistical analysis. *p < 0.05, **p < 0.01, ***p < 0.001. Source data are provided as a Source Data file.
THRAP3 knockdown enhances ferroptosis sensitivity of AML cells via inhibiting GIT2 Exon14 skipping
To further verify whether GIT2 Exon14 skipping was involved in THRAP3-mediated ferroptosis resistance, THP-1 cells were transfected with GIT2-L overexpression plasmid, sgTHRAP3 combined with or without GIT2-S overexpression plasmid. CCK-8 data indicated that loss of THRAP3 or GIT2-L overexpression reduced the viability of AML cells treated with or without RSL3, and overexpression of GIT2-S effectively restored the viability of THRAP3-deficient cells (Fig. 9A). THRAP3 knockdown or GIT2-L overexpression enhanced lipid peroxidation and MDA levels, which were counteracted by GIT2-S overexpression (Fig. 9B, C). TEM showed that THRAP3 deficiency or GIT2-L overexpression resulted in mitochondrial dysfunction, as evidenced by disrupted cristae and reduction in volume (condensation). However, the above ultrastructural injury was relieved by GIT2-S overexpression (Fig. 9D, E). Besides, THRAP3 depletion or GIT2-L overexpression enhanced iron and LIP levels, and reduced GSH/GSSG, and NADPH/NADP+ ratios without RSL3 treatment, and these changes were reversed by GIT2-S overexpression (Fig. 9F–I). sgTHRAP3 or GIT2-L overexpression decreased FTL, GSS, GCLC, GCLM levels, and increased TFRC level, which could be reversed by GIT2-S overexpression (Fig. 9J). Next, we further validated the involvement of GIT2 Exon14 skipping in THRAP3-induced ferroptosis resistance. Considering that HL-60 cells exhibited a lower level of GIT2-S, HL-60 cells were transfected with THRAP3 or GIT2-S overexpression plasmid together with or without shGIT2-S. As expected, THRAP3 or GIT2-S overexpression increased cell viability, reduced lipid peroxidation, MDA, iron, and LIP levels, attenuated mitochondrial dysfunction, raised GSH/GSSG, and NADPH/NADP+ ratios, up-regulated FTL, GSS, GCLC, GCLM, and down-regulated TFRC in HL-60 cells, which were neutralized by shGIT2-S transfection (Supplementary Fig. 11A–I). Therefore, these findings demonstrated that THRAP3 conferred ferroptosis resistance of AML cells by promoting GIT2 Exon14 skipping.
THP-1 cells were transfected with GIT2-L overexpression plasmid, sgTHRAP3 combined with or without GIT2-S overexpression plasmid in the presence or absence of RSL3. A Cell viability was assessed by CCK-8. B Lipid peroxidation was evaluated by C11-BODIPY 581/591 probe. C MDA level was detected. D, E Mitochondrial dysfunction was observed by TEM (scale bar = 500 nm). F–I Iron, LIP, GSH/GSSG and NADPH/NADP+ ratios were detected. J The protein abundance of FTL, GSS, GCLM, GCLC, and TFRC was determined by Western blotting. n = 3, biological replicates for (A–J). The data are represented as mean ± standard deviation (SD). Experiments were repeated independently for 3 times with similar results, and representative images from one experiment are shown in (B, D, J). One-way ANOVA followed by Tukey’s post hoc test was performed for statistical analysis. *p < 0.05, **p < 0.01, ***p < 0.001. Source data are provided as a Source Data file.
THRAP3 drives AML growth in vivo via promoting GIT2 Exon14 skipping
Finally, we determined whether THRAP3 facilitated AML progression via promoting GIT2 Exon14 skipping in nude mice in vivo. THRAP3 or GIT2-S overexpression accelerated tumor xenograft growth, increased Ki-67 expression and reduced 4-HNE expression, whereas GIT2-S down-regulation counteracted these changes (Fig. 10A, B and Supplementary Fig. 12). THRAP3 knockout or GIT2-L overexpression inhibited tumor xenograft growth, decreased Ki-67 expression and increased 4-HNE expression, whereas GIT2-S overexpression counteracted these changes (Fig. 10A, B and Supplementary Fig. 12). The survival of mice was decreased by THRAP3 or GIT2-S overexpression, which could be restored by GIT2-S silencing (Fig. 10C). In the metastasis model by tail vein injection, THRAP3 or GIT2-S overexpression elevated liver and spleen weight, together with percentage of CD45+Ki67+, whereas these changes were abolished by shGIT2-S (Fig. 10D–F). Therefore, these findings suggested that THRAP3-mediated GIT2 Exon14 skipping contributed to AML development in vivo.
A, B Xenograft tumor volume and weight were monitored. C The survival rate of mice was evaluated. HL-60 cells were injected into NOD/SCID mice via tail vein to establish a metastatic model. D, E The liver and spleen were collected and weighed. F The percentage of CD45+Ki67+ in liver and spleen was assessed by flow cytometry. n = 6 mice/group for A-F. The data are represented as mean ± standard deviation (SD). Experiments were repeated independently for 6 times with similar results, and representative images from one experiment are shown in (F). One-way ANOVA followed by Tukey’s post hoc test was performed for statistical analysis. *p < 0.05, **p < 0.01, ***p < 0.001. Source data are provided as a Source Data file.
Discussion
Although improvements have been made in AML therapies, including identification of therapeutic targets, chemotherapy drugs, and stem cell transplantation, the prognosis of AML patients remains poor23. Studies have proved that ferroptosis induction could effectively remove chemotherapy-resistant AML cells, thus delaying cancer recurrence and development24,25. Therefore, ferroptosis induction may be a promising anticancer strategy for AML. It has been recognized that dysregulation of iron metabolism and lipid peroxidation-related genes and pathways might affect ferroptosis sensitivity26. Here, we elucidated the regulation and mechanism of THRAP3 in ferroptosis sensitivity in AML cells. We found that abnormal high expression of THRAP3 promoted AML progression via repressing iron metabolism and GSH synthesis to reduce ferroptosis sensitivity of AML cells. Mechanistically, THRAP3 recruited SLU7 to facilitate GIT2 Exon14 skipping, thus resulting in AML cell ferroptosis resistance. Our findings uncover the mechanism of THRAP3 in ferroptosis resistance, which suggest THRAP3 as a potential therapeutic target for AML.
THRAP3 is a well-known regulator of AS, which possesses crucial biological functions. Vohhodina et al. suggested that loss of THRAP3 enhanced sensitivity of tumor cells to DNA damaging chemotherapy by the modulation of mRNA splicing and nuclear export27. Another study documented that THRAP3 was responsible for expression of HPV16 E6 and E7 oncoproteins by promoting E6*I/E7 mRNA splicing, thereby conferring the carcinogenic properties of human papillomaviruses28. Even so, the regulation of THRAP3 in AML has not been clarified. Our results demonstrated that THRAP3 was up-regulated in the BM samples of AML patients, which was associated with a poor prognosis. Overexpression of THRAP3 accelerated AML tumorigenesis in vivo, while THRAP3 knockdown exerted an opposite role. This study provided evidence that THRAP3 served as a contributor to AML development. As reported, TP53 gene was absent in HL-60, KG-1, NB-4, and THP-1 cell lines29. Besides, NB-4 cell line has the highest number of mutated genes related to homologous recombination repair (BRCA1, RAD51D, PAXIP1, and RBBP8) that is responsible for the accurate repair of DNA double-strand breaks29. Thus, these AML cell lines exhibit differences in genetic background, especially in DNA double-strand breaks-related genes. Consistent with this study, we found a difference in THRAP3 expression level in various AML cell lines. So far, whether THRAP3 expression level can be affected by driver mutations, genetic instability, and cell cycle progression in AML cells has not been determined. A previous study showed that the mRNA stability and expression of THRAP3 was regulated by METTL3-mediated m6A methylation in multiple myeloma cells30. The possible factors that influence THRAP3 expression in AML cells need to be further clarified.
Ferroptosis is a kind of cell death, featured by abnormal iron metabolism, lipid peroxidation, and GSH depletion. Recently, multiple enzymes and molecules have been identified to govern the complicated process of ferroptosis. For instance, TFRC is an iron transporter that contributes to the entrance of Fe2+ into LIP and following ROS production, thereby triggering ferroptosis31. FTL is the key iron storage protein, and its down-regulation causes imbalance of intracellular iron to promote ferroptosis32. GSS is responsible for GSH synthesis, and depletion of GSS induces ferroptosis via enhancing Fe2+ accumulation33. GSH synthesis can be catalyzed by GCLC and GCLM, two subunits of glutamate cysteine ligase, and deficiency of GCLC and GCLM reduces antioxidant capacity to promote ferroptosis34. Herein, we discovered that THRAP3 deficiency enhanced ferroptosis sensitivity of AML cells as evidenced by increasing iron, LIP and ROS levels and reducing GSH synthesis through up-regulation of TFRC, and down-regulation of FTL, GSS, GCLM, GCLC. These findings suggested that THRAP3 facilitated AML progression via increasing ferroptosis sensitivity of AML cells.
AS, a key biological process, maintains a wonderful balance among various isoforms, which exerts crucial biological functions. However, abnormal AS events may promote the expression of oncogenic alternative mRNA isoforms in AML cells35. AML cells may enhance malignant growth capacity via altering AS36. Our data revealed that abnormal AS events, especially exon skipping could be caused by THRAP3 depletion in AML cells. GIT2 is a member of the GIT family, which modulates multiple cellular processes, including inflammation, immune response, aging process, and tumorigenesis37,38,39,40. GIT2 can undergo extensive AS to generate diverse mRNA forms that possess different structures and functions41. So far, whether GIT2 undergo undergoes AS during AML progression has not been reported. This study discovered that abnormal GIT2 Exon14 skipping was enhanced in AML patients, especially in recurrent patients. Additionally, RNA-seq and RT-PCR results demonstrated that GIT2-S (without Exon14) level, as well as the ratio of GIT2-S/GIT2-L (with Exon14) were elevated by THRAP3 overexpression, but reduced by THRAP3 silencing. Ferroptosis can be regulated by the generation of various splicing isoforms via AS. For example, HMGA2 isoforms produced by AS affected ferroptosis sensitivity of prostate cancer cells42. The correlation of GIT2 with superoxide production has been identified43. Furthermore, a previous study also revealed that GIT2 took part in the regulation of oxidative stress44. However, the correlation of GIT2 Exon14 skipping with ferroptosis resistance of AML cells has not been reported. Herein, our data demonstrated that Exon14 skipping-mediated truncated GIT2 isoform facilitated ferroptosis resistance, while GIT2-L (before skipping) enhanced ferroptosis sensitivity of AML cells. Furthermore, THRAP3 contributed to GIT2 Exon14 skipping to result in ferroptosis resistance, thus accelerating AML progression. Our observations proved that THRAP3 conferred ferroptosis resistance by promoting Exon14 skipping of GIT2 in AML cells.
SLU7 is known as a splicing regulator, which regulates 3′ site choice during the step 2 in exon ligation45. Human SLU7 protein is made up of 3 functional domains, including zinc-knuckle motif, lysine-rich region, and nuclear localization signal. SLU7 is involved in the modulation of gene splicing during cancer development46,47. Thus, we speculated that SLU7 might contribute to THRAP3-mediated GIT2 Exon14 skipping in AML cells. Surprisingly, our findings indicated that THRAP3 N-terminal could directly bind to zinc-knuckle motif of SLU7 protein. Furthermore, genetic silencing of THRAP3 abolished GIT2 Exon14 skipping via interaction with SLU7 in AML cells. Therefore, SLU7 zinc-knuckle motif was responsible for THRAP3-mediated GIT2 Exon14 skipping.
There are several limitations in this study. Firstly, we investigated the influence of GIT2 Exon14 skipping on AML ferroptosis resistance; however, how GIT2 alternative splicing influences the translated protein remains unknown. NP_001317082.1 (708 AA) is the encoding protein for GIT2-S; NP_476510.1 (759 AA) is the encoding protein for GIT2-L. As compared with NP_476510.1 (759 AA), NP_001317082.1 (708 AA) is lack of GIT coiled-coil Rho guanine nucleotide exchange factor (GIT-CC domain). As reported, GIT-CC domain determined the complex formation between GIT and PIX proteins48. Thus, we speculated that loss of GIT-CC domain might influence the stability, biological function, and localization of GIT2 protein, which might be responsible for GIT2 Exon14 skipping-induced ferroptosis resistance of AML cells. Secondly, although GIT2 exon 14 skipping affected GSH synthesis, iron metabolism, and the expression of ferroptosis-related genes of AML cells, its potential mechanism is unclear. Thirdly, we analyzed the correlation between GIT2 Exon14 skipping and THRAP3 level. The low r2 value suggested that THRAP3 was not the only factor influenced GIT2 Exon14 skipping. Thus, we propose hypotheses that other potential factors might also affect GIT2 Exon14 skipping, which needs to be further elucidated. Fourthly, the growth of tumor is a complex biological process that is influenced by multiple factors. In addition to ferroptosis, other types of death may also involve in the regulation of tumor growth. Previous studies have illustrated that ferroptosis, as a lipid peroxidation-dependent cell death, may indirectly induce apoptosis via releasing oxidative stress products49,50,51. Whether THRAP3 regulates the crosstalk of different cell death pathways and the related mechanisms deserve to be further explored. Fifthly, apart from aberrant splicing, many regulatory mechanisms can affect protein levels, such as transcriptional regulation, translation efficiency, posttranslational modification, protein stability, and so on52,53. For the non-overlapping differentially expressed proteins and the differentially spliced genes, the specific mechanisms of their expression changes still require further investigation. These issues are the directions for our future experiments.
Taken together, this study discovered that THRAP3 high expression was responsible for ferroptosis resistance of AML cells. Moreover, we demonstrated that THRAP3 interacted with SLU7 zinc-knuckle motif to facilitate GIT2 Exon14 skipping, thus reducing ferroptosis sensitivity and driving AML progression. Although the possible mechanism for abnormal high expression of THRAP3 in AML remains to be explored, our results expound functions of THRAP3 in AML ferroptosis resistance and suggest THRAP3 as a promising therapeutic target for AML.
Methods
Ethics statement
All experimental procedures were approved by the Ethics Committee of the Third Xiangya Hospital of Central South University. The methods were carried out in accordance with the approved guidelines.
Clinical samples
Bone marrow (BM) samples were randomly collected from healthy donors (n = 12, 6 males/6 females, median age 35, range 25-48) and AML patients in the stages of new diagnosis (n = 82, 43 males/39 females, median age 55, range 21-75), complete remission (CR, n = 42, 28 males/14 females, median age 52, range 19-72) and relapse (n = 16, 11 males/5 females, median age 57, range 24-70) at the Third Xiangya Hospital of Central South University according to the Declaration of Helsinki. Written informed consent was collected from all participators. There was no compensation for participators. The Ethics Committee of the Third Xiangya Hospital of Central South University approved this study.
Isolation of bone marrow stromal cells (BMSCs) and bone marrow mononuclear cells (BMMNCs)
BMSCs were isolated from normal human BM samples. In brief, the BM samples were added with heparin and bone marrow solution, and centrifugated at 1500 × g for 5 min. The BMSCs in the cell pellets were cultured with α-MEM medium (Gibco, NY, USA) containing 10% fetal bovine serum (FBS, Gibco). BMMNCs were extracted from normal human BM samples via density gradient centrifugation using the Ficoll-Paque Plus (Amersham Biosciences, Sweden). BMMNCs were cultured with α-MEM medium supplemented with 10% FBS.
Cell culture and treatments
Normal BM stromal cell line HS-5 (Cat. No.: CRL-3611) and AML cell lines THP-1 (Cat. No.: TIB-202), HL-60 (Cat. No.: CCL-240) and KG-1 (Cat. No.: CCL-246) were purchased from American Type Culture Collection (USA). AML cell line NB4 (Cat. No.: CL-0676) was purchased from Procell (Wuhan, China). HS-5 cells were cultured with DMEM, THP-1, NB4, HL-60, and KG-1 cells were cultured with RPMI-1640 (Gibco) containing 10% FBS and mycoplasma test was regularly conducted using the MycoAlert-Plus Mycoplasma Detection Kit (Lonza, Basel, Switzerland). All cell lines were authenticated by STR genotyping. AML cells were treated with various concentrations of Erastin (1.25, 2.5, 5, 10, 20, 40, 80 μM, MCE, NJ, USA) or RSL3 (0.25, 0.5, 1, 2, 4, 8, 16 μM, MCE). AML cells were treated with ferroptosis inhibitor Fer-1 (10 μM, MCE) or DFO (0.5 μM, MCE), apoptosis inhibitor ZVAD-FMK (10 μM, MCE), necrosis inhibitor Nec-1 (2 μM, MCE), autophagy inhibitor 3-MA (1 mM, MCE), buthionine sulfoximine (BSO, 100 μM, MCE) or glutathione ethyl ester (GSH-EE, 5 mM, MCE).
Cell transfection and lentivirus infection
Small guide RNA (sgRNA) targeting THRAP3 or SLU7 and negative control sgRNA were designed and synthesized by Editgene (Guangzhou, China). Small interfering RNA targeting THRAP3 (siTHRAP3) and siNC, were provided by GenePharma (Shanghai, China). Lentiviruses carrying overexpression plasmids for THRAP3, SLU7, THRAP3△C359, THRAP3△N190, THRAP3△NC, SLU7△NLS (truncated nuclear localization signal), SLU7△Zinc-knk (zinc knuckle), SLU7△Leu (leucine-rich repeat), SLU7△Lys (lysine enrichment region) and vector, short hairpin RNA targeting SLU7 (shSLU7), shGIT2-L (before skipping), shGIT2-S (after skipping), shNC, GIT2-S, GIT2-L and vector were packaged by GenePharma.
THP-1 or HL-60 cells (5 × 105 cells/mL) were transfected with sgTHRAP3. siTHRAP3, or overexpression plasmids using the Lipofectamine 2000 (Thermo Fisher, MA, USA). For the selection of stable transfected cells, treatment with puromycin at a concentration of 1–10 μg/mL was performed. Lentivirus infection was conducted in the presence of polybrene (5 μg/mL), followed by selection of positive cells using 1 μg/mL puromycin. sgRNA, siRNA, or shRNA sequences are presented in Supplementary Table 1.
Antisense oligonucleotides (ASOs)
ASOs 1–3 targeting the binding sites of SLU7 to GIT2 pre-mRNA were designed using Soligo software, which were used to repress GIT2 exon14 skipping. AML cells were treated with 20 μM ASOs 1-3. GIT2 exon14 skipping was assessed after 48 h. The group exhibiting effective interference efficient for GIT2 exon14 skipping was selected. ASO sequences are listed in Supplementary Table 1.
Cell counting kit-8 (CCK-8)
Cell proliferation was evaluated using the CCK-8 kit (Cat. No.: HY-K0301, MCE). In short, CCK-8 reagent was added to the 96-well plates planted with AML cells (2 × 104 cells per well), followed by incubation for 2 h. The optical density at 450 nm was detected on a microplate reader (BioTek, WA, USA), which indicates the proliferation ratio.
Apoptosis detection
Apoptosis was determined using the Annexin V-FITC Apoptosis Staining/Detection kit (Abcam, UK). Briefly, AML cells were collected and resuspended in binding buffer, followed by staining with 5 μL of Annexin V and 5 μL of propidium iodide for 5 min in the dark. Apoptosis was detected using a flow cytometer (BD Biosciences, NJ, USA) using a minimum of 5000 events. The gating strategy was presented in Supplementary Fig. 13. The percentage of apoptotic cells was evaluated using the BD FACS DIVA 9 software (version 9.0.1).
Flow cytometry and fluorescence activated cell sorting (FACS)
The CD45 and Ki67 percentages in the liver and spleen tissues of nude mice were analyzed by flow cytometry. The liver and spleen tissues of nude mice were minced and subjected to partial enzyme digestion (DNAse, 10104159001, Roche, Basel, Switzerland; 10165921001, Pronase, Roche; 17100, Collagenase, Gibco), and then passed through a 100 μm nylon mesh. Single-cell suspensions were collected and stained with APC-mCD45 (1:40, ab210182, Clone: 104, Abcam) and Alexa Fluor® 488 conjugated-Ki-67 (1:1000, ab281847, Clone: SP6, Abcam). The stained cells were analyzed using a flow cytometer. The gating strategy was presented in Supplementary Fig. 13.
RNA-sequencing (RNA-Seq)
Total RNA was extracted from THP-1 cells after transfection with sgNC or sgTHRAP3 using the TRIzol (Thermo Fisher). The concentration and quality of RNAs were analyzed using the ND-2000 Nanodrop instrument (Thermo Fisher). After section using the oligo-dT beads, libraries were established using the NEBNext Ultra II RNA Library Prep (Illumina, CA, USA). Subsequently, paired-end RNA-Seq was conducted on the NovaSeq 6000 Sequencing System (Illumina). AS events were analyzed using rMATS (v4.1.0) based on splice-junction reads54. AS events were labelled significant if FDR < 0.05 and │Inclusion Level Difference│> 0.15.
Liquid chromatography and tandem mass spectrometry (LC-MS/MS)
The downstream proteins regulated by THRAP3 in THP-1 cells were analyzed by proteomics. Briefly, THP-1 cells were divided into control and KO groups with 3 biological replicates, which were transfected with sgNC or sgTHRAP3, respectively. Subsequently, the cells were lysed, digested by trypsin for 12 h, followed by labeling with Tandem Mass Tags. Then, tagged digests were fractionated and resuspended in 0.1% formic acid. The peptide fractions were separated with LC (PepMap C18 column, 3 μm particle, 100-Å pore; 75 μm 150 mm; Thermo Fisher) on the EASYnLC 1200 system (Thermo Fisher) at 300 nL/min with a 120-min gradient: 0–105 min: 0 to 40% B; 105-110 min: 40 to 70% B; 110–115 min: 70% to 100% B; 115–120 min: 100% B (solvent A: 0.1% vol/vol formic acid; solvent B: 80% vol/vol acetonitrile; column temperature: 50 °C). MS were conducted on the Q-Exactive HF Orbitrap mass spectrometer (Thermo Fisher) with the following settings: polarity, positive; data-dependent acquisition (DDA), top 15 ions; MS resolution, 60,000; mass range, 300-1,650 m/z; precursor dynamic exclusion, 30 s; maximum ion injection time, 20 ms; MS automatic gain control (AGC) target, 3e6; isolation window, 1.4 m/z; stepped normalized collision energy (NCE), 28, 30, 32; MS2 resolution, 60,000; MS2 maximum ion injection time, 100 ms; and MS2 AGC target, 2e5. MS data were interpreted using the Proteome Discoverer v2.4 requiring 1% FDR for identification of UniProt sequences.
Bioinformatics analysis
The Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/) was adopted to analyze AS events across three datasets: 17 healthy samples from GSE183252, 6 AML patients from GSE151594, and 54 AML patients from GSE49642. The screening criteria are as follow: FDR (corrected p value) <0.05; the difference value is set at 0.15.
The differential expression of THRAP3 was analyzed in 3,910 AML patients and 2,175 normal samples, as well as 416 relapsed AML patients and 2,128 primary AML patients, using data from the databases of TARGET, BEAT and LEUCEGENE. The expression levels of THRAP3 in 33 tumor types were analyzed using the GEPIA database (http://gepia2.cancer-pku.cn/#analysis). The differential screening criteria are as follow: log2FC > 1, padj <0.05 for up-regulation; log2FC <-1, padj <0.05 for down-regulation.
The correlation between THRAP3 expression and survival rate of AML patients was analyzed by TCGA database (https://portal.gdc.cancer.gov/). The effect of THRAP3 expression on the overall survival of AML patients from included datasets was drawn based on the optimal threshold. The optimal cut-off point was determined by the “cutp” function using the SurvMisc software (version 0.5.6), which selected the best threshold based on the survival results. To reduce the bias and overfitting, the p values at different cut-off points were adjusted by multiple test adjustments. In each dataset, the patients were divided into high expression group and low expression group based on the optimal cut-off value of THRAP3.
Gene ontology (GO) and Kyoto Encyclopaedia of Genes and Genomes (KEGG)
To analyze the biological significance of differentially expressed genes in THRAP3-knockout THP-1 cells, enrichment analysis of GO and KEGG was performed using the DAVID database (https://david.ncifcrf.gov/). The DAVID database can offer large gene lists from multiple biological perspectives, providing associated biological insights to the fullest extent. The p values (≤0.05) were ranked from the smallest to the largest, which indicated a stronger association. The terms were selected from the GO and KEGG pathway analysis results based on their p Values and visualized through the SangerBox platform (http://sangerbox.com/).
Lipid ROS detection
The lipid ROS was evaluated using the C11-BODIPY 581/591 (Cat. No.: D3861, Thermo Fisher). In short, the AML cells were incubated with 5 µM C11-BODIPY 581/591 reagent for 30 min at 37 °C. After rinsing with PBS, the stained cells were analyzed by flow cytometry. The gating strategy was presented in Supplementary Fig. 13.
Measurement of ROS production
The DCFH-DA probe (Cat. No.: S0033S, Beyotime) was used to determine the ROS level. AML cells were stained with 10 μM DCFH-DA reagent for 30 min, and detected on a flow cytometer.
Measurement of malondialdehyde (MDA), iron, glutathione (GSH)/oxidized glutathione (GSSG), and the reduced form of NADPH/nicotinamide adenine dinucleotide phosphate (NADP+)
The MDA level in AML cells was evaluated using the MDA Content Assay Kit (BC0025, Solarbio). Iron level was calculated using the Cell Iron Content Assay Kit (BC5315, Solarbio). Redox status was determined by GSH/GSSG and NADPH/NADP+ ratios using the GSH and GSSG Assay Kit (S0053, Beyotime) and NADPH/NADP+ assay kit (S0179, Beyotime), respectively.
Labile iron pool (LIP) assay
LIP was detected according to a study by Zhao et al.55. The AML cells were treated with 0.5 µM calcein-acetoxymethyl (Sigma-Aldrich, MO, USA) for 15 min at 37 °C. Subsequently, the cells were incubated with or without 100 µM deferiprone (Sigma-Aldrich) for 1 h at 37 °C. Finally, the cells were examined on a flow cytometer. LIP is calculated as Δfluorescence intensity (calcein-acetoxymethyl/deferiprone–calcein-acetoxymethyl alone).
Immunofluorescence staining
The colocalization of THRAP3 and SLU7 proteins was observed by immunofluorescence staining. AML cells were fixed in 4% paraformaldehyde and then treated with 0.1% Triton X-100 for 10 min, followed by blocking with 5% bovine serum albumin for 1 h. After washing, the cells were probed with primary antibodies against THRAP3 (sc-133250, 1:50, clone name: F-10, Santa Cruz, TX, USA) and SLU7 (sc-376985, 1:50, clone name: B-11, Santa Cruz) at 4 °C overnight. Thereafter, the cells were incubated with FITC Goat Anti-Rabbit IgG (H + L) (AS011, 1:50, ABclonal, Wuhan, China) and Cy3 Goat Anti-Mouse IgG (H + L) (AS008, 1:50, ABclonal) for 1 h. After nuclear staining with DAPI for 5 min, the stained cells were observed under a fluorescence microscopy (Olympus).
Transmission electron microscope (TEM)
The AML cells were seeded in 6-well plates, and then fixed with 2.5% glutaraldehyde overnight. After rinsing with 0.1 M PBS, the cells were treated with cacodylate-buffered tannic acid, fixed with 1% buffered osmium, and stained with 1% uranyl acetate. The cells were embedded in 812 resin and propylene oxide, followed by incubation for 24 h at 60 °C. The images were taken using a transmission electron microscope (Hitachi, Tokyo, Japan).
Yeast two-hybrid system
The specific region of THRAP3-SLU7 interaction was determined using yeast-two hybrid system as previously reported56. THRAP3 plasmid with various domains together with SLU7 plasmid were co-transfected into yeast cells. The cells were allowed to grow on SD plates without tryptophan and leucine as control or without tryptophan, leucine, adenine, and histidine, leucine, and tryptophan for 72 h at 30 °C.
RNA immunoprecipitation (RIP)
The binding of SLU7 to GIT2 pre-mRNA was validated using the RIP-Assay Kit (Cat. No.: Bes5101, BersinBio, Guangzhou, China). AML cells were lysed and supernatant was collected by centrifugation at 13,000 × g for 10 min. The cell lysates were immunoprecipitated with anti-SLU7 (2 µg, sc-376985, Clone name: B11, Santa Cruz) immobilized Protein A/G PLUS-Agarose beads at 4 °C for 3 h with rotation. After centrifugation, the immunoprecipitated RNAs were washed, purified, and detected by RT-qPCR.
Co-immunoprecipitation (Co-IP)
The endogenous interaction between THRAP3 and SLU7 proteins was validated in AML cells using Co-IP. Cells were lysed with RIPA Lysis Buffer containing protease inhibitor cocktail. Subsequently, the cell lysates were pre-cleaned with protein A/G beads at 4 °C for 4 h, followed by immunoprecipitation with anti-THRAP3 (2 µg, sc-133250, Clone name: F10, Santa Cruz), or anti-SLU7 (2 µg, sc-376985, Clone name: B11, Santa Cruz) antibody at 4 °C overnight. The immunoprecipitated proteins were washed with IP lysis buffer and measured by Western blotting. To investigate the specific binding domains between THRAP3 and SLU7 proteins, 293 T cells were transfected with THRAP3, THRAP3△C359, THRAP3△N190, THRAP3△NC, SLU7, SLU7△NLS, SLU7△Zinc-knk, SLU7△Leu, or SLU7△Lys plasmids. Cell lysates were subjected to Co-IP as above described.
Proximity ligation assay
The interaction between THRAP3 N-terminal and SLU7 proteins was assessed using the Duolink proximity ligation assay kit (DUO92007, Sigma-Aldrich). Cells were fixed, permeabilized, blocked, and incubated with primary antibodies against THRAP3 (sc-133250, 1:50, Clone name: F10, Santa Cruz) and SLU7 (ab181138, 1:100, Clone name: EPR11111, Abcam). Duolink probes were applied, followed by ligation and amplification. DAPI was used for counterstaining, and protein interaction was observed under a confocal microscope (Zeiss, Oberkochen, Germany).
Animal experiments
Male 4-5 weeks old NOD/SCID mice and male 4-6 weeks old BALB/c nude mice were purchased from Hunan SJA laboratory animal Co., Ltd, (Changsha, China). Mice were kept under standard housing conditions with a 12:12 h light-dark cycle, an ambient temperature between 20–24 °C and humidity between 45–65%. All mice were maintained on a normal chow diet containing 11.3% kcal fat (1010044, Xietong Shengwu, Nanjing, China). To establish orthotopic AML xenografts, NOD/SCID mice were randomly divided and injected with 1 × 106 HL-60 cells that were infected with lentiviruses carrying vector, THRAP3, shNC, or shGIT2-S via tail vein (n = 12 per group, 6 mice for survival analysis, and 6 mice for AML progression evaluation). Tumor cell progression was evaluated after the injection. The mice were euthanatized by cervical dislocation and then the peripheral blood, liver, and spleen samples were collected. To establish the subcutaneous AML xenografts, BALB/c nude mice were randomly divided and subcutaneously injected with 1 × 106 HL-60 cells stably transfected with sgNC, sgTHRAP3, shNC, shGIT2-L, shGIT2-S or infected with lentiviruses carrying vector, THRAP3, shNC, or shGIT2-S. When the tumor volume was more than 50 mm3, 100 mg/kg RSL3 or 5 mg/kg Fer-1 was injected into the tumors. Tumor volume was calculated by length × (width)2/2. All investigators were blinded to group allocation during the study. All mice were euthanatized by cervical dislocation at the end of the experiments, and tumors were obtained. All experimental procedures were approved by the Ethics Committee of the Third Xiangya Hospital of Central South University. The maximal tumor size permitted in this study was a diameter of 1.5 cm, and this limit was not exceeded in any of the experiments. Welfare monitoring was conducted and if mice exhibited signs of moribund conditions, including hunched posture, reduced activity, labored breathing and weight loss (>20%) or body condition score (BSC) ≤ 2, they were euthanatized as humane endpoint. All in vivo experiments were conducted using male mice. The decision to use a single sex was guided by the primary research objective to investigate the cell-intrinsic mechanism of THRAP3 in ferroptosis resistance, for which no prior evidence suggests a sex-dependent effect. The use of male mice aimed to avoid potential confounders associated with the estrous cycle, and was consistent with prior studies establishing AML animal models57,58.
Immunohistochemistry staining
Immunohistochemistry staining was performed on 4% paraformaldehyde-fixed, paraffins-embedded 4 μm sections of tumors, liver, and spleen tissues. The sections were dewaxed and rehydrated, followed by antigen retrieval using sodium citrate heated by microwave. The sections were probed with Ki-67 (ab279653, Clone name: B56, 1:1000, Abcam), or 4-HNE (ab48506, Clone name: HNEJ-2, 1:25, Abcam) antibody overnight at 4 °C. After incubation with HRP-conjugated secondary antibody (AS033, 1:500, ABclonal) for 30 min at 37 °C, the sections were developed with diaminobenzidine and photographed under a microscope. The percentage of positive staining was quantified using the Image J software.
Reverse transcription polymerase chain reaction (RT-PCR) or quantitative polymerase chain reaction (RT-qPCR)
Total RNA was isolated from AML cells using the TRizol (Thermo Fisher), followed by reverse transcription using the SuperScript VILO kit (Cat. No.: 11754250, Thermo Fisher). Subsequently, the cDNA was subjected to PCR amplification using the PowerUp™ SYBR™ Green Mix (Cat. No.: A25741, Thermo Fisher) and 2 × Accurate Master Mix (Cat. No.: A0211, Accurate Biology, Changsha, China). For RT-qPCR, the relative gene level was calculated using the 2-ΔΔCT method, with GAPDH as a control gene. The RT-qPCR primers are as follow: THRAP3 F: 5’- AGCCCAGAGATACACAGGAGA-3’, R: 5’-GGTTCCCGGGGACTTTTCAT-3’; GIT2-S (transcript ID: NM_001330153.2) F: 5’-CCGGTCTCGTACATGGACATG-3’, R: 5’- GAAGGCTTCAAACACTCCAGAGTG-3’; GIT2-L (transcript ID: NM_057169.5) F: 5’- GGTTGTGGCCTGTTTCCTGAG-3’, R: 5’-GTGGCATCAGACGAAGACACAG-3’ and GAPDH F: 5’-AGGTCGGAGTCAACGGATTT-3’, R: 5’- TGACGGTGCCATGGAATTTG-3’. For PCR, the PCR products were detected using agarose gel electrophoresis and quantified using the ImageJ software (National Institutes of Health, NY, USA). The primers for RT-PCR are as follow: GIT2 F: 5’-GGTTGTGGCCTGTTTCCTGAG-3’, R: 5’-GTGGCATCAGACGAAGACACAG-3’ and GAPDH F: 5’-AGGTCGGAGTCAACGGATTT-3’, R: 5’-TGACGGTGCCATGGAATTTG-3’.
Rapid Amplification of cDNA Ends (RACE)
We extracted total RNA from THP-1 cells. RNA integrity and concentration were determined using the Agilent RNA 6000 Nano II assays (Agilent Technologies, CA, USA). Total RNA (1 μg) was reversely transcribed using the modified oligo (dT) primer and the SMARTScribe Reverse Transcriptase (Clontech, CA, USA). cDNA was amplified using the SMARTer RACE 5′/3’ Kit (Clontech). RACE PCR products were separated by agarose gel electrophoresis, with bands of interest excised from the gel and purified using the Gel-Out Concentrator kit (A&A Biotechnology; Gdynia, Poland).
Western blotting
The cells and tissues were lysed by RIPA buffer (Cat. No.: R0010, Solarbio) supplemented with PMSF (1 mM, Solarbio). The BCA protein assay kit (Cat. No.: PC0020, Solarbio) was used for protein concentration detection. A total of 30 μg protein samples were loaded into sodium dodecyl sulfate polyacrylamide gel electrophoresis and blotted to PVDF membranes. The membranes were blocked for 1 h with 10% skim milk and probed with primary antibodies against THRAP3 (sc-133250, 1:1000, Clone name: F10, Santa Cruz), FTL (ab109373, 1:1000, Clone name: EPR5260, Abcam), GSS (ab133592, 1:1000, Clone name: EPR6563, Santa Cruz), GCLM (ab126704, 1:500, Clone name: EPR6667, Abcam), GCLC (ab207777, 1:500, Clone name: EPR20078, Abcam), TFRC (ab214039, 1:500, Clone name: EPR20584, Abcam), ACSL4 (ab155282, 1:500, Clone name: EPR8640, Abcam), GAPDH (AC001, 1:10000, ABclonal), β-actin (AC038, 1:10000, ABclonal) overnight at 4 °C. After incubation with the HRP Goat Anti-Rabbit IgG (AS014, 1:2000, ABclonal) or HRP Goat Anti-Mouse IgG (AS003, 1:2000, ABclonal) for 30 min, the membranes were developed by ECL Western Blotting Substrate (Solarbio) and quantified using Image J software. Uncropped and unprocessed blots were provided in Source Data file.
Statistical and reproducibility
Sample size of the mice was predetermined using power analyses. All experiments were conducted using at least 3 biological replicates. No data were excluded from the analyses. Randomization and blinding were used. The quantitative data are shown as mean ± standard deviation (SD). Statistical analyses were performed by two-tailed Student’s t test for two groups or one-way analysis of variance (ANOVA) followed by Tukey’s post hoc test for multiple groups using GraphPad Prism 8.0 software. Overall survival was evaluated using the Kaplan-Meier method. The correlation between THRAP3 and Exon14 skipped-GIT2 was evaluated by Pearson correlation analysis. P < 0.05 was considered as statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data supporting the findings of this study are available within the paper and its supplementary information files. Source data are provided with this paper. Furthermore, the RNA sequencing data generated in this study have been deposited in the SRA database under accession code PRJNA1348436. The proteomics data generated in this study have been deposited to the ProteomeXchange Consortium via the PRIDE partner (http://www.ebi.ac.uk/pride) repository with the dataset identifier PXD069724. The remaining data are available within the Article, Supplementary Information or Source Data file. Source data are provided with this paper.
References
Rajagopalan, A. et al. A gain-of-function p53 mutant synergizes with oncogenic NRAS to promote acute myeloid leukemia in mice. J. Clin. Invest. 133, e173116 (2023).
Austin, R. J. et al. Oncogenic drivers dictate immune control of acute myeloid leukemia. Nat. Commun. 14, 2155 (2023).
Kantarjian, H. et al. Acute myeloid leukemia: current progress and future directions. Blood Cancer J. 11, 41 (2021).
Wang, H. et al. TMIGD2 is an orchestrator and therapeutic target on human acute myeloid leukemia stem cells. Nat. Commun. 15, 11 (2024).
Ma, Z. et al. The Ferroptosis Landscape in Acute Myeloid Leukemia. Aging (Albany NY) 15, 13486–13503 (2023).
Stockwell, B. R. Ferroptosis turns 10: emerging mechanisms, physiological functions, and therapeutic applications. Cell 185, 2401–2421 (2022).
Zhang, J. et al. Ferroptosis in hematological malignancies and its potential network with abnormal tumor metabolism. Biomed. Pharmacother. 148, 112747 (2022).
Yu, X. et al. Inhibition of NRF2 enhances the acute myeloid leukemia cell death induced by venetoclax via the ferroptosis pathway. Cell Death Discov. 10, 35 (2024).
Yang, W. S. et al. Regulation of ferroptotic cancer cell death by GPX4. Cell 156, 317–331 (2014).
Lee, Y. & Rio, D. C. Mechanisms and Regulation of Alternative Pre-mRNA Splicing. Annu Rev. Biochem 84, 291–323 (2015).
Ghram, M. et al. The eukaryotic translation initiation factor eIF4E reprograms alternative splicing. EMBO J. 42, e110496 (2023).
Adamia, S. et al. A genome-wide aberrant RNA splicing in patients with acute myeloid leukemia identifies novel potential disease markers and therapeutic targets. Clin. Cancer Res 20, 1135–1145 (2014).
de Necochea-Campion, R., Shouse, G. P., Zhou, Q., Mirshahidi, S. & Chen, C. S. Aberrant splicing and drug resistance in AML. J. Hematol. Oncol. 9, 85 (2016).
Wong, A. C. H., Rasko, J. E. J. & Wong, J. J. We skip to work: alternative splicing in normal and malignant myelopoiesis. Leukemia 32, 1081–1093 (2018).
Zhang, J. et al. CircRAPGEF5 interacts with RBFOX2 to confer ferroptosis resistance by modulating alternative splicing of TFRC in endometrial cancer. Redox Biol. 57, 102493 (2022).
Liu, L. et al. HBV Enhances Sorafenib Resistance in Hepatocellular Carcinoma by Reducing Ferroptosis via SRSF2-Mediated Abnormal PCLAF Splicing. Int. J. Mol. Sci. 24, 3263 (2023).
Wahl, M. C., Will, C. L. & Luhrmann, R. The spliceosome: design principles of a dynamic RNP machine. Cell 136, 701–718 (2009).
Wang, Y. P., Ma, C., Yang, X. K., Zhang, N. & Sun, Z. G. Pan-cancer and single-cell analysis reveal THRAP3 as a prognostic and immunological biomarker for multiple cancer types. Front Genet 15, 1277541 (2024).
Lee, K. M., Hsu Ia, W. & Tarn, W. Y. TRAP150 activates pre-mRNA splicing and promotes nuclear mRNA degradation. Nucleic Acids Res 38, 3340–3350 (2010).
Yarosh, C. A. et al. TRAP150 interacts with the RNA-binding domain of PSF and antagonizes splicing of numerous PSF-target genes in T cells. Nucleic Acids Res 43, 9006–9016 (2015).
Feng, S. et al. Low-dose hypomethylating agents cooperate with ferroptosis inducers to enhance ferroptosis by regulating the DNA methylation-mediated MAGEA6-AMPK-SLC7A11-GPX4 signaling pathway in acute myeloid leukemia. Exp. Hematol. Oncol. 13, 19 (2024).
Zhang, B., Yang, L., Wang, X. & Fu, D. Identification of survival-related alternative splicing signatures in acute myeloid leukemia. Biosci Rep 41, BSR20204037 (2021).
Brivio, E. et al. Targeted inhibitors and antibody immunotherapies: Novel therapies for paediatric leukaemia and lymphoma. Eur. J. Cancer 164, 1–17 (2022).
Lyu, T., Li, X. & Song, Y. Ferroptosis in acute leukemia. Chin. Med J. (Engl.) 136, 886–898 (2023).
Yu, Y. et al. The ferroptosis inducer erastin enhances sensitivity of acute myeloid leukemia cells to chemotherapeutic agents. Mol. Cell Oncol. 2, e1054549 (2015).
Soula, M. et al. Metabolic determinants of cancer cell sensitivity to canonical ferroptosis inducers. Nat. Chem. Biol. 16, 1351–1360 (2020).
Vohhodina, J. et al. The RNA processing factors THRAP3 and BCLAF1 promote the DNA damage response through selective mRNA splicing and nuclear export. Nucleic Acids Res 45, 12816–12833 (2017).
Jonsson, J., Wang, L., Kajitani, N. & Schwartz, S. A novel HPV16 splicing enhancer critical for viral oncogene expression and cell immortalization. Nucleic Acids Res 52, 316–336 (2024).
Lapa, B. S. et al. AZD-7648, a DNA-PK inhibitor, induces DNA damage, apoptosis, and cell cycle arrest in chronic and acute myeloid leukemia cells. Int. J. Mol. Sci. 24, 15331 (2023).
Chen, C. J. et al. Metformin attenuates multiple myeloma cell proliferation and encourages apoptosis by suppressing METTL3-mediated m6A methylation of THRAP3, RBM25, and USP4. Cell Cycle 22, 986–1004 (2023).
Yu, W. et al. Sorting nexin 3 exacerbates doxorubicin-induced cardiomyopathy via regulation of TFRC-dependent ferroptosis. Acta Pharm. Sin. B 13, 4875–4892 (2023).
Hou, W. et al. Autophagy promotes ferroptosis by degradation of ferritin. Autophagy 12, 1425–1428 (2016).
Zhang, R. et al. Gamma-glutamylcysteine exerts neuroprotection effects against cerebral ischemia/reperfusion injury through inhibiting lipid peroxidation and ferroptosis. Antioxidants (Basel) 11, 1653 (2022).
Tang, X., Li, X., Zhang, D. & Han, W. Astragaloside-IV alleviates high glucose-induced ferroptosis in retinal pigment epithelial cells by disrupting the expression of miR-138-5p/Sirt1/Nrf2. Bioengineered 13, 8240–8254 (2022).
Rivera, O. D. et al. Alternative splicing redefines landscape of commonly mutated genes in acute myeloid leukemia. Proc. Natl. Acad. Sci. USA. 118, e2014967118 (2021).
Smith, M. A. et al. U2AF1 mutations induce oncogenic IRAK4 isoforms and activate innate immune pathways in myeloid malignancies. Nat. Cell Biol. 21, 640–650 (2019).
Lim, R. & Lappas, M. GIT2 deficiency attenuates inflammation-induced expression of pro-labor mediators in human amnion and myometrial cellsdagger. Biol. Reprod. 100, 1617–1629 (2019).
Siddiqui, S. et al. Genomic deletion of GIT2 induces a premature age-related thymic dysfunction and systemic immune system disruption. Aging (Albany NY) 9, 706–740 (2017).
Martin, B. et al. GIT2 Acts as a Systems-Level Coordinator of Neurometabolic Activity and Pathophysiological Aging. Front Endocrinol. (Lausanne) 6, 191 (2015).
Duan, B. et al. EGF-stimulated activation of Rab35 regulates RUSC2-GIT2 complex formation to stabilize GIT2 during directional lung cancer cell migration. Cancer Lett. 379, 70–83 (2016).
Premont, R. T., Claing, A., Vitale, N., Perry, S. J. & Lefkowitz, R. J. The GIT family of ADP-ribosylation factor GTPase-activating proteins. Functional diversity of GIT2 through alternative splicing. J. Biol. Chem. 275, 22373–22380 (2000).
Campbell, T. et al. Novel roles for HMGA2 isoforms in regulating oxidative stress and sensitizing to RSL3-Induced ferroptosis in prostate cancer cells. Heliyon 9, e14810 (2023).
Mazaki, Y. et al. Neutrophil direction sensing and superoxide production linked by the GTPase-activating protein GIT2. Nat. Immunol. 7, 724–731 (2006).
van Gastel, J. et al. The RXFP3 receptor is functionally associated with cellular responses to oxidative stress and DNA damage. Aging (Albany NY) 11, 11268–11313 (2019).
Ohrt, T. et al. Molecular dissection of step 2 catalysis of yeast pre-mRNA splicing investigated in a purified system. RNA 19, 902–915 (2013).
Jimenez, M. et al. Splicing events in the control of genome integrity: role of SLU7 and truncated SRSF3 proteins. Nucleic Acids Res 47, 3450–3466 (2019).
Urtasun, R. et al. Splicing regulator SLU7 preserves survival of hepatocellular carcinoma cells and other solid tumors via oncogenic miR-17-92 cluster expression. Oncogene 35, 4719–4729 (2016).
Schlenker, O. & Rittinger, K. Structures of dimeric GIT1 and trimeric beta-PIX and implications for GIT-PIX complex assembly. J. Mol. Biol. 386, 280–289 (2009).
Yu, B. et al. Erianin induces apoptosis of osteosarcoma cells through ferroptosis signaling pathway, and inhibits proliferation and migration of osteosarcoma cells. Cytotechnology 77, 156 (2025).
Doi, H. et al. Andrographolide drives dual apoptosis and ferroptosis via caspase-3 and FACL4 in T-ALL cell lines. Int. J. Hematol. 122, 843–855 (2025).
Li, M. & Liu, X. Ferulic acid/GRB2/NF-kappaB signaling pathway that alleviates ferroptosis-induced apoptosis of nucleus pulposus cells is a potential mechanism for intervertebral disc degeneration. J. Orthop. 65, 302–309 (2025).
Wu, Q. & Bazzini, A. A. Translation and mRNA Stability Control. Annu Rev. Biochem 92, 227–245 (2023).
Biasini, E. & Faccioli, P. Functional, pathogenic, and pharmacological roles of protein folding intermediates. Proteins 93, 1299–1307 (2025).
Salib, A. et al. MYCN and SNRPD3 cooperate to maintain a balance of alternative splicing events that drives neuroblastoma progression. Oncogene 43, 363–377 (2024).
Zhao, L. P. et al. beta-Elemene induced ferroptosis via TFEB-mediated GPX4 degradation in EGFR wide-type non-small cell lung cancer. J. Adv. Res. 62, 257–272 (2023).
Liu, Z. et al. SETDB1 interactions with PELP1 contributes to breast cancer endocrine therapy resistance. Breast Cancer Res 24, 26 (2022).
Liu, Y. et al. SLC25A21 correlates with the prognosis of adult acute myeloid leukemia through inhibiting the growth of leukemia cells via downregulating CXCL8. Cell Death Dis. 15, 921 (2024).
Wen, D., Xiao, H., Gao, Y., Zeng, H. & Deng, J. N6-methyladenosine-modified SENP1, identified by IGF2BP3, is a novel molecular marker in acute myeloid leukemia and aggravates progression by activating AKT signal via de-SUMOylating HDAC2. Mol. Cancer 23, 116 (2024).
Acknowledgements
This work was supported by National Natural Science Foundation of China (82270185) from Mh Y; Hunan Provincial Science and Technology Innovation Platform and Talent Program (2022RC3077) from Mh Y; Health Research Project of Hunan Provincial Health Commission (20232811, W20242008) from Mh Y; National Key Clinical Specialty Scientific Research Project (Z2023023) from Mh Y.
Author information
Authors and Affiliations
Contributions
D.W.: Conceptualization; Formal analysis; Visualization; Z.W.: Writing—Original Draft; Supervision; Methodology; S.L.: Investigation; Resources; Jing He: Formal analysis; Data Curation; H.Z.: Validation; Supervision; Pan Chen: Writing—Review & Editing; Funding acquisition. M.Y.: Writing - Review & Editing; Project administration.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Lynn Lee, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, D., Wu, Z., Liu, S. et al. THRAP3 promotes ferroptosis resistance in acute myelocytic leukemia through SLU7-mediated alternative splicing of GIT2. Nat Commun 17, 235 (2026). https://doi.org/10.1038/s41467-025-66931-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-66931-1