Abstract
To deepen the understanding of tissue-resident microbiota in colorectal cancer (CRC), we analyzed whole-genome and transcriptome data from 937 patients. We identified 249 genera and 361 species commonly present in both tumors and adjacent normal tissues (NATs). Distinct microbial signatures were associated with anatomical location, tumor stages, hypermutation status, mutations in CRC driver and DNA damage repair genes, as well as consensus molecular subtypes (CMSs). Notably, the presence of the pks island and elevated abundance of Enterobacteriaceae were linked to poor prognosis specifically in CMS2 tumors. Finally, microbial risk scores derived from taxa present in tumor or NATs predicted patient prognosis independently of established clinico-molecular factors. Prognostic taxa were strongly associated with tumor transcriptomic pathways related to hypoxia, immune response, and metabolic status. These findings revealed the heterogeneity of tissue-resident microbiota and their critical role in CRC progression, highlighting potential avenues for targeted intervention.
Similar content being viewed by others
Introduction
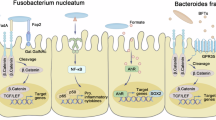
Colorectal cancer (CRC) is the third most prevalent cancer and the second leading cause of cancer-related mortality worldwide1. However, its substantial heterogeneity in clinicopathological characteristics, molecular profiles and prognosis, poses challenges in risk stratification and treatment optimization. Beyond host factors, the gut microbiome, exemplified by Fusobacterium nucleatum (Fn), enterotoxigenic Bacteroides fragilis (ETBF) and pks+ Escherichia coli (E. coli), actively participates in CRC carcinogenesis through cellular invasion, immune modulation, and DNA damage induction2,3.
Despite the technological advances, several challenges persist in the metagenomics field. Notably, the low microbial biomass of tumor tissues complicates the removal of putative contaminants and human-derived sequences, which can profoundly impact taxonomic annotation and downstream conclusions. Moreover, many prior studies often lack sufficient sequencing depth, negative controls, or standardized analysis pipelines, limiting reproducibility and cross-cohort comparability4,5. Recent reevaluations of tumor microbiota, including the retractions of earlier studies6,7,8, have underscored the importance of stringent contamination control and analytical frameworks that prioritize specificity over sensitivity for studying low-biomass samples.
While key carcinogenic bacteria have been identified, a detailed understanding of the CRC tissue-resident microbiome remains underexplored, including its anatomical distribution, interactions with host clinicopathological and genomic features, and its potential impact on patient prognosis.
Here, we conducted in-depth analyses of whole genome sequencing (WGS) data from 937 patients with CRC from a large prospective longitudinal study of primary cancers in Sweden9, to unravel the heterogeneity of the tissue-resident microbiome in tumors and normal adjacent tissues (NATs) and assess its prognostic implications.
Results
Identification of tissue-resident microbiota in CRC
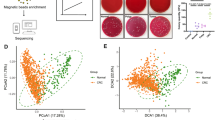
We implemented a rigorous pipeline (see Methods for detail) to detect, decontaminate, and quantify tissue-resident microbiota using high-depth WGS (~52× coverage) data from 1412 tissues (937 tumors and 475 NATs) and 462 blood samples from 937 treatment-naive patients enrolled in the U-CAN cohort9, including 775 from Uppsala University Hospital (UU) and 162 from Umeå University Hospital (UM) (Fig. 1a and Supplementary Data 1). Briefly, sequencing data were first aligned to the human genome (GRCh38) using Bowtie2, followed by reclassification of remaining reads using Kraken2 against a custom database including human (GRCh38 and T2T-CHM13v2.0) and microbial genomes (4644 from the UHGG catalog, and 17 Treponema species from NCBI). This two-step host removal pipeline ensured stringent exclusion of human reads in low-biomass samples. Subsequently, stringent quality-control filters were applied to remove putative contaminants (Supplementary Data 2A) and taxa with low prevalence and coverage. This resulted in 249 genera and 361 species, representing the CRC tissue-resident microbiota (Methods; Supplementary Fig. 1a and Supplementary Data 2B). To mitigate potential biases of different DNA extraction methods on microbial composition between centers, we designated UU as the discovery cohort, and UM as the internal validation cohort (Methods; Fig. 1a and Supplementary Fig. 1a−c). The AC-ICAM cohort (n = 246)10, with patient demographics and clinicopathological characteristics similar to the U-CAN cohort (Supplementary Data 1), was used as an external validation cohort.
a Schematic representation of the microbiome study design of 937 CRC patients from the Uppsala-Umeå Comprehensive Cancer Consortium (U-CAN) cohort. Patients included at Uppsala (UU, n = 775) and Umeå (UM, n = 162) University Hospitals served as discovery and internal validation cohort, respectively. Patients with primary colon cancer from the AC-ICAM (Atlas and Compass of Immune-Cancer-Microbiome Interactions in Colon Cancer) study (n = 246) served as external validation cohort. Tissue-resident microbes were identified using non-human reads from whole-genome sequencing (WGS) data, and were associated with host genomic alterations, gene expressions and patient survivals. Illustration elements (colon schematic and human outline silhouette) adapted from Servier Medical Art (https://smart.servier.com), CC BY 3.0. b Number of Kraken2-mapped prokaryotic reads from tumor tissues and paired blood samples (n = 462 pairs). c The proportions and taxonomic assignments of microbial reads classified as contaminants in tissue and paired blood (n = 462) samples from UU. d Number of annotated reads (left) and average genome coverage (right) of tissue-resident species across tumors (n = 775), normal adjacent tissues (NATs; n = 313) and blood (n = 462) samples from the UU cohort. e Average genome coverage (Y-axis) and number of mapped reads (X-axis) of tissue-resident prokaryotic species in tissue (n = 1088) and blood samples from the UU cohort. Each dot represents a different species. The name of the top 10 species by average Kraken2-mapped reads across tissue samples are shown. f Number of non-human reads from tumor tissue samples of the UU (n = 775), UM (n = 162) and AC-ICAM (n = 246) cohorts in the different processing steps. g Left: The average genome coverage of tissue-resident species in tumor tissue samples from the UU (n = 775), UM (n = 162) and AC-ICAM (n = 246) cohorts. Right: The top 50 most-covered species identified across three tumor datasets. In b, d, f, g, p-values were calculated using two-sided Wilcoxon rank-sum test. In all boxplots, boxes represent the interquartile ranges (IQRs) between the first and third quartiles, the center line represents the median, and the whiskers extend 1.5 times the IQR from the top and bottom of the box.
Consistent with the established notion that solid tissues harbor more abundant and diverse microbes than matched blood samples11, UU tumors had significantly higher prokaryotic read counts per sample (median 347,957) compared to matched blood samples (median 7097; Fig. 1b, and Supplementary Fig. 1a). Blood samples had a notably higher proportion of reads identified as putative contaminants (64.9% versus 4.8% of total prokaryotic reads in tumors), with the majority (84.2%) derived from Proteobacteria. After decontamination, only a limited microbial signal was retained in blood samples (median mapped reads= 960; median valid species=1, defined as taxa with ≥100 reads) (Fig. 1c and d, Supplementary Fig. 1b and Supplementary Data 2C, D). In contrast, tissues exhibited substantially better microbial genome coverage, enabling confident detection of previously reported CRC-related species such as Bacteroides fragilis, Prevotella nigrescens, Escherichia coli, Alloprevotella tannerae, and Fn subspecies animalis clade 2 (Fna C2)12,13, as well as gut commensals like Bacteroides dorei, Bacteroides uniformis, Prevotella copri, and Faecalibacterium prausnitzii (Fig. 1e).
Employing identical pipelines, we achieved highly consistent detection of tissue-resident microbes in AC-ICAM tumors, comparable to UU and UM tumors in terms of prokaryotic read counts, detected taxa, and genome coverage. Of note, 27 of the top 50 most-covered species overlapped across the three tumor datasets (Fig. 1f, g). These findings demonstrate the validity and reliability of our approach in identifying the tissue-resident microbiome in CRC.
Compared to paired WGS samples, the whole-transcriptome sequencing (WTS) samples detected much fewer species and reduced genome coverage, making accurate quantification of active microbes challenging (Supplementary Fig. 1d−f). Therefore, we focused solely on associating WGS-derived tissue-resident microbiota with CRC characteristics.
CRC tissue-resident microbial composition varies between NAT and tumor tissues, and across anatomical locations
With a robust framework for identifying and validating tissue-resident microbiota, we investigated variations in the tissue-resident microbial composition related to host factors. Tissues from the left colon and rectum were consolidated as left-sided colon due to their similar microbial composition, which distinctly differed from that of the right-sided colon tissues (Supplementary Fig. 2a−c). Notably, tumor location (right- vs. left-sided) emerged as the strongest covariate associated with tissue-resident microbiota in both tumors and NATs (Bray-Curtis dissimilarity, D-MANOVA, P = 1.19E-32; Supplementary Data 3), prompting our initial focus on location-specific microbial composition.
In the UU cohort, both right-sided tumors and NATs exhibited significantly higher prokaryotic read counts and lower alpha diversity compared to left-sided tissues (MaAsLin2, P < 0.05; Fig. 2a and b). Significant differences in the relative abundances of 167 microbial taxa (63 genera and 104 species) between the right- and left-sided tissues were consistently observed in tumors and NATs (MaAsLin2, BH-adjusted P < 0.05; Fig. 2c and Supplementary Data 4). Left-sided tissue samples were enriched with taxa from the Enterobacteriaceae (e.g., E. coli), Akkermansia (e.g., A. muciniphila), and Porphyromonas (e.g., P. somerae). Conversely, right-sided samples were predominantly enriched with Firmicutes members belonging to Lachnospiraceae (e.g., Blautia and Roseburia) and Ruminococcaceae (e.g., Faecalibacterium) (Fig. 2c, BH-adjusted P < 0.05). We confirmed these anatomical-location differences in mapped prokaryotic read counts, alpha diversity, and taxa abundances in both validation cohorts (Fig. 2a−c and Supplementary Fig. 2d−f). Specifically, 83.9% and 87.8% of location-specific taxa in UM and AC-ICMA tumors, respectively, were in concordance with those identified in UU tumors (MaAsLin2 BH-adjusted P < 0.05; Supplementary Fig. 2f and Supplementary Data 4). A random forest (RF) classifier trained with 5-fold cross-validation within UU samples achieved high performance in distinguishing right- from left-sided tissues across all datasets (AUC ranged from 86% to 91% for UU; and from 80% to 85% for validation cohorts; Fig. 2f and Supplementary Fig. 2g). We identified consistent location-specific differences in 28 taxa across at least two independent fecal metagenomic datasets of CRC patients. In the Yachidas_2019 cohort14, an RF model trained on fecal samples achieved an AUC of 0.71 in distinguishing right- from left-sided tumors (Supplementary Fig. 2h, i). These results suggest that fecal microbiota can partially reflect anatomical location microbial differences.
a Comparison of prokaryotic reads number between left- and right-sided tissue samples. Two-sided P-values were calculated with Wilcoxon rank-sum test. b Genus level microbial diversity in left- vs. right-sided tissue samples. Two-sided P-values were calculated with MaAsLin2, with adjustment for age, sex, and prokaryotic read count. For a and b: UU tumors: n (left)=386; n (right)=389; for UU NATs: n (left)=145, n (right)=168. For UM: n (left)=73, n (right)=89. c Enrichment of tissue-resident species between anatomical locations. Species displaying consistent differences between anatomical locations in both tumors (UU_T, n = 775; UM_T, n = 162) and NATs (UU_N, n = 313; UM_N, n = 162). MaAsLin2, BH-adjusted P < 0.05 (two-sided test). Green, right-side enriched specie; yellow, left-side enriched species. Colored outer bars indicate the corresponding phyla of presented species. d Genus-level microbial diversity in paired tumor-NAT samples across anatomical locations: all pairs (UU, n = 313; UM, n = 162), right-sided pairs (UU, n = 168; UM, n = 89), and left-sided pairs (UU, n = 145; UM, n = 73). Two-sided P-values were calculated with MaAsLin2. e Enrichment of tissue-resident species displaying significant and consistent differences between tumors and paired NATs in UU (168 UU_R and 145 UU_L) and UM (89 UM_R and 73 UM_L). MaAsLin2 (two-sided test), BH-adjusted P < 0.05 for UU, and P < 0.05 for UM (two-sided test). Blue, NAT-enriched; red, tumor-enriched. Colored outer bars indicate location-enriched species (yellow, left-sided; green, right-sided). f Two-sided 95% confidence interval for the area under the ROC curve (AUC) showing the ability of tissue-resident microbiota in distinguishing right from left-sided samples. Microbial based random forest (RF) models were generated using UU tumors (n = 775) with 5-fold cross validation (CV), and tested on UU_N (NATs, n = 313), UM_T (tumors, n = 162) and UM_N (NATs, n = 162). g Performance of RF models in distinguishing tumors from paired NATs. The training model was established on paired tissues of UU cohort (n = 313 pairs, 5-fold CV), and the testing set was 162 paired UM tissues. For the boxplots, boxes represent the interquartile range (IQR, Q1–Q3), the central line represents the median, and whiskers extend to 1.5 times of IQR.
Additionally, right-sided enriched taxa exhibited more pronounced associations with right-sided specific pathways in both NATs and tumors, consistently identified across the UU and validation cohorts (Supplementary Fig. 2j). This may reflect the elevated hypoxia often observed in the right-sided colon9, which may induce glycolysis, lipid metabolism, and extracellular matrix (ECM) remodeling15.
Compared to matched NATs, UU tumors exhibited significantly reduced alpha diversity and altered microbial composition, with 60.3% (140/232) of the differential taxa replicated in UM (MaAsLin2, BH-adjusted P < 0.05, Fig. 2d-e and Supplementary Data 5A). More than half of the tumor-enriched taxa (47/85, 55.3%) demonstrated significant anatomical location-specific differences in the UU cohort (Supplementary Data 5B). For instance, Porphyromonas spp. and several Enterobacteriaceae taxa were enriched in left-sided tissues, while A. tannerae, Prevotella intermedia and Clostridium perfringens were more abundant in right-sided tissues (Fig. 2e and Supplementary Data 5A). Noteworthy, established gastrointestinal cancer-associated bacteria12,16,17,18,19, including four Fn subspecies (Fnn, Fna C2, Fn. vincentii, and Fn. polymorphum), Streptococcus spp. (e.g., S. anginosus), Clostridium spp. (e.g., C. bolteae, C. citroniae, C. clostridioforme, and C. symbiosum)17, B. fragilis, Hungatella hathewayi, and Parvimonas micra, were consistently enriched in UU and UM tumors, while displaying no significant abundance differences between anatomical locations (MaAsLin2, BH-adjusted P > 0.05; Fig. 2e). Among the NAT-enriched taxa, a large portion (n = 106, 72.1%) were Firmicutes members, and 35.4% (n = 52) were enriched in right-sided tissues, including Blautia and Faecalibacterium (Fig. 2e and Supplementary Data 5A).
Furthermore, the differences in microbial composition between NATs and tumors were more pronounced in the right-sided tissues (Fig. 2d and Supplementary Data 5B). Consistently, the RF classifier trained with UU samples (5-fold CV) achieved higher accuracy in distinguishing tumors from NATs in right-sided cases (AUC = 82%) than in left-sided cases (AUC = 75%) for UM samples (Fig. 2g).
CRC tissue-resident microbiota is associated with somatic mutations
Next, we classified hypermutated (HM) UU tumors as those with a tumor mutational burden (TMB) of >23.2 mutations/Mb9. HM tumors (n = 180) were more prevalent in older (median 76 years vs. 71 years) and females patients compared to non-hypermutated (nHM, n = 595) tumors (Fig. 3a). As expected, HM was more prevalent in right-sided compared to left-sided colon, across all cohorts (Fig. 3b).
a Prevalence of somatic mutations in tumors of the UU cohort. The dotted line indicates the cutoff for hypermutated (HM) and non-hypermutated (nHM) tumors (23.2 mutation/Mb). Colored columns represent the samples (n = 775) annotated with HM or nHM, tumor locations, sex, and age groups. Two-sided Chi-square test P-values to show relationships between groups with mutational status and clinical phenotypes. b Discovery and validation of relationships between total mutations and 35 mutational status-associated taxa in UU (n = 775), UM (n = 162) and AC-ICAM (n = 167) cohorts. Top: Horizontal bars show distribution of HM and nHM samples between right- and left-sided tumors. Bottom: Differences in abundance of 35 taxa between HM and nHM identified using MaAsLin2. Associations between bacterial abundances and total number of single nucleotide variants (SNVs) were shown using Spearman’s correlation coefficients (rho). *, BH-adjusted P < 0.05; #, P < 0.05. c Tumor-resident taxa significantly associated with hypermutation status in the UU cohort. The X axis represents significance of differences between HM and nHM tumors, and the Y axis shows that between paired tumors and NATs. Dotted lines indicate BH-adjusted P = 0.05 for enrichment in NATs (lower) or tumors (upper), and enrichment in nHM (left) or HM (right) tumors. Circle sizes indicate average relative abundance of each taxon across 775 UU tumors. d Differences in microbial abundances between tumors with mutations (Mutant) or wild-type (WT) in individual driver genes or DNA damage response and repair (DDR) genes. Top bars, number of taxa consistently associated with gene mutations in UU and at least one validation cohort. Right bars, number of genes consistently associated with tumor-enriched taxa in UU and at least one validation cohort. Red, enrichment in Mutant; blue, enrichment in WT. In total, mutations in 41 divers and 18 DDR genes displayed significant associations with microbes in UU (BH-adjusted P < 0.05), with triangles indicating consistent observations validated in UM and circles in AC-ICAM (P < 0.05). For b−d, two-sided P values from MaAsLin2 models were determined with adjustment for age, sex, location, and Kraken2-mapped prokaryotic read count.
We identified 35 tumor-enriched taxa with significant differences between HM and nHM tumors after adjusting for age, sex, anatomical location and prokaryotic read counts. Notably, 26 of tumor-enriched taxa were enriched in HM tumors, including the four Fn subspecies and multiple oral-derived taxa18 such as Treponema, A. tannerae, Selenomonas, Parvimonas, P. stomatis and Solobacterium moorei (BH-adjusted P < 0.05 in UU; Fig. 3b and c). This enrichment pattern persisted in right- and left-sided tumors (Supplementary Data 6A), further confirming their independence from tumor location. The majority of HM-enriched taxa were positively associated with the number of single-nucleotide variants (SNVs) and were more abundant in patients with mutations in CRC drivers or DNA damage response and repair (DDR) genes compared to their wild-type (WT) counterparts (BH-adjusted P < 0.05 in UU; P < 0.05 in UM/AC-ICAM; Fig. 3b and d, and Supplementary Data 6A and B). Among tumor-enriched taxa, Treponema was the top genus associated with CRC drivers or DDR gene mutations, followed by Fn subspecies. Importantly, the genus Treponema was predominantly composed of oral-resident species, such as T. denticola (Supplementary Data 6E), and was previously detected in orodigestive tumors20. The CASP8 mutations, found in about 7% of U-CAN patients, were associated with the highest number of HM-enriched taxa (Fig. 3d and Supplementary Fig. 3a and b). Those genetic defects are known to increase susceptibility to microbial infection21. Similar patterns were observed in genes known for higher mutational frequencies in MSI tumors9, such as ACVR2A and BRAF (epithelial-to-mesenchymal transition)22,23, CD58 and BCL9 (tumor immune microenvironment modulation)24,25, ARID1A and ARID1B (hypermethylation)26, and MSH3, MSH6, and MLH3 (DNA mismatch repair)27 (Fig. 3d and Supplementary Data 6B). Conversely, APC mutations or loss of heterozygosity (LOH) events in tumor suppressor genes such as FLCN, MAP2K4 and TP53 were associated with lower abundances of HM-enriched taxa, especially Fusobacterium spp. (Fig. 3d, Supplementary Fig. 3c and d and Supplementary Data 6C).
Nine tumor-enriched taxa, largely from Enterobacteriaceae, including E. coli, were enriched in nHM tumors in UU but exhibited limited consistency across the validation cohorts (Fig. 3b and c), likely reflecting the high degree of tumor heterogeneity within this subgroup. Nonetheless, we detected the pks island in 17.4% of the UU tumors (135/775). The relative abundances of E. coli and pks island were strongly correlated, and both were enriched in left-sided and nHM tumors (Supplementary Fig. 4a and b and Supplementary Data 7). Three mutational signatures were overrepresented in pks+ tumors compared to pks- tumors, including the previously established colibactin-induced signature SBS8828, and two of unknown aetiology SBS93 and ID14 (P < 0.05, Supplementary Fig. 4c and d and Supplementary Data 8). The latter two were highly correlated (Spearman’s rho=0.69, Supplementary Fig. 4e). Additionally, SBS93 shared a similar mutational pattern with SBS88, characterized primarily by T > N single-base substitutions, and ID14 was dominated by 1 bp insertions (Supplementary Fig. 4f).
Despite the significant associations between microbiota and genomic variations observed in tumors across three CRC cohorts, a substantial proportion of CRC-associated taxa, including Fusobacterium spp., were consistently enriched in both HM and nHM tumors compared to their matched NATs (Supplementary Data 6D). This indicates that microbial dysbiosis occurs during CRC tumorigenesis, partially independent of tumor mutational status.
CRC molecular subtype-dependent microbial profiles with prognostic implications
The Consensus Molecular Subtype (CMS) is a gene expression-based classification that categorizes CRC by shared molecular signatures. Emerging evidence has reported enrichment of Fn in a CMS-dependent manner18,29. However, the CMS-dependent prognostic implications of overall CRC-enriched microbes remain inconclusive.
To address this gap, we characterized 85 tumor-enriched taxa in relation to CMS subtypes based on gene expression of U-CAN tumors (Supplementary Fig. 5a). Consistent with findings from the TCGA-COADREAD and AVANT cohorts18,29, four Fn subspecies and multiple CRC-associated oral taxa (35.3%, 30 out of 85) were significantly enriched in CMS1 tumors compared to CMS2-4 tumors (BH-adjusted P < 0.05, Fig. 4a and c), mirroring their enrichment in HM tumors compared to nHM tumors. While no tumor-enriched taxa were found to be specifically enriched in CMS2, CMS3 or CMS4 tumors (Fig. 4c, Supplementary Data 9, BH-adjusted P > 0.05). CMS2 cases (32%, 224 out of 709) were younger and had more left-sided tumors (73%), nHM tumors (99.5%), and a higher prevalence (25%) of the pks colibactin gene island (pks + ) than other subtypes, which may collectively influence the genetic variations30, expression patterns and prognosis of CMS2 tumors (Fig. 4a and b). Notably, the two newly identified pks+ associated mutational signatures in this study, SBS93 and ID14, were significantly enriched in patients with CMS2 tumors (Supplementary Fig. 5b). Also, higher levels of Enterobacteriaceae taxa (e.g., Citrobacter, Enterobacter, Escherichia and E. coli) were associated with shorter overall survival (OS) and recurrence-free survival (RFS) exclusively in overall CMS2 patients (adjusted HROS for E. coli = 1.82, 95% CI = 1.26−2.63, adjusted HRRFS for E.coli = 1.46, 95% CI = 1.08−1.96; Fig. 4d), and those with stage II or III CMS2 tumors (Supplementary Data 10). Moreover, within the CMS2 subtype, pks+ patients exhibited significantly worse survival compared to pks- subjects (UU: adjusted HROS = 2.77, 95% CI = 1.45−5.29, P = 0.002; adjusted HRRFS = 3.22, 95% CI = 1.82−5.69, P = 5.73e-5; Fig. 4e). Importantly, this CMS2-specific pattern was independent of age, anatomical location, tumor stage and grade, and was evident in validation cohorts, where higher Enterobacteriaceae abundance was associated with shorter OS (adjusted HROS UM+AC-ICAM = 1.58; adjusted HROS AC-ICAM-16S = 1.97; Fig. 4f and Supplementary Data 10). Conversely, Fusobacterium, Fnn, Fna C2, and Fnv were solely prognostic for CMS4 tumors (adjusted HROS for Fusobacterium = 1.77, 95% CI = 1.28−2.44, adjusted HRRFS for Fusobacterium = 1.53, 95% CI = 1.14−2.04; Fig. 4d), a subtype-dependent pattern previously observed in other CRC cohorts29.
a A total of 709 (91.48%, 709/775) tumors from the UU cohort count by CMS subtypes and the proportions of samples by anatomical locations, mutational, and the pks colibactin gene island (pks + ) status within each CMS subtype. b Age distribution among CMS subtypes for the UU cohort. Two-sided Wilcoxon rank-sum test. c Taxa enrichment in CMS1 versus other CMS subtypes. Left: color bar, phyla of CMS1 enriched taxa. Heatmap, taxa abundances between HM versus nHM, and CMS subtypes, colored by effect sizes from MaAsLin2, with adjustment for age, sex, anatomical location, and prokaryotic read counts. *, BH-adjusted P < 0.05. d Associations between tumor-enriched taxa and survival in CMS2 (upper, n = 224) and CMS4 (lower, n = 255) stage I-III patients. Adjusted HRs with two-sided P-value were calculated by multivariable Cox regression. Covariates include age, anatomical locations, tumor stage, grade, postoperative treatment, and prokaryotic read counts. Error bars represent 95% CI for HR. *, BH-adjusted P < 0.05. e CMS-dependent associations between presence of pks and survival in stages I-III patients from UU. Adjusted HRs and two-sided P values were calculated by multivariable Cox proportional hazard regression. f CMS2-dependent associations between the family Enterobacteriaceae and OS in stage I-III patients. Adjusted HRs with P-value for CMS2 and others were calculated by multivariable Cox regression (two-sided test). Covariates include age, anatomical locations, and tumor stage. Error bars represent 95 % CI for HR. g CMS2-dependent associations between UU tumor hypoxia levels and relative abundances of Escherichia coli (left) or presence of pks gene island. Left: P values calculated with two-sided partial Spearman’s correlation. Right: P values calculated with two-sided Wilcoxon rank-sum test. h CMS4-dependent associations between relative abundances of Fusobacterium spp. and expressions of CRC-related pathways and immune cells in UU cohort. *, BH-adjusted P < 0.05 (Two-sided partial Spearman’s correlation). In g, h, two-sided partial Spearman’s correlation adjusted for sex, age, tumor location and prokaryotic read counts. In b, g, p-values were calculated using two-sided Wilcoxon rank-sum test. Boxplots show the IQR, median (center line), and 1.5 times of IQR whiskers.
Notably, the associations between oncogenic pathways and microbes also tended to be specific to CMS subtypes and their prognostic relevant taxa. In CMS2 tumors, E. coli abundance was positively associated with hypoxia levels (Spearman’s rho=0.176, P = 0.0083; Fig. 4g). Correspondingly, only CMS2 pks+ tumors showed significantly higher hypoxia levels, which was not observed in other subtypes (Wilcoxon rank-sum test; Fig. 4g). We further observed significant heterogeneity in associations between Fn subspecies and the tumor immune microenvironment (TIME) across different CMS subtypes. Only in CMS4 tumors, Fn subspecies were positively associated with vascular endothelial growth factor (VEGF) and glutamine metabolism pathways, but were consistently and negatively associated with immune checkpoints (PD1 activation), complement activation pathways, and multiple immune response markers (e.g., T-lymphocytes scores, expression of CD8 + T cells and macrophages M2, and signature of CD8 + T effector)31 (Partial Spearman’s rank correlation, BH-adjusted P < 0.05; Fig. 4h, Supplementary Fig. 5a and Supplementary Data 11).
CRC tissue-resident microbes predict patient survival
Prognostic taxa were identified separately in UU tumors and NATs, and their independent associations with prognostic outcomes, including OS and RFS, were assessed in stage I-III patients from UM and AC-ICAM cohorts (Fig. 5a).
a Stage I-III patients and tissues from UU, UM, and AC-ICAM cohorts. Colon schematic adapted from Servier Medical Art (https://smart.servier.com), CC BY 3.0. b Heat tree of taxa associated with OS in NATs and tumors from the UU cohort. Significance was defined as BH-adjusted P < 0.05 in tumors and two-sided P < 0.01 in NATs using multivariate Cox models. c Overlaps of prognostic taxa identified in UU tumors and NATs. d Relative abundances of prognostic taxa between tumor vs. NATs, HM vs. nHM tumors and left- vs. right-sided samples in UU, colored by effect sizes from MaAsLin2. Diamonds denote prognostic taxa. Circles denote differential taxa between groups (BH-adjusted P < 0.05, two-sided test). e The associations between Akkermansia abundance and OS in NATs and tumors from the UU cohort. The optimal cutoff (black line) for Akklow (Akk <0.05) and Akkhigh groups was determined by grid search maximizing HR in NATs. f Kaplan–Meier curve for the estimated OS for Akklow and Akkhigh groups in UU NATs. g Forest plot displaying adjusted HRs and two-sided P-values between Akklow and Akkhigh groups for OS in NATs from the UU, UM and AC-ICAM (16S data) cohorts, and RFS in NATs from UU and UM cohorts, estimated by multivariate Cox models. Horizontal error bars represent 95 % CIs of the HR. h Spearman’s correlation between MRS-N and MRS-T across 288 paired stage I-III tissue samples from UU cohort. i. Kaplan–Meier curves for low- and high MRS groups (defined by median). Adjusted HRs and two-sided P-values for continuous and categorical MRS were determined by multivariate Cox models. j Concordance index (C-index) of combinatorial prognostic factors. Parenthesis indicates the C-index increase after adding MRS-T and MRS-N to the reference model. k Associations between prognostic taxa and expressions of CRC-related pathways and genes in tumors. Two-sided partial Spearman’s correlation. Red, positive; blue, negative. Significance was defined as BH-adjusted P < 0.05 in UU and P < 0.05 in UM and AC-ICAM.
In the UU cohort, 122 prognostic taxa in tumors (BH-adjusted P < 0.05) and 34 in NATs (P < 0.01) were identified after adjusting for clinicopathological factors (Supplementary Fig. 6a), with 21 taxa shared between both tissue types (Fig. 5b, c and Supplementary Data 12). The majority of tumor-derived prognostic taxa (90.2%, n = 110) were associated with prolonged OS, with over half enriched in right-side tissues (62.3%, n = 76) and primarily belonging to Lachnospiraceae (Blautia) and Ruminococcaceae (e.g., Ruminococcus and Faecalibacterium). In contrast, only a few favorable taxa in tumors were enriched in left-sided tissues, such as Bifidobacterium infantis and B. adolescentis (Fig. 5d). Most favorable taxa were consistently associated with better OS across all CMS subtypes (adjusted HRs <1, Supplementary Fig. 6b); and 15 taxa were consistently linked to favorable prognosis in both tumors and NATs, including known butyrate producers such as Roseburia inulinivorans, Faecalibacterium prausnitzii, and Fusicatenibacter saccharivorans (Fig. 5d and Supplementary Fig. 6c). In contrast, 6 taxa, including Clostridium bolteae, C. lavalense, C. symbiosum, and C. scindens were significantly associated with worse OS in both tissue types (Fig. 5b and d and Supplementary Data 12). Additionally, 26 of the tumor-derived prognostic taxa were validated in at least one of the validation cohorts (Supplementary Fig. 6c and d), including 19 taxa associated with favorable prognosis (e.g., Lachnospiraceae spp., and Faecalibacterium spp.), and 7 with unfavorable prognosis (e.g., Clostridium spp. and Eggerthella lenta).
Moreover, the abundance of 31 prognostic taxa differed significantly between stage I and stage IV UU tumors. In stage IV tumors, there was a notable depletion of taxa associated with favorable prognosis and an enrichment of those linked to poor prognosis, including Clostridium spp., suggesting their indicative roles in CRC progression (Supplementary Fig. 6e and Supplementary Data 13).
Unexpectedly, in stage I-III UU patients, a higher abundance of A. muciniphila in NATs was significantly associated with poor prognosis (P = 0.0074, adjusted HROS = 1.36; Fig. 5b and d), also seen with the genus Akkermansia (Akk) (Supplementary Data 14). An optimal cutoff of 0.05 for Akk abundance in NATs revealed that the median OS was 2.68 years for Akk high patients (≥0.05, n = 29) compared to 5 year for the Akk low patients (<0.05, n = 259) (unadjusted HROS = 2.04, P = 0.012; Fig. 5e and f). This Akk-specific pattern was confirmed in NATs from UM (adjusted HROS = 2.17; P = 0.146; adjusted HRRFS = 2.19; P = 0.146) and AC-ICAM cohorts (adjusted HROS = 2.83, P = 0.007) using the same abundance cutoff (Fig. 5g). Host characteristics were balanced between Akk high and Akk low groups in UU stage I-III patients, except for a higher proportion of left-sided tumors in the Akk high group (72% vs. 42%; Supplementary Data 14).
We calculated individual microbial risk scores for tumors (MRS-T) and NATs (MRS-N) based on prognostic taxa (Methods, Supplementary Data 15). The two scores were strongly correlated (Spearman’s rho= 0.68; Fig. 5h), and both were independently associated with worse survival in UU (adjusted HROS for MSR-T = 1.81; adjusted HRRFS for MSR-T = 1.58, adjusted HROS for MRS-N = 2.31; adjusted HRRFS for MRS-N = 1.98), and validated in the UM and AC-ICAM cohorts (Fig. 5i and Supplementary Data 16). MRS-T and MRS-N both enhanced the prognostic models based on individual host clinicopathological factors, including age, sex, BMI, tumor location, grade, stage, hypermutation status, MSI status, and CMS subtype (Supplementary Fig. 7a). Compared to the reference model using the host factors alone, MRS-T and MRS-N increased the consistency index (C-index) by 0.046 and 0.079, respectively (Fig. 5j). Similar improvements were observed in the validation cohorts (Fig. 5j). Using 249 paired tissue 16S rRNA gene amplicon datasets from stage I-III AC-ICAM patients, we identified 12 genera (including Akkermansia) for MRS-Namplicon and 14 for MRS-Tamplicon, and confirmed their prognostic performance (adjusted HROS for MRS-N = 1.98; adjusted HROS for MRS-T = 1.37) (Supplementary Fig. 7b−d and Supplementary Data 17). Additionally, the prognostic value of MRS-T was further observed in 98 U-CAN patients with stage IV tumors, independent of host factors (P < 0.05, adjusted HROS for stage IV = 2.01; adjusted HRRFS for stage IV = 2.04) (Supplementary Fig. 7e).
Further analyses revealed that a large portion of favorable taxa were significantly associated with the downregulation of oncogenic pathways including hypoxia, nuclear factor kappa B (NF-kB), and tumor necrosis factor alpha (TNF-α) signaling (Fig. 5k). These taxa were also linked to lower levels of proinflammatory cytokines and chemokines, such as IL1, IL6, CSF2, CSF3, CCL3, CCL4, CCL20, CXCL8, ICAM1, and PTX3. Additionally, pathways related to immune function (e.g., interleukin signaling and toll-like receptor cascades), cancer-related signal transduction, and the circadian clock were also downregulated in tissues enriched with these taxa (BH-adjusted P < 0.05; Fig. 5k). Conversely, unfavorable Clostridium taxa were associated with higher levels of hypoxia, IL6, and MAPK1/3 activation. These microbial associations with tumor gene expressions were broadly confirmed in the validation cohorts (Fig. 5k and Supplementary Data 18). Thus, in all cohorts, patients identified with high MRS-T values (above the median) exhibited lower abundances of favorable taxa and higher expressions of proinflammatory genes compared to patients with low MRS-T values (Supplementary Fig. 8 and Supplementary Data 19).
Discussion
Gut microbial dysbiosis is implicated in the carcinogenesis, treatment response, and prognosis of CRC2,10. Using in-depth bulk sequencing data from the U-CAN9 and the AC-ICAM cohorts10, along with clinical follow-up, we performed a thorough profiling of the presence and relevance of tissue-resident microbiota in primary CRC. We characterized distinct microbial composition across anatomical locations and identified consistent associations between microbial signatures, tumor genomics variations, transcriptome-based CMSs, and oncogenic pathways across UU, UM and AC-ICAM cohorts. We have also developed microbial risk scores (MRS-T and MRS-N) based on prognostic microbes, with the ability to predict survival independently of the host clinicopathological factors.
In view of the recent debate over the validity and reproducibility of microbiome analysis in low-biomass tumors, we developed an analytical workflow designed to prioritize specificity over sensitivity in microbial detection. To minimize potential contamination, we applied a two-step host read removal strategy32, used paired blood samples as negative controls11, and adopted a published blacklist of common reagent contaminants4. We further required sufficient mapped reads, prevalence and genome coverage for taxa inclusion, a principle also highlighted in a recent cancer microbiome benchmarking study based on bulk WGS datasets8. That study further underscored the importance of stringent decontamination to mitigate sequencing noise in cancer microbiome analyses8. Additionally, species-level annotation using only high-quality human gastrointestinal–derived genomes from the UHGG collection markedly reduced false-positive assignments from unrelated habitats. Together, these measures enhanced taxonomic resolution and cross-cohort consistency in microbial profiling across CRC tissues, particularly regarding location differences. As the field moves toward standardized best practices, continued efforts will be crucial to further strengthen the accuracy and interpretability for tumor microbiome.
Our study revealed significant location-dependent disparities in microbial composition and host-microbe interactions between right- and left-sided tissues, substantially expanding upon observations from prior studies28,33,34,35. A recent study reported distinct forms of O-glycosylated mucus in the proximal versus distal gut and their respective roles in modulating local microbial community36. Such differences in mucus architecture across anatomical sites may contribute to the location-specific microbial patterns observed in both tumor and normal tissues. Moreover, distinct molecular and genomic features in right-sided tumors might favor the colonization and persistence of specific microbes at this site. For example, right-sided tumors are frequently characterized by hypermutation, the CMS1 subtype, and mutations in CASP8, ACVR2A, BCL9, BRAF and MSH3 genes37,38,39, which were consistently accompanied by significant enrichment of Fn subspecies and Treponema. The co-aggregation of these oral-derived microbes11,18 may synergistically promote multi-bacterial biofilm formation and metabolic adaptions to right-sided tumor microenvironments. Conversely, location-specific microbes may influence tumor molecular characteristics. For instance, pks+ tumors were more prevalent in left-sided tumors and enriched with specific mutational signatures, including SBS93 and ID14, both of which were more prevalent in gastrointestinal cancers than in other cancer types40,41.
We confirmed and extended the prognostic role of tumor-resident microbiota in specific CMS subtypes. The CMS4-dependent prognostic impact of Fn subspecies was observed in both UU and TCGA-COADREAD29 cohorts. In CMS4, Fusobacterium spp. showed negative associations with PD1 signaling, complement activation, and CD8+ T cells. This aligns with the reported immunosuppressive role of Fn, which can inhibit T cell activities through Fap2-mediated TIGIT binding42 or via succinic acid secretion43. High tumor Fn load has also been linked to increased resistance to chemotherapy and immunotherapy in metastatic CRC43,44. However, the observed CMS-dependent Fusobacterium associations might reflect the heterogeneity and complex immunological roles of TMEs in tumor progression. To our knowledge, this is the first report linking pks-island and elevated Enterobacteriaceae abundance with poor prognosis specifically within CMS2 tumors. These tumors also exhibited increased hypoxia levels, suggesting a potential impact of key Enterobacteriaceae taxa (e.g., E. coli) on CRC progression beyond colibactin-induced DNA damage45. Given the critical roles of adhesins such as FimH and FmlH in facilitating colibactin-mediated genotoxicity46, further studies are needed to investigate strain-level functional diversity and identify factors contributing to the CMS2-specific prognostic impact.
We uncovered the prognostic values of tissue-resident microbes in patients with locoregional (stage I−III) CRC for both overall and recurrence-free survival, independent of host clinicopathological factors. Several tumor-enriched Clostridium spp. were associated with poorer prognosis in both tumor and NATs. These taxa have previously been shown to increase stepwise from healthy to adenoma to carcinoma in feces17, and in our study, from locoregional to metastatic tumors, suggesting their potential role in disease progression and translational potential. Mechanistically, Clostridium species may influence host immunity and metabolism through branched-chain amino acids production47, or deoxycholic acid biotransformation48. Conversely, elevated levels of taxa from Blautia, Roseburia (R. inulinivorans), Faecalibacterium (F. prausnitzii), and Fusicatenibacter (F. saccharivorans) were associated with prolonged survival, and with downregulation of hypoxia and key pro-inflammation pathways, such as NF-κB, TNFa, IL-6, and toll-like receptor signaling. Several of these favorable taxa are known to produce butyrate to boost the activity of CD8 + T cells49,50. These findings warrant further exploration into whether microbiome modulation, such as targeted depletion of Clostridium spp. or enrichment of beneficial Lachnospiraceae taxa, may offer therapeutic benefit for CRC patients. Additionally, given the invasiveness of tissue sampling, future paired tissue–stool studies are needed to assess the utility of stool-based microbial markers as non-invasive postoperative surveillance and risk stratification.
High Akkermansia abundance (≥5% relative abundance) was significantly associated with poor survival in NATs but not in tumors. Interestingly, patients with advanced non-small-cell lung cancer exhibited worse immunotherapy outcomes when fecal A. muciniphila levels were either absent or above 4.8%51. This non-linear dose-effect was recently validated in patients with melanoma and CRC52. A. muciniphila, a well-known mucin degrader53, may, when excessively abundant, increase intestinal permeability and inflammation, thereby altering tumor microenvironment and promoting resistance to immunotherapy. Despite its growing reputation as a next-generation probiotic54, these findings underscore its context-dependent roles in cancer patients55,56 and suggest that Akkermansia supplementation should be approached with caution in oncology settings.
Overall, our results advance the understanding of CRC heterogeneity by characterizing the tissue-resident microbiota landscape and their interplays with tumor characteristics and patient prognosis. This work provides a stringent analytical framework for low-biomass microbiome research, laying the foundation for future mechanistic exploration and the rational design of microbiome-based interventions in CRC and beyond.
Limitations of this study
Our study benefits from a prospective design, deep sequencing, and a large sample of patients with CRC with long-term follow-up. However, several limitations should be noted: (1) Despite achieving sufficient WGS depth for tumor tissues, taxonomic and functional characterization of tissue-resident microbiota, particularly low-abundance bacteria and non-bacterial microbes, remains limited due to the inherently low microbial biomass. Moreover, bulk WGS does not allow clear discrimination between intracellular and extracellular microbes, leaving uncertainty whether the microbial reads originated from active microbes or from those enveloped in host cells such as macrophages. Future work employing high-resolution imaging approaches and spatial transcriptomics will be required to resolve microbial localization and viability at cellular resolution. (2) This study exclusively included treatment-naïve CRC cases; hence, the impact of cancer therapies on tissue-resident microbes cannot be directly assessed. Nonetheless, our results demonstrate the prognostic value of NAT-resident microbiota, highlighting the potential utility of post-surgical microbiome modulation against tumor recurrence and progression. (3) Given the observational nature of this study, the reported associations should not be interpreted as evidence of direct causality. Although consistent enrichment patterns and clinical associations were observed across multiple cohorts, the underlying mechanisms—whether driven by microbial activity, tumor immune microenvironment, or host-microbe interactions—remain to be elucidated. Future integrative efforts combining human multi-omics with experimental models are warranted to establish causal links and functional roles of microbiota in CRC pathogenesis and prognosis.
Methods
Patient cohort
Samples used in this study are from patients diagnosed with CRC between 2004 and 2019, at the Uppsala University Hospital (UU) or the Umeå University Hospital (UM). Most patients included from June 2010 were drawn from the Uppsala-Umeå Comprehensive Cancer Consortium (U-CAN) biobank collections (Uppsala Biobank and Biobanken Norr)57. Sampling and analyses were performed under the ethical permits Uppsala EPN 2004-M281, 2010-198, 2007-116, 2012-224, 2015-419, 2018-490, EPM Dnr 2022-06182-01, and Umeå EPN 2016-219 and EPM 2019-566. Patients or guardians provided informed written consent for participation in this study as stipulated by the study protocols. Participants were not compensated.
Clinicopathological and follow-up data was extracted from the national quality registry, the Swedish Colorectal Cancer Registry (SCRCR), and completed from medical records. Patients who received chemotherapy and/or radiotherapy before sampling or surgery were excluded, leading to a final number of 937 treatment-naïve patients (UU = 775; UM = 162) retained in this study9. Follow-up for alive patients was minimum 3.9 year and median 8 years (data lock 14th June 2023), with 870 patients (93%) having complete 5-year follow up.
Tissue handling, sample collection and nucleic acids extraction
Tissue samples from UU were embedded and stored at −70 °C on collection day. Tissue samples from UM were frozen directly in segments and similarly stored as UU samples. Patient-matched normal DNA samples were obtained from blood (UU = 462) or normal adjacent tissue (NAT) (UU = 313; UM = 162), and normal RNA from 103 patient-matched colon or rectum tissue samples (UM = 8; UU = 95).
For UU samples, tissue DNA was extracted using the NucleoSpin Tissue kit (740952, Macherey-Nagel), RNA with the RNeasy Mini Kit (74106, Qiagen), and blood DNA with the NucleoSpin 96 Blood Core kit (740456, Macherey-Nagel). For UM tissue samples, both DNA and RNA were extracted with AllPrep DNA/RNA/miRNA Universal kit (80224, Qiagen). The tissue samples that used to extract DNA and RNA were completely consumed, no remaining material was available.
Host genomic features and molecular subtypes
Somatic genomic alterations (SNV, MNV, INDEL, SV and CNV), driver genes, hypermutated (HM) and non-hypermutated (nHM) status, mutational signatures (SBS, DBS, and ID), consensus molecular subtypes (CMS), single-sample pathway profiles, CMS-featured pathways, cancer-related pathways and tumor microenvironment of U-CAN tumors were identified and described in our previous study9. As described by Kim et al.58, HM thresholds were determined based on the distribution of somatic mutation counts specific to each dataset, using the total count of somatic variants in the whole genome for U-CAN (>23.2 mutations/Mb)9 and in the whole exome for AC-ICAM tumors (>56.35 mutations/Mb). CMS classification of the AC-ICAM tumors was obtained from Supplementary Source Data10 (Supplementary Data 1).
Methods for microbiota
Part 1 Detection, decontamination, and quantification of tissue-resident microbiota
We implemented a rigorous pipeline to detect, decontaminate, and quantify tissue-resident microbiota in CRC samples from the U-CAN study.
Step 1, Non-human sequencing data extraction and quality control
High-quality WGS (average 52× coverage) and WTS (average 30 million paired reads) data were produced by the DNBSEQ platform (MGI) with 100-bp paired-end mode9. Sequences were aligned to human genome reference GRCh38 as previously described9. For the WGS data, we processed 1,088 tissues (775 tumors and 313 paired NATs) and 462 blood samples from UU, and 324 tissues (162 tumors and paired NATs) from UM. For the WTS data, we processed 870 tissues (775 tumors and 95 NATs) for UU, and 170 tissues (162 tumors and 8 NATs) for UM. Extracted raw reads were subjected to quality controls as follows: (1) low-quality reads were removed using fastp59 (V0.20.1, parameters: --cut_front --cut_right --cut_front_window_size 4 --cut_front_mean_quality 20 --cut_tail_window_size 4 --cut_tail_mean_quality 20 --length_required 51); (2) human reads were identified by alignment with the GRCh38 genome (Bowtie2 v2.4.2, -very-sensitive) and removed; (3) non-human aligned paired reads were extracted from sequencing data (SAMtools v1.9, view -f 12)60 for each sample. After QC, these high-quality reads were obtained for tissue and blood samples (referred to as “input reads”, Supplementary Fig. 1a).
Step2, Microbiota detection using Kraken 2 and UHGG catalog
Taxonomic assignment of “input reads” was performed using Kraken 261 with a custom reference database included human genomes (GRCh38 and T2T-CHM13v2.0), the Unified Human Gastrointestinal Genome (UHGG) catalog (V1.0)62, comprising 4644 high-quality gut representative genomes), and 17 additional NCBI prokaryotic genomes from the genus Treponema. Reads classified as human by Kraken2 were re-removed at this stage to enhance stringency. For each individual sample, a taxon was considered present if it was supported by ≥10 mapped reads. The overall mapping rate against the UHGG was calculated by dividing Kraken2-mapped “prokaryotic reads” by total “input reads”.
Step 3, Removal of putative microbial contaminants using blood samples
To distinguish tissue-resident microbiota from potential contaminants, we used WGS data from 462 paired tumors and blood samples in UU, with blood samples as positive controls. Following the criteria detailed by Dohlman et al.11, we excluded 114 microbial genera identified as putative contaminants based on the two conditions: (1) prevalence in ≥20% of UU blood samples, and (2) significantly higher prevalence in blood compared to paired tissue samples (two-sided Fisher exact test, P < 0.05; Supplementary Data 2).
Step 4, Removal of putative microbial contaminants using a published blacklist
To further minimize contaminants, we excluded 24 additional genera frequently found in negative blank reagents as curated in a previous study4.
Step 5, Removal of microbial taxa with low-prevalence
To enable confident identification of tissue-resident taxa and to ensure statistical power, we excluded low-prevalent genera (<20% of occurrence in tissue samples). This yielded 249 genera shared between both UU and UM tissues, termed “common genera”.
Step 6, Removal of microbial species with low genome coverage
To guarantee accurate species-level assignment, high-quality representative species belonging to the common genera were extracted to create a Bowtie2 database (bowtie2-build, default parameters). Prokaryotic reads from WGS tissue samples were aligned to this custom database, and genome coverages were estimated using bedtools genomecov (V2.30.0, default settings). A total of 361 species with at least 5% genome coverage in ≥5% of WGS tissue samples were identified as tissue-resident species.
We further assessed genome coverage for these 361 species in 870 WTS tissue samples and 462 WGS blood samples. Despite comparable non-human read counts from paired WGS and WTS data of UU tumors, WTS data exhibited notably fewer detected species and lower genome coverage for tissue-resident species compared to WGS data (Supplementary Fig. 1d−f, Supplementary Data 2D). Application of RiboDetector63 (version 0.2.7, -e norna) revealed that, on average, 40% of prokaryotic reads in the total microbial RNA data were ribosomal RNA, making accurate quantification of active microbes challenging (Supplementary Fig. 1g). We therefore opted not to proceed with the WTS-based microbial analyses.
Prior to subsequent analyses, genus-level distance-based Redundancy Analysis (db-RDA)64 was performed on all U-CAN tissue samples to assess host-related confounding variables associated with tissue-resident microbiota. Significant disparities in microbial composition were observed between UU and UM cohorts, possibly due to difference in DNA extraction methods9. To account for batch effects, the UU cohort was designated as the discovery cohort and the UM cohort as the internal validation cohort. Additionally, non-human WGS data from 167 AC-ICAM tumors were collected and processed using the identical pipeline for identifying and quantifying tissue-resident microbes, and used as an external validation cohort10.
Part 2 Associations between host clinical features and tissue-resident microbiota
To deal with the zero-inflated compositional data, the relative abundances (RAs) of 249 tissue-resident genera and 361 species were transformed by centered log-ratio (CLR) for samples of the discovery cohort (UU) and validated cohorts (UM and AC-ICAM). Prior to transformation, zero values in the compositional matrix were imputed using the Bayesian multiplicative replacement method (R package zCompositions, v1.5.0-3)65.
Comparison of tissue-resident microbiota between the right- and left-sided CRC tissues
CRC tissue samples were categorized by colonic location of the primary tumor: right-sided colon (caecum, ascending, hepatic flexure and transverse colon), left-sided colon (splenic flexure, descending colon, sigmoid and rectosigmoid junction), and rectum (distal 15 cm from the anal verge). To mitigate impacts of disease status on microbial composition, all location-related analyses were conducted separately for tumors and NATs.
Based on the UU discovery cohort, we applied Permutational analysis of variance (PERMANOVA) to quantify the proportion of compositional variance of tissue-resident microbiota explained by host factors in NATs (n = 313) and tumors (n = 775). PERMANOVA was conducted using Bray-Curtis dissimilarity matrix at the genus level. Pseudo-F statistics were computed to quantify compositional variations for PERMANOVA. D-MANOVA66 was used to estimate asymptotic P values. A Benjamini-Hochberg (BH)-adjusted P < 0.05 was considered statistically significant (R function adonis2 in package vegan, v2.5-7).
We next performed MaAsLin267 on CLR-transformed abundance data of microbial genera and species to identify location-specific taxa after adjusting for age, sex, and prokaryotic read counts (R package MaAsLin2, version 1.14.1). Model coefficients (estimates) were calculated to indicate the strength and direction of taxa enrichment patterns. Due to similarities in microbial composition between the left colon and rectum, we merged samples from these two locations into a “left-sided” category (Supplementary Fig. 2a−c). Microbial differences were reassessed between the right- and left-sided colorectal samples in the UU cohort and validated in UM and AC-ICAM (tumors only) cohorts. A BH-adjusted P < 0.05 was considered statistically significant for the UU discovery cohort and at a P value of < 0.05 for validation cohorts.
We also conducted partial Spearman’s correlation analysis to assess associations between RAs of tissue-resident taxa and expression levels of host pathways (Reactome database) in tumors from UU, UM and AC-ICAM cohorts, separately. The covariates included age, sex, anatomical location, and prokaryotic read counts. A BH-adjusted P < 0.05 was considered statistically significant for the UU cohort, and a P value of < 0.05 for validation cohorts.
Assessment of location-specific microbial differences using fecal shotgun metagenomes of CRC patients
To determine whether tissue-derived, location-specific microbial features are detectable in fecal samples, we analyzed 538 fecal metagenomes from CRC patients with documented tumor anatomical locations across five independent cohorts14,68,69,70,71. Fecal microbes were identified and quantified using the same analytical pipeline established in the U-CAN cohort to ensure consistency. Microbial abundance differences were evaluated using Wilcoxon rank-sum test, and taxa with a P < 0.05 in at least two cohorts were considered as repeated location-specific features (Supplementary Fig. 2h)
Comparison of tissue-resident microbiota between tumors and NATs
Differential abundance analysis was performed on paired tumor and NAT tissues from UU (n = 313 pairs) and UM (n = 162 pairs) cohorts using MaAsLin2. The Shannon index and CLR-transformed abundances of genera and species were analyzed using linear mixed-effect regression models, with patient ID as a random effect, and tissue types (tumor vs. NAT), age, sex, anatomical location, and prokaryotic read counts as fixed effects. Considering significant location differences in microbial composition, we also analyzed paired tumor and NAT tissues separately from right- and left-sided CRC, adjusting for age, sex, and prokaryotic read counts. Significance was determined at BH-adjusted P value of <0.05 for UU and at P value of <0.05 for the validation cohort (UM).
Determination of clade-level taxonomy of Fn subspecies animalis (Fna)
To determine the clade-level taxonomy of Fna subspecies identified in our tumor samples, we evaluated average nucleotide identity (ANI) between the UHGG Fna reference genome and the recently identified Fna clades (Fna C1: GCF037889065.1 and Fna C2: GCF037889045.1)12. We found that the UHGG Fna genome GUT_GENOME096031 exhibited 97% ANI with Fna C2 but a much lower ANI (≤93%) with Fna C1 or other Fn subspecies, indicating that the Fna identified in U-CAN belongs to the Fna C2 clade.
Species-level identification of Treponema in CRC tumors
Although members of the genus Treponema are rarely detected in fecal samples from urban populations72,73, they were frequently enriched in CRC tumor tissues across cohorts10,34,74,75. To resolve the species-level taxonomy of Treponema detected in CRC tumor tissues, we mapped microbial reads mapped to a curated set of high-quality Treponema reference genomes, including two gut-resident species from the UHGG database (T. succinifaciens and T. berlinense) and 17 oral-resident species from NCBI (Supplementary Data 6E). Genome-wide ANI between gut- and oral-resident Treponema species was between 75 and 80%, suggesting their phylogenetic divergence. For species assignment, we quantified the proportion of Treponema-aligned reads that mapped to each reference genome. Over 90% of Treponema reads in tumor samples were assigned to oral residents, such as T. denticola, T. medium, and T. maltophilum (Supplementary Data 6E).
Microbiota-based random forest models for classifying right- vs. left-sided, and tumor vs. NATs tissues
We developed two independent random-forest (RF) models to evaluate how well tissue-resident microbiota distinguish anatomical locations (right-sided vs. left-sided) and tissue types (tumor vs. NAT), respectively. Models were built using CLR-transformed abundances from common tissue-resident prokaryotic taxa. The left-right classifier was trained on all UU tumors (n = 775) using 5-fold cross-validation (CV) and evaluated on multiple independent datasets: UU NATs (n = 313), UM tumors (n = 162) and NATs (n = 162), and AC-ICAM tumors (n = 167). Similarly, the tumor-NAT classifier was developed by training on all UU paired tissues (n = 313 paired tumors and NATs) using 5-fold CV, and tested on 162 independent paired tissues from the UM cohort. A separate RF model was applied to fecal samples of CRC patients in the Yachidas_2019 study (167 left-sided and 83 right-sided CRC tumors) using 5-fold CV14.
RF models were computed using the R package randomForest (v4.7-1.1). Model performance was evaluated based on the area under the receiver operating characteristic curve (AUC), implemented in the R package pROC (v1.18.0).
Part 3 Associations between host genetic alterations and tissue-resident taxa in CRC tumors
To assess associations between host genetic alterations and tumor-resident microbes, we applied MaAsLin2 to compare relative abundances of microbial taxa between hypermutated (HM) and non-hypermutated (nHM) tumors across UU, UM and AC-ICAM cohorts. The models included age, sex, colonic locations, and prokaryotic read counts as covariates, we further conducted partial Spearman’s correlation analysis between taxon abundances and the number of somatic mutations (single nucleotide variations, SNVs), adjusting for the same covariates as above.
We then extended the MaAsLin2 models to assess relationships between microbial abundances and genetic alterations in CRC driver or DNA damage response and repair (DDR) genes. Tumors were categorized into wild-type and mutant groups based on the presence of SNVs or loss of heterozygosity (LOH) in specific genes. To ensure adequate statistical power, we excluded genes with SNVs or LOH events observed in fewer than 20 tumors in the UU cohort. This resulted in the inclusion of 83 driver genes and 35 DDR genes for SNV analysis, and 91 driver genes and 69 DDR genes for LOH-based analysis. A BH-adjusted P < 0.05 was considered statistically significant for UU, and a P < 0.05 for two validation cohorts.
Identification of pks+ associated CRC mutation signatures
The colibactin-encoding pks gene island in CRC tumors was detected using shortBRED76 (V0.9.5), a validated tool for high-specificity, marker-based functional profiling in metagenomic data. Virulence protein sequences, including the 19 clb genes (clbA–clbS), were retrieved from the Virulence Factor Database77 (VFDB, release 2022.12.23), and ShortBRED-Identify was applied to generate gene-specific peptide markers using UniRef90 as the background to minimize non-specific matches. A tumor sample is considered pks-positive if at least one clb gene (among clbA–clbS) has ≥1 unique ShortBRED marker hit (Supplementary Data 7). Based on this approach, 17.4% (135/775) of UU tumors were classified as pks-positive (pks + ). On average, seven distinct pks genes were identified per pks+ tumor.
ShortBRED-Quantify was used to estimate gene relative abundance, by normalizing read counts (marker hits) to the effective length of the gene-specific peptide markers and sequencing depth (reported as “normalized counts” in RPKM). The relative abundance of the pks island was quantified by summing the gene-level abundances across all clb genes (Supplementary Data 7). Spearman’s rank correlation analysis was applied to assess relationships between E. coli abundances and pks RPKM, as well as between pks abundance and the activity of COSMIC mutational signatures within the 775 tumors. Partial Spearman’s correlation was used to determine whether specific mutational signatures were enriched in pks+ tumors in the full UU cohort and the nHM tumor subset adjusting for age, sex, location, and prokaryotic read counts. A P < 0.05 was considered statistical significance. Fisher’s exact test was conducted to compare the prevalence of individual COSMIC signatures between pks+ and pks- tumors.
Part4: Associations between host expression levels and tissue-resident taxa in CRC tumors
Microbial composition and host gene expression profiles in CMS subtypes
A total of 709 UU tumors were classified into four CMS subtypes (CMS1 = 138, CMS2 = 224, CMS3 = 92 and CMS4 = 255) as described previously9. Enrichment patterns of tissue-resident taxa and pks-island were investigated for each subtype as compared to all other subtypes using MaAsLin2 models, adjusting for sex, age, colonic location, and prokaryotic read counts. A BH-adjusted P < 0.05 was considered statistically significant.
CMS-dependent associations between RAs of tissue-resident taxa and host gene expression levels
A total of 14 cancer-related pathways (PROGENy)78, 36 previously-reported CMS-featured pathways9, two well-established scoring systems that evaluate the abundance and activity of T-lymphocytes (T-cell score79 and Immunoscore80), cytokines involved in immunosuppression (IL-6, IL-10, and CXCL12), pathways and signatures related to anti-tumor immune (CD8 + T effector81, TMEscoreA82), immunosuppression (TNFα78, T cell exhaustion, T cell regulatory83), and tumor immune escape (TGF-β78, CAF, MDSC, TAM83, TMEscoreB82), and 21 types of tumor-infiltrating immune cells (CIBERSORT)84 were included in analysis. Enrichment patterns of cancer pathways, CMS-featured pathways and immune cells were investigated for each subtype as compared to all other subtypes using two-sample T-test. Partial Spearman’s correlation analysis was conducted to assess associations between tissue-resident taxon abundances and host gene expression levels within each CMS subtype. Covariates included age, sex, colonic location, and prokaryotic read count. A BH-adjusted P < 0.05 was considered statistically significant.
Identification of CMS-dependent prognostic impact of tumor-enriched taxa and pks island
The identification of prognostic-related tissue microbes was conducted in stage I-III patients from three cohorts: UU (n = 689), UM (n = 150) and AC-ICAM (WGS = 149,16S rRNA amplicon=224). We first performed univariate Cox proportional hazard models to identify host clinical variables associated with 5-year overall survival (OS) and relapse-free survival (RFS) in UU patients (R function coxph in survival package, version 3.5-5). The clinical variables included age, sex, tumor stage (I vs. II and I vs. III), tumor grade (low vs. high), tumor locations (right- vs. left-sided), post-surgical treatment (yes vs. no), hypermutation status (HM vs. nHM), microsatellite instability status (MSS vs. MSI), total number of SNVs, and CMS subtypes. From this analysis, five variables (age, tumor stage, grade, location, and post-surgical treatment) were significantly associated with 5-year OS or RFS (P < 0.05) and were subsequently taken as covariates in microbiome -based multivariate survival models.
We then assessed the CMS-dependent prognostic effects of tumor-enriched taxa and the presence of pks-island in stage I-III UU tumors using multivariate Cox proportional hazard models. Covariates included the five variables from the univariate analyses, and prokaryotic read counts to account for microbial sequencing depth. Before modeling, microbial RAs were log10 transformed and normalized to Z-scores. Hazard ratios (HRs) with 95% confident intervals (95% CI) and P-values for both OS or RFS were estimated using the coxph function (R package survival, V 3.5-5). A BH-adjusted P < 0.05 was considered statistically significant. Forest plots were generated using the forest function in the R package forestploter (version 1.1.0). Kaplan–Meier curves were generated using ggsurvplot from R package survminer (version 0.4.9).
To validate the association between Enterobacteriaceae abundance and 5-year OS specifically within CMS2 subtype, we further applied the same survival models to WGS dataset from UM and AC-ICAM cohorts (stage I-III CMS2 tumors, n = 98, other CMS subtypes n = 178), and 16S rRNA amplicon data from AC-ICAM patients (stage I−III tumors, n = 55, others, n = 134). For validation analyses, P < 0.05 was considered statistically significant.
Part5: Assessing the prognostic impacts of tissue-resident microbiota in tumors and NATs
Identification of prognostic taxa independent of host variables
We next analyzed highly prevalent taxa (prevalence ≥ 50%) to identify microbes with prognostic value independent of clinicopathological factors. Multivariable Cox proportional hazard models were conducted separately tumor-resident and NAT-resident taxa in patients with stage I-III CRC (tumors=689, NATs=288), adjusting for five host prognostic factors (age, tumor stage, tumor grade, tumor location, and post-surgical treatment) and prokaryotic read counts.
In UU tumors, 122 taxa were identified as significant prognostic biomarkers at BH-adjusted P < 0.05. In UU NATs, 34 prognostic taxa were identified at a threshold of P < 0.01. Prognostic associations of the tumor-derived taxa were further investigated in each CMS subtype within UU cohort, and the overall UM and AC-ICAM cohorts (stage I-III patients only). A P < 0.05 was used for validation cohorts.
Construction of tumor- and NAT-derived microbial risk score (MRS-T and MRS-N)
To quantify the combined prognostic contribution of microbial communities, we constructed microbial risk scores in tumors (MRS-T) and NATs (MRS-N), respectively. The MRS was not developed using a machine learning model but a mathematically defined index that integrates both the number and relative abundances of prognostic taxa identified in the discovery cohort, using a previously developed formula85:
Where |N + | and |N-| represent the total number of prognostic taxa significantly associated with shorter (HR > 1) and longer survival (HR < 1), respectively. RN+ and RN- denote the richness (number of prognostic taxa) of taxa associated with shorter and longer OS/RFS in each sample. Pi and pj represent the relative abundances of taxa associated with shorter and longer OS in each sample. This approach is conceptually similar to established microbial diversity indices, which provide a simple and interpretable framework for integrating multiple prognostic taxa into a single quantitative index. Briefly, the MRS-T and MRS-N values were calculated for stage I-III patients in the UU cohort, and subsequently evaluated in independent cohorts, including stage I-III patients from UM and AC-ICAM cohorts, and stage IV patients from the U-CAN cohort.
Assessing the prognostic impacts of MRS-T and MRS-N
Multivariable Cox proportional hazard models were applied to estimate the independent HRs of MRS-T and MRS-N for OS and 5-year RFS in stage I-III patients from the UU (tumors=689, NATs=288) and UM cohorts (tumors=150, NATs=150), adjusting for the five host prognostic factors and prokaryotic reads counts. For the AC-ICAM cohort (stage I-III tumors=147), HRs of MRS-T for OS were adjusted for age, tumor location, stage, and prokaryotic read counts. For stage IV patients in U-CAN (n = 98; UU = 86 and UM = 12), multivariable models were adjusted for age, tumor location, grade, post-surgical treatment, and prokaryotic reads counts. A P < 0.05 was considered statistically significant.
The Improvement in prognostic accuracy conferred by MRS-T and MRS-N was evaluated using the Concordance index (C-index)86 (cindex function in dynpred package, v0.1.2), by comparing microbiome-based models with host-only models across UU, UM, and AC-ICAM cohorts. Spearman’s rank correlation analysis was further conducted to assess the relationship between paired MRS-T and MRS-N values in UU samples (N = 288).
Associations between prognostic taxa and host gene expressions in tumor samples
Partial Spearman’s correlation analysis was conducted across three cohorts to assess the associations between the RAs of tumor-derived prognostic taxa and host gene expression. Covariates included age, sex, tumor location, and prokaryotic read counts. A BH-adjusted P < 0.05 was considered statistically significant in UU while a P < 0.05 was applied in UM and AC-ICAM cohorts. Wilcoxon rank-sum test was applied to examine differences in proinflammatory cytokine and chemokine gene expression levels between high and low MRS-T groups, divided by the median value of MRS-T.
Construction and assessment of MRS-T and MRS-N based on 16S rRNA data from the AC-ICAM cohort
Next, 154 common genera (prevalence ≥ 10%) identified by 16S rRNA amplicon data from matched tumors and NATs of 209 stage I-III AC-ICAM patients10 were used as an independent validation dataset for generating MRS-T and MRS-N. Genus abundance was log-10 transformed. Multivariable Cox proportional hazard models were conducted separately for tumors and NATs, adjusting for age, tumor location and stage (coxph function in R survival package, V3.5-5). A P < 0.05 was considered as the significance threshold. In tumors, 14 genera were identified as prognostic biomarkers, and in NATs, 12 genera were identified. The 16S rRNA-based MRS-T and MRS-N were then constructed and their independent prognostic effects were assessed following the same methodology, with adjustments for relevant host clinical variables.
Statistics and reproducibility
No statistical method was used to pre-determine sample size. Samples were included if they passed predefined quality-control criteria, and no data were excluded otherwise. Two-sided tests were performed and P values were adjusted using the Benjamini–Hochberg method. Randomization and blinding were not applicable due to the observational design of the study.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The processed non-human reads from tumor and NAT tissues of 937 CRC patients in the U-CAN study are deposited at the European Nucleotide Archive (ENA, PRJEB86036) and the China National GeneBank Sequence Archive (CNSA87, CNP0007001) of the China National GeneBank DataBase (CNGBdb)88. Genomic variations and gene expression profiles from the U-CAN study are available at the European Variation Archive89 (EVA, PRJEB61514), and at the ArrayExpress90 (E-MTAB-12862), and available at the CNSA (CNP0004160). The raw WGS data can be accessed by directing to the U-CAN cancer biobank at Uppsala University (https://www.uu.se/forskning/u-can/)9. Non-human WGS reads from tumor tissues of 167 colon cancer patients in the AC-ICAM study10 are publicly available via the Sequence Read Archive (SRA) (PRJNA941834). A genus-level 16S rRNA amplicon-based abundance matrix for tumor and matched healthy colon tissues from 246 patients in the AC-ICAM study were retrieved from Figshare (https://figshare.com/articles/dataset/Supplementary_Data_AC-ICAM/16944775). Clinical and gene expression data for the AC-ICAM study are available from Supplementary Source Data of the study. Source data are provided with this paper.
Code availability
The source code for generating the main figures and constructing the Microbial Risk Score (MRS) is available to use on GitHub under the MIT License (https://github.com/rusher321/UCAN_Microbiome). The repository has been archived on Zenodo (https://doi.org/10.5281/zenodo.17588313, version v1.0)91.
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. Ca. Cancer J. Clin. 74, 229–263 (2024).
Wong, C. C. & Yu, J. Gut microbiota in colorectal cancer development and therapy. Nat. Rev. Clin. Oncol. 20, 429–452 (2023).
Wang, N. & Fang, J. Y. Fusobacterium nucleatum, a key pathogenic factor and microbial biomarker for colorectal cancer. Trends Microbiol. 31, 159–172 (2023).
Eisenhofer, R. et al. Contamination in low microbial biomass microbiome studies: issues and recommendations. Trends Microbiol. 27, 105–117 (2019).
Wang, Q. et al. Computational methods and challenges in analyzing intratumoral microbiome data. Trends Microbiol. 31, 707–722 (2023).
Poore, G. D. et al. Microbiome analyses of blood and tissues suggest cancer diagnostic approach. Nature 579, 567–574 (2020).
Hermida, L. C., Gertz, E. M. & Ruppin, E. Predicting cancer prognosis and drug response from the tumor microbiome. Nat. Commun. 13, 2896 (2022).
Gihawi, A. et al. The landscape of microbial associations in human cancer. Sci. Transl. Med. 17, eads6166 (2025).
Nunes, L. et al. Prognostic genome and transcriptome signatures in colorectal cancers. Nature 633, 137–146 (2024).
Roelands, J. et al. An integrated tumor, immune and microbiome atlas of colon cancer. Nat. Med. 29, 1273–1286 (2023).
Dohlman, A. B. et al. The cancer microbiome atlas: a pan-cancer comparative analysis to distinguish tissue-resident microbiota from contaminants. Cell Host Microbe 29, 281–298.e5 (2021).
Zepeda-Rivera, M. et al. A distinct Fusobacterium nucleatum clade dominates the colorectal cancer niche. Nature 628, 424–432 (2024).
Ternes, D. et al. Microbiome in colorectal cancer: how to get from meta-omics to mechanism?. Trends Microbiol. 28, 401–423 (2020).
Yachida, S. et al. Metagenomic and metabolomic analyses reveal distinct stage-specific phenotypes of the gut microbiota in colorectal cancer. Nat. Med. 25, 968–976 (2019).
Chen, Z., Han, F., Du, Y., Shi, H. & Zhou, W. Hypoxic microenvironment in cancer: molecular mechanisms and therapeutic interventions. Signal Transduct. Target. Ther. 8, 70 (2023).
Wirbel, J. et al. Meta-analysis of fecal metagenomes reveals global microbial signatures that are specific for colorectal cancer. Nat. Med. 25, 679–689 (2019).
Liang, J. Q. et al. A novel faecal Lachnoclostridium marker for the non-invasive diagnosis of colorectal adenoma and cancer. Gut 69, 1248–1257 (2020).
Younginger, B. S. et al. Enrichment of oral-derived bacteria in inflamed colorectal tumors and distinct associations of Fusobacterium in the mesenchymal subtype. Cell Rep. Med. 4, 100920 (2023).
Fu, K. et al. Streptococcus anginosus promotes gastric inflammation, atrophy, and tumorigenesis in mice. Cell 187, 882–896.e17 (2024).
Nieminen, M. T. et al. Treponema denticola chymotrypsin-like proteinase may contribute to orodigestive carcinogenesis through immunomodulation. Br. J. Cancer 118, 428–434 (2018).
Chun, H. J. et al. Pleiotropic defects in lymphocyte activation caused by caspase-8 mutations lead to human immunodeficiency. Nature 419, 395–399 (2002).
Zhang, H. et al. Activin A/ACVR2A axis inhibits epithelial-to-mesenchymal transition in colon cancer by activating SMAD2. Mol. Carcinog. 62, 1585–1598 (2023).
Ma, Z. et al. BRAFV600E mutation promoted the growth and chemoresistance of colorectal cancer. Am. J. Cancer Res. 13, 1486–1497 (2023).
Wu, B., Zhan, X. & Jiang, M. CD58 defines regulatory macrophages within the tumor microenvironment. Commun. Biol. 7, 1025 (2024).
Feng, M. et al. BCL9 regulates CD226 and CD96 checkpoints in CD8 + T cells to improve PD-1 response in cancer. Signal Transduct. Target. Ther. 6, 313 (2021).
Erfani, M. et al. Altered ARID1A expression in colorectal cancer. BMC Cancer 20, 350 (2020).
Wang, Z. et al. Comutations in DNA damage response pathways serve as potential biomarkers for immune checkpoint blockade. Cancer Res. 78, 6486–6496 (2018).
Pleguezuelos-Manzano, C. et al. Mutational signature in colorectal cancer caused by genotoxic pks + E. coli. Nature 580, 269–273 (2020).
Salvucci, M. et al. Patients with mesenchymal tumours and high Fusobacteriales prevalence have worse prognosis in colorectal cancer (CRC). Gut 71, 1600–1612 (2022).
Cornish, A. J. et al. The genomic landscape of 2,023 colorectal cancers. Nature 633, 127–136 (2024).
Bukhari, S. et al. Single-cell RNA sequencing reveals distinct T cell populations in immune-related adverse events of checkpoint inhibitors. Cell Rep. Med. 4, 100868 (2023).
Gihawi, A. et al. Major data analysis errors invalidate cancer microbiome findings. MBio 14, e0160723 (2023).
Dejea, C. M. et al. Microbiota organization is a distinct feature of proximal colorectal cancers. Proc. Natl. Acad. Sci. USA 111, 18321–18326 (2014).
Mouradov, D. et al. Oncomicrobial community profiling identifies clinicomolecular and prognostic subtypes of colorectal cancer. Gastroenterology 165, 104–120 (2023).
James, K. R. et al. Distinct microbial and immune niches of the human colon. Nat. Immunol. 21, 343–353 (2020).
Bergstrom, K. et al. Proximal colon-derived O-glycosylated mucus encapsulates and modulates the microbiota. Science 370, 467–472 (2020).
Loree, J. M. et al. Classifying colorectal cancer by tumor location rather than sidedness highlights a continuum in mutation profiles and consensus molecular subtypes. Clin. Cancer Res. 24, 1062–1072 (2018).
Baran, B. et al. Difference between left-sided and right-sided colorectal cancer: a focused review of literature. Gastroenterol. Res. 11, 264–273 (2018).
Guinney, J. et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 21, 1350–1356 (2015).
Islam, S. M. A. et al. Uncovering novel mutational signatures by de novo extraction with SigProfilerExtractor. Cell Genom. 2, 100179 (2022).
Everall, A. et al. Comprehensive repertoire of the chromosomal alteration and mutational signatures across 16 cancer types from 10,983 cancer patients. https://doi.org/10.1101/2023.06.07.23290970 (2023).
Gur, C. et al. Binding of the Fap2 protein of fusobacterium nucleatum to human inhibitory receptor TIGIT protects tumors from immune cell attack. Immunity 42, 344–355 (2015).
Jiang, S. S. et al. Fusobacterium nucleatum-derived succinic acid induces tumor resistance to immunotherapy in colorectal cancer. Cell Host Microbe 31, 781–797.e9 (2023).
Yu, T. C. et al. Fusobacterium nucleatum promotes chemoresistance to colorectal cancer by modulating autophagy. Cell 170, 548–563.e16 (2017).
Dziubańska-Kusibab, P. J. et al. Colibactin DNA-damage signature indicates mutational impact in colorectal cancer. Nat. Med. 26, 1063–1069 (2020).
Jans, M. et al. Colibactin-driven colon cancer requires adhesin-mediated epithelial binding. Nature 635, 472–480 (2024).
Ren, Y. M. et al. BCAA-producing Clostridium symbiosum promotes colorectal tumorigenesis through the modulation of host cholesterol metabolism. Cell Host Microbe https://doi.org/10.1016/j.chom.2024.07.012 (2024).
Cong, J. et al. Bile acids modified by the intestinal microbiota promote colorectal cancer growth by suppressing CD8 + T cell effector functions. Immunity 57, 876–889.e11 (2024).
Ye, L. et al. Repressed Blautia-acetate immunological axis underlies breast cancer progression promoted by chronic stress. Nat. Commun. 14, 6160 (2023).
Kang, X. et al. Roseburia intestinalis generated butyrate boosts anti-PD-1 efficacy in colorectal cancer by activating cytotoxic CD8 + T cells. Gut 72, 2112–2122 (2023).
Derosa, L. et al. Intestinal Akkermansia muciniphila predicts clinical response to PD-1 blockade in patients with advanced non-small-cell lung cancer. Nat. Med. 28, 315–324 (2022).
Derosa, L. et al. Custom scoring based on ecological topology of gut microbiota associated with cancer immunotherapy outcome. Cell 187, 3373–3389.e16 (2024).
Chelakkot, C. et al. Akkermansia muciniphila-derived extracellular vesicles influence gut permeability through the regulation of tight junctions. Exp. Mol. Med. 50, e450 (2018).
Cani, P. D., Depommier, C., Derrien, M., Everard, A. & de Vos, W. M. Akkermansia muciniphila: paradigm for next-generation beneficial microorganisms. Nat. Rev. Gastroenterol. Hepatol. 19, 625–637 (2022).
Jiang, Y. et al. Acetyltransferase from Akkermansia muciniphila blunts colorectal tumourigenesis by reprogramming tumour microenvironment. Gut 72, 1308–1318 (2023).
Routy, B. et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science 359, 91–97 (2018). (80-.).
Glimelius, B. et al. U-CAN: a prospective longitudinal collection of biomaterials and clinical information from adult cancer patients in Sweden. Acta Oncol. 57, 187–194 (2018).
Kim, J. et al. Somatic ERCC2 mutations are associated with a distinct genomic signature in urothelial tumors. Nat. Genet. 48, 600–606 (2016).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. Fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Wood, D. E., Lu, J. & Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 20, 257 (2019).
Almeida, A. et al. A unified catalog of 204,938 reference genomes from the human gut microbiome. Nat. Biotechnol. 39, 105–114 (2021).
Deng, Z. L., Münch, P. C., Mreches, R. & McHardy, A. C. Rapid and accurate identification of ribosomal RNA sequences via deep learning. Nucleic Acids Res. 50, E60 (2022).
McArdle, B. H. & Anderson, M. J. Fitting multivariate models to community data: A comment on distance-based redundancy analysis. Ecology 82, 290–297 (2001).
Palarea-Albaladejo, J. & Martín-Fernández, J. A. ZCompositions - R package for multivariate imputation of left-censored data under a compositional approach. Chemom. Intell. Lab. Syst. 143, 85–96 (2015).
Chen, J. & Zhang, X. D-MANOVA: fast distance-based multivariate analysis of variance for large-scale microbiome association studies. Bioinformatics 38, 286–288 (2021).
Mallick, H. et al. Multivariable association discovery in population-scale meta-omics studies. PLoS Comput. Biol. 17, e1009442 (2021).
Zeller, G. et al. Potential of fecal microbiota for early-stage detection of colorectal cancer. Mol. Syst. Biol. 10, 766 (2014).
Feng, Q. et al. Gut microbiome development along the colorectal adenoma-carcinoma sequence. Nat. Commun. 6, 6528 (2015).
Yu, J. et al. Metagenomic analysis of faecal microbiome as a tool towards targeted non-invasive biomarkers for colorectal cancer. Gut 66, 70–78 (2017).
Vogtmann, E. et al. Colorectal cancer and the human gut microbiome: reproducibility with whole-genome shotgun sequencing. PLoS ONE 11, e0155362 (2016).
Carter, M. M. et al. Ultra-deep sequencing of Hadza hunter-gatherers recovers vanishing gut microbes. Cell 186, 3111–3124.e13 (2023).
Maghini, D. G. et al. Expanding the human gut microbiome atlas of Africa. Nature 638, 718–728 (2025).
Galeano Niño, J. L. et al. Effect of the intratumoral microbiota on spatial and cellular heterogeneity in cancer. Nature 611, 810–817 (2022).
Zhang, X. et al. Tissue-resident Lachnospiraceae family bacteria protect against colorectal carcinogenesis by promoting tumor immune surveillance. Cell Host Microbe 31, 418–432.e8 (2023).
Kaminski, J. et al. High-Specificity Targeted Functional Profiling in Microbial Communities with ShortBRED. PLoS Comput. Biol. 11, e1004557 (2015).
Liu, B., Zheng, D., Zhou, S., Chen, L. & Yang, J. VFDB 2022: a general classification scheme for bacterial virulence factors. Nucleic Acids Res. 50, D912–D917 (2022).
Schubert, M. et al. Perturbation-response genes reveal signaling footprints in cancer gene expression. Nat. Commun. 9, 20 (2018).
Grasso, C. S. et al. Genetic mechanisms of immune evasion in colorectal cancer. Cancer Discov. 8, 730–749 (2018).
Mezheyeuski, A. et al. An immune score reflecting pro- and anti-tumoural balance of tumour microenvironment has major prognostic impact and predicts immunotherapy response in solid cancers. EBioMedicine 88, 104452 (2023).
Mariathasan, S. et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Zeng, D. et al. Tumor microenvironment evaluation promotes precise checkpoint immunotherapy of advanced gastric cancer. J. Immunother. Cancer 9, e002467 (2021).
Jiang, P. et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat. Med. 24, 1550–1558 (2018).
Chen, B., Khodadoust, M. S., Liu, C. L., Newman, A. M. & Alizadeh, A. A. Profiling tumor infiltrating immune cells with CIBERSORT. Methods Mol. Biol. 1711, 243–259 (2018).
Gupta, V. K. et al. A predictive index for health status using species-level gut microbiome profiling. Nat. Commun. 11, 4635 (2020).
Uno, H., Cai, T., Pencina, M. J., D’Agostino, R. B. & Wei, L. J. On the C-statistics for evaluating overall adequacy of risk prediction procedures with censored survival data. Stat. Med. 30, 1105–1117 (2011).
Wang, W. et al. The China National GeneBank Sequence Archive (CNSA) 2024 update. Hortic. Res. 12, uhaf036 (2025).
Chen, F. Z. et al. CNGBdb: China National GeneBank DataBase. Yi chuan = Hered. 42, 799–809 (2020).
Cezard, T. et al. The European Variation Archive: A FAIR resource of genomic variation for all species. Nucleic Acids Res. 50, D1216–D1220 (2022).
Sarkans, U. et al. The BioStudies database-one stop shop for all data supporting a life sciences study. Nucleic Acids Res. 46, D1266–D1270 (2018).
Shi, Z. et al. Tissue-resident microbiota impacts colorectal cancer progression and prognosis. Zenodo https://doi.org/10.5281/zenodo.17588313 (2025).
Acknowledgements
The U-CAN tumor biobank was funded by a grant from the Swedish Government (CancerUU) to Uppsala University, Umeå University, KTH Royal Institute of Technology, and Stockholm University (2010-ongoing). This study was also funded by the National Science and Technology Major Project (No. 2025ZD0551700 to K.W.) and the Guangdong Provincial Key Laboratory of Human Disease Genomics (2020B1212070028 to K.W.). Data handling and storage was enabled by resources in project sens2019031 provided by the National Academic Infrastructure for Supercomputing in Sweden (NAISS) at UPPMAX, funded by the Swedish Research Council through grant agreement no. 2022-06725.
Author information
Authors and Affiliations
Contributions
Z.S., H.R., C.L., F.L., and H.Z. conceived the study; C.L., K.W., T.S., I.L., B.G., R.P., S.Z., Y.Z. M.U. and H.Z., coordinated the study; Z.S., H.R., C.L., F.L., M.W., T.L., F.Y., L.N., K.H., A.I., T.Z., B.G., and H.Z performed data curation; Z.S., H.R., and F.L., analyzed data; K.W., T.S., C.L., and H.Z. supervised the study; Z.S., H.R., C.L., F.L., T.S., and H.Z. wrote the paper with input from all other authors. All authors approved the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shi, Z., Ren, H., Lin, C. et al. Tissue-resident microbiota impacts colorectal cancer progression and prognosis. Nat Commun 17, 346 (2026). https://doi.org/10.1038/s41467-025-67047-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67047-2