Abstract
Migrating cells dynamically adapt their morphogenetic programs in response to microenvironmental changes, requiring coordinated spatiotemporal integration of mechanical and biochemical signals. The plasma membrane, through membrane tension and actin dynamics modulation, is essential for cell motility. Caveolae, small plasma membrane invaginations, act as mechanosensors to buffer tension changes under mechanical stress. Recent evidence suggests a role for caveolae in cell migration. Here, we demonstrate that breast cancer cells exhibit a front-rear asymmetry in caveolae and caveolin-1 scaffolds, which is regulated by membrane tension and is crucial for persistent migration and cell directionality. RhoA-driven cell contraction relies on the spatiotemporally coordinated assembly of caveolae and recruitment of RhoA-GEFs at the cell rear. These results are supported by a physical model establishing a feedback loop between local membrane tension and contractility, through caveolae formation and disruption. Our findings underscore the importance of caveolae mechanosensing in regulating RhoA activation and guiding cell migration.
Similar content being viewed by others
Introduction
Cell migration enables effective navigation through complex environments and is crucial in various physiological processes, such as gastrulation during development, immune response, tissue homeostasis, and wound healing1,2. In pathological conditions, such as cancer, cell migration, and invasion play an essential function during the metastatic process, contributing to tumor progression. Recent studies indicate that the biochemical and mechanical microenvironment of solid tumors play an important role in the control of their progression3. The cell surface, comprising the plasma membrane and the dynamic underlying actin cortex, is a major mechanobiological actor4,5. Migrating cells are polarized and transiently undergo cyclic phases of protrusions at the front and contractions at the rear. The plasma membrane is designed to respond to changes in mechanical properties by enabling transformations in shape, geometry, and specialized functions such as the regulation of cell polarity and motility5,6. However, the mechanistic link between mechanosensing and polarization dynamics of migrating cells remains poorly understood.
Caveolae are small, specialized invaginations of the plasma membrane composed of the main structural protein caveolin-1 (Cav1), cavins, sphingolipids, and cholesterol, and are associated with signaling molecules7,8. Cav1 oligomerizes in the Golgi apparatus, forming the minimal 8S complex made of 11 Cav1 protomers9. Once the 8S complexes are targeted and inserted into the inner leaflet of the plasma membrane, cytosolic cavins are recruited to stabilize the caveolae invagination as a budded caveola9,10,11. Recent studies have identified structurally distinct subpopulations of Cav1 assemblies at the plasma membrane, in addition to bona fide caveolae. These include the 8S complex (also known as the S1A scaffold) as well as higher-order Cav1 oligomers, such as the S1B and S2 scaffolds9,11,12. Caveolae have traditionally been associated with various cellular functions, including membrane and lipid trafficking13, plasma membrane organization14, migration6,15,16, and invasion14,17. More recently, we have identified a new role for caveolae as both mechanosensors and mechanoprotectors. Budded caveolae act as a membrane reservoir by flattening out to accommodate increase in membrane tension, such as those caused by mechanical stress18. Additionally, this mechanical disassembly of caveolae can initiate mechanotransduction events19, some of which are disrupted in diseases like muscular dystrophies20 and cancer19.
In this study, we investigated the role of caveolae mechanosensing and dynamics in regulating breast cancer cell migration. Our findings demonstrate that the front-rear asymmetry of Cav1 scaffolds and caveolae is critical for maintaining cell directionality during migration. Specifically, membrane tension drives the asymmetric distribution of caveolar structures, leading to the preferential localization of budded caveolae at the cell rear, while Cav1 scaffolds are predominantly found at the cell front. This asymmetry is crucial because budded caveolae, rather than Cav1 scaffolds, facilitate the recruitment and activation of the RhoA GTPase exchange factor ARHGEF25 at the cell rear, which triggers cell contraction. Furthermore, the accumulation of caveolae, spatiotemporally coupled with RhoA-GEF, precedes changes in cell directionality. This process is regulated by caveolae mechanosensing, as an increase in membrane tension—induced by hypo-osmotic shock—rapidly disassembles caveolae into Cav1 scaffolds and inhibits RhoA activation at the cell rear, ultimately halting cell migration. These findings underscore the central role of caveolae mechanosensing and dynamics in cell migration.
Results
Caveolae front-rear asymmetry drives directional reorientation of migrating cells
Asymmetrical localization of caveolae, with fewer at the cell front and more at the cell rear, has been observed in polarized migrating cells21,22,23,24. To investigate the role of caveolae mechanosensing during cellular symmetry breaking and directional orientation, we first analyzed single cell migration in various experimental setups. Triple-negative breast cancer Hs578t cells, genome-edited to express Cav1-RFP (Cav1-RFP Hs578t), displayed an asymmetrical distribution of Cav1. The Cav1 signal was significantly lower in the front (F) compared to the rear (R). Caveolae front-rear asymmetry was consistently observed during cell migration, with arrows indicating direction of migration (Fig. 1a and Supplementary Fig. 1a). In contrast, the membrane-anchored CAAX motif used as a negative control for polarization remained uniformly distributed along the plasma membrane during migration (Supplementary Fig. 1d). We stained cells for cavin1 to visualize bona fide caveolae and observed a similar front-rear asymmetry, with higher accumulation of Cav1 and cavin1 at the rear of migrating cells (Supplementary Fig. 1b and Supplementary Movie 1). Knockdown of cavin1 (siCavin1; cells retaining only non-caveolar Cav1) disrupted Cav1 front-rear asymmetry (Fig. 1a and Supplementary Fig. 1c) and led to a 53% reduction in migration speed and a 23% decrease in directionality (Fig. 1b, c). Knockdown of Cav1 (siCav1; cells lacking both caveolae and non-caveolar Cav1) resulted in 60% decrease in migration speed and a 48% reduction in directionality (Fig. 1b, c). These results confirm that caveolae front-rear asymmetry is essential for efficient and directed cell migration.
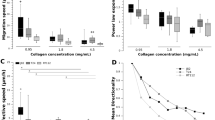
a Representative time-lapse images of Cav1-RFP Hs578t cells embedded in 3D collagen gels, expressing functional caveolae WT (siControl), non-caveolar Cav1 expression (siCavin1), or complete loss of caveolae (siCav1). White arrows indicate the direction of migration; red arrows indicate change in migration direction. b Quantification of migration speed (μm/min). c Directionality index (Euclidean/accumulated distance). c, d Data are presented as mean values +/- SEM., n = 34 (siControl), 36 (siCavin1), and 57 (siCav1) cells from three independent experiments. Statistical analysis: One-way ANOVA with Kruskal-Wallis test; ****P < 0.0001, *P < 0.01, ns = not significant. Time-lapse images and corresponding kymographs of Cav1-RFP Hs578t cells migrating on 1D line-micropatterns (d), 2D coverslips (e), and in 3D collagen gel under unipolar (f) or bipolar (g) Cav1 distribution. h Migration trajectory of Cav1-RFP MDA-MB-231 cells on Z-shaped micropatterns. Representative cell is shown from n = 75 cells, three independent experiments. Scale bars, 10 µm. F, front; R, rear. Source numerical data are provided.
We further examined the relationship between caveolae asymmetry and cell directionality. We monitored changes in caveolae distribution in cells forced to migrate on 1D line micropatterns (Fig. 1d and Supplementary Movie 2), randomly migrating on coverslips in 2D (Fig. 1e), and within 3D collagen gels (Fig. 1f and Supplementary Movie 3). When caveolae asymmetry was lost, as evidenced by Cav1 redistribution (Fig. 1d, f), cell migration stopped. Symmetry breaking was re-established by caveolae, which by forming a new rear allowed cell migration to resume in the opposite direction (Fig. 1d–f; red arrows). We observed a similar process in Cav1-RFP MDA-MB-231 cells that were forced to migrate and change their direction in Z-shaped micropatterns (Fig. 1h and Supplementary Movie 4). When cells changed direction, the loss of caveolae front-rear asymmetry was the first observable event, resulting in cessation of migration. The front-rear asymmetry was subsequently reestablished before the change of directionality occurred. These results confirm that caveolae front-rear asymmetry is essential for directional reorientation. This conclusion is quantitatively supported by kymographs of migrating cells (bottom panel Fig. 1d, f and Supplementary Fig. 1e,f), where white arrows indicate the initial direction of migration and red arrows indicate the reoriented direction. Additionally, we observed that some cells, following the initial establishment of caveolae front-rear asymmetry, assembled two distinct bi-polar pools of caveolae at opposing sides over time (Fig. 1g). When these caveolae pools became symmetrically distributed, the cells failed to migrate (kymograph Fig. 1g and Supplementary Fig. 1g), further underscoring the necessity of caveolae front-rear asymmetry for efficient and persistent cell migration.
Membrane tension regulates caveolae front-rear asymmetry and RhoA activation in migrating cells
Caveolae act as mechanosensors and serve as membrane reservoirs that flatten in response to elevated membrane tension, thereby protecting the plasma membrane from rupture18,25. In addition, caveolae function as mechanotransducers, as their mechanical disassembly is associated with the regulation of signaling events and gene transcription19,20. To investigate the role of caveolae mechanosensing in cell migration, we assessed membrane tension in Hs578t cells using fluorescence lifetime imaging microscopy (FLIM) with the Flipper-TR probe26. This probe intercalates into membranes and reports on membrane tension by measuring lipid packing and order—higher fluorescence lifetimes correspond to increased membrane tension. Under iso-osmotic conditions, Flipper-TR lifetime measurements revealed a front-rear gradient of membrane tension in migrating Hs578t cells, with lower tension at the cell rear and higher tension at the front (Fig. 2a–d), consistent with previous reports22,27,28,29. To perturb this gradient, we applied an acute hypo-osmotic shock (30 mOsm) to Cav1-RFP Hs578t cells, inducing a global increase in membrane tension. Under these conditions, Flipper-TR lifetimes indicated a uniform membrane tension across the cell, effectively abolishing the front-rear gradient (Fig. 2e–g). This was accompanied by a selective decrease in Cav1 intensity at the rear, while the front remained unaffected (Fig. 3a, b and Supplementary Movie 5).
a Representative Flipper-TR intensity images of a WT-Hs578t cell under isotonic (Iso) condition. White dashed boxes indicate front and rear regions of interest (ROIs). Color bar: Flipper-TR intensity (photons per pixel). Scale bar, 20 µm. b Fluorescence decay curves from front (green) and rear (red) ROIs of a representative cell. Experimental data (dots) were fitted with a two-exponential decay function (blue line) to calculate Flipper-TR lifetime (ns) and adjusted R² values. Insets show ROI locations. c Flipper-TR lifetimes at the front (green) and rear (red) of individual cells. Gray lines connect paired measurements from the same cell. b, c Data are presented as mean values +/- SEM of n = 44 cells from three independent experiments. Statistical analysis: One-tailed, Mann–Whitney test; ****P < 0.0001, **P < 0.001. d Per-cell differences in lifetime (Δ = Front - Rear); each blue circle represents one cell. e Representative Flipper-TR intensity images of a WT-Hs578t cell under isotonic (Iso, left) and hypotonic (Hypo, right) conditions. ROIs at the cell front and rear indicated by dashed boxes. Color bar as in panel a. Scale bar, 10 µm. f Flipper-TR lifetimes at the front and rear under Iso and Hypo conditions. (g) Per-cell differences in lifetime (Δ = Front - Rear) under Iso (dark blue circles) and Hypo (light blue circles). Gray lines link paired values; circles show per-cell differences. f, g Data are presented as mean values +/- SEM., n = 19 cells from three independent experiments. One-tailed, Mann–Whitney test; ****P < 0.0001. h Representative time-lapse images of WT-Hs578t cells co-expressing opto-PI3K (iSH2), SiR-actin, and Cav1-RFP. Insets show optogenetically activated ROIs. Timepoints span from -2 min (pre-activation) to +16 min (post-activation). i Quantification of normalized signal intensity and cell area in the activated ROI over time for actin (orange), Cav1 (gray), and cell area (blue). Data are presented as mean values +/- SEM of n = 17 cells from three independent experiments.
a Representative time-lapse images of Cav1-RFP Hs578t cells migrating in 3D collagen under isotonicity (Iso), 5 min after hypo-osmotic shock (Hypo), and 5 min after return to iso-osmotic conditions (Rec). White arrows indicate direction of migration. b Quantification of Cav1 fluorescence intensity. Mean ± SEM. n = 31 cells from three independent experiments. Statistical analysis: Two-tailed, one-way ANOVA with Šídák multiple comparisons; ****P < 0.0001; ns = not significant. c Time-lapse images of Cav1-RFP Hs578t cells migrating in 3D collagen under isotonicity (Iso), 30 min after hyper-osmotic shock (Hyper), and 30 min after return to iso-osmotic conditions (Rec). d Quantification of Cav1 fluorescence intensity. Mean ± SEM., n = 52 cells from three independent experiments. Two-tailed, one-way ANOVA with Šídák multiple comparisons; ****P < 0.0001; ns = not significant. e Representative RhoA biosensor (3x-rGBD, RhoA-bio) activity in WT-Hs578t, cavin1-KO, Cav1-KO, and Cav1-KO cells re-expressing wild-type Cav1-RFP (Cav1-KO+wtCav1). White arrows indicate direction of migration. f Quantification of RhoA-bio fluorescence intensity. Mean ± SEM., n = 44 (WT-Hs578t), 45 (cavin1-KO), 43 (Cav1-KO), and 39 (Cav1-KO+wtCav1) cells from three independent experiments. Two-tailed, one-way analysis of variance (ANOVA) with Dunn’s multiple comparisons; *P < 0.01; ns = not significant. g RhoA-bio activity under Iso, Hypo, and Rec conditions in the same genotypes. h Quantification of RhoA-bio fluorescence intensity. i Quantification of Cav1 fluorescence intensity. Data are presented as mean values +/- SEM., n = 44 (WT), 43 (Cav1-KO), and 39 (Cav1-KO+wtCav1) cells from three independent experiments. h, i Two-tailed, one-way ANOVA with Šídák multiple comparisons; ***P < 0.0001; **P < 0.001; *P < 0.01; ns = not significant. Scale bars, 10 μm (a, c), 20 μm (e, g). F, front; R, rear. Fluo., fluorescence. Source numerical data are provided.
To locally modulate membrane tension, we used an optogenetic approach to activate PI3 kinase (PI3K) at the cell rear in Cav1-RFP Hs578t cells expressing opto-PI3K (Fig. 2h, I; ref. 28.). Opto-PI3K activation of at the cell rear triggered Rac1-mediated filopodia formation (cyan line) and an increase in cell area within the opto-ROI (gray line), consistent with localized membrane tension increase. This was followed by a decrease in Cav1 intensity (orange line), indicative of caveolae disassembly. The loss of Cav1 asymmetry coincided with an immediate arrest of cell migration. Upon return to isotonic conditions (300 mOsm), membrane tension relaxed, Cav1 levels at the rear were restored, front-rear polarity re-established, and cell migration resumed (Fig. 3a, b and Supplementary Fig. 2b). Conversely, a global decrease in membrane tension induced by hyper-osmotic shock (850 mOsm, Hyper) increased intensity of Cav1 at both front and rear (Fig. 3c, d). However, this enhanced caveolae asymmetry did not accelerate migration, suggesting this process becomes limited at lower membrane tension and/or higher caveolae density (Supplementary Fig. 2c). Finally, using photoconvertible Cav1-Dendra2 expressed in WT-Hs578t cells, we observed that caveolae reassembled at the initial rear during the recovery phase (white arrowheads, bottom panel), ruling out caveolae endocytosis or trafficking as the mechanism for re-establishing caveolae front-rear asymmetry (Supplementary Fig. 2a). Together, these results demonstrate that caveolae front-rear asymmetry is regulated by membrane tension and plays a crucial role in directional cell migration.
Cell migration depends on a highly dynamic and coordinated biochemical cycle involving the Rho GTPase subfamily. Specifically, Rac1 and Cdc42 are localized at the leading edge of migrating cells, where they drive processes such as lamellipodia and filopodia formation. In contrast, RhoA regulates cytoskeletal contraction and focal adhesion turnover at the rear of the cell30,31. Caveolae and caveolin-1 have been implicated in RhoA-mediated contractility, as well as in cell migration and invasion15,16,32,33. We investigated the potential correlation between caveolae front-rear asymmetry and the levels and localization of active RhoA (RhoA-GTP) in migrating cells. Cav1-KO Hs578t cells exhibited an 80% reduction in overall levels of GTP-bound RhoA, while cavin1-KO Hs578t cells showed a 35% decrease, suggesting that the presence of caveolae is essential for the global activation of RhoA (Supplementary Fig. 3a). Immunofluorescence analysis further confirmed a significant decrease in RhoA-GTP levels in Cav1-KO and cavin1-KO Hs578t cells (Supplementary Fig. 3c). We monitored the dynamics of active RhoA in migrating WT and KO Hs578t cells by transiently expressing HaloTag-3xrGBD (RhoA-bio), a biosensor for active RhoA34,35 (Supplementary Fig. 3b). White arrows indicate localized sites of active RhoA associated with cell contraction (Supplementary Fig. 3d and Supplementary Movie 6). Notably, higher levels were localized at the rear of WT-Hs578t cell, establishing a front-rear polarity for active RhoA. In contrast, active RhoA polarization was absent in cavin1-KO and Cav1-KO cells but was restored upon expression of wild-type Cav1 in Cav1-KO cells (Fig. 3e, f). These findings confirm that bona fide caveolae and their front-rear polarity are essential for maintaining the polarity of active RhoA in migrating cells.
We further investigated whether mechanical stress affects the polarity of active RhoA. In WT-Hs578t cells, a global increase in membrane tension induced by hypo-osmotic shock led to a 16% reduction in active RhoA levels at the rear, with no significant change at the front (first panel, Fig. 3g, h). In Cav1-KO cells, the levels of active RhoA were insensitive to mechanical stress at either the front or rear (middle panel, Fig. 3g, h). Remarkably, in Cav1-KO cells re-expressing wild-type Cav1, the sensitivity of rear active RhoA to global membrane tension increase was restored at the rear (12% decrease at the rear, with no effect at the front), similar to WT cells (last panel, Fig. 3g, h). Analysis of Cav1 intensity under each condition, revealed a strict correlation between active RhoA and Cav1 levels at both the rear and the front of the cell (Fig. 3i). These results suggest that caveolae mechanosensing plays a crucial role in regulating active RhoA polarity in migrating cells.
Active RhoA preferentially interacts with Cav1 scaffolds
Co-immunoprecipitation, proximity ligation assay (PLA), and colocalization analyses revealed that Cav1 and active RhoA interact similarly in WT-Hs578t and cavin1-KO cells (Fig. 4a–d, f and Supplementary Fig. 4d). Co-immunoprecipitation experiments revealed an increased interaction between Cav1 and active RhoA under hypo-osmotic shock (Fig. 4e). Additionally, PLA experiments confirmed this effect with an increased proximity between active RhoA and Cav1 (Fig. 4g). These findings strongly suggest that active RhoA is more likely to interact with Cav1 when caveolae have been disassembled into scaffolds. Indeed, a higher level of colocalization between active RhoA and Cav1 was observed in cavin1-KO cells that present only non-caveolar Cav1, albeit at a lower expression level (Fig. 4a and Supplementary Fig. 4d).
a Co-immunoprecipitation of endogenous Cav1 from lysates of WT-Hs578t and Cav1- or cavin1- knocked out (KO) Hs578t cells to assess interaction with RhoA-GTP. Representative immunoblots shown. b Quantification of RhoA-GTP relative to total RhoA (input), RhoA-GTP in IP, RhoA relative to tubulin (input), and RhoA-GTP normalized to Cav1 signal. Data are presented as mean values +/- SEM., n = 4 independent experiments. Statistical tests: RhoA-GTP (input): Two-tailed, one-way ANOVA with Tukey’s multiple comparisons; *P < 0.01. ns = not significant; RhoA-GTP (IP): Two-tailed, Mann-Whitney test; *P < 0.01; RhoA (input): Two-tailed, one-way ANOVA with Dunn’s multiple comparisons; ns = not significant; RhoA-GTP/Cav1: Two-tailed, one-way ANOVA with Šídák multiple comparisons; ns = not significant. c GST-Rhotekin-RBD pulldown of active RhoA to detect associated Cav1 in WT, Cav1-KO, and cavin1-KO Hs578t cells. Representative immunoblots shown. d Quantification of RhoA-GTP relative to total RhoA, RhoA relative to tubulin, and RhoA-GTP relative to Cav1. Data are presented as mean values +/- SEM., n = 4 independent experiments. Statistical tests: RhoA-GTP/RhoA: Two-tailed, one-way ANOVA with Tukey’s multiple comparisons; ***P < 0.0001; *P < 0.01; ns = not significant; RhoA/tubulin: Two-tailed, one-way ANOVA with Šídák multiple comparisons; ns = not significant; RhoA-GTP/Cav1: Two-tailed, Mann-Whitney test; ns = not significant. e Co-immunoprecipitation of Cav1 and RhoA-GTP from WT-Hs578t under isotonic (Iso), hypo-osmotic (Hypo), and recovery (Rec) conditions. Representative immunoblots shown. Quantification of RhoA-GTP relative to Cav1 shown. Data are mean ± SEM., n = 4 independent experiments. Two-tailed, one-way ANOVA with Kruskal-Wallis test; *P < 0.01. f Proximity ligation assay (PLA) assay to detect endogenous Cav1-RhoA-GTP interactions in WT, Cav1-KO, and cavin1-KO Hs578t cells. Quantifications show median ± 95% CI; n = 326 cells from 3 independent experiments. Two-tailed, one-way ANOVA with Kruskal-Wallis test; ****P < 0.0001; **P < 0.001. g PLA analysis to detect endogenous Cav1-RhoA-GTP interactions in WT-Hs578t cells under Iso, Hypo, and Rec conditions. Quantifications as median ± 95% CI; n = 314 cells from 3 independent experiments. Two-tailed, one-way ANOVA with Kruskal-Wallis test; ****P < 0.0001; **P < 0.001; ns = not significant. Scale bars, 20 µm. Fluo., fluorescence. Source numerical data and unprocessed blots are provided.
The interaction of Cav1 with downstream effectors is often mediated by a conserved 20-amino acid sequence known as the scaffolding domain (CSD)12. To determine whether the Cav1-CSD is necessary for the interaction with active RhoA, we transiently expressed comparable levels of wt-Cav1RFP (wtCav1) and F92A/V94A CSD Cav1RFP (F92A/V94A CSD Cav1), a mutated form with an inactive CSD, in Cav1-KO Hs578t cells (Supplementary Fig. 4a). Co-immunoprecipitation, PLA, and colocalization analyses demonstrated that the interaction between active RhoA and both wtCav1 and F92A/V94A CSD Cav1 remained unchanged (Supplementary Fig. 4b–d). This indicates that active RhoA does not interact with Cav1 through the CSD, in line with previous studies36.
Non-caveolar Cav1 assemblies, organized into scaffolds of defined sizes, have been characterized using stochastic optical reconstruction microscopy (STORM) and single-molecule localization microscopy (SMLM) network analysis11. The smallest class, termed S1A scaffolds, correspond to the minimal 11-Cav1 protomer-8S complex, as resolved by cryo-electron microscopy9. S1B scaffolds represent dimers of S1A, while S2 scaffolds correspond to larger hemispherical structures formed by the association of several S1A scaffolds11. To examine the spatial contributions of caveolae and Cav1 scaffolds during cell migration, we applied the same computational network modeling of STORM Cav1 localizations in Hs578t cells. This analysis revealed that budded caveolae were predominantly localized at the cell rear, while Cav1 scaffolds, primarily from the S1A population, were concentrated at the cell front (Fig. 5a, b). Upon hypo-osmotic shock, this front-rear asymmetry was markedly perturbed: the caveolae pool at the rear decreased by 24%, while S1A scaffold at the front increased by 23%, relative to cells at rest (Supplementary Fig. 5a,b). These changes were reversed upon restoration of isotonic conditions, indicating a tension-dependent dynamic redistribution of Cav1 assemblies (Supplementary Fig. 5c,d). Importantly, the reduction in caveolae at the rear under elevated tension (−24%) correlated with a decrease in active RhoA in the same region (−16%) (Fig. 3g, h), suggesting a functional link between caveolae disassembly and RhoA signaling. In contrast, cavin1-KO Hs578t cells, which lack budded caveolae, exhibited only S1A and S1B scaffolds, with both structures symmetrically distributed along the cell axis (Fig. 5c, d), further supporting the role of caveolae asymmetry in regulating directional migration.
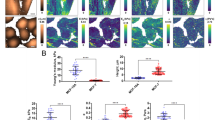
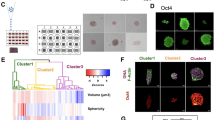
a Representative widefield image of a migrating WT-Hs578t cell under isotonic (Iso) condition, stained for Cav1 (cyan) and RhoA-GTP (magenta). b Quantifications of Cav1 structures in WT cells based on SuperResNET classification of cluster segmentation images, using parameters described in ref. 11. Cav1 structures are categorized as caveolae, S2, S1B, or S1A scaffolds. Cell edges are delineated by RhoA-GTP signal. Data are presented as mean values ± SEM., n = 6 cells from 3 independent experiments. Two-tailed, paired t-test; *P < 0.01; ns = not significant. c Same as (a), for a cavin1-KO Hs578t cell. Zoomed 2D dSTORM images of the cell front (F) and rear (R) are shown for both cell types. d Quantifications of Cav1 structures in cavin1-KO Hs578t cells. Data are presented as mean values ± SEM., n = 7 cells from 3 independent experiments. Two-tailed, paired t-test; ns = not significant. Cluster segmentation visualizes SuperResNET classification results. Scale bars, 10 µm (widefield), 5 µm (STORM), 2 µm (zoomed). e Time-lapse images of migrating WT-Hs578t, cavin1-KO, Cav1-KO, and Cav1-KO+wtCav1 cells expressing an active RhoA biosensor. f Morphodynamic map of a representative migrating cell; X-axis = time, Y-axis = cell contour. Blue indicates contractile regions (white circle), red indicates protrusions. g Quantification of contractile events per hour across genotypes. Data are presented as mean values ± SEM., n = 38 (WT), 46 (cavin1-KO), 42 (Cav1-KO), and 43 (Cav1-KO+wtCav1) cells from 3 independent experiments. Two-tailed, one-way ANOVA with Kruskal-Wallis test; ****P < 0.0001; ***P < 0.001; ns = not significant. Scale bars, 20 µm (e), 5 µm (f). Source numerical data are provided.
Altogether, these data suggest that active RhoA is more likely to interact with Cav1 scaffolds than with bona fide caveolae. We further establish this point by applying hypo-osmotic shock to WT-Hs578t, as this condition results in a significant conversion of caveolae into Cav1 scaffolds at the rear of the cell, while it has no significant effect on the amount of Cav1 scaffolds present at the front (Supplementary Fig. 5a,b). Cav1 and active RhoA showed a significantly higher level of colocalization at the rear, with an 87% increase, and a non-significant increase at the front (Supplementary Fig. 6a). Colocalization levels returned to initial levels when membrane tension homeostasis was restored (Supplementary Fig. 6a). Taken together, these data further support the role of caveolae mechanosensing in regulating RhoA activity.
Caveolae, but not Cav1 scaffolds, are required for efficient actomyosin contractility
During migration, the formation of stable lamellipodia at the cell’s leading edge front protrusions involves cycles of actin polymerization and depolymerization37. In contrast, RhoA-mediated actomyosin contraction generates the tension necessary to detach the cell rear, facilitating forward movement38. To better understand the spatiotemporal regulation of actomyosin contractility by RhoA, we performed a morphodynamic analysis of individual migrating cells during edge contraction to correlate caveolae and Cav1 scaffolds functions with cell contractility. We performed time-lapse imaging of WT-Hs578t, cavin1-KO, Cav1-KO, and Cav1-KO+wtCav1 cells expressing the active RhoA biosensor (Fig. 5e). Morphodynamic maps of the cell contour allowed us to visualize contraction activity and active RhoA over time39,40. These maps were constructed by measuring cell contour displacement between two consecutive time points, with displacement color-coded: blue indicating contraction and red indicating protrusion (Fig. 5f, Methods). Contraction events associated with active RhoA (white circles and blue spots) were analyzed across all conditions (Fig. 5f). The data show that caveolae significantly accelerate RhoA-mediated actomyosin contractility with 2.4 events per hour, compared to Cav1 scaffolds in cavin1-KO cells, which exhibit 0.9 events per hour (Fig. 5g). Furthermore, re-expression of wild-type Cav1 in Cav1-KO cells restored the contraction rate from 0.4 events per hour to 1.9 events per hour, confirming that budded caveolae are essential for efficient RhoA-mediated actomyosin contractility. Our data demonstrate that while active RhoA preferentially interacts with Cav1 scaffolds (Fig. 4), only budded caveolae significantly enhance actomyosin contractility (Fig. 5e–g).
Spatiotemporal regulation of ARHGEF25 and ARHGEF11 by caveolae
Guanine nucleotide exchange factors (GEF) act as upstream activators of RhoA GTPase31. To identify potential mediators of caveolae-regulated actomyosin contractility, we investigated the interaction between RhoA-specific GEFs and caveolae at contracting membrane sites in migrating cells. We selectively recruited RhoA GEFs locally through optogenetics. Among the ~ 14 RhoA-specific GEFs involved in migration that do not affect other Rho family members such as Rac1 and cdc4241, we selected ARHGEF25 and ARHGEF11 for their established roles in RhoA activation and promotion of cell contraction35,42.
Optogenetic stimulation of ARHGEF25 and ARHGEF11 was applied at the front of migrating cells, enabling visualization and quantitative analysis of Cav1 and GEF temporal dynamics at the newly formed rear of the cell. This setup allowed us to track the cellular changes preceding directional re-orientation. We measured ARHGEF25 intensity within the opto-ROI prior to photoactivation including only cells with comparable expression levels (Supplementary Fig. 7e,f). Upon blue-light photoactivation, all conditions—WT-Hs578t, cavin1-KO, cavin1-KO+cavin1, Cav1-KO, and Cav1-KO+wtCav1—displayed the characteristic contractile behavior associated with RhoA-induced actomyosin activity. Notably, WT-Hs578t cells exhibited a Cav1 intensity peak that coincided with ARHGEF25 recruitment within the opto-ROI (Fig. 6a and Supplementary Movie 7). In contrast, cells transfected with the control plasmid containing only the light-sensitive component (Lck-mTurquoise2-iLID) showed no activation of RhoA and exhibited reduced membrane movement throughout the experiment (Supplementary Fig. 7a,b), ruling out the effect of blue light alone. Additional random-ROIs, positioned away from the opto-ROI in all cells, confirmed the specificity of the optogenetic tool (Supplementary Fig. 7c,d).
a–e Representative time-lapse images of Hs578t cells expressing optogenetic constructs (Lck-mTurquoise2-iLID, SspB-HaloTag-ARHGEF25-DH (ARHGEF25), and Cav1-RFP), in the following conditions: (a) WT, (b) cavin1-KO, (c) cavin1-KO+cavin1, (d) Cav1-KO, (e) Cav1-KO+wtCav1. Insets show zoomed ROI corresponding to optogenetic activation. Timepoints span from -2 min (pre-activation) to +16 min (opto-activation). Normalized intensities of ARHGEF25 (orange), Cav1 (gray), and cell area (blue) in the opto-activated ROI are plotted over time. Data are presented as mean values ± SEM., n = 21 (WT), 16 (cavin1-KO), 22 (cavin1-KO+cavin1), 19 (Cav1-KO), and 23 (Cav1-KO+wtCav1) cells from 3 independent experiments. f Representative time-lapse images of WT- and cavin1-KO Hs578t cells expressing Halo-ARHGEF25 and Cav1-RFP. White arrow indicate direction of migration. g Quantification of ARHGEF25 and Cav1 fluorescence intensity at the front and rear. h Quantification at the cell periphery. Data are presented as mean values ± SEM., n = 32 (WT) and 43 (cavin1-KO) cells from 3 independent experiments. Statistical tests: Two-tailed, Wilcoxon matched-pairs signed-rank test (g); Two-tailed, Mann-Whitney t-test (h); ****P < 0.0001; ns = not significant. i Representative time-lapse images of random contracting events in WT- and cavin1-KO Hs578t cells expressing Halo-ARHGEF25 and Cav1-RFP. Plots show normalized intensity and cell area: ARHGEF25 (black), Cav1 (purple), and cell area (blue). Data are presented as mean values ± SEM., n = 90 (WT) and 42 (cavin1-KO) contraction events pooled from 3 independent experiments. Scale bars, 10 µm. F, front; R, rear. Fluo., fluorescence. Source numerical data are provided.
Within the opto-ROI, the contracting membrane decreased its area by approximately 20% within ~ 6 minutes (blue line, Fig. 6a). Cavin1-KO cells, lacking a Cav1 signal peak linked with ARHGEF25, required over 50 minutes to contract (Fig. 6b and Supplementary Movie 8). However, cavin1-KO cells re-expressing cavin1 exhibited a restored Cav1 peak concurrent with ARHGEF25 recruitment and achieved 20% decrease in membrane area in ~8 minutes (Fig. 6c). In contrast, Cav1-KO cells contracted at a significantly slow rate, taking over 70 minutes, despite effective ARHGEF25 recruitment via opto-activation (Fig. 6d). Re-expression of wtCav1 in Cav1-KO cells restored the Cav1 intensity peak coupled with ARFGEF25 (Fig. 6e). Cav1 intensity changes within the opto-ROI and membrane area contraction dynamics were compared across conditions Supplementary Fig. 7g,h).
To further investigate the spatiotemporal interaction between ARHGEF25 and Cav1, we performed measurements on randomly migrating cells. We generated morphodynamic maps of cell contours and analyzed individual contraction events to quantify ARHGEF25 and Cav1 intensities over time. Consistent with the optogenetic experiments, we found that in WT-Hs578t cells, ARHGEF25 and Cav1 were spatiotemporally coupled, with synchronous recruitment to contracting membrane regions. In contrast, cavin1-KO Hs578t cells showed no increase of Cav1 recruitment (Fig. 6i), despite ARHGEF25 membrane levels being similar to those in WT cells (Fig. 6f, h). Notably, ARHGEF25 displayed a polarized distribution in WT cells, with higher levels at the cell rear, whereas cavin1-KO cells displayed no front-rear polarity (Fig. 6g). We performed a parallel experiment using an optogenetic approach to recruit another RhoA-specific GEF, ARHGEF11, in WT, cavin1-KO, and Cav1-KO Hs578t cells (Supplementary Fig. 8a,b). In WT cells, Cav1 intensity increased prior to contraction, whereas no such increase was observed in cavin1-KO cells. Moreover, contraction kinetics were impaired: contraction times were two- and three-fold slower in cavin1-KO and Cav1-KO cells, respectively, compared to WT cells (Supplementary Fig. 8c,d). As a negative control, we optogenetically activated Tiam1, a Rac1-specific GEF, in WT-Hs578t cells. Photoactivation triggered local recruitment of Tiam1 and a corresponding increase in cell area within the opto-ROI, consistent with Rac1-mediated protrusion. However, Cav1 intensity remained unchanged during this response, indicating that Cav1 is not spatiotemporally coupled with Tiam1 (Supplementary Fig. 8e–f). Together, these results demonstrate that caveolae are selectively and dynamically coupled to specific RhoA-GEFs, such as ARHGEF25 and ARHGEF11, enabling rapid responses to cues for directional change and promoting efficient contraction. In contrast, Cav1 scaffolds alone are not coupled to RhoA-GEFs, resulting in delayed or impaired contractile responses and directional migration.
Caveolae assembly is a prerequisite for RhoA activation at contraction sites
Finally, we examined the temporal coupling between active RhoA and Cav1 at the membrane contraction edge of migrating cells. Contraction events were identified from morphodynamic maps as previously described. Our temporal intensity correlation analysis revealed that Cav1 intensity displayed a positive peak approximately 2 minutes before the appearance of active RhoA at the contracting sites in WT-Hs578t cells (Fig. 7a, b and Supplementary Movie 9). This Cav1 peak was absent at the contracting sites of cavin1-KO cells, which showed a slower contraction rate, as evidenced by the contracting area over time (Fig. 7c, d). Notably, Cav1-KO cells re-expressing wtCav1 exhibited a positive Cav1 intensity peak roughly 2 minutes before active RhoA appeared at the contraction sites (Fig. 7e, f). The contractile membrane area dynamics over time were comparable between WT and Cav1-KO+wtCav1 but were significantly delayed in cavin1-KO cells (Fig. 7b–d, f). Overall, these results confirm that caveolae play a critical role in sensing and executing migratory signals by accelerating RhoA activation through the recruitment of RhoA-GEFs at contracting sites. In contrast, Cav1 scaffolds are unable to recruit RhoA-GEFs, which significantly delays the contraction process.
a Representative time-lapse images of WT Hs578t cells expressing wild-type Cav1-RFP and a RhoA activity biosensor (RhoA-bio), showing a random contraction event. b Quantification of normalized intensity over time for RhoA-bio (purple), Cav1 (orange), and cell area (blue) at contraction sites. Data are presented as mean values ± SEM., n = 109 contraction events from 45 cells pooled from 3 independent experiments. c Time-lapse images of cavin1-KO cells under the same conditions. d Corresponding quantification of normalized RhoA-bio, Cav1, and cell area. Data are presented as mean values ± SEM., n = 68 contraction events from 45 cells from 3 independent experiments. e Time-lapse images of Cav1-KO cells re-expressing wtCav1 and RhoA-bio. f Data are presented as mean values ± SEM., n = 73 contraction events from 45 cells from 3 independent experiments. Scale bars, 10, µm. Fluo., fluorescence. Source numerical data are provided.
A physical model explains cell persistence through feedback between local membrane tension and contractility
To understand how the local regulation of contractility by membrane tension influences the persistence of cell movement, we developed a minimal physical model of cell motility (Fig. 8a). The model, described in detail in the Supplementary Information, is one-dimensional (1D) and designed to represent a cell moving along a linear track (Fig. 1d). It consists of three beads corresponding to the cell body and its two leading edges. These beads are connected by elastic springs that mimic local membrane tension (\({\sigma }_{m,i}\)), while the cell edges are subjected to active contractile forces \(({\sigma }_{i})\), with i = 1,2 representing the left and right edges, respectively. The motion of each bead follows a local force balance between the active force, membrane tension, and a friction force with the substrate proportional to the bead velocity. The cell stochastically switches between a quiescent state (\({\sigma }_{1}={\sigma }_{2}=0\)) and polarized states that promote motion in either direction (Left: \({\sigma }_{1}=-1,\,{\sigma }_{2}=1\); Right: \({\sigma }_{1}=1,\,{\sigma }_{2}=-1\)). Transitions to the quiescent state occur at a constant rate \({k}_{d}\), whereas transitions to polarized states depend on the local membrane tension via a feedback mechanism mediated by caveolae (Fig. 8b). In cells lacking this feedback such as Cav1-KO cells, transitions to contractile states occur independently and symmetrically at both cell edges at a baseline rate \({k}_{u0}\). In contrast, when feedback is present, the probability of edge contraction depends on local membrane tension, such that: \({k}_{u,i}={k}_{u,0}+{\delta k}_{u}{\sigma }_{m,i}\), where \({\delta k}_{u}\) quantifies the strength of the feedback. This parameter is reduced in cavin-1-KO cells compared to WT cells and is set to zero in Cav1-KO cells. If the calculated \({k}_{u,i}\) becomes negative due to high tension, it is capped at zero in simulations. Local membrane tension is directly related to the extension of the corresponding cell edge, and its relaxation occurs at a rate \({k}_{\gamma }\), determined by the ratio of the spring constant to the edge friction. We also allow for differential friction between the cell edges (with friction coefficient \({\gamma }_{g}\)) and the cell body (\(\gamma\)), defining the ratio \(\mu={\gamma }_{g}/\gamma\). As shown in Supplementary Information (SI 1), the tension difference between cell edges, which biases cell movement, relaxes at an effective rate \({k}_{\gamma }^{{\prime} }={k}_{\gamma }\,(1+2{\rm{\mu }})\).
a Model schematic. The motile cell is represented by three beads: a central bead (cell body) flanked by two side beads representing the leading and trailing edges\(\,(i=\{\mathrm{1,2}\})\). Each edge experiences stochastic active forces \({\sigma }_{i}\), local membrane tension \({\sigma }_{{mi}}\) (modeled as springs), and substrate friction. b Stochastic switching of actives forces. Active contractile forces alternate between high and low states. Feedback is implemented by coupling the transition rate from low to high contractility to membrane tension, such that increased \({\sigma }_{m,i}\) reduces the activation probability: \({k}_{u,i}={k}_{u0}-{\delta k}_{u}{\sigma }_{m,i}\). c Simulation results. Typical trajectories from stochastic simulations with and without feedback. Parameters were determined by experimental observations: the duration of contraction events gives \({k}_{d}\approx 0.1/\) min (Fig. 4f); the contraction frequency without feedback (siCav1) yields \({k}_{u0}\approx 0.5/\) hr (Fig. 4g); and the relaxation dynamics after optogenetic activation of contractility set \({k}_{{\rm{\gamma }}}\approx 0.1/\) min (Fig. 5a). The key parameter for directionality \({\delta k}_{u}\) is fitted to reproduce the contraction frequency with feedback in WT cells ≈2.5/hr. d Directionality as a function of contractile activity. The directionality of motion, measured as the ratio of end-to-end distance over total path length during observation time \({t}_{{obs}}\), increases with the number of contractions per hour in cells with feedback, consistent with experimental data. In contrast, without feedback (random walk), directionality decreases as \(\left\langle \,{x}^{2}\right\rangle /L\) ~ 1/\(\sqrt{{\rm{k}}{{t}}_{{obs}}}\). e Cartoon model. In WT-Iso conditions, the cell front exhibits fewer caveolae and more Cav1 scaffolds, preventing RhoA activation, whereas the rear accumulates caveolae colocalized with RhoA-GEFs, promoting RhoA-GTP-mediated contractility and persistent migration. Under mechanical stress (WT-Hypo) or in cavin1-KO cells, front-rear caveolae/Cav1 scaffold asymmetry is lost, leading to reduced GEF and RhoA activation and nondirectional migration. f Caveolae asymmetry and reorientation. (1) Migrating cells show enrichment of caveolae at the rear (R) and Cav1 scaffolds at the front (F), establishing front-rear polarity. (2) Changes in membrane tension or imposed directional shifts disrupt this asymmetry, leading to a temporary halt in migration. (3) Caveolae then reassemble, restoring asymmetry, which precedes (4) cell reorientation and the formation of a new front-rear axis.
We performed stochastic simulations of this model using parameters chosen to reproduce the contractile activity observed in WT, Cavin-1-KO and Cav1-KO cells (Fig. 5g). We then compute cell directionality as the net displacement (end-to-end distance) divided by the total path length over 4 hours, as in (Fig. 1c). Representative trajectories for cells with (WT) and without (Cav1-KO) feedback are shown in (Fig. 8c). The key indicator of the effect of feedback is the correlation between contractile activity and directionality. Directionality is defined as the ratio of root-mean-square (Euclidean) displacement to the total distance traveled over a given observation period \({t}_{{obs}}\). In a purely random walk—where steps in either direction occur at a constant rate \({k}_{u,0}\)—this ratio is \(\sqrt{\frac{ < {x}^{2} > }{L({t}_{{obs}})}}=\frac{1}{\sqrt{{k}_{u,0}{t}_{{obs}}}}\), and thus, directionality decreases with increasing contractile events (\({ \sim k}_{u,0}{t}_{{obs}}\)). This behavior is reproduced in simulations without feedback (\({\delta k}_{u}=0\)). Strikingly, introducing feedback reverses this relationship: directionality increases with contractile activity (Fig. 8d). This provides direct evidence that feedback between local membrane tension and contractility—mediated by caveolae—enhances short-term directional persistence. Additional analytical and stochastic simulation results are show in Supplementary Fig. 9.
The model also offers insight into the mechanism by which this feedback improves directionality. When a contractile event is initiated at one end of the cell—either stochastically or via optogenetic stimulation (Fig. 6)—a protrusion forms at the opposite end, driving cell movement. This leads to a local drop in membrane tension at the rear and an increase at the front (see also Supplementary Fig. 9): in the model (Fig. 8a), the rear spring compresses while the front spring extends. The cell returns to a quiescent state after a characteristic time of order \({1/k}_{d}\), yet the tension gradient persists for a time of order \({1/k}_{\gamma }^{{\prime} }\) governed by substrate friction. This residual tension gradient biases subsequent contractile events toward the same direction, thereby reinforcing directional persistence.
Discussion
Cells in migration experience various mechanical cues, such as substrate stiffness, topography, fluid shear stress, and other strains, in their environment43. For efficient migration, cells must rapidly adapt their morphogenetic program to constantly shifting microenvironments, necessitating precise spatiotemporal integration of mechanical and biochemical signals. However, it remains unclear how migrating cells perceive and process these mechano-chemical stimuli44. Recent studies suggest that caveolae and the RhoA signaling pathway regulate actomyosin contractility, especially at the cell rear22,45. Yet, the mechanisms by which caveolae interpret mechanical cues at both the cell front and rear, through dynamics and mechanosignaling initiation are not well understood.
In this study, we investigated how the mechanosensing and mechanotransducing functions of caveolae regulate cell migration and directional reorientation. Consistent with previous reports, we observed that caveolae exhibit front-rear asymmetry in WT-Hs578t and MDA-MB-231 triple-negative breast cancer cells migrating on uniformly stiff substrates21,23,24. This finding extends our understanding on cell migration, as previous studies reported caveolae asymmetry primarily on durotactic substrates22. Using super-resolution microscopy combined with 3D machine learning analysis, we demonstrated that bona fide caveolae predominantly localize at the rear of the migrating cell, whereas Cav1 scaffolds, recently identified as the 8S by cryo-electron microscopy9, are enriched at the front. Our comparative analysis of the cell front and rear revealed that maintaining an appropriate ratio of budded caveolae to Cav1 scaffolds is critical for efficient rear contractility, directional reorientation, and persistent migration. We further show that elevated membrane tension, induced by osmotic shock, significantly increases Cav1 scaffolds at the rear while reducing budded caveolae. These observations suggest that a sustained membrane tension gradient along the front-rear axis is essential for stabilizing caveolae at the rear and supporting their polarization in migrating cells. This is consistent with previous reports suggesting that lower membrane tension at the rear aids in contraction and substrate detachment, while higher tension at the front supports protrusion formation5. Although the existence of spatially regulated membrane tension gradients remains debated27,28, recent studies using the Flipper-TR fluorescent probe have detected tension asymmetries in adherent cells, with lower tension at the rear and higher tension at protrusions29. Interestingly, lower membrane tension was measured at the rear of cells rapidly migrating within a 3D matrix, while no tension gradient was observed in other setups where caveolae asymmetry was absent22. Our own Flipper-TR lifetimes measurements confirmed the presence of a front-rear membrane tension gradient in Hs578t cells. These findings provide a mechanistic explanation for the observed front-rear asymmetry of caveolae: low membrane tension at the rear favors the maintenance of budded caveolae, whereas elevated front tension promotes their disassembly into Cav1 scaffolds. Together, our results demonstrate that membrane tension regulates caveolae front-rear asymmetry and that this polarization plays a crucial role in guiding directional cell migration.
Caveolae’s mechanosensing properties have been increasingly linked to mechanotransduction pathways19,20,46. The Rho GTPases Rac1/Cdc42 and RhoA are core signaling regulators in both mesenchymal and amoeboid cell migration, integrating extracellular and intracellular cues to modulate the actin cytoskeleton and drive cell protrusion and contraction, respectively47. Our findings suggest that caveolae mechanosensing during migration regulates RhoA signaling by modulating RhoA activity at the cell’s leading and trailing edges. Caveolae are required at the cell rear for RhoA activation, while the front with high density of Cav1 scaffolds, exhibits significantly reduced RhoA activity. Increased membrane tension induces caveolae disassembly, enhancing the proportion of Cav1 scaffolds and RhoA-GTP interaction, which correlates with decreased actomyosin contractility. Similar inhibition of contractility was observed in cavin-KO cells, which lack caveolae and present only Cav1 scaffolds. Our findings imply that Cav1 scaffolds sequester RhoA-GTP in a non-functional state, preventing downstream interaction with actomyosin contractility effectors. At the front, Cav1 scaffolds locally inhibit RhoA, facilitating migration, while global inhibition, induced by high membrane tension or cavin-1 deficiency, halts migration. While proximity labeling studies have identified Cav1 as a potential interactor of active RhoA48, the specific Cav1 domain responsible for this interaction remains unknown. In this study, we provide evidence that the caveolin-1 scaffolding domain (CSD) does not mediate RhoA binding. The CSD has been proposed to bind aromatic-rich caveolin binding motifs present in effector proteins12 but its role remains debated8. Bioinformatic and structural analyses have suggested that the CSD is embedded within the plasma membrane, rendering it largely inaccessible for protein-protein interactions36. Interestingly, recent molecular dynamics simulations indicate that the accessibility of Cav1 binding domains, including the CSD, differs between Cav1 scaffolds and fully formed caveolae49. These findings suggest, in agreement with our data, that Cav1 scaffolds may present a more accessible conformation for interacting with RhoA compared to intact caveolae.
Directional cell migration is initiated by external cues that are converted by cellular sensing machineries into coordinated responses44. We found that prior to directional changes, caveolae and Cav1 scaffolds lose their front-rear polarity and adopt a bipolar distribution, coinciding with a transient arrest in migration. Caveolae subsequently reassemble at the new rear, re-establishing front-rear polarity that is crucial for directional reorientation. During directional signaling, Rho-GEFs are recruited to the plasma membrane, triggering cytoskeletal rearrangements50. Using optogenetics, we revealed a precise spatiotemporal sequence initiated by membrane tension: caveolae formation and recruitment of Rho-GEFs at the plasma membrane, followed by localized RhoA activation at sites of contraction. This cascade drives actomyosin contractility and facilitates the establishment of new front-rear polarity, thereby initiating directional migration. Recent work has shown that, in addition to the well-characterized short-range mutual inhibition between RhoA and Rac1, long-range mutual inhibition also contributes to the spatial polarization of these GTPases during migration51. Our findings demonstrate that caveolae dynamics differentially regulate RhoA and Rac1 activation in response to membrane tension gradients. This raises the intriguing possibility that caveolae-mediated mechanosensing actively contributes to the long-range coordination of RhoA and Rac1 signaling during directional migration.
To substantiate these findings, we developed a physical model that simulates cell edge dynamics, where phases of protrusion and retraction alternate stochastically (Fig. 7a–d). The model demonstrates that a feedback loop between local membrane tension and local contractility, through caveolae disruption and reassembly, enhances the persistence of cell motion (Fig. 7a, b). In the absence of external cues, cell motion is unbiased over long timescales (Fig. 7c). However, the model shows that the feedback between membrane tension and contractility provides short-term persistence, which quantitatively reproduces our experimental observations (Fig. 7d).
In summary, our findings suggest a feedback loop wherein caveolae assembly at the cell rear promotes RhoA-GEF and RhoA activation, contraction, and persistent migration, while Cav1 scaffolds at the cell front prevent RhoA-GEF downstream interactions, which enables protrusions (Fig. 8e). This regulatory process fails in cells lacking Cav1 or in conditions where Cav1 scaffolds predominate, such as high membrane tension or cavin-1 deficiency. Our study identifies caveolae’s front-rear asymmetry and mechanosensing as critical factors for directional reorientation, adding a previously uncharacterized regulatory layer to cell migration (Fig. 8f).
The hypothesis that caveolar Cav1 and non-caveolar Cav1 serve distinct cellular functions has been long proposed8,52. Using super-resolution microscopy and machine learning analysis, we identified Cav1 scaffolds as distinct from bona fide caveolae, each exerting differential regulation in cell migration. A positive feedback loop between caveolae and RhoA-GEFs could involve clustering, as lipid nanoclusters at the plasma membrane are known to enhance GEF recruitment53. It was recently found that distinct levels of ARHGEF17 clustering was correlated with the differential activation of RhoA and myosin filaments assembly and disassembly cycles at the front and back of the protrusive lamella in migrating fibroblasts54. Clustering-based RhoA-GEF by caveolae aligns with caveolae’s role in nanoscale lipid organization55.
Recent findings propose that migrating cells retain a “mechanical memory” of past morphological states, through actin cortex adjustments, facilitating rapid adaptation to mechanical environments56. Our observation that caveolae consistently reassemble at the rear after mechanical stress suggest that caveolae might contribute to this memory, aiding breast cancer cell navigation through varied mechanical settings encountered during tissue invasion. Mechanical forces are increasingly recognized as critical factors that influence tumor progression57, with caveolae mechanics emerging as an important factor in this process58,59. This property of caveolae may contribute to the mechanical regulation of cancer cell invasion, as recently illustrated by the interplay between caveolae mechanosensing and invadopodia formation17. Our findings are particularly relevant to tumor progression and metastasis, both of which are characterized by variations in Cav1 expression levels60, potentially affecting the ratio of caveolae to Cav1 scaffolds60.
Methods
Cell culture
Human triple negative breast cancer Hs578t cells, genome-tagged with Cav1-RFP including WT-Hs578t, Cav1-KO Hs578t and cavin1-KO Hs578t variants were cultured in Dulbecco’s modified Eagle’s medium + GlutaMAX (DMEM) (Gibco # 10566016), supplemented with 10% fetal calf serum, 5 mM pyruvate, 1% penicillin-streptomycin. Cells were maintained at 37 °C under 5% CO2. MDA-MB-231, genome tagged with Cav1-RFP, were grown in DMEM/F-12 (Gibco #11320033) supplemented with 10% fetal calf serum and maintained under identical conditions. Coverslips and FluoroDish were pre-coated with collagen before cell seeding.
Plasmids and antibodies
Plasmids
Cav1-RFP, cavin-1-eGFP, Cav1-Dendra2, and Cav1-HaloTag plasmids have been published18,19. WT-Cav1-myc-mRFP and Cav1-F92A-V94A-myc-mRFP (Cav1-CSD mutant) were a gift from Robert Nabi (University of British Columbia, Canada). HaloTag-3xrGBD (Addgene #176108) (RhoA biosensor), ARHGEF25 plasmid for optogenetics, pLV-SspB-HaloTag-p63DH (Addgene #176129), Lck-mTurquoise2-iLID (Addgene #176125), Tiam-1 GEF and pLV-SspB-HaloTag-DHPH-TIAM1 plasmids for optogenetics (Addgene#176114) were gifts from Dr. Dorus Gadella (University of Amsterdam, Netherlands). ARHGEF11 plasmid for optogenetics pll7-IRFP-SSPB-P2A-Venus-iLID-CAAX (control), pll7-ARGEF11-iRFP-sSspb-P2A-mVenus-iLID-CAAX and ARHGEF11 (DHPH)-CRY2-mCherry were gifts from Dr. Mathieu Coppey (Institut Curie, France). Opto-PI3K, pHR ISH2-EGFP-SspBMicro and iLID-BFP-CAAX plasmids for optogenetics were gifts from Dr. Orion D. Weiner, University of California, San Francisco.
Primary antibodies
Rabbit anti-Cav1 (Cell signaling 3238S; 1:2000 for WB, 1:500 IF), rabbit anti-cavin1 (abcam, ab48824; 1:1000 for WB), rabbit anti-cavin1 (Proteintech18892-1-AP; 1:1000 for WB), mouse anti-RhoA-GTP (New East Biosciences, 26904, 1:500 for WB and IF), rabbit anti-total RhoA (Invitrogen, PA599537, 1:1000 for WB), rabbit anti-HaloTag (Recombinant Antibody Platform, Institut Curie, A-P-R#57; 1:1000 for WB), mouse anti-RhoA antibody from RhoA-Pull down kit (Cytoskeleton, #ARH05, 1:500 for WB), mouse anti-α-tubulin (clone B512; Sigma, cat. no. T5168; 1:5000 for WB), mouse anti-actin B (Sigma A5316; 1:1000 for WB). For STORM acquisition rabbit anti-Cav1 and mouse anti-RhoA-GTP antibodies were used at 1:500 dilution for 48 hours at 4 °C.
Secondary antibodies
Mouse-Alexa 488 (Invitrogen, A21202), rabbit-Cy3 (Jackson ImmunoResearch, 111166045), were used at 1:200 for IF; mouse-HRP (Jackson ImmunoResearch, 715-035-151) and rabbit-HRP (Jackson ImmunoResearch, 711035152) were used at 1:5000 for WB; goat anti-rabbit (12005870 and 12004162) or anti-mouse (12005867 and 12004159) IgG StarBright Blue 520 or 700 (Bio-Rad) were used at 1:5000 for WB; For STORM acquisition, goat anti-mouse CF568 (Sigma SAB4600312, 1:200 dilution), donkey anti-rabbit Alexa647 (Jackson ImmunoResearch, 711-606-152, 1:200 dilution) for 24 hours at 4 °C.
siRNA and plasmid transfection
siRNA knock-down (KD) of caveolin-1 and cavin1 was performed using HiPerFect (Qiagen #301704) following the manufacturer’s protocol, with cells cultured for 48 hours post-transfection. All siRNA sequences were used at the final concentration of 20 nM. Control siRNA (SI03650325, 5′-AAUUCUCCGAACGUGUCACGU-3′), siPTRF/cavin1 (Hs_PTRF_6, SI04178496) were purchased from Qiagen. For siCav1, four oligomers were used (Eurogentec; 5′-CUAAACACCUCAACGAUGA-3′, 5′-GCAUCAACUUGCAGAAAGA-3′, 5′-GCAAAUACGUAGACUCGGA-3′, and 5′-GCAGUUGUACCAUGCAUUA-3′). Knockdown efficiency was validated by western blotting using specific antibodies, with tubulin as a loading control. Plasmids were transiently transfected using Lipofectamine LTX-Plus Reagent (Invitrogen #15338100) according to the manufacturer’s instructions. For one Fluorodish (35 mm diameter), 2 µg DNA was transfected using 2 µl LTX and PLUS reagent in a 1:1 ratio, and cells were cultured overnight prior to imaging. Cells expressing Halo-tag plasmids were stained with Janelia Fluor Halo Tag Ligand (#GA1120), at a final concentration of 100 nM for 1 hour at 37 °C. Cells were washed three times with 1X PBS, fresh DMEM supplemented with 10% FBS was added, and cells imaged.
Collagen networks and cell migration assay
For 3D collagen gel migration assays, 50 µl droplets of collagen I (Corning #354236) at a final concentration of 2.2 mg/ml were seeded with 20,000 genome-edited Cav1-RFP Hs578t (WT or KD) cells and polymerized on glass-bottom dishes at room temperature (RT) for 30 minutes. The polymerized gels were immersed in pre-warmed complete medium and incubated overnight. Cells were imaged the next day using a spinning disk confocal microscope at 20-minute intervals over a 4-hour period. The collagen droplets were prepared with a 10:1 ratio of unlabeled to Alexa488-labeled collagen. A Z-stack of 9 µm thickness, with 1 µm intervals, was acquired every 2 or 20 minutes, depending on the experiment. Manual tracking of cell migration was performed using the ImageJ Fiji plugins Manual Tracking. Migration parameters were quantified using “Chemotaxis and Migration Tool” (Ibidi), available as a plugin for ImageJ/Fiji (https://ibidi.com/chemotaxis-analysis/171-chemotaxis-and-migration-tool.html; At least 15 cells per condition, across three independent experiments, were analyzed. For preparing thin collagen fiber layers, 35 mm glass-bottom FluoroDishes (World Precision Instruments #FD3510) were coated with 50 μg/ml collagen I (Corning #354236) in 0.01% acetic acid for 1 hour at 37 °C. The dishes were then washed twice with PBS and immersed in complete DMEM medium prior to cell seeding. Imaging was performed at acquisition intervals of 1-2 minutes over a duration of 2-6 hours, depending on the experimental conditions.
Direction changes and Cav1 intensity correlation analysis
Cell segmentation in time-lapse movies was performed using SAM2 (Segment Anything Model v2)61; with the pre-trained “Hiera large” model, run via a Jupyter Notebook. For most movies, a single prompt point place at the center of the cell in the first frame was sufficient to achieve accurate segmentation. When necessary, up to four prompt points were added (always in the first frame) to improve segmentation quality. The resulting masks were exported and converted into Fiji ROIs. A custom Fiji macro62 was developed to measure Cav1 intensity within two fixed square ROIs positioned on the left and right sides of the cell. These ROIs were drawn in the first frame and propagated across all subsequent time points. Manual adjustment of the ROIs was performed as needed if significant cell displacement occurred during imaging. The directional movement of the cell along the X-axis was then compared to the relative Cav1 intensity between the left and right ROIs.
Flipper-TR measurement
Cells were incubated with the fluorescent membrane tension probe Flipper-TR (Spirochrome, cat. no. #CY-SC020) at a final concentration of 500 nM for 15 minutes at 37 °C in a 5% CO₂ atmosphere prior to imaging. Fluorescence lifetime imaging microscopy (FLIM) was performed on a Leica SP8 confocal microscope equipped with a 63x oil-immersion objective (NA 1.4). Flipper-TR was excited using a white light laser set at 488 nm, with power setting ranging from 10% to 25%, and a repetition rate of 40 MHz (pulse picker mode). Fluorescence emission was collected using a HyD SMD detector with a bandwidth set from 550 nm to 650 nm. Images were acquired at a scan speed of 100-200 Hz, and each frame was averaged over 10 repetitions to increase photon yield.
For hypotonic shock experiments, cells were first imaged under isotonic conditions. The imaging medium was then diluted with water to 80% (v/v) final concentration to induce hypotonic stress, and imaging was continued for up to 6 minutes following the medium change.
Fluorescence decay curves were obtained by pooling photon counts from manually defined ROIs at the front and rear of individual cells. Decay curves were fit using a biexponential model, and fluorescence lifetimes were extracted from the tail region of each curve. Following the methodology described in ref. 26, the longer lifetime component was used as a proxy for Flipper-TR lifetime, while the shorter component was excluded from further analysis.
Osmotic shocks
Isotonic conditions were maintained in DMEM supplemented with 10% FBS (300 mOsm) for the specified duration. A 90% hypo-osmotic shock was induced by diluting complete DMEM with deionized water (1:9 dilution, resulting in ~30 mOsm) for 5–10 minutes. Recovery conditions involved reintroducing DMEM supplemented with 10 % FBS (300 mOsm). Hyper-osmotic shock was induced by adding 600 mM sorbitol to DMEM supplemented with 10% FBS, achieving a final osmolarity of ~850 mOsm.
Spinning-disc confocal microscopy
Time-lapse images were acquired using an inverted Nikon Eclipse Ti-E microscope equipped with a Yokogawa CSU-X1 spinning disk confocal unit, integrated into Metamorph software (Gataca Systems). The system featured an sCMOS camera with 95% quantum efficiency and a pixel size of 11 µm. Appropriate emission filters were used for each fluorophore: DAPI (440 nm), GFP (ET525/50), mCherry (605/64) and Cy5 (ET708/75). Imaging was performed on a heated stage (37 °C) with controlled 5% CO2, using a 60X CFI Plan Apo oil objective.
Cell lysates preparation and western blotting
To validate knockdown (KD) efficiency and knockout (KO) cell lines, cells were lysed in a sample buffer containing 2% SDS, 10% glycerol, 4 mM DTT, and Tris (pH 6.8) with phenol Red. Lysates were loaded onto 4–20% Mini-PROTEAN TGX gels (BioRad) and separated by SDS-PAGE. Proteins were transferred to nitrocellulose membranes using the Trans-Blot SD Semi-Dry Transfer Cell (Bio-Rad #1703940). Membranes were blocked with 5% BSA for 1 hour at RT. Primary antibodies were diluted in 0.5% BSA, and the membranes were incubated overnight at 4 °C. The following day, membranes were washed three times with 1X TBS-Tween and incubated with secondary antibodies, either HRP-conjugated or fluorescent StarBright (BioRad). Chemiluminescence signals were detected using SuperSignal West Dura Extended Duration or SuperSignal West Femto substrate (Thermo Scientific Life Technologies). Imaging was performed on a ChemiDoc MP Imaging System (BioRad #12003154), and protein band quantification was conducted using ImageJ.
Indirect immunofluorescence assays
Cells were fixed with 4% paraformaldehyde for 15 minutes at RT, washed with PBS, and permeabilized with 0.1% saponin in PBS for 15 minutes. Blocking was performed with 5% BSA for 30 minutes at RT. Primary antibodies, Rabbit anti-Cav1 (BD Transduction Laboratories, cat. no. 610059; 1:200) and mouse anti-RhoA-GTP (New East Biosciences, cat.no. 26904, 1:200), were diluted in 0.5% BSA, and cells were incubated for 1 hour at RT. Appropriate secondary antibodies, along with Phalloidin-Alexa 647 (1:500, Invitrogen), were added for 1 hour at RT. Cells were mounted with Fluoromount-G containing DAPI (Invitrogen, cat.no. 00-4958-02) and imaged using a spinning disc confocal microscope.
Co-immunoprecipitation (Co-IP) assays
Caveolin-1 antibody Co-IP assay
1 × 106 cells (WT or KO) were seeded in a 100 mm dish and cultured overnight. The next day, cells were washed with cold PBS and lysed by scraping in 600 µl lysis buffer (25 mM Tris pH 7.4, 150 mM NaCl, 10 mM EDTA, 5% glycerol, 1% NP40, and 1X protease inhibitor cocktail (PIC, Roche)). Lysates were centrifuged at 10,600 g for 15 minutes to remove cell debris. A 100 µl aliquot of the lysate was mixed with 4 mM DTT and 50 µl sample buffer and labeled as the input sample. The remaining 500 µl lysate was pre-cleared with 25 µl protein A/G magnetic beads (Thermo Scientific, cat. no. 88802) for 1 hour at 4 °C with rotation. After pre-clearing, the lysate was incubated overnight at 4 °C with 1 µg/ml rabbit anti-Cav1 antibody (BD Transduction Laboratories, cat. no. 610059) with rotation. The lysate was further incubated with 25 µl protein A/G magnetic beads (Thermo Scientific) for 1 hour at 4 °C. Immunoprecipitated beads were washed three times with lysis buffer (without PIC) and eluted in 60 µl elution buffer according to the manufacturer’s instructions. Input (20 µl) and immunoprecipitated (IP) samples (20 µl) were analyzed by SDS-PAGE gel followed by immunoblotting.
RhoA-GTP pulldown assays
The RhoA pulldown assay was preformed using the RhoA Activation Assay Biochem Kit (Cytoskeleton, cat. no. BK036) according to the manufacturer’s instructions. Briefly, 2 × 106 cells were harvested in RhoA lysis buffer. Lysates were cleared by centrifugation at 10,600 g for 2 minutes at 4 °C. Supernatants were incubated with 50 µg Rhotekin-RBD protein beads per sample for 1 hour at 4 °C with rotation. Pulldown beads were washed, eluted, and analyzed by SDS-PAGE and immunoblotting. The ratio of active RhoA (RhoA-GTP) to total RhoA was quantified and compared between cell lines.
Proximity ligation assays (PLA)
The in situ interaction between endogenous Cav1 and RhoA-GTP was analysed using the NaveniFlex Cell MR kit (Ozyme, #NC.MR.100) according to the manufacturer’s instructions. Cells were fixed with 4% paraformaldehyde (PFA) for 10 minutes at RT, quenched with 50 mM NH4Cl for 10 minutes, and permeabilized with 0.1% Triton X-100 for 10 minutes at RT. Non-specific binding was blocked using the NaveniFlex Blocking buffer for 1 hour at 37 °C. Cells were then incubated with the indicated primary antibodies for 1hour at 370 °C. Subsequent steps, including incubation with DNA probes, ligation, amplification, and detection, were carried out using the NaveniFlex Cell MR kit. A single-antibody condition was used as a negative control for the PLA reaction. After washing, cells were mounted in Fluoromount with DAPI to stain nuclei. PLA signals, appearing as fluorescent dots, were imaged using a Nikon AX R confocal microscope equipped with a 40X oil immersion objective. Z-sections were acquired and processed for deconvolution. Cells boundaries were delineated to generate masks, and mean fluorescence intensities were quantified using ImageJ. Data are presented as the mean PLA intensity per cell surface area ± the median with a 95% confidence interval, based on three independent experiments.
dSTORM Imaging
Stochastic optical reconstruction microscopy (STORM) imaging was performed using an inverted microscope (Nikon TiE2) equipped with SAFe360 technology (Abbelight), a high-numerical aperture (NA) oil-immersion objective (100X, NA 1.5), and an sCMOS camera (Hamamatsu Fusion BT C15440-20UP). Total internal reflection fluorescence (TIRF) mode was used during the experiment. Cells were grown on 170 μm coverslips (Menzel-Gläser 18 mm, diameter #1.5) and mounted on glass slides with a 15 mm central hole filled with imaging buffer (Tris 50 mM, NaCl 10 mM, 10% glucose, 100 mM MEA (Sigma 30070), 70 U/mL glucose oxidase (Sigma G2133-10KU), 10 mg/mL catalase (Sigma C40-100MG) and sealed with Picodent Twinsil63. Samples were illuminated with a 637 nm laser (500 mW) and a 405 nm laser (100 mW) for fluorophore reactivation. Cells grown on line micropatterns were selected using widefield microscopy and imaged in dSTORM mode for 35,000 frames with 637 nm laser (21 ms per frame) over a field of view of 20 × 20 µm). Elongated cells were selected to minimize fluorophore bleaching and to ensure that both extremities were imaged simultaneously. Imaging both the front and rear of each cell allowed for statistical paired comparison between these regions. A total of 28 cells across four experimental conditions, were analyzed, distributed over three independent replicates: 8 cells in isotonic conditions, 6 cells in hypotonic conditions, 7 cells in recovery conditions for WT-Hs578t, and 7 cells for cavin1-KO Hs578t. This resulted in 52 distinct images, representing the front and rear regions of individual cells. Drift correction was performed using carboxylated nanodiamonds (ADÀMAS nano, 140 nm, Lower Brightness, -COOH NDNV140nmMd10ml) added to the sample (10 μL/mL). Post-processing was performed with AbbelightTM NEO software. Nanodiamonds were isolated in the images using the crop tool in NEO software, and local drift corrections were applied. The resulting drift correction was then propagated to the entire image.
dSTORM analysis, cluster segmentation, S1A, S1B, S2 and caveolae classification
Cav1 proteins were labeled using a rabbit anti-Cav1 antibody (Cell signaling, 3238S), followed by a donkey anti-rabbit Alexa647 secondary antibody (Jackson ImmunoResearch, 711-606-152). The localizations of Cav1 were processed, and the resulting clusters were classified using the 3D SMLM (single-molecule localization microscopy) network analysis pipeline11. For analysis, all Z-coordinates were assumed to be in the same plan (Z = 0). The SuperResNet platform was employed with the following parameters: Merging threshold: 12 nm (to correct for multiple blinking of single fluorophores); Proximity threshold: 80 nm (for network construction); Alpha: 1 (for noise filtering); Bandwidth: 120 nm (to segment localizations into blobs/clusters using the mean shift algorithm). A total of 28 features/descriptors, including shape, topology, network properties, size, and hollowness, were extracted for each segmented blob/cluster. The clusters or blobs feature from all conditions were grouped into four categories using the x-means clustering algorithm. Biological identities for these groups i.e., caveolae and S2, S1A, and S1B scaffolds, were assigned by comparing the group centers with previously characterized Cav1 groupings11. Assignments were based on the best match (highest similarity) to these prior classifications.
Cellpose and front-rear intensity measurements
Accurate cell segmentation was performed using the Cellpose algorithm64, a robust deep learning tool for cell detection. To improve segmentation precision, we trained a custom model based on the pre-trained cyto2 model, using manually annotated data from approximately 30 cells. Following segmentation, a Fiji (ImageJ) macro62 was used to compute the mean fluorescence intensity along the periphery of each cell, within a 2 µm-wide band inside the cell boundary. The macro separately measured the intensity in the front and rear regions of interest (ROIs), with each ROI defined as a 50-pixel area. ROIs were automatically determined by the macro, where the rear was defined as the leftmost part of the cell and the front as the rightmost part. For cells oriented differently or in cases where the automatic ROI placement was suboptimal, the macro allowed users to manually adjust the ROI positions. This flexibility ensured accurate intensity measurements for all analyzed cells.
Morphodynamic analysis
To perform cell segmentation during morphodynamic analysis, we utilized the Cellpose framework64, retrained using data from approximately 30 manually annotated cells based on the pre-trained cyto2 model. Accurate segmentation was essential, as this analysis relies on precise delineation of cell contours across all frames. Morphodynamic maps were generated following the methodology described in ref. 39 and adapted from ref. 40 to detect and quantify cell contractions and protrusions. The morphodynamic mapping algorithm, implemented in MATLAB, characterizes edge dynamics by measuring the displacement of cell contours between consecutive frames. Contours in each frame were represented by 100 equidistant markers, which were aligned across frames by applying a circular shift to minimize quadratic distances. Displacements between corresponding markers (representing consecutive segments on the same contour) were computed and visualized as color-coded maps. The X-axis of these maps represents time points, while the Y-axis represents the contour segment displacement. Color-coding was used to distinguish contractions (negative displacement, blue) from protrusions (positive displacement, red), using the Physics lookup table for visualization. These maps were exported as text files and subsequently analyzed using a Fiji macro62. The macro displayed the map, allowing users to identify contraction events (blue regions), by clicking on them using the Point tool. Selected points provided coordinates (X, Y), corresponding to the frame (X) and the specific contour segment (Y). The identified event was visualized as an 80 ×80-pixel square, which could be manually reviewed and adjusted for spatial and temporal accuracy. Intensity measurements within the defined square and across the entire cell were extracted for 10 frames preceding and 15 frames following each event, encompassing all imaging channels.
Optogenetic activation assays and analysis
Optogenetic activation experiments were performed on cells seeded in 35 mm glass-bottom FluoroDishes. Cells were transiently transfected with the specified marker and, when required, labeled with Halo-Tag fluorescent dye prior to imaging. Imaging was performed using an inverted spinning disk CSU-X1 (Yokogawa) microscope equipped with a FRAP/photoactivation module, integrated into Metamorph software (Gataca Systems), and paired with an sCMOS camera (95% quantum efficiency). ptogenetic activation was achieved using the iLas2 laser system, with targeted illumination controlled via Metamorph software. A targeted 80×80-pixel region was positioned at the leading edge of the cell using a 60X objective. Activation was performed using 4.5% power from a 491-nm laser, pulsed every 30 seconds within the defined opto-ROI. Pre-activation images were acquired for 2 minutes for each cell to establish baseline intensity values for subsequent quantification. For data analysis, opto-ROI information from Metamorph was imported into Fiji using the Metamorph.nd & ROI files importer plugin developed by F. Cordelières (available at https://imagejdocu.list.lu/plugin/inputoutput/nd_stacks_builder/start). A custom macro was used to calculate the mean fluorescence intensity within the opto-ROI and a random-ROI, which was symmetrically positioned relative to the centroid of the cell. Mean intensity values were measured at all time points for all fluorescent channels. The intensity measurements for ARHGEF25, Caveolin-1, and contraction-related ROI areas were plotted as functions of time and compared across experimental conditions.
Statistics and reproducibility
The number of cells analyzed and the number of independent experiments are detailed in the corresponding figure legends. For each experimental condition, at least three independent experiments were performed, with measurements obtained from distinct cells in each experiment. Statistical parameters and significance levels are presented within the figures and their legends. Data were considered statistically significant when p < 0.05. Statistical tests employed include Wilcoxon’s rank-sum test, Mann-Whitney test, one-way ANOVA, and paired t-test, as appropriate, and were performed using GraphPad Prism 8.0 software.
Data availability
The main data supporting the findings of this study are available within the article and its Supplementary Information. The relevant raw data from each figure (in the main manuscript and in the Supplementary Information) are represented by Excel file labeled “Source Data Fig.X and Source Data uncropped blots,” provided with this article. All other data supporting the findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
Code availability
All codes used for this study are available from the corresponding author on request.
References
Trepat, X., Chen, Z. & Jacobson, K. Cell migration. Compr. Physiol. 2, 2369–2392 (2012).
Merino-Casallo, F., Gomez-Benito, M. J., Hervas-Raluy, S. & Garcia-Aznar, J. M. Unravelling cell migration: defining movement from the cell surface. Cell Adh Migr. 16, 25 (2022).
Yu, H., Mouw, J. K. & Weaver, V. M. Forcing form and function: biomechanical regulation of tumor evolution. Trends Cell Biol. 21, 47 (2011).
Sitarska, E. & Diz-Muñoz, A. Pay attention to membrane tension: mechanobiology of the cell surface. Curr. Opin. Cell Biol. 66, 11 (2020).
Sens, P. & Plastino, J. Membrane tension and cytoskeleton organization in cell motility. J. Phys. Condens. Matter 27, 273103 (2015).
Zmurchok, C., Collette, J., Rajagopal, V. & Holmes, W. R. Membrane tension can enhance adaptation to maintain polarity of migrating cells. Biophys. J. 119, 1617–1629 (2020).
Lamaze, C., Tardif, N., Dewulf, M., Vassilopoulos, S. & Blouin, C. M. The caveolae dress code: structure and signaling. Curr. Opin. Cell Biol. 47, 117–125 (2017).
Parton, R. G. et al. Caveolae: the FAQs. Traffic 21, 181–185 (2020).
Porta, J. C. et al. Molecular architecture of the human caveolin-1 complex. Sci. Adv. 8, eabn7232 (2022).
Epand, R. M., Sayer, B. G. & Epand, R. F. Caveolin scaffolding region and cholesterol-rich domains in membranes. J. Mol. Biol. 345, 339–350 (2005).
Khater, I. M., Meng, F., Wong, T. H., Nabi, I. R. & Hamarneh, G. Super resolution network analysis defines the molecular architecture of caveolae and caveolin-1 scaffolds. Sci Rep 8, 9009 (2018).
Lim, J. E., Bernatchez, P. & Nabi, I. R. Scaffolds and the scaffolding domain: an alternative paradigm for caveolin-1 signaling. Biochem. Soc. Trans. 52, 947 (2024).
Pilch, P. F. & Liu, L. Fat caves: caveolae, lipid trafficking and lipid metabolism in adipocytes. Trends Endocrinol. Metab. 22, 318 (2011).
Parton, R. G. & Del Pozo, M. A. Caveolae as plasma membrane sensors, protectors and organizers. Nat. Rev. Mol. Cell Biol. 14, 98–112 (2013).
Grande-García, A. et al. Caveolin-1 regulates cell polarization and directional migration through Src kinase and Rho GTPases. J. Cell Biol. 177, 683 (2007).
Joshi, B. et al. Phosphorylated caveolin-1 regulates Rho/ROCK-dependent focal adhesion dynamics and tumor cell migration and invasion. Cancer Res 68, 8210–8220 (2008).
Monteiro, P. et al. A mechanosensitive caveolae-invadosome interplay drives matrix remodelling for cancer cell invasion. Nat. Cell Biol. 25, 1787–1803 (2023).
Sinha, B. et al. Cells respond to mechanical stress by rapid disassembly of caveolae. Cell 144, 402–413 (2011).
Torrino, S. et al. EHD2 is a mechanotransducer connecting caveolae dynamics with gene transcription. J. Cell Biol. 217, 4092–4105 (2018).
Dewulf, M. et al. Dystrophy-associated caveolin-3 mutations reveal that caveolae couple IL6/STAT3 signaling with mechanosensing in human muscle cells. Nat Commun 10, 1974 (2019).
Beardsley, A. et al. Loss of caveolin-1 polarity impedes endothelial cell polarization and directional movement. J. Biol. Chem. 280, 3541–3547 (2005).
Hetmanski, J. H. R. et al. Membrane tension orchestrates rear retraction in matrix-directed cell migration. Dev. Cell 51, 460 (2019).
Lentini, D. et al. Polarization of caveolins and caveolae during migration of immortalized neurons. J. Neurochem 104, 514–523 (2008).
Parat, M. O., Anand-Apte, B. & Fox, P. L. Differential caveolin-1 polarization in endothelial cells during migration in two and three dimensions. Mol. Biol. Cell 14, 3156 (2003).
Golani, G., Ariotti, N., Parton, R. G. & Kozlov, M. M. Membrane curvature and tension control the formation and collapse of caveolar superstructures. Dev. Cell 48, 523–538.e4 (2019).
Colom, A. et al. A fluorescent membrane tension probe. Nat. Chem. 10, 1118–1125 (2018).
Shi, Z., Graber, Z. T., Baumgart, T., Stone, H. A. & Cohen, A. E. Cell membranes resist flow. Cell 175, 1769–1779.e13 (2018).
De Belly, H. et al. Cell protrusions and contractions generate long-range membrane tension propagation. Cell 186, 3049–3061.e15 (2023).
García-Arcos, J. M. et al. Actin dynamics sustains spatial gradients of membrane tension in adherent cells. bioRxiv https://doi.org/10.1101/2024.07.15.603517 (2024).
Jaffe, A. B. & Hall, A. Rho GTPases: biochemistry and biology. Annu Rev. Cell Dev. Biol. 21, 247–269 (2005).
Mosaddeghzadeh, N. & Ahmadian, M. R. The RHO Family GTPases: mechanisms of regulation and signaling. Cells 10, 1831 (2021).
Boettcher, J. P. et al. Tyrosine-phosphorylated caveolin-1 blocks bacterial uptake by inducing Vav2-RhoA-Mediated cytoskeletal rearrangements. PLoS Biol. 8, 55–56 (2010).
Nakanishi, N. et al. MURC deficiency in smooth muscle attenuates pulmonary hypertension. Nat. Commun. 7, 12417 (2016).
Mahlandt, E. K. et al. Visualizing endogenous Rho activity with an improved localization-based, genetically encoded biosensor. J. Cell Sci. 134, jcs258823 (2021).
Seze, J. De, Bongaerts, M., Boulevard, B. & Coppey, M. Optogenetic control of a GEF of RhoA uncovers a signaling switch from retraction to protrusion. Elife 12, RP93180 (2023).
Byrne, D. P., Dart, C. & Rigden, D. J. Evaluating caveolin interactions: do proteins interact with the caveolin scaffolding domain through a widespread aromatic residue-rich motif?. PLoS One 7, e44879 (2012).
Ridley, A. J. Rho GTPases and cell migration. J. Cell Sci. 114, 2713–2722 (2001).
Ridley, A. J. et al. Cell migration: integrating signals from front to back. Science 302, 1704–1709 (2003).
MacHacek, M. et al. Coordination of Rho GTPase activities during cell protrusion. Nature 461, 99–103 (2009).
Vaidžiulytė, K. et al. Persistent cell migration emerges from a coupling between protrusion dynamics and polarized trafficking. Elife 11, e69229 (2022).
Lawson, C. D. & Ridley, A. J. Rho GTPase signaling complexes in cell migration and invasion. J. Cell Biol. 217, 447 (2018).
Mahlandt, E. K. et al. Opto-RhoGEFs, an optimized optogenetic toolbox to reversibly control Rho GTPase activity on a global to subcellular scale, enabling precise control over vascular endothelial barrier strength. Elife 12, RP84364 (2023).
Charras, G. & Sahai, E. Physical influences of the extracellular environment on cell migration. Nat. Rev. Mol. Cell Biol. 15, 813–824 (2014).
SenGupta, S., Parent, C. A. & Bear, J. E. The principles of directed cell migration. Nat. Rev. Mol. Cell Biol. 2021 22, 529–547 (2021).
Martin, E., Girardello, R., Dittmar, G. & Ludwig, A. Time-resolved proximity proteomics uncovers a membrane tension-sensitive caveolin-1 interactome at the rear of migrating cells. Elife 13, e85601 (2024).
McMahon, K. A. et al. Identification of intracellular cavin target proteins reveals cavin-PP1alpha interactions regulate apoptosis. Nat. Commun. 2019 10, 1–17 (2019).
Pertz, O. Spatio-temporal Rho GTPase signaling—where are we now?. J. Cell Sci. 123, 1841–1850 (2010).
Bagci, H. et al. Mapping the proximity interaction network of the Rho-family GTPases reveals signalling pathways and regulatory mechanisms. Nat. Cell Biol. 22, 120–134 (2020).
Doktorova, M. et al. Caveolin assemblies displace one bilayer leaflet to organize and bend membranes. bioRxiv https://doi.org/10.1101/2024.08.28.610209 (2025).
Petrie, R. J., Doyle, A. D. & Yamada, K. M. Random versus directionally persistent cell migration. Nat. Rev. Mol. Cell Biol. 10, 538–549 (2009).
De Belly, H. et al. Long range mutual activation establishes Rho and Rac polarity during cell migration. bioRxiv https://doi.org/10.1101/2024.10.01.616161 (2025).
Pol, A., Morales-Paytuví, F., Bosch, M. & Parton, R. G. Non-caveolar caveolins—duties outside the caves. J. Cell Sci. 133, jcs241562 (2020).
Nawrotek, A., Dubois, P., Zeghouf, M. & Cherfils, J. Molecular principles of bidirectional signalling between membranes and small GTPases. FEBS Lett. 597, 778–793 (2023).
Rao, V. et al. Feedback regulation by the RhoA-specific GEF ARHGEF17 regulates actomyosin network disassembly. bioRxiv 2024 08, 610052 (2024).
Ocket, E. & Matthaeus, C. Insights in caveolae protein structure arrangements and their local lipid environment. Biol. Chem. https://doi.org/10.1515/HSZ-2024-0046 (2024).
Kalukula, Y., Luciano, M., Charras, G., Brückner, D. B. & Gabriele, S. The actin cortex acts as a mechanical memory of morphology in confined migrating cells. bioRxiv 2024 08, 606589 (2024).
Hayward, M. K., Muncie, J. M. & Weaver, V. M. Tissue mechanics in stem cell fate, development, and cancer. Dev. Cell 56, 1833–1847 (2021).
Singh, V. & Lamaze, C. Membrane tension buffering by caveolae: a role in cancer?. Cancer Metastasis Rev. 39, 505–517 (2020).
Lamaze, C. & Torrino, S. Caveolae and cancer: a new mechanical perspective. Biomed. J. 38, 367 (2015).
Campos, A. et al. Cell intrinsic and extrinsic mechanisms of caveolin-1-enhanced metastasis. Biomolecules 9, 314 (2019).
Ravi, N. et al. SAM 2: Segment anything in images and videos. ArXiv (2024).
Schindelin, J. et al. Fiji - an Open Source platform for biological image analysis. Nat. Methods 9, 676–682 (2012).
Collot, M. et al. MemBright: a family of fluorescent membrane probes for advanced cellular imaging and neuroscience. Cell Chem. Biol. 26, 600–614.e7 (2019).
Pachitariu, M. & Stringer, C. Cellpose 2.0: how to train your own model. Nat. Methods 19, 1634–1641 (2022).
Acknowledgements
The core facilities of Institut Curie, the scientific and technical assistance from staff of the Cell and Tissue Imaging (PICT-IBiSA) and the Nikon Imaging Center at Institut Curie, member of the French National Research Infractucture France-BioImaging (ANR10-INBS-04) are acknowledged. We thank Yahongyang Li and Robert Nabi (UBC, Vancouver, Canada) for discussion with control STORM experiments analysis; Mathieu Coppey (Institut Curie, France) for help in setting up optogenetics, and Jacqueline Cherfils (Ecole Normale Supérieure, Paris-Saclay) for insightful discussions about Rho-GTPases. NeurImag is part of GIS IBISA and the national infrastructure France BioImaging supported by the French National Research Agency (ANR-10-INBS-04). We thank the Leducq Foundation for funding the LEICA SP8 confocal system which was used to carry out FLIM experiments. This work was supported by institutional grants from the Curie Institute, Institut National de la Santé et de la Recherche Médicale (INSERM), Centre National de la Recherche Scientifique (CNRS), and by specific grants from Agence Nationale de la Recherche (ANR-17-CE13-0020-02 and ANR-19-CE15-0020-02) and Institut National du Cancer (Inca 2021-094 and 2021-176) to C.L.; P.S. is co-funded by the European Union (ERC, PushingCell, project 101071793). V.S. was funded by postdoctoral fellowships from Fondation pour la Recherche Médicale (#SPF201909009115) and LABEX CellnScale.
Author information
Authors and Affiliations
Contributions
Conceptualization: C.L. Experimental design: V.S.; C.M.B.; C.L.; P.S. Investigation and data analysis: V.S.; V.B.; A.S.V.; C.M.B.; C.V.L.; P.B. Tools and resources: A.S.M. Funding acquisition: C.L.; P.S. Writing the original draft: V.S.; C.L.; P.S. Supervision: C.L.; P.S. All the authors approved the final version of the manuscript and agree on the content and conclusions.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Congying Wu, and the other anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Singh, V., Breton, V., Viaris de Lesegno, C. et al. Spatiotemporal coupling of caveolae mechanosensing and RhoA-GEFs regulates cell polarity and directional migration. Nat Commun 17, 398 (2026). https://doi.org/10.1038/s41467-025-67090-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67090-z