Abstract
Proteasome activator 28 subunit gamma (PA28γ) attracts considerable attention for its ability to regulate multiple molecules involved in tumor progression. However, its intrinsic activation and functional regulatory mechanisms in the regulation of tumor growth remain incompletely understood. Here, using head and neck squamous cell carcinoma (HNSCC) as a model, we identify phosphorylation at the T23 site of PA28γ, a modification that is highly expressed across pan-cancer types and associated with poor prognosis. Notably, phosphorylation-deficient PA28γ-T23 mutant mice exhibit resistance to carcinogen-induced HNSCC tumors, whereas the phosphomimetic PA28γ-T23 mutant promotes tumor formation and progression. We subsequently explore the underlying mechanism and find that casein kinase 2 (CK2) mediates PA28γ-T23 phosphorylation, which regulates the abundance of growth-related E4F transcription factor 1 (E4F1) through the PA28γ-proteasome pathway. These findings highlight PA28γ-T23 phosphorylation as a crucial pro-oncogenic signal and an unfavorable prognostic indicator across various cancers and underscore its potential as a target for early cancer intervention and treatment.
Similar content being viewed by others

Introduction
Proteasome activator 28 subunit gamma (PA28γ, also known as REGγ, PSME3, 11Sγ, or Ki antigen) was initially identified as a nuclear protein targeted by autoantibodies in the serum of patients with systemic lupus erythematosus1. PA28γ is a member of the PA28 family of proteasome activators and forms a homoheptamer that binds to the 20S catalytic core of the proteasome. Unlike typical ATP-dependent ubiquitination and degradation by the 26S proteasome, the PA28γ-proteasome promotes the degradation of model peptide substrates in an ATP- and ubiquitin-independent manner2. Notably, the PA28γ-proteasome degrades intact cellular proteins such as SRC-33, and subsequent studies identified multiple substrates including p21WAF1, p16INK4A, GSK3β, PTTG1, SirT1, c-Myc, and IκBε4,5,6,7,8,9,10, a function unique to PA28γ within the PA28 family.
Head and neck squamous cell carcinoma (HNSCC) is the most common malignancy in the head and neck region, with a steadily increasing incidence11. HNSCC frequently impairs craniofacial functions and can be life-threatening. While conventional surgical treatments remain standard, they may compromise facial aesthetics and impose physical and psychological burdens. Therefore, exploring effective molecular therapeutic targets is crucial. Our previous investigations indicated that PA28γ promotes HNSCC malignant progression. Furthermore, PA28γ-knockdown reversed the malignant phenotype and inhibited tumor growth in xenograft mouse models12,13. Amplification or overexpression of PA28γ has also been found in many human cancers, promoting malignant growth and correlating with poor patient outcomes14,15. Despite the growing evidence of PA28γ’s contribution to cancer progression, the molecular mechanisms regulating its function in tumors remain largely uncharacterized.
Protein phosphorylation is the most widespread covalent post-translational modification and plays critical roles in regulating tumor growth and therapeutic responses16,17. However, many phosphorylation sites and their functional significance remain to be explored. In the context of PA28γ, only a single phosphorylation site (S247) associated with apoptosis has been reported in human osteosarcoma cells18, and no phosphorylation events have been described so far in HNSCC or other cancers. Therefore, exploring phosphorylation sites on PA28γ that may modulate its role in HNSCC malignant progression is a critical and timely objective.
Equally important is identifying the kinase responsible for such phosphorylation events, as understanding the upstream regulators is essential for deciphering the mechanisms controlling PA28γ function in cancer. Among potential kinases, casein kinase 2 (CK2), a ubiquitous serine/threonine kinase, phosphorylates hundreds of substrates and represents a major contributor to the human phospho-proteome. CK2 is a heterotetramer composed of two catalytic subunits (CK2α and CK2α‘, encoded by CSNK2A1 and CSNK2A2) and two regulatory CK2β subunits (encoded by CSNK2B), which maintain substrate binding and enzyme stability19,20. CK2 interacts directly with cancer-associated proteins and is often essential for cancer cell survival21. Understanding CK2-mediated phosphorylation could provide insights into cancer progression and therapeutic potential.
Besides kinase-mediated phosphorylation, transcription factors also critically influence tumor progression by regulating gene expression programs that control cell proliferation and survival. One such factor is E4F transcription factor 1 (E4F1), a member of the GLI-Kruppel family of zinc finger proteins, which modulates the cell cycle and cancer growth22. Analysis of the TCGA database reveals that low E4F1 expression associates with poor prognosis in HNSCC patients, suggesting its potential as a protective prognostic factor22. Therefore, understanding molecular mechanisms regulating abnormal growth in HNSCC and the functional and physical connections among these molecules is vital for refining oncogenic networks and identifying therapeutic targets.
Here, we show phosphorylation at the T23 site of PA28γ (PA28γ-pT23) and investigate its abundance in various cancers, including HNSCC. We demonstrate that PA28γ-pT23 serves as a prognostic marker. Importantly, PA28γ-pT23 deficiency and continuous phosphorylation confer resistance and susceptibility, respectively, to carcinogen-induced tumors. We further reveal that CK2 mediates PA28γ-T23 phosphorylation, which controls the degradation of the growth-control factor E4F1 via the PA28γ-proteasome pathway. Overall, our data elucidate the pro-oncogenic role and molecular mechanism of the CK2-mediated PA28γ-pT23 axis in tumor initiation and progression, highlighting its potential as a target for early cancer intervention and therapy.
Results
PA28γ-T23 phosphorylation exhibits a crucial pro-oncogenic signal and unfavorable prognostic indicator in various cancers
To investigate the phosphorylation of PA28γ protein in HNSCC, we established HSC-3 cells expressing Flag-PA28γ, and the Flag-antibody-immunoprecipitated proteins were resolved by SDS–PAGE and visualized by Coomassie blue staining (Supplementary Fig. 1a). PA28γ-T23 was identified as the phosphorylation site of PA28γ by liquid chromatography-tandem mass spectrometry (LC-MS/MS) in HSC-3 cells (Fig. 1a). Subsequently, we constructed an unphosphorylatable alanine mutation, T23A, and a phosphomimetic aspartic acid mutation, T23D, which were transduced into HNSCC cells using lentiviral vectors. The overall phosphorylation levels of PA28γ dramatically decreased following the T23 mutation (Fig. 1b). Furthermore, given that the PhosphositePlus database shows repeated detection of phosphorylation at the S24 site adjacent to T23 in other study models, we introduced the S24A mutant into HNSCC cells to prevent any potential oversight. Our results indicated that the S24 mutation did not significantly affect the overall phosphorylation levels of PA28γ in HNSCC cells (Supplementary Fig. 1b). These findings, together with our LC-MS/MS data, suggest that T23 is the predominant phosphorylation site in the HNSCC model employed in this study. We then generated and validated a phospho-specific antibody against PA28γ phosphorylated at T23 (PA28γ-pT23). The specificity of this antibody for PA28γ-pT23, without cross-reactivity to the adjacent S24 site, was confirmed by dot blot assays and immunoblotting analyses using site-directed mutants (Supplementary Fig. 1c–e). Moreover, we observed that the absolute and normalized levels of PA28γ-pT23 were significantly higher in HNSCC tumor tissues compared to adjacent tissues (Fig. 1c).
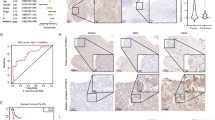
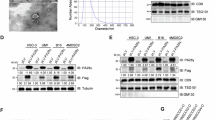
a Determination of PA28γ phosphorylation sites by liquid chromatography-tandem mass spectrometry (LC-MS/MS) analysis. The LC-MS/MS spectrum of modified “IT(phos)SEAEDLVANFFPK” is shown. b Immunoblot (IB) analysis of immunoprecipitation (IP) products and whole-cell lysates (WCLs) derived from HNSCC cells transduced with empty vector (EV), Flag-PA28γ WT, T23A, or T23D. c Human HNSCC tumors and paired adjacent normal tissues were extracted and collected to IB analysis (n = 4). Statistical analysis of the normalized level of PA28γ-pT23 is shown. A, adjacent tissue; T, tumor tissue. d–f IHC analyses of PA28γ-pT23 in the tissue microarray (TMA) containing 81 HNSCC patients with paired tumors and adjacent tissues. Assessment of their levels in tumors and adjacent tissues (d) as well as in different AJCC stages (e). Survival analysis with Kaplan-Meier curves was performed on their high and low levels (f). Scale bars, 100 μm. g, h IHC analyses of PA28γ-pT23 in the TMA containing 13 types of cancer with paired tumors and adjacent tissues. Representative samples (g) and statistical analysis (h) of PA28γ-pT23 levels in tumors and adjacent tissues are shown. Scale bars, 100 μm. i IHC analyses of PA28γ-pT23 in rectal cancer TMA (n = 90), pancreatic cancer TMA (n = 89), and breast cancer TMA (n = 130). Survival analysis with Kaplan-Meier curves was performed on PA28γ-pT23 high and low levels. Statistical significance was assessed by two-sided paired t-test in (c, d) unpaired t-test in (e, h) and log-rank (Mantel-Cox) test in (f, i). Source data are provided as a Source Data file.
We subsequently utilized an HNSCC tissue microarray containing 81 patients to detect the expression patterns of PA28γ-pT23 and its potential clinical prognostic implications (Supplementary Data 1), and found that absolute levels of PA28γ-pT23 were upregulated in tumor tissues compared with adjacent normal counterparts (Fig. 1d). Moreover, absolute levels of PA28γ-pT23 exhibited upregulation in tumor tissues of advanced American Joint Committee on Cancer (AJCC) stages, and its high levels were associated with poor survival (Fig. 1e, f and Supplementary Data 2). To further investigate whether a similar pattern exists for PA28γ-pT23 in other cancers, we performed immunohistochemistry (IHC) staining on tissue microarrays of pan-caner (13 cancer types, n = 91), rectal cancer (n = 90), pancreatic cancer (n = 89), and breast cancer (n = 130) (Supplementary Data 3–6). Of note, consistent with findings in HNSCC, PA28γ-pT23 is overexpressed in tumor tissues of the 11 cancer types compared with their corresponding adjacent tissues (Fig. 1g and h and Supplementary Fig. 1 f, g), especially in advanced AJCC stages (Supplementary Fig. 1h–j), and positively correlates with a poor survival outcome (Fig. 1i and Supplementary Data 7). Collectively, these results highlight that PA28γ-pT23 can serve as a crucial signal for promoting cancer progression and a poor prognosis marker in various cancer types, not just HNSCC.
Phosphorylation of PA28γ-T23 facilitates tumor initiation and progression
To investigate the impact of PA28γ-pT23 on tumorigenesis and progression in vivo, we generated T23 phosphorylation-deficient PA28γT23A and phosphomimetic PA28γT23D knock-in mice using CRISPR–Cas9 technology (Fig. 2a, b). Notably, both PA28γT23A and PA28γT23D mice exhibited comparable body size and weight compared with WT mice (Supplementary Fig. 2a, b), unlike the growth retardation observed in PA28γ-knockout mice23. Interestingly, proliferation marker staining showed no significant difference in Ki67-positive cells among the three groups, while Cyclin A2 staining was reduced in T23A and increased in T23D compared to WT (Supplementary Fig. 2c, d). This indicates that the T23 mutation affects certain intracellular pathways, but compensatory mechanisms within complex molecular networks likely prevent overt phenotypes in normally developing mice. We then established HNSCC models in WT and T23 mutant mice using the carcinogen 4-Nitroquinoline 1-oxide (4NQO) to disrupt tissue homeostasis and assess potential phenotypic differences under tumor-inducing conditions (Fig. 2c). In this model, we observed that PA28γT23A mice exhibited visibly fewer and smaller cauliflower-like or protruding lesions on the tongue and less weight loss compared with WT mice, while PA28γT23D mice had more lesions and greater weight loss (Fig. 2d–f and Supplementary Fig. 2e). Additionally, lower tumor invasion rates were observed in PA28γT23A mice compared with WT mice, while all PA28γT23D mice developed tumor invasion, with a higher proportion than the WT group (Fig. 2g). This suggests that PA28γ-pT23 could promote HNSCC formation and progression in vivo. Therefore, our results suggest that PA28γ-pT23 could sensitively impact tumor initiation and progression without affecting mouse development.
a Schematic diagram of gRNA target sequences and partial donor oligo sequences in CRISPR/Cas-mediated genome engineering for generating the C57BL/6 mouse model with a point mutation (T23A or T23D) at the PA28γ gene locus. The gRNA specifically targets the exon 2 region of the PA28γ gene, which encodes T23. The donor oligo introduced the T23A (ACA to GCA) or T23D (ACA to GAC) mutation into exon 2 by homology-directed repair. In addition, a synonymous mutation R21 (CGG to CGC) was introduced to prevent the binding and re-cutting of the sequence by gRNA after homology-directed repair. b Sanger sequencing results derived from genomic DNA of PA28γT23A and PA28γT23D mice. c Experimental schematic illustrating the strategy for inducing HNSCC in WT, PA28γT23A, and PA28γT23D mice (n = 16/group) using the carcinogen 4NQO. d Representative images of tongue lesions in 4NQO-induced HNSCC mouse models at week 24. The dotted circles indicate macroscopically prominent lesions. e The number (Left) and volume (Right) of macroscopic prominent lesions in the tongues were quantified in week 24 of the 4NQO-induced HNSCC experiment. Data shown represent the mean ± SD for n = 16 mice; statistical significance was assessed by two-sided unpaired t-test. f Survival analysis using the Kaplan-Meier method on over 10% body weight loss in WT, PA28γT23A, and PA28γT23D mice during weeks 16–24 of the 4NQO-induced HNSCC experiment. Statistical significance was assessed by the log-rank (Mantel-Cox) test. g Representative H&E staining of severe dysplasia or carcinoma in-situ, microinvasive carcinoma, and invasive carcinoma in tongue lesions (Left). Scale bars, 100 μm. The tongue lesions of three mouse groups were evaluated and compared (Right). Statistical significance was assessed by two-sided Fisher’s exact test. Source data are provided as a Source Data file.
CK2 phosphorylates PA28γ at T23
We next investigated the kinase responsible for phosphorylating the T23 site. First, we performed a bioinformatics analysis using DTU Health Tech (an online bioinformatics tool), which predicted 14 putative kinases involved in phosphorylating the T23 residue (Supplementary Data 8). LC-MS/MS analysis was also performed on the immunoprecipitated PA28γ and its potential interacting proteins (Supplementary Fig. S3a and Supplementary Data 9). The intersection of results from the bioinformatics analysis and the LC-MS/MS analysis revealed CK2 as a putative kinase for PA28γ-T23 (Supplementary Fig. S3b), with a potential interaction between PA28γ and CK2β (the regulatory subunit of CK2) suggested by the LC-MS/MS analysis. Co-immunoprecipitation assays demonstrated the interaction between CK2β and PA28γ proteins (Fig. 3a and Supplementary Fig. S3c, d), with immunofluorescence staining indicating both primarily localized in the nucleus (Fig. 3b).
a WCLs derived from HSC-3 cells were immunoprecipitated with control IgG, anti-PA28γ, or anti-CK2β antibody, followed by IB analysis of co-IP products and WCLs. b Immunofluorescence analyses of the cellular localization of endogenous PA28γ and CK2β in HNSCC cells. DAPI (blue), nucleus; Scale bars, 10 μm. c Schematic representation of the evolutionarily conserved PA28γ-T23 site in different species. Residues around the T23 site have a common characteristic of CK2 kinase substrates, namely the predominance of acidic amino acid residues (E/D) between n-4 and n + 7. α1–4 denotes helix-α1–4. d IB analysis of IP products and WCLs derived from HEK293T cells transfected with the indicated plasmids. e In vitro kinase assays were performed with recombinant His-PA28γ WT and T23A proteins purified from E. coli system as substrates and purified CK2 proteins containing CK2α and CK2β as the source of kinase. IB analysis of the indicated proteins in vitro kinase assay samples. f, g IB analysis of WCLs derived from HEK293T cells transfected with the subunits of CK2 (f) and HNSCC cells treated with 100ngml–1 EGF before harvesting (g). h, i HNSCC cells silenced with siRNA negative control (siNC) or siCK2 (h) and HSC-3 cells silenced with siNC or CK2α/α‘/β siRNA (i) were stimulated without or with 100ngml–1 EGF for 30 min before harvesting for IB analysis. j HEK293T cells transfected with the indicated plasmids were treated with 100ngml–1 EGF. WCLs were immunoprecipitated with the indicated antibodies, followed by IB analysis of co-IP products and WCLs. k In vitro kinase assays were performed with recombinant His-PA28γ proteins purified from E. coli system as substrates and purified CK2 proteins containing CK2α and CK2β as the source of kinase. 2 μM or 20 μM CK2 inhibitor TBB was added as indicated. l IB analysis of IP products and WCLs derived from HEK293T cells transfected with the indicated plasmids and silenced with siNC or #1, #2 NIP30 siRNA. m IB analysis of WCLs derived from HEK293T cells transfected with the indicated plasmids and silenced with siNC or #1, #2 NIP30 siRNA. Source data are provided as a Source Data file.
Notably, the T23 residue of PA28γ is highly conserved across different species, and the surrounding residues conform to the common features of CK2 kinase substrates, with a high occurrence of acidic amino acids E/D in the n-4 to n + 7 positions24 (Fig. 3c). We found that overexpression of CK2 in cells only upregulated the pT23 levels of PA28γ WT, but not T23A (Fig. 3d and Supplementary Fig. S3e). Moreover, CK2-knockdown resulted in a marked reduction of PA28γ-pT23 levels (Supplementary Fig. S3f). Consistent with the intracellular results, CK2 directly phosphorylated the T23 residue of PA28γ WT in vitro, but not T23A (Fig. 3e). As CK2 is a heterotetrameric holoenzyme, we next investigated the effects of overexpressing individual subunits of CK2 on pT23 levels. Interestingly, overexpression of any single subunit of CK2 did not have an observable impact on pT23 levels. However, overexpression of its regulatory subunit CK2β and at least one catalytic subunit of CK2 (either CK2α or CK2α‘) significantly increased the pT23 levels, demonstrating the dependence of pT23 regulation on the CK2 holoenzyme (Fig. 3f).
Furthermore, PA28γ-pT23 levels showed a time-dependent increase following stimulation with epidermal growth factor (EGF) – an activator of CK2 kinase25,26,27 (Fig. 3g). Consistent with the effect of CK2, EGF significantly increased PA28γ-pT23 levels in WT, but not the T23A mutation (Supplementary Fig. 3g). Notably, the EGF-induced T23 phosphorylation is more pronounced following CK2 overexpression compared with its control (Supplementary Fig. 3h). Upon CK2 knockdown, EGF failed to alter pT23 levels, suggesting the dependence of EGF-induced pT23 on CK2 kinase (Fig. 3h). Furthermore, we investigated the key subunits of CK2 involved in regulating pT23. We observed that knocking down CK2β or both catalytic subunits CK2α and CK2α‘ significantly blocked the EGF-induced upregulation of pT23 levels, while the knockdown of either the CK2α or CK2α‘ subunit had no apparent effect, highlighting the importance of CK2β subunit in pT23 regulation (Fig. 3i and Supplementary Fig. 3i). We further found that the interaction between CK2β and PA28γ was promoted by EGF stimulation (Fig. 3j and Supplementary Fig. 3j). In addition, the CK2 inhibitors 4,5,6,7-tetrabromobenzotriazole (TBB)28 and CX-494529 were found to dose-dependently reduce PA28γ-pT23 levels (Fig. 3k and Supplementary Fig. 3k, l). Notably, the reduction in PA28γ-pT23 levels was observed only upon CK2 inhibition, but not with the inhibition of GSK3, CaMK-II, or CK1 activity (Supplementary Fig. 4a), and this effect was not affected by the S24A mutation (Supplementary Fig. 4b). Furthermore, compared to PA28γ WT, the T23A mutation abolished both T23 phosphorylation and CK2-mediated T23 phosphorylation, while the S24A mutation had no significant effect on either (Supplementary Fig. 4c). These results also indicate that phosphorylation of the T23 site by CK2 is independent of the S24 site.
Given that CK2 serves as the kinase of NIP30, a major partner of PA28γ30,31, it is essential to determine whether the interaction between PA28γ and CK2, as well as PA28γ-T23 phosphorylation, is associated with NIP30. Following the knockdown or overexpression of NIP30, no alterations were observed in the interaction between PA28γ and CK2β (Fig. 3l and Supplementary Fig. 4d, e), and the upregulation of CK2-mediated PA28γ-T23 phosphorylation was also unaffected when NIP30 expression was altered (Fig. 3m and Supplementary Fig. 4f), indicating that the PA28γ-CK2 interaction and PA28γ-pT23 are independent of NIP30. Collectively, these data suggest that the T23 residue of PA28γ undergoes phosphorylation, which is mediated by CK2 kinase.
Phosphorylation of PA28γ at T23 attenuates E4F1 protein levels, thereby alleviating its transcriptional effect
To further investigate the molecular mechanisms by which PA28γ-T23 phosphorylation regulates tumor malignancy and its association with prognosis, we performed quantitative proteomic analysis on PA28γ WT and PA28γ-T23A groups, identifying 39 potentially differentially expressed proteins (Fig. 4a, Supplementary Fig. 5a and Supplementary Data 10). Given that E4F1 influences survival in HNSCC patients (Fig. 4b), serves as an independent prognostic risk factor (Supplementary Data 11), and is clearly associated with growth and proliferation, it was selected as the downstream focus of pT23. We next performed multiplex IHC staining on the tongues of WT and T23 mutant mice, revealing nuclear co-localization of PA28γ-pT23 and E4F1 in the tongue epithelial cells (Fig. 4c). Moreover, we quantified E4F1 levels in the 4NQO-induced mouse model. Consistent with our mass spectrometry findings, E4F1 levels in PA28γT23A mice were higher than those in WT mice, whereas PA28γT23D mice exhibited lower E4F1 levels compared to WT mice (Supplementary Fig. 5b). Concurrently, we found that PA28γT23A mice exhibited decreased expression of proliferation markers in dysplastic lesions and tumor tissues, including reduced Cyclin A2 staining intensity and fewer Ki67-positive cells compared with WT mice, whereas PA28γT23D mice showed the opposite trend (Supplementary Fig. 5b).
a Heatmap of 26 up-regulated and 13 down-regulated proteins from proteomic analysis between PA28γ WT and PA28γ-T23A-rescued HEK293T cells (both groups with three biological replicates). b Kaplan-Meier survival analysis of overall survival in HNSCC patients from the TCGA database based on E4F1 expression using Kaplan-Meier Plotter. Statistical significance was assessed by the log-rank (Mantel-Cox) test without adjustment for multiple comparisons. c Representative multiplex IHC images of tongue mucosal tissues from WT, PA28γT23A, and PA28γT23D mice with the indicated antibodies. Scale bars, 50 μm. d PA28γ WT, T23A or T23D were re-expressed in endogenous PA28γ-knockdown HNSCC cells to establish PA28γ WT, PA28γ-T23A, and PA28γ-T23D-rescued HNSCC cell lines. IB analysis of WCLs derived from HNSCC cells transduced with lentiviral vectors expressing shRNA against endogenous PA28γ and EV, Flag-PA28γ WT, T23A, or T23D. The upper band is exogenous Flag-PA28γ, the lower band is endogenous PA28γ as marked. O.E., overexpression; Endo., endogenous. e IB analysis of WCLs derived from the indicated HNSCC cells. f, g IB analysis of WCLs derived from PA28γ WT, PA28γ-T23D (f), or PA28γ-T23A-rescued (g) HEK293T cells transfected with the indicated plasmids. h, i IB analysis of WCLs derived from PA28γ-rescued HNSCC cells treated with 100ngml–1 EGF (h) or HSC-3 cells treated with 50 μM TBB (i) at indicated time points. Source data are provided as a Source Data file.
To better understand the impact of PA28γ-pT23 on E4F1, we conducted a series of experiments using our constructed PA28γ WT, PA28γ-T23A, and PA28γ-T23D-rescued cell lines (Fig. 4d). These cells stably expressed PA28γ WT, PA28γ-T23A or PA28γ-T23D, with simultaneous stable knockdown of endogenous PA28γ. Compared with PA28γ WT, the T23A exhibited an increase in E4F1 protein levels, whereas the T23D displayed a decrease, without affecting E4F1 mRNA expression (Fig. 4e and Supplementary Fig. 5c, d). Furthermore, the T23A mutation prolonged the half-life of the E4F1 protein compared to WT, while the T23D mutation shortened it (Supplementary Fig. 5e). Notably, overexpression and knockdown of E4F1 in HNSCC cells led to a decrease and increase in the mRNA and protein levels of the cell cycle regulator Cyclin A2, respectively (Supplementary Fig. 6a–d), supporting the findings from several studies that Cyclin A2 is a transcriptional repression target of E4F122. Interestingly, we observed a decrease in Cyclin A2 protein levels following the T23A mutation compared to WT, while an increase was noted following the T23D mutation, which was opposite to the changes in E4F1 (Fig. 4e, f). This suggests that T23 phosphorylation can mitigate the transcriptional effect of E4F1 on Cyclin A2 by reducing E4F1 protein abundance.
Notably, we observed that overexpression or knockdown of CK2 led to a decrease and increase in E4F1 levels in WT, respectively, but not in T23A and T23D-rescued cells (Fig. 4g and Supplementary Fig. 6e). Consistently, EGF stimulation led to a reduction of E4F1 levels in WT, but not in T23A-rescued cells (Fig. 4h). In contrast, treatment with TBB or CX-4945 resulted in a time-dependent increase in the abundance of E4F1 (Fig. 4i and Supplementary Fig. 6f). Furthermore, E4F1 has also been reported to serve as a transcriptional activator for several genes such as DLAT, DNAJC19, CHK1, and TTI232,33. To further assess the regulatory effect of PA28γ on E4F1, we examined the expression of these genes in PA28γ WT and T23D-rescued cells, as well as in cells with control or ectopic expression of PA28γ. The results showed a decrease in mRNA levels of E4F1 target genes when either the T23D mutation or PA28γ WT was upregulated (Supplementary Fig. 6g, h). These results indicate that elevated PA28γ-pT23 levels can reduce E4F1 protein amounts, thereby alleviating E4F1-mediated transcriptional effect.
CK2-mediated phosphorylation of PA28γ-T23 facilitates the interaction of PA28γ with E4F1
To understand how PA28γ-pT23 affects E4F1 protein levels, we further explored the potential mechanisms. Given their nuclear co-expression (Fig. 5a), we conducted co-immunoprecipitation assays. These assays revealed an interaction between endogenous as well as ectopically expressed PA28γ and E4F1 proteins (Fig. 5b and Supplementary Fig. 7a, b). Furthermore, to determine the specific region of E4F1 binding to PA28γ, we generated several truncations of the E4F1 protein. Co-immunoprecipitation and pull-down assays demonstrated that PA28γ specifically interacted with the aa357–784 of E4F1, rather than the aa1–357 responsible for DNA binding (Fig. 5c and Supplementary Fig. 7c, d).
a Immunofluorescence analyses of the cellular localization of endogenous PA28γ and E4F1 in HNSCC cells. DAPI (blue), nucleus; Scale bars, 10 μm. b WCLs derived from HNSCC cells were immunoprecipitated with control IgG, anti-PA28γ, or anti-E4F1 antibody, followed by IB analysis of co-IP products and WCLs. c Schematic representation of the various E4F1 truncations used in the PA28γ-binding assays tested for their capacity to bind PA28γ. The ubiquitin E3 ligase active region is represented by a green box, while the C2H2 and C2HC zinc finger motifs are highlighted in yellow and pink, respectively (Left). IB analysis of co-IP products and WCLs derived from HEK293T cells transfected with Flag-PA28γ and Myc-E4F1 truncations (Right). d IB analysis of co-IP products and WCLs derived from PA28γ-rescued HEK293T cells transfected with indicated constructs. e PA28γ-rescued HEK293T cells transfected with the indicated plasmids were stimulated without or with 50 μM TBB for 30 min before harvesting. WCLs were immunoprecipitated with Flag-beads, followed by IB analysis of co-IP products and WCLs. f HSC-3 cells transduced with control or Flag-PA28γ lentiviral vectors and silenced with siNC or siCK2 were stimulated without or with 100ngml–1 EGF for 30 min before harvesting. WCLs were immunoprecipitated with Flag-beads, followed by IB analysis of co-IP products and WCLs. g IB analysis of co-IP products and WCLs derived from PA28γ-rescued UM1 cells treated with 100ngml–1 EGF for the indicated time before harvesting. h, i HSC-3 cells were stimulated with 50 μM TBB (h) or 1 μM or 5 μM CX-4945 (i), and 100ngml–1 EGF before harvesting. WCLs were immunoprecipitated with anti-PA28γ antibody, followed by IB analysis of co-IP products and WCLs. j PA28γ-rescued HSC-3 cells were stimulated without or with 50 μM TBB and 100ngml–1 EGF for 30 min before harvesting. WCLs were immunoprecipitated with Flag-beads, followed by IB analysis of co-IP products and WCLs. k HEK293T cells were transfected with indicated plasmids and treated without or with 100ngml–1 EGF and 50 μM TBB for 30 min before being subjected to GST-PD assays and IB analyses. Source data are provided as a Source Data file.
We next investigated the potential impact of PA28γ-pT23 on the interaction of PA28γ with E4F1. Importantly, we observed that the T23A mutation reduced the PA28γ-E4F1 interaction, while the T23D mutation increased their interaction (Fig. 5d and Supplementary Fig. 7e, f). Considering the effects of CK2 kinase on T23 phosphorylation, we investigated its potential impact on the PA28γ-E4F1 interaction. We found that CK2 overexpression enhanced the interaction of E4F1 with PA28γ WT, but not with the PA28γ-T23 mutants (Fig. 5e and Supplementary Fig. 7g, h). Notably, CK2-knockdown prevented the influence of EGF on their interaction (Fig. 5f). Furthermore, EGF stimulation strengthened the interaction between PA28γ and E4F1 (Fig. 5g and Supplementary Fig. 7i, j), whereas TBB or CX-4945 weakened it (Fig. 5h–k). Taken together, these results indicate an interaction between PA28γ and E4F1, which can be enhanced by CK2-mediated phosphorylation of PA28γ-T23.
PA28γ promotes E4F1 degradation through the PA28γ-proteasome pathway
Given that PA28γ-pT23 promotes the PA28γ-E4F1 interaction and reduces E4F1 protein levels, we further investigated the impact of PA28γ protein on E4F1. We found that PA28γ reduced E4F1 protein levels in a dose-dependent manner (Fig. 6a), while E4F1 mRNA expression remained unaltered upon PA28γ upregulation (Fig. 6b and Supplementary Fig. 8a). In addition, PA28γ overexpression reduced the half-life of E4F1 protein, an effect that is dependent on the aa357–784 of E4F1, but not the aa1–357 (Fig. 6c–e). Conversely, the knockdown of PA28γ increased E4F1 levels and enhanced the stability of E4F1 (Supplementary Fig. 8b, c). Additionally, we observed that downregulation of PA28γ resulted in increased levels of its known downstream proteins p21WAF1, SRC3, SirT1, and p53, and knocking down E4F1 did not affect this effect (Supplementary Fig. 8d). The in vitro proteolytic assay also demonstrated that E4F1 does not influence the PA28γ-mediated degradation of the p21WAF1 protein (Supplementary Fig. 8e). Thus, it is evident that the effects mediated by PA28γ on its key targets such as p21WAF1, SRC3, SirT1, and p53 are independent of E4F1. These findings, coupled with the role of PA28γ as a proteasome activator, prompt our further investigation into its potential role in regulating E4F1 through the PA28γ-proteasome.
a IB analysis of WCLs derived from HEK293T cells transfected with His-E4F1 and increasing doses of Flag-PA28γ. b Quantitative real-time PCR analysis of the indicated genes in HEK293T cells transfected with EV and increasing doses of Flag-PA28γ. Endo., endogenous. c HSC-3 cells stably expressing EV or Flag-PA28γ were treated with 100 μgml−1 CHX for the indicated time before harvesting. Quantification of E4F1 levels relative to tubulin levels is shown. d, e HEK293T cells stably expressing EV or Flag-PA28γ were transfected with Myc-E4F1 truncations, and treated with 100 μgml−1 CHX for the indicated time before harvesting. Quantification of Myc-E4F1 levels relative to tubulin levels is shown. f HEK293T cells transfected with the indicated plasmids were treated without or with 10 μM MG132 for 6 h before harvesting, and WCLs were collected for IB analysis. g HEK293T cells stably expressing shRNA against endogenous PA28γ were transfected with the indicated plasmids, and WCLs were collected for IB analysis. h Purified PA28γ, E4F1 and 20S proteasome in the absence or presence of 100 nM proteasome inhibitor epoxomicin (Epox) were incubated as indicated for 45 min, followed by IB analysis. i Purified PA28γ WT, PA28γ-T23A, E4F1, p21 (as a positive control for E4F1) and 20S proteasome were incubated as indicated for 45 min, followed by IB analysis. Data in (b–e) represent the mean ± SD of three biological replicates; statistical significance was assessed by two-sided unpaired t-test. Source data are provided as a Source Data file.
We observed that the proteasome inhibitor MG132 blocked PA28γ-mediated downregulation of E4F1 (Fig. 6f). Interestingly, PA28γ WT and T23D, but not N151Y and K195R, which lack proteasome activation activity34,35, as well as T23A, caused a notable reduction in E4F1 levels (Fig. 6g and Supplementary Fig. 8f). Furthermore, compared with WT, both N151Y and T23A prolonged the half-life of E4F1 (Supplementary Fig. 8g). Importantly, we conducted in vitro proteolytic assays using the purified PA28γ WT protein with T23 phosphorylation (Supplementary Fig. 8h). The results demonstrated that E4F1 was degraded by the PA28γ-proteasome independently of ubiquitin and ATP, and this degradation process could be effectively inhibited by the proteasome inhibitor epoxomicin (Fig. 6h). Furthermore, compared with WT, T23A displayed a much weaker in vitro degradation capacity on E4F1 (Fig. 6i). These results suggest that the PA28γ-proteasome mediates E4F1 degradation. Collectively, our findings reveal that CK2-mediated PA28γ-pT23 enhances its binding with E4F1, resulting in increased degradation of E4F1 through the PA28γ-proteasome, ultimately reducing E4F1 abundance.
PA28γ-T23 phosphorylation promotes HNSCC cell proliferation and tumor growth through E4F1
Our previous study demonstrated that PA28γ-knockdown suppresses the proliferation and growth of HNSCC cells12. Moreover, E4F1, a protein involved in regulating the cell cycle and cell growth36,37, acts as a downstream effector of PA28γ. Interestingly, we observed that E4F1-knockdown rescued the decreased cell proliferation and clonogenic capacity (Fig. 7a and Supplementary Fig. 9a), as well as tumor growth inhibition caused by PA28γ downregulation (Fig. 7b–d and Supplementary Fig. 9b, c), indicating that E4F1 plays a crucial role in the PA28γ-mediated regulation of cancer cell growth.
a HSC-3 cells stably expressing shRNA against endogenous PA28γ were silenced with siNC or #1, #2 E4F1 siRNA. Cell numbers of cells were counted for 4 days in cell proliferation assays. b–d HSC-3 cells transduced with lentiviral EV, shPA28γ and/or shE4F1 vectors were injected into BALB/c nude mice (n = 6/group). The tumors derived from xenograft assays were dissected and photographed (b). Tumor sizes were monitored every 3 days starting one week after injection (c), and tumor masses were weighed at the endpoint (d). e, f Cell numbers of EV, PA28γ WT, PA28γ-T23A, and PA28γ-T23D-rescued HNSCC cells were counted for 4 days in cell proliferation assays (e), and the cells were seeded into 6-cm dishes in colony-formation assays (f). g–i EV, PA28γ WT, PA28γ-T23A, and PA28γ-T23D-rescued HSC-3 cells were injected into BALB/c nude mice (n = 6/group). The tumors derived from xenograft assays were dissected and photographed (g). Tumor sizes were monitored every 3 days starting one week after injection (h), and tumor masses were weighed at the endpoint (i). j–l PA28γ WT and PA28γ-T23A-rescued HSC-3 cells were silenced with siNC or #1, #2 E4F1 siRNA, and then collected for cell proliferation (j) and colony-formation (k) assays. Cells were also treated with propidium iodide staining, followed by flow cytometry analysis of DNA content (l). m, n PA28γ WT and PA28γ-T23D-rescued HNSCC cells silenced with siNC or siCK2 were collected for IB analysis (m) and colony-formation assays (n). Data in a, e, f, j–l and n represent the mean ± SD of three biological replicates; statistical significance was assessed by two-sided unpaired t-test. Source data are provided as a Source Data file.
Our investigation revealed the regulatory effect of PA28γ-pT23 on the growth regulator E4F1, prompting a subsequent exploration of their potential functions in tumor growth. Notably, T23A mutation or E4F1 overexpression effectively inhibited HNSCC cell and tumor growth, while the growth ability was found to be increased following T23D mutation or E4F1 knockdown (Fig. 7e–i and Supplementary Fig. 9d–l). Importantly, E4F1-knockdown rescued the decreased cell proliferation ability caused by the T23A mutation (Fig. 7j, k). Additionally, we observed that E4F1 overexpression in HNSCC cells suppressed cell growth by inhibiting the G1 to S phase transition of the cell cycle (Supplementary Fig. 9m). Interestingly, compared with WT, T23A-rescued cancer cells displayed G0/G1 arrest and the promotion of cell proliferation through E4F1-knockdown was achieved by relieving G0/G1 phase arrest, resulting in a significant increase in the S phase proportion (Fig. 7l). Furthermore, CK2-knockdown inhibited colony formation in PA28γ WT, but not T23D-rescued cancer cells (Fig. 7m, n), suggesting that CK2 depletion can suppress cancer cell growth by reducing PA28γ-pT23. These findings demonstrate that CK2-mediated PA28γ-pT23 promotes both in vivo and in vitro HNSCC cell growth through E4F1.
CK2-mediated PA28γ-T23 phosphorylation axis facilitates HNSCC progression
Given our findings on E4F1 as a downstream target of PA28γ-pT23, we subsequently validated its expression patterns in the HNSCC tissue microarray (TMA). E4F1 exhibited an inverse trend relative to PA28γ-pT23, with higher expression in adjacent normal tissues, decreased levels in advanced AJCC stages, and poor overall survival observed in patients with low E4F1 expression (Fig. 8a–c and Supplementary Data 12). These results suggest that E4F1 is associated with disease progression and patient outcomes in HNSCC.
a–c Multiplex IHC analyses of E4F1 in the TMA containing 81 HNSCC patients with paired tumors and adjacent tissues. Assessment of its levels in tumors and adjacent tissues (a), as well as in different AJCC stages (b). Survival analysis with Kaplan-Meier curves was performed on its high and low levels (c). Scale bars, 100 μm. d Correlation analyses among CK2β, PA28γ-pT23, E4F1, and Cyclin A2 were performed based on multiplex IHC staining scores from the HNSCC TMA. Representative tumor samples are shown. Scale bars, 100 μm. e Expression patterns of CK2β, PA28γ-pT23, E4F1, and Cyclin A2 in normal tissues, premalignant lesions, primary tumors, and metastatic lymph nodes were analyzed based on multiplex IHC staining scores. Representative samples are shown. Scale bars, 100 μm. f Expression levels of CK2β, PA28γ-pT23, E4F1, and Cyclin A2 in HNSCC tissues with or without lymph node metastasis. g Schematic illustration of this study. The study reveals the expression patterns of PA28γ-pT23 in cancers along with its clinical prognostic implications (Upper left panel), the sensitivity of T23 mutations to carcinogen-induced HNSCC in mice (Lower left panel), and the molecular mechanisms by which the CK2-mediated PA28γ-pT23 axis regulates cancer cell proliferation and growth (Right panel). Statistical significance was assessed by two-sided paired t-test in (a) unpaired t-test in (b, e and f) log-rank (Mantel-Cox) test in (c) and two-sided Pearson’s correlation test in (d). Source data are provided as a Source Data file.
Furthermore, we analyzed the expression correlations of proteins within the CK2-mediated PA28γ-T23 phosphorylation axis in the TMAs of HNSCC and other cancers. PA28γ-pT23 levels did not consistently correlate with CK2β expression (Fig. 8d and Supplementary Fig. 10), possibly reflecting coordinated regulation of PA28γ-pT23 by multiple CK2 subunits rather than CK2β alone. Consistent with the observation that PA28γ-pT23 reduces E4F1 levels, E4F1 expression was negatively correlated with both PA28γ-pT23 and Cyclin A2, while Cyclin A2 showed a positive correlation with PA28γ-pT23 (Fig. 8d and Supplementary Fig. 10). We also observed a progressive increase in CK2β, PA28γ-pT23, and Cyclin A2 levels from normal and premalignant tissues to primary tumors, and further to metastatic lymph nodes, whereas E4F1 expression showed a decreasing trend across these stages (Fig. 8e and Supplementary Data 13, 14). Moreover, in HNSCC cases with lymph node metastasis, CK2β, PA28γ-pT23, and Cyclin A2 levels were significantly elevated compared to tumors without lymph node involvement, whereas E4F1 levels were reduced (Fig. 8f and Supplementary Data 14). Collectively, these findings provide direct patient-derived evidence supporting that the CK2-mediated PA28γ-T23 phosphorylation axis promotes HNSCC progression.
Discussion
PA28γ has garnered significant interest due to its ability to regulate multiple molecules implicated in tumor growth, such as p53, SRC-3, p21WAF1, p16INK4A, IκBε, GSK3β, c-Myc, and SirT13,4,5,6,8,9,10,38, thereby impacting the malignant development of cancers. Overexpression of PA28γ is often closely associated with unfavorable prognostic outcomes in various cancers14. Furthermore, our previous findings suggested that PA28γ serves as a prognostic risk factor in HNSCC, promoting malignant tumor growth12,13. However, the intricate mechanisms underlying this process warrant further exploration.
Protein phosphorylation has been a focal point of research on the mechanisms of cancer pathogenesis and treatment strategies for an extended period16. Previous studies have indicated that phosphorylation is involved in various hallmark processes of cancer and is associated with the initiation and progression of various cancer types17,39. In our current study, we found that PA28γ can be phosphorylated at the T23 site. This modification is prevalent across pan-cancer types, including HNSCC, and its high expression is closely associated with poor patient prognosis, suggesting its potential as a prognostic marker. Notably, PA28γ-pT23 promoted the initiation and progression of carcinogen-induced tumors. This indicates that targeting this site could prevent tumor development at early precancerous stages, highlighting its potential as an early intervention target in cancer therapy. Furthermore, we identified a proliferative signaling axis based on the specific post-translational modification of PA28γ at T23, namely CK2-PA28γ-T23-E4F1, which functions in cell cycle control (Fig. 8g). Notably, we analyzed the expression patterns of proteins within this axis, as well as their correlations, across normal tissues, premalignant lesions, primary tumors, and metastatic lymph nodes, indicating that activation of this axis is closely associated with HNSCC progression. However, future validation in large, longitudinal clinical cohorts is required to establish whether activation of the pT23 axis serves as a risk factor for malignant transformation and/or lymph node metastasis. This will be valuable for laying the groundwork for potential clinical translation.
In our previous review, we emphasized E4F1 as a critical regulator of cell growth, associated with the development of various cancers. Survival analysis based on the TCGA database indicated that E4F1 affects patient prognosis in various cancers22. Notably, the role of E4F1 in cell cycle regulation is also intertwined with the well-known tumor suppressor gene TP53. TP53 is the most frequently mutated gene in pan-cancer and the most commonly mutated gene in HNSCC40, often resulting in loss of p53 function. Although p53 is reported to be indispensable for E4F1-mediated growth arrest41, in this study, altering E4F1 levels in TP53-mutated HNSCC cell lines also affected the cell cycle, consistent with some reports36,42, indicating that E4F1 can regulate cell proliferation in a p53-independent manner. Here, we also identified E4F1 as a substrate of the PA28γ−20S proteasome system. Structural prediction analysis using the AlphaFold Server (AlphaFold 3 model)43 indicates that the E4F1 protein contains extensive flexible regions, characterized by intrinsic disorder and a lack of rigidly folded structure. This observation is consistent with recent findings that the PA28γ−20S proteasome preferentially degrades unfolded proteins44. Future research is required to elucidate the tertiary structure of E4F1 and to clarify the structural mechanisms underlying its specific regulation by PA28γ-T23 phosphorylation. Moreover, previous studies indicated that overexpression of full-length E4F1 can inhibit Cyclin A2 transcription, leading to G0/G1 phase arrest22, consistent with our results. These findings refine the understanding of E4F1’s function and mechanism as a growth regulator in HNSCC, laying a theoretical foundation for potential therapeutic strategies targeting E4F1. Further investigations will be needed to determine other targets regulated by PA28γ-pT23 through E4F1.
In summary, our study identifies PA28γ-pT23 as a crucial prognostic marker for cancer patients and a potential target for early cancer prevention. Additionally, we elucidate that CK2-mediated phosphorylation of PA28γ at T23 regulates E4F1 levels, which is a critical intrinsic mechanism driving abnormal tumor proliferation. Therefore, targeting the CK2-mediated PA28γ-pT23 axis may represent a promising therapeutic strategy for treating cancers, including HNSCC.
Methods
Study approval
This study complies with all relevant ethical regulations and was approved by the Research Ethics Committee of West China Hospital of Stomatology, Sichuan University, China. Approval for animal experiments was obtained under protocol numbers WCHSIRB-D-2020-023 and WCHSIRB-D-2024-240. Approval for the use of human tissue samples was granted under protocol numbers WCHSIRB-D-2020-046, WCHSIRB-D-2024-115, and by the National Human Genetic Resources Sharing Service Platform (approval number 2005DKA21300).
All mice were housed in specific pathogen-free (SPF) conditions at an ambient temperature of 20–26 °C and humidity of 30–70%, with a 12 h light/dark cycle, and handled according to the Institutional Animal Care and Use Committee guidelines at the State Key Laboratory of Oral Diseases, Sichuan University. The maximal tumor burden permitted was 1500 mm³. Throughout the study, tumor sizes in all mice were carefully monitored and did not exceed this limit at any time. Animals were euthanized before reaching it to prevent distress, ensuring compliance with ethical standards. All human tissue samples obtained from TMAs were collected with prior written informed consent from the participants. Sex of human tissue samples was determined based on original clinical records associated with each sample. No sex assignment or modification was performed by the researchers.
Moreover, with the exception of breast cancer, whose tissue samples on the TMA were exclusively derived from female patients, sex was not incorporated as a variable in the experimental design or data analysis of other mouse models or human TMA samples due to the absence of prior evidence indicating sex-dependent differences in tumor susceptibility in the models used.
Cell culture, transfection, and transduction
HEK293T cells were obtained from ATCC (no. CRL-3216). HSC-3 cells were obtained from the JCRB Cell Bank (no. JCRB0623). UM1 cells were obtained from the Gutkind Lab, UC San Diego Moores Cancer Center. All cell lines were routinely confirmed to be free of mycoplasma contamination. The cells were maintained in DMEM medium supplemented with 10% fetal bovine serum (FBS), 100Uml−1 penicillin, and 100 μgml−1 streptomycin. EGF (Sigma, SRP3027), TBB (Sigma, T0826), CX-4945 (TargetMol, T2259), CHIR99021 (TargetMol, T22657), KN-93 (TargetMol, T2697), D4476 (TargetMol, T2449), CHX (Sigma, C7698), and MG132 (Sigma, M7449) were used at the indicated times and doses. Cell transfection was performed using Lipofectamine 2000 (Invitrogen, 11668019) for plasmid DNA and Lipofectamine RNAiMAX for siRNA (Invitrogen, 13778150) following the manufacturer’s instructions. For cell transduction with lentiviral vectors expressing short-hairpin RNA (shRNA) or plasmids, 50% confluent cells with fresh medium were added with lentivirus particles and polybrene (5 μgml−1) (Day 1). After incubation for 8–12 h at 37 °C in a 5% CO2 incubator, the medium containing lentivirus was replaced with fresh regular medium, and then selected with puromycin, hygromycin B, or geneticin, depending on the lentiviral vectors (Day 3) for 5–7 days to eliminate the non-transduced cells.
Plasmid, small interfering RNA (siRNA), and short hairpin RNA (shRNA) construction
Constructs of pCMV-His-CK2α (no. HG10630-NH), pCMV-HA-CK2α (no. HG10630-CY), pCMV-Flag-CK2α (no. HG10630-CF), pCMV-His-CK2α‘ (no. HG11018-NH), pCMV-His-CK2β (no. HG13106-NH), pCMV-His-E4F1 (no. HG20515-CH) and pCMV-HA-NIP30 (no. HG16370-NY) were purchased from Sino Biological (Beijing, China). pCMV-Myc-CK2β was purchased from Addgene (no. #27088). pcDNA3.1-Flag-PA28γ WT, pcDNA3.1-Flag-PA28γ mutants (T23A, T23D, S24A, N151Y and K195R), and pcDNA3.1-Myc-E4F1 (aa1–784, aa1–357, aa357–784, and aa184–784) were custom synthesized by GENEWIZ (Azenta Life Sciences, Suzhou, China). The lentiviral packaged plasmids including pLenti-Flag-PA28γ generated by cloning the corresponding complementary DNAs into pLenti-hygro vector and pLenti-Myc-E4F1 generated by cloning the corresponding complementary DNAs into pLenti-neo vector, as well as the corresponding virus particles were custom generated by VectorBuilder. The sequences of all plasmids were verified by DNA sequencing.
siRNAs targeting human CK2α (siCK2α, 5′-GGTGAGGATAGCCAAGGTT-3′), CK2α‘ (siCK2α‘, 5′-GATCCACACTTCAACGATA-3′), CK2β (siCK2β #1, 5′-CACCCAA GTCATCAAGACA-3′; #2, 5′-AGAGCCCAGTCAAGACGAT-3′), NIP30 (siNIP30 #1, 5′-GUCUGAGGCAGAACUAGAUTT-3′; #2, 5′-CCCUCGAUCUCUAUAUGAATT-3′), PA28γ (siPA28γ #1, 5′-GACAGAGATTGATGAGAAA-3′; #2, 5′-GGAAACAGTTGCAGAGCTA-3′) and E4F1 (siE4F1 #1, 5′-TGAAGCTACTGGTGAACAA-3′; #2, 5′-GCAAGCGCTACAAGACTAA-3′) were custom synthesized by RIBOBIO (Guangzhou, China).
The lentiviral shRNAs against human endogenous PA28γ (shPA28γ 3′-untranslated region, 5′-ACACCCAACCTGTCGTAATTT-3′) and E4F1 (shE4F1 #1, 5′-TAGAGACTT CACCCGTGATTC-3′; #2, 5′-AGGACGTGGTTGTCAGCAAAG-3′), as well as the corresponding virus particles were custom generated by VectorBuilder.
Antibodies
The primary antibodies were diluted with TBST buffer containing 5% non-fat milk for immunoblot (IB) analysis, PBS buffer containing 3% BSA for immunofluorescence (IF), and antibody diluent (Abcam, ab64211) for IHC. Anti-α-tubulin (IB, 1:1000; #2125), anti-PA28γ (IB, 1:1000; IP, 1:100; #2412), anti-Myc (IB, 1:1000; #2278), anti-p21 (IB, 1:1000; #2947), anti-p53 (IB, 1:1000; #2527) and anti-GAPDH (IB, 1:1000; #5174) antibodies were purchased from Cell Signaling Technology (CST). Anti-Cyclin A2 (IB, 1:2000; IHC, 1:500; ab181591) antibody was purchased from Abcam. Anti-CK2β (IB, 1:300; IP, 1:50; IF, 1:50; sc-12739), anti-CK2α (IB, 1:200; sc-12738), anti-CK2α‘ (IB, 1:200; sc-514403), anti-HA (IB, 1:200; sc-7392) and anti-E4F1 (IB, 1:100; IP, 1:50; sc-514718) antibodies were purchased from Santa Cruz Biotechnology. Anti-Flag antibody (IB, 1:2000; F1804, clone M2) was purchased from Sigma. Anti-His (IB, 1:10000; IP, 1:300; 66005-1-Ig), anti-GST (IB, 1:10000; 66001-2-Ig), anti-NIP30 (IB, 1:1000; 16830-1-AP), anti-SRC3 (IB, 1:3000; 29587-1-AP), anti-SirT1 (IB, 1:3000; 13161-1-AP), anti-pan-keratin (pan-K) (IHC, 1:3000; 26411-1-AP), anti-Ki67 (IHC, 1:5000; 27309-1-AP) and anti-CK2β (IHC, 1:200; 20234-1-AP) antibodies were purchased from Proteintech. Anti-E4F1 antibody (IHC, 1:100; bs-9724R) was purchased from Bioss (Beijing, China). Anti-PA28γ antibody (IF, 1:400; IHC, 1:400; PA5-21789) was purchased from Thermo Scientific. Anti-phospho-Ser/Thr/Tyr antibody (IB, 1:50; NB600-558) was purchased from Novus Biologicals. Anti-Flag agarose beads (651503) were purchased from BioLegend. The anti-PA28γ-pT23 antibody (IB, 1:1000; IHC, 1:500) derived from rabbits was generated by PTM Biolab (Hangzhou, China). The PA28γ aa18–29 (FRERI-T-SEAEDC) was the antigen sequence used for immunization.
Immunoblot and immunoprecipitation analyses
Proteins were extracted from cells using RIPA buffer (50 mM Tris (pH 7.4), 150 mM NaCl, 1% NP-40, 0.5% sodium deoxycholate and 0.1% SDS) or NP-40 buffer (50 mM Tris (pH 7.4), 150 mM NaCl and 1% NP-40) with protease inhibitor cocktail (Sigma, P8340) and phosphatase inhibitor cocktail (MedChemExpress, HY-K0021). The cell lysates were clarified by centrifugation (12000 g, 15 min, 4 °C), followed by protein quantification assays with the BCA protein assay kit (Thermo Scientific, 23227) and the Varioskan LUX multimode microplate reader (Thermo Scientific, VLB000GD1). For immunoblot analysis, WCLs with 1×SDS–PAGE sample loading buffer were heated at 95 °C for 5 min. The denatured proteins were then resolved by loading equal amounts of WCLs to the SDS-PAGE gels, running the gels, transferring the proteins from the gels to PVDF membranes, blocking the membranes with TBST buffer containing 5% non-fat milk, incubating the membranes with the corresponding primary and secondary antibodies, and scanning images with Amersham Imager 600. For immunoprecipitation analysis, cell lysates were incubated with anti-Flag agarose beads (4 h, 4 °C) or the corresponding antibody (4 h to overnight, 4 °C) followed by 1.5 h incubation with protein A/G agarose (Santa Cruz, sc-2003) after protein quantification. The beads were washed three times with 1×NETN buffer (20 mM Tris (pH 8.0), 150 mM NaCl, 0.5% NP-40 and 1 mM EDTA) to remove non-specific binding, followed by immunoblot analysis. The immunoblot band intensities were analyzed by ImageJ v1.48 and normalized to α-tubulin.
Peptide synthesis and dot blot assays
The PA28γ peptides used for dot blot assays were synthesized by PTM Biolab (Hangzhou, China). The sequences are listed below (the asterisk denotes phosphorylation):
unphosphorylated peptide (unphos, aa16–29): DSFRERITSEAEDC;
T23-phosphorylated peptide A (pT23, aa16–29): DSFRERIT*SEAEDC;
T23-phosphorylated peptide B (pT23, aa18–29): FRERIT*SEAEDC (for antibody generation);
S24-phosphorylated peptide (pS24, aa16–29): DSFRERITS*EAEDC;
T23/S24-phosphorylated peptide (pT23/S24, aa16–29): DSFRERIT*S*EAEDC.
For the dot blot assays, the uncrosslinked antigen peptides were spotted onto the PVDF membrane in gradients of 1 ng, 4 ng, 16 ng, and 64 ng. After drying at room temperature, the membrane was blocked with TBST containing 5% non-fat milk for 60 min and washed with 1×TBST. Primary antibody incubation was performed using a 5% non-fat milk dilution, followed by 2 h at room temperature and three washes with 1×TBST. In competitive inhibition assays, 2.5% BSA was used as a diluent, with the antibody and peptide mixed at a predetermined ratio and incubated overnight at 4 °C. The antibody-peptide mixture was applied to the membrane and incubated at room temperature for 2 h, followed by washing with 1×TBST. The corresponding rabbit secondary antibody was then incubated at room temperature for 1 h and subsequently washed three times with 1×TBST. Finally, the membrane was treated with substrate for detection and exposed for imaging.
Purification of His-tagged PA28γ proteins from bacteria
The plasmid containing the PA28γ gene (vector pET28a) was transformed into BL21(DE3) competent cells, which were then evenly spread onto LB agar plates and incubated overnight at 37 °C. The following day, a single colony was picked from the plate and inoculated into 4 mL of LB medium. The culture was grown to an optical density of 0.5-0.8, at which point isopropyl-β-D-thiogalactoside (IPTG) was added to a final concentration of 0.2 mM to induce protein expression at 16 °C. For larger scale cultures, the cells were grown to an optical density of 0.8, induced again with 0.2 mM IPTG, and incubated at 15 °C for 16 h. Cells were then harvested for protein purification. The cell pellet was resuspended in a buffer containing 20 mM Tris (pH 8.0), 300 mM NaCl, 20 mM Imidazole with 1% Triton X-100, 1 mM DTT, and 1 mM PMSF, then lysed by sonication. The lysate was centrifuged at 12000 g for 15 min at 15 °C to remove insoluble proteins and cell debris. Concurrently, a Ni-IDA affinity chromatography column was equilibrated with a buffer of 20 mM Tris (pH 8.0), 300 mM NaCl, and 20 mM Imidazole. The supernatant was loaded onto the equilibrated column and incubated on a shaker at 4 °C for 90 min. The flow-through was collected and sampled for analysis. The column was washed with the equilibration buffer until no impurities were detected. Finally, the target protein was eluted using buffers with increasing concentrations of imidazole, and the eluates were collected. Each fraction was analyzed by Coomassie blue staining, and fractions with high purity were pooled. These fractions were dialyzed against 1×PBS (pH 7.4) containing 5% glycerol, filtered through a 0.22 μm filter, aliquoted, and stored at −80 °C.
In vitro kinase assays
The bacterially purified His-PA28γ WT and T23A proteins (0.5 μg) from pET28a-His-PA28γ WT and T23A constructs prepared by elution using His-tag Ni-IDA resins were incubated with 1 μl of casein kinase II (New England Biolabs, P6010) and 200 μM ATP (CST, #9804) in the absence or presence of TBB with the indicated concentrations in 30 μl kinase buffer (25 mM Tris-HCl (pH 7.5), 5 mM β-glycerophosphate, 2 mM dithiothreitol, 0.1 mM Na3VO4 and 10 mM MgCl2) (CST, #9802) for 30 min at 30 °C. The reaction was subsequently terminated by adding SDS–PAGE sample loading buffer and heating at 95 °C for 5 min, followed by immunoblot analysis with the indicated antibodies.
GST and His pull-down assays
For in vitro interaction, the GST-E4F1 protein (Novus Biologicals, H00001877-P01) or bacterially purified His-PA28γ protein was incubated with corresponding cell lysates containing prey proteins together with resin in an interaction buffer. GST and His pull-down assays were performed using the GST protein interaction kit (Thermo Scientific, 21516) and His protein interaction kit (Thermo Scientific, 21277) following the manufacturer’s instructions, respectively. The bound proteins were boiled for 5 min in 1 × SDS–PAGE sample loading buffer and subjected to immunoblot analysis.
In vitro proteolytic assays
Recombinant human E4F1 protein (H00001877-P01) and recombinant human p21/CDKN1A protein (NBP2-22976) were purchased from Novus Biologicals. The human 20S proteasome protein (E-360) was purchased from R&D Systems. The PA28γ heptamers used herein were obtained from MerryBio (Nanjing, China). For vitro proteolytic assays, 50 ng of the protein substrate E4F1 or p21/CDKN1A was incubated with or without 20S proteasome (0.25 μg) and PA28γ heptamers (1 μg) in the absence or presence of epoxomicin (100 nM) in 50 μl proteolytic reaction buffer (20 mM HEPES (pH 7.5), 100 mM KCl, 0.5 mM EDTA and 10% glycerol) for 45 min at 30 °C4,10. The reaction was subsequently analyzed by immunoblot analysis with the indicated antibodies.
Immunofluorescence
Cells seeded into glass coverslips were washed three times with PBS and then fixed with 4% paraformaldehyde for 20 min at room temperature. After three washes with PBS, the cells were permeabilized with 0.5% Triton X-100 for 8 min at room temperature. After three washes with PBS, the cells were blocked with 3% BSA for 1 h and then incubated with the indicated antibodies overnight at 4 °C. The next day, after three washes with PBS, the cells were incubated with the secondary antibodies—anti-mouse Alexa Fluor 488 (1:400 in 3% BSA; Abcam, ab150113) or anti-rabbit Alexa Fluor 647 (1:400 in 3% BSA; Abcam, ab150083) and TRITC-F-actin antibody (100 nM; Yeasen, 40734ES75) for 1 h at room temperature in the dark conditions. After three washes with PBS, the coverslips were mounted onto glass slides with antifade mounting medium with DAPI (Vector Laboratories, H-1200-10). Fluorescence images were visualized with Olympus FV3000RS confocal laser scanning microscope and processed using the Olympus FV31S-SW v2.6 software.
Identification of PA28γ phosphorylation in HSC-3 cells
Flag immunoprecipitations were conducted using WCLs from HSC-3 cells stably expressing Flag-PA28γ to confirm PA28γ phosphorylation. The Flag immunoprecipitates were resolved by SDS–PAGE and subsequently stained with Coomassie blue. Gel pieces containing the Flag-PA28γ protein were excised for further analysis (no replicates; preliminary exploration). For in-gel tryptic digestion, gel pieces were destained in 50 mM NH4HCO3 containing 50% acetonitrile (v/v) until the gel pieces became transparent. Gel pieces were dehydrated with 100 µl of 100% acetonitrile for 5 min. The liquid was then removed, and the gel pieces were rehydrated in 10 mM dithiothreitol and incubated at 56 °C for 60 min. The gel pieces were again dehydrated with 100% acetonitrile, after which the liquid was removed, and the gel pieces were rehydrated with 55 mM iodoacetamide. Samples were incubated in the dark at room temperature for 45 min. Gel pieces were washed with 50 mM NH4HCO3 and then dehydrated with 100% acetonitrile. Gel pieces were rehydrated with 10 ng/μl of trypsin resuspended in 50 mM NH4HCO3 and kept on ice for 1 h. Excess liquid was removed, and the gel pieces were digested with trypsin at 37 °C overnight. Peptides were extracted with 50% acetonitrile/5% formic acid, followed by extraction with 100% acetonitrile. The peptides were dried completely and then resuspended in 2% acetonitrile/0.1% formic acid.
The tryptic peptides were dissolved in 0.1% formic acid (solvent A), directly loaded onto a homemade reversed-phase analytical column (15 cm length, 75 μm i.d.). The gradient was comprised of an increase from 6% to 23% solvent B (0.1% formic acid in 98% acetonitrile) over 16 min, 23% to 35% in 8 min and climbing to 80% in 3 min then holding at 80% for the last 3 min, all at a constant flow rate of 400 nl/min on an EASY-nLC 1000 UPLC system.
The peptides were subjected to NSI source followed by MS/MS in Q ExactiveTM Plus (Thermo) coupled online to the UPLC. The electrospray voltage applied was set at 2.0 kV. The m/z scan range for the full scan was 350 to 1800, with intact peptides detected in the Orbitrap at a resolution of 70,000. Peptides were then selected for MS/MS using NCE setting of 28, and the fragments were detected in the Orbitrap at a resolution of 17,500. A data-dependent procedure that alternated between one MS scan followed by 20 MS/MS scans with 15.0 s dynamic exclusion. Automatic gain control (AGC) was set to 5 × 104.
The resulting MS/MS data were processed using the MaxQuant search engine v.1.6.15.0. Tandem mass spectra were searched against the UniProt human protein database. Trypsin/P was specified as the cleavage enzyme, allowing for up to 2 missed cleavages. The mass error was set to 10ppm for precursor ions and 0.02 Da for fragment ions. Carbamidomethyl on Cys was specified as a static modification, while oxidation on Met was designated as a dynamic modification. Peptide confidence was set at high, and peptide ion score was set > 20.
Identification of potential interacting proteins of PA28γ
Flag immunoprecipitation was performed using whole cell lysates (WCLs) from HSC-3 cells stably expressing Flag-PA28γ to confirm potential interacting proteins. The immunoprecipitates were resolved by SDS–PAGE and visualized with Coomassie blue staining. Gel pieces containing the proteins were excised (no replicates; preliminary exploration), rinsed twice with ultrapure water, and treated with 50 μL of destain buffer (25 mM NH₄HCO₃, 50% ACN) for 5 min. After decolorization, the gel was washed three times with deionized water and allowed to sit overnight. The gel was dehydrated sequentially with dehydration buffer 1 (50% ACN) for 30 min and dehydration buffer 2 (100% ACN) for another 30 min before vacuum freeze-drying. Following freeze-drying, 50 μL of reduction buffer (10 mM DTT/25 mM NH₄HCO₃) was added and incubated at 55 °C for 1 h. Excess liquid was removed, and 50 μL of alkylation buffer (50 mM IAA/25 mM NH₄HCO₃) was added for 30 min in the dark. The gel was then treated with gel recovery buffer (25 mM NH₄HCO₃) for 10 min, followed by dehydration steps with both dehydration buffers. For digestion, 10 μL of digestion buffer (25 mM NH₄HCO₃) was added to swell the gel for 30 min. Then, 20 μL of trypsin (0.02 μg/μL in digestion buffer) was added, and the mixture was incubated at 37 °C for 16 h. The supernatant was collected, and 50 μL of peptide elution buffer (5% formic acid/67% ACN) was added to the remaining gel, incubated at 37 °C for 30 min, centrifuged, and combined with the supernatant. The solvent was evaporated, and the sample was prepared for desalting using a C18 column.
The dried peptide sample was re-dissolved in Nano-HPLC buffer A (0.1% formic acid in water), and the C18 column was activated with methanol and equilibrated with 100% buffer A. The peptide sample was loaded onto the column, centrifuged, and passed through the column again. Buffer A was used for washing, followed by elution with 40 μL of buffer B (80% acetonitrile, 0.1% formic acid) before evaporating the eluent.
Separation was performed using the EASY-nLC1200 Nano-HPLC system with buffer A and buffer B as described. The trap column (100 μm × 20 mm, RP-C18) was equilibrated with buffer A, and the sample was injected and adsorbed before separation on the analytical column (75 μm × 150 mm, RP-C18) at a flow rate of 300nL/min, with a 30-min blank solvent gradient for cleaning. The enzymatic digestion products were analyzed using a Q-Exactive Plus mass spectrometer (Thermo Scientific) over a 30-min duration. The parent ion scan range was set from 300 to 1800m/z in data-dependent acquisition (DDA) mode, collecting the top 10 fragment spectra (MS2) after each full scan. Fragmentation was performed using high-energy collision dissociation (HCD) with a normalized collision energy (NCE) of 27 and a dynamic exclusion time of 40 s. MS1 resolution was set to 70,000 with an AGC target of 3 × 106 and a maximum injection time of 50 ms; MS2 resolution was set to 17,500 with an AGC target of 5 × 105 and a maximum injection time of 50 ms.
The resulting MS/MS data were processed using Proteome Discover 2.3. Tandem mass spectra were searched against the UniProt human protein database. Trypsin/P was specified as the cleavage enzyme, allowing for up to 2 missed cleavages. The mass error was set to 10ppm for precursor ions and 0.02 Da for fragment ions. Carbamidomethyl on Cys was specified as a static modification, and acetylation on protein N-terminus, deamidation on Asn and Gln, and oxidation on Met were specified as dynamic modifications. Peptide confidence was set at high, and peptide ion score was set > 20.
Tandem Mass Tags (TMT)-based quantitative proteomic analyses
PA28γ WT and PA28γ-T23A-rescued cells (both groups with three biological replicates) were harvested and lysed in lysis buffer (8 M urea, 1% protease inhibitor cocktail) using an ultrasonic processor. Equal amounts of protein from each sample were taken for digestion. The protein samples were incubated with 20% trichloroacetic acid for 2 h at 4 °C. The precipitates obtained by centrifugation were washed three times with acetone and then sonicated in 200 mM tetraethylammonium bromide (TEAB), followed by the addition of trypsin (MS grade, 1:50 trypsin-to-protein mass ratio) for overnight digestion. The next day, the samples were reduced with 5 mM dithiothreitol for 30 min at 56 °C and alkylated with 11 mM iodoacetamide for 15 min at room temperature in the dark. The peptides were desalted using Strata X C18 (Phenomenex), followed by vacuum freeze drying.
The peptides were then dissolved with 0.5 M TEAB and labeled using a TMT 6-plex Kit (Thermo Scientific) following the manufacturer’s instructions. For sample labeling, we utilized different TMT tags to distinguish the samples. Specifically, the PA28γ WT group consisted of three biological replicates labeled with tags 126, 127, and 128, while the PA28γ-T23A group included three biological replicates labeled with tags 129, 130, and 131.
The sample was fractionated into 9 fractions by high pH reverse-phase HPLC using an Agilent 300Extend C18 column (5 μm particles, 4.6 mm ID, 10 × 250 mm length). The wavelength was set to 214 nm, and the column oven temperature was set to 35°C. The column was equilibrated for at least 30 min using 95% buffer A (2% ACN, pH 9.0 adjusted with ammonia) and 5% buffer B (98% ACN, pH 9.0 adjusted with ammonia). Applied the stepwise gradient method after the baseline became flat. Then, 1 ml buffer A was added to the peptide sample and vortex to dissolve it. The sample was centrifuged at 12000 g for 5 min and transferred to a new tube. It was centrifuged again to take the supernatant, and the sample was loaded onto the HPLC while initiating the sample separation method and starting the automatic collector. Samples were collected at 1 min/tube from the 11th to 64th tubes, totaling 54 tubes. Finally, the peptides were combined into 9 fractions and dried by vacuum centrifuging.
Subsequently, the peptides were analyzed by LC-MS/MS. The tryptic peptides (1 μg) were dissolved in solvent A and directly loaded onto a home-made reversed-phase analytical column (25 cm length, 100 μm i.d.). The mobile phase consisted of solvent A (0.1% formic acid, 2% acetonitrile/in water) and solvent B (0.1% formic acid, 90% acetonitrile/in water). Peptides were separated with the following gradient: 0–36 min, 7%–23% B; 36–52 min, 23%–32% B; 52–56 min, 32%–80% B; 56–60 min, 80% B, all at a constant flow rate of 500 nl/min on an EASY-nLC 1200 UPLC system (ThermoFisher Scientific). The separated peptides were analyzed in a Q Exactive HF-X with a nano-electrospray ion source. The electrospray voltage applied was 2100 V. Precursors and fragments were analyzed at the Orbitrap detector. The full MS scan resolution was set to 120000 for a scan range of 350-1600 m/z. The fragments were detected in the Orbitrap at a resolution of 30000, and the fixed first mass was set as 100.0 m/z. Up to 20 most abundant precursors were then selected for further MS/MS analyses with a 30.0 s dynamic exclusion. The HCD fragmentation was performed at a normalized collision energy (NCE) of 28%. The automatic gain control (AGC) target was set at 1 × 105, with an intensity threshold of 8.3 × 104 ions/s and a maximum injection time of 60 ms.
Finally, the resulting MS/MS data were processed using the MaxQuant search engine v.1.6.15.0. Tandem mass spectra were searched against the human SwissProt database (20387 entries) concatenated with a reverse decoy database. Trypsin/P was specified as the cleavage enzyme allowing up to 2 missing cleavages. The mass tolerance for precursor ions was set to 20ppm in the First search and 4.5ppm in the Main search, and the mass tolerance for fragment ions was set to 20ppm. Carbamidomethyl on Cys was specified as fixed modification. Acetylation on protein N-terminal, oxidation on Met, and deamidation (NQ) were specified as variable modifications. TMT 6-plex quantification was conducted. FDR was adjusted to <1%, and the minimum score for peptides was set to > 40.
To select proteins with altered abundance, we calculated the average relative quantification values for the two groups of samples and subsequently determined the fold change (FC) of the proteins based on these values. We then performed a t-test on the FC to obtain P values, with a significance threshold set at P < 0.05. To meet the assumption of normality, the data were log2-transformed. Finally, proteins were classified as significantly upregulated if FC > 1.2 and downregulated if FC < 1/1.2, both requiring P < 0.05 for significance.
Quantitative real-time PCR
Total RNA was extracted using TRIzol Reagent (Invitrogen) and cDNA synthesis was performed using the PrimeScript RT reagent kit (Takara, RR037) following the manufacturer’s instructions. Quantitative real-time PCR analysis was performed using the SYBR Select Master Mix (Applied Biosystems, 4472908) with the QuantStudio 6 Flex Real-Time PCR Systems (Applied Biosystems). Data were collected using QuantStudio Design & Analysis Software v1.5.1. Relative expression of the mRNA was normalized to a control gene (GAPDH). Data represent mean ± SD from three independent experiments.
The following primer pairs were used for quantitative real-time PCR: E4F1, 5′-AGGATGGCCGCTATGTGTG-3′ (forward) and 5′-GAGCTTGCACTCGTGGTCC-3′ (reverse); PA28γ, 5′-AAGGTTGATTCTTTCAGGGAGC-3′ (forward) and 5′-AGTGGATCTGAGTTAGGTCATGG-3′ (reverse); Cyclin A2, 5′-CGCTGGCGGTACTGAAGTC-3′ (forward) and 5′-GAGGAACGGTGACATGCTCAT-3′ (reverse); DLAT, 5′-CCGCCGCTATTACAGTCTTCC-3′ (forward) and 5′-CTCTGCAATTAGGTCACCTTCAT-3′ (reverse); DNAJC19, 5′-AGTGGTAGCAGTTGGACTGAC-3′ (forward) and 5′-GGCAGATTTTGGTAGGCTTTGAA-3′ (reverse); CHK1, 5′-CCAGATGCTCAGAGATTCTTCCA-3′ (forward) and 5′-TGTTCAACAAACGCTCACGATTA-3′ (reverse); TTI2, 5′-GGCAGGTAGCAAAAGCCCT-3′ (forward) and 5′-TCTTAGCAGTCTCAACTTTCCCT-3′ (reverse); GAPDH, 5′-GAGTCAACGGATTTGGTCGT-3′ (forward) and 5′-TTGATTTTGGAGGGATCTCG-3′ (reverse).
Cell proliferation and colony formation assays
For cell proliferation assays, cells seeded into a 96-well plate were tested by cell counting kit-8 (Sigma, 96992) for four days following the manufacturer’s instructions. The absorbance at 450 nm was measured using a microplate reader (Thermo Scientific). Data were collected using SkanIt Software RE for Microplate Readers v7.1. For colony formation assays, cells were seeded into 6-well plates (800 cells per well) and cultured for two weeks until the formation of visible colonies. Cells were fixed with 4% paraformaldehyde for 20 min and then stained with 0.5% crystal violet solution for 10–20 min. After the plates were washed and air-dried, the colonies were photographed and counted. Data represent mean ± SD from three independent experiments.
Propidium iodide (PI) staining and flow cytometry analyses
For cell cycle analyses, 1 × 106 cells were harvested, fixed with cold 70% ethanol for 12 h at 4 °C, and then incubated with PI and RNase A for 1 h at room temperature in the dark. Cells were then determined by Attune NxT flow cytometer (Invitrogen). Data were collected using Attune NxT Software v2.4 and analyzed with FlowJo v10.6.2. Results are presented as mean ± SD from three independent experiments.
Mouse xenograft assays
The indicated HSC-3 cells (1 × 106) were suspended in DMEM medium mixed with matrigel (Corning) and then subcutaneously injected into the flank of female nude mice (six mice per group; 5–6 weeks of age; BALB/c-nu mice purchased from GemPharmatech Co., Ltd, China). The tumor size was measured every three days with a digital caliper and the tumor volume was determined with the following formula: L × W2× 0.5 (L, the longest diameter; W, the shortest diameter). The mice were sacrificed at the end of the studies and the xenografted solid tumors were dissected, weighed, and photographed.
Generation of PA28γT23A or PA28γT23D knock-in mice
The C57BL/6 J mouse models with the point mutation (PA28γT23A or PA28γT23D) at the PA28γ gene locus created by CRISPR/Cas-mediated genome engineering were obtained from Cyagen Biosciences. The PA28γT23A (ACA to GCA) or PA28γT23D (ACA to GAC) in donor oligo was introduced into the exon 2 of the PA28γ gene (the location of T23) by homology-directed repair. The synonymous mutation PA28γ-R21 (CGG to CGC) was also introduced to prevent the binding and re-cutting of the sequence by sgRNA after homology-directed repair. Cas9, sgRNA (sgRNA-A1, CAGAGAGCGGATCACAAGTG-AGG and sgRNA-B1, AGGTTGATTCTTTCAGA GAG-CGG) and donor oligo (PA28γT23A, CATGCCCTTGTATCACTAGGTGTTAAGTTGCTCTCTCTGTTCATTGTAGGTTGATTCTTTCAGAGAGCGCATCGCAAGTGAGGTGAGTGTTGTAAAGGAGAAAAGTGGTGATGGTTAATAGCTGGGGGTGTCCCCA; PA28γT23D, CATGCCCTTGTATCACTAGGTGTTAAGTTGCTCTCTCTGTTCATTGTAGGTTGATTCTTTCAGAGAGCGCATCGACAGTGAGGTGAGTGTTGTAAAGGAGAAAAGTGGTGATGGTTAATAGCTGGGGGTGTCCCCA) were co-injected into fertilized eggs for knock-in mouse production.
The pups were genotyped by PCR with the specific primers (forward, 5′-GTCTGCTTATGAAATTGAGAGG-3′; reverse, 5′-ATGTCTGAGTGGATCTGAGTTAG-3′), followed by sequence analysis. The validated heterogeneous knock-in mice (termed founder mice) were backcrossed with WT C57BL/6 J mice (Jackson Laboratories) for a minimum of two generations. Then the heterogeneous knock-in offspring were crossed with each other to obtain WT and T23 knock-in homogenous mice for subsequent experiments.
4-Nitroquinoline N-oxide (4NQO)-induced HNSCC and evaluation
Treatment of the mice with 4NQO is known to cause HNSCC in the oral cavity45. The WT, PA28γT23A, and PA28γT23D knock-in mice (7–8 weeks old; n = 16/group, comprising 8 females and 8 males) were fed with water containing 4NQO (50μgml−1; Sigma, N8141) for 16 weeks (1–16 wk). White lesions were observed in the tongue of all mice in the 16th week. After 4NQO treatments were discontinued, tumors gradually appeared during 8 weeks of feeding with normal water, and then the mice were sacrificed and the tongues were dissected out at the 24th week, followed by hematoxylin-eosin (H&E) and IHC staining. The mouse weight and lesions of the tongue were monitored weekly. The number of cauliflower-like or protruding lesions was recorded, with volume calculations following the same method as mouse xenograft assays.
Patient materials of TMAs
The TMAs were obtained from West China Hospital of Stomatology, Sichuan University (China), including 81 paired HNSCC samples (tumors and adjacent tissues; 65 males and 16 females; age range 27–84 years; Supplementary Data 1), oral mucosal samples (normal tissues, premalignant lesions, and primary tumors; 35 males, 13 females, and 23 unknown; age range 31–77 years; Supplementary Data 13), and 161 HNSCC samples (primary tumors, metastatic, and non-metastatic lymph node tissues; 117 males and 44 females; age range 24–88 years; Supplementary Data 14). TMAs of other cancer types, including pan-caner (13 cancer types, n = 91; Supplementary Data 3), rectal cancer (n = 90; 58 males and 32 females; age range 31–90 years; Supplementary Data 4), pancreatic cancer (n = 89; 48 males and 41 females; age range 39–83 years; Supplementary Data 5), and breast cancer (n = 130; 130 females; age range 29–87 years; Supplementary Data 6) were obtained from the Shanghai Outdo Biotech Company (HOrgC180PG01-1, HRec-Ade180Sur-03, HPanA180Su10, and HBreD130Su09).
IHC staining and evaluation
IHC was performed using 4-μm-thick, formalin-fixed paraffin-embedded (FFPE) tissue sections. FFPE tissue sections were deparaffinized with xylene and rehydrated in graded ethanol (100%, 95%, 90%, 80%, and 75%). Sections were heated for 15–20 min at 95 °C with 10 mM sodium citrate buffer (pH 6.0) in a microwave oven for antigen retrieval. To block endogenous peroxidase activity and non-specific antigens, tissue sections were incubated with 3% hydrogen peroxide for 20 min at room temperature and 5% BSA for 30 min at 37 °C. Sections were then incubated with the indicated primary antibody at 4 °C overnight. After three washes with PBS, sections were incubated with biotin-labeled goat anti-mouse/rabbit IgG and streptavidin-biotin-complex (SA1020, Bosterbio, Wuhan, China,) at 37 °C for 30 min each. Sections were visualized by the DAB detection kit (GK600505, Gene Tech, Shanghai, China) and lightly counterstained with hematoxylin.
For multiplex IHC staining, several proteins in tissues were stained using the multiple fluorescence IHC kit (Panovue, 10082100100) following the manufacturer’s instructions.
The IHC signals were reviewed and scored by two independent experienced pathologists in a blinded manner. For mouse tissues and xenografted solid tumors, five high-power fields were randomly selected to compare staining intensity. The scores of human TMAs were evaluated based on the antibody staining intensity (negative, 0; weak, 1; moderate, 2; strong, 3) and the extensity (the percentage of positive cells: ≤ 25%, 1; 25–50%, 2; 50–75%, 3; > 75%, 4). The combined scores (intensity × extensity) were calculated for subsequent analysis. For the survival analysis of TMA cohorts, the expression levels were classified as low and high (score: ≤ 6 and > 6). The overall survival duration was defined as the time from the date of surgery to the date of death or the date on which the patient was last known to be alive.
Statistics and reproducibility
Statistical analyses were performed using GraphPad Prism v8.3.0. All the mean ± SD and the calculated P values are derived from repeated biologically independent experiments or samples. Statistical significance was calculated by a two-tailed Student’s t-test between two groups or one-way ANOVA with Tukey’s multiple comparison test among three groups. Values of P < 0.05 were considered statistically significant. Survival analyses included univariate analysis using the Kaplan-Meier method with log-rank test, and multivariate analysis using the Cox proportional hazards model. Pearson’s correlation test was used to evaluate the correlation of tissue protein expressions. The immunoblot, immunoprecipitation, immunofluorescence, quantitative real-time PCR, cell proliferation, colony formation, and cell cycle assays, along with the mIHC or HE staining of the same mouse tissues, were each independently repeated three times, and collectively, similar results were obtained. Due to the difficulty in obtaining TMAs, the IHC or mIHC staining results of TMAs were statistically analyzed based on a single staining outcome.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The LC-MS/MS data generated in this study have been deposited in the ProteomeXchange Consortium via the iProX partner repository with the dataset identifiers PXD051881 (https://www.iprox.cn/page/project.html?id=IPX0008726000), corresponding to Fig. 1a), PXD044901 (https://www.iprox.cn/page/project.html?id=IPX0006991000), corresponding to Supplementary Data 9) and PXD051872 (https://www.iprox.cn/page/project.html?id=IPX0008719000), corresponding to Supplementary Data 10). Survival analysis of differentially expressed genes from TMT-based quantitative proteomic analysis was conducted using TCGA data, with univariate and multivariate analyses performed on the Kaplan-Meier Plotter (http://kmplot.com/analysis/) and TIMER2.0 (http://timer.cistrome.org/) platforms, respectively (corresponding to Supplementary Data 11). The remaining data are available within the article, Supplementary Information or Source Data file. Source data are provided with this paper.
References
Tojo, T. et al. Precipitating antibody to a soluble nuclear antigen “Ki” with specificity for systemic lupus erythematosus. Ryumachi 21, 129–140 (1981).
Ma, C. P., Slaughter, C. A. & DeMartino, G. N. Identification, purification, and characterization of a protein activator (PA28) of the 20 S proteasome (macropain). J. Biol. Chem. 267, 10515–10523 (1992).
Li, X. et al. The SRC-3/AIB1 coactivator is degraded in a ubiquitin- and ATP-independent manner by the REGgamma proteasome. Cell 124, 381–392 (2006).
Li, X. et al. Ubiquitin- and ATP-independent proteolytic turnover of p21 by the REGgamma-proteasome pathway. Mol. Cell 26, 831–842 (2007).
Chen, X., Barton, L. F., Chi, Y., Clurman, B. E. & Roberts, J. M. Ubiquitin-independent degradation of cell-cycle inhibitors by the REGgamma proteasome. Mol. Cell 26, 843–852 (2007).
Li, L. et al. REGγ is critical for skin carcinogenesis by modulating the Wnt/β-catenin pathway. Nat. Commun. 6, 6875 (2015).
Ying, H. et al. Aberrant accumulation of PTTG1 induced by a mutated thyroid hormone beta receptor inhibits mitotic progression. J. Clin. Invest 116, 2972–2984 (2006).
Dong, S. et al. The REGγ proteasome regulates hepatic lipid metabolism through inhibition of autophagy. Cell Metab. 18, 380–391 (2013).
Li, S. et al. Regulation of c-Myc protein stability by proteasome activator REGγ. Cell Death Differ. 22, 1000–1011 (2015).
Xu, J. et al. The REGγ-proteasome forms a regulatory circuit with IκBɛ and NFκB in experimental colitis. Nat. Commun. 7, 10761 (2016).
Johnson, D. E. et al. Head and neck squamous cell carcinoma. Nat. Rev. Dis. Prim. 6, 92 (2020).
Li, J. et al. Associations between proteasomal activator PA28γ and outcome of oral squamous cell carcinoma: evidence from cohort studies and functional analyses. EBioMedicine 2, 851–858 (2015).
Liu, S. et al. PA28γ acts as a dual regulator of IL-6 and CCL2 and contributes to tumor angiogenesis in oral squamous cell carcinoma. Cancer Lett. 428, 192–200 (2018).
Lei, K. et al. PA28γ, an Accomplice to Malignant Cancer. Front Oncol. 10, 584778 (2020).
Funderburk, K. E., Kang, J. & Li, H. J. Regulation of life & death by REGγ. Cells 11, 2281 (2022).
Singh, V. et al. Phosphorylation: implications in cancer. Protein J. 36, 1–6 (2017).
Geffen, Y. et al. Pan-cancer analysis of post-translational modifications reveals shared patterns of protein regulation. Cell 186, 3945–3967.e3926 (2023).
Magni, M. et al. Chk2 and REGγ-dependent DBC1 regulation in DNA damage induced apoptosis. Nucleic Acids Res. 42, 13150–13160 (2014).
Venerando, A., Ruzzene, M. & Pinna, L. A. Casein kinase: the triple meaning of a misnomer. Biochem J. 460, 141–156 (2014).
Pinna, L. A. Protein kinase CK2: a challenge to canons. J. Cell Sci. 115, 3873–3878 (2002).
Borgo, C., D’Amore, C., Sarno, S., Salvi, M. & Ruzzene, M. Protein kinase CK2: a potential therapeutic target for diverse human diseases. Signal Transduct. Target Ther. 6, 183 (2021).
Sun, S., Zhong, B., Zeng, X., Li, J. & Chen, Q. Transcription factor E4F1 as a regulator of cell life and disease progression. Sci. Adv. 9, eadh1991 (2023).
Murata, S. et al. Growth retardation in mice lacking the proteasome activator PA28gamma. J. Biol. Chem. 274, 38211–38215 (1999).
Meggio, F. & Pinna, L. A. One-thousand-and-one substrates of protein kinase CK2? FASEB J. 17, 349–368 (2003).
Sommercorn, J., Mulligan, J. A., Lozeman, F. J. & Krebs, E. G. Activation of casein kinase II in response to insulin and to epidermal growth factor. Proc. Natl. Acad. Sci. USA 84, 8834–8838 (1987).
Ackerman, P. & Osheroff, N. Regulation of casein kinase II activity by epidermal growth factor in human A-431 carcinoma cells. J. Biol. Chem. 264, 11958–11965 (1989).
Ji, H. et al. EGF-induced ERK activation promotes CK2-mediated disassociation of alpha-Catenin from beta-Catenin and transactivation of beta-Catenin. Mol. Cell 36, 547–559 (2009).
Cozza, G., Pinna, L. A. & Moro, S. Protein kinase CK2 inhibitors: a patent review. Expert Opin. Ther. Pat. 22, 1081–1097 (2012).
Siddiqui-Jain, A. et al. CX-4945, an orally bioavailable selective inhibitor of protein kinase CK2, inhibits prosurvival and angiogenic signaling and exhibits antitumor efficacy. Cancer Res. 70, 10288–10298 (2010).
Jonik-Nowak, B. et al. PIP30/FAM192A is a novel regulator of the nuclear proteasome activator PA28γ. Proc. Natl. Acad. Sci. USA 115, E6477–e6486 (2018).
Gao, X. et al. The REGγ inhibitor NIP30 increases sensitivity to chemotherapy in p53-deficient tumor cells. Nat. Commun. 11, 3904 (2020).
Rodier, G. et al. The transcription factor E4F1 coordinates CHK1-dependent checkpoint and mitochondrial functions. Cell Rep. 11, 220–233 (2015).
Lacroix, M. et al. E4F1 controls a transcriptional program essential for pyruvate dehydrogenase activity. Proc. Natl. Acad. Sci. USA 113, 10998–11003 (2016).
Zhang, Z. et al. Identification of an activation region in the proteasome activator. REGalpha. Proc. Natl. Acad. Sci. USA 95, 2807–2811 (1998).
Liu, J. et al. Site-specific acetylation of the proteasome activator REGγ directs its heptameric structure and functions. J. Biol. Chem. 288, 16567–16578 (2013).
Fernandes, E. R., Zhang, J. Y. & Rooney, R. J. Adenovirus E1A-regulated transcription factor p120E4F inhibits cell growth and induces the stabilization of the cdk inhibitor p21WAF1. Mol. Cell Biol. 18, 459–467 (1998).
Fajas, L. et al. Cyclin A is a mediator of p120E4F-dependent cell cycle arrest in G1. Mol. Cell Biol. 21, 2956–2966 (2001).
Ali, A. et al. Differential regulation of the REGγ-proteasome pathway by p53/TGF-β signalling and mutant p53 in cancer cells. Nat. Commun. 4, 2667 (2013).
Wang, H., Yang, L., Liu, M. & Luo, J. Protein post-translational modifications in the regulation of cancer hallmarks. Cancer Gene Ther. 30, 529–547 (2023).
Kandoth, C. et al. Mutational landscape and significance across 12 major cancer types. Nature 502, 333–339 (2013).
Sandy, P. et al. p53 is involved in the p120E4F-mediated growth arrest. Oncogene 19, 188–199 (2000).
Fajas, L. et al. pRB binds to and modulates the transrepressing activity of the E1A-regulated transcription factor p120E4F. Proc. Natl. Acad. Sci. USA 97, 7738–7743 (2000).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Frayssinhes, J. A. et al. PA28gamma-20S proteasome is a proteolytic complex committed to degrade unfolded proteins. Cell Mol. Life Sci. 79, 45 (2021).
Bouaoud, J. et al. The 4-NQO mouse model: An update on a well-established in vivo model of oral carcinogenesis. Methods Cell Biol. 163, 197–229 (2021).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (82072999 to J.L., 82273320 to J.L., 82403113 to S.S., 82472686 to J.L., and 82401328 to B.Z.), the CAMS Innovation Fund for Medical Sciences (2019-I2 M-5-004 to Q.C.), the Natural Science Foundation of Sichuan Province (2023NSFSC1447 to B.Z.), the China Postdoctoral Science Foundation (2024M762252 to S.S.), and the Postdoctoral Fellowship Program of CPSF (GZB20230471 to S.S.).
Author information
Authors and Affiliations
Contributions
J.L. conceived and designed the study. S.S. and B.Z. performed most of the experiments and statistical analyses. Y.W. assisted with the multiplex IHC staining. Z.L. and Y.J. helped with in vitro assays. N.J. and Y.J. were involved in vivo studies. J.L. supervised the study. X.F. and R.L. provided scientific inputs. S.S. and B.Z. wrote the manuscript. J.L., H.Z., Q.C., C.L., X.Z. (Xikun Zhou), and X.Z. (Xin Zeng) reviewed and edited the manuscript. All authors commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Paolo Cascio, Jochen Hess and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sun, S., Zhong, B., Wang, Y. et al. Phosphorylation of PA28γ by CK2 kinase facilitates HNSCC tumor formation and progression. Nat Commun 17, 443 (2026). https://doi.org/10.1038/s41467-025-67131-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67131-7