Abstract
The mechanism underlying the establishment of human endometrial receptivity remains elusive, constituting a significant obstacle to advancing our knowledge of female infertility. Via integrating high-resolution spatiotemporal and single-cell transcriptomic profiling and in situ sequencing, we construct a spatiotemporal atlas of human endometrial receptivity at single-cell resolution. Our study depicts detailed spatial molecular topography governing the opening and closing of human endometrial receptivity. Notably, the results indicate that stromal-specific NR2F1, CEBPD and epithelial-specific SGK1, KLF5 and ELF3 are closely associated with the opening of implantation window. Furthermore, cholesterol metabolism is found to promote ciliogenesis. Remarkably, we identify stroma- and epithelium-specific factors and unravel a previously unappreciated role of FGFs-FGFR2 signaling pathway mediating stroma-epithelium crosstalk in epithelial differentiation during the onset of receptivity. This spatiotemporally resolved receptivity code provides insights into the female fertility, with potential implications for the intervention of female infertility.
Similar content being viewed by others
Introduction
Appropriate preparation of the human endometrium is a critical prerequisite for successful embryo implantation, whose deficiency contributes to female infertility and adverse pregnancy outcomes, affecting women worldwide at childbearing age. Despite significant advancements in assisted reproductive technology (ART), the incidence of embryo implantation failure remains high, ascribed to both seed (embryo) and soil (endometrium)1. While significant advances have been made to improve the development of the embryo, the development of a receptive endometrium has thus emerged as another important determinant of ART outcomes2.
As the site of embryo implantation and subsequent embryonic development in humans, the endometrium undergoes complex and dynamic alterations throughout the menstrual cycle, orchestrated by ovarian hormones3. The endometrial epithelium and stroma undergo profound proliferation after menstruation, contributing to their re-thickening under the influence of preovulatory estrogen. Following ovulation triggered by luteinizing hormone (LH), the endometrium transitions into the secretory phase, characterized by the expansion of glands and stromal edema, driven by elevated progesterone level4. The mid-secretory phase, occurring approximately 7–10 days post-LH surge, is considered a critical window for embryo implantation, known as the implantation window or endometrial receptivity5. In the absence of an implanted embryo, the endometrium transits into the refractory phase or late-secretory phase, in preparation for the subsequent menstruation. Therefore, elucidating the molecular mechanisms underlying the onset of receptivity is crucial for optimizing the timing of embryo transfer in in vitro fertilization and embryo transfer (IVF-ET).
Accompanied by the advancement of genetic models, several key regulators for endometrial receptivity have been identified, including leukemia inhibitory factor (Lif), signal transducer and activator of transcription 3 (Stat3), heart and neural crest derivatives expressed 2 (Hand2), and msh homeobox 1 (Msx1)6,7,8,9,10. However, the prediction of endometrial receptivity in humans remains contentious due to the lack of reliable markers specific to human endometrial receptivity11,12,13. Although single-cell RNA sequencing (scRNA-seq) has been applied to understand the gene signature of human endometrial receptivity in normal or Asherman’s Syndrome endometrium14,15 and spatial transcriptomics has been utilized to resolve the gene localization in proliferative and secretory endometrium using 10X Visium with 55 μm resolution16, the spatiotemporal code around the timing of implantation at high resolution remains to be untangled.
To comprehensively decrypt the spatiotemporal molecular and cellular landscapes of endometrial receptivity, we integrated spatiotemporal enhanced resolution transcriptomic sequencing (Stereo-seq, 500 nm resolution), high-throughput in situ sequencing (ISS), and scRNA-seq to decipher the black box of spatial transcriptomics of implantation. In this study, we constructed a spatial atlas of the human endometrium, spanning the transition from the opening to the closing of the implantation window. Our findings unravel that both stroma and epithelium undergo proliferation, differentiation, and degradation along specific pathways. Especially, we dissect out spatial and cell-type-specific markers for the opening of the implantation window, including stromal NR2F1, CEBPD, and epithelial SGK1. Notably, we find that cholesterol metabolism plays a critical role in ciliogenesis. Furthermore, fibroblast growth factors (FGFs) and interleukin 15 (IL-15), derived from the stroma, were found to be important for epithelial differentiation and the dynamic regulation of natural killer (NK) cells during the peri-implantation period, respectively. This study provides insights into the molecular mechanisms of endometrial receptivity establishment through the lens of spatial transcriptomics, providing valuable insights for refining therapeutic strategies in IVF-ET to combat human infertility.
Results
Single-cell spatial atlas of human endometrial receptivity
The human endometrium undergoes dynamic remodeling driven by fluctuating ovarian steroid hormones and is classically divided into proliferative (follicular) and secretory (luteal) phases within an average 28-day menstrual cycle5. A transient window of receptivity during the secretory phase is critical for embryo implantation and the establishment of pregnancy. However, precisely identifying this transient window remains challenging due to the difficulty in accurately determining the timing of ovulation, inter-individual differences in follicular dynamics, stochastic variation in hormonal responses, asynchronous follicle development rates, and so on. To address this, human chorionic gonadotropin (hCG) was administered to the preovulatory follicle to simulate natural cycles17. The endometrial biopsies were collected at 2 (LH2, pre-receptive phase), 7 and 9 (LH7 and LH9, receptive phase), and 12 days (LH12, refractory phase) after hCG injection. These samples were subjected to Stereo-seq and scRNA-seq to decode the spatial-temporal transcriptomic landscapes of receptivity at single-cell resolution (Fig. 1A, Supplementary Fig. 1A). Since progesterone receptor (PR) expression is reported to peak in the epithelium during the early secretory phase and diminish thereafter18,19, PR staining was employed to confirm the sample stages (Supplementary Fig. 1B).
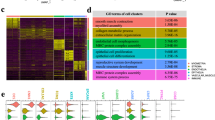
A Diagram illustrates the experimental workflow for single-cell transcriptome and spatial-transcriptome profiling in human endometrium across LH stages. B UMAP of the major cell types in human endometrium. Each dot represents a single cell, with distinct colors denoting individual cell populations (n = 2 per time point). C Violin plots showing marker genes in different cell types. D Line chart showing the percentage of different cell types over time. E Spatial visualization of all cell distributions within the human endometrium from LH2 to LH12. Cells are colored by their annotations (n = 1 per sample point). F Localization of major cell types within human endometrium, including stroma, epithelium, NK cells, and endothelium. Cells are colored by their annotations (n = 1 per sample point in Stereo-seq). G Multiplexed immunofluorescence staining of the main cell types of the endometrium (n = 2). Epithelial cells are identified by E-CADHERIN (yellow); stromal cells are marked by WT1 (red), HAND2 (hotpink) and PR (fuchsia); endothelial cells are labeled with CD31 (cyan); NK cells are marked by NCAM1 (blueviolet); T cells are identified by CD8 (bisque) and CD4 (darkorange); and pericytes are highlighted by MEF2C (darkturquoise). PCNA (light yellow) represents cell proliferation; C-CASPASE 3 (green) represents cell apoptosis. Nuclei are counterstained with DAPI (blue).
scRNA-seq was applied to interpret the single-cell transcriptomic atlas of human endometrium during the peri-receptive period. After quality control, 82,980 cells were retained for further analyses. The results of scRNA-seq revealed that the superficial endometrium consisted of a variety of cell types, including proliferating stroma (DNA topoisomerase II alpha, TOP2A), stroma (estrogen receptor 1, ESR1), endothelium (platelet and endothelial cell adhesion molecule 1, PECAM1), epithelium (paired box 8, PAX8), cilium (forkhead box J1, FOXJ1), pericyte (regulator of G protein signaling 5, RGS5), NK (granzyme B, GZMB), macrophage (CD14 molecule, CD14), and other lymphocyte (CD96 molecule, CD96) (Fig. 1B, C), which is aligned with previous works3,20 and listed in Supplementary Data 1. Among the major cell populations, the proportion of proliferating stromal cells decreased substantially, while NK cells significantly increased from LH2 to LH12 (Fig. 1D). After integrating our data with recently published scRNA-seq data covering LH3-LH1115, high concordance in major cell-type composition was noticed (Supplementary Fig. 1C–E).
To further depict the spatial distribution and dynamic organization of these distinct cell types around receptivity, Stereo-seq was applied to capture the spatial transcriptomic landscapes of the endometrium during the establishment of receptivity. To enhance the accuracy of spatial cell annotation, particularly for cell types with smaller populations and diffuse distributions, such as endothelial cells and immune cells, Stereo-seq was integrated with scRNA-seq to annotate cells in the endometrium. This combined approach mapped these cell types across 179,520 spatial spots in the endometrium with high precision (Fig. 1E, Supplementary Fig. 2A–C). Our spatial-temporal transcriptomic analysis revealed that stromal cells were the most abundant cell type. Consistent with scRNA-seq results, a notable increase in NK cells was observed at LH9 and LH12 (Fig. 1F). Meanwhile, multiplexed immunofluorescence (mIF) staining was performed to validate the major cell types in the endometrium (Fig. 1G). Together, these results provided a comprehensive depiction of the spatial distribution and dynamic changes in cell populations within the human endometrium during the peri-implantation window.
Molecular and spatial characterizations of stroma
Stromal cells represent the most abundant cell type in the endometrium, which are critical for the establishment of uterine receptivity and the maintenance of pregnancy. The localization of stromal cells was confirmed by the expression of HAND2 and platelet-derived growth factor receptor alpha (PDGFRα) (Supplementary Fig. 2D). Pericytes have been identified as a significant source of endometrial mesenchymal stem/stromal cells (MSCs), marked by high expression of sushi domain containing 2 (SUSD2), cluster of differentiation 146 (CD146), and platelet-derived growth factor receptor beta (PDGFRβ)21. In our study, the transcription factor myocyte enhancer factor 2 C (MEF2C) was specifically expressed in pericytes. Immunofluorescence co-localization analysis confirmed that MEF2C expression precisely overlapped with established pericyte markers NG2 and RGS5 (Supplementary Fig. 2E).
To better characterize the molecular heterogeneity within stromal cells, we re-clustered stroma population into four subclusters: proliferating stromal cells (proStr), marked by classical DNA replication genes, such as TOP2A and marker of proliferation Ki-67 (MKI67); undifferentiated stromal cells (undiffStr), enriched for Wnt pathway-related genes associated with maintaining undifferentiated state, including secreted frizzled related protein 4 (SFRP4) and patched 1 (PTCH1); differentiated stromal cells (diffStr), which highly expressed aldehyde dehydrogenase 1 family member A3 (ALDH1A3), nuclear receptor subfamily 2 group F member 1 (NR2F1); and degrading stromal cells (deStr), characterized by the expression of genes such as insulin-like growth factor binding protein 3 (IGFBP3), centrin-like protein (CTN4), and unc-51 like kinase 4 (ULK4) (Fig. 2A–C, Supplementary Fig. 3A). Cell cycle analysis of these subclusters revealed that proStr were predominantly enriched in the S and G2M phases (Fig. 2D). Immunofluorescence staining of WT1 transcription factor (WT1) and MKI67 confirmed significant proliferation in both the stroma and epithelium at LH2. Accompanied by elevated progesterone secreted from the corpus luteum after ovulation, the proliferation of both stroma and epithelium gradually ceased. However, cell proliferation continued in some WT1-negative cells, peaking at LH12, indicating the propagation of other cell types such as immune cells or endothelial cells (Fig. 2E). The subpopulations of stromal cells were further confirmed by integrating our data with recently published work (Supplementary Fig. 3B, C).
A UMAP visualization showing the re-clustering of stroma. proStr proliferating stromal cells, undiffStr undifferentiated stromal cells, diffStr differentiated stromal cells, deStr degrading stromal cells. B Heatmap showing the module correlations of stromal cell subclusters based on pairwise gene expression. C Expression of classical marker genes of stromal cell subclusters. D Cell cycle analysis of proStr, undiffStr, diffStr, and deStr in the endometrium. E Co-immunostaining of WT1 (green) and MKI67 (red) in human endometrium across the LH stages (n = 3). Nuclei are counterstained with DAPI (blue). F Stereo-seq visualization of different stroma subclusters within human endometrium. G Spatial expression of SFRP4, NR2F1, SERPING1, and THBS2 in human endometrium from LH2 to LH12 using Stereo-seq (bin20). H ISS of SFRP4, NR2F1, SERPING1, and THBS2 in human endometrium from LH2 to LH12 (n = 1).
To delineate the dynamic changes of stromal cells spatially, these subsets were mapped onto Stereo-seq data. Stromal cells at LH2 were primarily undiffStr and proStr, while the LH7 and LH9 endometrium were mainly characterized by diffStr, and deStr predominantly emerged at LH12 (Fig. 2F). Notably, SFRP4 was highly expressed at LH2, which was critical to maintain the undifferentiated status of stroma. Our Stereo-seq revealed that NR2F1 exhibited dynamic alterations during the natural cycle in human endometrium with prominent expression in stromal cells at LH7 and LH9, coinciding with the timing of embryo implantation. Interestingly, while NR2F2 exhibited high stromal expression levels comparable to NR2F1, its expression remained remarkably stable from LH2 to LH12 in both scRNA-seq and Stereo-seq analyses (Supplementary Fig. 3D, E). Serpin family G member 1 (SERPING1), a type of serine protease inhibitor involved in the regulation of the complement cascade22, was highly expressed from LH7 to LH12, while thrombospondin 2 (THBS2), a member of the thrombospondin family participating in negative regulation of angiogenesis, was predominantly expressed at LH12 (Fig. 2G). These results were further corroborated by high-throughput probe-based ISS (Fig. 2H, Supplementary Fig. 3F). Together, these studies revealed a detailed spatiotemporally dynamic code of distinct stromal cells across the phases of the endometrial cycle at high spatial resolution (Supplementary Fig. 4A).
The stromal determinant of receptivity establishment
To comprehensively elucidate the functional characteristics of the four stromal cell populations, genes specifically enriched in each population were subjected to the Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis. This analysis unraveled distinct functional hallmarks of each stromal subset: genes related to cell cycle regulation were predominantly enriched in proStr; undiffStr exhibited significant enrichment in the mTOR and Hedgehog signaling pathways; and diffStr were prominently characterized by enrichment in the extracellular matrix (ECM)-receptor interaction pathways (Fig. 3A, B, Supplementary Fig. 4B). Notably, deStr displayed marked enrichment of genes linked to autophagy and apoptosis pathways, such as B-cell lymphoma 2 (BCL2), growth arrest and DNA damage-inducible alpha and gamma (GADD45A and GADD45G), Sequestosome 1 (SQSTM1), along with canonical autophagy markers microtubule-associated protein 1 light chain 3 alpha and beta (MAP1LC3A and MAP1LC3B) and the apoptosis regulator myeloid cell leukemia 1 (MCL1) (Fig. 3A–C, Supplementary Fig. 4B). Differentially expressed gene (DEG) analysis between stromal cells elucidated that forkhead box O1 (FOXO1), a key gene critical for stromal cell differentiation and an early marker for stromal cell decidualization23, was obviously upregulated in diffStr compared to undiffStr (Fig. 3D, Supplementary Fig. 4C). Additionally, the augmented phospholipase A2 group IIA (PLA2G2A) and prostaglandin D2 synthase (PTGDS) in diffStr signified a potential role of stromal prostaglandin in receptivity opening (Fig. 3D). Meanwhile, there were substantial DEGs between diffStr and deStr, indicating that both the transitions from pre-receptivity to receptivity and from receptivity to the refractory state involved profound transformation of stromal cells (Fig. 3D, E). Furthermore, the comparison of stromal cells between LH2, LH7/9, and LH12 also manifested similar results. Cellular proliferation was the predominant feature of stromal cells at LH2. Upon transitioning to LH7/9, stromal cells exhibited marked upregulation of genes linked to the FOXO1 signaling pathway. Notably, as the process advanced to LH12, stromal cells displayed a distinct shift toward apoptosis, characterized by the elevated expression of cell death-associated genes (Supplementary Fig. 4D).
A The average gene expression patterns of stromal subclusters using scRNA-seq (left and middle), with corresponding representative KEGG signaling pathways (right). Data are shown as boxplots where midlines indicate medians, boxes indicate interquartile range, and whiskers indicate minimum/maximum range. ***P < 0.001. B Expression of specific representative genes in stromal subclusters from LH2 to LH12. The color bar represents the normalized expression of genes. C The network diagram showing genes in apoptosis and mitophagy. D Differentially expressed genes between undiffStr, diffStr, and deStr. E UpSet plots visualizing the number of differentially expressed genes among undiffStr, diffStr, and deStr. F Heatmap of expression and regulon activity of putative transcription factors. G Circular heatmap showing the representative transcription factors in different stromal subpopulations. H Spatial visualization of TCF4, CEBPD, and FOS in human endometrium across the LH stages using Stereo-seq (bin10). I UMAP visualization showing the IHH signaling pathway in stromal subclusters. J UMAP visualization showing the mitophagy signaling pathway in stromal subclusters. K UMAP visualization showing stromal trajectory. L Column diagram showing the trajectory score of stromal subclusters.
Given that diffStr encompassed stromal cells from both LH7 and LH9 stages, we also compared gene expression between these two stages. The number of DEGs between LH7 and LH9 diffStr was substantially smaller than in comparisons with DEGs between diffStr and other subtypes. Notable differences included lower expression of ArfGAP with GTPase domain, ankyrin repeat and PH domain 1 (AGAP1), Rho GTPase activating protein 28 (ARHGAP28) and thrombospondin type 1 domain containing 4 (THSD4) at LH7 and reduced expression of lipopolysaccharide binding protein (LBP), arachidonate 15-lipoxygenase type B (ALOX15B) and phospholipase A2 group IIA (PLA2G2A) at LH9 (Supplementary Fig. 5A–C).
To excavate the critical determinant of stroma transformation, regulon analysis was applied, leveraging Single-cell regulatory network inference and clustering (SCENIC). Both regulon activity and expression of the E2F family and forkhead box M1 (FOXM1) were highly enriched in stromal proliferation (Supplementary Fig. 5D), consistent with previous works24. Meanwhile, homeobox A10 (HOXA10) and transcription factor 4 (TCF4) were overtly decreased from proStr and undiffStr to diffStr (Fig. 3F), implying their roles in maintaining the undifferentiated state of stroma. HOXA10 was reported to be closely associated with endometrial receptivity25, while our results showed that although HOXA10 expression was high at LH2, it decreased markedly from LH7 onward in our Stereo-seq and scRNA-seq data (Supplementary Fig. 5E–I), consistent with previous reports showing peak expression in the proliferative and early secretory phases followed by a decline3. Interestingly, we unraveled the previously unappreciated role of CCAAT enhancer binding protein delta (CEBPD) in stroma differentiation (Fig. 3F, G). While CEBPB was consistently expressed and showed no dynamic change in both scRNA-seq and Stereo-seq compared with CEBPD (Supplementary Fig. 5J, K). The results of IHC confirmed the expression and dynamic changes of CEBPD in human endometrium (Supplementary Fig. 5L). The highly expressed and increased activation of AP1 family (JunD proto-oncogene, JUND; Jun proto-oncogene, JUN; Fos proto-oncogene, FOS) and X-box binding protein 1 (XBP1) at LH12 suggested their involvement in stromal degradation and the initiation of menstruation (Fig. 3G, Supplementary Fig. 5D). The dynamic spatial localization of TCF4, CEBPD, and FOS further supported the above speculations (Fig. 3H).
The signal pathway projection further confirmed that IHH and mitophagy were specifically enriched in undiffStr and diffStr, respectively (Fig. 3I, J). Trajectory analysis revealed that IHH-dominant undiffStr exhibited lower pseudotime scores, while deStr displayed the highest pseudotime scores, indicating the differential potential of stromal cells during transformation (Fig. 3K, L). This result is consistent with our previous work that the derailed IHH-PTCH pathway is important for the pathological causes of thin endometrium26. Intriguingly, insulin-like growth factor binding protein 1 (IGFBP1) and prolactin (PRL), two critical markers for stromal cell pre-decidualization and decidualization, only manifested minimal expression in terms of both cell ratio and gene level in stroma in later LH stages based on Stereo-seq and scRNA-seq (Supplementary Fig. 5M, N). Collectively, these results in the stroma provided a comprehensive spatial framework for understanding the molecular dynamics of stromal cells during receptivity onset.
Spatial identification and visualization of epithelium
Epithelium mediates the first direct interplay between the implanting embryo and the receptive endometrium. To extrapolate the diversity and dynamic roles of epithelial cells in endometrial receptivity establishment, we subclustered epithelial cells into seven distinct populations: (1) proliferating epithelial cells (proEpi) with high levels of proliferative genes such as MKI67 and TOP2A; (2) luminal epithelial cells (LE) expressing leucine rich repeat containing G protein-coupled receptor 5 (LGR5) and claudin 22 (CLDN22); (3) glandular epithelial cells (GLE) expressing hydroxysteroid 17-beta dehydrogenase 2 (HSD17B2) and laminin subunit beta 1 (LAMB1); (4) differentiated epithelial cells (diffEpi) expressing metallothionein 2A (MT2A) and metallothionein 1 M (MT1M); (5) degraded epithelial cells (deEpi) expressing DNA polymerase gamma 2 (POLG2); (6) senescent epithelial cells (seneEpi) expressing gremlin 2 (GREM2) and disks large MAGUK scaffold protein 2 (DLG2); and (7) Cilium (ciliated epithelial cells) characterized by expression of dynein axonemal heavy chain 12 (DNAH12), cadherin related family member 3 (CDHR3), and HYDIN axonemal central pair apparatus protein (HYDIN) (Fig. 4A, B, Supplementary Fig. 6A). The classification of epithelial subpopulations was further confirmed by projection of recently published data to our data (Supplementary Fig. 6B).
A UMAP visualization showing the re-clustering of epithelium. proEpi proliferating epithelial cells, GLE glandular epithelial cells, LE luminal epithelial cells, diffEpi differentiated epithelial cells, deEpi degraded epithelial cells, seneEpi senescent epithelial cells, Cilium ciliated epithelial cells. B Dot plot showing marker genes for different epithelial cells. The color bar represents the normalized expression of genes. The size of the plot represents the percentage of expressed cells. C Heatmap showing the specific genes in each epithelial subcluster. D Functional enrichment analysis of specific genes across epithelial subclusters. E Co-immunostaining of E-Cadherin (red) and acetylated Tubulin (green) in human endometrium at distinct LH phases (n = 3). Nuclei are counterstained with DAPI (blue). F Spatial visualization displaying the distribution pattern of epithelial subclusters. G The proportion of epithelial subclusters in the human endometrium across the LH stages. H The spatial distribution of CITED4 and DPP4 in human endometrium across the LH stages by ISS. I ISS of MSX1, DUSP2, SGK1, GPX3, and PAEP in human endometrium across the LH stages. J Spatial visualization of MSX1, DUSP2, SGK1, GPX3, and PAEP in human.
Meanwhile, to thoroughly elucidate the signature of epithelium, we analyzed subcluster-specific genes for functional enrichment. GLE cell-specific genes were highly enriched in “allograft rejection” and “viral infection”, implicating a role in maintaining the endometrial microenvironment via cytokine secretion. Under the influence of accumulated serum progesterone level, proEpi transitioned into diffEpi characterized by enriched FOXO signaling and MAPK resistance signaling (Fig. 4C, D), aligned with the previously reported genetic evidence that epithelial FOXO1 expression and its nuclear translocation are critical for embryo implantation in mice19. SeneEpi exhibited autophagy-related signatures, whereas deEpi were predominantly associated with apoptosis-related pathways. Interestingly, Cilium was associated with NOTCH signaling and motor proteins, which were implicated in ciliogenesis and movement. Immunostaining of E-Cadherin and acetylated Tubulin documented a significantly higher enrichment of Cilium at LH7 (Fig. 4E, Supplementary Fig. 6C).
After projecting these cells into Stereo-seq, proEpi and GLE were abundant during the early LH2 stage, with most LE transitioning into diffEpi by LH7. It was interesting to notice that most of the diffEpi turned into seneEpi with residual diffEpi localized in the same gland at LH9. By the end of LH cycle when the implantation window closed, almost all epithelium transitioned into seneEpi (Fig. 4F, G, Supplementary Fig. 6A). Luminal epithelium, which forms glandular structures through invagination, showed specific expression of genes such as prostaglandin-endoperoxide synthase 1 (PTGS1) and LGR5 in our study (Supplementary Fig. 6D). Immunostaining revealed that COX1, coded by PTGS1, was specifically expressed in LH2 luminal epithelium with also lower expression in glandular epithelium in later stages. While COX2, another member of the cyclooxygenase family, was more specifically localized in luminal epithelium (Supplementary Fig. 6E). Since double deletion of these two genes contributes to complete embryo implantation failure27, our results underscore the potential role of PTGS1 and PTGS2-derived prostaglandins in human embryo implantation.
Surprisingly, co-immunostaining of forkhead box A2 (FOXA2) and keratin 8 (CK8) suggested that FOXA2 was expressed unevenly in glandular epithelium, indicating the unexpected heterogeneity of glandular epithelium (Supplementary Fig. 6F). Indeed, we noticed that Cbp/p300 interacting transactivator with Glu/Asp rich carboxy-terminal domain 4 (CITED4) was gradually decreased in glandular epithelium accompanied with cumulatively increased dipeptidyl peptidase 4 (DPP4) from LH2 to LH12 with the co-expression of these two genes in the same cell at LH7 and LH9 (Fig. 4H, Supplementary Fig. 6G). A similar scenario was also observed for achaete-scute family bHLH transcription factor 5 (ASCL5) and GADD45A (Supplementary Fig. 6H, I). The increased expression of apoptosis-related GADD45A was further reinforced by cleaved Caspase 3 in LH12 epithelium (Supplementary Fig. 6J).
Apart from these genes, we also noticed that MSX1 was highly expressed in LH2 epithelium and gradually decreased to LH12 (Fig. 4I, J, Supplementary Fig. 7A, B). Dual specificity phosphatase 2 (DUSP2), a nuclear phosphatase that inactivates mitogen-activated protein kinase28, exhibited transiently high expression at LH7 (Fig. 4I, J, Supplementary Fig. 7A, B). Serum/glucocorticoid regulated kinase 1 (SGK1), a critical gene mediating rapid progesterone responses29,30, was found highly expressed at LH7 and LH9, heralding its potential role in receptivity opening and closing (Fig. 4I, J, Supplementary Fig. 7A, B). Besides, deregulated SGK1 was documented to be crucial for embryo implantation failure and human recurrent pregnancy loss (RPL)30. In line with previous works, the expressions of glutathione peroxidase 3 (GPX3) and progestagen-associated endometrial protein (PAEP), hallmarks associated with uterine milk in the secretory phase3,31, were persistently increased after the endometrium entered the secretory phase and thereafter (Fig. 4I, J, Supplementary Fig. 7A, B). Additionally, stage-specific gene expression was further revealed, including distal-less homeobox 5 (DLX5), PKHD1-like 1 (PKHD1L1) at LH2, disks large MAGUK scaffold protein 5 (DLG5), and metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) at LH7, C-X-C motif chemokine ligand 14 (CXCL14), DEP domain containing MTOR interacting protein (DEPTOR), and FOXO1 at LH9 (Supplementary Fig. 8A, B). Together, these findings highlighted the dynamic progression of epithelial cell states and gene expression patterns after progesterone exposure, shedding light on the molecular mechanisms underpinning receptivity establishment.
Epithelial cholesterol metabolism is critical for ciliogenesis
To further delineate the molecular characteristics of the epithelial subclusters, we conducted a comparative analysis of DEGs across these populations. The gene expression profiles revealed striking differences among the subclusters (Fig. 5A, Supplementary Fig. 8C). Leveraging SCENIC analysis, we further investigated transcriptional regulon activities, uncovering distinct patterns of transcription factor (TF) enrichment specific to each epithelial subtype. Notably, the expression and regulon activity of FOXO1, a critical epithelial regulator of endometrial receptivity19, were enriched in proEpi and diffEpi, highlighting its role in human embryo implantation. The seneEpi displayed enrichment of senescence-associated transcription factors, such as TP53 and JUN, while the proliferation-related TFs E2F1 and E2F8 were predominantly expressed and activated in proEpi cells (Fig. 5B). While integrins are widely reported as implantation markers, our data revealed that only ITGA3 and ITGB4 were markedly upregulated in the epithelium during receptivity, suggesting that the α3β4 heterodimer may be the predominant integrin involved in embryo attachment (Supplementary Fig. 8D–F).
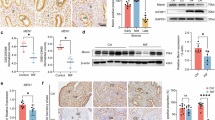
A Differentially expressed genes of epithelial subclusters. B Heatmap of expression and regulon activity of putative transcription factors in each epithelial subtype. C The spatial distribution of CREB3L1 presented by Stereo-seq (bin10). D Spatial visualization of SOX17, HOMER2, KLF5, and ELF3 using ISS in human endometrium across the LH stages. E Stereo-seq and ISS showing cholesterol-related gene in human endometrium at distinct LH phases. Immunostaining displaying SQLE (red) in human endometrium across the LH stages (n = 3). F The morphology, hematoxylin and eosin staining, and FOXJ1 (green) immunostaining of organoids derived from human endometrial epithelium under the treatment of hormones, cholesterol, and atorvastatin (n = 3). Nuclei are counterstained with DAPI (blue). CTL control, EPC estrogen, progesterone, and 8Br-cAMP, Cho cholesterol, Ato atorvastatin.
Interestingly, newly identified cyclic adenosine monophosphate responsive element-binding protein 3 like 1 (CREB3L1), a critical regulator for stromal cell decidualization32, was significantly enriched in proEpi, suggesting its involvement in epithelial differentiation during the proliferative phase (Fig. 5C). Beyond previously reported epithelial-specific SOX1733, we identified other unappreciated epithelial-specific transcription factors, including homer scaffold protein 2 (HOMER2), LH7 specific KLF transcription factor 5 (KLF5) and E74 like ETS transcription factor 3 (ELF3) (Fig. 5D, Supplementary Fig. 9A). LIF-STAT3 signaling pathway was reported to be critical for embryo implantation7,34,35,36,37, we therefore detected the expression of genes associated with LIF-STAT3 signaling pathway. It was interesting that the expression of LIF and its receptor IL6ST was increased at LH9, while interleukin 6 (IL6) remained low and LIFR showed no significant changes (Supplementary Fig. 9B). Furthermore, phosphorylated STAT3 (p-STAT3) was primarily localized in stroma at LH2 and LH7, significantly upregulated in the epithelium at LH9, and markedly decreased in both the epithelium and stroma by LH12 (Supplementary Fig. 9C). These results indicated that LIF-STAT3 played an important role in the establishment of endometrial receptivity in humans. Although both LH7 and LH9 belong to the mid-secretory phase, the transient activation of p-STAT3 in the LH9 epithelium implied that LH9 might represent the critical time point for the opening of the implantation window.
Additionally, FOXJ1, a central regulator of ciliogenesis, was exclusively expressed in Cilium, consistent with previous studies3. Cilia were reported to secrete extracellular vesicles, termed ectosomes, at their tip to transport bioactive enzymes, peptide products, or remove activated receptors from the cell surface38. Cholesterol had been reported as essential for ciliogenesis39. Intriguingly, we also found that cholesterol biosynthesis rate-limiting enzymes 3-hydroxy-3-methylglutaryl-CoA reductase (HMGCR) and squalene epoxidase (SQLE) were specifically expressed in epithelium. More importantly, the expression of HMGCR peaked at LH7 (Fig. 5E, Supplementary Fig. 10A). To validate this, primary epithelium-derived organoids were treated with cholesterol or the HMGCR inhibitor atorvastatin. Results showed that the expression of FOXJ1 and Ac-Tubulin was significantly increased in the presence of cholesterol (Fig. 5F, Supplementary Fig. 10B) and inhibited by atorvastatin (Fig. 5F). In a word, the epithelium underwent dynamic spatial-temporal changes of cell types across the LH cycle and was critical for embryo implantation.
The molecular characterizations of the endothelium in receptivity
Endothelial cells were classified into three subtypes according to the endothelium atlas40, including venous endothelial cells (C-C motif chemokine ligand 14, CCL14; ryanodine receptor 2, RYR2), arterial endothelial cells (gap junction protein alpha 4, GJA4; regulator of calcineurin 2, RCAN2) and proliferating endothelial cells (proEndo) (aurora kinase B, AURKB; kinesin family member 2 C, KIF2C) (Fig. 6A–C). Notably, the proportion of proEndo peaked at LH7 and LH9 and was validated by the expression of MKI67 (Fig. 6D). According to KEGG analysis, LH2 endothelial-specific genes were primarily enriched in “FOXO signaling pathway”, which was closely relevant to endothelial quiescence41. By LH12, pathways related to apoptosis and the AGE-RAGE signaling axis were prominently enriched (Fig. 6E). Interestingly, atypical chemokine receptor 1 (ACKR1), a critical mediator in the transport of chemokines42, was predominantly expressed at later LH stages, implicating the role of endothelial cells in facilitating intercellular communication during this phase (Fig. 6F). Furthermore, spatial transcriptomic analysis revealed distinct localization of endothelial cells expressing ACKR1, roundabout guidance receptor 4 (ROBO4) and kinase insert domain receptor (KDR) within the endometrium (Fig. 6G). In brief, our results unraveled the spatiotemporal characteristics of the endothelium in the receptivity window.
A UMAP visualization showing the re-clustering of endothelium. B The proportion of endothelial subclusters in the human endometrium across the LH stages. C Dot plot showing marker genes for different endothelial cells. The color bar represents the normalized expression of genes. The size of the plot represents the percentage of expressed cells. D Co-immunostaining of VE-Cadherin (red) and MKI67 (green) in human endometrium across the LH stages (n = 3). Nuclei are counterstained with DAPI (blue). The arrows represent proliferating endothelial cells. E Functional enrichment analysis of DEGs among vein, artery, and proEndo in human endometrium across the LH stages. The color bar represents the log-transformed p-value. The size of the plot represents the gene ratio. Significance based on over-representation test in clusterProfiler analysis with Benjamini–Hochberg-adjusted P-values. F Violin plots showing the expression of ABCG2, DMBT1, ACKR1, and KCNJ6 in human endometrium across the LH stages. G The localization of ACKR1, ROBO4, and KDR by ISS across the LH stages. H UMAP visualization showing the re-clustering of immune cells (left). The proportion of immune subclusters in the human endometrium across the LH stages (right). I Violin plots showing marker genes in different immune cell subclusters. J Co-immunostaining of CD56 (red) and MKI67 (green) in human endometrium at LH7 and LH12 (n = 3). Nuclei are counterstained with DAPI (blue). K Distribution of CD56+ NK cell subcluster (green) in human endometrium from LH2 to LH12 by Stereo-seq (cellbin). L Localization of RRM2 and GZMA in endometrium by ISS. M Functional enrichment analysis of DEGs among CD56 + NK cell subclusters in human endometrium across the LH stages. The color bar represents the log-transformed p-value. The size of the plot represents the gene ratio. Significance based on over-representation test in clusterProfiler analysis with Benjamini-Hochberg-adjusted P-values. N Expression of cell cycle and apoptosis-related genes in CD56 + NK cell subcluster from LH2 to LH12. The color bar represents the normalized expression of genes. O Spatial distribution of NK cells specific GZMA, EOMES, and macrophage CD14 in human endometrium across the LH stages.
Spatial resolution of the immune microenvironment
For immune cells annotated with Azimuth package43, we identified 12 immune cell subclusters within the endometrium, including mast cells (Mast), CD4 positive T cells (CD4_T), CD8 positive T cells (CD8_T), double-negative T cells (dnT), regulatory T cells (T_reg), proliferating natural killer T cells (pro_NKT), CD56 positive natural killer cells (CD56_NK), CD16 positive natural killer cells (CD16_NK), B cells (B), macrophage (Mac), and two types of dendritic cells: plasmacytoid dendritic cells (pDC) and conventional dendritic cells (cDC) (Fig. 6H, I). Co-immunostaining of MKI67 and CD56 revealed substantial accumulation of CD56+ NK cells, known for their robust cytokine production and rapid mediation of antibody-dependent cell-mediated cytotoxicity (ADCC), in the endometrium at LH9 (Fig. 6J, Supplementary Fig. 10C). Stereo-seq and ISS were also performed to verify the above changes of CD56+ NK subclusters (Fig. 6K, L). This accumulation of NK cells at LH12 likely correlated with processes involved in endometrial shedding. Functional enrichment analysis of DEGs across LH stages indicated that “DNA replication” was markedly upregulated in NK cells during LH7/LH9, as evidenced by high expression of cyclin A2 (CCNA2), cyclin-dependent kinase 6 (CDK6), and proliferating cell nuclear antigen (PCNA). Conversely, genes in the terms of “p53 signaling pathway” and “apoptosis” were increased at LH12, indicating a preparatory phase for endometrial shedding (Fig. 6M, N, Supplementary Fig. 10D). Additionally, macrophages, marked by CD14, were significantly enriched in the endometrium at LH12. This macrophage accumulation likely reflected their role in phagocytic clearance of cellular debris, contributing to tissue homeostasis and repair (Fig. 6O). Collectively, by integrating Stereo-seq, scRNA-seq, and ISS technologies, we obtained a comprehensive spatiotemporal map of endothelium and immune cells across the LH stages, highlighting the intricate coordination of the immune microenvironment in the maintenance of endometrial homeostasis and remodeling.
The dominant role of stroma in spatial communications
Stromal cells were the most abundant cell type, indicating their critical role in maintaining endometrial homeostasis. To elucidate the interactions between stromal cells and other cellular types, CellPhoneDB was applied to identify potential ligand-receptor pair-mediated communications. Ranking cellular interactions by the number of connections revealed stromal cells, endothelial cells, and epithelial cells as the most active participants in endometrial signaling (Fig. 7A). Specifically, intercellular crosstalk between the epithelium and stroma was predominantly mediated by signal molecules such as fibroblast growth factor 2 (FGF2), FGF7 and bone morphogenetic protein 4 (BMP4) (Fig. 7B). Among them, the FGF7-FGFR2 signaling axis emerged as particularly significant, playing a critical role in epithelial cell differentiation and maturation44. FGF7 was predominantly expressed in diffStr, whose receptor FGFR2 was specifically localized to epithelial cells, suggesting a key regulatory role of FGF7-FGFR2 signaling in driving epithelial differentiation and functional maturation (Fig. 7B, C). Consistently, spatial transcriptomic visualization demonstrated robust FGF7 expression in stromal compartments from LH7 to LH9, with FGFR2 being distinctly enriched in epithelial regions (Fig. 7D). Ligand-receptor pairs such as IHH_HHIP, WNT4_SFRP4, WNT9A_SFRP4, WNT7A_SFRP, and WNT5B_SFRP4 were primarily observed in LH2, indicating their roles in maintaining the undifferentiated state of the endometrium. Additionally, the E2-ESR1 axis was predominantly enriched in LH2, which is associated with the thickening of the endometrium by promoting cell proliferation (Fig. 7C).
A Abundance of connections between different cell types in the human endometrium across the LH stages is shown by CellphoneDB. B The interactions between epithelial cells (proEpi, diffEpi, deEpi, seneEpi, Cilium, GLE, and LE) and stromal cells (proStr, undiffStr, diffStr, and deStr). Prioritized ligands are defined by NicheNet, with Pearson correlation indicating the strength of the ligand-target gene relationship. Higher-ranking ligands are predictive of stronger interactions. Dot plots represent the expression of ligands in stromal cells and their corresponding receptors in epithelial cells. The heatmap shows the predicted activity of ligands on their target genes across different stromal cell types, as determined by NicheNet. C Bubble diagram showing highly expressed ligand-receptor pairs mediating interactions between stromal and epithelial cells in LH2 and LH9. The size of the dot represents the log-transformed expression of the indicated ligand and its receptor in different cell types, and the color bar represents their enrichment significance. D Spatial visualization of FGF2, FGF7, and FGFR2 using Stereo-seq (bin10). E Heatmap represents the interactions between ligands in four stromal cells (proStr, undiffStr, diffStr, and deStr) and their receptors in NK cells (pro_NK, CD56_NK, and CD16_NK). Dot plots respectively represent the average expression levels of ligands and receptors in stromal and NK cells. The color bar in the bottom left corner indicates the normalized gene expression levels, the color bar in the bottom middle represents the evaluated ligand activity, and the color bar in the bottom right corner denotes the interaction potential. F Spatial visualization showing ligands in stromal cells (IL-15) and their receptors in NK cells (IL2RB, IL2RG) using Stereo-seq (bin10). ISS verifies the expression of IL-15 (rose red). G A schematic illustration showing the interactions between stromal cells, epithelial cells, and NK cells in human endometrium across the LH stages.
IL-15 plays a critical role in endorsing the survival, proliferation, and cytotoxic activity of NK cells. In the endometrium, IL-15 was mostly expressed in diffStr and deStr. Ligand-receptor interaction analyses demonstrated the presence of IL-15-IL2RB-IL2RG signaling between stromal and NK cells, implying the role of stromal cells in supporting the proliferation and activation of CD56+ NK cells (Fig. 7E). Stereo-seq analysis confirmed that IL-15 was specifically localized to the stromal compartment, with its expression markedly increased from LH7 onward. Concurrently, its receptors, IL2RB and IL2RG, were distinctly upregulated in NK cells, underscoring a spatially coordinated regulatory network (Fig. 7F). Signals originating from the epithelium and NK cells that targeted stromal compartments were also examined (Supplementary Fig. 11A, B). In summary, stromal cells dynamically responded to changes in the endometrium by secreting cytokines such as FGF2 and FGF7, which promoted epithelial differentiation via FGFR2. Moreover, stromal cells produced IL-15 to modulate NK cell activity (Fig. 7G). Our spatial-temporal map of receptivity not only uncovers the cell-specific expression code of genes associated with receptivity opening, closing, and menstruation preparation, but also depicts the potential spatial crosstalk underlying this process (Fig. 8).
Spatiotemporal transcriptomic profiling reveals pronounced cellular heterogeneity of the endometrium across distinct LH stages, during which epithelial and stromal cells sequentially undergo proliferation, differentiation, and senescence/degradation, accompanied by the activation of specific signaling pathways. Notably, epithelial cholesterol is critical for ciliogenesis, and stromal-derived FGF2/7 and IL-15 coordinately regulate intercellular interactions and microenvironmental homeostasis, thereby facilitating the establishment of the implantation window.
Discussion
Gaining insights into the topographical transcriptomics of human endometrial receptivity remains a challenge due to the difficulty in catching the transient timing of the implantation window precisely. Previous studies have primarily focused on the mid-secretory phase, relying on B-ultrasonography or urine/serum LH levels to estimate the approximate ovulation timing. In this study, the exact ovulation timing is induced by a single hCG administration based on the size of the follicles. While previous studies have provided comprehensive resources of human endometrium by scRNA or 10× Visium14,16, a spatiotemporal landscape of human endometrial receptivity at single-cell resolution remains elusive. To fill this gap, we integrated Stereo-seq, scRNA-seq, and ISS in synchronized human endometrium to parse the spatiotemporal characteristics of the opening and closing of receptivity, along with an interactive website (https://db.cngb.org/stomics/datasets/STDS0000249/summary) for visualization. This spatiotemporal transcriptome dataset offers an invaluable resource to determine the pivotal cellular and molecular features underlying endometrial receptivity.
Previous studies using microarray and scRNA-seq have identified several genes, such as GPX3, PAEP, MAOA, and IL-15, that are closely associated with the opening of the implantation window11,15. Our study supported these studies well. Additionally, we constructed the implantation-associated spatial atlas and provided insightful information about the previously unappreciated markers. Among them, CEBPD and NR2F1 manifested special expression in the stromal cells, with SGK1 expression exclusively in the epithelium during the implantation window. Although the expression of GPX3 and PAEP increased from LH7, these two genes peaked at LH12, showing poor correlation with the implantation window. Besides, although IGFBP1 and PRL were reported as two critical markers for the pre-decidualization of stromal cells, our data clearly showed that their expression levels were very low and they were mainly localized in the epithelium before embryo implantation, implying that they are inappropriate to be the markers of the implantation window.
The LIF signaling pathway is essential for embryo implantation7,34,35,36,37. LIF exhibited higher expression in the secretory phase than the proliferative phase, and was mainly located in the epithelium45,46, which was also proven in our data. Infertile women show lower LIF level and disturbed LIF pathway47,48. Our data hinted that epithelial p-STAT3 is primarily activated at LH9 by LIF and IL6ST. Given that, administering LIF at the appropriate timing could be a promising strategy to enhance pregnancy outcomes for infertile patients. While HOXA10 was reported to be a critical marker of implantation, our data clearly demonstrated that HOXA10 mRNA was obviously decreased at the window of implantation, which was well supported by other scRNA-seq datasets3,15. In one word, our data refined the cognition of endometrial receptivity and established a valuable spatial code for the opening of the implantation window.
Aberrant MSX1 expression in human endometrium has been linked to infertility49. Our data support that lower MSX1 expression may serve as a potential marker for receptivity establishment. MSX1 is known as a transcriptional repressor; its direct target genes remain largely unexplored, which deserves further delving.
The endometrial cilium is believed to facilitate the transport of glandular secretions (“gland milk”) and the movement and reorganization of embryos31,50. Although the role of cilium in endometrium is gaining recognition, the genes and pathways driving ciliogenesis remain elusive. Our results demonstrate the crucial role of cholesterol in ciliogenesis. Cholesterol, a vital component of the plasma membrane, regulates the rigidity, fluidity, and permeability of the cell membrane and the formation of cilia via posttranslational modification of Hedgehog (HH) and Smoothened (SMO), ensuring proper Hedgehog signaling39,51. Given the fact of the vital role of cholesterol in ciliogenesis, the use of cholesterol-lowering statins during pregnancy preparation should be carefully evaluated for women with hypercholesterolemia, considering the potential impact on endometrial receptivity.
In stromal cells, the most notable change is the transformation from WNT pathway-dominant undiffStr to diffStr, accompanied by extensive endometrial remodeling for embryo implantation preparation. Our results suggest that PTGDS and PLA2G2A-derived prostaglandins are potential factors for stromal cell differentiation. Intriguingly, our previous study reveals that PLA2G2A is predominantly expressed in undecidualized stromal cells in first-trimester endometrium, without the expression of decidualization markers IGFBP1 and PRL52. As the most abundant cell type in the endometrium, stromal cells play a vital role in maintaining homeostasis within the endometrial niche through spatial crosstalk with other cell types. Our investigation unravels that the stroma is the primary source of FGFs in the endometrium, especially at LH7 and LH9. FGFs are also critical for mouse receptivity under the orchestration of HAND28. Distinct FGFs possess varying affinity to FGFRs, participating in various processes, including proliferation, epithelial-to-mesenchymal transition (EMT), and, particularly, differentiation8,44,53,54. Our study throws a previously unknown role of stromal cell-derived FGF2/7 in regulating epithelial differentiation for receptivity establishment by targeting epithelial FGFR2.
Our understanding of menstruation remains limited. We notice that after the endometrium transitions into refractory phase at LH12, approximately three days before menstruation, both stroma and epithelium switch to a senescence state. Endometrial epithelial senescence used to be believed to be related to advanced age and pathological conditions26,55. Meanwhile, NK accumulation increases significantly at later stage via proliferation under the influence of stroma-derived IL-15. It is plausible that the increased immune cells in the late stage facilitate the senescence of epithelial and stromal cells and simultaneously prepare for tissue repair following menstruation.
Current commercial endometrial receptivity array (ERA) identified 238 genes derived from microarray, which were further supported by recent randomized controlled trials11,12. With the advent of high-throughput sequencing, scRNA-seq, and spatial-transcriptome, the cell-type and spatial-specific signature provides new insight into endometrium dating. Especially, the spatiotemporally expressed genes peaked at LH7 and LH9 in this investigation, such as stromal NR2F1 and CEBPD and epithelial SGK1, ELF5, and KLF5, need to be paid more attention in receptivity evaluation. In all, these findings provide mechanistic insights and generate actionable hypotheses that may guide future clinical applications.
While the concordance across multiple experimental approaches (including Stereo-seq, scRNA-seq, ISS, immunostaining, and organoid assays) strengthens the robustness of our findings, we acknowledge that the relatively limited sample size (1 sample per time point for Stereo-seq and two samples per time point for scRNA-seq) may affect the generalizability of stage-specific conclusions and reduce statistical power. Future studies utilizing larger, independent cohorts will be essential to assess inter-individual variability and validate and extend our observations.
In conclusion, this study constructs a spatiotemporal transcriptional landscape of human endometrial receptivity, offering a valuable resource for investigating the cellular and molecular mechanisms underlying receptivity opening and closing.
Methods
Human endometrium sample collection
The study was approved by the Ethics Committee of Nanjing Drum Tower Hospital, Nanjing University Medical School (IRB reference number 2013-081-01) and was conducted according to all relevant ethical regulations. Written informed consent was obtained from all participants, and no financial compensation was provided. Detailed clinical information on these 21 subjects is provided in Supplementary Data 2.
To investigate the molecular signatures associated with endometrial receptivity, we recruited 40 females undergoing in vitro fertilization and embryo transfer (IVF-ET) cycles due to male factors. Inclusion criteria were as follows: (1) 23–35 years old, (2) regular menstrual cycle ranging from 21 to 35 days, (3) 18.5 kg/m² ≤body mass index (BMI) < 24 kg/m², (4) absence of other endocrine or immune-related disorders. Follicle development was monitored starting on day 10 of the natural menstrual cycle. Subcutaneous administration of 250 μg recombinant hCG (OVIDREL, Merck Serono) was performed on the day when the dominant follicle reached a diameter of ≥16 mm, which was designated as LH0. Ovulation occurred around 38 h later56. Endometrial biopsies were subsequently collected on LH2, LH7, LH9, and LH12 (n = 10 for each group). Of these, 21 subjects subsequently achieved a confirmed clinical pregnancy, and their endometrial samples were included in further high-throughput profiling and experimental validation. Eight samples, two per time point, were subjected to single-cell RNA sequencing (scRNA-seq) analysis. From these, two representative samples per time point were also analyzed using spatiotemporal enhanced resolution omics-sequencing (Stereo-seq, n = 4) and high-throughput in situ sequencing (ISS, n = 4). The remaining samples were used for experimental validation, including immunostaining and organoid experiments.
Culture of human endometrial epithelial organoids
The procedure of the development of human endometrial epithelial organoids was modified from a previous study57. Briefly, endometrial biopsy from three independent individuals was minced and digested with collagenase type I. After 30 min, the digestion was stopped by adding FBS, and the mixture was passed through a 100-μm cell strainer. Then, the strainer was inverted, and the gland structures were backwashed and pipetted up and down to partially dissociate cells. Four times the volume of ice-cold Matrigel was added and mixed thoroughly, and 20 μL of the cell-Matrigel mixture was plated on the preheated 48-well plate. After the cell-Matrigel mixture set, each drop was overlaid with 250 μL Expansion Medium (ExM).
To mimic the secretory phase epithelium, estrogen, progesterone (P), and 8-bromoadenosine 3′,5′-cyclic monophosphate (8Br-cAMP) were sequentially supplemented in the ExM for 4 days. Specifically, when the diameter of most organoids reached 150 μm, the medium was replaced with ExM supplemented with 10 nM E2, which was counted as day 1 (D1) of hormone stimulation. The medium was changed daily for 2 days. On D3, the medium was replaced with ExM supplemented with 10 nM E2, 1 μM P and 0.5 mM 8Br-cAMP, and was changed daily for a 2-day culture. To evaluate the impact of cholesterol metabolism on epithelial ciliogenesis, atorvastatin (8 μM) was added starting on D2, or cholesterol (50/250 μM) was added starting on D3. The expression changes of ciliogenesis-related markers FOXJ1 and acetylated α-tubulin were then assessed. To determine the effects of FGF2 and FGF7 on epithelial differentiation, FGF10 was removed from the original ExM to avoid interference, and FGF2/FGF7 (100 ng/mL) were added from D3. Chemicals, peptides, recombinant proteins, and commercial assays used in this paper are listed in Supplementary Table 1.
Immunofluorescence
Frozen sections from the human endometrium were fixed at room temperature with freshly prepared 4% paraformaldehyde (PFA) for 60 min, followed by 4 times washing in PBST buffer (PBS + 0.1%Triton X-100) for 5 min each time. Blocking with 5% bovine serum albumin (BSA) at room temperature for 1 h reduced the non-specific binding of antibodies. Primary antibodies were diluted with blocking solution in appropriate proportions according to the ratio recommended in the antibody instructions and incubated at 4 °C overnight. Secondary antibodies or DAPI at a 1:200 dilution in PBS buffer were incubated for 30 min at room temperature. The antibodies used in this paper are listed in the Supplementary Table 2.
Hematoxylin and eosin (H&E) staining
Frozen tissue sections of the human endometrium were used for H&E staining. Firstly, tissue sections were fixed with 4% PFA for 30 min and washed 3 times with PBS for 5 min each time. Then, sections were stained with hematoxylin for 3 min, immersed in PBS for 5 min, washed again with water for 40 min, and stained with eosin staining solution for 2 min. Next, tissue sections were dehydrated with 50% ethanol, 70% ethanol, 80% ethanol, 95% ethanol, and anhydrous ethanol for 10 s each sequentially, and then replaced with xylene twice for 5 min each time. Finally, the tissue sections were sealed with neutral resin.
10X Genomics single-cell RNA-seq of human endometrium
The human endometrium was rinsed with ice-cold DPBS and chopped using scalpels into small pieces and enzymatically digested in 5 mL DPBS containing 1 mg/mL collagenase type IV. The supernatant was diluted with DPBS and passed through a 40 μm cell sieve. The flow-through was centrifuged and resuspended in 1 mL of red blood cell lysis buffer for 2 min. The cells were collected by spinning down at 500 rpm for 5 min and resuspended in DPBS for subsequent 10× scRNA sequencing (Chromium GEM-X Single-Cell 3’ Reagent Kits v4).
Stereo-seq sample preparation and sequencing
For Stereo-seq, the human endometrium was embedded in Tissue-Tek O.C.T. Compound and snap-frozen on top of dry ice. The frozen samples were stored at −80 °C until further use. Frozen sections with a thickness of 10 μm were mounted onto a Stereo-seq chip (1 cm × 1 cm), which is composed of numerous 200 nm-diameter DNA nanoballs (DNBs) with 500 nm for the distance from the center to center of DNBs (500 nm resolution). Tissue sections adjacent to Stereo-seq were retained for H&E staining. The tissue sections were fixed with pre-cooled methanol at −20 °C for 30 min and then stained with ssDNA Assay Kit. Tissue permeabilization was performed using PR Enzyme for 18 min. Reverse transcription was conducted at 42 °C using the Gene Expression kit. For library construction, cDNA was fragmented and amplified by a Stereo-seq DNA library kit, after which sequencing libraries were constructed for the subsequent sequencing process.
Processing of Stereo-seq data
Paired-end FASTQ reads were generated by the MGI DNBSEQ-Tx sequencer, where the coordinate identity (CID) and molecular identifier (MID) were obtained from Read 1, and the cDNA sequence was included in Read 2. The raw FASTQ files were subjected to the Stereo-seq Analysis Workflow (SAW). Briefly, the preprocessed reads were aligned to the human GRCh38 reference genome using STAR. Low-quality reads were filtered out using HandleBam. The final expression matrix was constructed from exonic reads containing CID information, with genes as rows and DNBs as columns. The resulting GEF file from SAW was subjected to downstream analysis or visualization using different bin sizes. The size of these bins determines the resolution of the spatial transcriptome. A smaller bin size (e.g., bin10 representing 10 × 10 DNB bins) allows for more precise localization of gene expression patterns.
In situ sequencing (ISS)
In situ sequencing exploits a novel probing and barcoding approach for improving signal intensity and specificity. Frozen tissue sections of human endometrium were subjected to in situ sequencing performed by Genedenovo Biotechnology Co., Ltd (Guangzhou, China). In brief, tissue sections of human endometrium in 10 μm thickness were fixed in 4% PFA for 5 min, followed by washing twice with diethylpyrocarbonate-treated PBS. Tissue sections were dehydrated in an ethanol series and then were permeabilized with 0.1 M HCl at 37 °C for 5 min after 3 times washes. Tissue sections were incubated at 37 °C for 4 h in the probe hybridization mixtures containing RNA molecular markers. Next, the sections were incubated with the ligature mixture, circularization mixture including splint primers, and amplification mixture for rolling circle amplification (RCA). For sequencing, the ligation sequence, imaging, and stripping off were applied for each cycle until all cycles were achieved (four cycles at most).
Multiplexed immunofluorescence staining
The frozen tissues were sectioned from the human endometrium and attached to the slide provided in the FluoXpert mIF kit. The sections were fixed with 4% PFA for 15 min and washed 3 times with PBS, and then submerged in PBS and stored at 4 °C. Before the staining experiment, the tissue sections were re-fixed with 4% PFA for 15 min, and then washed 3 times with PBS and permeabilized with 0.5% Triton X-100 for 15 min at room temperature prior to flow cell assembly. Different antibodies were added to the FluoXpert mIF reagent cartridge. Multiplexed immunofluorescence was then performed with DNBSEQ-G400RS FluoXpert.
Data processing of scRNA-seq data
Single-cell libraries were sequenced on Illumina HiSeq X Ten instruments using 150 bp paired-end sequencing. The raw data were processed using CellRanger (v5.0.1) with default parameters and aligned to the GRCh38 genome reference and annotation. The downstream analyses, including preprocessing, normalization, identification of variable features, data scaling, dimensional reduction, clustering, and top marker identification, were mostly fulfilled by Seurat (v5.0.3). We filtered cells with >20% mitochondrial content (percent. mt), unique feature counts (nFeature_RNA) over 10,000 or under 500, or RNA counts (nCount_RNA) exceeding 30,000 or below 1000, resulting in 82,980 cells remaining. Batch effects across datasets from different libraries were addressed using the RunHarmony function. FindAllMarkers identified the marker genes of each cluster mostly. For subset analysis, we followed a similar approach: selecting the top 2000 variable features, scaling the data, recomputing PCA, clustering, and visualizing with UMAP. If necessary, RunHarmony was used to adjust for batch effects. The differential expression genes were calculated utilizing the FindMarkers function in Seurat with the Wilcoxon test (fold change <0.5 or >2 and p-value < 0.05) and visualized using the R package scRNAtoolVis. The DEGs of stromal cells, epithelial cells, and NK cells were provided in Supplementary Data 3.
Mapping cell types from scRNA-seq onto Stereo-seq
To visualize spatial distribution patterns of the identified cell types, we mapped them onto Stereo-seq data based on our scRNA-seq datasets (LH2, LH7, LH9, and LH12 endometrium). The FindTransferAnchors method in Seurat was applied to identify anchors between the scRNA-seq and Stereo-seq datasets. The TransferData function was then used to transfer scRNA-seq clusters to selected spots in the Stereo-seq data. Each spot was assigned a potential cell type based on the scRNA-seq cluster with the highest transfer score. Subcluster annotation for Stereo-seq was performed in the same manner as above.
Cell-type annotation using TACCO
To accurately annotate cell types in Stereo-seq spatial transcriptomic data, we applied TACCO (Transfer of Annotations to Cells and their COmbinations)58. scRNA-seq data from human endometrium were used as the reference for cell-type signatures. Both scRNA-seq and Stereo-seq data were normalized and log-transformed prior to analysis. TACCO assigns probabilistic cell-type labels to each spatial spot, accounting for spot composition and gene capture variability. The cell type with the highest probability was assigned as the primary annotation. This approach enabled reliable annotation of small or sparsely distributed cell populations, including epithelial subtypes and immune cells. Analyses were performed using TACCO v0.0.2 (https://github.com/simonwm/tacco) with default parameters.
Cell-cell interaction analysis by CellPhoneDB and NicheNet
To explore potential interactions between different cell types, cell-cell interaction analysis was conducted using CellPhoneDB, a public database of receptor-ligand interactions, including heteromeric complexes with subunit structures. Briefly, a log2-normalized count matrix was subsampled to 500 cells per cluster. Significant ligand-receptor pairs were identified after filtering. For each identified pair, the total number across clusters within the same cellular module was calculated, allowing cellular-module-specific ligand-receptor pairs to be determined based on enrichment scores. To identify potential ligands influencing latent phenotypes, NicheNet (v2.2.0) was used to investigate potential interactions and target genes between specified cell types.
Single-cell regulatory network inference and clustering (SCENIC) analysis
Following cluster annotation, we performed pySCENIC analysis using a three-step workflow based on human cisTarget resources. First, we converted the scRNA-seq gene expression matrix into a loom file using Python. Next, we used the pyscenic grn, pyscenic ctx, and pyscenic aucell functions to infer the gene regulatory network, predict regulon activity, and identify the key transcription factors and their target genes in each cell type. AUCell then scored the activity of each regulon in individual cells, producing a binarized activity matrix.
RNA velocity analysis
Unspliced and spliced RNA were extracted from the BAM file using velocyto, resulting in a data matrix that was preprocessed with scVelo. Preprocessing steps included normalization, feature gene selection, and PCA for dimensionality reduction. Default parameters were then used to estimate dynamic parameters and compute gene-wise RNA velocity vectors. These velocity vectors were subsequently projected onto the UMAP space for visualization and analysis. To explore dynamic gene expression along the velocity and latent time, a differential kinetic test was performed using the scv.tl.differential_kinetic_test function.
Statistics & reproducibility
No statistical method was used to predetermine sample size. Statistical analyses were performed using GraphPad Prism (v.9.0) (for experimental data), R (v.4.1.0), and RStudio (2021.09.1). Comparisons between groups were conducted using Student’s t-tests and Wilcoxon rank-sum tests, and ANOVA was used for continuous variables. P < 0.05 was considered to be statistically significant. For organoid experiments, three independent samples were utilized, and one-way ANOVA was applied for multiple group comparisons. Software and algorithms used in this paper are listed in the Supplementary Table 3.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw sequencing data reported in this paper have been deposited at the National Center for Biotechnology Information Sequence Read Archive under accession number PRJNA1210010 and the China National GeneBank under accession number STT0000132. The online visualization of spatial transcriptomics data in this paper have been deposited at the China National GeneBank under the dataset ID STDS0000249. Source data are provided with this paper.
Code availability
All original code is deposited at GitHub (https://github.com/Zluei/Human_endometrium). The combined RDS files are publicly accessible in Zenodo at https://doi.org/10.5281/zenodo.16851856 and https://doi.org/10.5281/zenodo.17176901.
References
Niederberger, C. et al. Forty years of IVF. Fertil. Steril. 110, 185–324.e185 (2018).
Achache, H. & Revel, A. Endometrial receptivity markers, the journey to successful embryo implantation. Hum. Reprod. Update 12, 731–746 (2006).
Wang, W. et al. Single-cell transcriptomic atlas of the human endometrium during the menstrual cycle. Nat. Med. 26, 1644–1653 (2020).
Noyes, R. W., Hertig, A. T. & Rock, J. Reprint of: Dating the endometrial biopsy. Fertil. Steril. 112, e93–e115 (2019).
Wang, H. & Dey, S. K. Roadmap to embryo implantation: clues from mouse models. Nat. Rev. 7, 185–199 (2006).
Stewart, C. L. et al. Blastocyst implantation depends on maternal expression of leukaemia inhibitory factor. Nature 359, 76–79 (1992).
Lee, J. H. et al. Signal transducer and activator of transcription-3 (Stat3) plays a critical role in implantation via progesterone receptor in uterus. FASEB J. 27, 2553–2563 (2013).
Li, Q. et al. The antiproliferative action of progesterone in uterine epithelium is mediated by Hand2. Science 331, 912–916 (2011).
Surveyor, G. A. et al. Expression and steroid hormonal control of Muc-1 in the mouse uterus. Endocrinology 136, 3639–3647 (1995).
Daikoku, T. et al. Uterine Msx-1 and Wnt4 signaling becomes aberrant in mice with the loss of leukemia inhibitory factor or Hoxa-10: evidence for a novel cytokine-homeobox-Wnt signaling in implantation. Mol. Endocrinol. 18, 1238–1250 (2004).
Diaz-Gimeno, P. et al. A genomic diagnostic tool for human endometrial receptivity based on the transcriptomic signature. Fertil. Steril. 95, 50–60 (2011).
Simon, C. et al. A 5-year multicentre randomized controlled trial comparing personalized, frozen and fresh blastocyst transfer in IVF. Reprod. Biomed. Online 41, 402–415 (2020).
Doyle, N. et al. Effect of timing by endometrial receptivity testing vs standard timing of frozen embryo transfer on live birth in patients undergoing in vitro fertilization: a randomized clinical trial. JAMA 328, 2117–2125 (2022).
Santamaria, X. et al. Decoding the endometrial niche of Asherman’s syndrome at single-cell resolution. Nat. Commun. 14, 5890 (2023).
Cao, D. et al. Time-series single-cell transcriptomic profiling of luteal-phase endometrium uncovers dynamic characteristics and its dysregulation in recurrent implantation failures. Nat. Commun. 16, 137 (2025).
Garcia-Alonso, L. et al. Mapping the temporal and spatial dynamics of the human endometrium in vivo and in vitro. Nat. Genet. 53, 1698–1711 (2021).
Ho, V. N. A. et al. Livebirth rate after one frozen embryo transfer in ovulatory women starting with natural, modified natural, or artificial endometrial preparation in Viet Nam: an open-label randomised controlled trial. Lancet 404, 266–275 (2024).
Mote, P. A., Balleine, R. L., McGowan, E. M. & Clarke, C. L. Colocalization of progesterone receptors A and B by dual immunofluorescent histochemistry in human endometrium during the menstrual cycle. J. Clin. Endocrinol. Metab. 84, 2963–2971 (1999).
Vasquez, Y. M. et al. FOXO1 regulates uterine epithelial integrity and progesterone receptor expression critical for embryo implantation. PLoS Genet. 14, e1007787 (2018).
Mareckova, M. et al. An integrated single-cell reference atlas of the human endometrium. Nat. Genet. 56, 1925–1937 (2024).
Gargett, C. E., Schwab, K. E. & Deane, J. A. Endometrial stem/progenitor cells: the first 10 years. Hum. Reprod. Update 22, 137–163 (2016).
Davis, A. E. 3rd, Lu, F. & Mejia, P. C1 inhibitor, a multi-functional serine protease inhibitor. Thromb. Haemost. 104, 886–893 (2010).
Christian, M. et al. Cyclic AMP-induced forkhead transcription factor, FKHR, cooperates with CCAAT/enhancer-binding protein beta in differentiating human endometrial stromal cells. J. Biol. Chem. 277, 20825–20832 (2002).
Gao, F., Bian, F., Ma, X., Kalinichenko, V. V. & Das, S. K. Control of regional decidualization in implantation: Role of FoxM1 downstream of Hoxa10 and cyclin D3. Sci. Rep. 5, 13863 (2015).
Gui, Y., Zhang, J., Yuan, L. & Lessey, B. A. Regulation of HOXA-10 and its expression in normal and abnormal endometrium. Mol. Hum. Reprod. 5, 866–873 (1999).
Lv, H. et al. Deciphering the endometrial niche of human thin endometrium at single-cell resolution. Proc. Natl. Acad. Sci. USA 119, e2115912119 (2022).
Aikawa, S. et al. Spatiotemporally distinct roles of cyclooxygenase-1 and cyclooxygenase-2 at fetomaternal interface in mice. JCI Insight 9, e181865 (2024).
Wu, M. H., Lin, S. C., Hsiao, K. Y. & Tsai, S. J. Hypoxia-inhibited dual-specificity phosphatase-2 expression in endometriotic cells regulates cyclooxygenase-2 expression. J. Pathol. 225, 390–400 (2011).
Jeong, J. W. et al. Identification of murine uterine genes regulated in a ligand-dependent manner by the progesterone receptor. Endocrinology 146, 3490–3505 (2005).
Salker, M. S. et al. Deregulation of the serum- and glucocorticoid-inducible kinase SGK1 in the endometrium causes reproductive failure. Nat. Med. 17, 1509–1513 (2011).
Kelleher, A. M., DeMayo, F. J. & Spencer, T. E. Uterine glands: developmental biology and functional roles in pregnancy. Endocr. Rev. 40, 1424–1445 (2019).
Ahn, J. I. et al. cAMP-response element-binding 3-like protein 1 (CREB3L1) is required for decidualization and its expression is decreased in women with endometriosis. Curr. Mol. Med 16, 276–287 (2016).
Wang, X. et al. SOX17 regulates uterine epithelial-stromal cross-talk acting via a distal enhancer upstream of Ihh. Nat. Commun. 9, 4421 (2018).
Heath, J. K. Cytokines. Can there be life without LIF?. Nature 359, 17 (1992).
Ernst, M. et al. Defective gp130-mediated signal transducer and activator of transcription (STAT) signaling results in degenerative joint disease, gastrointestinal ulceration, and failure of uterine implantation. J. Exp. Med. 194, 189–203 (2001).
Sun, X., Bartos, A., Whitsett, J. A. & Dey, S. K. Uterine deletion of Gp130 or Stat3 shows implantation failure with increased estrogenic responses. Mol. Endocrinol. 27, 1492–1501 (2013).
Fukui, Y. et al. Uterine epithelial LIF receptors contribute to implantation chamber formation in blastocyst attachment. Endocrinology 162, bqab169 (2021).
Nager, A. R. et al. An actin network dispatches ciliary GPCRs into extracellular vesicles to modulate signaling. Cell 168, 252–263.e214 (2017).
Maerz, L. Z. et al. Pharmacological cholesterol depletion disturbs ciliogenesis and ciliary function in developing zebrafish. Commun. Biol. 2, 31 (2019).
Kalucka, J. et al. Single-cell transcriptome atlas of murine endothelial cells. Cell 180, 764–779.e720 (2020).
Andrade, J. et al. Control of endothelial quiescence by FOXO-regulated metabolites. Nat. Cell Biol. 23, 413–423 (2021).
Girbl, T. et al. Distinct compartmentalization of the chemokines CXCL1 and CXCL2 and the atypical receptor ACKR1 determine discrete stages of neutrophil diapedesis. Immunity 49, 1062–1076.e1066 (2018).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e3529 (2021).
Aure, M. H. et al. FGFR2 is essential for salivary gland duct homeostasis and MAPK-dependent seromucous acinar cell differentiation. Nat. Commun. 14, 6485 (2023).
Hambartsoumian, E. Endometrial leukemia inhibitory factor (LIF) as a possible cause of unexplained infertility and multiple failures of implantation. Am. J. Reprod. Immunol. 39, 137–143 (1998).
Cullinan, E. B. et al. Leukemia inhibitory factor (LIF) and LIF receptor expression in human endometrium suggests a potential autocrine/paracrine function in regulating embryo implantation. Proc. Natl. Acad. Sci. USA 93, 3115–3120 (1996).
Wu, M., Yin, Y., Zhao, M., Hu, L. & Chen, Q. The low expression of leukemia inhibitory factor in endometrium: possible relevant to unexplained infertility with multiple implantation failures. Cytokine 62, 334–339 (2013).
Aghajanova, L. et al. Disturbances in the LIF pathway in the endometrium among women with unexplained infertility. Fertil. Steril. 91, 2602–2610 (2009).
Bolnick, A. D. et al. Reduced homeobox protein MSX1 in human endometrial tissue is linked to infertility. Hum. Reprod. 31, 2042–2050 (2016).
Cooke, P. S., Spencer, T. E., Bartol, F. F. & Hayashi, K. Uterine glands: development, function and experimental model systems. Mol. Hum. Reprod. 19, 547–558 (2013).
Maharjan, Y. et al. TMEM135 regulates primary ciliogenesis through modulation of intracellular cholesterol distribution. EMBO Rep. 21, e48901 (2020).
Du, L. et al. Single-cell transcriptome analysis reveals defective decidua stromal niche attributes to recurrent spontaneous abortion. Cell Prolif. 54, e13125 (2021).
Chung, D., Gao, F., Jegga, A. G. & Das, S. K. Estrogen mediated epithelial proliferation in the uterus is directed by stromal Fgf10 and Bmp8a. Mol. Cell. Endocrinol. 400, 48–60 (2015).
Qian, X. et al. N-cadherin/FGFR promotes metastasis through epithelial-to-mesenchymal transition and stem/progenitor cell-like properties. Oncogene 33, 3411–3421 (2014).
Loid, M. et al. Aging promotes accumulation of senescent and multiciliated cells in human endometrial epithelium. Hum. Reprod. Open 2024, hoae048 (2024).
Andersen, A. G., Als-Nielsen, B., Hornnes, P. J. & Franch Andersen, L. Time interval from human chorionic gonadotrophin (HCG) injection to follicular rupture. Hum. Reprod. 10, 3202–3205 (1995).
Turco, M. Y. et al. Long-term, hormone-responsive organoid cultures of human endometrium in a chemically defined medium. Nat. Cell Biol. 19, 568–577 (2017).
Mages, S. et al. TACCO unifies annotation transfer and decomposition of cell identities for single-cell and spatial omics. Nat. Biotechnol. 41, 1465–1473 (2023).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (82288102 to H.W., 82030040 to H.S., 82371680, 82171653 to G.Y., 82301897 to Y.L., 32171117 and 82430052 to W.D., 82222026 and 32270907 to S.K., 82301886 to H.B., 32300954 to Y.T.) and the National Key R&D Program of China (2022YFC2702500 to H.W., 2022YFC2704500 and 2022YFC2704600 to W.D.).
Author information
Authors and Affiliations
Contributions
H.S., H.W., G.Y., W.D., and H.B. designed and supervised the project; Y.L., Y.W., L.Z., Y.T., Y.S., and Y.H. performed experiments and prepared figures; W.D., L.Z., M.L., and M. Wang performed bioinformatics analyses; X.S., J.M., M. Wu, and N.K. collected clinical samples; W.D., Y.L., Y.W., L.Z., and S.K. wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Francesco DeMayo, Anna Wilbrey-Clark, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, Y., Wang, Y., Zhang, L. et al. Decrypting spatiotemporal code of human endometrial receptivity. Nat Commun 17, 801 (2026). https://doi.org/10.1038/s41467-025-67492-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67492-z