Abstract
Biomanufacturing provides a more sustainable alternative to fossil-based chemical manufacturing. 3-Hydroxypropionic acid (3HP) is a top Department of Energy value-added chemical and precursor to bioplastics, yet cost-effective microbial production remains elusive. Here, we establish the acid-tolerant yeast Issatchenkia orientalis as a robust host for low-pH 3HP biosynthesis. Genome-scale modeling identifies the β-alanine pathway as optimal, offering the highest theoretical yield and lowest oxygen requirement. Thermodynamic analysis confirms its favorability under acidic conditions. Using sequence similarity network analysis, we discover highly active aspartate 1-decarboxylase (PAND), β-alanine-pyruvate aminotransferase (BAPAT), and 3HP dehydrogenase (YDFG), which significantly improve the pathway efficiency. Next, to further elevate the production, pathway optimization through multi-copy PAND integration, byproduct elimination (knockouts of pyruvate decarboxylase and glycerol-3-phosphate dehydrogenase), and reinforcement of aspartate flux by overexpression of pyruvate carboxylase and aspartate amino transferase improves the titer to 29 g/L in shake flasks. Fed-batch fermentation at pH 4 with low-cost corn steep liquor medium further increases the production to 92 g/L with 0.7 g/g yield and 0.55 g/L/h productivity. Techno-economic analysis indicates that such performance could potentially enable a financially viable process for sustainable acrylic acid production. This work establishes I. orientalis as a next-generation platform for cost-effective 3HP production and paves the way toward industrial commercialization.
Similar content being viewed by others
Introduction
Biomanufacturing has been increasingly used to produce chemicals and materials from renewable plant biomass1,2,3. 3-Hydroxypropionic acid (3HP) is one of the top 12 sugar-based platform chemicals that were identified by the United States Department of Energy to have great potential for commercial applications4,5,6,7,8. For example, it can be converted to acrylic acid, 1,3-propanediol, methyl acrylate, acrylamide, ethyl 3HP, malonic acid, propiolactone, and acrylonitrile6,9,10. In particular, acrylic acid represents a large market opportunity (approximately US$20 billion), with a global annual demand of approximately 6.6 million tons in 2019 and is expected to grow by more than 5% every year11. Currently, both 3HP and acrylic acid are produced almost exclusively via chemical synthesis routes, which rely on fossil-derived feedstocks and involve energy-intensive processes that contribute to greenhouse gas emissions and environmental pollution9,12,13.
Both bacteria and yeasts have been extensively explored for the biological production of 3HP in the past few decades14. Bacteria such as Escherichia coli have been engineered to produce 3HP with high yields and titers (0.6 g/g yield with a titer of 72 g/L15). However, the inhibitory effects of low pH on bacterial growth necessitates the use of neutralization agents such as calcium carbonate to maintain fermentations at neutral pH, i.e., above the pKa of 3HP (4.5). As a result, an acidification step is necessary to recover 3HP, which leads to the formation of gypsum, rendering downstream processing complex and costly (Supplementary Fig. 1). In contrast, due to their tolerance to low pH, yeasts can be engineered to produce 3HP at low pH, which greatly simplifies the whole 3HP production process and eliminates the formation of gypsum. A previous techno-economic analysis of the theoretical fermentation space suggested a yield of >0.78 g/g combined with a titer of >80 g/L or a yield of >0.62 g/g combined with a titer of >60 g/L would be needed to achieve financial viability under neutral and low-pH regimes, respectively16. However, to the best of our knowledge, the highest titer and yield achieved using a yeast are 100.4 g/L (with a yield of 0.2 g/g)17 and 0.56 g/g (with a titer of 11.25 g/L)18, respectively.
To develop a cost-effective biological process for 3HP production, we seek to engineer a low-pH tolerant yeast Issatchenkia orientalis SD10819 capable of producing 3HP at high titer and yield and establish an end-to-end pipeline consisting of biomanufacturing facility design, simulation, techno-economic analysis (TEA), life cycle assessment (LCA), strain development, downstream processing, and scale-up. We previously developed a genetic toolbox for metabolic engineering of I. orientalis, including episomal plasmids, a library of promoters and terminators with varying strengths, a homologous recombination-based DNA assembly method20, a multiplex CRISPR-based gene deletion system21, and a landing pad system for multiple copy gene integration22. In this work, we leverage this toolbox to identify the rate limiting enzyme in the 3HP biosynthetic pathway, the PAND enzyme which catalyzes the transformation of aspartate to β-alanine. We then introduce up to eight copies of the PAND gene, block the ethanol and glycerol production pathways, and overexpress pyruvate carboxylase (PYC) and aspartate amino transferase (AAT) enzymes. The final strain can produce 29 g/L of 3HP with a yield of 0.58 g/g in shake flask fermentation. Next, we perform fed-batch fermentation at the lab scale, which results in a titer of 92 g/L and a 0.7 g/g glucose yield. Future efforts directed toward scaling up, integrating downstream processing (DSP), and utilizing alternative renewable feedstocks are expected to further enhance the economic feasibility of the process and substantially reduce environmental impacts compared with conventional fossil-based production routes.
Results
Selection of 3HP biosynthetic pathway in I. orientalis
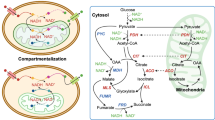
For host selection, we chose I. orientalis SD108 for 3HP production because of its high tolerance to 3HP (up to 125 g/L, Supplementary Fig. 2 at initial pH of 5.6). For pathway selection, we employed a genome-scale metabolic (GSM) model of I. orientalis23 augmented separately with reactions capable of producing 3HP by seven previously reported pathways24, including β-alanine and malonyl-CoA pathways. Systematic evaluation revealed that in context of an I. orientalis background, the β-alanine pathway has both the highest maximum theoretical yield of 3HP (i.e., carbon yield from glucose of 99.7% or 0.986 g 3HP/g glucose) as well as the most favorable (i.e., lowest) minimal oxygen uptake requirements at high yield (i.e., only 0.018 mmol/mmol glucose) (Fig. 1). The reactions of this pathway, provided in Table 1, starting from the conversion of pyruvate requires three non-native enzymes: PAND (aspartate 1-decarboxylase), BAPAT (β-alanine-pyruvate aminotransferase), and YDFG (3HP dehydrogenase). Subsequently, we performed thermodynamics analysis of this pathway since external pH is known to affect the intracellular pH of Saccharomyces cerevisiae25 and pH can impact thermodynamics. Table 1 gives the change in Gibbs free energy for each reaction step using dGPredictor, a recently developed moiety-based automated fragmentation tool that has improved goodness of fit over group contribution (GC) methods26, at pH 7 and 6 and an assumed ionic strength of 0.1 M (see Supplementary Table 1 for computations using eQuilibrator27). For pH 6, two non-native steps are substantially more negative (i.e., favorable) than at pH 7. Notably, though, for pH 6 the net change becomes more positive at the aspartate transaminase step and less negative at the pyruvate carboxylase step than they are at pH 7. By shifting the equilibrium through accumulation of reactants and fast removal of products (i.e., the latter by increased PAND levels), aspartate transaminase could nevertheless proceed in the desired direction. Therefore, we decided to assemble and express the β-alanine pathway in I. orientalis for 3HP production.
Pathway numbers, indicated by roman numerals, follow the numbering in Table 2 in a previous study7, i.e., I: pyruvate → lactate → lactoyl-CoA → acryloyl-CoA → 3HP-CoA → 3HP; II: pyruvate → acetyl-CoA→ malonyl-CoA → 3-oxopropanoate → 3HP; III: pyruvate → oxaloacetate→ aspartate → β-alanine → 3-oxopropanoate→ 3HP; IV: pyruvate → oxaloacetate→ aspartate → β-alanine → β-alanyl-CoA → acryloyl-CoA→ 3HP-CoA → 3HP; V: pyruvate → succinate → propionyl-CoA → acryloyl-CoA → 3HP-CoA → 3HP; VI: pyruvate → α-alanine → β-alanine → 3-oxopropanoate → 3HP; VII: pyruvate → α-alanine → β-alanine → β-alanyl-CoA → acryloyl-CoA → 3HP-CoA → 3HP. Pathway III corresponds to the β-alanine pathway in the current work, with superscript p indicating NADPH utilization. Biomass yield (left) and oxygen uptake (right) envelopes were generated using the genome-scale metabolic model iIsor85023 with reactions added as needed to enable 3HP production by the indicated pathway, and with the indicated gene deletions. Shading in the biomass yield denotes the corresponding minimal oxygen uptake rate.
Construction of efficient 3HP biosynthetic pathway in I. orientalis
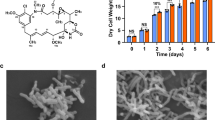
To design an optimal β-alanine pathway in I. orientalis, we used the sequence similarity network (SSN) analysis tool28 to identify up to 10 different genes for each of the three genes in the pathway. The template PAND, BAPAT and YDFG genes were from a previous report in which S. cerevisiae was engineered to produce 3HP via the β-alanine pathway29. At the beginning, we designed an in vivo strategy in which the screening strains were integrated with template BAPAT and template YDFG or template PAND and template YDFG or template PAND and template YDFG for screening candidate PAND or candidate BAPAT or candidate YDFG, respectively. Our first strategy could only allow us to effectively screen the best PAND, but not for BAPAT and YDFG (Supplementary Figs. 3, 4, and 5), which indicated that the first step was significantly rate-limiting due to limited supply of the precursors for BAPAT and YDFG. For the PAND gene, we also started another screening assay by directly measuring the intracellular β-alanine accumulation using the recombinant I. orientalis harboring different candidate PAND-encoding plasmids in SC-URA with 50 g/L glucose after two days. As shown in Fig. 2A, the hypothetical protein GE061_003755-Apolygus lucorum (i.e., P5), gave us the highest production of β-alanine, which was consistent with the result obtained from the in vivo test (Supplementary Fig. 3). For characterizing the other two enzymes, BAPAT and YDFG, we chose an in vitro enzymatic reaction, in which the cell lysate with enzymes produced from I. orientalis were directly mixed with the enzyme substrate (Fig. 2B). For BAPAT, we adopted a previously developed assay by Borodina et al.29, which used colorimetry to quantify the formation of an aldehyde product using Purpald as a purple-color-forming agent (Supplementary Fig. 6). We found that the protein aminotransferase class III-fold pyridoxal phosphate-dependent enzyme from Methanosarcina sp. MTP4 (i.e., B6), gave the best result (Fig. 2C). For YDFG, due to the lack of commercially available malonic semialdehyde and the high cost of NADPH, we decided to couple the best BAPAT enzyme and glucose dehydrogenase (GDH) to synthesize malonic semialdehyde and NADPH for the YDFG catalyzed reaction. The results shown in Fig. 2D indicate that the bifunctional NADP-dependent 3-hydroxy acid dehydrogenase/3-hydroxypropionate dehydrogenase YDFG from Erwinia mallotivora (i.e., Y3) produced the most 3HP.
A β-alanine accumulation in the strain overexpressing the PAND candidate genes. B in vitro assay for the BAPAT and YDFG genes. C OD540 measurement for in vitro BAPAT assay. D 3HP measurement for in vitro YDFG assay. Data present mean ± S.D. of three biologically independent samples (n = 3). Source data for this Figure is available in the Source Data file. Created in BioRender. Zhao, H. (2025) https://BioRender.com/vkz7w01.
Improving 3HP production via metabolic engineering
To improve the production of 3HP in I. orientalis, we explored various metabolic engineering strategies (Fig. 3). The strain containing an episomal plasmid, harboring β-alanine pathway consisting of the best genes selected from the section of construction of efficient 3HP biosynthetic pathway in I. orientalis, referred to as PBY, produced 2.5 g/L of 3HP (Fig. 3B, column 2) compared to 0.86 g/L (Fig. 3B, column 1) from the strain harboring the β-alanine pathway consisting of the original PAND, BAPAT and YDFG genes. We posited that introducing more copies of the β-alanine pathway integrated into the genome along with an episomal plasmid harboring PBY would increase production. The resulting strain pPBY/PBY3/ura3Δ (3 copies of the pathway integrated in the genome with PBY on an episomal plasmid) led to an increase in 3HP production, reaching 5.81 g/L (Fig. 3B, column 5). To identify the rate-limiting step, we introduced individual episomal plasmids for the PAND gene, the BAPAT gene, the YDFG gene, and all three genes (PBY) into the strain PBY2/ura3Δ (Fig. 3B, column 6 to column 8). Only the strain harboring the plasmid with the PAND gene could produce 3HP as the strain harboring the plasmid with all three genes, indicating PAND was the rate-limiting step. We further investigated the effect of the copy number of the PAND gene on 3HP production by integrating up to eight copies into the LPB/Δura3 strain, our landing pad strain22 that allows multiple gene integrations at a time. This also circumvented generating redundant expression of the BAPAT and YDFG genes. This led to a substantial increase in the 3HP titer from 3.2 g/L for strain pPBY/1 P/ura3Δ (1 copy of PAND, Fig. 3B, column 9) to 8.4 g/L for pPBY/8 P/ura3Δ (8 copies of PAND, Fig. 3B, column 16), despite a drop to 5.5 g/L with five to seven copies of PAND. qPCR analysis (Supplementary Method 1) confirmed that the expression level of PAND increased with increasing copy number (Supplementary Fig. 7, yellow bar), but the 3HP production did not continually increase. Further investigation of the PYC gene revealed its elevated expression levels at three and eight PAND copies (Supplementary Fig. 7, orange bar), suggesting that integration at the EG6 site activated PYC, boosting the production. The data indicated that the native aspartate flux supported only three to four PAND copies.
A 3HP production using the β-alanine pathway. GPD, glycerol-3-phosphate dehydrogenase; PDC, pyruvate decarboxylase; PYC, pyruvate carboxylase; PAND, aspartate decarboxylase; BAPAT, β-alanine aminotransferase; YDFG, 3HP dehydrogenase. OAA, oxaloacetate; ASP, aspartate; β-ala, β-alanine; MSA, malonic semialdehyde, 3HP, 3-hydroxypropionic acid. B 3HP production using various engineered strains. Episomal plasmids PBY, P, B and Y mean the strains harbored an episomal plasmid encoding PBY, P, B and Y, respectively. Horizontal hatch marks are for comparison of the original gene set (column 1) and new gene set from SSN (column 2). Right diagonal hatch marks are for discovering the copy number effect and rate-limiting step. Left diagonal hatch marks are for PAND copy effect. The empty bars reflect further metabolic engineering strategies, including knocking out the PDC and GPD genes, integrating all pathway genes, and overexpressing the PYC and AAT genes. Data present mean ± S.D. of at least three biologically independent samples (n = 3) unless specified. For column 3 and 4, data present mean ± S.D. of five biologically independent samples (n = 5). For column 5, data present mean ± S.D. of seven biologically independent samples (n = 7). Source data for this Figure is available in the Source Data file. Created in BioRender. Zhao, H. (2025) https://BioRender.com/vkz7w01.
To further enhance the 3HP production, we knocked out the PDC gene in the pPBY/8 P/ura3Δ strain to generate pPBY/8 P/ura3Δ/pdcΔ, and the 3HP titer significantly increased from 8.4 g/L to 17.4 g/L (Fig. 3B, column 17). Next, to further reduce the glycerol byproduct, we knocked out GPD (glycerol-3-phosphate dehydrogenase) in the PDC knock-out strain, leading to a production of 18.6 g/L (Fig. 3B, column 18). Both ethanol and glycerol were also analyzed for three strains, including pPBY/8 P/ura3Δ, pPBY/8 P/ura3Δ/pdcΔ, and pPBY/8 P/ura3Δ/pdcΔ/gpdΔ. With PDC knockout, the maximum ethanol accumulation reduced from 3.0 g/L to 0 g/L and surprisingly the maximum glycerol accumulation also reduced from 6.9 g/L to 3.6 g/L, which explained the significant increase of 3HP titer after PDC knockout. With double knockout of PDC and GPD, there was no accumulation of ethanol and glycerol (Supplementary Fig. 8). Predictions made using the GSM model to simultaneously knock out both GPD and PDC while using the β-alanine pathway had no decrease in the maximum theoretical carbon yield of 3HP or substantial change in oxygen uptake rate (Fig. 1). Based on the double knock-out strain, we introduced one copy of PYC-AAT to further enhance the carbon flux toward aspartate which is an important precursor in the β-alanine pathway. After integration of one copy of PYC-AAT, the strain pPBY/8P-AP/ura3Δ/pdcΔ/gpdΔ produced 22 g/L of 3HP (Fig. 3B, column 19). Next, considering that an industrially relevant strain should have all genes integrated into the chromosome to improve genetic stability, we decided to integrate the BAPAT and YDFG genes to the chromosome to generate the 1AP/ura3∆ strain and cultured it in synthetic complete medium (CSM medium), which produced 25 g/L of 3HP (Fig. 3B, column 20). Because the BAPAT and YDGF genes were provided from a two-copy episomal plasmid in the previous section, we transformed the episomal plasmid pPBY and empty episomal plasmid back to the 1AP/ura3∆ strain to evaluate the gene copy number effect from BAPAT and YDFG. However, the results showed that there is no difference in production (Supplementary Fig. 9). To further increase the yield, another copy of PYC-AAT was integrated, which resulted in a titer of 29 g/L (Fig. 3B, column 21). After integrating a second copy of PYC-AAT, the resulting strain, 2AP/ura3∆, consumed glucose slightly slower than the 1AP/ura3∆ strain (Supplementary Fig. 10A), but the pyruvate accumulation reduced from 2.6 g/L to 0.86 g/L (Supplementary Fig. 10B).
Fed-batch fermentation
Next, we performed fed-batch fermentation to further increase the titer and yield of 3HP production. During fed-batch fermentation, a pH of 4 was maintained, which eliminated the acidification step for downstream processing. Moreover, dissolved oxygen (DO) was maintained at 5% to create a microaerobic environment, based on the computational modeling studies as illustrated in Fig. 1, to increase 3HP production. We utilized CSL (Corn Steep Liquor) as the fermentation media to make this fermentation industrially feasible and lower cost. To culture the strain in CSL, we recovered the ura3 gene in the 2AP/ura3∆ strain to generate the 2AP strain since the 2AP/ura3∆ strain grew very slowly in pure CSL medium but with uracil supplement, 2AP/ura3∆ could grow similarly as the 2AP strain in pure CSL (Supplementary Fig. 11). The 2AP strain consumed 50 g/L of glucose, accumulated up to 15 g/L of pyruvate and produced 18 g/L of 3HP in CSM medium by day 3. From day 3 to day 5, the 2AP strain consumed pyruvate to produce 3HP, which reached 26 g/L and a yield of 0.52 g/g on day 5 (Fig. 4A–C). As for the CSL medium, the 2AP strain consumed 50 g/L glucose in 9 days and accumulated only 0.7 g/L pyruvate with a 3HP production of 32 g/L (Fig. 4A–C). Lower pyruvate accumulation in CSL explains why the 2AP strain produced more 3HP in CSL because pyruvate is an important precursor in the β-alanine pathway. Overall, although CSL medium reduced the glucose uptake rate, pyruvate did not accumulate and the 3HP yield was increased to 0.64 g/g glucose. The 2AP strain was used for all fed-batch fermentations. To demonstrate the maximum performance of 2AP, we performed the fed-batch fermentation in 300 mL DASbox system with CSL medium, DO set to 5%, pH of 4, and CO2 gassing, which resulted in a titer, yield, and productivity of 92 g/L, 0.7 g/g, and 0.55 g/L/h, respectively (Fig. 4D). Feeding was performed manually at each scale in pulse mode and was dependent on the concentration of glucose at the time of sampling (Supplementary Table 2). For the cell growth, the OD600 (optical density) along with the fermentation time was shown in Supplementary Fig. 12. Also, during the whole period of fermentation, no other metabolites at concentrations greater than 1 g/L were observed. We also attempted the preliminary scale-up study to 3 L scale and showed that the yield also reached 0.7 g/g with titer of 74.5 g/L. (Supplementary Fig. 13).
The profile of the 2AP strain for glucose consumption (A), pyruvate accumulation (B), and 3HP production (C) in both the CSM medium and the CSL medium in shake flask. Data present mean ± S.D. of three biologically independent samples (n = 3). D fed-batch fermentation a 300 mL. Glucose feeding began at 24 h post inoculation. The 2AP strain was used for fermentation. During fermentation, pyruvate, and glucose were monitored. No pyruvate accumulation or other metabolites greater than 1 g/L were observed. The cultivation was set at DO ranging from 5% to 15% and pH at 4 with CO2 gassing. For fed-batch fermentation, two independent biological replicates were performed (n = 2) and mean value from two replicates is presented. Source data for this Figure is available in the Source Data file. Created in BioRender. Zhao, H. (2025) https://BioRender.com/vkz7w01.
Financial viability and environmental benefits under uncertainty
Finally, we designed and simulated end-to-end biomanufacturing facilities for 3HP that could accept dextrose monohydrate as a feedstock (as well as corn, sugarcane, and corn stover), ferment to 3HP using I. orientalis, purify and concentrate the fermentation broth (to 30 wt% 3HP), catalytically upgrade 3HP to acrylic acid, and recover glacial acrylic acid of >99.5 wt% purity (Supplementary Fig. 14) at an annual production capacity of 134,000 metric tons of acrylic acid. We also performed sensitivity analyses using Spearman’s rank order correlation coefficients (Spearman’s ρ) to identify key drivers of production costs and life cycle carbon intensity (CI), and designed and simulated the biomanufacturing facility across the theoretical fermentation space (i.e., 2500 possible yield-titer combinations at the baseline productivity of 0.548 g/L/h as well as 100 possible productivity values at the baseline yield-titer combination of 0.694 g/g and 92 g/L) for each feedstock under both low-pH fermentation (i.e., fermentation pH of 4 and no acidulation required after fermentation) and neutral fermentation regimes (i.e., neutralization by base addition during fermentation and complete re-acidulation after fermentation) to prioritize research and development pathways to advance system sustainability.
For the simulated dextrose-based 3HP biomanufacturing based on experimental performance in the DASbox, the biomanufacturing facility could produce acrylic acid at an estimated minimum product selling price (MPSP) of $1.36/kg (baseline) with a range of $1.21 − 1.58/kg [5th-95th percentiles, hereafter in brackets] and a CI of 4.43 kg CO2-eq./kg [4.01–5.10 kg CO2-eq./kg; Fig. 5A, B]. Integrating the end-to-end pipeline with an on-site corn dry-grind process would substantially improve the biomanufacturing’s financial viability (MPSP of $1.13/kg [$1.01 − 1.27/kg], below the acrylic acid market price range in 100% of simulations) and reduce CI (3.26 kg CO2-eq./kg [2.78 − 3.92 kg CO2-eq./kg], below the fossil-based acrylic acid CI range in 98.5% of simulations). Biomanufacturing facilities designed to accept sugarcane could unlock even greater environmental benefits from the production of excess electricity by the turbogenerator, offsetting marginal grid impacts to achieve a CI of 2.12 kg CO2-eq./kg [1.49 − 2.76 kg CO2-eq./kg].
Uncertainties (box-and-whisker plots) for (A) minimum product selling price (MPSP) and (B) cradle-to-grave carbon intensity (CI) quantified as 100-year global warming potential per kg of acrylic acid produced by catalytic upgrading of 3HP produced via microbial conversion of sugars by I. orientalis. Box-and-whisker plots show results for four scenarios (markers report results for baseline values): (i) fermentation performance achieved in the DASbox using dextrose (dextrose, diamond markers); (ii) corn-based 3HP biomanufacturing under the same assumptions for fermentation performance as that achieved in the DASbox (corn, square markers); (iii) sugarcane based 3HP biomanufacturing under the same assumptions for fermentation performance as that achieved in the DASbox (sugarcane, triangle markers); and (iv) corn stover based 3HP biomanufacturing under the same assumptions for fermentation performance as that achieved in the DASbox (corn stover, pentagon markers; baseline values and distributions for all parameters included in the uncertainty analysis for each scenario are detailed in Supplementary Data 1). On box-and-whisker plots, whiskers, boxes, and the middle line represent 5th/95th, 25th/75th, and 50th percentiles, respectively, from 6000 Monte Carlo simulations for each scenario. C, E MPSP and (D, F) CI for dextrose-accepting biomanufacturing across 2500 fermentation yield-titer combinations at the baseline productivity (0.548 g/L/h) for neutral (middle panels; C, D) and low-pH (right panels; E, F) fermentation. Yield is shown as % theoretical maximum (%theoretical) scaled to the theoretical maximum yield of 1.00 g/g-glucose-equivalent (based on carbon balance). Diamond markers report results for baseline values for the dextrose scenario. Shaded gray and hatched regions show (A, C, E) market price range for acrylic acid ($1.40–1.65/kg)35 and (B, D, F) a range bounded by CI values reported for fossil-based acrylic acid36,37 (end-of-life impact of 1.83 kg CO2-eq./kg was added where the reported value was indicated as cradle-to-gate, assuming degradation of produced acrylic acid entirely by passive oxidation to CO2). For a given point on the contour plots (C, E, D, F), the x-axis value represents yield, the y-axis value represents titer, and the color and contour lines represent the value of MPSP (C, E) or CI (D, F). Tabulated data breaking down capital and material costs, utilities, and CI for each scenario are available online50. Source data for this Figure is available in the Source Data file.
Simulations across the theoretical fermentation space demonstrated improving yield in low-pH fermentation will have the most direct benefit to MPSP (Fig. 5C, E), which was consistent with MPSP being most sensitive to fermentation yield among 33 uncertain parameters (Spearman’s ρ of −0.69; Supplementary Fig. 15). Regardless of the feedstock used, the biomanufacturing facility’s MPSP benefited consistently from increased yield and titer values for both neutral and low-pH fermentation in the evaluated theoretical fermentation space (Fig. 5C,E; Supplementary Figs. 16–18), with potential minimum values of $1.39/kg for neutral fermentation and $1.02/kg for low-pH fermentation for dextrose-accepting facilities at the baseline productivity of 0.548 g/L/h. The benefits of a higher titer (e.g., lower heating demands from separation units) generally outweighed the increased expenses (e.g., larger fermentation reactors at a fixed 3HP productivity), but the magnitude of the net benefits diminished with increasing titer. Similarly, at higher yield values, further improvements to yield had diminishing benefits for MPSP. The relative benefits to MPSP from comparable relative improvements to yield and titer depended on the location in the theoretical fermentation space. For example, at the baseline fermentation yield (69.4% of the theoretical maximum) and titer (92.0 g/L), improving yield by 6.9% (a 10% relative increase) would decrease the MPSP by $0.06/kg, while improving titer by 9.2 g/L (a 10% relative increase) would decrease the MPSP by only $0.03/kg (Fig. 5E). The biomanufacturing facility’s MPSP, CI, and fossil energy consumption (FEC) were relatively insensitive to potential improvements to fermentation productivity (e.g., decreasing by $0.09/kg, 0.01 kg CO2-eq./kg, and 0.2 MJ/kg, respectively, with a 400% increase from the baseline productivity to 2.74 g/L/h; Supplementary Fig. 19). While the biomanufacturing facility’s CI and FEC were most substantially sensitive to 3HP titer (Spearman’s ρ of −0.78 and −0.73, respectively; Supplementary Fig. 15) and consistently benefited from titer improvements (Fig. 5F, Supplementary Fig. 20B), 3HP yield improvements at high yield values could actually increase CI and FEC. This was because the yield of I. orientalis cell mass was assumed to decrease with 3HP yield improvements, resulting in lower-energy wastes being diverted to the boiler for combustion. The resulting decrease in steam production necessitated the purchase of larger amounts of natural gas to satisfy the biomanufacturing facility’s heating, cooling, and power utility demands. For the simulated dextrose-based 3HP biomanufacturing based on experimental performance in the 3 L bioreactor (yield of 0.703 g/g, titer of 74.5 g/L, and productivity of 0.776 g/L/h), the MPSP did not change substantially ($1.39/kg) but the CI and FEC increased to 4.87 kg CO2-eq./kg and 54.06 MJ/kg, respectively, due to the decreased titer relative to that achieved in the DASbox (92.0 g/L). This demonstrates improving fermentation titer will have the most direct benefits to CI and FEC. However, high 3HP yields will be necessary to unlock the high titers needed for environmental benefits, with potential minima of 2.96 kg CO2-eq./kg and 31.8 MJ/kg for CI and FEC, respectively, for low-pH fermentation in dextrose-accepting biomanufacturing facilities (Fig. 5F, Supplementary Fig. 20 and Supplementary Method 2).
Discussion
Despite decades of effort, financially viable biomanufacturing of 3HP from renewable plant biomass remains a significant challenge. In this study, we engineered I. orientalis, which is renowned for its tolerance to low pH conditions and has previously been engineered to produce a variety of organic acids30,31,32,33, to overproduce 3HP to overcome this challenge. 3HP can be synthesized from at least four different metabolic intermediates: glycerol, lactate, malonyl-CoA, and β-alanine. The glycerol route is challenging in yeast because glycerol dehydratase is dependent on the co-factor vitamin-B12 which is not natively synthesized. The lactate route is likely to result in a mixture of lactate and 3HP. Previous examinations identified both thermodynamically favorable (i.e., β-alanine and malonyl-CoA) and unfavorable (i.e., acryloyl-CoA) pathway alternatives, but were performed at neutral pH24,29,34. The overall observations from our thermodynamic analysis suggest that increased PAND activity could offset the small reductions in favorability of the β-alanine pathway occurring through acidification of the intracellular space during 3HP production at low external pH. Moreover, the analysis of the GSM model highlights the high theoretical yields alongside low and nearly monotonic oxygen uptake unique to the β-alanine pathway, further suggesting that industrial processes employing micro-aerobic conditions can potentially reach high yields.
To achieve high-level 3HP production, we designed an optimal β-alanine pathway by screening pathway genes from different organisms. Although we successfully discovered a highly efficient PAND enzyme, PAND was still a significant rate-limiting step. Thus, by leveraging our recently developed landing pad strain22, we were able to integrate 8 copies PAND gene in two rounds of transformation and the resulting strain (i.e., 8 P strain) can reach a significantly higher titer (i.e., from 3.2 g/L to 8.4 g/L). We also increased the 3HP production by knocking out the PDC/GPD genes, integrating all pathway genes, and overexpressing the PYC-AAT genes. Knocking out the PDC/GPD genes reduced the byproduct formation, thus enhancing the yield. Integration of BAPAT and YDFG circumvented the use of an unstable episomal plasmid thus an enhancement of production was observed. To further pull the flux toward β-alanine, overexpression of PYC-AAT significantly increased 3HP production. Notably, we still have not encountered any cofactor limitations as only one NADPH is required to produce one molecule of 3HP through the β-alanine pathway, whereas the malonyl-CoA pathway requires two molecules of NADPH per molecule of 3HP. Our final strain achieved the highest titer, yield and productivity among the reported engineered yeast strains. In terms of titer, yield, and productivity, if using bacteria as hosts, a higher titer and productivity could be reached (Supplementary Table 3). The best heterologous bacterial host of 3HP production, E. coli, reached 72.2 g/L of 3HP with the high productivity of 1.64 g/L/hr. Although the yield was acceptable (0.6 g/g)15, the fermentation process is still not financially viable because 0.85 g/g yield was required to be economically viable in neutral pH fermentation16. In general, bacteria as a host can reach higher productivity due to faster bacterial growth, but fermentation requires the maintenance of pH at 7, which increases the expense of DSP. A better way to reach industrial scale production, therefore, is to perform low pH fermentation using yeast as a host. Many efforts have been made in yeast hosts previously. For yeast as a host, two main pathways were chosen, including the β-alanine pathway and the malonyl-CoA pathway. For the malonyl-CoA pathway when used in yeast as a host, the best titer was 100.4 g/L using a non-model yeast, Y. lipolytica, but the yield was low (0.2 g/g)17. A recent study used an engineered S. cerevisiae that produced 3HP at an industrially relevant yield of 0.6 g/g; however, no fed-batch fermentation was included in the study18. As for the β-alanine pathway in yeast, the pioneering work was conducted in S. cerevisiae29, however, the performance was lower than the published results using the malonyl-CoA pathway. Although these studies performed fermentation near acidic conditions (i.e., pH 5.0, pH 5.6 or no control), none of the existing studies performed their fermentations below the pKa value of 3HP (i.e., 4.5), which resulted in costly DSP due to acidification post fermentation. In our study, we performed the fermentation at pH 4 and showed the robustness of our final strain to produce 3HP at high titer and yield in a low pH condition. To the best of our knowledge, our study represents the highest reported 3HP yield among all engineered yeast hosts.
Furthermore, we performed TEA and LCA to determine the financial viability and environmental benefits of 3HP biomanufacturing facilities that leverage our developed fermentation process, purify the produced 3HP, and catalytically upgrade it to acrylic acid. Using dextrose as a substrate, the MPSP remained below the high end of the reported market price range ($1.65/kg)16,35 in 99.0% of simulations, while the facility’s CI (4.01–5.10 kg CO2-eq./kg) was comparable to reported values for fossil-based acrylic acid (4.07 − 5.96 kg CO2-eq./kg36,37) and the FEC was on the low end of the fossil-based range (Supplementary Fig. 21). Low-pH fermentation had consistently lower MPSP, CI, and FEC values than neutral fermentation at the same yield, titer, and productivity combinations (Fig. 5, Supplementary Figs. 19–21) due to reduced base and acid requirements (Supplementary Data 1), and continued improvements to 3HP yield and titer combined with on-site conversion of biomass feedstocks to sugars represent the greatest opportunities to reduce system costs and environmental impacts. Sugarcane can unlock further, substantial environmental benefits over dextrose, corn stover, and corn, enabling the sustainable production of bio-based 3HP and acrylic acid.
In conclusion, we engineered a low-pH tolerant non-model yeast I. orientalis strain enabling cost-effective production of 3HP from CSL, glucose and CO2 at low pH condition. We used TEA and LCA to assess this 3HP biomanufacturing process and found our process was financially viable at bench scale with the potential for reduced life cycle carbon intensity and fossil energy consumption relative to fossil-derived acrylic acid. Overall, this work establishes I. orientalis as a next-generation host and provides a clear roadmap for scaling and integrating DSP toward industrial 3HP and acrylic-acid production.
Methods
Strains, media, and materials
All the strains and plasmids were listed in Supplementary Table 4 and Supplementary Table 5, respectively. E. coli DH5α was used to maintain and amplify plasmids. I. orientalis SD108, and S. cerevisiae YSG50 were propagated in Yeast-Peptone-Adenine-Dextrose (YPAD) medium consisting of 1% yeast extract, 2% peptone, 0.01% adenine hemisulphate, and 2% glucose. Recombinant I. orientalis strains were grown in Synthetic Complete (SC) dropout medium lacking uracil (SC-URA). LB broth, bacteriological grade agar, yeast extract, peptone, yeast nitrogen base (w/o amino acid and ammonium sulfate), ammonium sulfate, and D-xylose were obtained from Difco (BD, Sparks, MD), while complete synthetic medium was purchased from MP Biomedicals (Solon, OH). All restriction endonucleases, Q5 DNA polymerase and Phusion polymerase were purchased from New England Biolabs (Ipswich, MA). The QIAprep spin mini-prep kit and RNA isolation mini kit were purchased from Qiagen (Valencia, CA), whereas Zymoclean Gel DNA Recovery Kit and Zymoprep Yeast Plasmid Miniprep Kits were purchased from Zymo Research (Irvine, CA). All other chemicals and consumables were purchased from Sigma (St. Louis, MO), VWR (Radnor, PA), and Fisher Scientific (Pittsburgh, PA). Oligonucleotides including gBlocks and primers (Supplementary Data 2–5) were all synthesized by Integrated DNA Technologies and Twist Biosciences (IDT, Coralville, IA; Twist Biosciences, San Francisco, CA). For plasmid construction, Gibson assembly or DNA assembly was used. For strain construction, the CRISPR method was used for gene integration/deletion and the lithium acetate-mediated method was used for transformation. Detailed plasmid construction, strain construction, and gene selection are described in the Supplementary Methods 3, 4 and 5, respectively.
PAND gene screening assay
Plasmids encoding different PAND genes were transformed into I. orientalis SD108/ura3Δ. Three colonies were picked into 2 mL SC-URA with 50 g/L of glucose and cultivated for one day and then transferred to 20 mL SC-URA and cultivated for two days at 30 °C with shaking at 250 rpm. All cells were collected and adjusted to OD600 of 100. The cells were disrupted by the glass beads method. For glass beads disruption, cells were lysed using FastPrep-24 5 G for 20 s (6.0 m/s) and the lysed cycle was repeated three times. In each interval, the sample was placed on ice for 2 min cooling. The cell extract was derived by FMOC first and then subjected to amino acid analysis to quantify the β-alanine production. For sample preparation for amino acid analysis, 370 µL of 500 mM Borate (pH 10) and 20 µL of FMOC (2.5 mg/mL in acetonitrile) were added to the 110 µL sample. The mixture was vortexed and stood at room temperature for at least 20 min, followed by centrifugation and filtering.
BAPAT gene screening assay
Plasmids encoding different BAPAT genes were transformed into I. orientalis SD108/ura3Δ. Three colonies were picked into 2 mL SC-URA with 50 g/L of glucose and cultivated for one day and then transferred to 20 mL SC-URA and cultivated for three days at 30 °C with shaking at 250 rpm. All cells were collected and adjusted to OD600 of 100. The concentrated cells were disrupted by the glass beads method. To set up the in vitro enzyme assay, in 1 mL reaction, 900 µL of solution containing 11.1 mM of pyruvate, 11.1 mM of β-alanine, 0.11 mM of PLP, and 55.6 mM of phosphate buffer (pH7) were mixed with 100 µL of cell extract, and the reaction mixtures were incubated at 37 °C for an hour. Then, 25% TCA was added to quench the reaction. To quantify the malonic semialdehyde production, 110 µL of the reaction mixture was mixed with 140 µL of 68 mM Purpald (4-amino-3-hydrazino-5-mercapto-1,2,4-triazole), and the purple product was measured at 540 nm using a plate reader (Molecular Devices, San Jose, CA).
YDFG gene screening assay
Plasmids encoding different YDFG genes were transformed into I. orientalis SD108/ura3Δ. Three colonies were picked into 2 mL SC-URA with 50 g/L of glucose for one day and then transferred to 20 mL SC-URA for three days at 30 °C with shaking at 250 rpm. All cells were collected and concentrated to OD600 of 100. Then, the cells were disrupted by the glass beads method. To set up the in vitro enzyme assay, in 1 mL reaction, 800 µl of solution containing 37.5 of mM pyruvate, 37.5 mM of β-alanine, 0.125 mM of PLP, 6.25 U of GDH, 2.5 mM of NADP+, 125 mM of glucose and 62.5 mM of phosphate buffer (pH 7) were mixed with 100 µL of YDFG-containing cell lysate and 100 µL of B6 cell lysate containing the BAPAT6 protein from Methanosarcina sp. MTP4. The reaction mixture was incubated at 37°C for 2 hours. Then, 25% TCA was added to quench the reaction. 3HP was measured by HPLC.
Tube, shake-flask fermentation and DASbox fed-batch fermentation
For tube fermentation (2 mL), three colonies were picked into SC-URA with 50 g/L glucose and cultured for 5 days. 3HP was quantified at day 5. For shake flask fermentation (20 mL) using plasmid-harboring strain, three colonies were picked into 2 mL SC-URA with 50 g/L of glucose for 3 days at 30 °C with shaking at 250 rpm. Then, the cells were subcultured to 20 mL SC-URA with 50 g/L of glucose and 10 g/L of CaCO3 and cultured at 30 °C with shaking at 250 rpm. For shake flask fermentation using all-integrated strains (i.e., 1AP and 2AP strain), three colonies were picked in YPAD for 1 day. Then, the cells were subcultured into 20 mL CSM with 50 g/L of glucose and 10 g/L of CaCO3. The initial OD600 was 0.2 and, the samples were collected every 24 hours for HPLC analysis. For fed-batch fermentation in the DASbox bioreactors (DASbox Bioreactors, Eppendorf, Hamburg, Germany, 100 mL working volume, Supplementary Table 6), single colonies of the I. orientalis strain were inoculated into the YPAD medium and cultured for 1 day. Then, 1 mL of cells was injected into 100 mL of CSL liquid medium in the bioreactor. The cells were cultivated at 30 °C and the DO was set to 5%. A cascade program was used to maintain DO by changing the stirrer speed between 400-1200 RPM, O2 concentration in the gas mix between 21-100% pure O2 and then O2 flow rate between 1–3 vvm while maintaining the pH at 4 using 4 M HCl and 28% ammonium hydroxide. When the DO dropped below 5%, CO2 was introduced at 0.2 vvm and conjointly aeration dropped to 0.2 vvm. Therefore, total aeration remained at 0.4 vvm and at this point DO was maintained using agitation only. The CO2 supply could provide an ample amount of bicarbonate for 3HP production.
Analytic methods
For quantification of glucose consumption and ethanol and glycerol productions, Agilent 1200 HPLC system equipped with a refractive index detector (Agilent Technologies, Wilmington, DE, USA) and Rezex ROA-Organic Acid H+ (8%) column (Phenomenex, Torrance, CA, USA) were used. The column and the detector were operated at 50 °C, and 0.005 N H2SO4 was used as the mobile phase at a flow rate of 0.6 mL/min. For quantification of 3HP production and pyruvate accumulation, Shimadzu HPLC with a UV-Vis detector and Aminex HPX-87H Column #1250140 (BioRad, USA) were used with injection of 50 µL sample. The column was operated at 60 °C with 0.005 N H2SO4 at a flow rate of 0.6 mL/min and 210 nm was used as the detection wavelength. For quantification of amino acids, Agilent 1260 HPLC system equipped with a DAD detector (262 nm was used as detecting wavelength) and Phenomenex Kinetex® 5 µm EVO C18 100 Å LC column (150 × 4.6 mm2; Phenomenex) were used with injection of 5 µL sample. The column was run at room temperature with a gradient elution process using solvent A (20 mM sodium acetate) and solvent B (acetonitrile). The elution process runs the following program: 18% B to 57% B (linear gradient, 0–13 min), 57% to 100% (linear gradient, 13–13.3 min), 100% B (isocratic elution, 13.3 to 14.3 min), 100% to 18% (linear gradient, 14.3–15.8 min), 18% B (isocratic elution, 15.8 to 18 min).
Genome-scale metabolic modeling, flux balance analysis, and thermodynamic analysis
The genome-scale metabolic model used for I. orientalis SD108 is iIsor85023, which follows nomenclature of the BiGG database38. To evaluate 3HP production, the model was augmented to separately include seven different 3HP pathways24, as well as the transport of 3HP from the cytosol to the extracellular environment and an exchange flux for extracellular 3HP. The BiGG database38, MetaNetX39, ChEBI40, and PubChem41 were used for information about the metabolites and reactions for pathway intermediates. In general, BiGG nomenclature for metabolite and reaction ids was used when available, with compatible ids created otherwise, as noted in the model files; within BiGG 3-hydroypropionic acid (3HP) has the id 3hpp, which was used in the model and simulations. Marvin was used for characterizing chemical structures including formula and charge information (Marvin 23.17.0, Chemaxon). In agreement with conventions of flux balance modeling, these values for each reaction in each pathway were computed using pH 7.2 for each intracellular species. The added 11 metabolites and 21 reactions are provided in YAML format in Supplementary Data 6, and the accompanying notes field of each reaction includes the associated pathway(s). The reaction set was also updated at the malonic semialdehyde reductase step for YDFG to use NADPH as a cofactor instead of NADH (which was used in previous simulations for S. cerevisiae42).
Flux balance analysis (FBA) was used for model predictions43. For the simulation of a given pathway, only the reactions associated with that pathway were added to the model. The glucose substrate uptake rate was set to 10 mmol g/DW/h glucose during simulations. The maximum oxygen uptake rate was limited to no more than 18.18 mmol g/DW/h during all simulations, as this is the minimal oxygen uptake that does not impact the maximum growth rate for a 10 mmol glucose g/DW/h basis. By convention exchange fluxes assume negative values for uptake, thus these constraints were set as \({v}_{{EX\_glc\_\_D\_e}}=-10\) and \(0\ge {v}_{{EX\_o}2{\_e}}\ge -18.18\). Model simulations of growth phenotype or 3HP formation were obtained using FBA with the objective of maximizing the flux of the biomass reaction (\({v}_{{biomass}}\)) or product exchange flux (\({v}_{{EX\_}3{hpp\_e}}\)) in the context of the indicated pathway. Yields are presented as carbon yields by normalizing to the carbons from glucose. Specifically, for each pathway p yield is computed as
where \({y}_{3{hpp},{glc}\_D}^{c}\) is the number of carbons in 3hpp (i.e., three) divided by those in glucose (i.e., six). Since \({v}_{{EX\_glc}\_{D\_e}}\) is fixed to –10, for maximum yield this expression can be simplified to
FBA calculations were implemented using GAMS 33.2.0 using IBM ILOG CPLEX solver. Adding non-native reactions and data processing were performed using the COBRApy package (version 0.29.0)44 and pandas45 within Jupyter notebooks46.
Thermodynamics analysis of 3HP metabolic pathways was performed by estimating the standard Gibbs energy change \({\triangle }_{r}{{G}^{{\prime} }}^{0}\)) of enzymatic reactions using dGPredictor26 and eQuilibrator27, with SMILES from structures determined using Marvin for compounds not available within KEGG47. The evaluations were performed at pH 7 to compare with previous studies and at pH 6 which corresponds as a lower estimate of the intracellular pH in I. orientalis using the value that S. cereivisiae can have in an external environment of pH 425.
3HP biomanufacturing design and simulation
All 3HP biomanufacturing facilities discussed in this study were designed, simulated, and evaluated using BioSTEAM48 and the thermodynamic package utilized was Thermosteam49. Briefly, influent and effluent streams of each unit are simulated in BioSTEAM and coupled with operating parameters and equipment cost algorithms for unit design and cost calculations. Component equipment for major units (Supplementary Data 7), breakdowns of system costs and environmental impacts (Supplementary Tables 7 and 8, and Supplementary Fig. 22), as well as baseline values and uncertainty distributions of key parameters (Supplementary Data 1) are included in the Supplementary Information. Simulations were run based on experimental fermentation performance at 300 mL and for multiple feedstocks (dextrose, corn, sugarcane, and corn stover). All Python scripts for BioSTEAM and the biomanufacturing facilities (including biomanufacturing facilities setup and system analyses) as well as a system report (including detailed process flowsheet, stream composition and cost tables, unit design specifications, and utilities for the baseline simulation) are available in the online repository (Supplementary Data 8)50. The dextrose-accepting biomanufacturing facilities in this study are comprised of three main (inside battery limits) processes (fermentation, separation, and catalytic upgrading) with outside-battery wastewater treatment and miscellaneous facilities (boiler, turbogenerator, cooling utility regeneration, heat exchanger network, clean-in-place, air distribution, fire water distribution, and process water distribution) with all design assumptions consistent with Bhagwat et al.16 other than for fermentation (based on experimental data obtained in this study). As done in previous studies that involved TEA and LCA to assess the production of platform chemicals16,51,52, the designed biomanufacturing facilities included catalytic upgrading of the platform chemical (3HP) to a commodity chemical (acrylic acid) as no established market yet exists for 3HP. Including catalytic upgrading to acrylic acid in the designed biomanufacturing facilities enabled meaningful comparisons to market price ranges and environmental impacts from conventional production of acrylic acid (detailed under Techno-economic analysis and Life cycle assessment). Further descriptions of design and modeling assumptions for these processes (including catalytic upgrading) and facilities are detailed in Bhagwat et al.16, and all baseline values, references, and uncertainty distributions of key parameters are included in the Supplementary Data 1.
For biomanufacturing facilities in this study designed to accept biomass feedstocks, there were additional processes included inside battery limits: (i) for corn-accepting biomanufacturing facilities, design assumptions for the corn dry grind process were consistent with Cortés-Peña et al.48; (ii) for sugarcane-accepting biomanufacturing facilities in this study, design assumptions for feedstock juicing and clarification were consistent with Bhagwat et al.53; and (iii) for corn stover-accepting biomanufacturing facilities in this study, design assumptions for corn stover pretreatment and saccharification were consistent with Cortés-Peña et al.48. For all biomanufacturing facilities, at each simulation, a heat exchanger network is newly synthesized using the Linnhoff-Hindmarsh pinch-based approach54 to offset heating and cooling utility demands. The production capacity for all biomanufacturing facilities was set to 134,000 metric tons of acrylic acid annually in the baseline case to be consistent with a previous study35. The baseline production capacity was 2% of the global acrylic acid market size and 14% of U.S. capacity for glacial acrylic acid production in 2020, a fraction of the projected growth projected from 2021 to 2025 in the global market for acrylic acid (a growth of approximately 1,230,000 metric ton/y), and comparable to the projected growth in that 5-year period in the U.S. market (approximately 91,000 metric ton/y)35. Further, a triangular distribution with bounds corresponding to ±20% of the baseline value was attributed to the annual production in the uncertainty analysis. Simplified block flow diagrams of the fermentation and separation processes (Supplementary Fig. 14) as well as baseline values and distributions for all parameters included in the uncertainty analysis (Supplementary Data 1) are detailed in the Supplementary Information, and a full process flowsheet is available online50.
Techno-economic analysis (TEA)
We performed TEA for the designed and simulated biomanufacturing facilities using BioSTEAM’s discounted cash flow rate of return analysis to calculate the minimum product selling price (MPSP, $/kg) of acrylic acid to achieve a net present value of zero with a 10% annual internal rate of return (a uniform distribution of 8 − 12% was assumed; a full list of baseline parameter values, uncertainty distributions, and literature references for the same are provided in Supplementary Data 1. All costs and prices shown are presented in 2019 U.S. dollars. To benchmark the MPSP of the produced acrylic acid, we assumed a market price range of $1.40 − 1.65/kg from reported 2015 − 2019 market prices35 to be consistent with Bhagwat et al.16. All biomanufacturing facilities were modeled as nth plant designs (i.e., it is assumed a successful industry has been established with mature technologies). Key construction (e.g., warehouse, site development), fixed operating (e.g., labor burden, property insurance), and financial (e.g., depreciation, taxes) parameters, as well as all parameters used in the cash flow analysis, followed assumptions made by Bhagwat et al.16. Tabulated breakdowns of the estimated revenue and capital and operating expenditures, as well as additional details on the design, utility requirements, purchase costs, and installed equipment costs can be found online50.
Life cycle assessment (LCA)
We performed LCA in Python using the simulated inventories for streams (input chemicals and output emissions) and utilities from BioSTEAM. The LCA scope included the operational phase of the biomanufacturing, including cradle-to-grave impacts for all raw materials, ancillary processes, and unit processes. The functional unit was set to 1 kg of produced acrylic acid to be consistent with the TEA. The sale of co-produced electricity was assumed to displace the impacts of marginal electricity production. Impacts resulting from infrastructure construction were excluded to be consistent with the U.S. renewable fuel standard (RFS)55. Final characterization and discussion of environmental impacts focused on carbon intensity (CI; quantified as 100-year global warming potential, GWP100) and fossil energy consumption (FEC). The impact assessment methodology used for CI was Intergovernmental Panel on Climate Change (IPCC) 201356. Life cycle inventory data were collected from ecoinvent 3.8 and some unit impacts were gathered from GREET 202036,37, and their sources were noted in the script50. The baseline impacts of feedstock farming (excluding credit for fixed carbon), harvest and collection, transportation, storage, and handling were considered. The amount of carbon fixed in the feedstock is equal to the sum of carbon in the product (acrylic acid) and in the biogenic portion of direct waste emissions from the biomanufacturing facility. To focus on the biomanufacturing facility, we assumed the end-of-life impacts associated with the product (acrylic acid) as well as non-gaseous wastes (e.g., unconsumed and non-combusted sugars and insoluble lignin in the brine, accounting for <0.0002% of the CI) would be exclusively from the oxidation of all carbon into CO2.
Uncertainty and sensitivity analyses
Uncertainty analyses were conducted using Monte Carlo simulation with Latin Hypercube Sampling (6,000 simulations) for 33 uncertain parameters each in the dextrose, corn, sugarcane, and corn stover scenarios. We assigned uncertainty distributions for parameters to be consistent with Bhagwat et al.16, except for the parameters describing the feedstock acquisition, juicing, and clarification processes (consistent with Bhagwat et al.53) and the fermentation process (based on experimental data obtained in this study). The full list of the evaluated parameters, their uncertainty distributions, and literature references are provided in Supplementary Data 1. The sensitivity of MPSP, CI, and FEC to all uncertain inputs was determined via Spearman’s rank order correlation coefficients (Spearman’s ρ) using Monte Carlo simulation results, and only parameters with p < 0.05 were discussed to ensure robustness (Supplementary Fig. 15). Files with comprehensive results of all analyses are available online50.
Statistics & reproducibility
No statistical method was used to predetermine sample size. Most experiments were performed with at least three independent biological replicates, and results are presented as mean ± S.D. The 300 mL scale fed-batch fermentation was conducted with two biological replicates, whereas the 3 L scale fed-batch fermentation and qPCR analysis were performed with one biological replicate. No data were excluded from the analyses. The experiments were not randomized, and the investigators were not blinded to allocation during experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data supporting the findings of this work are available within the paper and its Supplementary Information files or uploaded through public repositories. If specific data is believed to be missing, that data is available from the corresponding authors upon request. Plasmids used in this study will be available upon request. The corresponding author will respond to the request within two weeks. Source data are provided with this paper.
Code availability
The code analyzing 3HP pathways in the genome-scale metabolic model is available on https://github.com/maranasgroup/I_orientalis_organic_acids, with details of the added metabolites and reactions also included in Supplementary Information. All results for techno-economic analysis and life cycle assessment (including plots and raw data) and the software scripts used to generate the same are available at BioSTEAMDevelopmentGroup: 3-Hydroxypropionic acid biorefineries, 2024 (https://github.com/BioSTEAMDevelopmentGroup/Bioindustrial-Park/tree/master/biorefineries/HP), with details on component equipment for major units, breakdowns of system costs and environmental impacts, as well as baseline values and uncertainty distributions of key parameters included in the Supplementary Information.
References
Nielsen, J. & Keasling, J. D. Engineering cellular metabolism. Cell 164, 1185–1197 (2016).
Lee, S. Y. et al. A comprehensive metabolic map for production of bio-based chemicals. Nat. Catal. 2, 18–33 (2019).
Volk, M. J. et al. Metabolic engineering: methodologies and applications. Chem. Rev. 123, 5521–5570 (2023).
Werpy, T. & Petersen, G. Top value added chemicals from biomass: volume I -- results of screening for potential candidates from sugars and synthesis gas. https://www.osti.gov/biblio/15008859, https://doi.org/10.2172/15008859 (2004).
Bozell & Petersen, J. J. G. R. Technology development for the production of biobased products from biorefinery carbohydrates—the US Department of Energy’s “Top 10” revisited. Green Chem. 12, 539–554 (2010).
Vidra, A. & Németh, Á Bio-based 3-hydroxypropionic acid: a review. Period. Polytech. Chem. Eng. 62, 156–166 (2018).
Kumar, V., Ashok, S. & Park, S. Recent advances in biological production of 3-hydroxypropionic acid. Biotechnol. Adv. 31, 945–961 (2013).
Kildegaard, K. R. et al. Engineering and systems-level analysis of Saccharomyces cerevisiae for production of 3-hydroxypropionic acid via malonyl-CoA reductase-dependent pathway. Microb. Cell Factories 15, 53 (2016).
Moussa, M. et al. Reactive extraction of 3-hydroxypropionic acid from model aqueous solutions and real bioconversion media. Comparison with its isomer 2-hydroxypropionic (lactic) acid: Reactive extraction of 3-HP acid from aqueous solutions and bioconversion broths. J. Chem. Technol. Biotechnol. 91, 2276–2285 (2016).
Rathnasingh, C., Raj, S. M., Jo, J.-E. & Park, S. Development and evaluation of efficient recombinant Escherichia coli strains for the production of 3-hydroxypropionic acid from glycerol. Biotechnol. Bioeng. 104, 729–739 (2009).
IndustrialSectorPMR, A. Acrylic Acid: Growth of Coatings Industry – Key Influencer 2029. Site Title https://marketreserchfmi.wordpress.com/2021/10/01/acrylic-acid-growth-of-coatings-industry-key-influencer-2029/ (2021).
Dishisha, T., Pyo, S.-H. & Hatti-Kaul, R. Bio-based 3-hydroxypropionic- and acrylic acid production from biodiesel glycerol via integrated microbial and chemical catalysis. Microb. Cell Factories 14, 200 (2015).
Abraham, T. W. et al. Recovery of 3-hydroxypropionic acid. Patent No. 11,236,036 (U.S. Patent and Trademark Office, Washington, DC, 2022).
Wang, X., Cui, Z., Sun, X., Wang, Z. & Chen, T. Production of 3-hydroxypropionic acid from renewable substrates by metabolically engineered microorganisms: a review. Molecules 28, 1888 (2023).
Batista, R. S. et al. Glycerol as substrate and NADP+-dependent glyceraldehyde-3-phosphate dehydrogenase enable higher production of 3-hydroxypropionic acid through the β-alanine pathway in E. coli. Bioresour. Technol. 393, 130142 (2024).
Bhagwat, S. S. et al. Sustainable production of acrylic acid via 3-hydroxypropionic acid from lignocellulosic biomass. ACS Sustain. Chem. Eng. 9, 16659–16669 (2021).
Kang, F. et al. Expanding the genetic toolkit of Yarrowia lipolytica: dynamic promoter engineering enables high-titer biosynthesis of 3-hydroxypropionic acid. Bioresour. Technol. 432, 132656 (2025).
Qin, N. et al. Increased CO2 fixation enables high carbon-yield production of 3-hydroxypropionic acid in yeast. Nat. Commun. 15, 1591 (2024).
Xiao, H., Shao, Z., Jiang, Y., Dole, S. & Zhao, H. Exploiting Issatchenkia orientalis SD108 for succinic acid production. Microb. Cell Factories 13, 121 (2014).
Cao, M. et al. A genetic toolbox for metabolic engineering of Issatchenkia orientalis. Metab. Eng. 59, 87–97 (2020).
Tran, V., Fatma, Z., Song, X. & Zhao, H. Development of a CRISPR/Cas9-based tool for gene deletion in Issatchenkia orientalis. mSphere 4, 10–1128 (2019).
Fatma, Z., Tan, S.-I., Boob, A. G. & Zhao, H. A landing pad system for multicopy gene integration in Issatchenkia orientalis. Metab. Eng. 78, 200–208 (2023).
Suthers, P. F. et al. Genome-scale metabolic reconstruction of the non-model yeast Issatchenkia orientalis SD108 and its application to organic acids production. Metab. Eng. Commun. 11, e00148 (2020).
Jiang, X., Meng, X. & Xian, M. Biosynthetic pathways for 3-hydroxypropionic acid production. Appl. Microbiol. Biotechnol. 82, 995–1003 (2009).
Valli, M. et al. Intracellular pH distribution in Saccharomyces cerevisiae cell populations, analyzed by flow cytometry. Appl. Environ. Microbiol. 71, 1515–1521 (2005).
Wang, L., Upadhyay, V. & Maranas, C. D. dGPredictor: automated fragmentation method for metabolic reaction free energy prediction and de novo pathway design. PLOS Comput. Biol. 17, e1009448 (2021).
Beber, M. E. et al. eQuilibrator 3.0: a database solution for thermodynamic constant estimation. Nucleic Acids Res. 50, D603–D609 (2022).
Zallot, R., Oberg, N. & Gerlt, J. A. The EFI web resource for genomic enzymology tools: leveraging protein, genome, and metagenome databases to discover novel enzymes and metabolic pathways. Biochemistry 58, 4169–4182 (2019).
Borodina, I. et al. Establishing a synthetic pathway for high-level production of 3-hydroxypropionic acid in Saccharomyces cerevisiae via β-alanine. Metab. Eng. 27, 57–64 (2015).
Tran, V. G. et al. An end-to-end pipeline for succinic acid production at an industrially relevant scale using Issatchenkia orientalis. Nat. Commun. 14, 6152 (2023).
Tran, V. G. et al. Decompartmentalization of the yeast mitochondrial metabolism to improve chemical production in Issatchenkia orientalis. Nat. Commun. 16, 7110 (2025).
Lee, Y. G. et al. Self-buffering system for cost-effective production of lactic acid from glucose and xylose using acid-tolerant Issatchenkia orientalis. Bioresour. Technol. 399, 130641 (2024).
Tan, S. I., Liu, Z., Tran, V. G., Martin, T. A. & Zhao, H. Issatchenkia orientalis as a platform organism for cost-effective production of organic acids. Metab. Eng. 89, 12–21 (2025).
Matsakas, L., Hrůzová, K., Rova, U. & Christakopoulos, P. Biological production of 3-hydroxypropionic acid: an update on the current status. Fermentation 4, 13 (2018).
IHS Markit. Chemical Economics Handbook - Acrylic Acid and Esters (IHS Markit, London, England, 2020).
Wang, M. et al. Greenhouse gases, regulated emissions, and energy use in technologies model. Computer Software. (U.S. DOE, Office of Energy Efficiency and Renewable Energy, 2020).
Wernet, G. et al. The ecoinvent database version 3 (part I): overview and methodology. Int. J.Life Cycle Assess. 21, 1218–1230 (2016).
Norsigian, C. J. et al. BiGG Models 2020: multi-strain genome-scale models and expansion across the phylogenetic tree. Nucleic Acids Res. 48, D402–D406 (2020).
Moretti, S., Tran, V. D. T., Mehl, F., Ibberson, M. & Pagni, M. MetaNetX/MNXref: unified namespace for metabolites and biochemical reactions in the context of metabolic models. Nucleic Acids Res. 49, D570–D574 (2021).
Hastings, J. et al. ChEBI in 2016: improved services and an expanding collection of metabolites. Nucleic Acids Res. 44, D1214–D1219 (2016).
Kim, S. et al. PubChem 2023 update. Nucleic Acids Res. 51, D1373–D1380 (2023).
Ji, R. Y. et al. Metabolic engineering of yeast for the production of 3-hydroxypropionic acid. Front. Microbiol. 9, 2185 (2018).
Orth, J. D., Thiele, I. & Palsson, B. Ø What is flux balance analysis? Nat. Biotechnol. 28, 245–248 (2010).
Ebrahim, A., Lerman, J. A., Palsson, B. O. & Hyduke, D. R. COBRApy: COnstraints-based reconstruction and analysis for python. BMC Syst. Biol. 7, 74 (2013).
McKinney, W. Data structures for statistical computing in Python. scipy 445, 51–56 (2010).
Granger, B. E. & Pérez, F. Jupyter: thinking and storytelling with code and data. Comput. Sci. Eng. 23, 7–14 (2021).
Kanehisa, M., Furumichi, M., Sato, Y., Ishiguro-Watanabe, M. & Tanabe, M. KEGG: integrating viruses and cellular organisms. Nucleic Acids Res. 49, D545–D551 (2021).
Cortes-Peña, Y., Kumar, D., Singh, V. & Guest, J. S. BioSTEAM: a fast and flexible platform for the design, simulation, and techno-economic analysis of biorefineries under uncertainty. ACS Sustain. Chem. Eng. 8, 3302–3310 (2020).
Cortés-Peña, Y. Thermosteam: BioSTEAM’s premier thermodynamic engine. J. Open Source Softw. 5, 2814 (2020).
BioSTEAM Development Group. 3-Hydroxypropionic Acid Biorefineries. https://github.com/BioSTEAMDevelopmentGroup/Bioindustrial-Park/tree/master/biorefineries/HP (2024).
Lee, J. W. et al. Rewiring yeast metabolism for producing 2,3-butanediol and two downstream applications: techno-economic analysis and life cycle assessment of methyl ethyl ketone (MEK) and agricultural biostimulant production. Chem. Eng. J. 451, 138886 (2023).
Kim, M. S. et al. Sustainable potassium sorbate production from triacetic acid lactone in food-grade solvents. Green Chem. 27, 6087–6104 (2025).
Bhagwat, S. S. et al. Sustainable triacetic acid lactone production from sugarcane and sweet sorghum by fermentation and fractional crystallization. ChemRxiv (2024).
Seider, W. D. et al. Product and Process Design Principles: Synthesis, Analysis and Evaluation (John Wiley & Sons, 2016).
U.S. EPA. Lifecycle Analysis of Greenhouse Gas Emissions under the Renewable Fuel Standard. https://www.epa.gov/renewable-fuel-standard-program/lifecycle-analysis-greenhouse-gas-emissions-under-renewable-fuel (2025).
WG, I. The physical science basis. Contrib. Work. Group Fifth Assess. Rep. Intergov. Panel Clim. Change 1535, 2013 (2013).
Acknowledgements
This work was funded by the DOE Center for Advanced Bioenergy and Bioproducts Innovation (U.S. Department of Energy, Office of Science, Biological and Environmental Research Program under Award Number DE-SC0018420 to H.Z.). Any opinions, findings, and conclusions or recommendations expressed in this publication are those of the author(s) and do not necessarily reflect the views of the U.S. Department of Energy. We thank Dr. Somesh Mishra and Dr. Vijay Singh for providing corn steep liquor. The online tool BioRender (biorender.com) was used to create Figs. 2, 3 and 4 and Supplementary Figs. 1, 3, 4, 5 and 12. Computations for this research involving the genome scale model were performed on the Pennsylvania State University’s Roar Collab supercomputer. The authors of this work recognize the Penn State Institute for Computational and Data Sciences (RRID:SCR_025154) for providing access to computational research infrastructure within the Roar Core Facility (RRID: SCR_026424).
Author information
Authors and Affiliations
Contributions
S.I.T. and H.Z. conceived and designed the study. S.I.T. constructed and characterized all I. orientalis strains as well as performed all the gene candidate screening test. Z.F. performed the SSN analysis and selected the gene candidates. P.F.S. performed genome-scale modeling and thermodynamic analysis. T.A.M. and V.G.T. performed the 300 mL DASbox fed-batch fermentation. T.A.M. performed 3 L scale-up. S.S.B. performed biomanufacturing facility design, modeling, techno-economic analysis, and life cycle assessment. W. T. assisted with strain construction and characterization. S.I.T., T.A.M., P.F.S., S.S.B., C.D.M., J.S.G., and H.Z. wrote the manuscript with input from all other authors.
Corresponding authors
Ethics declarations
Competing interests
Huimin Zhao, Teresa Martin and Shih-I Tan have submitted a patent application to the US patent and trademark office pertaining to the aspect of the production of 3HP using Issatchenkia orientalis in this work (application number 63/766,638). The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Sebastian Riedel who co-reviewed with Lara Santolin, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tan, SI., Bhagwat, S.S., Martin, T.A. et al. High yield production of 3-hydroxypropionic acid using Issatchenkia orientalis. Nat Commun 17, 899 (2026). https://doi.org/10.1038/s41467-025-67621-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67621-8