Abstract
DNA base editors are transformative genome editing tools that enable nucleotide conversions without inducing double-stranded DNA breaks, making them promising for correcting genetic mutations. Current base editors, however, are limited by fixed editing windows and constrained location of deaminases. To address these constraints, we develop a modular system termed Plug-in Base Editor (Plug-in BE), which dynamically programs deaminase positioning via integrating various epitopes and antibody-fused deaminases. This system expands the editing capabilities of base editors by optimizing deaminase’s spatial interaction with DNA, leading to improvements in efficiencies, window restrictions, and safety profiles. We validate Plug-in BE’s versatility and high fidelity in cancer gene therapy and zebrafish embryo editing, demonstrating its potential as a powerful and adaptable tool for basic research and therapeutic applications. This innovation can generate a series of base editors without extensive protein evolution, positioning Plug-in BE as a significant advancement in the field of genome editing.
Similar content being viewed by others
Introduction
DNA base editors enable the conversion of target nucleotides without introducing double-stranded DNA breaks1, making them highly promising tools for correcting human inherited diseases. Adenine base editors (ABEs) mediate A-to-G conversion2, while cytosine base editors (CBEs) enable C-to-T conversion1. Additionally, glycosylase base editors (CGBEs) implement C-to-G conversion3,4. Through deaminase evolution and the incorporation of certain glycosylases, DNA base editors can achieve other mutation types, such as ACBE converting A-to-C5. Further, replacing SpCas9 with other Cas proteins has led to base editors matching specific PAMs6,7. However, current base editors exhibit a specific editing pattern at a fixed window, limiting their effectiveness in genome editing8,9.
During the optimization of base editors, positioning the deaminase at the N- or C-terminus of nCas9(D10A) has been shown to influence editing efficiency, editing window, byproducts, and Indels1. Further, in inlaid and circularly permuted base editors, these characteristics can be tuned by embedding deaminases into nCas9(D10A) or permuting its domains10,11,12. However, such recombination alters protein conformation and activity, thereby restricting the number of viable configurations and resulting in a relatively fixed editing pattern and editing window.
In this study, we present a modular system designed to dynamically program the spatial locations of deaminase within DNA base editors. By embedding epitopes at various positions within nCas9(D10A) and fusing deaminase with antibody, we can manipulate the deaminase’s spatial interaction with DNA, thereby altering its editing outcome, including editing efficiency, editing window, byproducts, Indels, and sequence motif preference. These modular components are combined in a plug-in manner, with CRISPR/Cas as a plug-in platform, epitopes as slots, and deaminases as plug-ins to generate a wide array of new base editors with improvements in efficiency, fidelity, and window restriction, which we term Plug-in Base Editor (Plug-in BE). The efficacy of Plug-in BEs has been further validated in both cancer gene therapy and zebrafish embryo editing, highlighting its potential as a powerful and safe tool for basic research and therapeutic applications.
Results
Generation of SunTag plug-in BE via programming deaminase’s spatial location
Current DNA base editors generally restrict the location of deaminases to the N/C-terminal of nCas9(D10A) (Fig. 1A, left). To overcome this limitation, we embedded an epitope at internal positions within nCas9(D10A), thereby enabling the recruitment of antibody-fused deaminases to specific spatial locations (Fig. 1A, right). Here, we developed 62 D10A-GCN4 variants by embedding GCN4 (a 19 amino acids epitope from SunTag system13) primarily within loops between α-helices and β-sheets, thereby minimizing disruption to the function of nCas9(D10A) (Fig. 1B). Based on SpCas9 structure (PDB, 5f9r), the chosen embedded positions are spatially close to the target DNA. Further, structural predictions using AlphaFold314 confirmed that the GCN4 embedment did not significantly alter the conformation of nCas9(D10A) (Fig. 1C), suggesting that D10A-GCN4 variants may retain their functionality.
A Schematic of DNA base editors. Base editors generally contain N/C-terminal deaminases (left). Plug-in BE contains a spatially programmable deaminase (right). B 62 embedding sites within nCas9(D10A) generate 62 D10A-GCN4 variants. Arrow-marked numbers denote GCN4 embedded behind a specific amino acid. 1048–1063 represents substituting amino acids 1048–1063 with GCN4. C Conformation comparison between SpCas9 (PDB, 5f9r) and four D10A-GCN4 variants predicted by AlphaFold3. GCN4 is highlighted in red. D10A-532GCN4 represents embedding GCN4 behind the 532nd amino acid of nCas9(D10A). Nomenclature rules for other variants are similar. D Schematics for BE3, BE4, SunTag Plug-in BE3 (D10A-GCN4 and AntiGCN4-BE3), and SunTag Plug-in BE4 (D10A-GCN4 and AntiGCN4-BE4). AntiGCN4 with heavy and light chain variable regions is depicted as 2 segments. E Editing product analysis of BE3, BE4, and three D10A-GCN4 at four gene loci by amplicon sequencing (n = 2 biological replicates). 231GCN4-BE3 represents combination of D10A-231GCN4 and AntiGCN4-BE3. Nomenclature rules for other groups are similar. Legend only C-to-T indicates reads with only the C-to-T conversion and no other mutations. F Editing efficiency analysis of SunTag Plug-in BE3 at VEGFA site 2 by Sanger sequencing (n = 2 biological replicates). Numbers on Y-axis of heat map represent D10A-GCN4. The editing efficiency for 62 variants is displayed in Supplementary Fig. 1A. G Editing windows illustrating the impact of GCN4 insertion positions on editing pattern. Window graph is obtained by merging the editing results at HEK site 3, ZAP70, and FANCF site 1. Numbers with different color represent D10A-GCN4 variants. Certain cytosines at X-axis coordinates are specifically labeled with motif. Editing window of BE3 serves as control group (n = 2 biological replicates). Data are mean ± SD (Blue group, n = 10 variants; Green group, n = 15 variants; Purple group, n = 8 variants; Yellow group, n = 12 variants; Gray group, n = 17 variants). H Editing product analysis of BE3 and high-efficiency variants at four loci by amplicon sequencing (n = 2 biological replicates). Source data are provided as a Source Data file.
To establish a mobile deaminase module, we fused AntiGCN4 (a 272 amino acids antibody from SunTag system13) with APOBEC1 and UGI(s)1,15, resulting in constructs named AntiGCN4-BE3 with UGI and AntiGCN4-BE4 with 2xUGI (Fig. 1D). These systems were collectively termed SunTag Plug-in BE3 and SunTag Plug-in BE4. Initially, we selected D10A-231GCN4, D10A-1055GCN4, and D10A-1246GCN4 for editing four gene loci in HEK293T cells. Overall, SunTag Plug-in BE3 improved editing efficiency by 20% to 40% over BE3, BE4, and SunTag Plug-in BE4, making it a promising candidate for further investigation (Fig. 1E)
Subsequently, we systematically assessed the editing efficiencies of all 62 D10A-GCN4 variants within SunTag Plug-in BE3 framework. Notably, 29 variants demonstrated higher efficiency than BE3 (Fig. 1F and Supplementary Fig. 1A). For instance, at the VEGFA site 2, D10A-170/231/1059/1068GCN4 achieved around 90% C-to-T efficiency, outperforming BE3’s 79%. Similar improvements were observed at the FANCF site 1, HEK site 3, and ZAP70.
Interestingly, the GCN4 embedding position within nCas9(D10A) obviously affected the editing window. For example, the editing windows of D10A-170/308/1033GCN4 expanded to the right at the VEGFA site 2 (Fig. 1F). Based on these observations, we classified 62 variants into five groups with distinct editing patterns, according to the editing results at the HEK site 3, ZAP70, and FANCF site 1 (Fig. 1G). The blue group with 10 variants exhibited the highest efficiency and specific rejection to the motifs CC4A, AC9G, GC13T, and GC14A, while the green group with 15 variants rejected CC8T. Remarkably, the variants reduced byproducts (Fig. 1H). For instance, D10A-308GCN4 reduced byproducts from 25% with BE3 to 4% at the FANCF site 1, while D10A-1246GCN4 reduced from 37% to 2% at the VEGFA site 2. These results demonstrate that the strategic spatial programming of deaminase in the modularized Plug-in BE not only enhances editing efficiency but, more importantly, improves editing fidelity.
SunTag Plug-in BE with ABE8eWQ, miniCGBE1, and Target AID
Inspired by the success of SunTag Plug-in BE3, we further incorporated ABE8eWQ16 to develop SunTag Plug-in ABE8eWQ (Fig. 2A). D10A-1/532/945/1068/1072GCN4 exhibited comparable editing efficiencies to ABE8eWQ (Supplementary Fig. 2 A). To be noteworthy, the editing window analysis revealed that SunTag Plug-in ABE8eWQ’s editing activity was largely restricted to A6 and A7 (Fig. 2B). Consequently, the contracted window improved single-base editing precision (Fig. 2C). D10A-532GCN4 specifically enhanced the only A8-to-G from 1% with ABE8eWQ to 42% at E21, while D10A-1055GCN4 increased the only A7-to-G from 1% with ABE8eWQ to 90% at OCT4. More importantly, the specific variants further reduced unwanted Indels products (Supplementary Fig. 2C). For example, D10A-532GCN4 reduced Indels from 1.4% of ABE8eWQ to 0.44% at OCT4. Together, these results demonstrate that embedding GCN4 within nCas9(D10A) can refine the activity of ABE8eWQ by narrowing the editing window, enhancing single-base editing precision, and reducing byproducts.
A Schematic of ABE8eWQ and SunTag Plug-in ABE8eWQ (D10A-GCN4 and AntiGCN4-ABE8eWQ). B Editing window analysis of SunTag Plug-in ABE8eWQ. Window graph is obtained by merging the edit results at HEK site 2, E21, ZAP70, and OCT4. Numbers with different color represent D10A-GCN4 variants. Editing window of ABE8eWQ serves as control group (n = 2 biological replicates). Data are mean ± SD (Blue group, n = 5 variants; Green group, n = 14 variants; Purple group, n = 10 variants; Yellow group, n = 23 variants; Gray group, n = 10 variants). C Assessment of single-base editing precision for selected variants (n = 2 biological replicates). The legend only A5-to-G indicates that only the fifth A on 20nt spacer is edited to G, and the spacer does not contain any other mutation. Other legend naming rules are analogies. D Schematic of miniCGBE1 and SunTag Plug-in miniCGBE1 (D10A-GCN4 and AntiGCN4-miniCGBE1). E Editing window analysis of SunTag Plug-in miniCGBE1. The Window graph was obtained by merging the editing results of FANCF site 4, EMX1, PPP1R12C, and HEK site 3. Certain cytosines at X-axis coordinates are specifically labeled with motif. Numbers with different color represent D10A-GCN4 variants. Editing window of miniCGBE1 serves as control group (n = 2 biological replicates). Data are mean ± SD (Blue group, n = 12 variants; Green group, n = 13 variants; Purple group, n = 3 variants; Yellow group, n = 25 variants; Gray group, n = 9 variants). F, G Editing product analysis of high-efficiency D10A-GCN4 variants of SunTag Plug-in miniCGBE1 at FANCF site 4 and PPP1R12C, and its editing precision analysis (n = 2 biological replicates). Source data are provided as a Source Data file.
To evaluate whether Plug-in BE could achieve C-to-G conversion, we developed SunTag Plug-in miniCGBE13 and assessed its performance in HeLa cells (Fig. 2D and Supplementary Fig. 3A, C). At FANCF site 4, D10A-1246/1248/1252/1260GCN4 achieved the highest editing efficiency, ranging from 44% to 53%, which was comparable to miniCGBE1’s 46% (Supplementary Fig. 3A). Further analysis revealed that the editing window of SunTag Plug-in miniCGBE1 contracted to C6 (Fig. 2E). In addition, the variants showed reduction in byproducts (Fig. 2F, G). For instance, at PPP1R12C site, whereas miniCGBE1 reached 23% editing efficiency but generated 40% byproducts, D10A-231/1068/1072GCN4 achieved 30% editing efficiency with only 10% byproducts. In summary, SunTag Plug-in miniCGBE1 reduces byproducts, providing a narrow editing window that enhances the editing precision and practical application of CGBEs.
We further integrated Target AID to develop SunTag Plug-in Target AID17. Interestingly, SunTag Plug-in Target AID exhibited a series of wider editing windows and complex editing patterns (Supplementary Fig. 4 and 5).
Taken together, SunTag Plug-in BE is compatible with different deaminases and embedding positions, enabling a wide range of conversions (C-to-T, A-to-G, and C-to-G), while also reprogramming the editing patterns to better achieve diverse editing requirements.
Enhancement of Plug-in BE through integration of MoonTag system
MoonTag system contains a smaller epitope, gp41 (15 amino acids), compared with GCN4 (19 amino acids), and a smaller antibody, Antigp41 (123 amino acids), compared with AntiGCN4 (272 amino acids)18. Given the compact nature of MoonTag system, we hypothesized that it might improve editing performance with Plug-in BE, and therefore investigated the editing outcomes of MoonTag Plug-in BEs (Fig. 3A, E, H).
A Schematic of MoonTag Plug-in BE3. B Editing efficiency of MoonTag Plug-in BE3 at VEGFA site 2 (n = 2 biological replicates). Numbers on Y-axis represent D10A-gp41. C Editing window of MoonTag Plug-in BE3. Window graph merges from editing results at HEK site 3, ZAP70, and FANCF site 1. Certain cytosines at X-axis are labeled with motif. Numbers with color represent D10A-gp41. Editing window of BE3 is control group (n = 2 biological replicates). Data are mean ± SD (Blue group, n = 14 variants; Green group, n = 9 variants; Purple group, n = 13 variants; Yellow group, n = 13 variants; Gray group, n = 13 variants). D Editing product of MoonTag Plug-in BE3 at four gene loci (n = 2 biological replicates). E Schematic of MoonTag Plug-in ABE8eWQ. F Editing window of MoonTag Plug-in ABE8eWQ. Window graph merges from editing results at four gene loci. Numbers with color represent D10A-gp41. Editing window of ABE8eWQ is control group (n = 2 biological replicates). Data are mean ± SD (Blue group, n = 9 variants; Green group, n = 12 variants; Purple group, n = 3 variants; Yellow group, n = 15 variants; Gray group, n = 23 variants). G Single-base editing for MoonTag Plug-in ABE8eWQ at four gene loci (n = 2 biological replicates). H Schematic of MoonTag Plug-in miniCGBE1. I Editing window of MoonTag Plug-in miniCGBE1. Window graph merges from editing results at FANCF site 4, EMX1, PPP1R12C, and HEK site 3. The 7th cytosine at X-axis is labeled with motif. Numbers with color represent D10A-gp41. Editing window of miniCGBE1 is control group (n = 2 biological replicates). Data are mean ± SD (Blue group, n = 11 variants; Green group, n = 6 variants; Purple group, n = 6 variants; Yellow group, n = 27 variants; Gray group, n = 12 variants). J, K Editing product of MoonTag Plug-in miniCGBE1 at FANCF site 4 and PPP1R12C (n = 2 biological replicates). Source data are provided as a Source Data file.
In MoonTag Plug-in BE3 system, D10A-gp41 variants showed further improvement of editing efficiencies than the D10A-GCN4 counterparts at identical embedding positions (Supplementary Fig. 6A). For instance, at FANCF site 1, D10A-719/768/945/1050/1068/1248gp41 achieved over 70% editing efficiency, while D10A-GCN4 resulted in efficiencies ranging from 50% to 60%. Furthermore, D10A-gp41 variants altered the sequence motif preference and editing window width (Fig. 3B, C). Similar to D10A-GCN4, D10A-gp41 reduced byproducts (Fig. 3D). To evaluate the DNA off-target effects of MoonTag Plug-in BE3, we performed amplicon sequencing at off-target sites of HEK site 3 and VEGFA site 2. Based on the on/off-target ratio, D10A-gp41 variants showed comparable editing specificity to BE3 (Supplementary Fig. 8E–H and Supplementary Note 2).
In MoonTag Plug-in ABE8eWQ system, 24 D10A-gp41 variants exhibited editing efficiency higher than SunTag Plug-in ABE system (Fig. 3F and Supplementary Fig. 2B). Unlike the contracted editing window of SunTag Plug-in ABE8eWQ, MoonTag Plug-in ABE8eWQ exhibited diverse window modes (Fig. 3F). Among the variants, 3 variants (purple group) expanded the editing window to A3 and A8. In addition, 12 variants (green group) had the narrowest editing window with the highest efficiency at A7, while another 9 variants (blue group) contracted their window to A5, A6, and A7. Due to the shifting editing window, MoonTag Plug-in ABE8eWQ enhanced the single-base editing precision (Fig. 3G). Moreover, D10A-gp41 variants further reduced the Indels of ABE8eWQ (Supplementary Fig. 2D). To evaluate off-target effects of MoonTag Plug-in ABE, we conducted amplicon sequencing at off-target sites of OCT4 and HEK site 2, and found that D10A-gp41 variants maintained comparable DNA-level editing specificity to ABE8eWQ (Supplementary Fig. 8A, B). Furthermore, transcriptome analysis confirmed that D10A-gp41 variants had no increase in RNA-level off-target effects compared to ABE8eWQ (Supplementary Fig. 8C, D and Supplementary Note 2).
Next, we expanded MoonTag Plug-in BE to two additional deaminases. MoonTag Plug-in miniCGBE1 showed comparable performance to SunTag Plug-in miniCGBE1 in editing efficiency, editing window, and product purity (Fig. 3H–K and Supplementary Fig. 6E). Surprisingly, MoonTag Plug-in Target AID outperformed SunTag Plug-in Target AID in editing efficiency (Supplementary Figs. 4, 5, and 6C).
Cas variants with expanded targeting capacity (e.g., the nearly PAM-less SpRY) can overcome PAM restrictions, thereby broadening the range of base editing. However, even with sgRNA positional adjustments (1–5 bp rightward shifts), SpRY-BE3 failed to efficiently achieve C9-to-T conversion with only 1.4%–16.9% efficiency at HEK site 3 (Supplementary Fig. 7A, B). In contrast, MoonTag Plug-in BE3 (D10A-1246gp41) achieved a higher C9-to-T efficiency of 38.9% without relying on sgRNA redesign (Supplementary Fig. 7B). This finding underscores that expanded PAM compatibility alone does not guarantee precise and efficient base editing, whereas the Plug-in BE system’s programmable deaminase positioning enables targeted edits in regions inaccessible to conventional Cas-fused base editors.
In summary, the compact nature of MoonTag enhances editing efficiencies and improves editing window control in Plug-in BE system, without compromising editing specificity. These findings suggest that MoonTag Plug-in BEs offer a promising strategy for achieving more precise and efficient genome editing.
Expansibility of Plug-in BE with ALFA system for modulating editing patterns
To further validate the expansibility of Plug-in BE as a general strategy for manipulating editing patterns, we developed ALFA Plug-in BE by integrating the ALFA system, which contains a 13 amino acids epitope ALAF and a 132 amino acids antibody Anti-ALFA19,20. Firstly, we constructed ALFA Plug-in CBE and tested D10A-231ALFA and D10A-1246ALFA (Fig. 4A). Notably, the editing windows varied among GCN4, gp41, and ALFA, even when the same embedding positions were used (Fig. 4B). At C15, D10A-231ALFA possessed 36% editing efficiency compared to D10A-231GCN4/gp41’s 5% to 10%. At C3 and C4, D10A-1246ALFA increased editing efficiency by 20% compared to D10A-1246GCN4/gp41. Additionally, D10A-231ALFA and D10A-1246ALFA also reduced byproducts compared to BE3 (Fig. 4C).
A Schematic of ALFA Plug-in BE3. B Comparison of Plug-in BE3 editing windows for D10A-ALFA, D10A-GCN4, and D10A-gp41 under the 231st and 1246th embedding positions (n = 2 biological replicates). Window graph is obtained by combining the editing results at FANCF site 1, HEK site 3, and ZAP70. 231ALFA represents D10A-231ALFA and AntiALFA-BE3. Other combinations are named in analogies. C Editing product analysis of Plug-in BE3 for D10A-ALFA, D10A-GCN4, and D10A-gp41 (n = 2 biological replicates). D Schematic of ALFA Plug-in ABE8eWQ. E Comparison of editing windows among ALFA, GCN4, and gp41 in Plug-in ABE8eWQ under the 945th, 1246th, and 1252nd embedding positions (n = 2 biological replicates). Window graph is obtained by combining the edit results at HEK site 2, E21, ZAP70, and OCT4. F, G Product analysis and precision analysis of Plug-in ABE8eWQ for D10A-ALFA, D10A-GCN4, and D10A-gp41 (n = 2 biological replicates). Source data are provided as a Source Data file.
Next, we evaluated ALFA Plug-in ABE system at embedding positions 945, 1246, and 1252 (Fig. 4D). As with the previous Plug-in BEs, ALFA’s embedment altered the editing window of ABE8eWQ (Fig. 4D). Notably, D10A-1252ALFA improved editing efficiency by 10% to 40% at A3, A4, and A5, compared to the D10A-GCN4/gp41 counterparts. Similar to GCN4 and gp41, D10A-945/1246ALFA reduced byproducts and improved single-base editing precision, compared with ABE8eWQ (Fig. 4F, G).
The ALFA system generated a distinct editing pattern compared to SunTag and MoonTag when using the same embedding position, demonstrating that the type of Protein-Tagging system can impact editing outcomes. Since ALFA system is an excellent protein purification system19,20, ALFA Plug-in BE could be purified and would possess promising potential for basic and clinical research.
Optimization of plug-in BE via reordering deaminases and antibodies
To examine whether the order of antibody-deaminase fusion influences editing outcomes, we rearrange the components within the Plug-in BE framework. Firstly, we established the four different configurations of antibodies and deaminases within Plug-in ABE (Fig. 5A). As expected, the rearrangement shifted the editing window, as exemplified by ABE8eWQ-1059GCN4, which expanded the window to A9 and improved efficiency from 6% (ABE8eWQ) to 33% (Fig. 5B). Importantly, strategic reordering optimizes editing efficiency (Fig. 5C). For instance, at E21 and HEK site 2, ABE8eWQ-1059GCN4 enhanced efficiency from 5% of 1059GCN4-ABE8eWQ to 36%.
A Schematic of different orderings in Plug-in ABE8eWQ. B Impact of different orderings in Plug-in ABE8eWQ framework on editing windows (n = 2 biological replicates). Window is obtained by combining the editing results at HEK site 2, E21, ZAP70, and OCT4. 532GCN4-ABE8eWQ represents the combination of D10A-532GCN4 and AntiGCN4-ABE8eWQ. Other combinations are named in analogies. C, D Product analysis and precision analysis for different orderings in Plug-in ABE8eWQ framework at four gene loci (n = 2 biological replicates). E Schematic of different orderings in Plug-in ABE9. F Impact of different orderings in Plug-in ABE9 framework on editing windows (n = 2 biological replicates). G, H Product analysis and precision analysis for different orderings in Plug-in ABE9 framework at four gene loci (n = 2 biological replicates). Source data are provided as a Source Data file.
To extend this strategy, we incorporated ABE921, known for its minimal editing window, into Plug-in ABE9 (Fig. 5E). Remarkably, reordering within Plug-in ABE9 resulted in more distinct changes to the editing window of ABE9 (Fig. 5F). For example, ABE9-1059GCN4, ABE9-1059gp41, and 1059gp41-ABE9 exhibited narrower editing windows than ABE9. Likewise, reordering within Plug-in ABE9 led to optimization in editing efficiency and single-base editing precision (Fig. 5G, H).
Consistent with the above results, in Plug-in CGBE1 system3, reordering of eUNG and R33A also improved editing efficiency (Supplementary Fig. 9). Collectively, these results demonstrate that the strategic reordering of deaminases and antibodies can enhance the performance of Plug-in BE systems, allowing for the customization of specific editing patterns.
Given the ability of the Plug-in BE system to generate a diverse range of base editors with distinct editing patterns, we have provided a comprehensive comparative summary of recommended Plug-in BE variants (Supplementary Data 4). This summary outlines the advantages, limitations, and optimal applications for each variant, highlighting key parameters including editing windows, editing efficiency, product purity, and sequence motif preferences.
Systematic characterization of plug-in BE using self-targeting library
To systematically evaluate the editing performance of Plug-in BE, we constructed a Self-Targeting library8 containing 30,230 sgRNA sequences along with the corresponding target sequences (Fig. 6A). In MoonTag Plug-in BE3 system, D10A-532/1055/1246/1252gp41 exhibited wider editing windows, and their efficiencies were approximately 1.5 to 2 times that of BE3 (Fig. 6B). Of note, D10A-1246gp41 possessed a high efficiency at C5, C6, and C14 to C16, while D10A-1055gp41 improved at C1 to C13. Overall, the variants displayed an improvement in the TCN and CCD motifs, and a reduction in the GCN motifs (Fig. 6C). In addition, the variants reduced byproducts C-to-G/A (Fig. 6D, E). Similarly, MoonTag Plug-in Target AID expanded the editing window, reduced byproducts, and altered motif preferences (Supplementary Fig. 10C–H).
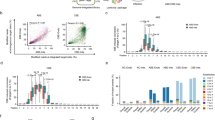
A Schematic of Self-Targeting library analysis. B Editing windows among BE3, D10A-532gp41, D10A-945gp41, D10A-1055gp41, D10A-1246gp41, and D10A-1252gp41 (n = 2 biological replicates). The Y-axis value represents the editing efficiency computed relatively to the highest (100%). 1246gp41 represents the combination of D10A-1246gp41 and Antigp41-BE3. Other combinations are named in analogies. C Sequence motif preference among BE3, D10A-532gp41, D10A-945gp41, D10A-1055gp41, D10A-1246gp41, and D10A-1252gp41 (n = 2 biological replicates). D, E Editing windows of byproducts C-to-G/A among BE3, D10A-532gp41, D10A-945gp41, D10A-1055gp41, D10A-1246gp41, and D10A-1252gp41 (n = 2 biological replicates). F Editing windows among ABE8eWQ, D10A-231gp41, D10A-532gp41, D10A-945gp41, and D10A-1055gp41. The Y-axis value represents the editing efficiency computed relatively to the highest (100%). 532gp41 represents the combination of D10A-532gp41 and AntiGCN4-ABE8eWQ. Other combinations are named in analogies. Data are mean ± SD (n = 3 biological replicates). G Sequence motif preference among ABE8eWQ, D10A-231gp41, D10A-532gp41, D10A-945gp41, and D10A-1055gp41. Data are mean ± SD (n = 3 biological replicates). Source data are provided as a Source Data file.
Library analysis of MoonTag Plug-in ABE8eWQ system further demonstrated the programmability of Plug-in BE, as different D10A-gp41 variants produced various editing windows (Fig. 6F). D10A-1055gp41 and D10A-231gp41 presented a right-shifted window with comparable editing efficiency, whereas D10A-532gp41 mainly improved the efficiency at A7 to A11. Moreover, the four variants exhibited different motif preferences (Fig. 6G), such as D10A-1055gp41 especially preferring TAC and GAC.
In summary, Self-Targeting library analysis fully demonstrates the detailed relationship between the spatial location of deaminase and editing outcome.
Advantages of Plug-in BE in gene therapy and zebrafish embryo editing
According to ClinVar data, nearly all of pathogenic single nucleotide variants (SNVs) consist of C-to-T, A-to-G, and C-to-G (Supplementary Fig. 11B, top). Importantly, Plug-in BE system expands the editable regions even further, thereby enabling the correction of additional SNVs, with the A-to-G corrections from 23.26% (ABE8eWQ) to 32.87% and the C-to-T correction from 43.53% (BE3/Target AID) to 77.73% (Supplementary Fig. 11B, down).
To demonstrate the therapeutic potential of Plug-in BE, we targeted the E6 and E7 oncogenes of HPV18 in HeLa cells to generate stop codons. The targeted cytosines of chosen sites are located at CCN motif and C11, which are typically edited inefficiently by BE3 and BE48. Remarkably, D10A-1246gp41 achieved an editing efficiency ranging from 10% to 37% at targeted cytosines, producing stop codons and reducing the cell growth rate (Supplementary Fig. 11C–E). Therefore, strategic adjustment of APOBEC1’s spatial position changes its motif preference and editing window, effectively overcoming the limitations of BE3 and BE4 in cancer gene therapy.
To further evaluate the in vivo editing potential of Plug-in BEs, we transcribed mRNAs along with sgRNAs targeting the zebrafish genes TWIST2 and TYR (Fig. 7A). The mRNAs included sequences encoding fluorescent proteins to facilitate the sorting of embryos expressing base editor (Fig. 7B). Firstly, we edited the TWIST2 site using the Plug-in ABE8eWQ system. Larvae edited by ABE8eWQ or D10A-231gp41 displayed phenotypes associated with TWIST2 dysfunction, including curved tails, body trunk malformations, and pericardial edema (Fig. 7C)22,23. Sanger sequencing showed that while ABE8eWQ primarily edited A6-to-G, D10A-231gp41 avoided the CA6G sequence and achieved A8-to-G with a 3.8-fold higher efficiency (Fig. 7C). Furthermore, amplicon sequencing of the top 5 larvae with the highest editing efficiency from Sanger sequencing revealed that D10A-231gp41 increased the only A8-to-G conversion from 10% (ABE8eWQ) to 59% of all mutational reads (Fig. 7D). Furthermore, we used the Plug-in BE3 system to edit the TYR. Sanger sequencing revealed that BE3 achieved approximately 21% C7-to-T, with only 3% C11-to-T, due to its restricted window. In contrast, D10A-1246gp41 achieved an average of 39% C11-to-T, generating a stop codon and resulting in albino larvae (Fig. 7F). To further validate these findings, we performed amplicon sequencing of the top 5 larvae with the highest editing efficiency from Sanger sequencing (Fig. 7G). As expected, sequence type analysis showed that BE3 generated 53.7% of reads with C7-to-G conversion, while almost no C11-to-T reads were detected. In contrast, D10A-1246gp41 produced over 75% of reads with C11-to-T conversion (Fig. 7H).
A schematic of the mRNA structure used for zebrafish embryo injection. An additional P2A-fluorescent protein is added to the C-terminal of protein to visualize protein expression. The sfGFP protein is green fluorescence and mCherry protein is red fluorescence. B Fluorescent embryos after RNAs microinjection were selected for subsequent DNA sequencing. A total of 137 injected zebrafish embryos showed detectable fluorescence at 6 h post-injection across all experiments. C Phenotype of zebrafish larvae with edited TWIST2 locus and corresponding editing results. 231gp41-ABE8eWQ represents the combination of D10A-231gp41 and Antigp41-ABE8eWQ. Data are means ± SD (ABE8eWQ, n = 22 biological replicates; 231gp41-ABE8eWQ, n = 35 biological replicates). D, E Editing product analysis (D) and representative editing profile (E) of ABE8eWQ and 231gp41-ABE8eWQ at TWIST2 locus by amplicon sequencing. Data are mean ± SD (n = 5 biological replicates). F Phenotype of zebrafish larvae with edited TYR locus and corresponding editing results. 1246gp41-BE3 represents the combination of D10A-1246gp41 and Antigp41-BE3. Data are mean ± SD (n = 40 biological replicates). G, H Editing product analysis (G) and representative editing profiles (H) of BE3 and 1246gp41-BE3 at TYR locus by amplicon sequencing. Data are mean ± SD (n = 5 biological replicates). Statistical significance was analyzed using two-sided unpaired student’s t test with 95% confidence interval. The exact p values are indicated in the graph. Source data are provided as a Source Data file.
Together, by overcoming the limitations of traditional base editors, Plug-In BE could achieve both efficient and safe editing in previously un-editable regions in gene therapy and zebrafish embryos with desired phenotypes. The great controllability and high fidelity of Plug-in BE could meet specific editing needs from both basic research and therapeutic applications.
Discussion
The editing capability of current DNA base editors is constrained by the limited spatial locations of deaminases1,2,3,4,12. In contrast, Plug-in BEs address the constraint and unleash the potential of base editors by strategically programming the spatial location of deaminase, resulting in the improvement of the editing efficiency, fidelity, and window.
In SunTag Plug-in BE3, many D10A-GCN4 variants outperform BE3, with higher efficiency and reduced byproducts. However, in SunTag Plug-in ABE8eWQ, only a few D10A-GCN4 variants maintained comparable efficiency to ABE8eWQ.To further improve Plug-in BE, we incorporated the smaller MoonTag system in Plug-in BE. For overall comparison, the efficiencies from Suntag Plug-in BEs and MoonTag Plug-in BEs were summarized as a dot plot (Supplementary Fig. 6). Interestingly, most of D10A-gp41 variants exhibited higher efficiency than D10A-GCN4 variants, while maintaining the high fidelity of Plug-in BE. The smaller size of gp41 epitope likely reduces its impact on the nCas9(D10A) function after embedding. Additionally, the gp41 antibody is only half the size of the GCN4 antibody, which may result in a shorter spatial distance between deaminase and DNA, contributing to higher editing efficiency. On the other hand, SunTag Plug-in BE performs better in reducing byproducts and indels, likely due to AntiGCN4’s larger size limiting excessive contact between deaminase and DNA.
By designing the editing window and motif preference, Plug-in BE addressed specific editing challenges that original base editors could not overcome. We successfully introduced the stop codons into HPV18’s E6 and E7 for cancer gene therapy using D10A-1246gp41 of Plug-in BE3, while both BE3 and BE4 could not edit the selected targets due to motif preference and window limitation8. In zebrafish TYR locus, D10A-1246gp41 of Plug-in BE3 efficiently edited C11-to-G, producing phenotype of albino larvae, while traditional BE3 could not edit C11 due to window restriction. The Plug-in BE’s ability to change motif preference and editing window means that modular combination can be programmed to not only reduce bystander editing but also achieve precise editing in gene therapy and live organisms.
Plug-in BE possesses the expansibility to be a general strategy for reprogramming editing patterns. The spatial location of deaminase is determined by adjusting the epitope embedding position within nCas9(D10A), generating base editors with diverse editing patterns. Interestingly, ALFA and MoonTag have similar sizes, but they produced different editing results, further suggesting that the type of Protein-Tagging system influences the editing pattern of deaminase. In Plug-in ABE8eWQ, Plug-in ABE9, and Plug-in CGBE1, changing the order of deaminases and antibodies yielded different editing outcomes. Thus, different orderings in the Plug-in BE framework create more opportunities for designed editing outcomes. Furthermore, inserting GCN4 into FrCas9 protein24 could also enhance base editing efficiency and broaden editing window (Supplementary Fig. 12). Together, programming epitope embedding positions, Protein-Tagging systems, Cas9 types and their orderings can create combinations of multiple editing patterns, and allow Plug-in BE to be customized for a wide range of editing tasks.
In conclusion, the Plug-in BE system uses CRISPR/Cas as a plug-in platform with many embedding epitopes as slots, Protein-Tagging systems, and deaminases as plug-ins, generating a series of DNA base editors with superior editing patterns and high-safety profiles (Supplementary Fig. 11A). Plug-in BE can serve as a modular and programmable DNA base editor toolbox that can be expanded with the increase of CRISPR protein, slots, and plug-ins, holding significant potential for both basic research and therapeutic applications.
Methods
Ethics statement
The experiments on zebrafish were approved by the Ethics Committee of Zhongnan Hospital of Wuhan University, Wuhan, China. Our experiments on zebrafish complies with the relevant ethical regulations.
Plasmid construction
The following plasmids with Addgene Catalog numbers were used for Plug-in BE plasmid constructions, BE3 (73021), BE4 (100802), ABE8eWQ (161815), ABE9 (194208), Target AID (131300), miniCGBE1 (140253), CGBE1 (140252), pST1374-scFv-APOBEC-UGI-GB1(113029), pST1374-GCN4-D10A (113022), MoonTag-Nb-GFP (128602), mEGFP-NbALFA (159986). KAPA HiFi HotStart Ready Mix (KAPA Biosystems, KK2602) and 2X MultiF Seamless Assembly Mix (ABclonal, RK21020) were adopted for plasmid construction according to the manufacturer’s protocol. During plasmid construction, pST1374-GCN4-D10A, which carries the resistance gene of blasticidin, served as the plasmid skeleton for the insertion of Plug-in BE elements. The DNA sequences of Plug-in BE variants are listed in Supplementary Data 1.
Cell culture and transfection
HEK293T cells (ATCC, CRL-3216) and HeLa cells (ATCC, CCL-2) were cultured in Dulbecco’s modified Eagle’s medium (Gibco, 11965118) supplemented with 1% penicillin-streptomycin (Gibco, 15140122) and 10% fetal bovine serum (Gibco, A5256701). The cell lines were passaged every 3 days and incubated at 37 °C with 5% CO2. The cell lines have been tested to exclude mycoplasma contamination. Cells were cultured in 24-well plates until the confluency was ~70%. Then, protein-expressing plasmids (500 ng) were co-transfected into cells with sgRNA plasmids (250 ng) using Hieff Trans® Liposomal Transfection Reagent (Yeasen, no. 40802ES08) according to the manufacturer’s protocol. All transfected cells were cultured for four days with blasticidin selection and then harvested for genomic DNA extraction. The sequences of sgRNA are listed in Supplementary Data 2, and the sequences of off-target sites are in Supplementary Data 3.
Detection of cell proliferation rate
The cells after trypsinization were seeded into 96-well plates. Adjust the cell density to 3000 cells per well. In the testing time points (24 h, 48 h, and 72 h), each well was added with 1/10 volume CCK8 (Abbkine, BMU106 - CN). The cells were then cultured in a 5% CO2 incubator at 37°C for 2 h. Finally, the absorbance value of cells was tested and recorded at 450 nm wavelength.
Genomic DNA extraction and PCR
Genomic DNA of transfected cells and zebrafish larvae was extracted with EasyPure® Genomic DNA Kit (Transgen, TEE101-01) according to the manufacturer’s protocol. The gene loci (200–600 bp) were amplified by PCR for Sanger sequencing. Primers are listed in Supplementary Data 2.
Editing efficiency analysis of Sanger sequencing data using EditR
PCR products were sequenced by Sanger sequencing technology (Sangon, Guangzhou). Sanger sequencing data were further quantified to obtain base editing efficiency using the online tool EditR (https://moriaritylab.shinyapps.io/editr_v10/)12.
Amplicon sequencing and data analysis
Genomic DNA was used for two-step PCR with KAPA HiFi HotStart Ready Mix. Primer pairs of first-step PCR with 25 cycles contained a gene-specific region and an outer Illumina-compatible adaptor sequence. Primer pairs of second-step PCR with 11 cycles had the outer-adaptor sequence to append a unique dual index. Then, the amplicon library was sequenced on MGISEQ-T7 platform. Base editing and indels efficiencies were determined by aligning reads to the genomic sequence using CRISPResso2. The percentage of editing product types, i.e., editing efficiency, was calculated as the desired editing reads / total sequencing reads. The percentage of single-base product types, i.e., editing precision, was calculated as the desired editing reads / total mutational reads. Amplicon primers are listed in Supplementary Data 2 and Supplementary Data 3.
mRNA sequencing
HEK293T cells were transfected with protein-expressing plasmids (500 ng) and sgRNA plasmid (250 ng). At 48 h post-transfection, RNA extraction was performed using TRIzol reagent (Invitrogen, 15596018CN). Subsequently, mRNA was reverse transcribed into cDNA using SuperScript III (Invitrogen, 18080051), and underwent high-throughput sequencing on the Illumina Novaseq-150PE platform. RNA-seq data were analyzed using RADAR (v1.0.0) for detection of RNA SNP.
Preparation of self-targeting library
The architecture and sequence of the Self-Targeting library with 30,230 sgRNAs were adopted from a previously reported article8. In brief, each oligonucleotide contains a full-length sgRNA with a corresponding target. The oligonucleotide pool was amplified by PCR with 10 cycles using KAPA HiFi HotStart Ready Mix, and then purified using magnetic beads (Cytiva, 65152105050250). The Lenti-gRNA-Puro plasmid (84752) was digested using BsmBI-v2 (NEB, R0739L). The pool PCR product was introduced into the digested Lenti-gRNA-Puro plasmid using 2X MultiF Seamless Assembly Mix. The recombinant plasmid was transformed into E. coli, and the colonies were scraped for plasmid purification. The lentivirus plasmid library was transfected into HEK293T cells for the construction of the lentivirus library, which was implemented by GURUN Biotechnology (Wuhan, China).
Characteristics of base editors through screening a self-targeting library
The Self-Targeting lentivirus library with an MOI of 1 was transduced into HEK293T cells for the construction of the Self-Targeting cell library. An sgRNA sequence cover at least 1000 cells. After 24 h transduction, the cell library was cultured with media containing 2.5 µg/mL puromycin for 10 days. Subsequently, the DNA base editor plasmid was transfected into the cell library. After 5 days of transfection, the genomic DNA of the cells was extracted for PCR amplification and amplicon sequencing.
Preparation of RNAs
mRNA preparation was performed using HiScribe T7 ARCA mRNA Kit (with tailing) (NEB, E2060S) according to the manufacturer’s protocol. The sgRNAs were modified with phosphorothioation and methoxy group, and synthesized by GenScript (Nanjing, China). The sequences of mRNAs are listed in Supplementary Note 1. The sequences of sgRNAs are listed in Supplementary Data 2.
Microinjection in zebrafish embryos
Wild-type AB strain zebrafish was purchased from China Zebrafish Resource Center, Wuhan, China. Zebrafish was maintained at 28 °C on a 14:10 hour light:dark cycle. Embryos were obtained from natural spawning of adult fish (3–6 months old, mixed male and female). Embryos at 1-cell stage were microinjected with a 2 nL mixture of mRNA (BE3 or ABE8eWQ, 100 ng/µL) and sgRNA (200 ng/µL), or Plug-in BE mRNAs (D10A-gp41 and Antigp41-deaminase, 100 ng/µL) and sgRNA (200 ng/µL). A total of 137 injected zebrafish embryos showed detectable fluorescence at 6 h post-injection across all experiments. Embryo genomic DNA was extracted at 72 h post-fertilization for assessing the base editing efficiency by Sanger sequencing. In each group, the top 5 embryos with the highest editing efficiency from Sanger sequencing were further sequenced by amplicon sequencing. Sex of zebrafish embryos was not considered in the study design because sexual differentiation in zebrafish occurs at approximately 3–4 weeks post-fertilization. The molecular mechanisms of base editing are not expected to differ between sexes at the embryonic stage. The animal experiments on zebrafish were approved by the Ethics Committee of Zhongnan Hospital of Wuhan University.
Statistics & reproducibility
No statistical method was used to predetermine sample size. No data were excluded from the analyses. The experiments were not randomized. The Investigators were not blinded to allocation during experiments and outcome assessment. The cultured cells from a homogeneous and well-mixed population were considered biologically equivalent. For every Plug-in BE combinations (a total of 533 combinations), we tested four genomic loci with two biological replicates (a total of 4264 biological replicates) to assess the base editing efficiency and ensure reproducibility. The Self-Targeting library cells contains 30,230 sgRNAs with >1000 cells per sgRNA, ensuring statistical robustness for identifying the editing pattern of Plug-in BE. Zebrafish embryos from the same clutch were collected sequentially and assigned to injection groups in the order of collection. A total of 137 zebrafish embryos were successfully injected with RNAs, resulting in detectable fluorescence at 6 h post-injection across all experiments (ABE8eWQ, n = 22; 231gp41-ABE8eWQ, n = 35; BE3, n = 40; and 1246gp41-BE3, n = 40). Statistical analysis was done using GraphPad Prism 8. Experiments were performed with at least two biological replicates. Error bars displayed on graphs represent the mean ± SD of at least three biological replicates. Statistical significance was analyzed using two-sided unpaired student’s t test with 95% confidence interval. The exact p value was represented on the graphs. All attempts at replication were successful.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The next-generation sequencing data generated in this study have been deposited in the NCBI Sequence Read Archive database under accession code PRJNA1221304. Source data are provided with this paper.
References
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Kurt, I. C. et al. CRISPR C-to-G base editors for inducing targeted DNA transversions in human cells. Nat. Biotechnol. 39, 41–46 (2021).
Yuan, T. et al. Optimization of C-to-G base editors with sequence context preference predictable by machine learning methods. Nat. Commun. 12, 4902 (2021).
Chen, L., et al. Adenine transversion editors enable precise, efficient A•T-to-C•G base editing in mammalian cells and embryos. Nat. Biotechnol. https://doi.org/10.1038/s41587-023-01821-9. (2023).
Richter, M. F. et al. Phage-assisted evolution of an adenine base editor with improved Cas domain compatibility and activity. Nat. Biotechnol. 38, 883–891 (2020).
Davis, J. R. et al. Efficient in vivo base editing via single adeno-associated viruses with size-optimized genomes encoding compact adenine base editors. Nat. Biomed. Eng. 6, 1272–1283 (2022).
Marquart, K. F. et al. Predicting base editing outcomes with an attentionbased deep learning algorithm trained on highthroughput target library screens. Nat. Commun. 12, 5114 (2021).
Arbab, M. et al. Determinants of base editing outcomes from target library analysis and machine learning. Cell 182, 463–480 e430 (2020).
Huang, T. P. et al. Circularly permuted and PAM-modified Cas9 variants broaden the targeting scope of base editors. Nat. Biotechnol. 37, 626–631 (2019).
Hu, J. et al. QBEmax is a sequence-permuted and internally protected base editor. Nat. Biotechnol. https://doi.org/10.1038/s41587-025-02641-9. (2025).
Liu, Y. et al. A Cas-embedding strategy for minimizing offtarget effects of DNA base editors. Nat. Commun. 11, 6073 (2020).
Tanenbaum, M. E., Gilbert, L. A., Qi, L. S., Weissman, J. S. & Vale, R. D. A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell 159, 635–646 (2014).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Komor, A. C. et al. Improved base excision repair inhibition and bacteriophage Mu Gam protein yields C:G-to-T:A base editors with higher efficiency and product purity. Sci. Adv. 3, eaao4774 (2017).
Jeong, Y. K. et al. Adenine base editor engineering reduces editing of bystander cytosines. Nat. Biotechnol. 39, 1426–1433 (2021).
Grünewald, J. et al. CRISPR DNA base editors with reduced RNA off-target and self-editing activities. Nat. Biotechnol. 37, 1041–1048 (2019).
Boersma, S. et al. Multi-color single-molecule imaging uncovers extensive heterogeneity in mRNA decoding. Cell 178, 458–472 e419 (2019).
Jedlitzke, B. & Mootz, H. D. A light-activatable photocaged variant of the ultra-high affinity ALFA-tag nanobody. ChemBioChem 23, e202200079 (2022).
Gotzke, H. et al. The ALFA-tag is a highly versatile tool for nanobody-based bioscience applications. Nat. Commun. 10, 4403 (2019).
Chen, L. et al. Engineering a precise adenine base editor with minimal bystander editing. Nat. Chem. Biol. 19, 101–110 (2022).
Zhao, Y., Shang, D., Ying, R., Cheng, H. & Zhou, R. An optimized base editor with efficient C-to-T base editing in zebrafish. BMC Biol. 18, 190 (2020).
Marchegiani, S. et al. Recurrent mutations in the basic domain of TWIST2 cause ablepharon macrostomia and Barber-Say syndromes. Am. J. Hum. Genet. 97, 99–110 (2015).
Cui, Z. et al. FrCas9 is a CRISPR/Cas9 system with high editing efficiency and fidelity. Nat. Commun. 13, 1425 (2022).
Acknowledgements
This work was supported by the Fundamental Research Funds for the Central Universities (grant no. 2042025YXA005 to D.H.); the National Natural Science Foundation of China (32171465 and 32371541 to Z.H. (Zheng Hu); 32500460 to C.Z.; 82102392 to R.T.; 82372672 to Z.H. (Zhiqiang Han)); the China Postdoctoral Science Foundation (2023M744121 to C.Z., 2023M734090 to R.T., and 2023M734091 to T.Z.); Joint Funds of Translational Medicine and Interdisciplinary Research of Zhongnan Hospital of Wuhan University (ZNJC202324 to D.H.); Support Project of Science and Technology Innovation Platform of Zhongnan Hospital of Wuhan University (PTXM2024016 to D.H.); Guangdong Special Support Plan Young Top-notch Talent (0820250 to L.Y.); the National key research and development program (2024YFC2707404 to Z.H. (Zhiqiang Han)).
Author information
Authors and Affiliations
Contributions
C.Z., R.T., T.Z., X.S., and L.Y. contributed to the experimental design; C.Z., R.T., T.Z., Z.L., X.S., and L.Y. performed the experiments; C.Z., Z.L., L.Y., X.S., and W.Z. analyzed data and drew figures; C.Z., R.T., T.Z., Z.L., X.S., and L.Y. drafted the manuscript; Z.H. (Zheng Hu), R.T., Z.H. (Zhiqiang Han), and D.H. managed the whole project; Z.H. (Zheng Hu), R.T., Z.H. (Zhiqiang Han), and D.H. revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
C.Z., R.T., T.Z., and W.Z. declare the following competing interest: C.Z., R.T., T.Z., and W.Z. are employees of Generulor Co., Ltd., a biotechnology company focused on genome editing technologies, while Generulor Co., Ltd. had no role in study design, data collection and analysis, and publication. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhong, C., Yu, L., Zhao, T. et al. A Plug-in system for reprogramming the editing patterns of base editors. Nat Commun 17, 910 (2026). https://doi.org/10.1038/s41467-025-67632-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67632-5