Abstract
Chronic stress is a risk factor for neuropsychiatric disorders, making the ability to adapt to repeated stress a crucial determinant of mental health. It remains unclear whether repeated stress exposure leads to habituation or new adaptive responses. Here, we investigated how the transcriptomic response to acute restraint stress changes – in the mouse ventral hippocampus – with repeated exposure. Using bulk RNA-seq across multiple timepoints, we observed widespread transcriptional habituation: stress-induced gene expression was blunted without the emergence of new response profiles or changes in baseline gene expression. Temporally resolved single-nucleus multi-omic profiling (RNA and chromatin accessibility) confirmed that the transcriptional habituation occurs across cell types, revealing cell type-specific patterns. We identify gene clusters whose expression habituates with repeated exposure, which appear primarily related to two distinct mechanisms: an early blunting of cAMP-associated genes chiefly linked to a reduced number of activated cells, and a shortened transcriptional response of corticosterone-associated genes, independent of the number of activated cells. Our study provides a comprehensive resource, accessible as an interactive app, that characterizes the dynamic transcriptomic response to stress in the mouse hippocampus and how it habituates with repeated exposure to stress.
Similar content being viewed by others
Introduction
In moments of acute stress, our bodies rapidly mobilize energy resources to ensure survival and initiate a complex molecular response to meet increased metabolic demands1. These widespread molecular changes resemble a symphony orchestrated by numerous neurotransmitters, peptides and hormones1,2. Temporally, the response can be broadly divided into fast and slow components. The fast component, occurring within milliseconds, is mediated by the release of neurotransmitters and peptides like glutamate, GABA, corticotropin releasing hormone (CRH), noradrenaline or dopamine. The slow component is mediated primarily by the hypothalamus-pituitary-adrenal (HPA) axis, with corticosterone (CORT) reaching the brain only several minutes after stress initiation3,4,5,6,7. At the transcriptional level, where our molecular understanding of the acute stress response is most detailed, gene expression unfolds in waves and concludes within several hours after stress initiation8,9. By manipulating individual signaling pathways, studies begin to reveal which stress-mediators contribute to these stress-induced changes4,10,11,12.
To prevent wear-and-tear, exhaustion, and the onset of stress-related disorders, the stress response must adapt when stress occurs repeatedly and becomes chronic13,14. However, adaptation to stress can come in many flavors. First, repeated stress exposure can lead to habituation, the reduced responsiveness to a repeated stimulus15,16. Habituation has been described, particularly for the HPA axis, across a variety of different stressors17,18,19,20. On a molecular level, stress-induced expression of immediate early genes (IEGs, like Fos or Arc) also typically habituates after chronic stress exposure21,22,23,24,25. Second, stress sensitization can occur when repeated presentation of the same stressor, or different stressors, triggers a more intense response than the first presentation. This process is often associated with heightened arousal in post-traumatic stress disorder, or with increased cortisol release from hypertrophied adrenal glands26,27,28,29,30. Third, chronic stress exposure can lead to the emergence of novel molecular and circuit adaptations that differ qualitatively from acute stress exposures, effects that have been studied particularly in the context of stress resilience14.
On a genome-wide level, it remains unknown how the acute stress response itself adapts after repeated stress exposure, and which mechanisms might be driving it. Most of the work on the transcriptomic consequences of chronic stress has focused on long-term alterations that manifest in the brain after extended periods of chronic stress31. Large gene expression screens have revealed widespread changes in stress-sensitive regions like ventral hippocampus (vHC), prefrontal cortex, hypothalamus or nucleus accumbens, but tissue was collected long after the termination of the last stressor32,33,34,35. To understand how the transcriptomic response to the stressor itself adapts when the stressor is presented repeatedly, we chose a widely used restraint stress model and dynamically profiled multi-omic changes across multiple time points after the first and subsequent stress exposures in the mouse hippocampus. We describe, in both sexes, how the transcriptional response to acute restraint stress unfolds over time. We show profound transcriptional habituation and blunted chromatin remodeling across different cell-types as the stressor becomes chronic, and we identify two distinct mechanisms associated with different temporal components of stress habituation.
Results
Transcriptomic dynamics after acute restraint stress in the hippocampus
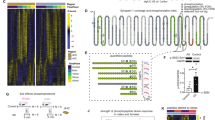
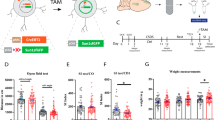
Acute restraint stress (ARS) is a commonly used stress model, which activates the HPA axis and leads to strong transcriptional responses in various brain regions25,36. Because the dynamic molecular changes triggered by ARS have never been characterized in detail, we exposed adult male and female mice to 1 h30 of restraint stress, and collected the ventral hippocampus (vHC) for bulk RNA-sequencing at baseline, 45 min, 1 h30, 3 h, 5 h30 and 24 h after stress initiation (Fig. 1A). Consistent with previously described transcriptional responses to various acute stressors9,37, we detected strong gene expression changes 45 min after stress initiation, which subsequently evolve in waves and resolve within several hours (Fig. 1B). While some differentially expressed genes were still detected 5h30 after stress initiation, none were detected 24 h afterwards (Fig. 1D). These dynamics and the overall return to baseline is also visible in a multi-dimensional scaling representation of the gene expression response (Fig. 1C, left). In previous work we showed that after an acute, short swim stress exposure, transcription had fully returned to baseline already 4 h after stress exposure9. To test whether the effects at 5 h30 represent lingering transcripts from earlier transcription, we looked at unspliced reads as a proxy for active transcription. We no longer observed significant group differences at 5 h30, suggesting that transcription had indeed returned to baseline at this point (Fig. 1C, right).
A Design of the time-course profiling of the transcriptomic response to ARS and CRS. B Relative gene expression of all genes affected by ARS. C Leading logFC multi-dimensional scaling of the stress-responsive genes for the full transcriptome (left) and the active transcription (unspliced reads, right). D Volcano plots showing significantly (FDR < 0.05 & >30% difference) expressed genes (in blue) at each time point following ARS. E Relative gene expression of all stress-responsive genes showing a general blunting of the transcriptional response after 10 and 20 days of CRS. F The lack of a global deviation in CRS:stress interaction coefficients across the two CRS durations indicates that overall, the two CRS durations elicit similar transcriptional habituation. Shown is the union of genes with a significant interaction after either 10 or 20 days CRS. The line is a fitted robust linear model, and reported are the Pearson correlation coefficient and associated p-value. G Key example genes showing similarly blunted response across the two CRS durations. The shaded area represents the standard error around the mean. H Volcano plot of the differential gene expression at baseline between CRS and “handling control” samples, showing no significant changes. The dashed line indicates a 0.05 FDR threshold. The animal numbers in each group are indicated in Supplementary Fig. 1. Some icons in this figure were created in BioRender, von Ziegler, L. (2025) https://BioRender.com/wmx3joz.
Global transcriptomic habituation in response to repeated stress
To examine how the molecular response to ARS changes after repeated stress exposure, we subjected male and female mice to chronic restraint stress (CRS). CRS is one of the most widely used chronic stress models, it leads to habituation and results in long-term behavioral, molecular, and structural changes in the hippocampus and other brain regions38. As the transition from acute to chronic stress is hard to define, we exposed mice to 10 or 20 days of daily 1 h30 restraint stress. The vHC was collected for bulk RNA-sequencing at baseline, 45 min, 1 h30, 3 h, 5 h30 and 24 h after the last stress initiation (Fig. 1A). Due to large cohorts, the experiments had to be conducted separately. Thus, additional handling control groups were added to both the 10 d and 20 d CRS experiments, for which tissue was collected at baseline, 45 min and 24 h after an ARS exposure. This allowed us to compare groups across cohorts, but it also enabled detailed within-cohort comparisons. Each group contained 8 mice, and each mouse served as a biological replicate (192 mice in total, a detailed schematic of the experimental design is provided in Supplementary Fig. 1).
Across both sexes and treatment durations (10 and 20 days of CRS), we observed a profound habituation of the transcriptional response to ARS in repeatedly stressed animals. Virtually all stress-responsive genes showed a reduced response in at least one time-point in CRS animals (Fig. 1E). Surprisingly, we did not observe any overall difference in the magnitude of habituation between 10 and 20 days of CRS, as indicated by a strong correlation between the CRS:ARS interaction fold-changes, which captures how each CRS treatment impacts the response to ARS (Fig. 1F and Supplementary Fig. 2A). This reveals that, at the level of gene expression, 10 days of CRS are sufficient to elicit strong transcriptional habituation of the acute stress response. Figure 1G shows the change in expression of prominent stress-responsive genes across the 3 treatment groups. Finally, we could not observe any significant expression differences between CRS and handling animals at baseline (Fig. 1H). Although surprising given the extensive literature reporting long-term changes following chronic stress exposure38, this indicates that the habituation after CRS is not the result of a change in baseline gene expression. Thus, we next explored which mechanisms could account for the blunted gene expression response.
Different mechanisms are associated with the temporal dynamics of habituation
The profiling of transcriptional changes over multiple time points allowed us to follow the temporal dynamics of stress habituation in unprecedented detail. To quantify these dynamics, we used the angle on the centered multidimensional scaling plot as a pseudo-temporal labeling of the samples across the global stress response (Fig. 2A). This revealed that—in comparison to the first stress exposure — the global transcriptional response in CRS animals (10 or 20 days) was reduced at 45 min. At 1 h 30 the molecular response peaked similarly in all groups, but returned to baseline much faster in CRS animals (Fig. 2B). To further explore these different temporal dynamics, we used spectral clustering to capture groups of genes with similar transcriptional dynamics across time (Supplementary Fig. 2B). Having previously shown that most transcriptional downregulation following acute stress is explained by RNA half-lives rather than direct transcriptional control9, we restricted the clustering to upregulated genes (see downregulated in Supplementary Fig. 2B and F). We resolved three main habituation patterns (Fig. 2C): Cluster 1 captured transcripts that were blunted already at 45 min. Cluster 2 captured transcripts that peaked normally in CRS animals, but returned to baseline faster. Cluster 3 captured transcripts that were normally upregulated at 3 h but failed to do so in CRS animals. In the unspliced RNA fraction, most genes in cluster 3 show comparable active transcription at 1 h 30 (Supplementary Fig. 2C), suggesting, also in this case, a faster return to baseline. Enrichment analyses comparing the genes in each cluster to all expressed genes or to all stress responsive genes revealed converging results (Fig. 2D and Supplementary Fig. 2E): The genes blunted already at 45 min (cluster 1) were especially enriched for CRH and Orexin receptor signaling pathways, as well as GO terms related to the vasculature (Fig. 2D). Genes in this cluster particularly included neuronal IEGs, such as Fos, but also early response genes from the vasculature such as Apold1 (Fig. 2E, left). Instead, genes that peaked normally at 1h30 but returned to baseline faster in CRS animals (cluster 2) were enriched for response to abiotic stimulus and oxygen levels, as well as response to corticosteroids (Fig. 2D). In line with this, one of the most specific glucocorticoid-responsive genes, Fkbp539,40, was normally upregulated at 1h30, but significantly blunted at later time points (Fig. 1G), and the glucocorticoid regulated kinase Sgk1 showed a similar pattern (Fig. 2E, center). The genes ablated at 3 h (cluster 3) did not show enrichment that passed multiple testing correction, yet some exemplary genes are shown (Fig. 2E, right panel). Testing whether the three clusters showed enrichment for curated gene targets of transcription factors (based on the collecTRI regulons41) yielded corroborating results (Supplementary Fig. 2D): Genes in cluster 1 were most prominently enriched for targets of the cAMP response element binding CREB1 (2.3-fold, FDR~5e-4), as well as other factors including the Serum Response Factor (SRF). In contrast, genes from cluster 2 were most strongly enriched for targets of the glucocorticoid receptor (GR) (3-fold, FDR~1e-7), with no target enrichment for cluster 3. To further confirm the involvement of GR signaling, we compared the response to ARS with the hippocampal gene expression response to an IP injection of the GR agonist dexamethasone (DEX)11. The vast majority of genes upregulated 3 h after ARS (which are all blunted in CRS animals, whether in cluster 2 or 3) were also upregulated by DEX injection (Fig. 2F and G). Conversely, we had previously shown that in vitro treatment of CA1/CA3 hippocampal slices with the cAMP activator forskolin42 largely recapitulates—in the absence of corticosterone signaling—many of the early gene expression changes seen in response to acute stress exposure1. Indeed, we replicate this in the current dataset, where most of the genes blunted at 45 min (cluster 1) were upregulated by forskolin in vitro (Fig. 2H). In contrast, this was true only for a small minority of those genes that return faster to baseline (cluster 2), and only for 1 gene from those genes with ablated expression at 3 h (cluster 3). These observations suggest that there are two mechanisms, a rapid, GR-independent early blunting that is related to cAMP signaling, and a slower, GR-dependent blunting that emerges at later time points.
A The angle on the multidimensional scaling (MDS) plot provides a pseudo-time labeling of the samples (shown are the animals without chronic stress). B Using the MDS angle of each sample as pseudo-time, one can observe a significant blunting of the early response in CRS animals, as well as a more rapid return to baseline. Box plot bars indicate the median, and the boxes the first and third quartiles. Both treatments pooled for testing using two-sided Wilcoxon tests (0 min W = 132 p = 1.0e-01; 45 min W = 325 p = 1.7e-05; 1 h 30 W = 83 p = 2.6e-01; 3 h W = 0 p = 2.7e-06; 5 h 30 W = 3 p = 2.9e-05; 24 h W = 205 p = 7.3e-01). Significance reported in the panel is after multiple testing correction through Holm’s method. C Main patterns of transcriptional habituation across upregulated genes. D Gene Ontology/Wikipathway over-representation analysis across the blunted clusters (Fisher’s exact test followed by FDR correction). E Example genes from each blunted cluster. The shaded area represents the standard error around the mean. F A comparison with DEX IP injection from Gerstner et al., 202211 shows that the changes observed at 3 h ARS are recapitulated by DEX IP injection. G Scatterplot of the logFC of the union of genes changed after 4 h DEX IP or 3 h ARS. The blue line is a fitted robust linear model, and reported are the Pearson correlation coefficient (‘cor’) and associated p-value. H Expression of the blunted clusters in hippocampal slices upon forskolin-induced LTP42 recapitulates the upregulated of the genes blunted at 45 min, but not of the other clusters, as expected given the absence of GR signaling. The animal numbers in each group are indicated in Supplementary Fig. 1.
Behavioral and neuroendocrine outcomes of habituation
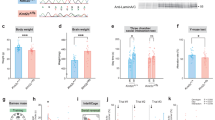
We were surprised that 10 days of restraint stress led to strong molecular habituation that was indistinguishable on the whole transcriptome level from 20 days of restraint. Thus, we asked whether behavioral changes, typically reported after 3 weeks of restraint stress exposure34, are also apparent after 10 days. Therefore, we compared the behavioral profiles of control, ARS and CRS male and female mice in the open-field test 45 min after the end of their first ARS exposure, or after 10 or 20 days of CRS exposure (Fig. 3A and B). In general, we observe an increase in distance moved and supported rearing upon both 10 d and 20 d CRS, indicating an increase in general activity compared to control animals (Supplementary Fig. 3A and C), with male and female animals generally responding similarly (Supplementary Fig. 3B and D). Additionally, we find a trend in CRS animals spending less time in the center of the arena. This suggests that behavioral changes emerge 10 days after stress exposure, which are similar to the changes observed after 20 days of stress exposure. Given the comparable behavioral response, and to increase power, we combined the animals of both experiments. This global analysis shows strong differences between those animals being stressed for the first time and those being repeatedly stressed. After CRS, mice are hyperactive (more movement and more supported rearing) and spend less time in the center, suggesting increased anxiety (Fig. 3C and D).
A Experimental Design of 10 d CRS experiment (n = 16/group). B Experimental Design of 20 d CRS experiment (n = 16/group). C 10 d and 20 d CRS normalized to the respective control groups (1-way ANOVA, Tukey’s multiple comparison, Distance: F = 14.94, Time in Center: F = 5.383, Supported Rears: F = 4.816, Unsupported Rears: F = 1.489). D Distance covered for female and male animals separately after 10 d and 20 d CRS (2-way ANOVA, Tukey’s multiple comparison). E Experimental Design blood collection (n = 19-20/group). F Males (left) and females (right) are tested separately using a 2-way ANOVA (Treatment*TimePoint) with Sidak’s multiple comparison. Males: Two-way repeated measures ANOVA showed significant effects of treatment (p = 0.015), timepoint (p < 0.0001), and interaction (p = 0.035). 95% CI for treatment difference: [5.71, 46.51]. Females: Two-way repeated measures ANOVA showed a significant effect of time (p < 0.0001), but no significant effect of treatment or time × treatment interaction. 95% CI for treatment difference: [−50.16, 27.31]. G Expression of the DEGs at 3 h following ARS in the original CRS experiment and the replication experiment also used for cort sampling. H Schematic representation of the response coefficient. I Response coefficients show a significant transcriptional habituation. Significance was assessed through a linear regression of the response coefficients on the group (homecage, 3 h ARS original, or 3 h ARS replication), the treatment (10 days CRS or handling), and their interaction. The significance of the following coefficients were tested using two-sided t-statistics: ARS_original:10dCRS \(\beta\)= −0.47 + /−0.1 t = −4.9 p = 5.7e-06; ARS_replication:10dCRS \(\beta\)=−0.29 + /−0.09 t = −3.2 p = 2.2e-03; ARS_original:20dCRS \(\beta\)=-0.48 + /−0.1 t = -5 p = 3.5e-06. The boxplot lines indicate the median and the boxes the inter-quartile range. Error bars in C, D, F represent means ± SEM. *adjusted p < 0.05, **adjusted p < 0.01, ***adjusted p < 0.001, ****adjusted p < 0.0001. Source data of behavior and CORT measurements are provided as a Source Data file. Some icons in this figure were created in BioRender, von Ziegler, L. (2025) https://BioRender.com/wmx3joz.
Given that habituation of the HPA axis has been well-documented, and because we see a clear involvement of GR in the blunting of certain clusters of genes (Fig. 2C and F), we measured corticosterone (CORT) levels in the blood in response to an ARS challenge presented either for the first time (ARS) or after 10 days of restraint stress (CRS-ARS) (Fig. 3E). In both sexes, CORT levels rose after restraint in both conditions, with only a modest blunting of the response in males and no detectable blunting in females (Supplementary Fig. 4A). To verify these findings, we repeated the experiment in an independent replication cohort in which we also collected the hippocampus after the final blood sampling to assess the impact of CRS on CORT and transcriptional blunting in the same animals (Supplementary Fig. 4B). This replication confirmed that after 10 days of CRS both males and females still mounted a strong CORT response (Supplementary Fig. 4B). When data from both cohorts were pooled, a moderate blunting of CORT levels was detectable only in males at the 45 min timepoint, but CORT levels peaked similarly at 90 min (Fig. 3F). RNA-seq confirmed that hippocampal gene expression changes were again blunted after CRS in both sexes (Fig. 3G), showing a disconnect between CORT levels and transcriptional blunting. In line with this, we found no correlation between CORT levels and expression of the GR-sensitive gene Fkbp5 (Supplementary Fig. 4C). To compare CORT levels with the overall strength of transcriptional habituation within each animal, we developed a response coefficient score by regressing each sample’s response on the average logFCs of the differentially-expressed genes at 3 h in the original ARS cohort (Fig. 3H and methods). This confirmed the strong habituation effect at the transcriptional level (Fig. 3I), but revealed no significant correlation between CORT levels and the transcriptional response coefficient (Supplementary Fig. 4D). Together, these results strengthen the conclusion that after 10 days of CRS, peripheral CORT levels peak normally in males and females, while the transcriptional response in these animals is strongly blunted.
Whole-tissue chromatin accessibility does not explain molecular habituation
To test whether changes in chromatin prevent stress-induced activation of gene expression after repeated stress exposure, we performed ATAC-seq of male and female mice that were either handled or stressed for 10 days both at baseline and 45 min after ARS initiation (Fig. 4A). Notably, analyses were performed in the same animals that were used for RNA sequencing, by using the contralateral hemisphere of the ventral hippocampus. We chose the 45 min time point, because earlier work had shown that within an hour after stress exposure clear chromatin remodeling can be observed43. Indeed, we observe differences (particularly increases) in peak accessibility, upon ARS (Fig. 4B and C). A differential promoter accessibility analysis yielded several genes with increased accessibility upon ARS (Fig. 4D), including known stress-responsive genes such as Fkbp5 or Nfkbia, and lesser-known genes like Phyhd1 (Fig. 4D and E). We correlated changes in promoter accessibility upon ARS with changes in RNA, using the union of genes that were significant in either ATAC-seq or RNA-seq. Since RNA changes persist much longer than changes in active transcription, and DNA accessibility relates to the latter, we focused on unspliced RNA at 45 min, and observed a weak, but significant correlation (Fig. 4F, left). Of note, some genes (e.g., Hif3a and Fkbp5) show an increase in promoter accessibility already at 45 min, while the effect on gene expression can only be observed at later time points, indicating that chromatin accessibility foreshadows changes in transcription. We therefore performed the same comparison with the transcriptome at the next time point (1 h 30), and observed a slightly higher correlation (Fig. 4F, right).
A Experimental design of the CRS experiment. B Heatmap of the relative accessibility in statistically significant peaks (log2FC are relative to the average of the Handling 0 min samples). C Volcano plot of the differentially-accessible peaks upon 45 min ARS (peaks with FDR < 0.05 are in blue). D Volcano plot of the differential promoter (2 kb upstream the TSS) accessibility 45 min after ARS. Significant (FDR < 0.05) genes are in blue, and the top ones are labeled. E Tn5 insertion coverage across samples in the two promoters most significantly affected by ARS at 45 min, Fkbp5 (top) and Phyhd1 (bottom). F Correlation between changes in RNA and changes in the corresponding promoter accessibility upon stress. In the left panel, the change in promoter accessibility 45 min after ARS is correlated to the change in active transcription (unspliced reads) at the same time point, while in the right panel it is correlated to changes in the full transcriptome at the next time point (1 h 30). Reported are the Pearson correlation coefficients and associated p-values. G Volcano plot of the differential motif accessibility upon ARS (labeled are those with FDR < 0.05). H Relative motif accessibility of the most relevant representative of each of the three families of significant motifs The lines indicate the median and the boxes the inter-quartile range (Handling+ARS-0: n = 8; CRS + ARS-0: n = 8; Handling+ARS-45: n = 7; CRS + ARS-45: n = 7). Some icons in this figure were created in BioRender, von Ziegler, L. (2025) https://BioRender.com/wmx3joz.
We next searched for transcription factors whose putative binding sites changed in accessibility (Fig. 4G and H). The most significant motif was of the glucocorticoid receptor (as well as the very similar motifs of other hormone receptors), which showed an increase in accessibility, followed by MEF2-related motifs, showing decreased accessibility. Finally, we also observed a weaker, but significant increase in accessibility of the JUND motif (Fig. 4G and H).
While we could identify major changes in accessibility upon ARS, we could not detect any change associated with CRS, either at baseline or in interaction with ARS. This means that 10 days of CRS do not alter ARS-induced chromatin remodeling. Even when focusing on the genes blunted at 45 min or 1 h 30, we could not observe any general pattern in the accessibility of their promoter in response to CRS. Thus, chromatin accessibility does not explain the habituation in gene expression after repeated stress exposure. However, accessibility changes occur faster and are more transient than RNA changes, thus it is possible that we have missed CRS-associated changes in accessibility and TF activity that have resolved at the 45 min time point. Additionally, it is possible that bulk ATAC-seq lacks the sensitivity to detect CRS-related changes, especially if they happen in a subset of cells. To address these two caveats, we turned to single-nuclei analysis of both transcriptome and chromatin accessibility across several time points.
Transcriptomic single-cell dynamics of the acute stress response
It has recently been shown that stress-induced changes have cell-type specific effects, and that the analysis of bulk tissue can obscure biologically meaningful changes44,45. To test how habituation to the acute stress response evolves over time across cell types, we exposed female mice to ARS after 10 days of handling (handling-ARS), or after 10 days of chronic restraint stress (CRS-ARS), and collected the vHC at 15 min, 45 min and 3 h after initiation of the last stress exposure (n = 3 mice per group) (Fig. 5A). We performed simultaneous single-nucleus RNA-seq and ATAC-seq on the same nuclei (10X Multiome). We sequenced a total of 215’349 nuclei across 23 samples, and after stringent quality filtering (see methods) we retained ~160’000 cells with a median of 7343 UMIs per cell. Non-neuronal cells included oligodendrocytes and oligodendrocyte progenitor cells (OPC), with a fraction of committed oligodendrocyte progenitors in-between (Fig. 5B). Microglia and astrocytes had prominent clusters, and we could also identify perivascular macrophages (PVM). Although lower in number, we could observe clear populations of pericytes, endothelial cells, smooth muscle cells, and some smaller subpopulations of vascular cells with unclear signatures, and we merged them as ‘vascular’ cells for downstream analysis.
A Experimental design of the CRS experiment and the multiomic pipeline. B UMAP of the neuronal cell populations in the transcriptomic space, colored and labeled by cluster. C UMAP of the neuronal populations in the ATAC space, showing a good segregation of the cell clusters identified from the RNA modality. D UMAP of the neuronal populations in the RNA space, in which the cells are colored by their Ventral-dorsal score based on the CA1/3 signature from ref. 46. Note that the signature is based on cells from the pyramidal layer, and might not be meaningful for other cellular populations (e.g., inhibitory neurons). E UMAP of the non-neuronal populations in the transcriptomic space. F UMAP of the non-neuronal populations in the ATAC space, coloring cells by their cluster from the RNA modality. G Overview of the global, dynamic response to ARS across broad cell classes. H Pseudo-bulk expression of genes found significantly changing upon ARS in more than one neuronal cell type. On top is the pattern of average scaled logFC of the displayed genes in Handling+ARS and CRS + ARS. I Pseudo-bulk expression of the genes significantly changing upon ARS in each broad class of non-neuronal cells (max 20 genes shown per cell type). On top is the pattern of average absolute scaled logFC of the displayed genes in Handling+ARS and CRS + ARS. n = 3mice per group per timepoint, except Handling+ARS at 45 min (n = 2mice). Some icons in this figure were created in BioRender, von Ziegler, L. (2025) https://BioRender.com/wmx3joz.
The different neuronal subpopulations are shown in Fig. 5C (marker expression is shown in Supplementary Fig. 5A). Inhibitory neurons clustered by known type (Sst, Pvalb, Vip, Ntng1 interneurons, and Lamp5 neurogliaform interneurons), while excitatory neurons clustered by region, first with a clear distinction between granule cells from the dentate gyrus, mossy cells from the hilus, and CA1 and CA3 excitatory neurons. Within the pyramidal layer (i.e., CA1-3), cells then clustered by relatively more dorsal vs ventral regions (Fig. 5C and D) based on signatures from Cembrowski et al.46. Dorsal scores were generally lower, and fewer cells had a high dorsal score, in line with the fact that only ventral hippocampi were analyzed (Supplementary Fig. 5B). This shows that the ventral-dorsal gene expression gradient is preserved – and can be resolved – even within the ventral portion. In addition, we identified a gradient of cells spanning from the CA1 to the pro-subiculum and the distal subiculum (Fn1 + ), as well as a separate population most likely from the posterior subiculum (with Ly6g6e and Cntnap4 expression). Another gradient especially characterized by high Camk2d expression extended from the ventral CA1 population. Based on correlations with the Allen brain atlas, they appear to be neurons from the hippocampus-tegmental amygdala (HATA) and/or hippocampus-inferior temporal cortex interfaces. These populations were the only ones to (mildly) vary in abundance across samples, consistent with their location on the border of the dissected tissue. Of note, the transcriptome-based cell type labels were also clearly segregated in the ATAC-seq embedding, both for non-neuronal (Fig. 5E) and neuronal (Fig. 5F) populations, corroborating the robustness of the cell type assignment.
To date, no dynamic profiling of the acute stress response on the single-cell level exists in the hippocampus. Therefore, we first examined the transcriptional changes to ARS on a pseudo-bulk level (Supplementary Fig. 6). Globally, the different broad cell types all responded to ARS early on (Fig. 5G). In neuronal cell types, the response peaked at 15 min (inhibitory neurons) or 45 min (excitatory neurons), before strongly decreasing at 3 h. In contrast, the response of all non-neuronal cell types was sustained at 3 h. In neurons, a particularly strong response was observed in CA1 pro-subiculum and VIP interneurons, which upregulated a number of major IEGs already 15 min after stress initiation, such as Fos/Fosb, Npas4, and Btg2 (Fig. 5H). With the exception of vascular cell types, which also responded very quickly, other non-neuronal populations were slower to respond (Fig. 5I). The significant genes in non-neuronal populations included a number of prominent stress-induced genes, such as Apold1 in vascular cells47, the cell cycle regulator Cdnkn1a, or the GR target Fkbp5 across multiple cell types. Of note, some changes, such as the strong upregulation of the cAMP/cGMP-degrading enzyme Pde10a specifically in astrocytes upon stress (Fig. 5I and Supplementary Fig. 6), had not been detected in the bulk transcriptome, presumably because the gene is also highly expressed in other glial populations at baseline, underlining the value of single-cell profiling.
Transcriptomic habituation to repeated stress occurs across cell-types
Visual inspection of the heatmaps in Fig. 5H and I suggests that across cell types, the response to ARS was generally blunted after CRS, with different temporal patterns emerging for different cell types. To investigate this systematically, we tested, within each cell type, which stress-induced gene expression profiles were altered by CRS at any time point. Several genes were significantly affected by CRS, with a clear pattern of ARS-responsive genes habituating in CRS animals (Fig. 6A). This was also the case for several prominent stress-responsive genes (Fig. 6B). Notably, while Npas4 upregulation appeared to be completely ablated in CRS animals at 45 min based on the bulk transcriptomic data, the single-cell data revealed an upregulation at 15 min also in CRS animals across multiple excitatory neuron subtypes (Supplementary Fig. 7B). However, in CRS animals this upregulation was weaker, and had already returned to baseline at 45 min across cell types, while it remained upregulated in animals who experienced the stressor for the first time (Fig. 6B and Supplementary Fig. 7B), consistent with the bulk data. In contrast to our bulk sequencing data, a number of genes appeared differentially expressed at baseline in CRS animals (Fig. 6C). While some of these candidates showed a variable expression pattern across ARS time-points (e.g., Cacna1e and Vwf), some looked more coherent, such as Pde1a in VIP interneurons, a gene that is involved in cGMP/cAMP homeostasis48. To ascertain these findings, we also specifically tested for differences at baseline, which revealed some statistically significant candidates (Supplementary Fig. 7A). While the identification of rare, cell-type specific baseline gene expression changes emphasizes the power of single-cell sequencing, these limited changes cannot explain the widespread habituation of stress-induced transcriptional activity across cell types.
A Relative expression of genes showing a significant impact of CRS on the stress response across cell types. B Key genes showing a significant habituation in CRS; shown are the cell types in which they are the most significant. C Candidate genes with a significant effect of CRS at baseline. Reported significance values are from a likelihood ratio test comparing the full model (ARS time points and CRS covariates, i.e., ~ARS*CRS) to a null model without CRS covariates (CRS p-value), and adjusting for multiple testing using Benjamini & Hochberg’s method. A significant CRS q-value means that CRS affects either the baseline gene expression or that the gene responds differently to ARS after CRS. Boxplot lines indicate the median and the boxes the inter-quartile range. A–C n = 3mice per group per timepoint, except Handling+ARS at 45 min (n = 2mice).
After habituation, fewer cells respond to stress
A blunted transcriptomic response to stress could be due to fewer cells responding, or to the same proportion of cells having a weaker response. cFOS or ARC stainings for instance indicate that only a fraction of hippocampal neurons respond to salient, stressful events36. Leveraging the power of single-cell transcriptomics, we estimated the recent activity of single neurons using an activity-dependent transcription (ADT) score based on genes that are differentially expressed upon ARS at the pseudo-bulk level (see methods). The ADT score indicates, for each cell, whether it looks more like a cell that does not respond to ARS (score of 0) or like the most stress-responsive cells of that type (score of 1). As expected, while only a minority of the cells showed ADT, these were unequally spread across subpopulations (Fig. 7A), with the highest activity in subiculum, pro-subiculum and CA1 neurons. Using an ADT score above 0.5 as a threshold for an activated neuron, we observed that the proportion of activated neurons doubled upon the first ARS exposure, a response that was blunted after CRS (Fig. 7B). In contrast, while the median ADT score of activated neurons did show a significant increase upon ARS, the difference was very small in magnitude (roughly 5%, Fig. 7C). Together, this indicates that most of the blunted response is due to fewer neurons being activated, rather than the same number of neurons being activated less strongly.
A UMAP of neuronal populations, colored by their activity-dependent transcription (ADT) score. Proportion of neurons activated (B) and median ADT score or activated neurons (C) across samples, indicating a decreased number of recruited neurons upon stress in CRS animals. D Proportion of activated neurons per pseudo-bulk sample across neuronal subpopulations, using the mean of the ADT and neuroestimator scores. E UMAP of non-neuronal populations, colored by their ADT score. F Proportion of activated cells across broad non-neuronal cell types. Reported p-values are from a likelihood ratio test, comparing the full model (ARS time points and CRS covariates, i.e., ~ARS*CRS) to a null model without the time points (ARS p-value) or without the CRS covariates (CRS p-value). A significant CRS q-value means that CRS affects either the baseline activation or that activation in response to ARS differs after CRS. The boxplot lines indicate the median and the boxes the inter-quartile range. All single-cell analyses: n = 3 mice per group per timepoint, except Handling+ARS at 45 min (n = 2mice).
We next repeated this analysis at the level of individual cell types, which showed that the habituation after CRS is cell-type specific, with the strongest blunting of ADT observed in VIP interneurons and in several excitatory cell populations (Supplementary Fig. 8). To corroborate these results, we also used an independent method to estimate neuronal activation from single-cell data, neuroestimator49. Although the estimated baseline activation levels across subpopulations varied depending on the methods, the main patterns upon ARS and CRS were largely reproduced for inhibitory cell types, with less congruence in excitatory populations (Supplementary Fig. 8). Figure 7D shows the proportions of each cell type activated based on the average of the single-cell scores from the two methods. Consistently, VIP interneurons showed the highest change in stress-induced activity. Strong responses in ADT were also observed in CA1 and subiculum populations, in mossy cells, as well as in other inhibitory neuron subtypes (Pvalb and Ntng1). Notably, the temporal dynamics varied across cell types, e.g., VIP interneurons were strongly activated already at 15 min, while other inhibitory neurons showed a more delayed activation. In all these populations, the strong ARS-induced change was blunted or completely absent after CRS, especially at 45 min (Fig. 7D).
Contrary to the neuroestimator method, our ADT approach is applicable to any cell type, and we thus used it to estimate the proportion of cells recruited across non-neuronal populations (Fig. 7E and F). Only a minority of vascular cells reacted to ARS, whereas the majority of astrocytes and oligodendrocytes strongly reacted upon first ARS exposure, with microglia and OPC showing intermediate activation. In all glial cell types, but not in vascular cells, the ARS-induced activation was blunted in CRS animals, especially at 3 h. This reduction was particularly strong for astrocytes, dropping roughly by half already from 45 min on. In contrast, oligodendrocytes seem to be recruited similarly in response to stress, but while their activity is sustained across 3 hours in handling animals, activity drops sharply at this time point in CRS mice.
Repeated stress shortens GRE accessibility across cell types
To investigate the mechanisms underlying the observed transcriptional changes following acute stress, and the blunting after chronic stress exposure, we next turned to the changes in chromatin accessibility across cell types. The strongest and most significant ARS-induced changes in motif accessibility consistently came from the glucocorticoid receptor and highly similar steroid hormone receptor motifs (Supplementary Fig. 9), with increased accessibility already at 15 min and extending to later time points (Fig. 8A). While the ARS-induced change in GR motif accessibility was similar between handling-ARS and CRS-ARS at 15 min across all cell types, we observed clear transcriptional habituation in the CRS-ARS group at 45 min in many cell types (Fig. 8A). After 3 h, GR motif accessibility completely returned to baseline in the CRS-ARS group, but not in handling-ARS animals (Fig. 8A). Notably, the increase in GR motif accessibility upon ARS was much weaker in interneurons, which also did not upregulate its target Fkbp5. Expression of the GR gene (Nr3c1) is low in interneurons, and across cell types we observed that the magnitude of the increase in GR motif accessibility in response to ARS was proportional to the expression of Nr3c1 (Fig. 8B and Supplementary Fig. 10A).
A GR motif accessibility across all cell types. B Overview of the relationship between GR (Nr3c1) expression (left in purple), relative changes in accessibility of Glucocorticoid Response Elements (GRE, center), and Fkbp5 mRNA expression (right) across cell types and treatment. C selection of top motifs responding in accessibility to ARS across neuronal cell types. To relate these to RNA, the expression of key IEGs is shown below in the corresponding samples. D–F Relative accessibility of selected motifs. G–J Relative accessibility of selected motifs in activated vs non-activated excitatory neurons (defined by an ADT threshold of 0.5). In A, D, E, F the reported significance values are from moderated F-statistics, comparing the full model (ARS time points and CRS covariates, i.e., ~ARS*CRS) to a null model without the time points (ARS p-value) or without the CRS covariates (CRS p-value). A significant CRS p-value means that CRS affects either the baseline motif accessibility or that the motif accessibility responds differently to ARS. Adjustment for multiple testing using Benjamini & Hochberg’s method. For G–J the models additionally included the activation state, and uncorrected p-values are reported because the motifs are hypothesis driven. All boxplot lines indicate the median and the boxes the inter-quartile range. For all single-cell analyses: n = 3 mice per group per timepoint, except Handling+ARS at 45 min (n = 2mice).
While the accessibility of glucocorticoid response elements (GRE) already increases 15 min after stress exposure, GR-responsive genes – like Fkbp5 – show transcriptional changes only at later time points. Across many cell types, Fkbp5 expression increased at 45 min, and expression was significantly blunted 3hrs after stress initiation (Supplementary Fig. 10B and Fig. 8B). These results suggest that, in both handling and CRS animals, glucocorticoids reach the brain and activate GREs already within 15 min after initiation of stress, but that following CRS, GRE accessibility is shortened, and accordingly GR-dependent transcription shuts down more rapidly. Further, it supports the hypothesis that much of the transcriptional habituation observed at later time points is GR-dependent, while the blunted response observed at 15 min is not.
Repeated stress induces differences in cAMP-associated TFs independent of GR activation
Apart from GR, we identified several other TF motifs that strongly respond to ARS (Supplementary Fig. 9; Fig. 8C–E), as well as candidate motifs showing baseline differences in CRS animals (Fig. 8F). In non-neuronal cell types, we particularly observed strong activation of CEBPA in microglia and NFIA in oligodendrocytes (Fig. 8D). In neurons, across stress-responsive neuronal cell types (i.e., especially CA1, pro-subiculum and VIP neurons), we could observe a strong increase in the accessibility of FOS/JUND motifs at all time points following ARS (Fig. 8C and E). However, this was not well coupled to changes in the accessibility of cAMP-response elements (CRE): while the CREB1 motif appeared to increase in accessibility at 15 min in Ntng1 and VIP interneurons (Fig. 8E), this was not the case in most excitatory neurons, where it instead showed a trend in the other direction (Fig. 8C). This was especially surprising given the clear recruitment of these neurons and the known rapid CREB-dependent transcription upon depolarization50,51. To confirm the expected involvement of CREB in this paradigm, we tested motif enrichment within regions accessible in neurons in proximity to genes that respond to stress rapidly (i.e., genes that are significantly increased by stress at 15 min in single-nuclei data or at 45 min in the spliced bulk transcriptome). Amongst several over-represented motifs, CREB1 showed the strongest enrichment (Supplementary Fig. 10C). A similar analysis using experimental binding data highlighted a number of factors (Supplementary Fig. 10D), but CREB1 was the only significant factor in the combined analysis (Supplementary Fig. 10E).
To further investigate the seemingly inconsistent CRE accessibility pattern, we looked at motif accessibility across activated (defined by ADT > = 0.5) and non-activated excitatory neurons (Fig. 8G–J). In activated neurons, we observed a strong increase in motif accessibility across several fast responding TFs like CREB1, SRF and JUN/FOS family members (Fig. 8G–I), confirming the expected increase in CRE accessibility in activated compared to non-activated neurons. The mild decrease in CRE accessibility in non-activated neurons (which are the majority) likely masks, in pseudobulk of the entire population (Fig. 8C), the increase from activated neurons. The accessibility of the TF’s respective motifs had differing temporal dynamics, which likely relate to how sustained the activity of the different TFs is following activation: SRF strongly spiked at 15 min but returned close to baseline already at 45 min (Fig. 8H), while JUND appeared more comparably active across activated neurons from different timepoints (Fig. 8I). Of note, the increase in SRF motif accessibility in activated neurons was significantly blunted in CRS animals, suggesting a difference in activation that cannot be attributed to fewer neurons being activated. This could be due to a difference in SRF response to neuronal activity or, given its fast spiking, a shorter neuronal activation.
In contrast to activity-dependent TFs, the temporal dynamics of GRE-accessibility were highly consistent between activated and non-activated neurons. Critically, in both populations we observed the faster return to baseline in CRS animals, again indicative of shortened GR retention. (Fig. 8J). Of note, activated neurons generally displayed a much lower GRE accessibility independently of the timepoint. This is not specific to GRE, but affected a large number of motifs, including lineage-specific TFs. This could be due to competition with activity-dependent transcription, and would be in line with similar observations after kainic acid treatment52. Together, these results indicate that the cAMP-signaling cascade implicated in early blunting of gene expression after stress habituation is linked to a reduced number of activated cells. In contrast, the shortened transcriptional response of corticosterone-associated genes, which is associated with a protracted blunting of the gene expression response after stress habituation, affects cells independently of their activation status. This confirms that the transcriptional habituation in the hippocampus of mice repeatedly exposed to restraint stress is driven by at least two, mechanistically distinct processes.
Discussion
Here, we explored the adaptation of the acute stress response on the level of gene expression and chromatin accessibility in the mouse hippocampus. While previous work had extensively profiled both the molecular changes induced by acute stress1,9, and the long-lasting changes after chronic stress31,33,34,53,54, it was unknown how the acute stress response adapts on a molecular level as a stressor becomes chronic. By profiling the transcriptomic response to acute restraint stress, we show that repeated restraint exposure for 10 or 20 days leads to strong transcriptional habituation across the vast majority of stress-responsive genes. Based on the temporal dynamics of this effect, we can dissociate a reduced activation of immediate early genes from a blunted and shortened response of glucocorticoid-sensitive genes.
Early GR-independent transcriptional habituation
Within minutes of an acute stress exposure, the transcriptional machinery becomes activated and gene expression changes can be observed4,55. These changes are pioneered by well-known IEGs, such as Fos, Npas4 and Apold1, which group together in a cluster that strongly habituates after repeated stress (cluster 1, Fig. 2C). Fos is known to strongly habituate to CRS on the transcript and protein level in various brain regions21,22,23,24. Both our bulk and snRNA-Seq results replicate Fos habituation at 45 min across cell types. Similarly, we find that virtually all stress-responsive IEGs show habituation at least at one time point. We observe that the members of cluster 1 are strongly enriched for CREB1 and SRF transcription factor activity, both crucial for IEG induction56. Indeed, members of cluster 1 are upregulated following forskolin treatment of hippocampal slices42 (Fig. 2H), but not in response to Dex, thus indicating that cAMP signaling is at least partially involved. Importantly, this correlation is not found for clusters 2 and 3, suggesting that upregulation of the targets in these clusters is largely cAMP-independent. Our single-nucleus ATAC-seq analysis further confirms the involvement of CREB- and SRF-dependent signaling.
What remains unclear, however, is whether after habituation some IEGs are blunted from the start, or whether their initial response is preserved but transcription shuts down faster. Npas4, for example, shows complete habituation 45 min after repeated stress in our bulk RNA-Seq, but we still find a significant – albeit weak – increase in Npas4 transcripts at 15 min in several cell types in our single-nucleus data. In a few cell types, however, Npas4 seems to be completely silenced across time points (Supplementary Fig. 7B). Due to the lack of a finer temporal resolution within the first 15 min of stress initiation, we cannot fully resolve the question whether IEG transcription is similarly initiated between ARS and CRS. However, our method for quantifying activity dependent transcription (ADT) allows us to make two critical observations: First, we find that, across cell types, the blunted transcriptional response to ARS after a history of CRS is chiefly due to fewer cells being activated, rather than the same proportion of cells showing a weaker activation. Second, activated neurons show a weaker increase in SRF motif accessibility at 15 min (Fig. 8), which could indicate either a shorter or weaker activation response.
GR-dependent transcriptional habituation
Acute restraint stress strongly induces HPA axis activation. While CORT levels in the blood rise within a few minutes, CORT is detected in the brain after 10 min5,6. On the bulk and single-cell level, our transcriptomic analyses reveal genes that peak similarly at 1 h 30 after stress onset, but shut down faster and return to baseline levels at the 3 h time point in animals with a history of chronic stress, while at that time point still being strongly upregulated after the first ARS exposure. Converging evidence shows that these transcripts are enriched for known GR target genes, such as Fkbp539,40, and they overlap with DEX-responsive genes from an independent analysis. In both our bulk and single-cell ATAC sequencing, the strongest ARS-induced changes were associated with an increase of GR motif accessibility, consistent with previous research43. After CRS exposure, this ARS-induced increase is largely blunted at 45 min and 3 h across cell types. However, we do observe a strikingly similar increase in GR accessibility at 15 min (Fig. 8A), thus indicating that indeed GR activation is initiated similarly to ARS, but shortened. Importantly, across neurons this late component of transcriptional habituation is independent of whether the cell was activated or not. CORT levels measured in the blood support the notion that the initial rise is similar in animals with or without a history of prior stress. However, at later time points, CORT dynamics paint a more complex picture, suggesting that CORT levels habituate slightly in males, but not in females. Notably, several reports find that the CORT levels habituate strongly in response to repeated restraint stress18,19,20, yet not all studies observe this habituation57,58 and we suspect that varying the time of day (unpredictable restraint stress) might have prevented HPA-axis habituation in our setup. While our results support previous reports of sex differences in HPA-axis regulation and a more pronounced CORT response in females59,60,61, they contrast with the fact that we observe no corresponding sex differences in overall transcriptomic habituation patterns. These data suggest that in males, a blunting of the HPA-axis response could account – at least to a small degree – for the blunted expression of GR-sensitive genes after repeated stress exposure. However, we think this link is weak for several reasons: (1) The CORT-blunting observed in males at a single time-point is a small effect, whereas the molecular habituation is strong and occurs across time points. (2) We observe no clear correlation between CORT levels and gene expression changes (Supplementary Fig. 4). (3) In females, where the transcriptomic blunting looks very similar to males, we detect no changes in CORT levels after repeated stress exposure. Notably, sex differences in glucocorticoid binding globulin have been reported60, thus it remains to be explored if this could contribute to differences in stress habituation. Another possibility, supported by the observation that the initial rise of GRE accessibility is comparable between groups, but returns to baseline faster in CRS animals, could be that the GR-dependent blunting is due to differences in the retention of GR in the nucleus, for instance through nuclear export or what has been proposed as the chaperon cycle62,63,64. More work is needed to reveal the mechanisms behind these unexpected effects.
Limitations, outlook and conclusion
As the acute stress response represents a dynamic system, and because the border between acute to chronic stress is blurry, we faced several challenges: (1) To capture temporal dynamics we had to collect samples at many time points across several hours; our data suggest that habituation of GR-dependent transcription is due to a more transient transcriptional activation, rather than a blunted increase in the first place. For GR-independent stress habituation that occurs much earlier after stress exposure, our single-nucleus data suggest that in some cell types the activation of IEGs might be completely abolished. However, we would need more time points during the first 15 min after stress initiation to firmly corroborate this claim. (2) We had to capture the gradual onset of adaptive changes over days to weeks. We find that both 10 and 20 days of CRS result in a very consistent behavioral phenotype as measured in the open field. Additionally, we do not find a difference in the transcriptional stress response between animals being exposed to 10 or 20 days of CRS. This indicates that the actual habituation process already happens earlier in time. Although some stress models find a clear reduction of the stress-induced CORT response already upon the second or third stress exposure20,65, we do not observe habituation of the HPA axis in our model after 10 or 20 days, and despite the fact that GR-signaling is clearly involved in the blunted transcriptional response, this is not due to changes in CORT levels in the blood. Thus, it remains to be tested after how many restraint stress exposures the transcriptional blunting in the hippocampus emerges. (3) Our work has only focused on the ventral hippocampus, thus we lack a brain-wide understanding of transcriptional habituation to repeated stress. It also remains unclear how brain regions other than the hippocampus contribute to the emerging behavioral phenotypes. Notably, the transcriptional habituation cannot be explained by a baseline shift, as we detect – across large numbers of samples – no significant differences in baseline gene expression levels between handled and chronically stressed animals. While this underscores that the transcriptional habituation can only be revealed when an acute stress challenge is imposed and the subsequent gene expression response is profiled dynamically across time, it remains unclear if baseline differences might be detectable in different brain regions. (4) Our study focuses on group effects and does not account for individual variance in the ability to habituate. Although the transcriptional and behavioral habituation itself was highly consistent across animals, the extent of habituation likely varies among individuals, highlighting a potential area for future investigation. (5) Due to the requirement of a chromatin accessibility readout, we profiled single nuclei instead of single cells, which restricts the RNA readout to the nucleus and excludes transcripts located in the cytoplasm and neuropil. While this approach prevents dissociation-induced transcriptional artifacts44, the resulting data is dominated by nascent transcripts66. This offers a clear snapshot of ongoing transcription, but lacks the cumulative representation of lingering transcripts seen in our bulk RNA data.
Using multi-omic bulk and single-cell sequencing, we identify two temporally distinct mechanisms that contribute to the emergence of genome-wide transcriptional blunting after habituation to a recurrent stressor. We have integrated these data with our previously published sequencing analysis of the acute stress response9, thus creating a searchable, open-source database. These data offer a detailed characterization of the transcriptional events that unfold in a stress-sensitive brain region as a stressor transitions from acute to chronic. These results lay the foundation to explore how habituation to repeated stress varies between individuals, and how the molecular changes associated with transcriptional blunting may relate to stress-related diseases.
Methods
Animals
C57BL/6J (C57BL/6JRj) mice were obtained from Janvier (France). Mice were maintained in a dedicated animal facility on a 12 h reversed light–dark cycle (lights off at 09:30 a.m.) with food and water ad libitum. The temperature in the animal rooms was maintained at 22–24 °C with a maximum deviation of ±1 °C. The relative humidity was kept at 50%, with a maximum deviation of ±10%. Male and female mice were housed, separated by sex, in groups of 2–5 animals per cage and used for experiments when 2–3 months old. All tests were conducted during the animals’ active (dark) phase. All experimental procedures were carried out in accordance with the guidelines of the Federal Veterinary Office of Switzerland and were approved by the Cantonal Veterinary Office in Zurich.
Stress paradigm
Mice were exposed to restraint stress for 1 h 30 in a 50 ml Falcon tube with a large air hole. All stress exposures were carried out between 10:00 a.m. and 5:00 p.m. Animals were single-housed 24 h prior to the final stress exposure to reduce confounding effects induced by cage mates on the day of the experiment61. For acute stress experiments, animals were exposed to one restraint session (1 h 30). The restraint tube was placed in the homecage for the duration of stress. For chronic stress experiments, animals were subjected to either 10 or 20 consecutive days of daily restraint stress at varying times of the day. Control animals and animals exposed to acute stress were gently handled daily for an equal number of days. Gentle handling involved lifting the animals by the tail, simulating the handling method used before restraint stress.
CORT measurement
To assess CORT levels, blood samples were collected via tail prick at 0 min, 15 min, 45 min, 90 min and 180 min after the onset of restraint. Blood samples were stored at RT for 30 min or 4 °C overnight to allow for clotting and centrifuged at 2500 g for 10 min at 4 °C. The serum supernatant was collected and stored at −80 °C until further analysis.
Corticosterone levels in the serum were measured using an ELISA kit (Invitrogen, EIACORT) according to the manufacturer’s instructions. A 1.5 μl serum sample was diluted 1:100 for the assay. Absorbance was measured at 450 nm with a reference at 560 nm using a plate reader, and final concentrations were determined based on the provided standards.
Tissue collection
At the indicated time point after initiation of stress, mice were euthanized by cervical dislocation and decapitation. When tissue was collected at 15 min, 45 min or 1 h 30 after stress initiation, mice were removed from the restraint tube at the indicated time point and immediately euthanized for brain collection. The brain was quickly dissected on a cool surface and isolated ventral hippocampi were snap-frozen in liquid nitrogen and stored at −80 °C until further processing.
Whole tissue RNA extraction
One snap-frozen ventral hippocampus per animal was used to obtain transcriptional profiles. Samples were processed in a multiple block design with every block containing one replicate of each condition and sex. Processing within blocks was randomized. Samples were homogenized in 500 µl Trizol (Invitrogen 15596018) using metal beads (Qiagen 69989) in a tissue-lyser bead mill (Qiagen, Germany) at 4 °C for 2 min at 20 Hz. RNA was extracted according to the manufacturer’s recommendations. RNA purity and quantity were determined with a UV/V spectrophotometer. RNA integrity was assessed with high sensitivity RNA screen tape on an Agilent Tapestation/Bioanalyzer, according to the manufacturer’s protocol.
Whole-tissue library preparation and sequencing
For experiment 1 (see Supplementary Fig. 1), library preparation and sequencing was performed at the Functional Genomics Center Zurich (FGCZ) of University of Zurich and ETH Zurich. The TruSeq stranded RNA kit (Illumina Inc.) was used according to the manufacturer’s protocol. For bulk sequencing library preparation, the TruSeq stranded RNA kit (Illumina Inc.) was used according to the manufacturer’s protocol. The mRNA was purified by polyA selection, chemically fragmented and transcribed into cDNA before adapter ligation. Single-end (100nt) sequencing was performed with HiSeq 4000. Samples within experiments were each run on one or multiple lanes and demultiplexed. A sequencing depth of ~20 M reads per sample was obtained.
For experiments 2 + 3 (see Supplementary Fig. 1), library preparation and sequencing was performed at Novogene, UK. Briefly, RNA samples were used for library preparation using NEBNext® Ultra RNA Library Prep Kit for Illumina®. Indices were included to multiplex multiple samples. Briefly, mRNA was purified from total RNA using poly-T oligo-attached magnetic beads. After fragmentation, the first strand cDNA was synthesized using random hexamer primers followed by the second strand cDNA synthesis. The library was ready after end repair, A-tailing, adapter ligation, and size selection. After amplification and purification, insert size of the library was validated on an Agilent 2100 and quantified using quantitative PCR (Q-PCR). Libraries were then sequenced on Illumina NovaSeq 6000 S4 flowcell with PE150 according to results from library quality control and expected data volume. Samples within experiments were each run on one or multiple lanes and demultiplexed. A sequencing depth of ~20 M paired reads per sample was obtained.
Bulk transcriptome analyses
Bulk RNA-seq was analyzed as described in ref. 9. Briefly, trimmed reads were used for quantification of the GENCODE M17 transcripts using kallisto 0.44, reads from protein-coding transcripts were summed to the gene level, and differential gene expression analysis was performed using edgeR 4.6.3. In addition to a FDR threshold of 0.05, an absolute logFC threshold of at least 0.15 was used to determine differentially-expressed genes (DEGs). Wherever surrogate variable (SV) analysis (using sva package version 3.50.067) identified more than 5 SVs, we restricted it to 5. To identify habituation patterns, sva-corrected, unit-variance-scaled logFCs of DEGs across samples from time points 45 min to 3 h were first extracted, and values ceiled between −5 to 5 to restrict outlier influences. Genes were then clustered using the Spectrum 1.1 package with k = 10.
The MDS was computed through the plotMDS function of the limma package version 3.58.1 using the sva-corrected expression of the acute stress DEGs across Handling samples, and the CRS samples were then projected onto this space. The MDS was recentered so that all time point medians were equidistant to the origin before calculating angles.
Cluster enrichment analysis was done using Fisher’s exact test (followed by FDR correction) on the KEGG, WIKIPATHWAYS and GO genesets of effective sizes ranging from 5 to 1000 from the MSigDB68, accessed from the msigdbr R package version 7.5.1. For the enrichment of motifs/ChIP peaks in early stress-responsive genes (Supplementary Fig. 9C–E), we first used liftover to translate the human brainTF peaks to mouse coordinates, and then overlapped the lifted neuronal peaks on the single-cell ATAC peaks. We then used Fisher’s exact test to test for enrichment of motifs or peaks in the broad promoter region (±10 kb around TSS) of stress-responsive genes, compared to expressed genes.
To make the data from the Cort profiling cohort (Fig. 3J) comparable to the other batches, we used the empirical Bayes batch effect correction (ComBat, as implemented in the sva package) on log(CPM) data, treating the Handling+3 h ARS samples of each batch as coming from the same group. We then computed response coefficients by regressing each sample’s logFCs of the genes significantly responding at 3 h onto the corresponding average logFCs in the original cohort using a robust linear model without intercept, and used the slope/coefficient to assess overall response.
Nuclei isolation for ATAC-Seq
Nuclei isolation was performed as previously described69. All steps were performed on ice. Briefly, one vHC was homogenized in lysis buffer (26 mM Sucrose, 30 mM KCl, 10 mM MgCl2, 20 mM Tricine-KOH (pH 7.8), 1 mM DTT, 0.5 mM Spermidine, 0.15 mM Spermine, 0.3% NP-40, Complete protease inhibitor in nuclease-free water) by 10 strokes with pestle A and 20 strokes with pestle B. The sample was filtered through a 70 μm Flowmi strainer and centrifuged for 5 min at 400 g at 4 °C. Supernatant was removed and additional lysis buffer was added to resuspend the pellet. The sample was then mixed with 50% Iodixanol solution (30 mM KCl, 26 mM Sucrose, 10 mM MgCl2, 20 mM Tricine-KOH (pH 7.8), 50% Iodixanol solution in nuclease-free water) and underlaid by 30% Iodixanol solution (50% Iodixanol solution in lysis buffer) and 40% Iodixanol (50% Iodixanol solution in lysis buffer) layers. The sample was centrifuged for 20 min at 3000 g at 4 °C with brakes off. The nuclei band was collected and mixed with ATAC buffer (10 mM NaCl, 10 mM Tris-HCl (pH7.5), 3 mM MgCl2, 0.1% Tween20 in nuclease-free water). The nuclei count was assessed and 50’000 nuclei were transferred to a new tube with ATAC buffer. The samples was centrifuged for 10 min at 500 g at 4 °C and supernatant was removed.
Preparation of ATAC libraries
Nuclei were resuspended in 50 μl of transposition mix (25 μl 2× TD buffer, 2.5 μl transposase (Diagenode, C01070012), 16.5 μl PBS, 0.5 μl 1% digitonin, 0.5 μl 10% Tween-20, and 5 μl nuclease-free water). Transposition reactions were incubated at 37 °C for 30 min while shaking at 1000 r.p.m. Tagmented DNA was purified using the MinElute Reaction Cleanup Kit (Qiagen, 28204). Purified DNA was amplified using the NEBNext high-Fidelity 2X master mix (New England Biolabs, Ipswich, Massachusetts, United States, M0541) and Diagenode primers (Diagenode, Marlborough, Massachusetts, United States, C01011036) at 72 °C for 5 min, 98 °C for 30 s and 5 cycles of 98 °C for 10 s, 63 °C for 30 s, and 72 °C for 1 min. A fraction of the amplification product was used to determine the number of additional cycles using qPCR, resulting in a total of 8–9 amplification cycles for all samples. A double-sided size selection (0.5X and 1.3X) was performed to remove large and very small library fragments. Insert size of the library was validated and quantified on an Agilent 2100 Bioanalyzer. Libraries were then sequenced on Illumina NovaSeq 6000 S4 flowcell with PE150 according to results from library quality control and expected data volume aiming for 50 Million read pairs per sample. Sequencing was performed at Novogene, UK.
Analysis of bulk ATAC sequencing
Reads were trimmed with trimmomatic and aligned to the GRCm38 genome using bowtie2 with --dovetail --no-mixed --no-discordant -I 15 -X 2000. Duplicates were marked using Picard. Peaks were called using Genrich 0.6.1 (https://github.com/jsh58/Genrich) with ATAC mode with a FDR threshold of 0.05, discarding PCR duplicates and excluding the mitochondria and ENCODE blacklisted regions. Quantification was performed by counting the number of insertions events (i.e., shifted fragment starts/ends) within peaks or promoter regions. Two samples (one from each of the two 45 min groups) were excluded because they had much lower coverage, lower sequencing quality, longer fragments, a different GC distribution from the other samples, and were very clear outliers on an MDS plot. Differential analysis was performed with edgeR glmFit using a model including Sex, Treatment, ARS time point, and a Treatment:TimePoint interaction. For promoter-based analysis, an additional surrogate variable based on the SVA package was included. Insertion coverage tracks were generated (and plotted) with the epiwraps package v0.99.102, using binWidth=10 L, shift=c(4 L,-5L), type = “ends”, extend=3 L, and includeDuplicates=FALSE.
Motif accessibility analysis was performed using chromVAR. For this purpose, peaks were resized to 300 bp and insertions were counted as described above. 2000 iterations were used, and the z-scores were quantile-normalized as recommended in ref. 70. Differential analysis was performed with limma using 1 surrogate variable.
Nuclei isolation for single-cell experiments
Nuclei isolation was performed similarly as for ATAC-Seq, with the addition of RNase inhibitors as previously described69. All steps were performed on ice. Briefly, one vHC was homogenized in lysis buffer (26 mM Sucrose, 30 mM KCl, 10 mM MgCl2, 20 mM Tricine-KOH (pH 7.8), 1 mM DTT, 0.5 mM Spermidine, 0.15 mM Spermine, 0.3% NP-40, Complete protease inhibitor, 0.2 U/μl RNAse inhibitor in nuclease-free water) by 10 strokes with pestle A and 20 strokes with pestle B. The sample was filtered through a 70 μm Flowmi strainer and centrifuged for 5 min at 400 g at 4 °C. Supernatant was removed and additional lysis buffer was added to resuspend the pellet. The sample was then mixed with 50% Iodixanol solution (30 mM KCl, 26 mM Sucrose, 10 mM MgCl2, 20 mM Tricine-KOH (pH 7.8), 50% Iodixanol solution in nuclease-free water) and underlaid by 30% Iodixanol solution (50% Iodixanol solution in lysis buffer) and 40% Iodixanol (50% Iodixanol solution in lysis buffer) layers. The sample was centrifuged for 20 min at 3000 g at 4 °C with brakes off. The nuclei band was collected and mixed with ATAC buffer (10 mM NaCl, 10 mMTris-HCl (pH7.5), 3 mM MgCl2, 0.1% Tween20, 0.2 U/μl RNase inhibitor in nuclease-free water) before the nuclei count was assessed. The sample was centrifuged for 5 min at 500 g at 4 °C and supernatant was removed. Diluted nuclei buffer (0.1 M DTT, 1 U/μl RNase inhibitor, 1X Nuclei buffer (10X) in nuclease-free water) was added to achieve the desired nuclei concentration.
Preparation of single-nucleus Multiome libraries
10X Multiome libraries were prepared following the Chromium Next GEM Single Cell Multiome ATAC + Gene Expression Reagent Kits User Guide (CG000338, Rev. F, 10X Genomics) with a targeted nuclei recovery of 10,000 nuclei (1 sample failed due to clogging). 7 cycles were performed for sample index PCR during ATAC library preparation. 6 cycles were used for cDNA amplification and a total of 13 cycles was performed for sample index PCR for gene expression library preparation. RNA integrity was assessed on an Agilent Tapestation, according to the manufacturer’s protocol. Samples of each library type were pooled. Both libraries (gene expressions and ATAC) were then sequenced on one Illumina X Plus 25B flow cell on individual lanes with an adjusted read configuration (150-10-24-150 bp). A sequencing depth of ~50,000 paired reads per nucleus was targeted. Library preparation and sequencing were performed at the Functional Genomics Center Zurich (FGCZ) of University of Zurich and ETH Zurich.
Single-cell analysis
Pre-processing and filtering
Pre-processing was performed using cellranger-arc 2.0.2 using the optimized mm10 transcriptome from ref. 71. A RNA quantification using alevin-fry72 was also performed to estimate splicing proportions. Decontamination was performed using cellbender 0.3.073 with custom initialization, and doublet identification with scDblFinder 1.19.074. For the RNA modality, scDblFinder was run using standard parameters twice (i.e., with two seeds) and the scores averaged, the AMULET method was used on the ATAC modality. Cells were excluded as doublets if they were called as such using both scDblFinder runs, or if their mean doublet score was above 0.2 and their amulet q-value below 0.01. Only cells with at least 500 fragments (after decontamination), less than 60% contamination, less than 1% mitochondrial reads, more than 20% unspliced fragments (before decontamination) and less than 50% of the fragments assigned to the top 10% features were used for further analysis.
Single-nucleus RNA-seq analysis
Cell annotation was performed based on the RNA modality. A first rough Seurat 5.0.1 clustering75 was performed to split neuronal and non-neuronal cells, which were then processed separately using Seurat.
For neurons, 3500 variable features were selected using the disp method. Prominent activity-dependent genes were removed to limit the influence of the activation status on cell annotation, and known markers were added (see repository for the exact list). Batch correction was performed using harmony 1.2.176. The first 30 dimensions of the harmony embedding were then used to identify the 50 shared nearest neighbors used for clustering at resolution 1.3 and UMAP projection. Cell clusters with a median proportion of unspliced reads below 50%, a median doublet score above 0.3, a median proportion of mitochondrial reads above 0.02, a median proportion of contamination above 0.3, or half of their markers being more highly expressed in empty droplets than in cells were removed as low-quality clusters. Clustering was then performed again on the cleaned object. The clusters were then compared to the Allen brain atlas annotation using SingleR77 for the purpose of annotation, in combination with known markers. In addition, we used the CA1/3 marker genes from Cembrowski et al.46 to distinguish ventral and more dorsal populations.
For non-neuronal cells, we first used geo-sketching78 to 6000 cells as implemented in sketchR 1.1.2 to increase sensitivity to rare cell types. We took the union of the top 2000 variable features identified using the disp and vst methods, as well as the top 1000 most expressed features in the sketched data and known markers. We then performed PCA on the entire dataset based on these features followed by harmony. The first 20 dimensions of the harmony embedding were then used for clustering and UMAP projection. Cell clusters with a median proportion of unspliced reads below 35% or a median doublet score above 0.2 were removed, as well as clusters of cycling cells based on the expression of Top2a, Ccnb2, Cdk1 and Ccnf. The remaining clusters were annotated manually based on known markers.
For ADT-based analysis (below) and visualization, only cells with at least 1000 fragments were used.
Pseudo-bulk differential analysis was performed using edgeR with a ~TimePoint*Treatment model. If the median abundance of the cluster across samples was higher than 50 cells, one surrogate variable estimated using the sva package was further included in the model. For each cell type, genes were filtered using filterByExpr with min.count=20, and only genes with an average log(CPM) above 1.5 were considered. We then harnessed our vast bulk RNA-seq dataset to improve power by estimated global FDR in the pseudobulk data with the IHW package v1.30.0 (10.1038/nmeth.3885), using the FDR from the bulk data as covariate (see in-depth demonstration in ref. 79). Specifically, we used the smallest FDR between the spliced and unspliced bulk data from dropping all experimental covariates (for baseline differences between CRS and Handling, the FDR from the baseline bulk comparison was used instead). Genes which were below filtering thresholds at the bulk level, and which consequently had no attached bulk FDR, were assigned a value of 2 to have their weights estimated separately. IHW was then applied independently for each contrast, across cell types of the same class (i.e., neuronal/non-neuronal). Finally, a minimum absolute logFC of 0.5 was used in addition to FDR to identify differentially-expressed genes.
ADT scoring
The Activity-Dependent Transcription (ADT) score was computed separately for broad cell classes (with neurons grouped as either excitatory or inhibitory). For each class, we first defined a set of stress-induced genes with global (i.e., across time points) FDR below 0.15 (0.05 for non-neurons), an absolute logFC above 0.2 and a log(CPM) above 1. Since the latest time point is chiefly due to GR, we additionally required the gene to have a FDR < 0.15 at either 15 min or 45 min. For each gene, we then computed the weighted average of its pseudobulk logFC at 15 min and 45 min, weighted by 1-sqrt(FDR) in the respective time point. These logFC estimates then served as the expected activity logFC, on which each cell was regressed. Specifically, we used Gamma-Poisson regression (using the glmGamPoi package 1.17.3 with global overdispersion and no normalization80 of the cell’s counts for those genes on two covariates: the scaled sum expression of the genes across all cells of that type, and the expected logFC. The coefficient assigned to the second covariate then represents a raw activity score, which is however strongly influenced by the proportion of cells of that type that are in an activated state. To address this issue, we then scale the score for each cell, setting 0 to the median score in baseline animals, and 1 to the 98th percentile of the score. An ADT score of 0.5 then represents the point where cells are more similar to the most strongly activated cells than to the median baseline cell. For neurons, an alternative activity scoring was performed using the neuroestimator package version 0.149, and to circumvent the biases of both methods, the two scores were averaged for each cell for the purpose of quantifying the proportions of cells activated.
Single-nucleus ATAC-seq analysis
Using the RNA-based clusters, fragments were merged for neuronal and non-neuronal cell types and peaks were called separately using MACS2 version 2.2.9.181 via the CallPeaks function of Signac 1.13.0 version82. Downstream analysis was performed on pseudobulks of the RNA-based clusters obtained using muscat version 1.16.083. For motif accessibility analysis, peaks were resized to 500 bp before running chromVAR on each cell type separately and performing differential analysis as discussed for bulk ATAC-seq. To avoid reporting the several sets of highly similar motifs showing a common pattern, we first applied hierarchical clustering of motifs using the Yule similarity of their matches across peaks, and cutting the dendrogram at a height of 0.5. For each cell type, we took the significant motifs from the same cluster and excluded the ones for which the corresponding gene had an expression below 2 logCPM in that cell type. We then reported the remaining ones in decreasing order of expression.
To assess differences in motif accessibility separately for activated and non-activated neurons (Fig. 8G), we split neurons based on an ADT threshold of 0.5 and compiled pseudobulk accessibility profiles. As activated neurons are fewer, we downsampled the counts of non-activated pseudobulk samples so that their median would match that of activated ones.
Open field test
Open-field testing took place inside sound insulated, ventilated multi-conditioning chambers (TSE Systems Ltd, Germany). The open field arena (45 cm × 45 cm × 40 cm [L × W × H]) consisted of four transparent plexiglas walls and a light gray PVC floor. Animals were tested for 10 min under dim lighting (4 lux). Video recordings were collected and distance, time in center, supported rears and unsupported rears were analyzed.
Pose estimation using DLC and behavior analyses
DeepLabCut 2.0.7 (DLC)84,85 was used to track 13 body points of each animal. Tracked points included nose, head center, neck, right ear, left ear, body center, body center left, body center right, left hip, right hip, tailbase, tail center and tail tip. The four corners of the OFT were tracked additionally to automatically detect the arena boundaries in each recording. X and Y coordinates of DLC tracking data were imported into R Studio (v3.6.1) and processed with the DLCAnalyzer package86. Furthermore, a previously trained supervised classifier86 was applied to quantify supported and unsupported rears in the OFT on a per-frame basis.
Data availability
The sequencing datasets generated in this study have been deposited to GEO under accession GSE292240. We additionally used previously-published datasets GSE190712 and GSE79790. Other data are provided as Supplementary Information/Source Data. Source data are provided with this paper.
Code availability
All code for analyzing sequencing experiments are available on Github https://github.com/ETHZ-INS/CRS_habituation, as well as on Zenodo https://doi.org/10.5281/zenodo.1746518287.
References
Floriou-Servou, A. et al. The acute stress response in the multiomic era. Biol. Psychiatry 89, 1116–1126 (2021).
Joëls, M. & Baram, T. Z. The neuro-symphony of stress. Nat. Rev. Neurosci. 10, 459–466 (2009).
Kitchener, P., Di Blasi, F., Borrelli, E. & Piazza, P. V. Differences between brain structures in nuclear translocation and DNA binding of the glucocorticoid receptor during stress and the circadian cycle. Eur. J. Neurosci. 19, 1837–1846 (2004).
Revest, J.-M. et al. The MAPK pathway and Egr-1 mediate stress-related behavioral effects of glucocorticoids. Nat. Neurosci. 8, 664–672 (2005).
Thoeringer, C. K. et al. The temporal dynamics of intrahippocampal corticosterone in response to stress-related stimuli with different emotional and physical load: an in vivo microdialysis study in C57BL/6 and DBA/2 inbred mice. Psychoneuroendocrinology 32, 746–757 (2007).
Droste, S. K. et al. Corticosterone levels in the brain show a distinct ultradian rhythm but a delayed response to forced swim stress. Endocrinology 149, 3244–3253 (2008).
Qian, X. et al. A rapid release of corticosteroid-binding globulin from the liver restrains the glucocorticoid hormone response to acute stress. Endocrinology 152, 3738–3748 (2011).
Reul, J. M. H. M. Making memories of stressful events: a journey along epigenetic, gene transcription, and signaling pathways. Front. Psychiatry 5, 5 (2014).
von Ziegler, L. M. et al. Multiomic profiling of the acute stress response in the mouse hippocampus. Nat. Commun. 13, 1824 (2022).
Revest, J.-M. et al. BDNF-TrkB signaling through Erk1/2MAPK phosphorylation mediates the enhancement of fear memory induced by glucocorticoids. Mol. Psychiatry 19, 1001–1009 (2014).
Gerstner, N. et al. DiffBrainNet: differential analyses add new insights into the response to glucocorticoids at the level of genes, networks and brain regions. Neurobiol. Stress 21, 100496 (2022).
Privitera, M. et al. Noradrenaline release from the locus coeruleus shapes stress-induced hippocampal gene expression. eLife 12, RP88559 (2024).
McEwen, B. S. & Akil, H. Revisiting the stress concept: implications for affective disorders. J. Neurosci. 40, 12–21 (2020).
Nestler, E. J. & Russo, S. J. Neurobiological basis of stress resilience. Neuron 112, 1911–1929 (2024).
Thompson, R. F. & Spencer, W. A. Habituation: a model phenomenon for the study of neuronal substrates of behavior. Psychol. Rev. 73, 16–43 (1966).
Grissom, N. & Bhatnagar, S. Habituation to repeated stress: get used to it. Neurobiol. Learn. Mem. 92, 215–224 (2009).
Armario, A., Castellanos, J. M. & Balasch, J. Adaptation of anterior pituitary hormones to chronic noise stress in male rats. Behav. Neural Biol. 41, 71–76 (1984).
Campmany, L., Pol, O. & Armario, A. The effects of two chronic intermittent stressors on brain monoamines. Pharmacol. Biochem. Behav. 53, 517–523 (1996).
Bhatnagar, S., Huber, R., Nowak, N. & Trotter, P. Lesions of the posterior paraventricular thalamus block habituation of hypothalamic-pituitary-adrenal responses to repeated restraint. J. Neuroendocrinol. 14, 403–410 (2002).
Babb, J. A., Masini, C. V., Day, H. E. W. & Campeau, S. Habituation of hypothalamic–pituitary–adrenocortical axis hormones to repeated homotypic stress and subsequent heterotypic stressor exposure in male and female rats. Stress Amst. Neth. 17, 224–234 (2014).
Melia, K., Ryabinin, A., Schroeder, R., Bloom, F. & Wilson, M. Induction and habituation of immediate early gene expression in rat brain by acute and repeated restraint stress. J. Neurosci. 14, 5929–5938 (1994).
Chen, X. & Herbert, J. Regional changes in c-fos expression in the basal forebrain and brainstem during adaptation to repeated stress: correlations with cardiovascular, hypothermic and endocrine responses. Neuroscience 64, 675–685 (1995).
Martinez, M., Phillips, P. J. & Herbert, J. Adaptation in patterns of c-fos expression in the brain associated with exposure to either single or repeated social stress in male rats. Eur. J. Neurosci. 10, 20–33 (1998).
Stamp, J. A. & Herbert, J. Multiple immediate-early gene expression during physiological and endocrine adaptation to repeated stress. Neuroscience 94, 1313–1322 (1999).
Girotti, M. et al. Habituation to repeated restraint stress is associated with lack of stress-induced c-fos expression in primary sensory processing areas of the rat brain. Neuroscience 138, 1067–1081 (2006).
Konarska, M., Stewart, R. E. & McCarty, R. Habituation and sensitization of plasma catecholamine responses to chronic intermittent stress: effects of stressor intensity. Physiol. Behav. 47, 647–652 (1990).
Ulrich-Lai, Y. M. et al. Chronic stress induces adrenal hyperplasia and hypertrophy in a subregion-specific manner. Am. J. Physiol.-Endocrinol. Metab. 291, E965–E973 (2006).
Anisman, H. Sensitization in relation to posttraumatic stress disorder. Biol. Psychiatry 70, 404–405 (2011).
Ursin, H. Brain sensitization to external and internal stimuli. Psychoneuroendocrinology 42, 134–145 (2014).
Belda, X., Fuentes, S., Daviu, N., Nadal, R. & Armario, A. Stress-induced sensitization: the hypothalamic–pituitary–adrenal axis and beyond. Stress 18, 269–279(2015).
Peña, C. J. et al. Early life stress alters transcriptomic patterning across reward circuitry in male and female mice. Nat. Commun. 10, 5098 (2019).
Bagot, R. C. et al. Ketamine and Imipramine reverse transcriptional signatures of susceptibility and induce resilience-specific gene expression profiles. Biol. Psychiatry 81, 285–295 (2017).
Bagot, R. C. et al. Circuit-wide transcriptional profiling reveals brain region-specific gene networks regulating depression susceptibility. Neuron 90, 969–983 (2016).
Nasca, C. et al. Role of the astroglial glutamate exchanger xCT in ventral hippocampus in resilience to stress. Neuron 96, 402–413.e5 (2017).
Brivio, E. et al. Sex shapes cell-type-specific transcriptional signatures of stress exposure in the mouse hypothalamus. Cell Rep. 42, 112874 (2023).
Cullinan, W. E., Herman, J. P., Battaglia, D. F., Akil, H. & Watson, S. J. Pattern and time course of immediate early gene expression in rat brain following acute stress. Neuroscience 64, 477–505 (1995).
Floriou-Servou, A. et al. Distinct proteomic, transcriptomic, and epigenetic stress responses in dorsal and ventral hippocampus. Biol. Psychiatry 84, 531–541 (2018).
McEwen, B. S. Stress and hippocampal plasticity. Annu. Rev. Neurosci. 22, 105–122 (1999).
Zannas, A. S., Wiechmann, T., Gassen, N. C. & Binder, E. B. Gene–Stress–epigenetic regulation of FKBP5: clinical and translational implications. Neuropsychopharmacology 41, 261–274 (2016).
Häusl, A. S. et al. The co-chaperone Fkbp5 shapes the acute stress response in the paraventricular nucleus of the hypothalamus of male mice. Mol. Psychiatry 26, 3060–3076 (2021).
Müller-Dott, S. et al. Expanding the coverage of regulons from high-confidence prior knowledge for accurate estimation of transcription factor activities. Nucleic Acids Res. 51, 10934–10949 (2023).
Chen, P. B. et al. Mapping gene expression in excitatory neurons during hippocampal late-phase long-term potentiation. Front. Mol. Neurosci. 10, 39 (2017).
Caradonna, S. G., Paul, M. R. & Marrocco, J. An allostatic epigenetic memory on chromatin footprints after double-hit acute stress. Neurobiol. Stress 20, 100475 (2022).
Waag, R. & Bohacek, J. Single-nucleus RNA-sequencing in brain tissue. Curr. Protoc. 3, e919 (2023).
Miranda, L., Bordes, J., Gasperoni, S. & Lopez, J. P. Increasing resolution in stress neurobiology: from single cells to complex group behaviors. Stress Amst. Neth. 26, 2186141 (2023).
Cembrowski, M. S., Wang, L., Sugino, K., Shields, B. C. & Spruston, N. Hipposeq: a comprehensive RNA-seq database of gene expression in hippocampal principal neurons. eLife 5, e14997 (2016).
Fan, Z. et al. The vascular gene Apold1 is dispensable for normal development but controls angiogenesis under pathological conditions. Angiogenesis 26, 385–407 (2023).
Delhaye, S. & Bardoni, B. Role of phosphodiesterases in the pathophysiology of neurodevelopmental disorders. Mol. Psychiatry 26, 4570–4582 (2021).
Bahl, E. et al. Using deep learning to quantify neuronal activation from single-cell and spatial transcriptomic data. Nat. Commun. 15, 779 (2024).
Sheng, M., Thompson, M. A. & Greenberg, M. E. CREB: a Ca2 + -regulated transcription factor phosphorylated by calmodulin-dependent kinases. Science 252, 1427–1430 (1991).
West, A. E., Griffith, E. C. & Greenberg, M. E. Regulation of transcription factors by neuronal activity. Nat. Rev. Neurosci. 3, 921–931 (2002).
Fernandez-Albert, J. et al. Immediate and deferred epigenomic signatures of in vivo neuronal activation in mouse hippocampus. Nat. Neurosci. 22, 1718–1730 (2019).
McEwen, B. S. The neurobiology of stress: from serendipity to clinical relevance1. Brain Res. 886, 172–189 (2000).
McEwen, B. S. Neurobiological and systemic effects of chronic stress. Chronic Stress 1, 2470547017692328 (2017).
Roszkowski, M. et al. Rapid stress-induced transcriptomic changes in the brain depend on beta-adrenergic signaling. Neuropharmacology 107, 329–338 (2016).
Bahrami, S. & Drabløs, F. Gene regulation in the immediate-early response process. Adv. Biol. Regul. 62, 37–49 (2016).
Makino, S., Smith, M. A. & Gold, P. W. Increased expression of corticotropin-releasing hormone and vasopressin messenger ribonucleic acid (mRNA) in the hypothalamic paraventricular nucleus during repeated stress: association with reduction in glucocorticoid receptor mRNA levels. Endocrinology 136, 3299–3309 (1995).
Umemoto, S., Kawai, Y., Ueyama, T. & Senba, E. Chronic glucocorticoid administration as well as repeated stress affects the subsequent acute immobilization stress-induced expression of immediate early genes but not that of NGFI-A. Neuroscience 80, 763–773 (1997).
Kitay, J. I. Sex differences in adrenal cortical secretion in the rat. Endocrinology 68, 818–824 (1961).
Galea, L. A. et al. Sex differences in dendritic atrophy of CA3 pyramidal neurons in response to chronic restraint stress. Neuroscience 81, 689–697 (1997).
Bohacek, J., Manuella, F., Roszkowski, M. & Mansuy, I. M. Hippocampal gene expression induced by cold swim stress depends on sex and handling. Psychoneuroendocrinology 52, 1–12 (2015).
Conway-Campbell, B. L. et al. The HSP90 molecular chaperone cycle regulates cyclical transcriptional dynamics of the glucocorticoid receptor and its coregulatory molecules CBP/p300 during ultradian ligand treatment. Mol. Endocrinol. Baltim. Md 25, 944–954 (2011).
Carrigan, A. et al. An active nuclear retention signal in the glucocorticoid receptor functions as a strong inducer of transcriptional activation. J. Biol. Chem. 282, 10963–10971 (2007).
George, A. A., Schiltz, R. L. & Hager, G. L. Dynamic access of the glucocorticoid receptor to response elements in chromatin. Int. J. Biochem. Cell Biol. 41, 214–224 (2009).
De Boer, S. F., Koopmans, S. J., Slangen, J. L. & Van Der Gugten, J. Plasma catecholamine, corticosterone and glucose responses to repeated stress in rats: effect of interstressor interval length. Physiol. Behav. 47, 1117–1124 (1990).
Bakken, T. E. et al. Single-nucleus and single-cell transcriptomes compared in matched cortical cell types. PloS ONE 13, e0209648 (2018).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Castanza, A. S. et al. Extending support for mouse data in the Molecular Signatures Database (MSigDB). Nat. Methods 20, 1619–1620 (2023).
Grandi, F. C., Modi, H., Kampman, L. & Corces, M. R. Chromatin accessibility profiling by ATAC-seq. Nat. Protoc. 17, 1518–1552 (2022).
Gerbaldo, F. E. et al. On the identification of differentially-active transcription factors from ATAC-seq data. PLOS Comput. Biol. 20, e1011971 (2024).
Pool, A.-H., Poldsam, H., Chen, S., Thomson, M. & Oka, Y. Recovery of missing single-cell RNA-sequencing data with optimized transcriptomic references. Nat. Methods 20, 1506–1515 (2023).
He, D. et al. Alevin-fry unlocks rapid, accurate and memory-frugal quantification of single-cell RNA-seq data. Nat. Methods 19, 316–322 (2022).
Fleming, S. J. et al. Unsupervised removal of systematic background noise from droplet-based single-cell experiments using CellBender. Nat. Methods 20, 1323–1335 (2023).
Germain, P.-L., Lun, A., Meixide, C. G., Macnair, W. & Robinson, M. D. Doublet identification in single-cell sequencing data using scDblFinder. F1000Research 10, 979 (2022).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Hie, B., Cho, H., DeMeo, B., Bryson, B. & Berger, B. Geometric sketching compactly summarizes the single-cell transcriptomic landscape. Cell Syst 8, 483–493.e7 (2019).
Germain, P.-L., Wang, J. & Robinson, M. D. Bulk-based hypothesis weighing increases power in single-cell differential expression analysis. Peer Community J. 5, e138.
Ahlmann-Eltze, C. & Huber, W. glmGamPoi: fitting Gamma-Poisson generalized linear models on single cell count data. Bioinformatics 36, 5701–5702 (2021).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol 9, R137 (2008).
Stuart, T., Srivastava, A., Madad, S., Lareau, C. A. & Satija, R. Single-cell chromatin state analysis with Signac. Nat. Methods 18, 1333–1341 (2021).
Crowell, H. L. et al. muscat detects subpopulation-specific state transitions from multi-sample multi-condition single-cell transcriptomics data. Nat. Commun. 11, 6077 (2020).
Nath, T. et al. Using DeepLabCut for 3D markerless pose estimation across species and behaviors. Nat. Protoc. 14, 2152–2176 (2019).
Lauer, J. et al. Multi-animal pose estimation, identification and tracking with DeepLabCut. Nat. Methods 19, 496–504 (2022).
Sturman, O. et al. Deep learning-based behavioral analysis reaches human accuracy and is capable of outperforming commercial solutions. Neuropsychopharmacology 45, 1942–1952 (2020).
Germain, P.-L. & Sonder, E. ETHZ-INS/CRS_habituation: Accepted paper. Zenodo https://doi.org/10.5281/zenodo.17465182 (2025).
Acknowledgements
The lab of J.B. is supported by the ETH Zurich, ETH Project Grant ETH-20 19-1, SNSF Grant 310030_204372, the Basel Research Centre for Child Health (BRCCH), the Swiss 3 R Competence Center, the Hochschulmedizin Zürich Flagship project STRESS, and ERA-NET NEURON - PROGRESS (SNSF: 31NE30_219119). E.S. was supported by the ETH Project Grant ETH-25 20-2 to P.-L.G. We thank Julia Bode and Anjana Madhavan for maintaining the animal colony, and Fabienne Rössler for help with data analysis. We thank the 3 R Hub at the ETH Zurich for their support with behavior testing, and the staff of the EPIC for animal care and their service to our animal facility.
Author information
Authors and Affiliations
Contributions
R.W. conceived experiments, conducted experiments and molecular screens, data analysis, graphs & figures, interpreted results, wrote manuscript; L.v.Z. performed data analysis (transcriptomics); E.S. performed data analysis (single-cell ATAC); O.S. helped with data analysis and behavior setups; J.L. provided practical help with experiments; S.F. performed corticosterone ELISA; R.L. performed corticosterone ELISA; K.G. provided supervision; P.L.G. conducted data analysis (transcriptomics, ATAC, single-cell transcriptomics), conceived experiments, generated graphs & figures, interpreted results, provided funding, wrote manuscript; J.B. conceived experiments, interpreted results, provided resources and funding, wrote manuscript. All authors have read the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Catherine Peña who co-reviewed with Rixing Lin; and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Waag, R., von Ziegler, L., Sonder, E. et al. Distinct mechanisms of transcriptomic habituation to repeated stress in the mouse hippocampus. Nat Commun 16, 11569 (2025). https://doi.org/10.1038/s41467-025-67825-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-67825-y