Abstract
The development and progression of cardiometabolic diseases and depression multimorbidity involves pathophysiological processes across multiple organs. Using multi-organ imaging data from 31,246 UK Biobank participants, we investigate the multi-organ manifestations and their phenotypic connections and shared genetic architecture underlying the multimorbidity. Phenotypic analyses identify seven abdominal, 16 cardiac, and 107 brain traits forming 1418 abdomen-heart-brain cliques, with liver volume, myocardial wall thickness, and white matter hyperintensity volume as central nodes. Genetic analyses reveal 43 distinct genomic loci (21 novel) shared by these cliques, with the most widely shared loci mapped to genes NUDC, ARID1A, and CRHR1. The 224 protein-coding genes mapped by these loci are enriched in 39 biological processes related to cardiometabolic and neuropsychiatric functions, with 15 genes expressed across liver-heart-brain axis tissues. Combining biochemical and multi-organ imaging indicators significantly improves multimorbidity prediction. These findings uncover multi-organ network underlying physical-mental multimorbidity and highlight the necessity of holistic management.
Similar content being viewed by others
Introduction
The prevalence of chronic disease multimorbidity has been steadily increasing with population aging, imposing substantial health and economic burdens worldwide1,2,3. Cardiometabolic diseases (CMDs) and depression represent one of the most common multimorbidity patterns, with the depression prevalence up to nearly 21% in people with cardiovascular disease (CVD)4 and 28% in those with type 2 diabetes (T2D)5, which are substantially higher than the 4% prevalence observed in the general population6. This relationship is bidirectional, as individuals with depression face 2–3 fold increased risk of developing CMDs compared to the general population7,8. Due to shared risk factors and overlapping pathophysiological mechanisms, including autonomic nervous system imbalance, hypothalamic-pituitary-adrenal (HPA) axis dysregulation, and chronic inflammation, CMDs-depression multimorbidity leads to increased mortality, higher healthcare costs, and diminished quality of life compared to single chronic conditions8,9,10,11.
Both CMDs and depression involve pathophysiological processes spanning multiple organ systems, including the liver, pancreas, heart, and brain12,13,14,15. The multimorbidity of these conditions exhibits significantly heterogeneous correlations with changes across organ systems compared to either condition alone7. This multi-system nature underscores the need to switch from an isolated single-organ perspective to an integrated approach that captures cross-organ interactions. Given that magnetic resonance imaging (MRI)-derived phenotypes have been widely validated in assessing structures and functions of multiple organs16,17,18, leveraging multi-organ MRI traits to comprehensively understand the organ-specific manifestations of CMDs-depression multimorbidity can provide valuable insights into the biological underpinnings of this complex condition.
Growing evidence has revealed the interconnectedness of multi-organ systems in human health, exemplified by connections between heart-brain19, eye-brain20, and heart-brain-liver21 axes that reflect shared genetic architecture and biological pathways identified through genome-wide association studies (GWAS). However, most existing multi-organ crosstalk has focused on general connections between organ traits in the overall population rather than examining disease-specific relationships that could provide direct clinical utility for pathophysiology understanding, disease prediction, and therapeutic targeting. While extensive single-disease studies have characterized imaging and genetic correlates of CVD22,23,24, T2D25,26, and depression27,28 independently—revealing organ-specific alterations in cardiac structure and function, metabolic tissues, and brain networks—this approach cannot capture the complex systemic alterations that characterize multimorbid states, where pathological processes may interact synergistically across organ systems. Elucidating the connections among organ traits associated with CMDs-depression multimorbidity and their shared genetic underpinnings holds promise for advancing holistic approaches to multimorbidity risk management and identifying potential therapeutic targets for intervention.
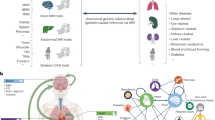
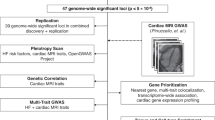
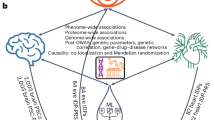
This study comprehensively investigated the multi-organ manifestations of CMDs-depression multimorbidity through integrated phenotypic, genetic, and predictive analyses across the abdomen-heart-brain axis. The specific hypotheses are formulated as follows: (1) Multi-organ MRI traits across abdominal, cardiac, and brain regions are significantly associated with CMDs-depression multimorbidity, reflecting the systemic nature beyond single-organ alterations; (2) Multimorbidity-associated imaging traits exhibit extensive cross-organ phenotypic connections, forming integrated abdomen-heart-brain networks that indicate interconnected pathophysiology underlying physical-mental comorbidity; (3) Cross-organ MRI traits share common genetic architecture, with pleiotropic variants influencing multiple organ systems; and (4) Prediction models incorporating multi-organ MRI traits significantly improve multimorbidity prediction beyond traditional risk factors (Fig. 1).
(2025) https://BioRender.com/q51m660). A Overview of the study design. B Description of specific abdominal, cardiac, and brain MRI traits. C Key analyses involved in each step. Abbreviations: CMDs cardiometabolic diseases, CPASSOC cross-phenotypic association analysis, DTI diffusion tensor imaging, GPA genetic analysis incorporating pleiotropy and annotation, LAVA local analysis of variant association, LDSC linkage disequilibrium score regression, NODDI neurite orientation dispersion and density imaging, TWAS transcriptome-wide association studies.
Results
Study sample characteristics
This study included three separate datasets for analyzing associations between MRI traits and CMDs-depression multimorbidity: an abdominal set (N = 25,344), a heart set (N = 27,705), and a brain set (N = 31,246), each containing the respective MRI scans (Fig. S1). The three-region set of 14,122 participants who had undergone all three types of MRI scans was used for estimating cross-organ pairwise correlations (Fig. S1). The demographic characteristics were similar across all four study samples. Specifically, in the three-region set, participants had a mean (SD) age of 62.9 (7.2) years, with 6363 (45.1%) being male, 13,063 (92.5%) being White, and 61 (0.4%) having prevalent CMDs-depression multimorbidity (Table 1).
Further demographic stratification by disease status revealed that the CMDs-only, depression-only, multimorbidity, and control groups each showed comparable baseline characteristics (age, sex distribution, ethnicity, and socioeconomic status [SES]) across the abdominal, cardiac, brain, and three-region datasets (Supplementary Data 1).
CMDs-depression multimorbidity was significantly enriched in all four datasets, with ERs ranging from 2.3 to 2.6 (all ERs > 2.0, all p < 0.001), showing non-random synergistic CMDs-depression co-occurrence (Supplementary Table 1).
Associations between imaging traits and multimorbidity
Seven abdominal, 16 cardiac, and 107 brain traits showed significant associations with CMDs-depression multimorbidity (Fig. 2, Supplementary Data 2). Among abdominal MRI traits, subcutaneous fat volume, visceral fat volume, liver proton density fat fraction (PDFF), pancreas PDFF, and liver volume were positively, while liver iron and pancreas volume were negatively associated with multimorbidity (Fig. 2A). The cardiac assessment revealed that left ventricle (LV) myocardial wall thickness, circumferential strain, and longitudinal strain were positively associated with multimorbidity, while stroke volume (including left atrium [LA], LV, and right ventricle [RV]) and LV radial strain showed negative associations (Fig. 2B). Among brain gray matter traits, ventricular volumes (including the third and inferior lateral ventricles) were positively associated with multimorbidity, while global gray matter volume, most regional cortical and subcortical volumes (including the caudal anterior cingulate, superior temporal, and superior frontal regions) showed negative associations (Fig. 2C, D). Among brain white matter traits, larger white matter hyperintensity volume was positively, while better white matter microstructural integrity, characterized by higher fractional anisotropy (FA), isotropic volume fraction (ISOVF), and mode of anisotropy (MO) values, and lower mean diffusivity (MD), intracellular volume fraction (ICVF), and orientation dispersion (OD) values, were negatively associated with multimorbidity. Specifically, these associations were observed in major white matter pathways (corpus callosum, corona radiata, and cerebral peduncle) and limbic tracts (cingulum cingulate gyrus, uncinate fasciculus, cingulum hippocampus, and fornix cres + stria terminalis) (Fig. 2E–J).
A cardiac B, regional cortical C and subcortical gray matter volumes D, and white matter microstructural properties including fractional anisotropy (E), mean diffusivity (F), mode of anisotropy (G), isotropic volume fraction (H), intracellular volume fraction (I), and orientation dispersion (J). Visualization methods differ by organ system due to anatomical mappability constraints. Abdominal and cardiac anatomical diagrams provide organ anatomy without trait-specific mapping, with trait associations represented through wordmap font sizes scaled to statistical significance (-log10 p values). Brain images display statistical significance through both anatomical color mapping (-log10 p values) and wordmap font sizes. Regional cortical volumes (Desikan-Killiany atlas) and subcortical gray matter volumes (aseg atlas) are mapped onto brain surfaces, while white matter microstructural properties (JHU ICBM-DTI-81 atlas) are rendered in Montreal Neurological Institute-152 space. The associations were estimated based on logistic regressions adjusted for age, sex, ethnicity, Townsend deprivation index, education level, body mass index, smoking status, drinking status, physical activity, and imaging site. Whole body surface and de-meaned scanner table coordinates were additionally adjusted in the brain analysis and heart analysis, respectively. Two-sided t-tests were used for all associations, and false discovery rate adjustment was applied for multiple comparisons. Complete results of all association analyses are presented in Supplementary Data 2. FA fractional anisotropy, ICVF intracellular volume fraction, ISOVF isotropic volume fraction, L left hemisphere, LA left atrium, LV left ventricle, MD mean diffusivity, MO mode of anisotropy, OD orientation dispersion, PDFF proton density fat fraction, R right hemisphere, RA right atrium, RV right ventricle.
Sensitivity analyses showed that significant associations remained between all 7 abdominal and 16 cardiac traits and CMDs-depression multimorbidity after additional adjustment for imaging parameters, while 92 brain traits remained significantly associated with multimorbidity (Supplementary Data 3). In addition, the associations remained significant for most organ traits in subgroup analyses. For CVD-depression multimorbidity, 5 abdominal, 10 cardiac, and 86 brain traits overlapped with CMDs-depression findings, while for T2D-depression multimorbidity, 7 abdominal, 14 cardiac, and 58 brain traits overlapped with the CMDs-depression associations (Supplementary Data 4–5).
Cross-organ phenotypic pairwise correlations between imaging traits
Multimorbidity-associated imaging traits showed widespread inter-organ correlations across the abdomen, heart, and brain regions. Among 2573 significant cross-organ imaging traits pairs, we identified 1418 interrelated abdomen-heart-brain cliques, comprising six abdominal traits, 16 cardiac traits, and 86 brain traits (Supplementary Data 6–9). Within this complex network, the most connected nodes for each region were liver volume, LV mean myocardial wall thickness of American Heart Association (AHA) segment 6, and total volume of white matter hyperintensities, respectively (Fig. 3A, B). The network exhibited high sparsity (0.84), low modularity (0.15), and moderate centrality measures (0.54 for closeness and 17.07 for degree) (Supplementary Table 2).
A Network of interconnected abdominal, cardiac, and brain traits underlying cardiometabolic diseases and depression multimorbidity. Node colors represent organ regions (green: abdomen, purple: heart, blue: brain) and node sizes indicate connectivity degrees. Two key functional subnetworks are highlighted: the Metabolic Load-Cardiac Remodeling-Brain Structural Abnormality Module (Module 1) and the Ectopic Lipid-Cardiac Dysfunction-White Matter Pathology Module (Module 2). Complete results of all network associations are presented in Supplementary Data 9. B Number of cardiac and brain traits significantly associated with abdominal traits. C Heatmap of standardized beta coefficients from linear regressions between abdominal and cardiac traits. D Heatmap of standardized beta coefficients from linear regressions of abdominal and cardiac traits with brain traits. Cross-organ associations were assessed using linear regressions adjusted for age, sex, ethnicity, Townsend deprivation index, education level, body mass index, smoking status, drinking status, physical activity, imaging site, and history of cardiovascular disease, type 2 diabetes, hypertension, hyperlipidemia, and depression. De-meaned scanner table coordinates and whole body surface were additionally adjusted in the brain analysis and heart analysis, respectively. Two-sided t-tests were used for all associations, and false discovery rate adjustment were applied for multiple comparisons. The abdominal, cardiac, and brain traits are labeled with abbreviations, and their full names are provided in Supplementary Data 31. Abbreviations: CC corpus callosum, Ctx cortex, IC internal capsule, ST Vol volume of supratentorial, SV stroke volume.
Cross-organ connections within abdomen-heart-brain cliques revealed two distinct functional subnetworks with specific directional correlation patterns (Fig. 3A, C, D and Supplementary Data 9). The Metabolic Load-Cardiac Remodeling-Brain Structural Abnormality Module comprised three components: metabolic load indicated by increased liver volume and visceral fat volume, cardiac remodeling represented by increased myocardial wall thickness, and brain structural abnormalities involving reduced gray matter volumes with compromised white matter integrity across major tracts (corpus callosum, corona radiata, internal capsule, and limbic pathways). Within this module, greater metabolic load correlated with more severe cardiac remodeling and more pronounced brain structural abnormalities, while more severe cardiac remodeling correlated with more pronounced brain structural abnormalities. The Ectopic Lipid-Cardiac Dysfunction-White Matter Pathology Module encompassed ectopic lipid accumulation in the liver and pancreas, cardiac dysfunction reflected by reduced stroke volume, and white matter pathology manifested as increased hyperintensity volume and compromised corpus callosum integrity. This module showed positive correlations between ectopic lipid accumulation and both cardiac dysfunction and white matter pathology, as well as positive correlations between cardiac dysfunction and white matter pathology.
In sensitivity analyses, 1234 abdomen-heart-brain cliques were retained following additional adjustment for imaging parameters, with central nodes remaining unchanged within each anatomical region (Supplementary Data 10). Network stability was further validated through the application of increasingly stringent false discovery rate (FDR) thresholds for clique definition, as evidenced by the consistent identification of liver volume as the central node, preservation of two core functional subnetworks, and maintenance of stable topological parameters across threshold levels, including high sparsity (0.82–0.84), low modularity (0.14–0.17), and moderate centrality measures (0.53–0.54 for closeness and 12.45–17.07 for degree) (Fig. S2, Supplementary Table 2, Supplementary Data 11–13).
Subgroup analyses revealed 609 and 707 overlapping abdomen-heart-brain cliques in CVD-depression and T2D-depression multimorbidity, respectively. The most connected nodes for each region remained consistent across different disease outcomes, with the exception that the ascending aorta minimum area emerged as the most central cardiac node in T2D-depression multimorbidity. The two central functional modules with specific directional correlation patterns observed in the main analysis were also evident in these subgroups (Supplementary Data 14, 15).
Genetic correlations and genetic overlap of abdomen-heart-brain cliques
Extensive genetic correlations and genetic overlap were observed across 776 trait pairs from abdomen-heart-brain cliques, of which 14 were identified from cross-trait linkage disequilibrium score regression (LDSC), 147 from local analysis of variant association (LAVA), and 776 from genetic analysis incorporating pleiotropy and annotation (GPA) (Supplementary Data 16). These trait pairs included 51 from the abdomen-heart axis, 161 from the abdomen-brain axis, and 564 from the heart-brain axis, involving six abdominal traits (liver traits, pancreas traits, and fat distribution), 16 cardiac traits (myocardial wall thickness, myocardial longitudinal and radial strain, and chamber volumes), and 83 brain traits (global and regional gray matter volumes, white matter hyperintensity volume, and white matter microstructural properties). The magnitude and direction of genetic relationships varied considerably across organ systems. Abdomen-heart pairs showed the strongest genetic correlations (ranging from −0.38 to 0.23) and the highest proportions of shared genetic variants (0.29% to5.27%). Heart-brain pairs exhibited moderate genetic correlations (−0.20 to0.20) with moderate shared genetic architecture (0.40% to3.46%), while abdomen-brain pairs showed more modest but consistently negative genetic correlations (−0.11 to −0.06) with lower proportions of shared genetic variants (0.26% to2.41%).
Shared genetic loci across abdomen-heart-brain cliques
A total of 78 unique genomic loci were identified as being shared by 216 abdomen-heart-brain cliques, encompassing six abdominal traits (liver traits, pancreas traits, and fat distribution), 14 cardiac traits (myocardial wall thickness, myocardial longitudinal strain, and chamber volumes), and 62 brain traits (global and regional gray matter volumes, white matter hyperintensity volume, and white matter microstructural properties) (Supplementary Data 17). The lead single-nucleotide polymorphisms (SNPs) across the genomic loci exhibited heterogeneous directional effects, with 33.2% displaying concordant associations and 66.8% showing discordant associations across all three traits within their respective cliques (Fig. S3). Sensitivity analyses revealed strong genetic concordance, with 58 genomic loci (shared by 198 cliques) overlapping with primary results after additional adjustment for imaging parameters (Supplementary Data 18). Subgroup analyses showed substantial consistency, with 49 loci (shared by 138 cliques) in CVD-depression multimorbidity and 56 loci (shared by 148 cliques) in T2D-depression multimorbidity overlapping with those identified in the main analysis (Supplementary Data 19–20).
Most candidate SNPs from the independent shared genomic loci were located in intronic (46.9%) and intergenic (41.1%) regions, with five SNPs in exonic regions, including two non-synonymous mutations (Fig. 4A, B). Of all candidate SNPs, 5.0% had combined annotation-dependent depletion (CADD) scores larger than 12.37, indicating pathogenicity, 7.4% had RegulomeDB scores less than 3, showing high regulatory potential, and 44.8% had chromatin state less than 8, suggesting presence in transcriptionally active regions (Supplementary Data 21).
A Manhattan plot of candidate SNPs identified by cross-phenotypic association analysis from each abdomen-heart-brain clique. Statistical significance was assessed using the two-sided SHet test. A genome-wide significance threshold of p < 5 × 10−8 was applied. Each point represents a SNP and the horizontal dashed line indicates the genome-wide significance threshold (5 × 10−8). For SNPs with multiple p values, only the smallest value is displayed. B Distribution of functional consequences of candidate SNPs in the genomic loci shared by each abdomen-heart-brain clique. C Ideogram of 43 distinct shared genomic loci identified by cross-phenotypic association analysis from each abdomen-heart-brain clique. Cytogenetic bands are labeled for each locus, with the numbers of cliques sharing each locus indicated in parentheses. The circles, rectangles, and triangles represent traits from the abdominal, cardiac, and brain categories, respectively. The colors within each shape indicate specific traits within each organ category. D The bubble plot shows the enriched GO terms of biological processes for protein-coding genes mapped by all candidate SNPs. Enrichment was tested using Fisher’s exact test, and p values were adjusted for multiple comparisons using the false discovery rate method. The colors of the dots indicate the magnitude of the -log10-transformed values of false discovery rate-adjusted p values, and the size of the dots represents the number of genes enriched in a given term. Abbreviations: BP biological process, CC cell component.
Integration of overlapping genomic positions yielded 43 distinct loci, including 21 novel loci not previously associated with abdominal, cardiac, or brain traits used in this study (Fig. 4C, Supplementary Data 22). Among these, 28 loci exhibited pleiotropic effects across multiple abdomen-heart-brain cliques. The three most widely shared loci included chr1:27262545-27262545 (lead SNP: rs182050989, mapped gene: NUDC) appearing in 33 cliques, chr1:27021913-27021913 (lead SNP: rs114165349, mapped gene: ARID1A) in 27 cliques, and chr17: 43546853-43805194 (lead SNP: rs1635298, mapped gene: CRHR1) in 25 cliques. These cliques were combinations of traits, including pancreas and liver volume, myocardial wall thickness, white matter hyperintensity volume, and white matter microstructural properties in regions such as the corpus callosum, corona radiata, and posterior thalamic radiation.
Furthermore, gene mapping of all candidate SNPs within genomic loci identified 224 protein-coding genes, which were enriched in 39 Gene Ontology (GO) biological processes (Fig. 4D, Supplementary Data 23–24). These processes primarily involved lipid metabolism and transport, mitochondrial functions, and membrane organization—biological pathways with relevance to both cardiometabolic and neuropsychiatric functions.
Shared genes across tissues
Transcriptome-wide association studies (TWAS) validated that 15 unique genes expressed in 24 tissues were shared by 16 abdomen-heart-brain cliques, spanning adipose tissue, whole blood, blood vessel, brain, heart, liver, and pancreas (Fig. 5, Supplementary Data 25). These cliques comprised liver volume, myocardial wall thickness, and global gray matter and white matter microstructural properties in different tracts, including corpus callosum, posterior thalamic radiation, and corona radiata. For instance, ARID1A, WNT3, and HSPA4 expressed in adipose tissue, heart, and brain, functioned as shared genes underlying the significant liver-heart-brain connections. CENPW, PIGV, and GPN2 expressed in the brain, heart, and blood vessels connected traits across the liver-heart-brain axis. SYTL1, TNFSF12, and NUDC, expressed in heart, brain, blood, and liver tissues, served as shared genetic links between liver, heart, and brain. RSPO3, MPDU1, and NMBR expressed in the brain and adipose tissue demonstrated connections of the same three organs.
From inner to outer layers, the plot represents significant genes, the tissues they are expressed in, and the cliques that share these genes. The width of each ring segment is proportional to the number of organ-related tissues and the number of abdomen-heart-brain cliques. Statistical details of transcriptome-wide association studies are presented in Supplementary Data 25. The abdominal, cardiac, and brain traits are labeled with abbreviations, and their full names are provided in Supplementary Data 31.
All 15 genes expressed in 24 organ-related tissues retained significance in sensitivity analyses after additional adjustment for imaging parameters (Supplementary Data 26). Of the 15 genes identified, 14 were also observed in the CVD-depression subgroup (excluding REEP3) and in the T2D-depression subgroup (excluding MPDU1) across the same 24 organ-related tissues (Supplementary Data 27, 28).
Prediction modeling
Following the procedures illustrated in Fig. S4, seven nested prediction models were constructed using Cox models for multimorbidity development and progression. The addition of multi-organ imaging traits sharing genetic architecture across the abdomen-heart-brain axis improved prediction accuracy for CMDs-depression multimorbidity incidence among individuals without both conditions compared to base models. Consistent improvements were observed for Royston’s D, integrated discrimination improvement (IDI), and net reclassification improvement (NRI). These performance improvements were comparable to those achieved by incorporating traditional SES factors, lifestyle factors, or biochemical factors, showing no significant C-statistic differences (Fig. 6A, B, Supplementary Data 29, 30).
A Model performance evaluated by Harrell’s C statistic across three scenarios, including the multimorbidity risk in individuals without both conditions, depression risk in cardiometabolic diseases patients, and cardiometabolic diseases risk in depression patients. Data are presented as estimated Harrell’s C statistic and 95% confidence intervals in forest plots. B Performance gains of models incorporating MRI traits quantified by ΔC statistic across the above three scenarios (n = 57,284 for multimorbidity risk, n = 6650 for depression risk in CMDs, n = 1344 for CMDs risk in depression). Data are presented as estimated ΔC statistic and 95% confidence intervals in forest plots. C Heatmap of standardized beta coefficients of imaging traits across the above three scenarios in MRI scores. Three composite scores were constructed using 10-fold cross-validated elastic net Cox regression: lifestyle score (body mass index, smoking status, alcohol consumption, and physical activity), biomedical score (HbA1c, total cholesterol, triglycerides, high-density lipoprotein cholesterol, low-density lipoprotein cholesterol, systolic blood pressure, and C-reactive protein), and MRI score (imaging traits from genetically-informed abdomen-heart-brain cliques). Cox models were used to construct seven prediction models including base model (age, sex, and ethnicity), socioeconomic status model (age, sex, ethnicity, Townsend deprivation index, and education), lifestyle model (socioeconomic status model adding the lifestyle score), biochemical model (socioeconomic status model adding the biochemical score), MRI model (socioeconomic status model adding the MRI score), Lifestyle-MRI model (socioeconomic status model adding the lifestyle score and the MRI score), and Biochemical-MRI model (socioeconomic status model adding the biochemical score and the MRI score). The abdominal, cardiac, and brain traits are labeled with abbreviations, and their full names are provided in Supplementary Data 31. Abbreviations: CI confidence interval, CMDs cardiometabolic diseases, SES socioeconomic status.
The combination of genetically-informed imaging traits and biochemical factors significantly enhanced multimorbidity risk prediction performance. The Biochemical-MRI prediction model outperformed the Biochemical model (ΔC-statistic: 0.05 [95% confidence intervals (CIs), 0.01–0.10]), with the C-statistic increasing from 0.70 (95% CI, 0.61–0.78) to 0.74 (95% CI, 0.67–0.81). Similar improvements were observed for predicting depression incidence in individuals with pre-existing CMDs (ΔC-statistic: 0.06 [95% CI, 0.02–0.13]), where the integration of biochemical factors and MRI traits achieved a C-statistic of 0.77 (95% CI, 0.66–0.88), superior to 0.72 (95% CI, 0.60–0.83) for the MRI model alone. Royston’s D statistic consistently reached peak performance when combing MRI and biochemical indicators across both development and progression outcomes (Fig. 6A, B, Supplementary Data 29, 30).
Across all imaging traits in all MRI-enhanced prediction model, liver volume, LV mean myocardial wall thickness, and global gray matter volume consistently emerged as the strongest predictors for both multimorbidity development and progression. Additionally, ICVF values of the retrolenticular part of the internal capsule tract showed high-ranking coefficients specifically for predicting multimorbidity risk in individuals with single pre-existing conditions (Fig. 6C).
Discussion
Our study provided novel insights into the phenotypic and genetic architecture of the multi-organ network in CMDs-depression multimorbidity using MRI-derived traits. We identified significant associations of multiple organ-specific traits (seven abdominal, 16 cardiac, and 107 brain traits) with multimorbidity and revealed extensive phenotypic connections along the abdomen-heart-brain axis. The shared genetic architecture of these cross-organ connections was evidenced by genetic correlations and overlap, common genomic loci and genes, and biological pathways. Moreover, the integration of these imaging traits and biochemical factors significantly improved the prediction of multimorbidity development and progression.
This study characterized the multi-organ networks along the abdomen-heart-brain axis underlying CMDs-depression multimorbidity, revealing the coordinated alterations of metabolic, cardiovascular, and neural dysfunction. In the abdomen, increased liver volume and ectopic fat accumulation in the liver and pancreas were strongly correlated with multimorbidity. Liver volume—a parameter reflecting hepatic fat accumulation but understudied in previous studies—emerged as a novel marker for multimorbidity with potential diagnostic utility. Cardiovascular assessment identified increased myocardial wall thickness and reduced stroke volume, and brain imaging revealed structural alterations concentrated in limbic networks and major white matter pathways (corpus callosum, corona radiata, and posterior thalamic radiation). The concurrent presence of cardiac alterations known from cardiovascular disease research29,30 and brain changes characteristic of depression31,32 underscores the simultaneous involvement of cardiovascular and neural substrates underlying multimorbidity pathophysiology. Of particular significance are the limbic system structural changes, which likely represent a critical neurobiological interface linking mood regulation and cardiovascular function, given its central role in emotional regulation and autonomic control of the heart-brain axis33,34. Importantly, these organ-specific alterations formed a densely interconnected network comprising 1418 cliques, with the correlation patterns crystallizing into two functional modules: the Metabolic Load-Cardiac Remodeling-Brain Structural Abnormality Module and the Ectopic Lipid-Cardiac Dysfunction-White Matter Pathology Module. Within each module, the systematic intercorrelation of metabolic dysfunction, cardiovascular impairment, and brain structural abnormalities indicates that CMDs-depression multimorbidity manifests as a synchronized system-wide disorder with coordinated perturbations rather than isolated organ dysfunction.
The coordinated metabolic-cardiac-neural alterations observed in our multi-organ network likely reflect two hypothetical pathophysiological processes underlying multimorbidity. On one hand, primary diseases may trigger multi-organ changes through behavior-mediated cascade effects and increase secondary disease risk. Depression as a primary disease may induce ectopic fat accumulation through behavioral changes (reduced physical activity, sleep disturbance, and poor dietary quality) while simultaneously affecting cardiovascular function through chronic stress pathways, ultimately increasing both diabetes and cardiovascular disease risk7,35,36,37,38. Conversely, CMDs may affect brain structural abnormalities through chronic disease-related psychological stress, promoting depression development7,35. On the other hand, shared physiological regulatory mechanisms may simultaneously act on multiple organ systems and multimorbidity development. Autonomic nervous system dysregulation33,34,39,40,41, chronic inflammatory7,42, and HPA axis hyperactivity39,43 states during multimorbidity may concurrently contribute to the observed lipid metabolism disorders, cardiac hypertrophy and dysfunction, and brain structural abnormalities. The multi-organ network could provide a biological window into the multimorbidity development and progression, underscoring the inadequacy of single-organ approaches to this complex condition and highlighting the need for holistic approaches in multimorbidity management.
Sequential genetic analyses uncovered the shared genetic architecture of multi-organ traits related to CMDs-depression multimorbidity, providing a biological foundation for their interconnected nature. While LDSC analysis showed limited significant genetic correlations, LAVA identified moderate regional associations, and GPA revealed substantial genetic overlap among abdomen-heart-brain cliques. This hierarchical pattern aligns with different methodological assumptions: LDSC captures genome-wide consistent allelic effects, LAVA detects regional pleiotropic signals, and GPA identifies individual variant pleiotropy. These findings suggest that cross-organ genetic relationships may be primarily driven by scattered pleiotropic variants with mixed effect directions, indicating that both spatially dispersed and agonistic-antagonistic genetic mechanisms operate simultaneously in the pathophysiology of multi-organ involvement in multimorbidity. Cross-phenotypic association analysis (CPASSOC) further detected 43 distinct shared loci across 216 abdomen-heart-brain cliques, revealing specific genetic determinants that link traits along the abdomen-heart-brain axis. Among these, the most widely shared loci mapped to genes with established multi-organ functions, particularly ARID1A and CRHR1. Specifically, ARID1A, a chromatin remodeling factor regulating transcriptional programs, exerts pleiotropic effects along the liver-heart-brain axis through modulating hepatic lipid metabolism44, cardiomyocyte proliferation affecting myocardial wall thickness45, and neurodevelopment with particular impact on corpus callosum integrity46,47. CRHR1 encodes the corticotropin-releasing hormone receptor, central to HPA axis regulation, a key neuroendocrine pathway underlying CMDs-depression multimorbidity39. HPA axis dysregulation may result in elevated cortisol levels, which can simultaneously affect the pancreas-heart-brain axis via insulin resistance induction48, cardiomyocyte hypertrophy promotion49,50, and white matter integrity compromise through disrupting myelin plasticity and axonal function51,52. Notably, we identified 21 novel loci associated with multiple organ traits, including five mapped to TMEM106B, ROBO1, PTCH1, and DENND1A—genes previously implicated in both depression53,54 and CMDs such as T2D26,55,56,57, coronary artery disease58, and stroke59. These novel loci establish a genetic bridge between CMDs-depression multimorbidity and specific organ alterations.
The 224 protein-coding genes mapped by 378 candidate SNPs within genomic loci were enriched in 39 GO biological processes, primarily involving lipid metabolism and transport processes related to liver, pancreas, and fat distribution, mitochondrial and regenerative processes crucial for myocardial contraction and remodeling, and membrane organization processes essential for white matter integrity. TWAS analysis further validated 15 unique genes identified by CPASSOC in organ-related tissues from the abdomen-heart-brain cliques. Two particularly noteworthy examples include WNT3 and RSPO3, which serve as a ligand and a signal enhancer of the WNT/β-catenin pathway —a signaling cascade implicated in both CMDs60 and depression61. This pathway also coordinates lipid metabolic processes in liver62, cardiomyocyte proliferation in heart63,64, and myelination and axon guidance in white matter tracts65,66. Another two key genes, TNFSF12 and HSPA4, members of TNF and HSP70 family, affect organ alterations through inflammatory responses67 and oxidative stress pathways68 across the liver-heart-brain axis, respectively. These findings provide valuable insights into shared genetic loci, gene expression patterns, and biological pathways linking organ-specific alterations to psychiatric and cardiometabolic conditions. Based on these molecular discoveries, targeted therapeutic interventions, including HPA axis modulators, lipid-targeting agents, mitochondrial enhancers, WNT pathway regulators, and anti-inflammatory treatments, represent potentially promising strategies for both multimorbidity and overall organ health management that warrant further investigations.
Our analysis also showcased the robust prediction power of the combination of biochemical factors and imaging traits with shared genetic architecture among the abdomen-heart-brain axis for multimorbidity risk and depression incidence in individuals with pre-existing CMDs. These findings underscore that traditional biochemical markers with genetically-informed multi-organ imaging provide enhanced risk stratification capabilities, supporting a precision medicine approach for early intervention in physical-mental multimorbidity. Within these developed prediction models, the identification of liver volume, cardiac wall thickness, and global gray matter volume as the strongest individual predictors suggests that prioritizing monitoring of these three markers among numerous available imaging parameters may enhance multimorbidity prevention and inform targeted interventions addressing metabolic dysregulation, cardiac dysfunction, and brain structural abnormalities. The retrolenticular internal capsule ICVF specifically emerged as a key predictor for multimorbidity progression, indicating that white matter integrity in this neural pathway may serve as a targeted biomarker for monitoring disease advancement in individuals with CMDs or depression.
This study has several limitations. First, the predominantly European ancestry of our cohort may limit the generalizability of these results to a more diverse population. Second, our analyses relied on the ongoing UK Biobank brain imaging study, which currently includes only a limited proportion of participants and captures a limited number of multimorbidity cases, although this is the largest MR imaging study ever. Additional data collection is expected to reveal more multimorbidity-related traits and multi-organ connections. Third, the cross-sectional nature of our analysis makes it challenging to establish causal relationships and temporal dynamics for organ-multimorbidity and organ-organ connections, which could be addressed by future longitudinal imaging studies. Fourth, the well-documented “healthy volunteer” selection bias of the UK Biobank study may limit its representativeness of the general European population. Fifth, the use of genetic data from the same cohort for both trait identification and genetic correlation analyses may introduce confounding from shared ancestry, population stratification, and regional socioeconomic factors. Sixth, employing the same cohort for both model training and evaluation may lead to over-optimistic performance estimates, limiting the generalizability of the prediction results. Seventh, causal inference could not be established in this observational study. Finally, misinformation associated with self-reported data on socioeconomic characteristics and lifestyle factors cannot be avoided.
In conclusion, this study delineated both organ-disease associations and phenotypic inter-organ connections in CMDs-depression multimorbidity. Shared genetic architecture among these organ-specific traits further provides evidence for their interconnected nature. The integration of multi-organ imaging traits and biochemical factors significantly improved the prediction of multimorbidity development and progression. These findings advance our understanding of physical-mental multimorbidity mechanisms within the multi-organ network, suggesting the importance of holistic and multi-system approaches for multimorbidity management.
Methods
Study design and participants
This study is embedded in the UK Biobank, a large-scale prospective cohort involving 502,536 participants aged 37–73 years from 22 assessment centers between 2006 and 201069. The data utilized in this analysis consisted of sociodemographic and behavioral assessments, medical conditions, genotype data, and multimodal neuroimaging. All participants provided electronically signed informed consent. Ethical approval for the UK Biobank was obtained from the North West Multi-center Research Ethics Committee. This research was conducted using the UK Biobank Resource under application number 101169, utilizing data provided by patients and collected by the National Health Service as part of their care and support.
Outcomes
The outcome of this study was prevalent multimorbidity of CMDs and depression, defined as the co-occurrence of at least one CMDs and depression. The diagnosis of diseases and deaths was ascertained through first occurrence data from hospital inpatient records and national death registries. CMDs comprised CVD and T2D. CVD was defined using the International Classification of Diseases, 10th revision (ICD-10) for coronary artery disease (I20-I25), stroke (I60-I64), and heart failure (I50), while T2D was identified with E11. Depression was defined using F32-F33. Mortality data were sourced from the NHS Information Centre (England and Wales) and the NHS Central Register (Scotland), with follow-up until October 30, 2023. To enhance the contrast between individuals with and without multimorbidity, participants who were diagnosed with CMDs only or depression only before their MRI assessment date were excluded from the analysis.
Abdominal MRI traits
All abdominal scans were performed using a 1.5T scanner (MAGENTOM Aera, Syngo MR D13, Siemens Healthineers, Erlangen, Germany). Four distinct groups of sequences, including the Dixon protocol with six separate series (neck-to-knees), a T1-weighted 3D acquisition sequence of the pancreas volume, a single-slice multi-echo acquisition sequence for liver fat and iron, and a single-slice multi-echo acquisition sequence for pancreas fat and iron, were utilized to derive abdominal traits. All imaging-derived phenotypes (IDPs) were obtained directly from the UK Biobank data portal as previously preprocessed and segmented measures70 (Supplementary Methods S2). Specifically, three liver traits (PDFF, volume, and iron), two pancreatic traits (PDFF and volume), visceral fat volume, and subcutaneous fat volume were used in the study (Supplementary Data 31).
Heart MRI traits
Cardiovascular magnetic resonance (CMR) images were acquired through clinical wide bore 1.5-Tesla scanners (MAGENTOM Aera, Syngo MR D13, Siemens Healthineers, Erlangen, Germany). Short-axis, long-axis, and aortic cine images were acquired to characterize cardiac and aortic structures and functions, with all cardiac IDPs sourced from the UK Biobank data portal following standardized processing and quality control procedures71 (Supplementary Methods S2). Following the procedures above, 82 CMR traits, consisting of 64 LV traits, four LA traits, four RV traits, four right atrium (RA) traits, three ascending aorta traits, and three descending aorta traits were included (Supplementary Data 31).
Brain MRI traits
Multimodal neuroimaging was collected using a standard 3 T scanner (Magnetom Skyra, Siemens Healthineers, Erlangen, Germany), with a standard Siemens 32-channel RF receive head coil17,72. T1-weighted structural images, T2-weighted fluid-attenuated inversion recovery (FLAIR) structural images, and diffusion-tensor images were applied. All brain IDPs were downloaded from the UK Biobank data portal that were derived based on established preprocessing and automated segmentation workflows72 (Supplementary Methods S2). Specifically, T1 images were processed using FreeSurfer to extract five global volumes of different tissue types and 107 brain volumes of cortical and subcortical areas predefined by the Desikan-Killiany atlas and the aseg atlas. T2 images were processed with FMRIB Software Library (FSL) to estimate white matter hyperintensity volume. All volumetric traits, including cortical volumes, subcortical volumes, and white matter hyperintensity volume, were normalized for head size by multiplying raw volumes by T1-based head size scaling factors calculated during transformation from native to standard space73. Diffusion-weighted images underwent preprocessing using FSL and were subsequently fed into diffusion-tensor-imaging fitting tool (DTIFIT) and neurite orientation dispersion and density imaging (NODDI) modeling tool Accelerated Microstructure Imaging via Convex Optimization. Based on the Johns Hopkins University ICBM-DTI-81 atlas, 144 tract-averaged DTI parameters, including FA, MD, and MO, and 144 tract-averaged NODDI parameters, including ISOVF, ICVF, and OD, were derived (Supplementary Data 31).
GWAS summary statistics for MRI traits
GWAS summary-level data for imaging-derived traits of the abdomen (N = 32,860), heart (N = 31,875), and brain (N = 33,224) were obtained among participants of European ancestry in the UK Biobank19,70,74. Each dataset was imputed using the 1000 Genomes reference75. SNPs with minor allele frequency > 1% were utilized.
Statistical analysis
ER analysis of CMDs-depression co-occurrence
Non-random CMDs-depression co-occurrence was assessed using permutation testing76. A binary matrix was constructed with participants as rows and diseases as columns. Disease columns were randomly shuffled 10,000 times while marginal disease frequencies were maintained to generate null distributions. ER was defined as observed versus expected multimorbidity rates under the null model, where ER > 1 indicates synergistic co-occurrence. p values were derived by comparing observed multimorbidity proportions against the null distribution, and confidence intervals were estimated via bootstrapping applied to the binary matrix.
Associations between imaging traits and multimorbidity
Logistic regressions were applied to estimate associations of imaging traits from the abdomen, heart, and brain with the CMDs-depression multimorbidity in three separate samples. Adjustments were made for age, sex, ethnicity, Townsend deprivation index (TDI), education level, body mass index (BMI), smoking status, drinking status, physical activity, and imaging site. De-meaned scanner table coordinates (X, Y, Z, and table position) and whole body surface were additionally adjusted in the brain analysis and heart analysis, respectively77. Due to low data missingness levels, missing covariate data were imputed using multiple imputation by chained equations (Supplementary Table 3). The detailed definitions of covariates are listed in Supplementary Table 4. Results were presented as odds ratios (ORs) with corresponding 95% CIs. Imaging traits with Benjamini-Hochberg FDR-adjusted p < 0.05 were selected for cross-organ pairwise correlation analysis.
Cross-organ pairwise correlations between imaging traits
Linear regressions were employed to quantify cross-organ pairwise correlations among the multimorbidity-associated imaging traits from abdomen, heart, and brain in the overall sample, adjusting for the aforementioned covariates and history of CVD, T2D, hypertension, hyperlipidemia, and depression (Supplementary Methods S1). All imaging traits were standardized prior to analysis, and results were presented as standardized beta coefficients with 95% CIs. Standardized beta coefficients represent the change in one organ trait per standard deviation change in another organ trait. p values were adjusted for multiple testing using FDR correction across all pairwise correlations. Abdomen-heart-brain cliques were defined as combinations of one abdominal trait, one cardiac trait, and one brain trait with full pairwise significance (FDR-adjusted p < 0.05), where each trait pair within the combination showed significant standardized beta coefficients from linear regressions. These cliques were subsequently selected for genetic analysis. The centrality of this network formed by these cliques was indicated by the degree of each node.
Genetic correlations of abdomen-heart-brain cliques
Genetic correlation and overlap of abdomen-heart-brain cliques were estimated using three complementary approaches based on GWAS summary statistics: (1) LDSC78,79 for global genetic correlation, (2) LAVA80 for local genetic correlation, and (3) GPA81 for genetic overlap. For LDSC analysis, precomputed LD scores were derived from approximately 1.2 million Hapmap3 SNPs in the 1000 Genomes Project of European ancestry with the MHC region (chromosome 6: 28,477,797-33,448,354) excluded82. For LAVA analysis, bivariate local genetic correlation analyses were performed for trait pairs with nominally significant univariate local genetic signals as the detection of reliable local genetic correlations requires adequate genetic variation within genomic regions. For GPA analysis, independent SNPs (r2 < 0.2 within a 250 kb window) were obtained by LD pruning based on 1000 Genomes Phase 3 European-ancestry genotypes to reduce the influence of LD, with genetic overlap between traits quantified by the proportion of genetic variants associated with both traits. Trait pairs were considered significant if they showed FDR-adjusted significant associations in either LDSC, LAVA, or GPA analyses.
Cross-trait meta-analysis of abdomen-heart-brain cliques
To determine the shared genetic variants of each abdomen-heart-brain clique, CPASSOC was performed to conduct cross-trait meta-analysis at the SNP level83. The test statistic SHet, an extension of SHom that allows for heterogeneous effects across traits and offers improved statistical power, was calculated to combine the GWAS summary statistics through a sample size weighted meta-analysis. To compute SHet, a correlation matrix of independent SNPs (r2 < 0.2 within a 250 kb window) without significant associations (−1.96 < Z < 1.96) is required, to account for trait correlations and sample overlap or relatedness across cohorts84. Significant shared genetic variants were defined as SNPs achieving genome-wide significance in the abdomen-heart-brain clique (pCPASSOC < 5 × 10−8) and suggestive significance in single trait GWAS data (psingle trait < 1 × 10−3).
Genomic locus definition
The independent genomic loci of the significant SNPs of each abdomen-heart-brain clique identified from CPASSOC were defined using functional mapping and annotation (FUMA)85. Briefly, independent significant SNPs were defined as those that achieved genome-wide significance and were independent of other SNPs (LD r2 < 0.6). Lead SNPs were selected from independent significant SNPs which were independent of each other (LD r2 < 0.1). Candidate SNPs were defined as all GWAS-tagged SNPs that were in LD (r2 ≥ 0.6) with the independent significant SNPs, establishing the boundaries of each locus. A single genomic locus was formed when the distance between two loci was less than 250 kb. Moreover, independent genomic loci across all abdomen-heart-brain cliques were merged into distinct loci when the genetic positions of two or more loci overlapped.
For all distinct loci, a locus was considered novel if (1) it was located more than 500 kb from any previously reported loci in original GWAS19,70,74, and (2) none of its candidate SNPs had been reported in the NHGRI-EBI GWAS Catalog86. For reported loci without specific position information, a 1000 kb distance from the lead SNP was used to define the locus border.
Functional annotation
Functional annotation and gene mapping of candidate SNPs from each abdomen-heart-brain clique were conducted using FUMA30. CADD scores, which predict the deleteriousness of SNPs on protein structure or function87, RegulomeDB scores, which evaluate the regulatory functionality88, and 15-core chromatin states, which estimate the transcriptional and regulatory effects at the SNP locus, were annotated for the associated SNPs89. Additionally, candidate SNPs were mapped to protein-coding genes using any of the three approaches: positional mapping, eQTL mapping, and 3D chromatin interaction mapping. To further uncover the biological pathways of the shared genes, pathway enrichment analysis was performed based on the GO database using the R package clusterProfiler. Pathways with FDR-corrected p < 0.05 were considered significant.
TWASs
TWASs were performed using S-PrediXcan to validate whether the expression levels of genes identified by CPASSOC of each abdomen-heart-brain clique showed significant associations with corresponding traits in relevant tissues90. Pre-computed eQTL prediction models of 24 tissues across seven categories (adipose tissue, whole blood, blood vessel, brain, heart, liver, and pancreas) were selected from GTEx v8. Significant TWAS genes were identified after the FDR correction and were then cross-referenced across traits to determine shared genes for each clique.
Prediction modeling
Separate prediction models were developed using Cox proportional hazard analysis to evaluate: (1) the risk of developing CMDs-depression multimorbidity in individuals free of both conditions (development); and (2) the risk of developing CMDs-depression multimorbidity in those with one pre-existing condition (two possible progressions) (Fig. S4). Three composite scores were constructed using 10-fold cross-validated elastic net Cox regression: lifestyle score (BMI, smoking status, alcohol consumption, and physical activity), biomedical score (HbA1c, total cholesterol, triglycerides, high-density lipoprotein cholesterol, low-density lipoprotein cholesterol, systolic blood pressure, and C-reactive protein)91, and MRI score (imaging traits from genetically-informed abdomen-heart-brain cliques identified through CPASSOC and validated by TWAS).
We constructed seven nested prediction models with progressively expanded sets of variables to assess the incremental predictive value of different risk factor domains. The seven models were base model (age, sex, and ethnicity), SES model (age, sex, ethnicity, TDI, and education), lifestyle model (SES model adding the lifestyle score), biochemical model (SES model adding the biochemical score), MRI model (SES model adding the MRI score), Lifestyle-MRI model (SES model adding the lifestyle score and the MRI score), and Biochemical-MRI model (SES model adding the biochemical score and the MRI score). Model performance was assessed using Harrell’s C statistic, while performance gains were quantified by the ΔC statistic with 95% CI derived from 200 bootstrap samples. We also employed Royston’s D, NRI, IDI, and Brier score for comprehensive performance evaluation.
Subgroup analyses
To evaluate both the generalizability of our findings and examine consistency and specificity between subgroups, we conducted separate analyses for CVD-depression and T2D-depression multimorbidities as key components within the broader CMDs spectrum.
Sensitivity analyses
To assess the robustness of our findings, two sensitivity analyses were conducted: (1) additional imaging parameters were incorporated as covariates in all regression models, including mean absolute head motion for diffusion imaging metrics (DTI and NODDI parameters), inverted signal-to-noise ratio for T1-derived volumetric measures, and acquisition dates across all imaging modalities (abdomen, heart, and brain); (2) stricter FDR thresholds (0.01, 0.005, and 0.001) were applied to define abdomen-heart-brain cliques, and variations in clique size, composition, and network statistics (sparsity, modularity, and centrality) were assessed.
General association analyses were performed using R version 4.2.0. LDSC was conducted using the “ldsc” software (v1.0.1, https://github.com/bulik/ldsc). LAVA was performed using “LAVA” tool (v0.1.0, https://github.com/josefin-werme/LAVA). GPA was conducted using the “GPA” R package (v1.1.0,https://github.com/dongjunchung/GPA). CPASSOC was performed using the FunctionSet.R file provided by the original authors. TWAS was conducted using the “S-PrediXcan” tool (https://github.com/hakyimlab/MetaXcan).
Statistics and reproducibility
This research was conducted as an observational cohort study using data obtained from the UK Biobank. No statistical method was used to predetermine sample size. Sample inclusion and exclusion criteria are detailed in Fig. S1 and the “Outcomes” section. Participants who were diagnosed with CMDs only or depression only before their MRI assessment date were excluded from the analysis. The study design did not involve any intervention, and therefore, traditional experimental procedures such as randomization and blinding are not applicable. The investigators were not blinded to allocation during experiments and outcome assessment as this was an observational study analyzing pre-existing data.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The UK Biobank data used in this study (phenotypic, neuroimaging, and genotype) are available under restricted access for approved research purposes. Access can be obtained by submitting a research application through the UK Biobank Access Management System at www.ukbiobank.ac.uk. This research was conducted under UK Biobank Application Number 101169. The raw UK Biobank participant-level data are protected and are not available due to data privacy laws and the terms of our data access agreement. GWAS summary statistics for the abdominal, cardiac, and brain imaging traits based on UK biobank participants are available at https://ftp.ebi.ac.uk/pub/databases/gwas/summary_statistics/GCST90016001-GCST90017000/, http://heartkp.org/, and https://open.win.ox.ac.uk//ukbiobank//big40/, respectively. Source data are provided with this paper.
Code availability
The code used for all analyses in this study is publicly available on GitHub at https://github.com/cecilia-wjx/MRI_network and has been archived on Zenodo with DOI92: https://doi.org/10.5281/zenodo.17669873. Figures created with BioRender are in compliance with BioRender’s Academic License Terms and are intended for publication purposes.
References
Chowdhury, S. R., Chandra Das, D., Sunna, T. C., Beyene, J. & Hossain, A. Global and regional prevalence of multimorbidity in the adult population in community settings: a systematic review and meta-analysis. EClinicalMedicine 57, 101860 (2023).
Skou, S. T. et al. Multimorbidity. Nat. Rev. Dis. Prim. 8, 1–22 (2022).
McPhail, S. M. Multimorbidity in chronic disease: impact on health care resources and costs. Risk Manag. Health. Policy 9, 143–156 (2016).
Whooley, M. A. & Wong, J. M. Depression and cardiovascular disorders. Annu Rev. Clin. Psychol. 9, 327–354 (2013).
Khaledi, M., Haghighatdoost, F., Feizi, A. & Aminorroaya, A. The prevalence of comorbid depression in patients with type 2 diabetes: an updated systematic review and meta-analysis on huge number of observational studies. Acta Diabetol. 56, 631–650 (2019).
Depressive disorder (depression). https://www.who.int/news-room/fact-sheets/detail/depression.
Nemeroff, C. B. & Goldschmidt-Clermont, P. J. Heartache and heartbreak—the link between depression and cardiovascular disease. Nat. Rev. Cardiol. 9, 526–539 (2012).
Tabák, A. G., Akbaraly, T. N., Batty, G. D. & Kivimäki, M. Depression and type 2 diabetes: a causal association?. Lancet Diab. Endocrinol. 2, 236–245 (2014).
Roth, G. A. et al. Global burden of cardiovascular diseases and risk factors, 1990–2019. J. Am. Coll. Cardiol. 76, 2982–3021 (2020).
GBD 2021 Diabetes Collaborators. Global, regional, and national burden of diabetes from 1990 to 2021, with projections of prevalence to 2050: a systematic analysis for the Global Burden of Disease Study 2021. Lancet 402, 203–234 (2023).
Correll, C. U. et al. Prevalence, incidence and mortality from cardiovascular disease in patients with pooled and specific severe mental illness: a large-scale meta-analysis of 3,211,768 patients and 113,383,368 controls. World Psychiatry 16, 163–180 (2017).
Tian, Y. E. et al. Heterogeneous aging across multiple organ systems and prediction of chronic disease and mortality. Nat. Med. 29, 1221–1231 (2023).
Priest, C. & Tontonoz, P. Inter-organ cross-talk in metabolic syndrome. Nat. Metab. 1, 1177–1188 (2019).
Tian, Y. E. et al. Evaluation of brain-body health in individuals with common neuropsychiatric disorders. JAMA Psychiatry 80, 567–576 (2023).
Xourafa, G., Korbmacher, M. & Roden, M. Inter-organ crosstalk during development and progression of type 2 diabetes mellitus. Nat. Rev. Endocrinol. 20, 27–49 (2024).
Littlejohns, T. J., Sudlow, C., Allen, N. E. & Collins, R. UK Biobank: opportunities for cardiovascular research. Eur. Heart J. 40, 1158–1166 (2019).
Miller, K. L. et al. Multimodal population brain imaging in the UK Biobank prospective epidemiological study. Nat. Neurosci. 19, 1523–1536 (2016).
Littlejohns, T. J. et al. The UK Biobank imaging enhancement of 100,000 participants: rationale, data collection, management and future directions. Nat. Commun. 11, 2624 (2020).
Zhao, B. et al. Heart-brain connections: phenotypic and genetic insights from magnetic resonance images. Science 380, abn6598 (2023).
Zhao, B. et al. Eye-brain connections revealed by multimodal retinal and brain imaging genetics. Nat. Commun. 15, 6064 (2024).
McCracken, C. et al. Multi-organ imaging demonstrates the heart-brain-liver axis in UK Biobank participants. Nat. Commun. 13, 7839 (2022).
Morales, M. A., Manning, W. J. & Nezafat, R. Present and future innovations in AI and cardiac MRI. Radiology 310, e231269 (2024).
Nelson, C. P. et al. Association analyses based on false discovery rate implicate new loci for coronary artery disease. Nat. Genet. 49, 1385–1391 (2017).
Malik, R. et al. Multiancestry genome-wide association study of 520,000 subjects identifies 32 loci associated with stroke and stroke subtypes. Nat. Genet. 50, 524–537 (2018).
Geijselaers, S. L. C., Sep, S. J. S., Stehouwer, C. D. A. & Biessels, G. J. Glucose regulation, cognition, and brain MRI in type 2 diabetes: a systematic review. Lancet Diab. Endocrinol. 3, 75–89 (2015).
Vujkovic, M. et al. Discovery of 318 new risk loci for type 2 diabetes and related vascular outcomes among 1.4 million participants in a multi-ancestry meta-analysis. Nat. Genet. 52, 680–691 (2020).
Zhang, F.-F., Peng, W., Sweeney, J. A., Jia, Z.-Y. & Gong, Q.-Y. Brain structure alterations in depression: psychoradiological evidence. CNS Neurosci. Ther. 24, 994–1003 (2018).
Howard, D. M. et al. Genome-wide meta-analysis of depression identifies 102 independent variants and highlights the importance of the prefrontal brain regions. Nat. Neurosci. 22, 343–352 (2019).
Gupta, S. et al. Left ventricular hypertrophy, aortic wall thickness, and lifetime predicted risk of cardiovascular disease. Dallas Heart Study JACC Cardiovasc. Imaging 3, 605–613 (2010).
De Marco, M. et al. Influence of left ventricular stroke volume on incident heart failure in a population with preserved ejection fraction (from the Strong Heart Study). Am. J. Cardiol. 119, 1047–1052 (2017).
Wu, G.-R. & Baeken, C. Normative modeling analysis reveals corpus callosum volume changes in early and mid-to-late first episode major depression. J. Affect Disord. 340, 10–16 (2023).
Coloigner, J. et al. White matter abnormalities in depression: a categorical and phenotypic diffusion MRI study. Neuroimage Clin. 22, 101710 (2019).
Grippo, A. J. & Johnson, A. K. Biological mechanisms in the relationship between depression and heart disease. Neurosci. Biobehav. Rev. 26, 941–962 (2002).
Rossi, A. et al. Heart–brain interactions in cardiac and brain diseases: why sex matters. Eur. Heart J. 43, 3971–3980 (2022).
Moulton, C. D., Pickup, J. C. & Ismail, K. The link between depression and diabetes: the search for shared mechanisms. Lancet Diab. Endocrinol. 3, 461–471 (2015).
Oppert, J.-M., Bellicha, A. & Ciangura, C. Physical activity in management of persons with obesity. Eur. J. Intern. Med. 93, 8–12 (2021).
Chaput, J.-P. et al. The role of insufficient sleep and circadian misalignment in obesity. Nat. Rev. Endocrinol. 19, 82–97 (2023).
Chao, A. M., Quigley, K. M. & Wadden, T. A. Dietary interventions for obesity: clinical and mechanistic findings. J. Clin. Investig. 131, e140065–e140065 (2021).
Joynt, K. E., Whellan, D. J. & O’Connor, C. M. Depression and cardiovascular disease: mechanisms of interaction. Biol. Psychiatry 54, 248–261 (2003).
Rapacciuolo, A. et al. Important role of endogenous norepinephrine and epinephrine in the development of in vivo pressure-overload cardiac hypertrophy. J. Am. Coll. Cardiol. 38, 876–882 (2001).
Floras, J. S. & Ponikowski, P. The sympathetic/parasympathetic imbalance in heart failure with reduced ejection fraction. Eur. Heart J. 36, 1974–1982 (2015).
Liu, L., Mei, M., Yang, S. & Li, Q. Roles of chronic low-grade inflammation in the development of ectopic fat deposition. Mediat. Inflamm. 2014, 418185 (2014).
Baumgartner, C., Krššák, M., Vila, G., Krebs, M. & Wolf, P. Ectopic lipid metabolism in anterior pituitary dysfunction. Front. Endocrinol. 14, 1075776 (2023).
Qu, Y.-L. et al. Arid1a regulates insulin sensitivity and lipid metabolism. EBioMedicine 42, 481–493 (2019).
Boogerd, C. J. et al. Cardiomyocyte proliferation is suppressed by ARID1A-mediated YAP inhibition during cardiac maturation. Nat. Commun. 14, 4716 (2023).
Liu, X., Dai, S.-K., Liu, P.-P. & Liu, C.-M. Arid1a regulates neural stem/progenitor cell proliferation and differentiation during cortical development. Cell Prolif. 54, e13124 (2021).
Doyle, D. Z. et al. Chromatin remodeler Arid1a regulates subplate neuron identity and wiring of cortical connectivity. Proc. Natl. Acad. Sci. USA 118, e2100686118 (2021).
Sjöstrand, M. & Eriksson, J. W. Neuroendocrine mechanisms in insulin resistance. Mol. Cell. Endocrinol. 297, 104–111 (2009).
Yang, J., Chen, Y., Li, X. & Xu, D. New insights into the roles of glucocorticoid signaling dysregulation in pathological cardiac hypertrophy. Heart Fail Rev. 27, 1431–1441 (2022).
Ren, R., Oakley, R. H., Cruz-Topete, D. & Cidlowski, J. A. Dual role for glucocorticoids in cardiomyocyte hypertrophy and apoptosis. Endocrinology 153, 5346–5360 (2012).
Miguel-Hidalgo, J. J. Role of stress-related glucocorticoid changes in astrocyte-oligodendrocyte interactions that regulate myelin production and maintenance. Histol. Histopathol. 38, 1–8 (2023).
Dai, J., Buijs, R. & Swaab, D. Glucocorticoid hormone (cortisol) affects axonal transport in human cortex neurons but shows resistance in Alzheimer’s disease. Br. J. Pharm. 143, 606–610 (2004).
Td, A. et al. Depression pathophysiology, risk prediction of recurrence and comorbid psychiatric disorders using genome-wide analyses. Nature med. 29, 1832–1844 (2023).
Wray, N. R. et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat. Genet. 50, 668–681 (2018).
Suzuki, K. et al. Genetic drivers of heterogeneity in type 2 diabetes pathophysiology. Nature 627, 347–357 (2024).
Mahajan, A. et al. Multi-ancestry genetic study of type 2 diabetes highlights the power of diverse populations for discovery and translation. Nat. Genet. 54, 560–572 (2022).
Tan, Y., He, Q. & Chan, K. H. K. Identification of shared genetic architecture between non-alcoholic fatty liver disease and type 2 diabetes: a genome-wide analysis. Front. Endocrinol. 14, 1050049 (2023).
Aragam, K. G. et al. Discovery and systematic characterization of risk variants and genes for coronary artery disease in over a million participants. Nat. Genet. 54, 1803–1815 (2022).
Mishra, A. et al. Stroke genetics informs drug discovery and risk prediction across ancestries. Nature 611, 115–123 (2022).
Foulquier, S. et al. WNT signaling in cardiac and vascular disease. Pharm. Rev. 70, 68–141 (2018).
Sanchez-Ruiz, J. A. et al. The Wnt signaling pathway in major depressive disorder: a systematic review of human studies. Psychiatry Res. 339, 115983 (2024).
Russell, J. O. & Monga, S. P. Wnt/β-catenin signaling in liver development, homeostasis, and pathobiology. Annu. Rev. Pathol. Mech. Dis. 13, 351–378 (2018).
Fan, Y. et al. Wnt/β-catenin-mediated signaling re-activates proliferation of matured cardiomyocytes. Stem Cell Res. Ther. 9, 338 (2018).
Brade, T., Männer, J. & Kühl, M. The role of Wnt signalling in cardiac development and tissue remodelling in the mature heart. Cardiovasc. Res. 72, 198–209 (2006).
Fancy, S. P. J. et al. Dysregulation of the Wnt pathway inhibits timely myelination and remyelination in the mammalian CNS. Genes Dev. 23, 1571–1585 (2009).
Zou, Y. Wnt signaling in axon guidance. Trends Neurosci. 27, 528–532 (2004).
van Loo, G. & Bertrand, M. J. M. Death by TNF: a road to inflammation. Nat. Rev. Immunol. 23, 289–303 (2023).
Ménoret, A., Chaillot, D., Callahan, M. & Jacquin, C. Hsp70, an immunological actor playing with the intracellular self under oxidative stress. Int J. Hyperth. 18, 490–505 (2002).
Sudlow, C. et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Liu, Y. et al. Genetic architecture of 11 organ traits derived from abdominal MRI using deep learning. eLife 10, e65554 (2021).
Bai, W. et al. A population-based phenome-wide association study of cardiac and aortic structure and function. Nat. Med. 26, 1654–1662 (2020).
Alfaro-Almagro, F. et al. Image processing and quality control for the first 10,000 brain imaging datasets from UK Biobank. Neuroimage 166, 400–424 (2018).
Smith, S. M., Alfaro-Almagro, F. & Miller, K. L. UK Biobank - Brain Imaging Documentation. Accessed October, https://biobank.ndph.ox.ac.uk/showcase/ukb/docs/brain_mri.pdf. (2024).
Smith, S. M. et al. An expanded set of genome-wide association studies of brain imaging phenotypes in UK Biobank. Nat. Neurosci. 24, 737–745 (2021).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Mulugeta, A., Hyppönen, E., Ala-Korpela, M. & Mäkinen, V.-P. Cross-sectional metabolic subgroups and 10-year follow-up of cardiometabolic multimorbidity in the UK Biobank. Sci. Rep. 12, 8590 (2022).
Alfaro-Almagro, F. et al. Confound modelling in UK Biobank brain imaging. NeuroImage 224, 117002 (2021).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Werme, J., van der Sluis, S., Posthuma, D. & de Leeuw, C. A. An integrated framework for local genetic correlation analysis. Nat. Genet. 54, 274–282 (2022).
Chung, D., Yang, C., Li, C., Gelernter, J. & Zhao, H. GPA: a statistical approach to prioritizing GWAS results by integrating pleiotropy and annotation. PLoS Genet. 10, e1004787 (2014).
Genomes Project Consortium et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Zhu, X. et al. Meta-analysis of correlated traits via summary statistics from GWASs with an application in hypertension. Am. J. Hum. Genet. 96, 21–36 (2015).
Li, X. & Zhu, X. Cross-phenotype association analysis using summary statistics from GWAS. Methods Mol. Biol. 1666, 455–467 (2017).
Watanabe, K., Taskesen, E., van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 8, 1826 (2017).
Liu, M. et al. Investigating the shared genetic architecture between depression and subcortical volumes. Nat. Commun. 15, 7647 (2024).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Boyle, A. P. et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 22, 1790–1797 (2012).
Roadmap Epigenomics Consortium et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Barbeira, A. N. et al. Integrating predicted transcriptome from multiple tissues improves association detection. PLoS Genet. 15, e1007889 (2019).
SCORE2 working group and ESC Cardiovascular risk collaboration. SCORE2 risk prediction algorithms: new models to estimate 10-year risk of cardiovascular disease in Europe. Eur Heart J 42, 2439–2454 (2021).
Wang J. et al. Multi-organ network of cardiometabolic disease-depression multimorbidity revealed by phenotypic and genetic analyses of MR Images. Zenodo. https://doi.org/10.5281/zenodo.17669873 (2025).
Acknowledgements
This study was supported by the AI4S Climber Initiative (awarded to VWZ and ML), Intern Discovery (awarded to VWZ and ML), Natural Science Foundation of Hubei Province (2025AFB479, awarded to XL), Health Commission Foundation of Hubei Province (WJ2025Q070, awarded to XL), National Natural Science Foundation of China (82572306, awarded to FL), and Innovative Research Team of High-Level Local Universities in Shanghai (awarded to VWZ). This research has been conducted using the UK Biobank Resource under Application Number 101169. This work uses data provided by patients and collected by the NHS as part of their care and support.
Author information
Authors and Affiliations
Contributions
J.W., M.L., F.L., and G.Y. contributed equally to the paper as joint first authors. V.W.Z., N.F., S.R., and X.L. are joint corresponding authors. V.W.Z., N.F., S.R., X.L., J.W., M.L., F.L., and G.Y. designed the research. J.W. and G.Y. performed statistical analyses. J.W., M.L., F.L., G.Y., V.W.Z., N.F., S.R., X.L., Z.Y., H.H., Z.Z., L.W., Y.W., W.F., S.S., M.C., X.J., Q.Y., and J.L. interpreted data, critically reviewed and revised the manuscript, and have read and approved the final version. V.W.Z. is the guarantor of this work and, as such, had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, J., Liu, M., Liu, F. et al. Multi-organ network of cardiometabolic disease-depression multimorbidity revealed by phenotypic and genetic analyses of MR images. Nat Commun 17, 1333 (2026). https://doi.org/10.1038/s41467-025-68092-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-68092-7