Abstract
Sexual maturation and egg production in schistosomes require sustained male-female pairing, yet the cellular dynamics underlying this process remain poorly understood. Here, we generate a high-resolution single-cell transcriptomic atlas of 104,671 cells from male and female Schistosoma japonicum across four developmental stages, identifying 76 distinct cell populations and mapping lineage trajectories in reproductive and neural systems. Notably, we discover a male-specific neuron subtype - N4.3 neuron located in the gynecophoric canal, and identify lsamp as a critical gene co-expressed with nrps in N4.3 neurons. Loss of lsamp impairs axonal integrity and disrupts the male ventral nerve network, reducing the targeted delivery of pheromone BATT precursor to N4.3 neurons, thereby hindering female reproductive maturation. This work maps schistosome male-female interactions at single-cell resolution, revealing a male ventral nerve network that facilitates intersex communication and promotes female reproductive development, offering strategies to disrupt parasite reproduction and combat schistosomiasis.
Similar content being viewed by others
Introduction
Schistosomiasis is a prevalent and devastating neglected tropical disease caused by parasitic flatworms called Schistosoma spp., which affects over 250 million people in disadvantaged areas1. Currently, no vaccine is available, and treatment relies on a single drug (praziquantel)2,3. Therefore, new therapeutic and preventive strategies are urgently needed. Eggs produced by mature female schistosomes drive the pathology and transmission of this disease. In a definitive mammalian host, one mature female parasite could produce hundreds to thousands of eggs daily that are deposited in the host tissues, e.g., the liver, where they induce the local granuloma and further lead to tissue fibrosis4,5. Eggs excreted with feces or urine hatch in the water environment, and then the released miracidia invade the intermediate snail host before spreading. Therefore, developing strategies for obstructing the egg production of these parasites will contribute to the control of this disease.
Schistosomes have a unique reproductive feature. The pairing with the male worm initiates the sexual development of a female worm and maintains her mature state6. Unlike females, the male worm can reach sexual maturity independently7. Continued investigations have been carried out to uncover the underlying mechanism of this male-induced female reproduction since the first report of the phenomenon in Schistosoma japonicum nearly a century ago8. Recently, a dipeptide pheromone (β-alanyl-tryptamine) synthesized and secreted by male worms has been identified in S. mansoni that stimulates female sexual development9. The discovery of this neuron-derived key molecule that can be transmitted from males to females highlights the importance of understanding the complex network during male-female interplay.
The direct physical connection between males and females occurs at their late juvenile stage when the male worm clasps the female through the gynecophoric canal to form a worm pair10. Thereafter, the coupled male and female continue to grow and turn to be sexually mature. Besides the extension of body length, a series of previous studies have observed dramatic gene expression changes in males and females upon pairing10,11,12,13. These changes are not only restricted to the sexual organs but also to other somatic tissues, such as neurons and muscles. Additionally, labor division is more apparent between the two sexes at their adult stage, which is represented by the distinct sex-associated gene expression patterns10,14,15,16,17. Recently, researchers have revealed that the cell types of S. mansoni expanded along with the growth and development of the parasite in the mammalian hosts18,19,20. Notably, single-cell RNA sequencing showed distinct cell clusters between the adult male and female and between the virgin female and mature female in S. mansoni21, which highly indicates a dynamic sex-related cellular differentiation during the male-female interaction. However, the whole picture of the molecular changes of males and females during their interaction at the single-cell level is still lacking.
In this study, we construct the dynamic single-cell atlas of male and female S. japonicum across 4 time points throughout the reproductive development during their interplay. We sequence 104,671 single-cell transcriptomes and characterize 76 molecularly distinct cell populations. Along with the male-female interaction, the cell clusters of the reproductive and nervous systems undergo the most dramatic changes in both sexes. Trajectory inference analysis reveals the detailed lineage of cells related to reproductive development in both sexes. We analyze and predict gene expression patterns and key regulatory factors involved in the maturation of late male and female germ cells. Furthermore, we notice a complex heterogeneity and sex-biased characteristics of the nervous system during the maturation of the parasite. Notably, we identify a lsamp gene expressed within a male-specific neuron population, which triggers female reproductive development by maintaining axonal integrity within the male gynecophoric canal in S. japonicum. Taken together, our datasets provide a comprehensive single-cell atlas of male and female S. japonicum across their maturation and reveal the key regulator of a male-specific neuron that is required for the stimulus of female sexual development, which will further prompt the innovation of strategies against this human parasite.
Results
Dynamic single-cell transcriptomes of male and female S. japonicum during their interplay
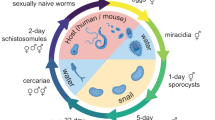
S. japonicum primarily undergoes pairing and sexual maturation between 14 and 26 dpi (days post-infection)10. To characterize the dynamic single-cell atlas of S. japonicum throughout the male-female pairing and subsequent maturation process, we collected worms at 4 time points (14, 18, 22, and 26 dpi) from a mouse model (Fig. 1a). Whole worms were separated by sex at each stage, except for those from 14 dpi (since their sex could not be distinguished morphologically at this stage). Subsequently, we dissociated these samples and isolated live single cells using fluorescence-activated cell sorting (FACS) (Supplementary Fig. 1a). Utilizing a 10× Genomics Chromium controller, we generated 7 single-cell RNA-seq (scRNA-seq) libraries (Fig. 1a and Supplementary Data 1). Through the analysis of these libraries using Cell Ranger, we obtained a total of 110,204 cells (Supplementary Data 1). After excluding cells with more than 30% mitochondrial reads or fewer than 500 detected genes, the final count was 104,671 cells, which were further clustered into 76 distinct cell clusters (Supplementary Fig. 1b and Data 1). Furthermore, we annotated the identities of these cell clusters by referring to cell markers defined in the single-cell sequencing of S. mansoni18,19,21,22 (Supplementary Data 2). In total, the S. japonicum single-cell atlas comprised 33 transcriptionally distinct neuron cell clusters, 14 neoblast clusters, 9 tegument cell clusters, 5 muscle cell clusters, 5 parenchyma clusters, 3 germline stem cell (GSC) clusters, 1 vitellaria cluster, 1 late male germ cell cluster, 1 Mehlis’ gland cell cluster, 1 gut cell cluster, 1 flame cell cluster, 1 esophageal gland cell cluster, and 1 unknown cluster (Supplementary Fig. 1b).
a Schematic workflow of the single-cell RNA sequencing (10× Genomics) for female and male S. japonicum at 14 dpi (days post infection, mixed sex), 18, 22, and 26 dpi. b UMAP visualization of 104,671 single-cell transcriptomes from seven samples, classified into 21 distinct cell types (including one unannotated “unknown” population).
Our single-cell atlas provided detailed lineage information for the parasite’s specific tissues. Interestingly, our data unexpectedly revealed many schistosome neural progenitor cells and neural precursor cells. In comparison, the single-cell atlas of S. mansoni depicts neural cells as scattered clusters lacking apparent lineage connections21. Based on their marked heterogeneity, we classified the neural cells into five categories: neural progenitor cells, neural precursor cells, kk7+ neurons, nonciliated neurons, and ciliated neurons (Fig. 1b). The tegument cells are divided into tegument progenitor, tegument progeny 1, tegument progeny 2, syncytial 1, and syncytial 2 (Fig. 1b). In addition to the well-defined cell markers in previous studies on S. mansoni21, we further identified the marker genes as shown in Supplementary Fig. 2 and Supplementary Data 3 and 4 through whole-mount in situ hybridization (WISH) and double fluorescence in situ hybridization (double-FISH), including new markers such as Sjc-0003010 and Sjc-0008393 for GSCs (Supplementary Fig. 3a), Sjc-0001566 (ccdc) for late male germ cells (Supplementary Fig. 3c), Sjc-0004922 (Tubulin alpha-1 chain) for neoblasts (Supplementary Fig. 3e), Sjc-0008047 (klf11) for neural progenitor cells (Supplementary Fig. 3f), Sjc-0005931 (acss2), and Sjc-0007590 (pcdh7) for neural precursor cells (Supplementary Fig. 3g), Sjc-0002588 (slc6a5) for parenchyma cells (Supplementary Figs. 3i and 4a, b), Sjc-0001890 (sgf1) for esophageal gland cells (Supplementary Fig. 3k), Sjc-0008478 and Sjc-0000277 (slc7a14) for syncytial of tegument cells (Supplementary Figs. 3m and 4a, c), Sjc-0008416 (cpb) and Sjc-0000095 (cacnb1) for muscle cells (Supplementary Figs. 3n and 4a, d).
We first analyzed the distribution of cells in each sample, which allowed us to identify developmental differences at the cellular level between male and female worms at various stages (Supplementary Fig. 5). We also examined the relative proportions of cells from different stages within the cell types of the combined S. japonicum dataset (Supplementary Fig. 6a). As expected, the proportion of the reproduction-related cell types gradually increased throughout the sexual maturation process in both sexes, such as GSCs, female vitellaria, and late male germ cells (Supplementary Fig. 6a and Data 5 and 6). To validate this, we performed WISH using the GSCs marker gene eled18. As shown in Supplementary Fig. 6b, the expression region of eled expanded significantly in both female and male sexual organs from 18 to 26 dpi, which was consistent with the single-cell data.
The proportion of neural progenitor cells and precursor cells decreased after 14 dpi in both sexes. In male worms, the populations of differentiated neural cells (e.g., nonciliated neurons and ciliated neurons) continuously expanded. In female worms, these populations did not show sustained expansion but rather exhibited an initial increase followed by a subsequent decline after 22 dpi, which is consistent with the weakening of neural activity in females, as revealed by bulk RNA-seq analysis10 (Supplementary Fig. 6a and Data 5).
Detailed lineage analysis of cells related to reproductive development in both sexes
The schistosome germ cells originate from a subset of stem cells at the asexual stage and eventually separate from somatic cell lineage at the juvenile stage18. Upon pairing, the committed GSCs in both sexes proliferate and differentiate into mature gametes. To demonstrate the cellular dynamics of the parasite’s reproductive system and the relationship between germline and somatic lineages, we subset the cells defined as stem cells (neoblasts) and those present in the reproductive organs, including GSCs, late male germ cells, late female germ cells, vitellaria, and Mehlis’ glands, from both female and male samples (18, 22, and 26 dpi), and performed subcluster analysis. Then, we identified 17 clusters with representative markers for each subtype based on well-defined markers from previous studies (Supplementary Fig. 7a and Data 2), such as the neoblast marker nanos-2 (Sjc-0005962)18,23, the vitellaria S1 cell marker vf1 (Sjc-0000108)24, the GSC markers nanos-1 (Sjc-0006622), and eled (Sjc-0009220)18,25 (Supplementary Fig. 7b, c).
We also validated newly identified tissue-specific markers using WISH, including Sjc-0000843 (ribc2, RIB43A-like with coiled-coils protein 2) for late male germ cells, Sjc-0003040 for late female germ cells, esg-2a (eggshell protein gene 2a26, Sjc-0006798) for vitelline cells, and Sjc-0000795 for Mehlis’ gland (Fig. 2a, Supplementary Fig. 7b, c, and Data 7). These markers can effectively characterize the developmental status of each organ. Interestingly, late female germ cells were also observed in male worms, albeit in relatively low numbers (Supplementary Fig. 7a, c). To confirm the presence of late female germ cells in males, we performed WISH using the late female germ cell marker gene Sjc-0004512 (clec, C-type lectin domain-containing protein) and observed a signal in the testes of male worms (Supplementary Fig. 7d). To further characterize clec expression within the testis, we conducted double-FISH with Sjc-0000843 (ribc2), a marker of late male germ cells. Based on testicular morphology visualized by DAPI staining (outlined by circles), both clec and ribc2 were expressed in the testes. Notably, weak ribc2 signals were also observed in clec⁺ germ cells (Supplementary Fig. 7e). Interestingly, although schistosomes are the only dioecious species within the class Trematoda, various forms of hermaphroditism have been documented, including males developing ovarian-like structures and females exhibiting testicular characteristics27,28,29. Our identification of cells expressing female germ cell markers in phenotypically normal male worms provides additional molecular insight into the evolution of sexual differentiation in schistosomes.
a (left) heatmaps showing tissue-specific markers and (right) in situ hybridization of candidate markers for characterizing the developmental status of reproductive organs: Sjc-0003040 (Ovary, Ov), Sjc-0000795 (Mehlis’ gland, Mg), Sjc-0006798 (esg-2a) (Vitellaria, Vi), and Sjc-0000843 (ribc2) (Testis, Te). The number of worms displaying the shown localization out of the total observed in 3 biological replicates is as follows: Sjc-0003040, n = 13/16 (22 dpi) and n = 15/17 (26 dpi); Sjc-0000795, n = 20/25 (22 dpi) and n = 23/24 (26 dpi); Sjc-0006798, n = 27/32 (22 dpi) and n = 24/26 (26 dpi); Sjc-0000843, n = 18/20 (22 dpi) and n = 20/20 (26 dpi). Scale bars = 100 μm. b RNA velocity analysis of cells related to the reproductive development in both S. japoncium sexes. c UMAP plots of female germ cells showing three subtypes (left) and Slingshot pseudotime trajectories of female germline differentiation (right) in S. japoncium. d UMAP plots of male germ cells showing three subtypes (left) and Slingshot pseudotime trajectories of male germline differentiation (right) in S. japoncium. e Expression dynamics of representative genes along female pseudotime, including Sjc-0003040, Sjc-0004774 (meiob), Sjc-0007561 (nuob), and Sjc-0008219 (horm-2). f Expression dynamics of representative genes along male pseudotime, including Sjc-0000843 (ribc2), Sjc-0004774 (meiob), Sjc-0007561 (nuob) and Sjc-0008219 (horm-2). g Expression of Sjc-0001604 (oc-1), Sjc-0006622 (nanos-1), Sjc-0009220 (eled), and Sjc-0009231 (boule) along the male pseudotime.
By performing RNA velocity analysis30, we inferred the possible developmental trajectories from GSCs to late germ cells, separately for female and male worms (Fig. 2b). To further investigate the differences in the germline lineage, we performed trajectory inference using Slingshot31, an expression-based method, on the GSCs, GSC progenies, and the germ cells of male and female worms. It revealed the cell trajectory originating from the GSCs population into late germ cells, in both sexes (Fig. 2c, d).
Subsequently, we analyzed the expression patterns of genes involved in the differentiation of late male or female germ cells. Temporally dynamic genes were grouped into six modules for each sex, each showing distinct expression trends and functional enrichments (Supplementary Fig. 8 and Data 8 and 9).
In females, 1596 genes formed modules associated with translation, RNA processing, DNA conformation change, vesicle organization, and muscle contraction (Supplementary Fig. 8a and Data 10). In males, 1,913 genes were grouped into modules linked to translation, protein processing, ERBB signaling, cilium organization, and microtubule-based movement (Supplementary Fig. 8b and Data 11). Shared dynamic genes between sexes were also identified (Supplementary Data 12).
Analysis of known and newly identified germline regulators showed that meiob, nuob, and horm-2 exhibited similar biphasic expression in both sexes—rising early then declining—indicating conserved roles in early-to-mid gametogenesis; In contrast, Sjc-0003040 (female) and ribc2 (male) were upregulated during late gametogenesis, suggesting sex-specific functions (Fig. 2e, f). Temporal profiling of established germline regulators revealed that nanos-1 was highly expressed in early stages, consistent with its role in germ stem cell maintenance; Conversely, oc-1 and boule promoted spermatogenic differentiation by suppressing self-renewal, while eled inhibited sperm differentiation32 (Fig. 2g). Together, these findings provide a pseudotemporal framework of germ cell development in S. japonicum, revealing that distinct gene modules coordinate sex-specific and stage-specific processes during gametogenesis.
Vitellaria is the largest reproductive organ in adult female schistosomes that occupies two-thirds of the body’s length. We found that both vitellaria S1 cells and GSCs express a certain level of eled (Sjc-0009220) (Supplementary Fig. 7b, c), suggesting a potential shared origin from a population of eled+ stem cells, as previously reported18.
Vitellaria maturation is crucial for egg production in schistosomes. Previous studies have shown that vitellaria development can be divided into stages S1–S4, all of which express vf1 (Sjc-0000108)24. In the single-cell transcriptomic analysis of S. mansoni, the vitellaria development process was further categorized into five stages: S1, S1 progeny, early vitellocytes, late vitellocytes, and mature vitellocytes21. However, the detailed differentiation trajectory of vitellaria cells has not been fully elucidated at the single-cell level. Here, we performed subclustering of vitelline cells and identified eight distinct subtypes, including S1(S1-0, and S1-1), S1 progeny(S1p-0, S1p-1, and S1p-2), early vitellocytes (EVi), and late vitellocytes (LVi-0, and LVi-1) (Supplementary Fig. 9a, b and Data 13). Trajectory analysis with Slingshot revealed two differentiation branches (Lineage 1 and Lineage 2; Supplementary Fig. 9c, d), among which only Lineage 1 showed high differentiation potential according to CytoTRACE33 (Supplementary Fig. 9e). Gene expression dynamics along Lineage 1 indicated that, in addition to the known marker vf1, p48 and the Sjc-0006798 (esg-2a, identified in this study) gradually increased in expression (Supplementary Fig. 9f), supporting Lineage 1 as the primary vitelline developmental pathway. This trajectory, annotated in the RNA velocity plot (Fig. 2b), traces the progression from S1 (vf1+ esg-2a−) to S1 progeny (vf1+ esg-2a+), and subsequently to EVi and LVi (Supplementary Fig. 10). Pseudotime-associated genes by tradeSeq34 (p < 1 × 10−5) were grouped into six co-expression modules (Supplementary Fig. 9g and Data 14) reflecting stage-specific processes: early–mid modules (C1, C3, C6) were enriched for DNA replication and cell cycle functions, whereas late modules (C2, C4) were enriched for eggshell formation–related pathways, including protein localization to the ER and purine metabolism (Supplementary Data 15).
Mehlis’ gland, a specialized organ unique to trematodes and cestodes, has been long speculated to play an essential role in eggshell formation and the release of eggshell granules35,36. However, research on its function in schistosomes remains limited. In our study, we identified candidate marker genes for the Mehlis’ gland, enriching its molecular characterization (Supplementary Data 7). Additionally, we observed that Mehlis’ gland cells emerge earlier than vitelline cells, appearing in large numbers by 18 dpi, while vitelline cells do not proliferate in large numbers until 22 dpi (Supplementary Fig. 11a, b). These findings were further validated through WISH experiments using the marker gene Sjc-0000795 for Mehlis’ gland, the marker gene esg-2a for vitellaria and the marker gene Sjc-0000278 (rreb1) for both vitellaria and Mehlis’ gland (Supplementary Fig. 11c).
Five neural lineages in S. japonicum
Unlike the scattered neuronal clusters revealed from the previously reported single-cell transcriptomic atlas of adult Schistosoma mansoni21, our datasets exhibit high connectivity among complex neuronal clusters (Fig. 1b and Supplementary Fig. 1b), which is likely to enable cell lineage reconstruction of the whole schistosome nervous system from transcriptomic data. We first reclustered all the neural cells of males and females, including neural progenitor cells, neural precursor cells, nonciliated neurons, and ciliated neurons, identifying 19 subclusters and summarizing them into 5 lineages, which were named N1 lineage, N2 lineage, N3 lineage, N4 lineage, and N5 lineage (Supplementary Figs. 12 and 13a).
Subsequently, we employed Slingshot to confirm and refine the reconstructed cell differentiation lineages and to infer pseudotime (Supplementary Figs. 12 and 13b), which revealed a clear alignment with sample collection times (Supplementary Figs. 12 and 13c). Additionally, we performed RNA velocity analysis on the male and female datasets separately using scVelo algorithms37. The velocity vectors were visualized on the UMAP embedding, revealing potential differentiation trajectories from neural progenitor cells toward mature cell types (Supplementary Figs. 12 and 13d).
To better understand the development trajectories leading up to different neuron populations, we performed CellRank38 analysis on both male and female neuron cells. In male S. japonicum, we identified 7 metastable states in the neural cells, with neural progenitor cells predicted as the initial state and N1.1, N2.2, N3.2, N4.2, N4.3, and N4.4 neuron inferred as six terminal states (Supplementary Fig. 12e). In female S. japonicum, we identified 7 metastable states in the neural cells, with neural progenitor cells predicted as the initial state and N1.1, N2.1, N3.1, N4.1, N4.2, and N5 neuron inferred as six terminal states (Supplementary Fig. 13e). In summary, we defined neural cell differentiation from an early neural progenitor state to several mature states and provided a mechanistic framework for characterizing worm neuronal cell fate decisions.
Male-enriched neuron clusters
To explore the neuronal diversity in S. japonicum, we performed a subcluster analysis of neuronal cells in both male and female parasites (Fig. 3a). From the integrated single-cell atlas of both sexes, we identified three neuron populations enriched in males: N2.2, N3.2, and N4.3 (Fig. 3a). The proportional distribution of neuronal subtypes in both sexes (Supplementary Fig. 14a) and the differential abundance analysis of neuronal cell types across sexes (Supplementary Fig. 14b) further confirmed the existence of these male-enriched neurons.
a UMAP subclustering of the nervous system in female (left) and male (right) S. japonicum. Three neuron populations enriched in males (N2.2, N3.2, and N4.3) are highlighted with circles. b FISH (Fluorescence in situ hybridization) localization of neural marker genes (nrps, Sjc-0001200, Sjc-0003639) in adult male worms. Scale bars = 200 μm. c GO (Gene Ontology) enrichment analysis of highly expressed genes in N2.2, N3.2, and N4.3 neurons. d Key driver genes (nrps, ddc, and lsamp) associated with the N4.3 neuron development. e Heatmaps showing three driver genes expressed in N4.3 neurons. f Localization of the lsamp gene in adult male worms: (left) in situ hybridization; (right) double-FISH colocalization of lsamp with nrps or ddc in N4.3 neurons. Scale bars = 200 μm (left) and 5 μm (right). g The expression profiles of nrps, ddc, and lsamp in female (red line) and male (blue line) S. japonicum based on previous RNA-seq data10. LOESS-smoothed curves are shown with 95% confidence bands (shaded areas), representing 95% confidence intervals calculated from the standard errors of the local regression fits. All of them exhibit male-enriched and pairing-induced expression patterns. Source data are provided as a Source Data file.
Using RNA probes targeting specific marker genes (Supplementary Fig. 15 and Data 16), we further validated the spatial distribution of male-enriched neuronal populations—N2.2 (Sjc-0003639+Sjc-0000814 (nrps)−), N3.2 (Sjc-0001200+), and N4.3 (nrps+)—as well as additional mature neuronal populations, including N2.1 (ano7, Sjc-000798+), N3.1 (Sjc-0006108+), N4.1 (bmp, Sjc-0004182+), N4.4 (fhl2, Sjc-0002320+), and N5 (Sjc-0000744+), in both sexes (Supplementary Figs. 16 and 17 and Data 16). As anticipated, all neurons expressed the pan-neuronal marker 7b2 (Supplementary Figs. 16 and 17). Notably, all three male-enriched neuronal subtypes were located within the gynecophoric canal, a specialized male structure that secures the female during pairing (Fig. 3b and Supplementary Fig. 16). Among these, the N4.3 population was exclusively distributed throughout the gynecophoric canal, whereas N2.2 and N3.2 populations were additionally detected in the head of both male and female parasites (Fig. 3b and Supplementary Fig. 16). To integrate these observations, we generated representations summarizing the spatial distribution patterns of all validated neuronal populations (Supplementary Fig. 18).
Interestingly, the marker gene Sjc-0000814 of N4.3 is nrps (nonribosomal peptide synthetase), which encodes the NRPS enzyme. In S. mansoni, the NRPS enzyme has been reported to synthesize β-alanyl-tryptamine (BATT), a dipeptide pheromone that passes from the males to induce female reproductive development9. Thus, our study clarified that the N4.3 neurons are the key populations that regulate female sexual development by expressing the nrps to produce BATT.
To elucidate the potential functions of these three male-enriched populations, we performed GO (Gene Ontology) enrichment analysis on the highly expressed genes within each neuronal subclass. These enriched genes indicated that N2.2 and N3.2 populations exhibited typical neuronal functions, such as synaptic transmission and neurotransmitter transport. Moreover, the N3.2 population showed more specific enrichments, particularly in ion transport (Fig. 3c, Supplementary Fig. 19, and Data 17 and 18). Unlike them, the N4.3 population emerged as a specialized neuronal subtype, expressing genes closely related to cilium organization, cell projection assembly, and microtubule bundle formation (Fig. 3c and Supplementary Fig. 19). This finding aligns with previous observations in S. mansoni, where SEM studies identified Sm-nrps+ cells with cilium-like protrusions, suggesting a conserved role of ciliated neurons in male-specific functions9. Collectively, these results indicate that these three types of neurons play different roles in male parasites. Considering the importance of the key enzyme coded by nrps in the N4.3 neuron that synthesizes the pheromone required for the female sexual development in schistosomes, we wondered whether there are essential genes maintaining the function of the N4.3 population.
Male lsamp is essential for female sexual development
To identify genes potentially driving the differentiation of N4.3 neurons, we analyzed the driver genes and pathways associated with the N4.3 neuron metastable state by correlating fate probabilities with gene expression using CellRank’s compute_lineage_drivers function38 (Supplementary Data 19 and 20). Among these, genes such as nrps (Sjc-0000814), ddc (Sjc-0006754, Aromatic L-amino acid decarboxylase), and lsamp (Sjc-0000373, limbic system-associated membrane protein) exhibited the highest correlations (Fig. 3d and Supplementary Data 19). Additionally, these three genes are specifically expressed in N4.3 neurons as indicated on the UMAP plot (Fig. 3e). The colocalization of nrps and ddc has been confirmed previously (Supplementary Fig. 16c). We performed double-FISH and found that lsamp is co-expressed with nrps and ddc in male N4.3 neurons (Fig. 3f and Supplementary Fig. 20). Additionally, lsamp and ddc also exhibited colocalization in the large cerebral ganglia (Fig. 3f). Nrps is a male-specific gene, and its expression is induced by pairing9. In S. japonicum, both ddc and lsamp displayed similar expression patterns to nrps, as reported in our previous study10 (Fig. 3g). We therefore reasoned that lsamp or ddc may play an important role in the N4.3 neuron that may further affect the male-female interaction.
By employing RNAi in vitro, we found the male worms treated with lsamp dsRNA lost their ability to initiate female sexual development (Fig. 4a, b). These female worms paired with lsamp RNAi males displayed reduced Fast Blue BB staining in their vitellaria and possessed poorly developed ovaries (Fig. 4b). In addition, they barely laid eggs (Fig. 4c). The qRT-PCR result indicated a strong reduction of the lsamp expression in the male worms following lsamp RNAi treatment (Fig. 4d). In contrast, treatment with ddc dsRNA did not affect female sexual development (Supplementary Fig. 21). The above results suggest that male lsamp is essential for female sexual development.
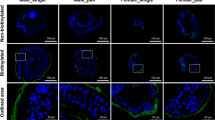
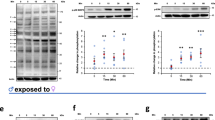
a Flowchart of in vitro RNAi interference and phenotypic observation. b Female worms paired with lsamp RNAi males exhibited reduced Fast Blue BB staining in vitellaria and underdeveloped ovaries on D28. Scale bars = 200 μm. c Egg production was significantly reduced in female worms paired with lsamp RNAi-treated males on D28. n = 3 biological replicates; p = 0.0006. d qRT-PCR validation of lsamp knockdown in male worms on D28. n = 9 (gfp RNAi) and n = 10 (lsamp RNAi); p < 0.0001. e FISH localization of N4.3 neurons (ddc+ cells) in the lsamp RNAi group and gfp RNAi group on D28 (left). Scale bars = 50 μm; Quantification of N4.3 neurons (ddc+ cells per 100 μm²) showed no significant change upon lsamp silencing on D28 (right, n = 8 biological replicates); p = 0.7582. f qRT-PCR analysis confirmed that nrps expression was unaffected by lsamp downregulation on D28. n = 10 (gfp RNAi) and n = 9 (lsamp RNAi); p = 0.7717. g LC-MS/MS results show levels of β-alanine remained unchanged in the lsamp RNAi group on D20 (n = 12 biological replicates); p = 0.9759. h LC-MS/MS results show levels of tryptamine remained unchanged in the lsamp RNAi group on D20 (n = 12 biological replicates); p = 0.6106. i LC-MS/MS results show β-alanyl-tryptamine (BATT) contents both in the worms (left) and the culture medium (right) were significantly reduced in the lsamp RNAi group on D20. Left: n = 12 (gfp RNAi) and n = 11 (lsamp RNAi); analyzed by non-parametric Mann–Whitney U test; p = 0. 0056. Right: n = 12 biological replicates; p = 0.0021. Statistical significance was analyzed by unpaired two-tailed Student’s t-tests, and data are presented as mean ± SEM (c–h and i, right). ns,not significant; *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001. Source data are provided as a Source Data file.
To determine whether the female phenotype resulting from the male lsamp knockdown was due to a reduction in the number of N4.3 neurons, we performed FISH experiments to quantify these cells. We selected the marker gene ddc to detect N4.3 neurons specifically. However, there was no significant difference in the lsamp silencing group compared to the control group (Fig. 4e).
Lsamp affects acetylated α-tubulin formation and vesicular transport within neurons
Since lsamp colocalized with nrps in N4.3 neurons (Fig. 3f and Supplementary Fig. 20), and nrps has been recognized as the functional gene encoding an enzyme to synthesize the male pheromone9, we therefore wondered whether the lsamp RNAi phenotype is caused by affecting the nrps expression. qRT-PCR results showed that the nrps expression was not affected in the lsamp knockdown males (Fig. 4f and Supplementary Fig. 22). According to Chen et al., the NRPS enzyme uses two simple precursors (β-alanine and tryptamine) to form the dipeptide pheromone BATT9. To explore why knocking down lsamp in male worms blocks female sexual development, we detected the levels of two precursors in worms (D20) using LC-MS/MS but found no changes between the gfp and lsamp RNAi groups (Fig. 4g, h). We then measured the amount of BATT, the product of the NRPS, that is made in males and can be released to the medium in vitro. Surprisingly, BATT contents were significantly reduced in the lsamp RNAi group, both in the male worms and the culture medium (Fig. 4i), even though the nrps level was not changed (Fig. 4f and Supplementary Fig. 22). These findings suggest that lsamp RNAi impairs the ability of male N4.3 neurons to synthesize BATT, consequently failing to stimulate female sexual development, not through affecting nrps expression.
To investigate the underlying mechanism of male lsamp in regulating BATT synthesis, we performed RNA-seq to overview the gene expression changes between paired lsamp RNAi and gfp RNAi males (D20). Silencing lsamp resulted in the downregulation of 50 genes and the upregulation of 19 genes (Supplementary Fig. 23a, b and Data 21 and 22). Our RNA-seq data confirmed that the expression of nrps and ddc remained unaffected by lsamp knockdown (Supplementary Fig. 23c), consistent with our previous results (Fig. 4f, and Supplementary Fig. 22). Since the upregulation of nrps is triggered by physical contact between males and females9, these results suggest that the knockdown of lsamp does not impair male worms’ ability to sense females. Furthermore, GO term analysis of the downregulated genes in lsamp RNAi males revealed significant enrichment in biological processes, including the generation of precursor metabolites and energy, and peptide biosynthesis, as well as molecular functions such as structural constituents of the ribosome and ATPase-coupled ion transmembrane transporter activity (Supplementary Fig. 23d, e and Data 23 and 24). KEGG pathway analysis of the downregulated genes showed that lsamp mainly affects pathways involved in neurodegenerative diseases such as Alzheimer’s disease and Parkinson’s disease (Fig. 5a and Supplementary Data 25), both of which are closely related to the disorder of microtubule acetylation and axon vesicular transport39,40,41,42,43. In addition, GO enrichment on the highly expressed genes in the N4.3 population indicated it as a specialized neuron that expressed genes highly related to cilium organization, cell projection assembly, and microtubule bundle formation (Fig. 3c and Supplementary Fig. 19). Previous studies in mice and nematodes have shown that lsamp homologues play a role in synapse formation, axon guidance, and neuronal stability44,45,46,47,48. We therefore hypothesized that lsamp may affect the processes of microtubule formation and axon vesicular transport in male schistosomes.
a KEGG pathway analysis of the downregulated genes set in lsamp RNAi worms, revealing significant involvement in neurodegenerative disease pathways (e.g., Alzheimer’s and Parkinson’s diseases), which are associated with microtubule acetylation and axon vesicular transport. b Immunofluorescence co-staining of acetylated α-tubulin and synapsin revealed severe degeneration of neural network functionality in the lsamp knockdown group. Scale bars = 20 μm. c The N4.3 neurons, labeled with nrps, ddc, and lsamp probes individually, exhibited colocalization with the neural axons labeled with acetylated α-tubulin. Scale bars = 10 μm. d The N4.3 neurons, labeled with nrps, ddc, and lsamp probes separately, showed colocalization with the synapsin-labeled neural axons, demonstrating clear overlap. Scale bars = 10 μm. e Development of ventral neural network in male schistosomes at 14, 18, 22, and 26 dpi, labeled with acetylated α-tubulin antibody, showing increasing robustness with maturation. Scale bars = 20 μm. f The diagram highlights the crucial role of lsamp in the maintenance of the ventral neural network in male S. japonicum. Representative immunofluorescence images from one of three independent experiments are shown in this figure (b–e).
Acetylation of α-tubulin, a post-translational modification of microtubules, is associated with microtubule stabilization and regulates cell morphology in nerve fibers39,49. It plays a crucial role in various neuronal processes, including axonal formation, dendritic growth, and neuronal migration39. And acetylated α-tubulin plays a pivotal role in the transport of synaptic vesicles within neurons. It stabilizes microtubules, which serve as “tracks” for motor proteins, facilitating the efficient movement of vesicles and ensuring the proper delivery of neurotransmitters to the synapse. In addition to classical neurotransmitters like dopamine, serotonin, and GABA, synaptic vesicles also transport other small molecules, such as the precursor β-alanine for BATT50,51,52. To further investigate the specific function of lsamp in schistosomes, we examined the acetylation status of microtubules and synapsin levels in mature male worms. As shown in Fig. 5b, acetylated α-tubulin antibody labeling revealed crisscrossed nerve fibers in the mature male gynecophoric canal. After knocking down the lsamp, the nerve fibers were markedly eliminated without affecting the nrps expression (Figs. 4f and 5b and Supplementary Fig. 22). These data suggest that LSAMP is essential for nerve fiber maintenance, likely by mediating appropriate axon guidance and target recognition. The observed alterations in microtubules may represent a secondary morphological consequence of impaired axon maintenance. Using a synapsin antibody, we detected disrupted synaptic protein distribution along nerve axons (Fig. 5b). Comparative co-staining of acetylated α-tubulin and synapsin revealed severe degeneration of neural network functionality in the lsamp knockdown group (Fig. 5b).
We next examined the spatial relationship between these neural fibers and N4.3 neurons. By labeling N4.3 neurons with three markers—ddc, nrps, and lsamp—we observed that N4.3 neurons were closely associated with neural fibers marked by acetylated α-tubulin (Fig. 5c and Supplementary Fig. 24a, b). Synaptic proteins along neural axons also exhibited substantial colocalization with N4.3 neurons, consistent with possible vesicular trafficking between these structures (Fig. 5d and Supplementary Fig. 24c). Together, these findings indicate that the loss of lsamp in N4.3 neurons disrupts the integrity of the surrounding ventral neural network in male schistosomes.
Lsamp guides the formation of the ventral neural network in males
LSAMP (Limbic system-associated membrane protein) is a glycosylated cell adhesion molecule of the IgLON family47,53. It is a GPI-anchored protein characterized by three immunoglobulin (Ig) domains (Supplementary Fig. 25). Previous studies in mice and nematodes have demonstrated that lsamp plays essential roles in neuronal development and function, including promoting neuronal growth and guiding axon targeting44,45,46,47,48,54. To investigate the potential role of lsamp in schistosomes, we examined the development of the ventral neural fiber network in male worms from 14 to 26 dpi, covering key stages from male-female pairing to full sexual maturity. At 14 dpi, when males begin pairing with females, only a few elongated nerve fibers were detected within the male gynecophoric canal. As the males developed, a characteristic crisscrossed neural fiber network became evident by 22 dpi and was maintained thereafter (Fig. 5e). Notably, the formation of this neural network coincided with a marked upregulation of lsamp expression (Fig. 3g). This temporal correlation is consistent with a potential role for lsamp in axon guidance and neural connectivity.
Overall, our study has profiled the dynamic single-cell atlas of male and female S. japonicum during pairing and sexual development. From a male-specific N4.3 neuron population, we identified the lsamp gene, which is crucial for female reproductive development and operates independently of a previously reported key regulator in males, nrps. Our observations indicate that lsamp expression is associated with the formation of ventral neural fibers in the male gynecophoric canal, which appear to connect to the N4.3 neurons (Fig. 5c–e and Supplementary Fig. 24). Based on these findings, we propose a model in which this nerve network may serve as a conduit, potentially facilitating the axonal transport of the BATT precursor—likely β-alanine rather than tryptamine, given that N4.3 neurons intrinsically express ddc required for tryptamine synthesis—to the N4.3 neurons for pheromone production and subsequent stimulation of female reproductive development (Fig. 5f).
Discussion
Our study performed single-cell RNA sequencing across four key developmental stages of male and female S. japonicum, covering the critical period from pairing to egg production. Here, we present a dynamic single-cell transcriptome atlas of S. japonicum males and females, systematically describing the proportional changes in various cell types during their developmental process. Notably, differentiation trajectories were inferred for reproductive and neuronal cells, showing the most prominent changes. A set of marker genes was identified that can be used to distinguish the reproductive status of male and female worms, and we predicted gene sets associated with gametogenesis in both sexes. Additionally, we clarified that N4.3 neurons are nrps-producing neurons and that lsamp, a highly expressed gene in N4.3 neurons, is essential for male-induced female reproductive development by maintaining axonal integrity and vesicle transport, and its deficiency ultimately leads to reduced BATT synthesis in males.
The sexual development of schistosomes is a fascinating biological process in which female worms rely on pairing with male worms to initiate reproductive maturation. Pairing behavior begins around 14 dpi, during which females become encapsulated within the male’s gynecophoric canal. By 26 dpi, most females reach sexual maturity and begin laying eggs10. Between 14 and 26 dpi, both male and female worms undergo dramatic growth and development, not only in their reproductive organs but also in other tissues. Bulk RNA-seq is limited in its ability to resolve the molecular changes occurring at the cellular level. In the present study, we used single-cell RNA sequencing to systematically profile gene expression dynamics underlying dimorphic gametogenesis in S. japonicum. Our analysis reveals that gametogenesis in S. japonicum is orchestrated by distinct, temporally coordinated gene modules that reflect the unique functional demands of oocyte and sperm production. In females, dynamic genes were enriched for processes like translation and vesicle organization, aligning with the biosynthetic needs of oocyte growth. In males, modules were distinctly associated with cilium function and motility, essential for spermatogenesis. This modularity underscores a fundamental divergence in developmental programming to achieve sexually dimorphic outcomes. Furthermore, we resolved the expression dynamics of key regulatory genes, confirming conserved roles for factors like nanos-1 in early development and identifying candidate factors such as Sjc-0003040 and ribc2 with sex-specific late-stage expression. The antagonistic patterns of regulators like oc-1 and eled illustrate the precise control over germ cell differentiation. Importantly, some unannotated genes within these clusters may represent Schistosoma-specific reproductive regulators that warrant further investigation.
Although our RNA velocity analysis of sex-related subsets revealed clear trajectories from GSCs to their progeny, it consistently showed convergence toward a central cell state identified as SSCs-2 in both sexes (Fig. 2b). The SSCs-2 population shares high expression of key markers (e.g., nanos-2, fgfrA, fgfrB, klf) with the multipotent δ'-neoblasts, which constitute a shared somatic stem cell pool in S. mansoni18. We therefore hypothesize that this convergence point may represent a similar, shared multipotent progenitor pool, capable of giving rise to multiple downstream cell types in both males and females, which requires further experimental validation.
Schistosome neural development remains poorly understood, especially the differentiation trajectories of neuronal cells. While single-cell RNA sequencing has been conducted on adult S. mansoni, it primarily captured mature neuron populations21, limiting lineage tracing analysis. In this study, we profiled single-cell data from S. japonicum at 14, 18, 22, and 26 dpi, identifying various neural cell stages, including neural progenitor cells, precursors, and mature neurons (Fig. 1). Pseudotime and RNA velocity analysis reconstructed clear lineage connections, providing a detailed neural development map (Supplementary Figs. 12 and 13). We further classified the nervous system of S. japonicum into five clear neuronal lineages, each exhibiting significant spatial heterogeneity (Supplementary Figs. 12, 13, 15–18). Intriguingly, some neurons exhibit sex-specific distribution, such as N4.3, N3.2, N2.2, N5, and N2.1 neurons (Fig. 3a, b and Supplementary Figs. 15–18). Our data reveal that male-specific N4.3 neurons express nrps (Fig. 3a, b, d, and e), a gene rapidly upregulated upon pairing. Nrps encodes NRPS, an enzyme synthesizing the male pheromone BATT, which stimulates female sexual development9,55. Additionally, we identified other driver genes expressed in N4.3 neurons, such as ddc (Sjc-0006754) and lsamp (Sjc-0000373). These findings highlight the robust heterogeneity of schistosome neurons and suggest that neural differentiation occurs at early developmental stages.
DDC is an enzyme involved in the synthesis of several monoamines, including serotonin (5-HT), tyramine, tryptamine, histamine, dopamine, and other catecholamines56,57. In planarians, inhibiting ddc expression results in the loss of ovaries and other female reproductive organs, including the yolk gland58. A recent study has shown that ddc is essential for BATT production and is co-expressed with nrps in planarians, where it likely catalyzes the conversion of tryptophan to tryptamine within BATT-producing cells to promote sexual organ development59. However, in this study, ddc knockdown (approximately 69% efficiency) in male schistosomes did not cause developmental arrest in female reproduction (Supplementary Fig. 21). The lack of an observable phenotype, despite significant knockdown, could be due to residual enzymatic activity being sufficient to maintain function, or possibly compensation through alternative pathways. We also acknowledge that the incomplete knockdown may have contributed to the absence of a pronounced phenotype. Future studies employing more potent interference strategies or conditional knockout approaches will be necessary to conclusively determine ddc’s functional role. Since biogenic amines, including 5-HT, are present in the culture medium, we also hypothesized that female schistosomes might sustain reproductive development by utilizing exogenous 5-HT. To test this, we cultured mature S. japonicum worm pairs using 5-HT-supplemented and 5-HT-free mAB169 (1640) medium. After 12 days, females in the 5-HT-free group exhibited impaired vitellaria and produced significantly fewer eggs, compared to the 5-HT-supplemented group (Supplementary Fig. 26). These findings indicate that 5-HT, a product of DDC activity, is essential for female reproductive development. Thus, while DDC and its product 5-HT are crucial for female sexual development, schistosome females can compensate for ddc knockdown by acquiring 5-HT from external sources under in vitro conditions.
Previous studies in mice and nematodes have shown that LSAMP plays key roles in synapse formation, axon guidance, and neuronal stability44,45,46,47,48, but there have been no studies on this gene in schistosomes. Although S. japonicum LSAMP shares only ~27% overall sequence similarity with its homologs in Drosophila, Caenorhabditis elegans, and Mus musculus, its functional domains are highly conserved (Supplementary Fig. 27a, b), suggesting that its biological roles may be preserved. In this study, we found that lsamp is essential for establishing the specialized neural network within the male gynecophoric canal. As a member of the IgLON family, LSAMP is a glycosylated cell adhesion molecule that functions as a GPI-anchored membrane protein44,46,47,53,60 (Supplementary Figs. 25 and 27c). In C. elegans, SAX-3 (LSAMP homolog) has been shown to localize on the surface of PVD neurons, guiding axon outgrowth47,48. Here, we observed lsamp expression in the BATT-producing N4.3 neurons in schistosomes. In the absence of lsamp, the intricate neural network connecting N4.3 neurons failed to form (Fig. 5b). Based on these observations and homology inference, we propose—though functional validation remains to be carried out—that LSAMP may serve as a selective adhesion molecule on the surface of N4.3 neurons, potentially maintaining axonal integrity and facilitating neuronal connectivity (Supplementary Fig. 27). Further experimental studies are needed to confirm its precise membrane localization and mechanistic role in schistosomes. Beyond structural organization, vesicular transport of amino acids is crucial for intracellular movement and distribution, particularly in neurotransmission61,62. Specific transport proteins facilitate the transfer of neurotransmitters into synaptic vesicles for release into the synapse62,63,64. The co-expression of ddc with nrps in N4.3 neurons—a pattern conserved in planarians59—strongly suggests that tryptamine is synthesized locally within these cells, rather than being transported through the neural network. This, in turn, implies that the primary function of the lsamp-dependent neural circuitry is likely to facilitate the delivery of the other key BATT precursor, β-alanine, rather than tryptamine. Although direct labeling of BATT precursor—β-alanine failed, our LC-MS results provided definitive evidence for the inhibition of its transport. Collectively, these findings reveal a sophisticated division of labor in the male-specific neural circuitry: while tryptamine synthesis occurs locally within N4.3 neurons, the lsamp-dependent network orchestrates the spatial delivery of β-alanine.
We observed a temporal synchronization between the formation of the male-specific neural network within the gynecophoric canal and the upregulation of lsamp expression, suggesting a potential role for lsamp in this process (Figs. 5e and 3g). We must acknowledge that the experimental evidence currently supporting this precise mechanism remains limited. While our colocalization analyses demonstrate spatial proximity between lsamp and the neural fibers in S. japonicum (Fig. 5c, d and Supplementary Fig. 24), confirming its direct function as a membrane protein in axon guidance and adhesion requires further investigation.
In conclusion, through the comprehensive single-cell transcriptomic analysis of S. japonicum, we decoded male-female interactions at single-cell resolution and delineated both conserved and sex-specific gene modules involved in gametogenesis. Furthermore, we uncovered a male-specific neural circuit that directly regulates female sexual development. Our study identified lsamp as a key regulator of axonal connectivity in pheromone-producing N4.3 neurons, which establish a specialized neural network essential for the transport of BATT precursor-β-alanine (Supplementary Fig. 28). These findings fundamentally reshape our understanding of parasite neurobiology by revealing how a primitive nervous system orchestrates reproductive processes, and they offer potential targets for disrupting the schistosome life cycle.
Methods
Ethics
All experiments related to animals were conducted in accordance with the guidelines for the Care and Use of Laboratory Animals of the Ministry of Science and Technology of the People’s Republic of China (2006398) and were approved by the Animal Care and Use Committee of Fudan University (Fudan IACUC 201802158S) to ensure ethical and responsible treatment of the animals.
S. japonicum parasites
The cercariae of S. japonicum were provided by the Chinese Center for Disease Control and Prevention. Single-sex infections of S. japonicum were obtained by infecting mice with male or female cercariae obtained from Oncomelania hupensis snails (Anhui strain) infected with single miracidia.
Animals
For all infection experiments in our study, six-week-old female C57BL/6J mice were purchased from Shanghai Jiesijie Laboratory Animal Co., Ltd (Shanghai, China). Female mice were selected to maintain consistent infection conditions and minimize stress related to aggressive behavior in social housing, which is crucial for reducing experimental variability in the schistosome-mouse model. Standard housing conditions were maintained on a 12-h light/12-h dark cycle, with ambient temperature at 22 ± 1 °C and relative humidity at 55 ± 10%. All mice were fed a standard, certified commercial rodent diet and had ad libitum access to sterile drinking water (filtered and autoclaved).
Fluorescence-activated cell sorting
Twenty female C57BL/6J mice were infected percutaneously through the abdomen with 80–200 cercariae. At each time point (14, 18, 22, and 26 dpi), five mice were sacrificed for the recovery of schistosomes10. Approximately 300 (14 dpi, days post infection), 150 (18 dpi), 100 (22 dpi), and 70 (26 dpi) male and female S. japonicum worms were collected respectively, and processed into single-cell suspensions for scRNA-seq.
Worms were collected from mice by perfusing the hepatic portal vein with 4 °C PBS (Cat# E607008, Sangon Biotech, Shanghai, China) and then rinsed twice in DMEM (5% FBS)10,55,65,66. Freshly perfused male and female worms were separated by incubation in a 0.25% solution of tricaine for approximately 5 min, except for the 14 dpi worms, which were processed as mixed-sex populations because their sex could not be distinguished morphologically under a light microscope. FACS sorting was performed as follows. Worms were washed twice in 8 mL PBS, then triturated for approximately 15 min in 4 mL 0.125% Trypsin-EDTA (in PBS) until the solution became turbid and no large pieces of worms were left21. The trypsin was inactivated by adding 8 mL 10% FBS DMEM. The dissociated worms were then centrifuged at 500 × g for 10 min at 4 °C. Next, the worm cells were resuspended in 1 mL of PBS with 10 μL of RQ1 DNAse (Cat# M6101, Promega, Madison, WI, USA) and incubated for 10 min at RT. The worm cells were centrifuged again at 500 × g for 10 min at 4 °C. Then the worm cells were resuspended in 1 mL of PBS containing Fixable Viability Dye eFluorTM 506 (1 μL/mL) (Cat# 65-0866-14, eBioscience, USA) and incubated for 30 min at 4 °C in the dark. The worm cell suspension was centrifuged at 500 × g for 10 min at 4 °C, then washed twice in PBS, and then filtered through a 40 μm cell strainer into a 12 × 75 mm FACS tube. Filtered cells were then sorted on a MoFlo XDP High-Speed Cell Sorter (BECKMAN COULTER) with 405/460/488/532/561/640 nm lasers. Sorts were performed with a 100 μm nozzle, and cells were sorted into sorting media (0.04% BSA in PBS, pH = 7.40). The sequential gating strategy was implemented using Summit Software (Beckman Coulter, v5.2) through four steps: (1) R1: initial events were gated on FSC-H versus SSC-H to identify the cell population and exclude debris; (2) R2: singlets were isolated using FSC-width versus SSC-log_height to exclude cell aggregates; (3) R3: a second doublet discrimination step was performed using SSC-width versus SSC-log_height to ensure a high-purity single-cell population; and (4) R4: viable cells were finally isolated by selecting the eFluor 506-negative population (identified via FL8-log_height versus SSC-log_height). Post-sort cell viability was further validated by trypan blue staining and manual counting under a light microscope to ensure high sample quality for downstream applications.
Single-cell RNA sequencing
FACS-sorted cells were centrifuged at 500 × g for 10 min at 4 °C and then resuspended in 0.04% BSA in PBS. Libraries were created using a Chromium Controller (10× Genomics) according to manufacturer guidelines and sequenced using a NovaSeq6000 (Illumina) to generate 150-bp paired-end reads. Raw data were processed and mapped to the S. japonicum genome (v3) using the Cell Ranger 3.0.0 (10× Genomics) pipeline.
Single-cell RNA-seq data processing
The Seurat package (v 3.1.5) was used to analyze the raw values of the matrix as follows67. Cells with greater than 30% mitochondrial reads or fewer than 500 genes, were excluded from the analysis. Mitochondrial genes were identified with the prefix “^ND|^C|^ATP”. Each of the 7 individual datasets was scaled and transformed, and variable genes were identified using the SCTransform function with the parameter “variable.features.n = 3000”. Integration anchors across samples were identified using the FindIntegrationAnchors function, and the datasets were subsequently integrated using IntegrateData from the Seurat R package. This sample-level integration approach minimizes batch effects while preserving biological variation across samples. Then we performed principal component analysis using variable genes and used the first 100 principal components (PCs) to perform UMAP to embed the dataset into two dimensions. For clustering, we identified 76 clusters using the FindNeighbors and FindClusters functions with the parameter “dims = 1:100, resolution = 2”.
To determine cellular identity, we used the function FindAllMarkers with the parameter “test.use = “wilcox”, only.pos = T, min.pct = 0.25, logfc.threshold = 0.25” to find DE genes for each cluster and compared to the known marker genes from the previous datasets18,19,21,68. GO (Gene Ontology) enrichment analysis was performed with the R package clusterProfiler69. The p-value was corrected for multiple hypothesis testing with the Benjamini–Hochberg false-discovery rate procedure (adjusted p-value). GO terms with corrected p-value < 0.05 were considered significant. R packages ggplot2 (https://ggplot2.tidyverse.org) and tidyverse (https://www.tidyverse.org) were used to generate figures and intermediate data preprocessing.
The density plots of cells from each sample over UMAP embedding were generated using a modified treecor_celldensityplot function from TreeCorTreat package (https://github.com/byzhang23/TreeCorTreat).
Differential abundance analysis
We tested for differences in cell-type abundances between female and male worms using the miloR package (v 1.4.0)70, which is a method for differential abundance analysis on KNN graphs from single-cell datasets. Briefly, we computed the k-nearest neighbor graph of cells in the integrated datasets based on latent embedding. Subsequently, cell neighborhoods were computed using miloR’s makeNhoods function with parameter “k = 25, prop = 0.2”. To leverage the variation in the number of cells between 18, 22, and 26 dpi samples, we counted the cells belonging to each sample in each neighborhood using the countCells function. We assigned to each neighborhood a cell-type label based on the majority voting of the cells belonging to that neighborhood. We then used the testNhoods function to test for enrichment of cells from the different sex datasets. Neighborhoods were enriched if the spatial FDR < 0.2 and |log2FC| > 1.
Subclustering analysis
To investigate cellular heterogeneity within reproductive system-associated and neuronal populations, we performed subclustering analysis using the Seurat package (v3.1.5)67. Reproductive system-associated cells (including neoblasts, germ cells, vitellaria, and Mehlis’ gland cells) and neuronal cells (including neural progenitor cells, neural precursor cells, nonciliated neurons, and ciliated neurons) were subset using the SubsetData function. Principal component analysis, dimensionality reduction, and clustering were then performed separately on each subset. We used 20 PCs and a resolution of 3 for reproductive system-associated cells, and 30 PCs with a resolution of 1 for neuronal cells, based on the variance explained in the Principal Component Analysis (PCA) elbow plots and the robustness of clustering results. Subclusters were annotated based on lineage-specific transcriptional signatures and established marker genes.
Trajectory analyses
To infer potential developmental trajectories within the reproductive-related or neuronal lineages, trajectory inference analysis was performed using the R package Slingshot (v 2.0.1)30. The vitelline, germline, and neuronal cell lineages identified in the integrated dataset were subsetted, and recomputed a UMAP embedding. Slingshot was then applied with default parameters, using neural progenitor cells, S1-0 cells, or GSCs as the starting clusters to infer cell trajectories and calculate pseudotime. The resulting trajectories were mapped onto the UMAP projection for visualization. To evaluate how pseudotime inferred from Slingshot corresponded to the actual sample collection time points, pseudotime distributions across samples were visualized using density plots.
In addition, to identify genes dynamically changing along vitelline and germline pseudotime trajectories, tradeSeq34 analysis was carried out on the Slingshot-derived trajectories. For each dataset, the “associationTest” function was used to identify genes whose expression patterns were associated with pseudotime, with a knots value of 6 for vitelline and 7 for germline lineages. A p-value cutoff of 1e−5 was applied to select significantly changing genes, and a more stringent cutoff (1e−10) was used to define a refined set of dynamic genes in both female and male datasets along the germline lineages. Heatmaps of these genes were generated using the R package ClusterGVis (v0.1.4) (https://github.com/junjunlab/ClusterGVis). To determine the optimal number of gene expression clusters, the “getClusters” function in ClusterGVis was applied, which implements the Elbow method to evaluate within-cluster sum of squares across a range of cluster numbers.
RNA velocity and cell rank analysis
To further characterize differentiation trajectories leading to distinct cell populations, we performed RNA Velocity analysis. In brief, we generated loom files of the spliced and unspliced RNA matrices using velocyto (v 0.17.17)30 with our CellRanger output. Then we applied scVelo (v 0.2.2) to estimate transient cell states and RNA velocities using a generalized dynamical model following the recommended workflow. Additionally, diffusion pseudotime71 was computed using scVelo to provide an alternative measure of cellular progression along developmental trajectories based on transcriptomic similarity and RNA velocity.
To further investigate cell fate decision, we used CellRank package (v 1.0.0)38 as described in (https://cellrank.readthedocs.io/en/stable/index.html) to identify initial and terminal states and characterize cell fate probabilities. Specifically, to infer putative driver genes for any of these trajectories, we correlate expression values with fate probabilities using the “cellrank.tl.lineage_drivers” function with the parameter “method = Fischer”.
CytoTRACE analysis
To evaluate the differentiation potential of cells within the vitelline lineages, we applied CytoTRACE33 with default parameters. The UMAP embedding was used to visualize CytoTRACE scores across cell clusters. A higher CytoTRACE score indicates greater stemness and a higher potential for differentiation into mature cell states.
RNA interference (RNAi)
Oligo sequences used to generate dsRNA templates and qPCR primers are listed in Supplementary Data 26. PCR primer synthesis and DNA sequencing were performed by Beijing Tsingke Biotech Co., Ltd. (Tianjin, China). Double-stranded RNA (dsRNA) was synthesized using the T7 High Yield RNA Synthesis Kit (Cat# 10623ES60, Yeasen, Shanghai, China) following the manufacturer’s protocols. Briefly, DNA templates for in vitro transcription were PCR-amplified from S. japonicum cDNA using gene-specific primers flanked with T7 promoter sequences (5’-TAATACGACTCACTATAGGGAGA-3’) at the 5’ ends. The PCR products were purified using a TIANgel Purification Kit (Cat# DP219, TIANGEN, Beijing, China) and utilized as templates for the transcription reaction. The mixture was incubated at 37 °C for at least 4 h. The synthesized dsRNA was subsequently purified, resuspended in nuclease-free water, and quantified using a NanoDrop ND-2000c spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). The integrity and quality of the dsRNA were verified by 1% agarose gel electrophoresis. The dsRNA of the gfp (green fluorescent protein) fragment was used as a negative control for all RNAi experiments55. Day 0 represents the first day of the experiment. Males were treated with 30 μg/mL dsRNA (on Day 0/2/4/6 with fresh medium) for one week in BM169 and then paired with virgin female parasites from Day 8 to Day 27 in ABC169 (fresh medium was replaced every other day). The eggs produced by the worms are retained when changing the medium. On Day 28, the egg number and the number of female parasites were counted to calculate egg/female/day.
Bulk RNA-sequencing and data analysis
We collected male worms from the gfp RNAi and lsamp RNAi groups on Day 20 for bulk RNA sequencing, with four biological replicates per group. Total RNA extraction, library construction, and RNA sequencing were performed by the Novogene Co., Ltd (Beijing, China). Briefly, RNA was extracted from worms using the AG RNAex Pro Reagent (Cat#AG21101, Accurate Biotechnology (Hunan) Co., Ltd., Changsha, China). The integrity and quality of total RNA were assessed using a Nanodrop ND-2000c spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA) and an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA). Pooled RNA was used to construct a library using Illumina TruSeq™ RNA sample prep Kit v2 (Illumina, USA) and the Ribo-Zero Plus rRNA Depletion Kit (Illumina, USA) for rRNA depletion, following the TruSeq RNA Sample Preparation Guide. The library was subsequently sequenced on the Illumina NovaSeq 6000 platforms to obtain 150 bp paired-end (PE) reads.
QC (Quality control) of the raw sequencing data was performed using the FASTQC program (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/). Low-quality reads and adapter sequences were trimmed using the Fastp tool (v 0.20.1) (parameters: -q 15 -u 40 -n 5 -l 15)72. The clean reads were mapped to the chromosome-level S. japonicum reference genome (SjV3) using HISAT2 (v 2.1.0) with default parameters73. The gene-level assignment was then performed to estimate the transcript abundance using featureCounts (1.6.4)74. PCA was performed using the prcomp function in the R stats package (v 3.6.0). Hierarchical clustering analysis was performed with pheatmap (https://cran.r-project.org/web/packages/pheatmap/index.html). Differential expression analysis was performed using DESeq275, and the differentially expressed genes (DEGs) were defined with adjusted p-value < 0.05 and |log2 FC| > 2. To identify the possible functions and pathways of DEGs, GO terms enrichment analysis and Gene Set Enrichment Analysis were performed using the clusterProfiler package69.
Parasite labeling and imaging
Colorimetric and fluorescence in situ hybridization were performed as follows. Riboprobes were synthesized using the Fluorescein RNA Labeling Mix (Cat# 11685619910, Sigma-Aldrich, St. Louis, MO, USA) or DIG RNA Labeling Mix (Cat# 11277073910, Sigma-Aldrich, St. Louis, MO, USA), with templates generated using primers listed in Supplementary Data 4. Typically, all probes were applied at a concentration of 100 ng/mL in the hybridization buffer, with the concentrations of certain probes adjusted to optimize localization performance. For WISH, anti-digoxigenin-AP (1:2000, Cat# 11093274910, Lot# 84080920, Sigma-Aldrich, St. Louis, MO, USA) was used, followed by color development with NBT/BCIP (Cat# 11681451001, Sigma-Aldrich, St. Louis, MO, USA). For FISH, anti-digoxigenin-POD (1:1000, Sigma-Aldrich, Cat# 11207733910, Lot# 77798000, St. Louis, MO, USA) or anti-fluorescein-POD (1:1000, Sigma-Aldrich, Cat# 11426346910, Lot# 57545020, St. Louis, MO, USA) were used. Signal amplification was performed using FITC-tyramide (Cat# B40953, Thermo Fisher Scientific, Waltham, MA, USA) or TAMRA-tyramide (Cat# B40957, Thermo Fisher Scientific, Waltham, MA, USA)23,66.
The immunofluorescence assays for detecting Synapsin levels in the parasite were performed using mouse monoclonal anti-Synapsin (1:100, Clone 3C11, Cat# 3C11, Lot# 10/1/21, Developmental Studies Hybridoma Bank, DSHB, Iowa City, IA, USA), and for detecting acetylated α-tubulin levels using mouse monoclonal anti-acetylated α-tubulin (1:500, Clone 6-11B-1, Cat# sc-23950, Lot# G2522, Santa Cruz, Dallas, Texas, USA)76,77. Subsequently, samples were incubated with Goat Anti-Mouse IgG H&L (Alexa Fluor® 488) secondary antibody (1:1000, Cat# ab150113, Lot# 1004120-39, Abcam, Cambridge, UK) or Goat Anti-Mouse IgG H&L (Alexa Fluor® 594) secondary antibody (1:1000, Cat# ab150116, Lot# GR3402482-1, Abcam, Cambridge, UK) for 1 h at room temperature.
For females collected on Day 28, vitellaria were labeled with 1% Fast Blue BB (Cat# F3378, Sigma-Aldrich, St. Louis, MO, USA) in PBSTx for 1 min at room temperature24,55.
All labeled parasites were counterstained with DAPI (1 μg/mL), cleared in 80% glycerol, and mounted on slides with 80% glycerol. Confocal imaging of fluorescently labeled samples was performed on Nikon A1 Laser Scanning Confocal Microscope or Olympus FV3000 Laser Scanning Confocal Microscope. Brightfield images were acquired on Olympus BX51 Microscope and Zeiss AxioZoom V16 Microscope.
qRT-PCR (quantitative real-time PCR)
All qPCR reactions were performed on a LightCycler® 96 Instrument (Roche, Switzerland) using 2× SYBR green qPCR master mix (Cat# 11201ES50, Yeasen, Shanghai, China) according to the manufacturer’s instructions. Each 20 μL qRT-PCR reaction mixture comprised 2 μL of cDNA(1:4), 10 μL 2× SYBR green master, 0.8 μL (5 μM) of each primer, and 6.4 μL ddH2O. The qRT-PCR cycle parameters were as follows: 95 °C for 3 min, followed by 40 cycles of 95 °C for 15 s, 60 °C for 30 s; melt curve analysis ranged from 60 °C to 95 °C to ensure that the specific product was amplified in each reaction. The 2−ΔΔCt method was used to calculate the relative fold change of the differentially expressed transcripts.
BATT synthesis and detection
The synthesis of BATT was conducted by the Chinese Center for Disease Control and Prevention. Concentrated stock solutions (100 mM) of BATT were prepared in sterile DMSO and stored at −20 °C. LC-MS/MS was employed for the detection of BATT in parasites (thirty male worms) and medium samples collected on Day 20. BATT detection was performed on a triple quadrupole mass spectrometer operating in multiple reaction monitoring employing negative ESI interface using QTRAP® 6500+ LC-MS/MS System (Sciex, Framingham, MA, USA), with the transitions set at m/z 232.4 → m/z 143.7, m/z 232.4 → m/z 185.1, m/z 232.4 → m/z 89.1, respectively9,55.
Quantification and statistical analysis
Quantification and statistical analysis were performed using GraphPad Prism software (v9.0, San Diego, CA, USA). The normality of data sets with an adequate sample size (n ≥ 5) was assessed using the D’Agostino-Pearson test (Supplementary Data 27). For data conforming to a normal distribution or with a sample size (n < 5), unpaired two-tailed Student’s t-tests were applied, and the results are presented as the mean ± SEM (Standard Error of the Mean). For data that did not pass the normality test, the non-parametric Mann–Whitney U test was applied (Supplementary Data 28). Statistical significance was defined as *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001; ns, not significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The scRNA-seq data and the bulk RNA-seq raw data for S. japonicum in this study can be accessed from NCBI Sequence Read Archive (SRA) using the accession number PRJNA1244463, The gene expression data for scRNA-seq have been deposited at the Gene Expression Omnibus (GEO) with the accession number GSE293642. Source data for the graphs within the Figures have been provided in Source Data.xlsx. Source data are provided with this paper.
References
McManus, D. P. et al. Schistosomiasis. Nat. Rev. Dis. Prim. 4, 13 (2018).
Molehin, A. J. Schistosomiasis vaccine development: update on human clinical trials. J. Biomed. Sci. 27, 28 (2020).
Bergquist, R., Utzinger, J. & Keiser, J. Controlling schistosomiasis with praziquantel: how much longer without a viable alternative? Infect. Dis. Poverty 6, 74 (2017).
Moore, D. V. & Sandground, J. H. The relative egg producing capacity of Schistosoma mansoni and Schistosoma japonicum. Am. J. Trop. Med. Hyg. 5, 831–840 (1956).
Schwartz, C. & Fallon, P. G. Schistosoma “eggs-iting” the host: granuloma formation and egg excretion. Front. Immunol. 9, 2492 (2018).
Popiel, I. & Basch, P. F. Reproductive development of female Schistosoma mansoni (Digenea: Schistosomatidae) following bisexual pairing of worms and worm segments. J. Exp. Zool. 232, 141–150 (1984).
Armstrong, J. C. Mating behavior and development of schistosomes in the mouse. J. Parasitol. 51, 605–616 (1965).
AE, S. Sex studies on Schistosoma japonicum. Q. J. Microsc. Sci. s2, 653 (1928).
Chen, R. et al. A male-derived nonribosomal peptide pheromone controls female schistosome development. Cell 185, 1506–1520.e1517 (2022).
Wang, J. et al. Dynamic transcriptomes identify biogenic amines and insect-like hormonal regulation for mediating reproduction in Schistosoma japonicum. Nat. Commun. 8, 14693 (2017).
Waisberg, M. et al. Microarray analysis of gene expression induced by sexual contact in Schistosoma mansoni. BMC Genomics 8, 181 (2007).
Sun, J. et al. Transcriptome profilings of female Schistosoma japonicum reveal significant differential expression of genes after pairing. Parasitol. Res. 113, 881–892 (2014).
Gobert, G. N., Moertel, L., Brindley, P. J. & McManus, D. P. Developmental gene expression profiles of the human pathogen Schistosoma japonicum. BMC Genomics 10, 128 (2009).
Fitzpatrick, J. M., Johansen, M. V., Johnston, D. A., Dunne, D. W. & Hoffmann, K. F. Gender-associated gene expression in two related strains of Schistosoma japonicum. Mol. Biochem. Parasitol. 136, 191–209 (2004).
Piao, X. et al. Global expression analysis revealed novel gender-specific gene expression features in the blood fluke parasite Schistosoma japonicum. PLoS ONE 6, e18267 (2011).
Hoffmann, K. F., Johnston, D. A. & Dunne, D. W. Identification of Schistosoma mansoni gender-associated gene transcripts by cDNA microarray profiling. Genome Biol. 3, RESEARCH0041 (2002).
Anderson, L. et al. Schistosoma mansoni egg, adult male and female comparative gene expression analysis and identification of novel genes by RNA-seq. PLoS Negl. Trop. Dis. 9, e0004334 (2015).
Wang, B. et al. Stem cell heterogeneity drives the parasitic life cycle of Schistosoma mansoni. eLife 7, e35449 (2018).
Diaz Soria, C. L. et al. Single-cell atlas of the first intra-mammalian developmental stage of the human parasite Schistosoma mansoni. Nat. Commun. 11, 6411 (2020).
Diaz Soria, C. L. et al. Single-cell transcriptomics of the human parasite Schistosoma mansoni first intra-molluscan stage reveals tentative tegumental and stem-cell regulators. Sci. Rep. 14, 5974 (2024).
Wendt, G. et al. A single-cell RNA-seq atlas of Schistosoma mansoni identifies a key regulator of blood feeding. Science 369, 1644–1649 (2020).
Nanes Sarfati, D., Li, P., Tarashansky, A. J. & Wang, B. Single-cell deconstruction of stem-cell-driven schistosome development. Trends Parasitol. 37, 790–802 (2021).
Collins, J. J. et al. Adult somatic stem cells in the human parasite Schistosoma mansoni. Nature 494, 476–479 (2013).
Wang, J., Chen, R. & Collins, J. J. Systematically improved in vitro culture conditions reveal new insights into the reproductive biology of the human parasite Schistosoma mansoni. PLoS Biol. 17, e3000254 (2019).
Wang, J. & Collins, J. J. 3rd Identification of new markers for the Schistosoma mansoni vitelline lineage. Int J. Parasitol. 46, 405–410 (2016).
Henkle, K. J. et al. The gene family encoding eggshell proteins of Schistosoma japonicum. Mol. Biochem. Parasitol. 42, 69–82 (1990).
Vogel, H. Hermaphrodites of Schistosoma mansoni. Ann. Trop. Med. Parasitol. 41, 266–277 (1947).
Hulstijn, M. et al. Hermaphrodites and supernumerary testicular lobes in Schistosoma mansoni (Trematoda: Schistosomatidae) analyzed by brightfield and confocal microscopy. J. Parasitol. 92, 496–500 (2006).
Loverde, P. T. Schistosomes: development, reproduction, and host relations. Paul F. Basch. Q. Rev. Biol. 67, 533–533 (1992).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Shariati, F. et al. Evaluation of the role of angiogenic factors in the pathogenesis of schistosomiasis. Exp. Parasitol. 128, 44–49 (2011).
Li, P. et al. Single-cell analysis of Schistosoma mansoni identifies a conserved genetic program controlling germline stem cell fate. Nat. Commun. 12, 485 (2021).
Gulati, G. S. et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science 367, 405–411 (2020).
Van Den Berge, K. et al. Trajectory-based differential expression analysis for single-cell sequencing data. Nat. Commun. 11, 1201 (2020).
Hanumantha-Rao, K. Histochemistry of Mehlis’ gland and egg-shell formation in the liver fluke Fasciola hepatica L. Experientia 15, 464–465 (1959).
Moczon, T., Swiderski, Z. & Huggel, H. Schistosoma mansoni: the chemical nature of the secretions produced by the Mehlis’ gland and ootype as revealed by cytochemical studies. Int. J. Parasitol. 22, 65–73 (1992).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Lange, M. et al. CellRank for directed single-cell fate mapping. Nat. Methods 19, 159–170 (2022).
Naren, P. et al. Microtubule acetylation dyshomeostasis in Parkinson’s disease. Transl. Neurodegener. 12, 20 (2023).
Deshpande, P. et al. N-Acetyltransferase 9 ameliorates Aβ42-mediated neurodegeneration in the Drosophila eye. Cell Death Dis. 14, 478 (2023).
Mondal, P. et al. Structure-based discovery of A small molecule inhibitor of histone deacetylase 6 (HDAC6) that significantly reduces Alzheimer’s disease neuropathology. Adv. Sci. 11, e2304545 (2024).
Mazzetti, S. et al. Linking acetylated α-Tubulin redistribution to α-Synuclein pathology in brain of Parkinson’s disease patients. npj Parkinson’s. Dis. 10, 2 (2024).
Esteves, A. R. et al. Acetylation as a major determinant to microtubule-dependent autophagy: relevance to Alzheimer’s and Parkinson disease pathology. Biochim. Biophys. Acta Mol. Basis Dis. 1865, 2008–2023 (2019).
Singh, K. et al. The combined impact of IgLON family proteins Lsamp and Neurotrimin on developing neurons and behavioral profiles in mouse. Brain Res. Bull. 140, 5–18 (2018).
Bregin, A. et al. Increased sensitivity to psychostimulants and GABAergic drugs in Lsamp-deficient mice. Pharmacol. Biochem. Behav. 183, 87–97 (2019).
Bregin, A. et al. Expression and impact of Lsamp neural adhesion molecule in the serotonergic neurotransmission system. Pharmacol. Biochem. Behav. 198, 173017 (2020).
Um, J. W. & Ko, J. Neural glycosylphosphatidylinositol-anchored proteins in synaptic specification. Trends Cell Biol. 27, 931–945 (2017).
Chen, C.-H., Hsu, H.-W., Chang, Y.-H. & Pan, C.-L. Adhesive L1CAM-robo signaling aligns growth cone F-actin dynamics to promote axon-dendrite fasciculation in C. elegans. Dev. Cell 48, 215–228.e215 (2019).
Matamoros, A. J. & Baas, P. W. Microtubules in health and degenerative disease of the nervous system. Brain Res. Bull. 126, 217–225 (2016).
Fykse, E. M. & Fonnum, F. Amino acid neurotransmission: dynamics of vesicular uptake. Neurochem. Res. 21, 1053–1060 (1996).
Van Onselen, R. & Downing, T. G. Uptake of β-N-methylamino-L-alanine (BMAA) into glutamate-specific synaptic vesicles: exploring the validity of the excitotoxicity mechanism of BMAA. Neurosci. Lett. 821, 137593 (2024).
Roseth, S. & Fonnum, F. A study of the uptake of glutamate, γ-aminobutyric acid (GABA), glycine and β-alanine in synaptic brain vesicles from fish and avians. Neurosci. Lett. 183, 62–66 (1995).
Pimenta, A. F. et al. The limbic system-associated membrane protein is an Ig superfamily member that mediates selective neuronal growth and axon targeting. Neuron 15, 287–297 (1995).
Cheng, S. et al. Family of neural wiring receptors in bilaterians defined by phylogenetic, biochemical, and structural evidence. Proc. Natl. Acad. Sci. USA 116, 9837–9842 (2019).
You, Y. et al. An improved medium for in vitro studies of female reproduction and oviposition in Schistosoma japonicum. Parasit. Vectors 17, 116 (2024).
Li, X., Weth, O., Haeberlein, S. & Grevelding, C. G. Molecular characterization of Smtdc-1 and Smddc-1 discloses roles as male-competence factors for the sexual maturation of Schistosoma mansoni females. Front. Cell. Infect. Microbiol. 13, 1173557 (2023).
Nikishin, D. A., Alyoshina, N. M., Semenova, M. L. & Shmukler, Y. B. Analysis of expression and functional activity of aromatic L-amino acid decarboxylase (DDC) and serotonin transporter (SERT) as potential sources of serotonin in mouse ovary. Int. J. Mol. Sci. 20, 3070 (2019).
Khan, U. W. & Newmark, P. A. Somatic regulation of female germ cell regeneration and development in planarians. Cell Rep. 38, 110525 (2022).
Issigonis, M., Browder, K. L., Chen, R., Collins, J. J. & Newmark, P. A. A niche-derived nonribosomal peptide triggers planarian sexual development. Proc. Natl. Acad. Sci. USA 121, e2321349121 (2024).
Smith, C. J. et al. Time-lapse imaging and cell-specific expression profiling reveal dynamic branching and molecular determinants of a multi-dendritic nociceptor in C. elegans. Dev. Biol. 345, 18–33 (2010).
Martineau, M. et al. Storage and uptake of D-serine into astrocytic synaptic-like vesicles specify gliotransmission. J. Neurosci. 33, 3413–3423 (2013).
Li, D. et al. Astrocyte VAMP3 vesicles undergo Ca2+ -independent cycling and modulate glutamate transporter trafficking. J. Physiol. 593, 2807–2832 (2015).
Kumar, R. et al. Fibroblast growth factor 2-mediated regulation of neuronal exosome release depends on VAMP3/cellubrevin in hippocampal neurons. Adv. Sci. 7, 1902372 (2020).
Sneeggen, M. et al. WDFY2 restrains matrix metalloproteinase secretion and cell invasion by controlling VAMP3-dependent recycling. Nat. Commun. 10, 2850 (2019).
Luo, F. et al. A chromosome-level genome of the human blood fluke Schistosoma japonicum identifies the genomic basis of host-switching. Cell Rep. 39, 110638 (2022).
Cheng, S. et al. Dynamic profiles of lncRNAs reveal a functional natural antisense RNA that regulates the development of Schistosoma japonicum. PLoS Pathog. 20, e1011949 (2024).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Xia, J. et al. A single-cell resolution developmental atlas of hematopoietic stem and progenitor cell expansion in zebrafish. Proc. Natl. Acad. Sci. USA 118, e2015748118 (2021).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Dann, E., Henderson, N. C., Teichmann, S. A., Morgan, M. D. & Marioni, J. C. Differential abundance testing on single-cell data using k-nearest neighbor graphs. Nat. Biotechnol. 40, 245–253 (2021).
Haghverdi, L., Büttner, M., Wolf, F. A., Buettner, F. & Theis, F. J. Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 13, 845–848 (2016).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Collins, J. J., King, R. S., Cogswell, A., Williams, D. L. & Newmark, P. A. An atlas for Schistosoma mansoni organs a nd life-cycle stages using cell type-specific markers and confocal microscopy. PLoS Negl. Trop. Dis. 5, e1009 (2011).
Creppe, C. et al. Elongator controls the migration and differentiation of cortical neurons through acetylation of α-tubulin. Cell 136, 551–564 (2009).
Acknowledgements
We would like to express our gratitude to the following individuals and institutions for their support and assistance: Bin Xu, Ying Wu, and Yang Hong from the National Institute of Parasitic Diseases, Chinese Center for Disease Control and Prevention, for their support and assistance with experimental materials. The amino metabolome and BATT were quantified by Professor Huiru Tang’s group as a paid service. Jingwei Zhang of Fudan University for his guidance and assistance in single-cell sample evaluation. This research was supported by National Natural Science Foundation of China (No. 31725025, and No. 31972699 to W.H.), the National Key Research and Development Program of China (No. 2021YFC2300800, and No. 2021YFC2300803 to J.P.W.), Science and Technology Leading Talent Team in Inner Mongolia Autonomous Region (No. 2022LJRC0009 to W.H.), Shanghai Natural Science Foundation Young Scientists Program (No. 25ZR1402030 to Y.M.Y.), National Natural Science Foundation of China Regional Innovation and Development Joint Fund (No. U22A20526 to W.H.), and the Young Scientists Fund of the National Natural Science Foundation of China (No. 32202831 to F.L).
Author information
Authors and Affiliations
Contributions
W.H. and Y.M.Y. conceptualized this project. W.H. and J.P.W. supervised the overall research and directed the bioinformatics analysis. Y.M.Y. performed all experiments, and S.Y.C. conducted the bioinformatics analysis of scRNA-seq and RNA-seq data. Y.M.Y. and J.P.W. drafted the manuscript, with Y.M.Y., J.P.W., S.Y.C., and W.H. participating in manuscript revision. Xu.C., Xi.C., C.Y., and M.Q.C. provided experimental assistance in WISH and FISH. F.L., X.L.W., Y.X.X., W.B.Y., L.X., and M.J.G., assisted in sample processing for scRNA-seq. C.S.S. provided negative Oncomelania hupensis snails. L.L.H. contributed to BATT synthesis. G.W.C. and S.L. provided assistance in sample collection. All authors critically reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Matthew Berriman who co reviewed with Frances Blow, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
You, Y., Cheng, S., Chen, X. et al. Dynamic single-cell transcriptomics reveals lsamp-guided neural network formation in male S. japonicum driving female reproduction. Nat Commun 17, 1602 (2026). https://doi.org/10.1038/s41467-026-68305-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68305-7