Abstract
Sex determination systems display striking evolutionary flexibility, yet the mechanisms underlying their transitions remain poorly understood. Using newly generated genome assemblies, we investigated the evolving sex-determining system in the amphioxus Branchiostoma belcheri. We identified two female-specific sex-determining regions (SDRs) on chromosome 13, both derived from independent transpositions of the autosomal gene tesD, which shows testis-specific expression in amphioxus species. CRISPR/Cas9 knockout experiments in Branchiostoma floridae confirmed that tesD functions as a male-determination gene, with loss of function producing an all-female phenotype. In B. belcheri, the older SDR (tesDwa) inserted into the coding region of twai, while the younger SDR (tesDwb), flanked by active Zator-1 transposons, inserted into the 3′ UTR of vps9c and later translocated to autosomes in ~10% of individuals. Transcriptomic analyses revealed that W-linked tesDwa and tesDwb produce antisense long non-coding RNAs that likely suppress tesD transcription in females, whereas autosomal tesDwb is not expressed and appears non-functional. The insertion sites and co-transcription with host genes suggest promoter hijacking. Together, these findings demonstrate that recurrent transpositions can generate new functional SDRs that coexist with older ones, driving dynamic turnover of sex determination in B. belcheri.
Similar content being viewed by others
Introduction
The mechanisms governing sex determination are essential for establishing gonochorism—the separation of males and females across species. While mammals and birds rely on heteromorphic sex chromosomes (e.g., XY in humans), most animals and plants use homomorphic chromosomes or environmental cues1,2,3. Recent genomic studies highlight the rapid evolution and frequent turnover of sex-determining systems and genes in species with homomorphic chromosomes4,5,6,7,8. However, the genetic and evolutionary forces driving these changes remain poorly understood and under debate7,9,10,11,12. Emerging discoveries of novel sex determination pathways, including the development of new sex chromosomes, present valuable opportunities to explore these mechanisms13,14,15,16. Broadening research to encompass a wider range of species, particularly those at key phylogenetic nodes, could offer critical insights into the prevalence of homomorphic sex chromosomes across diverse lineages, shedding light on the broader evolutionary dynamics of sex determination17,18.
Amphioxus has emerged as an important experimental model due to its pivotal phylogenetic position, linking invertebrates and vertebrates19,20,21,22. Previous studies have identified a female heterogametic system (ZW) in three species of Branchiostoma amphioxus: B. floridae (Bf), B. japonicum (Bj), and B. belcheri (Bb)23,24. The sex chromosomes in all three species are homomorphic. Despite this shared characteristic, the sex determination regions (SDRs) exhibit interspecific variation, locating on different chromosomes23. Comparative analysis of these species may offer a valuable opportunity to investigate the evolutionary dynamics and turnover of sex determination mechanisms. It may also provide clues on whether the key determinants or the “usual suspects” of sex in vertebrates had already emerged in chordate ancestors5,25,26.
In this study, we investigate the sex-determination system and its evolution in Bb using newly assembled genomes and sex chromosomes. We report that Bb possesses a polygenic sex-determination system, characterized by two variants of SDRs located at separate loci on chromosome 13. Both variants contain a duplicate of an autosomal gene, tesD, which was probably transposed via transposable elements (TEs) based on sequence structure analysis. Our knockout experiments in Bf demonstrate that tesD plays a critical and likely specific role in testis development in this species. Given the high sequence conservation and testis-specific expression of tesD across three Branchiostoma species, its role in male determination is likely conserved within this genus. In Bb, the two tesD duplicates (tesDwa and tesDwb) on the W chromosome are transcribed as unspliced long non-coding RNAs (lncRNAs) in a reverse orientation relative to tesD. These lncRNAs likely mediate female determination by repressing the transcription of autosomal tesD. Our analysis further indicates that tesDwb evolved more recently yet coexists with tesDwa in roughly half of the female population. Remarkably, tesDwb resides in an SDR that remains actively mobile due to TE activity and is found on different autosomes in approximately 10% of the population for both males and females. The transposed tesDwb copy and its associated SDR on autosomes are not transcribed and have likely lost their function. This highlights how TEs can drive the turnover of sex-determination systems and explains the persistence of homomorphic sex chromosomes in certain species.
Results
Characterization of two distinct SDRs in Bb
Previous research on Bb was limited by the assembly of a single haplotype (Z chromosome)23. To overcome this, we first sequenced a female Bb and performed haplotype phasing, followed by scaffolding of each haplotype (“Methods”, Supplementary Fig. 1a, b and Supplementary Table 1). This genome was designated Bb-a, and the two resulting haplotypes Bb-a-hap1 and Bb-a-hap2. Bb-a-hap1 and Bb-a-hap2 exhibited high similarity across all chromosomes, with only minor differences, such as a few short inversions (Supplementary Fig. 1c). While chromosome 3 was previously proposed as the putative sex chromosome, with tesD as the sole candidate sex-determining gene23, our analysis revealed complexities. Annotation and alignment of the two tesD loci on chromosome 3 showed 100% sequence identity in their coding regions (Supplementary Fig. 2). Furthermore, none of the four intronic single nucleotide polymorphisms (SNPs) within tesD displayed sex-specific association with prior genome-wide association studies (GWAS)23, suggesting that the two tesD alleles are not differentiated between the two haplotypes. We then conducted GWAS using previously published resequencing data23 and our newly assembled haplotypes as reference genomes (“Methods”, Supplementary Table 2). Using Bb-a-hap1, we recapitulated the identification of significant sex-linked SNPs at the tesD locus on chromosome 3, consistent with previous findings (Fig. 1A and Supplementary Fig. 3a). Surprisingly, both Bb-a-hap1 and Bb-a-hap2 analyses identified significant sex-linked SNPs at the pon2-like locus on chromosome 13 (Fig. 1A, B and Supplementary Fig. 3b, c), which were not found to be significant in the previous study23. A single significant SNP on chromosome 6 was identified; however, it was not fully associated with either male or female samples and was subsequently excluded from further analysis (Supplementary Fig. 3d, e). In addition, we applied the same GWAS pipeline using the reference assembly from the previous study23. This analysis similarly identified SNPs at the pon2-like locus (Supplementary Fig. 3f).
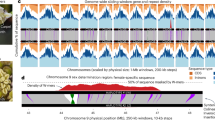
A GWAS analysis identified significant sex-linked SNPs on chromosomes Chr03, Chr06, and Chr13 using Bb-a-hap1 as the reference genome. The Y-axis displays the −log10 transformed p-values obtained in the GWAS. The p-values for the association of each SNP were calculated using two-sided p-values derived from the Wald Test implemented in EMMAX (“Methods”), with the test statistic assumed to follow a chi-square distribution with one degree of freedom. Adjustments for multiple comparisons were applied using a Bonferroni-type correction. The genome-wide significance threshold (dashed line) was set at p < 1/n, where n is the effective number of independent SNPs. P-values were calculated consistently across GWAS analyses performed in this figure (B) and Fig. 2A, B. B GWAS analysis identified significant sex-linked SNPs on Chr06 and Chr13 using Bb-a-hap2 as the reference genome. C Self-alignment dotplot of the region flanking mmp7-like and pon2-like on Bb-a-hap1-chr13. Blue dashed line boxes highlight multiple copies of an exon used as a marker to identify the SDR insertion site in Bb-a-hap2. The red arrow indicates the insertion site, as indicated by the red arrow in (F). D Alignment of the genomic regions flanking mmp7-like and pon2-like between the two haplotypes, showing the insertion of Bb-a-SDR (pink dashed line) into the labeled twai exon. E Annotation reveals that Bb-a-SDR contains a duplicate (tesDwa) of the autosomal tesD gene and three TEs. F Schematic representation of the Bb-a-SDR insertion into a twai exon, with the loss of the 3′ regions of twai in Bb-b-hap2. Source data are provided as a Source Data file.
Subsequent genome annotation and manual inspection revealed an additional copy of tesD (tesDwa, tesD on type a W chromosome) located 3′ of pon2-like on Bb-a-hap2-chr13. Alignment of tesDwa with an autosomal tesD showed an 11-bp deletion in the first exon of tesDwa, likely disrupting its open reading frame (Supplementary Fig. 4). Notably, many of the SNPs identified within this alignment were also found to be sex-linked in the GWAS analysis (indicated by red dots in Supplementary Fig. 4). This pattern strongly suggested that tesDwa functions as a female-specific genetic marker and confirmed that the corresponding chromosome 13 is the W chromosome.
The tesDwa is located between pon2-like and mmp7-like in Bb-a-hap2-chr13, a conserved collinearity across the two haplotypes. To determine whether this region harbors additional sex-specific differences, we conducted a comparative alignment of the homologous segments spanning pon2-like and mmp7-like of the two haplotypes. This analysis identified a potential female-specific SDR inserted into the W chromosome (Fig. 1C, D). The SDR is characterized by the presence of tesDwa flanked at its 3′ end by TEs (Fig. 1E, F). The SDR insertion site lies within an exon of a gene, designated twai (tesDw type a inserted), located between pon2-like and mmp7-like. Notably, the copy number of twai (and the corresponding insertion exon) varies allelically, ranging from a single copy in the Bb-b-hap2 haplotype (refer to subsequent text for details on the Bb-b assembly) to five copies in the Bb-PNAS haplotype (Bb assembly from Huang et al.23) (Fig. 1C, D and Supplementary Fig. 5a). The absence of additional degraded twai sequences at this locus on the Bb-a-hap2-chr13 suggested that the Bb-a-SDR likely inserted into an allele originally harboring a single twai copy, such as in Bb-b-hap2. Further analysis of this locus in the Bf genome revealed conserved collinearity of pon2-like and mmp7-like, along with six tandem twai duplications (Supplementary Fig. 5b). These findings pointed to a region of genomic instability, potentially facilitating the insertion of the Bb-a-SDR.
The high sequence similarity between the Z and W chromosomes flanking the Bb-a-SDR region suggested limited accumulation of female-specific sequences on the W chromosome since its divergence (Fig. 1D and Supplementary Fig. 5a). To explore this further, we analyzed resequencing data mapped to Bb-a-hap2 and measured normalized read coverage across this region in individual samples (“Methods”). This revealed Bb-a-tesDw as the only different region, present in approximately half (11/25) of the resequenced female individuals (Supplementary Fig. 6). The remaining females, except sample SRR12010276, carried the type b SDR (described below). However, the presence of a pseudogenized il18r1-like gene (Fig. 1D) and an accumulation of significant SNPs within the pon2-like locus (Supplementary Fig. 3b, c) suggested the occurrence of recombination suppression.
To further validate the identified Bb-a-SDR, we performed PCR analysis using primers designed to amplify the region between tesDwa and pon2-like in 20 male and 22 female Bb individuals (“Methods”). Consistent with our previous bioinformatics analyses (Supplementary Figs. 3a–c and 6), all males showed negative amplification, while 13 females showed positive and 9 females showed negative amplification (Supplementary Fig. 7). This result confirmed the presence of Bb-a-SDR at the 3′ end of pon2-like in a subset of females and its complete absence in males. The substantial proportion of females lacking Bb-a-SDR in the above two independent analysis suggested the potential existence of an alternative female sex-determining mechanism or another W chromosome within the population. While we cannot exclude the possibility of sex reversal, as previously suggested23, we explored the former hypothesis by sequencing and assembling the genome of a Bb female that tested negative for Bb-a-SDR in our PCR analysis.
The new genome, designated Bb-b, was assembled using the same methodology as Bb-a (Supplementary Fig. 8a, b and Supplementary Table 1). A whole-genome alignment between the two new haplotypes (Bb-b-hap1 and Bb-b-hap2) revealed no significant structural variations in chromosome 13 (Supplementary Fig. 8c). Subsequent GWAS analyses, conducted separately on Bb-b-hap1 and Bb-b-hap2, again identified significant SNPs on chromosomes 3 and 13 (Fig. 2A, B). These SNPs were consistently associated with the same females identified in the previous GWAS based on Bb-a haplotypes, while the remaining females exhibited similarities to males (Supplementary Figs. 3a–c and 9). Further investigation of chromosome 13 SNPs revealed their location is not at the tesDwa locus flanked by pon2-like, but rather at a distinct locus upstream of the panx3-like gene. At this locus, we identified a new tesD duplicate, designated tesDwb. Conserved collinearity flanking the panx3-like region was recognized between the two haplotypes (Fig. 2C, D). The primary distinction is an insertion of a unique Bb-b-SDR into the 3′ UTR of a VPS9 (InterPro: IPR003123) domain-containing gene, designated vps9c, in Bb-b-hap2 (Fig. 2E, F).
A GWAS analysis identified significant sex-linked SNPs on chromosomes Chr03, Chr06, Chr13, and Chr16 using Bb-b-hap1 as the reference genome. The Y-axis displays the −log10 transformed p values obtained in the GWAS. B GWAS analysis identified significant sex-linked SNPs on Chr03, Chr06, Chr12, and Chr13 using Bb-b-hap2 as the reference genome. C Self-alignment dotplot of the region flanking vps9c and panx2-like on Bb-b-hap1-chr13. D Alignment of the genomic regions flanking vps9c and panx2-like between the Bb-a-hap1 and Bb-b-hap2, showing the insertion of Bb-b-SDR (pinked dashed line) into the vps9c gene. Blue arrows indicate the tandem inverted repeats (TIRs) of the Zator-1 element, as the blue arrows in (E). E Annotation reveals that Bb-b-SDR contains a duplicate (tesDwb) of the autosomal tesD gene and three TEs. The SDR is flanked by two Zator-1 elements. F Schematic representation of the Bb-b-SDR insertion into the 3′ UTR of the vps9c gene. G Sequence alignment reveals the 33-bp TIRs and 3-bp TSDs flanking the Bb-b-SDR. Source data are provided as a Source Data file.
Sequence analysis and repeat annotation revealed that the Bb-b-SDR comprises two palindromic DNA type II transposons, classified as Zator-1 based on homology to Bf transposons27, tesDwb, and a hAT-hATw transposon identified by RepeatModeler28 (Fig. 2E). The two arms of each Zator-1 element are flanked by 33-bp TIRs (identical terminal inverted repeats) (Fig. 2G), discernible through sequence alignment (arrows in Fig. 2D). Notably, these TIRs exhibit perfect identity between Zator-1 elements in Bf and Bb27. TSDs (Target site duplications) could be identified as TAA (Fig. 2G), also consistent with those observed for Zator-1 in Bf27, suggesting that the transposition event was mediated by the Zator-1 arms and likely occurred recently. This hypothesis was further supported by the high sequence identity between tesD and tesDwb, which differ by only nine SNPs, with four residing within the CDS (Supplementary Fig. 10a). Three of these CDS SNPs result in missense mutations (Supplementary Fig. 10a) at amino acid residues highly conserved between BjtesD and BbtesD (Supplementary Fig. 10b), indicating that these mutations accumulated recently in tesDwb following its translocation to this locus. Nevertheless, none of these SNPs were resolved as significant in our GWAS analysis, indicating that they have not been fully fixed in females carrying the tesDwb.
In addition to the SNPs, we identified a 63-bp deletion at the 3′ end of tesDwb compared to tesD (Supplementary Fig. 10a). This deletion was predicted to interfere with short-read mapping in individuals lacking it. Indeed, mapping analysis at the tesDwb locus revealed that all females (except SRR12010276) lacking Bb-a-SDR carried this 63-bp deletion of Bb-b-SDR (Supplementary Fig. 11). However, three male samples also appeared to carry this deletion (arrow heads in Supplementary Fig. 11). To validate this observation, we performed PCR using primers specific to Bb-b-SDR on the same 42 individuals from our previous analysis on Bb-a-SDR. PCR results indicated that 11 females tested positive for Bb-b-SDR (Supplementary Fig. 12a), with 9 of these individuals testing negative for Bb-a-SDR in our earlier PCR analysis (Supplementary Fig. 7). Notably, three males in our samples also tested positive for Bb-b-SDR (Supplementary Fig. 12a), confirming that some males indeed carry Bb-b-SDR.
To determine the genomic location of Bb-b-SDR in these males, we designed primers spanning the vps9c gene and tesDwb (Supplementary Fig. 12b). This analysis revealed that the Bb-b-SDR in 9 females (excluding the 13 females carrying Bb-a-SDR) is located within the vps9c gene. In contrast, the Bb-b-SDR in two males (M2 and M6) and two females (F12 and F19, also carrying Bb-a-SDR) is located elsewhere (Supplementary Fig. 12b). This observation was consistent with our finding that Bb-b-SDR contains recognizable TE arms, suggesting it may remain actively transposing. Only one male (M1) carries Bb-b-SDR at the vps9c locus (Supplementary Fig. 12b), indicating potential genuine sex reversal in approximately 5% of Bb males.
To investigate the genomic locations of Bb-b-SDR in males M2 and M6, we sequenced and assembled their genomes using Nanopore technology (Supplementary Table 3). Using the tesD sequence as a query for BLAST searches, we identified contigs Bb-M2-ctg000660 and Bb-M6-ctg001040 that carry Bb-b-SDR in each respective genome. Alignment of Bb-M2-ctg000660 to Bb-b-hap2 did not reveal the corresponding chromosome for this contig. However, self-alignment of Bb-M2-ctg000660 and alignment to Bb-b-hap2-chr13 revealed that two tandem duplicates of the M2-Bb-b-SDR are located in a region rich in short tandem repeats (Supplementary Fig. 13a, b). These repeats are characterized by conserved telomeric motifs (TTAGGG)n23, indicating that the two M2-Bb-b-SDRs were inserted into the telomeric region of an unidentified chromosome. Alignment with the female Bb-b-SDR showed that another unknown repeat was inserted into the hAT-hATw transposon for both M2-Bb-b-SDRs (Supplementary Fig. 13c), which was in line with the larger band observed in the PCR validation for the M2 male compared to the females carrying Bb-b-SDR (Supplementary Fig. 12a). TSDs are also discernible at the ends of the two Zator-1 arms, but the sequences (TTA, Supplementary Fig. 13d) differ from the TSDs observed in the Bb-b-SDR inserted into the vps9c locus (TAA, Fig. 2G).
Alignment between Bb-M2-ctg000660 and Bb-b-hap2 demonstrated that contig Bb-M6-ctg001040 maps to chromosome 7 of the Bb-b-hap2 assembly (Bb-b-hap2-chr07, Supplementary Fig. 14a). Comparative genomic analysis of the region flanking the M6-Bb-b-SDR on Bb-M6-ctg001040 and its homologous region on Bb-b-hap2-chr07 revealed that the M6-Bb-b-SDR is inserted within an intron of a gene encompassing a methyltransferase FkbM domain (Pfam: PF05050), which we designated as fkbmc (Supplementary Fig. 14b, c). Notably, the TSDs and TIRs of the Zator-1 elements at these insertion sites are identical to those observed in the Bb-b-SDR inserted into the vps9c gene (Fig. 2E, G and Supplementary Fig. 14c, d).
To precisely determine the genomic location of Bb-b-SDR observed in three male resequencing samples (arrowheads in Supplementary Fig. 11), we mapped their reads to the Bb-b-hap2-chr13, Bb-M2-ctg000660, and Bb-M6-ctg001040 genomic sequences. Reads from all three male samples spanned the Bb-b-SDR insertion site on Bb-M2-ctg000660 (Supplementary Fig. 15a). Conversely, read mapping to Bb-b-hap2-chr13 and Bb-M6-ctg001040 failed to demonstrate coverage across the respective SDR insertion sites (Supplementary Fig. 15b, c). This finding strongly suggested that these three males possess their Bb-b-SDR at the telomeric region like the Bb-M2-ctg000660, consistent with our M2 male genome. Theoretically, the Bb-b-SDR found in M2 and M6 males should also be present in type “a” females. Our PCR validations confirmed this, revealing the presence of type “a” females (F12 and F19, Supplementary Fig. 7) carrying Bb-b-SDR, as evidenced in Supplementary Fig. 12a, b.
In summary, our genomic analyses revealed two distinct and potentially functional female-specific sex-determining regions (SDRs) within the Bb population: the Bb-a-SDR at the twai locus and the Bb-b-SDR at the vps9c locus. The two types of SDRs are present in approximately equal proportions in female individuals, as indicated by resequencing data and our PCR validations. In addition, Bb-b-SDR exhibits characteristics of an active DNA transposon, flanked by two Zator-1 elements, and demonstrates the capacity for on-going genomic translocation. However, definitive evidence showing the transposition is lacking and should be explored further. We identified at least two distinct translocation sites, one in the telomeric region of an unknown chromosome (M2-Bb-b-SDR) and the other in the intron of the fkbmc gene on chromosome 7 (M6-Bb-b-SDR). Approximately 5–10% of both male and female individuals harbor these translocated Bb-b-SDRs on autosomes, which possibly have lost their female-determining function, as explored in subsequent functional analyses.
TesD is required for testis development in Bf
Our preceding analyses indicated that duplicates (tesDwa/b) of tesD likely plays a crucial role in the evolution and dynamics of the sex-determining mechanism in Bb. TesD encodes a transcription factor containing a basic helix-loop-helix (bHLH) domain (Supplementary Fig. 10b). While BLAST searches did not reveal clear orthologs of tesD in vertebrates, it exhibited sequence similarity to MyoD-like genes in certain invertebrates. Our phylogenetic and synteny analysis suggested that tesD is likely conserved in Ambulacraria and amphioxus but was secondarily lost in vertebrates and tunicates (Supplementary Note 1). To our knowledge, no bHLH gene has been definitively identified as a master sex-determining gene in metazoans6,26,29. However, bHLH genes are known to participate in sex determination pathways in fruit fly30 and mammals31.
TesD exhibited high conservation and testis-specific expression in all three examined Branchiostoma species Bf, Bj, and Bb (6), suggesting its potential conserved role in testis development and male determination across these species. The collinearity of tesD is also highly conserved in the three species (Fig. 3a). To directly test the necessity of tesD for male development, we generated tesD mutants in the experimentally more amenable Bf using CRISPR/Cas9 (Fig. 3b). We specifically targeted Bf females carrying a previously identified 6-bp W-chromosome-specific marker to facilitate unambiguous determination of genetic sex in the mutants24. Remarkably, among F2 individuals of ZZ genotype, while all wild type (N = 21) and tesD heterozygotes (N = 27) developed testis, all tesD homozygotes (N = 7) developed ovaries (Fig. 3c). These results demonstrated that tesD is essential for testis development and represents a key developmental gene within the Branchiostoma testis differentiation pathway. To determine whether tesD is specifically required for testis development, and thus more likely to be a “usual suspect”25 involving master regulator of sex determination, we analyzed the survival rates of offspring from three independent crosses of heterozygous parents. The survival rate of homozygous offspring was 20% (Supplementary Table 4), closely aligning with the theoretical value of 25%, indicating the specificity of tesD in male development. Furthermore, homozygous mutants with ZW genotypes exhibited no additional phenotypic abnormalities and were capable of breeding with wild-type males.
a TesD is positioned within a conserved collinearity among three Branchiostoma species. b CRISPR/Cas9 sgRNA targeting the first exon was used to generate the tesD mutant in Bf. c Homozygous tesD mutants exhibit complete penetrance of sex-reversal in genetic males, with fully developed ovaries observed.
Reverse transcription of tesDwa/b likely suppresses expression of tesD through an unknown mechanism
Our genomic analysis identified tesDwa and tesDwb as the sole genes within their respective SDRs. Given the essential role of the autosomal tesD gene in male development, we hypothesized that tesDwa/b function as inhibitors of tesD, thereby directing female development. This proposed regulatory mechanism parallels observations in several Populus species and persimmons (Diospyros lotus)32,33.
At present, we are unable to delete specific genomic segments in any amphioxus species, including Bb. As a result, we adopted an omics-based strategy to investigate the roles of tesDwa/b in female sex-determination in Bb. Mapping of published RNA-seq data from male and female gonads (Supplementary Table 2) to Bb-a-hap2 revealed high expression of spliced tesD transcripts at the tesD locus in all ten male gonads (Supplementary Fig. 16a). Notably, three female gonads exhibited abundant unspliced reads spanning the tesD locus (Supplementary Fig. 16b). These reads contained SNPs diagnostic of tesDwb, a finding further supported by direct mapping of reads to the tesDwb locus (Bb-b-hap2) using the same female samples (Supplementary Fig. 16c). The remaining three female samples showed similar unspliced read mapping at the tesDwa locus of Bb-a-hap2, though being fewer compared to tesDwb (Supplementary Fig. 16d). In males, no unspliced reads were detected at tesDwa/b loci. To validate these findings and resolve transcriptional directionality, we conducted strand-specific RNA-seq on ovaries from three type a and three type b females, alongside testes from three males as controls (“Methods”). The results confirmed that tesDwa and tesDwb are transcribed exclusively in type a and type b ovaries, respectively (Fig. 4A, B). Control males exhibited canonical tesD expression, while all six females lacked spliced tesD transcripts (Supplementary Fig. 17). Intriguingly, tesDwa/b transcription occurred antisense to autosomal tesD mRNA, with both loci co-transcribed with adjacent sequences of the SDRs (Fig. 4A, B and Supplementary Fig. 16c, d). Additionally, RT-PCR of ovarian RNA from type a and type b females yielded products spanning the full-length tesDwa/b (Supplementary Fig. 18). Sanger sequencing confirmed these transcripts as potential lncRNAs, consistent with their antisense orientation and lack of splicing.
A TesDwa transcripts are present in the gonads of type a females (red dashed line box) but are absent in type b females. Blue reads represent transcription to the right, while red reads indicate transcription to the left. B TesDwb transcripts are present in the gonads of type b females (red dashed line box) but not in type a females. C ATAC-seq peaks (dashed-line box) are detected at the promoter region of the autosomal tesD in male gonads, but not in the gonads of either female type. Minor peaks in type b female gonads are attributed to misalignment of tesDwb reads. Bb-a-hap2 was used as the reference genome.
To examine the expression of W-linked tesDwa/b during embryonic development and across various tissues, we analyzed published transcriptomic data spanning several developmental stages and nine adult tissues from Bb34,35 (Supplementary Table 2). In the embryonic samples, tesDwa transcripts were present at the 1-cell stage and in adults, while tesDwb transcripts were observed at the late neurula, 1-gill stage, and in adults (Supplementary Fig. 19a, b). In adult tissues, tesDwa was detected in the skin, wheel organ, endostyle, and branchial arch, whereas tesDwb was found in the skin, wheel organ, endostyle, notochord, and gut (Supplementary Fig. 19c, d). Because these samples represent mixed genetic backgrounds, the absence of specific transcripts may reflect a lack of individuals carrying tesDwa or tesDwb. Notably, we consistently detected high levels of vps9c transcripts in all samples (Supplementary Fig. 19b, d). Given that tesDwb is inserted within the 3′ UTR of vps9c, it likely shares vps9c’s broad expression pattern as part of a composite transcript. In contrast to the testis-specific expression of autosomal tesD, the widespread expression of W-linked tesDwa/b may reflect a simpler inhibitory mechanism that does not depend on tissue-specific promoters/enhancers (see Discussion).
To compare with the expression of W-linked tesDwa/b, we also conducted strand-specific RNA-seq on testes from the two genome-sequenced males (M2 and M6) carrying tesDwb at loci other than vps9c. As expected, no unspliced reverse transcripts were detected at these tesDwb loci (Supplementary Fig. 20a, b), but canonical autosomal tesD transcripts were observed (Supplementary Fig. 20c), indicating that these tesDwb loci are likely non-functional. Additionally, we performed strand-specific RNA sequencing on whole-body samples from an M2 male and an M6 male, using their respective genome assemblies for reference. In both cases, we did not detect any tesDwb transcripts (Supplementary Fig. 21a, b). However, canonical autosomal tesD transcripts were present in both samples (Supplementary Fig. 21c). These results suggested that neither M2-tesDwb nor M6-tesDwb were transcribed in males, indicating they are unlikely to serve the same function as the W-linked tesDwa/b.
We further hypothesized that tesDwa/b lncRNAs undergo processing into small RNAs (e.g., siRNAs or miRNAs), which could mediate repression of tesD via RNA interference—a mechanism proposed in some Populus and Diospyros lotus32,33. To test this, we analyzed small RNA sequencing libraries from type b female ovarian tissues, using male testes as controls (“Methods”). However, only few small RNAs mapped to the tesDwb locus were detected, and the result was not consistent in the two females (Supplementary Fig. 22). Therefore, the inhibition of tesD in females is probably not through RNA interference by small RNAs. It should be noted that our analysis was limited to mature gonadal tissues, and we cannot exclude the possibility that the mechanisms tested here operate transiently during early gonad development.
Finally, we investigated chromatin accessibility at the tesD locus in both testes and ovaries (from type a and type b females) using ATAC-seq. Consistent with the RNA-seq data, the testes displayed greater differential accessibility peaks at the promoter region of the autosomal tesD (Fig. 4C). The slightly elevated peaks observed in type b female ovaries were attributed to the misalignment of some tesDwb reads (Supplementary Fig. 10), which only differ from the tesDw locus at SNP level. To verify this, we used Bb-b-hap2 as the reference genome and mapped the ATAC-seq reads of type-b female ovaries. Indeed, the results showed that these reads were primarily aligned to the Bb-b-tesDwb locus, with minimal mapping to the autosomal tesD locus (Supplementary Fig. 23), supporting that they originated predominantly from the Bb-b-tesDwb locus. These findings indicated that repression of tesD occurs not via post-transcriptional mechanisms but through epigenetic regulation of genomic DNA for transcription. While these lncRNAs likely mediate autosomal tesD repression, direct evidence of their binding to DNA or mRNA targets is lacking. Future studies should determine whether tesDwa/b-derived lncRNAs modulate chromatin accessibility to repress autosomal tesD expression in females.
Origin and evolution of the sex-determining loci in Bb
To elucidate the evolutionary routes of the two types of SDRs, we performed a detailed alignment analysis, leveraging the sequence conservation among tesD, tesDwa, and tesDwb. This analysis revealed distinct alignment patterns: tesDwa aligns as a single contiguous segment to the autosomal tesD locus, whereas the alignment between tesDwb and tesD is fragmented into three discrete segments (Supplementary Fig. 24a, b). A short ( ~ 60-bp) segment within tesDwb exhibits near-identity to a region 5′ tesD that is consistently interrupted by TEs across all four assembled alleles (Supplementary Fig. 24b). This observation strongly suggested that the ancestral autosomal tesD locus was not interrupted before being transposed to tesDwb, indicating a hotspot of TE activity recently. Indeed, the upstream region of tesD displays substantial variability in TE content among the four alleles (Supplementary Fig. 24b). The remaining segments of tesDwb, which are interrupted by an hATw TE following transposition, align contiguously to a corresponding segment of the autosomal tesD (Supplementary Fig. 24b).
To resolve the phylogenetic relationships among these loci, we compiled alignable sequences from tesD, tesDwa, and tesDwb (including M2-tesDwb and M6-tesDwb). Using Bf-tesD and Bj-tesD as outgroups, we reconstructed phylogenetic trees with both Maximum Likelihood (ML) and Bayesian Inference (BI) (“Methods”). Both methods yielded congruent topologies for Bb sequences, with all nodes receiving strong statistical support (bootstrap values = 100%; posterior probabilities ≥0.99; Supplementary Fig. 25). The trees revealed two distinct duplication events: tesDwa originated from a duplication of an ancient autosomal tesD, whereas tesDwb arose more recently from a tesD paralog closely related to the extant tesD. Notably, the two male tesDwb alleles (M2-tesDwb and M6-tesDwb) formed a monophyletic cluster, suggesting that the Bb-b-hap2-SDR may have transposed to these autosomes and lost its function in sex determination (Fig. 5). At present, we cannot exclude the possibility of additional, unidentified tesDwb copies at other loci within the population.
TesDwa originated as a duplicate of an ancient autosomal tesD copy, evolving independently after its translocation to the twai locus. TesDwb emerged more recently as a duplicate of an autosomal tesD copy closely resembling the extant tesD (Fig. S10) following a second transposition event. The Bb-b-SDR, containing tesDwb, was subsequently transposed to other autosomal loci, which are no longer transcribed.
Lastly, we analyzed three types of TEs in Bb-a-SDR and two types in Bb-b-SDR within their respective assemblies (Bb-a-hap2 and Bb-b-hap2). The TE divergence density plot revealed recent genomic accumulation for both TE types in Bb-b-SDR, with peaks at small divergences of 1.58% and 2.23% (Supplementary Fig. 26). In contrast, the three TE types in Bb-a-SDR showed density peaks at divergences of 13.68%, 18.25%, and 20.01%, reflecting an older burst (Supplementary Fig. 26). These findings supported that Bb-b-SDR is considerably younger than Bb-a-SDR.
Discussion
This study demonstrates that tesD, a bHLH transcription factor, is essential and specific for male sex determination in Bf, as evidenced by targeted gene disruption in this species. This function appears to be conserved across all three Branchiostoma species (Bb, Bj, and Bf), supported by its testis-specific expression in each species23. It remains possible that tesD function is specific to Bf. To confirm its functional conservation in Branchiostoma, future knockout experiments in other species will be necessary. Nevertheless, our results have expanded the list of instructive genes for sex determination across eukaryotes, highlighting the remarkable diversity of molecular mechanisms underlying sexual differentiation. Furthermore, our results contribute to the ongoing investigation of sex-determining genes and loci across chordates, offering new insights beyond previously characterized “usual suspects”25.
Using our newly assembled Bb genomes, we identified two distinct female-specific SDR variants on chromosome 13, each containing a duplicated tesD gene flanked by distinct TEs. These SDR variants likely arose through independent duplication and transposition events, with Bb-b-SDR retaining transpositional activity (Fig. 5). Both duplicates of tesD (i.e., tesDwa/b) are transcribed as reverse-oriented, unspliced lncRNAs in females, alongside their flanking TEs, suggesting TE-driven regulatory networks may contribute to female sex determination. Intriguingly, Bb-b-SDR (encompassing tesDwb) continues to transpose to new loci (e.g., outside vps9c), though transcription is not detected in these regions in males carrying it (Supplementary Fig. 19). The insertion loci appear to play a crucial role in determining whether SDRs are transcribed. Bb-a-hap2-SDR and Bb-b-hap2-SDR are inserted into the coding region of twai and the 3′ UTR of vps9c, respectively (Figs. 1F and 2F). In contrast, M2-Bb-b-SDR is inserted into a telomeric region (Supplementary Fig. 13b), while M6-Bb-b-SDR is located within an intron of the fkbmc gene (Supplementary Fig. 14c). Consequently, Bb-a-hap2-SDR and Bb-b-hap2-SDR likely utilize the transcriptional machinery or promoters of their host genes to produce their lncRNAs. This also implies that tesD silencing in females may occur via reverse-transcribed lncRNAs that interfere with canonical tesD expression. However, the precise mechanism—whether through chromatin modification, RNA interference, or another unknown mechanism—requires further investigation. The involvement of lncRNAs in sex determination and its evolutionary turnover may be more common than previously thought36,37.
Our findings illustrate an empirical example of de novo origin and establishment of a new SDR (Bb-b-SDR) within a single species, coexisting with the ancestral Bb-a-SDR. Notably, Bb-b-SDR has achieved around 50% frequency within the population, suggesting its potential towards fixation. However, its ongoing transposition activity also highlights the possibility of continued sex chromosome turnover. For instance, if Bb-b-SDR were inserted into the exon or 3′ UTR of another gene that is broadly expressed, it could potentially become functional—similar to the current W-linked Bb-b-SDR—and trigger a new sex chromosome turnover event. Interestingly, our GWAS analyses did not identify any significant SNPs fully fixed in the tesDwb of all type b females. This implies that the rise of Bb-b-SDR frequency occurred over a relatively short evolutionary timescale.
The ancestral SDR represents a stable system with full sex linkage, while about 5% of Bb males are likely true sex reversal, carrying Bb-b-SDR at the vps9 locus (M1 in Supplementary Fig. 12b). The coexistence of a stable ancestral SDR (Bb-a-SDR) with a newer, mobile and transposon-driven SDR (Bb-b-SDR) raises questions about the evolutionary forces shaping sex-determination systems. One possible explanation for this shift lies in the balance between the evolutionary advantages of stability and the potential benefits of innovation. A stable SDR, such as Bb-a-SDR, ensures reliable sex determination and enables the accumulation of co-adapted gene complexes, reducing the risk of developmental errors. However, over time, stable sex chromosomes can accumulate harmful mutations due to suppressed recombination, leading to genetic decay38,39. In contrast, a mobile SDR can create new sex-determining regions and replace deteriorating ones, helping preserve function and maintain homomorphic sex chromosomes. Our findings therefore suggest that such transposition-driven turnover may be a key factor underpinning the maintenance of homomorphic sex chromosomes in Bb10,17,18,40, balancing long-term reliability with the capacity for evolutionary novelty. Future studies should investigate how selective pressures, population structure, and genomic context influence the equilibrium between these contrasting evolutionary strategies.
A similar evolving sex-determination system has been documented in Chinook salmon (Oncorhynchus tshawytscha)41. In this species, a partial duplication of the immune-related gene irf9 produced the conserved salmonid sex-determining gene sdY42. This gene likely serves as an inhibitor of female development by binding to Foxl243, a key regulator of female sex determination44. Notably, sdY resides within a functional TE cassette that enables its repeated relocation across the genome45. This process likely also generates non-functional autosomal copies of sdY with additional mutation46, which exhibit no sex linkage47. This observation mirrors our findings in Bb, where approximately 10% of individuals carry a non-functional autosomal Bb-b-SDR that does not exhibit sex linkage, but this lack of functionality is likely attributable to their insertion sites that preclude transcription. Although the specific TEs involved differ between Bb and Chinook salmon, this “jumping sex locus” strategy may be a more general mechanism that keeps sex chromosomes “young” and homomorphic18,46.
Our data also suggests a hypothesis where homomorphic chromosomes can persist through balancing selection: frequent but transient transpositions generate new SDR candidates, while existing regions retain functionality with minimal genetic decay. This system may serve as a proxy for early vertebrate sex chromosome evolution, where ancestral homomorphic pairs transitioned to heteromorphic states. Population data and modeling are needed to test this hypothesis. Finally, the identification of tesDw’s transcriptional plasticity—hijacking promoters in different genomic contexts—suggests that lncRNA-mediated repression could be a more common mechanism in chordate sex determination. Future studies exploring the epigenetic and regulatory interactions between tesD duplicates and their host genes (twai and vps9c) will be critical to resolving the evolutionary trajectory of amphioxus sex chromosomes.
Methods
Bb sampling and laboratory maintenance
The Bb lancelets were collected from Huangcuo, Xiamen City, Fujian Province, and maintained in our aquarium according to our previous protocol48. In brief, they were cultured in seawater at 24 ~ 26 °C with 25‰ salinity and fed with Pyramimonas sp. and Isochrysis galbana. Prior to dissection and sample collection, the animals were euthanized using MgCl2 solution (64 mM).
Genome sequencing, assembly, and annotation of two types of Bb females
For the two female individuals used for genome sequencing, whole-genome sequencing was carried out using the PacBio HiFi platform. The freshly dissected Bb tissues, after gonads removal, was immediately flash-frozen in liquid nitrogen for 20 ~ 30 min, stored at −80 °C, and subsequently shipped on dry ice to Novogene (Tianjin, China) for sequencing. Genomic DNA was extracted from muscle using the TaKaRa DNA Extraction Kit (Dalian, China) according to the manufacturer’s instructions. DNA quality was assessed by 1% agarose gel electrophoresis, while concentration and purity were measured using a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific, MA, USA). A SMRTbell library was then constructed from 50 mg of high-quality DNA using the SMRTbell Express Template Prep Kit 2.0 (Pacific Biosciences, CA, USA), following standard protocols. Finally, sequencing was performed on a PacBio Revio system.
The initial contigs were assembled using HiFi reads with Canu (v2.2)49 under the parameters “correctedErrorRate = 0.035 -pacbio-hifi”. Given the high heterozygosity, Haplomerger250, a tool designed for haplotype phasing in highly heterozygous genomes such as amphioxus, was employed. To eliminate redundancy, we employed purge_dups (v1.2.5)51 to further filter these haplotypes. The assembled genome was then scaffolded to the chromosome level using published Hi-C data23 with Juicer and 3D-DNA pipeline52. Finally, synteny analysis was performed between the assembled genome and previously published genomes using MUMmer (v4.0.0rc1)53, and chromosome IDs were renamed accordingly based on synteny relationships. Genome completeness was assessed using BUSCO (metazoa_odb10)54.
Genome annotation comprised two main components: repeat annotation and gene annotation. Repeat annotation was conducted using both de novo prediction and homology-based approaches. For de novo prediction, RepeatModeler (v2.0.4)28 and LTR_FINDER (v1.07)55 were used to construct a species-specific repeat library. For the homology-based approach, the Deuterostomia repeat library was created by merging repeat sequences from RepBase (release 27.05) and Dfam 3.756. Finally, RepeatMasker (v4.1.5) was used to annotate repetitive elements in the genome using a combined repeat library that integrated both the species-specific and homology-based repeat datasets.
The annotation of protein-coding genes in the B. belcheri genome was conducted using a combination of ab initio, homology-based, and transcriptome-based prediction approaches. RNA sequencing reads (NCBI accession numbers SRR12011574 to SRR12011597) from 22 samples of adult and juvenile gonad and muscle tissues from a previous research23 were mapped to the genome using HISAT2 (v2.2.1)57, and the resulting alignment files (.bam) were merged using Samtools (v 1.15.1). For homology-based annotation, protein sequences from previously published B. belcheri and B. floridae genomes were incorporated. Finally, BRAKER3, the latest genome annotation pipeline in the BRAKER suite, was employed to predict protein-coding genes58.
Genome sequencing and assembly of M2 and M6 Bb males
Nanopore sequencing was performed on muscle tissue from two males (M2 and M6) that carry Bb-b-SDR (Supplementary Fig. 12a) but at a different locus from the vps9c locus (Supplementary Fig. 12a) using the PromethION platform. The freshly dissected Bb tissues, after gonads removal, was immediately flash-frozen in liquid nitrogen for 20 ~ 30 min, stored at −80 °C, and subsequently shipped on dry ice to Novogene (Tianjin, China) for sequencing.
Genomic DNA was extracted using the SDS method, and its integrity was assessed by agarose gel electrophoresis and pulsed-field gel electrophoresis to detect degradation, RNA contamination, or protein impurities, while DNA concentration was quantified using a Qubit fluorometer. A 1D sequencing library was prepared following the standard adapter-ligation protocol, including optional DNA fragmentation, end repair, and A-tailing, and ligation of sequencing adapters, motor proteins, and tether proteins. After quality control, the final library was sequenced on the PromethION platform, with sequencing parameters optimized based on the effective library concentration and data output requirements.
Base calling was conducted using a Recurrent Neural Network algorithm to convert electrical signals into raw sequencing reads, generating approximately 52× and 80× genome coverage. The reads were assembled using NextDenovo (v2.5.2)59 with the parameter read_cutoff = 1k. Contigs containing tesDwb were identified using BLAST. Repeat annotation was performed as described in previous methods. Gene annotation for the contigs was completed using AUGUSTUS (v3.5.0) with the pre-trained B. floridae model.
Genomic alignment analyses
For the whole genome alignment, the two haplotypes of Bb-a and Bb-b were aligned to each other using LASTz (v1.04.41) with default setting. The alignment dotplot was displayed with a resolution of 2 kb and >95% similarity. Flexidot60 was used to generate the alignment plots for the identification of both SDRs, through aligning the SDR-containing segment of the W chromosome with the homologous non-SDR segment of the Z chromosome. The autosomal tesD locus was aligned to the tesDwa/b loci to identify the alignable homologous sequences among them. The word size was set to 15-bp for all plots. The homologous sequences were also aligned using MAFFT61 to identify the exact insertion site in Figs. 1F and 2F, G and Supplementary Figs. 13c, d and S14c, d.
Identification of SDRs by GWAS
We conducted a sex association analysis using previously published Bb whole genome sequencing (WGS) data (25 female individuals and 20 male individuals, Supplementary Table 2)23. After quality control, reads were aligned to the two assembled Bb haplotype genomes using BWA-MEM (v0.7.17) with default setting62. We used MarkDuplicates in the Picard (v2.26.11) (https://broadinstitute.github.io/picard/) package to remove duplicates, and variants were called for each sample using GATK (v4.1.8.1)63. SNPs were filtered with the following parameters: QD < 2.0||FS > 60.0 || MQRankSum < −12.5 || RedPosRankSum < −8.0||SOR > 3.0||MQ < 40.0. After merging all samples, SNPs were further filtered in VCFtools (v 0.1.16)64 with the following parameters: –max-missing 0.9 –maf 0.05 –min-meanDP 5. We used Beagle (v 5.1)65 to impute genotypes and converted the data format with PLINK (v1.90b6.21)66. We performed genome-wide association studies (GWAS) with EMMAX (intel64)67 and used a significance threshold of 1/n (where n is the effective number of SNPs).
Analysis of normalized read coverage on Bb-a-hap2
The bam files generated in the GWAS analysis was used for this analysis. GATK:DepthOfCoverage (v4.1.8.1)63 was used to calculate the coverage depth of reads on chromosomes, and each sample sequencing depth was standardized individually. The results were visualized with a window size of 200-bp.
PCR-based genotyping analysis
The tail tip of each Bb individual was excised, mixed with Chelex 100 resin beads and Proteinase K, and lysed at 65 °C for 1 h and 95 °C for 10 min. The lysates were then used for PCR amplification to determine the genotype with primers (Bb-a-tesDwa-Fw: 5′-TTCTCTCTCTCCGTGGCG-3′/pon2-Rev: 5′-GGTAGTCTGGCTATAATGTT-3′, Bb-b-genotyping-Fw: 5′-CCGACAACTATGAGGGTACT-3′/Bb-b-genotyping-Rev2: 5′-CTGACCTCTGATGTTATATAGC-3′, Bb-b-genotyping-Fw: 5′-CCGACAACTATGAGGGTACT-3′/−63bp Rev2: 5′-GCTGTGAAACTAGTCAAAGTGC-3′).
Detection of tesDwa/b transcripts in female Bb
Freshly dissected Bb gonads used to extract total RNA by TRIzol reagent (Ambion). cDNA was synthesized using HiScript III RT SuperMix ( + gDNA wiper) kit (Vazyme). RT-PCR was subsequently performed using primers (Bb-a-hap2-SDR-RT-F: 5′-AACACCGGAGCCAGTATCAA-3′/Bb-a-hap2-SDR-RT-R: 5′-AAGACTCACGTGGGATGGAG-3′, Bb-b-3′ UTR-F: 5′-ACATCTACTAGTACGGCTCG-3′/−63bp Rev2: 5′-GCTGTGAAACTAGTCAAAGTGC-3′) with the obtained cDNA as the template to amplify the tesDwa and tesDwb transcripts, respectively.
Generation of tesD mutant in Bf
TesD mutants were generated using CRISPR/Cas9 system. One sgRNA (designated tesD-sgRNA) targeting the sequence (5′-AGAGAGAGAAGAGACGAT-3′) in the first coding exon of tesD was synthesized. The process of generating founders of tesD mutants was conducted as our previous study68. A pair of primers (BftesD-gRNA3-F1: 5′-TAGGTCACGGCGCTACAAAT-3′; BftesD-gRNA3-R1: 5′-CTGCGCACAGCTGAAAAATC-3′) and BsmBI enzyme were used for genotyping and mutation type analysis. F1 tesD+/− carrying −13 bp mutation were intercrossed to generate tesD−/−. Genotypic sex was identified by PCR, with the primers and amplification protocol performed according to our previously study24. Briefly, at juvenile or adult stages, a minimal tail tip tissue was collected from each individual and lysed with Animal Tissue Direct PCR Kit (Foregene Co.). The lysate was amplified with primer pair P-F3/P-R0, followed by BsmI restriction digestion for genotype confirmation.
Strand-specific RNA-seq and analysis
Gonads of male and female Bb individuals of different genetic backgrounds were carefully dissected under a stereo-microscope. While a portion of them were snap-frozen immediately in liquid nitrogen for 20 ~ 30 min and stored at −80 °C for other usages, the remaining were homogenized in pre-chilled TRIzol reagent on ice and then sent to Novogene (Tianjin, China) strand-specific transcriptome sequencing. fastp (v0.20.1)69 was utilized to filter the sequencing reads and Hisat2 (v2.2.1) was used to align the clean data to the assembled Bb haplotypes57. Samtools (v1.12) was used to convert sam files to bam. Visualization was performed using the First-of-pair strand mode in Integrative Genomics Viewer (IGV, v2.18.4)70.
Small-RNA sequencing and analysis
The dissected fresh Bb gonads were homogenized and lysed using TRIzol, stored at −80 °C, and subsequently shipped on dry ice to Novogene for sequencing. Total RNA was extracted from the gonads of Bb type b females and males using TRIzol, following the RNA-seq protocol. The extracted RNA served as the input material for RNA sample preparation. Briefly, 3′ and 5′ adaptors were ligated to the respective ends of small RNA molecules. First-strand cDNA synthesis was performed after hybridization with a reverse transcription primer. Subsequently, a double-stranded cDNA library was generated through PCR enrichment. Libraries with inserts ranging from 18 to 40 base pairs were purified, size-selected, and prepared for sequencing. After completing library construction, the inserted fragments were quantified to ensure effective concentration and high-quality libraries. The qualified libraries were pooled and sequenced on Illumina platforms.
Raw reads in fastq format were processed using fastp (v0.20.1) software to remove adapters, poly-N sequences, and low-quality reads. The resulting clean reads were aligned to the reference genome using Bowtie1 (v1.3.1)71 with the parameters: -p10 -k100 –best –strata. Samtools (v1.15.1) was utilized to convert sam files to bam format, and visualization was performed using IGV (v2.18.4)70.
ATAC sequencing and analysis
We collected gonadal tissues from male and female Bb individuals for ATAC-seq analysis, with each group comprising two biological replicates. The collected samples were washed with 0.04% BSA in 3× PBS and then lysed on ice for 7 min using a lysis buffer containing 10 mM Tris-HCl (pH 7.5), 10 mM NaCl, 3 mM MgCl₂, 0.5% NP-40, and 0.1% Tween 20. During lysis, gentle pipetting was performed until the tissues were completely disrupted, releasing the nuclei. The nuclei were then washed and resuspended in a pre-cooled wash buffer (10 mM Tris-HCl, pH 7.5; 10 mM NaCl; 3 mM MgCl₂; 0.1% Tween 20) to obtain a nuclear suspension. Subsequently, library construction was carried out according to the manufacturer’s instructions using the Chromatin Profile Kit for lllumina (Novoprotein, N248).
We processed sequencing reads using Cutadapt (v3.7)72 to remove adapter sequences and low-quality bases. The clean data were then aligned to the assembled Bb haplotype genome using Bowtie2 (v2.4.4)73. Subsequently, Picard (v2.26.11) (https://broadinstitute.github.io/picard/) was employed to eliminate PCR duplicates and reads mapped to mitochondrial DNA. Peak detection for each sample was performed using MACS2 (v2.2.7.1)74. To create a unified, non-redundant peak set, all individual peak files were merged using bedtools merge (v2.30.0)75. For visualization purposes, the preprocessed data were converted into appropriate formats using the bamCoverage tool from (v3.5.1)76, and both peak regions and gene structures were visualized with IGV (v2.18.4)70.
Phylogenetic analysis of tesDwa and tesDwb
Homologous sequences among Bf-tesD, Bj-tesD, Bb-tesD, tesDwa, and tesDwb were identified using Flexidot and MAFFT. Multiple sequence alignments were generated with MAFFT and subsequently trimmed using trimAl (v1.4.rev15)77 to prepare the data for phylogenetic analysis.
For phylogenetic reconstruction, the most likely tree was generated using RAxML (v8.2.12)78 with the following parameters: raxmlHPC-SSE3 -f a -x 12345 -c 1 -m GTRCATIX -# 1000 -p 12345. Additionally, BI was performed using MrBayes (v3.2.7a)79 under the parameters: Lset nst = 2 rates=equal; Prset statefreqpr=fixed(equal); mcmcp ngen = 1000000 printfreq = 1000 samplefreq = 1000 nruns = 2 diagnfreq = 1000 nchains = 4 savebrlens=yes. In both analyses, Bf-tesD was designated as the outgroup.
Divergence analysis of TEs harbored in Bb-a-SDR and Bb-b-SDR
The divergence values of the three types of TEs in Bb-a-SDR (DNA/CMC-EnSpm, LTR/Gypsy, LINE/CR1-Zenon) and two types of TEs in Bb-b-SDR (DNA/Zator-1, DNA/hAt-hATw) were extracted from RepeatMasker annotation results of their respective Bb-a-hap2 and Bb-b-hap2 assemblies and subsequently analyzed through density curve visualization. More specifically, rnd-5_family-61 of DNA/CMC-EnSpm, linear of LTR/Gypsy, linear of LINE/CR1-Zenon were retrieved from the annotation file of Bb-a-hap2; rnd-1_family-53 of DNA/harbinger and rnd-1_family-5 of DNA/Zator-1 were retrieved from the annotation file of Bb-b-hap2.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequencing data generated in this study have been deposited in the NCBI database under accession code PRJNA1240468. The assembled Bb-a, Bb-b, Bb-M2, and Bb-M6 genomes are deposited at [https://www.ncbi.nlm.nih.gov/datasets/genome/?taxon=7741]. Genome annotation files can be found at https://doi.org/10.6084/m9.figshare.28681217.v2. Repbase database (Release 16.10) [https://www.girinst.org/repbase/index.html] was used in this work for repeat annotations. A full list of accession IDs for public data analyzed in this study is available in the Supplementary Table 2. Source data are provided with this paper.
Code availability
All the data were analyzed by publicly available software and packages in this study. All involved software and packages in this study are described in the “Methods” section.
References
Bachtrog, D. et al. Sex determination: why so many ways of doing it? PLoS Biol. 12, e1001899 (2014).
Zhu, Z., Younas, L. & Zhou, Q. Evolution and regulation of animal sex chromosomes. Nat. Rev. Genet 26, 59–74 (2025).
Beukeboom, L. & Perrin, N. The Evolution of Sex Determination (Oxford University Press, 2014).
Kitano, J., Ansai, S., Takehana, Y. & Yamamoto, Y. Diversity and convergence of sex-determination mechanisms in teleost fish. Annu. Rev. Anim. Biosci. 12, 233–259 (2024).
Stöck, M. et al. A brief review of vertebrate sex evolution with a pledge for integrative research: towards sexomics. Philos. Trans. R. Soc. B: Biol. Sci. 376, 20200426 (2021).
Smith, S. H., Hsiung, K. & Böhne, A. Evaluating the role of sexual antagonism in the evolution of sex chromosomes: new data from fish. Curr. Opin. Genet. Dev. 81, 102078 (2023).
Vicoso, B. Molecular and evolutionary dynamics of animal sex-chromosome turnover. Nat. Ecol. Evol. 3, 1632–1641 (2019).
Pennell, M. W., Mank, J. E. & Peichel, C. L. Transitions in sex determination and sex chromosomes across vertebrate species. Mol. Ecol. 27, 3950–3963 (2018).
Blaser, O., Neuenschwander, S. & Perrin, N. Sex-chromosome turnovers: the hot-potato model. Am. Nat. 183, 140–146 (2014).
Perrin, N. Sex reversal: a fountain of youth for sex chromosomes? Evolution 63, 3043–3049 (2009).
van Doorn, G. S. & Kirkpatrick, M. Turnover of sex chromosomes induced by sexual conflict. Nature 449, 909–912 (2007).
Saunders, P. A., Neuenschwander, S. & Perrin, N. Sex chromosome turnovers and genetic drift: a simulation study. J. Evolut. Biol. 31, 1413–1419 (2018).
Myosho, T., Takehana, Y., Hamaguchi, S. & Sakaizumi, M. Turnover of sex chromosomes in Celebensis group medaka fishes. G3 Genes|Genomes|Genet. 5, 2685–2691 (2015).
Kabir, A. et al. Repeated translocation of a supergene underlying rapid sex chromosome turnover in Takifugu pufferfish. Proc. Natl. Acad. Sci. USA 119, e2121469119 (2022).
Roberts, R. B., Ser, J. R. & Kocher, T. D. Sexual conflict resolved by invasion of a novel sex determiner in Lake Malawi cichlid fishes. Science 326, 998–1001 (2009).
Li, X. et al. Divergent evolution of male-determining loci on proto-Y chromosomes of the housefly. Nat. Commun. 15, 5984 (2024).
Rodrigues, N., Studer, T., Dufresnes, C. & Perrin, N. Sex-chromosome recombination in common frogs brings water to the fountain-of-youth. Mol. Biol. Evol. 35, 942–948 (2018).
Han, W. et al. Ancient homomorphy of molluscan sex chromosomes sustained by reversible sex-biased genes and sex determiner translocation. Nat. Ecol. Evol. 6, 1891–1906 (2022).
Marlétaz, F. et al. Amphioxus functional genomics and the origins of vertebrate gene regulation. Nature 564, 64–70 (2018).
D’Aniello, S., Bertrand, S. & Escriva, H. Amphioxus as a model to study the evolution of development in chordates. eLife 12, e87028 (2023).
Holland, L. Z. et al. The amphioxus genome illuminates vertebrate origins and cephalochordate biology. Genome Res. 18, 1100–1111 (2008).
Holland, L. Z. & Holland, N. D. Chapter Four - Cephalochordates: A window into vertebrate origins. In Current Topics in Developmental Biology Vol. 141 (ed. Gilbert, S. F.) 119–147 (Academic Press, 2021).
Huang, Z. et al. Three amphioxus reference genomes reveal gene and chromosome evolution of chordates. Proc. Natl. Acad. Sci. USA 120, e2201504120 (2023).
Shi, C. et al. A ZZ/ZW sex chromosome system in cephalochordate amphioxus. Genetics 214, 617–622 (2020).
Herpin, A. & Schartl, M. Plasticity of gene-regulatory networks controlling sex determination: of masters, slaves, usual suspects, newcomers, and usurpators. EMBO Rep. 16, 1260–1274 (2015).
Pan, Q. et al. Evolution of master sex determiners: TGF-beta signalling pathways at regulatory crossroads. Philos. Trans. R. Soc. Lond. B Biol. Sci. 376, 20200091 (2021).
Bao, W., Jurka, M. G., Kapitonov, V. V. & Jurka, J. New superfamilies of eukaryotic DNA transposons and their internal divisions. Mol. Biol. Evol. 26, 983–993 (2009).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl. Acad. Sci. USA 117, 9451–9457 (2020).
Curzon, A. Y., Shirak, A., Ron, M. & Seroussi, E. Master-key regulators of sex determination in fish and other vertebrates—a review. Int. J. Mol. Sci. 24, 2468 (2023).
Salz, H. & Erickson, J. W. Sex determination in Drosophila: The view from the top. Fly 4, 60–70 (2010).
Bhandari, R. K., Schinke, E. N., Haque, M. M., Sadler-Riggleman, I. & Skinner, M. K. SRY induced TCF21 genome-wide targets and cascade of bHLH factors during sertoli cell differentiation and male sex determination in rats. Biol. Reprod. 87, 131 (2012).
Akagi, T., Henry, I. M., Tao, R. & Comai, L. A Y-chromosome–encoded small RNA acts as a sex determinant in persimmons. Science 346, 646–650 (2014).
Müller, N. A. et al. A single gene underlies the dynamic evolution of poplar sex determination. Nat. Plants 6, 630–637 (2020).
Huang, S. et al. Decelerated genome evolution in modern vertebrates revealed by analysis of multiple lancelet genomes. Nat. Commun. 5, 5896 (2014).
Zhang, Q.-L. et al. Selection of reliable reference genes for normalization of quantitative RT-PCR from different developmental stages and tissues in amphioxus. Sci. Rep. 6, 37549 (2016).
Pan, Q., Darras, H. & Keller, L. LncRNA gene ANTSR coordinates complementary sex determination in the Argentine ant. Sci. Adv. 10, eadp1532 (2024).
Kato, Y. et al. A 5′ UTR-Overlapping lncRNA activates the male-determining gene doublesex1 in the crustacean Daphnia magna. Curr. Biol. 28, 1811–1817.e1814 (2018).
Charlesworth, B., Harvey, P. H., Charlesworth, B. & Charlesworth, D. The degeneration of Y chromosomes. Philos. Trans. R. Soc. Lond. Ser. B: Biol. Sci. 355, 1563–1572 (2000).
Jay, P., Jeffries, D., Hartmann, F. E., Véber, A. & Giraud, T. Why do sex chromosomes progressively lose recombination? Trends Genet. 40, 564–579 (2024).
Charlesworth, D. When and how do sex-linked regions become sex chromosomes? Evolution 75, 569–581 (2021).
Bertho, S., Herpin, A., Schartl, M. & Guiguen, Y. Lessons from an unusual vertebrate sex-determining gene. Philos. Trans. R. Soc. B: Biol. Sci. 376, 20200092 (2021).
Yano, A. et al. An immune-related gene evolved into the master sex-determining gene in rainbow trout, Oncorhynchus mykiss. Curr. Biol. 22, 1423–1428 (2012).
Bertho, S. et al. The unusual rainbow trout sex determination gene hijacked the canonical vertebrate gonadal differentiation pathway. Proc. Natl. Acad. Sci. USA 115, 12781–12786 (2018).
Bertho, S. et al. Foxl2 and its relatives are evolutionary conserved players in gonadal sex differentiation. Sex. Dev. 10, 111–129 (2016).
Faber-Hammond, J. J., Phillips, R. B. & Brown, K. H. Comparative analysis of the shared sex-determination region (SDR) among salmonid fishes. Genome Biol. Evolution 7, 1972–1987 (2015).
Bertho, S. et al. A nonfunctional copy of the salmonid sex-determining gene (sdY) is responsible for the “apparent” XY females in Chinook salmon, Oncorhynchus tshawytscha. G3 Genes|Genomes|Genet. 12, jkab451 (2022).
Ayllon, F. et al. Autosomal sdY pseudogenes explain discordances between phenotypic sex and DNA marker for sex identification in Atlantic salmon. Front. Genet. 11, 544207 (2020).
Li, G., Yang, X., Shu, Z., Chen, X. & Wang, Y. Consecutive spawnings of chinese amphioxus, Branchiostoma belcheri, in captivity. PLoS ONE 7, e50838 (2012).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Huang, S., Kang, M. & Xu, A. HaploMerger2: rebuilding both haploid sub-assemblies from high-heterozygosity diploid genome assembly. Bioinformatics 33, 2577–2579 (2017).
Guan, D. et al. Identifying and removing haplotypic duplication in primary genome assemblies. Bioinformatics 36, 2896–2898 (2020).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95 (2017).
Marçais, G. et al. MUMmer4: a fast and versatile genome alignment system. PLOS Comput. Biol. 14, e1005944 (2018).
Manni, M., Berkeley, M. R., Seppey, M., Simão, F. A. & Zdobnov, E. M. BUSCO update: Novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. Mol. Biol. Evol. 38, 4647–4654 (2021).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res. 35, W265–W268 (2007).
Hubley, R. et al. The Dfam database of repetitive DNA families. Nucleic Acids Res. 44, D81–D89 (2015).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Gabriel, L. et al. BRAKER3: fully automated genome annotation using RNA-Seq and protein evidence with GeneMark-ETP, AUGUSTUS and TSEBRA. Genome research, 34, 769–777 (2024).
Hu, J. et al. NextDenovo: an efficient error correction and accurate assembly tool for noisy long reads. Genome Biol. 25, 107 (2024).
Seibt, K. M., Schmidt, T. & Heitkam, T. FlexiDot: highly customizable, ambiguity-aware dotplots for visual sequence analyses. Bioinformatics 34, 3575–3577 (2018).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv preprint arXiv. 1303.3997 (2013).
Poplin, R. et al. Scaling accurate genetic variant discovery to tens of thousands of samples. biorxiv, 201178 (2017).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Browning, B. L., Zhou, Y. & Browning, S. R. A one-penny imputed genome from next-generation reference panels. Am. J. Hum. Genet. 103, 338–348 (2018).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience https://doi.org/10.1186/s13742-015-0047-8 (2015).
Kang, H. M. et al. Variance component model to account for sample structure in genome-wide association studies. Nat. Genet. 42, 348–354 (2010).
Su, L., Shi, C., Huang, X., Wang, Y. & Li, G. Application of CRISPR/Cas9 nuclease in amphioxus genome editing. Genes 11, 1311 (2020).
Chen, S. Ultrafast one-pass FASTQ data preprocessing, quality control, and deduplication using fastp. iMeta 2, e107 (2023).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Feng, J., Liu, T., Qin, B., Zhang, Y. & Liu, X. S. Identifying ChIP-seq enrichment using MACS. Nat. Protoc. 7, 1728–1740 (2012).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012).
Acknowledgements
We thank Jian Li, Zhen Huang, and Qi Zhou for helpful discussions. We thank Mr. Funiu Qin for technical support. We are grateful to Luohao Xu for suggestions on bioinformatic analysis. This research was supported by the Natural Science Foundation of Fujian Province of China (No. 2022J06004) and National Science Foundation of China (No. 32270439, 32070458, and 32061160471) to G.L.; start-up funds from Xiamen University and Natural Science Foundation of Xiamen, China (No. 3502Z202473009) to Q.Q.; National Natural Science Foundation of China (No. 32200411) to C.X.; National Natural Science Foundation of China (No. 32300346) to C.S.
Author information
Authors and Affiliations
Contributions
Q.Q. and G.L. conceived and designed the study. H.L., J.L., C.S., Z.L., R.P., X.W., Y.P., C.X., and Y.W. performed the experiments on amphioxus. F.L., J.L., Y.Q., Q.Q. assembled the genome, conducted genome annotation, and performed genome and transcriptome analyses. Q.Q. prepared the initial manuscript. G.L., F.L., J.L., H.L., and Q.Q. revised the manuscript. Q.Q. and G.L. supervised this study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Qiaowei Pan, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, H., Liu, F., Li, J. et al. Evolutionary dynamics of sex determination in Branchiostoma belcheri driven by repeated transposition of a single novel gene. Nat Commun 17, 1616 (2026). https://doi.org/10.1038/s41467-026-68322-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68322-6