Abstract
The 2022 multi-country mpox (formerly monkeypox) outbreak, driven by mpox virus (MPXV) Clade IIb poses renewed threat to global public health. The cessation of smallpox vaccination has created large immunologically naïve cohorts, with uncertain implications for contemporary MPXV susceptibility. To assess whether residual vaccination-derived immunity influences exposure risk, we combine serological and phylodynamic analyses. Using a six-plex Luminex assay, we measure immunoglobulin G (IgG) binding to six MPXV antigens in 176 Nigerian adults comprising of 75 healthcare workers sampled in 2021 and 101 community volunteers sampled in 2023. At baseline, 24/176 (13.6%) were MPXV seropositive, predominantly born before 1980. Magnitude-breadth analysis scores were two-fold higher in pre-1980 cohort relative to post-1980 cohort. In 153 participants with follow-up samples (median 9 months), 5/153 (3%) showed evidence of exposure, with ≥2-fold increases in magnitude-breadth scores and antigen-specific responses against ≥4/6 antigens without reported clinical illness. Antigen-specific responses were strongest to B6R (11-fold), followed by M1R and A35R, with marked individual-level heterogeneity. Complementary phylodynamic reconstruction of 105 Nigerian MPXV genomes identified sporadic transmission against frequent dead-end infections. Together, these data show that residual smallpox immunity continues to shape mpox transmission and asymptomatic exposure contributes to under-detected spread, informing surveillance and targeted vaccination strategies.
Similar content being viewed by others
Introduction
Mpox is a zoonotic disease caused by the mpox virus (MPXV), a member of the Poxviridae family within the Orthopoxvirus genus1,2,3. Historically endemic to Central and West Africa, the recent global resurgence of MPXV in endemic and non-endemic regions resulted in its declaration as a Public Health Emergency of International Concern (PHEIC) in both 2022 and 20244. Unlike previous outbreaks, the 2022 outbreak, was driven by the MPXV Clade IIb, an evolutionarily distinct, pathogenically attenuated lineage with enhanced human to human transmissibility5,6,7,8,9,10,11,12. Thus, it has been associated with mild and asymptomatic disease13,14,15,16,17,18,19,20,21,22,23,24.
A key determinant of susceptibility to MPXV is residual immunity from historic smallpox vaccination25,26,27, discontinued globally in 1980 following eradication28. MPXV and variola virus (agent of smallpox) share multiple conserved structural and immunogenic viral proteins such as A29L, B6R, and M1R, implicated in viral entry and immune recognition29,30,31,32,33,34,35,36. Individuals born before 1980 likely received a smallpox vaccine during the WHO-led Smallpox Eradication Programme (1966–1980)37,38,39,40, although coverage was heterogenous ranging from 7% to 60%25. The vaccines used in these campaigns were based on live-attenuated vaccinia strains such as Lister and EM-63, which induced durable, broad and cross-protective immunity against Orthopoxviruses, including MPXV41,42,43.
Although historical smallpox vaccination conferred broad cross-protective immunity against Orthopoxviruses44,45,46,47,48,49,50,51,52,53, its global cessation has created a growing global cohort of immunologically naïve individuals54, particularly in Africa where vaccinia-based vaccination was never incorporated into routine national programmes. Waning immunity in the post-smallpox era has coincided with the emergence of MPXV transmission patterns, especially across sub-Saharan Africa where surveillance remains limited. Evidence from high-resource setting has demonstrated asymptomatic MPXV infections, particularly among high-risk groups4,13,14,15,16,17,18,19,20,21,22,55. However, whether this smallpox immunity gap has shaped transmission dynamics in West African populations remains unclear due to a paucity of serological and genomic data from these high-burden setting.
To address this gap, we analysed serum samples from two independent Nigerian SARS COV-2 vaccinee cohorts (healthcare workers from Lagos, 202156, and community participants from Abuja, 202357) to evaluate MPXV-specific antibody responses. In parallel, we analysed both time-series case counts and whole-genome MPXV sequences that were generated from the Nigerian epidemic during the same sampling period. This allowed us to contextualise local transmission patterns and infer epidemiological dynamics in real time. Together, these complementary approaches provide insight into the durability of smallpox-derived immunity and reveal evidence of undetected community exposure of MPXV during the SARS-CoV-2 pandemic in the absence of any Orthopoxvirus vaccination efforts.
Results
Study population and cross-sectional MPXV seropositivity
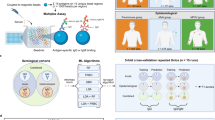
We analysed two independent cohorts originally established to assess SARS-CoV-2 vaccine responses in Nigeria (Fig. 1). The population comprised a total of n = 176 participants (timepoint 0; T0) stratified as follows: cohort 1: Healthcare workers (HCW, n = 75; enroled during the SARS-CoV-2 vaccination campaign in Lagos in January 2021) and cohort 2: general population (n = 101; enroled during community-level vaccination campaign in Abuja in January 2023). The median age was 37 years (IQR: 29-43) with balanced gender representation (Table 1). Follow-up samples were available for 153 participants (cohort 1: n = 75; cohort 2: n = 78), collected a median of 9 months after T0 (T1). This yielded a total of n = 329 samples. This dual-cohort design allows us to capture the potential impact of occupational exposure to MPXV and age-cohort differences on MPXV-specific antibody responses, particularly in the context of historical smallpox vaccination in Nigeria—a country central to the 2022 global outbreak.
a Study setting and cohorts. Participants were from two SARS-CoV-2 vaccine studies in Nigeria. Cohort 1 comprised healthcare workers (n = 75) in Lagos in 2021 originally enroled in the Lagos vaccine effectiveness study of ChAdOx1 (AstraZeneca). Cohort 2 comprised general community participants (n = 101) recruited in Abuja in 2023 through community outreach, as part of SARS-CoV-2 vaccine roll-out using Ad26.COV.S (Johnson and Johnson). The map shows Nigerian states of sampling. Baseline sera (T0, n = 176) were collected prior to SARS-CoV-2 vaccination and tested for IgG against six MPXV antigens (A29L, A35R, B6R, D6L, H3L, M1R). Paired follow-up sera (T1, n = 153) were available for all HCWs n = 75 and n = 78 general participants, yielding 329 total samples. b MPXV antigens tested. Cartoon schematic of an Orthopoxvirus virion showing the six antigens included in the Luminex assay. These represent IMV surface proteins (A29L, H3L, M1R), EEV glycoproteins (A35R, B6R), and a soluble virus-associated protein (D6L). Image in 1b was created in BioRender. Abdullahi, A. (2026) https://BioRender.com/jbph67x. IgG, immunoglobulin G; MPXV, mpox virus; IMV, intracellular mature virion; EEV, extracellular enveloped virion; HCWs, healthcare workers; T0, baseline timepoint; T1, follow-up timepoint.
At T0 across both cohorts, using binding antibodies above the antigen-specific cut-off for each target, we observed 48/176 (27%) with no reactivity, 46 (26%) with reactivity to one antigen, 37/176 (21%) with reactivity to two antigens and 21/176 (12%) recognising three antigens (Table 2; Supplementary Fig. 1). Applying a tiered algorithm to define seropositivity threshold and to eliminate the likelihood of false positivity from assay artefact or isolated cross-reactivity, we defined MPXV seropositivity as reactivity to at least four of six MPXV antigens. By this definition, 24/176 (13.6%) were seropositive. Seropositivity was strongly associated with birth cohort: 20/59 (34%) of individuals born before 1980 versus only 4/101 (4%) born after 1980 (Fig. 2; Supplementary Table 1). Of note, this birth cohort effect differed by occupational group. Among HCWs, all 13 seropositive cases were confined to those born before 1980, consistent with legacy smallpox vaccination. In contrast, in the general community cohort, while seropositivity was also enriched in those born before 1980 (7/11; 64%), we detected 4 younger individuals (born after 1980) meeting the ≥4-antigen threshold, suggesting possible previous exposure to circulating mpox or other Orthopoxviruses. We note further that only (20/59; 34%) of participants born before 1980 showed MPXV seropositivity suggesting possible low smallpox vaccination coverage of ≤34%.
Binary heat-map showing serum IgG reactivity to six MPXV proteins (A29L, A35R, B6R, D6L, H3L, M1R) in 24 participants who met the criteria for MPXV seropositivity defined as reactivity to ≥4 targets. Each row represents one individual: each column one antigen. Blue squares represent reactive (signal ≥ assay cut-off); light-green squares represent non-reactive (signal <assay cut-off). Seven participants showed reactivity to ≥5 targets and three showing reactivity to all six targets. The colour-coded side bar indicates individuals born before 1980 (orange squares) or in or after 1980 (grey). IgG, immunoglobulin G; MPXV, mpox virus.
Broad responses were uncommon; seven participants (six born prior to 1980) were reactive to five antigens and only three participants (all born prior to 1980) were reactive to all six antigens. We note that most individuals (67/116; 58%) born after 1980 had ≤1 reactive antigen (Table 2). Overall, occupational status exerted a lesser effect with 13/75 (17 %) of HCWs relative to 11/101 (11%) community participants meeting the ≥4-antigen seropositivity criteria. These baseline patterns highlight both the durability of smallpox-derived immunity in older cohorts and evidence of potential mpox exposure in younger community participants.
Evidence of moderate positive MPXV-specific IgG responses between MPXV-antigens
Following quantification of IgG-specific responses across our study population, we assessed pairwise correlations of log10-transformed binding antibody levels for all six antibodies measured in (n = 176) at T0. Correlation coefficients (ρ) were visualised using a matrix heatmap (Supplementary Fig. 2a), alongside pairwise scatter plots illustrating individual-level IgG responses across antigen pairs (Supplementary Fig. 2a). We observed moderate and weak positive correlations between multiple antigens, with the strongest correlation observed between the EEV-associated antigens A35R and B6R (ρ = 0.65; p < 0.0001). We also observed moderate correlations between A35R and M1R (ρ = 0.51; p < 0.0001), and between B6R and M1R (ρ = 0.53; p < 0.0001), suggesting weaker but partially positive pattern between these structurally distinct antigens. All other correlations were relatively weak (ρ range: 0.15–0.43) although statistically significant (p < 0.05 for all comparisons). These data indicate a heterogenous but measurable degree of serological correlation, with stronger coupling between certain antigen pairs, particularly those associated with the EEV. Such variation supports the utility of composite metrics to capture the breadth of MPXV-specific antibody reactivity which we apply in subsequent analyses. To exclude the possibility that patterns were diluted by vaccine-naive individuals, we performed sensitivity analysis restricted to individuals born before 1980 (n = 59), i.e. likely to retain residual immunity from smallpox vaccination. Results were broadly consistent, although a stronger correlation was observed between A35R and B6R (ρ = 0.81; p < 0.0001) with some variation in the strength of other pairwise associations. These sensitivity analyses are visualised in Supplementary Fig. 2b, which provides the full correlation matrix for the pre-1980 birth cohort. Notably, no significant correlations were observed for D6L with any other antigen, suggesting it likely elicits a distinct response profile.
Serological profiles suggest persistence of Orthopoxvirus residual immunity by birth cohort
We first compared MPXV-specific IgG binding responses between healthcare workers (HCWs) and community participants at baseline (T0; n = 176). To ensure comparability across antigens, we also normalised responses to their antigen-specific cut-off values, presenting both raw mean fluorescence intensity (MFI) and normalised signal-to-cut-off ratios in parallel. This was performed to provide additional perspective by scaling responses relative to assay thresholds. HCWs exhibited modestly higher geometric mean titres (GMTs) across four antigens—A35R (726 vs 503; p = 0.53), B6R (396 vs 255; p = 0.48), H3L (1904 vs 1522; p = 0.21), and M1R (401 vs 313 p = 0.0002), this corresponded to ∼1.2–1.6-fold higher responses relative to the general population, though these differences did not reach statistical significance (Fig. 3a). In contrast, responses to A29L (2208 vs 2550; p = 0.31) and D6L (1129 vs 1318; p = 0.56) were slightly lower in HCWs. These results suggest that occupational exposure may have a limited effect on MPXV seroreactivity in this setting.
Serum IgG binding responses to six MPXV antigens measured by Luminex. Antigens represent distinct viral components: A29L, H3L, and M1R (intracellular mature virion, IMV); A35R and B6R (extracellular enveloped virion, EEV); and D6L (soluble antigen). a, b Boxplots of raw and normalized mean fluorescence intensity (MFI) values stratified by a occupational status (healthcare workers [HCW], orange; general population [GenPop], blue) A29L (2208 vs 2550; p = 0.31); A35R (726 vs 503; p = 0.53); B6R (396 vs 255; p = 0.48); D6L (1129 vs 1318; p = 0.56); H3L (1904 vs 1522; p = 0.21) and M1R (401 vs 313 p = 0.0002) and b birth cohort (born before 1980, orange; born in or after 1980, blue); A29L (2948 vs 2161; p = 0.001); A35R (1572 vs 358; p < 0.0001); B6R (708 vs 202; p < 0.0001); D6L (1407 vs 1155; p < =0.24); H3L (2406 vs 1394; p = 0.0008) and M1R (452 vs 305; p < 0.0001). For each antigen, point estimates with 95% confidence intervals displayed above the boxplots indicate the proportion of participants seropositive. Boxes represent the 25th–75th percentiles (interquartile range), the centre line indicates the median, whiskers extend to the most extreme values within 1.5× the interquartile range, and individual points beyond this range are shown as outliers and the black x symbols denote the geometric mean titres (GMTs) for each group. c, d Cumulative magnitude-breadth plots comparing overall antibody responses across groups. Magnitude-breadth scores represent the sum of binding antibody intensities across all six antigens, capturing both the strength (magnitude) and diversity (breadth) of the humoral response. c compares distributions by occupational status (HCW vs GenPop; p = 0.55); d by birth cohort (born before vs after 1980; p < 0.0001). Statistical comparisons were made using the two-tailed Wilcoxon matched-pairs signed-rank test; *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. IgG, immunoglobulin G; MPXV, mpox virus; IMV, intracellular mature virion; EEV, extracellular enveloped virion; HCWs, healthcare workers; T0, baseline timepoint; T1, follow-up timepoint. Shaded bands represent pointwise exact binomial 95% confidence intervals around the cumulative proportion of participants exceeding each magnitude-breadth score, with the central curve representing the observed cumulative proportion.
Given the absence of responses by occupational status, we next stratified participants by birth cohort, reflecting the cessation of smallpox vaccination in 1980 across (n = 176) participants to evaluate the potential impact of residual smallpox immunity. Individuals born before 1980 showed consistently higher antibody levels across all six MPXV antigens - A29L (2948 vs 2161; p = 0.001), A35R (1572 vs 358; p < 0.0001), B6R (708 vs 202; p < 0.0001), D6L (1407 vs 1155; p = 0.24), H3L (2406 vs 1394; p = 0.0008), and M1R (452 vs 305; p < 0.0001), representing ∼1.2–4.4-fold higher titres compared with those born after 1980 (Fig. 3b). Almost all differences were statistically significant. Sensitivity analysis restricted to the general population cohort confirmed these patterns, with older participants showing markedly higher antibody responses but with a subset of younger individuals also meeting seropositivity criteria, confirming that the observed patterns are not driven by occupational biases in the HCW cohort (Supplementary Fig. 3). These findings are consistent with durable cross-reactive antibodies resulting from prior smallpox vaccination. However, the detection of MPXV-reactive IgG antibodies in some individuals born after 1980 suggests possible cryptic exposure to MPXV or cross-reactivity with related Orthopoxviruses.
To capture overall response patterns, we generated magnitude-breadth plots—analogous to Kaplan-Meier curves, which integrate antibody strength (magnitude) and the number of recognised MPXV antigens (breadth). Consistent with patterns observed in Fig. 3a, b, HCWs displayed a modest rightward shift in magnitude-breadth distribution compared with the community cohort, (GMT: 7070 vs. 8065; p = 0.55; Fig. 3c). In contrast, participants born before 1980 demonstrated significantly higher cumulative responses (GMT: 5812 vs. 12328, p < 0.0001; Fig. 3d), reinforcing the impact of residual smallpox immunity. To corroborate the serological patterns, we conducted supplementary follow-up in the HCW cohort, where call-response rates were high. We captured smallpox vaccination history and presence of a vaccination scar among the HCW seropositive individuals (n = 13). Among 11/13 respondent participants, 9/11 reported prior smallpox vaccination and had visible scars, two participants could not remember receiving a smallpox vaccine although one had a visible smallpox vaccine scar. These findings are consistent with the persistence of residual smallpox-derived immunity.
Immunological signatures reveal evidence of recent MPXV exposure
To investigate evidence of undetected MPXV exposure, we leveraged longitudinal serology, applying composite measures of antibody magnitude and breadth in the absence of any apparent or reported clinical illness. Using our stringent composite definition, we observed (n = 5/153; 3%) participants who demonstrated ≥2-fold increase into magnitude-breadth score between baseline (T0) and follow-up (T1; median 9 months), together with ≥2-fold increases in antibody levels against ≥4 of 6 MPXV antigens (Fig. 4a, left panel). These individuals showed marked increase in magnitude-breadth score (GMT: 3876 vs 24898) whereas participants not meeting exposure criteria displayed negligible difference (7570 vs 7066; Fig. 4b, right panel). None of these five participants reported mpox-compatible symptoms during follow-up as part of the parent SARS-CoV-2 vaccine studies, which captured vaccination-related adverse events. However, structured surveillance for mpox-specific symptoms was not undertaken. These data are consistent with evidence of sub-clinical and pauci-symptomatic MPXV exposure during the sampling interval in ostensibly healthy individuals which may be captured only through sensitive multiplex antibody profiling rather than routine clinical surveillance.
a Kaplan Meier curves showing magnitude-breadth score in participants with and without evidence of MPXV exposure, based on cumulative responses and antigen-specific boosting. b Paired magnitude-breadth scores from baseline (T0) and follow-up (T1) showing changes in exposed (navy blue) versus unexposed participants (dark orange). c Antigen-specific fold changes in raw mean fluorescence intensity (MFI) and normalized MFI across the six MPXV targets (A29L, A35R, B6R, D6L, H3L, M1R) for each exposed participant. Participants were classified as MPXV-exposed if they demonstrated a > 2-fold increase in magnitude-breadth score between T0 and T1, together with ≥2-fold increases in ≥4 of the six antigens tested. Analyses were restricted to participants with paired serum samples at T0 and T1 (n = 153). Of these, 5 participants met criteria for MPXV exposure and 148 were classified as unexposed. Each data point represents one individual participant. Participants were classified as MPXV-exposed if they demonstrated a ≥ 2-fold increase in magnitude-breadth scores between T0 and T1 together with ≥2-fold increases in at least 4 of the 6 antigens tested. To the small number of exposed individuals (n = 5), b, c are descriptive and no formal statistical testing was performed. IgG, immunoglobulin G; MPXV, mpox virus; T0, baseline timepoint; T1, follow-up timepoint; y.o, years old.
To explore potential factors associated with MPXV exposure, we examined antigen-specific boosting profiles among exposed individuals using raw MFI values and normalised MFI values (Fig. 4c). Most participants were born before 1980 (3/5; 60%), mostly males (4/5; 80%) and were all MPXV-seronegative at baseline. Across the six antigens, the largest median fold-increase was observed for B6R (11-fold) and M1R (6.2-fold) followed by A35R (5.2-fold), D6L (4.6-fold), A29L (4-fold) and H3L (2.7-fold). These results indicate that B6R, an EEV-associated antigen, elicited the largest median fold increase among exposed individuals, although the response varied across participants. M1R and A35R also demonstrated notable increases, and together with B6R, appear to represent consistently recognised targets of post-exposure boosting despite individual-level heterogeneity. This highlights the potential value of B6R, A35R, and M1R as antigenic markers for serological surveillance of Orthopoxvirus exposure. Overall, these data provide additional resolution into the heterogeneity of antibody response and boosting patterns following MPXV exposure or re-exposure.
Residual smallpox immunity explains slow mpox growth rate in Nigeria
To contextualise serological evidence of undetected exposure, we combined epidemiological and genomic analyses. We estimated the population dynamics of Clade IIb/sh2017/lineage A58 alone under a non-parametric Skygrid and parametric exponential model. While our analysis overlaps with the dataset reported by Parker and colleagues8, it differs in scope; whereas Parker et al. focused on phylogenetic reconstruction, we combined the genomic data with contemporaneous incidence data, enabling joint estimation of doubling time, effective population size, and renewal-based Rt dynamics. Our analysis remained consistent previous findings7,8 which indicated that the epidemic grew exponentially with an estimated doubling time of ≈2.5 years [95% HPD: 1.87–3.43] (Fig. 5a, b). Our coalescent reconstruction of the viral effective population size (Ne) using a Skygrid model indicates a transient expansion since 2015 preceding official case detection in 2017 by the Nigeria CDC. We then used case data to estimate the time varying reproductive number from 2017 onward.
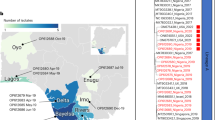
a The effective population size of the epidemic in Nigeria under a Skygrid and exponential coalescent model t. Shaded regions represent 95% highest posterior density (HPD) intervals around the posterior mean estimate. Vertical dashed lines indicate inferred breakpoints between epidemic phases. b The posterior distribution of the estimated doubling time of the epidemic since sustained human transmission. c Estimated reproductive number (Rt) over time, delineating two epidemic phases: initial phase (2017–2018) with Rt <1, and second phase (2021–2022) with Rt > 1. Solid lines represent the median posterior estimate of Rt, and shaded bands indicate the 95% confidence intervals. Grey background shading denotes alternating six-month time blocks to aid visualisation. d Map of Nigeria generated using Python coloured by regions to match c, showing the spatial grouping used in the analyses and where Sequences and cases were sampled. e Distribution of identical-sequence cluster sizes during the second epidemic phase, dominated by singletons. f, g Inferred transmission parameters under 5%, 10%, 50%, and 100% case-ascertainment scenarios. The estimated reproduction number is ≈ 1.2; 95% CIs: 1.11–1.26, 1.13–1.29, 1.11–1.35, 0.97–1.32 for 5%, 10%, 50%, 100%), while the dispersion parameter k hits the upper bound with only lower 95% limits identifiable (k ≥ 1.85, ≥ 1.62, ≥ 0.881, ≥ 0.493). The centre dot in f-g indicates maximum likelihood estimate, and the vertical lines indicate 95% likelihood profile confidence interval.
Renewal modelling59,60,61 of case data from 2017 to 2022 showed R < 1 during 2017–2018, consistent with limited human-to-human transmission a subsequent suppression in effective reproductive number (Fig. 5c, d) likely due to control measures rolled out by the Nigeria Centre for Diseases Control and Prevention (NCDC).10 In contrast, the second phase (2021–2022) shows sustained transmission across all Nigerian regions, with regional R values peaking at ≈1.3 (Southwest/Southeast), signalling exponential growth during this period, consistent with the timing of our cohort study and contemporaneous with observed serological boosting. Overdispersion analysis was performed using the method outlined by Tran-Kiem and Bedford62 for the 105 sequences sampled by Parker and colleagues8, which represented the second phase of the epidemic. This analysis revealed a median of seven distinct identical-sequence clusters, each typically comprising 1–3 sequences (Fig. 5e). This pattern indicates many transmission events leading to dead ends. Across ascertainment scenarios of 100%, 50%, and 10% respectively, R was consistently near-to-above unity, for 5–50% ascertainment the epidemic appears mildly supercritical (CIs above 1), while at 100% ascertainment the interval overlaps 1, consistent with near-critical transmission, which is consistent with the case data (Fig. 5f). Dispersion estimates varied with ascertainment (k ≥ 1.85 (5%), k ≥ 1.62 (10%), k ≥ 0.881 (50%), and k ≥ 0.493 (100%) (Fig. 5g). These patterns indicate frequent dead-end infections with modest transmission heterogeneity, and these bottlenecks in transmission likely reflect residual population immunity from historical smallpox vaccination, in line with our serological observations.
Discussion
Understanding how Orthopoxvirus immunity from historical smallpox vaccination shapes contemporary MPXV susceptibility is critical to estimating population-level risk and guiding rational vaccination strategies, particularly across resource-constrained settings. While smallpox vaccination historically conferred durable, cross-protective immunity to Orthopoxviruses, its global discontinuation post-elimination of smallpox has left a growing global cohort of immunologically naive individuals, especially those born after WHO eradication vaccination campaigns ended in 198040. In this context, our findings provide evidence of two complementary phenomena: (i) persistence of broad, durable residual immunity in older adults born before 1980, consistent with prior smallpox vaccination63,64 and (ii) a potentially underestimated yet significant threat65: serological signatures of recent MPXV exposure in ostensibly healthy adults without documented clinical illness24. To ensure robustness, we performed multiple sensitivity analyses for different comparator groups (pre- vs. post-1980 birth cohorts, healthcare workers vs. community participants, baseline seropositive vs. seronegative, and longitudinally boosted vs. non-boosted individuals). These analyses were consistent with our main findings.
Our baseline serology data shows residual vaccinia-derived immunity remains broad and antigen-diverse, with significantly higher IgG magnitude and breadth in individuals born before 1980 relative to younger counterparts, despite heterogeneity in historical vaccination coverage, (≈7–60% coverage during the WHO campaigns)25. In our cohort, 83% of MPXV-seropositive participants were born before 1980, compared with only 34% of the overall population, reinforcing the impact of historical vaccination. These datasets extend landmark immunological studies characterising durable Orthopoxvirus-specific antibody and T-cell responses persisting for 40–80 years post-vaccination66,67,68,69,70, now contextualised within a West African setting. Importantly, the retained breadth of responses spanned multiple Orthopoxvirus canonical targets of both intracellular mature virion form (A29L, H3L, M1R) and extracellular enveloped virion form (A35R, B6R), consistent with epitopes structural studies identify as dominant neutralising epitopes known to mediate protective neutralisation71,72.
Next, we observed antigen-specific antibody boosting in a subset of younger, post-1980 participants who lacked evidence of prior vaccination and were mostly male, supporting the occurrence of undetected MPXV exposure. While this likely reflects silent subclinical exposure rather than symptomatic infection, we acknowledge that serology alone cannot provide absolute proof of asymptomatic infection, and we therefore frame these findings as indicative of exposure rather than definitive evidence of silent transmission. In our pooled cohorts (HCWs and community participants), this corresponded to ≈3% of participants, a figure that should not be interpreted as a population-level prevalence but rather as a reflection of our study cohort. These observations dovetail previous reports of subclinical seroconversion from screening cohorts in Spain15 (6%, 7/113), France18 (6.5%, 13/200), Japan13 (0.4%, 5/1346), and Belgium17 (1.3%, 3/225). Animal data provide a mechanistic parallel: experiments in the UK showed that common brown rats (Rattus norvegicus) inoculated with Clade IIb MPXV remained clinically silent despite PCR positivity, yet seroconverted and transmitted infection to cage-mates without detectable viraemia73. This supports our observation of antibody boosting in the absence of reported clinical disease, highlighting the plausibility that cryptic rodent-human/human-rodent/human-human transmission cycles2,74,75,76 can sustain below the radar transmission that evade syndromic surveillance. The same cryptic transmission cycle has been observed for arenaviruses such as Lassa fever, which is peri-domestically maintained by the common African rat (Mastomys natalensis/couchi) reservoir host77,78,79,80. One recent meta-analysis estimated seroprevalence of Lassa fever up to 58% across the West African region81 with reported cross-sectional seroprevalence of 50% by age 582 and 82% in a general endemic West African population which exceeds reported case numbers; implying substantial subclinical spread with limited syndromic capture83. These data indicate higher than reported rates of exposure to heterogenous emerging infectious disease threats that can remain largely undetected84.
Our present observations are further validated by our country specific genomic data and previous phylogenetic analysis which revealed how MPXV may persist in the presence of partial immunity. Epidemiologically, a low dispersion parameter (k ≈ 0.4–0.8) at 50% and 100% ascertainment together with a long doubling time despite exponential growth suggests transmission concentrated in clustered contact networks, consistent with focal outbreaks in households or other close-contact groups10. The relative absence of large clusters (>3 linked cases) supports a pattern in which stochastic superspreading events, rather than sustained community-wide transmission, drive persistence. This dynamic aligns with modelling of the 2022 global outbreak and explains how MPXV can circulate at low reported incidence, undetected and yet still generate the boosting serological patterns we observed in our population59,62. In context, the dead-end infection of most mpox transmission means traditional case-based surveillance is unlikely to capture the full extent of mpox transmissions, and public health messaging must be context dependent85, particularly in settings where infections with the Clade IIb presents with mild and atypical symptoms. Furthermore, our focus on serological analysis of samples collected during the COVID-19 when healthcare resources were focused on responding to the pandemic strengthens the plausibility of missed mpox diagnosis.
Antigen-specific kinetics showed boosting of B6R, M1R, and A35R, with B6R exhibiting the largest fold increases across exposed participants, highlighting its prominence86 This EEV-specific envelope protein expressed in the extracellular virion form of MPXV exhibited the highest post-exposure boost (11-fold) and strongest inter-antigen correlation between antigen pairs. To a lesser extent, M1R (6.2-fold) and A35R (5.2-fold) were also exhibited notable post-exposure boosting. Structural and functional studies provide a mechanistic rationale for this hierarchy: cryo-EM resolution of B6R in complex with monoclonal antibodies has revealed multiple simultaneously accessible neutralising epitopes71, and B6-specific antibodies were shown to neutralise extracellular virus in a complement-dependent manner, significantly reducing viral load in lethal vaccinia mouse models. Additional work demonstrated that prophylactic administration of a B6R-specific monoclonal antibody (B7C9) within 24 h of lethal challenge reduced both viral burden and disease severity, further confirming its functional relevance87 Finally, complementary work with multivalent mRNA constructs incorporating B6R, A35R, and M1R reported 100% protection in mice against lethal vaccinia challenge32 highlighting the translational potential of this antigenic triad for surveillance and assay design. This triad may represent practical surrogate targets for MPXV exposure and could underpin the development of field-deployable multiplex assays to enhance detection of subclinical mpox in low-resource settings, building on recent advances88,89.
Our analysis was subject to limitations, and we note multiple caveats in our data interpretation. Firstly, genomic sampling was geographically skewed toward urban Southern Nigeria, likely under-representing rural transmission and although we assumed multiple underreporting factors in our genomic modelling, actual case detection rates could vary. Secondly, our population comprised of COVID-19 vaccinee cohorts, and their health-seeking behaviour may not fully reflect those of the wider population, introducing potential selection bias. Thirdly, our analysis measured only binding antibody and not functional neutralising antibody, although there is evidence supporting correlation between B6R, A35R, M1R and neutralisation. In addition, while our assay employed MPXV-specific antigens, the high degree of sequence similarity across Orthopoxvirus genus90 means that serological detection of anti-MPXV IgG cannot be attributed with absolute certainty to MPXV alone35. However, given the absence of a regional vaccinia vaccination program and the epidemiological context in Nigeria, recent MPXV exposure is the most plausible explanation in younger adults with legacy smallpox vaccination remaining the most likely explanation for reactivity in older adults. We note further that the utility of combining multiple antigens to define seropositivity and exposure type has been demonstrated91 We employed a slightly modified antigen panel adapted for the Luminex platform while ensuring overlap with validated and immunodominant antigenic targets. Our assay required reactivity to ≥4 of 6 MPXV antigens, a conservative approach relative to Yates and colleagues91 who used ≥3 of 5 antigens. This conservative approach, consistent with our prior work on hepatitis B serology in Africa was designed to minimise false positives90. Further, we supplemented our serological findings with follow-up interviews and professional verification of smallpox vaccination scars among seropositive HCWs, these data were limited to a small subset and are subject to recall bias. Nonetheless, the strong concordance between self-reported vaccination history, visible scars, and serological reactivity lends additional confidence to the interpretation of residual smallpox-derived immunity. Finally, structured collection of mpox-compatible clinical symptoms was not undertaken in the parent SARS-CoV-2 vaccine cohorts, which primarily captured SARS-CoV-2 vaccination events and COVID-like adverse events. Although none of the five participants with boosting responses reported mpox-like illness, we cannot exclude unrecognised or mild disease. The retrospective nature of the cohorts also limited assessment of behavioural or social drivers of exposure.
In conclusion, our study indicates that residual smallpox-derived immunity continues to shape the epidemiological landscape of mpox, with older adults born prior to the end of the smallpox eradication campaign retaining broad, multi-antigen reactive IgG profiles with younger adults showing a marked immunity gap. Evidence of antigen-specific antibody boosting in a small subset of ostensibly healthy participants suggests low-level MPXV exposure that symptoms-based surveillance is unlikely to detect65,91. Combining serology-based MPXV-specific targets such as B6R in combination with A35R and M1R may offer practical field-deployable tools91 for exposure surveillance and help inform targeted vaccination strategies, particularly in settings where systematic monitoring remains sparse. By combining serology with genomic and epidemiological analyses, we demonstrate that mpox transmission in Nigeria is characterised by slow epidemic growth, frequent dead-end infections, and modest transmission heterogeneity, consistent with the bottlenecks imposed by residual smallpox immunity. These findings suggest that while cryptic exposure occurs, large-scale expansion is constrained, highlighting the value of integrated sero-genomic approaches for understanding and monitoring epidemic persistence. Future work should (i) integrate multiplex assay with functional neutralisation testing and T-cell analysis to prospectively track immune dynamics following mpox vaccination with new third-generation mpox vaccines such as JYNNEOS to define real-world correlates of immunity (ii) explicitly evaluate the modifying impact of HIV infection and ART status92,93,94 on MPXV immune responses and (iii) evaluate the functional utility of fractional dosing strategies in real-world setting95.
Methods
All research conducted in this study complies with the relevant ethical regulations. Ethical approval was obtained from the Institutional Review Board of the Nigerian Institute of Medical Research (NIMR) (IRB-21-040) for the Lagos cohort and from the Federal Capital Territory Health Research Ethics Committee (FCT HREC) for the Abuja cohort (approval FHREC/2022/01/193/18-10-22). Written informed consent was obtained from all participants in the original Lagos and Abuja cohorts, and this consent covered the use of samples and data for the present secondary analysis.
Study population and sampling
We analysed convenience serum samples obtained from two previously established and published SARS-CoV-2 vaccine cohorts in Nigeria56,57 (Fig. 1a–b).
Cohort 1
Health care workers (HCWs) at the Nigerian Institute of Medical Research (NIMR) and Federal Medical Centre, Ebute Metta, were enroled in January 2021 as part of the NIMR SARS-CoV-2 vaccine effectiveness56. Participants received two-dose regimen of ChAdOx1 (AstraZeneca) during this period. All participants (n = 75) provided blood samples at baseline (T0, prior to vaccination; January 2021) and at follow-up (T1, October 2021). Paired sera were available for all participants (n = 75).
Cohort 2
HIV Negative adults were recruited in January at sites affiliated with Institute of Human Virology Nigeria (IHVN) in Abuja, Nigeria. Eligible participants were men and non-pregnant women >18 years old who had no previous SARS-CoV-2 vaccination and were confirmed HIV negative using the Nigerian national HIV rapid testing algorithm96,97. Recruitment was conducted through local community outreach and phone recall of registered patients. Participants received two-dose regimen of Ad26.COV.S (Johnson and Johnson) vaccine as part of the national vaccination programme. Blood samples were collected from (n = 101) at baseline (T0, prior to vaccination; January 2023) and at follow-up (T1, June 2023). Paired serum samples were available for 78/101 participants.
In total, 329 plasma samples (176 baseline and 153 follow-up) from 176 participants were included in the present analysis. Cohort demographics across both timepoints (T0 and T1) are shown in Supplementary Table 2, which also accounts for attrition between baseline and follow-up.
In addition, and to supplement serological analyses, we conducted follow-up interviews with a subset of healthcare worker (HCW) participants who were seropositive for MPXV antigens (n = 13). Telephone interviews were used to collect recall of smallpox vaccination history with consenting participants undergoing a brief clinical assessment by a trained healthcare professional to verify the presence of smallpox vaccination scars. Scar assessment was conducted privately, recorded as present or absent, and anonymised at data entry.
Laboratory methods and sample testing for MPXV-specific binding antibodies
To capture serological signatures of MPXV immunity, total IgG binding antibody to MPXV-specific antigens were measured in serum samples using a custom six-plex Luminex assay. Antigen selection was guided by prior work which identified A29L, H3L, M1R, A35R, B6R and D6L as core immunodominant antigens in response to infection and vaccination by mpox and vaccinia viruses98,99,100,101. This panel explicitly modelled on the multiplex approach91,102 while ensuring broad coverage and enable the application of a stringent conservative seropositivity threshold. The selected antigens represent distinct viral structural classes: A29L, H3L, and M1R associated with the intracellular mature virion (IMV) form; A35R and B6R associated with the extracellular enveloped virion (EEV); and D6L is a soluble viral protein. For this, six recombinant MPXV proteins were included: A29L (RayBiotech: Cat #230-30237), H3L (RayBiotech; Cat#230-30233), M1R (Sino Biological; Cat #40904-V07H), A35R (RayBiotech; Cat#230-30238), B6R (Sino Biological; Cat # 40902-V08H), and D6R (RayBiotech; Cat#230-01194) were covalently coupled to distinctive carboxylated bead sets (Luminex) to form a 6-plex assay103.
In addition, control analytes included BSA (negative binding control), IL-18BP (mpox-encoded virulence factor), LPS (positive binding control), and WHO MPXV reference standard (NIBSC 22/218). Briefly, beads were activated with 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide hydrochloride (Thermo Fisher Scientific) in the presence of N-hydroxysuccinimide (Thermo Fisher Scientific), according to the manufacturer’s instructions, to form amine-reactive intermediates. The activated bead sets were incubated with the corresponding proteins at a concentration of 50 μg ml−1 in the reaction mixture for 3 h at room temperature on a rotator. Beads were washed and stored in a blocking buffer (10 mM PBS, 1% BSA, 0.05% NaN3). Coupled bead sets were incubated with patient sera at 1/100 dilution for 1 h in 96-well filter plates (MultiScreenHTS; Millipore) at room temperature in the dark on a horizontal shaker. Fluids were aspirated with a vacuum manifold, and beads were washed three times with 10 mM PBS/0.05% Tween 20. Beads were incubated for 30 min with a PE-labelled anti-human IgG-Fc antibody (Leinco/Biotrend), washed as described above, and resuspended in 100 μl PBS/Tween. They were then analysed on a Luminex analyzer (Luminex/R&D Systems) using Exponent Software V31. Specific binding was reported as Mean Fluorescence Intensity. Cut-offs for antigen-specific reactivity were defined using infant samples (n = 14) from Nigeria (mothers were born after 1980, presumed MPXV-naïve). Antigen-specific thresholds were: A29L 3461.5, A35R 814.5, B6R 877.9, D6L 1718.6, H3L 1852.0, and M1R 1107.2. Specific binding was reported as Mean Fluorescence Intensity (Supplementary Fig. 4). We also report normalised Mean Fluorescence Intensity values, equal to, per antigen, the Mean Fluorescence Intensity divided by the relevant cut-off (i.e., values > 1 indicate reactivity and <1 non-reactivity).
Magnitude breadth analysis
To characterise the profile of MPXV-specific IgG responses across multiple antigens, we applied a magnitude-breadth analytical framework adapted from prior studies of polyclonal antibody responses in HIV104,105,106 and SARS-CoV-257,107 vaccine studies. This approach estimates the cumulative proportion of participants who retain detectable responses as antigen thresholds increase, effectively capturing both breadth (number of reactive targets) and magnitude (strength of signal) across the tested antigens. To compare distributions of magnitude-breadth scores visually, we used survival functions (also called complementary cumulative distribution functions)—these show the proportion of participants with scores above a given value for the full range of possible values. Curves were stratified by participant groups based on occupational exposure, birth cohort and MPXV exposure status. Groups of participants with higher magnitude responses across more antigens have a right-shifted curve, reflecting greater immunological breadth. Boxplots of magnitude-breadth scores are shown next to the curves. Differences in distributions were assessed using Mann-Whitney U tests.
Genomic dataset curation
We obtained 185 high-quality Clade IIb MPXV genomic sequences previously deposited to GenBank. This included all Clade IIb genomes from 2017 to 2022 sampled in Nigeria, sequences from non-endemic countries with a travel history to Nigeria. To select the sequences for the estimation of the cluster size and dispersion parameter we removed sequences sampled before 2021. We used 185 sequences to estimate for the sky grid model and 105 sequences for the cluster size estimation.
Transmission heterogeneity estimation of Nigerian mpox spread
To estimate the effective reproduction number (R) of mpox, we adapted the renewal equation framework62. Within this framework, the time varying effective reproduction number60 (defined as the average number of secondary infections generated by one infected individual) was modelled using a fourth-order spline with five evenly spaced knots. We assumed a discretized gamma distribution for the generation interval, (mean of 12.6 days and SD of 5.7 days)61. The observed case counts from the Nigeria CDC were fit using a Poisson likelihood to derive posterior estimates of R59.
To analyse the population dynamics of the mpox epidemic in Nigeria, we utilised the 185 genomic data obtained from GenBank8, to reconstruct epidemic dynamics using the BEAST software package v.10.57,8,92,93,94,108. Our analysis employed a non-parametric skygrid coalescent tree prior, incorporating 96 change points distributed over 10 years, as described108. We combined two independent Markov chain Monte Carlo (MCMC) runs, each consisting of 100 million states, with sampling occurring every 10,000 states. We used tracer to access that all parameters effective sample size values were above 200, as well as export the summary of the effective population size across the tree posterior. We have deposited the curated sequence list, metadata, and BEAST XMLs at: https://github.com/Ifeanyi-omah/Sero-Genomics_mpox.
To investigate whether a small number of individuals disproportionately contributed to transmission, we estimated the overdispersion parameter using the method outlined in Tran-Kiem and Bedford55, applying it to a subset of 105 Nigerian sequences generated from 2021 to 2023 (second phase of the epidemic). We used a probability of transmission before mutation of 0.6659. Given that ≈7% of confirmed cases during this study period were sequenced, we then used sampling proportions (the proportion of total cases represented by the genomic sample) 0.003510, 0.007021, 0.035104, 0.070209 which corresponds to case ascertainment rates of 5%, 10%, 50%, and 100% respectively. Given the APOBEC3-mediated mutations of mpox elevating the mutation rate, we specifically used identical sequences because they indicate very recent transmission events7,8 However, interpreting identical sequences must be approached cautiously and we acknowledge that identical sequences might represent closely linked transmission chains but may also mask subtle diversification occurring over slightly longer timescales of the genomic data.
Definition of terms and statistical analysis
We compositely defined MPXV exposure as a ≥ 2-fold increase in total binding antibody magnitude between baseline and follow-up, in combination with a ≥ 2-fold increase in antibody levels to at least 4/6 MPXV-specific antigens tested. This definition was designed to capture both the magnitude and breadth of the humoral immune response, providing a stringent immunological marker of recent exposure to MPXV or a closely related Orthopoxviruses and was informed by prior work in West African cohorts109 demonstrating that reliance on single antibody markers substantially underestimates true exposure and that multi-marker or functional antibody approaches offer superior discriminatory accuracy. Baseline characteristics of participants were summarised as counts and percentages for categorical variables, and medians with interquartile ranges (IQR) for continuous variables. Antibody responses were described using both the antigen-specific antibody levels (raw and normalised) and overall magnitude-breadth scores. Central tendencies were described by geometric mean titres (GMTs). Distributions of continuous measures between two groups were compared using the Mann-Whitney U test. Distributions were visualised using box plots (showing quartiles), with individual data points also shown; and survival functions (also called complementary cumulative distribution functions) which describe proportions of participants (y-axis) with measurements above each value (x-axis) in the range of possible values. All statistical analyses and data processing were performed by Python or R (version 4.2.2) or GraphPad Prism (v10.4.1).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated or analysed during this study are included in this published article and its Supplementary Information files. Source data for all figures and tables are provided with this paper and genomic dataset is available: https://github.com/Ifeanyi-omah/Sero-Genomics_mpox. Source data are provided with this paper.
References
Lu, J. et al. Mpox (formerly monkeypox): pathogenesis, prevention and treatment. Signal Transduct. Target Ther. 8, 458 (2023).
Lum, F.-M. et al. Monkeypox: disease epidemiology, host immunity and clinical interventions. Nat. Rev. Immunol. 22, 597–613 (2022).
Happi, C. et al. Urgent need for a non-discriminatory and non-stigmatizing nomenclature for monkeypox virus. PLoS Biol. 20, e3001769 (2022).
Zumla, A. et al. The 2024 public health emergency of international concern: a global failure to control Mpox. Am. J. Trop. Med Hyg. 112, 17–20 (2025).
Djuicy, D. D. et al. Molecular epidemiology of recurrent zoonotic transmission of mpox virus in West Africa. medRxiv 2024.06.18.24309115 https://doi.org/10.1101/2024.06.18.24309115 (2024).
Ndodo, N. et al. Distinct monkeypox virus lineages co-circulating in humans before 2022. Nat. Med 29, 2317–2324 (2023).
O’Toole, Á et al. APOBEC3 deaminase editing in mpox virus as evidence for sustained human transmission since at least 2016. Science (1979) 382, 595–600 (2023).
Parker, E. et al. Genomics reveals zoonotic and sustained human Mpox spread in West Africa. Nature https://doi.org/10.1038/s41586-025-09128-2 (2025).
Bunge, E. M. et al. The changing epidemiology of human monkeypox—A potential threat? A systematic review. PLoS Negl. Trop. Dis. 16, e0010141 (2022).
Yinka-Ogunleye, A. et al. Outbreak of human monkeypox in Nigeria in 2017–18: a clinical and epidemiological report. Lancet Infect. Dis. 19, 872–879 (2019).
Isidro, J. et al. Phylogenomic characterization and signs of microevolution in the 2022 multi-country outbreak of monkeypox virus. Nat. Med. 28, 1569–1572 (2022).
Sumner, R. P. et al. Attenuation of the 2022 global outbreak monkeypox virus relative to its clade IIb ancestor. bioRxiv 2025.02.27.640535 https://doi.org/10.1101/2025.02.27.640535 (2025).
Mizushima, D. et al. Prevalence of asymptomatic mpox among men who have sex with men, Japan, January–March 2023. Emerg. Infect. Dis. J. 29, 1872 (2023).
Accordini, S., Cordioli, M., Pomari, E., Tacconelli, E. & Castilletti, C. People with asymptomatic or unrecognised infection potentially contribute to monkeypox virus transmission. Lancet Microbe 4, e209 (2023).
Agustí, C. et al. Self-sampling monkeypox virus testing in high-risk populations, asymptomatic or with unrecognized Mpox, in Spain. Nat. Commun. 14, 5998 (2023).
Edouard, S. et al. Incidental diagnosis of mpox virus infection in patients undergoing sexually transmitted infection screening—findings from a study in France. Int. J. Infect. Dis. 143, 107009 (2024).
De Baetselier, I. et al. Retrospective detection of asymptomatic monkeypox virus infections among male sexual health clinic attendees in Belgium. Nat. Med. 28, 2288–2292 (2022).
Ferré, V. M. et al. Detection of monkeypox virus in anorectal swabs from asymptomatic men who have sex with men in a sexually transmitted infection screening program in Paris, France. Ann. Intern Med 175, 1491–1492 (2022).
Hampel, B. et al. Low prevalence of asymptomatic mpox in populations at high risk. Lancet Microbe 4, e856 (2023).
Mouro, M. & Duque, V. Asymptomatic transmission of human mpox. BMJ Case Rep. 17, e254305 (2024).
Matusali, G. et al. Asymptomatic Mpox virus infection in subjects presenting for MVA-BN vaccine. Clin. Infect. Dis. 77, 1483–1484 (2023).
Henry, S. et al. Serological response to Mpox and direct virus detection in asymptomatic patient prior to the first diagnosed case: a retrospective study of the 2022 montpellier epidemic. J. Med Virol. 97, e70365 (2025).
Ogale, Y. P. et al. Evidence of Mpox Virus Infection Among Persons Without Characteristic Lesions or Rash Presenting for First Dose of JYNNEOS Vaccine—District of Columbia, August 2022. Clin. Infect. Dis. 77, 298–302 (2023).
Reda, A., El-Qushayri, A. E. & Shah, J. Asymptomatic monkeypox infection: a call for greater control of infection and transmission. Lancet Microbe 4, e15–e16 (2023).
Taube, J. C., Rest, E. C., Lloyd-Smith, J. O. & Bansal, S. The global landscape of smallpox vaccination history and implications for current and future orthopoxvirus susceptibility: a modelling study. Lancet Infect. Dis. 23, 454–462 (2023).
Ge, Y. et al. Associations between the 2022 global mpox outbreak and multifaceted factors: A multi-geographical retrospective study. One Health 101224 https://doi.org/10.1016/j.onehlt.2025.101224 (2025).
Rimoin, A. W. et al. Major increase in human monkeypox incidence 30 years after smallpox vaccination campaigns cease in the Democratic Republic of Congo. Proc. Natl. Acad. Sci. 107, 16262–16267 (2010).
Gilchuk, I. et al. Cross-Neutralizing and Protective Human Antibody Specificities to Poxvirus Infections. Cell 167, 684–694.e9 (2016).
Freyn, A. W. et al. An mpox virus mRNA-lipid nanoparticle vaccine confers protection against lethal orthopoxviral challenge. Sci. Transl. Med 15, eadg3540 (2025).
Mucker, E. M. et al. Comparison of protection against mpox following mRNA or modified vaccinia Ankara vaccination in nonhuman primates. Cell 187, 5540–5553.e10 (2024).
Mengjun, L. et al. Bispecific antibodies targeting MPXV A29 and B6 demonstrate efficacy against MPXV infection. J. Virol. 0, e02320–e02324 (2025).
Kong, T. et al. Single-chain A35R-M1R-B6R trivalent mRNA vaccines protect mice against both mpox virus and vaccinia virus. EBioMedicine 109, (2024).
Tang, D. et al. Recombinant proteins A29L, M1R, A35R, and B6R vaccination protects mice from mpox virus challenge. Front Immunol Volume 14-2023, (2023).
Yu, H., Resch, W. & Moss, B. Poxvirus structural biology for application to vaccine design. Trends Immunol. https://doi.org/10.1016/j.it.2025.04.002 (2025).
Pauli, G. et al. Orthopox viruses: Infections in humans. Transfus. Med. hemotherapy 37, 351–364 (2010).
Manenti, A. et al. Evaluation of Monkeypox- and Vaccinia virus-neutralizing antibodies in human serum samples after vaccination and natural infection. Front Public Health Volume 11-2023, (2023).
WHO. Smallpox Eradication Programme - SEP (1966-1980). (2010).
Foege, W. Lessons and innovations from the West and Central African Smallpox Eradication Program. Vaccine 29, D10–D12 (2011).
WHO. Smallpox, Historic Milestone Underscores Urgent Need to Invest in Global Health Security and Universal Health Coverage.
Haeuser, E. et al. Global, regional, and national trends in routine childhood vaccination coverage from 1980 to 2023 with forecasts to 2030: a systematic analysis for the Global Burden of Disease Study 2023. The Lancet https://doi.org/10.1016/S0140-6736(25)01037-2 (2025).
Rosenthal, S., Merchlinsky, M., Kleppinger, C. & Goldenthal, K. Developing New Smallpox. Vaccines. Emerg. Infect. Dis. J. 7, 920 (2001).
Fenner, F. et al. Smallpox and Its Eradication (World Health Organization, 1988). https://iris.who.int/items/ba4ab312-1c43-4304-8235-969979499717.
Jacobs, B. L. et al. Vaccinia virus vaccines: Past, present and future. Antivir. Res 84, 1–13 (2009).
Edghill-Smith, Y. et al. Smallpox vaccine–induced antibodies are necessary and sufficient for protection against monkeypox virus. Nat. Med 11, 740–747 (2005).
Sanz-Muñoz, I. et al. Possible Mpox protection from smallpox. Vaccin.–Generated Antibodies Older Adults. Emerg. Infect. Dis. J. 29, 656 (2023).
Greenberg, R. N. et al. A randomized, double-blind, placebo-controlled phase II trial investigating the safety and immunogenicity of modified vaccinia ankara smallpox vaccine (MVA-BN®) in 56-80-Year-Old Subjects. PLoS ONE 11, e0157335 (2016).
Zaeck, L. M. et al. Low levels of monkeypox virus-neutralizing antibodies after MVA-BN vaccination in healthy individuals. Nat. Med. 29, 270–278 (2023).
Priyamvada, L. et al. Serological responses to the MVA-based JYNNEOS monkeypox vaccine in a cohort of participants from the Democratic Republic of Congo. Vaccine 40, 7321–7327 (2022).
Marchi, S. et al. Evaluation of monkeypox- and vaccinia-virus neutralizing antibodies before and after smallpox vaccination: a sero-epidemiological study. J. Med. Virol. 96, e29728 (2024).
Hubert, M. et al. Complement-dependent mpox-virus-neutralizing antibodies in infected and vaccinated individuals. Cell Host Microbe 31, 937–948.e4 (2023).
Galetti, S. C. et al. Historic smallpox vaccination and Mpox cross-reactive immunity: Evidence from healthcare workers with childhood and adulthood exposures. Vaccine 46, 126661 (2025).
Li, E. et al. Duration of humoral immunity from smallpox vaccination and its cross-reaction with Mpox virus. Signal Transduct. Target Ther. 8, 350 (2023).
Marchi, S. et al. Correlation between microneutralization test and a multiplexed immunoassay for evaluation of monkeypox and vaccinia virus antibodies before and after smallpox vaccination. Front Immunol. ume, 16–2025 (2025).
Wayengera, M. et al. Mpox epidemics: a call to restore humanity’s lost herd immunity to Orthopoxviruses. Viruses 17, (2025).
Satapathy, P. et al. Potentially asymptomatic infection of Monkeypox virus: A systematic review and meta-analysis. Vaccines (Basel) 10, 2083 (2022).
Abdullahi, A. et al. SARS-COV-2 antibody responses to AZD1222 vaccination in West Africa. Nat. Commun. 13, 6131 (2022).
Abdullahi, A. et al. SARS-CoV-2 infection imprints neutralising antibody responses in the absence of vaccination. ResearchSquare https://doi.org/10.21203/rs.3.rs-6347062/v1 (2025).
Ruis, C. et al. A systematic nomenclature for mpox viruses causing outbreaks with sustained human-to-human transmission. Nat. Med. 31, 2854–2858 (2025).
Paredes, M. I. et al. Underdetected dispersal and extensive local transmission drove the 2022 mpox epidemic. Cell 187, 1374–1386.e13 (2024).
Figgins, M. D. & Bedford, T. Inferring variant-specific effective reproduction numbers from combined case and sequencing data. medRxiv 2021.12.09.21267544 https://doi.org/10.1101/2021.12.09.21267544 (2024).
Guzzetta, G. et al. The decline of the 2022 Italian mpox epidemic: Role of behavior changes and control strategies. Nat. Commun. 15, 2283 (2024).
Tran-Kiem, C. & Bedford, T. Estimating the reproduction number and transmission heterogeneity from the size distribution of clusters of identical pathogen sequences. Proc. Natl. Acad. Sci. USA 121, e2305299121 (2024).
Henderson, D. A. The eradication of smallpox – An overview of the past. Future Vaccin. 29, D7–D9 (2011).
Crandell, J. et al. The impact of orthopoxvirus vaccination and Mpox infection on cross-protective immunity: a multicohort observational study. Lancet Microbe https://doi.org/10.1016/j.lanmic.2025.101098 (2025).
Mitjà, O., Watson-Jones, D., Choi, E. M., Jalloh, M. B. & Sahr, F. Clade IIb mpox outbreak in Sierra Leone. Lancet 405, 2274–2275 (2025).
Ennis, F. A., Cruz, J., Demkowicz, W. E. Jr, Rothman, A. L. & McClain, D. J. Primary induction of human CD8+ cytotoxic T lymphocytes and interferon-γ-producing T cells after smallpox vaccination. J. Infect. Dis. 185, 1657–1659 (2002).
Orr, N. et al. Clinical and immune responses after revaccination of israeli adults with the lister strain of vaccinia virus. J. Infect. Dis. 190, 1295–1302 (2004).
Hammarlund, E. et al. Duration of antiviral immunity after smallpox vaccination. Nat. Med 9, 1131–1137 (2003).
Hsieh, S.-M., Pan, S.-C., Chen, S.-Y., Huang, P.-F. & Chang, S.-C. Age distribution for T cell reactivity to vaccinia virus in a healthy population. Clin. Infect. Dis. 38, 86–89 (2004).
Taub, D. D. et al. Immunity from smallpox vaccine persists for decades: a longitudinal study. Am. J. Med 121, 1058–1064 (2008).
Zhao, R. et al. Two noncompeting human neutralizing antibodies targeting MPXV B6 show protective effects against orthopoxvirus infections. Nat. Commun. 15, 4660 (2024).
Giorgi, F. M., Pozzobon, D., Di Meglio, A. & Mercatelli, D. Genomic and transcriptomic analysis of the recent Mpox outbreak. Vaccine 42, 1841–1849 (2024).
Crossley, L. et al. Susceptibility and transmission of mpox virus infection in brown rats (Rattus norvegicus). J. Gen. Virol. 106, (2025).
Falendysz, E. A., Lopera, J. G., Rocke, T. E. & Osorio, J. E. Monkeypox virus in animals: Current knowledge of viral transmission and pathogenesis in wild animal reservoirs and captive animal models. Viruses 15, 905 (2023).
Forni, D., Cagliani, R., Molteni, C., Clerici, M. & Sironi, M. Monkeypox virus: the changing facets of a zoonotic pathogen. Infect., Genet. Evolution 105, 105372 (2022).
Patrono, L. V. et al. Monkeypox virus emergence in wild chimpanzees reveals distinct clinical outcomes and viral diversity. Nat. Microbiol. 5, 955–965 (2020).
Clark, J. et al. Domestic risk factors for increased rodent abundance in a Lassa fever endemic region of rural Upper Guinea. Sci. Rep. 11, 20698 (2021).
Olayemi, A. et al. New hosts of the lassa virus. Sci. Rep. 6, 25280 (2016).
Agbonlahor, D. E., Akpede, G. O., Happi, C. T. & Tomori, O. 52 years of lassa fever outbreaks in Nigeria, 1969–2020: an epidemiologic analysis of the temporal and spatial trends. Am. J. Trop. Med Hyg. 105, 974–985 (2021).
Bonwitt, J. et al. At Home with Mastomys and Rattus: Human-Rodent Interactions and Potential for Primary Transmission of Lassa Virus in Domestic Spaces. Am. Soc. Tropical Med. Hyg. 96, 935–943 (2017).
Doohan, P. et al. Lassa fever outbreaks, mathematical models, and disease parameters: a systematic review and meta-analysis. Lancet Glob. Health 12, e1962–e1972 (2024).
Mariën, J. et al. Serosurveillance Identifies an Endemic Hotspot of Lassa Fever in Faranah, Upper Guinea. J. Infect. Dis. jiaf308 https://doi.org/10.1093/infdis/jiaf308 (2025).
Garry, R. F. Lassa fever - the road ahead. Nat. Rev. Microbiol. 21, 87–96 (2023).
Herrera, B. B. et al. Pre-pandemic cross-reactive antibody and cellular responses against SARS-CoV-2 among female sex workers in Dakar, Senegal. Front Public Health 13, 1522733 (2025).
Ceccarelli, G., Francesco, B., Fabio, S., Marta, G. & Ciccozzi, M. Helminthiasis and mpox vaccination: challenges in Sub-Saharan Africa. Infect. Dis. 57, 301–303 (2025).
Cohen, M. E., Xiao, Y., Eisenberg, R. J., Cohen, G. H. & Isaacs, S. N. Antibody against extracellular vaccinia virus (EV) protects mice through complement and Fc receptors. PLoS ONE 6, e20597 (2011).
Ren, Z. et al. Identification of mpox M1R and B6R monoclonal and bispecific antibodies that efficiently neutralize authentic mpox virus. Emerg. Microbes Infect. 13, 2401931 (2024).
Laidlaw Stephen et al. Detection of mpox and other orthopoxviruses using a lateral flow device as a point-of-care diagnostic. Microbiol Spectr. 13, e02456–24 (2025).
Cavuto, M. L. et al. Portable molecular diagnostic platform for rapid point-of-care detection of mpox and other diseases. Nat. Commun. 16, 2875 (2025).
Abdullahi, A. et al. Hepatitis B Virus (HBV) infection and re-activation during Nucleos(t)ide reverse transcriptase inhibitor–sparing antiretroviral therapy in a high–HBV endemicity setting. Open Forum Infect. Dis. 5, ofy251 (2018).
Yates, J. L. et al. Development of a novel serological assay for the detection of mpox infection in vaccinated populations. J. Med Virol. 95, e29134 (2023).
Kayuni, S. A. et al. Prospective pilot study on the relationship between seminal HIV-1 shedding and genital schistosomiasis in men receiving antiretroviral therapy along Lake Malawi. Sci. Rep. 13, 14154 (2023).
Abdullahi, A. et al. Limited emergence of resistance to integrase strand transfer inhibitors (INSTIs) in ART-experienced participants failing dolutegravir-based antiretroviral therapy: a cross-sectional analysis of a Northeast Nigerian cohort. J. Antimicrobial Chemother. 78, 2000–2007 (2023).
Abdullahi, A. et al. A detailed characterization of drug resistance during darunavir/ritonavir monotherapy highlights a high barrier to the emergence of resistance mutations in protease but identifies alternative pathways of resistance. J. Antimicrobial Chemother. 79, 339–348 (2024).
Salako, A. et al. Safety and immunogenicity of fractional COVID-19 vaccine doses in Nigerian adults: a randomized non-inferiority trial. Sci. Rep. 15, 27614 (2025).
FMOH. Nigeria: National Guidelines for HIV Counselling and Testing. https://www.medbox.org/document/nigeria-national-guidelines-for-hiv-counselling-and-testing (2011).
Iriemenam, N. C. et al. Evaluation of the Nigeria national HIV rapid testing algorithm. PLOS Glob. Public Health 2, e0001077 (2022).
Cohn, H. et al. Mpox vaccine and infection-driven human immune signatures: an immunological analysis of an observational study. Lancet Infect. Dis. 23, 1302–1312 (2023).
Otter, A. D. et al. Monkeypox virus-infected individuals mount comparable humoral immune responses as Smallpox-vaccinated individuals. Nat. Commun. 14, 5948 (2023).
Jones, S. et al. Assessment of MpoxPlex, a high-throughput and multiplexed immunoassay: a diagnostic accuracy study. Lancet Microbe 6, (2025).
Fantin, R. F. & Coelho, C. H. Human antibody responses to circulating monkeypox virus emphasise the need for the first mpox-specific vaccine. Lancet Microbe 5, e204–e205 (2024).
Masirika, L. M. et al. Serological evidence of clade Ib Mpox transmission by sex workers and within household in South Kivu, DRC. Nat. Commun. 16, 7056 (2025).
Xiong, X. et al. A thermostable, closed SARS-CoV-2 spike protein trimer. Nat. Struct. Mol. Biol. 27, 934–941 (2020).
Montefiori, D. C. et al. Magnitude and breadth of the neutralizing antibody response in the RV144 and Vax003 HIV-1 vaccine efficacy trials. J. Infect. Dis. 206, 431–441 (2012).
Huang, Y., Gilbert, P. B., Montefiori, D. C. & Self, S. G. Simultaneous evaluation of the magnitude and breadth of a left- and right-censored multivariate response. Application HIV Vaccin. Dev. Stat. Biopharm. Res. 1, 81–91 (2009).
Gilbert, P. et al. Magnitude and breadth of a nonprotective neutralizing antibody response in an efficacy trial of a candidate HIV-1 gp120 vaccine. J. Infect. Dis. 202, 595–605 (2010).
Einhauser, S. et al. Longitudinal effects of SARS-CoV-2 breakthrough infection on imprinting of neutralizing antibody responses. EBioMedicine 110, 105438 (2024).
Gill, M. S. et al. Improving Bayesian population dynamics inference: a coalescent-based model for multiple loci. Mol. Biol. Evol. 30, 713–724 (2013).
Abdullahi, A. et al. Performance of SARS COV-2 IgG Anti-N as an independent marker of exposure to SARS COV-2 in an unvaccinated west African population. Am. J. Trop. Med Hyg. 109, 890–894 (2023).
Acknowledgements
We wish to thank the study participants who made this study possible. A.A. was supported by Cambridge-Africa award and Harvard Takemi Program in International Health and acknowledges receipt of support from Charité Centre for Global Health, Institute of International Health, Berlin, Germany through a public health intelligence fellowship. I.O was supported by the Wellcome Trust Hosts, Pathogens & Global Health program [Wellcome Trust, Grant number 218471/Z/19/Z] in partnership with Tackling infectious Disease to Benefit Africa. This research was funded in whole, or in part, by the Wellcome Strategic Core award: 227167/A/23/Z. For the purpose of open access, the author has applied a CC BY public copyright licence to any Author Accepted Manuscript version arising from this submission. RKG was supported by a Wellcome Trust Senior fellowship (WT108082AIA). This research was supported by the Hong Kong Jockey Club Global Health Institute (HKJCGHI), Hong Kong Special Administrative Region, China. This research was supported by the NIHR Cambridge Biomedical Research Centre (NIHR203312).
Author information
Authors and Affiliations
Contributions
Study conceptualization and design: A.A. and R.K.G. Methodology and investigation: A.A., I.O., R.K. F.I., M.E., C.K.K., S.O., H.W., E.J.O., E.P., O.F., A.M.I., U.A.S., L.C.G., E.O., A.H., O.S., R.A., S.H.A., M.C., C.H., R.D., B.L.S. Research data curation & analysis: A.A., I.O., R.K., M.E., S.O., A.M.I., L.C.G., O.S., C.H., R.D. Funding acquisition: A.A., B.L.S., A.A.b. and R.K.G. Writing (original draft): A.A. and I.O. Writing (review & editing): All authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Yap Boum, Miguel Paredes and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Abdullahi, A., Omah, I., Kassanjee, R. et al. Sero-genomic evidence for occult mpox exposure in healthy Nigerian adults. Nat Commun 17, 482 (2026). https://doi.org/10.1038/s41467-026-68335-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68335-1