Abstract
Plasma amyloid-β (Aβ) peptides, alone or in ratio with p-tau217, show strong potential as Alzheimer’s disease biomarkers. While immunoprecipitation-mass spectrometry (IP-MS) is the preferred method for plasma Aβ quantification, current assays are resource- and time-intensive. Here, we developed a streamlined IP-MS method using a cost-effective instrument that significantly improved the efficiency of an original assay by incorporating a single immunoprecipitation step, an optimized buffer system, and approximately 75% reductions in antibody and sample volume requirements. Technical validation revealed excellent dilution linearity (r²>0.99), high precision (< 10% variation), enhanced sensitivity, improved Aβ recovery, and markedly increased signal-to-noise ratios. In a large cohort of cognitively normal older adults (n = 317), the plasma Aβ1-42/Aβ1-40 ratio achieved stronger concordance with Aβ-PET and superior accuracies to identify abnormal scans (AUC 0.81 vs. 0.65 for the original assay). Notably, accuracies remained high even with plasma volumes as low as 100 μL. The improved IP-MS method enables robust and simplified plasma Aβ assessment in Alzheimer’s disease, with implications for prognosis, diagnosis and intervention trials.
Similar content being viewed by others
Introduction
Brain amyloid-β (Aβ) plaque deposition is a pathological hallmark, an antemortem diagnostic criterion, and a potential therapeutic target for Alzheimer’s disease (AD)1,2,3,4. Following the recent approval of anti-Aβ monoclonal antibody therapies for AD by the Food and Drug Administration (FDA), the importance of reliable yet accessible Aβ biomarkers for therapeutic trial selection and monitoring, as well as for routine clinical diagnostic and prognostic purposes continues to increase5,6,7. While positron emission tomography (PET) imaging of Aβ plaques, and cerebrospinal fluid (CSF) measurements of Aβ42/40 peptide ratio via immunoassays are widely used and accepted brain Aβ deposition biomarkers, their routine uses can be limited by high costs, invasiveness, and lack of widespread availability outside of major medical centers8,9,10.
High-performance blood-based biomarkers that accurately reflect Aβ pathology would offer minimally invasive yet affordable options for prognosis, clinical management, and to evaluate target engagement in therapeutic trial programs11,12,13,14,15. Plasma Aβ1-42/1-40 and p-tau217 are two of the leading plasma biomarker candidates for Aβ pathology, with their combined utility reflected in the plasma p-tau217/Aβ1-42 assay that has recently received FDA clearance for clinical use16,17,18,19. Plasma p-tau217 assays from different sources have shown high diagnostic utility and strong cross-assay correlations, suggesting that many could be interchangeable20,21. However, the same cannot be said for plasma Aβ assays, many of which show poor correlations and classification accuracies13. Plasma Aβ assays mostly employ either immunoassay or immunoprecipitation-mass spectrometry (IP-MS) methods8,22. The susceptibility of immunoassays to non-specific interference can compromise accuracy, particularly at the low plasma Aβ peptide levels typically found in early AD23. Conversely, sensitive IP-MS assays have demonstrated significantly stronger accuracies in distinguishing individuals with and without brain Aβ plaque pathology, evidenced by decreased plasma Aβ1-42/1-40 ratio8,23. Yet, the available IP-MS assays for plasma Aβ tend to be labor, sample, and resource intensive, requiring highly trained scientists with access to large sample volumes (often 1 mL and above) and state-of-art mass spectrometry instruments. These drawbacks affect the potential for assay scalability and widespread adoption, given the multiple sample processing steps that need to be meticulously followed and standardized8.
We previously adapted the IP-MS assay originally developed by Nakamura et al.11, referred to here as the Pittsburgh plasma Aβ assay version 1.0 (PAβ V1.0)18,24,25, which was implemented on a cost-effective benchtop MALDI-TOF instrument, the Bruker Microflex LT. However, considering the limited dynamic range of MS, the presence of strong background interference, including albumin and immunoglobulins, could impede the accurate detection of plasma Aβ peptides18,22,24,25,26. To address this limitation, the V1.0 assay used two rounds of immunoprecipitation (IP). Regrettably, this approach increases per-sample costs, reagent usage, and sample preparation time.
Here, we describe an improved IP-MS assay, referred to as the Pittsburgh Plasma Aβ assay version 2.0 (PAβ V2.0). This enhanced assay version streamlines the two rounds of IP into a single step, thereby significantly reducing pre-processing time, reagent and consumable needs. In addition, the modifications substantially increase the signal-to-noise ratio, allowing for only a small sample volume of 250 μL compared to ≥1 mL used by other IP-MS assays. We present the analytical performance of the PAβ V2.0 assay in comparison to the first-generation PAβ V1.0 method and immunoassay alternatives on the Quanterix Simoa platform. In addition, we tested the feasibility of further reducing sample volume from 250 μL to 100 μL. Finally, the clinical utility of the assay was evaluated in samples from three different cohorts, including two that exclusively recruited cognitively unimpaired participants, some of whom had preclinical evidence of AD.
Results
Comparison of detergents and blocking buffers to improve assay signal-to-noise ratio
To streamline the PAβ V1.0 assay into a single IP step, we experimented with various supplements in the IP binding buffer to reduce background interference. These included 10% v/v N4PE CSF sample diluent, 10% v/v SuperBlock, 10 μg/ml TruBlock, 0.5% v/v Triton100, and 0.5% Tween20, all tested following the Single IP protocol (see the Materials and Methods section).
Among the buffer systems tested, the N4PE CSF diluent demonstrated the best performance, effectively eliminating interference peaks while maintaining the highest signal-to-noise ratio. Use of the N4PE CSF diluent, but not with the other supplements, distinctively led to the elimination of the interference peak at 4153 mass-to-charge (m/z), which often obscures the Aβ1-38 internal standard (IS) and Aβ3-40 signals in the PAβ V1.0 assay (Supplementary Figs. 1, 2). Notably, SuperBlock and TruBlock resulted in worse signal-to-noise ratios than the original PAβ V1.0 assay, while the detergents, on the other hand, gave the lowest signal-to-noise ratios for all Aβ peptides (Supplementary Figs. 1, 2).
We further compared the single IP setup utilizing the N4PE CSF diluent with the original two IP-step PAβ V1.0 method, as well as with a single-IP configuration employing the same binding buffer as the original assay. As shown in the representative spectra (Fig. 1A), supplementing the IP binding buffer with the N4PE CSF diluent resulted in the cleanest spectra. The interference peaks observed in the single-step PAβ V1.0 assay at 3200 m/z to 3500 m/z, and at 6400 m/z and 6600 m/z, were reduced by using the N4PE CSF diluent. The remaining residual peak at 3900 m/z does not impact Aβ measurement due to its considerable separation in m/z. Furthermore, the single-step IP procedure using N4PE CSF diluent achieved a significantly higher signal-to-noise ratio, with mean values of 143.9 for Aβ1-40 and 9.5 for Aβ1-42, compared with 72.4 and 5.5, respectively for the original PAβ V1.0 assay, and 23.9 and 1.6, respectively for the PAβ V1.0 assay with single-step IP configuration (Figs. 1B, C). Similar improvements were observed for other Aβ peptides, including Aβ1-38, Aβ3-40, Aβ1-39, and APP669-711 (Fig. 1B, C).
A MALDI–TOF mass spectra of Aβ peptides derived from plasma samples utilizing the PAβ V1.0 assay original setup and buffers, 10% N4PE CSF diluent (the buffer system eventually selected for the PAβ V2.0 assay) or the original PAβ V1.0 assay modified to incorporate a single step of IP. Representative spectra from each experiment are presented. Apparent interference peaks were observed at 5771.1 m/z and 7746.8 m/z across all assays. In addition, another apparent interference peak at 6631.0 m/z was noted in all assay formats except the PAβ V1.0 assay. Interference peaks at 3200 m/z to 3500 m/z, and 6432.4 m/z were observed in the PAβ V1.0 assay with 1IP only. Nonetheless, the wide separation between these peaks and characteristic peaks for the Aβ peptides of interest (in the range of 4000–4850 m/z), presents no known challenge presently. The theoretical m/z values of the Aβ peptides are as follows: 4132.6 m/z for Aβ1–38, 4144.7 m/z for Aβ3-40, 4231.8 m/z for Aβ1-39, 4330.9 m/z for Aβ1–40, 4515.1 m/z for Aβ1–42, and 4689.4 m/z for APP669–711. Aβ1–38 IS at 4160.7 m/z, Aβ1–40 intern at 4383.3 m/z, and Aβ1–42 IS at 4569.3 m/z were utilized as ISs for the normalization of mass spectra. Notably, an interference peak was detected at 4153.4 m/z in samples processed using the original PAβ V1.0 assay with a single IP, but not in the other assays. B Signal to noise (S/N) ratios were compared across the three assay setups in triplicates, with asterisks indicating significant differences (*p = 0.032, **p = 0.008) as determined by the two-sided Wilcoxon Rank Sum test. Data were presented as mean values +/− standard deviation. C The averages and standard deviations of the S/N ratios are listed.
Due to its superior performance, we selected the single IP with N4PE CSF diluent-supplemented binding buffer as the improved version of the PAβ V1.0 assay, hereafter referred to as the Pittsburgh plasma Aβ version 2.0 method (PAβ V2.0 assay).
Analytical performance
We evaluated the analytical performance of the PAβ V2.0 assay relative to the original PAβ V1.0 method.
To assess assay linearity, we constructed standard curves using two-fold serial dilutions of a mixture of synthetic Aβ1-40 and Aβ1-42 in 6% bovine serum albumin (BSA)/phosphate-buffered saline (PBS). Separate seven-sample dilution series containing varying concentrations of Aβ1-40 (0-400 pg/ml) and Aβ1-42 (0-100 pg/ml) were included. The measured Aβ1-40 and Aβ1-42 peak areas were normalized using the Aβ1-38 IS or the analyte-specific IS (separately for Aβ1-40 and Aβ1-42) (Supplementary Fig. 3). Both the PAβ V1.0 and the PAβ V2.0 assay formats exhibited robust linearity across the tested concentration ranges with r2 > 0.99 for Aβ1-40 and Aβ1-42.
The inter-assay coefficients of variation (CVs) for both Aβ1-40 and Aβ1-42 were below 20% at their lowest non-zero concentrations—12.5 pg/mL for Aβ1-40 and 3.125 pg/mL for Aβ1-42. Accordingly, we established these values as the lower limits of quantification (LLOQ). In addition, since the linearity extended to the calibrator with the highest concentrations, the upper limits of quantification (ULOQs) were set at 400 pg/ml and 100 pg/ml for Aβ1-40 and Aβ1-42, respectively.
We compared Aβ peptide signals in plasma samples at three separate concentrations and calculated the matrix effect recovery following the formula outlined in the “Materials and Methods” section. Both assay formats showed similar matrix effects (Supplementary Tables 2 and Supplementary Fig. 3B). Recovery was improved with analyte-specific IS normalization (Supplementary Table 2 and Supplementary Fig. 3D).
Precision was evaluated at three concentrations using normalized peak areas for both intra- and inter-assay assessments (Supplementary Table 3). Similar %CVs were observed across both assays and normalization techniques, indicating strong reproducibility (%CV < 10%) for both PAβ assay versions.
Relationship between plasma sample dilution and normalized intensity
Strong linear associations were recorded between plasma sample dilution and the normalized intensity for the improved PAβ V2.0 assay (r2 > 0.99) (Fig. 2).
A The relationship between plasma sample dilution and normalized intensity of the PAβ V1.0 and PAβ V2.0 assays. Three technical replicates were performed for each volume. Data were presented as mean values +/− standard deviation. Both Aβ1-40 and Aβ1-42 were normalized using the Aβ1-38 IS. B The S/N ratios of measurements using plasma samples with various volumes (50 μl–250 μl) were compared in three technical replicates between the PAβ V1.0 and the PAβ V2.0 assays for Aβ1-40 and Aβ1-42. Data were presented as mean values +/− standard deviation.
IP recovery
To evaluate the proportion of plasma Aβ peptides retained after the IP procedures, we utilized Simoa immunoassays for the quantification of Aβ peptides pre- and post-IP in selected samples of low, medium, and high Aβ concentrations. The PAβ V2.0 assay showed improved recovery of Aβ peptides after IP vs. the original method (Supplementary Fig. 4).
Participant characteristics
We examined the clinical performance of the improved PAβ assay using baseline data from several well characterized independent cohorts. The Investigating Gains in Neurocognition in an Intervention Trial of Exercise study (IGNITE; NCT02875301) with participants recruited from three study sites: Boston (Northeastern University), Kansas City (University of Kansas Medical Center) and Pittsburgh (University of Pittsburgh), USA. Due to limited sample volume concerns, only specimen from n = 317 participants from IGNITE were included in this study. The Active Gains in Brain Using Exercise During Aging study from Granada, Spain (AGUEDA; NCT05186090; n = 76). The University of Pittsburgh Alzheimer’s Disease Research Center (Pitt-ADRC; MOD19110245-023; n = 30) in Pittsburgh, Pennsylvania, USA. All cohorts predominantly included cognitively unimpaired older adults, while Pitt-ADRC additionally included cognitively impaired participants. We utilized two grouping strategies to define centiloid (CL)27,28 groups: dichotomized using a single cutoff (CL > 24 [(Aβ-PET positive] and CL ≤ 24 [Aβ-PET negative]) and trichotomized using two cutoffs (CL < 12 (Aβ-PET negative), CL > 12 to < 24 (transition zone), and CL > 24 (Aβ-PET positive).
In the IGNITE cohort, the mean age was 69.5 (SD 3.6) years, with 229 (72.2%) females. Eighty-six participants (27.1%) were APOE ε4 carriers, with an average Aβ-PET CL of 7.91 (SD 29.0). Fifty-four (17%) were deemed to be Aβ-PET positive (CL > 24). The mean Montreal Cognitive Assessment (MoCA) score was 26.1 (SD 2.5). The average number of education years was 16.3 (SD 2.2). Aβ burden groups, defined using both dichotomized and trichotomized approaches, showed significant differences in APOE carriership and Aβ-PET CL levels (Table 1).
The mean age in AGUEDA was 71.4 (SD 3.9) years, with 44 (57.9%) females. Twelve participants (16.2%) were APOE ε4 carriers, 12 (15.7%) were Aβ-PET positive, and the average CL was 7.6 (SD 25.3). The mean Mini-Mental State Examination (MMSE) and MoCA scores were 29.0 (SD 1.1) and 25.8 (SD 2.2), respectively. We recorded no significant difference in these metrics according to Aβ-PET status (Table 1) for both cohorts. Similar to the IGNITE cohort, APOE carriership showed a significant association with the CL groups (Table 1).
In the Pitt-ADRC cohort, the mean age was 75.6 years (SD 7.8), with 16 (53.3%) females. Nine participants (30.0%) were APOE ε4 carriers, and eight (26.7%) were clinically diagnosed with probable AD. In terms of cognitive performance, the mean MMSE and MoCA scores were 24.7 (SD 6.3) and 22.9 (SD 7.3), respectively. Regarding Clinical Dementia Rating (CDR), 9 (30.0%), 16 (53.3%), 3 (10.0%), and 2 (6.7%) participants had scores of “disease absent” (CDR = 0), “questionable” (CDR = 0.5), and “disease present but mild” (CDR = 1), and “moderate” (CDR = 2), respectively. There were significant differences in MoCA, MMSE and CDR scores between the probable AD and normal control groups (Supplementary Table 1).
We also used a 49-sample sub-cohort from the IGNITE study to evaluate the PAβ V2.0 assay using two plasma input volumes (100 μl vs. 250 μl). The samples were selected to achieve a near-equal distribution of Aβ-PET positive and negative cases, matched by age, race, and sex. In this sub-cohort, the mean age was 70.9 years (SD 4.2), with 35 (71.4%) females. Fourteen individuals (28.6%) were APOE ε4 carriers, 21 (42.9%) were Aβ-PET positive, and the average CL value was 28.0 (SD 42.3). The mean MoCA score was 26.2 (SD 2.4), and participants had an average of 16.9 years of education (SD 1.9). (Supplementary Table 1).
Correlation of Aβ peptides measured using different PAβ assay versions
In the Pitt-ADRC cohort, strong correlations (r > 0.7) were observed for all Aβ peptides measured with the PAβ V2.0 vs. PAβ V1.0 assays, except Aβ1-42, which showed a moderate correlation (0.7> r > 0.5) (Supplementary Fig. 5A). Overall, similar results were observed in the other cohorts with mostly r > 0.5 correlations (Supplementary Figs. 5B, C and 6).
Comparison of normalization method on clinical performance of the PAβ V2.0 method
Strong correlations were observed in Aβ1-40 (r > 0.9) and Aβ1-42 (r > 0.8) in the Pitt-ADRC and AGUEDA cohorts. In the IGNITE cohort, Aβ1-40 demonstrated a strong correlation (r > 0.8), while Aβ1-42 showed a moderate correlation (0.6 >r > 0.5).
The clinical performance of the PAβ V2.0 assays in differentiating different CL groups was compared using common and analyte-specific ISs. In IGNITE, the AUCs were equivalent – 0.81 and 0.80 for the analyte-specific vs. common ISs. In AGUEDA, the area under the curve to distinguish Aβ-PET groups was significantly higher when using the analyte-specific ISs (AUC = 0.75) vs. using the common IS (AUC = 0.68; Delong test p = 0.036). The Pitt-ADRC cohort was excluded due to a lack of Aβ PET information. The PAβ V2.0 setup using analyte-specific ISs was selected for further use.
Accuracies to classify brain Aβ-PET uptake
In the AGUEDA cohort, the Aβ1-42/Aβ1-40 ratio was significantly lower in Aβ-PET-positive vs. Aβ-PET-negative groups for both the PAβ V2.0 and V1.0 assays with corresponding AUCs of 0.76 and 0.74, respectively (Supplementary Fig. 7). When examined according to the CL scale, there were significant differences in Aβ1-42/Aβ1-40 ratio between the Aβ-PET positive and negative groups for both assays but not between the transition and each of the positive or negative groups (Supplementary Fig. 8). Excluding the transition group marginally increased the AUC for the improved PAβ V2.0 assay (increased from 0.76 to 0.77) but not for the original PAβ V1.0 method.
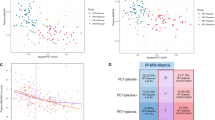
In IGNITE with a much larger sample size, the AUC for Aβ1-42/Aβ1-40 ratio in the improved PAβ V2.0 assay was significantly superior to the original method (0.81 vs. 0.65; Fig. 3A–C). The Youden index method identified a preliminary cutoff of 0.29 for the improved method. Using the CL scale and excluding the transition zone did not change the AUCs (Fig. 3D–F). Quanterix Simoa plasma Aβ42/Aβ40 ratio immunoassay gave a lower AUC of 0.74 (0.75 when excluding the transition group [Supplementary Fig. 9]).
A, B Box-and-whisker plots depict the separation of Aβ PET-positive and Aβ PET-negative groups using the original (PAβ V1.0) and the improved (PAβ V2.0 IS) assays, respectively. The plots show the median, 25th–75th percentiles (box), and whiskers extending to 1.5 × IQR, with individual data points overlaid. Group differences were evaluated using the two-sided Wilcoxon Rank Sum test, with p-values provided. The preliminarily derived optimal cutoff values for each assay, determined by the Youden method, are marked in red. C ROC curves with AUC values and confidence intervals showing the classification accuracies of the Aβ1-42/Aβ1-40 ratio measured with either the IP-MS assay to distinguish Aβ-PET positive vs. negative groups (positivity determined as a CL value of > 24). To better understand plasma Aβ1-42/Aβ1-40 ratio changes according to stepwise brain Aβ pathology, the same IGNITE cohort dataset was further divided into three CL groups (PET-negative, transition, PET-positive). The Box-and-whisker plots for the V1.0 assay (D), V2.0 assay (E) were generated using the same statistical tests and plotting format as in panels (A and B). The ROC curves to identify an abnormal Aβ PET scan with the transition (i.e., intermediate) group excluded (F) are also shown.
Furthermore, the AUC was assessed with adjustments for APOE ε4 status, age, and sex in various combinations for the AGUEDA and IGNITE cohorts. These adjustments resulted in only minor improvements in AUC performance, with increases less than 0.05 (Supplementary Table 4).
Together, the improved PAβ V2.0 assay showed superior biomarker performance vs. the original PAβ IP-MS assay and the Simoa immunoassay.
Concordance between plasma Aβ1-42/Aβ1-40 ratio and Aβ PET positivity
In the IGNITE cohort (including 17.0% and 83.0% Aβ-PET-positive and-negative cases, respectively [Table 1]), the PAβ V1.0 assay correctly classified 9.8% of participants as Aβ-PET-positive and 57.1% as Aβ-PET-negative, corresponding to 66.9% total accuracy. In contrast, the PAβ V2.0 assay demonstrated an enhanced 71.6% accuracy, correctly classifying 13.9% participants as Aβ-PET-positive and 57.7% as PET-negative classifications. Simoa immunoassays demonstrated the lowest concordance; 60% accuracy (15.8% for Aβ-PET-positive and 44.2% Aβ-PET-negative; Fig. 4). Notably, plasma-PET discordant cases for the PAβ V2.0 assay were mainly limited to those who were plasma Aβ-positive but Aβ-PET negative. This discordance may reflect early stages of AD, during which abnormal levels of soluble Aβ species become measurable in blood before abnormal levels of insoluble Aβ aggregates are detectable by PET15.
Concordance vs. Aβ PET is illustrated for Aβ1-42/Aβ1-40 measured with the (A) PAβ V1.0 assay and (B) the PAβ V2.0 assay, as well as (C) Aβx-42/Aβx-40 (i.e., peptides that do not necessarily start from amino acid 1) measured with Simoa multiplex immunoassays. The derived Aβ PET CL cutoff values are depicted on the y-axis, and the assay-specific plasma Aβ ratio cutoff values, determined by the Youden method, are marked on the x-axis. In each quadrant is highlighted the percentage of samples distributed across PET and assay cutoffs.
Biomarker performance of the PAβ V2.0 assay using 100 μl vs. 250 μl plasma volume
In the sub-cohort of IGNITE (n = 49), we evaluated the feasibility of further dropping the sample volume for the improved assay to 100 μl informed by results in Fig. 5. The characteristics of this sub-cohort are shown in Supplementary Table 1. While the measured plasma Aβ peptide levels and the Aβ1-42/Aβ1-40 ratio values were lower when using the 100 μl vs. 250 μl volume aliquots from identical sample tubes (Fig. 5A), we found no sigificant difference in their AUCs to separate Aβ-PET-positive and Aβ-PET-negative groups (0.80 for 100 μl and 0.84 for 250 μl, DeLong’s test p = 0.51; Fig. 5B). The cutpoint determined by the Youden method was 0.31 when utilizing 100 μl plasma.
This experiment was performed in a sub-cohort of the IGNITE study selected to reflect balanced demographics. A Box-and-whisker plots for the PAβ V2.0 IS assay using a 100 μl sample volume, and 250 μl sample volume, split into Aβ PET-positive and PET-negative groups, analyzing the Aβ1-42/Aβ1-40 biomarker. Differences were evaluated using the two-sided Wilcoxon Rank Sum test, with p-values provided. B ROC curves with AUC values for the same assay using 100 μl or 250 μl sample volumes illustrate the diagnostic accuracy of both volumes. The plots show the median, 25th–75th percentiles (box), and whiskers extending to 1.5 × IQR, with individual data points overlaid. C The correlation between the PAβ V1.0 and PAβ V2.0 assays was assessed for both 100 μl and 250 μl volumes, with Aβ1-40 and Aβ1-42 normalized using their respective ISs and other peptides normalized using Aβ1-38 IS. Spearman correlation was used to evaluate the strength of the relationships.
Correlations between the 100 μl and the original 250 μl sample volumes were also compared. Moderate correlations (0.7 >r > 0.5) were observed for Aβ1-38, Aβ1-42, Aβ1-40, and Aβ3-40, while lower correlations (0.5 >r > 0.3) were noted for Aβ3-40 and APP669-711 (Fig. 5C and Supplementary Fig. 10). These findings indicate that the PAβ V2.0 assay performs similarly when the sample volume is further reduced to 100 μl.
Discussion
Among plasma Aβ assays, IP-MS methods such as the assay from Nakamura et al. stand out for their performance but have their own limitations needing improvement. We adopted and enhanced this assay, resulting in the PAβ V2.0 assay with several improvements. Firstly, we successfully streamlined and hence decreased sample preparation time and pre-analytical processing steps, thus boosting throughput. Secondly, we selected to optimize and run the assay on a cost-effective table-top MS instrument – the Bruker Microflex LT – instead of more expensive LC-MS alternatives that are often used for IP-MS analysis. Thirdly, the assay demonstrated a substantially stronger signal to noise ratio, allowing for much lower sample volume requirements. Finally, and perhaps most importantly, the PAβ V2.0 assay recorded superior clinical accuracies to both the original PAβ V1.0 method and Simoa immunoassays alternatives. The resulting assay from these pioneering simplifications and enhancements to a well-known method is likely to enable widespread access to plasma Aβ assessments by IP-MS.
The PAβ V2.0 assay successfully streamlined the IP steps using a commercially available buffer – the N4PE CSF diluent. The high detergent, high salt content and the interference blocker mixture in the buffer helped reduce the background noise. This buffer was selected after comparing its performance against several detergents and blocking buffers. While other tested reagents exhibited lower signal to noise ratios compared with the PAβ V1.0 assay, the N4PE diluent demonstrated higher signal to noise ratio, supporting its selection for further use in the PAβ V2.0 assay.
The PAβ V2.0 assay maintained comparable analytical performance with a higher recovery rate compared to the PAβ V1.0 assay. We further tested the signal to noise of Aβ1-40 and Aβ1-42 in the PAβ V2.0 assay, utilizing a diluted sample volume of pooled plasma. The results demonstrated a higher signal to noise ratio and suggested the potential feasibility of decreasing the sample volume to 100 µl for the PAβ V2.0 assay. In addition, the PAβ V2.0 assay preserved similar clinical performance, with peptide concentrations showing strong correlation with those in the PAβ V1.0 assay.
In comparing the performance using analyte specific IS (separately for Aβ1-40 and Aβ1-42) vs. the common IS (Aβ1-38) for normalization, our results indicated that the analyte specific IS can slightly improve the matrix effect recovery of plasma Aβ peptides. However, this did not significantly change the analytical performance of the Aβ biomarkers. The correlation between these two normalization methods was strong. In the clinical performance analysis, using analyte specific IS in the PAβ V2.0 assay provided a minor improvement compared to the PAβ V2.0 assay with common IS. Our findings supported Nakamura et al.‘s approach, confirming that using a common IS in the MALDI-TOF based IP-MS Aβ assay did not significantly alter clinical performance. Due to this reason, we finalized the PAβ V2.0 assay using analyte specific IS for normalization.
The PAβ V2.0 assay demonstrated superior clinical performance in distinguishing Aβ PET-positive from Aβ PET-negative participants compared with the Simoa Aβ immunoassays when the CL-based transition group was excluded in the IGNITE cohort, indicating strong utility to identify incipient Aβ pathology in an asymptomatic cohort. Concordance analysis against Aβ PET CLs suggested that individuals first become abnormal for plasma Aβ1-42/Aβ1-40 ratio before becoming abnormal in Aβ PET, in agreement with previous results recorded for CSF Aβ1-42/Aβ1-4029,30. With adjustments for APOE ε4, age, and sex, the AUC showed only minor improvements. Based on these findings, we conclude that the PAβ V2.0 IS assay performs effectively without requiring such adjustments.
We evaluated the feasibility of using a reduced sample volume of 100 µl and observed comparable clinical performance to the initial 250 µl, highlighting the assay’s high sensitivity, which will be valuable in research cohort studies and clinical settings where sample volume may be a limiting factor.
The plasma IPMS Aβ assay described herein is important for several reasons. First, this MS-based assay provides specific, high-precision and high-throughput measurements with minimal non-specific binding and lower reagent costs. Furthermore, high-performance Aβ1-42 and the Aβ1-42/Aβ1-40 ratio assays directly reflect brain Aβ pathology even in the very early stages30. Hence, the assay will enable straightforward target engagement evaluation in anti-Aβ therapeutic programs instead of using non-Aβ measures such as p-tau217. In addition, the plasma Aβ measures can enhance the predictive accuracies of p-tau217 to identify preclinical AD participants in cross-sectional studies as well as predict who among these individuals with only subtle pathology will most likely decline cognitively in the near-term17,18,31. Moreover, the estimated reagent cost per sample is approximately $20, which is lower than that of conventional assays. The streamlined workflow also reduces total preparation time from roughly 6 h to about 3 h by eliminating the second immunoprecipitation step. Given these advantages, we believe the streamlined assay holds significant value in advancing AD diagnostics, prognosis and research.
The MS instrument utilized was a Bruker Microflex LT MALDI-TOF, widely adopted across numerous clinical facilities. Notably, it has received FDA approval for clinical microbiology diagnosis in humans32,33, and is widely available in many laboratories. In comparison to other MS instruments utilized in alternative IP-MS plasma Aβ assays, the Microflex is distinguished by its affordability and simplicity. Furthermore, it offers practical advantages, such as direct compatibility with a standard 110 V outlet, without necessitating the use of any special electrical modifications or a voltage converter. Its user-friendly interface also facilitates straightforward operation, enabling general laboratory technicians to operate the instrument proficiently without requiring specialized training in mass spectrometry.
This study has several notable strengths. Firstly, we describe in detail the technical development, analytical and clinical validation of an improved plasma Aβ assay by IP-MS. Secondly, we included multiple cohorts for the clinical performance assessments, including those characterized for biological evidence of disease using brain Aβ PET. The multi-site recruitment approach employed by the IGNITE cohort in particular enhances the generalizability of findings. Limitations of our study includes the limited comprehensive comparison of the streamlined assay with other plasma Aβ assays especially those using IP-MS technology.
A recent study found that longitudinal changes in plasma Aβ closely track—and may even precede—changes in brain Aβ15. This earlier change highlights the potential of plasma Aβ as a practical screening tool for clinical trial recruitment and for detecting at-risk individuals earlier than PET imaging. The study also indicated that PET-derived thresholds are not well suited for plasma soluble Aβ biomarker, as the two measurements reflect different concentration changes in diseased individuals: soluble Aβ in plasma declines while insoluble Aβ in the brain increases.
In conclusion, we report the successful development of a more resource-efficient and cost-effective IP-MS plasma Aβ assay. Compared with the in-house reproduced Nakamura et al. assay, the streamlined assay demonstrated improved clinical and analytical performance. The cost, time, and reagent savings, coupled with the utilization of a more affordable and widely available instrument, and the feasibility of using reduced sample volume, will enable research laboratories to conduct IP-MS analysis of Aβ in blood more effectively.
Methods
Immunoaffinity enrichment
Pittsburgh plasma Aβ assay V1.0
The PAβ V1.0 assay was developed at the University of Pittsburgh based on the method originally described by Nakamura et al.11. For each sample, 250 μl of binding buffer (100 mM Tris-HCl, pH 7.4 [Sigma #T2788-1L], 300 mM NaCl [Sigma #S7653-250G], 0.2% w/v n-dodecyl-β-D-maltoside [DDM; Sigma #D4641-1G], 0.2% w/v n-nonyl-β-D-thiomaltoside [NTM; Anatrace #148565-55-3]) containing 62.4 pg/ml of Aβ1-38 IS (Anaspec #AS-65220), was added to a 1.5 ml Eppendorf Protein LoBind Tube (ThermoFisher #13-698-794), followed by the addition of 250 μl plasma sample. To facilitate direct comparison with the PAβ V2.0 assay, 100 pg/ml Aβ1-40 IS (rPeptide #A-1101-2) and 30 pg/ml Aβ1-42 IS (rpeptide #A-1102-1) were also added to the binding buffer for the evaluation of analytical performance.
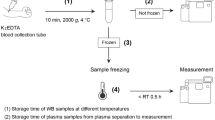
The samples were immunoprecipitated with 10 μl of 50 mg/ml Dynabeads (M-270 Epoxy; ThermoFisher #14301) coupled with 5 μg 6E10 anti-Aβ antibody (BioLegend #803003) for 1 h at 4 °C with rotation. The beads were coupled with the antibody following the protocol recommended by the manufacturer. After the IP, the supernatant was discarded, and the beads washed once with 0.5 ml of cold phosphate-buffered saline (PBS, Gibco #2537136). The washed beads were then transferred to a fresh Eppendorf tube using 0.5 ml of cold PBS and eluted with 25 μl of glycine elution buffer (50 mM glycine [pH 2.8, Sigma #G2879-100G], 0.1% DDM) after removing all liquid. The eluates were collected and transferred to fresh tubes containing 0.5 ml of the binding buffer (without any Aβ ISs) for a second round of IP. Following one hour of rotation at 4 °C, the beads were washed twice with 0.5 ml of cold HPLC-grade H2O (Fisher #7732-18-5) and transferred to a fresh Eppendorf tube by resuspending in 0.2 ml H2O. After complete removal of all liquid through vacuum aspiration, the beads were eluted using 6 μl of 3 mg/ml α-cyano-4-hydroxycinnamic acid matrix (Bruker #8201344) dissolved in TA50 (50% Acetonitrile [Fisher #75-05-8], 0.1% Trifluoroacetic acid [Alfa Aesar #UN2699], 1 mM ammonium dihydrogen phosphate [Sigma #204005]). The eluate was spotted four times with 1 μl each onto the MALDI target plate (Bruker #8280823) for MS analysis. A schematic illustration of the workflow for this assay is shown in Fig. 6A.
A The original PAβ version 1.0 (V1.0) assay requires two rounds of immunoprecipitation, with a total processing time of approximately 6 hours per plate. B The improved PAβ version 2.0 (V2.0) assay features a streamlined sample preparation procedure with a single round of immunoprecipitation, reducing processing time to approximately three hours per plate, offering significant savings in time, antibody and resources. Created in BioRender. Chen, Y. (2025) https://BioRender.com/lbr530d “.
Single IP procedure for detergents and blocking buffer tests
Similar to the first IP step of the PAβ V1.0 assay, we prepared 250 μl of the same assay binding buffer, either used as is or supplemented with one of the following detergents or blocking buffers: 10% v/v SuperBlock (Thermo #37535), 10 μg/ml TruBlock (Meridian #A66803H), 0.5% v/v Triton100 (Millipore #648462), 0.5% v/v Tween20 (BioRad #1610781), or 10% Quanterix Neurology Plex 4E CSF sample diluent (N4PE CSF diluent [Quanterix #103727]) for different tests.
This mixture was transferred to a 1.5 ml Eppendorf Protein LoBind tube with 62.4 pg/ml of Aβ1-38 IS, 100 pg/ml of Aβ1-40 IS, and 30 pg/ml of Aβ1-42. Subsequently, 250 μl of human plasma sample was added to the mixture. The sample was immunoprecipitated with 5 μl of 50 mg/ml Dynabeads coupled with 1.25 μg 6E10 Aβ antibody (BioLegend #803003) for 1 h at 4 °C with rotation. After IP, the supernatant was discarded, and the beads resuspended in 0.5 ml of the assay binding buffer with the corresponding supplement added as appropriate and transferred to another tube. The beads underwent an additional wash with 0.5 ml of the binding buffer with corresponding supplement, two washes with 0.5 ml of PBS and one wash with 0.5 ml of HPLC-grade H2O. Finally, the beads were transferred to a fresh Eppendorf tube using 0.2 ml of H2O. After removal of all liquid through vacuum aspiration, the beads were eluted using 6 μl of 3 mg/ml α-cyano-4-hydroxycinnamic acid matrix dissolved in TA50. The eluate was spotted four times with 1 μl each onto the MALDI target plate for analysis.
Screening of buffers and blockers for the PAβ V2.0 assay
We evaluated the effects of several buffer systems and heterophilic blocking agents for the PAβ V2.0. These included the 10% N4PE CSF diluent from Quanterix, the 10% v/v SuperBlock, 10 μg/ml TruBlock, 0.5% v/v Triton100 and 0.5% v/v Tween20. The results from the PAβ V2.0 assay were compared to those obtained using the PAβ V1.0 assay.
MALDI-TOF MS
After sample spotting, the MALDI target plate was air dried and then loaded into a benchtop MALDI- TOF mass spectrometer, Microflex LT (Bruker Daltonics), equipped with a 337 nm nitrogen laser to acquire mass spectra. The Microflex LT operated in linear mode with a pulsed positive ion extraction setting, utilizing an attenuator offset of 12%, an attenuator range of 30%, and an optimized laser power (63% in this case). An external mass calibration was performed using a peptide calibration mixture consisting of two calibration standards (Bruker #8222570, #8206355). The auto scan function was utilized, acquiring one spectrum for each spot through the combination of ion signals from 2500 laser shots, resulting in four spectra per sample. Aβ1-38 IS was employed to ensure spectrum quality in the auto scan function. Only the spectra, generated from every 50 shots, with Aβ1-38 IS S/N ratios greater than three, were collected34,35. After acquisition, the spectra underwent smoothing using the SavitzkyGolay algorithm with a width of 0.1 m/z and baseline subtraction using the TopHat algorithm. The peak intensity and S/N ratios were measured using FlexControl (v3.4, Bruker Daltonics). Subsequently, ClinPro Tools Software (v2.1, Bruker Daltonics) was employed for m/z alignment, peak detection, and peak area calculation.
Analytical assessment
Linearity analysis was conducted using a two-fold serial dilution of an Aβ peptide mixture, starting with concentrations of 400 pg/ml for Aβ1-40 (Anaspec, #AS-24235) and 100 pg/ml for Aβ1-42 (Anaspec, #AS-20276), in 6% BSA/PBS, diluting up to 64x. The analysis involved six replicates for each dilution, totaling 36 samples, which were evenly processed across two batches. The LLOQ was established as the lowest concentration measurable with a %CV under 20%36. The working range was defined as the range from the LLOQ to the highest concentration tested. To evaluate the plasma matrix effect, we assessed the recovery by comparing the results in plasma to those in 6% BSA/PBS at three different concentration levels (118.2 pg/ml, 53.6 pg/ml, and 21.4 pg/ml for Aβ1-40, 47.2 pg/ml, 23.0 pg/ml and 10.8 pg/ml for Aβ1-42). The average peak area of the replicates was used for recovery calculations. Both media were spiked with equal amounts of Aβ1-40 and Aβ1-42 prior to the IP procedures. Three replicates per concentration were tested. Recovery was calculated using the formula:
where P represents the average normalized peak area of three replicates. Intra- and inter-assay variability were determined by analyzing samples at three Aβ concentration levels across five batches, each containing six replicates per concentration.
The linearity, LLOQ, working range, matrix effect recovery and precision of Aβ1-40 and Aβ1-42 were normalized using either common IS (Aβ1-38 IS) or analyte specific IS (Aβ1-40 IS and Aβ1-42 IS), respectively.
Plasma dilution linearity
The effect of plasma dilution on normalized intensity for both the PAβ V1.0 and PAβ V2.0 assay formats were investigated by testing five separate amounts of a pooled plasma sample (50 μl to 250 μl), with three replicates each. All samples in this test were diluted to 250 μl prior to processing, and Aβ1-40 and Aβ1-42 levels were normalized using the Aβ1-38 IS only.
Simoa assay for IP recovery assessment
To quantify the proportion of Aβ peptides retained after the IP procedures, Simoa assays were utilized. These assays were performed using the Simoa Human Neurology 4-Plex E assay (N4PE) kit from Quanterix (103670) on an HD-X analyzer (Quanterix, Billerica, MA, USA). The IP recovery was evaluated at three concentration levels of low, medium, and high (27.4 pg/ml, 51.4 pg/ml, and 99.2 pg/ml for Aβ1-40; 7.0 pg/ml, 13.2 pg/ml, and 27.4 pg/ml for Aβ1-42) in triplicates. The average concentration of triplicates was used for recovery calculations. To monitor assay performance, QC samples at three different concentrations were analyzed at the beginning and end of each assay run. The average %CV for the QCs was below 10%.
Participants
This study included plasma samples from three cohorts. The first cohort was derived from the IGNITE study (ClinicalTrials.gov: NCT02875301). Participants were recruited at three study sites: Boston, Kansas City and Pittsburgh, USA37. The Aβ-PET was performed using the [18F] Florbetaben tracer, and Aβ burden was expressed in the CL scale. Participants were community-dwelling older adults (aged 65-80 years) without a history of neurological disease. Those with suspected cognitive impairment were excluded following a consensus conference involving geriatric neuropsychologists. Only participants classified as cognitively unimpaired based on the National Institute on Aging-Alzheimer’s Association (NIA-AA) 2011 criteria were included38. Detailed exclusion criteria are available in Erickson et al.18,37. In this study, only baseline samples from a subset of participants with available Aβ PET data and enough extra sample volume (n = 317) were used for analysis24. The study received Institutional Review Board approval at all participating sites, and written informed consent was obtained from all participants. Scientists were blinded to all participant information until the completion of data acquisition.
The second cohort was sourced from the AGUEDA project (NCT05186090). Participants (aged 65–80 years) were recruited from Granada, Spain, based on their classification as physically inactive and cognitively normal, assessed by the Spanish Telephone Interview for Cognitive Status modified (STICS-M), MMSE, and MoCA. Like the IGNITE, Aβ-PET was performed using the [18F] Florbetaben tracer, where Ab burden was expressed using the CL scale. Detailed information on eligibility criteria, participant selection methods, and recruitment procedures, as well as details about the study setting, locations, and data collection, can be found in a comprehensive description provided in the AGUEDA protocol39. Prior to enrollment in the AGUEDA trial, participants provided informed consent, and the trial was conducted in accordance with the approval of the Research Ethics Board of the Andalusian Health Service (CEIM/CEI Provincial de Granada; #2317-N-19). In this cross-sectional analysis, we focused on the baseline data.
For the third cohort, we enrolled participants from the University of Pittsburgh ADRC in Pittsburgh, Pennsylvania, USA. The participants (aged 60–92 years) in this ongoing study underwent annual clinical evaluation to assess their longitudinal brain health and potential development of cognitive impairment and dementia. Annual evaluations included neuroimaging, cognitive testing, and blood collection for use in plasma biomarker analysis outside of the clinical assessment. Neuropsychological evaluation and diagnoses were established through clinical assessments40,41. The battery of cognitive tests included the MoCA, MMSE, and the CDR scale. The current investigation was a prospective, blinded sub-study where participants were enrolled based on their order of clinical attendance and their informed consent to participate. This involved agreeing to provide an additional tube of blood for the project. The ADRC study was approved by the University of Pittsburgh Institutional Review Board (MOD19110245-023). Due to the limited sample size, this cohort was used for correlation analysis only.
Blood collection and processing procedures
At the University of Pittsburgh ADRC, blood samples were collected via venipuncture by nurses with extensive clinical experience and trained in ADRC procedures42. Blood collection was performed between 9:00 AM and 2:00 PM, with the time of the last meal recorded. For the AGUEDA cohort, fasting blood samples were collected at the Virgen de las Nieves University Hospital, Spain, between 8:00 AM and 10:00 AM after more than 8 h of fasting. In the IGNITE cohort, approximately 47 cc of fasting blood was collected between 8:00 AM and 10:00 AM across all sites, with participants following pre-draw restrictions on food, exercise, alcohol, nicotine, and specific medications.
In the ADRC cohort, blood was drawn using 10 mL lavender top EDTA tubes, inverted 8–10 times, and centrifuged at 2000 × g for 15 min at 4 °C to separate plasma, which was then aliquoted into cryovials and stored at − 80 °C. For the AGUEDA cohort, 4 mL lavender top EDTA tubes were used, with similar post-collection inversion and centrifugation at 2000 × g for 10 minutes at 4 °C before plasma was aliquoted and stored at − 80 °C. In the IGNITE cohort, blood was collected in 10 mL lavender top EDTA tubes, inverted 8–10 times, and centrifuged at 2000 × RPM for 20 min at room temperature. Plasma from IGNITE samples was promptly aliquoted into cryovials and stored at − 80 °C at the University of Pittsburgh.
Mass spectrometric and immunoassay experiments were performed in the Mass Spectrometry facility at the Biofluid Biomarker Laboratory, Department of Psychiatry, School of Medicine, University of Pittsburgh, Pittsburgh, PA, USA.
Correlation analysis
The correlation between levels normalized using a common IS and those normalized using analyte-specific ISs was evaluated for Aβ1-42 and Aβ1-40 measured by the PAβ V2.0 assay to find the optimized normalization method.
The correlation between the PAβ V1.0 and PAβ V2.0 assays was assessed by analyzing the normalized peak areas of multiple Aβ biomarkers, including Aβ1-42, Aβ1-40, Aβ1-39, Aβ3-40, Aβ1-38, and APP669-711, across all cohorts. Aβ1-42 and Aβ1-40 from the PAβ V2.0 assay were normalized with analyte-specific ISs, while the other biomarkers and all analytes in the PAβ V1.0 assay were normalized using the Aβ1-38 IS.
In addition, the correlation between the PAβ V2.0 and Simoa assays was evaluated for Aβ1-42 and Aβ1-40 for the IGNITE cohort, with both biomarkers in the PAβ V2.0 assay normalized using analyte-specific ISs.
For a clearer illustration, outliers exceeding upper quartile (Q3) + 3 × interquartile range (IQR) or falling below lower quartile (Q1) − 3 × IQR were excluded from this analysis. The version without any exclusion is illustrated as Supplementary Fig. 6. Correlation strength was interpreted based on previously published guidelines43.
Clinical performance assessment
The clinical assessment of the PAβ V2.0 Aβ1-42/Aβ1-40 biomarker was initially compared between the common IS and the analyte-specific ISs to finalize the normalization method.
Subsequently, two different Aβ biomarkers were evaluated: Aβ1-42/Aβ1-40 from the PAβ V1.0 assay, Aβ1-42/Aβ1-40 from the PAβ V2.0 assay, normalized with analyte-specific ISs. Biomarker performance was assessed in the AGUEDA and IGNITE cohorts, using Aβ PET imaging results and CL scales. In the IGNITE cohort, the biomarkers were further assessed for cutoff and compared with Aβ1-42/Aβ1-40 from the Simoa assay.
The IP-MS assay performance over multiple batches was evaluated using pooled QC plasma samples. Two levels of QCs were tested in duplicates to ensure assay performance. The intra- and inter-assay %CVs were determined to be less than 20% for the cohorts.
The Simoa assay was conducted on an HD-X analyzer following the manufacturer’s guidelines at the Biofluid Biomarker Laboratory. Aβ peptides were measured using the Neurology 2-Plex A kit (N2PA, Quanterix #14712) for the IGNITE cohort. To ensure assay reliability, QC samples at three concentrations were tested at the beginning and end of each assay run. The average %CV for QCs across all runs was below 10%.
Clinical performance assessment using 100 μl sample volume
Fifty-two samples were selected from the IGNITE cohort, ensuring that race, age, sex, and Aβ-PET results were evenly selected according. Each sample was processed with a starting volume of 100 μl, diluted to 250 μl with binding buffer, and prepared using the single IP procedure with a 10% N4PE CSF diluent buffer.
For the samples with signal to noise ratio of Aβ1-42 smaller than three using auto scan, manual scan was utilized. The manual scan has the same instrument setting and laser setting as the auto scan with laser shots goes up to 8000 for each spot. No spectrum quality criteria applied. After acquisition, the spectra underwent the same smoothing, baseline subtraction, alignment, peak detection, and peak area calculation as the auto scan.
Three samples (5.7%) failed processing and were excluded from the analysis, resulting in forty-nine samples. Two levels of QCs were tested in duplicates to ensure assay performance. The intra- and inter-assay %CV were determined to be less than 10%.
For both assays, Aβ1-39, Aβ3-40, Aβ1-38, and APP669-711 were normalized using the Aβ1-38 ISs, while Aβ1-42 and Aβ1-40 in the PAβ V2.0 assay were normalized using their respective analyte-specific ISs.
Statistical analyses
For participant demographic characteristics, continuous variables were summarized using means and standard deviations, while categorical variables were reported as numbers and percentages. Differences across cohorts for continuous variables were examined using the Wilcoxon Rank Sum test or Kruskal-Wallis test, depending on the number of groups involved. Categorical variables were analyzed using Fisher’s exact tests. For S/N ratio comparison between different assays, the Wilcoxon Rank Sum test was used. For clinical assessments, box and whisker plots were generated using Aβ PET imaging results and CL scales over the cohorts. The Wilcoxon Rank Sum test was used to assess the disease discriminating performance of biomarkers across cohorts based on the Aβ PET imaging results. The same test was also applied to compare differences between the three CL scale groups in pairwise comparisons. The optimal cut-offs were determined using Youden’s Index. For correlations, Spearman correlation analysis was conducted to evaluate the strength of the association between Aβ peptide measurements from the two different assays. For all the tests, a p-value less than 0.05 was considered statistically significant. All analyses were performed using R statistical software (version 4.2.1, R Foundation for Statistical Computing, Vienna, Austria), available at [http://www.r-project.org/].
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data supporting this study contain sensitive human participant information and cannot be deposited in a public repository due to institutional review board restrictions and the terms of participant consent. Specific de-identified data can be shared with qualified and identifiable investigators for the purpose of replicating the results and procedures in the study. Requests can be made to the corresponding author (T.K.K.), who will direct them to the respective cohort principal investigators where necessary. Requests will be reviewed by the investigators and respective institutions to ensure that data sharing requests conform to US and EU legislation on data protection, intellectual property and confidentiality obligations. Data request to the Pittsburgh ADRC can be made directly at https://www.adrc.pitt.edu/for-researchers/adrc-data-resources/. Source data are provided in this paper.
Code availability
R scripts used for data visualization and figure generation are available from the corresponding author upon reasonable request.
References
Zhang, Y. et al. Amyloid β-based therapy for Alzheimer’s disease: challenges, successes and future. Signal Transduct. Target. Ther. 8, 248 (2023).
Hampel, H. et al. The amyloid-β pathway in Alzheimer’s disease. Mol. Psychiatry 26, 5481–5503 (2021).
Dubois, B. et al. Alzheimer disease as a clinical-biological construct—an international working group recommendation. JAMA Neurol.81, 1304–1311 (2024).
Jack, C. R. et al. Revised criteria for diagnosis and staging of Alzheimer’s disease: Alzheimer’s Association Workgroup. Alzheimer’s Dementia 20, 5143–5169 (2024). p.
Cummings, J. L. Maximizing the benefit and managing the risk of anti-amyloid monoclonal antibody therapy for Alzheimer’s disease: Strategies and research directions. Neurotherapeutics 22, e00570 (2025).
Cummings, J. et al. Lecanemab: appropriate use recommendations. J. Prev. Alzheimer’s Dis. 10, 362–377 (2023).
Rashad, A. et al. Donanemab for Alzheimer’s disease: a systematic review of clinical trials. Healthcare 11, 32 (2022).
Karikari, T. K. et al. Blood phospho-tau in Alzheimer disease: analysis, interpretation, and clinical utility. Nat. Rev. Neurol. 18, 400–418 (2022).
Gonzalez-Ortiz, F. et al. Plasma phospho-tau in Alzheimer’s disease: towards diagnostic and therapeutic trial applications. Mol. Neurodegener. 18, 18 (2023).
Balogun, W. G. et al. Plasma biomarkers for neurodegenerative disorders: ready for prime time? Curr. Opin. Psychiatry 36, 112–118 (2023).
Nakamura, A. et al. High performance plasma amyloid-β biomarkers for Alzheimer’s disease. Nature 554, 249–254 (2018).
Keshavan, A. et al. Population-based blood screening for preclinical Alzheimer’s disease in a British birth cohort at age 70. Brain 144, 434–449 (2021).
Janelidze, S. et al. Head-to-head comparison of 8 plasma amyloid-β 42/40 assays in Alzheimer disease. JAMA Neurol. 78, 1375 (2021).
Li, Y. et al. Validation of plasma amyloid-β 42/40 for detecting Alzheimer disease amyloid plaques. Neurology 98, e688–e699 (2022).
Burnham, S. C. et al. Longitudinal evaluation of the natural history of amyloid-β in plasma and brain. Brain Commun. 2, fcaa041 (2020).
Wang, J. et al. Diagnostic accuracy of plasma p-tau217/Aβ42 for Alzheimer’s disease in clinical and community cohorts. Alzheimer’s Dementia 21, e70038 (2025).
Zhong, X. et al. Plasma p-tau217 and p-tau217/Aβ1-42 are effective biomarkers for identifying CSF- and PET imaging-diagnosed Alzheimer’s disease: Insights for research and clinical practice. Alzheimer’s Dementia 21, e14536 (2025).
Olvera-Rojas, M. et al. Influence of medical conditions on the diagnostic accuracy of plasma p-tau217 and p-tau217/Aβ42. Alzheimer’s Dementia 21, e14430 (2025).
Sehrawat, A. et al. Pittsburgh plasma p-tau217: Classification accuracies for autosomal dominant and sporadic Alzheimer’s disease in the community. Alzheimer’s Dementia 21, e70409 (2025).
Warmenhoven, N. et al. A comprehensive head-to-head comparison of key plasma phosphorylated tau 217 biomarker tests. Brain 148, 416–431 (2024).
Schindler, S. E. et al. Head-to-head comparison of leading blood tests for Alzheimer’s disease pathology. Alzheimer’s Dementia 20, 8074–8096 (2024).
Chong, J. R. et al. Blood-based high sensitivity measurements of beta-amyloid and phosphorylated tau as biomarkers of Alzheimer’s disease: a focused review on recent advances. J. Neurol. Neurosurg. Psychiatry 92, 1231–1241 (2021). p.
Brand, A. L. et al. The performance of plasma amyloid beta measurements in identifying amyloid plaques in Alzheimer’s disease: a literature review. Alzheimer’s Res. Therapy 14, 195 (2022).
Sewell, K. R. et al. Blood biomarkers differentiate AD-related versus non-AD-related cognitive deficits. Alzheimer’s Dementia 21, e14619 (2025).
Chen, Y. et al. Effect of blood collection tube containing protease inhibitors on the pre-analytical stability of Alzheimer’s disease plasma biomarkers. J. Neurochem. 168, 2736–2750 (2024).
Dey, K. K. et al. Deep undepleted human serum proteome profiling toward biomarker discovery for Alzheimer’s disease. Clin. Proteom. 16, 16 (2019).
Salvadó, G. et al. Centiloid cut-off values for optimal agreement between PET and CSF core AD biomarkers. Alzheimer’s Res. Ther. 11, 27 (2019).
La Joie, R. et al. Multisite study of the relationships between antemortem [11C]PIB-PET Centiloid values and postmortem measures of Alzheimer’s disease neuropathology. Alzheimer’s Dementia 15, 205–216 (2019). p.
Palmqvist, S. et al. Detailed comparison of amyloid PET and CSF biomarkers for identifying early Alzheimer disease. Neurology 85, 1240–1249 (2015).
Palmqvist, S. et al. Cerebrospinal fluid analysis detects cerebral amyloid-β accumulation earlier than positron emission tomography. Brain 139, 1226–1236 (2016).
Niimi, Y. et al. Combining plasma Aβ and p-tau217 improves detection of brain amyloid in non-demented elderly. Alzheimer’s Res. Ther. 16, 115 (2024).
Rocca, M. F. et al. Utility of platforms Viteks MS and Microflex LT for the identification of complex clinical isolates that require molecular methods for their taxonomic classification. PLoS ONE 14, e0218077 (2019).
Food, U. & D. Administration, 510 (k) substantial equivalence determination decision summary. AlloMap® Molecular Expression Testing. K, 73482. (2013).
Krutchinsky, A. N. & Chait, B. T. On the mature of the chemical noise in MALDI mass spectra. J. Am. Soc. Mass Spectrom. 13, 129–134 (2002).
Busse, K. et al. The signal-to-noise ratio as a measure of HA oligomer concentration: a MALDI-TOF MS study. Carbohydrate Res. 341, 1065–1070 (2006).
Iino, T. et al. Quantification of amyloid-β in plasma by simple and highly sensitive immunoaffinity enrichment and LC-MS/MS assay. J. Appl. Lab. Med. 6, 834–845 (2021).
Erickson, K. I. et al. Investigating gains in neurocognition in an intervention trial of exercise (IGNITE): protocol. Contemp. Clin. Trials 85, 105832 (2019).
Sperling, R. A. et al. Toward defining the preclinical stages of Alzheimer’s disease: recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement 7, 280–292 (2011).
Solis-Urra, P. et al. Active gains in brain using exercise during aging (AGUEDA): protocol for a randomized controlled trial. Front. Hum. Neurosci. 17, 1168549 (2023).
Lopez, O. L. et al. Research evaluation and diagnosis of probable Alzheimer’s disease over the last two decades: I. Neurology 55, 1854–1862 (2000).
Lopez, O. L. et al. Research evaluation and diagnosis of possible Alzheimer’s disease over the last two decades: II. Neurology 55, 1863–1869 (2000).
Zeng, X. et al. Alzheimer blood biomarkers: practical guidelines for study design, sample collection, processing, biobanking, measurement and result reporting. Mol. Neurodegener. 19, 40 (2024).
Asuero, A. G., Sayago, A. & González, A. The correlation coefficient: An overview. Crit. Rev. Anal. Chem. 36, 41–59 (2006).
Acknowledgements
This study was supported by the Joseph A. Massaro Alzheimer’s Research Fund of The Pittsburgh Foundation (AD2018-97918), the National Institutes of Health (P30 AG066468, P01 AG025204, and R01 AG083874). The AGUEDA study was supported by grants RTI2018-095284-J-I00, PID2022-137399OB-I00 and CNS2024-154835 funded by MCIN/AEI/10.13039/501100011033/ and “ERDF A way of making Europe”. The IGNITE study was funded by the National Institutes of Health (R01 AG053952) awarded to K.I.E., J.M.B., A.F.K., E.M. and R35 AG072307 awarded to KIE. Use of REDcap for the project was sponsored by the Clinical and Translational Science Institute at the University of Pittsburgh (UL1-TR-001857) and the University of Kansas Medical Center (P30 AG072973; UL1 TR002366). TKK and members of the Karikari Laboratory were further supported by NIH/NIA (R01 AG083874, U24AG082930, P30 AG066468, RF1AG077474, R01 AG083156, R37 AG023651, R01 AG025516, R01 AG073267, R01 AG075336, R01 AG072641, P01 AG025204), NIH/NINDS (U01 NS131740, U01 NS141777), NIH/NIMH (R01 MH108509), Aging Mind Foundation (DAF2255207), DoD (HT94252320064), the Anbridge Charitable Fund, and a professorial endowment from the Department of Psychiatry, University of Pittsburgh. MOR was supported by the Spanish Ministry of Science, Innovation and Universities (FPU 22/02476). PSU was supported by MICIU/AEI /10.13039/501100011033 and FEDER, UE (PID2022-137399OB-I00). IEC was supported by a grant RYC2019-027287-I funded by MCIN/AEI/10.13039/501100011033/ and “ESF Investing in your future”. The content of this article is solely the responsibility of the authors and does not necessarily represent the official views of the funders. The authors are thankful to the participants in the IGNITE, AGUEDA and ADRC cohorts whose dedication and commitment were fundamental to the successful implementation of this study. An earlier version of this work was included in the doctoral dissertation of Yijun Chen, “Innovative Pre-Analytical Strategies and Method Development for Blood-Based Biomarkers of Alzheimer’s Disease” (University of Pittsburgh, 2025) in ProQuest. The current manuscript represents a revised and extended version of that work.
Author information
Authors and Affiliations
Contributions
Y.C. designed the study, performed experiments, processed and analyzed data, optimized sample preparation and methods, assisted with clinical coordination and data acquisition, and conducted statistical analysis with support from X.Z. and T.K.K. X.Z., M.O.-R., and K.R.S. processed and analyzed data, contributed to clinical coordination and data acquisition, performed statistical analysis, and assisted with study design and interpretation of PET imaging data. A.S. and J.G. performed experiments and contributed to sample preparation and method optimization. T.A.P., M.D.I., A.D.C., V.L.V., O.L.L., E.M.T.-I., P.S.-U., M.G.-R., L.E.O., A.F.K., C.H.H., J.M.B., A.L.M., E.M., and C.K. contributed to clinical coordination and data acquisition and assisted with the interpretation of imaging and clinical data. I.E.-C., K.I.E., N.A.Y., and T.K.K. supervised the study. T.K.K. conceptualized the project, oversaw the study, and guided manuscript preparation. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
Y.C., X.Z., N.A.Y., and T.K.K. are named inventors on US and worldwide patents filed by the University of Pittsburgh on the streamlined plasma Aβ method described in this manuscript. Disclosure title: “Method for the Quantification of Plasma Amyloid-Beta Biomarkers in Alzheimer’s Disease;” US Application No.: 63/672,952; PCT Serial No.: PCT/IB2025/057270; Current stage: International search report and written opinion of the international searching authority. The specific aspects of the manuscript covered in the patent application include the streamlined plasma Aβ method and its improved clinical performance. TKK has served as an adhoc consultant and/or advisory board member for Quanterix Corporation, SpearBio Inc., Neurogen Biomarking LLC., Alzheon, Siemens Healthineers and Neurogen Biomarking LLC., outside the submitted work. T.K.K. is an inventor on patents and provisional patents regarding biofluid biomarker methods, targets and reagents/compositions, that may generate income for the institution and/or self should they be licensed and/or transferred to another organization. T.K.K. has received royalties from Bioventix for the transfer of specific antibodies and blood biomarker assays to third-party organizations. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Soumya Mukherjee and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, Y., Zeng, X., Olvera-Rojas, M. et al. Streamlined resource-efficient plasma amyloid-beta mass spectrometry assay has improved biomarker performance in preclinical Alzheimer’s disease. Nat Commun 17, 1673 (2026). https://doi.org/10.1038/s41467-026-68372-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68372-w