Abstract
Pediatric obesity is linked to multi-organ inflammation and an increased risk of cardiometabolic and steatotic liver disease. To identify circulating biomarkers of cardiometabolic risk, we performed proximity extension assay proteomics, to quantify 149 inflammation- and cardiovascular-related proteins in a cross-sectional study of 4024 children and adolescents (2377 with obesity and 1647 with normal weight). We identified protein signatures linked to obesity, dyslipidemia, insulin resistance, hyperglycemia, hypertension, and related cardiometabolic phenotypes. Using machine learning, a three-protein panel (CDCP1, FGF21, HAOX1) combined with liver enzymes improved prediction of steatotic liver disease versus liver enzymes alone (receiver operating characteristic–area under the curve (ROC–AUC) = 0.83 vs. 0.77; DeLong’s test, P < 0.05). During a 1-year non-pharmacological obesity intervention (n = 184), reductions in adiposity were associated with decreased inflammatory cytokines (including CDCP1, FGF21), which correlated with improvements in cardiometabolic risk profiles. Here we show that circulating proteomic signatures may mediate obesity-related cardiometabolic risk in youth.
Similar content being viewed by others
Introduction
The World Health Organization has declared childhood obesity as one of the most serious public health challenges of the twenty-first century, primarily due to the increasing global prevalence and the degree of associated cardiometabolic complications later in life, such as type 2 diabetes (T2D), metabolic dysfunction-associated steatotic liver disease (MASLD), and cardiovascular disease (CVD)1. The clustering of cardiometabolic risk factors, such as chronic low-grade inflammation, dysmetabolism, and hypertension, can emerge early in life and may serve as early indicators of future cardiometabolic comorbidities2. Steatotic liver disease (SLD) is a concerning complication of obesity in childhood, as it is associated with an increased risk of progression to cirrhosis in adulthood and an independent predictor of T2D risk3.
Pediatric obesity is highly heterogeneous, varying in clinical manifestation, progression, and treatment response. A key contributor to this heterogeneity is individual differences in systemic inflammation4. Traditionally measured inflammatory markers, such as C-reactive protein (CRP), provide limited insights into the molecular pathways underpinning obesity-related metabolic dysfunction4. Advancements in targeted proteomics, such as the proximity extension assay (PEA), have enabled high-throughput quantification of low-abundance proteins, including cytokines, chemokines, and CVD-related proteins5. This approach may help to unravel molecular pathways specific to inflammatory and metabolic dysfunction, underpinning pediatric obesity. Despite this potential, this area remains relatively unexplored6. Large-scale studies on protein-disease associations have been conducted mainly in adult populations7,8, despite evidence that cardiometabolic abnormalities emerge early on in life. Proteomic studies performed in pediatric cohorts remain limited in sample size and differ in terms of methodology9 and scope10.
In this study, we applied PEA-based plasma proteomics alongside comprehensive phenotyping in children and adolescents from The HOLBAEK Study. Our aim was to investigate how circulating protein markers mediate the relationship between overweight/obesity and inflammation-related cardiometabolic risk. The cross-sectional analysis included 2377 children and adolescents with overweight/obesity and 1647 with normal weight11,12, along with replication in 45825 adults from the UK Biobank (UKB). Additionally, the study features an intervention cohort of 184 children and adolescents with overweight/obesity who underwent The Holbaek obesity treatment, which is a family-centered, individualized, non-pharmacological approach, for 1 year at the Children’s Obesity Clinic at The Holbaek Hospital in Denmark.
Our analysis identified distinct obesity-associated plasma proteomic signatures linked to inflammation and metabolic disturbances from an early age, with similar patterns observed in adults from the UKB cohort. Specifically, we uncovered protein marker patterns associated with dyslipidemia, insulin resistance, hyperglycemia, hypertension, and related metabolic traits. Using machine learning, we identified a three-protein panel that improved the prediction of SLD over liver enzymes alone. Notably, the obesity intervention reduced adiposity, which was associated with decreases in circulating cytokines, highlighting the potential for early interventions aimed at mitigating inflammation-related cardiometabolic dysfunction.
Results
Study design
The study population was derived from The HOLBAEK Study11,12. It is composed of two cohorts: (1) an obesity clinic cohort, including children and adolescents with a BMI ≥ 90th percentile (BMI SDS ≥ 1.28)13 who participated in a comprehensive, multidisciplinary non-pharmacological obesity management program at the Children’s Obesity Clinic, Holbaek Hospital; and (2) a population-based cohort, recruited from the school system across 11 municipalities across Zealand, Denmark.
Participant assessments included anthropometry, whole-body dual-energy X-ray absorptiometry (DXA) scanning14, proton magnetic resonance spectroscopy (1H-MRS)15, and blood biochemistry12,16,17,18,19,20,21,22,23,24,25. PEA-based proteomics (Target 96 Cardiovascular II and Inflammation panels, Supplementary Data 1) were performed on 4024 children and adolescents with baseline examinations and available blood samples (overweight/obesity, n = 2377, and normal weight individuals from the population-based cohort, n = 1647). As well as 184 participants who underwent obesity management and had a median follow-up period of 1.1 years (Target 96 Inflammation panel)25. We utilized the adult UKB cohort, with available PEA-based plasma proteomics, for external validation. An overview of the study design is presented in Fig. 1.
Plasma proteomic profiles were measured using Olink Target 96 CVDII and INF panels in a cross-sectional study of 2377 children and adolescents with overweight/obesity and 1647 with normal weight. We assessed associations between circulating protein markers with age, sex, and overweight/obesity, as well as with cardiometabolic risk profiles. We evaluated the predictive performance of circulating protein biomarkers for detecting steatotic liver disease. Additionally, the Olink INF panel was measured in a subset of 184 children and adolescents undergoing non-pharmacological obesity treatment. We examined changes in INF proteins in response to BMI SDS reduction and investigated longitudinal associations between protein marker changes and cardiometabolic risk profiles. Created in BioRender. Stinson, S. (2025) https://BioRender.com/yloww5o. BMI body mass index, CVDII cardiovascular II, INF inflammation, SDS standard deviation score.
Study participant characteristics and cardiometabolic risk profiles
In the cross-sectional study, the group with overweight/obesity exhibited significantly higher cardiometabolic risk variables compared to the normal weight group (Supplementary Data 2). Specifically, the overweight/obesity group differed in terms of anthropometric measures, liver-related measures (liver fat % and liver enzymes), lipid profiles, markers of general inflammation, glycemic indicators, and blood pressure. Compared to their normal-weight counterparts, the overweight/obesity group had a higher prevalence of SLD, defined as liver fat ≥ 5.0% (16.3% vs. 0%), elevated ALT levels (32.0% vs. 9.5%), dyslipidemia (21.7% vs. 5.9%), insulin resistance (44.5% vs. 6.8%), hyperglycemia (16.8% vs. 7.0%), and hypertension (14.9% vs. 5.6%), underscoring the presence of significant cardiometabolic risk within this group.
Associations of age, sex, and puberty with protein markers
Our analysis revealed significant associations between age and circulating protein levels, with 130 out of 149 proteins (87.2%) showing age-related differences independent of sex and BMI SDS (P < 5% FDR, Fig. 2a, Supplementary Data 3). Of these, 63 proteins were positively and 67 were negatively associated with age. Notably, circulating AGRP, LEP, and SPON2 levels increased with age. Conversely, proteins such as DNER, MMP12, and TNFRSF9 levels decreased with age.
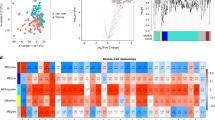
Significant associations of a 130 out of 149 plasma proteins with age, adjusted for sex and BMI SDS. b 83 out of 149 proteins with sex, adjusted for age and BMI SDS. c 36 out of 149 proteins with puberty (Tanner stage), adjusted for age, sex, and BMI SDS. d 122 out of 149 proteins with overweight/obesity compared to normal weight, adjusted for sex and age. The volcano plots (a–d) show the estimated effect sizes from multiple linear regressions (n = 2377 with overweight/obesity and 1647 with normal weight for age, sex, and BMI SDS as predictors; n = 1704 with overweight/obesity and 1169 with normal weight for puberty as a predictor). Gray circles indicate non-significant (NS) associations, while red and blue represent positive and negative associations. Statistical significance was assessed using two-sided P values adjusted for multiple comparisons using the false discovery rate (FDR) method at a 5% threshold. e Venn diagram showing the number of FDR significant proteins associated with age, sex, puberty, and overweight/obesity, highlighting overlapping and unique protein associations. f Scatter plot comparing β-values for protein associations with overweight/obesity between The Holbaek Study and the UK Biobank (UKB, n = 45,825), demonstrating concordance across cohorts, with examples of exceptions, including OPG. Source data are provided as a Source data file.
Sex-specific differences were observed for 83 out of 149 proteins (55.7%), independent of age and BMI SDS (Fig. 2b, Supplementary Data 3). Of these, 31 proteins were significantly higher in girls, and 52 were significantly higher in boys. Notably, girls exhibited higher levels of LEP, GH, and SERPINA12, while boys displayed elevated levels of TNFRSF9, PGF, and DNER.
Puberty-specific differences were observed for 36 out of 149 proteins (24.2%), independent of age, sex, and BMI SDS (Fig. 2c, Supplementary Data 3). Of these, 22 proteins were significantly higher in puberty/post-puberty (Tanner stage = 2–5) and 14 were significantly higher in pre-puberty (Tanner stage = 1). Notably, puberty/post-puberty was associated with higher levels of PGF, TRANCE, and BOC, while pre-puberty was associated with higher levels of FS, CTRC, and ADM.
Associations of overweight/obesity with protein markers
Overweight/obesity was associated with significant alterations in circulating protein levels, impacting 122 out of 149 proteins (81.9%), independent of age and sex (Fig. 2d, Supplementary Data 3). Of these, 81 proteins were positively associated, and 41 were negatively associated with overweight/obesity compared to normal weight. Notably, LEP, IL1ra, and ADM levels were higher with overweight/obesity, and PTX3, GH, and GDF2 levels were higher with normal weight.
We found that 43 out of 149 proteins (29%) were significantly associated with age, sex, and overweight/obesity, while 19 out of 149 proteins (12.8%) were significantly associated with all four factors, including puberty (Fig. 2e; Supplementary Fig. 1).
To explore these associations further across the life course, we compared protein markers linked to overweight/obesity in the pediatric HOLBAEK cohort with those observed in adulthood, utilizing the UKB (Fig. 2f, Supplementary Data 4). The direction of effects was largely consistent across the two cohorts. Notably, some proteins displayed significant associations in The HOLBAEK Study but not in the UKB. For example, OPG was inversely associated with overweight/obesity in The HOLBAEK Study (β = −0.25, P = 1.27E-15) but showed a non-significant association in the UKB (β = 0.00, P = 0.61).
Influence of overweight/obesity on age-associated protein markers
We examined age-related differences in circulating protein markers across weight groups using partial least squares-discriminant analysis (PLS-DA) (Fig. 3a, b). Age groups were categorized into: Group 1 (girls < 9 years, boys < 10 years), Group 2 (girls 9–15 years, boys 10–16 years), and Group 3 (girls >15 years, boys >16 years)26. The PLS-DA models showed good discriminatory performance across age groups for participants with overweight/obesity and normal weight (Q² = 0.214 and 0.270, respectively; permutation p = 0.001 for both). Key contributors included proteins such as TNFRSF9, MMP12, and DNER in both weight groups, with LEP exhibiting higher importance in the group with obesity.
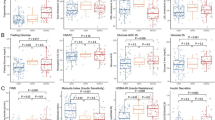
PLS-DA score plot showing plasma protein profiles across three age groups in children and adolescents with a overweight/obesity and b normal weight. c Associations between age and top 26 proteins with significant age × overweight/obesity interactions (FDR P < 1.0E-04). Linear regression models included an interaction term for age × overweight/obesity, adjusted for sex. β-coefficients (center) with 95% CIs (error bars) are shown separately for the overweight/obesity (light blue; n = 2377) and normal weight (brown; 1647) groups. d Box plots of the NPX levels for the four proteins with the largest age-related differences between overweight/obesity and normal weight, visualized across the three age groups. In box plots, the center line represents the median, the bounds of the box indicate the interquartile range (IQR), and the whiskers represent the minimum and maximum values within 1.5 × IQR. Comparisons between overweight/obesity and normal weight groups within each age group were performed using two-sided t-tests. Exact P values are: OSM: age group 1, P = 0.63; age group 2, P = 1.65E-46; age group 3, P = 4.00E-24. LEP: age group 1, P = 3.87E-194; age group 2, P = 1.31 E-296; age group 3, P = 1.44E-64. IL1ra: age group 1, P = 2.72E-93; age group 2, P = 2.42E-277; age group 3, P = 6.63E-76. HGF: age group 1, P = 1.02E-11; age group 2, P = 5.80E-59; age group 3, P = 5.86E-28. Three asterisks (***) indicate a significant difference between groups (P < 0.001); ns not significant. Sample sizes: n = 586, 1435, and 343 for age groups 1, 2, and 3 in the overweight/obesity group; and n = 486, 863, and 294 for age groups 1, 2, and 3 in the normal weight groups, respectively. Source data are provided as a Source data file.
While these results confirm age-related proteomic shifts in both groups, we next examined whether these trajectories differ by weight status by including an interaction term (age × [overweight/obesity vs. normal weight]) in regression models for each protein, adjusting for sex. We identified significant interactions (P < 5% FDR) for 90 out of 149 proteins (60.4%), indicating that the relationship between age and protein levels also differs according to weight status (Supplementary Data 5). The top 26 interactions (FDR P < 1.0E-04) are illustrated in Fig. 3c. Notably, several proteins showed amplified age-related increases in the overweight/obesity group compared to the normal weight group. For example, OSM exhibited a positive age-related trend in the overweight/obesity group (β = 0.22), yet a slight negative trend in the normal weight group (β = −0.04, PInteraction = 6.23E-18), same with HGF (βoverweight/obesity = 0.14 vs. βnormal weight = −0.03, PInteraction = 2.32E-09). Similarly, LEP (βoverweight/obesity = 0.33 vs. βnormal weight = 0.18, PInteraction = 1.37E-13) and IL1ra (βoverweight/obesity = 0.05 vs. βnormal weight = −0.11, PInteraction = 7.96E-11) showed stronger age-related increases.
To pinpoint when these age × overweight/obesity interactions emerge, we examined differences in these protein markers across the three age groups according to overweight/obesity or normal weight (Fig. 3d). Most markers were higher across all three age groups. Notably, OSM displayed a unique pattern, with significant differences only between age Groups 2–3, suggesting a potential age-related increase during puberty, which may be influenced by weight status.
Protein marker associations with cardiometabolic risk profiles
Of the 149 proteins analyzed, 128 proteins were significantly associated with at least one dichotomized cardiometabolic risk feature (P < 5% FDR), independently of age, sex, and BMI SDS (Supplementary Data 6). Specifically, 118 proteins were linked to a higher prevalence of cardiometabolic conditions, including SLD (liver fat ≥ 5%), elevated ALT, dyslipidemia, insulin resistance, hyperglycemia and hypertension; while 16 proteins associated with a lower prevalence of these risk features (Fig. 4a, Supplementary Fig. 2). Several proteins had divergent direction of effects across the cardiometabolic risk features, including proteins such as FS and FGF19.
a Top 27 plasma proteins associated with cardiometabolic risk features (P < 5% FDR), identified through logistic regression analysis, adjusted for age, sex, and BMI SDS. b Associations of top proteins with cardiometabolic traits, tested with linear regression, adjusted for age, sex, and BMI SDS. c Associations between five proteins and liver enzymes with significant overweight/obesity interactions (P < 5% FDR). Linear regression models included an interaction term for overweight/obesity vs. normal weight, adjusted for age and sex. β-coefficients (center) with 95% CIs (error bars) are shown separately for the overweight/obesity (light blue; n = 2356) and normal weight (brown; n = 1619) groups. d Discriminant accuracy of three proteins and liver enzymes for diagnosing steatotic liver disease (SLD; liver fat ≥ 5.0%). The protein panel was identified via LASSO regression for feature selection, followed by logistic regression for final classification. Analysis was performed on 425 participants, with controls down-sampled to match the 76 cases of SLD. Each ROC curve is accompanied by its corresponding 95% CI (shaded area), and the mean AUC values with their respective 95% CIs are provided for each curve. AUC area under the curve. Source data are provided as a Source data file.
For continuous phenotypes, all 149 proteins had at least one significant association (P < 5% FDR) with a cardiometabolic trait, independently of age, sex, and BMI SDS (Fig. 4b; Supplementary Data 7; Supplementary Fig. 2). For liver-related traits, examples of top significant positive predictors of ALT levels include HAOX1, ACE2, and CDCP1. For lipid-related traits, PRSS8, FGF21, and AMBP emerged as the strongest positive predictors for plasma triglyceride levels. For glycemic-related traits, LEP and PRSS8 emerged as the strongest positive predictors of insulin; on the other hand, FS was the strongest negative predictor. In general, FS was inversely associated with glycemic traits, yet positively associated with most other cardiometabolic risk factors. For inflammation-related traits, OSM, CEACAM8, and LOX1 were the most notable positive predictors of WBC. For proglucagon hormones, we observed FGF19, CCL20, and CCL11 among the strongest positive predictors for total glucagon-like peptide-1 (GLP-1). For blood pressure-related traits, FGF21, IL1ra, and LEP were identified as the strongest positive predictors of diastolic blood pressure (DBP).
Sex-specific associations of protein markers with cardiometabolic traits
We investigated sex differences in the associations between circulating proteins and cardiometabolic traits, revealing significant sex interactions (P < 5% FDR) for 64 out of 149 proteins (43.0%) across 13 of 21 cardiometabolic traits, independently of age and BMI SDS (Supplementary Data 8). Notably, several proteins exhibited opposite directions of effect between girls and boys. For example, TNFRSF9 was positively associated with ALT levels in girls (β = 0.14) but negatively in boys (β = −0.15, PInteraction = 8.61E-19). Similarly, FGF21 (βgirls = -0.05 vs. βboys = 0.15, PInteraction = 8.35E-11), CD4 (βgirls = 0.14 vs. βboys = -0.07, PInteraction = 2.13E-10) and TWEAK (βgirls = 0.04 vs. βboys = -0.15, PInteraction = 1.24E-10) showed sex-dependent associations with ALT levels.
Influences of overweight/obesity on protein marker-cardiometabolic associations
To investigate the role of overweight/obesity in modulating the associations between protein levels and cardiometabolic traits, we assess interaction effects (protein × [overweight/obesity vs. normal weight]), independent of age and sex (Supplementary Data 9). Significant interactions were observed for 127 out of 149 proteins (85.2%) across 19 out of 21 cardiometabolic traits (P < 5% FDR). For liver-related traits, the strongest interactions with overweight/obesity on ALT levels were observed for HAOX1, ACE2, LEP, FGF21, and CDCP1. Notably, FGF21 showed opposite direction of effects between weight groups (Fig. 4c). For lipid-related traits, LEP, ADM, and PTX3 had the most significant interactions with overweight/obesity for triglycerides. For glycemic-related traits, the strongest interactions with overweight/obesity were observed for LEP, ADM, and SCF with fasting serum insulin levels. For inflammation-related traits, LEP, FGF21, and IL1ra had the most significant interactions with overweight/obesity for WBC levels. For proglucagon hormones, LEP, FGF21, and HGF had the most significant interactions with overweight/obesity for fasting glucagon. For blood pressure-related traits, LEP, ADM, and DNER emerged as the strongest interaction with overweight/obesity for DBP.
Circulating protein marker panel for prediction of steatotic liver disease
To improve non-invasive detection of SLD, defined as liver fat exceeding 5%, we applied LASSO regression for initial feature selection, followed by logistic regression for final classification. This approach identified a three-protein panel consisting of CDCP1, FGF21, and HAOX1, achieving a mean cross-validated area under the receiver operating characteristic curve (ROC-AUC) of 0.81 (95% CI: 0.79–0.83) using five-fold cross-validation repeated ten times (Fig. 4d). Notably, combining this protein panel with traditional liver enzymes (ALT, AST, and GGT) significantly increased the AUC from 0.77 (95% CI: 0.75–0.79) to 0.83 (95% CI: 0.81–0.84), as confirmed by DeLong’s test (P < 0.05). These findings underscore the added predictive value of circulating protein biomarkers in assessing SLD risk.
Mediation effect of protein markers on cardiometabolic traits
To investigate whether the effect of obesity on cardiometabolic risk is explained in part by certain protein markers, we performed mediation analysis, adjusted for age and sex. Out of 149 proteins analyzed, 122 significantly mediated the effect of overweight/obesity on 21 cardiometabolic traits (P < 5% FDR), with a median proportion of 4.5%. Notably, 38 proteins exhibited substantial mediation effects, >20% of the total effect. The top ten proteins included LEP, IL1ra, Gal9, IL18R1, CDCP1, HAOX1, HGF, ACE2, PARP1, and GH, which mediated the associations between obesity on a broad range of cardiometabolic traits, highlighted by hierarchical clustering (Supplementary Fig. 3 and Supplementary Data 10).
Protein marker changes in response to individualized pediatric obesity treatment
We evaluated protein marker changes in response to personalized obesity treatment among 184 children and adolescents (84 boys) with overweight/obesity, with a median age of 11.6 years (IQR: 9.9–13.5). Participants with complete data on BMI SDS and blood samples at baseline and follow-up were included, with a median follow-up duration of 1.1 years (IQR: 1.0–1.2). Of these, 151 showed a reduction in BMI SDS, while 33 maintained or increased their BMI SDS. The median decrease in BMI SDS was −0.39 (IQR: −0.76 to −0.07), accompanied by significant improvements in body fat percentage, lipid levels, glucose, HbA1c, and blood pressure (all P < 0.05; Supplementary Data 11), as previously reported25.
We next explored the relationship between BMI SDS reduction and changes in inflammatory (INF) protein levels, independently of age, sex, and treatment duration. A total of 14 out of 64 INF proteins were significantly associated with BMI SDS reduction (P < 5% FDR). Notably, IL18R1, FGF21, IL10RB, and CDCP1 showed the largest decrease following BMI SDS reduction (Fig. 5a, Supplementary Data 12).
a BMI SDS reduction in individuals undergoing obesity treatment (n = 184) was associated with changes in 14 INF proteins (P < 5% FDR), assessed using linear regression, adjusted for age, sex, and treatment duration. The β-coefficients (center) with 95% CIs (error bars) are shown for each protein. BMI SDS reduction was calculated as the difference between baseline BMI SDS and follow-up values. b Changes in INF proteins in individuals undergoing obesity treatment (n = 184) were associated with changes in cardiometabolic traits, tested by linear regression, adjusted for age, sex, treatment duration, baseline BMI SDS, and changes in BMI SDS. Changes in INF proteins and cardiometabolic traits were calculated as the difference between follow-up and baseline values (P < 5% FDR). Source data are provided as a Source data file.
Longitudinal analyses revealed 62 significant associations between 47 out of 64 INF proteins and eight out of 17 cardiometabolic risk factors (P < 5% FDR), independently of age, sex, treatment duration, baseline BMI SDS, and changes in BMI SDS (Fig. 5b, Supplementary Data 13). For liver-related traits, decreases in IL18R1, CDCP1, CXCL11, and CXCL9 levels were linked to reductions in ALT levels, and a suggestive association of decreased CDCP1 with reductions in liver fat %, yet not statistically significant (P = 0.092). For lipid-related traits, changes in CX3CL1 levels were negatively associated with changes in LDL-C and TC. Numerous INF protein changes were associated with changes in glycemic-related traits. Decreases in TNFSF14, ST1A1, CD40, Flt3L, and VEGFA levels were linked to reductions in homeostasis model assessment of insulin resistance (HOMA-IR) levels. For inflammation-related traits, decreases in HGF, OSM, TGFa, ENRAGE, and MMP10 were associated with reductions in WBC levels. No INF proteins were significantly associated with changes in blood pressure-related traits, after multiple testing corrections.
Mediation effect of changes in protein markers on cardiometabolic traits
We investigated whether changes in protein markers mediate the relationship between BMI SDS reduction and changes in cardiometabolic traits, independently of age, sex, and treatment duration (Supplementary Fig. 4, Supplementary Data 14). Changes in 18 proteins significantly mediated the association between BMI SDS reduction and changes in 12 out of 17 cardiometabolic risk factors (P < 5% FDR). Notably, IL18R1 and CDCP1 were key positive mediators in the relationship between BMI SDS reduction and improvements in liver- and glycemic-related traits. Interestingly, CDCP1 exhibited inverse mediation effects on lipid levels, suggesting a complex role in lipid metabolism.
Discussion
Systemic inflammation and disrupted metabolic pathways characterize pediatric obesity, contributing to increased risk of developing future cardiometabolic comorbidities2. Our study highlights the profound impact of obesity on circulating protein markers in children and adolescents, revealing strong associations with cardiometabolic risk profiles. Although prior studies in pediatric populations have been limited, advances in proteomics technology and the availability of large-scale biobanks now enable the identification of protein markers associated with cardiometabolic dysfunction. Recent work from our group has mapped proteomic changes related to age, sex, and BMI SDS in The HOLBAEK Study using mass-spectrometry (MS)-based approaches, quantifying approximately 1200 plasma proteins in 3147 children and adolescents9. The present study further advances this field, utilizing PEA technology to capture lower-abundant proteins, such as inflammatory markers, which remain challenging to quantify using MS methods5. Additional pediatric biobanks include the ALSPAC population-based cohort, of which 92 proteins have been measured in non-fasted plasma samples using the Olink Target Inflammation panel in 3005 9-year-olds, yet the results of this study are not yet published10.
Our findings also align with studies performed in adults, including those from the BAMSE cohort (2074 adults; mean age of 22.5 years)27 and the UKB (46,825 adults; mean age of 56.5 years)8. In the present study, we demonstrated consistent directions of effects of adiposity associated with circulating plasma proteins across childhood (HOLBAEK) and adulthood (UKB). Strikingly, some proteins exhibited age-specific associations, present only in childhood and adolescence, particularly those related to factors such as bone metabolism. For example, OPG, a known marker of bone strength28 was inversely associated with overweight/obesity in The HOLBAEK Study, but was non-significant in the UKB, highlighting certain proteins that may have age-specific effects during growth and development.
In the present study, we identified key protein signatures associated with cardiometabolic risk patterns. For liver-related phenotypes, we observed HAOX1, ACE2, and CDCP1 to be among the strongest positive predictors of plasma ALT levels, proteins previously associated with adult MASLD29,30,31. For lipid traits, proteins such as PRSS8 were significantly associated with triglyceride levels. Notably, variants in the PRSS8 gene have previously been implicated in lipid metabolism through genome-wide association studies32. We observed a divergent pattern across cardiometabolic traits for FS (follistatin), which was inversely associated with glycemic traits, yet positively associated with most other cardiometabolic risk factors. Folkersen et al. previously highlighted an inverse causal effect of FS levels on fasting glucose and insulin through Mendelian randomization analysis33. For traditional inflammation-related phenotypes, we observed significant associations between hs-CRP and WBC with markers such as LOX1, a biomarker for ischemic stroke and a potential drug target34. Moreover, prior work from our group found LOX1 to be associated with inflammation in pediatric obesity23. For proglucagon hormones, we observed strong interactions between overweight/obesity and proteins, including LEP, FGF21, and HGF, on fasting glucagon levels, which may highlight the potential role of the liver-α-cell axis in obesity-related metabolic dysregulation35. We also observed significant links between several inflammatory proteins, such as FGF19, CCL20, and CCL11, and fasting GLP-1 levels, which aligns with the theory that GLP-1 has more diverse functions, beyond simply stimulating insulin secretion, including modulating response to inflammation36. For blood pressure-related traits, we observed certain proteins, including FGF21 and IL1ra, significantly associated with DBP SDS. FGF21 levels are associated with hypertension in adults37, and IL1ra antagonists are in clinical trials to reduce blood pressure38.
Heterogeneity in treatment response remains a critical challenge in treating pediatric obesity1. Over 80% of participants exhibited reductions in the degree of obesity, while approximately 20% were non-responders, highlighting the complexity of obesity management. Our data revealed significant decreases in the levels of 14 inflammatory proteins as a result of BMI SDS reduction, which correlated with improvements in cardiometabolic risk factors during the intervention. Notably, reductions in IL18R1, FGF21, and CDCP1 were linked to improved glycemic traits and liver enzyme profiles, suggesting potential pathways for targeted therapeutic strategies. However, the persistence of elevated levels of certain proteins indicates that lifestyle interventions alone may be insufficient for all patients. Pharmacological approaches, such as GLP-1 receptor agonists, could supplement personalized obesity management by targeting specific inflammatory pathways, when lifestyle management alone proves insufficient39.
To further elucidate the putative biological mechanisms underlying these associations, mediation analysis revealed that several key proteins, including LEP, IL1ra, Gal9, IL18R1, CDCP1, HAOX1, and HGF, partially explained the relationship between obesity and multiple cardiometabolic traits. Notably, IL18R1 and CDCP1 played a significant role in linking BMI SDS reduction to improvements in liver function and glycemic control, suggesting they may be important mediators of metabolic adaptation during obesity treatment. Interestingly, CDCP1 demonstrated opposing mediation effects on lipid traits, indicating it may have a more nuanced role in lipid metabolism.
A key finding of our study was the identification of a three-protein panel: CDCP1, FGF21, and HAOX1, that outperformed traditional liver enzymes in predicting SLD (defined by ≥5.0% liver fat), with an ROC-AUC of 81% compared to 77% for liver enzymes alone. The combination of the protein panel with liver enzymes further improved diagnostic accuracy to 83%. CDCP1 is a protein primarily expressed in skeletal muscle and colon, primarily functions in cell adhesion and migration, and has emerged as a particularly promising biomarker, with previous studies reporting elevated levels in adults with metabolic dysfunction-associated steatohepatitis (MASH)29 and in pediatric obesity40. In the present study, CDCP1 was strongly associated with overweight/obesity and cardiometabolic risk factor clustering, and CDCP1 levels were decreased during the intervention, which was associated with improvements in liver and glycemic-related traits. Similarly, FGF21 and HAOX1, both primarily expressed in the liver, were also strongly linked to overweight/obesity and liver disease markers in the present study41,42, underscoring their relevance as potential non-invasive biomarkers for SLD and cardiometabolic health in a pediatric population.
Although the observed improvements in AUC were moderate, even incremental gains can have clinical significance in a pediatric setting, where options for non-invasive biomarkers for SLD are limited. Furthermore, the clinical utility of proteomics-based risk scores has been demonstrated in adult populations, underscoring the potential of similar approaches in children and adolescents43,44. Additional validation of this three-protein panel in external pediatric cohorts, ideally with biopsy-confirmed MASLD, is warranted. Previous work from our group identified a circulating lipid panel predictive of SLD25 with similar predictive performance to the present protein-based model, suggesting that combining data across several omics layers could further improve risk prediction. Importantly, ceramides, key lipids associated with cardiometabolic risk, correlated with the three proteins in our predictive panel, highlighting potential for interconnected proteomic-lipidomic pathways in metabolic regulation. Future research integrating proteomics, lipidomics, and genomics may help disentangle causal relations and reveal underlying mechanisms of obesity-related cardiometabolic risk.
This study has limitations that warrant consideration. Our cohort was predominantly of European ancestry, limiting generalizability to more diverse populations. Additionally, the targeted proteomics approach restricted the analysis to proteins captured by the PEA panels, potentially overlooking other relevant biomarkers. We did not account for genetic predispositions45, lifestyle factors, such as diet and physical activity46, or treatment adherence, all of which may influence changes in circulating protein levels in response to the intervention. Recent evidence suggests that lifestyle factors explain only a small fraction of protein variation compared to non-modifiable factors, including genetics46. However, understanding these sources of individual variability could help identify subgroups more likely to benefit and improve personalized treatment strategies. The relatively small number of SLD cases warrants cautious interpretation of the predictive model’s performance. Furthermore, the longitudinal analysis, with only two time points and a relatively short follow-up (median 1.1 years), and the absence of a control group, limit our ability to establish temporality, whether changes in protein levels due to the obesity management intervention precede changes in cardiometabolic risk. Determining causal relationships would require follow-up studies using methods such as Mendelian randomization. Lastly, we have compared proteomics data between The HOLBAEK Study and UKB, which used two slightly differing methodologies, Olink Explore 3072 vs. the Target 96 platforms. Additional cohort-specific differences may exist, including intake of medications, which is more common in adults and may affect circulating protein levels. Nevertheless, our study’s strengths lie in the inclusion of deeply phenotyped cohorts of children and adolescents from both clinical and population-based settings, with integration of both cross-sectional and intervention data, as well as comparison to an adult cohort.
In conclusion, our study highlights the potential of proteomic profiling to identify circulating biomarkers implicated in the pathophysiology of pediatric obesity and its associated cardiometabolic comorbidities. The three-protein panel for SLD offers a promising non-invasive diagnostic tool, paving the way for earlier detection and possible intervention. Importantly, obesity treatment may modulate circulating inflammatory protein levels and reduce cardiometabolic risk, though a subset of patients may require adjunctive pharmacotherapy. Future research should focus on validating these findings in diverse populations and explore further integration across various omics layers.
Methods
Study population
Both cohorts from The HOLBAEK Study were enrolled between January 2009 and April 2019. Among 4133 children and adolescents with available plasma proteomics in the cross-sectional study, participants were excluded based on diagnosed type 1 diabetes mellitus or T2D (n = 7); intake of medications including statins, insulin, metformin and liraglutide (n = 8); meeting T2D criteria based on blood sample (fasting plasma glucose ≥7.0 mmol l−1 and/or HbA1c ≥ 48 mmol mol−1) (n = 8); the interval between visit and blood sample collection >90 days (n = 30); underweight (BMI <5th percentile [BMI SDS < −1.64]) (n = 53). As a result, 4024 participants (54.6% girls, 45.4% boys) were included in the cross-sectional analysis.
Children and adolescents with overweight or obesity (BMI SDS ≥ 1.28) enrolled in the multidisciplinary, family-based and individual-centered obesity clinic cohort received comprehensive management using an evidence-based treatment protocol which comprises a range of recommendations on nutrition, including meal exercises, picky eating, exercise, inactivity, border setting, promoting growth, development and improved physical, mental and social thriving11. The intervention study included 200 children and adolescents with overweight or obesity (56.0% girls, 44.0% boys), who were followed for a median of 1.1 years (IQR 1.0–1.2), and 184 had available blood samples at follow-up. Their proteomic profiles were available at both baseline and follow-up.
Anthropometric measurements
In the obesity clinic cohort, anthropometric data were obtained at clinical examinations, whereas the population-based group was assessed in a mobile laboratory by trained medical professionals22. Weight, height, waist, and waist-hip ratio were measured. BMI SDS was calculated according to a Danish reference13, waist SDS and waist-hip SDS were calculated according to age- and sex-specific reference values47. For systolic blood pressure (SBP) and DBP, mean values for the last two measurements of BP were calculated and converted to BP SDS, based on age-, sex-, and height-specific reference values from the American Academy of Pediatrics48.
Puberty stage
Tanner stage49,50 was evaluated by trained medical professionals in the obesity clinic cohort and self-reported using a questionnaire with picture pattern recognition in the population-based cohort. The puberty stage was then classified as either pre-pubertal = Tanner stage 1 or pubertal = Tanner stage 2–5.
Proteomics
PEA was performed using the Target 96 Cardiovascular II (CVDII) and Target 96 Inflammation (INF) panels from Olink Proteomics on EDTA plasma at baseline23,24,25, and the INF panel at follow-up. The targeted panels were designed by Olink to capture a broad spectrum of circulating proteins with biological relevance to CVD and inflammation, informed by prior literature, and optimized for assay performance. PEA technology uses nucleic acid labeling of antibodies in combination with qPCR, producing normalized protein expression values as an arbitrary unit on a log2 scale. Overall, 85 markers from CVDII and 64 markers from INF for a total of 149 proteins were included, as >80% of individuals were above the limit of detection (LOD; Supplementary Data 1). Each Protein was inverse-rank normalized, including NPX data below the LOD, prior to analyses and association testing.
DXA examination
Whole-body DXA scans were performed and total body fat percentage was quantified in the overweight/obesity (n = 1330), normal weight (n = 113) groups and 124 children and adolescents with overweight or obesity who received the obesity management, using a GE Lunar Prodigy (DF+10031, GE Healthcare) until October 2009 and thereafter using a GE Lunar iDXA (ME+200179, GE Healthcare)14.
1H-MRS examination
Liver fat content was quantified in the overweight/obesity (n = 479) and normal weight (n = 34) groups, as well as 96 children and adolescents with overweight/obesity who received obesity management, using a 3 T Achieva MR imaging system (Philips Medical Systems)15. Data processing was performed by a trained magnetic resonance physicist.
Biochemical analyses
Venous blood samples were collected after an overnight fast. Fasting biochemical measurements include in plasma: alanine transaminase (ALT), aspartate transaminase (AST), γ-glutamyl transferase (GGT), lactate dehydrogenase (LDH) and bilirubin16, high-density lipoprotein cholesterol (HDL-C), low-density lipoprotein cholesterol (LDL-C), total cholesterol (TC), triglycerides (TG)20, glucose17, glucagon19 and GLP-118, in serum: insulin, C-peptide17, high-sensitivity C-reactive protein (hs-CRP)22, and in whole blood: hemoglobin A1c (HbA1c)17 and white blood cell count (WBC)22.
Defining cardiometabolic risk features
SLD was defined using the histological cutoff of ≥5.0% liver fat in adults51. We also defined high ALT (above 24.5 U/L in girls and above 31.5 U/L in boys), which was found to be the optimal cutoff for diagnosing SLD by our group16. Hyperglycemia was defined as fasting plasma glucose ≥5.6 to 6.9 mmol/L and/or HbA1c ≥ 39 to 47 mmol/mol, according to the American Diabetes Association guidelines for prediabetes52. Insulin resistance was defined based on HOMA-IR value above the 90th percentile of previously published age- and sex-specific population-based reference values from our group17. HOMA-IR was calculated as (insulin mU/L × glucose mM)/22.5. Dyslipidemia was defined as values above the 95th percentile according to pediatric guidelines, corresponding to: TC ≥ 200 mg/dL (5.2 mM), LDL-C ≥ 130 mg/dL (3.4 mM), TG ≥ 100 mg/dL (1.1 mM) for 0–9 years or ≥130 mg/dL (1.5 mM) for 10–19 years, or HDL-C < 40 mg/dL (1.0 mM)53. Hypertension was defined as a SBP and/or DBP above the 95th percentile for children aged 1–12 years, and above 130/80 mm Hg for 13 years and older48.
Statistical analyses
Statistical analyses were performed using R software version 4.4.154. Descriptive summary data are expressed as median (IQR) for continuous variables or frequencies and percentages for categorical variables. The Wilcoxon rank-sum test (for continuous variables) and the chi-squared test (for categorical variables) were used to test for differences in characteristics between two groups.
Multiple linear regressions were used to examine the association of age, sex, puberty, or overweight/obesity versus normal weight with each protein individually. PLS-DA was performed to examine how protein profiles differ between the three age groups, separately for overweight/obesity versus normal weight groups, using the ropls v.1.36.0 R package55. Ten-fold cross-validation and 300 permutations were used. The first two component scores were plotted in a score plot, where each point represents an individual.
The effect of obesity on the association between continuous age and individual circulating protein markers was tested by a corresponding interaction model including an interaction term (age × overweight/obesity vs. normal weight), adjusting for sex. Cardiometabolic traits were log-transformed except for BMI SDS, SBP SDS, and DBP SDS. The associations of protein biomarkers with cardiometabolic risk features and cardiometabolic traits were examined using multiple logistic and linear regressions adjusted for age, sex, and BMI SDS when pooling the overweight/obesity and normal weight groups. The respective interactions between sex and protein biomarkers (protein × sex) and obesity (protein × overweight/obesity vs. normal weight) were examined by linear regression for cardiometabolic traits, adjusted for age and either BMI SDS or sex. The reported estimates (β or OR) are based on a 1-s.d. unit increase in independent variables. Multiple testing correction was performed based on FDR at 5% to be considered statistically significant. In all figures, only those proteins with at least one outcome association reaching FDR significance were included.
Changes in protein profiles before and after obesity management were assessed while adjusting for age and sex. The effects of BMI SDS reduction on protein profiles were analyzed using linear regressions controlling for age, sex, and treatment duration. The associations between changes in protein profiles and changes in continuous cardiometabolic traits were examined using linear regressions controlling for age, sex, treatment duration, baseline BMI SDS, and change in BMI SDS. BMI SDS reduction was calculated as the difference between BMI SDS at baseline and BMI SDS at follow-up. Changes in protein profiles and cardiometabolic traits were calculated as the difference between the values at follow-up and those at baseline.
Prediction model
Feature selection for SLD (defined as liver fat above 5%) was performed using LASSO regression implemented in the glmnet v.4.1-8 R package56. We applied five-fold cross-validation repeated ten times, identifying a protein panel with the highest mean ROC-AUC with logistic regression. The discriminative performance of three clinically used liver enzymes (ALT, AST, and GGT) was also evaluated individually and in combination with the protein panel, using the same cross-validation method. To mitigate imbalanced class distribution, a down-sampling approach was applied to the majority class within each cross-validation fold. The statistical comparison of AUCs was conducted using DeLong’s test. These analyses were performed using the caret v.6.0.9457 and pROC v.1.18.558 R packages.
Mediation
Mediation analyses were performed using the mediation 4.5.0 R package59. In the cross-sectional study, mediation analysis was performed to explore the mediating role of overweight/obesity on cardiometabolic traits. Bootstrapping with 1000 iterations was employed to estimate direct, indirect, and total effects across overweight/obesity-protein-trait triangles, adjusted for age and sex, at a significance level of P < 5% FDR. The proportion of effect mediated by overweight/obesity through the protein was determined by dividing its indirect effect by the total effect.
In the intervention study, mediation analysis was performed to examine the mediation effect of protein changes on the associations between BMI SDS reduction and changes in cardiometabolic traits, adjusting for baseline age, sex, and treatment duration, at a significance level of P < 5% FDR. Bootstrap confidence intervals were used to assess the statistical significance of the mediation effects.
Comparison with the UK Biobank
We obtained access to UKB data under accession number 32683, which has received ethical approval from the National Health Service National Research Ethics Service (ref 11/NW/0382), with informed consent obtained from all participants. Olink Explore 3072 PEA was measured quantifying up to 2923 proteins in baseline plasma samples8. We included British individuals with available Olink data, where self-report ethnicity was confirmed by genetics data (UKB field 22006). Proteins common to both The HOLBAEK Study and UKB were matched according to UniProt IDs. We compared effect sizes for associations between protein biomarkers and overweight/obesity (defined as BMI ≥ 25 kg/m2 vs. normal weight: BMI > 18.5 and <25 kg/m2 in UKB) between the two cohorts. Prior to linear regression analysis in the UKB, each protein was rank-inverse normal transformed. Models were adjusted for age, sex, center, fasting time, Olink batch, and time to assay.
Ethics
According to the Declaration of Helsinki, written informed consent was obtained from all participants. An informed oral assent was given by the participant if the participant was younger than 18, and then the parents gave informed written consent. The study was approved by the Ethics Committee of Region Zealand, Denmark (SJ-104) and by the Danish Data Protection Agency (REG-043-2013). The HOLBAEK Study is registered at ClinicalTrials.gov (NCT00928473).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All results from statistical and bioinformatics analysis are provided in Supplementary Data 1–14. The mean levels of proteins are available in the GitHub repository at https://github.com/sarastinson/olink_holbaek/tree/main/median_data. Source data are provided with this paper. In accordance with the General Data Protection Regulation (GDPR, https://gdpr-info.eu/), individual-level clinical and proteomics data generated in this study cannot be made publicly available to maintain patient confidentiality. However, proteomics datasets can be requested from the authors by contacting T.H. at torben.hansen@sund.ku.dk. The obesity management protocol can be requested from J.-C.H. at jhom@regionsjaelland.dk. Access to the data can be granted through the Danish Data Protection Agency and the ethics committee for the Region Zealand, Denmark, subject to obtaining proper approvals and adherence to patient information and data-processing agreements. The response time for data requests is within 1 month. Accessing and processing the data requires compliance with the following conditions: (1) a data-processing agreement must be signed between the data controller and processor; (2) data may only be processed for statistical and scientific studies; (3) personal data must be deleted, anonymized and destroyed at the end of the investigation; and (4) data must not be shared with unauthorized third parties or individuals. Source data are provided with this paper.
Code availability
The code used for data analysis is available on GitHub: https://github.com/sarastinson/olink_holbaek60.
References
Jebeile, H., Kelly, A. S., O’Malley, G. & Baur, L. A. Obesity in children and adolescents: epidemiology, causes, assessment, and management. Lancet Diabetes Endocrinol. 10, 351–365 (2022).
Caprio, S., Santoro, N. & Weiss, R. Childhood obesity and the associated rise in cardiometabolic complications. Nat. Metab. 2, 223–232 (2020).
Stefan, N., Yki-Järvinen, H. & Neuschwander-Tetri, B. A. Metabolic dysfunction-associated steatotic liver disease: heterogeneous pathomechanisms and effectiveness of metabolism-based treatment. Lancet Diabetes Endocrinol. 13, 134–148 (2025).
Soták, M., Clark, M., Suur, B. E. & Börgeson, E. Inflammation and resolution in obesity. Nat. Rev. Endocrinol. 21, 45–61 (2025).
Joshi, A., Rienks, M., Theofilatos, K. & Mayr, M. Systems biology in cardiovascular disease: a multiomics approach. Nat. Rev. Cardiol. 18, 313–330 (2020).
McCafferty, C., Chaaban, J. & Ignjatovic, V. Plasma proteomics and the paediatric patient. Expert Rev. Proteom. 16, 401–411 (2019).
Eldjarn, G. H. et al. Large-scale plasma proteomics comparisons through genetics and disease associations. Nature 622, 348–358 (2023).
Sun, B. B. et al. Plasma proteomic associations with genetics and health in the UK Biobank. Nature 622, 329–338 (2023).
Niu, L. et al. Plasma proteome variation and its genetic determinants in children and adolescents. Nat. Genet. https://doi.org/10.1038/s41588-025-02089-2 (2025).
Goulding, N. et al. Inflammation proteomics datasets in the ALSPAC cohort. Wellcome Open Res. 7, 277 (2024).
Holm, J.-C. et al. Chronic care treatment of obese children and adolescents. Int. J. Pediatr. Obes. 6, 188–196 (2011).
Lausten-Thomsen, U. et al. Reference values for serum total adiponectin in healthy non-obese children and adolescents. Clin. Chim. Acta 450, 11–14 (2015).
Nysom, K., Mølgaard, C., Hutchings, B. & Michaelsen, K. F. Body mass index of 0 to 45-y-old Danes: reference values and comparison with published European reference values. Int. J. Obes. Relat. Metab. Disord. 25, 177–184 (2001).
Nielsen, T. R. H. et al. Childhood obesity treatment; effects on BMI SDS, body composition, and fasting plasma lipid concentrations. PLoS ONE 13, e0190576 (2018).
Chabanova, E., Fonvig, C. E., Bøjsøe, C., Holm, J.-C. & Thomsen, H. S. H. MRS assessment of hepatic fat content: comparison between normal- and excess-weight children and adolescents. Acad. Radiol. 24, 982–987 (2017).
Johansen, M. J. et al. The effect of overweight and obesity on liver biochemical markers in children and adolescents. J. Clin. Endocrinol. Metab. 105, dgz010 (2020).
Frithioff-Bøjsøe, C. et al. Glucose metabolism in children and adolescents: population-based reference values and comparisons to children and adolescents enrolled in obesity treatment. Pediatr. Diabetes 20, 538–548 (2019).
Stinson, S. E. et al. Fasting plasma GLP-1 is associated with overweight/obesity and cardiometabolic risk factors in children and adolescents. J. Clin. Endocrinol. Metab. 106, 1718–1727 (2021).
Stinson, S. E. et al. Hyperglucagonemia in pediatric adiposity associates with cardiometabolic risk factors but not hyperglycemia. J. Clin. Endocrinol. Metab. 107, 1569–1576 (2022).
Nielsen, T. R. H. et al. Dyslipidemia and reference values for fasting plasma lipid concentrations in Danish/North-European White children and adolescents. BMC Pediatr. 17, 116 (2017).
Frithioff-Bøjsøe, C. et al. Leptin, adiponectin, and their ratio as markers of insulin resistance and cardiometabolic risk in childhood obesity. Pediatr. Diabetes 21, 194–202 (2020).
Lund, M. A. V. et al. Low-grade inflammation independently associates with cardiometabolic risk in children with overweight/obesity. Nutr. Metab. Cardiovasc. Dis. 30, 1544–1553 (2020).
Stinson, S. E. et al. High plasma levels of soluble lectin-like oxidized low-density lipoprotein receptor-1 are associated with inflammation and cardiometabolic risk profiles in pediatric overweight and obesity. J. Am. Heart Assoc. 12, e8145 (2023).
Stinson, S. E. et al. The interplay between birth weight and obesity in determining childhood and adolescent cardiometabolic risk. EBioMedicine 105, 105205 (2024).
Huang, Y. et al. Lipid profiling identifies modifiable signatures of cardiometabolic risk in children and adolescents with obesity. Nat. Med. 1, 12 (2024).
Juul, A. et al. Pubertal development in Danish children: comparison of recent European and US data. Int. J. Androl. 29, 247–255 (2006).
Klevebro, S. et al. Inflammation-related plasma protein levels and association with adiposity measurements in young adults. Sci. Rep. 11, 11391 (2021).
Pollock, N. K. Childhood obesity, bone development, and cardiometabolic risk factors. Mol. Cell. Endocrinol. 410, 52–63 (2015).
Jia, X. et al. Identification and multicentric validation of soluble CDCP1 as a robust serological biomarker for risk stratification of NASH in obese Chinese. Cell Rep. Med. 4, 101257 (2023).
Jacobs, A. K. et al. Hepatic angiotensin-converting enzyme 2 expression in metabolic dysfunction-associated steatotic liver disease and in patients with fatal COVID-19. World J. Gastroenterol. 30, 3705–3716 (2024).
Gianmoena, K. et al. Epigenomic and transcriptional profiling identifies impaired glyoxylate detoxification in NAFLD as a risk factor for hyperoxaluria. Cell Rep. 36, 109526 (2021).
Li, Z. et al. Integrating mouse and human genetic data to move beyond GWAS and identify causal genes in cholesterol metabolism. Cell Metab. 31, 741–754.e5 (2020).
Folkersen, L. et al. Genomic and drug target evaluation of 90 cardiovascular proteins in 30,931 individuals. Nat. Metab. 2, 1135–1148 (2020).
Hofmann, A., Brunssen, C., Wolk, S., Reeps, C. & Morawietz, H. Soluble LOX-1: a novel biomarker in patients with coronary artery disease, stroke, and acute aortic dissection? J. Am. Heart Assoc. 9, e013803 (2020).
Hædersdal, S., Andersen, A., Knop, F. K. & Vilsbøll, T. Revisiting the role of glucagon in health, diabetes mellitus and other metabolic diseases. Nat. Rev. Endocrinol. 19, 321–335 (2023).
Hammoud, R. & Drucker, D. J. Beyond the pancreas: contrasting cardiometabolic actions of GIP and GLP1. Nat. Rev. Endocrinol. 19, 201–216 (2022).
Greenhill, C. Link between FGF21 and blood pressure. Nat. Rev. Endocrinol. 14, 380–380 (2018).
Urwyler, S. A. et al. IL (Interleukin)-1 receptor antagonist increases Ang (Angiotensin [1-7]) and decreases blood pressure in obese individuals. Hypertension 75, 1455–1463 (2020).
Xie, Y., Choi, T. & Al-Aly, Z. Mapping the effectiveness and risks of GLP-1 receptor agonists. Nat. Med. 1, 12 (2025).
Manell, H. et al. Biomarker screening in children and adolescents reveals that CUB domain-containing protein 1 is associated with obesity and that hepatocyte growth factor is associated with weight gain. Obes. Med. 39, 100481 (2023).
Geng, L., Lam, K. S. L. & Xu, A. The therapeutic potential of FGF21 in metabolic diseases: from bench to clinic. Nat. Rev. Endocrinol. 16, 654–667 (2020).
Jones, J. M., Morrell, J. C. & Gould, S. J. Identification and characterization of HAOX1, HAOX2, and HAOX3, three human peroxisomal 2-hydroxy acid oxidases. J. Biol. Chem. 275, 12590–12597 (2000).
Niu, L. et al. Noninvasive proteomic biomarkers for alcohol-related liver disease. Nat. Med. 28, 1277–1287 (2022).
Sanyal, A. J. et al. Defining the serum proteomic signature of hepatic steatosis, inflammation, ballooning and fibrosis in non-alcoholic fatty liver disease. J. Hepatol. 78, 693–703 (2023).
Thielemann, R. et al. Obesity- and age-dependent genetic regulation of the plasma proteome in children and adolescents. Preprint at medRxiv https://doi.org/10.1101/2025.03.18.25324169 (2025).
Carrasco-Zanini, J. et al. Mapping biological influences on the human plasma proteome beyond the genome. Nat. Metab. 6, 2010–2023 (2024).
Sharma, A. K., Metzger, D. L., Daymont, C., Hadjiyannakis, S. & Rodd, C. J. LMS tables for waist-circumference and waist-height ratio Z-scores in children aged 5-19 y in NHANES III: association with cardio-metabolic risks. Pediatr. Res. 78, 723–729 (2015).
Flynn, J. T. et al. Clinical practice guideline for screening and management of high blood pressure in children and adolescents. Pediatrics 140, e20171904 (2017).
Marshall, W. A. & Tanner, J. M. Variations in pattern of pubertal changes in girls. Arch. Dis. Child. 44, 291–303 (1969).
Marshall, W. A. & Tanner, J. M. Variations in the pattern of pubertal changes in boys. Arch. Dis. Child. 45, 13–23 (1970).
Kleiner, D. E. et al. Design and validation of a histological scoring system for nonalcoholic fatty liver disease. Hepatology 41, 1313–1321 (2005).
American Diabetes Association. 2. Classification and Diagnosis of Diabetes: Standards of Medical Care in Diabetes-2021. Diabetes Care 44, S15–S33 (2021).
Expert Panel on Integrated Guidelines for Cardiovascular Health and Risk Reduction in Children and Adolescents & National Heart, Lung, and Blood Institute. Expert panel on integrated guidelines for cardiovascular health and risk reduction in children and adolescents: summary report. Pediatrics 128 S213–S256 (2011).
The R Project for Statistical Computing. https://www.r-project.org/.
Thévenot, E. A., Roux, A., Xu, Y., Ezan, E. & Junot, C. Analysis of the human adult urinary metabolome variations with age, body mass index, and gender by implementing a comprehensive workflow for univariate and OPLS statistical analyses. J. Proteome Res. 14, 3322–3335 (2015).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Kuhn, M. Building predictive models in R using the caret package. J. Stat. Softw. 28, 1–26 (2008).
Robin, X. et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinforma. 12, 1–8 (2011).
Tingley, D., Yamamoto, T., Hirose, K., Keele, L. & Imai, K. Mediation: R Package for Causal Mediation Analysis. J. Stat. Softw. 59, 1–38 (2014).
Stinson, S. E. et al. Identification of modifiable plasma protein markers of cardiometabolic risk in children and adolescents with obesity. GitHub repository, https://github.com/sarastinson/olink_holbaek; https://doi.org/10.5281/zenodo.17205894 (2025).
Acknowledgements
The authors thank all volunteers and their parents who participated in The HOLBAEK Study. The authors also thank colleagues from the Hansen Group at Novo Nordisk Foundation Center for Basic Metabolic Research and The Children’s Obesity Clinic for fruitful discussions. The authors thank A. Frost Bjerre, B. Holløse, O. Troest, T. Larsen, A. Forman, and T. Hvidtfeldt Lorentzen for their technical assistance. The authors thank L. Skovborg Just and L. Ryborg for managing the GALAXY and MicrobLiver consortia. The authors acknowledge the funding agencies that supported this study: the Innovation Fund Denmark (grant no. 0603-00484B), the Novo Nordisk Foundation (grant no. NNF15OC0016544 to T.H.), and the Novo Nordisk Foundation Challenge Program (grant no. NNF15OC0016692 to the MicrobLiver consortium (T.H. and A.K. in this study)). S.E.S. received funding from the Novo Nordisk Foundation Copenhagen Bioscience PhD Program (grant no. NNF18CC0033668) and DD2 Research Grant (grant no. NNF23OC0099658). R.T. is funded by the NNF Copenhagen Bioscience PhD Program (grant no. NNF0069781). M.A.V.L. was supported by the Danish Heart Foundation (grant no. 18-R125-A8447 and 21-R149-A10071). D.B. was funded by the NNF Copenhagen Bioscience PhD Program (grant no. NNF17CC0026760). L.A.H. was supported by the Danish Cardiovascular Academy, which is funded by the Novo Nordisk Foundation (grant no. NNF20SA0067242) and the Danish Heart Foundation (grant no. PhD2023009-HF). C.E.F. was supported by the BRIDGE-Translational Excellence Program (grant no. NNF18SA0034956), Steno Diabetes Center Sjælland, and the Region Zealand Health Scientific Research Foundation (grant no. R32-A1191). The European Union’s Horizon 2020 research and innovation program (grant no. 668031 to the GALAXY consortium (A.K., T.H., and M.T. in this study)). The authors also thank the Novo Nordisk Foundation for supporting the Novo Nordisk Foundation Center for Basic Metabolic Research (grant nos. NNF18CC0034900 and NNF23SA0084103).
Author information
Authors and Affiliations
Contributions
The manuscript was drafted by S.E.S., Y.H., R.T., E.S., and T.H. S.E.S. performed the bioinformatics analysis and generated figures for the manuscript. Y.H., R.T., and E.S. contributed to bioinformatics analysis and results interpretation. J.-C.H. and T.H. designed and coordinated the clinical cohort. M.A.V.L., C.E.F., and L.A.H. recruited participants in clinical cohorts, collected samples, and clinical data. D.B., L.Ä., and T.I.A.S. supported statistical analysis. M.A.V.L., H.B.J., M.T., A.K., O.P., and M.C. contributed to data coordination and management. J.-C.H. and T.H. developed the present project concept and protocol and supervised the project. All authors reviewed the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
H.B.J. is currently employed at Novo Nordisk Foundation. All other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Roya Kelishadi, Johannes Mueller-Reif, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Stinson, S.E., Huang, Y., Thielemann, R. et al. Identification of modifiable plasma protein markers of cardiometabolic risk in children and adolescents with obesity. Nat Commun 17, 1718 (2026). https://doi.org/10.1038/s41467-026-68415-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68415-2