Abstract
Sexual commitment in Plasmodium parasites is essential for malaria transmission, yet the signaling events initiating sexual conversion in only a subpopulation of parasites remain unclear. We discovered a single valine(V2163) to leucine(L2163) mutation in a transcription factor required for P. falciparum gametocytogenesis (AP2-G) that abrogates sexual differentiation. AP2-G.L2163 does not bind the ap2-g consensus motif, GnGTAC, or stimulate gene transcription, including autoregulation. The GDV1-dependent expression of AP2-G.L2163 demonstrates GDV1’s critical role in the initial activation of the silent ap2-g locus in the absence of functional AP2-G. While AP2-G.V2163 is required for MSRP1 expression, providing a marker that discriminates early from late sexually committed schizonts. Together this work demonstrates that V2163 in AP2-G plays a critical role in DNA binding, highlighting the functional importance of this specific region for malaria transmission as well as the key role of GDV1 in the initial activation of ap2-g expression and induction of sexual differentiation.
Similar content being viewed by others
Introduction
Malaria transmission is a global public health concern which accounted for 263 million clinical cases and more than half a million of deaths in 20231. A critical step in human-to-human transmission via mosquito is the production of sexual stage Plasmodium parasites, called gametocytes. Sexual differentiation begins in a subpopulation of blood stage parasites during the asexual replication cycle, but the percent of parasites that commit to sexual development varies in vitro and in vivo2,3. In contrast to relatively low in vitro gametocyte-commitment rates (GCR) (<30%)2 in P. falciparum laboratory adapted strains and clinical isolates, in vivo symptomatic or asymptomatic malaria studies show dramatic variability in GCRs (0-78%)3,4. The factors underlying this quantitative variability in gametocyte production remain undefined and make it difficult to target and eliminate the infectious reservoir. This GCR variation coupled, with the high prevalence of asymptomatic malaria infections, which can be as high as 60-80% in some regions5,6,7, and the inability of commonly used antimalarials to kill mature gametocytes8 are major challenges to malaria control programs. Defining the molecular mechanisms underlying the regulation of sexual commitment should facilitate the design of new transmission blocking strategies.
Two genes have been found to be required to initiate sexual commitment in P. falciparum, gametocyte development protein 1 (gdv1: PF3D7_0935400)9 and a plant-like apetala 2 transcription factor (ap2-g: PF3D7_1222600)10. GDV1 and AP2-G are required for ap2-g expression3,11, while AP2-G also activates downstream genes needed for the progression of sexually committed rings to continue gametocyte development10,12. In asexually committed blood stage parasites, the ap2-g locus is epigenetically silenced by histone 3 lysine 9 trimethylation (H3K9Me3)13 and this histone modification is stabilized by heterochromatin protein 1 (HP1: PF3D7_1220900)11,14. As the blood stage parasite matures, the start of DNA replication marks entry into the schizont stage of the life cycle. During schizogony, it has been proposed that in the subpopulation of schizonts that express GDV1, the ap2-g locus is de-repressed11, but the induction and regulation of this decision has not been established. There is evidence that once GDV1 is expressed, it interacts with HP-1 and this interaction has been proposed to contribute to ap2-g de-repression by removing HP111, but the detailed mechanism that confers specificity to the release of HP-1 only at the ap2-g locus and only in some parasites remains to be elucidated. Once expressed, AP2-G induces its own expression, termed autoregulation, by binding to the core AP2-G DNA recognition motif GnGTAC that is present in the 5’ flanking region of ap2-g10,12. This motif is also present upstream of a number of other genes, some of which are upregulated in the presence of AP2-G in schizonts and/or the next generation of blood stage parasites, called rings, initiating gametocyte differentiation. The reasons different AP2-G-dependent genes are expressed at different times after AP2-G expression remains unclear as does the regulation of initial expression of GDV1 and AP2-G.
In higher eukaryotes, de-repression of epigenetically silent genes has been described through post-translation modifications15. Phosphorylation of histone 3 (H3) serine 10 has been demonstrated to play a critical role in release of HP1 from lysine 9 trimethylated H316,17,18. This change allows RNA polymerase access to transcribe the previously silenced genes. In P. falciparum, several studies have consistently shown phosphorylation of H3 serine 29 and 3319,20,21 in trophozoite and schizont stages, which is when sexual commitment is also thought to be initiated. It is possible that phosphorylation of H3 serine 29 and/or 33 or both could play critical roles in releasing HP1 from ap2-g locus bound H3K9Me3.
We sought to evaluate whether serine/threonine kinase (STK2: PF3D7_0214600), which has a nuclear localization signal, could be one of the kinases playing a role in the phosphorylation of H320,22. We used reverse genetics to define the role of STK2 in gametocytogenesis but, unexpectedly, whole genome sequencing of the stk2 knockout parasites revealed a single valine2163 to leucine2163 mutation in ap2-g. We show that the replacement of valine with leucine in the presence and absence of stk2 abrogates AP2-G binding to the upstream core GnGTAC motif, autoregulation, downstream gene expression and gametocyte production. Here we have identified a highly robust system that can be utilized to study the transcriptional activation of ap2-g by an upstream regulator, GDV1, and monitor the activation of ap2-g specific targets.
Results
Determining the genetic locus underlying a complete loss of gametocytogenesis
In the course of evaluating kinases for a role in gametocytogenesis, we generated a serine/threonine kinase (STK2: PF3D7_0214600) knockout line (3D7.stk2∆) in a gametocyte-competent P. falciparum 3D7 strain (Supplementary Fig. 1a–c). Asexual growth was normal consistent with the previous piggyBac transposon insertion mutagenesis prediction that stk2 is dispensable during asexual growth23. In marked contrast, 3D7.stk2∆ clones did not produce immature or mature gametocytes (Fig. 1a, b, Supplementary Fig. 1d). To determine whether the block was at sexual commitment or maturation of sexually committed rings, we quantified gametocyte-committed ring stage parasites using the validated early gametocyte biomarker ap2-g4. Ap2-g expression levels were significantly reduced in 3D7.stk2∆ ring stage parasites (Fig. 1c), suggesting a block in sexual commitment similar to that observed in a previously reported gametocyte deficient line, F12 which has a premature stop codon in ap2-g at bp 392310, and a gametocyte inducible line NF54.gdv1.3xha.fkbp in the absence of Shield1 (Shld1-). In the gametocyte inducible line gdv1 is tagged with an in-frame ligand-dependent degradation domain (FKBP). In the absence of Shield 1 (Shld1) ligand the GDV1-FKBP chimeric protein is targeted for proteasomal degradation24 and therefore not available to activate ap2-g expression and sexual commitment. The presence of Shld1 blocks degradation of GDV1-FKBP and sexual commitment proceeds normally. To confirm the phenotype and define the cellular localization of STK2, we used the original double crossover plasmid to generate an inducible STK2 reporter line (NF54.stk2.gfp.fkbp) as well as a standard knock out (NF54.stk2∆) in a gametocyte competent Pf strain NF54. Surprisingly, the NF54.stk2∆ and NF54.stk2.gfp.fkbp lines with and without Shld1 all produced gametocytes and had GCRs similar to the parental line NF54 (Fig. 1d, e). These results strongly suggest that the loss of gametocytogenesis in 3D7.stk2∆ clones was independent of stk2.
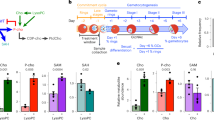
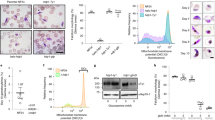
a Gametocyte conversion rate (GCR) of 3D7.stk2Δ clones (B4, F6) and the parental line (3D7). b Day 6 thin smear of n-acetylglucosamine treated cultures showing the presence or absence of gametocytes (arrows) in the parent 3D7 and 3D7.stk2Δ clone (B4), respectively. c Ap2-g expression fold changes in ring stage parasites from NF54, 3D7, 3D7.stk2Δ clones (B4, F6), ap2-g truncated line F12, and NF54.gdv1.3xha.fkbp in the absence (GDV1-) or presence (GDV1 + ) of Shld1. d GCR phenotype of the parent NF54 and STK2 inducible NF54.stk2.gfp.fkbp clones (A, B & C) in the presence (+) or absence (-) of Shld1. e GCR phenotype of the parent NF54 and NF54.stk2Δ clones (D, E & F). f Ap2-g mutation (g.6487 G > T) resulting in p.V2163L amino acid substitution was present in all the sequence reads from the 3D7.stk2Δ line (gray bars). The 3D7 reference sequence is shown at the bottom just above the corresponding amino acid sequence in all 3 reading frames. The arrow indicates the third reading frame encodes AP2-G. g Protein sequence alignment of the AP2-G AP2 domain derived from five diverse malaria parasite species, P. vivax (Pv), P. knowlesi (Pk), P. falciparum (Pf), P. berghei (Pb) and P. yoellii (Py), indicates conservation of the mutated valine (black box). h Schematic of AP2-G (aa 1 - 2432) with the AP2 DNA binding domain indicated by a green box and SNPs previously identified in the ap2-g locus in lab-adapted lines (red arrow), the present study (red dotted arrow) or field isolates (diamonds22). For graphs (a, c–e) independent data points are indicated by circles and the bar and error bar represent the mean and standard error of the mean, respectively. a, d, e experiments were performed in biological triplicates and repeated independently thrice whereas in (c) experiments were repeated independently 7 times for the parental 3D7, 8 times for clone B4, 7 times for clone F6, 4 times for F12 and 3 times for NF54 and GDV1 (+/-). A Mann-Whitney test (a, c) or Kruskal-Wallis test followed by Dunn’s multiple comparison (d, e) was used for statistical analysis between groups/conditions. The groups compared were parental wt vs stk2Δ clones (a), wt vs mutant ap2-g (c, d), intact (+) or knocked down stk2 (-) (d), or parental wt vs knocked out stk2 (e). Source data are provided as a Source Data file.
The gdv1 and ap2-g loci previously associated with gametocyte production9,10, remained intact in the 3D7.stk2∆ line and were free from the known ap2-g nonsense (F12) or frameshift (GNP-4) mutations10, or nonsense mutations (K561*, Q578*) in gdv125,26 (Supplementary Fig. 2). To extend our search, we sequenced the whole genome of the 3D7.stk2∆ line. Whole genome analysis confirmed the lack of deletions or mutations conferring premature termination of gdv1 or ap2-g. However, ap2-g had a missense mutation (g.6487 G > T) (Fig. 1f) that resulted in a switch from valine (V) to leucine (L) in the AP2 domain (V2163L). This V2163L mutation was the only missense SNP identified in 3D7.stk2∆ not present in wild type NF54 (Supplementary Table 1). The AP2-G AP2 domain is 50 aa long (AP2-G2162-2210)22,27 and highly conserved across the Plasmodium species infecting human, rodent and monkeys with all these species having a valine at the position corresponding to V2163 (Fig. 1g). Additionally, in the global P. falciparum isolate repositories analyzed to date, no mutations have been reported in the AP2-G AP2 domain (PlasmoDB22, MalariaGEN28) (Fig. 1h). These results suggest that the AP2-G AP2 domain is under strong selective pressure in the field as expected for a gene required for transmission. The 3D7.stk2∆ line was therefore designated as 3D7.stk2∆//ap2-g.L and wild type (wt) or mutant ap2-g designated ap2-g.V or ap2-g.L, respectively.
The AP2-G V2163L substitution is sufficient for a loss of gametocytogenesis
To confirm that ap2-g.V2163L prevents gametocytogenesis, we swapped the V allele with an L allele in a gametocyte competent P. falciparum strain, NF54, using a forward mutation editing approach (Fig. 2a), generating transgenic mutant lines (NF54.ap2-g.L and NF54.ap2-g.L.gfp.fkbp). While both lines have a V to L replacement, the NF54.ap2-g.L.gfp.fkbp line also has GFP.FKBP inserted in frame to regulate AP2-G.L. The whole genomes of both lines were sequenced and compared to the parental line, NF54, to rule out the presence of any unintended mutations. None of the lines expressing AP2-G.L produced immature or mature gametocytes (Fig. 2b, c), which is the same phenotype shown by original 3D7.stk2∆//ap2-g.L clones (Fig. 1a, b). To strengthen our hypothesis, we also tested whether reversing the L2163 mutation back to V can restore gametocytogenesis in the 3D7.stk2∆//ap2-g.L line, as well as the NF54.ap2-g.L.gfp.fkbp line (Fig. 2d). Correcting the mutation to V restored the rate of gametocytogenesis to that observed in the original parental lines (NF54 or 3D7) (Fig. 2e, f). This forward and reverse mutation editing approach firmly supports the valine to leucine mutation, not the stk2 knockout or another mutation, as the cause of the loss of gametocytogenesis in the 3D7.stk2∆//ap2-g.L line.
a Schematic of the allele swapping strategy showing the exchange of wild type (wt) ap2-g allele V with the mutant allele L in strain NF54. b Day 6 (D6) gametocyte conversion rate (GCR) analysis of wt and mutated NF54 parasites, NF54.ap2-g.L clones 32 L& 35 L were generated using CRISPR-Cas9 and NF54.ap2-g.L.gfp.fkbp: clones 17 L & 25 L were generated using homologous recombination without CRISPR-Cas9 and grown in the presence of Shld1. c D6 Giemsa-stained smear of wt NF54 or mutated NF54.ap2-g.L clone 32 L. d Schematic of allele swapping strategy for the reversion of mutant allele L into wild type V. e) Day 6 GCR analysis of 3D7 or NF54 derived lines with wt (V) or mutated (L) ap2-g, 3D7.stk2Δ//ap2-g.L clone B4L, 3D7.stk2Δ//ap2-g.V clones 9 V & 10 V and NF54.ap2-g.L.gfp.fkbp clone 17 L and NF54.ap2-g.V clone 11 V. f D6 Giemsa stained smear of 3D7, L mutated line (3D7.stk2Δ//ap2-g.L clone B4L), and mutation corrected line (3D7.stk2Δ//ap2-g.V clone 9 V). Gametocytes are indicated by an arrow. Blue, red, or black circles indicate mutated (Ap2-g.L), mutation corrected (Ap2-g.V), or parental with wt ap2-g lines, respectively. Each circle indicates an independent data point, and the bar and error bar represent the mean and standard error of the mean, respectively. Experiments were performed in three biological replicates and repeated 3 times independently. A Mann-Whitney test was used for statistical analysis between AP2-G mutant (L) or wt (V) allele parasite lines. Source data are provided as a Source Data file.
V2163L switch disrupts DNA binding
AP2 domain of AP2-G was described earlier22,27, and V2163 is located at the N-terminus of the domain (Fig. 1h). MOTIF search (https://www.genome.jp/tools/motif/) analysis also predicted the same amino acids as an AP2 domain (aa 2162 to 2210, i-Evalue = 0.00053) in which V2163 is the second aa (Fig. 3a). Changing V2163 to L to mimic the observed mutation shifted the start of the predicted AP2 domain downstream by 11 aa decreasing its length (Fig. 3a) and increasing the independent Expectation Value (i-Evalue = 0.0023) indicating reduced confidence in the motif prediction. Eleven aa shorter AP2 domains were also predicted when V2163 was substituted by any of the other aa (Fig. 3a, Supplementary Fig. 3), except isoleucine which was predicted to cause only a six aa shift (Fig. 3a). Consistent with the MOTIF search predicted AP2 domain, AlphaFold 3D structure modeling29,30 also predicted the structure of the same amino acids (aa 2162-2210) with a high confidence (Fig. 3b–d). MutationExplorer was then used to evaluate changes to the AP2-G structure following the leucine 2163 mutation. The analysis predicted a minimal overall structural destabilization (Total energy difference with parent = −25.5) with localized changes in stability within (Fig. 3e, f) and in close proximity to the AP2 domain (Supplementary Table 2), suggesting the possibility of a structural alteration in the mutated AP2 domain that could alter DNA binding. These computational predictions are consistent with V2163 being critical for DNA binding and functional activity31. To confirm the in-silico prediction, we generated recombinant proteins encoding the AP2 domain (AP2-G2150-2220, 60 aa) containing mutated (AP2-G.L) or wild type amino acid (AP2-G.V) as well as P. berghei wild type PbAP2-G. The recombinant proteins were purified from the soluble fraction of E. coli extracts and tested in an electrophoretic mobility shift assay (EMSA) for binding to a biotinylated oligonucleotide (GTGTACAC) that includes the core PfAP2-G DNA recognition motif GnGTAC10,12. In contrast to the wild type recombinant PfAP2-G domain (AP2-G.V) and PbAP2-G domain, the mutant recombinant protein (PfAP2-G.L) did not bind the GTGTACAC oligo (Fig. 3g). Both the in vitro and in silico results suggest that mutated AP2-G.L2163 would not bind to the core GnGTAC motif in genomic DNA and therefore no longer act as a functional transcription factor to activate gene expression. An alteration in DNA binding is also consistent with the inability of AP2-G.L to support ap2-g transcription and gametocyte production in the 3D7.stk2∆//ap2-g.L line (Fig. 1a–c).
a Motif search (https://www.genome.jp/tools/motif/) computational prediction of the amino acids (aa) contributing to the AP2 domain (red text) in AP2-G with the indicated amino acid at position 2163 (green box) and the independent expectation value (i-Evalue). b–f AlphaFold30,31 3D modeling predicts the structure of the domain (aa 2098-2245, green) that includes the AP2 domain (aa 2162-2210, purple) (c) with high confidence in the context of the full length wt AP2-G.V (d). The predicted local distance difference test (pLDDT) was used to evaluate confidence (b) and color code the structure, very high (dark blue >90%), high (cyan: 70-90%), low (yellow: 50-70%), very low (orange: 50%) (d, f). The dashed gray box indicates the amino acids in the recombinant AP2-G used in the electrophoretic mobility shift assay (EMSA) (f). e MutationExplorer58 analysis predicted local energy changes in full length AP2-G.L compared to wt AP2-G.V. The red bubble indicates the mutation sphere and residues that are stabilized or destabilized, respectively. f The AlphaFold prediction for recombinant mutant AP2-G.L (aa 2150-2220) used in the EMSA color coded based on pLDDT model confidence. The pink square indicates the mutant L2163 and the circles correspond to residues, I2154 and R2201 that are predicted to be destabilized using MutationExplorer. g EMSA analysis using a light shift EMSA assay kit with biotinylated probe GTGTACAC and recombinant proteins with either a wild type or mutated AP2 domain, P. falciparum recombinant AP2-G AP2 domain, aa 2150-2220 (Pf-WT or Pf-V2163L) or P. berghei wild type AP2-G AP2 domain (Pb-WT)63. Shifted and unshifted bands are indicated by respective arrow. The EMSA experiment was performed once at 2 different recombinant protein concentrations. The uncropped gel is included in the Source Data file.
GDV1 activates ap2-g in the absence of AP2-G or its autoregulation
Parasite lines expressing AP2-G.L provide tools to investigate the initiation of ap2-g expression in the absence of AP2-G autoregulation and interrogate the separate roles of GDV1 and AP2-G in this initiation process. Ap2-g expression is highly dynamic during schizogony, therefore first we did an RNA time-course using a gametocyte inducible line (NF54.gdv1.3xha.fkbp//ap2-g.V) expressing wild type AP2-G to identify the best time to compare expression between lines. As described previously, in the gametocyte inducible line GDV1-FKBP is stabilized by Shld1 ligand, and without Shld1 GDV1-FKBP gets degraded, as a result ap2-g is not expressed and there is no sexual commitment. Prior to the experiments, parasites were maintained in the absence of Shld1 to eliminate contamination with gametocyte-committed parasites produced during earlier asexual cycles that also express ap2-g and could interfere in the accurate determination of ap2-g expression initiation. After Shld1 addition to half the parasite cultures, ap2-g transcription was monitored every other hour starting from 28 hours post invasion (hpi) through 42 hpi in both Shld1 (+) and (-) conditions (Fig. 4a). This time course covers the trophozoite to schizont developmental transition. The ap2-g RNA time-course revealed that in the presence of Shld1-stabilized GDV1, ap2-g transcripts first increased over baseline (~two-fold) at 32 hpi and this increase was maintained until 36 hpi. After 36 hpi, ap2-g transcripts rose dramatically, until plateauing at 40 hpi. Six days later stage III-IV gametocytes were detected in the culture (Fig. 4b). In contrast, in the absence of Shld1 there was no increase in ap2-g transcript (Fig. 4a), and gametocytes were not produced (Fig. 4b).
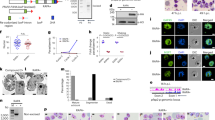
a A time-course using the NF54.gdv1.3xha.fkbp line in the absence and presence of Shld1 showing ap2-g transcript analysis every 2 hours from 28 to 42 h post invasion (hpi) and their day 6 gametocyte conversion rate (GCR) (b). Dashed and continuous lines indicate Shld1 (-) or (+), respectively. c Schizont stage ap2-g expression analysis from Shld1-regulated gdv1 lines with wt (ap2-g.V//gdv1.fkbp) or mutant (ap2-g.L//gdv1.fkbp) ap2-g and also wt (ap2-g.V.fkbp// wt gdv1) and mutant (ap2-g.L.fkbp//wt gdv1) ap2-g Shld1-regulated lines with wt gdv1. (+) or (-) indicate presence or absence of Shld1. d, e Ap2-g and msrp1 gene expression analysis in sorbitol synchronized ring stage parasites (6-10 hpi) in wt ap2-g parasite line (NF54.gdv1.3xha.fkbp) in the presence (GDV1(+)) or absence (GDV1(-)) of Shld1, ap2-g mutation corrected parasite line, 3D7.stk2Δ//ap2-g.V (AP2-G.V), ap2-g mutated parasite lines, NF54.ap2-g.L (AP2-G.L), 3D7.stk2Δ//ap2-g.L (B4L) and F12 with a premature stop codon in ap2-g. f Schematics showing ap2-g initial expression, autoregulation and AP2-G dependent downstream genes in the different engineered lines in the presence or absence of Shld1. Dotted circles indicate unknown potential co-factors, GTAC indicates the AP2-G binding DNA motif (GnGTAC), 1) depicts initial transcription, 2) depicts autoregulation that requires a functional AP2-G, and 3) depicts AP2-G dependent gene expression. Yellow circles indicate the FKBP domain that targets the attached protein for degradation in the absence of Shld1 ligand. Red or blue pentagons indicate AP2-G.V or AP2-G.L, respectively. a, b The time course experiment was performed once. a Error bars are derived from technical replicates. c–e Each circle indicates an independent data point; experiments were performed independently 2 to 6 times without replicates. c: Experiments were independently repeated six times for AP2-G.V//Gdv1.fkbp shld1 (+)/(-), three times for AP2-G.L//Gdv1.fkbp(+)/(-), AP2-G.L.fkbp//Gdv1.wt (+)/(-), and F12, and four times for AP2-G.V.fkbp//Gdv1.wt (+)/(-). d, e: Experiments were independently repeated three times for GDV1( + )/(-), F12, and AP2-G.L, four times for AP2-G.V, and two times for B4. The bar and error bar represent mean and standard error of the mean, respectively. An ANOVA test followed by Bonferroni’s multiple group comparison (c) was used to compare wt and mutant ap2-g expression in GDV1(+/-) conditions. Mann-Whitney test (d, e) was used for statistical analysis between wt and mutant ap2-g parasite lines as well as GDV1(-). Source data are provided as a Source Data file.
From the time course, 40-44 hpi was selected to compare ap2-g expression in lines with wt gdv1 and wt or mutant ap2-g tagged with fkbp (ap2-g.V.fkbp//wt gdv1, ap2-g.L.fkbp//wt gdv1), allowing Shld1-mediated regulation of AP2-G. In addition, fkbp-tagged gdv1 lines expressing non-tagged wt or mutant ap2-g (gdv1.3xha.fkbp//ap2-g.V or gdv1.3xha.fkbp//ap2-g.L) were included to compare the effect of the presence or absence of GDV1 on wt or mutant AP2-G expression and activity. Ap2-g transcription at schizont stage was completely disrupted by GDV1 knockdown in the absence of Shld1, regardless of whether the line expressed wt or mutated ap2-g (Fig. 4c). In contrast, in the fkbp-tagged ap2-g wt or mutant lines expressing wt GDV1, as well as a gametocyte-deficient line, F12, which expresses wt GDV1 but has a truncated AP2-G lacking the AP2 domain10, ap2-g was still expressed (~16 fold or more) in the presence or absence of Shld1. However, ap2-g expression was higher in ap2-g.V.fkbp line (Shld1 + ) than the ap2-g.L.fkbp line (Shld1 + ) or either line in the absence of Shld1. These results are consistent with wt GDV1 initiating ap2-g expression in all these lines, while only Shld1-stabilized wt AP2-G.V stimulated AP2-G mediated autoregulation. Ap2-g transcript levels were also higher in the presence of Shld1 in the gdv1.fkbp//ap2.g.V line than the gdv1.fkbp//ap2-g.L line (Fig. 4c) again indicating that during schizogony wt AP2-G enhances its own transcription.
In the next generation, only parasites expressing both AP2-G.V and GDV1 [3D7.stk2∆//ap2-g.V, NF54-gdv1.3xha.fkbp//Ap2-g.V (Shld1 + )] expressed all of the AP2-G dependent early gametocyte genes4 tested (Fig. 4d, e, Supplementary Fig. 4a–g) that contain at least one copy of the upstream AP2-G DNA recognition motif, GnGTAC. In contrast, none of these early gametocyte genes were expressed by parasite lines expressing AP2-G.L (3D7.stk2∆//ap2-g.L, NF54.ap2-g.L), truncated AP2-G (F12) or those lacking GDV1 [NF54.gdv1.3xha.fkbp//ap2-g.V (Shld1-)]. These results clearly demonstrate that after GDV1 initiates ap2-g expression AP2-G.V, not AP2-G.L, stimulates ap2-g autoregulation during schizogony and downstream gene expression in the progeny consistent with the need for DNA binding to mediate these effects (Fig. 4f).
Quantification of ap2-g.V & ap2-g.L and gdv1 translation
A nanoluciferase (nLuc) reporter was used to quantify and compare the time course of ap2-g.V, ap2-g.L, and gdv1 translation using a series of transgenic lines (3D7.stk2∆//ap2-g.V.fkbp.p2a.nLuc, 3D7.stk2∆//ap2-g.L.fkbp.p2a.nLuc and NF54.gdv1.p2a.nLuc). The P2A peptide that causes ribosomal skipping32,33 was included upstream of nLuc to allow the production of two separate proteins from a single RNA. We first confirmed a linear increase in nLuc signal with increasing parasite number (Supplementary Fig. 5). The assay was then used to monitor nLuc activity from 20 to 44 hpi and again 9 and 22 h later (53 hpi & 66 hpi) in the next generation rings. The time course data shows that signal from gdv1-linked nLuc was first detected at 26 hpi, which is prior to ap2-g-linked nLuc activity, as expected for an upstream regulator. Gdv1-linked nLuc activity increased rapidly until 34/35 hpi, then started to plateau (Fig. 5a). This early GDV1 activity is also supported through live imaging of a GDV1-tdTomato reporter line (683.gdv1.3xha.tdTom.fkbp) in which the GDV1-tdTomato fluorescent signal was first detected at 28-30 hpi which corresponds to the late trophozoite to early schizont transition prior to the first nuclear division (Fig. 5b). As schizogony progressed, GDV1 expression continued and a punctate pattern around each nucleus was observed (Supplementary Fig. 6). In contrast to gdv1-linked nLuc, ap2-g.V-linked nLuc activity was first detected at 31 hpi and continued to increase until 41 hpi after which activity plateaued, consistent with its transcription time-course (Fig. 4a). The time course of Ap2-g.L-linked nLuc signal was similar to ap2-g.V-linked nLuc activity during schizogony but levels were 2-3 fold lower in magnitude than ap2-g.V-linked nLuc activity, even throughout the plateau at 39-44 hpi (Fig. 5a). In the next generation ring stage parasites (53 & 66 hpi) ap2-g.V-linked nLuc activity remained high, which is in marked contrast with the ap2-g.L-linked nLuc line that had ~12 fold less nLuc activity and no gametocyte conversion (Fig. 5a, Supplementary Fig. 7). The increased ap2-g.V-linked nLuc activity in comparison to ap2-g.L-linked nLuc activity in schizonts (38-40 hpi) and next generation rings stage parasites (10-15 hpi) was confirmed using additional ap2-g.L (NF54.ap2-g.L.p2a.nLuc, 3D7.stk2∆//ap2-g.L.fkbp.p2a.nLuc) and ap2-g.V (NF54.ap2-g.V.p2a.nLuc, 3D7.stk2∆//ap2-g.V.fkbp.p2a.nLuc) parasite lines with NF54 or 3D7 backgrounds. Consistently there was a significant increase in nLuc activity in schizonts (3-4 fold: Fig. 5c) and rings (4-10 fold: Fig. 5d) stages in ap2-g.V over ap2-g.L parasite lines with the same parent as well as a significant increase in gametocyte conversion rate in the ap2-G.V lines (Fig. 5e). This pattern is consistent with an increase in AP2-G.V, not AP2-G.L, protein expression due to AP2-G.V-dependent autoregulation. The nLuc activity observed in ap2-g.L lines in schizont stage strongly supports wt GDV1 driving the initial ap2-g.V or ap2-g.L transcription, that is then augmented in the ap2-g.V lines due to autoregulation. The markedly lower levels of ap2-g.L-linked nLuc activity in the next generation rings are consistent with the decrease in GDV1-linked nLuc activity in rings (Fig. 5a) as well as the lack of functional AP2-G.V to maintain expression in the absence of GDV1. The findings also highlight the importance of AP2-G.V expression in gametocyte maturation.
a Time course analysis of ap2-g and gdv1-linked NanoLuc luciferase (nLuc) activity from early trophozoites (20 hpi) to the next generation of rings (53, 66 hpi). nLuc activity was monitored in ap2-g.V (3D7.stk2Δ//ap2-g.V.fkbp.p2a.nLuc), ap2-g.L (3D7.stk2Δ//ap2-g.L.fkbp.p2a.nLuc) and gdv1 (NF54.gdv1.p2a.nLuc) nLuc lines. b Live imaging of GDV1.tdTomato expression using the 683.gdv1.tdTomato.fkbp (Shld1 + ) line stained with Hoechst to localize the nucleus, which is representative of the 100 parasite positive red blood cells analyzed. Fold change in nLuc activity in 40-44 hpi schizonts (c) and in the next generation 5-10 hpi rings (d) and the corresponding gametocyte conversion rate (GCR) at day 4 (e) in ap2-g.L [NF5.ap2-g.L.p2a.nLuc (L1) and 3D7.stk2Δ//ap2-g.L.fkbp.p2a.nLuc (L2)] and in ap2-g.V [NF54.ap2-g.V.p2a.nLuc (V1) and 3D7.stk2Δ//ap2-g.V.fkbp.p2a.nLuc (V2, V3)] parasite lines. f Ap2-g linked nLuc signal from NF54.gdv1.3xha.fkbp//ap2-g.V.p2a.nLuc without Shld1 (GDV1-), NF54.ap2-g.L.p2a.nLuc (AP2-G.L), and NF54.ap2-g.V.p2a.nLuc (AP2-G.V) reporter lines. g GDV1 regulated ap2-g-linked nLuc activity and GCR quantification using the NF54.gdv1.3xha.fkbp//ap2-g.V.p2a.nLuc line. The X-axis indicates the time points when Shld1 was removed from the culture well and the dashed box indicates the developmental phase that requires GDV1 for gametocyte production. Assay details are provided in Supplementary Fig. 9b. c–f Red and blue circles indicate ap2-g.V and ap2-g.L, respectively. Each circle indicates an independent data point, and the bar and error bar represent mean and standard error of the mean, respectively. a, g Experiments were performed once with two biological replicates, c–f Experiments were repeated independently 4 to 8 times without replicates. c Experiments were repeated seven times for L1, L2, V2, and V3, and five times for V1, d experiments were repeated five times for L1 and V1, and seven times for L2, V2, and V3, e experiments were repeated five times for L1 and V1, eight times for L2, and seven times for V2 and V3, f experiments were repeated four times for each condition. Mann-Whitney test (c–e) or ANOVA followed by Turkey’s multiple comparison (f) was used for statistical analysis between groups with AP2-G.L or.V (c–e) or between GDV1(-), AP2-G.V or AP2.G.L parasite lines (f). Source data are provided as a Source Data file.
To directly assess the timing of the requirement for GDV1 in ap2-g transcription initiation, we generated NF54.gdv1.3xha.fkbp//ap2-g.V.p2a.nLuc transgenic reporter that allows monitoring of ap2-g activity by regulating GDV1 with Shld1. As expected in the absence of Shld1 there was no ap2-g.V-linked nLuc activity (Fig. 5f) and following the addition of Shld1 at ring stages (16hpi) ap2-g.V-linked nLuc activity began at ~33 hpi and started to plateau at 40 hpi (Supplementary Fig. 8a). To determine when GDV1 was no longer needed for ap2-g.V expression, Shld1 was added to tightly synchronized cultures in a 24 well plate at 16 hpi and then removed from replicate wells at defined time points from 30 hpi to 44 hpi (Supplementary Fig. 8b). Ap2-g.V- linked nLuc activity was assessed in all wells at 44 hpi (schizonts) and 56 hpi (rings). The GCR of each culture was also measured (Supplementary Fig. 8b). Early GDV1 removal (<40 hpi) resulted in lower ap2-g.V-linked nLuc levels in both schizonts (44 hpi: Fig. 5g) and the next generation rings (56 hpi: Supplementary Fig. 8c), as well as lower gametocyte production. Levels of ap2-g.V-linked nLuc activity gradually increased after 32-34 hpi reaching a plateau at ~40 hpi, after which time removing GDV1 had little effect (Fig. 5g). This controlled GDV1 exposure indicates it is required until ~40 hpi, for maximal ap2-g.V expression during the trophozoite-schizont transition.
Dynamics of AP2-G autoregulation and downstream gene induction
Ap2-g.mScarlet (mSc) reporter lines (3D7.stk2∆//ap2-g.L.mSc or 3D7.stk2∆//ap2-g.V.mSc) were generated to compare AP2-G.V and AP2-G.L protein expression patterns and identify sexually committed schizonts. Live imaging and flow cytometry show a subpopulation of schizonts (9-11%) positive for AP2-G.V.mSc which is similar to the 8-10% day 4 GCR (Fig. 6a, Supplementary Fig. 9a-b), suggesting the positive schizonts progressed to gametocytes. In contrast, none of the ap2-g.L.mSc lines were positive and there was no GCR (Fig. 6a, Supplementary Fig. 9a-b). It is possible that the fluorescence assays are not sensitive enough to detect the amount of AP2-G.L generated from the low transcript levels induced by GDV1 in the absence of AP2-G autoregulation or, alternatively, that the DNA binding activity of AP2-G.V is required for protein expression.
a Live imaging showing expression of AP2-G.mScarlet (mSc) in a schizont stage parasite from mutated (3D7.stk2Δ//ap2-g.L.mSc: bottom panel) or mutation corrected (3D7.stk2Δ//ap2-g.V.mSc: top panel) lines that were stained with Hoechst to detect DNA. b Live imaging of double reporter lines showing AP2-G and MSRP1 expression in the mutation corrected line (3D7.stk2Δ//ap2-g.V.mSc//msrp1. mNeonGreen (mNG): top panel), not the mutated line (3D7.stk2Δ//ap2-g.L.mSc//msrp1.mNG: bottom panel). Both parasite lines were stained with Hoechst to detect DNA. The images are representative of the 100 parasite positive red blood cells analyzed. c Quantification of AP2-G.mSc and MSRP1.mNG positive schizonts in the mutated (left panel) and mutation corrected (right panel) lines shown in (b). d Retro analysis of flow cytometer data shows AP2.G.mSc positive schizonts (top panel) have less Hoechst staining, consistent with younger schizonts with less DNA, whereas schizonts positive for both AP2.G.mSc and MSRP1.mNG (bottom panel) have higher Hoechst staining suggesting they are older. e Flow cytometric quantification of single positive (AP2-G.mSc + /MSRP1mNG-: dotted line) or double positive (AP2-G.mSc + /MSRP1.mNG + : continuous line) schizonts from 37 to 46 hpi using 3D7.stk2Δ//ap2-g.V.mSc//msrp1.mNG parasites. Details of flow cytometric quantification are provided as Supplementary Fig. 11. Source data are provided as a Source Data file.
To extend the expression profile of sexually committed schizonts beyond ap2-g, the RNA time course of msrp1 (ID# PF3D7_1335000) another gene expressed in sexually committed schizonts was tested. Msrp1 transcription increases over baseline only in the AP2-G.V line and the time course was slightly later than ap2-g RNA (Supplementary Fig. 10a), consistent with earlier reports that it is AP2-G regulated3,4,12. Protein expression was tracked by tagging MSRP1 with mNeon Green (mNG) in 3D7 and NF54 parental lines (3D7.stk2∆//ap2-g.L.mSc//msrp1.mNG, 3D7.stk2∆//ap2-g.V.mSc//msrp1.mNG, NF54.msrp1.mNG//ap2-g.V.mSc and NF54.msrp1.mNG//ap2-g.L.mSc). AP2-G.V.mSc and MSRP1.mNG were co-expressed in a subpopulation of schizonts, while no signal from either protein was detected in the AP2-G.L lines using fluorescence microscopy (Fig. 6a, b, Supplementary Fig. 10d (lower panels)). AP2-G.V.mSc expression was punctate and perinuclear (correlation coefficient with DNA = 0.56 ± 0.08). MSRP1 was present throughout cytoplasm and tended to be localized to the merozoite periphery in mature schizonts (46-48 hpi) (Fig. 6b (top panel), Supplementary Fig. 10b). Using flow cytometry AP2-G+MSRP1+ double positive schizonts were also only found in the AP2-G.V lines and they were in the same proportion as the D4 GCR (Fig. 6c, Supplementary Fig. 10e) and a similar correlation was found between MSRP1+ schizonts and D4 GCR in the NF54.msrp1.mNG line (Supplementary Fig. 10c) suggesting MSRP1 can be used as another marker of sexual commitment. In some experiments, low background fluorescence (<3×103 fluorescence intensity) was detected in the AP2-G.L lines using flow cytometry, but this variation is likely due to differences in the background fluorescence of different RBC preparations.
Analysis of MACS-purified schizonts from a dual reporter line (NF54.ap2-g.V.mSc//msrp1.mNG) indicated schizonts single positive for AP2-G.mSc, not MSRP1.mNG, had less DNA, suggesting they were younger than schizonts double positive for AP2-G.mSc and MSRP1.mNG (Fig. 6d). In these dual reporter lines there was also no MSRP1.mNG single positive schizonts, only double positive AP2-G.mSc / MSRP1.mNG schizonts, consistent with MSRP1 expression being dependent on AP2-G. The time course of AP2-G.V.mSc and MSRP1.mNG (NF54.ap2-g.V.mSc//msrp1.mNG) expression was further evaluated every hour from 28-46 hpi to understand the dynamics of sexual commitment (Fig. 6e, Supplementary Fig. 11). AP2-G.mSc single positive schizonts were first observed from 39-40 hpi, typically in 4-5 nuclear schizonts, and none of the schizonts with less than three nuclei were positive. Approximately 2 hrs later (41-42 hpi), these AP2-G positive schizonts turned positive for MSRP1.mNG. Again, no single positive MSRP1.mNG schizonts were detected (Supplementary Fig. 11). AP2-G and MSRP1 expression was much later than GDV1.tdTom, which was detected prior to nuclear division at 28-29hpi (Fig. 5b, Supplementary Fig. 6). These distinct MSRP1.mNG expression patterns allow differentiation of early and late sexually committed schizonts and demonstrate the distinct dynamics of AP2-G.V and downstream gene induction.
Discussion
A single valine to leucine mutation at the beginning of the AP2 DNA binding domain in AP2-G disrupted DNA binding and gametocyte production demonstrating the critical role of this AP2-G transcription factor for sexual differentiation. In contrast, we found the initiation of ap2-g expression during early schizogony requires GDV1, not functional AP2-G. Whereas, later in mature schizonts and the next generation ring stages AP2-G.V-dependent regulation of itself or another factor plays a major role in maintaining ap2-g expression and driving downstream gene expression. These regulatory differences allowed the development of stage specific reporters that differentiate the initial induction of ap2-g, early AP2-G autoregulation followed in 2 hours by the expression of the downstream gene, msrp1. The lines provide tools to further assess the underlying molecular mechanisms and screen for factors regulating malaria transmission.
Our detailed assessment of the time-course of sexual commitment demonstrates that gdv1 translation is first detected at 26 hpi and increases rapidly until 34/35 hpi when it begins to plateau as wt and mutant (L) ap2-g transcription and translation are initiated at 30-31 hpi. The early expression of GDV1 is consistent with its proposed role in de-repressing the ap2-g locus that has been epigenetically silenced by H3K9Me3 and HP1 binding upstream of the ap2-g promoter11,14. The role of GDV1 prior to AP2-G autoregulation is also supported by the almost complete block in ap2-g (V and L) transcripts when GDV1 is knocked down in the absence of Shld1. In contrast, ap2-g transcripts are still generated in GDV1 expressing lines in the absence of functional AP2-G, either due to knock down or changing AP2-G V to L. Defining this early period of ap2-g activation prior to the requirement for AP2-G.V expression can be used to focus further investigation of the mechanisms involved in de-repression, such as HP1 and histone 3 lysine 9 trimethylation removal and the participation of additional players to fully understand the initiation of sexual commitment in a subpopulation of parasites. AP2-I12,34, AP2-G535 and a recently identified AP2-P36 have been reported to bind upstream of ap2-g and could also contribute to its regulation.
After 38 hpi, the fluorescent detection of AP2-G.V coupled with the higher expression level of ap2-g in AP2-G.V expressing lines in comparison to lines lacking AP2-G, or those expressing AP2-G.L or truncated AP2-G suggests the expression increase requires functional AP2-G and is likely due to autoregulation. Additionally, during this phase (>38 hrs), knocking down GDV1 had little effect on ap2-g.V-linked nLuc expression or sexual conversion consistent with the requirement for wt AP2-G to activate sexual differentiation. AP2-G.V is also required for MSRP1.mNG expression in sexually committed schizonts and allows separation of AP2-G.V.mSc single positive parasites prior to MSRP1.mNG expression to evaluate early changes associated with AP2-G.V autoregulation. The effect of AP2-G.V autoregulation in maintaining ap2-g transcription is even more pronounced in the next cycle rings and early stage I gametocytes and suggests AP2-G.V is likely to be the primary activator of continued sexual differentiation and downstream gene transcription. In P. berghei, AP2-G is primarily detected in a subpopulation ring, not schizont, stages that progressed to gametocyte production, not asexual replication, consistent with the importance of AP2-G in early gametocyte formation37,38. In addition to demonstrating the importance of V2163 to AP2-G function and dissecting the roles of GDV1 and AP2-G.V in P. falciparum sexual commitment, these lines provide sensitive tools to directly quantify sexual commitment in schizont stages and allow small molecule library screens39,40 for compounds that block sexual differentiation. Additionally, the inability of AP2-G.L to stimulate AP2-G dependent gene expression allows the mutated lines to be used to focus the analysis on the regulation of the ap2-g locus without interference from AP2-G autoregulation or downstream gene expression. Both fluorescent and nLuc reporter lines can also be used to screen plasma/serum samples to identify human factors that regulate sexual conversion which vary dramatically in malaria patients3,4,26.
In contrast to the essential role of gdv1 and ap2-g in sexual differentiation, pfstk2 appears to be dispensable for asexual or sexual growth. This finding is consistent with previous piggyBac transposon-based insertion mutagenesis23 in P. falciparum monitoring asexual growth and the analysis of sexual development following knockout in P. berghei41. Interestingly, pbstk2 which is expressed in mosquito and liver stages, has been shown to play a critical role in the maturation of liver stage schizonts41. The pfstk2-KO or inducible transgenic lines expressing AP2-G.V generated in the course of our work could be used to investigate the role of pfstk2 in P. falciparum liver stage development.
The functional importance of a single valine located at the beginning of the predicted AP2 DNA recognition domain of AP2-G is clearly demonstrated by this work. Simply changing valine to leucine, a similar hydrophobic amino acid, blocked gametocyte production, the auto-upregulation of ap2-g transcription, translation and protein production as well as the up-regulation of downstream genes expressed in sexually committed schizonts or the next generation of ring stage parasites. The critical nature of this site in AP2-G is further supported by in silico analysis predicting an alteration in the AP2 DNA recognition site when V2163 is replaced by any other amino acid and more directly by the inability of recombinantly expressed AP2 domain of AP2-G. V2163L (aa 2150-2220) to bind an oligonucleotide containing the AP2-G binding motif, GnGTAC. We also found that the same V2163L mutation was present in some previously reported P. falciparum gametocyte non-producer piggyBac mutants, including clone IGM 2E442. Together, the data strongly suggests changing valine to leucine at position 2163 disrupts DNA binding, thereby blocking the ability of AP2-G.L to activate transcription of itself or downstream genes and abrogating gametocytogenesis. Further structural analysis of the mutant AP2-G protein would provide additional insights into the molecular basis of the loss of DNA-protein binding.
The lack of missense mutations among the thousands of clinical isolates sequenced to date (PlasmoDB22, MalariaGEN28) also supports the importance of V2163 in sexual differentiation, which is required for person-to-person transmission through the community. Conversely, in vitro culture depends only on asexual growth for propagation. Therefore, in vitro, a mutation that inhibits ~10% of schizonts from committing to gametocytes would increase asexual growth by ~10%, providing selective pressure for the mutant parasite in in vitro culture. This in vitro growth advantage of sterile parasite lines always needs to be considered when screening transgenic lines and highlights the importance of complementation and whole genome sequencing when analyzing phenotypes. There are multiple examples which suggest that gametocytogenesis phenotypes are selected for in vitro cultures, including deletion of parts of chromosome 9 containing gdv19 or mutations causing premature termination of GDV125,26 or AP2-G10. Interestingly, as opposed to all other known mutation/deletions conferring gametocyte deficiency through gdv1 or ap2-g, V2163L is the first mutation that alters AP2-G function without causing premature termination consistent with the key role played by V2163 in protein function. Outside the AP2 domain of ap2-g there are a large number of mutations in ap2-g in worldwide geographical isolates (PlasmoDB22, MalariaGEN28) that have not been functionally evaluated.
In Plasmodium, a conservative valine to leucine substitution with a strong phenotype, such as drug resistance, is rare but not unprecedented. A V to L mutation has been reported for P. falciparum cytochrome b where V259L is selected with atovaquone drug pressure in multiple studies43,44,45,46 and in P. vivax dhfr V64L is associated with antifolate resistance in clinical isolates47. A loss of gene function due to a single valine to leucine mutation has been also demonstrated in various organisms such as a V875L in cifb reduces cytoplasmic incompatibility in Wolbachia bacteria48, a V865L substitution in a human androgen receptor reduces sensitivity to androgen hormone49, and a V321L substitution in neuregulin1 in the hippocampus disrupts its nuclear localization, resulting in a complete loss of the function50,51.
In summary, the data strongly support the vital role of valine2163 in the AP2 domain of AP2-G for DNA binding, sexual commitment and early gametocytogenesis. In contrast, GDV1 is required for the initial activation of ap2-g transcription from 30-39 hpi and functional AP2-G during this phase is not essential. The definition of the time course and the development of lines with inducible GDV1 expression in the presence and absence of functional AP2-G allows the direct evaluation of the signaling pathways involved in ap2-g activation. AP2-G and MSRP1 reporter lines also clearly differentiate between early and late stages of sexual commitment during schizogony providing new tools to assess the effect of factors to modulate gametocyte conversion and transmission through the community.
Methods
Parasite culture
Plasmodium falciparum parasite strains (NF54, 3D7) or transgenic or mutant lines, were cultured in complete RPMI medium containing RPMI 1640, 25 mM HEPES, 100 μg/ml of hypoxanthine, and 0.3 mg/ml of glutamine (KD Biomedical, Columbia, MD) supplemented with 25 mM NaHCO3 (pH 7.3), 5 μg/ml of gentamicin, and 10% human serum (Interstate Blood Bank, Memphis, TN) or 0.5% Albumax II (Gibco, USA) in an atmosphere of N2:O2:CO2 90:5:5. Sorbitol treatment (5%, 10–30 min at 37 °C) or MACS® LS columns (Miltenyi Biotec, Auburn, CA) were used to synchronize parasites in ring or schizont stages, respectively. Parasitemia [(parasite-infected RBCs/total RBCs) *100] was evaluated through microscopy of culture smears stained with 10% Giemsa for 15 m (Sigma-Aldrich, St Louis, MO). Light microscope images were captured using QC Capture 2.9.13 software.
P. falciparum gametocyte phenotyping
The gametocyte conversion rate (GCR) of a single cycle was determined by quantifying parasitemia of a synchronized ring stage culture (Day 0) then adding N-acetylglucosamine (50 mM) (NAG) to block further asexual growth. The plate/flask was then cultured for five or seven days (D4 or D6) with daily media changes without RBC supplementation and the D4 or D6 gametocytemia (stage II-III or III-IV) determined (Supplementary Fig. 1d). Parasites were quantified using Giemsa stained culture smears or flow cytometry, which is described in the next section. The GCR of each sample was calculated using the formula (D4 or D6 gametocytemia / D0 ring parasitemia) x 100.
Flow cytometry and data analysis
Parasite cultures and uninfected RBCs (negative control) were resuspended at 0.02% hematocrit (~2,000 cell/µl) in 1x SA buffer (154 mM NaCl, 9.27 mM glucose, 10 mM Tris-HCl, pH 7.4) with 500 nM MitoProbe DiIC1(5) (Cat: M34151) and incubated at 37 °C in the dark for 30 min prior to analysis on an Accuri C6 flow cytometer (BD Biosciences, Franklin Lakes, NJ) or Attune Nxt flow cytometer (Invitrogen, USA). The forward and side scatter signals were used to select the total RBC population and then the DiIC1(5) (Ex 640 nm/Em 675 ± 25 nm) was used to identify Pf infected RBC.
A Cytek Aurora Spectral Analyzer flow cytometer (USUHS, Biomedical Instrumentation Center facility) was used to identify parasite infected RBC positive for mScarlet (mSc) or mNeonGreen (mNG) or both fluorescent protein signals. Hoechst 33342 DNA stain (Invitrogen, USA, Cat: H3570), 1:500 dilution, 10 minutes staining) was used to quantitate and identify the stage of infected RBCs (iRBCs). The proportion of mSc and or mNG positive iRBCs observed in total or stage-specific iRBCs was determined using FlowJo v10.10.0. A retro gating strategy was used to visualize and quantify only AP2-G.mSc positive iRBCs or MSRP1.mNG positive iRBCs from MACS-column purified parasites having a range of younger to mature schizonts based on the Hoechst 33342 signal. BD Accuri C6 Software, Attune software v5.3.0, and SpectralFlo v3.03 were used to analyze Accuri, Attune Nxt flow, and Cytek Aurora flow cytometers, respectively.
Plasmid engineering (CRISPR & non-CRISPR) and transgenic line generation
The stk2 knockout lines (3D7.stk2∆.wr or NF54stk2∆.wr) were generated using a double crossover plasmid (pCC1) in the absence of Cas9. To generate this plasmid, two homologous fragments were introduced in the plasmid using SacII + SpeI and NcoI+AvrII restriction enzymes. T4-DNA ligase was used to ligate the restriction enzyme-digested PCR products into the plasmid. The inducible stk2 line was generated using a single crossover plasmid (p1605.stk2.gfp.fkbp.wr) (Supplementary Fig. 12a) to tag the C-terminus of stk2 with green fluorescent protein (GFP) followed by the FKBP-destabilization domain (FKBP)3. This single crossover plasmid was also used to generate an ap2-g mutated line (NF54.ap2-g.L.gfp.fkbp.wr). To generate the single crossover plasmid construct, a single homologous region (600–1500 bp) of the target gene was PCR amplified, digested with XhoI + AvrII restriction enzyme and ligated into XhoI + AvrII digested plasmid using T4-DNA ligase. Primer DNA sequences for stk2 and ap2-g genes are given in Supplementary Table 3.
E. coli electrocompetent cells (DH10B) were transformed with the ligation reaction and grown overnight on ampicillin (100 µg/ml) containing LB plates. Ampicillin-resistant insert positive colonies were assessed using restriction enzyme digestion or by PCR and then further verified by DNA sequencing using Geneious prime® 2025.2.2 software. DNA sequence-confirmed colonies were expanded, and plasmid was purified using the NucleoBond Xtra Plasmid Maxi prep kit (Macherey Nagel, USA). Purified plasmid reconstituted in 1x cytomix buffer containing KCl (120 mM), CaCl2 (0.15 mM), K2HPO4/KH2PO4 (10 mM, pH 7.6), HEPES (25 mM, pH 7.6), EGTA (2 mM, pH 7.6), and MgCl2 (5 mM) was stored at -20 °C. 100–150 µg plasmid DNA per transformation was used to transform parasites using established transformation protocols3. Approximately 5% rings were transformed (Day 0) and on D2 appropriate drug was added for selection: WR99210 (2.5 nM) (Jacobus Pharmaceuticals, Plainsboro, NJ) or Blasticidin (2.5 nM) (Gibco, USA). Resistant parasites from three independent transformations were selected and grown for 3 weeks without drug before reapplying the appropriate drug. The resistant parasites obtained were screened for the presence of single or double-crossover chromosomal integration (Supplementary Fig. 1b and 12b) using PCR with primers listed in Supplementary Table 3 and selected for cloning by limiting dilution. Transformed lines were maintained in the appropriate drug/s.
CRISPR/Cas9 gene editing was used to generate the majority of the gdv1, ap2-g and msrp1 transgenic lines. Single plasmid (pDC-cam-cas9-gfp.wr)52 or two-plasmid based CRISPR/Cas9 gene editing was employed to transform gametocyte competent laboratory strains (3D7, NF54) or gametocyte deficient precursor transgenic lines (3D7.stk2∆-WR, NF54.ap2-g-L-gfp-fkbp-WR). pDC-cam-cas9.gfp.wr has a repair template (flanked by EcoRI+AatII), a guide RNA sequence and Cas9 expression cassettes52 and this plasmid is also called a single-guide plasmid (SGP). We used this SGP plasmid to replace ap2-g.V2163 with ap2-g.L2163 (pDC-cam.ap2-g.L2163.wr) or vice versa ap2-g.L2163 with ap2-g.V2163 (pDC-cam.ap2-g.V2163.bsd), and also to tag the C-terminus of gdv1 with 3xha.fkbp (pDC-cam.cas9.gdv1.3xha.fkbp-WR) or p2a.NanoLuc (pDC-cam.cas9.gdv1.p2a.NanoLuc-BSD) or tdTomato.fkbp (pDC-cam.cas9.gdv1.tdTomato.fkbp-BSD), and the msrp1 with mNeonGreen (pDC-cam.cas9.msrp1.mNeonGreen-DSM) (Supplementary Fig. 12c-d). To tag the c-terminus of ap2-g with a number of different reporters, we modified the SGP and generated a two-guide plasmid (TGP) by replacing Cas9 with a 2nd gSeq cassette using infusion cloning (Supplementary Fig. 12d-e). Using this TGP, we tagged the c-terminus of ap2-g with fkbp (pDC-cam.ap2-g(L or V).fkbp), mScarlet (pDC-cam.ap2-g(L or V).mSc), fkbp.p2a.NanoLuc (pDC-cam.ap2-g (L or V).fkbp.p2a.NanoLuc) or p2a.NanoLuc (pDC-cam.ap2-g(L or V).p2a.NanoLuc). The WR drug resistance marker was replaced with either BSD or DSM1 depending on the requirement. When TGP was used for the transformation, if the parasite line used for transformation lacked Cas9 expression a second plasmid pUF-DSM53 was used as the Cas9 source.
Guide RNA sequence (20 nt) with NGG PAM was designed using e-CRISP web-based tools54 that provide only a few but very efficient guide sequences per gene. The gRNA seq was digested with BbsI, a type IIS restriction enzyme, ligated into the plasmid using T4-DNA ligase and Stellar competent cells were used for the transfection. All other modifications (single or multiple donor fragments, drug resistance markers or the second gRNA cassette) were done using infusion cloning. To use this system, PCR fragments generated with infusion cloning compatible primers and then infused with the appropriate restriction enzymes digested plasmid. Insert positive colonies were identified and the NucleoBond Xtra Plasmid Maxi kit was used to purify plasmid for transformation as described above in the non-CRISPR plasmid section. The DNA sequences of the InFusion PCR primers and gRNAs used are listed in Supplementary Table 3. Fresh, uninfected RBCs were transformed using 100–150 ug of each plasmid, washed with RPMI, mixed with 0.2–1% MACS purified schizont stage parasites and incubated in triplicates in a 12 well culture plate. On D3 (ring stage) appropriate drug/s were applied for three generations/cycles (~ 6 days) to select transformed parasites and then removed. Parasites recovered after drug/s selection were tested for editing using PCR (Supplementary Fig. 12c) and confirmed by DNA sequencing. Transformed parasites were cloned after genotype verification using a limiting dilution method and clones were further tested by DNA sequencing.
Quantitative reverse transcriptase-polymerase chain reaction (RT-qPCR)
RNA was extracted from parasites preserved in 1x RNA Shld1 (Zymo Research, USA) using RNA Micro kit (Zymo Research, USA), as per the manufacturer’s instructions. RNA was eluted in 36 µl RNAse-free water and purified RNA (4.0 µl or 8.0 µl) was additionally treated with ezDNAse™ before cDNA synthesis using SuperScript™ IV VILO™ Master Mix (Invitrogen, USA). A 1:20 dilution of cDNA was made and 2 µl from this dilution was used for each RT-qPCR well. RT-qPCR (QuantStudio7 Pro) was performed using gene-specific primers designed using the web-based tool Primer3Plus v3.3.055 (Supplementary Table 3) and Fast SYBR Green PCR Master Mix (Applied Biosystems, USA) as described earlier3. Primer efficiency was tested by serial dilution and ranged from 85 to 105%. All samples were run in duplicate and tested for both the gene of interest and the control constitutive gene, arginyl tRNA (schizont stage) or skeleton binding protein 1 (sbp1) (ring stage) on the same plate. Data was analyzed using QuantStudio™ Design & Analysis Software v2.6.0, and the ΔCt values were determined by subtracting the mean cycle threshold (Ct) value for arginyl tRNA or sbp1 from the mean Ct of each test gene. The relative quantity (2-∆∆Ct) was calculated for each gene using RNA from NF54-gdv1-3xha-fkbp parasites grown in the absence of Shld1 or gametocyte deficient lines (3D7.stk2∆ or NF54-ap2-g-L).
Whole genome sequencing
Whole Genome Sequencing was conducted using a short-read workflow. Genomic DNA was used as input for library preparation after mechanical shearing and subsequent end-repair, a-tailing and ligation with unique dual index adapters. Sequencing libraries were assessed for size distribution and absence of adapter dimers before sequencing on an Illumina NovaSeq 6000 using a 151 + 8 + 8 + 151 bp read parameter for run conditions. An average 26,329,252 sequences per samples were generated and subsequently mapped using the Burrows-Wheeler Alignment56 against P. falciparum 3D7 reference genome (PlasmoDB)22. The average percentage of mapped reads per isolate was 78% and an average 74x coverage was achieved across the genome. The Genome Analysis Toolkit57 was then used to identify single nucleotide polymorphism (SNP) variants per isolate (Supplementary Table 1). Whole genome DNA Sequencing data generated in this study have been deposited in the BioProject/SRA under the accession codes PRJNA1260958: SRR33482688, SRR33482687, SRR33482686, SRR33482685, SRR33482684, SRR33482683, SRR33482682.
In-silico prediction of AP2 domain and structural visualization of AP2-G-DNA interaction
MOTIF search (https://www.genome.jp/tools/motif/), a web-based tool was used to predict the AP2 domain in-silico. This web tool has definitions of known protein domains, including AP2 domains derived from plants to higher eukaryotes. The independent Expectation Value (i-Evalue) represents the number of motifs with a similar statistical score that would be expected to occur by chance in a random dataset of the same size. Lower i-Evalue indicates that the motif is highly unlikely to have occurred randomly, suggesting it is a biologically significant pattern. This tool predicts a domain based on amino acid sequence similarity. We provided wild type or mutated amino acid sequences of AP2-G as an input and for each sequence an AP2 domain was predicted with a significance score. To visualize the 3D structure, an AlphaFold29,30 predicted AP2-G structure was used (AF-Q8I5I9-F1-model_v4.pdb), which was generated using full length AP2-G. Domains were identified using the predicted local distance difference test (pLDDT) model confidence score (pLDDT=70 or higher). For modeling the impact of AP2-G.L2163 mutation on the 3D structure, the MutationExplorer58 webtool was used. To do this, the predicted 3D structure (AF-Q8I5I9-F1-model_v4.pdb) was loaded to MutationExplorer and L2163 mutation was introduced. This tool compares the global and individual residue specific energy changes from wild type AP2-G. The structure stabilizing (positive) and de-stabilizing (negative) energies were recorded with a threshold of >0.5.
Electrophoretic mobility shift assay
Plasmodium falciparum (Pf-AP2-G-L2163, PfAP2-G.V2163wt) and P. berghei (PbAP2-Gwt) recombinant proteins were prepared as described earlier59. These recombinant proteins were expressed as glutathione S-transferase (GST: pGEX-4T1, GE Life Sciences) fusion proteins and purified from the soluble fraction of sonicated bacterial material. Electrophoretic mobility shift assay (EMSA) was performed using Light Shift EMSA kits (Thermo Scientific)10 using 2 μg of protein and 20 fmol of biotinylated probe59. Briefly, probes and recombinant DNA-binding domains were incubated at RT for 20 min before loading on a 0.5x TBE-buffered 6% polyacrylamide gel. Complexes were separated at 100 V until the dye front had migrated 3/4 of the gel length. Probes were then transferred overnight onto Nylon membranes, immobilized by UV (302 nm) cross-linking for 15 min, and visualized using strepdavidin-HRP conjugate and luminol detection solution (BioRad).
Parasite live imaging
MACS-column purified schizont stage parasites were used for visualizing mScarlet or mNeonGreen or tdTomato fluorescent protein. Live parasites re-suspended in 1xSA were stained with Hoechst 33342 DNA stain (1:500 dilution, 10 m) and washed once in 1xSA before imaging. Live imaging was performed using Zeiss inverted fluorescence microscope (Axio Observer 7) under 630x magnification and ZEN v3.0 (blue edition) in-built analysis software. For co-localization, Z-Stack images were deconvolved using ZEN v3.0 in built software and the weighted correlation coefficient was used for the DAPI (DNA) and mSc (AP2-G) signals as well as the mSc (AP2-G) and mNG (MSRP1) signals. A total of five schizont’s Z-Stack images were analyzed, and ~15 layers of images through the center of each schizont were included in correlation coefficient analysis. The mean and standard error of the correlation coefficients for each schizont was used to determine colocalization.
Protein quantification using nLuc system
Nanoluciferase (nLuc) bioluminescence signal was measured using a Nano-Glo® Luciferase Assay System (Cat# N1110, Promega, USA) and the Cytation 5 Biotek imaging system as per manufacturer’s instruction. Gen 5 v3.08 software was used to quantitate bioluminescence signals. Tightly synchronized parasite cultures (Sorbitol-MACS-sorbitol) were used to measure nLuc signals at defined time points or developmental stages (schizonts or rings). Nano-substrate was used in 1:50 dilution in the assay buffer (30 µl) with an equal volume of culture (30 µl) making final dilution of 1:100. The number of parasites in each comparison group was quantified using flow cytometry or hemocytometer and used to normalize the bioluminescence signal. In the time-course experiment, a background signal subtraction was applied. Mean nLuc signal obtained from 16-22 hpi was subtracted from each data-point measured later from 24 to -44 hpi. In generation 2, the ring stage nLuc signal was compared between samples containing the same number of parasites without a background subtraction. For the fold change analysis (Fig. 5c, d), nLuc signal was presented relative to nLuc signal observed in ap2-g.L2163 line.
Data analysis
GraphPad Prism 10.6.1 statistical package was used to evaluate differences between parasite lines. All comparisons between wt and stk2∆ clones, Shld1 (+) and Shld1 1 (-) or ap2-g-L2163 and ap2-g-V2163 groups were analyzed using a Kruskal-Wallis test followed by Dunn’s multiple comparison. ANOVA test followed by Bonferroni’s or Turkey’s multiple comparison was performed after evaluating the normal distribution of data using a Quantile-Quantile plot. Two groups were compared using Mann-Whitney test. All statistical tests were two-sided and the p value of each comparison is indicated. Correlations between groups or conditions were assessed using Microsoft excel (Microsoft Excel 2013) to determine the Pearson correlation coefficient.
Plasmids and parasite lines source, and authentication
P. falciparum wt (Nf54, 3D7) and mutant F12 strains were obtained from B.E.I Resources (USA) and Prof Manuel Llinas’s lab (Pennsylvania State University, USA), respectively. Plasmid with tdTomato (pGEMT-PT2A-iRFP670-Tdtomato-GFP: Addgene plasmid # 111817)33 or mScarlet (pmScarlet_C1: Addgene plasmid # 85042)60 or mNeonGreen (ER-mNeonGreen: Addgene plasmid # 137804)61 or NanoLuciferease (pUAS-NanoLuc: Addgene plasmid # 87696)62 were obtained from Addgene Inc (USA).
Genomic DNA from the parasite lines was analyzed for the presence or absence of the appropriate chromosomal insertions/modifications/alleles using targeted and whole genome sequencing. The parasite lines were reassessed periodically throughout the study for the genotype stability using targeted sequencing of PCR amplicons spanning the modified region of the genome. Wild type lines (NF54 and 3D7) were also tested to confirm that they continued to produce gametocytes. Giemsa-stained culture smears of P. falciparum infected and uninfected red blood cells were assessed for normal morphology by light microscopy. The following lines were analyzed by whole genome sequencing - Wild type parasite (NF54), gametocyte deficient line 3D7.stk2∆, ap2-g mutation introduced (NF54.ap2-g.L2163 and NF54-Ap2g-L2163-gfp.fkbp) or mutation corrected (3D7.stk2∆.ap2-g.v2163 and NF54.ap2-g.V2163) transgenic lines.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated during this study are included in this published article or the supplementary information file, source data file, or public repositories for whole genome sequences. Source data are provided with this paper. Whole genome DNA Sequencing data generated in this study have been deposited in the NCBI BioProject/SRA (https://www.ncbi.nlm.nih.gov/bioproject/?term=PRJNA1260958) under the accession codes PRJNA1260958: SRR33482688, SRR33482687, SRR33482686, SRR33482685, SRR33482684, SRR33482683, SRR33482682. Source data are provided with this paper.
References
World Malaria Report 2024. World Health Organization (2024).
Stewart, L. B. et al. Plasmodium falciparum sexual commitment rate variation among clinical isolates and diverse laboratory-adapted lines. Microbiol. Spectr. 10, e0223422 (2022).
Usui, M. et al. Plasmodium falciparum sexual differentiation in malaria patients is associated with host factors and GDV1-dependent genes. Nat. Commun. 10, 2140 (2019).
Prajapati, S. K. et al. The transcriptome of circulating sexually committed Plasmodium falciparum ring stage parasites forecasts malaria transmission potential. Nat. Commun. 11, 6159 (2020).
Hofmann, N. et al. Ultra-sensitive detection of Plasmodium falciparum by amplification of multi-copy subtelomeric targets. PLoS Med. 12, e1001788 (2015).
Geiger, C. et al. Declining malaria parasite prevalence and trends of asymptomatic parasitaemia in a seasonal transmission setting in North-Western Burkina Faso between 2000 and 2009-2012. Malar. J. 12, 27 (2013).
Adjah, J., Fiadzoe, B., Ayanful-Torgby, R. & Amoah, L. E. Seasonal variations in Plasmodium falciparum genetic diversity and multiplicity of infection in asymptomatic children living in southern Ghana. BMC Infect. Dis. 18, 432 (2018).
Ippolito, M. M. et al. The relative effects of artemether-lumefantrine and non-artemisinin antimalarials on gametocyte carriage and transmission of Plasmodium falciparum: a systematic review and meta-analysis. Clin. Infect. Dis. 65, 486–494 (2017).
Eksi, S. et al. Plasmodium falciparum gametocyte development 1 (Pfgdv1) and gametocytogenesis early gene identification and commitment to sexual development. PLoS Pathog. 8, e1002964 (2012).
Kafsack, B. F. et al. A transcriptional switch underlies commitment to sexual development in malaria parasites. Nature 507, 248–252 (2014).
Filarsky, M. et al. GDV1 induces sexual commitment of malaria parasites by antagonizing HP1-dependent gene silencing. Science 359, 1259–1263 (2018).
Josling, G. A. et al. Dissecting the role of PfAP2-G in malaria gametocytogenesis. Nat. Commun. 11, 1503 (2020).
Salcedo-Amaya, A. M. et al. Dynamic histone H3 epigenome marking during the intraerythrocytic cycle of Plasmodium falciparum. Proc. Natl. Acad. Sci. USA 106, 9655–9660 (2009).
Flueck, C. et al. Plasmodium falciparum heterochromatin protein 1 marks genomic loci linked to phenotypic variation of exported virulence factors. PLoS Pathog. 5, e1000569 (2009).
Sales-Gil R., Vagnarelli P. How HP1 post-translational modifications regulate heterochromatin formation and maintenance. Cells 9, (2020).
Papamokos, G. V. & Kaxiras, E. How to evict HP1 from H3: using a complex salt bridge. Biophys. Chem. 300, 107062 (2023).
Hirota, T., Lipp, J. J., Toh, B. H. & Peters, J. M. Histone H3 serine 10 phosphorylation by Aurora B causes HP1 dissociation from heterochromatin. Nature 438, 1176–1180 (2005).
Fischle, W. et al. Regulation of HP1-chromatin binding by histone H3 methylation and phosphorylation. Nature 438, 1116–1122 (2005).
Treeck, M., Sanders, J. L., Elias, J. E. & Boothroyd, J. C. The phosphoproteomes of Plasmodium falciparum and Toxoplasma gondii reveal unusual adaptations within and beyond the parasites’ boundaries. Cell Host Microbe 10, 410–419 (2011).
Ganter, M. et al. Plasmodium falciparum CRK4 directs continuous rounds of DNA replication during schizogony. Nat. Microbiol. 2, 17017 (2017).
Pease, B. N. et al. Global analysis of protein expression and phosphorylation of three stages of Plasmodium falciparum intraerythrocytic development. J. Proteome Res 12, 4028–4045 (2013).
Amos, B. et al. VEuPathDB: the eukaryotic pathogen, vector and host bioinformatics resource center. Nucleic Acids Res. 50, D898–D911 (2022).
Zhang, M. et al. Uncovering the essential genes of the human malaria parasite Plasmodium falciparum by saturation mutagenesis. Science 360, (2018).
Armstrong, C. M. & Goldberg, D. E. An FKBP destabilization domain modulates protein levels in Plasmodium falciparum. Nat. Methods 4, 1007–1009 (2007).
Tiburcio, M. et al. A 39-amino-acid C-terminal truncation of GDV1 disrupts sexual commitment in Plasmodium falciparum. mSphere 6, (2021).
Llora-Batlle, O. et al. Conditional expression of PfAP2-G for controlled massive sexual conversion in Plasmodium falciparum. Sci. Adv. 6, eaaz5057 (2020).
Balaji, S., Babu, M. M., Iyer, L. M. & Aravind, L. Discovery of the principal specific transcription factors of Apicomplexa and their implication for the evolution of the AP2-integrase DNA binding domains. Nucleic Acids Res. 33, 3994–4006 (2005).
MalariaGen et al. Pf7: an open dataset of Plasmodium falciparum genome variation in 20,000 worldwide samples. Wellcome Open Res. 8, 22 (2023).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Varadi, M. et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444 (2022).
Lindner, S. E., De Silva, E. K., Keck, J. L. & Llinas, M. Structural determinants of DNA binding by a P. falciparum ApiAP2 transcriptional regulator. J. Mol. Biol. 395, 558–567 (2010).
Donnelly, M. L. L. et al. The ‘cleavage’ activities of foot-and-mouth disease virus 2A site-directed mutants and naturally occurring ‘2A-like’ sequences. J. Gen. Virol. 82, 1027–1041 (2001).
Liu, Z. et al. Systematic comparison of 2A peptides for cloning multi-genes in a polycistronic vector. Sci. Rep. 7, 2193 (2017).
Santos, J. M. et al. Red blood cell invasion by the malaria parasite is coordinated by the PfAP2-I transcription factor. Cell Host Microbe 21, 731–741 e710 (2017).
Shang, X. et al. A cascade of transcriptional repression determines sexual commitment and development in Plasmodium falciparum. Nucleic Acids Res. 49, 9264–9279 (2021).
Subudhi, A. K. et al. DNA-binding protein PfAP2-P regulates parasite pathogenesis during malaria parasite blood stages. Nat. Microbiol. 8, 2154–2169 (2023).
Yuda, M., Kaneko, I., Murata, Y., Iwanaga, S. & Nishi, T. Mechanisms of triggering malaria gametocytogenesis by AP2-G. Parasitol. Int. 84, 102403 (2021).
Kent, R. S. et al. Inducible developmental reprogramming redefines commitment to sexual development in the malaria parasite Plasmodium berghei. Nat. Microbiol. 3, 1206–1213 (2018).
Kato, N. et al. Diversity-oriented synthesis yields novel multistage antimalarial inhibitors. Nature 538, 344–349 (2016).
Baniecki, M. L., Wirth, D. F. & Clardy, J. High-throughput Plasmodium falciparum growth assay for malaria drug discovery. Antimicrob. Agents Chemother. 51, 716–723 (2007).
Jillapalli, R. et al. A Plasmodium berghei putative serine-threonine kinase 2 (PBANKA_0311400) is required for late liver stage development and timely initiation of blood stage infection. Biol Open 8, (2019).
Ikadai, H. et al. Transposon mutagenesis identifies genes essential for Plasmodium falciparum gametocytogenesis. Proc. Natl. Acad. Sci. USA 110, E1676–E1684 (2013).
Nguyen, W. et al. 7-N-substituted-3-oxadiazole quinolones with potent antimalarial activity target the cytochrome bc(1) complex. ACS Infect. Dis. 9, 668–691 (2023).
Antonova-Koch, Y. et al. Open-source discovery of chemical leads for next-generation chemoprotective antimalarials. Science 362, (2018).
Hayward, J. A. et al. A screen of drug-like molecules identifies chemically diverse electron transport chain inhibitors in apicomplexan parasites. PLoS Pathog. 19, e1011517 (2023).
Calit, J. et al. Pyrimidine Azepine Targets the Plasmodium bc (1) Complex and Displays Multistage Antimalarial Activity. JACS Au 4, 3942–3952 (2024).
Prajapati, S. K., Joshi, H., Dev, V. & Dua, V. K. Molecular epidemiology of Plasmodium vivax anti-folate resistance in India. Malar. J. 10, 102 (2011).
Beckmann J. F., Van Vaerenberghe K., Akwa D. E., Cooper B. S. A single mutation weakens symbiont-induced reproductive manipulation through reductions in deubiquitylation efficiency. Proc Natl Acad Sci USA 118, (2021).
Kazemi-Esfarjani, P. et al. Substitution of valine-865 by methionine or leucine in the human androgen receptor causes complete or partial androgen insensitivity, respectively with distinct androgen receptor phenotypes. Mol. Endocrinol. 7, 37–46 (1993).
Walss-Bass, C. et al. A novel missense mutation in the transmembrane domain of neuregulin 1 is associated with schizophrenia. Biol. Psychiatry 60, 548–553 (2006).
Rajebhosale, P. et al. Neuregulin1 nuclear signaling influences adult neurogenesis and regulates a schizophrenia susceptibility gene network within the mouse dentate gyrus. bioRxiv (2024).
Adjalley, S. & Lee, M. C. S. CRISPR/Cas9 Editing of the Plasmodium falciparum Genome. Methods Mol. Biol. 2470, 221–239 (2022).
Ghorbal, M. et al. Genome editing in the human malaria parasite Plasmodium falciparum using the CRISPR-Cas9 system. Nat. Biotechnol. 32, 819–821 (2014).
Heigwer, F., Kerr, G. & Boutros, M. E-CRISP: fast CRISPR target site identification. Nat. Methods 11, 122–123 (2014).
Untergasser, A. et al. Primer3-new capabilities and interfaces. Nucleic Acids Res 40, e115 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet 43, 491–498 (2011).
Philipp, M. et al. MutationExplorer: a webserver for mutation of proteins and 3D visualization of energetic impacts. Nucleic Acids Res 52, W132–W139 (2024).
Campbell, T. L., De Silva, E. K., Olszewski, K. L., Elemento, O. & Llinas, M. Identification and genome-wide prediction of DNA binding specificities for the ApiAP2 family of regulators from the malaria parasite. PLoS Pathog. 6, e1001165 (2010).
Bindels, D. S. et al. mScarlet: a bright monomeric red fluorescent protein for cellular imaging. Nat. Methods 14, 53–56 (2017).
Anna O. Chertkova MM, Marten Postma, Nikki van Bommel, Sanne van der Niet, Kevin L. Batenburg, Linda Joosen, Theodorus W.J. Gadella Jr., Yasushi Okada, Joachim Goedhart. Robust and Bright Genetically Encoded Fluorescent Markers for Highlighting Structures and Compartments in Mammalian Cells. bioRxiv (2017).
Zhang, W. et al. Optogenetic control with a photocleavable protein, PhoCl. Nat. Methods 14, 391–394 (2017).
Sinha, A. et al. A cascade of DNA-binding proteins for sexual commitment and development in Plasmodium. Nature 507, 253–257 (2014).
Acknowledgements
This investigation received financial support from Public Health Service grants from the National Institute of Allergy and Infectious Diseases (NIAID), National Institutes of Health (NIH), KCW (R01 AI069314 and R01 AI103638) and ML [R01 AI076276 and R01-AI125565 with support from the Center for Quantitative Biology (P50 GM071508)], the Intramural Research Program of the NIAID, NIH (BJM) and intramural research funds from the Uniformed Services University of the Health Sciences, USA (KCW: C073384316). We thank Drs. Kateryna Lund and Sintayehu Gebreyohannes (USU-BIC facility) for flow cytometry support. We extend thanks to former lab members Christen Dona and Van Anh Nguyen for their support generating a transgenic line (683.gdv1.3xha.tdTom.fkbp) as well as Will Valiant for preparing a plasmid (p1605.stk2.gfp.fkbp.wr). The single plasmid CRISPR system (pDC-cam-cas9-gfp.wr) was a gift from Prof Marcus Lee (University of Dundee, UK). All Addgene plasmids containing fluorescent protein were gifts from the Li Qian lab (UNC Chapel Hill, USA, tdTomato), Dorus Gadella Lab (University of Amsterdam, Netherlands, mScarlet and mNeonGreen) and Robert Campbell Lab (University of Tokyo, Japan, NanoLuciferase). The opinions and assertions expressed herein are those of the author(s) and do not reflect the official policy or position of the Uniformed Services University of the Health Sciences (USUHS), or Henry M. Jackson Foundation for the Advancement of Military Medicine, Inc. (HJF), or the Department of Defense (DOD).
Author information
Authors and Affiliations
Contributions
Study design: S.K.P., K.C.W., data collection: S.K.P., J.X.D., B.J.M., T.D., M.C.B., A.E.W., D.H., and B.F.C.K.; data analysis and interpretation: S.K.P., J.X.D., B.J.M., D.H., K.C.W., M.D.W., C.L.D., and M.L.; statistical analysis: S.K.P. and K.C.W.; paper writing and critical review: S.K.P., K.C.W., D.H., C.L.D., M.L., and B.F.C.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Abhisheka Bansal and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Prajapati, S.K., Dong, J.X., Morahan, B.J. et al. A single valine to leucine switch disrupts Plasmodium falciparum AP2-G DNA binding and reveals GDV1’s role in ap2-g activation. Nat Commun 17, 1719 (2026). https://doi.org/10.1038/s41467-026-68416-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68416-1