Abstract
Innate lymphoid cells (ILCs) are crucial for cancer immunosurveillance. While mouse type 1 ILCs (ILC1s) control acute myeloid leukemia (AML) by targeting leukemia stem cells (LSCs), the role of human ILC1s in AML remains largely undefined. Here, we find that ILC1s in AML patients are impaired, with reduced total ILC1 numbers and diminished function. In contrast, healthy donor (HD) ILC1s-derived TNFα inhibits the leukemic transition from CD34+CD38+ to CD34−CD38+ cells and blocks the differentiation of LSCs (CD34+CD38−) into immunosuppressive, macrophage-like leukemia-supporting cells. HD ILC1-derived IFNγ partially suppresses the differentiation of CD34−CD38+ to CD34−CD38− cells. These combined effects limit human leukemogenesis in vivo. We also identify a human ILC1 subset as Lin−CD127+CD161−CRTH2−CD117− (CD161− ILC1s) that can be generated from umbilical cord blood CD34+ hematopoietic stem cells. This method could provide a reliable source of ILC1s for potential adoptive transfer therapies in AML, offering a therapeutic approach to prolong disease-free survival in AML.
Similar content being viewed by others
Introduction
Acute myeloid leukemia (AML) is a devastating disease, with a median 5-year survival is 30–35% for patients younger than age 65 treated with standard chemotherapy1. Although allogeneic stem cell transplantation has been shown to be curative in some cases, the treatment-related mortality and the risk for disease relapse due to the possible persistence of leukemia stem cells (LSCs, defined by a CD34+CD38− phenotype) remain relatively high2,3. LSCs possess self-renewal capacity and are believed to be the driving force behind leukemia initiation, maintenance, and relapse2,4. They have the potential to differentiate into more mature leukemic progenitor cells (CD34+CD38+) and terminally differentiated blasts (myeloid blasts or leukemic cells; CD34−CD38+), which contribute to disease progression5,6. CD34+CD38+ leukemic progenitor cells have progressed beyond the undifferentiated LSC stage7. These cells exhibit increased differentiation compared to LSCs but are still considered immature with self-renewal capacity and the ability to propagate the disease. CD34−CD38+ cells represent a more terminally differentiated leukemic population, having lost some of the stem cell-like properties compared to LSCs and CD34+CD38+ leukemic progenitor cells5,8. While they are more differentiated, CD34−CD38+ cells still possess some self-renewal capacity and the ability to give rise to more mature leukemic cells, perpetuating the disease9. LSCs can also differentiate into CD34⁻CD38⁻ cells, a subset of leukemic cells that are highly resistant to standard therapies. Higher levels of these cells are often associated with a poorer prognosis and a higher likelihood of disease relapse after treatment5,10. Due to the complexity and heterogeneity of LSC differentiation and their relatively lower sensitivity to treatment compared to later stages of AML, understanding the differentiation pathways and properties of these cell populations, as well as developing safer and more effective therapeutic approaches, is crucial for improving the clinical outcome of patients with AML.
Innate lymphoid cells (ILCs) play a critical role in mediating immune responses and regulating tissue homeostasis and inflammation11,12,13,14,15,16,17,18,19,20,21,22. Unlike adaptive immune cells, ILCs lack antigen-specific receptors and instead are activated by cytokine signals23. ILCs are classified into three main subsets based on their cytokine production profiles and transcription factor (TF) requirements: Group 1 ILCs, including ILC1s and natural killer (NK) cells that produce interferon-gamma (IFNγ) and tumor necrosis factor-alpha (TNFα), and are primarily involved in immune responses against intracellular pathogens such as viruses and certain bacteria as well as tumors24,25,26. These cells express the TF T-BET. Group 2 ILCs, known as ILC2s, secrete type 2 cytokines such as IL-4, IL-5, IL-9, and IL-13, which play a pivotal role in defense against extracellular parasites like helminths, in allergic inflammation, and defense against tumors22,27,28. These cells are characterized by the expression of the TF GATA3. Group 3 ILCs, or ILC3s, produce IL-17 and IL-22, contributing significantly to mucosal immunity, tumor immunity, and the maintenance of barrier integrity, particularly within the gastrointestinal tract29,30,31,32. These cells express the TF RORγt as a defining feature of their phenotype. We recently reported that ILC1s from healthy mice contribute to the control of murine AML by eliminating LSCs and inhibiting their differentiation into myeloid blasts33. However, the potent anti-cancer properties and mechanisms of human ILC1s in AML remain to be further explored.
In this study, we demonstrate that human ILC1s are quantitatively and qualitatively impaired in AML patients, thereby playing a pivotal role in influencing the fate of LSCs. Specifically, we demonstrate that ILC1s isolated from healthy donors (HD) can hinder the differentiation of LSCs into leukemic progenitor cells or a more differentiated population of leukemic cells via their secretion of IFNγ or TNFα. Additionally, these ILC1s can limit the differentiation of LSCs into immunosuppressive, macrophage-like leukemia-supporting cells via their secretion of TNFα, while ILC1s from AML patients cannot. These regulatory functions of HD ILC1s converge to limit human leukemogenesis in vivo. Our study also reveals the inherent heterogeneity within the human ILC1 population, comprising both CD161+ and CD161− subsets. Despite this diversity, both subsets demonstrate similar expression of crucial TFs and effector molecules. We also demonstrate how umbilical cord blood (UCB) CD34+ cells can efficiently generate ILC1s with promising anti-AML activity. Our study shows that human ILC1s are defective in AML patients, yet human HD ILC1s or those generated from UCB CD34+ cells can halt the progression of AML and have the potential to be manufactured ex vivo for improved treatment of AML in vivo.
Results
ILC1 population and their functional roles are disrupted in the setting of AML
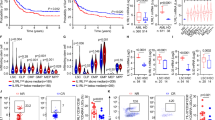
We have reported that DNAM-1(+) and IL-7 receptor (R) (+)-ILC1s isolated from healthy mice can recognize CD155/CD112 and IL-7, respectively, on mouse LSCs and induce IFNγ that in turn promotes apoptosis and blocks their differentiation into myeloid blasts33. We have also reported that human ILC1s (defined as Lin−CD127+CD161+/−CRTH2−CD117−, hereafter referred to as ILC1s) promote apoptosis of human LSCs (defined by CD34+CD38−) at a high ratio of ILC1s to LSCs33. Whether human ILC1s control LSC fate by affecting the differentiation of human LSCs remains largely undefined. Human ILC1s can be defined as Lin−CD127+, with “Lin−” defined as depletion of CD3, CD4, CD8, CD14, CD15, CD16, CD19, CD20, CD33, CD34, CD203c, FceRI, and CD56 positive cells. Following analysis of ILC1s in the peripheral blood (PB) of healthy donors (HDs) and AML patients, we observed a highly significant increase in the percentage of ILC1s among total ILCs (CD127+) in AML patients relative to HDs (Fig. 1a and Supplementary Fig. 1a). However, within total lineage-negative cells from the same patient population, there was a notable decrease in both the percentage (Fig. 1b) and absolute number of the ILC1 subset compared to HDs (Fig. 1c). This finding is further supported by our assessment of T-BET expression (encoded by TBX21), a critical TF of human ILC1s. Data showed that the number of T-BET+ ILC1s was decreased in AML, while T-BET expression on a per cell basis is equivalent between HDs and AML patients (Fig. 1d and Supplementary Fig. 1b).
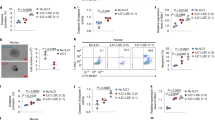
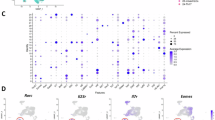
a Representative flow cytometry plots of the percentages (left) and statistics (right) of ILC1s among total ILCs (CD127+) in peripheral blood (PB; n = 6 each). b Percentages of total ILC1s among total lineage-negative cells in PB (n = 6 each). c An absolute number of total ILC1s in PB (n = 6 each). d Total number of T-BET+ ILC1s in the PB (n = 6 each). e A UMAP analysis of human ILCs identified four distinct clusters (ILC1, ILC2a, ILC2b, and ILCp) from the PB (n = 2 HDs; n = 4 AML patients). Cells are color-coded according to the defined subsets. f Dot plot analysis displaying expression of selected and previously described genes encoding specific cell surface markers, cytokine receptors, and transcriptional factors used to annotate clusters (n = 2 HDs, n = 4 AML patients). g Volcano plots show DEGs between HD ILC1s (n = 2) and AML ILC1s (n = 4). h Graphics displaying the functional score in HD ILC1s (n = 2) and AML ILC1s (n = 4) associated with signaling pathways for type 2 IFN production, TNF superfamily cytokine production, cell killing, and cytolysis. i, j Representative plots and summary data of HD ILC1s and AML ILC1s producing IFNγ (i, n = 6 HDs, n = 4 AML patients) and TNFα (j, n = 3 HDs, n = 3 AML patients). k, l Representative plots (left) and statistics (right) of the expression of DNAM-1 (k) and NKG2D (l) in HD ILC1s (n = 6) and AML ILC1s (n = 5). Data are representative of two (i–l) and three (a–d) independent experiments, and are presented as mean ± s.d.; P values were calculated by two-tailed Student’s t-test (a–d, i–l) or by two-tailed Wilcoxon Rank-Sum Test (g, h; an adjusted P value was used). Boxplots display the median and interquartile range (25th percentile –75th percentile) with whiskers representing the upper- and lower-quartile (1.5× the 75th and 25th percentile values). Source data are provided as a Source Data file.
We also analyzed 10× Genomics high-throughput droplet-based single-cell RNA sequencing (scRNA-seq) that we previously performed22. In our scRNA-seq, we initially employed the EasySep™ Human Pan-ILC Enrichment Kit to isolate total ILCs. Following this, we performed cell sorting using a comprehensive set of lineage markers34,35,36 (CD3, CD4, CD8, CD14, CD15, CD16, CD19, CD20, CD33, CD34, CD203c, FcεRI, and CD56), in addition to CD127. This strategy of utilizing specific surface markers ensured the exclusion of T cells (including CD4+ T cells, CD8+ T cells, and other T cell lineage cells), B cells, NK cells, myeloid cells, and CD34+ hematopoietic precursors, as well as nonhematopoietic cells from the sorted population. We also excluded cells that expressed CD4 and CD8 during the analysis. This exclusion of CD4 and CD8 differs from an earlier report37. By selecting Lin−CD127+ cells, we were able to focus on ILCs, ensuring that no other known immune cell types were present in our analysis. The examination of transcriptomic profiles in sorted total ILCs identified the ILC1 population (cluster 8; Supplementary Fig. 1c) from HDs and AML patients using previously reported ILC1 markers34 (Fig. 1e, f and Supplementary Fig. 1c, d). This population expressed ILC1-specific markers, IL7R, GZMK, IKZF3, IL2RB, LEF1, and GPR183, without the expression of NK cell markers such as FGFBP2, FCGR3A, KIR2DL1, KIR2DL3, and GZMH and without expression of T cell markers, CD4 and CD8 (Supplementary Fig. 1e). Compared to AML ILC1s, HD ILC1s exhibited higher expression of cytotoxicity-associated genes, including NKG7, PRF1, CST7, and CTSW38,39 (Fig. 1g). Gene ontology (GO) pathway analysis for ILC1s from AML patients demonstrated a decrease in pathways associated with cell killing, cell activation, and cytokine production when compared to HD ILC1s (Supplementary Fig. 1f). Additionally, compared to HD ILC1s, AML ILC1s showed a lower functional score of signaling pathways for type 2 IFN production, TNF superfamily cytokine production, cell killing, and cytolysis (Fig. 1h). These findings collectively indicate a dysfunctional state of ILC1s in AML patients. The results were validated by flow cytometry, revealing a significant reduction in the production of IFNγ and TNFα in ILC1s from AML patients compared to HD ILC1s (Fig. 1i, j). Further, ILC1s from AML patients exhibited decreased surface expression of activation receptors DNAM-1 and NKG2D relative to HD ILC1s (Fig. 1k, l). Finally, a negative correlation was observed between the percentage of ILC1s and the percentage of leukemic blasts in AML patients (R = −0.9024, P = 0.0004) (Supplementary Fig. 2a, b). Collectively, these data suggest that the functional role(s) of human ILC1s become dysregulated in the context of AML. A high level of human ILC1s appears to be associated with a lower AML blast count, suggesting that ILC1s could have a role in controlling AML.
HD ILC1 TNFα reduces the differentiation of CD34+CD38+ cells into CD34−CD38+ cells, while HD ILC1 IFNγ reduces the differentiation of CD34−CD38+ cells into CD34−CD38− cells
As noted above, HD human ILC1s induce cell death of human LSCs at a high ILC1:LSC ratio of 4:133. We also explored the impact of HD ILC1s on LSCs at a low effector-to-target ratio (E:T) of one ILC1 to four LSCs (1:4), as well as a culture of LSCs alone with recombinant human (rh) IFNγ (50 ng/ml) or rhTNFα (20 ng/ml) for 3 days. Our results show that compared to LSC alone, (1) coculturing LSCs with HD ILC1s or treatment with rhIFNγ or rhTNF did not significantly change the absolute number and percentage of LSCs during coculture (Fig. 2a, b and Supplementary Fig. 3a); (2) coculturing LSCs with HD ILC1s or rhTNFα, but not rhIFNγ, significantly increased the number of LSC-derived CD34+CD38+ cells (Fig. 2a, c), while the percentage of LSC-derived CD34+CD38+ cells was elevated only in the rhTNFα-treated group (Supplementary Fig. 3b); (3) coculturing LSCs with HD ILC1s or rhIFNγ or rhTNFα significantly reduced the absolute number of LSC-derived CD34−CD38+ cells (Fig. 2a, d). The percentage of LSC-derived CD34−CD38+ cells was higher in the ILC1 and rhIFNγ-treated groups but reduced in the rhTNFα-treated group (Supplementary Fig. 3c); and (4) coculturing LSCs with HD ILC1s or rhIFNγ or rhTNFα significantly decreased the number of LSC-derived CD34−CD38− cells (Fig. 2a, e). The percentage of LSC-derived CD34−CD38− cells was decreased only in the HD ILC1 and rhIFNγ-treated groups but not in the rhTNFα-treated group (Supplementary Fig. 3d).
a–e Human LSCs (CD34+CD38−) from the blood of AML patients were co-cultured for 3 days without or with human HD ILC1s or with human recombinant IFNγ (50 ng/ml) or with TNFα (20 ng/ml). Representative flow cytometry plots (a) and summary data showing the number of CD34+CD38− (b), CD34+CD38+ (c), CD34−CD38+ (d), and CD34−CD38− (e) cells (n = 8 HDs; n = 8 AML patients). f–j Human LSCs were co-cultured for 3 days without or with human HD ILC1s, or HD ILC1s with anti-IFNγ (20 μg/ml), or HD ILC1s with anti-TNFα (20 μg/ml). Representative flow cytometry plots (f) and summary data showing the number of CD34+CD38− (g), CD34+CD38+ (h), CD34−CD38+ (i), and CD34−CD38− (j) cells (n = 8 HDs; n = 8 AML patients). Data are representative of three (a–j) independent experiments and are presented as mean ± s.d.; P values were calculated by one-way ANOVA model (g–j) or after Log10 transformation due to big variations, followed by one-way ANOVA model (b–e). Source data are provided as a Source Data file.
However, compared to HD ILC1s cocultured with LSCs, HD ILC1s cocultured with LSCs and neutralizing mAb to TNFα or IFNγ did not significantly change the absolute number or percentage of CD34+CD38− LSCs during coculture (Fig. 2f, g and Supplementary Fig. 3e). The addition of a neutralizing mAb to TNFα but not to IFNγ, suppressed the increased number of CD34+CD38+ cells from LSCs induced by HD ILC1s; however, the percentage of CD34+CD38+ cells was unaffected (Fig. 2f, h and Supplementary Fig. 3f). In contrast, HD ILC1s cocultured with LSCs and neutralizing mAb to IFNγ but not TNFα reversed the decrease in number and the increase in percentage of CD34−CD38+ cells that were originally differentiated from LSCs cocultured with HD ILC1s (Fig. 2f, i and Supplementary Fig. 3g). These data highlight the distinct roles of HD ILC1-derived IFNγ and TNFα in the regulation of LSC differentiation. However, the absence of any notable impact via the neutralization of either cytokine on the differentiation of CD34−CD38− cells from LSCs in the presence of ILC1s suggests that other factors may regulate this specific cell population (Fig. 2f, j and Supplementary Fig. 3h).
HD ILC1-derived TNFα rather than IFNγ reduces the differentiation of immunosuppresive, macrophage-like leukemia-supporting cells from LSCs
Several reports have described that CD11b+CD206+ immunosuppresive, macrophage-like leukemia-supporting cells inhibit effective immune responses and contribute to a favorable environment for LSCs by secreting factors that promote leukemia progression40,41,42. However, it remains unclear whether LSCs can directly differentiate into CD11b+CD206+ immunosuppressive, macrophage-like leukemia-supporting cells. To investigate this, we cocultured HD ILC1s with LSCs at the E:T of 1:4 for 3 days. By Wright-Giemsa staining and flow cytometry, we observed that ILC1s blocked the differentiation of LSCs into CD11b+CD206+ macrophage-like leukemia-supporting cells. The addition of a neutralizing mAb to TNFα but not to IFNγ reversed this effect (Fig. 3a–c). This was further supported by the observation that rhTNFα, but not rhIFNγ, reduces the differentiation of LSCs into these supportive cells (Fig. 3d, e). Importantly, the inhibitory effect of ILC1s on LSC differentiation into immunosuppressive, macrophage-like leukemia-supporting cells was significantly diminished when cocultured with ILC1s from AML patients (Supplementary Fig. 4a,b), consistent with the lower TNFα production by ILC1s from AML patients (Fig. 1j). These findings underscore the dysfunction of ILC1s in AML patients and highlight the importance of TNFα signaling along with IFNγ in regulating LSC differentiation. The colony-forming unit (CFU) assay further confirmed that HD ILC1s impede the differentiation of human LSCs into munosuppresive, macrophage-like leukemia-supporting cells, as indicated by decreased numbers of total colonies and granulocyte-macrophage progenitor (CFU-GM) colonies after coculture with HD ILC1s compared to culture without HD ILC1s, while there was no significant difference in the number of granulocytes (CFU-G) colonies (Fig. 3f, g and Supplementary Fig. 4c).
a Human LSCs were co-cultured without or with human HD ILC1s for 3 days. Representative images of Wright-Giemsa staining of cells (20× magnification, scale bar, 100 µm, n = 3). b, c Human LSCs were co-cultured without or with HD ILC1s, HD ILC1s plus anti-IFNγ, or HD ILC1s plus anti-TNFα for 3 days. Representative flow cytometry plots (b) and summary data of the absolute cell number (c; left) and percentage (c; right) of CD11b+CD206+ cells (n = 7 HDs; n = 7 AML patients). d, e Human LSCs were co-cultured without or with IFNγ or TNFα for 3 days. Representative flow cytometry plots (d) and summary data of the absolute cell number (e; left) and percentage (e; right) of CD11b+CD206+ cells (n = 4 AML patients). f, g Human LSCs were co-cultured without or with HD ILC1s for 3 days. Total colony-forming cells (f) and colony-forming myeloid progenitors (CFU-GM and CFU-G; g) were counted at each round of plating (n = 3 HDs; n = 3 AML patients). (h) Identification of FLT3-ITD mutation by PCR. A representative 3% agarose gel electrophoresis of PCR products. The 329 bp fragment indicates the wild type. The presence of both a band at 329 bp and a fragment larger than 329 bp (>329) indicates FLT3-ITD mutation56 (n = 2 AML patients). Data are representative of two (a, d, e, and h) and three (b, c, f, and g) independent experiments and are presented as the mean ± s.d.; P values were calculated by one-way ANOVA (c and e) and two-tailed unpaired Student’s t test (f and g). For (f), P values were calculated after the Log10 transformation due to large variations. Source data are provided as a Source Data file.
To provide definitive evidence that these munsuppresive, macrophage-like leukemia-supporting cells are, in fact, derived from LSCs, we sorted LSCs from two AML patients with the FMS-like tyrosine kinase 3-internal tandem duplication (FLT3-ITD) mutation and induced their differentiation into CD11b+CD206+ immunosuppressive macrophage-like leukemia-supporting cells. The CD11b+CD206+ cells were then sorted, and the presence of the FLT3-ITD was detected by polymerase chain reaction (PCR) using primers specific for the mutation, thus demonstrating their derivation from the LSC43,44 (Fig. 3h, Lane 1 and Lane 2).
Collectively, these data indicate that produced by HD ILC1 TNFα contributes to reducing the differentiation of LSCs into CD11b+CD206+ immunosupressibe, macrophage-like leukemia-supporting cells in vitro, whereas this same effect is impaired in ILC1s from AML patients.
HD ILC1s prolong the survival of mice bearing human LSCs in vivo
We also performed the in vivo transplantation experiment, in which human CD34+CD38− LSCs and human HD ILC1s were administrated intravenously (i.v.) into NOD.Cg-Prkdcscid Il2rgtm1Wjl Tg(CMV-IL3,CSF2,KITLG)1Eav/MloySzJ (NSG-SGM3) mice. On each of days 1–7, the mice received an intraperitoneal (i.p.) injection of human IL-15 (hIL-15) (Fig. 4a). The mice were monitored for over 600 days. We found that the injection of human HD ILC1s reduced the LSC engraftment in NSG-SGM3 mice and suppressed the progression of AML, as evidenced by a significantly decreased number of CD45+CD33+ blast cells, a significantly decreased number of CD34+CD38− LSCs, and significantly prolonged survival of NSG-SGM3 mice, all compared to mice that did not receive an injection of HD ILC1s but received daily i.p. injections of hIL-15 from days 1 to 7 (Fig. 4b–d). These data indicate that human HD ILC1s can suppress the differentiation of human LSCs and the development of AML in vivo. Taken together, these findings strongly suggest that human HD ILC1s have a role in protecting against AML and provide a rationale for using HD ILC1s as a cellular-based therapy to extend disease-free survival in AML.
a Schematic illustration of the design and procedures for (b–d). 0.5 × 104 human LSCs were sorted from the blood of AML patients were i.v. injected into NSG-SGM3 mice on day 0. One day later, 5 × 104 human HD ILC1s were sorted from the blood of human healthy donors and i.v. injected into these mice. Three weeks later, the numbers of human CD34+CD38− LSCs and CD45+CD33+ cells were analyzed. Images were created with Adobe Photoshop. b Summary data of the absolute cell number of human CD34+CD38− LSCs in the blood of NSG-SGM3 mice (n = 5 individual mice). c Summary data of the absolute cell number of human CD45+CD33+ cells in the blood of NSG-SGM3 mice (n = 5 individual mice). d Survival of the mice transplanted with LSCs and treated without or with HD ILC1s isolated from PBMCs (n = 5 individual mice in each group). Survival data were analyzed by Kaplan–Meier survival analysis and log-rank test. All non-survival data are shown as mean ± s.d. P values were calculated by two-taild Student’s t-test (b and c). Source data are provided as a Source Data file.
CD161− ILC1s exhibit a phenotype similar to CD161+ ILC1s
ILC1s have conventionally been characterized as CD161-expressing cells. However, isolation and ex vivo expansion of Lin−CD161+ ILC1s from either HDs or AML patients does not result in a homogeneous population of ILC1s. Through extensive flow cytometric analyses, we found that ILC1s are heterogeneous in human PB using ILC1-specific surface markers (Lin, CD127, CRTH2, and CD117) that were reported previously35,45 along with an anti-CD161 mAb (Supplementary Fig. 5). The heterogeneous ILC1s in the blood of HDs included the conventional CD161+ ILC1s (hereafter referred to as CD161+ ILC1s) and Lin−CD127+CD161−CRTH2−CD117−cells (hereafter referred to as CD161− ILC1s). In our hands, the percentage of CD161− ILC1s was higher than CD161+ ILC1s among total ILCs isolated from HD PB (Fig. 5a). Moreover, both CD161+ and CD161− ILC1s exhibited comparable expression levels of T-BET and EOMES (Note: human EOMES+ ILC1s were also previously reported in human ILC1s34) (Fig. 5b), comparable production of IFNγ and TNFα (Fig. 5c), and comparable expression of the cytolytic molecules GZMA and GZMB (Note: human GZMA- and GZMB-expressing ILC1s were also previously reported11 (Fig. 5d, e).
a Total ILCs were isolated from PBMCs of HDs to quantify and display the proportion of CD161+ and CD161− ILC1s (left-bottom quadrants highlighted in color in the right panel) within Lin− (Lin: surface CD3, CD4, CD8, CD14, CD15, CD16, CD19, CD56, CD33, CD34, CD203c, CD123, FcεRIα, viability marker) cells. b Representative flow cytometry plots (left) and summary data (right) for the expression of EOMES and T-BET in CD161+ and CD161− ILC1s (n = 4 HDs). c Representative flow cytometry plots (left) and summary data (right) of the expression of IFNγ and TNFα in CD161+ and CD161− ILC1s (n = 4 HDs). d, e Representative flow cytometry plots (left) and summary data (right) of the expression of GZMA (d) and GZMB (e) in CD161+ and CD161− ILC1s (n = 4 HDs). Data are representative of two independent experiments (a–e) and are shown as mean ± s.d. P values were calculated by two-tailed Student’s t-test (b–e). NS: not significant. Source data are provided as a Source Data file.
We further expanded the total ILCs on either DL1 or DL4-transfected OP9 (hereafter referred to as DL1 and DL4) stromal cells in the presence of IL-2, IL-7, and IL-15 and found that there were far fewer CD161+ ILC1s and a far greater number of CD161− ILC1s regardless of whether they were co-cultured with DL1 or DL4 stromal cells (Supplementary Fig. 6a–d). The CD161− ILC1s were further identified by the expression of IFNγ, T-BET, and EOMES (Supplementary Fig. 6e, f), indicative of an ILC1 lineage.
Efficient generation of ILC1s from umbilical cord blood CD34+ cells confers anti-AML activity
Using the same method recently reported for ILCs differentiated from umbilical cord blood (UCB) CD34+ hematopoietic stem cells (HSCs)46, we obtained ILC1s that were almost entirely CD161−, regardless of co-culture on either DL1-expressing or DL4-expressing OP9 stromal cells, as demonstrated by the phenotype and the expression of IFNγ, EOMES, and T-BET (Fig. 6a–c). These cells were referred to as UCB CD34+ cell-derived CD161− ILC1s. We also observed that both UCB freshly isolated CD161+/− ILC1s and UCB CD34+ cell-derived CD161+/−ILC1s expressed NKG2D and CD200R1 (Fig. 6d). However, unlike NK cells, they do not express KIR (including KIR2DL4, KIR2DL5, and KIR2DL1/S1/S3/S5). Additionally, unlike UCB CD34 cell-derived CD161+/−ILC1s, UCB-freshly isolated CD161+/− ILC1s do not express NKG2A (Fig. 6d). Furthermore, both UCB freshly isolated CD161− ILC1s and UCB CD34+ cell-derived CD161−ILC1s expressed GZMB and perforin (Supplementary Fig. 7). A nearly 700-fold increase from UCB CD34+ cells to CD161− ILC1s (>97% purity) was observed on day 28 (Fig. 6e). The cytotoxicity against LSCs was comparable between UCB CD34+ cell-derived CD161− ILC1s and the CD161+ ILC1s isolated from UCB mononuclear cells (UBMCs; Fig. 6f, g). Neither UCB CD34+ cell-derived CD161− ILC1s nor UBMC-derived CD161+ ILC1s displayed high cytotoxicity against HSCs isolated from HD PB (Fig. 6f, g). Injection of UCB CD34+ cell-derived CD161− ILC1s significantly extended the survival of mice bearing human LSCs when compared to the group injected with LSCs alone (Fig. 6h, i). Importantly, this effect was abolished upon administration of anti-IFNγ (Fig. 6j,k). Furthermore, we also compared the anti-tumor effect of ILC1s and another innate effector cell type (NK cells). Our results demonstrated that injections of both UCB CD34+ cell-derived CD161− ILC1s and UCB CD34+ cell-derived NK cells delayed the progression of AML compared to the control group without these injections. However, the injection of UCB CD34+ cell-derived CD161− ILC1s showed a trend towards better prolongation of survival than the injection of UCB CD34+ cell-derived NK cells in our LSC model (Fig. 6j, k). Taken together, the results suggest that ILC1s derived from UCB CD34+ cells exhibit anti-AML efficacy both in vitro and in vivo.
a Schematic of culture conditions. UCB CD34+ HSCs were cultured on either OP9-DL1 or OP9-DL4 cells in the presence of Flt3-L, SCF, IL-3, IL-7, and IL-15 for 14 days. Created in BioRender. Ma, R. (2024) https://BioRender.com/w45e333. b Percentage of CD161+ and CD161− ILC1s, ILC2s, and ILC3s within Lin− cells. c Representative expression of IFNγ, T-BET, and EOMES within CD161− ILC1s on day 14. d Representative expression of NKG2A, NKG2D, KIR2DL4, KIR2DL5, KIR2DL1/S1/S3/S5, and CD200R in ILC1s. e The fold change of harvested ILC1s vs. pre-seeded UCB CD34+ HSCs cultured on OP9-DL1 cells in the presence of Flt3-L, SCF, IL-7, IL-15, and IL-3 (n = 4 HDs). f, g Representative flow cytometry plots (f) and summary data of (g) the percentages of apoptotic cells identified by Annexin V and DAPI in human LSCs (n = 4 HDs; n = 4 AML patients). h Schematic of the design and procedures for (i). Images were created with Adobe Photoshop. i Survival of the mice transplanted with LSCs and treated without or with UCB CD34+ cell-derived CD161− ILC1s (n = 6 mice). Survival data were analyzed by Kaplan–Meier survival analysis and log-rank test. j Schematic of the design and procedures reported in (k) showing 5 × 104 LSCs were i.v. injected into NSG-SGM3 mice on day 0. One day later, mice were randomized to receive (1) no ILC1 or NK cell infusion; (2) 1.5 × 105 UCB CD34+ cell-derived CD161− ILC1s; (3) 1.5 × 105 UCB CD34+ cell-derived NK cells, or (4) 1.5 × 105 UCB CD34+ cell-derived CD161− ILC1s plus anti-IFNγ (500 μg given intraperitoneally every 2 days for 8 days). Images were created with Adobe Photoshop. Survival data (n = 5 mice) were analyzed by Kaplan–Meier survival analysis and log-rank test. Data are shown as the means ± s.d. NS: not significant. Source data are provided as a Source Data file.
Discussion
In AML, LSCs play a pivotal role in disease relapse and the failure to achieve a cure in many cases. They are often resistant to conventional chemotherapy and can remain dormant or evade immune surveillance, allowing them to survive initial treatment, thereby eventually leading to disease relapse47. This selective pressure can lead to the expansion of resistant LSC populations and the eventual relapse of the disease48. Furthermore, the process of leukemic differentiation, where LSCs give rise to more mature leukemic cells, adds another layer of complexity to AML treatment49. Therefore, the inability to eradicate LSCs is a major challenge in achieving success with therapy for AML.
The results presented in this preclinical study shed light on a pivotal role human ILC1s play in orchestrating the fate of human LSCs within the context of AML. We build upon our previous report focused on mouse ILC1s in AML33, where we demonstrated the ability of ILC1s to regulate the differentiation of mouse LSCs into terminal myeloid blasts via IFNγ. This study reveals that human ILC1s can similarly inhibit the differentiation of human LSCs into terminal myeloid blasts (CD34−CD38+ cells). However, unlike their mouse counterparts, this study also reveals that human ILC1s suppress the differentiation of LSCs into immunosuppressive, macrophage-like leukemia-supporting cells through the secretion of TNFα. These dual functions combine to significantly impede the progression of human AML. It’s noteworthy that immunosuppressive macrophages have been implicated in promoting the aggressive development of leukemia40,50. Therefore, in utilizing HD ILC1s for the treatment of AML, there would be a three-part strategy: apoptosis of LSCs33, suppression of differentiation, and disruption of the tumor-supportive microenvironment facilitated by immunosuppressive macrophages.
Earlier studies in AML demonstrated a decrease in the percentage of total ILCs within the lineage-negative cell subsets, coupled with an increase in the percentage of ILC1s among CD127+ cells35. However, the absolute number of ILC1s in the blood of AML patients remained unknown, leaving a gap in our understanding of ILC1 homeostasis in this disease. Here, we provide a comprehensive analysis of ILC1s in AML patients, demonstrating a significant decrease in both the percentage and the absolute number of PB ILC1s when compared to HDs. Furthermore, our study shows a negative correlation between the percentage of ILC1s and the percentage of leukemic blasts in AML patients. These data suggest that a reduction of ILC1 cell numbers could be a mechanism utilized by AML to enhance progression of the disease. Indeed, HD ILC1s significantly improved the survival of mice engrafted with LSCs from AML patients, suggesting that administration of allogeneic ex vivo-expanded ILC1s from HDs may have a positive impact on prolonging relapse-free survival of AML patients.
Characterizing human ILC1s has proven to be challenging due to the lack of unique surface markers, particularly for distinguishing ILC1s in peripheral blood. Recent studies have indicated that bona fide human ILC1s are primarily found in tissues rather than in peripheral blood37. However, these studies did not analyze Lin−CD3−CD4−CD8− cells, which are critical for accurate identification of ILC1 populations. In our analysis, we specifically excluded cells expressing T cell markers, CD4 and CD8, to ensure the integrity of our ILC population assessment. By filtering out these T cell lineage cells, we were able to isolate a distinct population expressing key ILC1 markers, without concomitant expression of NK cell markers and any T cell markers. Our findings suggest that bona fide ILC1s are indeed present in human peripheral blood, providing further validation of the existence of ILC1s outside of tissue environments.
Our study also reveals the heterogeneity inherent within the human ILC1 population. Through comprehensive flow cytometric analyses, we identified distinct subsets within the circulating ILC1 population, including a CD161+ subset known as conventional ILC1s46,51,52, and a CD161− subset. Expanding ILC1s ex vivo is challenging due to their limited abundance within PBMCs and low proliferative capacity. While a recent study provided a system for generating human CD161+ ILC1s from UCB CD34+ HSCs, it did not examine their expansion and efficacy against tumors in vitro or in vivo, or an analysis of the CD161− ILC1 subset46. However, our study reveals that CD161− ILC1 subsets can also be generated from UCB CD34+ HSCs. This subset exhibits strong anti-AML efficacy both in vitro and in vivo. Despite our findings suggesting the potential biological benefits of ILC1s for the treatment of AML, the practical aspects of their utilization in clinical practice must be carefully weighed. The manufacturing of UCB-expanded ILC1s is more complex and more resource-intensive compared to conventional NK cells. This complexity translates into longer production times and higher costs, which could present challenges in a clinical setting. However, if the ILC1s are effectors that kill AML LSCs, we should consider developing cost-effective and less time-consuming manufacturing approaches of ILC1s. The use of induced pluripotent stem cells (iPSCs) may offer a promising alternative to circumvent these limitations. iPSCs can be used to produce standardized, off-the-shelf NK cells, CAR NK cells, or CAR macrophages with improved anti-tumor activity53,54. This approach circumvents many of the challenges seen with blood-derived NK cells and/or T cells for adoptive cell therapy, such as the need for donor collection and the variability in cell quality. By enabling the routine derivation of uniform ILC1s from iPSCs, this strategy could provide a scalable and standardized solution for cancer treatment, potentially overcoming the current barriers to ILC1 engineering. Furthermore, the development of iPSC-derived ILC1s could facilitate the creation of personalized therapies that are tailored to the specific needs of individual patients, thereby enhancing the efficacy and safety of cancer treatments. Our results suggest that in the future, it may be feasible to provide a readily available, allogeneic, off-the-shelf supply of CD161− ILC1s from iPSCs, leading to improved therapeutic outcomes in AML and possibly other hematologic malignancies. Thus, our findings provide a foundation for developing a potential adoptive cell therapy.
In conclusion, our study highlights the quantitative and qualitative defects of ILC1s in AML. The observed alterations in ILC1 phenotype and function have important implications for understanding the immune dysregulation in AML and may inform the development of potential immunotherapeutic approaches targeting not only the LSC but also the immune microenvironment in AML. Further investigation into the precise mechanisms underlying the dysregulation of ILC1s in AML is warranted to fully elucidate their therapeutic potential in the management of this disease.
Methods
Human samples
Peripheral blood (PB) samples were obtained from healthy donors (HDs) and acute myeloid leukemia (AML) patients at the City of Hope National Medical Center (COHNMC). All HDs and AML patients signed an informed consent form approved by the City of Hope Institutional Review Board. Human specimens were collected from individuals with AML registered at the COHNMC who consented to an Institutional Review Board (IRB)-approved protocol (IRB 18067); healthy donor specimens were collected from individuals who consented to IRB 06229. Recruitment in this cohort was independent of sex, as any patient with newly diagnosed, relapsed, or refractory AML was eligible. All participants in this study were patients with relapsed or refractory acute myeloid leukemia. Eligibility criteria included being at least 18 years old and providing written informed consent in accordance with national legal and regulatory requirements before any project-specific procedures. Information on race, ethnicity, or other socio-economical parameters was not collected for this cohort. Healthy donors and patients with AML were not compensated for their blood cell or bone marrow donations. Detailed characteristics of the study population can be found in Supplementary Tables 1, 2. Given the limited size of the cohort, we did not conduct any analyses stratified by sex. Patients who met the inclusion criteria outlined above were recruited for this study, so we do not anticipate any self-selection or other biases. Mononuclear cells were isolated using Ficoll separation. LSCs were sorted using a BD FACSAriaTM Fusion (BD Biosciences). ILCs were isolated using EasySep™ Human Pan-ILC Enrichment Kit (STEMCELL) or were sorted using a BD FACSAriaTM Fusion. UCB was obtained from StemCyte, Inc. (Baldwin Park, California). CD34+ cells were isolated using the CD34 MicroBead Kit (Miltenyi Biotec) and then sorted using a BD FACSAriaTM.
Cells and cell culture
OP9 cells were purchased from the American Type Culture Collection (ATCC; Cat# CRL-2749). Using this base cell line, human DLL (delta-like canonical Notch ligand)1 and DLL4 genes were overexpressed to create OP9-DL1 and OP9-DL4 cell lines, respectively22. OP9-DL1 and OP9-DL4 were maintained in MEMα GlutaMAX media supplemented with 20% FBS, 100 U/ml penicillin, 0.1 mg/ml streptomycin, and 50 μM β-Mercaptoethanol. Human LSCs were cultured in StemSpan™ Serum-Free Expansion Medium II with penicillin (100 U/mL), streptomycin (100 mg/mL), stem cell factor (SCF, 20 ng/mL), thrombopoietin (TPO, 20 ng/mL), Flt3-L (20 ng/mL), IL-3 (10 ng/mL), and IL-6 (10 ng/mL). Human ILC1s were cultured in MEMα GlutaMAX media. The media was supplemented with 10% human AB serum, IL-2 (500 IU/mL), IL-7 (20 ng/mL), and IL-15 (20 ng/mL). Cultures were incubated at 37 °C in a humidified atmosphere of 5% CO2.
Flow cytometry
ILC1s from human peripheral blood were identified using surface staining with a live/dead cell viability cell staining kit and the following monoclonal antibodies: lineage (FITC-anti-CD3, Cat# 561802, Clone: HIT3a; anti-CD4, Cat# 555346, Clone: RPA-T4; anti-CD8, Cat# 555634, Clone: HIT8a; anti-CD14, Cat# 555397, Clone: M5E2; anti-CD15, Cat# 555401, Clone: HI98; anti-CD16, Cat# 555406, Clone: 3G8; anti-CD19, Cat# 555412, Clone: HIB19; anti-CD20, Cat# 555622, Clone: 2H7; anti-CD33, Cat# 555626, Clone: HIM3-4; anti-CD34, Cat# 555821, Clone: 581; anti-CD203c, Cat# MA5-28586, Clone: NP4D6; anti-FceRI Cat# 334608, Clone: AER-37 (CRA-1)), anti-CD56 (FITC, Cat# 562794, Clone: B159; AF700, Cat# 557919, Clone: B159; BV421, Cat# 562751, Clone: NCAM16.2, or BUV395, Cat# 563555, Clone: NCAM16.2), anti-CD127 (APC, Cat# 558598, Clone: HIL-7R-M21), anti-CRTH2 (PE-Cy7, Cat# 350118, Clone: BM16), and anti-c-Kit (PE, Cat# 555714, Clone: YB5.B8 or BV711, Cat# 313230, Clone: 104D2). Human LSCs were identified by lineage (FITC-anti-CD3, anti-CD4, anti-CD8, anti-CD14, anti-CD19, anti-CD20, anti-CD11b, anti-CD56, and anti-CD235a, Cat# 349104, Clone: HI264), anti-CD45 (BV510, Cat# 368526, Clone: 2D1; or BUV395, Cat# 563791, Clone: HI30), anti-CD34 (PE, Cat# 555822, Clone: 581; FITC, at# 555821, Clone: 581), and anti-CD38 (BV605, Cat# 303532, Clone: HIT2; PE-Cy7, Cat# 560677, Clone: HIT2). To examine intracellular cytokine production, intracellular staining for IFN-γ, TNFα, GZMA, GZMB, perforin, EOMES, and T-BET was performed using a Fix/Perm kit (eBiosciences), followed by staining with an anti-IFN-γ antibody (BV786, Cat# 5563731, Clone: 4S.B3; BV421, Cat# 564791, Clone: 4S.B3), an anti-TNFα antibody (APC, Cat# 502912, Clone: MAb11; BV650, Cat# 563418, clone: MAb11), an anti-GZMA antibody (Pacific Blue, Cat# 507207, Clone: CB9 or APC, Cat# IC29051A, Clone: 356412), an anti-GZMB antibody (PE, Cat# 561142, Clone: GB11), perforin (APC, Cat# 30811, Clone: dG9), anti-EOMES antibody (BUV395, Cat# 567171, Clone: X4-83 or PE, Cat# 566749, Clone: X4-83), or an anti-T-BET antibody (APC, Cat# 644814, Clone: 4B10), respectively. All mouse antibodies were used at 1:200. All human antibodies were used at 1:50, except for CD34, CD38, CRTH2, and CD117, which were used at 1:100. All analyses were performed on a Fortessa X-20 flow cytometer (BD Biosciences), and sorting was performed using a BD FACSAriaTM Fusion.
Single-cell RNA-seq data analysis
The single-cell RNA-seq data were analyzed following the methodology outlined in our previous report, utilizing the same analysis approach as reported22. Briefly, single-cell RNA sequencing data were processed and analyzed using the Scanpy framework. Cells with fewer than 300 detected genes or with a mitochondrial UMI fraction >15% were removed, and genes detected in fewer than three cells were excluded. Potential doublets were identified and filtered out using Scrublet. The remaining count matrices were normalized using scanpy.pp.normalize_total with ‘target_sum = 1e4’. To enrich for innate lymphoid cells (ILCs) and minimize T-cell contamination, we retained cells exhibiting positive expression of IL7R (CD127) and negative expression of CD4 and CD8A. Highly variable genes (HVGs) were identified using scanpy.pp.highly_variable_genes. To reduce technical variation, the effects of total UMI counts and mitochondrial transcript percentages were regressed out using scanpy.pp.regress_out on the HVG matrix. Principal component analysis was then performed using scanpy.tl.pca, and the top 30 components were retained. Batch effects were corrected using BBKNN, followed by Leiden clustering (scanpy.tl.leiden, resolution = 1). Uniform Manifold Approximation and Projection (UMAP) embeddings were generated using scanpy.tl.umap with default parameters.
Gene-set activity scores were computed using scanpy.tl.score_genes in Scanpy. Gene signatures were obtained from the MSigDB database. Statistical significance of score differences between groups was assessed using the Mann–Whitney U test.
Differential expression analysis was carried out using the Wilcoxon rank-sum test implemented in the scanpy.tl.rank_genes_groups. Genes with an adjusted p-value < 0.05 and an absolute log fold change >0.25 were considered significantly differentially expressed. Differentially expressed gene lists were subsequently subjected to Gene Ontology (GO) enrichment analysis to identify overrepresented biological processes.
In vitro LSCs and ILC1s co-culture assays
Human LSCs from AML patients were labeled with 5 mM CellTrace™ Violet dye (CTV) and co-cultured with or without ILC1s (Lin−CD127+CRTH2−CD117−) isolated from the PB of HD in the presence of human IL-12 (10 ng/ml) and IL-15 (100 ng/ml). For coculture assays with cytokines and antibodies, human LSCs were cocultured with or without IFNγ (50 ng/ml), TNFα (20 ng/ml), anti-IFNγ antibody (20 µg/ml), or anti-TNFα (20 µg/ml) antibody. For all coculture assays, after three days of coculture, cells were harvested and analyzed using flow cytometry. Annexin V and 7-amino-actinomycin D (7-AAD, BD Biosciences) were used to identify dead cells following the manufacturer’s instructions.
In vitro colony-forming unit assay
Human LSCs were isolated from the blood of AML patients and were cocultured with or without ILC1s isolated from the blood of HDs for 3 days. Cells were then plated into human methylcellulose complete media (R&D, HSC003) supplied with recombinant human SCF (50 ng/ml), human recombinant IL-3 (10 ng/ml), IL-6 (10 ng/ml), recombinant human GM-CSF (10 ng/ml), and recombinant human EPO (3 IU/ml). Cultures were incubated at 37 °C in a humidified atmosphere of 5% CO2 for 14 days. Colony numbers were counted using a microscope (Zeiss AxioCam 702).
Analysis of FLT3-ITD mutation
Detection of the FLT3-ITD mutation was performed using a conventional PCR technique as outlined by Kiyoi et al. in 199755. Specific primers were designed to amplify both the FLT3-ITD mutant and wild-type FLT3 at exons 14 and 15. The forward primer 11 F (5′-GCAATTTAGGTATGAAAGCCAGC-3′) and the reverse primer 12 R (5′-CTTTCAGCATTTTGACGGCAACC-3′) were used to produce a 329 bp wild-type product. The presence of an additional PCR product longer than 329 bp indicated the FLT3-ITD mutant type56. The PCR products were analyzed using 3% agarose gel electrophoresis.
In vivo LSC transplantation assay
Mice were housed in the City of Hope Animal Facility with a 12-h light/12-h dark cycle and temperatures of ~18–23 °C with 40–60% air humidity. Mouse care and experimental procedures were performed in accordance with federal guidelines and protocols approved by the Institutional Animal Care and Use Committee at City of Hope under protocol numbers 18008, 20003, and 24049. Tumor-bearing mice were monitored twice per week and at more frequent intervals depending on the status of the mice. Mice exhibiting evidence of distress, discomfort, pain, lethargy, inability to properly groom, or inability to obtain food and/or water were killed immediately via CO2 inhalation. Tumor-bearing mice with 20% weight loss from the age-matched controls without receiving tumor cell inoculation were killed.
For the human LSC engraftment experiments related to Fig. 4, 0.5 × 104 human LSCs were isolated from the peripheral blood of AML patients and then transplanted via intravenously (i.v.) injection into sublethally irradiated (200 cGy) 6–8-week-old female NOD.Cg-Prkdcscid Il2rgtm1Wjl Tg(CMV-IL3, CSF2, KITLG)1Eav/MloySzJ (NSG-SGM3) mice purchased from the Jackson laboratory. One day later, 5 × 104 human ILC1s isolated from the PBMCs of healthy donors were injected via the tail vein into these mice. A control group had no ILC1 injection. Recombinant human IL-15 (2 µg/mouse) was injected intraperitoneally (i.p.) into recipient mice daily for 7 days. Engraftment of human CD45+CD33+ and CD34+CD38− cells in the peripheral blood of mice was monitored at three weeks by flow cytometry.
For the human LSC engraftment experiments related to Fig. 6h, i, 0.5 × 104 LSCs were injected i.v. into sublethally irradiated (200 cGy) 6–8-week-old female NSG-SGM3 mice on day 0. One day later, 5 × 104 UCB CD34+ cell-derived CD161−ILC1s were i.v. injected into these mice. A control group had no CD161− ILC1 injection. Recombinant human IL-15 (2 µg/mouse) was injected i.p. into recipient mice daily for 7 days.
For the human LSC engraftment experiments related to Fig. 6j, k, 5 × 104 LSCs were i.v. injected into sublethally irradiated (200 cGy) 6–8-week-old female NSG-SGM3 mice on day 0. One day later, mice were randomized to receive (1) no ILC1 or NK cell infusion; (2) 1.5 × 105 UCB CD34+ cell-derived CD161− ILC1s; (3) 1.5 × 105 UCB CD34+ cell-derived NK cells; or (4) 1.5 × 105 UCB CD34+ cell-derived CD161− ILC1s plus anti-IFNγ (500 μg given intraperitoneally every 2 days for 8 days). Recombinant human IL-15 (2 µg/mouse) was injected i.p. into recipient mice daily for 7 days.
In vitro ILC1 induction from UCB CD34+ HSCs
CD34+ cells from UCBs were sorted by FACS and were plated onto approximately 80–90% confluent OP9-DL1 or OP9-DL4 stromal cell monolayers in MEMα GlutaMAX media (Thermo Fisher Scientific). Media was supplemented with 10% human AB serum, and SCF (20 ng/ml), Flt3-L (20 ng/ml), IL-7 (20 ng/ml), IL-15 (20 ng/ml), and IL-3 (5 ng/ml) in the first week. One week later, IL-3 was removed from the media. Two weeks later, Flt3-L was reduced to 5 ng/ml. Media and cytokines were refreshed every 3-4 days by replacing half of the media containing 1× concentration of cytokines. At the end of weeks 1, 2, and 3, cells were replated onto fresh OP9-DL1 or OP9-DL4 in six-well plates.
In vitro cytotoxicity assay
NK cells were first enriched from PBMCs using the RosetteSep Human NK Cell Enrichment Kit to remove non-NK cells and red blood cells. Next, total ILCs were enriched to remove non-ILCs using the EasySep™ Human Pan-ILC Enrichment Kit. The CD161+ ILC1s and CD161 − ILC1s were sorted from the enriched pan-ILCs to be cocultured with HSCs sorted from HD blood and LSCs from the blood of AML patients at various ratios.
Statistics & reproducibility
For continuous endpoints, Student’s t test was used to compare two independent conditions, and one-way ANOVA models were used to compare three or more independent conditions. Paired t test or one-way ANOVA models with repeated measures were used to compare two or more donor-matched groups. For survival data, survival functions were estimated by the Kaplan–Meier method and compared by log-rank tests. All tests were two-sided. P values were adjusted for multiple comparisons by Holm’s procedure. No statistical methods were used to predetermine sample sizes, but our sample sizes are similar to those reported in previous publications33. No data were excluded from the analyses. Experimenters were blinded to observe the survival of mice. Otherwise, blinding was not performed, such as during in vitro experiments, where experimenters were required to know the conditions of each well. Data distribution was assumed to be normal, but this was not formally tested. Data are presented as mean ± s.d. Prism software v.10 (GraphPad, CA, USA), and SAS v.9.4 (SAS Institute, NC, USA) were used to perform statistical analyses.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data are provided with this paper. An uncropped version of the gel is found in the Source Data file. The single-cell data used in this study are accessible in GEO under accession code GSE247205. Source data are provided with this paper.
References
Kantarjian, H. et al. Acute myeloid leukemia: current progress and future directions. Blood Cancer J. 11, 41 (2021).
Lapidot, T. et al. A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature 367, 645–648 (1994).
Bonnet, D. & Dick, J. E. Human acute myeloid leukemia is organized as a hierarchy that originates from a primitive hematopoietic cell. Nat. Med 3, 730–737 (1997).
Ng, S. W. et al. A 17-gene stemness score for rapid determination of risk in acute leukaemia. Nature 540, 433–437 (2016).
Taussig, D. C. et al. Leukemia-initiating cells from some acute myeloid leukemia patients with mutated nucleophosmin reside in the CD34(-) fraction. Blood 115, 1976–1984 (2010).
Sarry, J. E. et al. Human acute myelogenous leukemia stem cells are rare and heterogeneous when assayed in NOD/SCID/IL2Rγc-deficient mice. J. Clin. Invest 121, 384–395 (2011).
Hope, K. J., Jin, L. & Dick, J. E. Acute myeloid leukemia originates from a hierarchy of leukemic stem cell classes that differ in self-renewal capacity. Nat. Immunol. 5, 738–743 (2004).
Goardon, N. et al. Coexistence of LMPP-like and GMP-like leukemia stem cells in acute myeloid leukemia. Cancer Cell 19, 138–152 (2011).
Quek, L. et al. Genetically distinct leukemic stem cells in human CD34- acute myeloid leukemia are arrested at a hemopoietic precursor-like stage. J. Exp. Med 213, 1513–1535 (2016).
Ishikawa, F. et al. Chemotherapy-resistant human AML stem cells home to and engraft within the bone-marrow endosteal region. Nat. Biotechnol. 25, 1315–1321 (2007).
Kansler, E. R. et al. Cytotoxic innate lymphoid cells sense cancer cell-expressed interleukin-15 to suppress human and murine malignancies. Nat. Immunol. 23, 904–915 (2022).
Weizman, O. E. et al. ILC1 Confer Early Host Protection at Initial Sites of Viral Infection. Cell 171, 795–808.e712 (2017).
Nabekura, T., Riggan, L., Hildreth, A. D., O’Sullivan, T. E. & Shibuya, A. Type 1 Innate Lymphoid Cells Protect Mice from Acute Liver Injury via Interferon-γ Secretion for Upregulating Bcl-xL Expression in Hepatocytes. Immunity 52, 96–108.e109 (2020).
Tsou, A. M. et al. Neuropeptide regulation of non-redundant ILC2 responses at barrier surfaces. Nature 611, 787–793 (2022).
Jarick, K. J. et al. Non-redundant functions of group 2 innate lymphoid cells. Nature 611, 794–800 (2022).
Schneider, C. et al. A Metabolite-Triggered Tuft Cell-ILC2 Circuit Drives Small Intestinal Remodeling. Cell 174, 271–284.e214 (2018).
Serafini, N. et al. Trained ILC3 responses promote intestinal defense. Science 375, 859–863 (2022).
Lyu, M. et al. ILC3s select microbiota-specific regulatory T cells to establish tolerance in the gut. Nature 610, 744–751 (2022).
Horn, V. & Sonnenberg, G. F. Group 3 innate lymphoid cells in intestinal health and disease. Nat. Rev. Gastroenterol. Hepatol. https://doi.org/10.1038/s41575-024-00906-3 (2024).
Spits, H. & Mjösberg, J. Heterogeneity of type 2 innate lymphoid cells. Nat. Rev. Immunol. 22, 701–712 (2022).
Jacquelot, N., Seillet, C., Vivier, E. & Belz, G. T. Innate lymphoid cells and cancer. Nat. Immunol. 23, 371–379 (2022).
Li, Z. et al. Therapeutic application of human type 2 innate lymphoid cells via induction of granzyme B-mediated tumor cell death. Cell 187, 624–641.e623 (2024).
Diefenbach, A., Colonna, M. & Koyasu, S. Development, differentiation, and diversity of innate lymphoid cells. Immunity 41, 354–365 (2014).
Klose, C. S. & Artis, D. Innate lymphoid cells as regulators of immunity, inflammation and tissue homeostasis. Nat. Immunol. 17, 765–774 (2016).
Spits, H. & Di Santo, J. P. The expanding family of innate lymphoid cells: regulators and effectors of immunity and tissue remodeling. Nat. Immunol. 12, 21–27 (2011).
Li, Z. et al. Purinergic receptor P2Y(6) is a negative regulator of nk cell maturation and function. J. Immunol. 207, 1555–1565 (2021).
Moro, K. et al. Innate production of T(H)2 cytokines by adipose tissue-associated c-Kit(+)Sca-1(+) lymphoid cells. Nature 463, 540–544 (2010).
Neill, D. R. et al. Nuocytes represent a new innate effector leukocyte that mediates type-2 immunity. Nature 464, 1367–1370 (2010).
Sonnenberg, G. F. & Artis, D. Innate lymphoid cell interactions with microbiota: implications for intestinal health and disease. Immunity 37, 601–610 (2012).
Abramson, J., Dobeš, J., Lyu, M. & Sonnenberg, G. F. The emerging family of RORγt(+) antigen-presenting cells. Nat. Rev. Immunol. 24, 64–77 (2024).
Goc, J. et al. Dysregulation of ILC3s unleashes progression and immunotherapy resistance in colon cancer. Cell 184, 5015–5030.e5016 (2021).
Monticelli, L. A. et al. Innate lymphoid cells promote lung-tissue homeostasis after infection with influenza virus. Nat. Immunol. 12, 1045–1054 (2011).
Li, Z. et al. ILC1s control leukemia stem cell fate and limit development of AML. Nat. Immunol. 23, 718–730 (2022).
Mazzurana, L. et al. Tissue-specific transcriptional imprinting and heterogeneity in human innate lymphoid cells revealed by full-length single-cell RNA-sequencing. Cell Res. 31, 554–568 (2021).
Trabanelli, S. et al. CD127+ innate lymphoid cells are dysregulated in treatment naive acute myeloid leukemia patients at diagnosis. Haematologica 100, e257–e260 (2015).
Falquet, M. et al. Dynamic single-cell regulomes characterize human peripheral blood innate lymphoid cell subpopulations. iScience 26, 107728 (2023).
Jaeger, N. et al. Diversity of group 1 innate lymphoid cells in human tissues. Nat. Immunol. 25, 1460–1473 (2024).
Rebuffet, L. et al. High-dimensional single-cell analysis of human natural killer cell heterogeneity. Nat. Immunol. 25, 1474–1488 (2024).
Tang, F. et al. A pan-cancer single-cell panorama of human natural killer cells. Cell 186, 4235–4251.e4220 (2023).
Al-Matary, Y. S. et al. Acute myeloid leukemia cells polarize macrophages towards a leukemia supporting state in a Growth factor independence 1 dependent manner. Haematologica 101, 1216–1227 (2016).
Mussai, F. et al. Acute myeloid leukemia creates an arginase-dependent immunosuppressive microenvironment. Blood 122, 749–758 (2013).
Miari, K. E., Guzman, M. L., Wheadon, H. & Williams, M. T. S. Macrophages in acute myeloid leukaemia: significant players in therapy resistance and patient outcomes. Front Cell Dev. Biol. 9, 692800 (2021).
Kiyoi, H. et al. Prognostic implication of FLT3 and N-RAS gene mutations in acute myeloid leukemia. Blood 93, 3074–3080 (1999).
Whitman, S. P. et al. Absence of the wild-type allele predicts poor prognosis in adult de novo acute myeloid leukemia with normal cytogenetics and the internal tandem duplication of FLT3: a cancer and leukemia group B study. Cancer Res 61, 7233–7239 (2001).
Simoni, Y. et al. Human innate lymphoid cell subsets possess tissue-type based heterogeneity in phenotype and frequency. Immunity 46, 148–161 (2017).
Hernández, D. C. et al. An in vitro platform supports generation of human innate lymphoid cells from CD34(+) hematopoietic progenitors that recapitulate ex vivo identity. Immunity 54, 2417–2432.e2415 (2021).
Vetrie, D., Helgason, G. V. & Copland, M. The leukaemia stem cell: similarities, differences and clinical prospects in CML and AML. Nat. Rev. Cancer 20, 158–173 (2020).
Stelmach, P. & Trumpp, A. Leukemic stem cells and therapy resistance in acute myeloid leukemia. Haematologica 108, 353–366 (2023).
Thomas, D. & Majeti, R. Biology and relevance of human acute myeloid leukemia stem cells. Blood 129, 1577–1585 (2017).
Weinhäuser, I. et al. M2 macrophages drive leukemic transformation by imposing resistance to phagocytosis and improving mitochondrial metabolism. Sci. Adv. 9, eadf8522 (2023).
Bernink, J. H., Mjösberg, J. & Spits, H. Human ILC1: To Be or Not to Be. Immunity 46, 756–757 (2017).
Bernink, J. H. et al. Human type 1 innate lymphoid cells accumulate in inflamed mucosal tissues. Nat. Immunol. 14, 221–229 (2013).
Shah, Z. et al. Human anti-PSCA CAR macrophages possess potent antitumor activity against pancreatic cancer. Cell Stem Cell 31, 803–817.e806 (2024).
Li, Y., Hermanson, D. L., Moriarity, B. S. & Kaufman, D. S. Human iPSC-Derived Natural Killer Cells Engineered with Chimeric Antigen Receptors Enhance Anti-tumor Activity. Cell Stem Cell 23, 181–192.e185 (2018).
Kiyoi, H. et al. Internal tandem duplication of FLT3 associated with leukocytosis in acute promyelocytic leukemia. Leukemia Study Group of the Ministry of Health and Welfare (Kohseisho). Leukemia 11, 1447–1452 (1997).
Ei Ei Aung, N., Yamsri, S., Teawtrakul, N., Kamsaen, P. & Fucharoen, S. FLT3 Gene Mutations in Acute Myeloid Leukemia Patients in Northeast Thailand. Med Sci. Monit. Basic Res 28, e937446 (2022).
Acknowledgements
This work was supported by grants from the National Institutes of Health (CA210087, CA265095, and CA163205 to M.A.C.; NS106170, AI129582, CA247550, CA264512, CA266457, and CA223400 to J.Y.) and the Leukemia and Lymphoma Society (1364-19 to J.Y.). Images were created with BioRender.com and Adobe Photoshop.
Author information
Authors and Affiliations
Contributions
Conception and design: Z.L., J.Y., and M.A.C. Development of methodology: Z.L., R.M., and N.L. Acquisition of data (provided animals, acquired and managed patients, provided facilities, etc.): Z.L., R.M., N.L., H.T., and G.M. Analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis): Z.L. and J.Z. Writing, review, and/or revision of the manuscript: Z.L., R.M., N.L., J.Y., and M.A.C. Administrative, technical, or material support (i.e., reporting or organizing data, constructing databases): J.Z. Study supervision and funding acquisition: J.Y. and M.A.C. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Dong-Yeop Shin, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Z., Ma, R., Tang, H. et al. Human type-1 innate lymphoid cells control leukemia stem cell differentiation and limit acute myeloid leukemia development. Nat Commun 17, 2377 (2026). https://doi.org/10.1038/s41467-026-68582-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68582-2