Abstract
The transition between motility-associated acute infection and biofilm-associated chronic infection is crucial for bacterial proliferation and survival within hosts. This process of transition can be influenced by various host and environmental cues, however, whether bacteria could actively modulate the switch remains poorly understood. Here we identify a novel enoyl-CoA hydratase/isomerase, PA0744 (designated as DspII), which is co-upregulated with the previously characterized enoyl-CoA hydratase/isomerase DspI in a cell density-dependent manner to induce chronic-to-acute virulence switch in Pseudomonas aeruginosa. The two proteins form a heterocomplex that synthesizes the DSF-family quorum sensing signal cis−2-decenoic acid (CDA), which positively regulates the expression of the phosphodiesterase RbdA, and subsequently, activates biofilm dispersion and swarming motility through the c-di-GMP/FleQ pathway. Intriguingly, the DspII-DspI complex also synergistically antagonizes the GacA-upregulated sRNAs, RsmY and RsmZ, leading to overproduction of type III secretion system (T3SS) and augment of acute pathogenesis. Notably, the affinity of DspII-DspI interaction and consequently T3SS gene expression are fine-tuned by CDA levels. This study unveils the novel enzymatic role of the DspII-DspI complex in CDA biosynthesis as well as its non-enzymatic role in T3SS gene regulation, which presents a simple yet sophisticated bacterial signal-driven auto-regulatory mechanism that governs the chronic-to-acute virulence switch in P. aeruginosa.
Similar content being viewed by others
Introduction
Bacterial pathogenesis progresses through multiple stages, which initiate with colonization at host infection sites by planktonic, hypervirulent pathogenic cells, resulting in the early stage of acute infection1. This early stage is characterized by the expression of acute virulence factors such as the type III secretion system (T3SS) that facilitate host invasion and immune evasion2,3. However, upon encountering unfavorable conditions or certain environmental cues, such as immune response or other bio-stresses, the pathogens may reprogram their virulence either mounting attack by boosting virulence factor production4, or retracting by switching from acute to chronic infection5. The transition from acute to chronic infection is marked by the attenuation of motility (e.g., flagellar biosynthesis) and the activation of biofilm-associated factors (e.g., exopolysaccharides (EPSs)), leading to the formation of a sessile, matrix-encased community known as biofilm6. Within this protective biofilm structure, bacteria establish persistent infections that are extraordinary resistant to both host immune defenses and antimicrobial treatments7,8. Notably, this is a reversible process and bacterial cells within biofilms are common source of new infections, as they could also be activated under certain environmental conditions to switch back from chronic persistence to acute infection mode9.
Pseudomonas aeruginosa, a versatile opportunistic pathogen capable of causing serious acute and chronic infections in humans10, serves as an exemplary model for studying the environmental cues and mechanisms that influence the virulence switch. Diverse host and environmental cues such as oxygen levels, nutrient availability, metal ion concentrations, antibiotics, and other factors11,12,13,14,15, could influence the transition between acute and chronic infections. Central to this switch is the second messenger cyclic di-GMP (c-di-GMP). Elevated intracellular c-di-GMP content promotes biofilm formation, shifting the planktonic-associated acute infection to biofilm-associated chronic persistence, whereas depleted intracellular c-di-GMP level promotes biofilm dispersal, switching the pathogen from chronic persistence to acute infection mode16.
cis-2-decenoic acid (CDA), a self-produced fatty acid-based quorum sensing (QS) signaling molecule belonging to the diffusible signal factor (DSF) family, has been demonstrated to induce biofilm dispersion and reactivate motility in P. aeruginosa, thereby promoting the chronic-to-acute virulence switch17. The enoyl-CoA hydratase/isomerase DspI is essential for CDA biosynthesis, and recently, it was reported that CDA is recognized by the sensor/response regulator hybrid DspS and then induces the chronic-to-acute virulence switch by decreasing the intracellular c-di-GMP content9,18,19. Despite these advances, it remains largely unclear how CDA is synthesized and whether DspI is the sole synthase for CDA. In addition, detailed molecular mechanisms underlying the regulation of virulence switch by CDA are largely unknown. In this study, we identified a novel enoyl-CoA hydratase/isomerase, PA0744 (designated as DspII), which complexes with DspI for CDA biosynthesis. Intriguingly, we further uncovered a non-enzymatic role of the DspII-DspI complex in regulating T3SS gene expression by directly binding to a response regulator GacA and the promoter of a small RNA RsmZ. More interestingly, formation of the DspII-DspI complex and interaction between the complex and DNA promoter are auto-modulated by CDA. These findings unveil the novel dual-functional roles of the DspII-DspI complex in modulating chronic-to-acute virulence switch and the sophisticated regulatory network.
Results
A novel enoyl-CoA hydratase/isomerase PA0744 is potentially involved in chronic-to-acute virulence switch
Systematic genomic analysis of P. aeruginosa PAO1 revealed 12 genes encoding probable enoyl-CoA hydratase/isomerases (Supplementary Fig. 1A). Except PA1748 which did not show significant similarity with the previously characterized enoyl-CoA hydratase/isomerase DspI (PA0745), the other 10 uncharacterized ones share about 26% ~ 40% similarity with DspI (Supplementary Fig. 1A and 1B). Given the established role of DspI in CDA biosynthesis20,21, we hypothesized potential functional redundancy among these proteins in synthesizing CDA and regulating biofilm dispersion. We generated 12 mutants with single gene deletion and determined their function in biofilm dispersion by comparing the production of biofilm with the PAO1 wild-type (WT) strain. In addition to DspI, we found that the loss of another enoyl-CoA hydratase/isomerase, PA0744, led to a substantial accumulation of biofilm production (Supplementary Fig. 1C), suggesting that PA0744 is also essential for biofilm dispersion and potentially required for the chronic-to-acute virulence switch.
PA0744 is encoded by the gene immediately downstream of dspI (PA0745), sharing about 27.85% similarity with DspI at the peptide sequence level (Supplementary Fig. 1A and 1B), and notably, they constitute a contiguous cluster with three metabolic genes PA0743, PA0746, and PA0747, which encodes probable 3-hydroxyisobutyrate dehydrogenase, probable acyl-CoA dehydrogenase, and probable aldehyde dehydrogenase, respectively (Figs. 1A, B). In addition, an adjacent gene PA0748 is predicted to encode a probable local transcriptional regulator for the gene cluster of PA0743-PA0747 (Fig. 1A, B). To verify if PA0743-PA0747 genes are transcribed together, we performed reverse transcriptase PCR (RT-PCR) and confirmed polycistronic transcription of PA0743-PA0747 (Fig. 1C). Given the close genetic relationship of these genes, we were curious whether PA0743, PA0746, PA0747, and PA0748 contributed to the chronic-to-acute virulence switch as well. To answer this, we knocked out each of these genes and compared their production of biofilm and ability of swarming compared to the WT strain. Unexpectedly, only the mutants ΔPA0744 and ΔdspI displayed increased biofilm accumulation and impaired swarming motility, while deletions of adjacent genes PA0743, PA0746, PA0747, and PA0748 did not alter biofilm or swarming profiles (Fig. 1D, E), indicating functional specificity of PA0744 and dspI in modulating the chronic-to-acute virulence switch within the operon. Furthermore, the double-deletion mutant ΔPA0744ΔdspI showed similar levels of biofilm production and swarming motility with the single-deletion mutants ΔPA0744 and ΔdspI, respectively, and complementation of the deleted gene(s) fully restored biofilm dispersion and swarming motility in all the mutants ΔPA0744, ΔdspI, and ΔPA0744ΔdspI (Fig. 1F, G), suggesting that PA0744 and DspI probably function in the same pathway.
A A diagram showing the gene cluster from PA0743 to PA0747 as well as the gene PA0748 encoding a probable transcriptional regulator. Blue arrows indicate the location of primers used in (C). B Functional summary of genes shown in (A). C RT-PCR analysis using the primer pairs F1/R, F2/R, F3/R and F4/R shown in (A) to detect the transcription pattern of the gene cluster PA0743 to PA0747. gDNA (left panel) and cDNA (right panel) were used as templates. Representative gel image shown of 2 independent experiments. D, E Biofilm biomass (D) and swarming motility (E) were measured in PAO1 WT, ΔPA0743, ΔPA0744, ΔPA0745 (ΔdspI), ΔPA0746, ΔPA0747 and ΔPA0748 strains. F, G Biofilm biomass (F) and swarming motility (G) were measured in the PAO1 WT, ΔPA0744, ΔdspI, ΔPA0744ΔdspI, and their gene complemented strains. EV represents empty vector. For panels D to G: data are presented as mean values ± SEM of n = 3 biological replicates. Statistical analysis was conducted based on the one-way analysis of variance (ANOVA) with Dunnett’s multiple comparisons test compared to the WT group (D, E) or WT(EV) group (F, G). ***, p < 0.0001; n.s., no significance. For panel D the p value was calculated as (from left to right): 0.9998, < 0.0001, < 0.0001, 0.7449, 0.9502, 0.9999; For panel E the p value was calculated as (from left to right): > 0.9999, < 0.0001, < 0.0001, 0.9446, 0.9997, 0.4293; For panel F the p value was calculated as (from left to right): < 0.0001, 0.9996, < 0.0001, 0.9998, < 0.0001, 0.9999; For panel G the p value was calculated as (from left to right): < 0.0001, 0.9997, < 0.0001, 0.9557, < 0.0001, 0.4368. Source data are provided as a Source Data file.
PA0744 (DspII), co-expressed with DspI in a cell density-dependent manner, is essential for CDA biosynthesis
Given that CDA is a QS signal, we investigated whether PA0744 and dspI are expressed in a cell density-dependent manner. First, reverse transcription-quantitative PCR (RT-qPCR) assay showed that the expression of PA0744 and dspI was reciprocally regulated, i.e., loss of one almost fully inactivated the expression of the other gene (Fig. 2A). Moreover, promoter activity assay using a lacZ fusion showed that both PA0744 and DspI were essential for the expression of the whole PA0743-PA0747 operon (Fig. 2B). Time-course RT-qPCR assay during the growth of PAO1 revealed the synchronous upregulation of PA0744 and dspI along with the increase of cell density (Fig. 2C), confirming the cell density-dependent co-expression of PA0744 and dspI.
A Relative expression of PA0744 and dspI was measured by RT-qPCR in the PAO1 WT(EV), ΔPA0744(EV), ΔdspI(EV), ΔPA0744(PA0744), and ΔdspI(dspI) strains. N.D., not detected. The 50S ribosomal protein gene rplU was used as internal control. B β-galactosidase activity of the PPA0747-lacZ transcriptional fusions was measured in the PAO1 WT(EV), ΔPA0744(EV), ΔdspI(EV), ΔPA0744(PA0744), and ΔdspI(dspI) strains. C Relative expression of PA0744 and dspI in the PAO1 WT strain was measured by RT-qPCR at different growth stages with different OD600 values of the cell culture compared to the samples collected at OD600 of 0.3. (D) Biofilm biomass was measured in PAO1 WT, ΔPA0744, and the mixed cultures containing ΔPA0744 and WT in different ratios (1:1, 1:2, 1:3). E CDA production in the PAO1 WT(EV), ΔPA0744(EV), ΔPA0744(PA0744), ΔdspI(EV), and ΔdspI(dspI) strains was measured by LC-MS. N.D., not detected. F Biofilm biomass was measured in the PAO1 WT(EV), ΔPA0744(EV), and ΔPA0744(PA0744) strains when the strains were cultured with exogenous addition of CDA at concentrations of 1 nM, 50 nM, 500 nM, 5 μM, and 50 μM. G LC-MS quantification of CDA in the PAO1 WT, ΔdspII, ΔdspI, ΔdspIIΔdspI strains and strains with complemented expression of dspII or/and dspI. H, I Biofilm biomass (H) and swarming motility (I) were measured in the PAO1 WT, ΔdspII, ΔdspI, ΔdspIIΔdspI strains and the strains with complemented expression of dspII or/and dspI. J Domain structures of DspI and DspII. The conserved enzymatic residues of enoyl-CoA hydratase/isomerases (Glu126 and Glu146 in DspI and Glu143 in DspII) were labeled. K CDA production in ΔdspII and ΔdspI mutants with the complementation of their enzymatically dead variants. L, M Biofilm biomass (L) and swarming motility (M) were measured in ΔdspII and ΔdspI mutants with the complementation of their enzymatically dead variants. For panels A to I and K to M: data are presented as mean values ± SEM of n = 3 biological replicates. Statistical analysis was performed using two-way ANOVA with Tukey’s multiple comparisons test (A, F), one-way ANOVA with Tukey’s multiple comparisons test (B, D), or one-way ANOVA with Dunnett’s multiple comparisons test compared to the WT(EV) group (E, G, H, I) or the indicated groups (L, M). ***, p < 0.0001. Source data are provided as a Source Data file.
Owing to the same enzymatic annotation and the physiological function of PA0744 and DspI, we speculated that PA0744 participates in CDA biosynthesis. As expected, accumulation of biofilm in the ΔPA0744 strain was prevented with the co-culturing of the WT strain (Fig. 2D), implying the importance of PA0744 in producing a diffusible signal for cell-to-cell communication. The signal was subsequently confirmed as CDA because CDA was no longer detected by LC-MS in the mutant ΔPA0744 and exogenous CDA restored biofilm dispersion in ΔPA0744 (Fig. 2E, F). Considering the same function of PA0744 as DspI in CDA biosynthesis and biofilm dispersion, we renamed PA0744 as DspII (Dispersion inducer II) hereafter.
Because deletion of dspII and dspI respectively resulted in almost complete repression of the other gene (Fig. 2A), we next expressed DspI in the ΔdspII mutant and expressed DspII in the ΔdspI mutant to ensure that DspI and DspII were normally produced in ΔdspII and ΔdspI, respectively, to further clarify the contribution of DspII and DspI to CDA biosynthesis. However, CDA biosynthesis was not restored in both reconstructed strains (Fig. 2G). To further dissect the functional interdependence, we expressed DspII, DspI, and DspII-DspI in the mutant ΔdspIIΔdspI which serves as a pure background without any production of either DspII or DspI. As shown in Fig. 2G, expression of DspII or DspI alone in the double mutant ΔdspIIΔdspI was incapable of CDA biosynthesis, whereas the presence of both DspII and DspI restored CDA biosynthesis in ΔdspIIΔdspI. Consistently, biofilm dispersion and swarming motility were re-activated by the co-presence of DspII and DspI in ΔdspIIΔdspI (Fig. 2H, I). These data indicate that DspII and DspI function in the same pathway or as a complex for CDA biosynthesis and the chronic-to-acute virulence switch.
Domain architecture analysis using InterPro22 showed that DspII contains an N-terminal ClpP/crotonase domain and an additional C-terminal ClpP/crotonase domain while DspI contains only one (Fig. 2J). Conserved glutamate residues for the activity of enoyl-CoA hydratase/isomerases were predicted in both DspII and DspI. DspI was predicted to contain two active glutamate residues Glu126 and Glu146, while DspII has one active glutamate residue Glu143 (Fig. 2J). To verify the enzymatic activity of these residues, we generated DspII and DspI variants, DspIIE143A, DspIE126A, and DspIE146A, and found that these variants were incapable of recovering CDA biosynthesis in ΔdspII and ΔdspI mutants (Fig. 2K). Moreover, biofilm dispersion and swarming motility in the ΔdspII and ΔdspI mutants cannot be re-activated with the complementation of their enzymatically dead variants (Fig. 2L, M), indicating that these glutamate residues are essential for the enzymatic activity of DspII and DspI to synthesize CDA and regulate the chronic-to-acute virulence switch.
CDA induces biofilm dispersion and swarming motility by inversely regulating Psl and flagellar biosynthesis
We moved to explore detailed mechanisms about how CDA could induce the chronic-to-acute virulence switch by shifting the sessile lifestyle to the motile lifestyle. Given that Psl and Pel are two important EPSs stabilizing the biofilm structure in non-mucoid P. aeruginosa strains23, we evaluated the biosynthesis of Psl and Pel in mutants ΔdspII and ΔdspI by measuring the promoter activity of their synthetic operons with fluorescent reporters. We found that activities of the psl promoter in both ΔdspII and ΔdspI mutants were substantially higher than the WT strain, particularly at the late growth stages with high cell density (Fig. 3A). In contrast, the pel promoter in both ΔdspII and ΔdspI mutants showed a decreased activity compared to the WT strain (Fig. 3B). Consistently, RT-qPCR assay showed significantly increased expression levels of the pslA gene in both ΔdspII and ΔdspI mutants compared to the WT strain (Fig. 3C). These results suggested that the loss of DspII or DspI leads to the continuous biosynthesis of Psl, the primary EPS used to develop biofilm and maintain its structure in PAO124, which prevents biofilm dispersion. To confirm this, we deleted pslA to disrupt Psl biosynthesis in the ΔdspII and ΔdspI mutants, which consequently abolished the accumulation of biofilm (Fig. 3D). In contrast, biofilm still accumulated in ΔdspII and ΔdspI mutants which were unable to synthesize Pel (Fig. 3D). Regarding to swarming motility, RT-qPCR assay showed that the flagellar synthetic genes flgB and flgF were significantly repressed in the ΔdspII and ΔdspI mutants (Fig. 3E), meaning that DspII and DspI activate swarming motility by upregulating flagellar biosynthesis. We thus compared the flagellar morphology of ΔdspII, ΔdspI, and WT strains under a transmission electron microscope (TEM), which showed that the average flagellar lengths of the ΔdspI and ΔdspII cells were significantly shorter than the WT cells (Figs. 3F, G). Collectively, these data demonstrated that CDA induces biofilm dispersion and swarming motility by negatively regulating Psl biosynthesis and positively regulating flagellar biosynthesis.
A, B Fluorescence intensities of the PpslA-gfp (A) and PpelA-gfp (B) transcriptional fusions were measured at different time points during growth in the PAO1 WT, ΔdspII, and ΔdspI strains. C Relative expression of pslA was determined by RT-qPCR in PAO1 WT, ΔdspII, and ΔdspI strains. D Biofilm biomass was measured in the PAO1 WT, ΔdspII, and ΔdspI strains as well as the strains with additional deletion of pslA or pelA and complementation of pslA. E Relative expression of flgB and flgF was determined by RT-qPCR in PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔdspII(dspII), and ΔdspI(dspI) strains. F Representative images showed the flagellar length of PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔdspII(dspII), and ΔdspI(dspI) strains. Bacterial cells were observed under TEM and flagellar length was measured by Image J. Representative images shown of 10 biological replicates. G Flagellar length was measured for the PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔdspII(dspII), and ΔdspI(dspI) strains. For panels A to E: data are presented as mean values ± SEM of n = 3 biological replicates. For panel G: data are presented as means ± SEM of n = 10 biological replicates. Statistical analysis was performed using one-way ANOVA with Dunnett’s multiple comparisons test compared to the WT group (C) and WT(EV) group (G), one-way ANOVA with Tukey’s multiple comparisons test (D), or two-way ANOVA with Dunnett’s multiple comparisons test compared to the WT(EV) group (E). Source data are provided as a Source Data file.
CDA induces biofilm dispersion and swarming motility through the c-di-GMP/FleQ pathway
The second messenger c-di-GMP has been identified as a critical intracellular molecule that orchestrates acute-to-chronic virulence switch when its content elevates in response to specific external cues25. We first evaluated c-di-GMP contents in ΔdspII, ΔdspI, and WT strains by measuring the expression of a c-di-GMP-responsive indicator CdrA26, which suggested that c-di-GMP contents in two mutants were dramatically higher than that in the WT strain (Fig. 4A). LC-MS quantification confirmed that c-di-GMP contents were increased in the ΔdspII and ΔdspI mutants (Fig. 4B). c-di-GMP is synthesized by diguanylate cyclase (DGC) and degraded by phosphodiesterase (PDE)27. To validate if reduced abilities of biofilm dispersion and swarming in ΔdspII and ΔdspI mutants were ascribed to increased c-di-GMP levels, we expressed an exogenous PDE W909_14950 (14950), which is from Dickeya oryzae EC1 and has been well-characterized and demonstrated to function efficiently in P. aeruginosa to lower c-di-GMP levels without nonspecific effects28, in both mutants and found that biofilm dispersion and swarming motility were recovered (Figs. 4C, D).
A Relative expression of cdrA was measured by RT-qPCR in PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔdspII(dspII), and ΔdspI(dspI) strains. B c-di-GMP content in the PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔdspII(dspII), and ΔdspI(dspI) strains was measured by LC-MS. C, D Biofilm biomass (C) and swarming motility (D) were measured in the PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔdspII(14950) and ΔdspI(14950) strains. E Fluorescence intensity of the PrbdA-gfp transcriptional fusion was measured at different time points during growth in the PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔdspII(dspII), and ΔdspI(dspI) strains. F Relative expression of rbdA was measured by RT-qPCR in PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔdspII(dspII), and ΔdspI(dspI) strains. G, H Biofilm biomass (G) and swarming motility (H) were measured in the PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔdspII(rbdA), and ΔdspI(rbdA) strains. I A diagram showing the domains and the phosphodiesterase activity of RbdA. RbdA contains both GGDEF and EAL domains as well as two accessory domains, PAS and PAC. J, K Biofilm biomass (J) and swarming motility (K) were measured in the PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔdspII(rbdAΔEAL), and ΔdspI(rbdAΔEAL) strains. L Fluorescence intensity of the PpslA-gfp transcriptional fusion was measured at different time points during growth in PAO1 WT, ΔfleQ strains with the treatment of CDA. M Fluorescence intensity of the PpslA-gfp transcriptional fusion was measured at different time points during growth in PAO1 WT, ΔdspII, ΔdspI strains and strains with additional deletion of fleQ. N, O Biofilm biomass (N) and swarming motility (O) were measured in the PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔdspII(rbdA), and ΔdspI(rbdA), strains and the strains with additional deletion of fleQ. For panels A–H, J–O: data are presented as mean values ± SEM of n = 3 biological replicates. Statistical analysis was performed using one-way ANOVA with Tukey’s multiple comparisons test. ***, p < 0.0001. Source data are provided as a Source Data file.
There are at least 42 DGCs and PDEs in P. aeruginosa29. We next moved to further explore how c-di-GMP content was increased in ΔdspII and ΔdspI mutants. We constructed promoter-gfp fusions for all the DGC and PDE genes and compared their expression in the ΔdspI and WT strains. The results identified 2 putative DGCs (PA0290 and PA2072) and 2 PDEs (PipA and RbdA) with more than 20% higher or lower expression, which might potentially increase the c-di-GMP levels in the ΔdspI mutant (Supplementary Data 1). Upregulation of PA0290 and PA2072 and downregulation of PipA and RbdA were further confirmed in the ΔdspII mutant (Fig. 4E, F; and Supplementary Fig. 2A to 2C). However, deletion of the DGC enzymes PA0290 or PA2072 and overexpression of the PDE enzyme PipA in ΔdspII and ΔdspI mutants did not prevent biofilm accumulation (Supplementary Fig. 2D–F). In contrast, overexpression of RbdA restored biofilm dispersion and swarming motility in the ΔdspII and ΔdspI mutants (Figs. 4G, H). RbdA is a transmembrane PDE which contains multiple domains (Fig. 4I). We further overexpressed RbdAΔEAL, a RbdA variant lacking the EAL domain and the phosphodiesterase activity, in the ΔdspII and ΔdspI mutants. As shown in Fig. 4J, K, the variant did not restore biofilm dispersion and swarming motility of two mutants any longer. Together, these results indicate that CDA decreases c-di-GMP content and induces the chronic-to-acute virulence switch by upregulating the PDE RbdA in P. aeruginosa.
The second messenger c-di-GMP fulfills its physiological functions through interactions with specific proteins named effectors30. At least 12 effectors have been already identified in P. aeruginosa29. To determine which effector connects the chronic-to-acute virulence switch with the CDA-repressed c-di-GMP levels, we deleted 12 candidate effector genes in the WT, ΔdspII, and ΔdspI strains and measured biofilm production. Among 12 candidates, only the deletion of fleQ abolished the accumulation of biofilm in both the ΔdspII and ΔdspI mutants, whereas deletion of other effector genes had no effect on the biofilm production in these two mutants (Supplementary Fig. 3). We treated WT and ΔfleQ strains with CDA and found that CDA inhibited the activity of the psl promoter only in the WT strain but not the ΔfleQ mutant (Fig. 4L). Furthermore, deletion of fleQ abolished the upregulation of Psl synthetic genes and downregulation of flagellar synthetic genes in the ΔdspII and ΔdspI mutants (Fig. 4M and Supplementary Fig. 4). These results suggest the essential role of the c-di-GMP effector FleQ in mediating CDA’s induction on the chronic-to-acute virulence switch. The involvement of the c-di-GMP/FleQ pathway was further confirmed by the overexpression of RbdA in the backgrounds of ΔdspIIΔfleQ and ΔdspIΔfleQ, which no longer restored biofilm dispersion and swarming motility (Fig. 4N, O).
DspII and DspI synergistically upregulate T3SS gene expression and acute pathogenesis
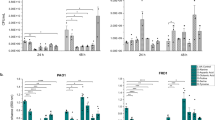
To further validate the chronic-to-acute virulence switch, we evaluated the pathogenicity of ΔdspII and ΔdspI mutants by performing cytotoxicity assay using the A549 cell line and acute infection assay using the model of Galleria mellonella31,32. It was shown that deletion of dspII or dspI led to attenuated cytotoxicity (Fig. 5A) and increased survival of G. mellonella (Fig. 5B and Supplementary Fig. 5), confirming that DspII and DspI not only induce biofilm dispersion and swarming motility but also contribute to acute pathogenesis in P. aeruginosa. Since T3SS is the key virulence factor causing acute infection, we examined T3SS gene expression in the ΔdspII and ΔdspI mutants. RT-qPCR assay showed that the mRNA levels of selected T3SS genes, including exsA, exsC, exoS, and pcrV, were significantly reduced in ΔdspII and ΔdspI mutants compared to the WT strain (Fig. 5C). T3SS gene expression is activated by the master transcription factor ExsA, which is transcribed from two promoters, i.e., PexsCEBA and PexsA, under the control of various transcription factors33 (Fig. 5D). Thus, we focused on ExsA and found that reduced expression of ExsA in both the ΔdspII and ΔdspI mutants was confirmed at the protein level using western blot assay (Fig. 5E). We also measured the activity of promoters, PexcCEBA and PexsA, and found that upregulation of exsA was due to activated transcription of PexsCEBA instead of PexsA by DspII and DspI (Fig. 5F, G). Moreover, we found that DspII and DspI did not influence T3SS gene expression, i.e., activity of PexcCEBA and ExsA production, when ExsA is not or saturated expressed (Fig. 5H, I), which demonstrated that DspII and DspI positively regulate T3SS gene expression through the master activator ExsA.
A Relative cytotoxicity was assessed by monitoring LDH release from the A549 cells which were infected by the strains of PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔdspII(dspII), and ΔdspI(dspI). B Acute pathogenesis of PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔdspII(dspII), and ΔdspI(dspI) strains was measured using the infection model of Galleria mellonella. C Relative expression of exoS, exsA, exsC, and pcrV in the ΔdspII and ΔdspI mutants compared to the PAO1 WT strain was determined by RT-qPCR. D A diagram showing the expression and function of ExsA. ExsA is transcribed from two promoters, PexsCEBA and PexsA, which resulted in polycistronic and monocistronic mRNA, respectively. ExsA positively regulates T3SS genes expression and activates its own transcription through PexsCEBA. E ExsA protein expression in PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔdspII(dspII), and ΔdspI(dspI) strains was measured using western blot assay. The β-RNA polymerase (β-RNAP) was used as a loading control. Representative blots shown of 2 independent experiments. F, G β-galactosidase activities of the PexsA-lacZ (F) and PexsCEBA-lacZ (G) transcriptional fusion were measured in the PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔdspII(dspII), and ΔdspI(dspI) strains. (H) β-galactosidase activity of the PexsCEBA-lacZ transcriptional fusion was measured in the PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔexsA(EV), ΔexsAΔdspII(EV), ΔexsAΔdspII(EV), ΔexsA(exsA), ΔexsAΔdspII(exsA), and ΔexsAΔdspI(exsA) strains. I ExsA protein expression in PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔexsA(EV), ΔexsAΔdspII(EV), ΔexsAΔdspII(EV), ΔexsA(exsA), ΔexsAΔdspII(exsA), and ΔexsAΔdspI(exsA) strains was measured using western blot assay. Representative blots shown of 2 independent experiments. J β-galactosidase activity of the PexsCEBA-lacZ transcriptional fusion was measured in the PAO1 WT, ΔdspII, ΔdspI, ΔdspIIΔdspI strains and strains with complemented expression of dspII or/and dspI. K β-galactosidase activity of the PexsCEBA-lacZ transcriptional fusion was measured in the PAO1 WT, ΔdspII, and ΔdspII(dspII) strains when they were grown with exogenous addition of CDA at a final concentration of 10 μM, 50 μM, and 100 μM. L β-galactosidase activity of the PexsCEBA-lacZ transcriptional fusion was measured in ΔdspII and ΔdspI mutants with the complementation of their enzymatically dead variants. For (A, C, F–H, J–L) data are presented as mean values ± SEM of n = 3 biological replicates. For panel B: data are presented as mean values ± SEM of n = 10 biological replicates. Statistical analysis was performed using Student’s t-test (A), one-way ANOVA with Tukey’s multiple comparisons test (F–H, J, L), and two-way ANOVA with Tukey’s multiple comparisons test (C, K). ***, p < 0.0001. Source data are provided as a Source Data file.
Like biofilm dispersion, we wondered if DspII and DspI regulate T3SS interdependently and whether the regulation is controlled by CDA. We next measured the activity of PexcCEBA in strains including ΔdspII(dspI), ΔdspI(dspII), ΔdspIIΔdspI(dspI), and ΔdspIIΔdspI(dspII), which only produce DspII or DspI. It was shown that the activity of PexcCEBA in these strains remained repressed at the level as that in ΔdspII, ΔdspI, and ΔdspIIΔdspI strains (Fig. 5J). Only the strains producing both DspII and DspI displayed as high activity of PexcCEBA as that in the WT strain (Fig. 5J), meaning that upregulation of T3SS gene expression also requires the cooperation of DspII and DspI. Intriguingly, exogenous supplementation of CDA was incapable of recovering the T3SS gene expression and complementation of enzymatically dead variants DspIIE143A, DspIE126A, and DspIE146A significantly induced T3SS gene expression in both ΔdspII and ΔdspI mutants (Fig. 5K, L), indicating that DspII and DspI might activate T3SS gene expression through a novel CDA-independent mechanism that is different from the regulation of biofilm dispersion and swarming motility. Moreover, it was fascinating to notice that the activity of PexcCEBA in the WT strain was reduced with the increased concentration of exogenous CDA (Fig. 5K), showing an inhibitory effect of CDA on the DspII and DspI-induced T3SS gene expression.
DspII and DspI activate T3SS gene expression by antagonizing the GacA-upregulated small RNAs RsmY and RsmZ
Considering the contribution of c-di-GMP to inhibit T3SS gene expression as we recently found34, we initially hypothesized that the increased c-di-GMP content in ΔdspII and ΔdspI mutants led to the downregulation of T3SS genes. However, decreasing c-di-GMP content by overexpressing PDEs 14950 and RbdA or additional deletion of fleQ did not restore the level of T3SS gene expression in ΔdspII and ΔdspI mutants (Supplementary Fig. 6), meaning that DspII and DspI activate T3SS gene expression bypassing the c-di-GMP/FleQ pathway. Another important pathway modulating T3SS gene expression in P. aeruginosa is the Gac/Rsm pathway. In this pathway, the two-component system GacS/GacA positively regulates the expression of two small RNAs (sRNAs) named RsmY and RsmZ, and subsequently, the sRNAs sequester RsmA which is a posttranscriptional regulator involved in the activation of T3SS gene translation, leading to the reduced translation of T3SS proteins including ExsA35,36 (Fig. 6A). The expression of the response regulator GacA was not changed with the deletion of dspII or dspI (Fig. 6B), while the expression of two sRNAs RsmY and RsmZ was obviously increased (Fig. 6C, D). This result suggests that DspII and DspI might repress RsmY and RsmZ to liberate free RsmA for the upregulation of T3SS genes. We then generated ΔrsmZ, ΔrsmY, and ΔrsmZΔrsmY mutants. It was shown that T3SS gene expression was still reduced by deleting dspII and dspI in the genetic backgrounds of ΔrsmZ or ΔrsmY, while the expression was no longer reduced in the genetic background of ΔrsmZΔrsmY (Fig. 6E, F). Since RsmY and RsmZ repress T3SS gene expression by sequestering RsmA, we next overexpressed RsmA in the ΔdspII and ΔdspI mutants and the results showed that T3SS gene expression was reactivated (Fig. 6G, H). These results together demonstrate that DspII and DspI repress the expression of RsmY and RsmZ, which releases RsmA to elevate T3SS gene expression in P. aeruginosa.
A A diagram showing the regulation of the Gac/Rsm pathway on T3SS gene expression. RsmA, an RNA binding protein, stimulates the translation of T3SS genes including ExsA. Two small RNAs, RsmY and RsmZ, repress the activity of RsmA by sequestering RsmA from target mRNAs. Expression of RsmY and RsmZ is positively regulated by the two-component system GacS/GacA, which is inversely regulated by two additional kinases LadS and RetS. B Relative expression of gacA in the ΔdspII and ΔdspI mutants compared to the PAO1 WT strain was determined by RT-qPCR. C, D Fluorescence intensities of the PrsmY-gfp (C) and PrsmZ-gfp (D) transcriptional fusions were measured in the PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔdspII(dspII), and ΔdspI(dspI) strains. E, F β-galactosidase activity of the PexsCEBA-lacZ transcriptional fusion (E) and ExsA protein expression (F) were measured in the PAO1 WT, ΔdspII, ΔdspI, ΔrsmZ, ΔdspIIΔrsmZ, ΔdspIΔrsmZ, ΔrsmY, ΔdspIIΔrsmY, ΔdspIΔrsmY, ΔrsmZΔrsmY, ΔdspIIΔrsmZΔrsmY, and ΔdspIΔrsmZΔrsmY strains. G, H β-galactosidase activity of the PexsCEBA-lacZ transcriptional fusion (G) and ExsA protein expression (H) were measured in the PAO1 WT(EV), ΔdspII(EV), ΔdspI(EV), ΔdspII(rsmA), and ΔdspI(rsmA) strains. I, J Fluorescence intensities of the PrsmY-gfp (I) and PrsmZ-gfp (J) transcriptional fusions were measured in the PAO1 WT, ΔdspII, ΔdspI, ΔgacA, ΔdspIIΔgacA and ΔdspIΔgacA strains. K, L β-galactosidase activities of the PexsCEBA-lacZ transcriptional fusion (K) and ExsA protein expression (L) were measured in the PAO1 WT, ΔdspII, ΔdspI, ΔgacA, ΔdspIIΔgacA and ΔdspIΔgacA strains. For B–E, G, I–K: data are presented as mean values ± SEM of n = 3 biological replicates. For F, H, L representative blots shown of 2 independent experiments. Statistical analysis was performed using one-way ANOVA with Tukey’s multiple comparisons test (B, G, K) or Dunnett’s multiple comparisons test (E). Source data are provided as a Source Data file.
Because GacA is a critical transcriptional activator of RsmY and RsmZ and its expression was consistent among WT, ΔdspII, and ΔdspI strains, we were curious about the connection of GacA, DspII, and DspI in controlling RsmY and RsmZ expression. Deletion of gacA dramatically reduced the transcription of both RsmY and RsmZ (Fig. 6I, J). In addition, deletion of dspII or dspI did not increase the transcription of RsmY and RsmZ any longer in the ΔgacA background (Fig. 6I, J), meaning that DspII and DspI repress RsmY and RsmZ in a GacA-dependent manner. Consistently, T3SS gene expression was no longer reduced when dspII and dspI were deleted in the ΔgacA background (Fig. 6K, L), which confirmed the GacA-dependent regulation of T3SS gene expression by DspII and DspI.
DspII and DspI antagonize the GacA-activated transcription of RsmZ and RsmY by directly interacting with the RsmZ promoter and the GacA protein
Our findings suggested that DspII and DspI only diminished the transcription of RsmY and RsmZ when GacA is present to first drive their transcription. GacA activates transcription of RsmY and RsmZ by binding at their promoters. Given that DspII and DspI activate T3SS gene expression independently of CDA, we hypothesized that two proteins might be potential roadblocks to impede GacA-induced transcription of RsmY and RsmZ by directly binding at their promoters. We purified DspII and DspI (Supplementary Fig. 7) and performed electrophoretic mobility shift assay (EMSA) to examine the binding potentials of two proteins at the promoters of rsmY and rsmZ. DspII and DspI did not interact with the promoter of rsmY (PrsmY) (Supplementary Fig. 8A and 8B), while it was surprisingly found that both DspII and DspI interacted with the promoter of rsmZ (PrsmZ) (Supplementary Fig. 8A and 8B; Figs. 7A, B), albeit DNA-binding motif was not predicted in both proteins. This result indicates that DspII and DspI repress the transcription of RsmZ by directly binding at its promoter.
A, B EMSA showing the specific binding of DspII (A) and DspI (B) to the rsmZ promoter (PrsmZ). The recA promoter was selected as a negative control. Red triangles indicated the protein-bound probe, blue triangles indicated the free probe. C A schematic diagram showing the sequence of rsmZ promoter and the fragments used for EMSA. D The binding abilities of DspI and DspII to the rsmZ promoter fragments R1 to R5 indicated in (C) were examined by EMSA. E The binding abilities of DspII and DspI to the fragments of PrsmZ−123, 143,163 and 184 indicated in (C) were examined by EMSA. F The binding abilities of DspII and DspI to the rsmZ promoter fragments F1 to F5 indicated in (C) were examined by EMSA. G EMSA detecting the binding affinity of DspII, DspI and their enzymatically dead variants to the rsmZ promoter. Stars in the same color indicate that the proteins were used in the same concentration. (H) Pull-down assay showed the direct interaction between GacA and DspII or DspI proteins. For A, B, D–H representative results shown of 2 independent experiments. Source data are provided as a Source Data file.
To further characterize the binding details of the DspII-PrsmZ and DspI-PrsmZ complexes, we performed multiple rounds of EMSA which tested the binding of DspII or DspI with truncated promoter probes. A 320-bp DNA sequence containing the promoter PrsmZ was used as a full-length probe (Fig. 7C). The R fragments were truncated from the 3’ end, resulting in R1 (103 bp), R2 (184 bp), R3 (225 bp), R4 (272 bp), and R5 (320 bp) (Fig. 7C). As shown in Fig. 7D, both DspII and DspI bound to fragments R2 to R5 but not the fragments R1, suggesting that R2 contained the binding site of DspII and DspI. Therefore, R2 was further truncated into 123-bp, 143-bp, 163-bp, and 184-bp fragments. EMSA showed that DspII and DspI bound to the 143-bp, 163-bp, and 184-bp fragments (Fig. 7E), indicating that the sequence between 123-143 contained the binding site. To determine whether additional binding sites existed upstream of the 123-143 region, we truncated the full-length probe from the 5’ end, resulting in F1 (95 bp), F2 (176 bp), F3 (225 bp), F4 (285 bp), and F5 (320 bp). EMSA results showed that DspII and DspI bound to F3, F4, and F5 but not F1 and F2 (Fig. 7F), meaning that there was no binding site upstream of the 123-143 region. Taken together, the binding site of DspII and DspI reside in the region of 5’-TTAAGTAACTTATTGAAATTA-3’, which partially overlaps with the binding site of MvaT and locates at downstream of the binding site of GacA35,37 (Fig. 7C). This result not only determines the binding details of DspII-PrsmZ and DspI-PrsmZ complexes but also explains that DspII and DspI act as roadblocks to antagonize the GacA-dependent upregulation of RsmZ.
Since CDA exhibited an inhibitory effect on DspII/DspI-induced T3SS gene expression (Fig. 5K), we moved to explore whether CDA influenced the binding affinity of DspII and DspI at the promoter PrsmZ. It was shown that the binding between DspII or DspI and PrsmZ became weaker when the CDA concentration was increased (Supplementary Fig. 8C), implying that high concentration of CDA inhibits T3SS gene expression partially due to the reduced binding affinity of DspII and DspI with the promoter of rsmZ. Given that the enzymatically dead DspII and DspI variants cannot fully restore T3SS gene expression in the ΔdspII and ΔdspI mutants (Fig. 5L), we wondered whether these enzymatically active sites participate in DNA binding. We purified these variants and performed EMSA to examine their binding affinity with PrsmZ. As expected, decreased binding affinity of these variants with PrsmZ was observed compared to the wild-type DspII and DspI (Fig. 7G). These results indicate that all three enzymatically active residues in DspII and DspI are important for DNA binding. Possibly owing that DspI contains one more enzymatically active residue than DspII, it displayed higher DNA-binding affinity than the DspII protein as we observed in all the EMSA results.
To further understand how DspII and DspI antagonize the GacA-dependent upregulation of RsmY, we first tried to examine whether co-presence of DspII and DspI can bind at the promoter of RsmY since upregulation of T3SS gene expression requires the cooperation of DspII and DspI. However, EMSA did not show any shifted PrsmY in the co-presence of DspII and DspI (Supplementary Fig. 9A). Because the DNA sequence that is recognized by DspII and DspI in PrsmZ is not present in PrsmY, it is reasonable why DspII and DspI cannot bind to PrsmY. We next investigated whether DspII and DspI affect the binding affinity of GacA at PrsmY but the EMSA result showed that neither DspII nor DspI affects the DNA-binding affinity of GacA (Supplementary Fig. 9B). We moved to explore if DspII or DspI can potentially interact with GacA and thus reduce the transcription activity of PrsmY. Interestingly, both DspII and DspI were found to directly interact with GacA using a pull-down assay (Fig. 7H). This result indicated that DspII and DspI antagonize the GacA-activated transcription of RsmY by directly interacting with the GacA protein.
DspII interacts with DspI and the binding affinity is associated with CDA concentrations
Given the interdependence of DspII and DspI in synthesizing CDA, inducing biofilm dispersion and swarming motility, and activating T3SS gene expression, we hypothesized that DspII and DspI might interact directly. First, we conducted bacterial two-hybrid assay to detect potential interactions between DspII and DspI. It showed that co-presence of DspII and DspI enabled the normal growth of colonies on the selective medium (Fig. 8A), suggesting that DspII interacts with DspI. A pull-down assay was also performed, confirming the physical interaction between DspII and DspI (Fig. 8B). We further investigated the effect of CDA on DspII and DspI interaction. As shown in Fig. 8C, the binding affinity between DspII and DspI was enhanced with an increasing concentration of CDA below 10 μM, while the binding affinity was decreased when the concentration of CDA was higher than 10 μM. These findings demonstrated that DspII interacts with DspI to form a complex and CDA modulates the affinity of interaction in a concentration-dependent manner.
A Bacterial two-hybrid assay suggested direct interaction between DspII and DspI. Interaction between LGF2 and GallP served as positive control. B Pull-down assay showed the interaction between the DspII and DspI proteins. C Pull-down assay was performed to examine the interaction between the DspII and DspI proteins with or without the presence of the signal CDA. D Structural model of the DspII-DspI complex. E Representative amino acids predicted for the interaction between DspII and DspI. F Pull-down assay showed the abolished interaction between DspII and the DspI# variant (#: R57A/R96A/E231A/R232A). G CDA production was measured in the ΔdspI mutant with the complementation of the DspI# variant. H–J Biofilm biomass (H), swarming motility (I), and β-galactosidase activity of the PexsCEBA-lacZ transcriptional fusion (J) were measured in the ΔdspI mutant with the complementation of the DspI# variant. For G–J data are presented as mean values ± SEM of n = 3 biological replicates. For B, C, F representative blots shown of 2 independent experiments. Statistical analysis was performed using one-way ANOVA with Tukey’s multiple comparisons test. Source data are provided as a Source Data file.
To understand the interaction details of the DspII-DspI complex, we employed HDOCK and PLIP servers38,39 to predict the interaction model and the result showed three types of interactions including hydrophobic interactions, hydrogen bonds, and salt bridges (Fig. 8D and Supplementary Data 2). To validate the interaction between DspII and DspI, we selectively generated substitutions of four amino acids residues involved in the three types of interactions in the DspI protein (Fig. 8E). The resultant DspI# (DspIR57A/R96A/E231A/R232A) showed a dramatically decreased binding affinity with DspII compared to the wild-type DspI (Fig. 8F). In addition, complementation of the DspI# variant in the ΔdspI mutant was unable to re-activate the biosynthesis of CDA, biofilm dispersion, swarming motility, and T3SS gene expression (Fig. 8G–J). Together, these results demonstrated that the interaction between DspII and DspI is essential for both the enzymatic and non-enzymatic functions to mediate the chronic-to-acute virulence switch.
Discussion
The transition from chronic to acute virulence represents a pivotal adaptive strategy for bacterial pathogens, facilitating their dynamic response to host environments. In P. aeruginosa, the DSF-family QS molecule CDA serves as an endogenous inducer of this virulence switch, making it a promising target for anti-infective therapies. While DspI has been recognized for over a decade as the primary enzyme involved in CDA production18, our current study identifies DspII as a functionally equivalent and an indispensable partner with DspI in CDA biosynthesis and mediating the chronic-to-acute virulence switch (Fig. 9). Notably, DspII and DspI exhibit coordinated, cell density-dependent upregulation and interact physically to form a DspII-DspI complex that is essential for CDA biosynthesis. The synthesized CDA is likely detected by DspS, a known CDA sensor9, which subsequently upregulates the PDE RbdA and reduces intracellular c-di-GMP levels, thereby inhibiting Psl biosynthesis while simultaneously activating flagellar biosynthesis. These inversely regulated effects enable P. aeruginosa cells to regain motility and disperse from the overcrowded community to new host niches. Intriguingly, beyond the enzymatic role in CDA biosynthesis, the DspII-DspI complex demonstrates a non-enzymatic function in transcriptional regulation. The complex effectively suppresses the transcriptions of sRNAs RsmZ and RsmY, which are initiated by the response regulator GacA. This suppression leads to overproduction of the key acute virulence factor T3SS. The dual functionality of the DspII-DspI complex, serving both as a metabolic enzyme and a non-enzymatic transcriptional regulator, underscores its multifunctional nature in orchestrating the chronic-to-acute virulence switch. This sophisticated regulatory mechanism ensures that acute virulence factors are primed during biofilm dispersal, enabling rapid host invasion following bacterial dissemination.
DspII and DspI are synchronously upregulated in a cell density-dependent manner. They interact physically to form a DspII-DspI complex that is essential for CDA biosynthesis. The interaction between DspII and DspI is coordinated by CDA concentration. Sub-threshold levels of CDA enhance the affinity of the DspII-DspI complex to promote CDA production and the dispersal process. In contrast, high CDA concentrations destabilize DspII-DspI interaction, preventing further CDA production. The synthesized CDA is likely detected by DspS (Kalia et al., mBio, 2023), which subsequently upregulates the PDE RbdA and reduces intracellular c-di-GMP levels. Decrease of c-di-GMP content leads to the downregulation of Psl biosynthesis while the activation of flagellar biosynthesis through the effector FleQ, which enables P. aeruginosa cells to regain motility and disperse from the biofilm to new host niches. Simultaneously, the DspII-DspI complex plays a non-enzymatic role in transcriptional regulation when the environmental CDA concentration is low in the dispersed cells. The complex suppresses the transcriptions of sRNAs, RsmZ and RsmY, which are initiated by the response regulator GacA. This suppression leads to the overproduction of the key acute virulence factor T3SS and increased cytotoxicity and pathogenicity of the pathogens, promoting the establishment of new infections.
The DSF-family signal for interspecies, intraspecies, and inter-kingdom communication are characterized by an unsaturated fatty acid backbone40. Originally identified in Xanthomonas campestris pv. campestris (Xcc) as a critical regulator of virulence factor production and biofilm dispersion41, DSF-family signal is now recognized as evolutionarily conserved across diverse bacterial species. In P. aeruginosa, the DSF analog CDA has been established as a key mediator of biofilm dispersion and motility. Although the enoyl-CoA hydratase/isomerase DspI was previously recognized as an essential enzyme for CDA biosynthesis18, the mechanism of CDA biosynthesis remains largely unclear. The PAO1 genome encodes 12 putative enoyl-CoA hydratases/isomerases with similarity from non-significant to about 40% compared with DspI, suggesting potential functional redundancy or diversification. Here, by systematically investigating all these enzymes, we reported, for the first time, the discovery of DspII (PA0744) as a novel enoyl-CoA hydratase/isomerase that complexes with DspI to regulate biofilm dispersion via CDA biosynthesis. Notably, PA4980, a previously proposed candidate for CDA biosynthesis42, showed no impact on biofilm dispersion upon deletion, excluding its role in this regulatory pathway. Despite their genomic proximity and shared function as enoyl-CoA hydratase/isomerases, it is interesting to note that DspII and DspI exhibit low sequence identity, which suggests that they originated from an ancient gene duplication event. Analysis on their distribution and evolution reveals that the clustered dspII and dspI genes are confined to three specific clades, one Alphaproteobacteria clade and two distinct Gammaproteobacteria clades, suggesting that the dspII-dspI gene cluster was formed on at least three independent occasions in evolutionary history (Supplementary Fig. 10). The repeated and independent formation of this gene cluster implies a strong selective advantage, potentially reflecting a pre-existing functional synergy between two proteins that was stabilized and enhanced through coordinated, localized regulation.
Similar to the DSF signaling pathway in other bacterial species, which depends on the second messenger c-di-GMP to regulate bacterial physiology and virulence43,44, the CDA signaling system in P. aeruginosa orchestrates critical behaviors in coordination with the c-di-GMP signaling pathway. In this study, we established that DspII and DspI are essential for CDA biosynthesis, which subsequently triggers biofilm dispersion and swarming motility. These phenotypic changes are linked to a reduction of intracellular c-di-GMP levels, primarily caused by the upregulation of the PDE RbdA. Notably, our screening result revealed that, apart from RbdA, several other DGCs and PDEs including those known to modulate c-di-GMP content exhibited expression changes (Supplementary Data 1). This finding highlights the intricate regulatory network through which CDA influences intracellular c-di-GMP dynamics via global regulation with RbdA as the main player. Future investigations could explore the involvement of additional DGCs and PDEs targeted by CDA, as their collective actions may contribute to the chronic-to-acute virulence switch. FleQ, a master regulator that inversely controls EPS and flagellar biosynthesis, plays a pivotal role in mediating the switch between planktonic and biofilm lifestyles in P. aeruginosa45. Here, we discovered that FleQ is essential for biofilm accumulation by specifically activating the biosynthesis of Psl in response to elevated c-di-GMP levels observed in CDA-deficient ΔdspI and ΔdspII mutants. These findings delineate the signaling cascade by which CDA governs the transition from a sessile to a motile state through c-di-GMP and its downstream effector FleQ. This work provides detailed molecular-level insights into how CDA could induce biofilm dispersion, offering a deeper understanding of its role in bacterial adaptation and pathogenesis.
It was previously shown that disruption of dspI inhibits biofilm dispersion and attenuates pathogenicity by diminishing the production of several secreted virulence factors such as pyoverdine19. Additionally, the mutant ΔdspI has been shown to exhibit reduced cytotoxicity towards A549 cells19, a phenotype primarily attributed to T3SS which facilitates the direct injection of effector proteins into host cells. However, whether DspII or DspI and their product CDA regulate T3SS gene expression is not known. Intriguingly, our study revealed that both DspII and DspI, independent of CDA, play a critical role in activating T3SS gene expression. This discovery uncovers a previously unrecognized non-enzymatic function of DspII and DspI, wherein they synergistically promote T3SS gene expression by directly antagonizing the GacA-upregulated transcription of sRNAs, RsmY and RsmZ. Although both DspII and DspI are found to bind the region of 5’-TTAAGTAACTTATTGAAATTA-3’ which is located at downstream of the GacA-binding site in the PrsmZ promoter, more specific binding sequence for each protein requires further investigation since both DspII and DspI can bind to this region individually. Nonetheless, these findings highlight a novel regulatory layer in the control of acute virulence in P. aeruginosa, expanding our understanding of the multifunctional roles of DspII and DspI beyond their enzymatic activity in CDA biosynthesis.
Our findings demonstrate that direct binding between DspII and DspI plays a crucial role in both CDA biosynthesis and T3SS gene regulation. Interestingly, this protein-protein interaction exhibits dynamic responsiveness to CDA concentrations through a sophisticated feedback mechanism. During the initial stages of biofilm dispersal, sub-threshold levels of CDA enhance the affinity of the DspII-DspI complex, thereby amplifying CDA production to trigger the dispersal process. In contrast, high CDA concentrations destabilize DspII-DspI interaction, establishing an auto-regulatory negative feedback loop that curtails further CDA biosynthesis. The regulatory influence of CDA extends beyond biofilm dynamics, as high concentrations within the biofilm community seem to not only initiate dispersal but also repress T3SS gene expression in resident cells. This dual function promotes metabolic efficiency by preventing the energy-costly production of acute virulence factors in non-dispersing populations. Following dispersal from the biofilm and subsequent drop of environmental CDA levels, the high protein levels of DspII and DspI in the departed cells activate their T3SS gene expression to establish new infections. This intricate regulatory mechanism, balancing biofilm dispersal initiation with virulence modulation, represents an evolutionary adaptation that enables precise resource allocation and facilitates the pathogen’s strategic transition between chronic persistence and acute invasion.
Disarming the pathogen by blocking its ability to switch to the acute, invasive state could suppress or prevent disease severity. Our finding that the DspII-DspI complex possesses dual functions offers two potential avenues for intervention. First, small-molecule inhibitors could be developed to block the enzymatic activity of the complex or CDA-degrading enzymes could be applied, thereby lowering CDA content and preventing biofilm dispersion to cause new infections. Second, and perhaps more promisingly, agents that disrupt the protein-protein interaction between DspII and DspI could simultaneously abolish both CDA biosynthesis and T3SS gene activation, effectively locking the pathogen in a chronic, less harmful state. This dual inhibition would not only prevent the dissemination of biofilm cells but also ensure that escaped cells are deficient in expressing the key acute virulence factor T3SS. Given that our enzymatically dead variants retained their regulatory function on T3SS gene expression, targeting the complex’s structural integrity might offer a more comprehensive blockade of the virulence switch. Future work will focus on high-throughput screening for compounds that disrupt the DspII-DspI interaction and evaluating their efficacy in preventing acute infections.
In summary, this study, for the first time to our knowledge, reveals a sophisticated, multilayered regulatory network in which DspII and DspI serve as a linchpin that integrates QS signal biosynthesis, transcriptional regulation, and virulence factor expression. By synthesizing CDA, decreasing c-di-GMP levels, and repressing sRNAs transcription, the DspII-DspI partner enables P. aeruginosa to actively shift from chronic persistence to acute invasion. The discovery of their non-enzymatic transcriptional regulatory activity adds a novel dimension to the functionality of QS signal synthases. These findings not only deepen our understanding of bacterial adaptive strategies but also highlight potential therapeutic targets to prevent chronic-to-acute virulence switch, offering new avenues to combat P. aeruginosa infections.
Methods
Statistics & Reproducibility
Experiments were generally conducted with three biological replicates unless otherwise noted. Three bacterial colonies were randomly selected from a solid plate for each experiment. Blinding was not applicable because bacterial colonies were randomly selected for each experiment. No data were excluded from the analyses. Data were analyzed using GraphPad Prism (version 8.0.2). Statistical analyses were conducted based on one-way analysis of variance (ANOVA), two-way ANOVA, or Student’s t-test as indicated in the figure legends. A p value of 0.05 or less was considered significant for all analyses.
Bacterial strains, plasmids, primers, and growth conditions
Strains and plasmids used in this study are listed in Supplementary Data 3. Primers used in this study are summarized in Supplementary Data 4. Strains were normally cultured in Luria-Bertani (LB) medium (10 g/L Tryptone, 5 g/L Yeast extract and 5 g/L NaCl) or on LB-agar plates at 37 °C. Antibiotics were added as following concentrations when necessary: gentamicin, 50 μg/ml; kanamycin, 50 μg/ml; ampicillin, 100 μg/ml.
Construction of gene-deletion mutants and gene complementation or overexpression
For gene deletion, 500–1000-bp upstream and downstream regions of target genes were amplified and ligated to pK18mobsacB which was pre-digested with BamHI and HindIII. Using pRK2013 as the helper plasmid, constructed plasmids for gene deletion were delivered into PAO1 or its derivatives by tri-parental mating. Desired mutants were selected on LB plate containing 10% sucrose and confirmed by PCR and sequencing. For gene complementation, the open reading frames of target genes and their promoter sequences were amplified from the PAO1 genome and cloned into the plasmid pUC18T-mini-Tn7T-Gm which was pre-digested with BamHI and HindIII. The constructed plasmids were introduced into PAO1 mutants with the help of the plasmid pTNS2 by electroporation. Desired colonies were selected on Pseudomonas Isolation Agar (PIA) plates with gentamicin and confirmed by PCR and sequencing. For gene overexpression, target genes were ligated into pUCP18 which was pre-digested with HindIII and BamHI. Constructed plasmids were introduced into corresponding strains by tri-parental mating.
Biofilm assay
Biofilm biomass was measured as previously described46. Briefly, 100 μl bacterial culture with OD600 of 0.02 was inoculated in LB medium in 96-well polypropylene microtiter dishes. After incubation under static condition for 16 h, cell density was determined, and the liquid culture was removed carefully. 0.1% crystal violet was added to each well and incubated for 15 min at room temperature ( ~ 25 °C). After staining, wells were washed carefully to remove the unbound crystal violet. Biofilms were dissolved in 95% ethanol and A570 of each well was measured using a microplate reader (BioTek, United States). Biofilm biomass was normalized by cell density (OD600). All experiments were conducted with three independent biological repeats.
Swarming motility assay
Swarming motility was performed as described previously47. Plates used for swarming assay were prepared with 8 g/L nutrient broth, 5 g/L glucose, and 5 g/L bacto-agar. The overnight cultures were diluted to OD600 of 1.0 and 1 μl culture was spotted at the center of swarming plates. After incubation at 37 °C for 16-20 h, diameters of swarming zones were measured. All experiments were conducted with three independent biological repeats.
Transmission electron microscopy (TEM)
TEM observation of bacterial flagellum was performed as preciously described48. Bacterial cells were collected from swarming plates and diluted in the MilliQ water. Cells were stained with 2% aqueous solution of phosphotungstic acid (pH 7.4) for 20 min before fixing in parlodion (Mallinckrodt, Inc., United States) carbon-coated grid (300 mesh). After that, samples were examined by the transmission electron microscope Talos F200S (Thermo Fisher Scientific, United States).
Promoter activity assay
Promoter activity was assessed by measuring GFP fluorescence or β-galactosidase activity. Target gene promoters were amplified and ligated to pPROBE-NT-gfp or mini-CTX-lacZ vectors. The resulting constructs were transformed into P. aeruginosa strains and verified by colony PCR. For GFP fluorescence assay, overnight bacterial cultures were diluted to an OD600 of 0.02 and then sub-cultured in 96-well plates under static conditions. Cell densities (OD600) and fluorescence were measured every 4 h using a microplate reader (BioTek, United States). Results were shown as the ratio of fluorescence and OD600 at different time points. For β-galactosidase activity assays, bacterial cells were harvested at an OD600 of 1.5 via centrifugation and then resuspended in Z buffer (60 mM Na2HPO4, 40 mM NaH2PO4, 10 mM KCl, 1 mM MgSO4, 50 mM β-mercaptoethanol). A 100 μl aliquot of bacterial suspension was diluted in 500 μl Z buffer, followed by the addition of 20 μl chloroform and 20 μl 0.1% SDS. Cells were lysed by thorough vortexing and incubated in a 30 °C water bath for 20 min. Subsequently, 100 μl of 4% (w/v) o-nitrophenyl-β-D-galactoside (ONPG) was then added to the suspension, which was then incubated for 10 min. The reaction was terminated with 250 μl of 1 M Na₂CO₃. The promoter activity in Miller Units was calculated and relative promoter activity was presented.
RT-PCR and RT-qPCR
Overnight bacterial cultures were 1:50 diluted with fresh LB medium and grown when OD600 reached 1.5. Total RNA was extracted using the SV Total RNA separation and purification system (Promega, United States) following manufacturer’s instructions. After determining the RNA quality and concentration, first-strand cDNAs were reverse transcribed from 1 μg RNA using HiScript III 1st Strand cDNA Synthesis Kit (with gDNase) (Vazyme, China). Subsequently, RT-PCR was performed using 2×Phanta Max Master Mix (Vazyme, China) with RNA or cDNA as templates. For RT-qPCR analysis, the ChamQ Universal SYBR qPCR master mix (Vazyme, China) was used, and the amplification was monitored in the ABI QuantStudio 6 Flex real-time system. Relative gene expression level was calculated by using the 2-ΔΔCT method and the 50S ribosomal protein gene rplU was selected as the reference to standardize all the samples49. All experiments were conducted with three independent biological repeats.
CDA quantification using LC-MS
CDA quantification was carried out as reported previously with slight modifications17. The overnight culture was 1:50 diluted with fresh LB medium and incubated for 20 h. Supernatant was collected, and 1/1000 volume of acetic acid was added before CDA extraction with an equal volume ethyl acetate. The organic phase was dried by vacuum freeze and residues were dissolved with 1 ml methanol. Samples were analyzed using Q Exactive Focus Hybrid Quadrupole-Orbitrap mass spectrometer (Thermo Fisher Scientific, United States) in negative ionization mode. CDA separation was achieved by a 2.1×100-mm HSS T3 1.8 μm column (Waters, United States) using a gradient system consisting of methanol and MilliQ water with 1/1000 volume of formic acid. CDA was quantified using selected-ion monitoring (SIM) mode. All measurements were performed with three independent biological repeats.
c-di-GMP quantification using LC-MS
The quantification of c-di-GMP was carried out following a previously reported protocol with minor adjustments50. The overnight culture was diluted 1:100 into fresh LB medium and incubated for 20 h. 10 ml cell culture were pelleted and resuspend in 1 ml sterilized MilliQ water. After perchloric acid was added, cell suspensions were vortexed thoroughly and then incubated on ice for 30 min. After centrifugation, supernatants were transferred to a new tube and 1/5 volume of 2.5 M KHCO3 was added. The samples were centrifuged to remove perchlorate salt precipitates. Supernatant was collected for LC-MS analysis. LC-MS was undertaken using an Q Exactive Focus Hybrid Quadrupole-Orbitrap mass spectrometer (Thermo Fisher Scientific, United States) with a 2.1 × 100-mm HSS T3 1.8 μm column (Waters, United States). All measurements were performed with three independent biological repeats.
Bacterial two-hybrid assay
Bacterial two-hybrid assay was carried out to detect protein-protein interaction according to protocol of BacterioMatch II Two-Hybrid System Vector Kit (Agilent Technologies, United States). The ORFs of dspI and dspII were cloned into pBT and pTRG, respectively. The recombinant pBT-dspI and pTRG-dspII plasmids were co-transformed into the reporter strain XL1-Blue MRF Kan. The overnight bacterial cultures were serially diluted using M9+ His dropout broth and spotted on nonselective medium and selective medium (5 mM 3-AT plus 12.5 μg/ml streptomycin). Plates were incubated at 37 °C for 24 h.
Pull-down assay
Pull-down assay was carried out using Pierce GST Protein Interaction Pull-Down Kit (Thermo Fisher Scientific, United States) according to the manufacturer’s instructions. Briefly, bacterial cells were collected by centrifugation and resuspended with lysis buffer. The purified GST-tagged protein was incubated with glutathione agarose for 4 h at 4 °C and then washed gently to remove unbound proteins. The glutathione agarose was then incubated with cell lysate containing His-tagged protein at 4 °C for 16 h. The column was washed five times, and the proteins were eluted using 10 mM glutathione elution buffer. The eluted samples were incubated with loading buffer and heated at 95 °C for 10 min prior to western blot analysis.
Protein purification
Plasmids for protein expression were delivered into BL21(DE3). For protein expression, 1 ml bacterial overnight culture was inoculated into 1 L LB medium. When bacterial cells grown to OD600 of 0.5-0.8, IPTG was added to induce protein expression, and the cell culture was incubated at 16 °C overnight. Cell pellets were harvested and lysed by sonication. After centrifugation, the supernatant was filtered with a 0.22-μm filter prior to incubation with glutathione beads (Smart-Lifescience, China) or Ni NTA beads (Smart-Lifescience, China) for 30 min. After washing with PBS, proteins were eluted by 10 mM glutathione elution buffer or 200 mM imidazole elution buffer. The GST-tag and His-tag of the purified proteins were further digested using PreScission Protease (Smart-Lifescience, China), respectively, following the manufacture’s construction.
Electrophoretic gel mobility shift assay (EMSA)
DNA sequences of target promoter regions were amplified by PCR using the primers listed in Supplementary Data 4. DNA fragments were labeled with biotin using a Biotin 3′ End DNA Labeling Kit (Thermo Fisher Scientific, United States). Protein-DNA interaction was analyzed using LightShift Chemiluminescent EMSA Kit (Thermo Fisher Scientific, United States) following the manufacture’s construction. Briefly, purified protein and the labeled probes were added into the binding reactions and incubated at 25 °C for 30 min. The reaction mixtures were mixed with 5×Loading Buffer and loaded onto the polyacrylamide gel. Following electrophoresis, DNA probes were transferred to the nylon membrane (Millipore, United States) and crosslinked using UV irradiation. Membranes were sequentially washed with Blocking Buffer, Wash Buffer, and Substrate Equilibration Buffer as specified in the manufacturer’s protocol. Protein-DNA complexes were visualized using Working Substrate Solution and imaged with a cooled CCD camera (Tanon, China). For competitive EMSA assays, 50 ng or 150 ng of unlabeled PrsmZ DNA fragments were added to the reaction mixtures.
Western blot analysis
Overnight bacterial cultures were 1:50 diluted in fresh LB medium and grown at 37 °C when OD600 reached 1.5. 10-ml aliquots of bacterial cultures were centrifuged, and pellets were suspended in 200 µl RIPA lysis buffer (Biosharp, China) and followed by incubation at 25 °C for 30 min. After centrifugation, the supernatants were mixed with 5× SDS-PAGE loading buffer and heated at 98 °C for 10 min. Proteins were separated using SmartPAGETM 4-20% Precast Protein Gel (Smart-Lifescience, China) and transferred to a PVDF membrane (Millipore, United States) via electroblotting. Subsequently, Membranes were blocked for 1 h at room temperature with PBST (PBS containing 1% Tween-20) containing 5% skim milk, then probed with anti-ExsA, anti-β-RNAP, anti-GST, and anti-His primary antibodies, respectively. After washing, membranes were incubated with HRP-conjugated goat anti-mouse secondary antibody (Abbkine, United States). Protein bands were visualized using the ECL kit according to the manufacture’s protocol and imaged with a cooled CCD camera (Tanon, China).
Cytotoxicity assay
Bacterial cytotoxicity was assessed by measuring the activity of lactate dehydrogenase (LDH) which is released from the human lung epithelial cell A54951. Briefly, 1.5 × 104 A549 cells were seeded into a 96-well plate containing DMEM medium (Gibco, United States) supplemented with 10% FBS (Gibco, United States) and cultured overnight at 37 °C under 5% CO2 and 95% humidity. P. aeruginosa strains were cultured in LB medium to an OD600 of 1.5. Bacterial cells were centrifuged and resuspended in DMEM with 1% FBS. A549 cells were washed twice with PBS buffer and infected with bacteria at a multiplicity of infection (MOI) of 50 for 4 h. Cytotoxicity was quantified by CytoTox 96® Non-Radioactive Cytotoxicity Assay kit (Promega, United States) according to the manufacturer’s instructions.
Galleria mellonella killing assay
The acute virulence of P. aeruginosa was evaluated using a G. mellonella infection model adapted from a previous study52. Overnight bacterial cultures were 1:50 diluted in fresh LB medium and grown to an OD600 of 1.5. Bacterial cells were pelleted, resuspended in PBS, and adjusted to 1×105 CFU. Ten G. mellonella larvae were injected with 20 μl bacterial suspension into the secondary right proleg using a syringe and then incubated at 30 °C. The survival rates were monitored every 2 h.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data are provided with this paper.
References
Zhou, J., Ma, H. & Zhang, L. Mechanisms of virulence reprogramming in bacterial pathogens. Annu. Rev. Microbiol. 77, 561–581 (2023).
Galán, J. E. & Collmer, A. Type III secretion machines: bacterial devices for protein delivery into host cells. Science 284, 1322–1328 (1999).
Coburn, B., Sekirov, I. & Finlay, B. B. Type III secretion systems and disease. Clin. Microbiol. Rev. 20, 535–549 (2007).
Lee, J. et al. A cell-cell communication signal integrates quorum sensing and stress response. Nat. Chem. Biol. 9, 339–343 (2013).
Sousa, A. M. & Pereira, M. O. Pseudomonas aeruginosa diversification during infection development in cystic fibrosis lungs-a review. Pathogens 3, 680–703 (2014).
Flemming, H. C. et al. Biofilms: an emergent form of bacterial life. Nat. Rev. Microbiol. 14, 563–575 (2016).
Costerton, J. W., Stewart, P. S. & Greenberg, E. P. Bacterial biofilms: a common cause of persistent infections. Science 284, 1318–1322 (1999).
Liu, H. Y., Prentice, E. L. & Webber, M. A. Mechanisms of antimicrobial resistance in biofilms. npj Antimicrob. Resist. 2, 27 (2024).
Kalia, M., Amari, D., Davies David, G. & Sauer, K. cis-DA-dependent dispersion by Pseudomonas aeruginosa biofilm and identification of cis-DA-sensory protein DspS. mBio 14, e02570–23 (2023).
Qin, S. et al. Pseudomonas aeruginosa: pathogenesis, virulence factors, antibiotic resistance, interaction with host, technology advances and emerging therapeutics. Signal Transduct. Target. Ther. 7, 199 (2022).
Broder, U. N., Jaeger, T. & Jenal, U. LadS is a calcium-responsive kinase that induces acute-to-chronic virulence switch in Pseudomonas aeruginosa. Nat. Microbiol. 2, 16184 (2016).
Cao, P. et al. A Pseudomonas aeruginosa small RNA regulates chronic and acute infection. Nature 618, 358–364 (2023).
Moscoso, J. A., Mikkelsen, H., Heeb, S., Williams, P. & Filloux, A. The Pseudomonas aeruginosa sensor RetS switches Type III and Type VI secretion via c-di-GMP signalling. Environ. Microbiol. 13, 3128–3138 (2011).
Nadal et al. The multiple signaling systems regulating virulence in Pseudomonas aeruginosa. Microbiol. Mol. Biol. Rev. 76, 46–65 (2012).
Wang, X. et al. An antibiotic-responsive regulator orchestrates chronic-to-acute virulence switch in Pseudomonas aeruginosa. Nucleic Acids Res 53, gkaf471 (2025).
Valentini, M. & Filloux, A. Biofilms and cyclic di-GMP (c-di-GMP) signaling: Lessons from Pseudomonas aeruginosa and other bacteria. J. Biol. Chem. 291, 12547–12555 (2016).
Davies David, G. & Marques Cláudia, N. H. A fatty acid messenger is responsible for inducing dispersion in microbial biofilms. J. Bacteriol. 191, 1393–1403 (2009).
Amari Diana, T., Marques Cláudia, N. H. & Davies David, G. The putative enoyl-coenzyme a hydratase DspI is required for production of the Pseudomonas aeruginosa biofilm dispersion autoinducer cis−2-decenoic acid. J. Bacteriol. 195, 4600–4610 (2013).
Liu, L. et al. Structural and functional studies on Pseudomonas aeruginosa DspI: implications for its role in DSF biosynthesis. Sci. Rep. 8, 3928 (2018).
Davies, D. G. & Marques, C. N. H. A Fatty Acid Messenger Is Responsible for Inducing Dispersion in Microbial Biofilms. J. Bacteriol. 191, 1393–1403 (2009).
Amari, D. T., Marques, C. N. H. & Davies, D. G. The putative enoyl-coenzyme A hydratase DspI is required for production of the biofilm dispersion autoinducer -2-decenoic acid. J. Bacteriol. 195, 4600–4610 (2013).
Blum, M. et al. InterPro: the protein sequence classification resource in 2025. Nucleic Acids Res 53, D444–D456 (2025).
Irie, Y. et al. Pseudomonas aeruginosa biofilm matrix polysaccharide Psl is regulated transcriptionally by RpoS and post-transcriptionally by RsmA. Mol. Microbiol. 78, 158–172 (2010).
Colvin, K. M. et al. The Pel and Psl polysaccharides provide Pseudomonas aeruginosa structural redundancy within the biofilm matrix. Environ. Microbiol. 14, 1913–1928 (2012).
Wang, Z., Song, L., Liu, X., Shen, X. & Li, X. Bacterial second messenger c-di-GMP: emerging functions in stress resistance. Microbiol. Res. 268, 127302 (2023).
Borlee, B. R. et al. Pseudomonas aeruginosa uses a cyclic-di-GMP-regulated adhesin to reinforce the biofilm extracellular matrix. Mol. Microbiol. 75, 827–842 (2010).
Ha, D. G. & O’Toole George, A. c-di-GMP and its effects on biofilm formation and dispersion: a Pseudomonas aeruginosa review. Microbiol. Spectr. 3, MB-0003−2014 (2015).
Zhou, T. et al. The Two-component system FleS/FleR represses H1-T6SS via cyclic di-GMP signaling in Pseudomonas aeruginosa. Appl. Environ. Microbiol. 88, e01655–21 (2022).
Banerjee, P. et al. Molecular and structural facets of c-di-GMP signalling associated with biofilm formation in Pseudomonas aeruginosa. Mol. Asp. Med. 81, 101001 (2021).
Feng, Q., Zhou, J., Zhang, L., Fu, Y. & Yang, L. Insights into the molecular basis of c-di-GMP signalling in Pseudomonas aeruginosa. Crit. Rev. Microbiol. 50, 20–38 (2024).
Miyata, S., Casey, M., Frank Dara, W., Ausubel Frederick, M. & Drenkard, E. Use of the Galleria mellonella caterpillar as a model host to study the role of the type III secretion system in Pseudomonas aeruginosa pathogenesis. Infect. Immun. 71, 2404–2413 (2003).
Huang, J., Zhang, Y., Sheng, S., Zhang, L. H. & Xu, Z. Protocol for examining the T3SS-mediated cytotoxicity of Pseudomonas aeruginosa using the A549 cell line. STAR Protoc. 5, 103301 (2024).
Williams McMackin E. A., Djapgne, L., Corley J. M. & Yahr T. L. Fitting pieces into the puzzle of Pseudomonas aeruginosa type III secretion system gene expression. J. Bacteriol. 201, e00209–19 (2019).
Huang, J., Xu, Z., Zhou, T., Zhang, L. H. & Xu, Z. Suppression of Pseudomonas aeruginosa type III secretion system by a novel calcium-responsive signaling pathway. iScience 27, 109690 (2024).
Brencic, A. et al. The GacS/GacA signal transduction system of Pseudomonas aeruginosa acts exclusively through its control over the transcription of the RsmY and RsmZ regulatory small RNAs. Mol. Microbiol. 73, 434–445 (2009).
Intile Peter, J., Diaz Manisha, R., Urbanowski Mark, L., Wolfgang Matthew, C. & Yahr Timothy, L. The AlgZR Two-component system recalibrates the RsmAYZ posttranscriptional regulatory system to inhibit expression of the Pseudomonas aeruginosa type III secretion system. J. Bacteriol. 196, 357–366 (2014).
Wang, C., Ye, F., Kumar, V., Gao, Y. G. & Zhang, L. H. BswR controls bacterial motility and biofilm formation in Pseudomonas aeruginosa through modulation of the small RNA rsmZ. Nucleic Acids Res 42, 4563–4576 (2014).
Yan, Y., Tao, H., He, J. & Huang, S. Y. The HDOCK server for integrated protein–protein docking. Nat. Protoc. 15, 1829–1852 (2020).
Schake, P., Bolz, S. N., Linnemann, K. & Schroeder, M. PLIP 2025: introducing protein–protein interactions to the protein–ligand interaction profiler. Nucleic Acids Res 53, W463–W465 (2025).
He, Y. W. et al. DSF-family quorum sensing signal-mediated intraspecies, interspecies, and inter-kingdom communication. Trends Microbiol. 31, 36–50 (2023).
Wang, L. H. et al. A bacterial cell–cell communication signal with cross-kingdom structural analogues. Mol. Microbiol. 51, 903–912 (2004).
Rahmani-Badi, A., Sepehr, S., Fallahi, H. & Heidari-Keshel, S. Dissection of the cis-2-decenoic acid signaling network in Pseudomonas aeruginosa using microarray technique. Front. Microbiol. 6, 383 (2015).
Wang, M. et al. The cis-2-dodecenoic acid (BDSF) quorum sensing system in Burkholderia cenocepacia. Appl. Environ. Microbiol. 88, e02342–21 (2022).
Yang, C. et al. Burkholderia cenocepacia integrates cis-2-dodecenoic acid and cyclic dimeric guanosine monophosphate signals to control virulence. Proc. Natl. Acad. Sci. USA 114, 13006–13011 (2017).
Oladosu Victoria, I., Park, S. & Sauer, K. Flip the switch: the role of FleQ in modulating the transition between the free-living and sessile mode of growth in Pseudomonas aeruginosa. J. Bacteriol. 206, e00365–23 (2024).
An, S., Wu, J. & Zhang, L. H. Modulation of Pseudomonas aeruginosa biofilm dispersal by a cyclic-Di-GMP phosphodiesterase with a putative hypoxia-sensing domain. Appl. Environ. Microbiol. 76, 8160–8173 (2010).
Rashid, M. H. & Kornberg, A. Inorganic polyphosphate is needed for swimming, swarming, and twitching motilities of Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 97, 4885–4890 (2000).
Liu, Z. et al. CzcR Is Essential for Swimming Motility in Pseudomonas aeruginosa during Zinc Stress. Microbiol. Spectr. 10, e02846–22 (2022).
Zhou, T., Huang, J., Liu, Z., Xu, Z. & Zhang, L. H. Molecular mechanisms underlying the regulation of biofilm formation and swimming motility by FleS/FleR in Pseudomonas aeruginosa. Front. Microbiol. 12, 707711 (2021).
Hickman, J. W. & Harwood, C. S. Identification of FleQ from Pseudomonas aeruginosa as a c-di-GMP-responsive transcription factor. Mol. Microbiol. 69, 376–389 (2008).
Lin, Q. et al. tRNA modification enzyme MiaB connects environmental cues to activation of Pseudomonas aeruginosa type III secretion system. PLOS Pathog. 18, e1011027 (2022).
Si, M. et al. Manganese scavenging and oxidative stress response mediated by type VI secretion system in Burkholderia thailandensis. Proc. Natl. Acad. Sci. USA 114, E2233–E2242 (2017).
Acknowledgements
We thank Prof. Yinyue Deng (Sun Yat-sen University) for professional discussions and providing acetyl phosphate. We thank Prof. Haibo Wang (Chinese Academy of Sciences) for the instructions on protein purification. This work was supported by the National Natural Science Foundation of China (No. U22A20480 to L.Z. and No. 32370188 to Z.X.), the Guangzhou Basic and Applied Basic Research Foundation (No. 2025A04J5440 to Z.X.), the Guangdong Provincial Pearl River Talents Program (No. 2023QN10N029 to Z.X.), and the State Key Laboratory of Green Pesticide (No. GPLSCAU202406 to Z.X.).
Author information
Authors and Affiliations
Contributions
L.Z. and Z.X. designed the study; J.H., T.Z., X.Z., Y.X., W.Z., L.L., and C.C. performed research; J.H., L.Z., and Z.X. analyzed data; J.H., L.Z., and Z.X. wrote and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no conflicts of interest with the contents of this article.
Peer review
Peer review information
Nature Communications thanks Maria Tomas, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Huang, J., Zhou, T., Zhou, X. et al. Dual-functional quorum sensing signal synthases DspII and DspI coordinate virulence switch in Pseudomonas aeruginosa. Nat Commun 17, 1926 (2026). https://doi.org/10.1038/s41467-026-68622-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68622-x