Abstract
While numerous studies have reported that intratumor bacteria modulate tumor biological characteristics and therapeutic outcomes, the role of intracellular bacteria in gastrointestinal tumor (GIT) progression remains largely unexplored. Here, we show, through bioinformatics prediction and consecutive patient sample validation, that Stenotrophomonas is associated with tumor progression and poor prognosis in GIT. Using various murine tumor models, we find that intracellular Stenotrophomonas promotes GIT progression by decreasing IFNβ secretion and inhibiting MHC-II+ macrophages, thereby impairing the activity of effector CD8+ T cells. Mechanistically, Stenotrophomonas entry into tumor cells induces endoplasmic reticulum stress, degrades STING protein, and downregulates the type I IFN pathway response. Additionally, Stenotrophomonas contributes to resistance to immunotherapy in GIT, which can be reversed by levofloxacin. Our findings indicate that intracellular Stenotrophomonas within tumor cells plays an important role in promoting GIT progression and poor prognosis and could serve as a potential target to improve outcomes for patients with GIT.
Similar content being viewed by others
Introduction
Gastrointestinal tumors (GIT), including adenocarcinomas of the esophagogastric junction, gastric cancer, and colorectal cancer, are common malignancies associated with high mortality rates worldwide. GIT, which interfaces with the external environment, frequently interacts with diverse microbiomes. Bacteria are closely associated with tumor progression and metastasis, as well as the modulation of immunotherapy and chemotherapy responses in GIT1,2,3,4,5,6,7,8,9,10,11,12. Bacterial interactions with GIT progression occur through both indirect mechanisms within the tumor microenvironment (TME) and direct intracellular pathways in tumor or immune cells. In indirect interactions, bacteria secrete proteins or directly bind to cell receptors on tumor or immune cells. For instance, C. maltaromaticum enhances intestinal vitamin D production, and S. thermophilus secretes β-galactosidase, both inhibiting tumor progression. Conversely, S. anginosus activates the TMPC-ANXA2-MAPK axis, bacteria lacking ureolytic function suppress macrophage function, mycobacteria indirectly induce de novo Treg cells, and C. jejuni’s cytolethal distending toxin promotes tumor occurrence or progression1,2,3,4,6,7. These intracellular bacteria are associated with tumor subtypes and responses to immunotherapy13. Furthermore, intracellular bacteria can regulate cellular cytoskeleton and viability through mechanical stress, promoting breast cancer metastasis14. Notably, Stenotrophomonas (SP), a common opportunistic pathogen, is a multidrug-resistant bacterium widely colonizing the human body15. It possesses migratory abilities, inducing mastitis through upregulation of the calcium-ROS-AMPK-mTOR-autophagy pathway16. SP has been detected intracellularly in hepatic stellate cells, promoting senescence-associated secretory phenotype and cirrhosis via the TLR4-driven NF-κB signaling pathway, thus contributing to hepatocarcinogenesis17. Additionally, SP upregulates HDAC5 expression, promoting lung cancer progression18. While most previous research has focused on intratumor rather than intracellular bacteria, further investigation of intracellular bacteria in GIT is warranted to elucidate their role in cancer progression and treatment response.
Recent studies on TME regulation have predominantly focused on macrophages and their immunosuppressive role in inhibiting T cell function19,20,21,22. However, the paradigm of macrophage function has evolved. While initially considered solely as cancer cell killers and cytotoxic T cell activators, recent research has shifted towards identifying the regulatory mechanisms governing macrophage functional plasticity23,24. The functional state of macrophages is intricately linked to the activation of the type I interferon (IFN) signaling pathway, which induces macrophages to secrete tumor necrosis factor-alpha (TNFα) and interleukin-12, activating cytotoxic T cells and promoting tumor cell elimination, while also reprogramming CD206+ macrophages into nitric oxide synthase-expressing or MHC-II+ macrophages to enhance antigen processing and presentation capabilities23,24,25. The activation of the type I IFN signaling pathway in tumor-associated macrophages is regulated by a complex network of molecules, including toll-like receptors, MYD88, MAVS, STING, IFNβ26,27,28,29. Macrophages-mediated antigen processing and cytotoxic T cell activation are impaired in the absence of IFNβ stimulation, resulting in an M2-like phenotype that exhibits immunosuppressive properties and promotes tumor progression21,26,30.
The Stimulator of Interferon Gene (STING) protein emerges as a critical mediator in bacterial defense, modulating host innate immune responses and augmenting adaptive immunity through type I IFN pathway activation31,32,33,34,35,36,37,38,39,40. Within the TME, microbiota-derived STING agonists, such as cyclic di-AMP, activate the type I IFN pathway in monocytes, inducing substantial IFNβ production that shifts macrophages from pro-tumor to anti-tumor states, activates dendritic cells (DC), recruits natural killer (NK) cells, and enhances overall antitumor responses41,42. S. typhimurium with cGAS expression activates the STING pathway via cGAMPs in macrophages or DCs and promotes antitumor immunotherapy43. Conversely, Lachnospiraceae-secreted butyric acid inhibits STING activation and IFNβ production in intratumor DCs, impairing the antitumor effects of ionizing radiation44. There remains a lack of research specifically investigating the role of intracellular bacteria in modulating STING protein and immune responses within the TME.
The involvement of intracellular bacteria in GIT and their impact on immune-regulation remains insufficiently defined. In this study, we analyze intracellular bacteria in GIT progression, identifying SP that degrade STING in tumor cells and its resulting mitigated immune response. By impacting macrophages, SP would eventually damage the killing function of CD8+ T cells and lead to poor prognosis of cancer patients. These results highlight their broader implications as viable targets for therapeutic intervention in GIT that may inform the development of more effective treatment strategies for this patient population.
Results
Intracellular SP is associated with tumor progression in GIT
The majority of studies have focused on the intersection of gut microbes and tumor therapy45,46,47,48,49. Recent studies have reported the presence of intracellular bacteria in various tumor types. However, investigations of intracellular bacteria in GIT remain limited, despite the direct contact of GIT with numerous bacteria, which increases the likelihood of bacterial infiltration into tumor cells13,14,50,51,52. To investigate the enrichment of intracellular bacteria in GIT, we analyzed five public datasets using a bulk microbiome pipeline, employing the Kraken2Uniq method to match bacterial reference sequences (Fig. S1)13,53,54,55. Across all GIT subtypes, including adenocarcinoma of the gastroesophageal junction (AEG), gastric cancer (GC), and colorectal cancer (CRC), we observed abundant bacterial reads (Fig. 1A). We performed normalized comparisons to mitigate false positives and account for partial read degeneracy during sequence matching, which may result from the limited length of high-throughput sequencing reads. To standardize differences in sequencing library size, we applied two normalization metrics, reads per million microbiome reads and reads per million total reads, based on cell line data. By comparing the normalized read distributions of each species between cell lines and tumor samples, species showing similar or higher abundance in cell lines were classified as contaminants. This approach enabled the identification of several bacterial species unaffected by contamination (Fig. S2). Bacillus cereus, SP, Klebsiella pneumoniae, and Pseudomonas sp. CIP-10 is present at significantly higher abundances compared to other bacterial species in GIT (Fig. 1B). We conducted survival analysis and found that SP is significantly associated with poor prognosis (Fig. 1C). Furthermore, fluorescence in situ hybridization (FISH) confirmed the presence of SP within the GIT of our selected cohort (Fig. 1D and S3B). In the high-abundance group, SP is also significantly correlated with poor prognosis in our cohort, which includes patients with GC and AEG (Fig. 1E).
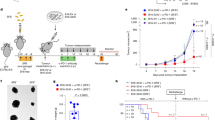
A Relative abundance of the top 10 microbiomes (species level) in multi-cohort datasets. GSE126304-AEG (n = 46), PRJNA788008-AEG (n = 83), GSE156451-CRC (n = 47), GSE184336-GC (n = 231), GSE122401-GC (n = 80), GSE179252-GC (n = 38). B Bar plot showing the number of samples detected for each microbiome. C Survival plot showing prognosis in patients from PRJNA788008. The bacterial abundance is divided into high (total reads before 2/3) and low groups (total reads after 1/3). D Representative images showing SP stained with SP-specific FISH probe in clinical samples from patients with AEG. Red, SP; blue, cell nucleus. Scale bar, 25 μm, 100 μm. E Kaplan-Meier OS analysis stratified by SP abundance in clinical AEG and GC (n = 115). F Construct design of SP-GFP. G Flow cytometry validation of SP-GFP in tumor cells. H TEM showing intracellular SP. Red arrow, SP. Scale bar, 500 nm. I Schematic diagram showing the workflow of immune-competent mouse model. J Tumor volume and tumor weight in C57 mouse implanted MC38 cells. Mock, group without SP; SP, group with coculture SP; Data are mean ± SEM from n = 5. K Tumor volume and tumor weight in C57 mouse implanted B16F10 cells. Mock, group without SP; SP, group with coculture SP; Data are mean ± SEM from n = 5. L Tumor volume and tumor weight in BALB/c mouse implanted CT26 cells. Mock, group without SP; SP, group with coculture SP; Data are mean ± SEM from n = 6. M Tumor volume and tumor weight in C57 mouse implanted Hepa1-6 cells. Mock, group without SP; SP, group with coculture SP; Data are mean ± SEM from n = 5. N Tumor volume and tumor weight in C57 mouse implanted MC38 cells. Based on Fig. S6C, mixed MC38 cells and SP-GFP are sorted by fluorescence-activated cell sorting (FACS); Negative, group with SP-GFP- MC38 cell; Positive, group with SP-GFP+ MC38 cell; Data are mean ± SEM from n = 10. P values were determined by unpaired two-tailed t-tests unless otherwise specified. *p < 0.05, **p < 0.01, ***p < 0.001. Source data are provided as a Source data file.
The detection of SP genomic signatures following the tissue dissociation process, which would be expected to remove extracellular bacteria, suggests that the SP population we identified was physically protected within host cells. This observation provided the critical rationale for our subsequent hypothesis that SP is an intracellular bacterium. To further confirm whether SP is located in tumor cells, we engineered a recombinant strain of SP expressing green fluorescent protein (Fig. S4A). We found that CD45- cells could be detected in SP-infected tissues, whereas T cells, DCs, NK cells, B cells, and macrophages were hardly detectable (Fig. S4A). Those cells were sorted from the tumor by flow cytometry, and quantitative real-time PCR (qPCR) analysis showed that CD45− cells exhibited a high abundance of SP in the SP group, while no other immune cells were detected in vivo (Fig. S4B). To confirm the identity of CD45⁻ cells, EpCAM was used as a tumor cell marker, which validated that CD45⁻EpCAM⁺ cells displayed a higher abundance of SP (Fig. S4C). We further conducted in vitro experiments, where qPCR and flow cytometry demonstrated SP infection in tumor cells (Figs. 1F, G and S4D). In addition, immunofluorescence (IF) indicated that SP is nearly localized within the cytoplasm of tumor cells (Fig. S5). Consistent with these findings, transmission electron microscopy (TEM) revealed that SP is present within tumor cells, primarily localized in the cytosol (Fig. 1H). Finally, we constructed a SP-infected tumor cell model, injected it into mice, and the tumor was dissociated and cultured on a bacterial plate, where SP could be cultured (Fig. S6A). Tumor areas with high abundance of SP tended to exhibit increased hypoxia (Fig. S6B). These data suggest that SP is an intracellular bacterium, predominantly distributed in the cytoplasm of tumor cells.
We next sought to validate that SP infection leads to poor prognosis in GIT. To investigate this, we constructed an SP-infected tumor model (Fig. 1I). We observed that tumors harboring SP exhibited accelerated progression compared to the mock group in MC38 cells (Fig. 1J). Consistently, intracellular SP significantly promoted tumor progression in B16F10, Hepa1-6, and CT26 cells (Fig. 1K–M). To address potential interference from extracellular SP in the simple mixed coculture system, we developed a more precise approach. We cocultured MC38 cells with GFP-labeled SP for 4 h and then isolated the SP-containing tumor cells using fluorescence-activated cell sorting (FACS) (Fig. S6C). Following FACS, we obtained two distinct populations: a positive group consisting of SP-GFP+ MC38 cells and a negative group comprising SP-GFP- MC38 cells. Positive and negative MC38 cells isolated by FACS were engrafted into the right flank of C57 mice to establish tumor model. The positive group exhibited significantly enhanced tumor progression compared to the negative group (Fig. 1N). These results provide compelling evidence that intracellular SP promotes tumor progression.
Intracellular SP induces an immunosuppressive TME by impairing MHC-II+ macrophages
Intracellular bacteria establish symbiotic relationships with host cells by modulating immune responses, enabling their survival within the host13,14,18,50,51,52,56. To investigate how intracellular SP promotes GIT progression by suppressing tumor immunity, we used multiplex immunofluorescence to analyze the TME patterns in AEG with high and low intracellular SP groups (Fig. 2A). The results revealed that the proportions of macrophages (CD68+ cells), Treg cells (FoxP3+CD4+ cells), and CD8+ T cells were similar between the high and low SP groups; however, MHC-II+ macrophages were significantly decreased in the high SP group among AEG patient samples, suggesting that the presence of high levels of SP suppresses MHC-II+ macrophage function (Fig. 2B). These findings suggest that intracellular SP may inhibit macrophage function. Furthermore, intracellular SP did not promote tumor progression in MC38 cell engrafted NCG mice, indicating that its effects are dependent on host immunity (Fig. 2C, D). To further explore this, we constructed a C57 mouse model with depleted CD4+ and CD8+ T cells through the administration of anti-CD4 and anti-CD8 antibodies (Fig. 2E). Flow cytometry and quantification revealed a high deletion rate of CD4+ and CD8+ T cells in the mouse spleen (Fig. S7A and S7B). In mice with deleted CD4+ and CD8+ T cells, intracellular SP similarly promoted tumor progression compared to the group without T cell deletion (Fig. 2F). Collectively, these results indicate that intracellular SP primarily inhibits tumor progression by modulating macrophage function, which in turn suppresses T cell activity. As anticipated, the tumor-promoting effect of intracellular SP was attenuated in macrophage-depleted mice (Figs. 2G and S7C). Consistent with these findings, APC (Min/+) mice administered SP via intragastric gavage developed more adenomas compared to PBS-treated controls (Fig. 2H–J). Moreover, compared to the PBS group, the SP-treated group exhibited a lower proportion of MHC-II⁺ macrophages and a higher proportion of CD206⁺ macrophages in APC(Min/+) mice (Figs. 2K and S7D). Collectively, our data indicate that intracellular SP promotes GIT progression by suppressing macrophage function, thereby inducing an immunosuppressive TME.
A Representative images showing intratumor macrophages (CD68+), MHC-II+ macrophages (HLA_DR+CD68+), Treg cells (FoxP3+CD4+), and CD8+ cells (CD8+) in AEG patients (n = 5). CD68, orange; CD8, red; HLA_DR, green; FoxP3, pink; CD4, yellow. DAPI, blue. B Percentage of macrophages, MHC-II+ macrophages, Treg cells, and CD8+ cells between low and high SP in AEG patients. Data are mean ± SEM from n = 5. C Tumor volume and tumor weight in NCG mouse implanted MC38 cells. Control, group without mixing SP, SP, group with mixing SP; Data are mean ± SEM from n = 10. D Tumor volume and tumor weight in NCG mouse implanted MC38 cells. Mixed MC38 cells and SP-GFP are sorted by FACS; Negative, group with SP-GFP- MC38 cell; Positive, group with SP-GFP+ MC38 cell; Data are mean ± SEM from n = 10. E Schematic of anti-CD4 and anti-CD8 treatment in C57 mice based on control, group without mixing SP, SP, group with mixing SP. F Tumor volume and weight in C57 mice with/without SP after CD4 and CD8 depletion; Data are mean ± SEM from n = 10. G Tumor volume and weight in C57 mice with or without SP after macrophage depletion by clodronate; Data are mean ± SEM from n = 8. H Schematic diagram for SP (1 × 108 CFU/ per mouse) or 100 μL PBS/ per mouse in APC (Min/+) 6 weeks mouse, once per week, for Fig. 2G-K; n = 4. I Colorectum images (left) and the numbers of tumor (right) in APC (Min/+), red row, point to adenoma; n = 4. J Representative HE staining of Colorectum in control and SP group. scale bars, 500 μm. Blue row, points to dysplastic gland. K IF staining of cell nucleus (DAPI, blue), F4/80 (red), and MHC-II+ (green) in tumor tissues from APC (min/+). Percentage of MHC-II+ macrophages is quantified and compared between PBS and SP groups. Scale bar: 50 μm. Data are mean ± SEM from n = 4. P values were determined by unpaired two-tailed t tests unless otherwise specified. *p < 0.05, **p < 0.01, ***p < 0.001 (For Fig. 2F, one-way ANOVA was used). Source data are provided as a Source data file.
Intracellular SP within tumor cells decreases macrophage function by type I IFN related pathway
To further characterize macrophage regulation by intracellular SP in tumor cells, we performed single-cell RNA sequencing (scRNA-seq) of CD45+CD11b+ cells from the TME of MC38-engrafted mice using FACS. MC38 cells with or without intracellular SP were implanted in C57 mice, and tumor tissues were harvested after 7 days for scRNA-seq analysis. Unsupervised clustering identified 23 distinct clusters, classified into seven types of immune cells, including NK cells, monocytes and macrophages (Mono/Macro), fibroblasts, dendritic cells, CD8⁺ T cells, CD4⁺ T cells, and B cells (Fig. S8A–D). Consistent with SP regulating macrophage function, scRNA-seq analysis revealed that the proportion of DCs was much lower than that of the Mono/Macro cluster, and the antigen presentation module score in DCs was similar between the control and SP groups, while macrophage function in the SP group showed a suppressive trend compared to the control group (Fig. S8E and S8F). We focused on the Mono/Macro cluster, which was the most abundant in the scRNA-seq data, and re-clustering revealed 10 distinct clusters (Figs. 3A, S8G and S8H). Five of these clusters exhibited M1-like functions: the c3_Cd74 cluster indicated immune activation; the c4_H2-Aa cluster was associated with antigen presentation; the c7_S100a8 cluster and the c8_Il1b cluster displayed pro-inflammatory characteristics; and the c10_Isg15 cluster was linked to type I IFN-related responses (Fig. 3B and S8I). Three clusters exhibited M2-like functions: the c1_Mrc1 cluster displayed regulatory properties; the c2_Spp1 cluster was associated with angiogenesis; and the c5_Pf4 cluster also indicated angiogenesis. Additionally, the c9_Plac8 cluster exhibited IFN-related characteristics from monocytes, while the c6_Mki67 cluster displayed proliferative features.
A UMAP (Uniform manifold approximation and projection) plot indicates the cell distribution and clusters of macrophages. B Dot plot indicates the marker genes and cell function of each cluster, based on MonoMacro (macrophage) clusters. C Streamline of RNA velocity projected in UMAP plot indicates the different transition direction in each cluster between control (n = 2) and SP (n = 2). D Dot plot showing the Ro/e (ratio of observed to expected) of samples from control (n = 2) and SP (n = 2). P value was calculated by Fisher’s exact test, two-sided. E Flowchart indicates the difference of cell composition between control (n = 2) and SP (n = 2). F Schematic diagram for direct coculture macrophages and MC38 cells. G qPCR (left) and ELISA (right) evaluating macrophage function in direct coculture (BMDM: MC38 = 2: 1). Macrophages isolated via FACS from the direct coculture were used for qPCR analysis. Data are mean ± SEM from n = 3. H Flow cytometry image (left) and quantification (right) of macrophage function in direct coculture (BMDM: MC38 = 2: 1). Data are mean ± SEM from n = 3. I Schematic diagram for separately coculture (left) and qPCR (right); Transwell chamber is 0.45 μm. Data are mean ± SEM from n = 3. J Flow cytometry image (left) and quantification (right) of macrophage function in separate coculture (BMDM: MC38 = 2: 1). Data are mean ± SEM from n = 3. K Density line plot indicates the distribution of signature score in cell clusters. Left, signature score for type I interferon; right, signature score for antigen presentation. P value is calculated between c4_H2-Aa and other clusters using Wilcox-rank sum test, two-sided. L Density line plot indicates the distribution of signature score in cell clusters. Left, signature score for M2; right, signature score for M1. P value was calculated between c4_H2-Aa (as control group) and other clusters using Wilcox-rank sum test, two-sided. P values were determined by unpaired two-tailed t tests unless otherwise specified. *p < 0.05, **p < 0.01, ***p < 0.001. Source data are provided as a Source data file.
RNA velocity analysis of each cluster between control and SP groups revealed transitions from M1-like macrophages (c8_Il1b) to M2-like macrophages (c5_Pf4) in SP-treated samples (Fig. 3C). Quantification of cluster changes following SP group showed a significant decrease in M1-like macrophages and an increase in M2-like macrophages, with c4_H2-Aa, an antigen processing cluster, notably reduced (Figs. 3D, E, S8J and S8K). These results suggest SP suppresses macrophage function. To validate this finding, we established a co-culture model to assess macrophage function influenced by SP-treated MC38 cells (Fig. 3F). Compared to macrophages co-cultured with untreated MC38 cells, those exposed to SP-treated MC38 cells exhibited increased ARG1 and decreased CD86 expression, as measured by qPCR, and reduced IFNβ secretion, as determined by enzyme-linked immunosorbent assay (ELISA) (Fig. 3G). Flow cytometry analysis of co-cultured macrophages revealed a significant decrease in CD206-MHC-II+ macrophages and an increase in CD206+MHC-II− macrophages, confirming SP-mediated inhibition of macrophage function via tumor cells (Fig. 3H). Similar results were observed with varying tumor cell to macrophage ratios (Fig. S9A–D).
Previous studies have reported that IFNβ secretion is a host cell defense response to bacterial attack57,58,59,60. We hypothesize that SP inhibits macrophages by reducing tumor cell IFNβ secretion through weakening the cell defense response. To rule out direct macrophage interaction, we employed a 0.45 µm transwell chamber co-culture model, which demonstrated macrophage inhibition by SP, regardless of the varying MC38 cell to macrophage ratios (Figs. 3I, J and S9E–H). These findings indicate that SP in tumor cells leads to macrophage inhibition.
We assessed the response of type I IFN in tumor and immune cells using qPCR (samples from Fig. S4A), and found that SP infection in tumor cells suppressed the expression of interferon-stimulated genes (ISGs), while immune cells showed no significant differences in vivo (Fig. S9I). Furthermore, in vitro experiments demonstrated that SP infection in tumor cells also suppressed ISGs expression, while immune cells sustained high ISGs expression (samples from Figs. S4D and S9J). This contrasting pattern of ISGs expression suppressed in tumor cells yet sustained in immune cells supports the model that SP compromises macrophage function primarily by blunting type I IFN production within tumor cells, rather than through direct action on immune cells. These findings also indicate that SP in tumor cells leads to macrophage inhibition.
Reassessment of macrophage clusters by gene signature in scRNA-seq revealed that c3_Cd74 and c4_H2-Aa clusters positively correlated with type I IFN pathway and antigen presentation, while c1_Mrc1 and c6_Mki67 clusters negatively correlated in the SP group (Fig. 3K). Furthermore, c3_Cd74 and c4_H2-Aa clusters exhibited M1-like macrophage characteristics, while c5_Pf4 and c1_Mrc1 clusters displayed M2-like macrophage features in the SP group (Fig. 3L). Notably, SP treatment significantly decreased c3_Cd74 and c4_H2-Aa clusters while increasing c1_Mrc1, c6_Mki67, and c5_Pf4 clusters (Fig. 3D, E). Since DCs were the primary antigen-presenting cells in the human body, SP was neither able to affect the proportion of DC recruitment, nor to suppress MHC-II expression on DCs, whereas MHC-II expression on macrophages was significantly decreased in vivo (Fig. S10). In vitro experiments, the SP-conditioned medium (SP-CM) group with IFNa2b treatment increased MHC-II expression on DCs (Fig. S11A, B). However, the SP-CM group did not reduce MHC-II+ DC compared to the CM group, and the SP-CM-cGAMP group was unable to increase MHC-II+ DC compared to the CM-cGAMP group, regardless of whether DCs were IFNAR1 knockout (Fig. S11B). These results suggest that SP has a limited effect on DC function. Nevertheless, the SP-CM group reduced MHC-II+ macrophages and could not be rescued by cGAMP treatment compared to the CM or CM-cGAMP groups (Fig. S11C). In contrast, the SP-CM with IFNa2b treatment group rescued MHC-II+ macrophages compared to the CM group in vitro (Fig. S11C). When macrophages were IFNAR1 knockout, these effects were abolished in vitro (Fig. S11C). This suggests that SP-mediated macrophage inhibition may decrease IFN-related macrophages through an IFNβ-dependent mechanism.
Intracellular SP impairs MHC-II+ macrophage via decreasing tumor cell secretion IFNβ
Consistent with our hypothesis, IFNAR1 is required for IFNβ-mediated macrophage activation. Therefore, we generated an IFNAR1-KO mouse model and found no significant difference in tumor progression between the SP-treated and control group (Fig. 4A). Furthermore, administration of human IFNa2b, a clinically approved drug belonging to the type I IFN family with homology to mouse IFNs, failed to promote tumor progression in the SP with IFNa2b group compared to the SP group61,62,63,64 (Fig. 4B). We harvested tumors from the experiment described in Fig. 4B to assess changes in the immune TME during SP-induced tumor progression (Fig. S12A). While the overall proportion of macrophages was comparable among the three groups, an intriguing observation was that the frequency of MHC-II⁺ macrophages was significantly lower in the SP group than in the control group, and this reduction was rescued by IFNα2b supplementation (Fig. 4C, D). Moreover, the proportions of NK cells, NKT cells, CD8+ T cells, CD4+ T cells, and Tregs, as well as macrophage activation markers CD86 and CD206, showed no significant differences between groups (Fig. S12B, C and S13A–D). However, TNFα+ macrophages were suppressed in the SP group (Fig. S13E). This finding aligns with our scRNA-seq results, which revealed that SP impairs the c4_H2-Aa macrophage cluster involved in antigen processing. Although the proportion of CD8+ T cells did not differ among the three groups, the cytotoxic function of CD8+ T cells, as measured by GranzB+ and IFNγ+ expression, was markedly inhibited in the SP group (Fig. S13F). These results indicate that SP impairs MHC-II+ macrophages, leading to suppressed CD8+ T cell cytotoxic function.
A Tumor volume (left) and tumor weight (right) in a C57 mouse. IFNAR1-KO, knock out the gene of IFNAR1 in C57 mouse; Data are mean ± SEM from n = 5. B Schematic diagram for supplement IFNa2b in C57 mouse (left), tumor volume (right); Data are mean ± SEM from n = 5. C Proportion (left) and flow cytometry image (right) of macrophages in tumor tissues. Data are mean ± SEM from n = 5. D Proportion (left) and flow cytometry image (right) of MHC-II+ macrophage. Data are mean ± SEM from n = 5. E Proportion of IFNγ+, KI67+, and GranzB+CD8+ T cells based on wild type (WT) macrophage; Data are mean ± SEM from n = 3. F Proportion of IFNγ+, KI67+, and GranzB+CD8+ T cells based on IFNAR1 knockout (IFNAR1-KO) macrophage; Data are mean ± SEM from n = 3. G Schematic diagram for anti-MHC-II in C57 mouse (left), tumor volume and weight (right); data are mean ± SEM from n = 5. H Flow cytometry image (left) and Proportion (right) of IFNγ+ and GranzB+CD8+ T cells in tumor tissues (from Fig. 4G). P values were determined by unpaired two-tailed t-tests unless otherwise specified (one-way ANOVA was used to multiple group comparisons). *p < 0.05, **p < 0.01, ***p < 0.001. Source data are provided as a Source data file.
To further confirm that SP suppresses CD8+ T cell cytotoxic function, we co-cultured macrophages pretreated with CM with CD8+ T cells (Fig. S14). Macrophages pretreated with SP-CM significantly inhibited the expression of IFNγ+, GranzB+ and Ki67+CD8+ T cells compared to the CM group (Fig. 4E). Additionally, SP-CM with cGAMP treatment failed to rescue CD8+ T cell cytotoxic function compared to the CM with cGAMP treatment (Fig. 4E). However, macrophages pretreated with SP-CM and IFNa2b treatment enhanced the expression of IFNγ+ and Ki67+CD8+ T cells compared to the SP-CM group (Fig. 4E). Furthermore, IFNAR1-KO macrophages pretreated with SP-CM, with or without IFNa2b or cGAMP treatment, had little effect on CD8+ T cell cytotoxic function (Fig. 4F). To further confirm the critical role of MHC-II⁺ macrophages in this process, mice were treated with anti-MHC-II antibodies (Fig. 4G). As expected, when MHC-II function was inhibited, SP no longer exerted a significant effect on tumor progression or CD8+ T cell activity (Fig. 4G, H). In summary, our findings demonstrate that intracellular SP inhibits macrophage function by decreasing tumor cell secretion of IFNβ, resulting in immune suppression and limiting STING function.
Intracellular SP decreases IFNβ secretion via STING protein degradation
While we have established that intracellular SP induces immune suppression through the type I IFN pathway, the precise mechanism by which SP decreases IFNβ secretion requires further elucidation. To investigate this, we first conducted bulk RNA-seq analysis in the HCT116 cell. Results revealed significant downregulation of ISGs such as IFIT1, IFNB1, CXCL10, and ISG15 in the SP group compared to the control group, confirming that intracellular SP inhibits the type I IFN pathway response (Fig. 5A). To further characterize SP’s inhibitory function, we treated tumor cells with various IFN agonists in the presence or absence of SP (Fig. S15A). Notably, the expression of ISGs was markedly suppressed in the SP group treated with cGAMP, a STING agonist, while other IFN agonists showed no significant effect (Fig. 5B). In contrast, poly (I:C), which activates interferon responses through MAVS recognition, showed no significant differences in WB analysis, further highlighting that the observed effects are specifically induced by STING (Fig. 5C). Additionally, ELISA revealed reduced IFNβ secretion by tumor cells in the SP-treated, cGAMP-stimulated group (Fig. 5D). Collectively, these data support our conclusion that intracellular SP suppresses the type I IFN pathway by promoting STING protein degradation.
A Heatmap showing the decreased expression of majority of ISGs compared SP to Control group (each group, n = 4) in HCT116 cells; SP treated 4 h. B qPCR testing the expression of CXCL10, ISG15, IFNB1, and IFIT1 in HCT116 cells. PAM3, LPS, and cGAMP treat HCT116 for 3 h. PAM3, Pam3CSK4, stimulates TLR2 ligand; LPS, stimulates TLR4 ligand; poly (I:C), stimulates MAVS; cGAMP, 2′3′-cGAMP, stimulates STING. Data are mean ± SEM from n = 4. C WB analysis in HCT116 pretreated with SP for 48 h, followed by a 3 h stimulation with cGAMP (left) and poly (I:C) (right); p-IRF3, IRF3 phosphorylate; p-TBK1, TBK1 phosphorylate; t-IRF3, total IRF3; t-TBK1, total TBK1. D ELISA analysis IFNβ secretion in HCT116 treated with cGAMP for 3 h after with or without infected SP for 48 h. Data are mean ± SEM from n = 3. E WB analysis STING protein in different cell lines treated with SP for 48 h in different MOI. F IF shows SP-GFP within HCT116 cells. ER, light blue; SP-GFP, green; Nucleus, DAPI, red. Scale bar = 15 μm, 5 μm. G ELISA analysis of IFNβ secretion in STING-KO HCT116. Data are mean ± SEM from n = 3. H Flow cytometry analysis of MHC-II⁺ macrophage populations following coculture with wild-type or STING-KO HCT116 cells. Data are mean ± SEM from n = 3. I Flow cytometry analysis of CD206⁺ macrophage populations following coculture with WT or STING-KO HCT116 cells. Data are mean ± SEM from n = 3. J Tumor volume (left) and tumor weight (right) in C57 mouse. STING-KO, knock out the gene of STING in MC38 cells. WT wild type; Data are mean ± SEM from n = 5. P values were determined by unpaired two-tailed t-tests unless otherwise specified (one-way ANOVA was used to multiple group comparisons). *p < 0.05, **p < 0.01, ***p < 0.001. Source data are provided as a Source data file.
To further validate the finding above, we examined STING protein expression in OE19, MC38, SKGT4, and HCT116 cells treated with varying multiplicities of infection (MOI) of SP for 48 h. WB analysis confirmed a decrease in STING protein levels across four cell lines while cGAS protein level showed no significant difference (Figs. 5E and S15B). Intriguingly, IF revealed colocalization of SP with the endoplasmic reticulum (ER), where STING protein is typically localized in tumor cells (Fig. 5F and S15C). qPCR results showed that SP with different MOI did not affect the gene expression of STING in HCT116 (Fig. S15D). This observation provides compelling evidence for the association of SP with STING degradation.
To corroborate these findings in vivo, we utilized STING knockout (STING-KO) MC38 cells in a C57 mouse model (Fig. S15E). As expected, no significant differences were observed in IFNβ secretion, and flow cytometry analysis revealed that the proportions of MHC-II⁺ and CD206⁺ macrophages also remained unchanged in SP treatment (Fig. 5G–I). Moreover, SP failed to promote tumor progression in STING-KO MC38 cells compared with wild-type controls (Fig. 5J). This result further emphasizes the critical role of STING in SP-mediated tumor progression.
Intracellular SP activates ERAD pathway degrading STING protein
To elucidate the mechanism by which SP, localized near the ER, degrades STING protein in tumor cells, we investigated major cellular protein degradation pathways65,66. While previous studies have reported that STING degradation typically occurs through the lysosomal pathway following its physiological function, abnormal cellular states such as ER stress can trigger alternative degradation pathways, particularly the ER-associated degradation (ERAD) pathway mediated by the proteasome67,68,69,70,71,72. Enrichment analysis of proteasome and lysosome-related pathways revealed that SP treatment in HCT116 cells activates the proteasome pathway while inhibiting the lysosomal pathway compared to the control group (Fig. 6A). Furthermore, TEM showed ER swelling in SP-treated tumor cells across multiple cell lines, indicating SP-induced ER stress (Figs. 6B and S16A). IF corroborated these findings, confirming ER swelling in SP-treated HCT116 cells compared to control group (Fig. 6C, E). These results collectively demonstrate that SP induces ER stress, potentially leading to STING protein degradation.
A GSEA bubble plot of GO enrichment in Mock and SP group (n = 4) in HCT116 cells. B TEM showing SP (red row) and ER (yellow row) in HCT116 cells (based on Fig. 1F). C Representative of IF showing ER treated 24 h with or without SP (MOI = 0.01) in HCT116 cells; Nucleus, DAPI, red; Light blue, ER; SP is without GFP marker; White row, points to ER stress (ER swelling); Scale bar = 15 μm. D Representative of IF showing p-STING in HCT116. p-STING, STING phosphorylate; Nucleus, DAPI, red; Vehicle, without treating 4 h with cGAMP. Mock, without treating SP; SP, treating 24 h with SP (MOI = 0.01); SP is without GFP marker. E, F Quantification of ER stress and p-STING in HCT116 corresponding to (C, D). Data are mean ± SEM from n = 4. G WB analysis p-PERK, PERK, IRE1a, ATF6, and XBP1s protein with SP (MOI = 0.01) for 48 hours in HCT116 cells. H–J WB analysis of STING protein with CHX, MG132, CB5083, BafA1 or chloroquine after SP treatment. K Co-IP analysis interaction STING with SEL1L, treated by SP (MOI = 0.01) and MG132 (25 µM), for 48 h in HCT116 cells; IP, STING. L WB analysis STING protein, treated by SP (MOI = 0.01) and SEL1L-KO, for 48 h in HCT116 cells. SEL1L-KO, SEL1L knockout in HCT116 cells. M WB analysis p-PERK, PERK, IRE1a, ATF6, and XBP1s protein with SP (MOI = 0.01) for 48 h in WT or SEL1L-KO HCT116 cells. N ELISA analysis IFNβ secretion in WT or KO HCT116 cells treated with cGAMP for 3 h after infected SP (MOI = 0.01) for 48 h. Data are mean ± SEM from n = 3. O Schematic diagram for coculture SP with SEL1L-KO MC38 in vivo. P Proportion (left) and flow cytometry image (right) of MHC-II+ macrophage. Data are mean ± SEM from n = 3 P values were determined by unpaired two-tailed t-tests unless otherwise specified (one-way ANOVA was used to multiple group comparisons). *p < 0.05, **p < 0.01, ***p < 0.001. Source data are provided as a Source data file.
To determine whether SP-induced STING degradation occurs via the proteasomal or lysosomal pathway, we examined phosphorylated STING (p-STING) levels, which typically indicate normal lysosomal degradation following STING activation. IF analysis revealed lower p-STING expression in SP-treated tumor cells compared to controls, particularly upon cGAMP stimulation (Fig. 6D, F). The expression of the marker proteins (PERK, IRE1a, ATF6 and XBP1s) for ER stress was increased in the SP group (Fig. 6G). To distinguish between inhibition of STING synthesis and enhanced degradation, we employed the proteasomal inhibitor MG132 and the protein synthesis inhibitor cycloheximide (CHX). Treatment with MG132 rescued STING protein levels, confirming that SP induces STING degradation via the proteasome pathway (Fig. 6H). Other proteasome inhibitor CB-5083 effectively rescued SP-induced STING degradation in HCT116 cells, whereas lysosome inhibitors (BafA1 and chloroquine) showed no rescue effects (Fig. 6I, J).
A previous study reported that the ERAD pathway can degrade STING protein via SEL1L in the proteasome70. Co-immunoprecipitation (Co-IP) experiments suggested the interaction between SEL1L and STING protein in SP-treated tumor cells compared to control (Fig. 6K). Moreover, Co-IP revealed that MG132 increased the interaction between STING and SEL1L during SP treatment (Fig. 6K). Corroborating this finding, knockdown or knockout of SEL1L expression resulted in reduced STING degradation in the SP-treated group (Figs. 6L; S16B and S16C). Although SEL1L knockout failed to degrade the STING protein, it was shown to induce ER stress, demonstrating that SP-mediated degradation of STING protein through ER stress depended on SEL1L (Fig. 6M). ELISA results demonstrated that SP failed to suppress cGAMP-induced IFNβ secretion in SEL1L-knockout HCT116 cells, suggesting that SEL1L is essential for SP-mediated inhibition of STING-dependent type I interferon signaling (Fig. 6N). Intriguingly, STING protein levels were also reduced, and STING rarely colocalized with the ER in SP-treated cells even without cGAMP stimulation, suggesting that intracellular SP decreases basal STING protein levels rather than inhibiting STING activation (Figs. S16D and 16E). To further confirm the relationship between STING and the ER, we examined STING protein expression and its association with SP in the presence of ER stress attenuators (Fig. S16F). As anticipated, treatment with ER stress attenuators rescued the SP-induced STING degradation, supporting our conclusion.
We next investigated whether SEL1L is essential for SP-mediated suppression of macrophage function via IFNβ reduction. In coculture experiments, SP treatment in SEL1L-knockout MC38 cells failed to reduce the proportion of MHC-II+ macrophages or elevated CD206+ macrophages across varying coculture ratios of BMDMs and tumor cells, demonstrating that SEL1L is required for SP-dependent macrophage suppression (Fig. S17A). Consistent with the coculture experiment findings, in vivo experiments using SEL1L-knockout MC38 tumors revealed no significant difference in macrophage MHC-II expression between SP-treated and untreated groups (Fig. 6O, P). Notably, DC proportions and functionality remained unaffected by SP or SEL1L knockout, suggesting that SP mainly impairs macrophage function within the TME (Fig. S17B)
These results indicate that SP-induced STING degradation primarily targets basal STING levels and depends on the SEL1L-mediated proteasomal pathway. In conclusion, our findings demonstrate that intracellular SP activates the ERAD pathway via the SEL1L-proteasome axis to degrade STING protein, thereby modulating the type I IFN response in tumor cells.
Intracellular SP impairs immunotherapy efficacy by reducing MHC-II+ macrophages
We next investigated whether intracellular SP within tumor cells influences the efficacy of immunotherapy. The effect of anti-PD-1 therapy was significantly impaired in the SP with anti-PD-1 group (Fig. 7A). To elucidate the underlying mechanisms, we conducted flow cytometry analysis of tumors harvested from the experiment described in Fig. 7A. The proportions of CD45+ cells and macrophages were similar across all four groups (Figs. 7B and S18A, B). Compared to the control group, NKT cells were decreased in the SP and SP with anti-PD-1 groups, while NK cell populations remained similar (Fig. S18C). Assessment of macrophage function revealed that the SP and SP with anti-PD-1 groups secreted less TNFα compared to the control group, while the proportions of CD206+ or CD86+ macrophages showed no significant differences among groups (Fig. S18D–F). Notably, the proportion of MHC-II+ macrophages was similar between the SP and SP with anti-PD-1 groups (Fig. 7C). These results indicate that SP, even in combination with anti-PD-1 therapy, impairs macrophage function, particularly MHC-II+ macrophages, explaining the lack of therapeutic effect of anti-PD-1 in the SP group. The proportions of CD4+ T cells, CD8+ T cells, and Tregs showed no significant differences among the four groups, except for a reduction in CD8+ T cell infiltration in the SP with anti-PD-1 group (Fig. S19A and S19B). IFNγ+ CD8+ T cells were similar across all groups and Ki67+ CD8+ T cells were similar between the SP and SP with anti-PD-1 group (Fig. 7D, E). These findings suggest that T cell cytotoxic function is impaired in the SP and SP with anti-PD-1 groups, likely due to the inhibition of MHC-II+ macrophages. Importantly, the addition of IFNa2b to SP or SP with anti-PD-1 groups failed to promote tumor progression and restored the anti-tumor effect of anti-PD-1 (Fig. 7F). The therapeutic efficacy of the oxaliplatin plus anti-PD-1 combination in SP group was reduced compared to the anti-PD-1 alone group (Fig. 7G). In summary, these findings indicate that gastrointestinal tumors harboring intracellular SP are associated with poor treatment outcomes, primarily due to impaired immunotherapy efficacy. Overall, tumors containing intracellular SP exhibit resistance to immunotherapy by suppressing MHC-II⁺ macrophages, thereby weakening T cell cytotoxic function.
A Schematic diagram for anti-PD-1 antibody treatment in C57 mouse (left) and tumor volume or weight (right); SP group (MOI = 0.01); Data are mean ± SEM from n = 5. B Proportion of CD45+ cells and macrophages (CD45+CD11b+F4/80+) in tumor tissues; Data are mean ± SEM from n = 5. C Proportion (left) and flow cytometry image (right) of MHC-II+ macrophages in tumor tissues; Data are mean ± SEM from n = 5. Proportion (D) and flow cytometry image (E) of IFNγ+, KI67+, and GranzB+CD8+ T cells in tumor tissues; Data are mean ± SEM from n = 5. B–E Those tumors are collected for flow cytometry from (A). F The treatment regimen for anti-PD-1 therapy in C57 mice implanted with SP-infected MC38 cells, with supplemental IFNα2b. After 8 days of MC38 cell implantation, daily intraperitoneal (i.p.) injections of 10 μg of IFNα2b per mouse were administered. Anti-PD-1 treatment began 6 days after implantation, with i.p. injections of 100 μg per mouse administered three times in total. Tumor volume (left) and tumor weight (right) were measured; Data are mean ± SEM from n = 5. G Schematic diagram for oxaliplatin and anti-PD-1 treatment in C57 mouse (left) and tumor volume or weight (right); Oxp, oxaliplatin; Data are mean ± SEM from n = 5 One-way ANOVA was used to multiple group comparisons; 7 G’s p values were determined by unpaired two-tailed t tests unless otherwise. *p < 0.05, **p < 0.01, ***p < 0.001. Source data are provided as a Source data file.
Ablation of intracellular SP by levofloxacin eliminates therapeutic resistance
Given that GIT with SP leads to therapeutic resistance and poor prognosis compared to tumors without SP, we investigated potential therapeutic strategies. Previous studies have reported that levofloxacin can ablate SP73,74. We observed significant suppression of SP growth under various concentrations of levofloxacin, with an optimal concentration of 5 µg/mL (Fig. S20A). After mixing SP with tumor cells for 24 h, the addition of levofloxacin inhibited intracellular SP across different cell lines (Fig. S20B). Results showed that levofloxacin decreased SP within tumor cells and alleviated ER swelling (Fig. 8A). We further conducted mouse rescue experiments, demonstrating that levofloxacin treatment mitigated SP-promoted MC38 tumor progression in C57 mice compared to the SP group without levofloxacin (Fig. 8B).
A TEM for SP (red row) and ER (yellow row) in infected-SP tumor cells, in both control and 5 μg/mL levofloxacin (Lev) treated. B Schematic diagram (left) illustrating levofloxacin treatment in C57 mice, with SP infection. After 4 days of MC38 cell implantation, levofloxacin was administered via intragastric gavage (i.g.) at 50 mg/kg per dose, every other day, for a total of 5 doses. Tumor volume and weight were recorded; Data are mean ± SEM from n = 5. C Schematic diagram (left) illustrating anti-PD-1 treatment in C57 mice, with or without levofloxacin treatment. After 4 days of MC38 cell implantation, levofloxacin was administered via i.g. at 50 mg/kg per dose, every other day, for a total of 5 doses. Tumor volume and weight were recorded; Data are mean ± SEM from n = 5. D Proportion (left) and flow cytometry image (right) of macrophage or MHC-II+ macrophage based on C tumor tissue; Data are mean ± SEM from n = 5 P values were determined by unpaired two-tailed t tests unless otherwise specified (one-way ANOVA was used to multiple group comparisons). *p < 0.05, **p < 0.01, ***p < 0.001. Source data are provided as a Source data file.
qPCR analysis of tumor tissues from Fig. 8B revealed a marked reduction in SP abundance following levofloxacin administration (Fig. S20C). As expected, levofloxacin reversed anti-PD-1 therapeutic resistance in tumors with SP (Fig. 8C). Flow cytometry analysis of tumors from the experiment in Fig. 8C revealed that levofloxacin treatment combined with anti-PD-1 therapy restored MHC-II expression in macrophages compared to anti-PD-1 monotherapy (Figs. 8D and S20D). These results indicate that levofloxacin may serve as a potential treatment strategy against SP-positive GIT.
Discussion
The complex association between microbiota and tumors has been extensively confirmed, with implications for tumor oncogenesis, progression, TME, therapeutic efficacy, and patient prognosis17,18,41,42,43,44,45,46,47,48,49,50,51,52,56. Advancements in detection techniques have deepened our understanding, progressing from gut microbiota to intratumor bacteria and finally to intracellular bacteria47,48,50. While some studies have demonstrated that intracellular bacteria contribute to tumor metastasis in breast cancer and impair antitumor immunity in esophageal squamous cell carcinoma, the presence and impact of intracellular bacteria in GIT remained largely unexplored14,50. Here, we have demonstrated that SP resides within tumor cells and is associated with poor prognosis in GIT. Moreover, GIT harboring intracellular SP exhibited TME suppression and impaired response to immunotherapy. Importantly, we found that levofloxacin could alleviate tumor progression and therapy resistance caused by intracellular SP. These findings suggest that intracellular SP may explain the poor therapeutic response and prognosis observed in some GIT patients, emphasizing the potential of intracellular SP as both a risk factor and a therapeutic target (Fig. 9).
A proposed model illustrating the association between intracellular SP, STING signaling suppression, altered macrophage function, and tumor progression in gastrointestinal tumors.
SP commonly colonizes various parts of the human body and has been associated with tumor progression15,17,18. Previous studies have reported notable enrichment of SP in cholangiocarcinoma compared to paired normal tissues75. Another study revealed that lung cancer with a high abundance of SP exhibited enhanced progression via upregulation of the gene expression of HDAC518. Consistent with these findings, our research demonstrates that SP directly promotes tumor progression.
To date, little has been known about how intracellular bacteria regulate the TME. Deborah et al. conducted a comprehensive analysis of the tumor microbiome across 1526 tumors and their adjacent normal tissues in seven cancer types, and confirmed that intratumor bacteria are primarily intracellular in cancer and immune cells, affecting the response to immunotherapy13. Furthermore, Fu et al. validated the presence of intracellular microbiota within tumor cells in breast cancer14. The presence of intracellular bacteria within tumor cells is increasingly recognized as a common occurrence14,17,50,51,52. In our results, intracellular SP promoted GIT progression primarily through direct intracellular colonization rather than systemic immune modulation. Consistent with this mechanism, SP infection enhanced adenoma formation and reduced the proportion of MHC-II⁺ macrophages in APC (Min/+) mice, indicating localized immune suppression driven by intracellular infection and blockade of IFN signaling. Defective IFN pathways and altered ERAD processes within tumor cells may facilitate SP persistence and STING degradation, thereby promoting immune escape. Our findings align with those of Liu et al., who reported that SP within hepatic stellate cells induced cell senescence and hepatocarcinogenesis17. In summary, we have identified and validated the presence of intracellular SP in GIT tumor cells and demonstrated its role in promoting tumor progression.
Intratumor bacteria have emerged as important regulators of the TME, particularly in their cross-talk with macrophages3,13,51,56,76,77,78,79. One study has shown that tumor-associated bacteria can impair toll-like receptor signaling, leading to T cell anergy and an increase in myeloid-derived suppressor cells and M2 macrophage differentiation76. Moreover, butyrate produced by intratumor bacteria has been shown to inhibit HDAC2 and increase H3K27 acetylation at the H19 promoter, inducing M2 macrophage polarization77. These results validate that certain bacteria can promote tumor progression by inhibiting macrophage anti-tumor functions. While compelling evidence supports the role of macrophages in mediating SP tumorigenic potential through interactions between STING and IFNβ, we also evaluated other potential factors, including other immune cell populations. NKT cells are unique lymphocytes that bridge innate and adaptive immunity, leveraging features of both T cells and NK cells80. Intriguingly, we found that the numbers of macrophages, CD4+ T cells, NK cells, NKT cells, and CD8+ T cells did not change significantly after SP infection. In summary, our findings indicate that intracellular SP suppresses macrophage function and promote tumor progression in GIT.
STING protein primarily recognizes bacterial DNA and protects against bacterial infections in host cells32,33,42. However, numerous studies have shown that bacteria often degrade or antagonize STING protein to survive within host cells31,44. Consistent with these findings, our results demonstrate that SP induces the degradation of STING protein and inhibits IFN pathway. Notably, we found that intracellular SP primarily degrades nascent STING protein. This aligns with the findings of Ji et al., who revealed that the SEL1L-HRD1 protein complex targets nascent STING protein for proteasomal degradation70. Previous studies have shown that abnormal degradation of STING protein can be driven by ER stress67,70. It is important to note that there are other mechanisms involved in the inactivation or degradation of STING protein68,69,71,72. For instance, Gentili et al. reported that an endosomal sorting complex required for transport (ESCRT) complex containing HGS, VPS37A, and UBAP1 promotes STING degradation and terminates STING-mediated signaling through STING oligomerization, increasing its ubiquitination by UBE2N and forming a platform for ESCRT recruitment at the endosome68. Zhang et al. found that Gal-9 carbohydrate recognition domain 1 interacts with the STING C-terminus, enhancing TRIM29-mediated K48-linked ubiquitination of STING and accelerating its degradation71. Some researchers have even identified AK59 as a STING degradation drug that leverages HERC4, a HECT-domain E3 ligase69. Our results provide new insights into how SP degrades nascent STING protein in the proteasome through ER stress. In addition, SP-mediated suppression of STING represents a distinct immune evasion strategy among tumor-associated bacteria. Whereas Fusobacterium nucleatum activates β-catenin or inhibits cGAS-STING signaling and Salmonella Typhimurium enhances immunity via inflammasome activation, SP reduces STING protein abundance, diminishing IFN production and MHC-II+ macrophage activation. This suppression contributes to immune escape and resistance to immunotherapy81,82,83. Moreover, we found that non-STING-dependent type I IFN pathways, including MAVS and TLR4 ligands, failed to impair the expression of ISGs. Through extensive experiments, we have demonstrated that SP within tumor cells exists in GIT, induces STING protein degradation, decreases IFNβ secretion, suppresses the anti-tumor function of macrophages, and promotes tumor progression.
Although our study reveals a clear role for intracellular bacteria in promoting tumor progression, some limitations remain. Firstly, the decontamination strategy used in sequencing-based bacteria discovery may result in false positive findings due to the absence of dedicated controls and reagent blanks. Multiple technologies including FISH could be utilized to cross-validate these findings. Secondly, the fate of intracellular SP (multiplication or death) during tumor cell proliferation, and whether it lives in symbiosis with tumor cells remains to be fully elucidated. Thirdly, our attempts to culture live SP from fresh human tumor samples have been unsuccessful to date, possibly due to the complexity of the GIT microbiome or the potential death of intracellular SP. Fourthly, our experiment based on mixing SP with tumor cells may limit the generalizability of our results. Although extensive washing steps were implemented in the SP infected tumor cell model, residual extracellular SP persists as a notable technical limitation, allowing only for partial minimization of its interference rather than complete elimination. Fifthly, macrophage functions triggered by intracellular SP may be involved in other aspects of SP-driven progression beyond decreasing IFNβ secretion. Further exploration is needed, such as investigating whether intracellular SP promotes tumor cell proliferation, as suggested by trends observed in NCG mice. Finally, Levofloxacin effectively cleared intracellular SP and restored antitumor immunity, although its broader use requires careful consideration of microbiota disruption and antibiotic resistance84,85,86. These findings highlight the potential of targeted antimicrobial or STING-stabilizing strategies and the value of microbial profiling for identifying SP-positive patients.
In summary, our study supports the notion that intracellular SP promotes tumor progression and may help explain the poor response of some GIT patients to clinical treatments. Additionally, intracellular SP may serve as a potential prognostic marker and therapeutic target in GIT.
Methods
Cell culture
OE19 (cat: CL-0754, Procell), SKGT4 (cat: CL-0874, Procell), HCT116 (CL-0096, Procell), DLD1 (cat: CL-0074, Procell), B16F10 (cat: CL-0319, Procell), CT26 (cat: CL-0071, Procell) and Hepa1-6 (cat: CL-0105, Procell) and MC38 (cat: iCell-m032, iCell) are purchased from company. DMEM or RPMI-1640 basic medium consisting of 10% fetal bovine serum (FBS, Procell, 164210-50) and 1% Penicillin-Streptomycin (Gibco, 15140122) cultures all cell line under 37 °C with 5% CO2 atmosphere.
Bacteria culture
SP (cat: BNCC342633, BNCC) is purchased from BNCC, but SP-GFP is a generous gift from Professor Xiaoyu Hu at Jilin University16. SP was revived on LB (Beef powder, 3.0 g; peptone, 10 g; sodium chloride, 5.0 g; distilled water, 1.0 L; if solid medium, adding agar, 20.0 g) medium at 37 °C for 24 h. An oscillating incubator at 180 round/minute in the aerobic environment cultures SP within liquid LB.
We determine the numbers of SP by OD600. We use PBS to gradient-dilute SP on Eppendorf (EP) tube and test each EP tubes the value of OD600 by ultra micro spectrophotometer (KAIAO TECHNOLOGY, k5800). Each EP tubes are further gradient-diluted, and calculate the numbers of SP by microscope and the CFU of SP by inoculation plate. Linear fitting calculates 1 OD600 approximately equal to 1 × 109 CFU/mL87. Because we test the cultured SP directly and chose to underestimate the number of SP to reduce the disturbance caused by bacterial excess cells, so 1 CFU is viewed as 1 SP.
Mouse experiments
All mouse experiments are conducted in Guangzhou LingFu TopBiotech. Co., LTD under permit number SYXK (Yue) 2023-0315, are conducted in line with the regulations for the Administration of Affairs Concerning Experimental Animals issued by the Ministry of Science and Technology of the People’s Republic of China. C57, nude, and BALB/c mice, aged 4-8 weeks, were purchased from Guangdong medical laboratory animal center. NCG and APC (min/+) mice, female, aged 4-8 weeks, are purchased from GemPharmatech (Guangdong, China).
For most experiments assessing tumor volume changes, tumor cells were pre-incubated with SP at an MOI of 0.01 for 4 h prior to injection. 1 × 106 MC38 cells are subcutaneously injected to C57 mouse with age-matched female to construct subcutaneous GIT murine model. Other murine tumor model including CT26, B16F10 and Hepa1-6, are subcutaneously injected with similar conditions in corresponding mouse. NCG mouse is subcutaneously injected with 5 × 105 MC38 cells. For the spontaneous colorectal cancer murine model, APC (min/+) mice is used to illustrate SP promoting GIT progression. SP (1 × 108 CFU/ per mouse) or PBS (100 μL/per mouse) are administered by gavage in APC (min/+) 4-8 weeks mouse, on once per week. For treatment with antibiotics (oral administration), the control group is gavaged with 200 μL PBS per mouse while the treatment group with the same volume of levofloxacin (5 mg/mL) per mouse). For depletion of macrophages, mice were intraperitoneally injected 200 μL clodronate liposome (cat: 40337ES08, YEASEN) once 3 days before xenograft MC38 cells, and 100 μL once a week after xenograft MC38 cells in C57 mouse. For deletion of CD4+ and CD8+ T cells, anti-CD4 and anti-CD8 antibodies (respectively 10 mg/kg per mouse) were intraperitoneally injected in C57 mouse. We recorded tumor volume every two or three days using a vernier caliper (tumor volume = 1/2 × length × width2). At the experimental endpoint, mice were euthanized by cervical dislocation following anesthesia.
qPCR assay
EZ-press RNA Purification Kit (cat: B0004D, EZBioscience) was used for total RNA extraction. The reverse transcribed system based on Hiscript III Reverse Transcriptase (cat: R302-01, Vazyme), and 1 μg of RNA in 20 μL system was applied to reverse cDNA. RuiBiotech Co synthesized all primers. BIORAD CFX96 Touch with a 2× ChamQ Universal SYBR qPCR Master Mix (Vazyme) was conducted. qPCR was performed in the reaction conditions: 95 °C for 30 second (s), followed by 40 cycles of 95 °C for 10 s and 60 °C for 30 s. The data was presented as a fold change normalized to the control group. In technical duplicates, we conducted all qPCR for each sample with three biological replicates. Primers are provided in the Supplementary Table 1.
Western blot
The proteins were extracted from the cells using RIPA lysis buffer (cat: CW2333, Beyotime Biotechnology) with a protease inhibitor cocktail (cat: B14001, Selleck). Pierce BCA Protein Assay Kit (cat: 23227, Thermo Scientific) was used to measure the protein concentration of each sample. Total protein (30 μg) was resolved using a 10% sodium dodecyl sulphate-polyacrylamide gel (SDS-PAGE) electrophoresis and transferred onto polyvinylidene difluoride (PVDF) membranes. 5% nonfat milk in PBST blocked the PVDF membrane at room temperature for 1 h and incubated with primary antibodies overnight at 4 °C. Membranes were incubated with secondary antibodies for 1 h. After washing with PBST, proteins were visualized by Omni-ECL™Femto Light Chemiluminescence Kit (cat: SQ201, EpiZyme) in a chemiluminescence imaging system ImageQuant LAS 500 (Cytiva, USA). Some proteins were detected by through Odyssey CLX. The primary antibodies included anti-β-tubulin (cat: ARG62347, Arigo), anti-GAPDH (cat: 2118, Cell Signaling Technology), anti-IRF3 (cat: 4302, Cell Signaling Technology), anti-phosphorylated IRF3 (cat: 4947, Cell Signaling Technology), anti-TBK1 (cat: 3504S, Cell Signaling Technology), anti-phosphorylated TBK1 (cat: 5483S, Cell Signaling Technology), anti-STING (cat: 13647, Cell Signaling Technology) and anti-phosphorylated STING (cat: 50907S, Cell Signaling Technology), anti-SEL1L (cat: 29801-1-AP, Proteintech), anti-Phospho-PERK (cat: 3179 T, Cell Signaling Technology), anti-PERK (cat: 3192 T, Cell Signaling Technology), anti-cGAS (cat:79978, Cell Signaling Technology), anti-IRE1α (cat: 3294, Cell Signaling Technology), anti-ATF-6 (cat: 65880, Cell Signaling Technology), anti-XBP-1s(cat: 40435, Cell Signaling Technology). The secondary antibodies included Goat anti-Mouse IgG(H + L)-HRP (cat: RM3001, Beijing Ray Antibody biotech), Goat anti-Rabbit IgG(H + L)-HRP (cat: RM3002, Beijing Ray Antibody biotech), Anti-Mouse IgG (cat: ab150116, ab150113, ab150080, ab150077, Abcam). The working dilutions of all antibodies are listed in Supplementary Table 2.
Elisa
The cell supernatant was collected for testing IFNβ secretion through LumiKine™ Xpress hIFN-β 2.0 ELISA kit (cat: luex-hifnbv2, Invivogen) and LumiKine™ Xpress mIFN-β 2.0 ELISA kit (cat: luex-mifnbv2, Invivogen). OD450 was read using Multiskan GO Luminometer (Thermofisher Scientific, USA).
Flow cytometry analysis and cell sorting
Single cell suspensions were stained with fluorochrome-conjugated antibodies against markers based on the manufacturer’s instructions. Flow cytometry was performed on the CytoFLEX (Beckman, USA) and FACSymphony S6 (Becton Dickinson, USA). All data were analyzed by FlowJo V10 or CytExpert software.
Mix tumor cells were SP-GFP cocultured MC38 tumor cells, and were sorted out negative tumor cell (without intracellular SP) and positive tumor cell (with intracellular SP) by FACSymphony S6.
Tumor tissues were harvested and minced into fragments of 8 mm3 and digested for 30 minutes at 37 °C with 1 mg/mL of collagenase IV (cat: 40510ES60, Yeason) and 0.2 mg/mL of DNase I (cat: D5025, Lablead). A 70 mm cell strainer filtered with the dissociated tissues. Single-cell suspensions were incubated in an Fc block (cat: 553141, Becton Dickinson) and Zombie Aqua™ Fixable Viability Kit (cat: 423101, Biolegend) for 15 min at room temperature, stained for surface markers at 4 °C for 30 min, washed with Cell Staining Buffer (cat: 420201, Biolegend) once. The intracellular staining used the Intracellular Fixation & Permeabilization Buffer Set (cat: 88-8824-00, Invitrogen&trade). The intranuclear staining used the eBioscience™ Foxp3/Transcription Factor Fixation/Permeabilization Concentrate and Diluent Kit (cat: 00-5521-00, Invitrogen&trade). These antibodies including Brilliant Violet 510™ anti-mouse CD45 (cat: 103108, BioLegend), APC anti-mouse/human CD11b (cat: 101228, BioLegend), Brilliant Violet 785™ anti-mouse F4/80 (cat: 123141, BioLegend), FITC anti-mouse NK-1.1 (cat:108706, BioLegend), APC/Cy7 anti-mouse CD3 (cat: 100221. BioLegend), PE/Cy7 anti-mouse CD4 (cat: 100422, BioLegend), Alexa Fluor700 Rat Anti-Mouse CD8a (cat:557959, Becton Dickinson), Alexa Fluor® 647 anti-mouse/rat/human FOXP3 (cat: 320014, BioLegend), Pacific Blue™ anti-human/mouse Granzyme B (cat: 515408, BioLegend), Brilliant Violet 605™ anti-mouse IFN-γ (cat: 505840, BioLegend), PE anti-mouse Ki-67 (cat: 103105, BioLegend), PE/Cyanine7 anti-mouse CD206 (cat: 141720, BioLegend), Brilliant Violet 421™ anti-mouse CD86 (cat: 105123, BioLegend), Alexa Fluor® 647 anti-mouse TNF-α (cat: 506314, BioLegend), Alexa Fluor® 700 (or other channel) anti-mouse I-A/I-E (cat: 107622, 107616, 107625, 107613, BioLegend), and APC anti-mouse CD326 (Ep-CAM) (cat: 118213, BioLegend) were used to stain.
Immunofluorescence staining
Immunofluorescence staining was performed by Absin Biotech (Shanghai, China). All image acquisition and quantitative analyses were performed by Absin Biotech (Shanghai, China), blinded to sample grouping and experimental purpose.
To confirm the localization of SP within tumor cells, cocultured SP-GFP tumor cells grown on confocal dishes were stained with AF647-Phalloidin (Cat. A22287, Invitrogen) recognizing F-actin and counterstained with DAPI. For mechanistic studies, tumor cells with or without SP-GFP coculture were labeled with ER-Tracker™ Blue-White DPX (Cat. E12353, Invitrogen) to visualize the endoplasmic reticulum, fixed and permeabilized using the corresponding buffer, and stained for STING and phospho-STING with primary and fluorescent secondary antibodies. All fluorescence images were captured using a Zeiss LSM 900 laser confocal microscope.
SP testing by FISH
16S rRNA was used to identify whether SP located in tumor by FISH. The FISH probe of SP (5′-GTCGTCCAGTATCCACTGC-3′) and the protocol of operation was referred to a previous study17. The intensity of the fluorescence signal was quantified using imageJ.
Transmission Electron Microscopy (TEM)
The cells were collected by electron microscope fixative (car: G1102, Servicebio). After a serial procession of agarose pre-embedding, fix, dehydrate, resin penetration and embedding, polymerization, ultrathin section and staining, the samples were observed under TEM (HT7800, Hitachi) and take images.
Macrophages induction
Bone marrow cells were extracted from the femurs and tibias of C57 mice and depleted of erythrocytes by Red Blood Cell Lysis Buffer (cat: G2015-500ML, Servicebio). Seeding in 6-well plates at a density of 2 × 106 cells/well cultured for 7 days in DMEM medium containing 10% FBS and 20 ng/mL macrophage colony stimulating factor (M-CSF).
HE staining
FFPE was treated by dewaxing and hydration, rewarming and fixing, and hematoxylin staining. Then, we put sections into hematoxylin solution for 5 min, treat the section with hematoxylin differentiation solution, and place the sections in 95% ethanol for 1 min, eosin dye for 15 s. Lastly, the section was conducted by dehydration and sealing.
Immunoprecipitation cell lysis buffer
The cells were washed with ice-cold PBS and lysed in immunoprecipitation buffer (cat: G2038-100ML, Servicebio) with a protease inhibitor mixture (cat: G2007, Servicebio) for 30 min at 4 °C. The lysates were centrifuged at 12,000 rpm for 10 min at 4 °C, and the protein was determined with the BCA kit. The proteins incubated with Protein A/G beads (cat: IP05, Millipore) shaken at 4 °C for 1 h, and then incubated 1 μg antibody rotated overnight at 4 °C. Immunoprecipitates were washed thoroughly with immunoprecipitation buffer and immunoblotted with the antibodies.
Microbiome detection
The microbiome abundance in each dataset was detected by the Kraken software suite (Kraken2 version 2.1.2, Bracken version 2.9)88. Raw reads were quality-controlled by fastp (version 0.23.4) to generate clean reads89. The “Standard” collection (version 26/9/2022) from Kraken index database which includes archaea, bacteria, viral, plasmid, human and UniVec_Core was used to annotate the clean reads. Annotated reads were then normalized to reads per million microbiome reads and reads per million total reads. Relative abundance of each microbiome was estimated by Bracken. The background microbiome data from the SAHMI was used to detect the enriched microbiomes in each dataset90,91. During our analysis, in addition to the microbiome genomes, the human genome, universe vectors and other sequences were also included in the database which may reduce the potential false positive rate. The background microbiome data from SAHMI was used in our analysis to reduce the potential false positive findings, which may reduce our findings if the background data had a high positive false rate. Six publicly available datasets were included in this study: GSE126304 (n = 46) and PRJNA788008 (n = 83) for AEG, GSE156451 (n = 47) for CRC, GSE122401 (n = 80), GSE179252 (n = 38) and GSE184336 (n = 231) for GC.
Single cell RNAseq analysis
The isolated single cells were loaded to the single-cell chip for GEM generation using the 10x Chromium controller to construct single cell library. The single-cell 3′ cDNA libraries were constructed using the Chromium Single Cell 3′ Reagent Kit (V3 chemistry, 10x Genomics) and sequenced on Illumina NovaSeq 6000 platform.
The raw single cell fastq data were processed by Cell Ranger pipelines (10x Genomics, version 6.0.1) to generate the count matrix. Reference genome (version refdata-gex-mm10-2020-A) was used for read alignment92. Then quality control was performed by filtering cells expressing less than 1000 genes or more than 6000 genes. Cells with more than 10% transcripts from mitochondrial genes were also filtered out. Doublets were identified by R package DoubletFinder (version 2.0.3) and filtered from the count matrix93.
After quality control, R package Seurat (version 4.0.4) was applied for downstream analysis. Gene expression was normalized, and highly variable genes were identified by SCTransform function with default parameters. Based on highly variable genes, principal component analysis was performed by RunPCA function94. Function RunHarmony from R package harmony (version 0.1.0) was applied to remove the potential batch effects between batches or samples. After batch correction, we used the Elbow function to identify the significant corrected components for cell clustering95. The identified 34 PCs with resolution 1.2 and 16 PCs with resolution 0.6 were used for major cell clustering and the macrophage clustering respectively. For visualization, we performed the dimension reduction to generate the Uniform Manifold Approximation and Project (UMAP) via the RunUMAP function. During the major cell clustering, we annotated the cell clusters manually with canonical cell type-specific markers, including B cells (Ms4a1, Cd79a, Cd79b), T cells (Cd3d, Cd3e, Cd3g), CD4 T cells (Cd4), CD8 T cells (Cd8a), NK cells (Ncr1, Ncam1, Prf1, Klrd1), DC (Clec9a, Cd209a, Xcr1, Ccr7, Fscn1, Ccl22), monocytes or macrophages (Itgam, Lyz2, Cd63, Csf1r, Cd14), fibroblast (Dcn, Col3a1), endothelial cells (Eng, Kdr), epithelial cells (Krt8, Epcam). During the cell clustering of macrophages or monocytes, cell clusters were annotated based on the cluster markers and their functionality.
RNA velocity
To detect the potential cell transition between each cluster from different groups, we performed RNA velocity analysis via velocyto.py (version 0.17.17) and scVelo (version 0.3.1)96,97. Briefly, loom files containing the spliced and unspliced transcript count matrix were generated by velocyto.py with parameter “run10x”. Generated loom files were then loaded into scVelo and preprocessed by scVelo.pp.filter_and_normalize function. The moments of normalized spliced and unspliced counts were calculated using the scVelo.pp.moments function with default parameters. RNA velocity was estimated using the scVelo.tl.velocity function with “dynamical” model and scVelo.tl.velocity_graph function was used to build the velocity graph. For visualization, the estimated RNA velocities were projected into the UMAP coordinates generated by RunUMAP from cell clustering step by the scVelo.pl.velocity_embedding_stream function.
Sample or group preference analysis
The ratio of observed to expected (Ro/e) were used to infer sample or group preference on cells in each cluster. For cells from each sample in each cluster, we constructed a contingency table to calculate the observed and expected number of cells. P value was calculated via the fisher.test function from R package stats (version 4.0.4) to indicate the significance of Ro/e.
Gene module score
The function “AddModuleScore” from R package Seurat were applied to calculate the gene module score of given gene sets to illustrate the functionality of cell clusters. In this study, four gene signature were used and converted to corresponding mouse genes during analysis: type 1 interferon (CXCL10, IFI16, IFI27, IFI30, IFI6, IFIH1, IFIT1, IFIT2, IFIT3, IFITM1, IFITM2, IFITM3, IFNA1, IFNA2, IFNA4, IFNAR1, IFNAR2, IFNB1, IFNE, IFNW1, IRF1, IRF2, IRF3, IRF5, IRF7, IRF9, ISG15, ISG20, JAK1, JAK2, OAS1, OAS2, SOCS1, STAT1, STAT2, STAT3, TMEM173, TYK2), antigen presentation (CD70, CD74, CD80, CD83, CD86, TNFSF9, HLA-A, HLA-B, HLA-C, HLA-DPA1, HLA-DPB1, HLA-DQA1, HLA-DQA2, HLA-DQB1, HLA-DQB2, HLA-DRA, HLA-DRB1, HLA-DRB5, B2M), M1 signature (IL23, TNF, CXCL9, CXCL10, CXCL11, CD86, IL1A, IL1B, IL6, CCL5, IRF5, IRF1, CD40, IDO1, KYNU, CCR7) and M2 signature (IL4R, CCL4, CCL13, CCL20, CCL17, CCL18, CCL22, CCL24, LYVE1, VEGFA, VEGFB, VEGFC, VEGFD, EGF, CTSA, CTSB, CTSC, CTSD, TGFB1, TGFB2, TGFB3, MMP14, MMP19, MMP9, CLEC7A, WNT7B, FASL, TNFSF12, TNFSF8, CD276, VTCN1, MSR1, FN1, IRF4)98.
RNA seq analysis and enrichment analysis
Raw fastq reads were quality-controlled by fastp. The clean reads were mapped to GRCh38 or mm10 reference genome to generate the expression matrix via STAR (version 2.7.2a) with default parameters99. Based on the expression matrix, differentially expressed genes were identified by R package DESeq2 (version 1.30.1)100. Gene set enrichment analysis (GSEA) on Gene ontology (GO) was performed by R package fgsea (version 1.16.0) according to the rank of log2FoldChange of each gene101.
Survival analysis
To estimate the effects of microbiomes or genes on prognosis, Kaplan–Meier survival analysis on single factor was performed via R package survival (version 3.2.7). During analysis, samples were grouped into high or low group based on the abundance of candidate microbiome or the expression level of candidate gene. The P value was calculated by the log-rank method in survival analysis.
Clinical data collection
All studies are performed in accordance with the Declaration of Helsinki principles under a protocol approved by the Ethics Committee of Guangdong Provincial People’s Hospital and written informed consent was obtained from all subjects.
We collect AEG, GC, and CRC paraffin-embedded tissue samples and clinical data from our hospital. GC patients derive from our previous study102. AEG patients are included with complete clinical data, paraffin-embedded tissue, and follow up information from 2011 to 2019 years. CRC is randomly selected 64 patients from 2012 to 2019 years. Combination AEG with GC is conducted survival analysis by SP high or low groups, excluding CRC patient because CRC patient prognosis is too different than AEG and GC.
Statistics and reproducibility
All experiments were independently repeated at least three times with similar results. Representative images shown in micrographs (FISH, TEM, IF) reflect reproducible findings across independent biological replicates.
Ethics statement
All patients consented to an institutional review broad-approved protocol that allows comprehensive analysis of tissues samples (Ethics committee of Guangdong Provincial People’s Hospital).
All animal-related procedures received prior approval from the Ethics Committee of Guangdong Provincial People’s Hospital. Animals were housed and monitored under institutional ethical oversight, and all handling followed national and institutional guidelines for humane care, welfare protection, and regulatory compliance. No animal manipulation was initiated before formal protocol authorization. According to the Ethics Committee, the size of subcutaneous tumors should not exceed 2000 mm3. The experiment should be terminated in time when the tumors develop ulceration, necrosis, or severe infection.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequencing data of public data sets are available in the GEO database under the accession number: GSE126304 and PRJNA788008 for adenocarcinoma of esophagogastric junction, GSE156451 for colorectal cancer, GSE122401, GSE179252 and GSE184336 for gastric cancer. The raw sequencing data generated in this study are available in the NCBI SRA database under the accession number: PRJNA1177527. Source data are provided with this paper.
References
Li, Q. et al. Carnobacterium maltaromaticum boosts intestinal vitamin D production to suppress colorectal cancer in female mice. Cancer Cell 41, 1450–1465.e1458 (2023).
Fu, K. et al. Streptococcus anginosus promotes gastric inflammation, atrophy, and tumorigenesis in mice. Cell 187, 882–896.e817 (2024).
Chen, H. et al. Urea cycle activation triggered by host-microbiota maladaptation driving colorectal tumorigenesis. Cell Metab. 35, 651–666.e657 (2023).
Artola-Borán, M. et al. Mycobacterial infection aggravates Helicobacter pylori-induced gastric preneoplastic pathology by redirection of de novo induced Treg cells. Cell Rep. 38, 110359 (2022).
Younginger, B. S. et al. Enrichment of oral-derived bacteria in inflamed colorectal tumors and distinct associations of Fusobacterium in the mesenchymal subtype. Cell Rep. Med. 4, 100920 (2023).
Li, Q. et al. Streptococcus thermophilus Inhibits Colorectal Tumorigenesis Through Secreting β-Galactosidase. Gastroenterology 160, 1179–1193.e1114 (2021).
He, Z. et al. Campylobacter jejuni promotes colorectal tumorigenesis through the action of cytolethal distending toxin. Gut 68, 289–300 (2019).
Si, W. et al. Lactobacillus rhamnosus GG induces cGAS/STING- dependent type I interferon and improves response to immune checkpoint blockade. Gut 71, 521–533 (2022).
Noto, J. M. et al. Iron deficiency linked to altered bile acid metabolism promotes Helicobacter pylori-induced inflammation-driven gastric carcinogenesis. J. Clin. Investig. https://doi.org/10.1172/jci147822 (2022).
Roelands, J. et al. An integrated tumor, immune and microbiome atlas of colon cancer. Nat. Med. 29, 1273–1286 (2023).
Liu, Y. et al. Peptostreptococcus anaerobius mediates anti-PD1 therapy resistance and exacerbates colorectal cancer via myeloid-derived suppressor cells in mice. Nat. Microbiol. 9, 1467–1482 (2024).
Zepeda-Rivera, M. et al. A distinct Fusobacterium nucleatum clade dominates the colorectal cancer niche. Nature 628, 424–432 (2024).
Nejman, D. et al. The human tumor microbiome is composed of tumor type-specific intracellular bacteria. Science 368, 973–980 (2020).
Fu, A. et al. Tumor-resident intracellular microbiota promotes metastatic colonization in breast cancer. Cell 185, 1356–1372.e1326 (2022).
Brooke, J. S. Stenotrophomonas maltophilia: an emerging global opportunistic pathogen. Clin. Microbiol. Rev. 25, 2–41 (2012).
He, Z. et al. Enterogenic Stenotrophomonas maltophilia migrates to the mammary gland to induce mastitis by activating the calcium-ROS-AMPK-mTOR-autophagy pathway. J. Anim. Sci. Biotechnol. 14, 157 (2023).
Liu, B. et al. Hepatic stellate cell activation and senescence induced by intrahepatic microbiota disturbances drive progression of liver cirrhosis toward hepatocellular carcinoma. J. Immunother. Cancer https://doi.org/10.1136/jitc-2021-003069 (2022).
Shen, J. et al. Stenotrophomonas maltophilia promotes lung adenocarcinoma progression by upregulating histone deacetylase 5. Front. Microbiol. 14, 1121863 (2023).
Tharp, K. M. et al. Tumor-associated macrophages restrict CD8(+) T cell function through collagen deposition and metabolic reprogramming of the breast cancer microenvironment. Nat. Cancer https://doi.org/10.1038/s43018-024-00775-4 (2024).
Jang, H. J. et al. Tobacco-induced hyperglycemia promotes lung cancer progression via cancer cell-macrophage interaction through paracrine IGF2/IR/NPM1-driven PD-L1 expression. Nat. Commun. 15, 4909 (2024).
Vanmeerbeek, I. et al. Targeting conserved TIM3(+)VISTA(+) tumor-associated macrophages overcomes resistance to cancer immunotherapy. Sci. Adv. 10, eadm8660 (2024).
Martinez-Terroba, E. et al. Overexpression of Malat1 drives metastasis through inflammatory reprogramming of the tumor microenvironment. Sci. Immunol. 9, eadh5462 (2024).
Sun, L. et al. Activating a collaborative innate-adaptive immune response to control metastasis. Cancer Cell 39, 1361–1374.e1369 (2021).
Sharma, G. et al. A dietary commensal microbe enhances antitumor immunity by activating tumor macrophages to sequester iron. Nat. Immunol. 25, 790–801 (2024).
Nasir, I. et al. Tumor macrophage functional heterogeneity can inform the development of novel cancer therapies. Trends Immunol. 44, 971–985 (2023).
Ivashkiv, L. B. & Donlin, L. T. Regulation of type I interferon responses. Nat. Rev. Immunol. 14, 36–49 (2014).
Hopfner, K. P. & Hornung, V. Molecular mechanisms and cellular functions of cGAS-STING signalling. Nat. Rev. Mol. Cell Biol. 21, 501–521 (2020).
Hoebe, K. et al. Identification of Lps2 as a key transducer of MyD88-independent TIR signalling. Nature 424, 743–748 (2003).
Okabe, Y., Sano, T. & Nagata, S. Regulation of the innate immune response by threonine-phosphatase of Eyes absent. Nature 460, 520–524 (2009).
Ho, W. S. et al. PP2Ac/STRN4 negatively regulates STING-type I IFN signaling in tumor-associated macrophages. J. Clin. Investig. https://doi.org/10.1172/jci162139 (2023).
Manzanillo, P. S., Shiloh, M. U., Portnoy, D. A. & Cox, J. S. Mycobacterium tuberculosis activates the DNA-dependent cytosolic surveillance pathway within macrophages. Cell Host Microbe 11, 469–480 (2012).
Burdette, D. L. & Vance, R. E. STING and the innate immune response to nucleic acids in the cytosol. Nat. Immunol. 14, 19–26 (2013).
Patel, D. J., Yu, Y. & Xie, W. cGAMP-activated cGAS-STING signaling: its bacterial origins and evolutionary adaptation by metazoans. Nat. Struct. Mol. Biol. 30, 245–260 (2023).
Wang, L. et al. STING agonist-loaded nanoparticles promotes positive regulation of type I interferon-dependent radioimmunotherapy in rectal cancer. Adv. Sci. 11, e2307858 (2024).
Ohkuri, T. et al. STING contributes to antiglioma immunity via triggering type I IFN signals in the tumor microenvironment. Cancer Immunol. Res. 2, 1199–1208 (2014).
Marzio, A. et al. EMSY inhibits homologous recombination repair and the interferon response, promoting lung cancer immune evasion. Cell 185, 169–183.e119 (2022).
Kosaka, A. et al. CD47 blockade enhances the efficacy of intratumoral STING-targeting therapy by activating phagocytes. J. Exp. Med. https://doi.org/10.1084/jem.20200792 (2021).
Ghosh, M., Saha, S., Li, J., Montrose, D. C. & Martinez, L. A. p53 engages the cGAS/STING cytosolic DNA sensing pathway for tumor suppression. Mol. Cell 83, 266–280.e266 (2023).
Li, S. et al. STING-induced regulatory B cells compromise NK function in cancer immunity. Nature 610, 373–380 (2022).
Long, Y. et al. GPR162 activates STING dependent DNA damage pathway as a novel tumor suppressor and radiation sensitizer. Signal Transduct. Target. Ther. 8, 48 (2023).
Lam, K. C. et al. Microbiota triggers STING-type I IFN-dependent monocyte reprogramming of the tumor microenvironment. Cell 184, 5338–5356.e5321 (2021).
Yang, L., Li, A., Wang, Y. & Zhang, Y. Intratumoral microbiota: roles in cancer initiation, development and therapeutic efficacy. Signal Transduct. Target. Ther. 8, 35 (2023).
Waanders, L. et al. Ectopic expression of cGAS in Salmonella typhimurium enhances STING-mediated IFN-β response in human macrophages and dendritic cells. J. Immunother. Cancer https://doi.org/10.1136/jitc-2022-005839 (2023).
Yang, K. et al. Suppression of local type I interferon by gut microbiota-derived butyrate impairs antitumor effects of ionizing radiation. J. Exp. Med. https://doi.org/10.1084/jem.20201915 (2021).
Liu, J. L. et al. Dysfunctional circadian clock accelerates cancer metastasis by intestinal microbiota triggering accumulation of myeloid-derived suppressor cells. Cell Metab. 36, 1320–1334.e1329 (2024).
Liu, Z. et al. Colorectal cancer microbiome programs DNA methylation of host cells by affecting methyl donor metabolism. Genome Med. 16, 77 (2024).
Nakatsu, G., Andreeva, N., MacDonald, M. H. & Garrett, W. S. Interactions between diet and gut microbiota in cancer. Nat. Microbiol. 9, 1644–1654 (2024).
Blake, S. J., Wolf, Y., Boursi, B. & Lynn, D. J. Role of the microbiota in response to and recovery from cancer therapy. Nat. Rev. Immunol. 24, 308–325 (2024).
Cao, C., Yue, S., Lu, A. & Liang, C. Host-gut microbiota metabolic interactions and their role in precision diagnosis and treatment of gastrointestinal cancers. Pharmacol. Res. 207, 107321 (2024).
Li, Y. et al. Intracellular Fusobacterium nucleatum infection attenuates antitumor immunity in esophageal squamous cell carcinoma. Nat. Commun. 14, 5788 (2023).
Schorr, L., Mathies, M., Elinav, E. & Puschhof, J. Intracellular bacteria in cancer-prospects and debates. npj Biofilms Microbiomes 9, 76 (2023).
Robinson, W. et al. Identification of intracellular bacteria from multiple single-cell RNA-seq platforms using CSI-Microbes. Sci. Adv. 10, eadj7402 (2024).
Greathouse, K. L. et al. Interaction between the microbiome and TP53 in human lung cancer. Genome Biol. 19, 123 (2018).
Davis, N. M., Proctor, D. M., Holmes, S. P., Relman, D. A. & Callahan, B. J. Simple statistical identification and removal of contaminant sequences in marker-gene and metagenomics data. Microbiome 6, 226 (2018).
de Goffau, M. C. et al. Human placenta has no microbiome but can contain potential pathogens. Nature 572, 329–334 (2019).
Pham, T. H. M. et al. Salmonella-driven polarization of granuloma macrophages antagonizes TNF-mediated pathogen restriction during persistent infection. Cell Host Microbe 27, 54–67.e55 (2020).
Corrales, L., McWhirter, S. M., Dubensky, T. W. Jr. & Gajewski, T. F. The host STING pathway at the interface of cancer and immunity. J. Clin. Investig. 126, 2404–2411 (2016).
Borden, E. C. Interferons α and β in cancer: therapeutic opportunities from new insights. Nat. Rev. Drug Discov. 18, 219–234 (2019).
McNab, F., Mayer-Barber, K., Sher, A., Wack, A. & O’Garra, A. Type I interferons in infectious disease. Nat. Rev. Immunol. 15, 87–103 (2015).
Chin, E. N., Sulpizio, A. & Lairson, L. L. Targeting STING to promote antitumor immunity. Trends Cell Biol. 33, 189–203 (2023).
He, Q., Hao, T., She, D. & Lv, P. IFNα2b in combination with oncolytic adenovirus enhances antitumor activity against melanoma. Int. Immunopharmacol. 165, 115435 (2025).
Harper, M. S. et al. IFN-α treatment inhibits acute Friend retrovirus replication primarily through the antiviral effector molecule Apobec3. J. Immunol. 190, 1583–1590 (2013).
Hu, B. et al. IFNα potentiates anti-PD-1 efficacy by remodeling glucose metabolism in the hepatocellular carcinoma microenvironment. Cancer Discov. 12, 1718–1741 (2022).
Lazear, H. M., Schoggins, J. W. & Diamond, M. S. Shared and distinct functions of type I and type III interferons. Immunity 50, 907–923 (2019).
Ding, W. X. & Yin, X. M. Sorting, recognition and activation of the misfolded protein degradation pathways through macroautophagy and the proteasome. Autophagy 4, 141–150 (2008).
Christianson, J. C., Jarosch, E. & Sommer, T. Mechanisms of substrate processing during ER-associated protein degradation. Nat. Rev. Mol. Cell Biol. 24, 777–796 (2023).
Gonugunta, V. K. et al. Trafficking-mediated STING degradation requires sorting to acidified endolysosomes and can be targeted to enhance anti-tumor response. Cell Rep. 21, 3234–3242 (2017).
Gentili, M. et al. ESCRT-dependent STING degradation inhibits steady-state and cGAMP-induced signalling. Nat. Commun. 14, 611 (2023).
Mutlu, M. et al. Small molecule induced STING degradation facilitated by the HECT ligase HERC4. Nat. Commun. 15, 4584 (2024).
Ji, Y. et al. SEL1L–HRD1 endoplasmic reticulum-associated degradation controls STING-mediated innate immunity by limiting the size of the activable STING pool. Nat. Cell Biol. 25, 726–739 (2023).
Zhang, C. X. et al. Galectin-9 promotes a suppressive microenvironment in human cancer by enhancing STING degradation. Oncogenesis 9, 65 (2020).
Zhao, Y. et al. A specific module of ESCRT regulates STING activity termination by controlling STING degradation. Sci. Bull. 69, 1000–1005 (2024).
Tamma, P. D. et al. Infectious Diseases Society of America guidance on the treatment of AmpC β-lactamase-producing enterobacterales, carbapenem-resistant Acinetobacter baumannii, and Stenotrophomonas maltophilia Infections. Clin. Infect. Dis. 74, 2089–2114 (2022).
Ko, J. H. et al. Fluoroquinolones versus trimethoprim-sulfamethoxazole for the treatment of Stenotrophomonas maltophilia infections: a systematic review and meta-analysis. Clin. Microbiol. Infect. 25, 546–554 (2019).
Chng, K. R. et al. Tissue microbiome profiling identifies an enrichment of specific enteric bacteria in opisthorchis viverrini associated cholangiocarcinoma. EBioMedicine 8, 195–202 (2016).
Pushalkar, S. et al. The pancreatic cancer microbiome promotes oncogenesis by induction of innate and adaptive immune suppression. Cancer Discov. 8, 403–416 (2018).
Ma, Y. et al. Intratumor microbiome-derived butyrate promotes lung cancer metastasis. Cell Rep. Med. 5, 101488 (2024).
Basu, S. et al. Execution of macrophage apoptosis by PE_PGRS33 of Mycobacterium tuberculosis is mediated by Toll-like receptor 2-dependent release of tumor necrosis factor-alpha. J. Biol. Chem. 282, 1039–1050 (2007).
Mirji, G. et al. The microbiome-derived metabolite TMAO drives immune activation and boosts responses to immune checkpoint blockade in pancreatic cancer. Sci. Immunol. 7, eabn0704 (2022).
Godfrey, D. I., Stankovic, S. & Baxter, A. G. Raising the NKT cell family. Nat. Immunol. 11, 197–206 (2010).
Jiang, S. S. et al. Fusobacterium nucleatum-derived succinic acid induces tumor resistance to immunotherapy in colorectal cancer. Cell Host Microbe 31, 781–797.e789 (2023).
Duong, M. T., Qin, Y., You, S. H. & Min, J. J. Bacteria-cancer interactions: bacteria-based cancer therapy. Exp. Mol. Med. 51, 1–15 (2019).
Casasanta, M. A. et al. Fusobacterium nucleatum host-cell binding and invasion induces IL-8 and CXCL1 secretion that drives colorectal cancer cell migration. Sci. Signal. https://doi.org/10.1126/scisignal.aba9157 (2020).
Xu, J. Y., Liu, M. T., Tao, T., Zhu, X. & Fei, F. Q. The role of gut microbiota in tumorigenesis and treatment. Biomed. Pharmacother. Biomed. Pharmacother. 138, 111444 (2021).
Ge, Y. et al. Gut microbiota influence tumor development and Alter interactions with the human immune system. J. Exp. Clin. Cancer Res. CR 40, 42 (2021).
Kim, J. & Lee, H. K. The role of gut microbiota in modulating tumor growth and anticancer agent efficacy. Mol. cells 44, 356–362 (2021).
Sun, Q. et al. Synergistic between autotrophic and heterotrophic microorganisms for denitrification using bio-S as electron donor. Environ. Res. 231, 116047 (2023).
Lu, J. et al. Metagenome analysis using the Kraken software suite. Nat. Protoc. 17, 2815–2839 (2022).
Chen, S. Ultrafast one-pass FASTQ data preprocessing, quality control, and deduplication using fastp. iMeta 2, e107 (2023).
Ghaddar, B. et al. Tumor microbiome links cellular programs and immunity in pancreatic cancer. Cancer Cell 40, 1240–1253.e1245 (2022).
Ghaddar, B., Blaser, M. J. & De, S. Denoising sparse microbial signals from single-cell sequencing of mammalian host tissues. Nat. Comput. Sci. 3, 741–747 (2023).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. Doubletfinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337.e324 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e3529 (2021).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Azizi, E. et al. Single-cell map of diverse immune phenotypes in the breast tumor microenvironment. Cell 174, 1293–1308.e1236 (2018).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Korotkevich, G. et al. Fast gene set enrichment analysis. bioRxiv https://doi.org/10.1101/060012 (2021).
Zheng, J. et al. KDM3A ablation activates endogenous retrovirus expression to stimulate antitumor immunity in gastric cancer. Adv. Sci. https://doi.org/10.1002/advs.202309983 (2024).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (82173829 to F. Xing; 32370836 and 82573900 to Y. Li), Excellent Young Talent Program of Guangdong Provincial People’s Hospital (grant no. KY012023333 to F. Xing). We thank Prof. Xiaoyu Hu (Jilin University) for providing SP-GFP, Prof. Juli Liu, Prof. Wei Yang and Prof. Xin Li (Southern medical University) for research ideas guidance and uplift.
Author information
Authors and Affiliations
Contributions
Y. Li, HL. Feng, KL. Chen and ZH. Xi designed this study; HL. Feng, KL. Chen, JQ. Guo, F. Ling, BJ. Xia, JL. Zhou, YH. Zhang, and YC. Zhang performed the experiments; JL. Zhou and ZH. Xi conducted single cell and bulk RNA-seq analysis; DD. Zhu, YF. Liu, WX. Zhang, JT. Lin, QS. Huang, ZJ. Lyu, JJ. Wang and ZD. Huang collected tissue samples and the clinical data; HL. Feng, KL. Chen and ZH. Xi analyzed and interpreted the data; HL. Feng drafted manuscript; Y. Li, F. Xing, TY. Hou, and JB. Zheng edited the manuscript; Y. Li, F. Xing, TY. Hou, and JB. Zheng supervised the study; X. Chen, YQ. Huang and ZD Huang assisted in revising the manuscript. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Feng, H., Chen, K., Xi, Z. et al. Stenotrophomonas promotes gastrointestinal tumor progression via STING degradation in tumour cells and mitigated immune response. Nat Commun 17, 2042 (2026). https://doi.org/10.1038/s41467-026-68649-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68649-0