Abstract
Age is a major risk factor for lung disease. We characterized the changing cellular, transcriptional, and genomic landscape of human lung aging using single-cell RNA sequencing. We find that lung aging is cell-type dyssynchronous, with alveolar epithelial and endothelial cells exhibiting the greatest transcriptional changes. Among alveolar epithelial cells, aging is associated with a decreased relative proportion of surfactant-expressing SPChigh AT2 cells. Among alveolar capillary cells, we observed loss of differentiation and capillary function. Analysis of somatic mutations called from single-cell data revealed an increase with aging, with alveolar epithelial and endothelial cell types exhibiting greater mutation burdens. Transcriptional entropy was increased with aging and was an independent predictor of age. Notably, cells expressing commonly accepted senescence signatures did not increase with age. Our results reveal cell type dyssynchrony in human lung aging with age-related changes concentrated in alveolar epithelial and endothelial cells.
Similar content being viewed by others
Introduction
Age is a substantial risk factor in nearly all lung diseases1,2. Acute diseases such as pneumonia and ARDS, and chronic diseases such as bronchiectasis, chronic obstructive lung disease (COPD), and idiopathic pulmonary fibrosis (IPF) are both more common and more lethal in aged individuals3. While there has been significant progress in understanding the role of aging-related mechanisms in advanced lung disease4,5,6 and in describing the physiological effects of aging in the lung7,8,9, the cellular and molecular mechanisms that underlie the lung’s aging response remain poorly understood.
There is limited knowledge about human lung cellular aging10. Pulmonary stem cell exhaustion and epithelial cell senescence are associated with advanced age and are implicated in the pathogenesis of age-related diseases such as IPF4. Mucociliary clearance and ciliary beat frequency decrease with age and increase predisposition to pneumonia in the elderly11. Changes in ECM composition with age, including increased collagen and decreased elastin and laminin, have also been repoprted12,13.
Recent advances in sequencing technologies and the availability of larger datasets such as the Genotype-Tissue Expression (GTEx) Project14 have enabled the identification of novel markers of aging and senescence in the lung and other organ systems15,16. Chow et al. identified changes in bulk and single-cell RNAseq expression and relative cell-type proportions with age in the context of SARS-CoV-2 susceptibility17. Angelidis et al. developed a mouse atlas of lung aging using single-cell transcriptomics and mass spectrometry-based proteomics and showed altered cellular proportions of airway epithelial cells and changes in ECM composition18. Jia et al. showed an age-associated increase in expression of inflammation-related genes in myeloid cells and developed a lung aging prediction score19. Most recently, Liu et al. analyzed differential gene expression in a compendium of single-cell datasets and reported loss of cell proliferation in AT2 cells and dysregulated epithelial to myeloid signaling20. To date, no comprehensive study of single-cell transcriptomic changes in human lung aging and its relations to somatic mutation accumulation and cellular senescence has been reported.
In this work, we developed a large-scale single-cell RNAseq atlas of human lung aging that revealed age-associated changes in cell-type proportions and gene expression. Deconvolution of established bulk RNAseq and weighted gene co-expression network analysis identified molecular programs that validated signatures associated with aging and cellular senescence. Moreover, we called somatic mutations from our single-cell dataset to determine the role of DNA damage accumulation in aging and senescence21. We show that somatic mutation accumulation in aging is associated with increasingly stochastic gene expression, which can be modeled using entropy and information theory.
Results
Generation of a single-cell RNAseq atlas of human lung aging
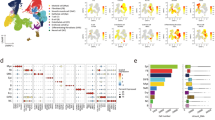
To identify signatures of aging and senescence in transcriptomic data from lungs, we developed a large-scale single-cell RNAseq atlas of human aging (Fig. 1A). We performed scRNAseq on 32 human lungs. This dataset was then integrated with control data consisting of 28 lungs from the previously published IPF Cell Atlas to increase cell count and age representation and ensure consistency of annotations22. The resulting integrated dataset consisted of 199,400 cells from 60 donors (Supplementary Data 1). These cells were annotated into 25 distinct cell-type populations (Fig. 1B). We confirmed cell-type annotations by analyzing the expression of marker genes for each cell type. The resulting dataset has broad coverage of different ages, with a relatively uniform age distribution between 11 and 80 years (Fig. 1C). Our data have been deposited on GEO and are available using accession GSE281219.
A Single-cell RNAseq data (and bulk RNAseq validation data) were used to determine cell-type proportions and cell-type-specific signatures. Signatures associated with aging and cellular senescence were identified and validated using immunohistochemistry. Somatic mutations were called from RNAseq datasets, and transcriptional changes were identified. B Integrated single-cell RNAseq dataset included 25 cell types from two datasets. C Cell-type marker genes for each annotated cell type (top). Age distribution, sex distribution, and sources of data (bottom). Source data are provided as a Source Data file. Created in BioRender. https://BioRender.com/aedhfrp.
Transcriptional alteration in AT2 cells and capillary cells
Differential gene expression (DGE) was performed using a general linear mixed effect model (GLMM) that accounts for random effects due to sample and cell-level variation (Supplementary Data 2, 3). Aged lungs were defined as those from donors aged 60 or older. To quantitatively determine which cell types experienced the greatest degree of transcriptional change with aging, we calculated the number of significantly regulated genes (p < 0.05) as a fraction of the total number of genes tested for each cell type. Genes that were expressed in less than 5% of cells of each type were excluded. The cell types that exhibited the greatest proportion of differentially expressed genes were general capillary (gCap) cells (21.5% DGEs), alveolar type II (AT2) cells (19.5%), and smooth muscle cells (SMCs) (17.3%) (Fig. 2A). In contrast, goblet cells exhibited the lowest proportion of differentially expressed genes (5.5%). Differential gene expression was also performed using a pseudobulk likelihood ratio test. Very strong concordance was observed between the two models for all tested cell types (Fisher's Exact p < 0.05) (Supplementary Data 4).
A Left panel: Beta coefficients for differentially expressed genes with p < 0.05 are plotted. Genes that remain significant after FDR correction are overlaid in blue. Right panel: proportion of tested genes that are differentially expressed, ordered by cell type. B Co-expression networks were developed in this study. The x-axis shows module size, and the y-axis shows the FDR-corrected p-value for the correlation between the module eigengene and age. Significance (color) is the −log10 transformed p-value multiplied by the sign of the correlation coefficient. C Bar plots labeled with the top REACTOME term for age-associated modules. The top panel shows positively correlated genes, and the bottom panel shows negatively correlated genes. The length of the bar represents the correlation of the module eigengene and age. The color of the bar corresponds to the -log10 transformed p-value multiplied by the sign of the correlation coefficient. The color corresponds to the color bar in (B). Source data are provided as a Source Data file.
Age-associated gene co-expression networks
To validate single-cell signatures of lung aging in bulk RNAseq data, we developed gene co-expression networks (Fig. 2B). Gene co-expression network analysis was performed using Weighted Correlation Network Analysis (WGCNA)23. Using a bulk RNAseq dataset from the Genotype-Tissue Expression (GTEx) Project, genes were grouped into 133 modules ranging from 30 to 4181 genes in size. Modules were summarized by calculating the first principal component, or module eigengene, of gene expression. Of these, 30 of 133 were significantly correlated with age (16 positive, 14 negative) (Fig. 2B). The molecular function of age-associated co-expression networks was assessed by enrichment of REACTOME pathways. Positively correlated modules were enriched for signal transduction, extracellular matrix (ECM) organization, and immunoregulatory interactions. Negatively correlated modules included the TCA cycle and respiratory electron transport, cell cycle, and metabolism of RNA (Fig. 2C).
To determine whether co-expression modules corresponded to specific cell types, we performed deconvolution of bulk RNAseq data using our group’s previously published single-cell RNAseq dataset as a reference22. Cell-type proportions for all samples were correlated with the module eigengene for each module, revealing patterns of cell-type specificity (Supplementary Fig. 1A). To assess whether this cell-type deconvolution was successful, we calculated the expression of each module in our single-cell RNAseq dataset. Agreement between both methods was present for the majority of cell types, with the exception of cells with low representation in our bulk RNAseq dataset (macrophages, B/T cells, peribronchial cells, and ciliated cells).
Strikingly, the module (6) most strongly associated with age was expressed primarily in AT2 cells, confirming the finding from single-cell data that AT2 cells are among the cell types most transcriptionally affected in aging (Supplementary Fig. 1A).
To identify specific genes that drive module function, hub genes were identified for age-associated modules. Hub genes were defined as those that had a high module membership (correlation of gene expression to ME) and high correlation with age based on expression in GTEx RNAseq data. Notable hub genes included IGFBP7 in the ECM module, QDPR in the mitochondrial module, and HHIP in the cholesterol biosynthesis module (Supplementary Fig. 1B). GLB1, which encodes SA-B-Gal, one of the gold standard markers for cellular senescence24, and NEU1, which encodes a lysosomal sialidase implicated in age-related neurodegeneration25, were hub genes for the lysosome module. Association of hub genes with aging was confirmed directly in our single-cell RNAseq dataset (Supplementary Fig. 1C).
Frequency of alveolar epithelial cells is altered in aging
We sought to determine how the cell-type composition of the human lung changes with age. Cell-type proportions for each sample were normalized to the total number of cells for each sample. We observed a significant decrease in the proportion of AT2 cells in aged individuals, consistent with previous published findings17,26 (Wilcoxon p = 0.048). Other cell-type proportions did not change significantly (Supplementary Fig. 2A).
To validate these cell-type proportion changes, we used cell-type proportion estimates from our deconvoluted bulk RNAseq dataset. This confirmed the loss of AT2 cells with aging (Wilcoxon p = 0.049) (Supplementary Fig. 2B). Hence, AT2 cells are transcriptionally altered and decline in proportion in the aged human lung.
SPChigh AT2 cells and surfactant are reduced in aged lungs
One of the strongest transcriptional signatures we observed was a prominent age-associated decrease in the expression of surfactant genes (SFTPA1, SFTPA2, SFTPB, SFTPC) in both bulk and single-cell data (Fig. 3A). When examining single-cell data for AT2 cells independently, we observed two subpopulations that formed independent clusters (Fig. 3B, C), similar to what has been reported previously by our group and others27,28,29. Travaglini et al. characterized two subtypes: bulk AT2 (AT2B) cells that have high expression of surfactants and correspond with a functional phenotype, and signaling AT2 (AT2S) cells that have lower levels of surfactant and correspond with a more stem-like progenitor phenotype29. In our dataset, we confirmed the existence of analogous subtypes, which we termed SPClow (reduced expression of surfactants) and SPChigh AT2 cells (Fig. 3D, Supplementary Fig. 3A). Moreover, SPClow AT2 cells exhibited greater expression of Wnt/β-catenin signaling genes (CTNNB1, GSK3B, TCF12, AXIN1, APC), EGF receptor signaling (EGFR, ERBB4, JAK1, STAT3), Hippo signaling (YAP1, STK3, DLG1/DLG2), and cell proliferation markers (HMMR, TOP2A) (Supplementary Fig. 3B).
A Log-normalized expression of surfactant genes in young versus aged. B Cell-type proportions as a fraction of total epithelial cell counts in young versus aged. Samples with fewer than 5 AT1 cells are excluded. C UMAP representation of AT2 cells, revealing SPCˡᵒʷand SPCʰⁱᵍʰ subpopulations. D Expression levels of AT2 marker SFTPC and SPCʰⁱᵍʰ marker HHIP. E Proportion of total AT2 cells that are SPCˡᵒʷ in young versus aged. Samples with less than 5 AT1 cells were excluded (n = 29, 20Y/9A). F Proportion of cells expressing HHIP in SPCˡᵒʷ vs SPCʰⁱᵍʰ subpopulations based on scRNAseq data (n = 59, 30 high/29 low). G Representative immunohistochemistry images for dual SPC/HHIP staining. The left panel is a young subject, and the right panel is an aged subject. SPC was stained with DAB (brown), and HHIP was stained with AP Red (pink). Red arrowheads indicate SPC-positive cells and blue arrowheads indicate dual-positive, co-stained cells. H The proportion of dual-positive cells among SPC-positive cells in young versus aged subjects (n = 12, 6Y/6A). Source data are provided as a Source Data file.
Aged samples had a significantly higher proportion of SPClow AT2 cells (mean 0.47 versus 0.14, Wilcoxon p = 0.0062) (Fig. 3E). HHIP was a robust marker gene for the SPChigh AT2 cell population (FDR p < 0.001) (Fig. 3F). Hence, the proportion of HHIP-expressing, surfactant-producing SPChigh AT2 cells decreased with age.
This finding was supported by the results of bulk RNAseq co-expression network analysis. The co-expression module that was most strongly negatively correlated with age (r = −0.31, FDR p = 2e-12) was enriched for cholesterol biosynthesis and was expressed in AT2 cells (Supplementary Fig. 1). This module contained HHIP as a hub gene, a gene that has been implicated in the pathogenesis of COPD and is expressed at lower levels in COPD subjects based on human single-cell RNAseq27. These findings suggest that HHIP is a central gene in a molecular program that leads to reduced surfactant production in AT2 cells.
We performed immunohistochemistry in aged and young FFPE sections. HHIP was co-stained with SPC. We observed co-localization of HHIP and SPC, confirming that HHIP is primarily expressed in AT2 cells (Fig. 3G). Next, the proportion of SPC-positive cells that also stained positive for HHIP was quantified. We observed a significant difference between young and aged subjects (mean 0.55 versus 0.31, Wilcoxon p = 0.032) (Fig. 3H).
Taken together, these results indicate that AT2 cells not only decrease in number with age and exhibit reduced expression of gene networks central to surfactant production, but also that the subpopulation functionally involved in secreting surfactant is not renewed, as reflected by relative accumulation of SPClow AT2. These findings potentially explain the lung’s increased predisposition to injury with age.
Aging lung capillary dysfunction and increased autophagy
Lung endothelial cells were among the cell types that exhibited significant transcriptional alterations in aging. To better characterize these changes, we analyzed lung endothelial cells collectively. The major lung endothelial cell populations30 (arterial, venous, aerocyte, gCap, and lymphatic) clustered together to form independent populations in our dataset (Fig. 4A).
A UMAP representation of endothelial cell subpopulations. B Expression levels for differentially expressed genes in aerocytes and gCap cells. Expression values are averaged per sample and scaled from 0 to 1. C Upset plot showing the number of differentially expressed genes that are shared across a combination of cell types, before FDR correction. D Top KEGG/Reactome enrichment terms for differentially expressed genes in capillary cells. Terms are ordered and colored by enrichment p-value, with increased signatures in yellow and decreased signatures in purple. E Violin plots showing expression levels of selected capillary endothelial cell marker genes in young versus aged samples. Asterisks indicate significance levels from the Wilcoxon Rank-Sum test. Source data are provided as a Source Data file.
When analyzing differential gene expression, the two capillary cell populations - aerocytes and gCap cells - exhibited overlapping signatures. Among gCap cells and aerocytes, 380 genes (180 exclusively) were differentially expressed in the same direction (221 increased, 159 decreased) in both cell types (Fig. 4B, C, Supplementary Data). Differential expression analysis among gCap cells and aerocytes revealed an increase in expression of genes related to ubiquitin, proteolysis, and Ras/MAPK signaling pathways following FDR correction. Autophagy-related genes included ubiquitin ligases (ABLIM1, MIB1, RNF19A), ubiquitin regulatory proteins (UBXN7, UBE2H, UBE2K), F-box proteins – which form ubiquitin ligase complexes (FBXL17, FBXO42), and general autophagy genes (NPEPPS, VMP1, ULK1, RICTOR). Genes implicated in the Ras/MAPK pathway included SOS1, SOS2, MAP3K20, MAP4K4, and MAPK8. In contrast, mitochondrial genes were consistently decreased in capillary cells (MT-CO1, MT-CO2, MT-CO3, MT-ND1, MT-ND5, MT-CYB). Enrichment analysis on the differentially expressed genes confirmed an increase in expression of genes related to terms such as “Ubiquitin mediated proteolysis” and “MAPK signaling pathway” and a decrease in terms such as “Oxidative phosphorylation” and “Aerobic respiration” (Fig. 4D). Markers of normal capillary endothelial cell function were also decreased (VIPR1, IL7R, FCN3, HPGD) (Fig. 4E). Hence, lung capillary cell aging is characterized by increased expression of autophagy genes, mitochondrial dysfunction, and loss of endothelial subpopulation differentiation markers.
In the bulk RNAseq data, the co-expression network containing CDKN2A was primarily expressed in endothelial cells and was increased with age (r = 0.26, p < 0.05). Interestingly, this module also contained many ubiquitin ligase complex genes, including the F-box gene FBXW7 as a hub gene. Hence, bulk co-expression analysis validated the signatures of lung endothelial aging we found using single-cell data.
Transcriptional changes in mesenchymal and immune cell types
Among mesenchymal cells, SMCs exhibited the greatest number of differentially expressed genes, with an increase in genes related to extracellular matrix remodeling (ITGA4, FGF1, COL4A6, LTBP1, LTBP2) (Supplementary Fig. 4A). Other genes included genes involved in regulation of transcription (CTBP2, SP1, GLI2, MECP2, MAML3) and zinc finger family genes (ZNF44, ZNF83, ZNF138, ZNF148). In myofibroblasts, genes involved in inflammation and innate immunity (UBASH3B, DNASE2, FAM135A) were increased (Supplementary Fig. 4B).
Immune cells exhibited fewer differentially expressed genes. Macrophages exhibited increased expression of ubiquitination genes (CUL1, PIAS1, HUWE1, CDC27) and a robust decrease in mitochondrial function genes (TUFM, MPC1, NDUF genes, mitochondrial ribosomal subunits), not dissimilar to endothelial cells. However, the significance of these genes did not reach the FDR threshold for significance. In alveolar macrophages, genes that were increased are involved in transcriptional regulation and genome stability (BRCA2, SP140, HMGN5) and immune system regulation (NLRC5, STAT1, IFI16). Genes that decreased included LRRC25 and NFKBIA, negative regulators of NF-KB signaling. Notably, canonical SASP markers such as IL6, IL8, CXCL1, CCL2, and TNF were not differentially expressed in any immune cell type.
Stratification of aging signatures by age and smoking
To determine whether aging signatures exhibit distinct temporal patterns, we performed differential expression at multiple time points across the lifespan. Specifically, identified gene signatures that emerge in early aging (>40 years) versus later aging (>60 years). Several genes were significantly increased in later adulthood that were not increased in early adulthood (Supplementary Fig. 5A, Supplementary Data 5). In AT2 cells, these included genes involved in genomic stability (DNMT3A, BRCA2, TP53) and inflammation (IRF3). In gCap cells, ubiquitin ligases (FBXL17, FBXO42, ABLIM1, MIB1, ANAPC1) and Ras/MAPK pathway genes all increased specifically in later aging. Very few genes were specifically increased in early aging (fewer than 10 for each cell type after FDR correction). Notably, aging signatures were not significantly affected by smoking (Supplementary Data 5 and Supplementary Fig. 5).
Somatic mutation burden increases with age
Accumulation of somatic mutations has been implicated in both cellular aging and senescence31,32. We used SComatic33 to call somatic mutations in our single-cell data. Somatic mutation burden was calculated for each sample-cell-type combination. Cell types without sufficient sequencing coverage to call mutations in at least five samples were excluded from analysis. Somatic mutation burden was highest in AT1 cells (7.2 mutations/MB), aerocytes (6.1), and general capillary (gCap) cells (6.0) (Supplementary Fig. 6A). The majority of mutations were exonic, with a similar distribution in young and aged samples (Supplementary Fig. 6B). Among exonic variants, mutation types were similar in young (59% synonymous, 36% nonsynonymous, 5% other) and aged (60% synonymous, 38% nonsynonymous, 2% other). Enrichment analysis for genes with nonsynonymous mutations revealed enrichment of terms including “Antigen processing and presentation”, “Ribosome”, “Phagosome”, and “Protein processing in endoplasmic reticulum.” (Supplementary Fig. 6C). To model the association between mutations and age, we used a GLMM that accounts for cell type. Notably, mutation burden was strongly positively correlated with age in our dataset (p < 0.001) (Fig. 5A).
A Plot of age versus mutation burden for each sample-cell-type combination in scRNAseq data. Samples with fewer than 10 mutations are excluded. GAM curves are fitted for each cell type. B Gene expression for genes that correlate with the sample somatic mutation burden. The top positively (top) and negatively correlated (bottom) genes are included. Expression values are averaged across cells for each sample. C Differentially expressed genes in AT2, and gCap cells. Expression values are averaged for cells with low mutation versus high mutation burden. D Entropy for select cell types, scaled to range from 0 to 1, in young versus aged cells. E Entropy as a function of whole-sample mutation burden (n = 30). Entropy values of cells were averaged for each sample. Entropy values were then scaled to range from 0 to 1. Source data are provided as a Source Data file.
To identify genes that correlated with somatic mutation burden, we applied a GLMM model that accounted for cell-type composition in our dataset. Genes that were positively correlated with mutation burden included ubiquitin ligase genes (SEL1L, ANAPC1, UBE4A, USP25, USP33) and DNA damage response genes (RAD50, MTREX, IFI16, SAMHD1, CBL, APC) (Fig. 5B, Supplementary Data 6). Genes associated with mitochondria were negatively correlated with somatic mutation burden (ETFB, COQ10A, GFER, MDH2, SCO2, IDH3B, COX4I1). Hence, somatic mutations are associated with damage response pathways, increased ubiquitination gene expression, and mitochondrial dysfunction.
We next sought to determine the association between somatic mutation burden and gene expression at cell-type resolution. We chose to focus on AT2 cells and gCap cells, which had high sequencing coverage and strong age-associated transcriptional changes. These cell types exhibited distinct signatures (Fig. 5C, Supplementary Data 7). gCap cells exhibited decreased expression of mitochondrial genes (MT-ND2, MT-ND1, NDUFA3, ATP9B, ATP5MG) and increased ubiquitination (USP24, CUL1, CUL3, CBL, SHPRH), mirroring the changes seen in aging. In AT2 cells, mutation burden correlated with increased expression of genes involved in chromatin organization (INO80, PSIP1, METTL3, NOC3L, unadjusted p < 0.05). Both cell types exhibited features of increased DNA damage repair. These results suggest that different cell types in the lung exhibit distinct changes with somatic mutation accumulation.
To validate the age-associated somatic mutation accumulation in the lungs, we analyzed somatic mutation data from a previously published GTEx study34. As in single-cell data, somatic mutation burden was positively correlated with age and associated with similar genes (Supplementary Fig. 6).
Transcriptional entropy and noise in aged human lung cells
The loss of marker gene expression with increasing age and somatic mutation burden accumulation suggests a possible loss of transcriptional regulation35,36. We reasoned that transcriptomes from cells with lower expression of phenotypic genes would exhibit greater uncertainty, or entropy, when modeled as a probability distribution37. To quantify this, we calculated transcriptional entropy for specific lung cell types. As an additional measure of transcriptional dysregulation, we calculated transcriptional noise for each cell as was previously performed18. Both entropy and transcriptional noise were increased with aging in most cell types (Fig. 5D, Supplementary Fig. 7A, 7B) except immune cells, including lymphocytes and myeloid cell types. Notably, cell types with higher mutation burden and gene expression changes also had greater differences in transcriptional entropy (capillary, AT1, AT2). Globally, entropy was strongly positively correlated with mutation burden (r = 0.51, p = 0.004) (Fig. 5E).
Transcriptional entropy and chronological age
To determine whether gene expression signatures and transcriptional entropy can predict chronological age, we applied a previously described method for elastic net38. polyEN takes cell-type-specific gene expressions as input feature columns and predict individual’s chronological age. The model was applied to the three cell types (AT1, AT2, gCap) for which we observed a high level of transcriptional changes, mutation burden, and difference in entropy. None of the parameters was particularly effective in determining an individual’s age, but entropy achieved the best performance with moderate correlation between the actual and predicted age and lower error (Pearson correlation coefficient (PCC) = 0.283, mean absolute error (MAE) = 12.32) compared to gene expression only (PCC = 0.280, MAE = 13.57), expression and entropy (PCC = 0.263, MAE = 16.42), or baseline (predicted age defined as the mean of all samples) (Supplementary Fig. 7C). When performed for each cell type individually, the combined gene expression and entropy model applied to gCap cells was the best predictor of subject age (PCC = 0.501, MAE = 11.30) (Supplementary Fig. 7D). Moreover, while entropy was a better predictor of age in gCap cells, gene expression was superior for both epithelial cell types. This suggests that the effects of gene expression changes and increased stochasticity might affect aging differently in different cell types. However, validations on additional cohorts are required.
SenMayo does not correlate with age
Because the SenMayo gene signature outperformed all other gene lists in its ability to detect senescent cells15, we evaluated SenMayo as a potential gene signature to study senescence in the lung. High SenMayo cells were identified using a SenMayo score threshold that was manually set to a level above background noise. These cells were then plotted using a UMAP representation, showing that they cluster together (Fig. 6A). High SenMayo cells were primarily in fibroblast, macrophage, and endothelial cell clusters. To confirm that the SenMayo score was characterizing senescent cells and not cells with an inflammatory phenotype, we also examined expression of canonical senescence markers in this dataset. Expression of CDKN1A, a known marker of senescence not included in the SenMayo list, was found to be increased in high SenMayo scoring cells (Fig. 6B). Notably, the SenMayo score was not significantly increased with age in any cell type in our dataset (p > 0.05) (Fig. 6C).
A High SenMayo cells cluster together on a UMAP plot. High SenMayo cells are colored pink. B High SenMayo cells have higher expression of independent senescent markers such as CDKN1A. C SenMayo signature in young versus aged cells, averaged per sample. D Venn diagram of genes that are positively correlated (top) and negatively correlated (bottom) with the SenMayo score in AT2 and gCap cells. E Expression profile for genes that are positively correlated with the SenMayo score in AT2 and gCap cells. Cells (columns) are ordered by cell type and by SenMayo score. Genes (rows) are ordered by cell type and by beta value for association with SenMayo. F CDKN2A expression in samples with high or low mutation burden (n = 31, 16 low/15 high). G Heatmap showing enrichment of top 3 REACTOME terms for each co-expression network. H Cell-type specificity of senescence co-expression network eigengenes. The left panel shows the expression score for each module in each cell type in single-cell RNAseq data. The right panel shows the correlation values between cell-type proportions and module eigengene in the deconvoluted bulk RNAseq dataset. I Plot of age correlation versus global mutation burden for co-expression modules. The x-axis shows correlation values between module expression and age correlation. The y-axis shows correlation values between module expression and global mutation burden. Source data are provided as a Source Data file.
SenMayo identifies heterogeneous signatures of senescence
We identified genes that correlated with the SenMayo score for each of the two cell types with the greatest age-associated transcriptional changes: AT2 and gCap cells. Interestingly, different cell types showed different transcriptional profiles associated with SenMayo (Fig. 6D, E, Supplementary Data 8). Genes associated with senescence in AT2 cells included chemokines CXCL2 and CXCL3, EGFR family gene AREG, and immune regulator CD83. In gCap cells, genes included serine proteinase inhibitors SERPINE1 and SERPINB9 as well as endothelial gene EDN1, which has been implicated in senescence and fibrosis. These genes may represent markers of senescence molecular programs in the different cell types of the lung. Notably, mutation burden was also associated with significantly increased expression of senescence marker CDKN2A (Fig. 6F).
Next, we were interested in determining whether we could identify similar signatures in senescence-associated co-expression modules. Modules were assessed for enrichment of six established senescence gene signatures39,40. Fisher’s Exact testing was performed to compare enrichment of all six gene lists across all modules. Strong overlap was observed across known senescent gene lists (Supplementary Fig. 8A, B). Of the 30 co-expression networks that correlated with age, six modules were also enriched for the SenMayo gene signature (FET p < 0.05), accounting for 59 of the 125 SenMayo genes (Supplementary Fig. 8C). These modules had distinct functional enrichment for REACTOME terms (Fig. 6G). Additionally, they exhibited cell-type-specific expression, both in single-cell data and based on deconvoluted proportions (Fig. 6H, Supplementary Fig. 8D). The module that included the greatest number of SenMayo genes (23 genes, including CXCL1/CXCL2/CXCL3, IL1A/B, IL6, TNF, SERPINE1, and VEGFA) also included p21/CDKN1A, a hallmark of senescence, as a hub gene (Supplementary Fig. 8E). Deconvolution correlated this module (15) with endothelial and myeloid cell types, with enrichment for cytokine/interleukin signaling pathways. Two modules (19, 20) correlated with mesenchymal cell types and were enriched for ECM pathways. Genes included IGF1 (19), IGF binding-proteins, and matrix metalloproteinases. Finally, the module most strongly correlated with age (2, r = 0.26, p < 0.05) contained p16/CDKN2A and FBXW7 as hub genes. Other genes in this module were associated with DNA damage response and oncogene-induced senescence. These include ATM (a gene which is activated by DNA double-strand breaks), MSH2, as well as known oncogenes such as KRAS and RAF141.
To validate the association between somatic mutation burden and senescence gene expression, we plotted age correlation values against somatic mutation estimates for each co-expression module (Fig. 6I). Two senescence modules (2,15) were among the most correlated with somatic mutation burden (133 total modules). Interestingly, these were the two modules that contained CDKN1A and CDKN2A, supporting the association of DNA damage with senescence. Among senescence-associated gene modules, module 2 had the strongest correlation with mutation burden and with age (Fig. 6I). In addition to CDKN2A/p16, this module also contained DNA damage response-associated genes, including ATM, ATR, HMGB1, and MSH2. These results suggest that there are specific gene networks that link DNA damage accumulation with cellular senescence.
Discussion
In this study, we identified aging signatures using a large, integrated single-cell RNAseq dataset and validated these using several independent datasets: bulk RNAseq data from the GTEx project and FFPE lung tissue sections from the LTRC. To our knowledge, this represents the largest single-cell atlas of human lung aging to date. We harnessed the sequencing depth of this dataset to call somatic mutations at cell-type resolution, calculated transcriptional entropy, and developed a method to identify cells expressing a senescence gene signature. AT2 cells and gCap cells experienced the greatest degree of transcriptional change with aging. Expression of surfactant genes declined in AT2 cells, a result of a decrease in the relative proportion of SPChigh AT2 cells in the alveolar epithelium. In the lung microvasculature, capillary cells exhibit increased transcription of autophagy genes and decreased mitochondrial genes and genes related to normal cell function. Our analysis of somatic mutations implicated the same cell types and revealed that mutation accumulation is associated with increased DNA damage response pathways, with epithelial and endothelial cell types exhibiting distinct changes. Finally, we showed that the SenMayo signature identifies high-CDKN1A expressing cells but does not increase with age. Using this signature, we identified heterogeneous markers of senescence in different cell types. Together, the results of this multi-omic analysis revealed cell-specific transcriptional changes associated with aging and senescence, and their relation to somatic mutation accumulation. This dataset is available as an online atlas of human lung aging (http://lungaging.com) for data sharing and exploration.
Our integrated analysis allowed us to obtain novel observations regarding the cellular changes that occur in the alveolar microenvironment with age. Variation in the degree of transcriptional change among cell types suggests that aging of the lung is dyssynchronous, as evidenced by the finding that the epithelium and microvasculature are disproportionately affected in aging. AT2 cells exhibited the greatest degree of change with aging. Analysis of cell-type proportions in both single-cell and bulk RNAseq data also showed an overall loss of AT2 cells with age. This aligns with previous studies that have shown a decrease in AT2 cells with age, particularly in the context of severe COVID-1917 and pulmonary fibrosis13. Analysis of differential gene expression in single-cell data showed reduced expression of surfactant genes in the aged. Similarly, using gene co-expression analysis, we identified a module associated with surfactant production in AT2 cells that was negatively associated with age. HHIP, one of the genes most strongly associated with susceptibility to COPD in human GWAS studies and functionally with maintaining normal lung function and alveolar structures in mice42,43,44, was a hub gene for this module. This finding suggested an age-related change in AT2 cell subpopulations because HHIP is primarily expressed in one subpopulation identified as SPChigh AT2 cells27,28. When examining single-cell data, AT2 cells were comprised of two major groups: SPChigh AT2 cells that have high expression of surfactants and correspond with a functional alveolar maintaining phenotype, and SPClow AT2 cells that have lower levels of surfactant and correspond with a more stem-like progenitor phenotype29. Our scRNAseq data, as well as our immunohistochemistry validation, confirmed decreased expression of HHIP with age, as well as a decrease in the proportion of SPChigh AT2 cells in the aged lung. Our findings are consistent with previous studies that showed reduced alveolar epithelial differentiation with aging45. Our analysis extends this and uncovers both a specific decline in SPChigh AT2 with age, as well as a general decline in surfactant gene networks among alveolar epithelial cells. The decline in surfactant transcriptional programs and the relative decline in SPChigh AT2 cells may have important implications in explaining the increased predisposition of the aged lung to alveolar injury and diseases such as ARDS and pneumonia10,46.
Our analysis also identified disproportionally high levels of differentially expressed genes in the lung microvasculature. We identified several hallmarks of aging in both the capillary cell populations of the lung: gCap cells and aerocytes. These cells increased the expression of genes related to autophagy as well and decreased transcription of mitochondrial genes, indicating possible loss of proteostasis and mitochondrial dysfunction in these cells. Moreover, these cells had reduced expression of their marker genes, suggesting a loss of normal cellular identity. We observed a gene signature unique to capillary endothelial cells consisting of ubiquitin ligases and mTOR signaling genes that was increased with aging. FBXW7, a component of the SCF ubiquitin ligase, also emerged as a hub gene in the co-expression network that was most highly correlated with age, which was expressed in endothelial cells. Prior studies have shown that FBXW7 expression increases downstream of p53 activation, suggesting that aged capillary cells might be upregulating ubiquitination in response to DNA damage accumulation47. The high level of somatic mutations in endothelial cells in our dataset supports this hypothesis.
Harnessing the large size and sequencing depth of this dataset, we were able to call somatic mutations directly from single-cell RNAseq data. Our estimated mutation burden ranged from 3.1 to 7.2 mutations/MB, which is on the same order of magnitude as the rate found for skin epithelial cells but slightly higher than other cell types in a paper validating the same mutation-calling method33. Other studies that called somatic mutations from bulk sequencing data of bronchial epithelial cells and other tissues reported lower mutation rates than approximated in this study31,48,49. This difference may be because such analyses may underestimate mutation burden when they occur in a subset of cells50. Additionally, the higher mutation burden we report may reflect the lung’s direct exposure to oxygen and environmental mutagens. Indeed, others have shown higher mutation rates in the skin and lungs compared to other organs51,52. Cell types present in the alveolar unit, including epithelial cells and gCap cells, were among the cells with the greatest somatic mutation burden, suggesting that epithelial and endothelial cell types also experience dyssynchronous patterns of genomic damage, in addition to transcriptional change, during aging. A possible explanation for the high rate of mutation accumulation in these cell types may be due to the environmental exposure of cells present in the alveolus. Mutation burden was positively correlated with age across cell types, consistent with previous findings. Differential gene expression revealed significant phenotypic changes, including increased DNA damage response genes and mitochondrial dysfunction. We showed that AT2 cells and gCap cells exhibited distinct signatures. The increase in DNA damage response pathways in samples with high mutation burden suggests that the observed differentially expressed genes likely represent a response to environmental mutagens rather than a regulated response to the number of mutations. Impressively, performing the same analysis on somatic mutation data derived from bulk RNAseq and validated with whole-exome sequencing revealed strong overlap with the signatures identified using single-cell data and confirmed our scRNAseq results. Taken together, our results extend on previous reports of increased somatic mutations with aging in bronchial epithelial cells31 by providing cell-type resolution of somatic mutations in the lung and characterization of the associated transcriptional changes.
Aging and genomic instability are thought to be associated with transcriptional stochasticity or dysregulation36. The loss of marker gene expression we observed with increasing aging and somatic mutation burden accumulation supports this idea36. Hence, we sought to quantify metrics such as transcriptional entropy and noise and determine their relation to aging. We modeled transcriptomes from cells as a probability distribution in order to quantify the statistical surprise of a cell’s transcriptome, which is inversely related to entropy. This can be interpreted as the degree of specialization of a cell37. This was compared with established methods of computing transcriptional noise. Interestingly, we observed the highest differences in metrics of transcriptional dysregulation in the same cell types that had the highest number of differentially expressed genes, such as capillary cells and AT2 cells, consistent with the finding of greater transcriptional change and mutation burden in these cell types. More work is needed to determine whether these changes are driven by loss of regulation or increased stochasticity. Interestingly, we find that entropy alone is superior to gene expression in predicting chronological age. This suggests that dysregulation of gene expression may contribute more to age-related phenotypes than coordinated changes in gene expression. However, when applying age prediction models to individual cell types, we observed that while entropy performs better in chronological age prediction in some cell types (gCap), gene expression performs better in others (AT1, AT2), suggesting that these changes affect different cell types to a different degree. Interestingly, age prediction was most accurate in gCap cells, supporting the finding that gCap cells had greater changes in gene expression and entropy. This also has implications for the development of lung aging biomarkers. Previous work in the human pancreas showed that pancreatic cells exhibit increased transcriptional noise and accumulate somatic mutations with age35. Here, we show similar trends in the human lung and identify cell-type-specific differences in these patterns. Our analysis points to an outsized level of age-associated genomic alterations and transcriptional entropy in subsets of lung cell types.
Our results suggest the existence of heterogeneous signatures of cellular senescence. We report differences in the genes that are associated with the senescent signature in AT2 and gCap cells, suggesting that different cell types might exhibit different phenotypes of cellular senescence and respond to different triggers. Specifically, we observed genes related to the inflammatory and immune-modulatory aspects of SASP in AT2 cells (CXCL2, CXCL3, CD83) and more genes related to ECM remodeling and vascular senescence in gCap cells (SERPINE1, SERPINB9, EDN1). In bulk RNAseq data, the top senescence-associated module included CDKN1A as a hub gene and was closely co-expressed with interferon signaling genes STAT3, IL4R, IL1R1, CCL2, and IL6, suggesting its central role in connecting cell cycle and SASP production. Another module (2) was expressed in most cell types and contained CDKN2A (p16). Enrichment analysis showed that this module contained a wide variety of genes involved in DNA repair and DNA damage response. This supports the idea of specific gene networks that link cellular damage response and cellular senescence21. Notably, CDKN1A and CDKN2A were associated with different modules in this analysis, indicating their likely contributions to different senescence signaling pathways, as has been previously reported53. Our findings suggest that p16 and p21 have different roles in the induction and maintenance of cellular senescence in the lung. Whereas p21 appears to be more directly related to the SASP, p16 is linked with DNA damage response and oncogene induction. Indeed, CDKN2A expression was associated with higher mutation burden in single-cell data, suggesting that mutation accumulation plays a role in activating senescence gene expression. We validated this by analyzing somatic mutations called from bulk RNAseq. This confirmed the increase of somatic mutations with age, as well as the association between mutation burden and senescence gene signature. Our results link somatic mutation accumulation with expression of senescence genes. Similar analysis has been performed on single-cell human pancreas data, showing a correlation between mutation load and CDKN2A expression in endocrine cells35. These findings provide support to the hypothesis that accumulation of macromolecule damage, including DNA damage, leads to both age-associated phenotypes and cellular senescence in the aged human lung21.
Our study has some limitations. Single-cell RNAseq is inherently sparse and subject to biases in transcript coverage, which affect cell inferences. We addressed this by validating our findings using bulk RNAseq data, showing that gene co-expression networks recapitulate the transcriptional signatures seen in single-cell RNAseq. Somatic mutation calling from single-cell data is, of course, not as accurate as DNA sequencing33. Hence, variants could only be confidently detected at highly expressed loci, mutation burden was extrapolated from a limited portion of the genome compared to whole-genome sequencing, and amplification errors may have contributed to higher mutation burden estimates. Some cell types also did not have enough sequencing depth to call somatic mutations, limiting our analysis. Moreover, the mutation callset achieved using this method was not large enough to identify mutation hotspots. We additionally analyzed the somatic mutation data from bulk RNA to validate gene expression signatures. However, this data consists of clonally amplified somatic mutations, making them distinct from those performed on single-cell RNAseq data. Our analysis also did not include methylation data, which could have provided additional validation for our age prediction model. Finally, our analysis also did not address the spatial and regional organization of aging changes within the lung. Further studies that sample multiple regions and use spatial profiling methodologies will be needed to further characterize and describe the temporal and spatial dynamics of these changes and define the role of these changes in the enhanced disease predisposition and reduced resilience.
In summary, our study represents the most comprehensive multi-omic study of lung aging to date; we have identified novel cellular and molecular signatures associated with aging, somatic mutation accumulation, and cellular senescence, in specific cell types in the aged human lung. We show that these alterations appear to be dyssynchronous, with epithelial cells and endothelial cells of the alveolar niche exhibiting greater effects. Further studies will be needed to further characterize and describe the temporal and spatial dynamics of these changes and define the role of these changes in the enhanced disease predisposition and reduced resilience.
Methods
Single-cell RNAseq
Single-cell RNAseq was performed on tissue from lung explants not suitable for transplant from 32 organ donors at Baylor College of Medicine. Tissue was collected under the Chronic Lung Disease Tissue Repository at Baylor College of Medicine (IRB H-46832), which required written consent. Samples were obtained from a single lung lobe, with the specific lobe varying among study subjects. Samples were collected from cryopreserved human lung samples in 10% DMSO. Cryopreservation durations ranged from several months to years prior to single-cell RNA sequencing library preparation. Lungs were not perfused before cryopreservation. Samples were provided without any identifying information, and the study was determined as non-human subject research by the Yale IRB committee (IRB Protocol ID: 200002456). IRB review and approval were therefore waived.
After thawing at 37 °C, the tissue was minced into small pieces, incubated for 45 min at 37 °C with DMEM + Elastase, DNAase, liberase and 1% penicillin+streptomycin+glutamine. Digestion was stopped with fetal bovine tissue to 10%. The digested tissue was passed on a 100-micron mesh and centrifuged at 300 g for 10 min. Cell pellets were resuspended with MACS medium, and later on, the cells were barcoded using the 10× Chromium single-cell platform. Following amplification, size selection, fragmentation, and addition of sequencing adapters, cDNA libraries were created per the standard 10× protocol (Single-Cell 5′ Reagent Kits v2, 10x Genomics, USA). Libraries were sequenced using the Illumina NextSeq 500 system. Cell Ranger (10X Genomics, USA) was used to generate FASTQ files from sequencing data. Adapter sequences were trimmed from FASTQ data. Finally, reads were demultiplexed, aligned, and counted.
This dataset was integrated with scRNAseq data reported in our group’s previously reported IPF Cell Atlas22. Healthy controls consisted of 38 samples from 28 subjects. This dataset consisted of a total of 96083 cells, with an age range from 20 to 80 years. The two scRNAseq datasets were integrated using reciprocal PCA integration54 in Seurat.
scRNAseq data analysis was performed using the standard Seurat pipeline. Gene expression values were normalized by the “NormalizeData” method. The top 2000 variable genes were identified by the “FindVariableFeatures” function. After scaling gene expressions, a linear dimensional reduction was performed using the variable genes by the “RunPCA” function, which generated 30 principal components. The “RunUMAP” function was applied for nonlinear dimensional reduction and cluster visualization. Clustering was performed using the “FindNeighbors” function, which uses a K-nearest neighbor (KNN) graph. The “FindClusters” function was subsequently applied to optimize modularity by the Louvain algorithm.
To identify marker genes of each cell type, the “FindAllMarkers” function from Seurat was applied to identify differentially expressed genes using a Wilcoxon Rank-Sum test. Only significantly upregulated genes (FDR < 0.05) with 0.1 log fold change and 1% minimum expression fraction were retained as marker genes.
The SenMayo score was calculated using the “AddModuleScore” function in Seurat, which averages the expression of each gene list, subtracted by the expression of random control feature sets. Correlation between the SenMayo score and age was calculated by determining the average SenMayo score per cell type, per subject, and correlating with subject age.
Differential gene expression
To identify transcriptional changes, we applied a general linear mixed effects model (GLMM). To ensure biologically interpretable results, we only tested genes with counts greater than zero in at least 50% of samples and 5% of total cells. This filtering method was performed independently for each cell type. glmmTMB55 was used to develop a model using a negative binomial distribution that accounts for variance due to differences in samples or assays. Hence, our model accounts for the sample as a random effect. We also included an offset term to weigh cells with greater sequencing coverage (higher UMIs) accordingly. To determine age-associated transcriptional changes, samples with an age of less than 20 years were excluded.
To further ensure the robustness and reproducibility of this set of differentially expressed genes, we further validated differentially expressed genes using a pseudobulk likelihood ratio test. Samples with fewer than 20 cells of the cell type being tested were excluded.
Enrichment of KEGG and REACTOME pathways was determined using gprofiler256 in R.
Bulk RNAseq datasets
Lung-specific bulk RNAseq data was downloaded from the Genotype-Tissue Expression (GTEx) Portal (https://gtexportal.org/home/datasets). GTEx consists of samples from 54 non-diseased tissue sites across nearly 1000 individuals. Tissue was collected from postmortem/organ procurement cases. The data consisted of 572 samples with an age range of 20–79 years. Because the age of GTEx subjects was reported in 10-year ranges, the mean value of these ranges was used for subsequent analysis. The normalized gene expressions were log2-transformed.
A second bulk lung RNAseq dataset from the Lung Tissue Research Consortium (LTRC) was processed in parallel with the original dataset and used to assess for module preservation across datasets. This dataset was downloaded from GEO GSE47460. The same parameters for data pre-processing were applied to this dataset. This dataset consisted of 91 samples, with an age range from 32 to 87 years.
Cellular deconvolution
The GTEx bulk RNAseq dataset was deconvoluted to determine cell-type proportions using MuSiC57 in R. MuSiC uses support vector regression on gene expression profiles using reference gene expression signatures. Our single-cell RNAseq dataset was used for this purpose. The cell-type composition was determined for each sample from the signatures in the original expression profiles. The resulting cell-type proportions were correlated with the module eigengenes, revealing cell-type specificity. This allowed for direct comparison of gene and co-expression network between bulk RNAseq datasets and single-cell RNAseq data.
Weighted gene co-expression network analysis
WGCNA was conducted using R/Bioconductor to identify modules in the normalized bulk RNAseq dataset. Analysis parameters were adjusted: sign of correlations between neighbors (TOMtype and networkType = ‘signed’), and module detection parameter (deepSplit = 2). Modules were identified by number in order of decreasing module size. Module eigengene (ME) was calculated as the first principal component of gene expression for the module. Module association with age was calculated using the module eigengenes (the first principal component of the expression profile). Pearson correlation adjusted for multiple comparisons by FDR. Module membership, a measure of the association of a gene to its module, was determined by Pearson correlation of gene expression to ME and used to rank module connectivity.
Module age correlation and cell-type specificity were validated by calculating expression in the single-cell dataset (expression of each gene list subtracted by expression of random control feature sets). Modules were then assessed for the presence of genes from the SenMayo gene list15 using Fisher’s Exact Test (FET).
Hub genes were defined as those that had a high module membership (correlation of gene expression to ME) and high correlation with age based on expression in GTEx RNAseq data. In order to validate these hub genes, the correlation of hub gene expression with age was calculated in the single-cell RNAseq dataset. Expression values were calculated for each cell-type individually, and only cells with a minimum expression value were included in the analysis.
Somatic mutations
Somatic mutations were detected in single-cell RNAseq data using SComatic33. Mutation calling was performed using the default parameters of the SComatic pipeline. BAM files containing sequencing reads were split into cell-type-specific files using existing cell-type annotations (see “Single-cell RNAseq” section). Base count matrices were created for each cell type and merged together for subsequent analysis. Mutations were called using a beta-binomial distribution test to determine whether variant allele counts were significant. Following this, candidate somatic mutations were subject to a variety of filters to reduce false positives.
Mutational burden was determined for each cell type per sample. This was calculated by dividing the number of detected somatic mutations by the number of callable sites for a given cell type. Correlation with age was determined using a GLMM model using cell type as a covariate.
Differential expression was performed using a GLMM model similar to that used to identify age-associated genes (see “Differential gene expression” section). To identify genes that were associated with somatic mutation accumulation globally, an additional random effect term to account for cell type was added.
Bulk somatic mutation data was acquired from Garcia-Nieto et al.34. Mutations in this study were calculated by mapping raw RNAseq reads to the reference genome Hg19 and deploying a comprehensive mutation-calling pipeline. False positive mutation calls were minimized by accounting for sequencing errors, RNA editing events, germline variants, and other sources of error. Somatic mutation calling was validated using matched whole-exome DNA sequencing data from the same samples. Somatic mutation burden for the present study was calculated using the sum of somatic mutation counts for each sample34. Mutation burden estimates per cell type were approximated by correlating cell-type proportion estimates from deconvolution with total sample mutation burden.
Bulk RNAseq gene expression was correlated with mutation burden using the DESeq2 package in R58. Pre-filtering was performed to include genes with at least 100 transcripts in at least 100 samples. Differential expression analysis was performed using the default DESeq2 model, which applies a generalized linear model to the raw count data.
Entropy and transcriptional noise
The dataset was randomly downsampled so that a maximum of 10,000 cells of each type and an equal number of young and old cells were included. Raw count matrices were downsampled so that all cells have equal total UMIs. Only cell types for which there were at least 10 young and 10 old cells were included.
Using the philentropy package in R, K–L divergence was calculated for each cell, comparing raw counts for a given cell versus a probability distribution where each gene count = 1. Increasing entropy was defined as decreasing the K–L divergence from a uniform distribution.
Similarly, transcriptional noise was calculated for each cell type following previous work18. Downsampling was performed as described above for entropy. Genes were grouped into 10 equal bins according to mean expression. The top and bottom bins were excluded, and the remaining top 10% of genes with the lowest coefficient of variation for each bin were retained. Following square-root normalization, we calculated a Euclidean distance between each cell transcriptome and the corresponding cell-type mean for each age. Euclidean distance was calculated using the distancevector function from the hopach package in R. Both metrics were calculated for every cell in the integrated dataset.
Chronological age prediction model
Only cells with available entropy values were used for chronological age prediction. For each subject, we computed mean entropies for cells annotated as AT1, AT2 (SPChigh and SPClow), or gCap and used them as four different input feature columns to predict age. We adapted a recently published method, polyEN38, to our prediction task. polyEN can take cell-type-specific gene expressions as input feature columns and predict an individual’s chronological age using an elastic net model.
Here, we considered input feature columns as the mean entropy of each cell type from each subject (4 columns in total for each subject). In addition to entropy features, we also computed the first 2 principal components (PCs) from the gene expressions for the selected four cell types (8 columns in total for each subject). These expression PCs were included as additional feature columns to predict an individual’s chronological age.
We used either entropy features alone, or expression PC features alone, or entropy + expression features together to predict chronological age. We additionally included a baseline method, which uses the average age from all subjects as the predicted age. The performance for all methods was then evaluated by a leave-one-out test. Among all donors (60 donors), we filtered the donors by using only the donors that have at least three out of four columns (cell types) available for entropy values. We used the Pearson correlation coefficient (PCC) and the mean absolute error (MAE) between the predicted chronological age and the true chronological age as evaluation metrics. We filtered the subjects by selecting subjects having at least two non-zero entropy columns among the four cell-type-specific entropy columns.
Immunohistochemistry
FFPE samples were provided by the LTRC and were derived from subjects undergoing thoracic surgery. These subjects were diagnosed as being controls or having interstitial lung disease or COPD as determined by clinical history, CT scan, and surgical pathology. There was no intervention, as these are cross-sectional data. This dataset included 582 total subjects (254 have interstitial lung disease, 220 have COPD, and 108 are controls).
FFPE samples from healthy control human subjects were used for IHC. Samples were split into young and aged groups for immunohistochemistry, with 6 FFPE samples in each group. Age ranged from 44–53 years for young samples to 66–78 years for aged samples (See Table 1). FFPE blocks were processed as 5-micron-thick sections.
FFPE slides were deparaffinized in xylene and rehydrated using a series of gradually decreasing ethanol concentrations. 10-min microwave antigen retrieval was performed using a pH 6.0 sodium citrate antigen retrieval buffer, followed by a 10-min period of blocking endogenous enzyme activity using Bloxall (Vector Laboratories). SFTPC was detected using mouse anti-SFTPC polyclonal antibody diluted 1:200 (Santa Cruz Biotechnology, #518029), and HHIP was detected using rabbit anti-HHIP polyclonal antibody diluted 1:1500 (ABClonal Science, #A3260). Slides were incubated with primary antibody at 4 °C for 16 h, and later with HRP and AP Red secondary antibodies for 1 h each. Immunoreactive signal was visualized with either DAB solution (Vector Laboratories) or AP Red (Vector Laboratories). Slides were then counterstained with hematoxylin, dehydrated, and mounted.
To quantify staining, digital images of slides (40× magnification) were viewed using NIS-Elements (Nikon Instruments). Positive-stained cells were counted using color deconvolution in FIJI (National Institutes of Health). 5 fields per sample were obtained, and cells were counted with a minimum size threshold of 300 pixels.
Statistics
Log-normalized expression values are shown in all figures. All p-values in this study were multiple test-corrected using an FDR threshold of 5%. Seurat’s FindMarkers function uses the Wilcoxon Rank-Sum test score to pick statistically significant differential genes among conditions. Comparison of differentially expressed genes between tests was performed using a one-sided Fisher's Exact Test.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data supporting the findings from this study are available in the manuscript and its supplementary information. Source data are provided with this paper. Single-cell RNAseq data generated for this study is available from NCBI Gene Expression Omnibus (GSE281219). Single-cell RNAseq data from the IPF Cell Atlas22 (GSE136831) and LTRC bulk RNAseq data (GSE47460) are also available on GEO. Lung-specific bulk RNAseq data was downloaded from the Genotype-Tissue Expression (GTEx) Portal (https://gtexportal.org/home/datasets). Matched bulk RNAseq somatic mutation data is available as supplementary data from Garcia-Nieto et al.34. Source data are provided with this paper.
Code availability
The code used to generate the results presented in this manuscript is available as a public repository (https://github.com/rubendeman/Lung-Aging-Cell-Atlas).
References
Bowdish, D. M. E. The aging lung is lung health good health for older adults?. Chest 155, 391–400 (2019).
Schneider, J. L. et al. The aging lung: physiology, disease, and immunity. Cell 184, 1990–2019 (2021).
Cho, S. J. & Stout-Delgado, H. W. Aging and Lung Disease. Annu. Rev. Physiol. 82, 433–459 (2020).
Chilosi, M., Poletti, V. & Rossi, A. The pathogenesis of COPD and IPF: distinct horns of the same devil? Resp. Res. https://doi.org/10.1186/1465-9921-13-3 (2012).
Bartling, B. Cellular senescence in normal and premature lung aging. Z. Gerontol. Geriatr. 46, 613–622 (2013).
Budinger, G. R. S. et al. The intersection of aging biology and the pathobiology of lung diseases: a Joint NHLBI/NIA Workshop. J. Gerontol. A Biol. Sci. Med. Sci. 72, 1492–1500 (2017).
Zeleznik, J. Normative aging of the respiratory system. Clin. Geriatr. Med. https://doi.org/10.1016/S0749-0690(02)00063-0 (2003).
Copley, S. J. Morphology of the aging lung on computed tomography. J. Thorac. Imag. 31, 140–150 (2016).
Schröder, T. H., Storbeck, B., Rabe, K. F. & Weber, C. The aging lung: clinical and imaging findings and the fringe of physiological state. Rofo 187, 430–439 (2015).
Brandenberger, C. & Muhlfeld, C. Mechanisms of lung aging. Cell Tissue Res. 367, 469–480 (2017).
Bailey, K. L. et al. Aging causes a slowing in ciliary beat frequency, mediated by PKCε. Am. J. Physiol. Lung Cell. Mol. Physiol. 306, L584–L589 (2014).
Godin, L. M. et al. Decreased laminin expression by human lung epithelial cells and fibroblasts cultured in acellular lung scaffolds from aged mice. PLoS ONE https://doi.org/10.1371/journal.pone.0150966 (2016).
Lee, S. et al. Molecular programs of fibrotic change in aging human lung. Nat. Commun. https://doi.org/10.1038/s41467-021-26603-2 (2021).
Lonsdale, J. et al. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Saul, D. et al. A new gene set identifies senescent cells and predicts senescence-associated pathways across tissues. Nat. Commun. https://doi.org/10.1038/s41467-022-32552-1 (2022).
Xu, P. et al. The landscape of human tissue and cell type specific expression and co-regulation of senescence genes. Mol. Neurodegener. https://doi.org/10.1186/s13024-021-00507-7 (2022).
Chow, R. D., Majety, M. & Chen, S. D. The aging transcriptome and cellular landscape of the human lung in relation to SARS-CoV-2. Nat. Commun. https://doi.org/10.1038/s41467-020-20323-9 (2021).
Angelidis, I. et al. An atlas of the aging lung mapped by single cell transcriptomics and deep tissue proteomics. Nat. Commun. https://doi.org/10.1038/s41467-019-08831-9 (2019).
Jia, M. X. et al. Transcriptional changes of the aging lung. Aging Cell https://doi.org/10.1111/acel.13969 (2023).
Liu, X. et al. Transcriptomics analysis identifies the decline in the alveolar type II stem cell niche in aged human lungs. Am. J. Resp. Cell Mol. Biol. 71, 229–241 (2024).
Ogrodnik, M., Salmonowicz, H. & Gladyshev, V. N. Integrating cellular senescence with the concept of damage accumulation in aging: relevance for clearance of senescent cells. Aging Cell 18, e12841 (2019).
Adams, T. S. et al. Single-cell RNA-seq reveals ectopic and aberrant lung-resident cell populations in idiopathic pulmonary fibrosis. Sci. Adv. https://doi.org/10.1126/sciadv.aba1983 (2020).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. https://doi.org/10.1186/1471-2105-9-559 (2008)
Lee, B. Y. et al. Senescence-associated beta-galactosidase is lysosomal beta-galactosidase. Aging Cell 5, 187–195 (2006).
Peng, Q., Gao, L., Cheng, H. B., Wang, J. S. & Wang, J. Sialidase NEU1 may serve as a potential biomarker of proliferation, migration and prognosis in melanoma. World J. Oncol. 13, 222–234 (2022).
Hegab, A. E. et al. High fat diet activates adult mouse lung stem cells and accelerates several aging-induced effects. Stem Cell Res. 33, 25–35 (2018).
Sauler, M. et al. Characterization of the COPD alveolar niche using single-cell RNA sequencing. Nat. Commun. https://doi.org/10.1038/s41467-022-28062-9 (2022).
McDonough, J. E. et al. Low surfactant type II alveolar epithelial cells are an enriched cell population in the aged lung. Am. J. Resp. Crit. Care 201, A4390 (2020).
Travaglini, K. J. et al. A molecular cell atlas of the human lung from single-cell RNA sequencing. Nature https://doi.org/10.1038/s41586-020-2922-4 (2020).
Schupp, J. C. et al. Integrated single-cell atlas of endothelial cells of the human lung. Circulation 144, 286–302 (2021).
Huang, Z. Q. et al. Single-cell analysis of somatic mutations in human bronchial epithelial cells in relation to aging and smoking. Nat. Genet. 54, 492 (2022).
Cagan, A. et al. Somatic mutation rates scale with lifespan across mammals. Nature 604, 517 (2022).
Muyas, F. et al. De novo detection of somatic mutations in high-throughput single-cell profiling data sets. Nat. Biotechnol. https://doi.org/10.1038/s41587-023-01863-z (2023).
Garcia-Nieto, P. E., Morrison, A. J. & Fraser, H. B. The somatic mutation landscape of the human body. Genome Biol 20, https://doi.org/10.1186/s13059-019-1919-5 (2019).
Enge, M. et al. Single-cell analysis of human pancreas reveals transcriptional signatures of aging and somatic mutation patterns. Cell 171, 321–330.e14 (2017).
Vijg, J. & Dong, X. Pathogenic mechanisms of somatic mutation and genome mosaicism in aging. Cell 182, 12–23 (2020).
Teschendorff, A. E. & Enver, T. Single-cell entropy for accurate estimation of differentiation potency from a cell’s transcriptome. Nat. Commun. https://doi.org/10.1038/ncomms15599 (2017).
Song, Q. et al. Predicting lung aging using scRNA-Seq data. PLoS Comput. Biol. https://doi.org/10.1371/journal.pcbi.1012632 (2024).
Avelar, R. A. et al. A multidimensional systems biology analysis of cellular senescence in aging and disease. Genome Biol. https://doi.org/10.1186/s13059-020-01990-9 (2020).
Hernandez-Segura, A. et al. Unmasking transcriptional heterogeneity in senescent cells. Curr. Biol. 27, 2652 (2017).
Negrini, S., Gorgoulis, V. G. & Halazonetis, T. D. Genomic instability - an evolving hallmark of cancer. Nat. Rev. Mol. Cell Biol. 11, 220–228 (2010).
Lao, T. T. et al. Haploinsufficiency of Hedgehog interacting protein causes increased emphysema induced by cigarette smoke through network rewiring. Genome Med. https://doi.org/10.1186/s13073-015-0137-3 (2015).
Lao, T. T. et al. Hhip haploinsufficiency sensitizes mice to age-related emphysema. Proc. Natl. Acad. Sci. USA 113, E4681–E4687 (2016).
Zhou, X. B. et al. Identification of a chronic obstructive pulmonary disease genetic determinant that regulates. Hum. Mol. Genet. 21, 1325–1335 (2012).
Watson, J. K. et al. Distal lung epithelial progenitor cell function declines with age. Sci. Rep. https://doi.org/10.1038/s41598-020-66966-y (2020).
Schousboe, P. et al. Reduced levels of pulmonary surfactant in COVID-19 ARDS. Sci. Rep. https://doi.org/10.1038/s41598-022-07944-4 (2022).
Yeh, C. H., Bellon, M. & Nicot, C. FBXW7: a critical tumor suppressor of human cancers. Mol. Cancer https://doi.org/10.1186/s12943-018-0857-2 (2018).
Yoshida, K. et al. Tobacco smoking and somatic mutations in human bronchial epithelium. Nature 578, 266 (2020).
Xu, H. et al. Landscape of human protein-coding somatic mutations across tissues and individuals. Preprint at bioRxiv. https://doi.org/10.1101/2025.01.07.631808 (2025).
Luquette, L. J., Bohrson, C. L., Sherman, M. A. & Park, P. J. Identification of somatic mutations in single cell DNA-seq using a spatial model of allelic imbalance. Nat. Commun. https://doi.org/10.1038/s41467-019-11857-8 (2019).
Ren, P. J., Dong, X. & Vijg, J. Age-related somatic mutation burden in human tissues. Front. Aging https://doi.org/10.3389/fragi.2022.1018119 (2022).
Yun, J. H. et al. Clonal somatic mutations in chronic lung diseases are associated with reduced lung function. Am. J. Resp. Crit. Care 208, 1196–1205 (2023).
Kumari, R. & Jat, P. Mechanisms of cellular senescence: cell cycle arrest and senescence associated secretory phenotype. Front. Cell Dev. Biol. https://doi.org/10.3389/fcell.2021.645593 (2021).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Brooks, M. E. et al. glmmTMB balances speed and flexibility among packages for zero-inflated generalized linear mixed modeling. R. J. 9, 378–400 (2017).
Kolberg, L. et al. g:Profiler-interoperable web service for functional enrichment analysis and gene identifier mapping (2023 update). Nucleic Acids Res. 51, W207–W212 (2023).
Wang, X. R., Park, J., Susztak, K., Zhang, N. R. & Li, M. Y. Bulk tissue cell type deconvolution with multi-subject single-cell expression reference. Nat. Commun. https://doi.org/10.1038/s41467-018-08023-x (2019).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. https://doi.org/10.1186/s13059-014-0550-8 (2014).
Acknowledgements
The study was funded by NIH NHLBI grants R01HL127349, R01HL141852, U01HL145567, and UH2HL123886 to N.K. R.D.M. is an MD/PhD student with support from the Yale Medical Scientist Training Program (MSTP) grant (T32GM136651). E.P.M. is a Pepper Scholar with support from the Claude D. Pepper Older Americans Independence Center at Yale School of Medicine (P30AG021342), Department of Veterans Affairs, Veterans Health Administration, VISN 1 Career Development Award, and NIA R03AG074063. The work on this study was supported in part by grant T32GM086287 from NIGMS to MSBR. The opinions expressed are those of the authors and do not necessarily represent the thoughts or opinions of NIGMS, NIH, or the United States government.
Author information
Authors and Affiliations
Contributions
R.D.M., J.E.M., I.O.R., and N.K. conceived and designed the project. R.D.M., R.R., and S.A. performed immunohistochemistry experiments. T.S.A., J.C., B.M., A.W., F.P., R.C., and C.C. contributed to single-cell RNA sequencing. R.D.M., J.E.M., T.S.A., F.N., I.O.R., and E.P.M. contributed to the analysis of single-cell RNAseq data. Q.S. and Z.B.J. performed age prediction modeling. M.S.B.R., X.Y., and N.K. supervised analysis of single-cell RNAseq data. B.M.V., W.A.W., L.N., I.O.R., and N.K. supervised sequencing experiments. R.D.M. and N.K. wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
N.K. served as a consultant to Boehringer Ingelheim, Nuvectis, Baobab, GlaxoSmithKline, Pliant, Samumed, Three Lake Partners, Astra Zeneca, Augmanity, CSL Behring, Galapagos, Fibrogen, and Thyron over the last 3 years, reports Equity in Pliant, and grants from Astra Zeneca and BMS. L.N. is the founder, President, and CEO of Humacyte Global Inc, a publicly traded regenerative medicine company. MSBR holds stock in and serves as a consultant for Humacyte Inc. The remaining authors declare no conflicts of interest.
Peer review
Peer review information
Nature Communications thanks Jianxin Wang and the other anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
De Man, R., McDonough, J.E., Adams, T.S. et al. Single-cell atlas of human lung aging identifies cell type dyssynchrony and increased transcriptional entropy. Nat Commun 17, 2095 (2026). https://doi.org/10.1038/s41467-026-68810-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68810-9